code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sympy import *
init_printing(use_unicode=True)
from sympy.codegen.ast import Assignment
C = Matrix( symarray('C', (2,2)) )
R = Matrix( symarray('R', (2,2)) )
n = Matrix( symarray('n', (2)) )
t = Matrix( symarray('t', (2)) )
R[0,0]=t[0]
R[1,0]=t[1]
R[0,1]=n[0]
R[1,1]=n[1]
C
simplify(transpose(R)*C*R)
# +
TensRot = Matrix( symarray('r', (3,3)) )
TensRot[0,0]=t[0]*t[0]
TensRot[0,1]=t[1]*t[1]
TensRot[0,2]=t[0]*t[1] + t[1]*t[0]
TensRot[1,0]=n[0]*n[0]
TensRot[1,1]=n[1]*n[1]
TensRot[1,2]=n[0]*n[1] + n[1]*n[0]
TensRot[2,0]=t[0]*n[0]
TensRot[2,1]=t[1]*n[1]
TensRot[2,2]=t[0]*n[1] + t[1]*n[0]
TensRot
# +
InvTensRot = Matrix( symarray('r', (3,3)) )
InvTensRot[0,0]=t[0]*t[0]
InvTensRot[0,1]=n[0]*n[0]
InvTensRot[0,2]=t[0]*n[0] + t[0]*n[0]
InvTensRot[1,0]=t[1]*t[1]
InvTensRot[1,1]=n[1]*n[1]
InvTensRot[1,2]=t[1]*n[1] + t[1]*n[1]
InvTensRot[2,0]=t[0]*t[1]
InvTensRot[2,1]=n[0]*n[1]
InvTensRot[2,2]=t[0]*n[1] + t[1]*n[0]
InvTensRot
# -
simplify(R*C*transpose(R))
print(latex(InvTensRot))
print(latex(TensRot))
# +
theta = Symbol("theta")
n[0] = -sin(theta)
n[1] = cos(theta)
t[0]= n[1]
t[1]= -n[0]
# +
InvTensRot[0,0]=t[0]*t[0]
InvTensRot[0,1]=n[0]*n[0]
InvTensRot[0,2]=t[0]*n[0] + t[0]*n[0]
InvTensRot[1,0]=t[1]*t[1]
InvTensRot[1,1]=n[1]*n[1]
InvTensRot[1,2]=t[1]*n[1] + t[1]*n[1]
InvTensRot[2,0]=t[0]*t[1]
InvTensRot[2,1]=n[0]*n[1]
InvTensRot[2,2]=t[0]*n[1] + t[1]*n[0]
TensRot[0,0]=t[0]*t[0]
TensRot[0,1]=t[1]*t[1]
TensRot[0,2]=t[0]*t[1] + t[1]*t[0]
TensRot[1,0]=n[0]*n[0]
TensRot[1,1]=n[1]*n[1]
TensRot[1,2]=n[0]*n[1] + n[1]*n[0]
TensRot[2,0]=t[0]*n[0]
TensRot[2,1]=t[1]*n[1]
TensRot[2,2]=t[0]*n[1] + t[1]*n[0]
# +
sigma = Matrix( symarray('\sigma', (3)) )
sigma[0]=1
sigma[1]=3
sigma[2]=2
(TensRot*sigma).subs(theta,20*pi/180).evalf()
# -
(Transpose(TensRot)*sigma).subs(theta,20*pi/180).evalf()
# +
(InvTensRot*sigma).subs(theta,-50*pi/180).evalf()
# -
(Transpose(TensRot)*TensRot).subs(theta,20*pi/180).evalf()
(InvTensRot*TensRot).subs(theta,20*pi/180).evalf()
# +
# EXTRA
# +
import numpy as np
import matplotlib.pyplot as plt
import math
def rotMatrix(angle):
c = np.cos(np.radians(angle))
s = np.sin(np.radians(angle))
return np.array([[c, -s], [s, c]])
v = np.array([0,1])
v30 = rotMatrix(30).dot(v)
plt.arrow(0,0,v[0],v[1], head_width=0.1, head_length=0.1)
plt.arrow(0,0,v30[0],v30[1],head_width=0.1, head_length=0.1)
v = np.array([1,0])
v30 = rotMatrix(30).dot(v)
plt.arrow(0,0,v[0],v[1], head_width=0.1, head_length=0.1)
plt.arrow(0,0,v30[0],v30[1],head_width=0.1, head_length=0.1)
plt.axis([-2,2,-2,2])
plt.grid
plt.axis('square')
plt.show()
# -
| PythonCodes/Utilities/Sympy/RotationMatrix.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # List
# # Basic Lists
# ## Bullets
#
# - b1
# - b2
# - b3
# ## Bullets
#
# - b1
# - b2
# - b3
# ## List
#
# 1. l1
# 1. l2
# 1. l3
# ## Numbered
#
# 1. a numbered list
# 1. and a second item
# 1. and a third item
# # Nested Lists
# ## Bullets
#
# - b1
# - c1
# - c2
# - b2
# - d1
# - d2
# - b3
# ## Bullets
#
# - this is
# - a list
# - with a nested list
# - and some subitems
# - and here the parent list continues
# ## Mixed Lists
#
# Currently mixed lists require 4 spaces of indentation
#
# 1. bla
#
# - foo1
# - foo 2
#
# 1. bla2
#
# - foo
#
# 1. bla3
#
#
# Using 2 spaces of indentation is returned as a block_quote and is an issue that needs to be fixed
# for compliance with RST spec.
#
# 1. bla
#
#
# > - foo1
# - foo 2
#
#
#
# 1. bla2
#
#
# > - foo
#
#
#
# 1. bla3
# ## Malformed Lists that seem to work in HTML
#
# - first item
# - second item
#
# - a sub item that is spaced with four spaces rather than two
# ## Complex Lists with Display Math
#
# Here is a computational algorithm:
#
# 1. Start with a guess for the value for $ \Phi $, then use the
# first-order conditions and the feasibility conditions to compute
# $ c(s_t), n(s_t) $ for $ s \in [1,\ldots, S] $ and
# $ c_0(s_0,b_0) $ and $ n_0(s_0, b_0) $, given $ \Phi $
# - these are $ 2 (S+1) $ equations in $ 2 (S+1) $ unknowns
# 1. Solve the $ S $ equations for the $ S $ elements of $ \vec x $
#
# $$
# u_{c,0}
# $$
#
# 1. Find a $ \Phi $ that satisfies
#
# $$
# u_{c,0} b_0 = u_{c,0} (n_0 - g_0) - u_{l,0} n_0 + \beta \sum_{s=1}^S \Pi(s | s_0) x(s) \tag{1}
# $$
#
# by gradually raising $ \Phi $ if the left side of [(1)](#equation-bellman2cons)
# exceeds the right side and lowering $ \Phi $ if the left side is less than the right side
# 1. After computing a Ramsey allocation, recover the flat tax rate on
# labor.
# ## Complex Lists with Code Blocks
#
# - foo, and then in your REPL run
# + [markdown] hide-output=false
# ```julia
# @assert x == 2
# ```
#
# -
#
# - and another item in the list. Hopefully this list item can be continued in the next markdown block in the notebook
#
# - and a sublist looks like this
# - second elements
#
#
#
# **Note:** This doesn’t work well with enumerated lists as the markdown engine will start a new
# list in the subsequent markdown cells (within Jupyter). So when you have mixed code blocks in lists
# then it is best to use unordered lists.
| tests/base/ipynb/lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DtL9-HYd33kZ"
# # **Padding and Stride**
# + id="wmBSs5023wIx"
import torch
import torch.nn as nn
# Takes convolution operation and applies it to X
def comp_conv2d(conv2d, X):
# Add two extra empty dimensions to X
X = X.reshape((1, 1) + X.shape)
Y = conv2d(X)
return Y.reshape(Y.shape[2:])
# + [markdown] id="HU0EfPZ74Dsp"
# ## **Padding**
# + id="vHN5b0wP4HU_" outputId="0b273876-a955-4540-bc88-02011196c5a0" colab={"base_uri": "https://localhost:8080/", "height": 34}
conv2d = nn.Conv2d(1, 1, kernel_size=3, padding=1)
X = torch.randn(size=(8, 8))
# Padding of 1 leaves shape unchanged for 3x3 convolution
comp_conv2d(conv2d, X).shape
# + [markdown] id="_qg3wvIp4WAY"
# ## **Stride**
# + id="K4KvgjcM4YE9" outputId="6b9ec385-e0f5-4dbd-9413-d4cba3ca4948" colab={"base_uri": "https://localhost:8080/", "height": 34}
conv2d = nn.Conv2d(1, 1, kernel_size=3, padding=1, stride=2)
comp_conv2d(conv2d, X).shape
# + [markdown] id="jkFFeoId4eDb"
# ## **Asymmetric kernels, padding and strides**
# We can use different strides, different kernel sizes and different padding for height and
# width. This can be used, e.g. to adjust the size to a desired shape (4:3 to 1:1).
# + id="c4CTk-AU4oSk" outputId="991abe4a-2588-4de2-9d78-066878016aa5" colab={"base_uri": "https://localhost:8080/", "height": 34}
# pad only vertically and use different strides on the 8x8 image
conv2d = nn.Conv2d(1, 1, kernel_size=(3, 5), padding=(0, 1), stride=(3, 4))
comp_conv2d(conv2d, X).shape
| L11 Convolutional Networks/L11_5_Padding_and_Stride.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# The standard fare:
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
from astropy.io import fits
import astropy.stats as stat
from photutils import aperture_photometry, CircularAperture, CircularAnnulus, DAOStarFinder
# -
# # Lab 9 - Aperture Photometry with Python Photutils
#
# ## 9.1 - Getting Started
#
# The goal of this lab is to teach you how to extract aperture photometry for all of the stars in a given image. Although the extraction itself is automated, there is some art to setting the parameters for extraction, and this lab is designed to teach you to choose these in an intelligent, data-driven way.
#
# To start we need to read in a raw image to work with.
#as an example, let's load in a randomly-selected raw Smith R band image
image = fits.getdata('M52-007_R.fit')
# We will be using a python package called photutils, which is based on an old IRAF package of the same name. One of the key functions within this package is the DAOStarFinder function, which we'll just call "the star finder" here.
#
# The star finder will extract sources, defined as some multiple (which you provide) of the background/sky level, so first we need to get a reasonable estimate of the background level. This is done using the function mad_std from the astropy.stats package, as below.
from astropy.stats import mad_std
bkg_sigma = mad_std(image)
print(bkg_sigma)
print(np.std(image))
# ### Exercise 1. Use the resources (docstrings, wikipedia, other functions) at your disposal to answer the following questions in the cell below.
# 1. What does the mad_std function do?
# 2. How is it different from the more typical np.std function? How different are the answers in these two cases, and why?
# ***Your answers go here***
# ## 9.2 - Extracting Stars
#
# The star finder requires two parameters to extract stars in the image:
#
# 1) The threshhold, which we will define as some number of background levels, above which we will call something a star
# 2) An estimate for the FWHM of point-sources (stars) in the image
#
# ### Exercise 2
# To start, estimate the FWHM of the stars in your image using pyraf's imexam functions, as you did in Lab 8. Measure the FWHM for at least 10 stars and average them. In the cell below, paste the imexam output and calculate the average of the measurements in the cell below that. Insert your calculated average FWHM in the third cell below.
# ***insert imexam output here***
# +
#insert calculation of average FWHM here
# -
#FWHM= this is a placeholder. INSERT YOUR VALUE IN PLACE OF 10 BELOW
FWHM=10
# We also need to set the threshhold (described above) for star finder, which we define as some multiple (nsigma) of the background level. To start, let's set nsigma to 10, meaning that in order for somehting to be considered a star, it must have at least 10x the detector counts of the background level.
#
# The next several lines below set up the parameters for the star finder (by specifying the FWHM and threshhold parameters), apply the star finder to the image, and then extract and print the number of sources.
nsigma=10
daofind = DAOStarFinder(fwhm=FWHM, threshold=nsigma*bkg_sigma)
sources = daofind(image)
nstars=len(sources)
nstars
# To check how well we're doing here, we need to be able to see how the sources that were automatically extracted line up with apparent sources in the image. To do so, we are going to write the information that star finder extracted from the sources it found into a DS9 region file, so that we can load it with the image. DS9 region files are text files where each line contains the following basic info:
# regiontype xcen ycen FWHM
#
# The code below writes the relevant outpt from daofind into a text file with this format.
xpos = np.array(sources['xcentroid'])
ypos = np.array(sources['ycentroid'])
f = open('M52_R.reg', 'w') #you will need to change the first input if you want to write to a different filename later
for i in range(0,len(xpos)):
f.write('circle '+str(xpos[i])+' '+str(ypos[i])+' '+str(FWHM)+'\n')
f.close()
# To display this region file, you should open the science image in DS9, then click Region --> Load Regions and navigate to the .reg file above. When you load it, you will see green circles appear on top of all of the stars that the Star Finder has extracted. Place a screenshot of this overlay in the cell below.
# ***DS9 screenshot goes here***
# ### Exercise 3
#
# Using the **median-combined V and R images that you generated for Homework 9**, answer the following questions. Include code, screenshots, etc. to support your argument, and add cells below as needed to do calculations, generate new region files, etc.
# 1) How many sources can you extract at V and R for nsigma=10 from your median-combined images? How much does the number of sources vary between the two wavelengths and why, do you think?
# 2) How different are the number of extracted stars in the raw R image you used in the example vs. the reduced and median combined R image that you generated for your homework? Name at least one potential reason why they are different, and find an example in the images that shows it.
# *Hint: An example really means a source that was identified in one image and not the other. Remember you can load multiple images in DS9, and can load a separate region file in each. Zoom in on your discrpant source. To match the zoom level and location between the two images, select Frame --> Match --> Frame --> Physical.*
# 3) For one of your images (V or R), discuss how the number of extracted sources changes when you change the nsigma threshhold. Make an argument based on the images for what you think the most reasonable limit is for this data.
# #### 1. Problem 1 explanation and images go here
# +
#problem 1 code goes here
# -
# #### 2. Problem 2 explanation and images go here
# +
#problem 2 code goes here
# -
# #### 3. Problem 3 explanation and images go here
# +
#problem 3 code goes here
# -
# ### 9.3 - Aperture Photometry
#
# The next step is to actually extract the photometry, and here too there is some art to choosing parameters. Although photutils will extract the photometry for each star in an automated way, you need to intelligently choose the parameters based on your data.
#
# The tunable parameters are:
# 1. the aperture radius inside of which to count the flux from the star
# 2. the inner and outer radius of the sky aperture. The annulus defined by these two numbers needs to be large enough to get a good measurement of the background level, but small enough to generally avoid confusion with other nearby sources.
#
# We'll start with some potentially reasonable values for these parameters.
aprad = 8
skyin=10
skyout=15
# ### Exercise 4
#
# For each of the next two cells, write a comment describing what each line is doing in the line above it.
#add comments to this cell
starapertures = CircularAperture((xpos,ypos),r=aprad)
skyannuli = CircularAnnulus((xpos,ypos),r_in=skyin,r_out=skyout)
phot_apers = [starapertures, skyannuli]
phot_table = aperture_photometry(image,phot_apers)
phot_table
#add comments to this cell
bkg_mean = phot_table['aperture_sum_1']/skyannuli.area()
bkg_starap_sum = bkg_mean * starapertures.area()
final_sum = phot_table['aperture_sum_0']-bkg_starap_sum
phot_table['background subtracted star counts'] = final_sum
phot_table
# ### Exercise 5 -
#
# Spend the rest of the lab time investigating what changes about the photometric measurement (background subtracted sky counts caluclated column) when you adjust the tunable parameters (aperture radius and inner/outer sky annulus) and report your findings below. You may wish to examine only a handful of stars in the table and avoid pr (to print just a few rows of a table, see the example below), but make sure the rows you select include stars with a range of brightnesses. You may also wish to make separate versions/copies of the table with different aperture parameters so that you can compare without overwriting. Think in particular about crowded fields and see if you can derive the best parameters for this case as well (identify things in the table with very close values for xcenter and ycenter to find these crowded regions).
#example of printing just a few rows in a table - chosen to be one bright star, one faint
phot_table[6:8]
#
| Labs/Lab9/Lab9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Open Data from various sources
# +
# import os
# os.environ['http_proxy'] = "usw04px100-restrictedproxy.tycoelectronics.net:80"
# os.environ['https_proxy'] = "usw04px100-restrictedproxy.tycoelectronics.net:80"
# -
#pip install knoema
import knoema
knoema.get('WBPEP2018Oct', timerange='1960-2019', frequency='A', Country='CN', Series='SP.POP.TOTL')
| 2019_conovarius/Population_Knoema.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9bOI4wnm9n31" colab_type="code" colab={}
# !git clone https://github.com/luweishuang/ConveRT-pytorch.git
# %cd ConveRT-pytorch
# !pip install -r requirements.txt
# + id="TyMFXSKr-bK2" colab_type="code" colab={}
# !python train.py
| docs/train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Analyzing a random sequence
#
# In this notebook, we will implement and query the following probabilistic model of a random sequence in SPPL.
# 1. For $i = 0$, $X[0]$ is a standard normal.
# 2. For $i > 0$
# - if $X[i-1]$ is negative, then $X[i]$ is a standard normal
# - otherwise, let $W[i]$ be the outcome of a fair coin:
# - if $W[i] = 0$, then $X[i] = 0.5*(X[i-1])^2 + X[i-1]$
# - othewrise, $X[i]$ is a standard normal.
# %load_ext sppl.magics
# **Defining the model in SPPL.**
# +
# %%sppl random_sequence
from fractions import Fraction
X = array(3)
W = array(3)
X[0] ~= norm()
for i in range(1, 3):
if X[i-1] > 0:
X[i] ~= norm(loc=0, scale=1)
W[i] ~= atomic(loc=0)
else:
W[i] ~= bernoulli(p=0.5)
if W[i] == 0:
X[i] ~= Fraction(1,2)*X[i-1]**2 + X[i-1]
else:
X[i] ~= norm(loc=0, scale=1)
# -
# **SPML programs are represented as sum-product expressions.<br/>
# We can view the SPE that corresponds to the `random_sequence` model.**
# %sppl_to_graph random_sequence
# It is informative to consider the distribution encoded by the graphical structure above.
#
# 1. `+` (sum) nodes indicate a probabilistic mixture model over its children, with weights along the edges.
# 2. `x` (proudct) nodes indicate that the children are independent.
# 3. Leaf nodes indicate names and distributions of the base random variables in the model.
# - Dashed edges and shaded nodes indicate deterministic transforms of random variables, with the symbol shown along the edge and the expression shown in the node label.
# **Executing the model returns a namespace `ns` that contains all the variables in the model.**
# ns = %sppl_get_namespace random_sequence
random_sequence = ns.random_sequence
# **Suppose we observe that X[1] > 0.<br/>
# What is the posterior probability that X[0] > 0?**
#
# We can use `condition` and `logprob` queries from **SPQL** to answer this question.
# +
# Condition `random_sequence` on the observation X[1] > 0.
random_sequence_given_X1 = random_sequence.condition(ns.X[1] > 0)
# Print the prior and posterior probabilities of X[0] > 0.
print('Pr(X[0]>0)\t\t= %1.2f' % (random_sequence.prob(ns.X[0]>0),))
print('Pr(X[0]>0 | X[1]>0)\t= %1.3f' % (random_sequence_given_X1.prob(ns.X[0]>0),))
# -
# **Suppose we observe that X[2] > 0.<br/>
# What is the posterior probability that X[0] > 0?**
#
# We can use `condition` and `logprob` queries from **SPQL** to answer this question.
# +
# Condition `random_sequence` on the observation X[2] > 0.
random_sequence_given_X2 = random_sequence.condition(ns.X[2] > 0)
# Print the prior and posterior probabilities of X[0] > 0.
print('Pr(X[0]>0)\t\t= %1.2f' % (random_sequence.prob(ns.X[0]>0),))
print('Pr(X[0]>0 | X[2]>0)\t= %1.3f' % (random_sequence_given_X2.prob(ns.X[0]>0),))
# -
# **Notice that observing (X[1]>0) shifts the probability of (X[0]>0) from 0.5 to 0.647,<br/>
# whereas observing (X[2]>0) shifts the probability of (X[0]>0) from 0.5 to 0.544.**
#
# Intuitively, X[1] contains 'more information' about X[0] as compared to X[2], since X[2] occurs later in the randoms sequence.<br/>
# We can use `mutual_infomration` to make this intuition quantitative.<br/><br/>
# The observation X[1]>0 has 10x more information about X[0]>0 as compared to the observation X[2]>0.
print('MI(X[0]>0 : X[1]>0)\t= %1.3f' % (random_sequence.mutual_information(ns.X[0]>0, ns.X[1]>0),))
print('MI(X[0]>0 : X[2]>0)\t= %1.3f' % (random_sequence.mutual_information(ns.X[0]>0, ns.X[2]>0),))
# **Note that the output of `random_sequence.condition(X[1] > 0)` is just another SPE.**
#
# Look how much smaller the conditioned SPE is!<br/>
# In effect, we have analytically (symbolically) eliminated all the exections of `random_sequence` where X[1] < 0.
# %sppl_to_graph random_sequence_given_X1
# **Similarly, the output of `random_sequence.condition(X[2] > 0)` is also another SPE.**
#
# In this case the SPE has the same structure, but the numeric probabiliites on the edges have changed.<br/>
# In general `.condition` can grow or shrink the underlying SPE, depending on the complexity of the query and SPE.
# %sppl_to_graph random_sequence_given_X2
| examples/random-sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# +
import pandas as pd
import fastai
df = pd.read_csv("downloads/40k_balanced_pm_acl.csv").sample(frac=1)
df = df.dropna(subset=["labels"])
df["text_nomask"] = df["text"] #backup
print(len(df))
df_test = df[:300]
df = df[300:]
fastai.__version__, len(df)
# -
df["text_length"]= df["text"].map(lambda t: len(t.split(" ")))
df = df[df["text_length"] > 7]
# +
from octoflow.core import replace_outof_vocab_words, vocab30k
df["text"] = df["text"].map(lambda t: replace_outof_vocab_words(t, vocab30k))
df_test["text"] = df_test["text"].map(lambda t: replace_outof_vocab_words(t,vocab30k))
# -
df["text"] = df["text"].astype(str)
df["labels"] = df["labels"].astype(int)
df = df[(df["labels"] == 1) | (df["labels"] == 0)]
df["lab"] = df["labels"] #worst fastai bug in history
# +
from fastai.text.all import *
dls = TextDataLoaders.from_df(df, text_col='text', label_col='lab')
dls.show_batch()
learn = text_classifier_learner(dls, AWD_LSTM, drop_mult=0.5, metrics=[accuracy, FBeta(beta=1)])
# -
# +
learn.fine_tune(2, 1e-2)
learn.show_results()
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_top_losses(3)
interp.plot_confusion_matrix()
learn.predict("We discuss the implications of a more 'civic-minded journalism.")
# -
learn.fine_tune(2, 1e-2)
learn.export('oct15_40k.pkl')
learn.save("oc15_40k")
# ## Sanity Check: Tough Cases
# 0 = non-problem
# 1 = problem
learn.predict("Anaphylaxis is a life-threatening emergency of which reliable epidemiological data are lacking")
#('1', TensorText(1), TensorText([0.0353, 0.9647]))
# +
from fastai.vision.all import *
def predict_batch(self, item, rm_type_tfms=None, with_input=False):
dl = self.dls.test_dl(item, rm_type_tfms=rm_type_tfms, num_workers=0)
ret = self.get_preds(dl=dl)
return ret
learn.predict_batch = predict_batch
# -
predicts = df_test["text"].map(learn.predict)
df_test["predicted"] = [int(p[0]) for p in predicts]
df_test["confidence_0"] = [round(p[2][0].item(),2) for p in predicts]
df_test["confidence_1"] = [round(p[2][1].item(),2) for p in predicts]
df_test.to_csv("downloads/oct15_40k_test300.csv")
print("accuracy in test set:", len(df_test[df_test["predicted"] == df_test["labels"]])/len(df_test))
predicts2 = df_test["text_nomask"].map(learn.predict)
df_test["predicted2"] = [int(p[0]) for p in predicts2]
print("accuracy in test set with no masking:", len(df_test[df_test["predicted2"] == df_test["labels"]])/len(df_test))
# That's very weird ... masking doesn't effect predictions
| 03_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings("ignore")
# data processing
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_absolute_error, mean_squared_error
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
# arima et arixmax
from pmdarima import auto_arima
# regression regularisée.
from numpy import arange
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RepeatedKFold
from sklearn.linear_model import Lasso,Ridge
# methodes d'ensembles
import xgboost as xgb
import lightgbm as lgb
from hyperopt import hp, fmin, tpe, Trials, STATUS_OK
## deep learning
from keras import optimizers
from keras.utils.vis_utils import plot_model
from keras.models import Sequential, Model
from keras.layers.convolutional import Conv1D, MaxPooling1D
from keras.layers import Dense, LSTM, RepeatVector, TimeDistributed, Flatten
import tensorflow as tf
adam = tf.optimizers.Adam()
# Tensorflow est une bibliothèque open-source développée par l’équipe Google Brain qui l’utilisait initialement en interne. Elle implémente des méthodes d’apprentissage automatique basées surle principe des réseaux de neurones profonds (deep learning).
# Keras est une librairie Python qui encapsule l’accès aux fonctions proposées par plusieurs librairies de machine learning, en particulier Tensorflow.
# Une'optimisation d'Adam est une méthode de descente de gradient stochastique basée sur l'estimation adaptative des moments de premier et de second ordre.
# +
sales_df=pd.read_pickle('intermed/sales_full.sav')
sales_df.head()
exogenous_features=['Is_holidays','seasonal_week_index','blackFriday2013', 'blackFriday2012','blackFriday2011',
'quarter', 'month',
'dayofyear','dayofmonth',
'event_value','isMonthEnd','isMonthStart',
'total_amt_mean_lag3','total_amt_mean_lag7','total_amt_mean_lag10',
'total_amt_std_lag3','total_amt_std_lag7','total_amt_std_lag10',
'PRCP','TAVG','TMAX' ,'TMIN']
# +
df1=sales_df.groupby(['year','seasonal_week_index']).sum()['total_amt'].reset_index()
col_lag=['total_amt_mean_lag3','total_amt_mean_lag7','total_amt_mean_lag10','total_amt_std_lag3','total_amt_std_lag7','total_amt_std_lag10','PRCP', 'TAVG', 'TMAX', 'TMIN']
df2=sales_df.groupby(['year','seasonal_week_index']).mean()[col_lag].reset_index()
temp=df1.merge(df2, on =['year','seasonal_week_index'])
col_calendar=['Is_holidays','blackFriday2013', 'blackFriday2012','blackFriday2011',
'quarter', 'month' , 'dayofyear','dayofmonth', 'weekofyear', 'event_value','isMonthEnd','isMonthStart']
calendar=sales_df.groupby(['year','seasonal_week_index']).max()[col_calendar].reset_index()
sales_df=temp.merge(calendar, on =['year','seasonal_week_index'])
# +
#centree et reduire les exogènes
sales_df[exogenous_features]= StandardScaler(with_mean=True,with_std=True).fit_transform(sales_df[exogenous_features])
# -
df_train = sales_df.iloc[:140,:]
df_test = sales_df.iloc[140:,:]
df_test
# ## Traditional Algorithms
# ### ARIMA
model = auto_arima(df_train.total_amt, trace=True, error_action="ignore", suppress_warnings=True)
model.fit(df_train.total_amt)
df_test["Forecast_ARIMA"] = model.predict(n_periods=len(df_test))
df_test[["total_amt", "Forecast_ARIMA"]].plot(figsize=(14, 7))
# ### ARIMAX
# +
# SARIMAX Model
sxmodel = auto_arima(sales_df[['total_amt']], exogenous=sales_df[exogenous_features],
start_p=1, start_q=1,
test='adf',
max_p=5, max_q=5,
start_P=0, seasonal=True,
d=None, D=1, trace=True,
error_action='ignore',
suppress_warnings=True,
stepwise=True)
sxmodel.summary()
# -
df_test["Forecast_ARIMAX"] = sxmodel.predict(n_periods=len(df_test), exogenous=df_test[exogenous_features])
df_test[["total_amt", "Forecast_ARIMAX"]].plot(figsize=(14, 7))
# ## Statistical Learning
# ### LightGBM
params = {"objective": "regression"}
dtrain = lgb.Dataset(df_train[exogenous_features], label=df_train.total_amt)
dvalid = lgb.Dataset(df_test[exogenous_features])
model_lgb = lgb.train(params, train_set=dtrain)
forecast = model_lgb.predict(df_test[exogenous_features])
df_test["Forecast_LGM"] = forecast
df_test[["total_amt","Forecast_LGM"]].plot(figsize=(14, 7))
# ### XGBoost
X_train, y_train = df_train[exogenous_features], df_train['total_amt']
X_test, y_test = df_test[exogenous_features], df_test['total_amt']
df_train[exogenous_features]
reg = xgb.XGBRegressor(n_estimators=1000)
reg.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_test, y_test)],
early_stopping_rounds=50, #stop if 50 consequent rounds without decrease of error
verbose=False) # Change ver
xgb.plot_importance(reg, height=0.9)
df_test["Forecast_XGB"] = reg.predict(df_test[exogenous_features])
df_test[["total_amt","Forecast_XGB"]].plot(figsize=(14, 7))
# ### XGBOOSTING tunning
# +
# Define the function hyperopt must optimise
def objective(params):
num_round = int(params['n_estimators'])
xgb_reg = xgb.train(params, X_train_matrix, num_round)
y_pred = xgb_reg.predict(X_test_matrix)
rmse = np.sqrt(mean_squared_error(y_test, y_pred))
return rmse
# Make DMatrix for calling function, to optimise speed of training
X_train_matrix = xgb.DMatrix(X_train, label=y_train)
X_test_matrix = xgb.DMatrix(X_test)
# +
# Définir l'espace d'optimisation
space = {
'verbosity': 0, # Valid values are 0 (silent), 1 (warning), 2 (info), 3 (debug).
'max_depth': hp.choice('max_depth', np.arange(4, 10, 1, dtype=int)), # Maximum depth of a tree. L'augmentation de cette valeur rendra le modèle plus complexe et plus susceptible de sur-ajuster. 0 n'est accepté dans la politique de croissance guidée par perte que lorsque tree_method est défini sur hist et qu'il n'indique aucune limite de profondeur. Méfiez-vous que XGBoost consomme agressivement de la mémoire lors de la formation d'un arbre profond. intervalle: [0,∞]
'n_estimators': hp.choice('n_estimators', np.arange(5, 50, 1, dtype=int)),
'colsample_bytree': hp.quniform('colsample_bytree', 0.3, 0.7, 0.05), # colsample_bytree, colsample_bylevel, colsample_bynode [default=1]. Il s'agit d'une famille de paramètres pour le sous-échantillonnage des colonnes. colsample_bytree est le rapport de sous-échantillon des colonnes lors de la construction de chaque arbre. Le sous-échantillonnage a lieu une fois pour chaque arbre construit. Tous les paramètres colsample_by* ont une plage de (0, 1], la valeur par défaut de 1, et spécifient la fraction de colonnes à sous-échantillonner.
'min_child_weight': hp.choice('min_child_weight', np.arange(0, 5, 1, dtype=int)), # Minimum sum of instance weight (hessian) needed in a child. Si l'étape de partition de l'arborescence aboutit à un nœud feuille avec la somme du poids de l'instance inférieure à min_child_weight, le processus de construction abandonnera le partitionnement supplémentaire. Dans le cas de régression linéaire, cela correspond simplement au nombre minimum d'instances nécessaires pour chaque nœud. Plus min_child_weight est grand, plus l'algorithme sera conservateur. range: [0,∞]
'subsample': hp.quniform('subsample', 0.3, 1, 0.05), # Subsample ratio of the training instances. Le régler sur 0,5 signifie que XGBoost échantillonnerait au hasard la moitié des données d'entraînement avant de faire pousser des arbres, ce qui évitera le surajustement. Le sous-échantillonnage se produira une fois dans chaque itération de boosting. intervalle: (0,1]
'gamma': hp.quniform('gamma', 140, 180, 2), # Réduction minimale des pertes requise pour créer une partition supplémentaire sur un nœud feuille de l'arbre. Plus le gamma est grand, plus l'algorithme sera conservateur. intervalle = [0,∞]
'learning_rate': hp.quniform('learning_rate', 0.1, 0.6, 0.05),# alias: eta. Default is 0.3. Range is [0,1]. Réduction de taille de pas utilisée dans la mise à jour pour éviter le surajustement. Après chaque étape de boosting, nous pouvons directement obtenir le poids des nouvelles features, et eta réduit le poids des features pour rendre le processus de boosting plus conservateur.
'objective': 'reg:squarederror',
'lambda': hp.quniform('lambda', 10, 20, 1), # regularisation L2 des terms poids.. L'augmentation de cette valeur rendra le modèle plus conservateur.
'alpha': hp.quniform('alpha', 50, 60, 1), # regularisation L1 des terms poids. L'augmentation de cette valeur rendra le modèle plus conservateur.
'eval_metric': 'rmse',
'X_train_matrix': X_train_matrix,
'X_test_matrix': X_test_matrix,
'y_test': y_test
}
# +
# Exécuter l'optimisation hyperopttrials = Trials()
best = fmin(
fn=objective, # objective function
space=space, # parameter space
algo=tpe.suggest, # surrogate algorithm
max_evals=200, # no. of evaluations
trials=Trials() # trials object that keeps track of the sample results (optional). In order to run with trails the output of the objective function has to be a dictionary including at least the keys 'loss' and 'status' which contain the result and the optimization status respectively.
)
# Imprimer les paramètres optimisés
print("Best = ", best)
# -
reg = xgb.XGBRegressor(alpha=54.0,
reg_lambda = 19,
colsample_bytree= 0.30000000000000004,
gamma= 154,
learning_rate= 0.55,
max_depth=5,
min_child_weight= 4,
n_estimators= 20,
subsample= 0.55)
reg.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_test, y_test)],
eval_metric="rmse",
early_stopping_rounds=50, #stop if 50 consequent rounds without decrease of error
verbose=False) # Change ver
xgb.plot_importance(reg, height=0.9)
df_test["Forecast_XGB_tune"] = reg.predict(df_test[exogenous_features])
df_test[["total_amt","Forecast_XGB_tune"]].plot(figsize=(14, 7))
# ## Ridge and Lasso Regression
# +
# load the dataset
data = sales_df.values
X, y = data[:, :-1], data[:, -1]
# define model
model_ridge = Ridge()
model_lasso = Lasso()
# define model evaluation method
cv = RepeatedKFold(n_splits=10, n_repeats=3, random_state=1)
# define grid
grid = dict()
grid['alpha'] = arange(0, 1, 0.001)
# define search
search_ridge = GridSearchCV(model_ridge, grid, scoring='neg_mean_absolute_error', cv=cv, n_jobs=-1)
search_lasso = GridSearchCV(model_lasso,grid, scoring='neg_mean_absolute_error', cv=cv, n_jobs=-1)
# perform the search
results_ridge = search_ridge.fit(X, y)
results_lasso = search_lasso.fit(X, y)
# summarize
print('MAE Ridge: %.3f' % results_ridge.best_score_)
print('Config: %s' % results_ridge.best_params_)
print('MAE Lasso: %.3f' % results_lasso.best_score_)
print('Config: %s' % results_lasso.best_params_)
# -
results_ridge
# +
# define model
model_lasso= Lasso(alpha=0.07)
model_ridge= Ridge(alpha=0.003)
# fit model
model_lasso.fit(X_train, y_train)
model_ridge.fit(X_train, y_train)
#forecast
df_train["Forecast_Lasso_train"] = model_lasso.predict(df_train[exogenous_features])
df_test["Forecast_Lasso"] = model_lasso.predict(df_test[exogenous_features])
df_train["Forecast_Ridge_train"] = model_lasso.predict(df_train[exogenous_features])
df_test["Forecast_Ridge"] = model_ridge.predict(df_test[exogenous_features])
# -
#df_test[["Forecast_Lasso","Forecast_Ridge"]].plot(figsize=(14, 7))
df_test[["total_amt","Forecast_Ridge"]].plot(figsize=(14, 7))
df_test[["total_amt","Forecast_Lasso"]].plot(figsize=(14, 7))
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return round(np.mean(np.abs((y_true - y_pred) / y_true)) ,3)
mean_absolute_percentage_error(df_test["total_amt"],df_test["Forecast_Ridge"])
mean_absolute_percentage_error(df_test["total_amt"],df_test["Forecast_Lasso"])
mean_absolute_percentage_error(df_test["total_amt"],df_test["Forecast_XGB"])
mean_absolute_percentage_error(df_test["total_amt"],df_test["Forecast_XGB_tune"])
mean_absolute_percentage_error(df_test["total_amt"],df_test["Forecast_LGM"])
# ## Deep Leraning Algorithms
epochs = 40 # epochs est le nombre maximum d’itérations ;
batch = 256 # batch_size correspond aux nombre d’observationsque l’on fait passer avant de remettre à jour les poids synaptiques
#lr = 0.0003
# Dans cette section, nous passons à un perceptron multicouche. Nous créons toujours une
# structure Sequential, dans lequel nous ajoutons successivement deux objets Dense :
# 1. le premier fait la jonction entre la couche d’entrée (d’où l’option input_dim indiquant le nombre de variablesprédictives) et la couche cachée qui comporte (units = 3) neurones
#
# ==> fonction d’activation Sigmoides
#
# 2. le second entre cette la dernière couche cachée et la sortie à un seul neurone (units = 1).
#
# ==> fonction d’activation RelU
# ### MLP
# Ici notre modèle aura des caractéristiques d'entrée égales 0 window size
#
# Le problème avec les modèles MLP est que le modèle ne prend pas l'entrée comme des données séquencées,il ne fait que recevoir des INPUTS et ne les traite pas comme des données séquencées, cela peut être un problème puisque le modèle ne le fera pas voir les données avec le modèle de séquence qu'il a.
# +
#instanciation du modèle
model_mlp = Sequential()
##architecture
model_mlp.add(Dense(50, activation="sigmoid", input_dim=X_train.shape[1]))
model_mlp.add(Dense(1,activation='relu'))
#compilation - algorithme d'apprentissage
model_mlp.compile(loss='mse', optimizer=adam)
model_mlp.summary()
# -
#Apprentissage
mlp_history = model_mlp.fit(X_train.values, y_train, validation_data=(X_test.values, y_test), epochs=epochs, verbose=2)
# ### CNN
#
# Les CNN sont moins couteux en termes de calcul que les RNN : CNN apprend par lots tandis que les RNN s'entraînent de manière séquentielle.
# Les CNN ne supposent pas que l'historique est complet : contrairement aux RNN, les CNN apprennent des modèles dans la fenêtre temporelle. Si vous avez des données manquantes, les CNN devraient être utiles.
# D'une certaine manière, les CNN peuvent regarder vers l'avenir : les modèles RNN n'apprennent des données qu'avant le pas de temps qu'ils doivent prédire. Les CNN (avec shuffling) peuvent voir les données d'un point de vue plus large
#
# Il existe quatre types de couches pour un réseau de neurones convolutif :
# 1. **la couche de convolution** : Son but est de repérer la présence d'un ensemble de features dans les images reçues en entrée. Pour cela, on réalise un filtrage (layer) par convolution ,
# 2. **la couche de pooling** :Ce type de couche est souvent placé entre deux couches de convolution : elle reçoit en entrée plusieurs feature maps, et applique à chacune d'entre elles l'opération de pooling. En cas d'images, consiste à réduire la taille des images, tout en préservant leurs caractéristiques importantes. La couche de pooling permet de réduire le nombre de paramètres et de calculs dans le réseau. On améliore ainsi l'efficacité du réseau et on évite le sur-apprentissage.Les valeurs maximales sont repérées de manière moins exacte dans les feature maps obtenues après pooling que dans celles reçues en entrée
# Ainsi, la couche de pooling rend le réseau moins sensible à la position des features : le fait qu'une feature se situe un peu plus en haut ou en bas, ou même qu'elle ait une orientation légèrement différente ne devrait pas provoquer un changement radical dans la classification de l'image.
# 3. **la couche Flatten lorsque nous est pas dans un 2D** comme c est le cas avec les imaes
# 4. **la couche fully-connected** : constitue toujours la dernière couche d'un réseau de neurones, convolutif ou non – elle n'est donc pas caractéristique d'un CNN. Ce type de couche reçoit un vecteur en entrée et produit un nouveau vecteur en sortie. Pour cela, elle applique une combinaison linéaire puis éventuellement une fonction d'activation aux valeurs reçues en entrée
# La couche convolutivet être capable d'identifier la séquence temporelles dans les données
X_train_series = X_train.values.reshape((X_train.shape[0], X_train.shape[1], 1))
X_valid_series = X_test.values.reshape((X_test.shape[0], X_test.shape[1], 1))
print('Train set shape', X_train_series.shape)
print('Validation set shape', X_valid_series.shape)
X_train_series
# ### CNN
#instanciation du modèle
model_cnn = Sequential()
##architecture
model_cnn.add(Conv1D(filters=64, kernel_size=2, activation='relu', input_shape=(X_train_series.shape[1], X_train_series.shape[2])))
model_cnn.add(MaxPooling1D(pool_size=2))
model_cnn.add(Flatten())
model_cnn.add(Dense(50, activation='relu'))
model_cnn.add(Dense(1))
#compilation - algorithme d'apprentissage
model_cnn.compile(loss='mse', optimizer=adam)
model_cnn.summary()
cnn_history = model_cnn.fit(X_train_series, y_train, validation_data=(X_valid_series, y_test), epochs=epochs, verbose=2)
# ### LSTM
# LSTM voit réellement les données d'entrée comme une séquence
#
# Il est donc capable d'apprendre des modèles à partir de données séquencées (en supposant qu'elles existent) mieux que les autres,en particulier les modèles à partir de longues séquences.
model_lstm = Sequential()
model_lstm.add(LSTM(50, activation='relu', input_shape=(X_train_series.shape[1], X_train_series.shape[2])))
model_lstm.add(Dense(1))
model_lstm.compile(loss='mse', optimizer=adam)
model_lstm.summary()
lstm_history = model_lstm.fit(X_train_series, y_train, validation_data=(X_valid_series, y_test), epochs=epochs, verbose=2)
# ### CNN-LSTM
# "The benefit of this model is that the model can support very long input sequences that can be read as blocks or subsequences by the CNN model, then pieced together by the LSTM model."
#
# For more informations : https://machinelearningmastery.com/how-to-get-started-with-deep-learning-for-time-series-forecasting-7-day-mini-course/
subsequences = 2
timesteps = X_train_series.shape[1]//subsequences
X_train_series_sub = X_train_series.reshape((X_train_series.shape[0], subsequences, timesteps, 1))
X_valid_series_sub = X_valid_series.reshape((X_valid_series.shape[0], subsequences, timesteps, 1))
print('Train set shape', X_train_series_sub.shape)
print('Validation set shape', X_valid_series_sub.shape)
model_cnn_lstm = Sequential()
model_cnn_lstm.add(TimeDistributed(Conv1D(filters=64, kernel_size=1, activation='relu'), input_shape=(None, X_train_series_sub.shape[2], X_train_series_sub.shape[3])))
model_cnn_lstm.add(TimeDistributed(MaxPooling1D(pool_size=2)))
model_cnn_lstm.add(TimeDistributed(Flatten()))
model_cnn_lstm.add(LSTM(50, activation='relu'))
model_cnn_lstm.add(Dense(1))
model_cnn_lstm.compile(loss='mse', optimizer=adam)
cnn_lstm_history = model_cnn_lstm.fit(X_train_series_sub, y_train, validation_data=(X_valid_series_sub, y_test), epochs=epochs, verbose=2)
cnn_lstm_history = model_cnn_lstm.fit(X_train_series_sub, y_train, validation_data=(X_valid_series_sub, y_test), epochs=epochs, verbose=2)
mlp_train_pred = model_mlp.predict(X_train.values)
mlp_valid_pred = model_mlp.predict(X_test.values)
print('Train mape:', mean_absolute_percentage_error(y_train,mlp_train_pred))
print('Test mape:', mean_absolute_percentage_error(y_test,mlp_valid_pred))
# cnn_train_pred = model_cnn.predict(X_train_series)
# cnn_valid_pred = model_cnn.predict(X_valid_series)
# print('Train mape:', mean_absolute_percentage_error(y_train,cnn_train_pred))
# print('Test mape:', mean_absolute_percentage_error(y_test,cnn_valid_pred))
lstm_train_pred = model_lstm.predict(X_train_series)
lstm_valid_pred = model_lstm.predict(X_valid_series)
print('Train mape:', mean_absolute_percentage_error(y_train,lstm_train_pred))
print('Test mape:', mean_absolute_percentage_error(y_test,lstm_valid_pred))
cnn_lstm_train_pred = model_cnn_lstm.predict(X_train_series_sub)
cnn_lstm_valid_pred = model_cnn_lstm.predict(X_valid_series_sub)
print('Train mape:', mean_absolute_percentage_error(y_train,cnn_lstm_train_pred))
print('Test mape:', mean_absolute_percentage_error(y_test,cnn_lstm_valid_pred))
# ## Conclusion
# Comme on devrait s y attrendre les modèles linéaires et modèles ensemblent sont très performantes par rapport aux algorithmes de deep learning. Cela est du en partie à la taille du datasets
#
# Entre les modèles ensemblistes et les modèles linéaires, nous choisirons les modèles linéaires vu la faciliter dans d'une part l'interprétabilité (possibilt" d'avoir l' ADN des ventes et d'autre part le deploiement du modèles.
#
# ==> Continuons avec du Ridge
| 03_Forecasting/03_modeling/modeling_test_weekly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LIME (Local Interpretable Model-agnostic Explanations)
#
# The original dataset contains 1000 entries with 20 categorial/symbolic attributes prepared by Prof. Hofmann. In this dataset, each entry represents a person who takes a credit by a bank. Each person is classified as good or bad credit risks according to the set of attributes.
#
# This version of dataset can be found at https://www.kaggle.com/uciml/german-credit
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import MinMaxScaler
from sklearn.impute import SimpleImputer
from sklearn.model_selection import train_test_split
# Read data
app_train = pd.read_csv("data/credits.csv")
app_train
# -
# ## Preprocess and balancing data
#
# I will not go too much into preprocessing, but it is quite important to mention that the original data was highly imbalanced, so I needed discard lots of data to restore balance.
# +
# Iterate through the columns to encode features
for col in app_train:
if app_train[col].dtype == 'object':
# If 2 or fewer unique categories
if len(list(app_train[col].unique())) <= 2:
le = LabelEncoder()
# Train on the training data
le.fit(app_train[col])
# Transform both training and testing data
app_train[col] = le.transform(app_train[col])
# Split data
X = app_train.drop(["TARGET"], axis=1)
y = app_train["TARGET"].to_numpy()
# one-hot encoding of categorical variables
X = pd.get_dummies(X)
features = X.columns
imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
# Scale each feature to 0-1
scaler = MinMaxScaler(feature_range = (0, 1))
# Transform training data
X = imputer.fit_transform(X)
X = scaler.fit_transform(X)
# Balancing data
X_0 = X[y == 0]
X_1 = X[y == 1]
y_0 = y[y == 0]
y_1 = y[y == 1]
X_0 = X_0[0:X_1.shape[0], :]
y_0 = y_0[0:X_1.shape[0]]
X = np.vstack([X_0, X_1])
y = np.hstack([y_0, y_1])
# Train test split data and convert to pytorch's tensors
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
print(f"Size of training data: {X_train.shape}. Testing data {X_test.shape}")
# -
# # Cross validation with 10 splits
#
# Cross Validation is a very useful technique for assessing the effectiveness of your model, particularly in cases where you need to mitigate overfitting. I would recommend this technique unless you are training a neural network.
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import KFold
from sklearn.metrics import accuracy_score
def cross_validation(X, y):
'''
K-Folds Cross-Validation
'''
kf = KFold(n_splits=10, shuffle=True, random_state=42)
results = []
for train_index, test_index in kf.split(X):
classifier = RandomForestClassifier(n_estimators=50, n_jobs=-1, random_state=42)
_X_train, _X_test = X[train_index], X[test_index]
_y_train, _y_test = y[train_index], y[test_index]
classifier.fit(_X_train, _y_train)
y_hat = classifier.predict(_X_test)
results.append(accuracy_score(_y_test, y_hat))
return np.mean(results), np.std(results)
results = cross_validation(X_train, y_train)
print(f"Cross validation on training with Accuracy of {results[0]} ± {results[1]}")
# -
# # Train the RBF on the whole training dataset
#
# For LIME to work, we need a pair of complex and simple model. First of all we need to train the complex model.
#
# The final result looks quite good, a bit of performance optimization could yield even better accuracy, but state of the art result is not our target, right now.
# +
from sklearn.metrics import confusion_matrix
import seaborn as sns
# Train complex model on whole dataset
classifier = RandomForestClassifier(n_estimators=50, n_jobs=-1, random_state=42)
classifier.fit(X_train, y_train)
# Validate result
validation = confusion_matrix(classifier.predict(X_test), y_test)
df_cm = pd.DataFrame(validation, index = ["true", "false"],
columns = ["true", "false"])
plt.figure(figsize=(10, 10))
plt.title("Confusion matrix of complex's model predictions")
sns.heatmap(df_cm, annot=True,cmap='Blues', fmt='g')
plt.show()
# -
# # Examine the model with LIME
#
# Our model works well, but which feature does it actually use for its decision? What is important and what not? Let us find out
# +
from sklearn.linear_model import LogisticRegression
# Graphic's parameters
size=15
params = {'legend.fontsize': 'large',
'figure.figsize': (20,8),
'axes.labelsize': size,
'axes.titlesize': size,
'xtick.labelsize': size*0.75,
'ytick.labelsize': size*0.75,
'axes.titlepad': 25}
plt.rcParams.update(params)
def lime(x, f, N, variance):
"""
Lime
x: sample which we want to examine locally
f: complex model, could be any,
N: how many samples do we wish to examine around the original
variance: how much variability do we want around the original
returns parameters of the simple model
"""
covariance_matrix = np.identity(x.shape[0]) * variance
# take sample from multivariate gauss
x_equiv = np.random.multivariate_normal(mean=x, cov=covariance_matrix, size=N)
# Predict sample with the complex model
y_hat = f.predict(x_equiv).astype(np.float64).reshape(x_equiv.shape[0])
# Fit simple model with input x and prediction y_hat of complex model
model = LogisticRegression(solver='saga', fit_intercept=False, random_state=42, max_iter=1000)
model.fit(x_equiv, y_hat)
# Return simple's model parameters
return list(zip(list(model.coef_[0, :]), list(features)))
# Pick a sample from test dataset
sample = X_test[1, :]
# Let lime examine how our complex model worked on this sample
linear_model = lime(sample, classifier, 1000, 10)
linear_model = sorted(linear_model, key=lambda item: np.abs(item[0]), reverse=True)[:20]
coefs = [feature[0] for feature in linear_model][::-1]
names = [feature[1] for feature in linear_model][::-1]
# Graphic plots
plt.figure(figsize=(15, 15))
plt.title("Features' importance for prediction of the sample")
plt.barh(names, coefs, label="Feature's coefficient of linear model", alpha=0.75)
plt.xlabel("Coefficients' values")
plt.legend()
plt.rcParams['axes.facecolor'] = 'red'
plt.rcParams['savefig.facecolor'] = 'white'
plt.savefig("output.png")
plt.show()
# -
# # Verify LIME's output with ONOVA
#
# Just to be sure
# +
from scipy import stats
models = []
for i in range(0, 100):
linear_model = lime(sample, classifier, 1000, 10)
coefs = [feature[0] for feature in linear_model]
models.append(coefs)
stats.f_oneway(*models)
# -
# # Interpreting LIME's output
#
# The variable `SK_ID_CURR` plays a great role for the decision of our RBF. The newer the customer, the less chance his/her application will get approved.
#
# `FLAG_OWN_CAR` and `OWN_CAR_AGE` give away how important your car is for your credit institute, basicly you should own a car, but it should not be too old.
#
# We now try to remove `SK_ID_CURR` and see how the model performs. As the graphics below show, the performance of the model dropped significantly.
#
# Conclusion: This very complex model depends too much on a single variable and could as well replaced by a much simpler model.
# +
# Set every ID to zero
X_train[:, 0] = 0
classifier = RandomForestClassifier(n_estimators=50, n_jobs=-1, random_state=42)
classifier.fit(X_train, y_train)
validation = confusion_matrix(classifier.predict(X_test), y_test)
df_cm = pd.DataFrame(validation, index = ["true", "false"],
columns = ["true", "false"])
plt.figure(figsize=(10, 10))
plt.title("Confusion matrix of complex's model predictions without `SK_ID_CURR`.")
sns.heatmap(df_cm, annot=True,cmap='Blues', fmt='g')
plt.show()
| Explain Random Forrest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
from jenga.tasks.income import IncomeEstimationTask
task = IncomeEstimationTask(seed=42)
# -
model = task.fit_baseline_model()
f"The ROC AUC score on the test data is {task.get_baseline_performance()}"
task.test_data
# ### Missing values in the 'age' column
# +
from jenga.corruptions.generic import MissingValues
age_corruption = MissingValues(column='age', fraction=0.8, na_value=-1.0)
corrupted_test_data = age_corruption.transform(task.test_data)
y_pred = model.predict_proba(corrupted_test_data)
f"The ROC AUC score on the corrupted test data is {task.score_on_test_data(y_pred)}"
# -
# Cleaning via mean imputation
mean_age = np.mean(task.train_data.age.values)
mean_age
clean_test_data = corrupted_test_data.copy(deep=True)
clean_test_data.loc[clean_test_data['age'] < 0, 'age'] = mean_age
clean_test_data
# +
y_pred_cleaned = model.predict_proba(clean_test_data)
f"The ROC AUC score on the corrupted test data is {task.score_on_test_data(y_pred_cleaned)}"
# -
# ### Missing values in 'marital_status'
# +
from jenga.corruptions.generic import MissingValues
marital_status_corruption = MissingValues(column='marital_status', fraction=0.8, na_value=np.nan)
# +
corrupted_test_data = marital_status_corruption.transform(task.test_data)
y_pred = model.predict_proba(corrupted_test_data)
f"The ROC AUC score on the corrupted test data is {task.score_on_test_data(y_pred)}"
# -
corrupted_test_data
# Cleaning via mode imputation
task.train_data['marital_status'].value_counts()
clean_test_data = corrupted_test_data.copy(deep=True)
clean_test_data['marital_status'].fillna('Married-civ-spouse', inplace=True)
clean_test_data
# +
y_pred_cleaned = model.predict_proba(clean_test_data)
f"The ROC AUC score on the corrupted test data is {task.score_on_test_data(y_pred_cleaned)}"
| src/jenga/notebooks/impute-missing-values-siks-lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sergiomarchena16/lab_02_spam/blob/main/lab2_SPAM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Gi_d4qQqv0AE"
# # Laboraotrio 2 - Deteccion de SPAM (NLP)
# + [markdown] id="w-4wyaG8Sgzg"
# ## PARTE 1: INGENIERIRA DE CARACTERISTICAS
# + [markdown] id="czpM1v8sQZzT"
# ### importaciones
# + id="g2f6AkrrvG3w"
import pandas as pd
import numpy as np
import nltk
import unicodedata
import re
from sklearn.feature_extraction.text import CountVectorizer
import re
from sklearn.feature_extraction.text import TfidfVectorizer
# + [markdown] id="fmJFg-9wmnBR"
# ### Datasets y arreglos necesarios
# + id="orq1VuJUvUyt"
dataset1 = pd.read_csv('/content/drive/MyDrive/UVG /2022/SECURITY DATA SCIENCE/Lab02_SPAM/completeSpamAssassin.csv')
dataset2 = pd.read_csv('/content/drive/MyDrive/UVG /2022/SECURITY DATA SCIENCE/Lab02_SPAM/enronSpamSubset.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 693} id="rDE26cbjwRhZ" outputId="adf8e2f6-e8f7-499a-d4c0-9d391f805dfc"
dataset1
# + colab={"base_uri": "https://localhost:8080/", "height": 780} id="WBu0Ae8nwmRi" outputId="7d0fce87-1bbd-4e59-dafe-f3604847362a"
dataset2
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="IB0rPYtxwwMm" outputId="5fb783b2-aaba-4df9-c033-58adc79ea189"
# comparing Unnamed 0 and Unnamed 0.1 to see if they are the same
dataset2[['Unnamed: 0', 'Unnamed: 0.1']]
# + colab={"base_uri": "https://localhost:8080/"} id="sos_SAXuxE6Q" outputId="a6cf5975-24c5-4fbf-9d4b-cc433f623d4b"
dataset2['Unnamed: 0'].describe(), dataset2['Unnamed: 0.1'].describe()
# they are the same indeed :)
# + colab={"base_uri": "https://localhost:8080/", "height": 780} id="FwQDVX1qyKJ_" outputId="22db6b81-5fef-4bf4-e2a6-4ee4135518da"
# merging the 2 datasets
frames = [dataset1, dataset2]
df = pd.concat(frames)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 745} id="6ueNmzXfyXd1" outputId="7db44d74-8512-49cc-f01e-7a6f395427c7"
# creating the final df with body and label only
df_final = df[['Body','Label']]
df_final
# + colab={"base_uri": "https://localhost:8080/"} id="Enk6VU3HKkkC" outputId="9f855fac-99c5-431d-a75d-4174e4b38434"
# info about the final dataset
df_final.info()
# + id="Nwuk7EJvzQGV" colab={"base_uri": "https://localhost:8080/", "height": 745} outputId="3c4cc6da-ca8e-44fb-9950-98376fc8c601"
# resetting the index
df_final = df_final[['Body', 'Label']]
df_final
# + colab={"base_uri": "https://localhost:8080/"} id="CtqTa7amtLhT" outputId="2dbffd41-bb17-4d6c-8dd5-8f81f0f31be5"
# final description of the final df
df_final.info()
# + [markdown] id="qrbWajdFu-Q_"
# ### Preprocesamiento
# + colab={"base_uri": "https://localhost:8080/"} id="URbKYGZst3d7" outputId="48e12c4a-cd0f-46e4-ab05-482704afd8e9"
import nltk
nltk.download('punkt')
nltk.download('wordnet')
nltk.download('stopwords')
nltk.download('averaged_perceptron_tagger')
# + colab={"base_uri": "https://localhost:8080/", "height": 406} id="CwEhG1bUvNtA" outputId="eedbb6e7-2481-40af-8d01-646fcef1bddb"
# lowercase all body sentences
df_final['Body'] = df_final['Body'].str.lower()
df_final.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 406} id="sMBejmzNM8bA" outputId="df2e8f02-aa04-4843-9602-1d17fb91bd26"
df_final['Body'] = df_final['Body'].str.strip()
df_final.head()
# + id="jwYsAj4GNaUl"
# remove accents function
def removerAcentos(texto):
texto = unicodedata.normalize('NFKD', str(texto)).encode('ascii','ignore').decode('utf-8','ignore')
return texto
# + id="38RUvNFMNpqV"
# remove accents
df_final['Body'] = df_final['Body'].apply(removerAcentos)
# + id="0kN8eCh1xdtJ"
# remove special characters, numbers and symbols function
def removerCaracteresEspecialesNumerosSimbolos(texto, removerDigitos = False):
patron = r'[^a-zA-Z0-9\s]' if not removerDigitos else r'[^a-zA-Z\s]'
texto = re.sub(patron, '', texto, re.I|re.A)
return texto
# + id="yN_wxAbJ05fa" colab={"base_uri": "https://localhost:8080/", "height": 406} outputId="5eb59f31-ef75-42b6-d223-0ba3cecee96c"
df_final['Body'] = df_final['Body'].apply(removerCaracteresEspecialesNumerosSimbolos)
df_final.head()
# + colab={"base_uri": "https://localhost:8080/"} id="31GA6v-oOTap" outputId="1467c655-cb17-4840-8b35-ec3166d6b1b1"
# !pip install contractions
# !pip install textsearch
# + id="rZ0BnfUx1sG_" colab={"base_uri": "https://localhost:8080/", "height": 406} outputId="837fe51b-3817-4e09-8a00-c61bdeac011a"
import contractions
df_final['Body'] = df_final['Body'].apply(contractions.fix)
df_final.head()
# + id="RfctbyFfOLfi"
# removing stop words
stop_words = nltk.corpus.stopwords.words('english')
def normalize_document(doc):
# tokenize document
tokens = nltk.word_tokenize(doc)
# filter stopwords out of document
filtered_tokens = [token for token in tokens if token not in stop_words]
# re-create document from filtered tokens
doc = ' '.join(filtered_tokens)
return doc
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="yjVK7s6GOw8r" outputId="0a796e6b-13b9-40a5-d715-fbde2bbe6b10"
df_final['Body'] = df_final['Body'].apply(normalize_document)
df_final.head()
# + [markdown] id="7hcD6Du7QMLq"
# ### Representacion de Texto
# + colab={"base_uri": "https://localhost:8080/"} id="KWEXoiPAO-p9" outputId="fa13e1fa-86d9-46ad-d2c3-c6851f6c012a"
cv = CountVectorizer(min_df=0.1, max_df=0.9)
cv_matrix = cv.fit_transform(df_final['Body'])
cv_matrix = cv_matrix.toarray()
cv_matrix
# + colab={"base_uri": "https://localhost:8080/"} id="exIA5tKQQQVp" outputId="24bc6137-e178-4e74-a26d-5427ed4085be"
# Palabras únicas del corpus
vocabulario = cv.get_feature_names()
# Mostrar el vector
df_cv = pd.DataFrame(cv_matrix, columns=vocabulario)
df_cv.head
# + id="-phoC79KQpws"
# bag of words n = 1
df_cv.to_csv('BoW.csv', index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="1R6TR9zmRI0w" outputId="12fe237b-eebd-4152-e6cc-bcccc3f9c7cd"
bv = CountVectorizer(ngram_range=(1,2), min_df=0.1, max_df=0.9)
bv_matrix = bv.fit_transform(df_final['Body'])
bv_matrix = bv_matrix.toarray()
vocabulario = bv.get_feature_names()
df_bv = pd.DataFrame(bv_matrix, columns=vocabulario)
df_bv.head()
# + id="Zi3n0wzYRyx2"
# bi-gram
df_bv.to_csv('bi-gram.csv', index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="8G9gaZfMSDBu" outputId="eebc8d80-14a2-463c-e7da-9828014ded9a"
tv = TfidfVectorizer(min_df=0.1, max_df=0.9, use_idf=True)
tv_matrix = tv.fit_transform(df_final['Body'])
tv_matrix = tv_matrix.toarray()
vocabulario = tv.get_feature_names()
df_tv = pd.DataFrame(np.round(tv_matrix, 2), columns=vocabulario)
df_tv.head()
# + id="isMmHiwpSNdu"
# TF-ID
df_tv.to_csv('TF-ID.csv', index=False)
# + id="Jjv57sdbSNUf"
# + [markdown] id="DIQ67Zt8SnzJ"
#
# + [markdown] id="4wc1vpl5SpDk"
# ## PARTE 2: IMPLEMENTACION DEL MODELO
# + [markdown] id="pZ3lVpENSwJG"
# ### Separacion de los datos
# + [markdown] id="KP_-omBuS8Xh"
# #### importaciones y carga de datasets
# + id="qa-0sKZlSs_u"
# libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn import metrics, model_selection, tree
from sklearn.naive_bayes import MultinomialNB
# + id="xznDY8tFTKQd"
# DATASETS
df1 = pd.read_csv('/content/drive/MyDrive/UVG /2022/SECURITY DATA SCIENCE/Lab02_SPAM/completeSpamAssassin.csv')
df2 = pd.read_csv('/content/drive/MyDrive/UVG /2022/SECURITY DATA SCIENCE/Lab02_SPAM/enronSpamSubset.csv')
bow = pd.read_csv('BoW.csv')
bigram = pd.read_csv('bi-gram.csv')
tf_id = pd.read_csv('TF-ID.csv')
# + id="Z0MlkCz-S_wf"
# original df
frames = [df1, df2]
df_final = pd.concat(frames)
# + [markdown] id="e9pJRHXUUSgV"
# #### Train and Test
# + id="WtkcAoseT4B-"
# train test split for every dataset (70% train, 30% test)
bow_train, bow_test, bow_target_train, bow_target_test = model_selection.train_test_split(bow, df_final['Label'], test_size=0.3, random_state=123)
bigram_train, bigram_test, bigram_target_train, bigram_target_test = model_selection.train_test_split(bigram, df_final['Label'], test_size=0.3, random_state=123)
tf_id_train, tf_id_test, tf_id_target_train, tf_id_target_test = model_selection.train_test_split(tf_id, df_final['Label'], test_size=0.3, random_state=123)
# + [markdown] id="mcBT3cUjUWwR"
# ### Implementacion de los modelos
# + [markdown] id="9wsA0SvqUis0"
# #### MODELO 1 - BOW
# + colab={"base_uri": "https://localhost:8080/"} id="iBMeCbKNULMC" outputId="0f8ce6af-0cb1-48a2-a639-2ececaf6ccb5"
# MULTINOMIAL NAIVE BAYES FOR BOW
nb_bow = MultinomialNB()
nb_bow.fit(bow_train, bow_target_train)
# + colab={"base_uri": "https://localhost:8080/"} id="dW18VBalVD4U" outputId="2277269b-cf81-4ea1-be49-6859d60f2242"
# PREDICTION AND METRICS
pred_bow = nb_bow.predict(bow_test)
print(metrics.accuracy_score(bow_target_test, pred_bow))
print('Matriz de confusion', metrics.confusion_matrix(bow_target_test, pred_bow))
print(metrics.classification_report(bow_target_test, pred_bow, target_names=['0', '1']))
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="DwDDfphRVH1O" outputId="a094752b-5729-4b02-86da-7b5f7aeb751b"
#PLOT
metrics.plot_roc_curve(nb_bow, bow_test, bow_target_test)
plt.show()
# + [markdown] id="8yXBQAqfVW_E"
# #### MODELO 2 - BIGRAMS
# + colab={"base_uri": "https://localhost:8080/"} id="I53EQWvMVROU" outputId="011fb53a-b024-48ef-9760-037d2284505e"
# NAIVE BAYES FOR BIGRAMS
nb_bigram = MultinomialNB()
nb_bigram.fit(bigram_train, bigram_target_train)
# + colab={"base_uri": "https://localhost:8080/"} id="5VZJfUADXbEF" outputId="7904e768-e2a4-4735-a629-3935649df084"
# prediction for bigrams
pred_bigram = nb_bigram.predict(bigram_test)
print(metrics.accuracy_score(bigram_target_test, pred_bigram))
print('Matriz de confusion', metrics.confusion_matrix(bigram_target_test, pred_bigram))
print(metrics.classification_report(bigram_target_test, pred_bigram, target_names=['0', '1']))
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="dWL0VfYHXuNg" outputId="126b60d8-e3a3-4ecd-8a81-5ea6dc7c6969"
# figure
metrics.plot_roc_curve(nb_bigram, bigram_test, bigram_target_test)
plt.show()
# + [markdown] id="_Lq2DiTXX8yf"
# #### MODELO 3 - TF-ID
# + colab={"base_uri": "https://localhost:8080/"} id="kl915guKX15C" outputId="0771b47e-50e6-4e09-eb51-9ddd09c4c1c9"
# NAIVE BAYES FOR TF-ID
nb_tf_id = MultinomialNB()
nb_tf_id.fit(tf_id_train, tf_id_target_train)
# + colab={"base_uri": "https://localhost:8080/"} id="i6Hv5ogIYE-W" outputId="275d8a3a-0885-4bb8-c8da-dfcfcf9dfa31"
# prediction for tf-id
pred_tf_id = nb_tf_id.predict(tf_id_test)
print(metrics.accuracy_score(tf_id_target_test, pred_tf_id))
print('Matriz de confusion', metrics.confusion_matrix(tf_id_target_test, pred_tf_id))
print(metrics.classification_report(tf_id_target_test, pred_tf_id, target_names=['0', '1']))
# + colab={"base_uri": "https://localhost:8080/", "height": 334} id="xRiNarlgYaRP" outputId="5febfa8e-f6e0-4e2e-d6b5-5c13911abe02"
# figure
metrics.plot_roc_curve(nb_tf_id, tf_id_test, tf_id_target_test)
plt.show()
| lab2_SPAM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright 2015 The TensorFlow Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
"""A deep MNIST classifier using convolutional layers.
See extensive documentation at
https://www.tensorflow.org/get_started/mnist/pros
"""
# Disable linter warnings to maintain consistency with tutorial.
# pylint: disable=invalid-name
# pylint: disable=g-bad-import-order
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import argparse
import sys
import tempfile
import numpy as np
from tensorflow.examples.tutorials.mnist import input_data
import matplotlib.pyplot as plt
import tensorflow as tf
import matplotlib.pyplot as plt
FLAGS = None
def deepnn(x):
"""deepnn builds the graph for a deep net for classifying digits.
Args:
x: an input tensor with the dimensions (N_examples, 784), where 784 is the
number of pixels in a standard MNIST image.
Returns:
A tuple (y, keep_prob). y is a tensor of shape (N_examples, 10), with values
equal to the logits of classifying the digit into one of 10 classes (the
digits 0-9). keep_prob is a scalar placeholder for the probability of
dropout.
"""
# Reshape to use within a convolutional neural net.
# Last dimension is for "features" - there is only one here, since images are
# grayscale -- it would be 3 for an RGB image, 4 for RGBA, etc.
with tf.name_scope('reshape'):
x_image = tf.reshape(x, [-1, 28, 28, 1])
# First convolutional layer - maps one grayscale image to 32 feature maps.
with tf.name_scope('conv1'):
W_conv1 = weight_variable([5, 5, 1, 32])
b_conv1 = bias_variable([32])
h_conv1 = tf.nn.relu(conv2d(x_image, W_conv1) + b_conv1)
# Pooling layer - downsamples by 2X.
with tf.name_scope('pool1'):
h_pool1 = max_pool_2x2(h_conv1)
# Second convolutional layer -- maps 32 feature maps to 64.
with tf.name_scope('conv2'):
W_conv2 = weight_variable([5, 5, 32, 64])
b_conv2 = bias_variable([64])
h_conv2 = tf.nn.relu(conv2d(h_pool1, W_conv2) + b_conv2)
# Second pooling layer.
with tf.name_scope('pool2'):
h_pool2 = max_pool_2x2(h_conv2)
# Fully connected layer 1 -- after 2 round of downsampling, our 28x28 image
# is down to 7x7x64 feature maps -- maps this to 1024 features.
with tf.name_scope('fc1'):
W_fc1 = weight_variable([7 * 7 * 64, 1024])
b_fc1 = bias_variable([1024])
h_pool2_flat = tf.reshape(h_pool2, [-1, 7*7*64])
h_fc1 = tf.nn.relu(tf.matmul(h_pool2_flat, W_fc1) + b_fc1)
# Dropout - controls the complexity of the model, prevents co-adaptation of
# features.
with tf.name_scope('dropout'):
keep_prob = tf.placeholder(tf.float32)
h_fc1_drop = tf.nn.dropout(h_fc1, keep_prob)
print (np.shape(h_fc1_drop))
# Map the 1024 features to 10 classes, one for each digit
with tf.name_scope('fc2'):
W_fc2 = weight_variable([1024, 10])
b_fc2 = bias_variable([10])
y_conv = tf.matmul(h_fc1_drop, W_fc2) + b_fc2
return y_conv, keep_prob
def conv2d(x, W):
"""conv2d returns a 2d convolution layer with full stride."""
return tf.nn.conv2d(x, W, strides=[1, 1, 1, 1], padding='SAME')
def max_pool_2x2(x):
"""max_pool_2x2 downsamples a feature map by 2X."""
return tf.nn.max_pool(x, ksize=[1, 2, 2, 1],
strides=[1, 2, 2, 1], padding='SAME')
def weight_variable(shape):
"""weight_variable generates a weight variable of a given shape."""
initial = tf.truncated_normal(shape, stddev=0.1)
return tf.Variable(initial)
def bias_variable(shape):
"""bias_variable generates a bias variable of a given shape."""
initial = tf.constant(0.1, shape=shape)
return tf.Variable(initial)
def plot(title, accuracy_arr):
plt.figure(figsize=(10,10))
plt.plot(accuracy_arr)
plt.title(title)
plt.xlabel('Batch Iterations')
plt.ylabel('Accuracy')
print("Average Accuracy {0}".format(accuracy_arr.mean()))
def main(_):
# Import data
mnist = input_data.read_data_sets(FLAGS.data_dir, one_hot=True)
# Create the model
x = tf.placeholder(tf.float32, [None, 784])
# Define loss and optimizer
y_ = tf.placeholder(tf.float32, [None, 10])
accuracy_iter_train = int(2000/100)
accuracy_iter = 400
accuracy_arr = np.zeros(accuracy_iter)
accuracy_arr_train = np.zeros(accuracy_iter_train)
# Build the graph for the deep net
y_conv, keep_prob = deepnn(x)
with tf.name_scope('loss'):
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=y_,
logits=y_conv)
cross_entropy = tf.reduce_mean(cross_entropy)
with tf.name_scope('adam_optimizer'):
train_step = tf.train.AdamOptimizer(1e-4).minimize(cross_entropy)
with tf.name_scope('accuracy'):
correct_prediction = tf.equal(tf.argmax(y_conv, 1), tf.argmax(y_, 1))
correct_prediction = tf.cast(correct_prediction, tf.float32)
accuracy = tf.reduce_mean(correct_prediction)
graph_location = tempfile.mkdtemp()
print('Saving graph to: %s' % graph_location)
train_writer = tf.summary.FileWriter(graph_location)
train_writer.add_graph(tf.get_default_graph())
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for i in range(2000):
batch = mnist.train.next_batch(50)
if i % 100 == 0:
accuracy_arr_train[int(i/100)] = accuracy.eval(feed_dict={
x: batch[0], y_: batch[1], keep_prob: 1.0})
print('step %d, training accuracy %g' % (i, accuracy_arr_train[int(i/100)]))
train_step.run(feed_dict={x: batch[0], y_: batch[1], keep_prob: 0.5})
plot("Train Accuracy after every 100 batch", accuracy_arr_train)
for i in range(accuracy_iter):
batch = mnist.test.next_batch(10)
accuracy_arr[i] = accuracy.eval(feed_dict={
x: batch[0], y_: batch[1], keep_prob: 1.0})
print('test accuracy %g' % accuracy_arr[i])
plot("Test Accuracy", accuracy_arr)
if __name__ == '__main__':
parser = argparse.ArgumentParser()
parser.add_argument('--data_dir', type=str,
default='/tmp/tensorflow/mnist/input_data',
help='Directory for storing input data')
FLAGS, unparsed = parser.parse_known_args()
tf.app.run(main=main)
# -
| ML_Assignment6/MNIST_conv2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data ingestion and ELT from object store
#
# NPS can load and unload data from object stores like Amazon S3 and IBM Cloud object store. This works by [using Netezza External Tables to read from and write to object store](https://www.ibm.com/support/knowledgecenter/SS5FPD_1.0.0/com.ibm.ips.doc/postgresql/load/c_load_loading_cloud.html).
#
# Lets take a look at a few examples; lets target an AWS S3 bucket. Prerequisites -
#
# - `NZ_USER`, `NZ_PASSWORD`, `NZ_HOST` environment variables are set to point to the right Netezza instance
# - `AWS_ACCESS_KEY_ID`, `AWS_SECRET_ACCESS_KEY` and `AWS_REGION` environment variables are set with the correct credentials
# - `BUCKET` environment variable has the correct bucket name
#
# _Note:_ One can configure the ACLs for the bucket on AWS like [this](https://www.ibm.com/support/knowledgecenter/en/SSTNZ3/com.ibm.ips.doc/postgresql/admin/adm_nps_cloud_provisioning_prereq_aws.html) to balance security and need for NPS to read/write to it.
#
#
# ## The sample data
#
# Lets use the [publically available Covid case data](https://github.com/owid/covid-19-data/tree/master/public/data) to highlight this NPS use case. For this example, the data has been put in an object store bucket (`$BUCKET`) under `/covid/owid-covid-data.csv`
import os, nzpy
import pandas as pd
con = nzpy.connect(user=os.environ['NZ_USER'], password=os.environ['NZ_PASSWORD'], host=os.environ['NZ_HOST'],
port=5480, database='system')
# There are two examples here.
#
# - Load data into the database and do some analysis
# - Use transient external tables to read (on the fly) data and perform some analysis
#
# The schema for the table is [published by OWID already](https://github.com/owid/covid-19-data/blob/master/public/data/owid-covid-codebook.csv).
# +
# Setup the table schema based on the json mentioned above
schema = '''
iso_code varchar (16),
continent varchar(48),
location varchar(48),
covid_date date,
total_cases numeric(32, 20),
new_cases numeric(32, 20),
new_cases_smoothed numeric(32, 20),
total_deaths numeric(32, 20),
new_deaths numeric(32, 20),
new_deaths_smoothed numeric(32, 20),
total_cases_per_million numeric(32, 20),
new_cases_per_million numeric(32, 20),
new_cases_smoothed_per_million numeric(32, 20),
total_deaths_per_million numeric(32, 20),
new_deaths_per_million numeric(32, 20),
new_deaths_smoothed_per_million numeric(32, 20),
new_tests numeric(32, 20),
total_tests numeric(32, 20),
total_tests_per_thousand numeric(32, 20),
new_tests_per_thousand numeric(32, 20),
new_tests_smoothed numeric(32, 20),
new_tests_smoothed_per_thousand numeric(32, 20),
tests_per_case numeric(32, 20),
positive_rate numeric(32, 20),
tests_units varchar(32),
stringency_index numeric(32, 20),
population numeric(32, 20),
population_density numeric(32, 20),
median_age numeric(32, 20),
aged_65_older numeric(32, 20),
aged_70_older numeric(32, 20),
gdp_per_capita numeric(32, 20),
extreme_poverty numeric(32, 20),
cardiovasc_death_rate numeric(32, 20),
diabetes_prevalence numeric(32, 20),
female_smokers numeric(32, 20),
male_smokers numeric(32, 20),
handwashing_facilities numeric(32, 20),
hospital_beds_per_thousand numeric(32, 20),
life_expectancy numeric(32, 20),
human_development_index numeric(32, 20)'''
# Read data on the fly and lets see if all is working well.
df = pd.read_sql(f'''
select unique(continent)
from external 'owid-covid-data.csv' ({schema})
using (
remotesource 'S3'
delim ','
uniqueid 'covid'
accesskeyid '{os.environ["AWS_ACCESS_KEY_ID"]}'
secretaccesskey '{os.environ["AWS_SECRET_ACCESS_KEY"]}'
defaultregion '{os.environ["AWS_REGION"]}'
bucketurl '{os.environ["BUCKET"]}'
skiprows 1
) where continent is not null and continent != '' ''', con)
df.columns = [c.decode().lower() for c in df.columns]
df
# -
# Ingest the data and do analysis and visualization
table = 'covid'
with con.cursor() as cur:
# drop any old table
r = cur.execute(f'select 1 from _v_table where tablename = ^{table}^')
if r.fetchall():
cur.execute(f'drop table {table}')
# create a table to load data
cur.execute(f'create table {table} ({schema})')
print(f"Table {table} created")
# load data from object store
cur.execute(f'''
insert into {table}
select * from external 'owid-covid-data.csv' ({schema})
using (
remotesource 'S3'
delim ','
uniqueid 'covid'
accesskeyid '{os.environ["AWS_ACCESS_KEY_ID"]}'
secretaccesskey '{os.environ["AWS_SECRET_ACCESS_KEY"]}'
defaultregion '{os.environ["AWS_REGION"]}'
bucketurl '{os.environ["BUCKET"]}'
skiprows 1
)''')
print(f"{cur.rowcount} Rows loaded")
# Get a week over week trend
df = pd.read_sql_query('''
select continent,
this_week(covid_date) as wk,
max(new_cases) as total
from covid
where
continent is not null and
continent != ''
group by wk, continent
order by wk, continent
''', con,
parse_dates = {b'WK': '%Y-%m-%d'})
df.columns = [c.decode().lower() for c in df.columns]
df.total = df.total.astype(float)
df.head()
# +
# Lets visualize the same
from mizani.formatters import date_format
from plotnine import *
( ggplot(df, aes(x='wk', y='total', color='continent')) + geom_line() + geom_point() +
labs(y = "Total cases", x = "Week") + facet_wrap('continent') +
scale_x_datetime(labels=date_format('%b %-d')) +
theme(axis_text_x=element_text(rotation=60, hjust=1))
)
# -
| nzpy/nzobjectstore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="Qhl-XDGhkAHl" colab_type="code" colab={}
# We import some necessary packages
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] id="nfNykyP0j_KG" colab_type="text"
# # Plotting Using Package ``matplotlib``
#
# In Python, one can use package ``matplotlib`` to handle most plotting tasks. Its interface is very similar to plotting in Matlab. For an instance, this is the code for plotting a simple sine function:
# + id="Ap-eBzgnj_Kl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 286} outputId="ed5d0bed-3c20-4458-dff5-3f7341e75432"
# We assign values from a certain range to x.
x = np.arange(0, 8, 0.1)
# We compute sin(x) for each x.
y = np.sin(x)
# We plot the resulting relationship.
plt.plot(x, y)
# + [markdown] id="d0dpZ43Mj_K6" colab_type="text"
# Axes can be labelled using ``plt.xlabel`` and ``plt.ylabel``.
# + id="cNhWFL4Vj_K-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="2b0b99f5-e6ba-4754-cced-78fb8bcbd0bd"
x = np.arange(0, 8, 0.1)
y = np.sin(x)
plt.plot(x, y)
# new code:
plt.xlabel("x")
plt.ylabel("y")
plt.legend(['sin(x)'])
# + [markdown] id="VxFlGlEEj_LJ" colab_type="text"
# We can also do other things, such as adding a grid using function ``plt.grid``:
# + id="R8a5hkZLj_LN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="b21c9a70-5269-4373-970d-e52eead2b27d"
x = np.arange(0, 8, 0.1)
y = np.sin(x)
plt.plot(x, y)
plt.xlabel("x")
plt.ylabel("y")
plt.legend(['sin(x)'])
# new code:
plt.grid(ls='--')
# + [markdown] id="16897hmgj_LW" colab_type="text"
# We can also specify other properties of the plot when calling ``plt.plot``, e.g. the line color (such as 'r' for red) or line style (such as '--' for a dashed line):
# + id="UglCv2Cij_LZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="ce41acec-cab6-4956-8ada-fe48737f231e"
x = np.arange(0, 8, 0.1)
y = np.sin(x)
plt.plot(x, y)
plt.xlabel("x")
plt.ylabel("y")
plt.legend(['sin(x)'])
plt.grid(ls='--')
# new code:
z = np.exp(0.1*x)
plt.plot(x, z, 'r--')
# + [markdown] id="SfxW9hSCj_Lk" colab_type="text"
# We update the legend – expression ``np.exp`` can be written using LaTeX notation, in which case it displays in the proper mathematical format:
# + id="SkGEguxCj_Lm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="228a16ad-caef-4fce-f9ae-ee90892b5959"
x = np.arange(0, 8, 0.1)
y = np.sin(x)
plt.plot(x, y)
plt.xlabel("x")
plt.ylabel("y")
plt.legend(['sin(x)'])
plt.grid(ls='--')
z = np.exp(0.1*x)
plt.plot(x, z, 'r--')
# new code:
plt.legend(['$\sin(x)$', '$e^{0.1x}$'])
# + [markdown] id="2b0tzNcfj_Lu" colab_type="text"
# # Further Functionality
#
# Package ``matplotlib`` also contains a huge number of other function and features – it really is very flexible. More information on these topics can be found in its [documentation](http://matplotlib.org/index.html). It is also possible to study the following [simple tutorial](http://matplotlib.org/users/pyplot_tutorial.html). There is also a [gallery of plots](http://matplotlib.org/gallery.html) for inspiration.
# + id="tuqlSghOj_Lx" colab_type="code" colab={}
| EN/p8_matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font size=-1>Licensed under the Apache License, Version 2.0 (the \"License\");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at [https://www.apache.org/licenses/LICENSE-2.0](https://www.apache.org/licenses/LICENSE-2.0)
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an \"AS IS\" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.</font>
# +
import yaml
# Set `PATH` to include the directory containing TFX CLI.
# PATH=%env PATH
# %env PATH=/home/jupyter/.local/bin:{PATH}
# -
# !python -c "import tfx; print('TFX version: {}'.format(tfx.__version__))"
# # Continuous training with TFX and Cloud AI Platform
# This lab walks you through a TFX pipeline that uses **Cloud Dataflow** and **Cloud AI Platform Training** as execution runtimes. The pipeline implements a typical TFX workflow as depicted on the below diagram:
#
#
# ## Understanding the pipeline design
# The pipeline source code can be found in the `pipeline` folder.
# %cd pipeline
# !ls -la
# The `config.py` module configures the default values for the environment specific settings and the default values for the pipeline runtime parameters.
# The default values can be overwritten at compile time by providing the updated values in a set of environment variables.
# !tail -n 15 config.py
# The `pipeline.py` module contains the TFX DSL defining the workflow implemented by the pipeline.
#
# The `preprocessing.py` module implements the data preprocessing logic the `Transform` component.
#
# The `model.py` module implements the training logic for the `Train` component.
#
# The `runner.py` module configures and executes `KubeflowDagRunner`. At compile time, the `KubeflowDagRunner.run()` method conversts the TFX DSL into the pipeline package in the [argo](https://argoproj.github.io/argo/) format.
#
# The `features.py` module contains feature definitions common across `preprocessing.py` and `model.py`.
#
# ## Building and deploying the pipeline
#
# You will use TFX CLI to compile and deploy the pipeline. As explained in the previous section, the environment specific settings can be provided through a set of environment variables and embedded into the pipeline package at compile time.
#
# ### Configure environment settings
#
# Update the below constants with the settings reflecting your lab environment.
#
# - `GCP_REGION` - the compute region for AI Platform Training and Prediction
# - `ARTIFACT_STORE` - the GCS bucket created during installation of AI Platform Pipelines. The bucket name starts with the `hostedkfp-default-` prefix.
# - `ENDPOINT` - set the `ENDPOINT` constant to the endpoint to your AI Platform Pipelines instance. Then endpoint to the AI Platform Pipelines instance can be found on the [AI Platform Pipelines](https://console.cloud.google.com/ai-platform/pipelines/clusters) page in the Google Cloud Console.
#
# 1. Open the *SETTINGS* for your instance
# 2. Use the value of the `host` variable in the *Connect to this Kubeflow Pipelines instance from a Python client via Kubeflow Pipelines SKD* section of the *SETTINGS* window.
# +
GCP_REGION = 'us-central1'
ENDPOINT = '315252b57cfb9312-dot-us-central2.pipelines.googleusercontent.com'
ARTIFACT_STORE_URI = 'gs://hostedkfp-default-l2iv13wnek'
PROJECT_ID = !(gcloud config get-value core/project)
PROJECT_ID = PROJECT_ID[0]
# -
# ### Compile the pipeline
#
# You can build and upload the pipeline to the AI Platform Pipelines instance in one step, using the `tfx pipeline create` command. The `tfx pipeline create` goes through the following steps:
# - (Optional) Builds the custom image to that provides a runtime environment for TFX components,
# - Compiles the pipeline DSL into a pipeline package
# - Uploads the pipeline package to the instance.
#
# As you debug the pipeline DSL, you may prefer to first use the `tfx pipeline compile` command, which only executes the compilation step. After the DSL compiles successfully you can use `tfx pipeline create` to go through all steps.
# #### Set the pipeline's compile time settings
#
# The pipeline can run using a security context of the GKE default node pool's service account or the service account defined in the `user-gcp-sa` secret of the Kubernetes namespace hosting Kubeflow Pipelines. If you want to use the `user-gcp-sa` service account you change the value of `USE_KFP_SA` to `True`.
#
# Note that the default AI Platform Pipelines configuration does not define the `user-gcp-sa` secret.
USE_KFP_SA=False
DATA_ROOT_URI = 'gs://workshop-datasets/covertype/small'
CUSTOM_TFX_IMAGE = 'gcr.io/{}/lab-02-tfx-image'.format(PROJECT_ID)
PIPELINE_NAME = 'tfx_covertype_continuous_training'
MODEL_NAME = 'tfx_covertype_classifier'
RUNTIME_VERSION = '2.1'
PYTHON_VERSION = '3.7'
# %env PROJECT_ID={PROJECT_ID}
# %env KUBEFLOW_TFX_IMAGE={CUSTOM_TFX_IMAGE}
# %env ARTIFACT_STORE_URI={ARTIFACT_STORE_URI}
# %env DATA_ROOT_URI={DATA_ROOT_URI}
# %env GCP_REGION={GCP_REGION}
# %env MODEL_NAME={MODEL_NAME}
# %env PIPELINE_NAME={PIPELINE_NAME}
# %env RUNTIME_VERSION={RUNTIME_VERSION}
# %env PYTHON_VERIONS={PYTHON_VERSION}
# %env USE_KFP_SA={USE_KFP_SA}
# !tfx pipeline compile --engine kubeflow --pipeline_path runner.py
# ### Deploy the pipeline package to AI Platform Pipelines
#
# After the pipeline code compiles without any errors you can use the `tfx pipeline create` command to perform the full build and deploy the pipeline.
#
# !tfx pipeline create \
# --pipeline_path=runner.py \
# --endpoint={ENDPOINT} \
# --build_target_image={CUSTOM_TFX_IMAGE}
# If you need to redeploy the pipeline you can first delete the previous version using `tfx pipeline delete` or you can update the pipeline in-place using `tfx pipeline update`.
#
# To delete the pipeline:
#
# `tfx pipeline delete --pipeline_name {PIPELINE_NAME} --endpoint {ENDPOINT}`
#
# To update the pipeline:
#
# `tfx pipeline update --pipeline_path runner.py --endpoint {ENDPOINT}`
# ### Create and monitor a pipeline run
# After the pipeline has been deployed, you can trigger and monitor pipeline runs using TFX CLI or KFP UI.
#
# To submit the pipeline run using TFX CLI:
# !tfx run create --pipeline_name={PIPELINE_NAME} --endpoint={ENDPOINT}
# To list all active runs of the pipeline:
# !tfx run list --pipeline_name {PIPELINE_NAME} --endpoint {ENDPOINT}
# To retrieve the status of a given run:
# +
RUN_ID='[YOUR RUN ID]'
# !tfx run status --pipeline_name {PIPELINE_NAME} --run_id {RUN_ID} --endpoint {ENDPOINT}
# -
| workshops/tfx-caip-tf21/lab-02-tfx-pipeline/lab-02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import the modules
import pandas as pd
import numpy as np
from scipy import stats
import math
from statsmodels.stats.weightstats import ztest
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error, mean_squared_error
from sklearn.metrics import r2_score
# ### Load the dataset
#
# - Load the train data and using all your knowledge of pandas try to explore the different statistical properties of the dataset.
# read the dataset and extract the features and target separately
train = pd.read_csv('train.csv')
train
train.info()
# +
random_key = 6
X = train[['Age','Overall','Potential','Wage (M)']]
y = train['Value (M)']
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.3, random_state=random_key)
# -
X_train
# ### Visualize the data
#
# - Check for the categorical & continuous features.
# - Check out the best plots for plotting between categorical target and continuous features and try making some inferences from these plots.
# - Check for the correlation between the features
# +
# Code Starts here
def show_boxplot(col_data, x_label, y_label, title, fig_size=(7, 7), show_outliers=True):
"""
Shows boxplot with means
Params:
-------
col_data: list or numpy array
x_label: str
y_label: str
title: str
fig_size: tupe of (int, int)
show_outliers: bool
"""
fig = plt.figure(figsize=fig_size)
plt.boxplot(col_data, showmeans=True, showfliers=show_outliers)
plt.title(title, fontsize=21, color='navy')
plt.xlabel(x_label)
plt.ylabel(y_label)
plt.show()
for col in X_train.select_dtypes(include=np.number).columns:
x_label = col
y_label = 'Distribution'
data = X_train[col]
title = f'Distribution for {col}'
show_boxplot(col_data=data, x_label=x_label, y_label=y_label, title=title)
# Code ends here
# -
sns.heatmap(X_train.corr())
# +
upper_threshold = 0.5
lower_threshold = -0.5
# List the correlation pairs
correlation = train.corr().unstack().sort_values(kind='quicksort')
# Select the highest correlation pairs having correlation greater than upper threshold and lower than lower threshold
corr_var_list = correlation[((correlation>upper_threshold) | (correlation<lower_threshold)) & (correlation!=1)]
print(corr_var_list)
# -
# ### Model building
#
# - Separate the features and target and then split the train data into train and validation set.
# - Now let's come to the actual task, using linear regression, predict the `Value (M)`.
# - Try improving upon the `r2_score` (R-Square) using different parameters that give the best score. You can use higher degree [Polynomial Features of sklearn](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html) to improve the model prediction.
#
#
# +
# Code Starts here
linreg = LinearRegression()
logreg = LogisticRegression()
#y = np.log(y_train)
linreg.fit(X_train,y_train)
y_pred = linreg.predict(X_test)
# display predictions
print('Mean Absolute Error :',(mean_absolute_error(y_test,y_pred)))
print('R-Square :',r2_score(y_test,y_pred))
# Code ends here
print('-'*20)
#Polynomial Feature
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(4)
X_train_2 = poly.fit_transform(X_train)
X_test_2 = poly.transform(X_test)
model = LinearRegression()
model.fit(X_train_2, y_train)
y_pred_2 = model.predict(X_test_2)
r2 = r2_score(y_test,y_pred_2)
print("R-Square :", r2)
mae = mean_absolute_error(y_test, y_pred_2)
print('Mean Absolute Error :', mae)
# -
# ### Prediction on the test data and creating the sample submission file.
#
# - Load the test data and store the `Id` column in a separate variable.
# - Perform the same operations on the test data that you have performed on the train data.
# - Create the submission file as a `csv` file consisting of the `Id` column from the test data and your prediction as the second column.
# +
# Code Starts here
test = pd.read_csv('test.csv')
Id = test['Id']
test = test.drop(["Name","Nationality","Club","Position",'Id'],axis=1)
test_poly = poly.transform(test)
y_pred_1 = model.predict(test_poly)
y_pred_1 = y_pred_1.flatten()
id_1=pd.DataFrame({'Id':id,'Value (M)':y_pred_1})
id_1.to_csv("submission.csv", encoding='utf-8', index=False)
# Code ends here
| FIFA_Project_Student-Template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import torch
import numpy as np
from scipy.stats import pearsonr
from sklearn.metrics import roc_auc_score, average_precision_score
from matplotlib import pyplot as plt
import seaborn as sns
import pandas as pd
from IPython.core.display import display, HTML
from bertviz import head_view, model_view
from scripts.evaluate_explanations import evaluate_word_level, evaluate_sentence_level
from utils import aggregate_pieces, read_qe_files
from model.xlm_roberta import load_checkpoint
from explain.explain_attn import get_valid_explanations
# +
def detokenize(pieces):
return ''.join(pieces).replace('▁', ' ').replace('</s>', ' </s>')
def colorize_twoway(words, color_array, max_width_shown=1800):
template_pos = '<span class="barcode"; style="color: black; background-color: rgba(0, 255, 0, {}); display:inline-block; font-size:12px;">  {}  </span>'
template_neg = '<span class="barcode"; style="color: black; background-color: rgba(255, 0, 0, {}); display:inline-block; font-size:12px;">  {}  </span>'
colored_string = ''
f = lambda w: w.replace('<', 'ᐸ').replace('>', 'ᐳ')
for word, color in zip(words, color_array / color_array.abs().max()):
if color > 0:
colored_string += template_pos.format(color, f(word))
else:
colored_string += template_neg.format(-color, f(word))
html_text = '<div style="100%">{}</div>'.format(colored_string)
display(HTML(html_text))
def plot_attention_map(words, attn_weights, vmax=1, ax=None):
df = pd.DataFrame(attn_weights.numpy(), columns=words, index=words)
sns.heatmap(df, vmax=vmax, cmap="Greens", square=True, cbar=False, ax=ax)
# +
device = torch.device("cpu")
# 1) Load Checkpoint
checkpoint = 'experiments/lightning/version_02-08-2021--23-36-52/epoch=4-step=30624.ckpt'
model = load_checkpoint(checkpoint, output_norm=False, norm_strategy='weighted_norm')
model.eval()
model.zero_grad()
model.to(device)
# 2) Prepare TESTSET
data = read_qe_files('data/ro-en/dev')
e_mt_gold = [sample["tgt_tags"] for sample in data]
e_src_gold = [sample["src_tags"] for sample in data]
# +
idx = 10
layer_id = 18
head_id = 3
reduction = 'sum'
sample = data[idx]
mt_word_tags = torch.tensor(e_mt_gold[idx])
src_word_tags = torch.tensor(e_src_gold[idx])
with torch.no_grad():
batch, targets = model.prepare_sample([sample], cuda=False)
pred_score, attn = model.forward(**batch, return_attentions=True)
pred_score = pred_score.view(-1).detach().cpu().item()
attn = torch.stack(attn).detach().cpu()
input_ids = batch['input_ids']
attn_mask = batch['attention_mask']
fs_mask = batch['first_sentence_mask']
eos_idx = fs_mask.squeeze().sum()
fp_mask = batch['first_piece_mask']
gold_score = targets['score'].squeeze().item()
tokens = model.tokenizer.tokenizer.convert_ids_to_tokens(input_ids.squeeze())
seq_len = attn.shape[-1]
mt_attn = attn[:, :, :, :eos_idx, :eos_idx]
src_attn = attn[:, :, :, eos_idx:, eos_idx:]
mt_tokens = tokens[:eos_idx]
src_tokens = tokens[eos_idx:]
attn_sum = (attn[layer_id, 0, head_id] * attn_mask.unsqueeze(-1).float()).sum(1)
attn_avg = attn_sum / attn_mask.sum(-1).unsqueeze(-1).float()
explanations = get_valid_explanations(attn_avg, attn_mask, fs_mask, fp_mask)
e_mt, e_src, e_mt_fp_mask, e_src_fp_mask = explanations[0]
e_mt = torch.from_numpy(e_mt)
e_src = torch.from_numpy(e_src)
e_mt_fp_mask = torch.from_numpy(e_mt_fp_mask)
e_src_fp_mask = torch.from_numpy(e_src_fp_mask)
ag_e_mt = aggregate_pieces(e_mt, e_mt_fp_mask, reduction=reduction)
ag_e_src = aggregate_pieces(e_src, e_src_fp_mask, reduction=reduction)
# -
print('pred score:', pred_score)
print('gold score:', gold_score)
print('')
print('pred:')
# colorize_twoway(mt_tokens, e_mt)
colorize_twoway(detokenize(mt_tokens).split(), ag_e_mt)
print('gold:')
colorize_twoway(detokenize(mt_tokens).split(), torch.tensor([0] + mt_word_tags.tolist() + [0]))
print('')
print('pred:')
# colorize_twoway(src_tokens, e_src)
colorize_twoway(detokenize(src_tokens).split(), ag_e_src)
print('gold:')
colorize_twoway(detokenize(src_tokens).split(), torch.tensor([0] + src_word_tags.tolist() + [0]))
plt.figure(figsize=(10,10))
plot_attention_map(tokens, attn[18,0,0], vmax=None)
# +
# not very useful afterall
# head_view(src_attn, src_tokens)
| notebooks/investigate_attention.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Population_Data_Visualisation
# +
## Importing important libraries
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rc('xtick', labelsize=12)
matplotlib.rc('ytick', labelsize=30)
matplotlib.rcParams.update({'font.size': 28})
import math
import datetime as dt
import os
import sys
# -
# ## Utility Functions
# +
## Visulalization function
def Visualize(dataset,List_of_count_to_print,title1,ylab,vx=50,vy=30,w=.80):
df = dataset
n = 0
for i in List_of_count_to_print:
filter1 = df['Country'] == i
df = df[filter1]
labels = df['Date']
conf = df['Confirmed']
Recov = df['Recovered']
Death = df['Deaths']
#high = max(conf)
#low = min(conf)
x = np.arange(len(labels)) # the x label locations
width = w # the width of the bars
fig, ax = plt.subplots(figsize=(vx,vy))
rects1 = ax.bar(x - width, conf, width, label='confirmed')
rects2 = ax.bar(x , Recov, width, label='Recovered')
rects3 = ax.bar(x + width , Death, width, label='Death')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel(ylab)
ax.set_title(title1)
ax.set_xticks(x)
plt.xticks(rotation=90)
#plt.ylim([math.ceil(low-0.5*(high-low)), math.ceil(high+0.5*(high-low))])
ax.set_xticklabels(labels)
ax.legend()
n = n + 1
plt.show()
## function to Check the List of Countries avaialable
def count_avalaible(dataframe,country_coul_rep = 'Country'):
x = 0
for i in set(dataframe.loc[:,country_coul_rep]):
print(i,end=' | ')
x = x + 1
if(x > 6):
x = 0
print()
print("\n\n##Total No of Countries = " + str(len(set(dataframe.loc[:,country_coul_rep]))))
# -
# ## Loading Population_Data Data
Population_Data_Countires_Wise_descreet = pd.read_csv('../../Population_Data/WPP2019_TotalPopulationBySex.csv')
Population_Data_Countires_Wise_descreet
# +
## Check the List of Countries avaialable
##
full_pop_data = Population_Data_Countires_Wise_descreet
##
## Columns renaming for Uniformity
Population_Data_Countires_Wise_descreet = Population_Data_Countires_Wise_descreet.rename(columns={'Location': 'Country'})
count_avalaible(Population_Data_Countires_Wise_descreet,'Country')
# -
Population_Data_Countires_Wise = pd.read_csv('../../Population_Data/WPP2019_PopulationBySingleAgeSex_1950-2019.csv')
Population_Data_Countires_Wise
# +
## Check the List of Countries avaialable
##
full_pop_data = Population_Data_Countires_Wise
##
## Columns renaming for Uniformity
Population_Data_Countires_Wise = Population_Data_Countires_Wise.rename(columns={'Location': 'Country'})
count_avalaible(Population_Data_Countires_Wise,'Country')
# +
## Analysing the data Structure
Country_to_look_for = 'India'
Year_to_look_for = 2019
ylab = "Population"
xlab = "Year"
filter1 = Population_Data_Countires_Wise['Country'] == Country_to_look_for
Population_Data_Countires_Wise_country_specific = Population_Data_Countires_Wise[filter1]
filter2 = Population_Data_Countires_Wise_country_specific['Time'] == Year_to_look_for
Population_Data_Countires_Wise_country_specific = Population_Data_Countires_Wise_country_specific[filter2]
Population_Data_Countires_Wise_country_specific
#Population_Data_Countires_Wise ## Uncomment this to view for all countires at once
# +
## Visualisation
df = Population_Data_Countires_Wise_country_specific
labels = df['AgeGrp']
prev_PopMale = df['PopMale']
prev_PopFemale = df['PopFemale']
prev_PopTotal = df['PopTotal']
title1 = 'Population in ' + str(Country_to_look_for) + ' and year = ' + str(Year_to_look_for)
#high = int(max(prev_2018))
#low = 0
x = np.arange(len(labels)) # the x label locations
width = .50 # the width of the bars
fig, ax = plt.subplots(figsize=(40,30))
rects1 = ax.bar(x-width/2, prev_PopMale, width, label='PopMale')
rects2 = ax.bar(x, prev_PopFemale, width, label='PopFemale')
rects3 = ax.bar(x+width/2, prev_PopTotal, width, label='PopTotal')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel(ylab)
ax.set_xlabel(xlab)
ax.set_title(title1)
ax.set_xticks(x)
#ax.set_yticks(y)
plt.xticks(rotation=90)
#plt.ylim([math.ceil(low-0.5*(high-low)), math.ceil(high+0.5*(high-low))])
ax.set_xticklabels(labels)
ax.legend()
plt.show()
# -
# ## Cleaning Population DATA(Preprocessing)
Population_Data_Countires_Wise = Population_Data_Countires_Wise.replace('United States of America', 'US')
Population_Data_Countires_Wise_descreet = Population_Data_Countires_Wise_descreet.replace('United States of America', 'US')
# +
## Column match
print('-----------------------------------------------------------------')
countries = ['Afghanistan','Italy' , 'Kuwait', 'India', 'South Africa' ,'US',
'United Kingdom','Sri Lanka', 'Chile' , 'Norway', 'New Zealand' ,'Switzerland',
'Australia', 'Canada', 'China','Slovenia','North Macedonia']
k = 0
match = []
for i in set(Population_Data_Countires_Wise.loc[:,'Country']):
if(i in countries):
k +=1
match.append(i)
print(i)
print(k)
print("-------Not Matching --------------------")
for i in countries:
if(i not in match ):
print(i)
# +
## Column match
print('-----------------------------------------------------------------')
countries = ['Afghanistan','Italy' , 'Kuwait', 'India', 'South Africa' ,'US','Bangladesh', 'Brazil',
'United Kingdom','Sri Lanka', 'Chile' , 'Norway', 'New Zealand' ,'Switzerland','Ireland','Argentina',
'Australia', 'Canada', 'China','Slovenia','North Macedonia','Zimbabwe','Sweden','Netherlands','Pakistan']
k = 0
match = []
for i in set(Population_Data_Countires_Wise_descreet.loc[:,'Country']):
if(i in countries):
k +=1
match.append(i)
print(i)
print(k)
print("-------Not Matching --------------------")
for i in countries:
if(i not in match ):
print(i)
# -
Population_Data_Countires_Wise_descreet_processed = Population_Data_Countires_Wise_descreet[
Population_Data_Countires_Wise_descreet['Time'] == 2020 ]
Population_Data_Countires_Wise_descreet_processed = Population_Data_Countires_Wise_descreet_processed[ Population_Data_Countires_Wise_descreet_processed['Variant'] == 'Medium']
Population_Data_Countires_Wise_descreet_processed
# ## Writing the cleaned data in Cleaned Folder
Population_Data_Countires_Wise.to_csv('../Pre_Processed_Data/Population_Data_Countires_Wise_Processed.csv')
Population_Data_Countires_Wise_descreet_processed.to_csv('../Pre_Processed_Data/Population_Data_Countires_Wise_Descreet_Processed.csv')
# ## Processing population Data to Extract Age data for countries
Age_data_country_Wise = Population_Data_Countires_Wise
#Age_data_country_Wise
Age_data_country_Wise = Population_Data_Countires_Wise
Age_data_country_Wise = Age_data_country_Wise[
Age_data_country_Wise['Time'] == 2019 ]
Age_data_country_Wise = np.array(Age_data_country_Wise)
Age_data_country_Wise = Age_data_country_Wise[:,[1,4,6,9,10,11]]
#pd.DataFrame(Age_data_country_Wise)
# +
## Processing the Age data to combine population into categories
# 0-16
# 17-45
# 45-60
# 60-100 or 60+
def age_grouping(input_data,countries):
Complete_Data = input_data
Complete_Data[:,[2,3,4,5]] = np.array(Complete_Data[:,[2,3,4,5]]).astype(int)
selected_countries = np.zeros(shape=(1,5))
data = Complete_Data[:,[0,2,5]]
age_val = [0,0,0,0] ## contains values in order age0_16 , age17_45 , age46_60 , age61_above
filter1 = []
for i in countries:
filter1.clear()
age_val = [0,0,0,0]
for j in range(0,len(data[:,0])):
if(data[j,0] == i):
filter1.append(True)
else:
filter1.append(False)
Complete_Data = data[filter1]
for j in range(0,len(Complete_Data[:,0])):
if(Complete_Data[j,0] == i):
if(Complete_Data[j,1] in range(0,16)):
age_val[0] += int(Complete_Data[j,2])
if(Complete_Data[j,1] in range(17,45)):
age_val[1] += int(Complete_Data[j,2])
if(Complete_Data[j,1] in range(45,60)):
age_val[2] += int(Complete_Data[j,2])
if(Complete_Data[j,1] in range(60,100)):
age_val[3] += int(Complete_Data[j,2])
selected_countries = np.append(selected_countries,[str(i),int(age_val[0]),int(age_val[1]),int(age_val[2]),int(age_val[3])])
## Start from here reshape the values
selected_countries = selected_countries.reshape((int(len(selected_countries)/5)),5)
selected_countries = selected_countries[1:,[0,1,2,3,4]]
#print(selected_countries[:,[0,1,2,3,4]])
return selected_countries
# -
#countries = list(Population_Data_Countires_Wise.loc[:,'Country']) # Use when its required to calculate population clasification for all
# the countries
Population_Data_Age_Wise = pd.DataFrame(age_grouping(Age_data_country_Wise,countries))
Population_Data_Age_Wise
## Storing data into a Dataset Population_Data_Age_Wise
Population_Data_Age_Wise.to_csv('../Pre_Processed_Data/Population_Data_Age_Wise.csv')
# ## Visualisation After Cleaning
# +
## Visualisation
df = Population_Data_Countires_Wise_country_specific
labels = df['AgeGrp']
prev_PopMale = df['PopMale']
prev_PopFemale = df['PopFemale']
prev_PopTotal = df['PopTotal']
title1 = 'Population in ' + str(Country_to_look_for) + ' and year = ' + str(Year_to_look_for)
#high = int(max(prev_2018))
#low = 0
x = np.arange(len(labels)) # the x label locations
width = .50 # the width of the bars
fig, ax = plt.subplots(figsize=(40,30))
rects1 = ax.bar(x-width/2, prev_PopMale, width, label='PopMale')
rects2 = ax.bar(x, prev_PopFemale, width, label='PopFemale')
rects3 = ax.bar(x+width/2, prev_PopTotal, width, label='PopTotal')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel(ylab)
ax.set_xlabel(xlab)
ax.set_title(title1)
ax.set_xticks(x)
#ax.set_yticks(y)
plt.xticks(rotation=90)
#plt.ylim([math.ceil(low-0.5*(high-low)), math.ceil(high+0.5*(high-low))])
ax.set_xticklabels(labels)
ax.legend()
plt.show()
# +
## Visualisation
## Analysing the data Structure
Country_to_look_for = 'India'
Year_to_look_for = 2020
ylab = "Population"
xlab = "Year"
filter1 = Population_Data_Countires_Wise_descreet['Country'] == Country_to_look_for
Population_Data_Countires_Wise_country_specific = Population_Data_Countires_Wise_descreet[filter1]
Population_Data_Countires_Wise_country_specific
#Population_Data_Countires_Wise ## Uncomment this to view for all countires at once
## Visualisation
df = Population_Data_Countires_Wise_country_specific
labels = df['Country']
prev_PopMale = df['PopMale']
prev_PopFemale = df['PopFemale']
prev_PopTotal = df['PopTotal']
title1 = 'Population in ' + str(Country_to_look_for) + ' and year = ' + str(Year_to_look_for)
#high = int(max(prev_2018))
#low = 0
x = np.arange(len(labels)) # the x label locations
width = .50 # the width of the bars
fig, ax = plt.subplots(figsize=(80,30))
rects1 = ax.bar(x-width/2, prev_PopMale, width, label='PopMale')
rects2 = ax.bar(x, prev_PopFemale, width, label='PopFemale')
rects3 = ax.bar(x+width/2, prev_PopTotal, width, label='PopTotal')
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_ylabel(ylab)
ax.set_xlabel(xlab)
ax.set_title(title1)
ax.set_xticks(x)
#ax.set_yticks(y)
plt.xticks(rotation=90)
#plt.ylim([math.ceil(low-0.5*(high-low)), math.ceil(high+0.5*(high-low))])
ax.set_xticklabels(labels)
ax.legend()
plt.show()
# -
| Experiment Scripts/Data_Visualisation_Code/.ipynb_checkpoints/Population_Data_Visualisation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <span style = "color:black" > Meteorite Landings on Earth! ~ by <NAME> </span>
# ### <span style = "color:red" > Introduction to the dataset being explored! </span>
#
# <span style = "color:blue"> The Meteoritical Society collects information about meteorites that have come down to Earth from space. This dataset is the sample data, which comprises 1000 rows from a total of 45,000 rows in the full dataset. It includes data on over 1000 meteorites that have struck our globe, such as their name, id, nametype, recclass, mass, fall, position, year, reclat, reclong, and geolocation. </span>
#
# <span style = "color:blue"> The dataset contains the following columns: </span>
#
# . name: Meteorite name
#
# . id: Meteorite's unique identifier
#
# . nametype: 1. valid: a typical meteorite / 2. relict: a meteorite that has been highly degraded by weather on Earth
#
# . recclass: Meteorite Classficcation
#
# . mass: Meteorite's mass in grams
#
# . fall: 1. Fell: the meteorite's fall was observed / 2. Found: the meteorite's fall was not observed
#
# . year: Year of Falling or observing of Meteorite
#
# . reclat: Meteorite's landing lattitude
#
# . reclong: Meteorite's landing longitude
#
# . GeoLocation: reclat and reclong as comma separated tuple enclosed in parentheses
#
#
# <span style="color:blue"> The name of the dataset is Earth_Meteroite_Landings and is available on NASA's website under Space-Science -> Meteorite-Landings section. </span>
#
# <span style="color:blue"> There are 2 versions of the dataset that is available on NASA's website, one is with 45,716 rows which is the full dataset and can be found here https://data.nasa.gov/Space-Science/Meteorite-Landings/gh4g-9sfh/data , for our analysis I've selected a sample dataset having 1000 entires which can be found here https://data.nasa.gov/resource/gh4g-9sfh.json </span>
#
# <span style="color:blue"> This dataset is publicly available on the NASA website and we are allowed to explore it.
# Privacy policy around using this dataset can be found here https://www.nasa.gov/about/highlights/HP_Privacy.html </span>
#
# <span style="color:blue"> The size of the file (sample data used for analysis) is 242 KB and has 1000 rows of entries.<span>
#
#Let us import the libraries for our analysis
import json
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import ipywidgets
#Reading the dataset in for the analysis
meteor_landing = pd.read_json("Earth_Meteorite_Landings.json")
meteor_landing
#dropping extra fields which are of no use
meteor_landing = meteor_landing.drop([":@computed_region_cbhk_fwbd", ":@computed_region_nnqa_25f4"], axis=1)
#Checking out the first 5 rows of our dataset using the head() function
meteor_landing.head()
#Let's use info() to know data type and non-null values of columns in our dataset
meteor_landing.info()
# +
#drop rows where mass is 'nan'
meteor_landing_new = meteor_landing.drop(meteor_landing[meteor_landing.mass == 'nan'].index)
#drop rows where year is <NA>
meteor_landing_new = meteor_landing_new.drop(meteor_landing_new[meteor_landing_new.year == '<NA>'].index)
#Getting values in the form of YYYY-MM-DD from year field using Panda's to_datetime function
meteor_landing_new["year"] = pd.to_datetime(meteor_landing_new['year'], errors = 'coerce')
meteor_landing_new
# -
#Checking the datatype of the year column
meteor_landing_new["year"].dtype
#Checking for the minimum year value present in the dataset
meteor_landing_new['year'].min()
#Checking for the maximum year value present in the dataset
meteor_landing_new['year'].max()
# <span style="color:blue"> This tells us that our dataset has data between year 1688 and 2013. <span>
#Count of total number of unique years in the dataset
meteor_landing_new["year"].nunique()
#Count of total number of unique fall category in the dataset
meteor_landing_new["fall"].nunique()
#Displaying unique fall category in the dataset
meteor_landing_new["fall"].unique()
#List the total number of unique recclass present
meteor_landing_new["recclass"].nunique()
#List all the unqiue recclass present in the dataset
meteor_landing_new["recclass"].unique()
#Lets drop all the rows where mass and year is zero
meteor_landing_new.drop(meteor_landing_new[meteor_landing_new['mass']==0].index, inplace=True)
meteor_landing_new.drop(meteor_landing_new[meteor_landing_new['year']==0].index, inplace=True)
#This will clean the remaining Null, NA, 0 values for all the fields in the dataset
meteor_landing_new = meteor_landing_new.dropna()
#Check the datatype of all the fields in the dataset
meteor_landing_new.info()
meteor_landing_new
#Lets take a look at the min and max mass of meteorites fallen in our dataset
meteor_landing_new["mass"].min(), meteor_landing_new["mass"].max()
# <span style="color:blue">Now that we have cleaned our dataset and we only have non-null values, we can now start the analysis of our dataset. In our dataset we have Longitude as reclong and Latitude as reclat, we also have mass of meteorites given in the dataset. Let us plot a scatter plot of meteorites with x-axis having Longitudes and y-axis with Latitudes and choose the density of the colors based on mass of meteorites within rainbow colormap.</span>
# +
#Plotting Longitude and Latitude on x and y axis and color coding the scatters based on the mass value of meteorites
#Colormap chosen for this visualization is rainbow
fig, ax = plt.subplots(figsize=(10,5))
scatter = plt.scatter(meteor_landing_new['reclong'], meteor_landing_new['reclat'],
c=np.log10(meteor_landing_new['mass']), cmap='rainbow') # we know there are problems with rainbow!
plt.colorbar() #Adding legend to the scatterplot with values in log10 base for viewers to interpret colors
plt.show()
# -
# <span style='color:blue'>Let's also plot a histogram based on the Longitudes and count of meteorites falling in those longitudal areas. </span>
#Plotting histograms based on the rows available for each longitudes
fig, ax = plt.subplots(figsize=(10,5))
meteor_landing_new.hist(column='reclong', ax=ax)
plt.show()
# ### Contextual Visualization 1
# <span style="color:blue">In our dataset we have meteorites which fall in different classifications of meteorites denoted by the column 'reclass'. Let us try to visualize using a bar plot the frequency of top 10 most common meteorites in our dataset. </span>
#lets now check the top 10 years where most number of meteorites data is available
top_10_meteor_class = meteor_landing_new.recclass.value_counts().rename_axis('recclass').reset_index(name='counts')
top_10_meteor_class = top_10_meteor_class.nlargest(10, 'counts', keep='first')
top_10_meteor_class = top_10_meteor_class.set_index('recclass')
top_10_meteor_class['counts'].plot(kind='bar', grid=True, figsize=(8,5),
ylim=(5, 250), title='Count of top 10 Meteorites types', color = 'red')
plt.show()
# <span style="color:blue">From the above bar plot we can see that the most common class of meteorites in our dataset in 'L6' followed by 'H5' and 'H6' and so on. Let us narrow down our dataset even further to only top 10 most frequent occuring meteorites to do our analysis. Starting with creating a list of these top 10 meteor class and using this list creating a subset of the primary dataset.</span>
#
# <span style="color:blue">Let us look at the images of top 3 meteorites L6, H5 and H6 individually.
# </span>
# <span style="color:blue"> Firstly, lets look at how L6 class of meteorite, which is the most frequent occuring class looks? The image below is taken taken from https://www.mindat.org/photo-1098795.html website and the copyright is in the name of '<NAME>' </span>
# 
# <span style="color:blue"> Now, lets look at how the second most frequent class of meteorite 'H5' appears to be in real? The image below is taken taken from https://www.mindat.org/photo-597283.html website and the copyright is in the name of '<NAME>' </span>
# 
# <span style="color:blue"> Now, lets look at how the third most frequent class of meteorite 'H6' photograph from the vault? The image below is taken taken from https://www.mindat.org/photo-1206566.html website and the copyright is in the name of '<NAME>' </span>
# 
# +
#Creating a list of top 10 most frequent occuring meteorite class from our primary dataset
options = ['L6', 'H5', 'H6', 'L5', 'H4','LL6','Eucrite-mmict','LL5','L4','CM2']
#Building subset of primary dataset with top 10 most frequent occuring meteor class.
meteorite_new_top10 = meteor_landing_new[meteor_landing_new['recclass'].isin(options)]
meteorite_new_top10
# -
#Meteorite minimum mass in subset dataset
meteorite_new_top10['mass'].min()
#Meteorite maximum mass in subset dataset
meteorite_new_top10['mass'].max()
# <span style="color:blue">Based on our new subset of data, let us build an interactive visualization based on frequency. The end-user will have the option to select either the categorical column i.e. 'reclass' and see the frequency of meteorites, or select the numerical columns i.e. 'reclong' or 'reclat'. From drop-down end-user can select either reclass, reclong or reclat for visualizing the data. If end-user selects reclong or reclat, there is another option of selecting the number of bins for plotting the histograms for the numerical columns. </span>
#Checking the data types of the field in the dataset to use under if-else condition when plotting visualization
meteorite_new_top10.dtypes
#Building a slider widget to increase the binning based on user input
bin_widget = ipywidgets.IntSlider(min=1, max=100, step=1)
bin_widget
meteorite_new_top10.head()
#Creating list of columns present in the csv file to filter the categorical and numerical columns
mylist=list(meteorite_new_top10.columns)
mylist
#Creating list of columns which user can pick to draw the histogram
hist_list=[mylist[3],mylist[7],mylist[8]]
# +
#Counting rows under each meteor class.
meteorite_new_top10_temp = meteorite_new_top10.groupby("recclass").agg({"id":"count"}).reset_index().rename(columns={"id":"total_count"})
meteorite_new_top10_temp
# -
@ipywidgets.interact(hist_list = hist_list, bin_widget = bin_widget)
def meteor_hist(hist_list, bin_widget):
if meteorite_new_top10[hist_list].dtype == 'float64': #if field is numerical plot histogram
fig, ax = plt.subplots(figsize=(10,5))
meteorite_new_top10.hist(column=hist_list, ax=ax, bins=bin_widget)
ax.set_ylabel("Frequency")
plt.title("%s Histogram"%hist_list.capitalize(), fontdict={'fontsize' : 30}, color = 'red')
else:
#creating meteorite_new_top10_temp to get the count for number of rows for each category based on 'id' field
meteorite_temp = meteorite_new_top10.groupby(hist_list).agg({"id":"count"}).reset_index().rename(columns={"id":"total_count"})
fig, ax = plt.subplots(figsize=(15,7))
meteorite_temp.plot.bar(x=hist_list,y="total_count", ax=ax) #plot bar graph for categorical variables
ax.set_ylabel("Frequency")
plt.title("%s and its %s bar plot"%(hist_list.capitalize(), "total_count".capitalize()),fontdict={'fontsize' : 20}, color = 'red')
# <span style="color:blue"> In the above visualization, the drop-down interactive option 'hist_list' provides user to select either of the 3 options which include one categorical variable 'recclass' and other 2 options are numerical variable i.e. reclong and reclat. There is an added interactivity which is 'bin_widget' which can be adjusted for the 2 numerical variables. Since, recclass falls under categorical variable type, bin_widget won't have any effect on it's bar plot if chosen as a variable.
#
# 1. If the user selects 'reclass' as the variable then the bar_plot will show the top 10 frequently occuring meteorites along with their count on the y-axis.
#
# 2. If the user selects 'reclat' as the variable then the histogram will show the frequency of the meteorites distributed over the latitudes. The user also has the freedom to increase/decrease the binning feature of the histogram in this case.
#
# 3. If the user selects 'reclong' as the variable then the histogram will show the frequency of the meteorites distributed over the longitudes. The user also has the freedom to increase/decrease the binning feature of the histogram in this case as well.
#
# </span>
#
# ### Contextual Visualization 2
# <span style="color:blue">I found a very interesting visualization built using SAS Visual Analytics on similar dataset with more than 30,000 meteorites entries. This visualization is built by '<NAME>' who is Distinguished Software Developer in the SAS Business Intelligence Research and Development division. The dataset used for the visualization can be found here https://data.nasa.gov/view/ak9y-cwf9. </span>
#to import image data we need Image Reader from PIL
from PIL import Image
#importing the visualization image in the notebook
meteor_viz = Image.open("Meteorite_SAS.png")
#Let plot our image now in the notebook
fig = plt.figure(figsize=(15,20))
plt.imshow(meteor_viz)
plt.show()
# <span style="color:blue">
# At first glance, the visualization appears to be extremely remarkable, and it also gives me an idea of how effective SAS Visual Analytics can be. <NAME> has published a blog post on how he went about developing and designing this interactive dashboard, which you can read here: https://blogs.sas.com/content/sascom/2017/03/28/design-meteorite-infographic-using-nasa-data-sas/ . Let's take a closer look at the intricacies of the visualization construct for the sake of analysis.
# <br/>
#
# Falko has utilized a dark-theme to show the dataset, which seems suitable given that we are analyzing meteorites that are extraterrestrial impacts on Earth. Between 2,500 BCE and 2012, the visualization offers the user a good indication of where meteorites have fallen. I really admire the work put in in creating two bar charts on opposite sides of the visualization. The one on the left is a bar chart that displays the most often occurring meteorites, while the one on the right is a bar chart that shows the distribution of meteorites based on their mass. The underlying geo-map display is filtered by both of these charts. The 'fall' column of the dataset, which has two values - fell (orange) and found (blue), determines the color of the scatters on the geo-map. The term "fell" refers to when a meteorite is witnessed descending, whereas "found" refers to when a meteorite has already fallen and was not observed directly.
# On the very bottom the author has included a line chart which shows us the frequency of meteorites over the years.
#
# By looking at this visualization, the end-user will get a good notion of the context of the visualization. The visualization's header includes provides basic information on the dataset as well as a link to the dataset.
# </span>
# <span style = "color:black">
# About the Author
# <br/>
#
# <NAME> is a graduate student studying in Information Management at the University of Illinois at Urbana-Champaign. He has about 5 years of professional work experience as a Data Architect at Adobe Inc. He has always been inspired by the vast amount of data available to us and how we may represent it using a variety of data visualization tools and programming languages. During his master's degree, he aspires to go deeper into the realm of data visualization and have a better understanding of principles for creating end-user friendly visualizations and dashboards.
# </span>
# ### Dataset and References
#
# https://data.nasa.gov/resource/gh4g-9sfh.json
#
# https://www.w3schools.com/python/matplotlib_pie_charts.asp
#
# https://www.nasa.gov/about/highlights/HP_Privacy.html
#
# https://realpython.com/visualizing-python-plt-scatter/
#
# https://realpython.com/python-histograms/
#
# <NAME>. (2017, March 28). How to design a meteorite infographic using NASA data and SAS. Retrieved from https://blogs.sas.com/content/sascom/2017/03/28/design-meteorite-infographic-using-nasa-data-sas/
#
# https://www.mindat.org/photo-1098795.html copyright is in the name of '<NAME>'.
#
# https://www.mindat.org/photo-597283.html copyright is in the name of '<NAME>'.
#
# https://www.mindat.org/photo-1206566.html copyright is in the name of '<NAME>'.
#
| keshri_pratham-final_project_part3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### 使用Tensorflow作用于推荐系统
# ### 1 数据准备 - 以Movielens 为例
"Movieslens 数据格式: user,item,rating,timestamp"
# ### 2 数据预处理部分
# +
# from __future__ import absolute_import,division,print_function => print ("this python3.x ") 需要按照python3 语法规则
#coding:utf-8
import numpy as np
import pandas as pd
"function 1:"
def read_data_and_process(filename, sep = "\t"):
col_names = ["user","item","rate","timestamp"]
df = pd.read_csv(filepath_or_buffer = filename, sep = sep, header = None, names = col_names, engine = 'python')
"逐级递减"
df['user'] -= 1
df['item'] -= 1
for col in ('user','item'):
df[col] = df[col].astype(np.int32)
df['rate'] = df['rate'].astype(np.float32) #"转换数据类型by astype"
return df
"随机生成一个batch一个batch的数据来保证每次训练模型拿到的数据均不同,避免过拟合的出现"
class ShuffleBatchData(object): # object
def __init__ (self,inputs,batch_size = 10):
self.inputs = inputs
self.batch_size = batch_size
self.num_cols = len(self.inputs)
self.len = len(self.inputs[0])
self.inputs = np.transpose( np.vstack([np.array(self.inputs[i] for i in range(self.num_cols))] ) )
def __len__ (self):
return self.len
def __iter__(self):
return self
def __next__(self):
return self.next()
"随机生成一个batch size个下标, 并取出相应的样本"
def next(self):
ids = np.random.randint(0,self.len,(self.batch_size, ) )
out = self.inputs[ids,:]
return [out[:,columns_index] for columns_index in range(self.num_cols)]
"顺序产生一个epoch的数据,用于测试中...."
class OneEpochTestDataProcesing(ShuffleBatchData):
def __init__(self,inputs,batch_size = 10):
super(ShuffleBatchData,self).__init__(inputs,batch_size = batch_size)
if batch_size>0:
self.idx_group = np.array_split(np.arange(self.len), np.ceil(self.len/batch_size) )
else:
self.idx_group = [np.arange(self.len)]
self.group_id = 0
def next(self):
if self.group_id > len(self.idx_group):
self.group_id = 0
raise StopIteration
out = self.inputs[self.idx_group[self.group_id],:]
self.group_id += 1
return [out[:,i] for i in range(self.num_cols)]
# -
# ### 3 搭建model
# +
import tensorflow as tf
"使用矩阵分解搭建网络结构"
def interface_svd(user_batch,item_batch,user_num,item_num,dim = 5, device = "/cpu:0"):
with tf.device("/cpu:0"):
"初始化几个偏执项"
global_bias = tf.get_variable("global_bias",shape = [])
w_bias_user = tf.get_variable('embd_bias_user',shape = [user_num])
w_bias_item = tf.get_variable('embd_bias_item',shape = [item_num])
"bias向量"
bias_user = tf.nn.embedding_lookup(w_bias_user,user_batch,name = "bias_user")
bias_item = tf.nn.embedding_lookup(w_bias_item,item_batch,name = "item_user")
w_user = tf.get_variable("embd_user",shape = [user_num,dim], initializer = tf.truncated.normal_initializer(stddev = 0.02) )
w_item = tf.get_variable("embd_item",shape = [item_num,dim], initializer = tf.truncated.normal_initializer(stddev = 0.02))
"user向量与item向量"
embd_user = tf.nn.embedding_lookup(w_user,user_batch, name = "embedding_user")
embd_item = tf.nn.embedding_lookup(w_item,item_batch,name = "embedding_item")
"以上的都是tensorflow 规定的向量初始化过程"
with td.device(device):
"按照公式对user 向量和item 向量求和"
infer = tf.reduce_sum(tf.multiply(embd_user,embd_item),1)
"按照公式加上几个偏置项"
infer = tf.add(infer,global_bias)
infer = tf.add(infer,bias_user)
infer = tf.add(infer,bias_item,name = 'svc_inference')
"加上正则化项"
regularizer = tf.add(tf.nn.l2_loss(embd_user),tf.nn.l2_loss(embd_item),name = 'svd_regularizer')
return infer, regularizer
"模型迭代"
def optimization(infer,regularizer,rate_batch,learning_rate = 0.001,reg = 0.10,device = "./cpu:0"):
global_step = tf.train.get_global_step()
assert global_step is not None
"选择合适的optimizer 优化"
with tf.device(device):
cost_l2 = tf.nn.l2_loss(tf.subtract(infer,rate_batch))
penalty = tf.constant(reg,dtype = tf.float32, shape = [], name = 'l2')
cost = tf.add(cost_l2, tf.multiply(regularizer,penalty))
train_optimizer = tf.train.AdamOptimizer(learning_rate).minimize(cost,global_step = global_step)
return cost,train_optimizer
# -
# ### 4.在实际数据上训练模型
#
# +
import time
from collections import deque
import numpy as np
import tensorflow as tf
from six import next
from tensorflow.core.framework import summary_pb2
np.random.seed(123)
"模型常量定义"
BATCH_SIZE = 2000
USER_NUM = 6040
ITEM_NUM = 3952
DIM = 15 # "factor 维度"
"最大迭代轮数"
EPOCH_MAX = 200
"使用cpu做训练"
DEVICE = "./cpu:0"
"截断"
def clip(x):
return np.clip(x,1,5)
"方便可视化做的summary"
def make_scalar_summary(name,value):
return summary_pb2.Summary(value = [summary_pb2.Summary.Value(tag = name,simple_value = value )])
"通过调用上面定义的函数获取数据:"
def get_data():
df = read_data_and_process("./movielens/ml-1m/ratings.dat",sep = "::")
rows = len(df)
"permutation - 生成随机队列"
df = df.iloc[np.random.permutation(rows)].reset_index(drop = True)
split_index = int(rows * 0.9)
df_train = df[0:split_index]
'reset_index : We can use the `drop = True` parameter to avoid the old index being added as a column'
df_test = df[split_index:].reset_index(drop = True )
print(df_train.shape,df_test.shape)
return df_train,df_test
"实现训练过程"
def svd(train,test):
samples_per_batch = len(train) # "batch size"
"将数据一个batch一个batch喂到模型里面训练"
iter_train = ShuffleBatchData([train["user"],train["item"], train["rate"] ],batch_size = BATCH_SIZE)
"测试数据"
iter_test = OneEpochTestDataProcesing([test["user"], test["item"], test["rate"]], batch_size = -1)
"user and item batch "
user_batch = tf.placeholder(tf.int32,shape = [None], name = "id_user")
item_batch = tf.placeholder(tf.int32,shape = [None],name = "id_item")
rate_batch = tf.placeholder(tf.int32,shape = [None])
"构建Graph 和训练"
infer, regularizer = interface_svd(user_batch,item_batch,USER_NUM,ITEM_NUM,DIM,DEVICE)
global_step = tf.contrib.get_or_create_global_step()
cost,train_optimizer = optimization(infer,regularizer,rate_batch,learning_rate = 0.001,reg = 0.10,device = DEVICE)
"初始化所有变量"
init_op = tf.global_variables_initializer()
"开始迭代"
with tf.Session() as sess:
sess.run(init_op)
summary_writer = tf.summary.FileWriter(logdir = '/InternalData/log', graph = sess.graph )
print "{} {} {} {}".format("epoch","train_error","val_error","elapsed_time")
errors = deque(maxlen = samples_per_batch)
start = time.time()
for i in range(EPOCH_MAX * samples_per_batch):
users,items,rates = next(iter_train)
_,pred_batch = sess.run([train_optimizer,infer], feed_dict = {user_batch:users,
item_batch:items,
rate_batch:rates})
pred_batch = clip(pred_batch)
errors.append(np.power(pred_batch - rates,2) )
if i%samples_per_batch == 0:
train_errors = np.sqrt(np.mean(errors))
test_errors = np.array([])
for users,items,rates in iter_test:
pred_batch = sess.run(infer, feed_dict = {user_batch:users,
item_batch:items})
pred_batch = clip(pred_batch)
test_errors = np.append(test_errors,np.power(pred_batch - rates,2 ))
end = time.time()
test_errors_sqrt = np.sqrt(np.mean(test_errors))
print "{:3d} {:f} {:f} {:f}".format(i//samples_per_batch,train_errors,test_errors_sqrt,end - start)
train_err_summary = make_scalar_summary("training_error",train_errors)
test_err_summary = make_scalar_summary("test_error",test_errors_sqrt)
summary_writer.add_summary(train_err_summary,i)
summary_writer.add_summary(test_err_summary,i)
start = end
# -
"获取数据集"
train_datasets, test_datasets = get_data()
"SVD训练数据"
svd(train_datasets,test_datasets)
help(pd.read_csv)
# +
filename = "./RawData/ReleasaeFilesCRC12.5.csv"
col_names = ["Baseline","Date"]
DF = pd.read_csv(filename, sep = ",", header = None, names = col_names, engine = 'python')
DF['Baseline'] = 1
DF['Date'] = "2019"
for col in ("Baseline","Date"):
DF[col] = DF[col].astype(np.float32)
# -
help(pd.DataFrame.reset_index)
DataFrame
| RecommendSystem/.ipynb_checkpoints/RecommendSystem-in-using-TensorFlow-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + dc={"key": "3"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 1. TV, halftime shows, and the Big Game
# <p>Whether or not you like American football, the Super Bowl is a spectacle. There is always a little something for everyone. For the die-hard fans, there is the game itself with blowouts, comebacks, and controversy. For the not so die-hard fans, there are the ridiculously expensive ads that are hilarious, gut-wrenching, thought-provoking, and sometimes weird. And of course, there are the halftime shows with the biggest musicians in the world entertaining us by <a href="https://youtu.be/ZD1QrIe--_Y?t=14">riding a giant mechanical tiger</a> or <a href="https://youtu.be/mjrdywp5nyE?t=62">leaping from the roof of the stadium</a>. It is a grand show! In this notebook, we're going to explore how some of these elements interact with each other. After exploring and cleaning the data, we're going to answer questions like:</p>
# <ul>
# <li>What are the most extreme game outcomes?</li>
# <li>How does the score difference affect television viewership?</li>
# <li>How have viewership, TV ratings, and advertisement costs evolved?</li>
# <li>Who are the most prolific musicians in terms of halftime show performances?</li>
# </ul>
# <p><img src="https://assets.datacamp.com/production/project_691/img/left_shark.jpg" alt="Left Shark Steals The Show">
# <em><a href="https://www.flickr.com/photos/huntleypaton/16464994135/in/photostream/">Left Shark Steals The Show</a>. <NAME>erry performing at halftime of Super Bowl XLIX. Photo by <NAME>. Attribution-ShareAlike 2.0 Generic (CC BY-SA 2.0).</em></p>
# <p>The dataset we'll use was <a href="https://en.wikipedia.org/wiki/Web_scraping">scraped</a> and polished from Wikipedia. It is made up of three CSV files, one with <a href="https://en.wikipedia.org/wiki/List_of_Super_Bowl_champions">game data</a>, one with <a href="https://en.wikipedia.org/wiki/Super_Bowl_television_ratings">TV data</a>, and one with <a href="https://en.wikipedia.org/wiki/List_of_Super_Bowl_halftime_shows">halftime musician data</a> for all 52 Super Bowls through 2018.</p>
# + dc={"key": "3"} tags=["sample_code"]
# Load packages
library(tidyverse)
# Load the CSV data
super_bowls <- read_csv("datasets/super_bowls.csv")
tv <- read_csv("datasets/tv.csv")
halftime_musicians <- read_csv("datasets/halftime_musicians.csv")
# Display the first six rows of each tibble
head(super_bowls)
head(tv)
head(halftime_musicians)
# + dc={"key": "10"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 2. Taking note of dataset issues
# <p>From the quick look at the Super Bowl game data, we can see that the dataset appears whole except for missing values in the backup quarterback columns (<code>qb_winner_2</code> and <code>qb_loser_2</code>), which make sense given most starting QBs in the Super Bowl (<code>qb_winner_1</code> and <code>qb_loser_1</code>) play the entire game.</p>
# <p>From the visual inspection of TV and halftime musicians data, there is only one missing value displayed, but I've got a hunch there are more. The first Super Bowl was played on January 15, 1967, and I'm guessing some data (e.g., the number of songs performed) probably weren't tracked reliably over time. Wikipedia is great but not perfect.</p>
# <p>Looking at a summary of the datasets shows us that there are multiple columns with null values.</p>
# + dc={"key": "10"} tags=["sample_code"]
# Summary of the TV data
summary(tv)
# Summary of the halftime musician data
summary(halftime_musicians)
# + dc={"key": "17"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 3. Combined points distribution
# <p>In the TV data, the following columns have a lot of missing values:</p>
# <ul>
# <li><code>total_us_viewers</code> (amount of U.S. viewers who watched at least some part of the broadcast)</li>
# <li><code>rating_18_49</code> (average % of U.S. adults 18-49 who live in a household with a TV that were watching for the entire broadcast)</li>
# <li><code>share_18_49</code> (average % of U.S. adults 18-49 who live in a household with a TV <em>in use</em> that were watching for the entire broadcast)</li>
# </ul>
# <p>In halftime musician data, there are missing numbers of songs performed (<code>num_songs</code>) for about a third of the musicians.</p>
# <p>There are a lot of potential reasons for missing values. Were the data ever tracked? Would the research effort to fill in the gaps be worth it? Maybe. Watching every Super Bowl halftime show to get song counts could be pretty fun. But we don't have time to do that now! Let's take note of where the datasets are not perfect and start uncovering some insights.</p>
# <p>We'll start by visualizing the distribution of combined points for each Super Bowl. Let's also find the Super Bowls with the highest and lowest scores.</p>
# + dc={"key": "17"} tags=["sample_code"]
# Reduce the size of the plots
options(repr.plot.width = 5, repr.plot.height = 4)
# Plot a histogram of combined points
ggplot(super_bowls, aes(combined_pts)) +
geom_histogram(binwidth = 5) +
labs(x = "Combined Points", y = "Number of Super Bowls")
# Display the highest- and lowest-scoring Super Bowls
super_bowls %>%
filter(combined_pts> 70 | combined_pts < 25)
# + dc={"key": "24"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 4. Point difference distribution
# <p>Most of the combined scores are between 40 and 50 points, with the extremes being roughly equal distance away in opposite directions. At the highest combined scores of 74 and 75, are two games featuring dominant quarterback performances. One happened last year - Super Bowl LII when <NAME> Patriots lost to <NAME>' underdog Eagles 33 to 41, for a combined score of 74.</p>
# <p>On the other end of the spectrum, we have Super Bowl III and VII, which featured tough defenses that dominated the games. We also have Super Bowl IX in New Orleans in 1975, whose 16-6 score can be attributed to inclement weather. Overnight rain made the field slick, and it was cold (46 °F / 8 °C), making it hard for the Steelers and Vikings to do much offensively. This was the second-coldest Super Bowl ever and the last to be played in inclement weather for over 30 years. The NFL realized people like points, I guess.</p>
# <p><em>UPDATE: In Super Bowl LIII in 2019, the Patriots and Rams broke the record for the lowest-scoring Super Bowl with a combined score of 16 points (13-3 for the Patriots).</em></p>
# <p>Now let's take a look at the point difference between teams in each Super Bowl.</p>
# + dc={"key": "24"} tags=["sample_code"]
# Reduce the size of the plots
options(repr.plot.width = 5, repr.plot.height = 4)
# Plot a histogram of point differences
ggplot(super_bowls, aes(difference_pts)) +
geom_histogram(binwidth = 2) +
labs(x = "Point Difference", y = "Number of Super Bowls")
# Display the closest game and largest blow out
super_bowls %>%
filter(difference_pts == min(difference_pts) | difference_pts == max(difference_pts))
# + dc={"key": "31"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 5. Do blowouts translate to lost viewers?
# <p>The vast majority of Super Bowls are close games. Makes sense. Both teams are the best in their conference if they've made it this far. The closest game ever was the Buffalo Bills' 1-point loss to the New York Giants in 1991, which is best remembered for <NAME>'s last-second missed field goal attempt that went <em><a href="https://www.youtube.com/watch?v=RPFZCGgjDSg">wide right</a></em>, kicking off four Bills Super Bowl losses in a row. Poor Scott. The biggest point spread so far is 45 points (!) when Hall of Famer, <NAME>, led the San Francisco 49ers to victory in 1990, one year before the closest game ever.</p>
# <p>I remember watching the Seahawks crush the Broncos by 35 points (43-8) in 2014, which sucked to watch in my opinion. The game was never really close. I'm pretty sure we changed the channel at the end of the third quarter. Let's combine the game data and TV data to see if this is a universal phenomenon. Do large point differences translate to lost viewers? We can plot <a href="https://en.wikipedia.org/wiki/Nielsen_ratings">household share</a> <em>(average percentage of U.S. households with a TV in use that were watching for the entire broadcast)</em> vs. point difference to find out.</p>
# + dc={"key": "31"} tags=["sample_code"]
# Filter out Super Bowl I and join the game data and TV data
games_tv <- tv %>%
filter(super_bowl != 1) %>%
inner_join(super_bowls, by = "super_bowl")
# Create a scatter plot with a linear regression model
ggplot(games_tv, aes( x = difference_pts, y = share_household )) +
geom_point() + # scatter plot
geom_smooth(method = "lm") + # adds regression line
labs(x = "Point Difference", y = "Viewership (household share)")
# + dc={"key": "38"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 6. Viewership and the ad industry over time
# <p>The downward sloping regression line and the 95% confidence interval for that regression <em>suggest</em> that bailing on the game if it is a blowout is common. Though it matches our intuition, we must take it with a grain of salt because the linear relationship in the data is weak due to our small sample size of 52 games.</p>
# <p>Regardless of the score, I bet most people stick it out for the halftime show, which is good news for the TV networks and advertisers. A 30-second spot costs a pretty <a href="https://www.businessinsider.com/super-bowl-commercials-cost-more-than-eagles-quarterback-earns-2018-1">\$5 million</a> now, but has it always been that much? And how has the number of viewers and household ratings trended alongside advertisement cost? We can find out using line plots that share a "Super Bowl" x-axis.</p>
# + dc={"key": "38"} tags=["sample_code"]
# Convert the data format for plotting
games_tv_plot <- games_tv %>%
gather(key = "category", value = "value", "avg_us_viewers", "rating_household", "ad_cost") %>%
mutate(cat_name = case_when(category == "avg_us_viewers" ~ "Average number of US viewers",
category == "rating_household" ~ "Household rating",
category == "ad_cost" ~ "Advertisement cost (USD)",
TRUE ~ as.character(category)))
# Plot the data
ggplot(games_tv_plot, aes( x = super_bowl, y = value)) +
geom_line() +
facet_wrap(~ cat_name, scales = "free", nrow = 3) +
labs(x = "Super Bowl", y = "") +
theme_minimal()
# + dc={"key": "45"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 7. Halftime shows weren't always this great
# <p>We can see that the number of viewers increased before advertisement costs did. Maybe the networks weren't very data savvy and were slow to react? Makes sense since DataCamp didn't exist back then.</p>
# <p>Another hypothesis: maybe halftime shows weren't as entertaining in the earlier years? The modern spectacle that is the Super Bowl has a lot to do with of big halftime acts. I went down a YouTube rabbit hole, and it turns out that older halftime shows were not quite the spectacle they are today. Some examples:</p>
# <ul>
# <li><a href="https://youtu.be/6wMXHxWO4ns?t=263">Super Bowl XXVI</a> in 1992: A Frosty The Snowman rap performed by children.</li>
# <li><a href="https://www.youtube.com/watch?v=PKQTL1PYSag">Super Bowl XXIII</a> in 1989: An Elvis impersonator who did magic tricks and didn't even sing one Elvis song.</li>
# <li><a href="https://youtu.be/oSXMNbK2e98?t=436">Super Bowl XXI</a> in 1987: Tap dancing ponies. Okay, that was pretty awesome actually.</li>
# </ul>
# <p>It turns out that <NAME>'s Super Bowl XXVII performance, one of the most watched events in American TV history, was when the NFL realized that the having big-name halftime acts brought in more viewers. Let's look at the halftime acts before <NAME> brought the NFL and entertainment industry together.</p>
# + dc={"key": "45"} tags=["sample_code"]
# Filter and diplay halftime musicians before and including Super Bowl XXVII
( pre_MJ <- halftime_musicians %>%
filter(super_bowl <= 27) )
# + dc={"key": "52"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 8. Who has the most halftime show appearances?
# <p>Now that's a lot of marching bands! There was also the American jazz clarinetist, <NAME>, and Miss Texas 1973 played the violin. Nothing against those performers - they are just simply not <a href="https://www.youtube.com/watch?v=suIg9kTGBVI">Beyoncé</a>. To be fair, no one is.</p>
# <p>Let's find all the musicians who performed at the Super Bowl more than once and count their performances.</p>
# + dc={"key": "52"} tags=["sample_code"]
# Display the musicians who performed more than once
halftime_musicians %>%
count(musician, sort = TRUE) %>%
filter(n > 1)
# + dc={"key": "59"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 9. Who performed the most songs in a halftime show?
# <p>The world-famous <a href="https://www.youtube.com/watch?v=RL_3oqpHiDg">Grambling State University Tiger Marching Band</a> takes the crown with six appearances. Beyoncé, <NAME>, Nelly, and <NAME> are the only post-Y2K musicians with multiple appearances (two each).</p>
# <p>Now let's look at the number of songs performed in a halftime show. From our previous inspections, the <code>num_songs</code> column has a lot of missing values:</p>
# <ul>
# <li>A lot of the marching bands don't have <code>num_songs</code> entries.</li>
# <li>For non-marching bands, there is a lot of missing data before Super Bowl XX.</li>
# </ul>
# <p>Let's filter out marching bands by using a string match for "Marching" and "Spirit" (a common naming convention for marching bands is "Spirit of [something]"). We'll only keep data from Super Bowls XX and later to address the missing data issue, and <em>then</em> let's see who performed the most number of songs.</p>
# + dc={"key": "59"} tags=["sample_code"]
# Remove marching bands and data before Super Bowl XX
musicians_songs <- halftime_musicians %>%
filter(!str_detect(musician, "Marching"),
!str_detect(musician, "Spirit"),
super_bowl > 20)
# Plot a histogram of the number of songs per performance
ggplot(musicians_songs, aes(num_songs)) +
geom_histogram(binwidth = 1) +
labs(x = "Number of songs per halftime show", y = "Number of musicians")
# Display the musicians with more than four songs per show
musicians_songs %>%
filter(num_songs > 4) %>%
arrange(desc(num_songs))
# + dc={"key": "66"} deletable=false editable=false run_control={"frozen": true} tags=["context"]
# ## 10. Conclusion
# <p>Most non-band musicians do 1 to 3 songs per halftime show. It's important to note that the duration of the halftime show is fixed (roughly 12 minutes) so songs per performance is more a measure of how many hit songs you have (cram as many hit songs in as you can!). Timberlake went off in 2018 with 11 songs! Wow! <NAME> comes in second with a ten song medley in 1996.</p>
# <p>In this notebook, we loaded, cleaned, and explored Super Bowl game, television, and halftime show data. We visualized the distributions of combined points, point differences, and halftime show performances using histograms. We used line plots to see how advertisement cost increases lagged behind viewership increases. And, we discovered that blowouts appear to lead to a drop in viewership.</p>
# <p>This year's Big Game will be here before you know it. Who do you think will win Super Bowl LIII?</p>
# <p><em>UPDATE: <a href="https://en.wikipedia.org/wiki/Super_Bowl_LIII">Spoiler alert</a>.</em></p>
# + dc={"key": "66"} tags=["sample_code"]
# 2018-2019 conference champions
patriots <- "New England Patriots"
rams <- "Los Angeles Rams"
# Who will win Super Bowl LIII?
super_bowl_LIII_winner <- patriots
paste("The winner of Super Bowl LIII will be the", super_bowl_LIII_winner)
# -
| data-science-projects/Super bowl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Traning YOLOv5 to detect t-shirts of Olympique de Marseille (OM) and Paris Saint-Germain (PSG)
#
# ## About YOLO
#
# YOLO (you only look once) is an object detection model proposed by [Redmon et al. in 2015](https://pjreddie.com/media/files/papers/yolo_1.pdf). From your first version, the YOLO architecture had modifications in your backbone and, the way to predict the bounding box coordinates. Nowadays the YOLO family counts with five different versions, the first three versions proposed by Redmon [YOLOv1](https://pjreddie.com/media/files/papers/yolo_1.pdf), [YOLO9000](https://arxiv.org/pdf/1612.08242.pdf) and [YOLOv3](https://arxiv.org/pdf/1804.02767.pdf). The YOLOv4 was proposed by [<NAME> et al.](https://arxiv.org/pdf/2004.10934.pdf) The Yolov5 was proposed by <NAME>, there's no article for this version, but the repository with the version, [here](https://github.com/ultralytics/yolov5).
#
# YOLO is an unified model, it means, the model considers one single Convolutional Neural Network to detect objects and classify them. Before YOLO, the models of object detection were built in two steps, first the detection of all possibles bounding boxes and then, the classification. These models as Faster R-CNN are very accurate, on the other hand, very slow. The representation of YOLO model can be seen in the follow image.
# 
#
# The image come from the first [YOLO paper](https://pjreddie.com/media/files/papers/yolo_1.pdf). As we can see, YOLO considers the image as a grid cell. Each cell on the grid has two functions, predict a class and, if the center of the object is in the cell, predict all possibles bounding boxes. The detection of the object and class is a regression problem in this model, the predictions of bounding boxes and class are made in the fully-connected layers.
#
# From the [second version](https://arxiv.org/pdf/1612.08242.pdf), YOLO changed the way to calculate the bounding boxes, YOLO considers anchors boxes using k-means, removing the fully-connected layers. The class prediction and objectness are calculated according with each anchor box. To calculate the bounding boxes, we have
#
# $b_x = \sigma(t_x) + c_x$
#
# $b_y = \sigma(t_y) + c_y$
#
# $b_w = p_we^\left(t_w\right)$
#
# $b_h = p_he^\left(t_h\right)$.
#
# The network predicts 5 coordinates $t_x$, $t_y$, $t_w$ and $t_h$, over the condition if the cell is offset from the top left corner of the image by $(c_x, c_y)$ and if, the bound box has width $p_w$ and height $p_h$. The term $\sigma$ represents the sigmoid function. Above an illustration from the [paper](https://arxiv.org/pdf/1612.08242.pdf).
# 
#
# The [third version](https://arxiv.org/pdf/1804.02767.pdf) of YOLO considers a new backbone for the model, using the DarkNet-56 (until YOLOv4 the DarkNet framework was considered), implementing residuals layers. The model also makes predictions across scales, considering grids of cells of different sizes, to take care about small, medium and larger objects. The output of the model is a tensor with four coordinates for the box, the objectness and the class predictions.
#
# The [fourth version](https://arxiv.org/pdf/1612.08242.pdf) of YOLO are influenced by state of art models as BoF (bag of freebies) and BoS (bag of specials). The BoF improve the accuracy of the detector. The BoS increase a bit the inference cost, it can significantly improve the object detection metrics.
#
# The [last version](https://github.com/ultralytics/yolov5) of YOLO is different from the previous versions. The model was developed using PyTorch, different to others versions that use DarkNet framework. The model, as YOLOv4, considers [CSP backbone](https://openaccess.thecvf.com/content_CVPRW_2020/papers/w28/Wang_CSPNet_A_New_Backbone_That_Can_Enhance_Learning_Capability_of_CVPRW_2020_paper.pdf) and [PA-NET](https://arxiv.org/pdf/1803.01534.pdf) at the neck of the network (segmetation). Additionally, the model introduces mosaic data augmentation and auto learning bounding box anchors.
#
# This project considers the last version of YOLO. Why? To experiment the model. Before, I've trained on DarkNet framework, using YOLOv3, the results are good, the YOLO models are fast and accurate. Using the YOLOv5 the training process is directly, with few line commands. You can easily have access to train script and know about all parameters. In addition, the output model is lightweight.
#
# ## Dataset
#
# The dataset for this model is composed by $320$ images of each team. The annotations was made with LabelImg. Even if the model resizes the images along the training process, before start it, all images was resized into $(640x640)$, conserving the aspect ratio.
#
# **Class balance**
#
# Among the $640$ image, we have $920$ annotated t-shirt. For the OM, we have $457$ instances and for the PSG $472$.
#
# **Background images**
#
# Additionally, we consider $64$ image as background, $10\%$ of the dataset. The background images do not contain objects to be predicted.
#
# If you want to train the model, you must select the images from the original [dataset](https://github.com/IgorMeloS/OMxPSG-T-Shirt-Detection/tree/main/dataset). For the labeling, you can use any tool of your preference, but remember to save the annotations into YOLO format.
# ## Importing Torch
import torch
torch.cuda.is_available()
torch.cuda.empty_cache() # doesn’t increase the amount of GPU memory available for PyTorch.
# ## Small YOLO v5 architecture
#
# YOLOv5 has some size of architecture, for this experimental project, we run the model with the small architecture composed by $24$ convolutional layers and the detection layer. The reason to use this model is due the time running and limitation of GPU memory. More lager models give better results. In the the next cells, we can see the model architecture and learning parameters.
# %cat yolov5/models/yolov5s.yaml
# %cat yolov5/data/hyps/hyp.scratch.yaml
# ## Training process
#
# To train the YOLOv5, first of all, you must download the model from [GitHub] and install the requirements.
#
# The train script found in the yolov5 folder requires some arguments.
#
# - --img (input image size)
# - --batch (batch size)
# - --epochs (number of epochs)
# - --data (file in yaml format with the dataset directory, name and number of classes)
# - --project (directory to save the model)
# - --workers (number of data loaders)
#
# It's recommended to check the train script to know all possible arguments. The model was trained over $150$ epochs. The batch size is $16$ due to a limitation of GPU memory, but if you can increase, it's better.
#
# The data.yaml file must be allocated inside the yolov5 folder, this contains the follow informations
#
# train: ../dataset/train/images
# val: ../dataset/valid/images
#
# nc: 2
# names: ['OM', 'PSG']
#
# The annotations file must be allocatted in the follow maner ../dataset/train/labels.
#
# Other arguments can be passed, for example, if you want to store to save check-points you can pass --save-period, the artifact will be saved on the [Wandb](https://wandb.ai/home) platform. During the training process you can follow the metrics curve evolution with Wandb, but it's optional.
#
# The training set is composed by $495$ images (including background, the validation set is composed by $220$ images.
# !cd yolov5 && python train.py --img 640 --batch 16 --epochs 150 --data data.yaml --project ../results --weights yolov5s.pt --workers 1
# ## Results
#
# At the end of the training process, the will be saved into the results folder. There are some outputs, as a csv file with the metrics evolution, images with some plots, as confusion matrix, for example.
#
# When we lead with object detection, we seek to get the best results for the mAP_0.5 and mAP_0.5:0.95 metrics. To measure this metrics, we take account two other metrics, precision and recall. Beyond these metrics, when we work with YOLO, it's important to see the evolution of the loss function for the Box predictions, Objectness and Class predictions. Now, we present these results.
import pandas as pd
import matplotlib.pyplot as plt
results = pd.read_csv('results/exp/results.csv')
results.columns
# ### Loss Functions
#
# To get more details how these functions are calculated, [see](https://pjreddie.com/media/files/papers/yolo_1.pdf).
plt.figure(figsize=(22,10))
plt.subplot(2,2,1)
plt.plot(results[' train/box_loss'], label = 'train')
plt.plot(results[' val/box_loss'], label = 'validation')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('box_loss')
plt.title('Box loss functions')
plt.axis((0, 150, 0.01,0.11))
plt.subplot(2,2,2)
plt.plot(results[' train/obj_loss'], label = 'train')
plt.plot(results[' val/obj_loss'], label = 'validation')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('objectness_loss')
plt.axis((0, 150, 0.005,0.033))
plt.title('Objectness loss functions')
plt.subplot(2,2,3)
plt.plot(results[' train/cls_loss'], label = 'train')
plt.plot(results[' val/cls_loss'], label = 'validation')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('class_loss')
plt.axis((0, 150, 0.00,0.033))
plt.title('Class loss functions')
plt.savefig("Loss functions.jpg")
plt.show()
# ### mAP_0.5 and m_AP_0.5:0.95
#
# About the metrics, there's this [blog](https://blog.paperspace.com/mean-average-precision/) that explains perfectly. So, I passthrough.
plt.figure(figsize=(22,10))
plt.subplot(2,2,1)
plt.plot(results[' metrics/precision'], label = 'precision')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('precision')
plt.axis((0, 150, 0.0, 1))
plt.title('Precision evolution')
plt.subplot(2,2,2)
plt.plot(results[' metrics/recall'], label = 'recall')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('recall')
plt.axis((0, 150, 0.0, 1))
plt.title('Recall evolution')
plt.subplot(2,2,3)
plt.plot(results[' metrics/mAP_0.5'], label = 'mAP_0.5', color = 'red')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('mAP_0.5')
plt.axis((0, 150, 0.00,1))
plt.title('mAP_0.5 evolution')
plt.subplot(2,2,4)
plt.plot(results['metrics/mAP_0.5:0.95'], label = 'mAP_0.5:0.95', color = 'red')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('mAP_0.5:0.95')
plt.axis((0, 150, 0.00,0.75))
plt.title('mAP_0.5:0.95 evolution')
plt.savefig('metrics.jpg')
plt.show()
# ## Conclusions
#
# The experience with YOLOv5 was agreeable in terms of results. Firstly, we don't have a large dataset and, we don't have many instances by class, even so, the model shows good results.
#
# Looking the loss function results, we can see that our curves are closely. Even if, the results for the Box loss show a difference between the training and validation, the gap among these curves is not enough large to be considered in over-fitting, the same analysis for the Objectness loss. The curves for the class prediction loss show the same behavior, indicating that the model was able to generalize the predictions.
#
# The mAP_0.5 result at the end of training was $0.948$, a good value, indicating that IoU (Intersection over Union) between ground truth and the predict object give a high score. The evolution of mAP_0.5:0.95 also presents good performance.
#
# The model was trained using the weights from YOLOv5, to improve the results, we must grab more images and use a more larger architecture. The test of this model can be found [here](https://github.com/IgorMeloS/OMxPSG-T-Shirt-Detection/blob/main/OMxPSG_YOLO/testing.ipynb).
| OMxPSG_YOLO/training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Ordered Logit Example
#
# Let's suppose, completely hypothetically, that we are not a big fan of Stata or simply want to learn the mechanics behind an ordered logit model by coding it up ourselves.
#
# In this notebook we show you how estimagic can help you to implement such a model very easily. Implementing a logit model consists of four basic steps:
#
# 1. Processing the user input into inputs for the likelihood function
# 2. Writing the likelihood function of an ordered logit model
# 3. Maximizing the likelihood function
# 4. Calculating standard errors
#
# The first two have to be done by the user, the last two are done by estimagic.
#
# To be very clear: Estimagic is not a package to estimate logit models or other models that are implemented in Stata, statsmodels or anywhere else. Its purpose is to estimate parameters with custom likelihood or method of simulated moments functions. We just use an orederd logit model as an example of a very simple likelihood function.
#
# The example we will use to test our model is taken from the [Stata Documentation](https://stats.idre.ucla.edu/stata/dae/ordered-logistic-regression/).
# +
import numpy as np
import pandas as pd
from patsy import dmatrices
from scipy import stats
from estimagic import maximize
from estimagic.inference.likelihood_inference import do_likelihood_inference
# -
# ### Process the user input
#
# First we have to take the formula and dataset, extract all relevant information about the model and construct the inputs for the likelihood function.
#
# We will need four inputs:
#
# 1. A DataFrame with start parameters for the optimization.
# 2. An array with the dependent variable.
# 3. A 2d array with explanatory variables.
# 4. Constraints for the optimization that keep the cutoffs increasing.
def ordered_logit_processing(formula, data):
"""Process user input for an ordered logit model."""
# extract data arrays
y, x = dmatrices(formula + " - 1", data, return_type="dataframe")
y = y[y.columns[0]]
# extract dimensions
num_choices = len(y.unique())
beta_names = list(x.columns)
num_betas = len(beta_names)
num_cutoffs = num_choices - 1
# set-up index for params_df
names = beta_names + list(range(num_cutoffs))
categories = ["beta"] * num_betas + ["cutoff"] * num_cutoffs
index = pd.MultiIndex.from_tuples(zip(categories, names), names=["type", "name"])
# make params_df
np.random.seed(5471)
start_params = pd.DataFrame(index=index)
start_params["value"] = np.hstack(
[
np.random.uniform(low=-0.5, high=0.5, size=len(x.columns)),
np.arange(num_cutoffs) * 2,
]
)
start_params["group"] = start_params.index.get_level_values("type")
# make constraints
constr = [{"loc": "cutoff", "type": "increasing"}]
# turn pandas objects into numpy arrays
y_arr = y.to_numpy().astype(int)
x_arr = x.to_numpy()
return start_params, y_arr, x_arr, constr
# ### Calculate the Likelihood
#
# Next, we want to evaluate the likelihood function, given parameters and data. There are more efficient ways of calculating the likelihood for an ordered logit but this one was chosen for brevity and readability.
def ordered_logit_loglike(params, y, x):
"""Likelihood function of an orderd logit model."""
# parse the parameter vector into its quantities
beta = params.loc["beta", "value"].to_numpy()
cutoffs = params.loc["cutoff", "value"].to_numpy()
# calculate deterministic part of utilities
xb = x.dot(beta)
# evaluate likelihood
upper_cutoffs = np.hstack([cutoffs, np.inf])[y]
lower_cutoffs = np.hstack([-np.inf, cutoffs])[y]
upper_cdf = stats.logistic.cdf(upper_cutoffs - xb)
lower_cdf = stats.logistic.cdf(lower_cutoffs - xb)
contributions = np.log(upper_cdf - lower_cdf)
res = {"contributions": contributions, "value": contributions.sum()}
return res
# If you have never programmed an estimator before, you migt be surprised how much code is spent on processing compared to calculating the actual likelihood function. This will almost always be the case -
# at least if you try to make your estimator flexible and user friendly. Estimagic is there to shorten this type of code as much as possible.
#
# Another peculiarity you might notice is that the likelihood function does not just return a scalar value, but also the likelihood contributions of each individual. This is because some optimizers (e.g. bhhh) can actually use the information on the contributions. Morover, you will need the contributions to calculate standard errors by the outer product of gradients.
#
# All estimagic functions (whether for numerical differentiation, standard error calculation or optimization) will simply pick from the dictionary what they need!
#
#
# ### Maximizing the likelihood
# +
data = pd.read_pickle("ologit.pickle")
formula = "apply ~ pared + public + gpa"
start_params, y, x, constraints = ordered_logit_processing(formula, data)
res = maximize(
criterion=ordered_logit_loglike,
params=start_params,
algorithm="scipy_lbfgsb",
constraints=constraints,
criterion_kwargs={"y": y, "x": x},
logging="ordered_logit.db",
)
# -
params = res["solution_params"]
params
# ### Calculate standard errors
# +
from estimagic.decorators import numpy_interface
numpy_interface(ordered_logit_loglike, params=params, constraints=constraints)
# +
inference = do_likelihood_inference(
loglike=ordered_logit_loglike,
params=params,
loglike_kwargs={"x": x, "y": y},
n_samples=10_000,
constraints=constraints,
)
inference["summary"].round(3)
# -
# ### Compare to STATA's results
stata_results = pd.read_csv("stata_ologit_results.csv")
stata_results.round(3)
# This looks pretty good! The parameter estimates line up perfectly. I actually had to try three optimizers to get at least one differenet digit which makes the result more credible. Other optimizers hit it on all digits.
#
# <div class="alert alert-danger">
# Note that standard error calculation, especially in combination with constraints is still considered experimental in estimagic.
# </div>
#
#
# ### Use the dashboard for monitoring the optimization
#
# Often you may want to monitor an optimization to see how far the algorithm has moved away from the start values or see how the
# algorithm arrived at its solution after it has finished.
#
#
# Both can be done using the estimagic dashboard.
#
# To use the dashboard, we need to activate logging
# which we had deactivated up until now.
# To activate logging, simply supply a database path to
# `ordered_logit`.
#
# To start the dashboard, make sure you have the estimagic environment
# installed and activated.
#
# Then all you need to do is navigate to the path's directory in your
# command line, start the cell below and enter the following into
# your command line after the optimization has started:
#
# ``estimagic dashboard {db_path}``
#
# This should open a page in your browser where you can press
# "Start Updating from Database" to start watching the optimization.
# +
db_path = "./logging.db"
res = maximize(
criterion=ordered_logit_loglike,
params=start_params,
algorithm="scipy_lbfgsb",
constraints=constraints,
criterion_kwargs={"y": y, "x": x},
logging=False,
)
| docs/source/getting_started/ordered_logit_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.7 64-bit
# name: python3
# ---
import os
import os.path as path
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tensorflow.keras import layers, models, optimizers, regularizers
from tensorflow.keras.models import load_model
current_dir = os.path.join(os.getcwd())
file = os.path.join(path.dirname(path.dirname(current_dir)), "generate_data\data_cwa.csv")
myData = pd.read_csv(file, delimiter=',', usecols=['cwa','credit','time','difficulty', 'score'])
my_data_copy = myData
myData.shape
myData["score"] = myData["score"].values / 100
myData["cwa"] = myData["cwa"].values / 100
myData["credit"] = myData["credit"].values / 10
myData ["difficulty"] = myData['difficulty'].values / 5
myData["time"] = myData["time"].values / 6
# +
df = pd.DataFrame(myData)
df = df.sample(frac=1)
myData = df
# +
targets = myData[['time']].values
myData.drop(('time'), axis=1, inplace=True)
data = myData.values
print(targets.shape)
print(data.shape)
# +
# num_train = int(0.5 * len(data))
# num_val = int(0.25 * len(data))
# num_test = int(0.25 * len(data))
# +
# train_data = data[0 : num_train]
# test_data = data[num_train: num_train + num_test]
# val_data = data[num_train + num_test:]
# train_targets = targets[0 : num_train]
# test_targets = targets[num_train: num_train + num_test]
# val_targets = targets[num_train + num_test:]
train_data = data[0 : 2000]
test_data = data[2000: 3000]
val_data = data[3000:4000]
train_targets = targets[0 : 2000]
test_targets = targets[2000: 3000]
val_targets = targets[3000 :4000]
print(len(train_data) + len(test_data) + len(val_data))
print(len(train_targets) + len(test_targets) + len(val_targets))
# +
model = models.Sequential()
model.add(layers.Dense(64, activation="relu", input_shape=(train_data.shape[1],)))
# model.add(layers.Dropout(0.5))
model.add(layers.Dense(64, activation="relu"))
# model.add(layers.Dropout(0.5))
model.add(layers.Dense(64, activation="relu"))
# model.add(layers.Dropout(0.5))
model.add(layers.Dense(1))
model.summary()
# -
model.compile(
optimizer=optimizers.RMSprop(learning_rate=2e-4),
loss="mse",
metrics=['mae']
)
history = model.fit(train_data,
train_targets,
epochs=50,
batch_size=100,
validation_data=(val_data, val_targets)
)
acc = history.history['mae']
val_acc = history.history['val_mae']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(1, len(acc) + 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss, 'bo', label='Training loss')
# b is for "solid blue line"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
test_mse_score, test_mae_score = model.evaluate(test_data, test_targets)
model.save('time_prediction.h5')
predicted = model.predict([[0.8771, 0.4, 0.6, 0.82]])
predicted
| app/prediction/cwa_prediction/time_prediction/time_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.5
# language: julia
# name: julia-1.0
# ---
# # Getting Started with JuMP
# **Originally Contributed by**: <NAME>
# This tutorial is aimed at providing a quick introduction to writing JuMP code. It assumes familiar with basic optimization and
# the notion of an [AML](https://en.wikipedia.org/wiki/Algebraic_modeling_language).
# ## What is JuMP?
# JuMP ("Julia for Mathematical Programming") is an open-source modeling language that is embedded in Julia. It allows users to
# users formulate various classes of optimization problems (linear, mixed-integer, quadratic, conic quadratic, semidefinite,
# and nonlinear) with easy-to-read code. These problems can then be solved using state-of-the-art open-source and commercial solvers.
# JuMP also makes advanced optimization techniques easily accessible from a high-level language.
# ## Installing JuMP
# JuMP is a package for Julia. From Julia, JuMP is installed by using the built-in package manager.
import Pkg
Pkg.add("JuMP")
# ## A Complete Example
# Let's try to solve the following linear programming problem by using JuMP and GLPK (a linear and mixed integer programming
# solver). We will first look at the complete code to solve the problem and then go through it step by step.
# $$
# \begin{align*}
# & \min & 12x + 20y \\
# & \;\;\text{s.t.} & 6x + 8y \geq 100 \\
# & & 7x + 12y \geq 120 \\
# & & x \geq 0 \\
# & & y \geq 0 \\
# \end{align*}
# $$
# +
using JuMP
using GLPK
model = Model(GLPK.Optimizer)
@variable(model, x >= 0)
@variable(model, y >= 0)
@constraint(model, 6x + 8y >= 100)
@constraint(model, 7x + 12y >= 120)
@objective(model, Min, 12x + 20y)
optimize!(model)
@show value(x);
@show value(y);
@show objective_value(model);
# -
# ## Step by Step JuMP Code
# Once JuMP is installed, to use JuMP in your programs, we just need to write-
using JuMP
# We also need to include a Julia package which provides an appropriate solver. We want to use GLPK.Optimizer here which is
# provided by the GLPK.jl package.
using GLPK
# A model object is a container for variables, constraints, solver options, etc. Models are created with the Model() function.
# The model can be created with an optimizer attached with default arguments by calling the constructor with the optimizer type, as follows:
model = Model(GLPK.Optimizer);
# A variable is modelled using `@variable(name of the model object, variable name and bound, variable type)`. The bound can be a
# lower bound, an upper bound or both. If no variable type is defined, then it is treated as real.
@variable(model, x >= 0)
@variable(model, y >= 0);
# A constraint is modelled using `@constraint(name of the model object, constraint)`.
@constraint(model, 6x + 8y >= 100)
@constraint(model, 7x + 12y >= 120);
# The objective is set in a similar manner using `@objective(name of the model object, Min/Max, function to be optimized)`
@objective(model, Min, 12x + 20y);
# To solve the optimization problem, we call the optimize function.
optimize!(model)
# Let's now check the value of objective and variables.
@show value(x);
@show value(y);
@show objective_value(model);
| notebook/introduction/getting_started_with_JuMP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import flowiz as f
import glob
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
files = glob.glob('flo/*.flo')
files
# Convert from [.flo] file
img = f.convert_from_file(files[0]) # load flo from file
plt.imshow(img)
# Convert from [.flo] array
floArray = f.read_flow(files[0]) # some random flo array
floArray[0]
plt.imshow(f.convert_from_flow(floArray))
# +
# Two channel image [u,v] with horizontal and vertical flows converted to UINT8
uv = f.convert_from_flow(floArray, mode='UV')
f, axarr = plt.subplots(1,2)
f.set_size_inches(15, 5)
axarr[0].set_title('Horizontal Flow (U)')
axarr[0].imshow(uv[...,0],cmap=plt.get_cmap('binary'))
axarr[1].set_title('Vertical Flow (V)')
axarr[1].imshow(uv[...,1],cmap=plt.get_cmap('binary'))
# -
import os
os.popen('python -m flowiz flo/*.flo -o png/ -v mp4/ -r 2 ').read().split('\n')
| demo/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
Car.objects.all()
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import os
import datetime as dt
cwd=os.getcwd()
print(cwd)
os.path.join(cwd,'data','week11 dsi202 - Car.csv')
# +
path=os.path.join(cwd,'data','week11 dsi202 - Car.csv')
df_car=pd.read_csv(path)
path=os.path.join(cwd,'data','week11 dsi202 - Customer.csv')
df_customer=pd.read_csv(path)
# -
df_car.info()
df_customer.info()
#customer,car,cost,start,stop
df_rent=pd.DataFrame(columns=['customer','car','cost','start','stop'])
df_rent=pd.DataFrame(columns=['customer','car','cost','start','stop'])
df_rent.start.astype('datetime64',copy=True)
for i in range(100):
year=2018
month=np.random.randint(low=1,high=13)
day=np.random.randint(low=1,high=32)
hour=np.random.randint(low=0,high=24)
minute=np.random.randint(low=0,high=60)
try:
d=dt.datetime(year,month,day,hour,minute)
except:
print('pass:%s, %s, %s, %s, %s'%(year,month,day,hour,minute))
continue
delta = dt.timedelta(
hours = np.random.randint(low=12,high=168),
minutes = np.random.randint(low=0,high=60),
)
df_rent=df_rent.append({
'customer':np.random.randint(low=1,high=23),
'car':np.random.randint(low=1,high=17),
'cost':np.random.randint(low=200,high=5000),
'start':str(d),
'stop':str(d+delta)
}, ignore_index=True)
df_rent.head()
df_rent.to_csv('data/rent.csv')
# # Note
print(dt.datetime(2019,10,25,7,15,55,0))
str(dt.datetime(2019,10,25,7,15,55,0))
d=dt.datetime(2019,10,25,7,15,55,0)
d.strftime("%d/%m/%y")
now = dt.datetime.now()
now
delta = dt.timedelta(hours = 12, minutes=10)
now+delta
np.random.rand(5)
x=np.random.rand(1000)
hist, bin_edges=np.histogram(x,bins=20)
plt.bar(bin_edges[1:], hist)
x=np.random.randn(1000)*2
hist, bin_edges=np.histogram(x,bins=20)
plt.bar(bin_edges[:-1], hist)
| dsi202/week11/projectname/Import Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='logbook'></a>
# # Summary 2020-12-15
# + slideshow={"slide_type": "skip"}
# # %load imports.py
"""
These is the standard setup for the notebooks.
"""
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from jupyterthemes import jtplot
jtplot.style(theme='onedork', context='notebook', ticks=True, grid=False)
import pandas as pd
pd.options.display.max_rows = 999
pd.options.display.max_columns = 999
pd.set_option("display.max_columns", None)
import numpy as np
import os
import matplotlib.pyplot as plt
from collections import OrderedDict
#plt.style.use('paper')
#import data
import copy
from mdldb.run import Run
from sklearn.pipeline import Pipeline
from rolldecayestimators.transformers import CutTransformer, LowpassFilterDerivatorTransformer, ScaleFactorTransformer, OffsetTransformer
from rolldecayestimators.direct_estimator_cubic import EstimatorQuadraticB, EstimatorCubic
from rolldecayestimators.ikeda_estimator import IkedaQuadraticEstimator
import rolldecayestimators.equations as equations
import rolldecayestimators.lambdas as lambdas
from rolldecayestimators.substitute_dynamic_symbols import lambdify
import rolldecayestimators.symbols as symbols
import sympy as sp
from sympy.physics.vector.printing import vpprint, vlatex
from IPython.display import display, Math, Latex
from sklearn.metrics import r2_score
from src.data import database
from mdldb import tables
# + [markdown] slideshow={"slide_type": "skip"}
# ## Nomenclature
# | Variable | Explain |
# |---|---|
# |$\pi$| example |
# + [markdown] slideshow={"slide_type": "skip"}
# Here is a cell link: [Logbook](#logbook)
# + [markdown] slideshow={"slide_type": "skip"}
# # Abstract
#
# Many cost-efficient computation methods have been developed over the years to analyze various aspects of ship hydrodynamics such as: resistance, propulsion and seakeeping. Getting the best possible accuracy with the lowest possible computational cost is an important factor in a ship’s early design stage. Potential flow-based analysis partly presents such a solution for seakeeping, with good accuracy for heave and pitch, but not for roll where the roll damping contains both inviscid and viscouseffects. Roll motion is, however, often a critical degree of freedom that needs to be analyzed since large roll motions can result in cargo shifting or even capsizing. The viscous part of roll damping can be assessed with high accuracy by means of experimental model tests or URANS calculations, but these are generally too expensive in the early design stage of ships. Many semi-empirical formulas to determine viscous damping were therefore developed during the 1970s, where Ikeda’s method is one of the most widely used. The viscous damping from this method is normally combined with inviscid roll damping from strip theory.
#
# With today’s computational power, more advanced potential flow methods can be used in the seakeeping analysis to enhance the accuracy in the predictions, but still at relatively low computational cost. This paper investigates the feasibility of combining 3D unsteady fully nonlinearpotential flow (FNPF) theory solved by means of a Boundary ElementMethod (BEM) together with the viscous contributions from Ikeda’smethod.
#
# The approach of substituting the inviscid part from Ikeda’s method using strip theory with FNPF is investigated by conducting roll decay simulations. The results estimated by the proposed approach are compared with both the classic strip theory approach and roll decay model tests.
# **It is found that potential improvements to the modelling of roll damping can be achieved by introducing FNPF analysis in the Ikeda’s method.**
# + [markdown] slideshow={"slide_type": "slide"}
# # Abstract
# + [markdown] slideshow={"slide_type": "fragment"}
# * **Important**: Good accuracy at low computational cost.
# + [markdown] slideshow={"slide_type": "notes"}
# (URANS or model test too expensive)
# + [markdown] slideshow={"slide_type": "fragment"}
# * → seakeeping: Potential flow
# + [markdown] slideshow={"slide_type": "fragment"}
# * heave/pitch good!
# + [markdown] slideshow={"slide_type": "fragment"}
# * **NOT** roll!
# + [markdown] slideshow={"slide_type": "fragment"}
# * **Solution**: Potential flow + semi empirical viscous roll damping
# + [markdown] slideshow={"slide_type": "fragment"}
# * Potential flow:
# * Strip theory (milli seconds)
# * Nonlinear 3D methods (hours)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
# The roll damping can be divided into various components <cite data-cite="7505983/4AFVVGNT"></cite>:
# $$B_{44} = B_F + B_E + B_L + B_W + B_{BK}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# Viscous: $ B_{visc} = B_F + B_E + B_L + B_{BK} $ (Ikeda's method, Simplified Ikeda)
# + [markdown] slideshow={"slide_type": "fragment"}
# Inviscid: $ B_{invisc} = B_W $ (Potential flow)
# + [markdown] slideshow={"slide_type": "fragment"}
# $$B^{Ikeda} = B_{invisc}^{1D} + B_{visc}$$
# + [markdown] slideshow={"slide_type": "fragment"}
# $$B^{Motions} = B_{invisc}^{3D} + B_{visc}$$
# + [markdown] slideshow={"slide_type": "slide"}
# ### Problems with $B_W$ in Simplified Ikeda
# 
# + slideshow={"slide_type": "skip"}
data = {
'KVLCC2' : {
'type':'tanker',
'test data':True,
'B_W': 'small',
'bilge keel':False,
'publish geom':True,
'publish test':True,
},
'DTC' : {
'type':'container',
'test data':True,
'B_W': '?',
'bilge keel':True,
'publish geom':True,
'publish test':True,
},
'Wallenius' : {
'type':'PCTC',
'test data':True,
'B_W': 'medium',
'bilge keel':True,
'publish geom':False,
'publish test':True,
},
}
test_cases = pd.DataFrame(data=data).transpose()
# + slideshow={"slide_type": "skip"}
def background_colorer(val):
return 'background-color: %s' % get_color(val)
def text_colorer(val):
return 'color: %s' % get_color(val)
def get_color(val):
color = 'none'
if isinstance(val, bool):
if val:
color = 'green'
else:
color = 'red'
return color
# + [markdown] slideshow={"slide_type": "slide"}
# ## Possible test cases:
# + slideshow={"slide_type": "-"}
test_cases.style.applymap(background_colorer).applymap(text_colorer)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Test case: KVLCC2
# [04.3_KVLCC2_Ikedas_model_tests](../../notebooks/04.3_KVLCC2_Ikedas_model_tests.ipynb)
# + [markdown] cell_style="center" slideshow={"slide_type": "slide"}
# <img src="../../reports/figures/KVLCC2_B_e_0.0.png" width="1100">
# + [markdown] slideshow={"slide_type": "notes"}
# * B_W very small!
# * B_E is dominating
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="../../reports/figures/KVLCC2_B_e_15.5.png" width="1100">
# + [markdown] slideshow={"slide_type": "notes"}
# * B_E decreased, B_L is now dominating
# * B_W is a bit larger but still minor
# + [markdown] slideshow={"slide_type": "slide"}
# # Abstract conclusion:
# #### "It is found that potential improvements to the modelling of roll damping can be achieved by introducing FNPF analysis in the Ikeda’s method"
# + [markdown] slideshow={"slide_type": "fragment"}
# # Is this what we are aiming for?
# + [markdown] slideshow={"slide_type": "slide"}
# ## References
# <div class="cite2c-biblio"></div>
| reports/presentation_2020-12-15/summary_2020-12-15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Answering Descriptive and Exploratory Questions About my Project
# +
# Import packages
import igraph as ig
import numpy as np
import math
import os
from subprocess import Popen, PIPE
# Initializing dataset names
dnames = list(['../data/desikan/MRN114', '../data/desikan/KKI2009', '../data/desikan/SWU4'])
print "Datasets: " + ", ".join(dnames)
print "D = " + str(len(dnames))
# Getting graph names
fs = list()
for dd in dnames:
fs.extend([root+'/'+file for root, dir, files in os.walk(dd) for file in files])
# fs
# -
# **Descriptive** <br />
# - *What is N<sub>i</sub> for all i?*
# Get lengths of sublists and total list
print "N_i for each dataset (same order as above): " +\
", ".join([str(len(filter(lambda x: dd in x, fs))) for dd in dnames])
print "Total N = " + str(len(fs))
# - *What is |V|?*
# We know that |V| is the same for all graphs, so here we really only need to load in 1
graph = ig.Graph.Read_GraphML(fs[0])
V = graph.vcount()
print "|V| = " + str(V)
# - *Do the graphs G<sub>n<sub>i</sub></sub> contain any values of A that cannot be processed traditionally (i.e. inf, NaN)?*
# +
# We actually need the graphs in memory now, it seems. I'll make a janky function for this
# in case I want to do it again later for some reason.
def loadGraphs(filenames, rois, printer=False):
A = np.zeros((rois, rois, len(filenames)))
for idx, files in enumerate(filenames):
if printer:
print "Loading: " + files
g = ig.Graph.Read_GraphML(files)
tempg = g.get_adjacency(attribute='weight')
A[:,:,idx] = np.asarray(tempg.data)
return A
A = loadGraphs(fs, V)
# Parallel index for datasets
c = 0
d_idx = []
for dd in dnames:
d_idx.append([c for root, dir, files in os.walk(dd) for file in files])
c += 1
d_idx = np.concatenate(d_idx)
A.shape
# -
# Now that my graphs are here, let's count NaNs and Infs in the set of them
nans= np.count_nonzero(np.isnan(A))
infs= np.count_nonzero(np.isinf(A))
print "Our data contains " + str(nans) + " NaN values"
print "Our data contains " + str(infs) + " Inf values"
# - *How sparse, |E<sub>n<sub>i</sub></sub>| / |V<sub>n<sub>i</sub></sub>|x|V<sub>n<sub>i</sub></sub>|, are the graphs?*
# First I'll want to binarize my adjacency matrix, then I can do population
# sparsity by summing all edges and diving by total number of possible edges.
# Alternatively, I could've done this per graph and averaged, or per dataset
# and averaged. I chose this one because I did.
bin_graph = 1.0*(A > 0)
sparsity = np.sum(bin_graph) / (V*V*len(fs))
print "The fraction of possible edges that exist in our data is: " + str(sparsity)
# **Exploratory** <br />
# - *What is mean(|E|) for each dataset i?*
# +
# This was computed across all graphs for each data set
bin_graph = 1.0*(A > 0)
for idx in np.unique(d_idx):
print 'Mean edge degree for dataset: ' + dnames[idx] + ' is: ' + \
str(np.sum((bin_graph[:,:,d_idx == idx]))/np.sum(d_idx == idx))
# -
# - *What is the average graph, where average here means the graph consiting of edges and weights corresponding to the average weight of a given potential edge across the all the datasets?*
# +
import matplotlib.pyplot as plt
# %matplotlib inline
font = {'weight' : 'bold',
'size' : 14}
import matplotlib
matplotlib.rc('font', **font)
A_bar = (np.mean(A,axis=2))
plt.figure(figsize=(6, 6))
plt.imshow(A_bar)
plt.xticks((0, 34, 69), ('1', '35', '70'))
plt.yticks((0, 34, 69), ('1', '35', '70'))
plt.xlabel('Node')
plt.ylabel('Node')
plt.title('Mean Connectome')
plt.savefig('../figs/mean_connectome.png')
plt.show()
# -
# - *What is the distribution of max(A)-min(A) (i.e. dynamic range) for each dataset i?*
# +
# first find min(A), max(A) for all data
for idx in np.unique(d_idx):
A_ds = A[:,:,d_idx == idx]
fig = plt.figure()
ax = fig.add_subplot(111)
plt.hist(np.log(np.ravel(A_ds)+1), bins=30) #adding 1 to prevent divide by 0
plt.title('Edge Weights of ' + dnames[idx].split('/')[-1] + ' Dataset')
plt.xlabel("Value (log_e)")
plt.ylabel("Frequency")
ax.set_yscale('log')
plt.savefig('../figs/'+dnames[idx].split('/')[-1]+'_ew_initial.png')
plt.show()
# -
| code/descriptive_and_exploratory_answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Plotting results from How_many.ipynb
#
# %pylab inline
import numpy as np
N = np.arange(0, 170, 4)
N1 = np.arange(0, 170, 4)
N_1 = N1
# +
beta_DB_rev = np.loadtxt('rev_beta_DB_20150907_regperm.txt')
beta_DB_rev_N = np.loadtxt('rev_beta_DB_20150907_regperm.txt')
#N_1 = np.arange(11, 11+len(beta_DB_rev)*4, 4)
#N_1 = np.arange(0, 70, 10)
# +
plt.figure(figsize=(11,8))
plt.plot(N_1, beta_DB_rev, 'o-', lw = 3, ms = 10)
plt.plot(N1, beta_DB_rev_N, 'o-', lw = 3, ms = 10)
plt.xlabel('N',fontsize = 30)
plt.ylabel(r'$\beta$',fontsize = 30)
plt.xlim(0,120)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(37, 0, 1, lw = 3)
plt.title('DB Scan: Permute Width',fontsize = 30)
#plt.savefig("DB_rev.pdf")
# -
# FROM DIU
# +
#k_new = np.array([ 0., 0. , 0.142 , 0.565, 0.845, 0.935])# 0. 0. 0. 0. ])
#Nk_new=np.arange(11, 11+len(k_new)*10, 10)
#print len(k_new), len(Nk_new)
k_mean = np.loadtxt('beta_K_20150907.txt')
N_k = np.arange(0, len(k_mean)*4, 4)
plt.figure(figsize=(11,8))
plt.plot(N_k, k_mean, 'o-', lw = 3, ms = 10)
#plt.plot(Nk_new, k_new, 'o-', lw = 3, ms = 10)
plt.xlabel('N',fontsize = 30)
plt.ylabel(r'$\beta$',fontsize = 30)
plt.title('K-Mean: Permute DM',fontsize = 30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(39, 0, 1, lw = 3)
plt.xlim(0,120)
#plt.savefig("kmean.pdf")
# +
db_scan = np.loadtxt('beta_DB_20150907_regperm.txt')
N_db = np.arange(0, 170, 4)
plt.figure(figsize=(11,8))
plt.plot(N_db, db_scan, 'o-', lw = 3, ms = 10)
plt.xlabel('N',fontsize = 30)
plt.ylabel(r'$\beta$', fontsize = 30)
plt.title('DB Scan: Permute DM',fontsize = 30)
plt.xlim(0,120)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(60, 0, 1, lw = 3)
#plt.savefig("DB.pdf")
# +
##[ 0. 0. 0. 0.005 0.017 0.016 0.016 0.014 0.043 0.122
## 0.241 0.373 0.55 ] np.arange(0, 50, 4)
#np.arange(48+4, 100, 4)
hierarc = [ 0. ,0., 0. , 0.005, 0.017, 0.016, 0.016, 0.014, 0.043, 0.122, 0.241, 0.373, 0.55, \
0.691, 0.743, 0.794, 0.865, 0.88, 0.891, 0.912, 0.926, 0.93, 0.914 , 0.933, 0.933, \
0.936, 0.935, 0.948, 0.955, 0.962, 0.959]#np.loadtxt('beta_Hier_20150907.txt')## ADD EXTRAS
N_h = np.arange(0, len(hierarc)*4, 4)
print len(hierarc), len(N_h)
plt.figure(figsize=(11,8))
plt.plot(N_h, hierarc, 'o-', lw = 3, ms = 10)
plt.xlabel('N', fontsize=30)
plt.ylabel(r'$\beta$',fontsize=30)
plt.xlim(0,120)
plt.title('Hierarchical: Permute DM',fontsize=30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(60, 0, 1, lw = 3)
plt.xlim(0,120)
#plt.savefig("Hier.pdf")
# +
rev_hierar = [0.0, 0. , 0. , 0. , 0. , 0.001 , 0.014 , 0.045 , 0.109 , 0.17, 0.276, \
0.375, 0.476, 0.57, 0.63, 0.695, 0.757, 0.81, 0.812, 0.852,0.873 ,0.874, 0.899 , \
0.896, 0.897, 0.932, 0.898, 0.912, 0.921, 0.93, 0.931, 0.934]#np.loadtxt('rev_beta_Hier_20150907.txt')
N_rh = np.arange(0, len(rev_hierar)*4, 4)
print N_rh
plt.figure(figsize=(11,8))
plt.plot(N_rh, rev_hierar, 'o-', lw = 3, ms = 10)
plt.xlabel('N', fontsize=30)
plt.ylabel(r'$\beta$',fontsize=30)
plt.title('Hierarchical: Permute Width',fontsize=30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(60, 0, 1, lw = 3)
plt.xlim(0,120)
# +
#rev_k_new = np.array([ 0. , 0. , 0.125, 0.541, 0.833, 0.935,])# 0. 0. 0. 0. ])
rev_k_mean = np.loadtxt('rev_beta_K_20150907.txt')
N_rk = np.arange(0, len(rev_k_mean)*4, 4)
plt.figure(figsize=(11,8))
plt.plot(N_rk, rev_k_mean, 'o-', lw = 3, ms = 10)
plt.xlabel('N',fontsize = 30)
plt.ylabel(r'$\beta$',fontsize = 30)
plt.title('K-means: Permute Width',fontsize = 30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(35, 0, 1, lw = 3)
plt.xlim(0,120)
#plt.savefig("kmean_rev.pdf")
# -
print N_h
# +
plt.figure(figsize=(11,8))
for i in range(0, len(N)):
plt.plot([N[i]]*1000, pvals_H[i], 'o',ms = 10, alpha = 0.3, )
plt.hlines(0.05, -1, 120, lw = 3)
plt.xlim(-1, 120)
plt.xlabel('N', fontsize = 30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.ylabel('P-values', fontsize = 30)# for Hierarchical with DM Permuatation')
#plt.savefig("pvals_v_N_H.pdf")
# +
plt.figure(figsize=(11,8))
plt.plot(N_rk, rev_k_mean, 'o-', lw = 3, ms = 10, label = 'K-Means: Width', alpha = 0.8)
plt.plot(N_k, k_mean, 'o-', lw = 3, ms = 10, label = 'K-Means: DM', alpha = 0.8)
plt.plot(N_rh,rev_hierar, 'o-', lw = 3, ms = 10, label = 'Hierarchical: Width', alpha = 0.8)
plt.plot(N_h, hierarc, 'o-', lw = 3, ms = 10, label = 'Hierarchical: DM', alpha = 0.8)
plt.plot(N_db, beta_DB_rev, 'o-', lw = 3, ms = 10, label ='DB Scan: Width', alpha = 0.8)
plt.plot(N_db, db_scan, 'o-', lw = 3, ms = 10, label = 'DB Scan: DM', alpha = 0.8)
plt.xlabel('$N$',fontsize = 30)
plt.ylabel(r'$\beta$',fontsize = 30)
#plt.title('K-means: Permute Width',fontsize = 30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
plt.hlines(0.9, -1, 120, lw = 3)
plt.vlines(45, 0, 1, lw = 3)
plt.vlines(90, 0, 1, lw = 3)
#plt.vlines(70, 0, 1, lw = 3)
plt.xlim(0,120)
plt.legend(numpoints = 1, loc = 4)
#plt.savefig("all_beta-20151008.png")
# +
plt.figure(figsize=(11,8))
plt.axhline(0.9,color = 'k', lw = 3)# -1, 130, lw = 3)
plt.vlines(45, 0, 1.1, lw = 3)
plt.vlines(90, 0, 1.1, lw = 3)
err_rk = np.sqrt(np.array(rev_k_mean)*(1.0- np.array(rev_k_mean))/1000.0)
plt.errorbar(N_rk, rev_k_mean, yerr = err_rk,fmt='b-', lw = 4, ms = 10, label = 'K-Means: Width', alpha = 0.7)
err_k = np.sqrt(np.array(k_mean)*(1.0-np.array(k_mean))/1000.0)
plt.errorbar(N_k, k_mean,yerr = err_k, fmt='r-', lw = 4, ms = 10, label = 'K-Means: DM', alpha = 0.7)
rev_hierar = np.array(rev_hierar)
plt.errorbar(N_rh, rev_hierar, yerr = np.sqrt(rev_hierar*(1.0-rev_hierar)/1000.0), fmt='m-', lw = 4, ms = 10, label = 'Hierarchical: Width', alpha = 0.7)
hierarc = np.array(hierarc)
plt.errorbar(N_h, hierarc,yerr = np.sqrt(hierarc*(1.0-hierarc)/1000.0), fmt='y-', lw = 4, ms = 7, label = 'Hierarchical: DM', alpha = 0.7)
beta_DB_rev = np.array(beta_DB_rev)
plt.errorbar(N_1, beta_DB_rev,yerr = np.sqrt(beta_DB_rev*(1.0-beta_DB_rev)/1000.0), fmt='c-', lw = 4, ms = 7, label ='DB Scan: Width', alpha = 0.77)
db_scan = np.array(db_scan)
plt.errorbar(N_db, db_scan,yerr = np.sqrt(db_scan*(1.0-db_scan)/1000.0), fmt='g-', lw = 4, ms = 7, label = 'DB Scan: DM', alpha = 0.7)
plt.xlabel('$N$',fontsize = 30)
plt.ylabel(r'$\beta$',fontsize = 30)
#plt.title('K-means: Permute Width',fontsize = 30)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
#plt.hlines(0.9, -1, 120, lw = 3)
#plt.vlines(60, 0, 1, lw = 3)
#plt.xlim(10,173)
plt.ylim(0,1.01)
plt.legend(numpoints = 1, loc = 4, fontsize = 18)
plt.xlim(0,120)
plt.savefig("/Users/karaponder/FRB/20151011_beta_ERR.pdf")
# -
x = np.linspace(0.01, 1, 1000)
y = 1.0/x**(1./2.)
plt.plot(x,y)
plt.ylim(0,10)
plt.title(r"$1/\sqrt{x}$ Distribution")
#plt.savefig('one_over_xsqu.pdf')
N = np.arange(0, 170, 10)
beta_GMM100_rev = np.loadtxt('rev_beta_GMM_100.txt')
beta_GMM100 = np.loadtxt('beta_GMM_100.txt')
plt.figure(figsize=(11,8))
plt.plot(N, beta_GMM100, 'o-', lw = 3, ms = 10, label = 'Permute DM')
plt.plot(N, beta_GMM100_rev, 'o-', lw = 3, ms = 10, label = 'Permute Width')
plt.xlabel('N',fontsize = 30)
plt.ylabel(r'$\beta$',fontsize = 30)
plt.xlim(0,160)
plt.ylim(0, 1)
plt.xticks(fontsize = 20)
plt.yticks(fontsize = 20)
#plt.vlines(37, 0, 1, lw = 3)
plt.title('GMM, 100 Permutations',fontsize = 30)
plt.legend(numpoints = 1, loc = 4)
plt.hlines(0.9, -1, 161, lw = -3, color = 'k')#, -1, 120, lw = 3)
plt.savefig("GMM2_100.png")
# + active=""
# plt.figure(figsize=(11,8))
#
# plt.axhline(0.9,color = 'k', lw = 3)# -1, 130, lw = 3)
# plt.vlines(45, 0, 1, lw = 3)
# plt.vlines(90, 0, 1, lw = 3)
# #print np.array(rev_k_mean)*1000
#
# err_rk = np.sqrt(np.array(rev_k_mean)*(1.0- np.array(rev_k_mean))/1000.0)
# plt.errorbar(N_rk, rev_k_mean, yerr = err_rk,fmt='o-', lw = 3, ms = 7, label = 'K-Means: Width', alpha = 0.8)
#
# ##err_rk_N = np.sqrt(np.array(rev_k_new)*(1.0- np.array(rev_k_new))/1000.0)
# ##plt.errorbar(Nk_new, rev_k_new, yerr = err_rk_N,fmt='o-', lw = 3, ms = 7, label = 'K-Means: Width; GP')
#
# err_k = np.sqrt(np.array(k_mean)*(1.0-np.array(k_mean))/1000.0)
# plt.errorbar(N_k, k_mean,yerr = err_k, fmt='o-', lw = 3, ms = 7, label = 'K-Means: DM', alpha = 0.8)
#
# #err_r_N = np.sqrt(np.array(k_new)*(1.0- np.array(k_new))/1000.0)
# #plt.errorbar(Nk_new, k_new, yerr = err_r_N,fmt='o-', lw = 3, ms = 7, label = 'K-Means: GP')
#
# rev_hierar = np.array(rev_hierar)
# plt.errorbar(N_rh, rev_hierar, yerr = np.sqrt(rev_hierar*(1.0-rev_hierar)/1000.0), fmt='o-', lw = 3, ms = 7, label = 'Hierarchical: Width', alpha = 0.8)
#
# #rev_hierar_N = np.loadtxt('rev_beta_Hier_20150818.txt')
# #plt.errorbar(N1, rev_hierar_N, yerr = np.sqrt(rev_hierar_N*(1.0-rev_hierar_N)/1000.0), fmt='o-', lw = 3, ms = 7, label = 'Hierarchical: Width; GP')
#
# hierarc = np.array(hierarc)
#
# #EXTRA_H = np.array([ 0.857 , 0.873 , 0.875, 0.896, 0.907])
# #hierarc = np.hstack((hierarc, EXTRA_H))
# #N_EXTRA_H = np.arange(120, 170, 10)
# #N_h = np.hstack((N_h, N_EXTRA_H))
# plt.errorbar(N_h, hierarc,yerr = np.sqrt(hierarc*(1.0-hierarc)/1000.0), fmt='o-', lw = 3, ms = 7, label = 'Hierarchical: DM', alpha = 0.8)
#
# #hierar_N = np.loadtxt('beta_Hier_20150818.txt')
# #plt.errorbar(N1, hierar_N, yerr = np.sqrt(hierar_N*(1.0-hierar_N)/1000.0), fmt='o-', lw = 3, ms = 7, label = 'Hierarchical: DM; GP')
#
# #plt.errorbar(N_EXTRA_H +10, EXTRA_H ,yerr = np.sqrt(EXTRA_H *(1.0-EXTRA_H )/1000.0), fmt='co-', lw = 3, ms = 7)
#
# beta_DB_rev = np.array(beta_DB_rev)
# plt.errorbar(N_1, beta_DB_rev,yerr = np.sqrt(beta_DB_rev*(1.0-beta_DB_rev)/1000.0), fmt='o-', lw = 3, ms = 7, label ='DB Scan: Width', alpha = 0.8)
#
# db_scan = np.array(db_scan)
# plt.errorbar(N_db, db_scan,yerr = np.sqrt(db_scan*(1.0-db_scan)/1000.0), fmt='o-', lw = 3, ms = 7, label = 'DB Scan: DM', alpha = 0.8)
#
# #DB_N = np.loadtxt('beta_DB_20150818.txt')
# #rev_DB_N = np.loadtxt('rev_beta_DB_20150818.txt')
#
# #plt.errorbar(N1, rev_DB_N,yerr = np.sqrt(rev_DB_N*(1.0-rev_DB_N)/1000.0), fmt='o-', lw = 3, ms = 7, label ='DB Scan: GP')
# #plt.errorbar(N1, DB_N,yerr = np.sqrt(DB_N*(1.0-DB_N)/1000.0), fmt='o-', lw = 3, ms = 7, label ='DB Scan: DM; GP')
#
# #db_regperm_newclus = np.array([0, 0.797, 0.845, 0.921, 0.919])
# #N_db_rp_nc = np.array([11,21,31,41, 51])
# #plt.errorbar(N_db_rp_nc, db_regperm_newclus,yerr = np.sqrt(db_regperm_newclus*(1.0-db_regperm_newclus)/1000.0), fmt='o-', lw = 3, ms = 7, label ='DB Scan: DM; New Clustering')
#
# #plt.plot(N, beta_GMM100, 'o-', color = 'k', lw = 3, ms = 10, label = 'GMM Permute DM')
# #plt.plot(N, beta_GMM100_rev, 'o--',color = 'k', lw = 3, ms = 10, label = 'GMM Permute Width')
#
# plt.xlabel('N',fontsize = 30)
# plt.ylabel(r'$\beta$',fontsize = 30)
# #plt.title('K-means: Permute Width',fontsize = 30)
# plt.xticks(fontsize = 20)
# plt.yticks(fontsize = 20)
# #plt.hlines(0.9, -1, 120, lw = 3)
# #plt.vlines(60, 0, 1, lw = 3)
# plt.xlim(10,173)
# plt.ylim(0,1.01)
# plt.legend(numpoints = 1, loc = 4, fontsize = 18)
#
# plt.xlim(0,120)
#
# #plt.axvline(80, color = 'k', lw = 3)
# #plt.axvline(160, color = 'k', lw = 2)
# #plt.axvline(110, color = 'k', lw = 2)
# #plt.savefig("/Users/karaponder/FRB/20150820_beta_ERR_DBscan_GP.pdf")
| Plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''torch'': conda)'
# language: python
# name: python3
# ---
from mapmaker import AltitudeMap, Map
p = AltitudeMap(
name='s7777', seed=7777,
land_level=0.5, sea_level=0.2, noise_level=0.5,
continent_number=3, slope=5, width_range=(0.25,0.75), height_range=(0.25,0.75),
perlin_cells=(20,10),
longtitude_range=200, latitude_range=80, resolution=1
)
print(p)
p.plot('all')
m = Map(p, name='多块大陆', seed='2222')
m.polish(times=3)
print(m)
m.plot()
| advanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import schedule
import time
def job_test():
print("I'm working...")
schedule.every(10).seconds.do(job_test)
# +
# does this work?
# -
| schedule.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **This script takes the best models and features from Phase 1 of the model selection process and undertakes a deeper dive in reviewing/selecting the optimal models. It also adapts wrangling steps to a Pipeline.**
#
# To-Do:
# - Use balanced sample of 50K observations DONE
# - Align features across scripts DONE for outcome variable - PENDING for features (scale, reformat, normalize)
# - Substitute dummies for label encoding, create a pipeline - PENDING
# - Add CV; randomize splits - PENDING
# - Review validation curve, precision versus recall
# - Robustness checks for county_type versus MSA binary
# - Holdout sets
# - Review learning curve
# - Grid search
# - Analyses on all years
# +
# %matplotlib inline
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns; sns.set()
# -
# **Import 2017 sample of 50,000 observations balanced for 1/0 Action Taken, removing two action_taken_name values.** Note import warning:"Columns (29,30,39,40) have mixed types. Specify dtype option on import or set low_memory=False."
filepath = os.path.abspath(os.path.join( "..", "fixtures", "hmda2017sample_balanced.csv"))
DATA = pd.read_csv(filepath, low_memory=False)
# **Drop features with missing data, locality information, and features resulting in model leakage**
# +
DATA = DATA.drop(DATA.columns[0], axis=1)
DATA = DATA.drop(['purchaser_type_name',
'preapproval_name',
'rate_spread',
'sequence_number',
'respondent_id',
'state_name',
'state_abbr',
'county_name',
'edit_status_name',
'denial_reason_name_3',
'denial_reason_name_2',
'denial_reason_name_1',
'co_applicant_race_name_5',
'co_applicant_race_name_4',
'co_applicant_race_name_3',
'co_applicant_race_name_2',
'census_tract_number',
'application_date_indicator',
'applicant_race_name_5',
'applicant_race_name_4',
'applicant_race_name_3',
'applicant_race_name_2',
'agency_name'],
axis=1)
# -
DATA['locality_type'] = None
# +
DATA['locality_type'] = None
cloc = DATA.columns.get_loc('locality_type')
i = 0
for x in DATA['msamd_name'].isna():
if x == True:
DATA.iat[i, cloc]= 0
else:
DATA.iat[i, cloc]= 1
i+=1
pd.crosstab(DATA['msamd_name'],DATA['locality_type'], margins=True, dropna=False) #dropna=False
# -
DATA.describe(include='all')
print("DATA dimensions: {}".format(DATA.shape))
# **Write the initial script using subset of features which are already int or float, plus the target**
#
# **NOTE: discard file closed, call 'application approved but not accepted" a 1 or discard, discard 'application withdrawn by applicant'. Concern about overfitting if we leave too much stuff in.**
DATA['action_taken'] = DATA.action_taken_name.apply(lambda x: 1 if x in ['Loan purchased by the institution', 'Loan originated'] else 0)
pd.crosstab(DATA['action_taken_name'],DATA['action_taken'], margins=True)
# **ACTION: look at imputing income using hud household median income rather than mean**
DATA_targ_numeric = DATA[['action_taken',
'tract_to_msamd_income',
'population',
'minority_population',
'number_of_owner_occupied_units',
'number_of_1_to_4_family_units',
'loan_amount_000s',
'hud_median_family_income',
'applicant_income_000s', 'locality_type'
]]
#resolve missing values in applicant_income_000s
DATA_targ_numeric.fillna(DATA_targ_numeric.mean(), inplace=True)
DATA_targ_numeric.info()
DATA_basefile = DATA_targ_numeric
# **Use one-hot encoding via Pandas, concatenate to the rest of the data frame.**
#
# Reference link:
# https://stackoverflow.com/questions/37292872/how-can-i-one-hot-encode-in-python
DATA = DATA.drop(['action_taken_name', 'msamd_name'], axis=1)
DATA.columns
non_categorical_features = ['action_taken',
'tract_to_msamd_income',
'population',
'minority_population',
'number_of_owner_occupied_units',
'number_of_1_to_4_family_units',
'loan_amount_000s',
'hud_median_family_income',
'applicant_income_000s',
'locality_type'
]
# +
for categorical_feature in list(DATA.columns):
if categorical_feature not in non_categorical_features:
DATA[categorical_feature] = DATA[categorical_feature].astype('category')
dummies = pd.get_dummies(DATA[categorical_feature], prefix=categorical_feature)
DATA_basefile = pd.concat([DATA_basefile, dummies], axis=1)
DATA_basefile.info(verbose=True)
# -
tofilepath = os.path.abspath(os.path.join( "..", "fixtures", "hmda2017sample_alltest_state_localitytest.csv"))
DATA_basefile.to_csv(tofilepath, index=False)
# +
# Determine the shape of the data
print("{} instances with {} features\n".format(*DATA_basefile.shape))
# Determine the frequency of each class
print(pd.crosstab(index=DATA['action_taken'], columns="count"))
# -
# ## Classification
#
# +
from sklearn import metrics
from sklearn.naive_bayes import GaussianNB, BernoulliNB, MultinomialNB
from sklearn.linear_model import LogisticRegressionCV, LogisticRegression, SGDClassifier
from sklearn.svm import LinearSVC, NuSVC, SVC
from sklearn import tree
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import BaggingClassifier, ExtraTreesClassifier, RandomForestClassifier
from yellowbrick.classifier import ClassificationReport
# -
X = DATA_basefile[DATA_basefile.columns[1:]]
y = DATA_basefile['action_taken']
def score_model(X, y, model, **kwargs):
"""
Test various estimators.
"""
#NOTE: for capstone add X_test, X_train, Y_test, Y_train for capstone code.
#Bake into model to see if it does cross validation, if not there do CV.
scores = {'precision':[], 'recall':[], 'accuracy':[], 'f1':[]}
# Instantiate the classification model and visualizer
model.fit(X, y, **kwargs)
expected = y
predicted = model.predict(X)
# Append our scores to the tracker
scores['precision'].append(metrics.precision_score(expected, predicted, average="binary"))
scores['recall'].append(metrics.recall_score(expected, predicted, average="binary"))
scores['accuracy'].append(metrics.accuracy_score(expected, predicted))
scores['f1'].append(metrics.f1_score(expected, predicted, average="binary"))
# Compute and return F1 (harmonic mean of precision and recall), Precision, Recall, Accuracy
print("{}".format(model.__class__.__name__))
print("Validation scores are as follows:\n")
print(pd.DataFrame(scores).mean())
# Try them all!
models = [
GaussianNB(),
MultinomialNB(),
BernoulliNB(),
tree.DecisionTreeClassifier(),
LinearDiscriminantAnalysis(),
LogisticRegression(solver='lbfgs', max_iter=6000),
LogisticRegressionCV(cv=3, max_iter=6000),
BaggingClassifier(),
ExtraTreesClassifier(n_estimators=100),
RandomForestClassifier(n_estimators=100),
LinearSVC(max_iter=6000)
]
for model in models:
score_model(X, y, model)
# +
def visualize_model(X, y, estimator):
"""
Test various estimators.
"""
# Instantiate the classification model and visualizer
visualizer = ClassificationReport(
model, classes=[1,0],
cmap="Blues", size=(600, 360)
)
visualizer.fit(X, y)
visualizer.score(X, y)
visualizer.poof()
for model in models:
visualize_model(X, y, model)
# +
from yellowbrick.features import FeatureImportances
model = RandomForestClassifier(n_estimators=10)
viz = FeatureImportances(model, size=(1080, 720))
viz.fit(X, y)
# Note: the FeatureImportances visualizer is a model visualizer,
# not a feature visualizer, so it doesn't have a transform method!
viz.poof()
# +
from yellowbrick.features import Rank2D
# Instantiate the visualizer with the Pearson ranking algorithm
visualizer = Rank2D(algorithm='pearson', size=(1080, 720))
visualizer.fit(X, y)
visualizer.transform(X)
visualizer.poof()
| notebooks/01m_Phase_2_ML_FullFeatureSet-BestInitialScaling_ak.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Crossword Puzzle
#
# <br>
#
# 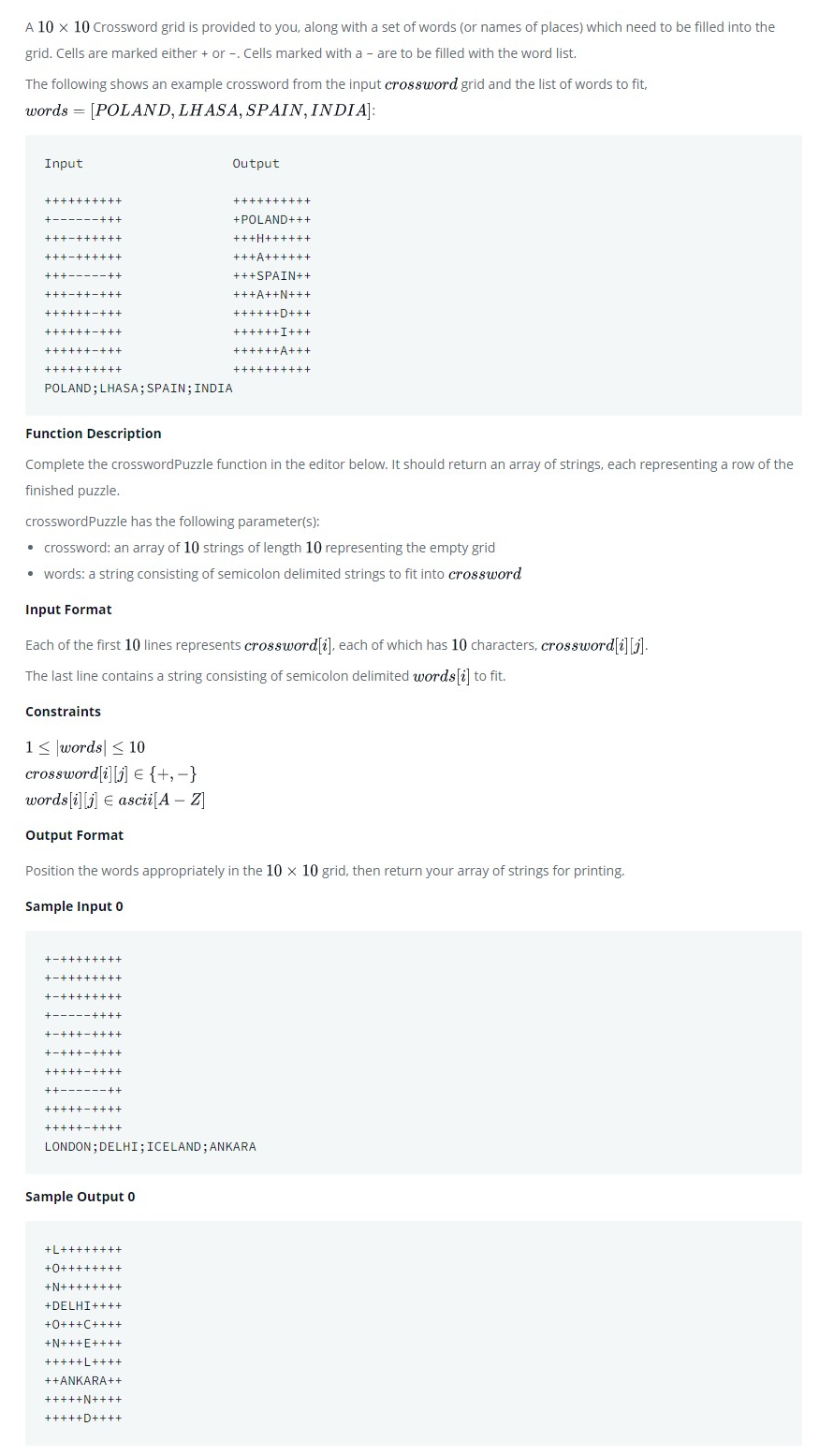
# +
# #!/bin/python3
import math
import os
import random
import re
import sys
# Complete the crosswordPuzzle function below.
from copy import deepcopy
def checkWords(crossword, k, i, j, direct):
if k == len(words):
return answer.append(crossword)
if words[k] == ';':
return findEmpty(crossword, k + 1)
if not (0 <= i < 10 > j >= 0) or crossword[i][j] == "+":
return
if crossword[i][j] != '-' and crossword[i][j] != words[k]:
return
copy_crossword = deepcopy(crossword)
copy_crossword[i][j] = words[k]
if direct == 1:
checkWords(copy_crossword, k + 1, i, j + 1, 1)
elif direct == 2:
checkWords(copy_crossword, k + 1, i + 1, j, 2)
def findEmpty(crossword, k):
for i in range(10):
for j in range(10):
if crossword[i][j] != '+':
checkWords(crossword, k, i, j, 1)
checkWords(crossword, k, i, j, 2)
def crosswordPuzzle(crossword, words):
global answer
crossword = [list(s) for s in crossword]
answer = []
findEmpty(crossword, 0)
return ["".join(s) for s in answer[0]]
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
crossword = []
for _ in range(10):
crossword_item = input()
crossword.append(crossword_item)
words = input()
result = crosswordPuzzle(crossword, words)
fptr.write('\n'.join(result))
fptr.write('\n')
fptr.close()
| Interview Preparation Kit/13. Recursion and Backtracking/Crossword Puzzle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
# + active=""
# div.output_prompt { visibility: hidden; }
# *{ font-family: "Palatino Linotype", "Book Antiqua", Palatino, serif !important }
# .input,#header { display: none; }
# -
# # Modified salinity water flooding
# The simplest model for the modify salinity water flooding assumes that the relative permeability of water and oil (mostly oil) is altered due to the change of the salinity of the injected water. Here, I follow the procedure described by Pope (SPE-7660) to find the fractional flow solution for the secondary and tertiary modified salinity water flooding. I will compare the analytical solution with a numerical solution (finite volume, upwind, fine mesh). Then, I will present a special case for the tertiary water flooding where the Pope's approach seems to give erroneous results. I then try to present an alternative solution procedure.
# ## Secondary modified salinity water flood
# First, we inject the low salinity water to a reservoir that is not water flooded. The injected brine pushes the formation brine, and the formation brine pushes oil. Moreover, the injected brine mobilizes more oil, since it changes the transport properties of oil and water in the reservoir, i.e., has a different sets of relative permeability curves and fractional flow curve. Let's define the relative permeability curves, and visualize them.
include("FractionalFlow.jl")
using PyPlot
FF = FractionalFlow
fluids_hs = FF.oil_water_fluids(mu_water=1e-3, mu_oil=2e-3)
fluids_ls = FF.oil_water_fluids(mu_water=1e-3, mu_oil=2e-3)
rel_perms_hs = FF.oil_water_rel_perms(krw0=0.4, kro0=0.9,
swc=0.15, sor=0.25, nw=2.0, no = 2.0)
rel_perms_ls = FF.oil_water_rel_perms(krw0=0.3, kro0=0.95,
swc=0.15, sor=0.15, nw=2.0, no = 2.0)
core_flood = FF.core_flooding(u_inj=1.15e-5, pv_inject=5.0, p_back=1e5, sw_init=0.2, sw_inj=1.0, rel_perms=rel_perms_hs)
core_props = FF.core_properties()
ls_res_sec = FF.low_sal_water_flood(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood);
FF.print_relperm(rel_perms_hs, title="high salinity (formation brine)")
FF.print_relperm(rel_perms_ls, title="low salinity (injected brine)")
FF.print_core_properties(core_props, title = "core properties")
FF.print_fluids(fluids_hs, title = "viscosity")
FF.print_core_flood(core_flood, title="core flooding condition")
figure()
FF.visualize(rel_perms_hs, label = "HS")
FF.visualize(rel_perms_ls, label = "LS")
figure()
FF.visualize(rel_perms_hs, fluids_hs, label="HS")
FF.visualize(rel_perms_ls, fluids_ls, label="LS")
tight_layout()
figure()
FF.visualize_solution(ls_res_sec)
figure()
FF.visualize_profiles(ls_res_sec)
title("Secondary low-sal");
# Here, I compare the above solution with the numerical solution of the same problem:
t_sec, pv_num, rec_fact, xt_num, sw_num, c_old, c_out_sal =
FF.forced_imb_implicit_upwind(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood);
plot(xt_num/(core_props.length)/pv_num[end], sw_num, label = "Sw-numerical")
plot(xt_num/(core_props.length)/pv_num[end], 1-c_old, label = "tracer-numerical")
plot(ls_res_sec.saturation_profile_xt[:,1], ls_res_sec.saturation_profile_xt[:,2], label = "Sw-analytical")
plot(ls_res_sec.tracer_profile_xt[:,1], ls_res_sec.tracer_profile_xt[:,2], label = "tracer-analytical")
xlabel("x/t [-]")
ylabel("Water saturation [-]")
title("Secondary low sal water flood")
legend();
# ## Tertiary low-salinity water flooding
# We assume that the reservoir is flooded with the formation brine (i.e, high salinity) and has reached the residual oil saturation. It means that the initial saturation of the reservoir is 1-$S_{or}^{HS}$. For this system, the solution looks like the following:
# +
core_flood = FF.core_flooding(u_inj=1.15e-5, pv_inject=5.0, p_back=1e5,
sw_init=1-rel_perms_hs.sor, sw_inj=1.0, rel_perms=rel_perms_hs)
ls_res_ter = FF.low_sal_water_flood(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood)
# figure()
# FF.visualize(rel_perms_hs, label = "HS")
# FF.visualize(rel_perms_ls, label = "LS")
# figure()
# FF.visualize(rel_perms_hs, fluids_hs, label="HS")
# FF.visualize(rel_perms_ls, fluids_ls, label="LS")
# tight_layout()
figure()
FF.visualize_solution(ls_res_ter)
figure()
FF.visualize_profiles(ls_res_ter)
title("Tertiary low-sal");
# -
t_sec, pv_num, rec_fact, xt_num, sw_num, c_old, c_out_sal =
FF.forced_imb_implicit_upwind(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood);
plot(xt_num/(core_props.length)/pv_num[end], sw_num, label = "Sw-numerical")
plot(xt_num/(core_props.length)/pv_num[end], 1-c_old, label = "tracer-numerical")
plot(ls_res_ter.saturation_profile_xt[:,1], ls_res_ter.saturation_profile_xt[:,2], label = "Sw-analytical")
plot(ls_res_ter.tracer_profile_xt[:,1], ls_res_ter.tracer_profile_xt[:,2], label = "tracer-analytical")
xlabel("x/t [-]")
ylabel("Water saturation [-]")
title("Tertiary low sal water flood")
legend();
# Again, we can see that the solution matches reasonably with the numerical results.
# ## A special case
# One special case happens when the high salinity residual oil saturation is very close to the low salinity residual oil saturation. In this case, the line that connects the intersection of the high salinity fractional flow curve and the tangent line from the origin to the low salinity fractional curve to the initial reservoir saturation (i.e., 1-$S_{w,or}^{HS}$, cuts through the low salinity fractional flow curve, as shown below. In this case the above procedure for finding the analytical solution procedure does not work:
rel_perms_hs = FF.oil_water_rel_perms(krw0=0.4, kro0=0.9,
swc=0.15, sor=0.20, nw=2.0, no = 2.0)
core_flood = FF.core_flooding(u_inj=1.15e-5, pv_inject=5.0, p_back=1e5,
sw_init=1-rel_perms_hs.sor, sw_inj=1.0, rel_perms=rel_perms_hs)
ls_res_ter = FF.low_sal_water_flood(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood)
FF.print_relperm(rel_perms_hs, title = "high salinity rel perm")
figure()
FF.visualize(rel_perms_hs, label = "HS")
FF.visualize(rel_perms_ls, label = "LS")
figure()
FF.visualize(rel_perms_hs, fluids_hs, label="HS")
FF.visualize(rel_perms_ls, fluids_ls, label="LS")
tight_layout()
figure()
FF.visualize_solution(ls_res_ter)
figure()
FF.visualize_profiles(ls_res_ter)
title("Secondary low-sal");
t_sec, pv_num, rec_fact, xt_num, sw_num, c_old, c_out_sal =
FF.forced_imb_implicit_upwind(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood);
plot(xt_num/(core_props.length)/pv_num[end], sw_num, label = "Sw-numerical")
plot(xt_num/(core_props.length)/pv_num[end], 1-c_old, label = "tracer-numerical")
plot(ls_res_ter.saturation_profile_xt[:,1], ls_res_ter.saturation_profile_xt[:,2], label = "Sw-analytical")
plot(ls_res_ter.tracer_profile_xt[:,1], ls_res_ter.tracer_profile_xt[:,2], label = "tracer-analytical")
xlabel("x/t [-]")
ylabel("Water saturation [-]")
title("Tertiary low sal water flood")
legend();
# ## The correct solution
# To construct the correct saturation profile, I assumed that we should follow the same procedure, but interpret the results in the right way. It means that the low salinity shock front saturation is represented by the intersection of the red line (see the solution procedure above) and the low salinity fractional flow curve (orange curve). This interpretation produces the following results:
# +
ls_res_ter = FF.single_ion_adsorption_water_flood_single_shock(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood, 0.0)
figure()
FF.visualize_solution(ls_res_ter)
figure()
FF.visualize_profiles(ls_res_ter)
title("Secondary low-sal");
# -
t_sec, pv_num, rec_fact, xt_num, sw_num, c_old, c_out_sal =
FF.forced_imb_implicit_upwind(core_props, fluids_ls, fluids_hs, rel_perms_hs,
rel_perms_ls, core_flood);
plot(xt_num/(core_props.length)/pv_num[end], sw_num, label = "Sw-numerical")
plot(xt_num/(core_props.length)/pv_num[end], 1-c_old, label = "tracer-numerical")
plot(ls_res_ter.saturation_profile_xt[:,1], ls_res_ter.saturation_profile_xt[:,2], label = "Sw-analytical")
plot(ls_res_ter.tracer_profile_xt[:,1], ls_res_ter.tracer_profile_xt[:,2], label = "tracer-analytical")
xlabel("x/t [-]")
ylabel("Water saturation [-]")
title("Tertiary low sal water flood")
legend();
# The above solution assumes that there is no adsorption of ions on the rock surface. However, it is quite easy to include the adsorption in the equations and in the solution procedure.
# If the above solution is correct, it explains several issues with the observations in the lab. for instance, sometimes the extra oil recovery observed before the breakthrough of the injected low salinity water. Sometimes, the tertiary recovery is extremely slow and needs the injection of many pore volumes of water. It is explained by the above fractional flow solution very well. I can probably think of other issues that I have observed in the lab.
| analytical/.ipynb_checkpoints/formulating_the_question_low_sal_frac_flow-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pfp_2021]
# language: python
# name: conda-env-pfp_2021-py
# ---
# # Experimentation I - Introduction to PsychoPy I
#
# [<NAME> (he/him)](https://peerherholz.github.io/)
# Habilitation candidate - [Fiebach Lab](http://www.fiebachlab.org/), [Neurocognitive Psychology](https://www.psychologie.uni-frankfurt.de/49868684/Abteilungen) at [Goethe-University Frankfurt](https://www.goethe-university-frankfurt.de/en?locale=en)
# Research affiliate - [NeuroDataScience lab](https://neurodatascience.github.io/) at [MNI](https://www.mcgill.ca/neuro/)/[McGill](https://www.mcgill.ca/)
# Member - [BIDS](https://bids-specification.readthedocs.io/en/stable/), [ReproNim](https://www.repronim.org/), [Brainhack](https://brainhack.org/), [Neuromod](https://www.cneuromod.ca/), [OHBM SEA-SIG](https://ohbm-environment.org/), [UNIQUE](https://sites.google.com/view/unique-neuro-ai)
#
# <img align="left" src="https://raw.githubusercontent.com/G0RELLA/gorella_mwn/master/lecture/static/Twitter%20social%20icons%20-%20circle%20-%20blue.png" alt="logo" title="Twitter" width="32" height="20" /> <img align="left" src="https://raw.githubusercontent.com/G0RELLA/gorella_mwn/master/lecture/static/GitHub-Mark-120px-plus.png" alt="logo" title="Github" width="30" height="20" /> @peerherholz
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Objectives 📍
#
# * get to know the [PsychoPy](https://www.psychopy.org/) library
# * learn basic and efficient usage of its `module`s and `function`s to create _simple_ experiments
# + [markdown] slideshow={"slide_type": "slide"}
# ## Important note
#
# The main content of this section will be presented via a mixture of [slides]() and [VScode]() which is further outlined in the [respective section of the course website]() and [slides](). As noted there, `jupyter notebooks` aren't the best way to work on and run experiments using `PsychoPy`, instead we need to switch to `IDE`s for this part of the course. Specifically, we will use `VScode` for this.
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/ipynb_IDE.png" alt="logo" title="jupyter" width="800" height="300" />
#
#
#
# This `notebook` is thus not intended as the main resource and you shouldn't try to test/run the experiment via the here included `code cells`. Rather, this is meant to be an add-on resource that presents some of the content and especially `code` in a more condensed form. We hope it will be useful/helpful for you.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Outline
#
# Within this notebook we will go through the basic and required steps to create a new experiment using `PsychoPy`, including:
#
# 1. Prerequisites
# 1.1 Computing environment
# 1.2 Folders & Files
# 2. `PsychoPy` basics
# 2.1 The general idea
# 2.2 Input via `dialog boxes`
# 2.3 Presenting instructions
# 3. `PsychoPy`'s working principles
# 3.1 `draw`ing & `flip`ping
# 3.2 `trial`s
# 4. Input/output
# 4.1 collecting responses
# 4.2 saving data
# 5. A very simple experiment
# + [markdown] slideshow={"slide_type": "slide"}
# ## Prerequisites
#
# Starting new experiments in `PsychoPy` follows the same guidelines as starting new projects in general. This includes the following:
#
# - create and store everything in a dedicated place on your machine
#
# - create and use a dedicated computing environment
#
# - document everything or at least as much as possible
#
# - test and save things in very short intervals, basically after every change
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/experiment_prereq.png" alt="logo" title="jupyter" width="800" height="200" />
#
#
#
# ### Computing environments
#
# As addressed during the first weeks of the course, `computing environments` are essential when programming, not only in `Python`. This refers to reproducibility, transfer/sharing of code and many other factors. Lucky for us `Python` makes it easy to `create` and `manage` `computing environments`, for example using [conda]().
#
# We can thus also use it to create a new `computing environment` specifically dedicated to creating and running a new experiment using `PsychoPy`. Here we will name it `psychopy` and include/install a few dependencies we already know:
# + language="bash"
#
# conda create -n psychopy psychopy
#
# conda activate psychopy
#
# pip install jedi psychtoolbox pygame pyo pyobjc
# python-vlc ujson
#
# -
# With these few steps we have our (initial) `computing environment` ready to go!
#
# Let's continue with creating `folders` and `files` we need.
#
# ### Folders & files
#
# As mentioned above, it's a good idea to keep things handy and organized. Obviously, this also holds true for running experiments using `PsychoPy`. While there are several ways we could do this, at the minimum we need a dedicated `folder` or `directory` somewhere on our machine within which we will store all corresponding information and `files`. Creating a new `directory` as no biggie using `bash`, we can simply use `mkdir` and specify the wanted `path` and `name`. For the sake of simplicity, let's put everything in a folder called `psychopy_experiment` on our `Desktop`s.
# + language="bash"
# mkdir /path/to/your/Desktop/psychopy_environment
# -
# Now we will change our `current working directory` to this new folder
# + language="bash"
# cd /path/to/your/Desktop/psychopy_environment
# -
# and once there, create a new `python script`, i.e. an empty `python file`. For this we can use `bash`s `touch` `function` followed by the desired `filename`. Keeping things simple again, we will name it `experiment.py` (notice the file extension `.py`).
# + language="bash"
# touch experiment.py
# -
# Within this, currently empty, `python file`/`script` we will put our `python code` needed to `run` the `experiment`.
#
# ### VScode setup
#
# Once more: please remember that we're switching to an `IDE` for this part of the course, specifically `VScode`, as `jupyter notebooks` aren't the most feasible way to implement/test/run `experiment`s via `PsychoPy`. Therefore, please open `VScode` and within it open the folder we just created (`File` -> `Open Folder`). Next, click on the `experiment.py` file which should open in the `editor` window and finally also open a `terminal` via `Terminal` -> `New Terminal` and activate the `computing environment` we created above. With that your setup is ready for our `PsychoPy` adventure and should look roughly like below:
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/vscode_setup.png" alt="logo" title="jupyter" width="800" height="400" />
# ## `PsychoPy` basics
#
# It's already time to talk about `PsychoPy`, one of the `python libraries` intended to run `experiments` and acquire `data`. For more information regarding different software and options, their advantages and drawbacks, please consult the [slides]().
#
# <img align="center" src="https://www.psychopy.org/_static/psychopyLogoOnlineStrap_h480.png" alt="logo" title="jupyter" width="600" height="300" />
#
# Make sure to check the [`PsychoPy` website](https://www.psychopy.org/), [documentation](https://psychopy.org/documentation.html) and [forum](https://discourse.psychopy.org/).
#
#
#
# ### What is `PsychoPy`
#
# - Psychology software in `Python`, i.e. a `Python library`, i.e. completely written in `Python`
# - 2002-2003: <NAME> began work on this for his own lab (visual neuroscience)
# - 2003-2017: a purely volunteer-driven, evenings and weekends project
# - 2017-now: still open source and free to install but with professional support
#
#
# ### Idea/goals of `PsychoPy`
#
# - allow scientists to run as wide a range of experiments as possible, easily and via standard computer hardware
# - precise enough for psychophysics
# - intuitive enough for (undergraduate) psychology (no offence)
# - flexible enough for everything else
# - capable of running studies in the lab or online
#
#
# ### Things to check/evaluate
#
# - computer hardware settings & interfaces
# - rapid software development
# - always check version
# - set `version` in `experiment`
# - use `environments` for `experiments`
# - don’t change version in running `experiments`
#
# First things first: do you have a working `PsychoPy` installation?
#
# We can simply check that via starting an `ipython` session from our `terminal`:
# + language="bash"
#
# ipython
# -
# and from within there then try `import`ing `PsychoPy`:
import psychopy
# ### The general idea
#
# If you don't get an `import error`, at least the basic `installation` should be ok!
#
# Cool, we are now ready to actually do some `coding`! As said before we will do that in our `experiment.py` `script`. While the transition from `jupyter notebooks` to `python scripts` might seem harsh at first, it’s actually straight-forward: the steps we conducted/`commands` we `run` in an incremental fashion will also be indicated/`run` in an incremental fashion here, just within one `python script` line-by-line.
#
# So, what's the first thing we usually do? That's right: `import`ing `modules` and `function`s we need. Comparably to `jupyter notebook`, we will do that at the beginning of our `script`. Please note, that we will go through a realistic example of `coding` `experiments` in `python` and thus might not all `modules`/`functions` we will actually need when we start. Thus, we will add them at the beginning as we go along!
#
# However, we actually haven't checked out what `modules`/`functions` `PsychoPy` has. Let's do that first.
#
#
# - [psychopy.core](https://psychopy.org/api/core.html): various basic functions, including timing & experiment termination
# - [psychopy.gui](https://psychopy.org/api/gui.html): various basic functions, including timing & experiment termination
# - [psychopy.event](https://psychopy.org/api/event.html): handling of keyboard/mouse/other input from user
# - psychopy.[visual](https://psychopy.org/api/visual/index.html)/[sound](https://psychopy.org/api/sound/index.html): presentation of stimuli of various types (e.g. images, sounds, etc.)
# - [psychopy.data](https://psychopy.org/api/data.html): handling of condition parameters, response registration, trial order, etc.
# - many more …: we unfortunately can’t check out due to time constraints
#
# Nice, looks like a decent collection of useful/feasible `modules`/`functions`. The question now is: which ones do we need to implement and `run` our `experiment`? Wait a minute...we genuinely didn't even talk about the `experiment` yet...
#
# Let’s assume we have obtained some data regarding favorite `movies`, `snacks` and `animals` from a group of fantastic students (obviously talking about you rockstars!) and now want to test how each respectively provided item is `perceived`/`evaluated` by our entire sample: how would we do that?
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/experiment_outline.png" alt="logo" title="jupyter" width="800" height="300" />
#
# As you can see, we need all of them!
# ### Input via dialog boxes
#
# Many `experiments` start with a `GUI dialog box` that allow users/participant to input certain information, for example `participant id`, `session`, `group`, `data storage path`, etc. . We can implement this crucial aspect via the [psychopy.gui module](https://psychopy.org/api/gui.html). Initially, we need to `import` it and thus need to start a new section in our `python script` and after that, we can define the `GUI dialog box` we want to create via a `dictionary` with respective `key-value pairs`.
#
# Please note: at this we will start populating our `experiment.py` `script`. Thus, you should copy paste the respective content of the `code cells` into your `experiment.py` `script` you opened in `VScode`. As we will go step-by-step, the `code`/`script` will get longer and longer.
# +
#===============
# Import modules
#===============
from psychopy import gui, core # import psychopy modules/functions
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
# -
# That’s actually all we need to test our `GUI dialog box`. In order to do that, we need to `run`/`execute` our `python script` called `experiment.py`. This is achieved via typing `python experiment.py` in the `VScode terminal` and pressing `enter` this will `run`/`execute` the `python script` `experiment.py` via the `python` installed in our `conda environment` called `psychopy`. Again, please don't `run`/`execute` `code` in this `jupyter notebook`!
python experiment.py
# If everything works/is set correctly you should see a `GUI dialog box` appearing on your screen asking for the information we indicated in our `experiment.py` `python script` (chances are the layout on your end looks a bit different than mine, that’s no biggie). After entering all requested information and clicking `ok` the `GUI dialog box` should close and no errors should appear.
#
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/gui_example.png" alt="logo" title="jupyter" width="350" height="200" />
# The next aspect we should take care of is the `data handling`, i.e. defining a `data filename` and `path` where it should be saved. We can make use of the `exp_info dictionary` right away and extract important information from there, for example, the `experiment` name and `participant ID`. Additionally, we will obtain the `date` and `time` via the `psychopy.core module`. We will also create a unique `filename` for the resulting `data` and check if the set `data path` works out via the `os` `module`.
# +
#===============
# Import modules
#===============
from psychopy import gui, core # import psychopy modules/functions
import os # import os module
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
# -
# ### Presenting instructions
#
# After having set some crucial backbones of our `experiment`, it’s time to actually start it. Quite often, `experiments` start with several messages of `instructions` that explain the `experiment` to the `participant`. Thus, we will add a few here as well, starting with a common `“welcome” text message`. To display things in general but also text, the [psychopy.visual module](https://psychopy.org/api/visual/index.html) is the way to go. What we need to do now is define a general `experiment window` to utilize during the entire `experiment` and a `text` to be displayed on it.
# +
#===============
# Import modules
#===============
from psychopy import gui, core, visual # import psychopy modules/functions
import os # import os module
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
#===============================
# Creation of window and messages
#===============================
# Open a window
win = visual.Window(size=(800,600), color='gray', units='pix', fullscr=False) # set size, background color, etc. of window
# Define experiment start text
welcome_message = visual.TextStim(win,
text="Welcome to the experiment. Please press the spacebar to continue.",
color='black', height=40)
# -
# ## PsychoPy working principles
#
# Here, we came across one of `PsychoPy`’s core working principles: we need a `general experiment window`, i.e. a place we can display/present something on. You can define a variety of different `windows` based on different `screens`/`monitors` which should however be adapted to the `setup` and `experiment` at hand (e.g. `size`, `background color`, etc.). Basically all `experiments` you will set up will require to define a `general experiment window` as without it no `visual stimuli` (e.g. `images`, `text`, `movies`, etc.) can be displayed/presented or how `PsychoPy` would say it: `drawn`
#
# Speaking of which: this is the next core working principle we are going to see and explore is the difference between `draw`ing something and showing it.
#
# ### `Draw`ing & `flip`ping
#
# In `PsychoPy` (and many other comparable software) there’s a big difference between `draw`ing and showing something. While we need to `draw` something on/in a `window` that alone won’t actually show it. This is because `PsychoPy` internally uses `“two screens”` one `background` or `buffer` `screen` which is not seen (yet) and one `front screen` which is (currently) seen. When you `draw` something it’s always going to be `draw`n on the `background`/`buffer` `screen`, thus “invisible” and you need to `flip` it to the `front screen` to be “visible”.
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/draw_flip.png" alt="logo" title="jupyter" width="800" height="400" />
#
# Why does `PsychoPy` (and other comparable software) work like that? The idea/aim is always the same: increase performance and minimize delays (as addressed in the [slides]()). `Draw`ing something might take a long time, depending on the stimulus at hand, but `flip`ping something already drawn from the `buffer` to the `front screen` is fast(er). It can thus ensure better and more precise timing. This can work comparably for `images`, `sounds`, `movies`, etc. where things are set/`draw`n/pre-loaded and presented exactly when needed.
# +
#===============
# Import modules
#===============
from psychopy import gui, core, visual # import psychopy modules/functions
import os # import os module
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
#===============================
# Creation of window and messages
#===============================
# Open a window
win = visual.Window(size=(800,600), color='gray', units='pix', fullscr=False) # set size, background color, etc. of window
# Define experiment start text
welcome_message = visual.TextStim(win,
text="Welcome to the experiment. Please press the spacebar to continue.",
color='black', height=40)
#=====================
# Start the experiment
#=====================
# display welcome message
welcome_message.draw() # draw welcome message to buffer screen
win.flip() # flip it to the front screen
# -
# Let’s give it a try via `python experiment.py`. If everything works/is set correctly you should see the `GUI dialog box` again but this time after clicking `OK`, the `text` we defined as a welcome message should appear next.
#
python experiment.py
# However, it only appears very briefly and in contrast to our `GUI dialog box` we don’t need to press anything
# to advance. This is because we didn’t tell `PsychoPy` that we want to `wait` for a distinct `key press` before we advance further, we need the [psychopy.event module](https://psychopy.org/api/event.html). Through its `.waitKeys()` `function` we can define that nothing should happen/we shouldn't advancing unless a certain `key` is pressed. While we are at it, let’s add a few more messages to our `experiment`. One will be presented right after the welcome message and explain very generally what will happen in the `experiment`. Another one will be presented at the end of the experiment and display a general “that’s it, thanks for taking part” message. The syntax for creating, `draw`ing and presenting these message is identical to the one we just explored, we only need to change the `text`.
# +
#===============
# Import modules
#===============
from psychopy import gui, core, visual, event # import psychopy modules/functions
import os # import os module
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
#===============================
# Creation of window and messages
#===============================
# Open a window
win = visual.Window(size=(800,600), color='gray', units='pix', fullscr=False) # set size, background color, etc. of window
# Define experiment start text
welcome_message = visual.TextStim(win,
text="Welcome to the experiment. Please press the spacebar to continue.",
color='black', height=40)
# Define trial start text
start_message = visual.TextStim(win,
text="In this experiment you will rate different movies, snacks and animals on a scale from 1 to 7. Please press the spacebar to start.",
color='black', height=40)
# Define experiment end text
end_message = visual.TextStim(win,
text="You have reached the end of the experiment, thanks for participating.",
color='black', height=40)
#=====================
# Start the experiment
#=====================
# display welcome message
welcome_message.draw() # draw welcome message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# display start message
start_message.draw() # draw start message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
#======================
# End of the experiment
#======================
# Display end message
end_message.draw() # draw end message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# -
# Let’s give it a try via `python experiment.py`. If everything works/is set correctly you should see the `GUI dialog box` and after clicking `OK`, the `text` we defined as a welcome message should appear next, followed by the start message and finally the end message. In all cases, the `experiment` should only advance if you press `spacebar` or quit when you press `escape`.
python experiment.py
# Having this rough frame of our `experiment`, it’s actually time to add the `experiment` itself: the `evaluation` of our `movies`, `snacks` and `animals`.
#
# ### `trials`
#
# Quick reminder: our `experiment` should collect responses from `participants` regarding our `list`s of `movies`, `snacks` and `animals`, specifically their respective `rating`. Thus we need to add/implement two aspects in our `experiment`: the **presentation of stimuli** and their **rating**/
#
# Starting with the `presentation of stimuli`, we will keep it simple for now and present them via `text`. However, any ideas how we could begin working on this? That’s right: we need to define `list`s with our stimuli!
#
# ```
# movies = [‘Forrest Gump’, ‘Interstellar’, ‘Love Actually’, …]
# ```
#
#
# Within that process, we can already think about the next step: quite often `experiments` `shuffle` the order of `stimuli` across participants to avoid sequence/order effects. We will do the same and implement that via the [numpy.random module](https://numpy.org/doc/stable/reference/random/index.html), specifically its [.shuffle()](https://numpy.org/doc/stable/reference/random/generated/numpy.random.shuffle.html) `function` which will allow us to randomly `shuffle` our previously created `list`.
#
# ```
# rnd.shuffle(movies)
# movies
# [‘Interstellar’, ‘Love Actually’, ‘Forrest Gump’, …]
# ```
#
#
# After that we need to bring our `shuffle`d `stimuli list` into the format required by `PsychoPy`. Specifically, this refers to the definition of `experiment trials`, i.e. `trials` that will be presented during the `experiment`, including their properties (e.g. `content`, `order`, `repetition`, etc.). In `PsychoPy` this is achieved via the [data.TrialHandler()](https://psychopy.org/api/data.html#psychopy.data.TrialHandler) `function` for which we need to convert our `shuffle`d `stimuli list` into a `list` of `dictionaries` of the form `“stimulus”: value`.
#
# ```
# stim_order = []
#
# for stim in movies:
# stim_order.append({'stimulus': stim})
#
# stim_order
#
# [{‘stimulus’:‘Interstellar’},
# {‘stimulus’:‘Love Actually’},
# {‘stimulus’:‘Forrest Gump’},…]
# ```
#
# The result is then provided as `input` for the [data.TrialHandler()](https://psychopy.org/api/data.html#psychopy.data.TrialHandler) `function`.
#
# With that we can simply `loop` over the `trials` in the `trials object` and during each `iteration` `draw` and `flip` the respective `value` of the `dictionary key` `“stimulus”` to present the stimuli of our `list` “movies” one-by-one after one another.
#
# ```
# for trial in trials:
#
# # display/draw respective stimulus within each iteration, notice how the stimulus is set "on the fly"
# visual.TextStim(win, text=trial['stimulus'], bold=True, pos=[0, 30], height=40).draw()
# ```
#
#
# Additionally, we want to display the question `“How much do you like the following?”` above the respective stimulus to remind participant about the task. Within each `iteration` of our `for-loop` we will also allow participants to `quit` the `experiment` by pressing `“escape”` via the `event.getKeys()` `function` as we don’t want to wait for a `key` to be `pressed` but want to do something whenever a certain `key` is `pressed`.
# +
#===============
# Import modules
#===============
from psychopy import gui, core, visual, event, data # import psychopy modules/functions
import os # import os module
import numpy.random as rnd # import random module from numpy
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
#===============================
# Creation of window and messages
#===============================
# Open a window
win = visual.Window(size=(800,600), color='gray', units='pix', fullscr=False) # set size, background color, etc. of window
# Define experiment start text
welcome_message = visual.TextStim(win,
text="Welcome to the experiment. Please press the spacebar to continue.",
color='black', height=40)
# Define trial start text
start_message = visual.TextStim(win,
text="In this experiment you will rate different movies, snacks and animals on a scale from 1 to 7. Please press the spacebar to start.",
color='black', height=40)
# Define experiment end text
end_message = visual.TextStim(win,
text="You have reached the end of the experiment, thanks for participating.",
color='black', height=40)
#==========================
# Define the trial sequence
#==========================
# Define a list of trials with their properties:
# create empty list
stim_order = []
# convert list into list of dictionaries with key ('stimulus') - value pairing
for stim in movies:
stim_order.append({'stimulus': stim})
# use dictionary to create trials object via data.TrialHandler, specifying further settings
trials = data.TrialHandler(stim_order, nReps=1, extraInfo=exp_info,
method='sequential') # create
#=====================
# Start the experiment
#=====================
# display welcome message
welcome_message.draw() # draw welcome message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# display start message
start_message.draw() # draw start message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# Run through the trials, stimulus by stimulus
for trial in trials:
# display/draw task question to remind participants
visual.TextStim(win, text='How much do you like the following?', pos=[0, 90], italic=True).draw()
# display/draw respective stimulus within each iteration, notice how the stimulus is set "on the fly"
visual.TextStim(win, text=trial['stimulus'], bold=True, pos=[0, 30], height=40).draw()
# after everything is drawn, flip it to the front screen
win.flip()
# if participants press `escape`, stop the experiment
if event.getKeys(['escape']):
core.quit()
#======================
# End of the experiment
#======================
# Display end message
end_message.draw() # draw end message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# -
# Let’s give it a try via `python experiment.py`. If everything works/is set correctly you should see the `GUI dialog box` and after clicking `OK`, the `text` we defined as a welcome message should appear next, followed by the start message. Subsequently, you should see all of our `stimuli` one after another and the same question above them every `trial`. Finally, you should see the end message.
python experiment.py
# While this is already great, the same thing as during our initial tests with the instruction screens happened: the `text`, i.e. our `stimuli`, is only very briefly shown on screen and disappears before we can do anything. That’s because we didn’t specify that we want to collect `responses` before moving on to the next `stimulus` yet. We need to add the `rating` to our `experiment`, specifically the `trials`.
#
# ## Input/output
#
# `PsychoPy` offers quite a bit of possible options to collect `responses`: simple yes/no questions, rating scales, visual analog scales, voice recordings, etc. and store outputs (`files`, different levels of detail, etc.).
#
# ### Collecting responses
#
# For the `experiment` at hand a simple `rating scale` (yes, a `Likert scale` to make it Psychology, hehe) should be sufficient. As with the other components we have explored so far, we need to implement/add this via two steps: `defining`/`creating` a `rating scale` and `draw`ing/presenting it.
#
# We can easily define and tweak a rating scale via `PsychoPy`’s [visual.RatingScale()](https://psychopy.org/api/visual/ratingscale.html#psychopy.visual.RatingScale) `function` which allows us to set the `range` of `values`, `labels`, `size`, etc..
#
# ```
# ratingScale = visual.RatingScale(win,
# scale = None,
# low = 1,
# high = 7,
# showAccept = True,
# markerStart = 4,
# labels = ['1 - Not at all', '7 - A lot'],
# pos = [0, -80])
# ```
#
#
# We then need to `draw` it and indicate that we want to `wait` until a `rating` was conducted before we advance to the next `trial`.
#
# ```
# while ratingScale.noResponse:
# ```
#
# Additionally, we are going to display a small helpful message describing the `rating` and make sure that the `rating scale` is reset back to its `default status` before the next `trial` starts.
#
# ```
# ratingScale.reset()
# ```
#
#
# Even though `participants` could already perform the `rating` of the `stimuli`, we don’t track and collect the respective `responses` yet. These need to be obtained from the `rating scale` before we `reset` it at the end of the `trial`. As indicated before `visual.RatingScale()` creates an `object`/`class`/`data type` with many inbuilt `functions`, this includes `.getRating()` and `.getRT()` to collect the provided `rating` and corresponding `response time`:
#
# ```
# rating = ratingScale.getRating()
# rt = ratingScale.getRT()
# ```
#
# We can then store both `values` per `trial` in the `trials object` via its `.addData()` `function`:
#
# ```
# trials.addData('rt', rt)
# trials.addData('rating', rating)
# ```
# +
#===============
# Import modules
#===============
from psychopy import gui, core, visual, event, data # import psychopy modules/functions
import os # import os module
import numpy.random as rnd # import random module from numpy
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
#===============================
# Creation of window and messages
#===============================
# Open a window
win = visual.Window(size=(800,600), color='gray', units='pix', fullscr=False) # set size, background color, etc. of window
# Define experiment start text
welcome_message = visual.TextStim(win,
text="Welcome to the experiment. Please press the spacebar to continue.",
color='black', height=40)
# Define trial start text
start_message = visual.TextStim(win,
text="In this experiment you will rate different movies, snacks and animals on a scale from 1 to 7. Please press the spacebar to start.",
color='black', height=40)
# Define experiment end text
end_message = visual.TextStim(win,
text="You have reached the end of the experiment, thanks for participating.",
color='black', height=40)
#==========================
# Define the trial sequence
#==========================
# Define a list of trials with their properties:
# create empty list
stim_order = []
# convert list into list of dictionaries with key ('stimulus') - value pairing
for stim in movies:
stim_order.append({'stimulus': stim})
# use dictionary to create trials object via data.TrialHandler, specifying further settings
trials = data.TrialHandler(stim_order, nReps=1, extraInfo=exp_info,
method='sequential') # create
#================================
# Define a rating scale
#================================
ratingScale = visual.RatingScale(win,
scale = None, # This makes sure there's no subdivision on the scale
low = 1, # This is the minimum value I want the scale to have
high = 7, # This is the maximum value of the scale
showAccept = True, # This shows the user's chosen value in a window below the scale
markerStart = 4, # This sets the rating scale to have its marker start on 5
labels = ['1 - Not at all', '7 - A lot'], # This creates the labels
pos = [0, -80]) # set the position of the rating scale within the window
#=====================
# Start the experiment
#=====================
# display welcome message
welcome_message.draw() # draw welcome message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# display start message
start_message.draw() # draw start message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# Run through the trials, stimulus by stimulus
for trial in trials:
# wait until a rating was conducted before advancing to the next trial
while ratingScale.noResponse:
# display/draw task question to remind participants
visual.TextStim(win, text='How much do you like the following?', pos=[0, 90], italic=True).draw()
# display/draw respective stimulus within each iteration, notice how the stimulus is set "on the fly"
visual.TextStim(win, text=trial['stimulus'], bold=True, pos=[0, 30], height=40).draw()
# display/draw the rating scale
ratingScale.draw()
# display/draw help message regarding rating scale
visual.TextStim(win, text='(Move the marker along the line and click "enter" to indicate your rating from 1 to 7.)',
pos=[0,-200], height=14).draw()
# after everything is drawn, flip it to the front screen
win.flip()
# if participants press `escape`, stop the experiment
if event.getKeys(['escape']):
core.quit()
# get the current rating
rating = ratingScale.getRating()
# get the response time of the current rating
rt = ratingScale.getRT()
# add the current rating to the trials object
trials.addData('rt', rt)
# add the response time of the current rating to the trials object
trials.addData('rating', rating)
# reset the rating scale
ratingScale.reset()
#======================
# End of the experiment
#======================
# Display end message
end_message.draw() # draw end message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# -
# Let’s give it a try via `python experiment.py`. If everything works/is set correctly you should see the `GUI dialog box` and after clicking `OK`, the `text` we defined as a welcome message should appear next, followed by the start message. Subsequently, you should see all of our `stimuli` one after another and the same question above them every `trial`, this time not advancing until you provided a `rating`. Finally, you should see the end message.
python experiment.py
# Our `experiment` works as expected but we don’t get any `output files`. The reason is once again simple: we actually didn’t tell `PsychoPy` that we would like to `save` our `data` to an `output file`. Importantly, our `data` is stored in the `trials object`.
#
# ### Saving data
#
# Because things work like a charm and we’re using `Python`-based tools, the `trials object` has an `in-built function`, called `.saveAsWideText()`, that easily allows this by indicating the desired `filename`, `type` and `delimiter`.
#
# ```
# trials.saveAsWideText(data_fname + '.csv', delim=',')
# ```
#
# +
#===============
# Import modules
#===============
from psychopy import gui, core, visual, event, data # import psychopy modules/functions
import os # import os module
import numpy.random as rnd # import random module from numpy
#========================================
# Create GUI dialog box for user input
#========================================
# Get subject name, age, handedness and other information through a dialog box
exp_name = 'PfP_2021' # set experiment name
exp_info = {
'participant': '', # participant name as string
'age': '', # age name as string
'left-handed':False, # handedness as boolean
'like this course':('yes', 'no'), # course feedback as tuple
'data path': '', # data path as string
}
dlg = gui.DlgFromDict(dictionary=exp_info, title=exp_name) # create GUI dialog box from dictionary
# If 'Cancel' is pressed, quit experiment
if dlg.OK == False:
core.quit()
#=================================================
# Data storage: basic information, filename & path
#=================================================
# Get date and time
exp_info['date'] = data.getDateStr() # get date and time via data module
exp_info['exp_name'] = exp_name # set experiment name
# Check if set data path exists, if not create it
if not os.path.isdir(exp_info['data path']):
os.makedirs(exp_info['data path'])
# Create a unique filename for the experiment data
data_fname = exp_info['participant'] + '_' + exp_info['date'] # create initial file name from participant ID and date/time
data_fname = os.path.join(exp_info['data path'], data_fname) # add path from GUI dialog box
#===============================
# Creation of window and messages
#===============================
# Open a window
win = visual.Window(size=(800,600), color='gray', units='pix', fullscr=False) # set size, background color, etc. of window
# Define experiment start text
welcome_message = visual.TextStim(win,
text="Welcome to the experiment. Please press the spacebar to continue.",
color='black', height=40)
# Define trial start text
start_message = visual.TextStim(win,
text="In this experiment you will rate different movies, snacks and animals on a scale from 1 to 7. Please press the spacebar to start.",
color='black', height=40)
# Define experiment end text
end_message = visual.TextStim(win,
text="You have reached the end of the experiment, thanks for participating.",
color='black', height=40)
#==========================
# Define the trial sequence
#==========================
# Define a list of trials with their properties:
# create empty list
stim_order = []
# convert list into list of dictionaries with key ('stimulus') - value pairing
for stim in movies:
stim_order.append({'stimulus': stim})
# use dictionary to create trials object via data.TrialHandler, specifying further settings
trials = data.TrialHandler(stim_order, nReps=1, extraInfo=exp_info,
method='sequential') # create
#================================
# Define a rating scale
#================================
ratingScale = visual.RatingScale(win,
scale = None, # This makes sure there's no subdivision on the scale
low = 1, # This is the minimum value I want the scale to have
high = 7, # This is the maximum value of the scale
showAccept = True, # This shows the user's chosen value in a window below the scale
markerStart = 4, # This sets the rating scale to have its marker start on 5
labels = ['1 - Not at all', '7 - A lot'], # This creates the labels
pos = [0, -80]) # set the position of the rating scale within the window
#=====================
# Start the experiment
#=====================
# display welcome message
welcome_message.draw() # draw welcome message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# display start message
start_message.draw() # draw start message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# Run through the trials, stimulus by stimulus
for trial in trials:
# wait until a rating was conducted before advancing to the next trial
while ratingScale.noResponse:
# display/draw task question to remind participants
visual.TextStim(win, text='How much do you like the following?', pos=[0, 90], italic=True).draw()
# display/draw respective stimulus within each iteration, notice how the stimulus is set "on the fly"
visual.TextStim(win, text=trial['stimulus'], bold=True, pos=[0, 30], height=40).draw()
# display/draw the rating scale
ratingScale.draw()
# display/draw help message regarding rating scale
visual.TextStim(win, text='(Move the marker along the line and click "enter" to indicate your rating from 1 to 7.)',
pos=[0,-200], height=14).draw()
# after everything is drawn, flip it to the front screen
win.flip()
# if participants press `escape`, stop the experiment
if event.getKeys(['escape']):
core.quit()
# get the current rating
rating = ratingScale.getRating()
# get the response time of the current rating
rt = ratingScale.getRT()
# add the current rating to the trials object
trials.addData('rt', rt)
# add the response time of the current rating to the trials object
trials.addData('rating', rating)
# reset the rating scale
ratingScale.reset()
#======================
# End of the experiment
#======================
# Save all data to a file
trials.saveAsWideText(data_fname + '.csv', delim=',')
# Display end message
end_message.draw() # draw end message to buffer screen
win.flip() # flip it to the front screen
keys = event.waitKeys(keyList=['space', 'escape']) # wait for spacebar key press before advancing or quit when escape is pressed
# -
# If you now try it again, everything should work as before bu after finishing the `experiment` you should see a new `file` within the indicated `data path` containing all `information` stored about the `experiment`: `trials`, `stimuli`, `responses`, `reaction times`, etc. .
# ## A very simple experiment
#
# Believe it or not folks, with that we already created our first working `PsychoPy` `experiment`. Using only a small amount of, _very readable_, `code`, we can obtain `ratings` for our `stimuli`. Obviously, this is a very simple `experiment` but nevertheless a good start, showcasing a lot of the core things you should know to start using `PsychoPy` for `experiments`. All the things addressed here are usually also part of much more complex `experiments`, as well as build their basis.
#
# <img align="center" src="https://raw.githubusercontent.com/PeerHerholz/Python_for_Psychologists_Winter2021/master/lecture/static/experiment_outline.png" alt="logo" title="jupyter" width="800" height="350" />
# ## Outro
#
# As usually: awesome work folks! The transition from basic `Python` aspects to applied together is definitely no cake-walk, especially when simultaneously switching to `python scripts` from `jupyter notebooks` but you did a great job!
# Thank so much for sticking with us throughout this!
#
# <img align="center" src="https://media4.giphy.com/media/7zWYE1ostmPWZdygj3/giphy.gif?cid=ecf05e475wkt0wsob0pgmaggnreymvom43vxe6ainhr1dzh0&rid=giphy.gif&ct=g" alt="logo" title="jupyter" width="600" height="300" />
| lecture/experiments/intro_psychopy_I.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from openml.datasets import list_datasets, get_datasets
import pandas as pd
from sklearn.model_selection import train_test_split
from supervised.automl import AutoML
import os
from prediction_feature.mljar import add_dataset
# + pycharm={"name": "#%%\n"}
dataset_dataframe = list_datasets(output_format="dataframe", number_classes='2..20').query('NumberOfInstances < 100000 & NumberOfFeatures < 1000')
dataset_dataframe
# + pycharm={"name": "#%%\n"}
dataset_ids = dataset_dataframe['did']
# + pycharm={"name": "#%%\n"}
dataset_dataframe_list = get_datasets(dataset_ids=dataset_ids)
# + pycharm={"name": "#%%\n"}
dataset = dataset_dataframe_list[8]
X, y, _, _ = dataset.get_data(
target=dataset.default_target_attribute)
print(X)
print('y: ')
print(y)
print('\n\n\n\n')
dataset_id = dataset.dataset_id
print(dataset_id)
dataset_dataframe.loc[dataset_id, dataset_dataframe.columns[6:]]
# + pycharm={"name": "#%%\n"}
for i in range(len(dataset_dataframe_list[:5])):
add_dataset(dataset_dataframe_list[i], dataset_dataframe)
# + pycharm={"name": "#%%\n"}
# + pycharm={"name": "#%%\n"}
| src/cumulator/prediction_feature/utils/openml_datasets_retrieval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Problem Statement
# The task is to predict whether a potential promotee at checkpoint in the test set will be promoted or not after the evaluation process.
# +
import pandas as pd
import numpy as np
import seaborn as sb
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
from sklearn import model_selection, metrics
from sklearn.grid_search import GridSearchCV
import matplotlib.pylab as plt
# %matplotlib inline
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 12, 4
# -
train_data='D:/My Personal Documents/Learnings/Data Science/Data Sets/WNS Analytics/train_LZdllcl.csv'
test_data='D:/My Personal Documents/Learnings/Data Science/Data Sets/WNS Analytics/test_2umaH9m.csv'
train_result='D:/My Personal Documents/Learnings/Data Science/Data Sets/WNS Analytics/train_result.csv'
train=pd.read_csv(train_data)
test=pd.read_csv(test_data)
test_results = pd.read_csv(train_result)
target = 'is_promoted'
IDcol = 'employee_id'
train['source']='train'
test['source']='test'
data = pd.concat([train, test],ignore_index=True)
print (train.shape, test.shape, data.shape)
data.apply(lambda x: sum(x.isnull()))
data.apply(lambda x: len(x.unique()))
plt.figure(figsize=(12,6))
sb.countplot(x='department', hue='education',data=data)
sb.countplot(x='is_promoted',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='department',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='education',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='awards_won?',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='no_of_trainings',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='gender',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='previous_year_rating',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='KPIs_met >80%',data=data)
plt.figure(figsize=(12,6))
sb.countplot(x='is_promoted', hue='recruitment_channel',data=data)
plt.figure(figsize=(12,6))
sb.distplot(data[data.is_promoted==0]['avg_training_score'])
def age_trans(age):
if(age<30):
return 'Young'
elif(age>=30 and age<40):
return 'Middle Age'
elif(age >=40):
return 'Senior'
data['age']=data['age'].apply(age_trans)
def rating_trans(rating):
if(rating>=4.0):
return 'High'
elif(rating==3.0):
return 'Medium'
elif(rating < 3.0):
return 'low'
data['previous_year_rating']=data['previous_year_rating'].apply(rating_trans)
data.previous_year_rating[data.previous_year_rating.isnull()]=3.0
data.education[data.education.isnull()]="Bachelor's"
data=pd.get_dummies(data,columns=['department','education','gender','recruitment_channel','age','previous_year_rating','region'])
train=data[data.source=='train']
train.drop('source',axis=1,inplace=True)
test=data[data.source=='test']
test.drop(['source','is_promoted'],axis=1,inplace=True)
train.head()
# +
def modelfit(alg, dtrain, dtest, predictors,useTrainCV=True, cv_folds=5, early_stopping_rounds=50):
if useTrainCV:
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(dtrain[predictors].values, label=dtrain[target].values)
xgtest = xgb.DMatrix(dtest[predictors].values)
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds,
metrics='auc', early_stopping_rounds=early_stopping_rounds)
alg.set_params(n_estimators=cvresult.shape[0])
#Fit the algorithm on the data
alg.fit(dtrain[predictors], dtrain[target],eval_metric='auc')
#Predict training set:
dtrain_predictions = alg.predict(dtrain[predictors])
dtrain_predprob = alg.predict_proba(dtrain[predictors])[:,1]
#Print model report:
print ("\nModel Report")
print ("Accuracy : %.4g" % metrics.accuracy_score(dtrain[target].values, dtrain_predictions))
print ("AUC Score (Train): %f" % metrics.roc_auc_score(dtrain[target], dtrain_predprob))
# Predict on testing data:
# dtest['predprob'] = alg.predict_proba(dtest[predictors])[:,1]
# results = test_results.merge(dtest[['employee_id','predprob']], on='employee_id')
# print ('AUC Score (Test): %f' % metrics.roc_auc_score(results[target], results['predprob']))
# feat_imp = pd.Series(alg.get_booster().get_fscore()).sort_values(ascending=False)
#feat_imp.plot(kind='bar', title='Feature Importances')
# plt.ylabel('Feature Importance Score')
# -
predictors = [x for x in train.columns if x not in [target, IDcol]]
xgb1 = XGBClassifier(
learning_rate =0.1,
n_estimators=202,
max_depth=5,
min_child_weight=1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb1, train, test, predictors)
pred=xgb1.predict(test[predictors])
result=pd.Series(pred.tolist()).astype(int)
result.to_csv('D:/My Personal Documents/Learnings/Data Science/Data Sets/WNS Analytics/final_result.csv')
result.value_counts()
# ### Below codes were used for tuning the parameters
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test6 = {
'reg_alpha':[1e-5, 1e-2, 0.1, 1, 100]
}
gsearch6 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=5,
min_child_weight=6, gamma=0.1, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test6, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch6.fit(train[predictors],train[target])
gsearch6.grid_scores_, gsearch6.best_params_, gsearch6.best_score_
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test5 = {
'subsample':[i/100.0 for i in range(75,90,5)],
'colsample_bytree':[i/100.0 for i in range(75,90,5)]
}
gsearch5 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=177, max_depth=4,
min_child_weight=6, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test5, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch5.fit(train[predictors],train[target])
# +
gsearch5.grid_scores_, gsearch5.best_params_, gsearch5.best_score_
# -
#Grid seach on subsample and max_features
#Choose all predictors except target & IDcols
param_test2 = {
'max_depth':[4,5,6],
'min_child_weight':[4,5,6]
}
gsearch2 = GridSearchCV(estimator = XGBClassifier( learning_rate=0.1, n_estimators=140, max_depth=5,
min_child_weight=2, gamma=0, subsample=0.8, colsample_bytree=0.8,
objective= 'binary:logistic', nthread=4, scale_pos_weight=1,seed=27),
param_grid = param_test2, scoring='roc_auc',n_jobs=4,iid=False, cv=5)
gsearch2.fit(train[predictors],train[target])
gsearch2.grid_scores_, gsearch2.best_params_, gsearch2.best_score_
| notebooks/WNS Analytics Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/akshit7165/Minor-Project-Lane-Obj-Detection/blob/main/Lane_object_detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="WZllmYhBl9-j" outputId="4edb3d5d-35f3-4d3e-b0a4-0ff490ef1b56"
# ! git clone https://github.com/akshit7165/Minor-Project-Lane-Obj-Detection.git
# ! mv Minor-Project-Lane-Obj-Detection/* ./
# ! mkdir videos
# ! mkdir images
# ! mkdir images/from_video
# ! mkdir images/detection
# # ! wget -P ./model_data/ https://pjreddie.com/media/files/yolov3.weights
# ! wget -P ./model_data/ https://s3-ap-southeast-1.amazonaws.com/deeplearning-mat/backend.h5
# !pip install git+git://github.com/minwook-shin/pytube --force
# + colab={"base_uri": "https://localhost:8080/"} id="flaA5psKppXa" outputId="41e4d9d6-b72a-4299-cbea-58f99175cc2b"
# !pip install pytube
from pytube import YouTube
import numpy as np
import cv2
from google.colab.patches import cv2_imshow
# + id="0fSGSgiRqsKQ"
def download_file(link, file) :
video = "./videos/" #to_do
yt = YouTube(link)
yt.streams.filter(progressive=True, file_extension='mp4').order_by('resolution').first().download(output_path=video, filename=str(file))
print(link, ' Downloaded to ',video+ str(file))
return video+ str(file)+".mp4"
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="ov9cIyvcGpx6" outputId="0fe8bfcb-b905-4346-be1c-1bc1631d7fe5"
# download_file("https://youtu.be/PA0O9Rapv38","nice_road")
download_file("https://www.youtube.com/watch?v=N0gzsIzzPJ4","nice_road")
# + colab={"base_uri": "https://localhost:8080/", "height": 408} id="1TyMYguSnHph" outputId="76b1397b-ea63-496c-db6b-404412ffa579"
# !pip install LANE_DETECTION
from lane_detection import LANE_DETECTION
video_reader = cv2.VideoCapture("/content/videos/nice_road")
fps = video_reader.get(cv2.CAP_PROP_FPS)
nb_frames = int(video_reader.get(cv2.CAP_PROP_FRAME_COUNT))
frame_h = int(video_reader.get(cv2.CAP_PROP_FRAME_HEIGHT))
frame_w = int(video_reader.get(cv2.CAP_PROP_FRAME_WIDTH))
pers_frame_time = 398# seconds
pers_frame = int(pers_frame_time *fps)
video_reader.set(1,pers_frame)
ret, image = video_reader.read()
ld = LANE_DETECTION( image,fps,
yellow_lower = np.uint8([ 20, 50, 110]),
yellow_upper = np.uint8([35, 255, 255]),
white_lower = np.uint8([ 0, 140, 0]),
white_upper = np.uint8([255, 255, 100]),
lum_factor = 110,
lane_start=[0.2,0.5])
cv2_imshow(image)
cv2_imshow(cv2.imread("./images/detection/perspective_lines.jpg"))
cv2_imshow(cv2.imread("./images/detection/vanishing_point.jpg"))
cv2_imshow(cv2.imread("./images/detection/edges.jpg"))
cv2_imshow(cv2.imread("./images/detection/mask.jpg"))
cv2_imshow(cv2.imread("./images/detection/masked_regions.jpg"))
cv2_imshow(cv2.imread("./images/detection/lane_width.jpg"))
# proc_img = ld.process_image(image)
# cv2_imshow(proc_img)
# + colab={"base_uri": "https://localhost:8080/"} id="TAMKTPqz5d3u" outputId="f2cd3f2e-444c-48d3-eb67-9200754f15d1"
image.shape
# + colab={"base_uri": "https://localhost:8080/"} id="v-HZThpynLeo" outputId="59fc7368-8efe-4a0d-e5a0-5162e8320896"
from frame import FRAME
file_path = "/content/videos/nice_road"
video_out = "/content/videos/output.mp4"
frame = FRAME(
ego_vehicle_offset = .0,
yellow_lower = np.uint8([ 20, 50, 110]),
yellow_upper = np.uint8([35, 255, 255]),
white_lower = np.uint8([ 0, 140, 0]),
white_upper = np.uint8([180, 255, 100]),
lum_factor = 110,
max_gap_th = 0.45,
YOLO_PERIOD = 2,
lane_start=[0.2,0.5] ,
verbose = 3)
frame.process_video(file_path, 2,\
video_out = video_out,pers_frame_time =398,\
t0 =398 , t1 =698)
# + id="tArbvmMoY5Lk"
| Lane_object_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# importing the libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# importing the data
df = sns.load_dataset('iris')
df.head()
df2 = df.copy()
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()
encoded = encoder.fit_transform(df['species'])
df['Species'] = encoded
df.drop(['species'],axis=1,inplace=True)
df.head()
# +
fig = plt.figure(figsize=(8,5))
ax = fig.add_subplot(111, projection = '3d')
x = df['sepal_length']
y = df['sepal_width']
z = df['petal_length']
ax.set_xlabel("sepal_length")
ax.set_ylabel("sepal_width")
ax.set_zlabel("petal_length")
ax.scatter(x, y, z, c=df['Species'])
plt.title("Plot before reducing the dimensions")
# -
# scaling the values present in the dataset
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_features = scaler.fit_transform(df)
scaled_df = pd.DataFrame(scaled_features, columns = df2.columns)
scaled_df[:5]
X = scaled_df[['sepal_length','sepal_width','petal_length']]
y = df2['species']
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
lda = LinearDiscriminantAnalysis(n_components=2)
x_lda = lda.fit_transform(X,y)
component_df = pd.DataFrame(data=x_lda, columns=['component_value 1','component_value 2'])
component_df[:5]
# checking the shape of the reduced data
component_df.shape
reduced_df = pd.concat([component_df,df2['species']],axis=1)
reduced_df[:5]
plt.figure(figsize=(6,5))
sns.scatterplot(data = reduced_df, x="component_value 1", y="component_value 2", hue="species", palette='Set1')
plt.title('Plot after reducing the dimensions')
# <h2>ANALYSIS</h2>
# * Let's create a ML model and train with original data and reduced data
# * Then we compare with the amount of time the model takes to train and also the accuracy
# Original data with 3 dimensions
df2.head()
# reduced data with 2 dimensions
reduced_df.head()
# <h3>Analysis for Original Data</h3>
X = df2[['sepal_length','sepal_width','petal_length']]
y = df2['Species']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3, random_state=42)
# let's consider a decision tree model
from sklearn.tree import DecisionTreeClassifier
original_model = DecisionTreeClassifier().fit(X_train, y_train)
original_pred = original_model.predict(X_test)
# model evaluation
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report(y_test,original_pred))
print(confusion_matrix(y_test, original_pred))
# <h3>Analysis for Reduced Data</h3>
X = reduced_df[['component_value 1','component_value 2']]
y = reduced_df['species']
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.3, random_state=42)
# let's consider a decision tree model
from sklearn.tree import DecisionTreeClassifier
reduced_model = DecisionTreeClassifier().fit(X_train, y_train)
reduced_pred = reduced_model.predict(X_test)
# model evaluation
from sklearn.metrics import classification_report, confusion_matrix
print(classification_report(y_test,reduced_pred))
print(confusion_matrix(y_test, reduced_pred))
# * we see that the reduced data is performing well
| Machine Learning/Principal Component Analysis/ML assign-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# Lanugages do not strictly follow the rules of concept. People tends to bend the rules to satisfy their need in communication.
#
# Instead of understanding the whole rules, we should ask ourselves "What are common patterns that occur in languages we use". To achieve the goal, we use mathematical tool call **Statistics**
# ## Terminology
#
# **Corpus**: A body of texts is call *corpus* in latin and the collection of *corpus* is namely called *corpora*.
#
# **Grammaticality**: An metrics is used to judge the structure of a sentence is well-formed.
#
# **Conventionality**: People frequently express in a certain way though there are other approaches exist.
#
# **word token**: The entity of word
#
# **word type**: Each word represents its own *type*.
# ## Resources
#
# There are several corpora we often use in Statistical NLP Task:
#
# * Brown Corpus
# * Balanced Corpus
# * Lancaster-Oslo-Bergen
# * Susanne Corpus
# * Penn Treebank
# * Canadian Hansard
# * WordNet
#
#
#
# ## Zipf's Law
#
# Zipf's Law shows that the product of words' frequency and their ranking $r$ is a constant:
#
# $$f \cdot r = k$$
#
# Simply to say, the frequency of word rank 100th is three times higher than the word rank 300th. However, this law is not really a Law rather an emprical result of word type statistic. The high rank words and the low rank ones bend the zip law severely.
# ## Mandelbrot's Law
#
# The Mandelbrot discovers the Zipf's Law cannot fit the words especially for the words in high and low rank so he addresses a more general approach:
#
# $$f = P (r + \rho)^{-B}$$
# or
# $$\log{f} = \log{P} - B \log(r + \rho)$$
#
# $B$ and $\rho$ are the parameters we tune for fitting the data. If $B = 0$ and $\rho = 0$, the Mandelbrot's approach will become exactly the same as Zipf's Law.
| chap1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Learning Rate Finder
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai import *
from fastai.docs import *
# -
# Learning rate finder plots lr vs loss relationship for a [`Learner`](/basic_train.html#Learner). The idea is to reduce the amount of guesswork on picking a good starting learning rate.
#
# **Overview:**
# 1. First run lr_find `learn.lr_find()`
# 2. Plot the learning rate vs loss `learn.recorder.plot()`
# 3. Pick a learning rate before it diverges then start training
#
# **Technical Details:** (first [described]('https://arxiv.org/abs/1506.01186') by <NAME>)
# >Train [`Learner`](/basic_train.html#Learner) over a few iterations. Start with a very low `start_lr` and change it at each mini-batch until it reaches a very high `end_lr`. [`Recorder`](/basic_train.html#Recorder) will record the loss at each iteration. Plot those losses against the learning rate to find the optimal value before it diverges.
# ## Choosing a good learning rate
# For a more intuitive explanation, please check out [Sylvain Gugger's post](https://sgugger.github.io/how-do-you-find-a-good-learning-rate.html)
data = get_mnist()
def simple_learner(): return Learner(data, simple_cnn((3,16,16,2)))
learn = simple_learner()
# First we run this command to launch the search:
# + hide_input=true
show_doc(Learner.lr_find)
# -
learn.lr_find()
# Then we plot the loss versus the learning rates. We're interested in finding a good order of magnitude of learning rate, so we plot with a log scale.
learn.recorder.plot()
# Then, we choose a value that is an order of magnitude before the mimum: the minimum value is on the edge diverging so it is too high. An order of magnitude before, a value that's still aggressive (for quicker training) but still safer from exploding. (In this example case 1e-1 is a good choice).
#
# Let's start training with this optimal value:
simple_learner().fit_one_cycle(2, 1e-1)
# Picking the minimum isn't a good idea because training will diverge.
learn = simple_learner()
simple_learner().fit_one_cycle(2, 1e-0)
# Picking a value to far below the minimum isn't optimal because training is too slow.
learn = simple_learner()
simple_learner().fit_one_cycle(2, 1e-2)
# + hide_input=true
show_doc(LRFinder)
# + hide_input=true
show_doc(LRFinder.on_train_end)
# + hide_input=true
show_doc(LRFinder.on_batch_end)
# + hide_input=true
show_doc(LRFinder.on_train_begin)
# + hide_input=true
show_doc(LRFinder.on_epoch_end)
| docs_src/callbacks.lr_finder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
me_m = pd.read_csv("data/Portfolios_Formed_on_ME_monthly_EW.csv",
header=0, index_col=0, parse_dates=True, na_values=-99.99)
me_m.head()
# -
cols = ['Lo 20', 'Hi 20']
returns = me_m[cols]
returns
returns = returns/100
returns.plot()
annualized_vol = returns.std()*np.sqrt(12)
annualized_vol
n_months = returns.shape[0]
return_per_month = (returns+1).prod()**(1/n_months) - 1
return_per_month
annualized_return = (return_per_month + 1)**12-1
annualized_return = (returns+1).prod()**(12/n_months) - 1
annualized_return
returns.index
returns.index = pd.to_datetime(returns.index, format="%Y%m")
returns.index
quiz_returns = returns["1999":"2015"]
quiz_returns
quiz_returns.index = quiz_returns.index.to_period('M')
quiz_returns.head()
quiz_returns.info()
quiz_rets = quiz_returns["1999":"2015"]
quiz_rets
quiz_n_months = quiz_rets.shape[0]
quiz_annualized_return = (quiz_rets+1).prod()**(12/quiz_n_months) - 1
quiz_annualized_return
quiz_n_months = quiz_rets.shape[0]
quiz_n_months
annualized_vol = quiz_returns.std()*np.sqrt(12)
annualized_vol
import edhec_risk_kit as erk
quiz_rets
quiz_rets.info()
quiz_rets["Lo 20"]
erk.drawdown(quiz_rets["Lo 20"])["Drawdown"].idxmin()
erk.drawdown(quiz_rets["Hi 20"])["Drawdown"].min()
erk.drawdown(quiz_rets["Hi 20"])["Drawdown"].idxmin()
hfi = erk.get_hfi_returns()
hfi
hfi["2009":]
erk.semideviation(hfi["2009":]).idxmin()
erk.semideviation(hfi["2009":])
erk.skewness(hfi["2009":]).idxmin()
erk.semideviation(hfi["2009":])
erk.kurtosis(hfi["2009":]).idxmin()
| course1/Quiz1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Statistics of the lengths
#
# We want to check that all the transformations we have done so far are sane so that we can work with a cleaned up dataset.
# +
import pandas as pd
df = pd.read_json("../data/processed/data.json")
df.head()
# -
# Let's look at some statistics of the extracted text lengths and the ratio.
df.describe()
# Whereas *count*, *mean*, *min* and *max* are self-explanatory, *std* stands for
# *standard deviation*. The rows with percentages are the 25%-, 50%-, and
# 75%-*quantiles*, respectively. They were defined in [my Blog post on means and
# medians](https://paul-grillenberger.de/?p=21). Here's a short refresher: The 25%-quantile is a value such that 25%
# of the data is smaller than or equal to it and the other 75% of the data is
# greater than or equal to it. The 50%-quantile is also known as the median.
#
# The minimum
# of 61 characters in Description_Length looks reasonable but a Containment
# Procedure with 0 characters? This has to be investigated. Before we do so, let
# us look at the same statistics but grouped by each label.
df.groupby("Label").describe().stack()
# This is where it starts to get interesting! As *safe* SCPs are much easier to contain than *euclid* ones which in turn are easier to contain than *keter* SCPs, we expect that the Containment Procedures are easier to describe for safe ones and need more elaborate descriptions for keter ones. On average, this is reflected in the mean length of Containment Procedures (579 for safe, 833 for euclid and 1108 for keter).
# Let us turn to the problematic cases of zero lengths.
df.loc[(df["Procedures_Length"] == 0) | (df["Description_Length"] == 0)]
# Thankfully, this is a single outlier. Investigating the article on the SCP Foundation web page and inspecting the html yields that the label "Special Containment Procedures" sits in its own `p` element so that we were not able to crawl this article correctly.
#
# Let us ignore the outlier.
df = df.loc[df["Procedures_Length"] > 0]
# Finally, let us compute correlations between our features and the target. The correlation coefficient may be computed for number-valued random variables. Thankfully, the *nominal* labels safe, euclid, and keter, carry *ordinal* information. That is to say, we can order them by their *containment complexity*.
# To make this even more explicit, let us assign numbers to the three labels. A safe label will be converted to -1, a euclid label to 0 and a keter label to 1 so that the order of the containment complexity is reflected by $\mathrm{safe} < \mathrm{euclid} < \mathrm{keter}$. However, the magnitude of this conversion is still open for discussion. Alternatively, we could have choosen $10^{100}$ for keter and this would have influenced the correlation coefficients. But let's stick to our simple way of converting for now.
# +
COMPLEXITY = {
"SAFE": -1,
"EUCLID": 0,
"KETER": 1
}
def compute_complexity(label):
return COMPLEXITY[label]
df["Complexity"] = df["Label"].apply(compute_complexity)
df.corr()
# -
# As it turns out, Complexity and Procedures_Length are positively correlated which is precisely what we have observed through the statistics that we have grouped by label. We also see that Description_Length is only very weakly correlated with Complexity: That is to say that there is no reason why, say, a safe SCP should not have a long description or why a keter SCP could not be described in a short manner.
| notebooks/statistics_of_the_lengths.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Metrolyrics (Pagination, scraping 1x per row)
# ## <NAME>
#
# http://www.metrolyrics.com/rem-lyrics.html (or whatever other musician you'd like!)
#
# Remember, we're scraping multiple pages of search results, so the URL will be different!
#
# Scrape all pages of search results for your musician, and save as a CSV file. Include the following fields:
#
# - Song title
# - URL
# - Popularity
# - Year
#
# Bonus: Make the popularity a normal number (e.g., 6)
import pandas as pd
import requests
from bs4 import BeautifulSoup
response = requests.get('http://www.metrolyrics.com/mariah-carey-lyrics.html')
doc = BeautifulSoup(response.text)
#print(doc.prettify())
div = doc.find('div', attrs={'class': 'content'})
# +
Mariah = doc.find_all(class_="title hasvidtable")
rows = []
for diva in Mariah:
row = {}
row['Song'] = diva.get('title')[13:-7]
row['URL'] = diva.get('href')
row['Year'] = diva.findNext('td').text.strip()
row['Popularity'] = diva.findNext('span').get('class')[1][7:] #Makes popularity a normal number
if "mariah-carey" in row['URL']:
rows.append(row)
rows
# -
# # Making Links, Building Loops
for page_num in range(1, 6): #start num and last num + 1
url = f"http://www.metrolyrics.com/mariah-carey-alpage-{page_num}.html"
print(url)
# +
print("Making empty list")
rows = []
for page_num in range(1, 6): # 1-10
url = f"http://www.metrolyrics.com/mariah-carey-alpage-{page_num}.html"
print("Now scraping", url)
#Download the appropriate page
response = requests.get(url)
doc = BeautifulSoup(response.text)
#Scraping
Mariah = doc.find_all(class_="title hasvidtable")
for diva in Mariah:
row = {}
row['Song'] = diva.get('title')[13:-7]
row['URL'] = diva.get('href')
row['Year'] = diva.findNext('td').text.strip()
row['Popularity'] = diva.findNext('span').get('class')[1][7:] #Makes popularity a normal number
if "mariah-carey" in row['URL']:
rows.append(row)
# Don't create dataframe until end
print("Building Pandas DataFrame")
df = pd.DataFrame(rows)
df.head()
# -
df.sort_values(by='Year').head() #The 'Year' data on this site is incorrect.
#Mariah first started releasing solo music in 1990
df.to_csv("Metrolyrics.csv", index=False)
# # Metrolyrics, Part 2: Scrape the lyrics pages
#
# Then, open your search results csv, and scrape the following field:
#
# - Lyrics
#
# Merge with your original song information and save as a new CSV file
#
# Tip: If you use .find for your lyrics, they'll have a bunch of ads inside! You can use the ingredients/directions trick from above, or you can clean them with regex.
df_Mariah = pd.read_csv("Metrolyrics.csv")
df_Mariah.head()
# +
rows = []
url = f"http://www.metrolyrics.com/one-sweet-day-lyrics-mariah-carey.html"
print("Now scraping", url)
#Download the appropriate page
response = requests.get(url)
doc = BeautifulSoup(response.text)
page = doc.find_all('div', id='lyrics-body-text')
for lyrics in page:
row = {}
inner_list = []
for words in lyrics.find_all(class_="verse"):
inner_list.append(words.text.strip())
song_lyrics = ' '.join(inner_list)
row['lyrics'] = song_lyrics
for key, value in row.items():
row[key] = value.replace('\n', ' ')
rows.append(row)
rows
# -
def scrape_page(row):
url = f"{row['URL']}"
print("Scraping:", url)
response = requests.get(url)
doc = BeautifulSoup(response.text)
page = doc.find_all('div', id='lyrics-body-text')
for lyrics in page:
row = {}
inner_list = []
for words in lyrics.find_all(class_="verse"):
inner_list.append(words.text.strip())
song_lyrics = ' '.join(inner_list)
row['lyrics'] = song_lyrics
for key, value in row.items():
row[key] = value.replace('\n', ' ')
return pd.Series(row)
lyrics_df = df_Mariah.apply(scrape_page, axis=1) #take df, go through every row, scrape page
#lyrics_df.head()
lyrics_df.shape
lyrics = df_Mariah.merge(lyrics_df, left_index=True, right_index=True)
lyrics.head()
lyrics.to_csv("Lyrics.csv", index=False)
| 09-homework/MetroLyrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Script to create zonal mean of polygons (in shp file)
import os
from time import time
import xarray as xr
import rioxarray
import fiona
import rasterio
import rasterio.mask as msk
import numpy as np
from datetime import date, datetime
from dateutil.rrule import rrule, DAILY
import pandas as pd
# #### Function to perform zonal mean on polygon and output data in csv file
def zonal_stats(shape_path, infile, parameter,
dict={"Year": [], "DOY": [], "parameter": [], "YYYYDOY": [], 'ID': [], 'name': [], 'mean': []}):
# === Zonal Stats ===
with fiona.open(shape_path, 'r') as shp:
features = [feature for feature in shp]
with rasterio.open(infile) as src:
for f in features:
src_shp = [f['geometry']]
outimage, out_transform = msk.mask(src, src_shp, crop=True)
ws_id = f['id']
ws_name = f['properties']['Name']
ws_mean = np.nanmean(outimage)
print(ws_mean)
ws_year = infile.split('.')[0][-7:-3]
ws_doy = infile.split('.')[0][-3:]
dict['name'].append(ws_name)
dict['mean'].append(ws_mean)
dict['Year'].append(ws_year)
dict['DOY'].append(ws_doy)
dict['YYYYDOY'].append('{}_{}'.format(ws_year,ws_doy))
dict['parameter'].append(parameter)
dict['ID'].append(ws_id)
return dict
# +
path_mos_dat = 's3://dev-et-data/enduser/DelawareRiverBasin/Run09_13_2020/ward_sandford_customer' # /year
path_mos_ppt = 's3://dev-et-data/in/USA_data/precip_gridmet_tiffs' # /year
shape_path = 'Small Watersheds 128_Aggregated.shp'
csv_output = '.' #"." represent the current dir in Linux or type '/home/jupyter-kagone/postprocess'
parameter = 'prec'
a = datetime(2000, 1, 1)
b = datetime(2000, 1, 3)
#b = datetime(2019, 12, 31)
for dt in rrule(DAILY, dtstart=a, until=b):
#print(dt.strftime("%Y-%m-%d")) # this returns string
year = dt.strftime("%Y")
month = dt.strftime("%m")
day = dt.strftime("%d")
doy = dt.timetuple().tm_yday
print('Day: {} = {}-{}-{}'.format(doy, year, month, day))
in_file = os.path.join(path_mos_ppt, year, '{}_{}{:03d}.tif'.format(parameter, year, doy))
print(in_file)
if dt == a:
outdict = zonal_stats(shape_path, in_file, parameter)
elif dt < b:
outdict = zonal_stats(shape_path, in_file, parameter, dict=outdict)
else:
finaldict = zonal_stats(shape_path, in_file, parameter, dict=outdict)
#csvdf = pd.DataFrame(finaldict, columns=['Year', 'DOY', 'YYYYDOY', 'parameter', 'ID', 'name', 'mean'])
csvdf = pd.DataFrame(finaldict, columns=['YYYYDOY', 'ID', 'mean'])
print(csvdf)
#csvdf = csvdf.pivot(index=None, columns='ID', values='mean')
csvdf = pd.pivot_table(csvdf, values='mean', index='YYYYDOY', columns='ID').reset_index()
csvdf[['YYYY','DOY']] = csvdf.YYYYDOY.apply(lambda x: pd.Series(str(x).split("_")))
shp_list = list(range(53))
listf = []
listf.append('YYYYDOY')
listf.append('YYYY')
listf.append('DOY')
for elem in shp_list:
listf.append('{}'.format(elem))
#print(listf)
csvdf = csvdf[listf]
#csvdf.to_csv(os.path.join(csv_output, '{}_zonalmean_2000_2019a.csv'.format(parameter)))
print('done')
# -
| postprocess-avgs/other_fun_schitt/zonalstats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # The following blocks of codes are for pruning purpose.
# ### The difference between this and the original pruning experiment is that the training data is all randomly generated
# +
import os
from copy import deepcopy
import torch
import torch.nn as nn
from torchvision import transforms
import cv2
import numpy as np
import matplotlib.cm as cm
from src.utils.plotting import make_matching_figure
from pathlib import Path
import torch_pruning as tp
from src.loftr.backbone import build_backbone
from src.loftr.backbone.resnet_fpn import BasicBlock
from einops.einops import rearrange
from src.loftr.utils.position_encoding import PositionEncodingSine
from src.loftr.loftr_module import LocalFeatureTransformer, FinePreprocess
from src.loftr.utils.coarse_matching import CoarseMatching
from src.loftr.utils.fine_matching import FineMatching
from src.loftr import LoFTR, default_cfg
# -
# The default config uses dual-softmax.
# The outdoor and indoor models share the same config.
# You can change the default values like thr and coarse_match_type.
_default_cfg = deepcopy(default_cfg)
_default_cfg['coarse']['temp_bug_fix'] = True # set to False when using the old ckpt
original_backbone = build_backbone(_default_cfg)
from collections import OrderedDict
backbone_weight = OrderedDict()
remain_weight = torch.load("./weights/indoor_ds_new.ckpt")['state_dict']
for k in list(remain_weight.keys()):
if k.startswith('matcher.backbone.'):
backbone_weight[k.replace('matcher.backbone.', '', 1)] = remain_weight.pop(k)
original_backbone.load_state_dict(backbone_weight)
new_backbone = deepcopy(original_backbone)
# +
def prune_model(model):
model.cpu()
DG = tp.DependencyGraph().build_dependency( model, torch.randn(1, 1, 480, 640) )
def prune_conv(conv, amount=0.2):
#weight = conv.weight.detach().cpu().numpy()
#out_channels = weight.shape[0]
#L1_norm = np.sum( np.abs(weight), axis=(1,2,3))
#num_pruned = int(out_channels * pruned_prob)
#pruning_index = np.argsort(L1_norm)[:num_pruned].tolist() # remove filters with small L1-Norm
strategy = tp.strategy.L1Strategy()
pruning_index = strategy(conv.weight, amount=amount)
plan = DG.get_pruning_plan(conv, tp.prune_conv, pruning_index)
plan.exec()
block_prune_probs = [0.05, 0.05, 0.1, 0.1, 0.1, 0.1]
blk_id = 0
for m in model.modules():
if isinstance( m, BasicBlock ):
prune_conv( m.conv1, block_prune_probs[blk_id] )
prune_conv( m.conv2, block_prune_probs[blk_id] )
blk_id+=1
return model
transform = transforms.Compose([
transforms.RandomCrop((480,640)),
transforms.RandomHorizontalFlip(p=0.5)
])
def get_random_img(img_list):
img = cv2.imread(img_list[np.random.randint(0,len(img_list))], cv2.IMREAD_GRAYSCALE)
#img = cv2.resize(img, (640, 480))
img = torch.from_numpy(img)[None][None] / 255. #return an image tensor
img = transform(img)
return img
# -
#prune model
#prune_model(new_backbone)
new_backbone = torch.load('./temp_backbone/untrain.pth')
# +
#create the retrain img list
img_list = []
for path in Path('/home/cvte-vm/Datasets/ScanNet/scannet_test_1500').rglob('*.jpg'):
img_list.append(str(path))
for path in Path('/home/cvte-vm/Datasets/Megadepth/megadepth_test_1500').rglob('*.jpg'):
img_list.append(str(path))
#knowledge distillation
#freeze original model
for name, param in original_backbone.named_parameters():
param.requires_grad = False
original_backbone = original_backbone.cuda()
original_backbone.eval()
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(new_backbone.parameters(), lr=0.02, betas=(0.9, 0.999), eps=1e-08, weight_decay=0)
lr_scheduler = torch.optim.lr_scheduler.ExponentialLR(optimizer, gamma=0.95, last_epoch=-1, verbose=False)
num_epoch = 4
num_step = 300
batch_size = 8
best_model_wts = deepcopy(new_backbone.state_dict())
def iterative_pruning(prune_time, criterion, optimizer, scheduler, original_backbone, new_backbone):
for time in range(prune_time):
print()
print('**************************')
print(f"starting the {time} prune")
#prune model
#prune_model(new_backbone)
new_backbone = new_backbone.cuda()
new_backbone.train()
epoch_loss_old = 100
num_param = sum(p.numel() for p in new_backbone.parameters())
print(f"total parameters for backbone is now {num_param} after pruning, now start retrain")
#start retrain
for i in range(num_epoch):
print(f"now learning rate becomes {optimizer.param_groups[0]['lr']}")
running_loss = 0.0
for j in range(num_step):
#first prepare data batch
img = get_random_img(img_list) #first img in a batch
#for bs in range(batch_size-1):
#img1 = get_random_img(img_list)
#img = torch.cat([img, img1], dim=0) #concatenate in batch dimension, now img is a batch
img = torch.randint(0,256,(8,1,480,640))/255.0
#print(img*255)
img = img.cuda()
#finding loss
optimizer.zero_grad()
(layer2_label, layer4_label) = original_backbone(img) #soft label from teacher
(layer2_student, layer4_student) = new_backbone(img) #student prediction
loss1 = criterion(layer2_student, layer2_label)
loss2 = criterion(layer4_student, layer4_label)
total_loss = loss1+loss2
total_loss.backward()
optimizer.step()
if j%10 == 0:
print('step'+str(j)+' loss is {:.4f} '.format(total_loss))
#calculating loss to check training result
running_loss += total_loss.item() * batch_size
epoch_loss = running_loss/(num_step*batch_size)
print('*******epoch loss is {:.4f} '.format(epoch_loss))
if epoch_loss < epoch_loss_old: #save if loss gets smaller
epoch_loss_old = epoch_loss
best_model_wts = deepcopy(new_backbone.state_dict())
if epoch_loss < 0.07: #good enough, start next prune
torch.save(new_backbone, '/home/cvte-vm/Deep_Feature_Extract/LoFTR/temp_backbone/backbones'+str(time)+'.pth')
torch.save(new_backbone.state_dict(), '/home/cvte-vm/Deep_Feature_Extract/LoFTR/temp_backbone/backbones'+str(time)+'.dict')
for g in optimizer.param_groups:
g['lr'] = 0.0003*(2**time)
break
lr_scheduler.step() #decay the learning rate for next epoch
for g in optimizer.param_groups: #reset learning rate for next prune
g['lr'] = 0.0003*(2**time)
if epoch_loss > 0.1: #if the loss cannot be optimized anymore, then stop pruning
print(f"can only prune {time+1} time, cannot continue")
new_backbone.load_state_dict(best_model_wts)
return new_backbone
else: #when 0.07<loss<0.10, you still save the model
torch.save(new_backbone, '/home/cvte-vm/Deep_Feature_Extract/LoFTR/temp_backbone/backbones'+str(time)+'.pth')
torch.save(new_backbone.state_dict(), '/home/cvte-vm/Deep_Feature_Extract/LoFTR/temp_backbone/backbones'+str(time)+'.dict')
new_backbone.load_state_dict(best_model_wts)
return new_backbone
# -
new_backbone = iterative_pruning(1, criterion, optimizer, lr_scheduler, original_backbone, new_backbone)
torch.save(new_backbone, '/home/cvte-vm/Deep_Feature_Extract/LoFTR/temp_backbone/random2.pth')
torch.save(new_backbone.state_dict(), '/home/cvte-vm/Deep_Feature_Extract/LoFTR/temp_backbone/random2.dict')
# # The following blocks of code is only for visualization, you don't need to run it
for m in original_backbone.modules():
#print(m)
if isinstance( m, BasicBlock ):
print(m)
print('............................')
#original
pytorch_total_params = sum(p.numel() for p in original_backbone.parameters())
pytorch_total_params
#new
pytorch_total_params = sum(p.numel() for p in new_backbone.parameters())
pytorch_total_params
#4882176 for first prune
#4111846 for second prune
#3528448 for third prune
#3258448 for fourth prune
#3103365 for fifth prune
#2772528 for sixth prune
#2520887 for seventh prune
#2338495 for eighth prune
#2193625 for ninth prune
#2080770 for tenth prune
pytorch_total_params1 = sum(p.numel() for p in matcher.parameters())
pytorch_total_params1
#original weight
for name, param in original_backbone.named_parameters():
print(name)
print(param)
print('..................')
#new weight
for name, param in new_backbone.named_parameters():
print(name)
print(param)
print('..................')
#original
original_backbone
new_backbone
torch.rand(1,2,3)
random_backbone = torch.load('./temp_backbone/random.pth')
random_backbone
seventh_backbone = torch.load('./temp_backbone/seventh_prune.pth')
seventh_backbone
untrain_backbone = torch.load('./temp_backbone/untrain.pth')
untrain_backbone
(torch.randint(0,255,(8,1,480,640))/255.0).shape
torch.randint(0,2,(4,4))
2**6/127
| .ipynb_checkpoints/pruning_experiment2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import networkx as nx
import pandas as pd
import seaborn as sns
import numpy as np
cmap = sns.cm.rocket_r
import matplotlib.pyplot as plt
import pandas as pd
from collections import defaultdict
import matplotlib.patches as patches
# %matplotlib inline
# -
G = nx.read_gpickle('../../data/processed/SP_multiGraph_Job_Edu_Level.gpickle')
dist_df = pd.read_csv('distritos.csv', sep=';')
dist_df = dist_df[dist_df['cod_municipio'] == 36].copy()
# +
centro_zones = ['Bela Vista','Bom Retiro','Cambuci','Consolação',
'Liberdade','República','Santa Cecília','Sé']
centro_codes = dist_df[dist_df['distrito'].isin(centro_zones)].sort_values(by='cod_distrito')['cod_distrito'].unique()
centro_nomes = dist_df[dist_df['distrito'].isin(centro_zones)].sort_values(by='cod_distrito')['nome'].values
tmp = dist_df[dist_df['cod_distrito'].isin(centro_codes)]['distrito']
print(tmp[~tmp.isin(centro_zones)])
pd.Series(centro_zones)[~pd.Series(centro_zones).isin(tmp)]
# +
leste_zones = ['Aricanduva', 'Carrão', 'Vila Formosa', 'Cidade Tiradentes', 'Ermelino Matarazzo', 'Ponte Rasa', 'Guaianases',
'Lajeado', 'Itaim Paulista', 'Vila Curuçá', 'Cidade Líder', 'Itaquera', 'José Bonifácio', 'Parque do Carmo', 'Água Rasa',
'Belém', 'Brás', 'Mooca', 'Pari', 'Tatuapé', '<NAME>', 'Cangaíba', 'Penha', '<NAME>', 'Iguatemi', 'São Mateus',
'São Rafael', '<NAME>', 'São Miguel', '<NAME>', 'São Lucas', 'Sapopemba', 'Vila Prudente', 'Cachoeirinha',
'Casa Verde']
leste_codes = dist_df[dist_df['distrito'].isin(leste_zones)].sort_values(by='cod_distrito')['cod_distrito'].unique()
leste_nomes = dist_df[dist_df['distrito'].isin(leste_zones)].sort_values(by='cod_distrito')['nome'].values
tmp = dist_df[dist_df['cod_distrito'].isin(leste_codes)]['distrito']
print(tmp[~tmp.isin(leste_zones)])
pd.Series(leste_zones)[~pd.Series(leste_zones).isin(tmp)]
# +
norte_zones = ['Limão', 'Brasilândia', 'Freguesia do Ó', 'Jaçanã', 'Tremembé', 'Anhanguera', 'Perus', 'Jaraguá', 'Pirituba',
'São Domingos', 'Mandaqui', 'Santana', 'Tucuruvi', 'Vila Guilherme', 'Vila Maria', 'Vila Medeiros']
norte_codes = dist_df[dist_df['distrito'].isin(norte_zones)]['cod_distrito'].unique()
norte_nomes = dist_df[dist_df['distrito'].isin(norte_zones)].sort_values(by='cod_distrito')['nome'].unique()
tmp = dist_df[dist_df['cod_distrito'].isin(norte_codes)]['distrito']
print(tmp[~tmp.isin(norte_zones)])
pd.Series(norte_zones)[~pd.Series(norte_zones).isin(tmp)]
# +
oeste_zones = ['Butantã','Morumbi','Raposo Tavares','Rio Pequeno','Vila Sônia','Barra Funda', 'Jaguara','Jaguaré',
'Lapa','Perdizes','Vila Leopoldina','Alto de Pinheiros','Itaim Bibi', 'Jardim Paulista','Pinheiros']
oeste_codes = dist_df[dist_df['distrito'].isin(oeste_zones)]['cod_distrito'].unique()
oeste_nomes = dist_df[dist_df['distrito'].isin(oeste_zones)].sort_values(by='cod_distrito')['nome'].unique()
tmp = dist_df[dist_df['cod_distrito'].isin(oeste_codes)]['distrito']
print(tmp[~tmp.isin(oeste_zones)])
pd.Series(oeste_zones)[~pd.Series(oeste_zones).isin(tmp)]
# +
sul_zones = ['Campo Limpo','Capão Redondo','Vila Andrade','Cidade Dutra','Grajaú','Socorro','Cidade Ademar','Pedreira','Cursino',
'Ipiranga','Sacomã','Jabaquara','Jardim Ângela','Jardim São Luís','Marsilac','Parelheiros','Campo Belo','Campo Grande',
'Santo Amaro','Moema','Saúde','Vila Mariana']
sul_codes = dist_df[dist_df['distrito'].isin(sul_zones)]['cod_distrito'].unique()
sul_nomes = dist_df[dist_df['distrito'].isin(sul_zones)].sort_values(by='cod_distrito')['nome'].unique()
tmp = dist_df[dist_df['cod_distrito'].isin(sul_codes)]['distrito']
print(tmp[~tmp.isin(sul_zones)])
pd.Series(sul_zones)[~pd.Series(sul_zones).isin(tmp)]
# -
all_codes = np.concatenate([centro_codes, oeste_codes, sul_codes, leste_codes, norte_codes])
all_codes.size
all_nomes = np.concatenate([centro_nomes, oeste_nomes, sul_nomes, leste_nomes, norte_nomes])
all_nomes.size
zona_dist_dict = {}
for i, row in dist_df[['zona', 'cod_distrito']].iterrows():
zona_dist_dict[row['zona']] = row['cod_distrito']
def make_dist_dic(relations=['home', 'school', 'work', 'neighbor']):
dist_dic = {}
for d in dist_df[dist_df['cod_municipio'] == 36]['cod_distrito'].unique():
dist_dic[int(d)] = defaultdict(int)
for x,y,z in G.edges(data=True):
if z['edge_type'] in relations:
home_x = zona_dist_dict[int(G.nodes[x]['home'])]
home_y = zona_dist_dict[int(G.nodes[y]['home'])]
dist_dic[home_x][home_y] += 1
dist_dic[home_y][home_x] += 1
return dist_dic
# +
def make_df(region_codes, region_dic):
region_dic = { new_key: dist_dic[new_key] for new_key in region_codes}
print(len(region_dic.keys()))
df = pd.DataFrame.from_dict(region_dic).T
df = df[list(region_codes)]
df = df.dropna(how='all').T.fillna(0).sort_index()
df = df.reindex(sorted(df.columns), axis=1)
return df
def make_heatmap(df, labels, name, squares, vmax=1500):
fig, ax = plt.subplots(figsize=(12,10))
fig = sns.heatmap(df, annot=False, cmap=cmap, cbar=False, vmax=vmax)
print(df.shape)
# Create a Rectangle patch
origin = 0
for i in range(len(squares)):
rect = patches.Rectangle((origin, origin), squares[i], squares[i],
linewidth=.7,edgecolor='b',facecolor='none')
origin += squares[i]
ax.add_patch(rect)
plt.tick_params(
axis='both', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
left=False, # ticks along the top edge are off
labelbottom=False,
labelleft=False) # labels along the bottom edge are off
#ax.set_yticklabels(labels, rotation=0)
#ax.set_xticklabels(labels, rotation=45, ha="right", rotation_mode="anchor")
#ax.set_aspect('equal','box')
plt.savefig(name)
plt.show()
# -
squares = [len(x) for x in [centro_codes, oeste_codes, sul_codes, leste_codes, norte_codes]]
dist_dic = make_dist_dic(['school'])
df_all = make_df(all_codes, dist_dic)
make_heatmap(df_all, [], name='region_heatmap_school_squares_no_legend.pdf', squares=squares, vmax=500)
dist_dic = make_dist_dic(['home', 'neighbor'])
df_centro = make_df(centro_codes, dist_dic)
make_heatmap(df_centro, centro_nomes, vmax=750, name='all_heatmap_centro.pdf', squares = [2, 4, 2])
dist_dic = make_dist_dic(['work'])
df_leste = make_df(leste_codes, dist_dic)
make_heatmap(df_leste, leste_nomes, vmax=750, name='all_heatmap_works_leste.pdf')
df_norte = make_df(norte_codes)
make_heatmap(df_norte, norte_nomes)
df = pd.DataFrame.from_dict(zones_dic).T.fillna(0)
df = df.reindex(sorted(df.columns), axis=1)
sns.heatmap(df.loc[:288, :288], annot=False, cmap=cmap, vmax=100)
def make_heatmap(people_dict, title, filename):
df = pd.DataFrame.from_dict(people_dict).T.fillna(0)
df = df.sort_values(by='age').reset_index(drop=True)
df['age'] = pd.cut(df.age, bins=bins, labels=labels)
df = df.set_index(df.age).drop('age', axis=1)
df.columns = pd.cut(df.columns.to_list(), bins=bins, labels=labels)
df = df.groupby(df.columns, axis=1).sum()
df = df.groupby('age').mean()
df = df.reindex(sorted(df.columns, reverse=True), axis=1).T
df.rename(columns = {df.columns[-1]: str(df.columns[-1]) + '+'}, inplace=True)
df.rename(index = {df.index[0]: str(df.index[0]) + '+'}, inplace=True)
sns.heatmap(df, annot=False, vmin=0, vmax=10, cmap=cmap)
plt.title(title, fontsize=18)
plt.ylabel('Age of Contact', fontsize=16)
plt.xlabel('Age of Person', fontsize=16)
plt.savefig(f'{filename}.png', dpi=300)
make_heatmap(people, 'All Relations', 'heatmap_no_restriction')
# +
people_no_schools = {}
for i,person in enumerate(G.nodes()):
people_no_schools[i] = defaultdict(int)
people_no_schools[i]['age'] = G.nodes[person]['age']
for x,y,values in G.edges(person, data=True):
if values['edge_type'] != 'school':
people_no_schools[i][G.nodes[y]['age']] += 1
# -
np.unique([z['edge_type'] for x,y,z in G.edges(data=True)])
make_heatmap(people_no_schools, 'No School Relations', 'heatmap_no_schools')
# +
people_no_work = {}
for i,person in enumerate(G.nodes()):
people_no_work[i] = defaultdict(int)
people_no_work[i]['age'] = G.nodes[person]['age']
for x,y,values in G.edges(person, data=True):
if values['edge_type'] != 'work':
people_no_work[i][G.nodes[y]['age']] += 1
# -
make_heatmap(people_no_work, 'No Work Relations', 'heatmap_no_work')
for z in range(1,340):
l = [(x,y,()) for x,y,v in G.edges(data=True) if v['edge_type'] == 'work' and v['zone'] == z]
print(len(l))
edges = [(x,y,()) for x,y,v in G.edges(data=True) if v['edge_type'] == 'work' and v['zone'] == 1]
s = nx.MultiGraph(edges)
s.nodes()
nx.write_gml(nx.relabel_nodes(s, lambda x: str(x)), 'subgraph_work.gml')
import json
with open('work_subgraph.json', 'w') as f:
f.write(json.dumps(nx.cytoscape_data(nx.relabel_nodes(s, lambda x: str(x)))))
nx.draw(nx.relabel_nodes(s, lambda x: str(x)), node_size=15)
| notebooks/Modelagem Grafo SP/Validando modelagem - Contatos por Zona.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solutions
# ## Problem 1
# Implement the Min-Max scaling function ($X'=a+{\frac {\left(X-X_{\min }\right)\left(b-a\right)}{X_{\max }-X_{\min }}}$) with the parameters:
#
# $X_{\min }=0$
#
# $X_{\max }=255$
#
# $a=0.1$
#
# $b=0.9$
# Problem 1 - Implement Min-Max scaling for grayscale image data
def normalize_grayscale(image_data):
"""
Normalize the image data with Min-Max scaling to a range of [0.1, 0.9]
:param image_data: The image data to be normalized
:return: Normalized image data
"""
a = 0.1
b = 0.9
grayscale_min = 0
grayscale_max = 255
return a + ( ( (image_data - grayscale_min)*(b - a) )/( grayscale_max - grayscale_min ) )
# ## Problem 2
# - Use [tf.placeholder()](https://www.tensorflow.org/api_docs/python/io_ops.html#placeholder) for `features` and `labels` since they are the inputs to the model.
# - Any math operations must have the same type on both sides of the operator. The weights are float32, so the `features` and `labels` must also be float32.
# - Use [tf.Variable()](https://www.tensorflow.org/api_docs/python/state_ops.html#Variable) to allow `weights` and `biases` to be modified.
# - The `weights` must be the dimensions of features by labels. The number of features is the size of the image, 28*28=784. The size of labels is 10.
# - The `biases` must be the dimensions of the labels, which is 10.
# +
features_count = 784
labels_count = 10
# Problem 2 - Set the features and labels tensors
features = tf.placeholder(tf.float32)
labels = tf.placeholder(tf.float32)
# Problem 2 - Set the weights and biases tensors
weights = tf.Variable(tf.truncated_normal((features_count, labels_count)))
biases = tf.Variable(tf.zeros(labels_count))
# -
# # Problem 3
# Configuration 1
# * **Epochs:** 1
# * **Batch Size:** 50
# * **Learning Rate:** 0.01
#
# Configuration 2
# * **Epochs:** 1
# * **Batch Size:** 100
# * **Learning Rate:** 0.1
#
# Configuration 3
# * **Epochs:** 4 or 5
# * **Batch Size:** 100
# * **Learning Rate:** 0.2
| Code/TF-NN-Lab/Solution_TF_NN_Lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Evaluation methods in NLP
# -
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Spring 2022"
# + [markdown] slideshow={"slide_type": "slide"}
# ## Contents
#
# 1. [Overview](#Overview)
# 1. [Your projects](#Your-projects)
# 1. [Set-up](#Set-up)
# 1. [Data organization](#Data-organization)
# 1. [Train/dev/test](#Train/dev/test)
# 1. [No fixed splits](#No-fixed-splits)
# 1. [Cross-validation](#Cross-validation)
# 1. [Random splits](#Random-splits)
# 1. [K-folds](#K-folds)
# 1. [Baselines](#Baselines)
# 1. [Baselines are crucial for strong experiments](#Baselines-are-crucial-for-strong-experiments)
# 1. [Random baselines](#Random-baselines)
# 1. [Task-specific baselines](#Task-specific-baselines)
# 1. [Hyperparameter optimization](#Hyperparameter-optimization)
# 1. [Rationale](#Rationale)
# 1. [The ideal hyperparameter optimization setting](#The-ideal-hyperparameter-optimization-setting)
# 1. [Practical considerations, and some compromises](#Practical-considerations,-and-some-compromises)
# 1. [Hyperparameter optimization tools](#Hyperparameter-optimization-tools)
# 1. [Classifier comparison](#Classifier-comparison)
# 1. [Practical differences](#Practical-differences)
# 1. [Confidence intervals](#Confidence-intervals)
# 1. [Wilcoxon signed-rank test](#Wilcoxon-signed-rank-test)
# 1. [McNemar's test](#McNemar's-test)
# 1. [Assessing models without convergence](#Assessing-models-without-convergence)
# 1. [Incremental dev set testing](#Incremental-dev-set-testing)
# 1. [Learning curves with confidence intervals](#Learning-curves-with-confidence-intervals)
# 1. [The role of random parameter initialization](#The-role-of-random-parameter-initialization)
# 1. [Closing remarks](#Closing-remarks)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Overview
#
# This notebook is an overview of experimental methods for NLU. My primary goal is to help you with the experiments you'll be doing for your projects. It is a companion to [the evaluation metrics notebook](evaluation_metrics.ipynb), which I suggest studying first.
#
# The teaching team will be paying special attention to how you conduct your evaluations, so this notebook should create common ground around what our values are.
#
# This notebook is far from comprehensive. I hope it covers the most common tools, techniques, and challenges in the field. Beyond that, I'm hoping the examples here suggest a perspective on experiments and evaluations that generalizes to other topics and techniques.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Your projects
#
# 1. We will never evaluate a project based on how "good" the results are.
# 1. Publication venues do this, because they have additional constraints on space that lead them to favor positive evidence for new developments over negative results.
# 1. In CS224u, we are not subject to this constraint, so we can do the right and good thing of valuing positive results, negative results, and everything in between.
#
# 1. We __will__ evaluate your project on:
# 1. The appropriateness of the metrics
# 1. The strength of the methods
# 1. The extent to which the paper is open and clear-sighted about the limits of its findings.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Set-up
# -
# %matplotlib inline
from collections import defaultdict, Counter
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
from scipy import stats
from sklearn.datasets import make_classification
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
import sst
from torch_rnn_classifier import TorchRNNClassifier
from torch_shallow_neural_classifier import TorchShallowNeuralClassifier
import utils
utils.fix_random_seeds()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Data organization
# + [markdown] slideshow={"slide_type": "slide"}
# ### Train/dev/test
#
# Many publicly available datasets are released with a train/dev/test structure. __We're all on the honor system to do test-set runs only when development is complete.__
#
# Splits like this basically presuppose a fairly large dataset.
#
# If there is no dev set as part of the distribution, then you might create one to simulate what a test run will be like, though you have to weigh this against the reduction in train-set size.
#
# Having a fixed test set ensures that all systems are assessed against the same gold data. This is generally good, but it is problematic where the test set turns out to have unusual properties that distort progress on the task. Ideally, every task would have dozens of test sets, so that we could report average performance and related statistics. The difficulty and expense of creating so many test sets means that this ideal is rarely if ever realized.
# + [markdown] slideshow={"slide_type": "slide"}
# ### No fixed splits
#
# Many datasets are released without predefined splits. This poses challenges for assessment, especially comparative assessment: __for robust comparisons with prior work, you really have to rerun the models using your assessment regime on your splits__. For example, if you're doing [5-fold cross-validation](#K-folds), then all the systems should be trained and assessed using exactly the same folds, to control for variation in how difficult the splits are.
#
# If the dataset is large enough, you might create a train/test or train/dev/test split right at the start of your project and use it for all your experiments. This means putting the test portion in a locked box until the very end, when you assess all the relevant systems against it. For large datasets, this will certainly simplify your experimental set-up, for reasons that will become clear when we discuss [hyperparameter optimization](#Hyperparameter-optimization) below.
#
# For small datasets, carving out dev and test sets might leave you with too little data. The most problematic symptom of this is that performance is highly variable because there isn't enough data to optimize reliably. In such situations, you might give up on having fixed splits, opting instead for some form of cross-validation, which allows you to average over multiple runs.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Cross-validation
#
# In cross-validation, we take a set of examples $X$ and partition them into two or more train/test splits, and then we average over the results in some way.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Random splits
#
# When creating random train/test splits, we shuffle the examples and split them, with a pre-specified percentage $t$ used for training and another pre-specified percentage (usually $1-t$) used for testing.
#
# In general, we want these splits to be __stratified__ in the sense that the train and test splits have approximately the same distribution over the classes.
#
# #### The good and the bad of random splits
#
# A nice thing about random splits is that you can create as many as you want without having this impact the ratio of training to testing examples.
#
# This can also be a liability, though, as there's no guarantee that every example will be used the same number of times for training and testing. In principle, one might even evaluate on the same split more than once (though this will be fantastically unlikely for large datasets). NLP datasets are generally large enough that this isn't a pressing concern.
#
# The function `utils.fit_classifier_with_hyperparameter_search` hard-codes a strategy of using random splits by using the [StratifiedShuffleSplit](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.StratifiedShuffleSplit.html#sklearn.model_selection.StratifiedShuffleSplit) utility. The benefit of decoupling the train/test ratio from the number of splits outweights the concerns about split composition.
#
# #### Random splits in scikit-learn
#
# In scikit-learn, the function [train_test_split](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) will do random splits. It is a wrapper around [ShuffleSplit](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.ShuffleSplit.html#sklearn.model_selection.ShuffleSplit) or [StratifiedShuffleSplit](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.StratifiedShuffleSplit.html#sklearn.model_selection.StratifiedShuffleSplit), depending on how the keyword argument `stratify` is used. A potential gotcha for classification problems: `train_test_split` does not stratify its splits by default, whereas stratified splits are desired in most situations.
# + [markdown] slideshow={"slide_type": "slide"}
# ### K-folds
#
# In K-fold cross-validation, one divides the data into $k$ folds of equal size and then conducts $k$ experiments. In each, fold $i$ is used for assessment, and all the other folds are merged together for training:
#
# $$
# \begin{array}{c c c }
# \textbf{Splits} & \textbf{Experiment 1} & \textbf{Experiment 2} & \textbf{Experiment 3} \\
# \begin{array}{|c|}
# \hline
# \textrm{fold } 1 \\\hline
# \textrm{fold } 2 \\\hline
# \textrm{fold } 3 \\\hline
# \end{array}
# &
# \begin{array}{|c c|}
# \hline
# \textbf{Test} & \textrm{fold } 1 \\\hline
# \textbf{Train} & \textrm{fold } 2 \\
# & \textrm{fold } 3 \\\hline
# \end{array}
# &
# \begin{array}{|c c|}
# \hline
# \textbf{Test} & \textrm{fold } 2 \\\hline
# \textbf{Train} & \textrm{fold } 1 \\
# & \textrm{fold } 3 \\\hline
# \end{array}
# &
# \begin{array}{|c c|}
# \hline
# \textbf{Test} & \textrm{fold } 3 \\\hline
# \textbf{Train} & \textrm{fold } 1 \\
# & \textrm{fold } 2 \\\hline
# \end{array}
# \end{array}
# $$
#
# #### The good and the bad of k-folds
#
# * With k-folds, every example appears in a train set exactly $k-1$ times and in a test set exactly once. We noted above that random splits do not guarantee this.
#
# * A major drawback of k-folds is that the size of $k$ determines the size of the train/test splits. With 3-fold cross validation, one trains on 67% of the data and tests on 33%. With 10-fold cross-validation, one trains on 90% and tests on 10%. These are likely to be __very__ different experimental scenarios. This is a consideration one should have in mind when [comparing models](#Classifier-comparison) using statistical tests that depend on repeated runs. This is a large enough drawback to k-folds that I generally favor random splits, [as discussed just above](#Random-splits).
# + [markdown] slideshow={"slide_type": "slide"}
# #### K-folds in scikit-learn
#
# * In scikit-learn, [KFold](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.KFold.html#sklearn.model_selection.KFold) and [StratifiedKFold](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.StratifiedKFold.html#sklearn.model_selection.StratifiedKFold) are the primary classes for creating k-folds from a dataset. As with random splits, the stratified option is recommended for most classification problems, as one generally want to train and assess with the same label distribution.
#
# * The methods [cross_validate](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_validate.html#sklearn.model_selection.cross_validate) and [cross_val_score](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html#sklearn.model_selection.cross_val_score) are convenience methods that let you pass in a model (`estimator`), a dataset (`X` and `y`), and some cross-validation parameters, and they handle the repeated assessments. These are great. Two tips:
# * I strongly recommend passing in a `KFold` or `StratifiedKFold` instance as the value of `cv` to ensure that you get the split behavior that you desire.
# * Check that `scoring` has the value that you desire. For example, if you are going to report F1-scores, it's a mistake to leave `scoring=None`, as this will default to whatever your model reports with its `score` method, which is probably accuracy.
# + [markdown] slideshow={"slide_type": "slide"}
# #### Variants
#
# K-folds has a number of variants and special cases. Two that frequently arise in NLU:
#
# 1. [LeaveOneOut](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.LeaveOneOut.html#sklearn.model_selection.LeaveOneOut) is the special case where the number of folds equals the number of examples. This is especially useful for very small datasets.
#
# 1. [LeavePGroupsOut](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.LeavePGroupsOut.html#sklearn.model_selection.LeavePGroupsOut) creates folds based on criteria that you define. This is useful in situations where the datasets have important structure that the splits need to respect – e.g., you want to assess against a graph sub-network that is never seen on training.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Baselines
#
# Evaluation numbers in NLP (and throughout AI) __can never be understood properly in isolation__:
#
# * If your system gets 0.95 F1, that might seem great in absolute terms, but your readers will suspect the task is too easy and want to know what simple models achieve.
#
# * If your system gets 0.60 F1, you might despair, but it could turn out that humans achieve only 0.80, indicating that you got traction on a very challenging but basically coherent problem.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Baselines are crucial for strong experiments
#
# Defining baselines should not be an afterthought, but rather central to how you define your overall hypotheses. __Baselines are essential to building a persuasive case__, and they can also be used to illuminate specific aspects of the problem and specific virtues of your proposed system.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Random baselines
#
# Random baselines are almost always useful to include. scikit-learn has classes [DummyClassifier](http://scikit-learn.org/stable/modules/generated/sklearn.dummy.DummyClassifier.html#sklearn.dummy.DummyClassifier) and [DummyRegressor](http://scikit-learn.org/stable/modules/generated/sklearn.dummy.DummyRegressor.html#sklearn.dummy.DummyRegressor). Each of them has a keyword argument `strategy` that allows you to specify a range of different styles of random guessing. I highly recommend using these in your work, for two central reasons:
#
# 1. They will probably fit into your overall modeling pipeline.
# 2. It's usual conceptually easy to describe these baselines but it can be tricky and error-prone to implement them – and the scikit-learn folks probably already did it for you flawlessly!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Task-specific baselines
#
# It is worth considering whether your problem suggests a baseline that will reveal something about the problem or the ways it is modeled. Two recent examples from NLU:
#
# 1. As disussed briefly in [the NLI models notebook](nli_02_models.ipynb#Other-findings), [<NAME>](https://leonidk.com/) observed [in his 2016 NLU course project](https://leonidk.com/stanford/cs224u.html) that one can do much better than chance on SNLI by processing only the hypothesis, ignoring the premise entirely. The exact interpretation of this is complex (we explore this a bit [in our NLI unit](nli_02_models.ipynb#Hypothesis-only-baselines) and [in our NLI bake-off](nli_wordentail.ipynb)), but it's certainly relevant for understanding how much a system has actually learned about reasoning from a premise to a conclusion.
#
# 1. [Schwartz et al. (2017)](https://www.aclweb.org/anthology/W17-0907) develop a system for choosing between a coherent and incoherent ending for a story. Their best system achieves 75% accuracy by processing the story and the ending, but they achieve 72% using only stylistic features of the ending, ignoring the preceding story entirely. This puts the 75% – and the extent to which the system understands story completion – in a new light.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Hyperparameter optimization
#
# In machine learning, the __parameters__ of a model are those whose values are learned as part of optimizing the model itself.
#
# The __hyperparameters__ of a model are any settings that are set by a process that is outside of this optimization process. The boundary between a true setting of the model and a broader design choice will likely be blurry conceptually. For example:
#
# * The regularization term for a classifier is a clear hyperparameter – it appears in the model's objective function.
# * What about the method one uses for normalizing the feature values? This is probably not a setting of the model per se, but rather a choice point in your experimental framework.
#
# For the purposes of this discussion, we'll construe hyperparameters very broadly.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Rationale
#
# Hyperparameter optimization is one of the most important parts machine learning, and a crucial part of building a persuasive argument. For one angle on why, it's helpful to imagine that you're in an ongoing debate with a very skeptical referee:
#
# 1. You ran experiments with models A, B, and C. For each, you used the default hyperparameters as given by the implementations you're using. You found that C performed the best, and so you reported that in your paper.
# 1. Your reviewer doesn't have visibility into your process, and maybe doesn't fully trust you. Did you try any other values for the hyperparameters without reporting that? If not, would you have done that if C hadn't outperformed the others? There is no way for the reviewer (or perhaps anyone) to answer these questions.
# 1. So, from the reviewer's perspective, all we learned from your experiments is that there is some set of hyperparameters on which C wins this competition. But, strictly speaking, this conveys no new information; we knew before you did your experiments that we could find settings that would deliver this and all other outcomes. (They might not be __sensible__ settings, but remember you're dealing with a hard-bitten, unwavering skeptic.)
#
# Our best response to this situation is to allow these models to explore a wide range of hyperparameters, choose the best ones according to performance on training or development data, and then report how they do with those settings at test time. __This gives every model its best chance to succeed.__
#
# If you do this, the strongest argument that your skeptical reviewer can muster is that you didn't pick the right space of hyperparameters to explore for one or more of the models. Alas, there is no satisfying the skeptic, but we can at least feel happy that the outcome of these experiments will have a lot more scientific value than the ones described above with fixed hyperparameters.
# + [markdown] slideshow={"slide_type": "slide"}
# ### The ideal hyperparameter optimization setting
#
# When evaluating a model, the ideal regime for hyperparameter optimization is as follows:
#
# 1. For each hyperparameter, identify a large set of values for it.
# 2. Create a list of all the combinations of all the hyperparameter values. This will be the [cross-product](https://en.wikipedia.org/wiki/Cartesian_product) of all the values for all the features identified at step 1.
# 3. For each of the settings, cross-validate it on the available training data.
# 4. Choose the settings that did best in step 3, train on all the training data using those settings, and then evaluate that model on the test set.
#
# This is very demanding. First, The number of settings grows quickly with the number of hyperparameters and values. If hyperparameter $h_{1}$ has $5$ values and hyperparameter $h_{2}$ has $10$, then the number of settings is $5 \cdot 10 = 50$. If we add a third hyperparameter $h_{3}$ with just $2$ values, then the number jumps to $100$. Second, if you're doing 5-fold cross-validation, then each model is trained 5 times. You're thus committed to training $500$ models.
#
# And it could get worse. Suppose you don't have a fixed train/test split, and you're instead reporting, say, the result of 10 random train/test splits. Strictly speaking, the optimal hyperparameters could be different for different splits. Thus, for each split, the above cross-validation should be conducted. Now you're committed to training $5,000$ systems!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Practical considerations, and some compromises
#
# The above is untenable as a set of laws for the scientific community. If we adopted it, then complex models trained on large datasets would end up disfavored, and only the very wealthy would be able to participate. Here are some pragmatic steps you can take to alleviate this problem, in descending order of attractiveness. (That is, the lower you go on this list, the more likely the skeptic is to complain!)
#
# 1. [Bergstra and Bengio (2012)](http://www.jmlr.org/papers/v13/bergstra12a.html) argue that __randomly sampling__ from the space of hyperparameters delivers results like the full "grid search" described above with a relatively few number of samples. __Hyperparameter optimization algorithms__ like those implemented in [Hyperopt](http://hyperopt.github.io/hyperopt/) and [scikit-optimize](https://github.com/scikit-optimize/scikit-optimize) allow guided sampling from the full space. All these methods control the exponential growth in settings that comes from any serious look at one's hyperparameters.
#
# 1. In large deep learning systems, __the hyperparameter search could be done on the basis of just a few iterations__. The systems likely won't have converged, but it's a solid working assumption that early performance is highly predictive of final performance. You might even be able to justify this with learning curves over these initial iterations.
#
# 1. Not all hyperparameters will contribute equally to outcomes. Via heuristic exploration, it is typically possible to __identify the less informative ones and set them by hand__. As long as this is justified in the paper, it shouldn't rile the skeptic too much.
#
# 1. Where repeated train/test splits are being run, one might __find optimal hyperparameters via a single split__ and use them for all the subsequent splits. This is justified if the splits are very similar.
#
# 1. In the worst case, one might have to adopt hyperparameters that were optimal for other experiments that have been published. The skeptic will complain that these findings don't translate to your new data sets. That's true, but it could be the only option. For example, how would one compare against [Rajkomar et al. (2018)](https://arxiv.org/abs/1801.07860) who report that "the performance of all above neural networks were [sic] tuned automatically using Google Vizier [35] with a total of >201,000 GPU hours"?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Hyperparameter optimization tools
#
# * scikit-learn's [model_selection](http://scikit-learn.org/stable/modules/classes.html#module-sklearn.model_selection) package has classes `GridSearchCV` and `RandomizedSearchCV`. These are very easy to use. (We used `GridSearchCV` in our course code, in `utils.fit_classifier_with_hyperparameter_search`.)
#
# * [scikit-optimize](https://github.com/scikit-optimize/scikit-optimize) offers a variety of methods for guided search through the grid of hyperparameters. [This post](https://roamanalytics.com/2016/09/15/optimizing-the-hyperparameter-of-which-hyperparameter-optimizer-to-use/) assesses these methods against grid search and fully randomized search, and it also provides [starter code](https://github.com/roamanalytics/roamresearch/tree/master/BlogPosts/Hyperparameter_tuning_comparison) for using these implementations with sklearn-style classifiers.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Classifier comparison
#
# Suppose you've assessed two classifier models. Their performance is probably different to some degree. What can be done to establish whether these models are different in any meaningful sense?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Practical differences
#
# One very simple step one can take is to simply count up how many examples the models actually differ on.
#
# * If the test set has 1,000 examples, then a difference of 1% in accuracy or F1 will correspond to roughly 10 examples. We'll likely have intuitions about whether that difference has any practical import.
#
# * If the test set has 1M examples, then 1% will correspond to 10,000 examples, which seems sure to matter. Unless other considerations (e.g., cost, understandability) favor the less accurate model, the choice seems clear.
#
# Even where the numbers suggest a practical difference, we might still wonder whether the difference is stable across different runs, and thus we might still want to gather more information.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Confidence intervals
#
# If you can afford to run the model multiple times, then reporting confidence intervals based on the resulting scores could suffice to build an argument about whether the models are meaningfully different.
#
# The following will calculate a simple 95% confidence interval for a vector of scores `vals`:
# -
def get_ci(vals):
if len(set(vals)) == 1:
return (vals[0], vals[0])
loc = np.mean(vals)
scale = np.std(vals) / np.sqrt(len(vals))
return stats.t.interval(0.95, len(vals)-1, loc=loc, scale=scale)
# It's very likely that these confidence intervals will look very large relative to the variation that you actually observe. You probably can afford to do no more than 10–20 runs. Even if your model is performing very predictably over these runs (which it will, assuming your method for creating the splits is sound), the above intervals will be large in this situation. This might justify bootstrapping the confidence intervals. I recommend [scikits-bootstrap](https://github.com/cgevans/scikits-bootstrap) for this.
#
# __Important__: when evaluating multiple systems via repeated train/test splits or cross-validation, all the systems have to be run on the same splits. This is the only way to ensure that all the systems face the same challenges.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Wilcoxon signed-rank test
#
# NLPers always choose tables over plots for some reason, and confidence intervals are hard to display in tables. This might mean that you want to calculate a p-value.
#
# Where you can afford to run the models at least 10 times with different splits (and preferably more like 20), [Demšar (2006)](http://www.jmlr.org/papers/v7/demsar06a.html) recommends the [Wilcoxon signed-rank test](https://en.wikipedia.org/wiki/Wilcoxon_signed-rank_test). This is implemented in scipy as [scipy.stats.wilcoxon](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.wilcoxon.html). This test relies only on the absolute differences between scores for each split and makes no assumptions about how the scores are distributed.
#
# Take care not to confuse this with [scipy.stats.ranksums](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.ranksums.html), which does the Wilcoxon rank-sums test. This is also known as the [Mann–Whitney U test](https://en.wikipedia.org/wiki/Mann–Whitney_U_test), though SciPy distinguishes this as a separate test ([scipy.stats.mannwhitneyu](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.mannwhitneyu.html#scipy.stats.mannwhitneyu)). In any case, the heart of this is that the signed-rank variant is more appropriate for classifier assessments, where we are always comparing systems trained and assessed on the same underlying pool of data.
#
# Like all tests of this form, we should be aware of what they can tell us and what they can't:
#
# * The test says __nothing__ about the practical importance of any differences observed.
#
# * __Small p-values do not reliably indicate large effect sizes__. (A small p-value will more strongly reflect the number of samples you have.)
#
# * Large p-values simply mean that the available evidence doesn't support a conclusion that the systems are different, not that there is no difference in fact. And even that limited conclusion is only relative to this particular, quite conservative test.
#
# All this is to say that these values should not be asked to stand on their own, but rather presented as part of a larger, evidence-driven argument.
# + [markdown] slideshow={"slide_type": "slide"}
# ### McNemar's test
#
# [McNemar's test](https://en.wikipedia.org/wiki/McNemar%27s_test) operates directly on the vectors of predictions for the two models being compared. As such, it doesn't require repeated runs, which is good where optimization is expensive.
#
# The basis for the test is a contingency table with the following form, for two models A and B:
#
# $$\begin{array}{|c | c |}
# \hline
# \textrm{number of examples} & \textrm{number of examples} \\
# \textrm{where A and B are correct} & \textrm{where A is correct, B incorrect}
# \\\hline
# \textrm{number of examples} & \textrm{number of examples} \\
# \textrm{where A is correct, B incorrect} & \textrm{where both A and B are incorrect} \\\hline
# \end{array}$$
#
# Following [Dietterich (1998)](http://sci2s.ugr.es/keel/pdf/algorithm/articulo/dietterich1998.pdf), let the above be abbreviated to
#
# $$\begin{array}{|c | c |}
# \hline
# n_{11} & n_{10}
# \\\hline
# n_{01} & n_{00} \\
# \hline
# \end{array}$$
#
# The null hypothesis tested is that the two models have the same error rate, i.e., that $n_{01} = n_{10}$. The test statistic is
#
# $$
# \frac{
# \left(|(n_{01} - n_{10}| - 1\right)^{2}
# }{
# n_{01} + n_{10}
# }$$
#
# which has an approximately chi-squared distribution with 1 degree of freedom.
#
# An implementation is available in this repository: `utils.mcnemar`.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Assessing models without convergence
#
# When working with linear models, convergence issues rarely arise. Typically, the implementation has a fixed number of iterations it performs, or a threshold on the error, and the model stops when it reaches one of these points. We mostly don't reflect on this because of the speed and stability of these models.
#
# With neural networks, convergence takes center stage. The models rarely converge, or they converge at different rates between runs, and their performance on the test data is often heavily dependent on these differences. Sometimes a model with a low final error turns out to be great, and sometimes it turns out to be worse than one that finished with a higher error. Who knows?!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Incremental dev set testing
#
# The key to addressing this uncertainty is to __regularly collect information about dev set performance as part of training__. For example, after every epoch, one could make predictions on the dev set and store that vector of predictions, or just whatever assessment metric one is using. These assessments can provide direct information about how the model is doing on the actual task we care about, which will be a better indicator than the errors.
#
# All the PyTorch models for this course accept a keyword argument `early_stopping`. The behavior should closely resemble that of `sklearn.neural_network` models. If `early_stopping=True`, then part of the dataset given to the `fit` method is reserved for incremental testing. The amount can be controlled with `validation_fraction` (default: `0.10`). After every epoch, this data will be used to evaluate the model using its `score` method. The parameters of the best model are stored. If an improvement of more than `tol` (default: `1e-5`) isn't seen within `n_iter_no_change` steps (default: `10`), then optimization stops, and the parameters of the numerically best model seen are used as the final model.
#
# It's important to see just how different this dev set performance can be from the training loss. In particular, the training loss can continue to go down even as the model grows worse and worse in evaluations on held-out data. This is a common form of __over-fitting__.
# + [markdown] slideshow={"slide_type": "slide"}
# #### Dataset to illustrate
# -
SST_HOME = os.path.join('data', 'sentiment')
def unigrams_phi(text):
return Counter(text.lower().split())
train = sst.build_dataset(
sst.train_reader(SST_HOME),
phi=unigrams_phi)
dev = sst.build_dataset(
sst.dev_reader(SST_HOME),
phi=unigrams_phi,
vectorizer=train['vectorizer'])
# #### Model without early stopping
# + slideshow={"slide_type": "slide"}
mod_no_stopping = TorchShallowNeuralClassifier(
early_stopping=False)
# -
_ = mod_no_stopping.fit(train['X'], train['y'])
print(classification_report(dev['y'], mod_no_stopping.predict(dev['X']), digits=3))
# + [markdown] slideshow={"slide_type": "slide"}
# #### Model with early stopping
# -
mod_stopping = TorchShallowNeuralClassifier(
early_stopping=True,
n_iter_no_change=50)
_ = mod_stopping.fit(train['X'], train['y'])
print(classification_report(dev['y'], mod_stopping.predict(dev['X']), digits=3))
# + [markdown] slideshow={"slide_type": "slide"}
# #### Errors vs. incremental performance
# +
scores = mod_stopping.validation_scores
errors = mod_no_stopping.errors[: len(scores)]
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
ax1 = pd.Series(scores).plot(ax=ax1)
ax1.set_xlabel("Epochs")
ax1.set_ylabel("Macro F1 score")
ax2 = pd.Series(errors).plot(ax=ax2)
ax2.set_xlabel("Epochs")
_ = ax2.set_ylabel("Error")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Learning curves with confidence intervals
#
# I frankly think the best response to all this is to accept that incremental performance plots like the above are how we should be assessing our models. This exposes all of the variation that we actually observe.
#
# In addition, in deep learning, we're often dealing with classes of models that are in principle capable of learning anything. The real question is implicitly how efficiently they can learn given the available data and other resources. Learning curves bring this our very clearly.
#
# We can improve the curves by adding confidence intervals to them derived from repeated runs. Here's a plot from a paper I wrote with <NAME> ([Dingwall and Potts 2018](https://arxiv.org/abs/1803.09901)):
#
# <img src="fig/diagnosis-curve.png" />
#
# I think this shows very clearly that, once all is said and done, the Mittens model (red) learns faster than the others, but is indistinguishable from the Clinical text GloVe model (blue) after enough training time. Furthermore, it's clear that the other two models are never going to catch up in the current experimental setting. A lot of this information would be lost if, for example, we decided to stop training when dev set performance reached its peak and report only a single F1 score per class.
# + [markdown] slideshow={"slide_type": "slide"}
# ## The role of random parameter initialization
#
# Most deep learning models have their parameters initialized randomly, perhaps according to some heuristics related to the number of parameters ([Glorot and Bengio 2010](http://proceedings.mlr.press/v9/glorot10a.html)) or their internal structure ([Saxe et al. 2014](https://arxiv.org/abs/1312.6120)). This is meaningful largely because of the non-convex optimization problems that these models define, but it can impact simpler models that have multiple optimal solutions that still differ at test time.
#
# There is growing awareness that these random choices have serious consequences. For instance, [<NAME> (2017)](https://www.aclweb.org/anthology/D17-1035) report that different initializations for neural sequence models can lead to statistically significant results, and they show that a number of recent systems are indistinguishable in terms of raw performance once this source of variation is taken into account.
#
# This shouldn't surprise practitioners, who have long struggled with the question of what to do when a system experiences a catastrophic failure as a result of unlucky initialization. (I think the answer is to report this failure rate.)
#
# The code snippet below lets you experience this phenomenon for yourself. The XOR logic operator, which is true just in case its two arguments have the same value, is famously not learnable by a linear classifier but within reach of a neural network with a single hidden layer and a non-linear activation function ([Rumelhart et al. 1986](https://www.nature.com/articles/323533a0)). But how consistently do such models actually learn XOR? No matter what settings you choose, you rarely if ever see perfect performance across multiple runs.
# + slideshow={"slide_type": "slide"}
def xor_eval(n_trials=10):
xor = [
([1.,1.], 1),
([1.,0.], 0),
([0.,1.], 0),
([0.,0.], 1)]
X, y = zip(*xor)
results = defaultdict(int)
for trial in range(n_trials):
model = TorchShallowNeuralClassifier(
hidden_dim=2,
max_iter=500,
eta=0.01)
model.fit(X, y)
preds = tuple(model.predict(X))
result = 'correct' if preds == y else 'incorrect'
results[result] += 1
return results
xor_eval(n_trials=10)
# + [markdown] slideshow={"slide_type": "slide"}
# For better or worse, the only response we have to this situation is to __report scores for multiple complete runs of a model with different randomly chosen initializations__. [Confidence intervals](#Confidence-intervals) and [statistical tests](#Wilcoxon-signed-rank-test) can be used to summarize the variation observed. If the evaluation regime already involves comparing the results of multiple train/test splits, then ensuring a new random initializing for each of those would seem sufficient.
#
# Arguably, these observations are incompatible with evaluation regimes involving only a single train/test split, as in [McNemar's test](#McNemar's-test). However, [as discussed above](#Practical-considerations,-and-some-compromises), we have to be realistic. If multiple run aren't feasible, then a more heuristic argument will be needed to try to convince skeptics that the differences observed are larger than we would expect from just different random initializations.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Closing remarks
#
# We can summarize most of the above with a few key ideas:
#
# 1. Your evaluation should be based around a few systems that are related in ways that illuminate your hypotheses and help to convey what the best models are learning.
#
# 1. Every model you assess should be given its best chance to shine (but we need to be realistic about how many experiments this entails!).
#
# 1. The test set should play no role whatsoever in optimization or model selection. The best way to ensure this is to have the test set locked away until the final batch of experiments that will be reported in the paper, but this separation is simulated adequately by careful cross-validation set-ups.
#
# 1. Strive to base your model comparisons in multiple runs on the same splits. This is especially important for deep learning, where a single model can perform in very different ways on the same data, depending on the vagaries of optimization.
| evaluation_methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Q) How to add a particular attribute to each and every object of a class?
#
# ### Ans) Using init()
# +
class Student:
def __init__(self): #self - the object we're creating
self.name = input()
self.rollNumber = input()
# -
s1 = Student()
s1.__dict__
s2 = Student()
s2.__dict__
# ### Aliter: Set the name and roll number in the init() itself
s3 = Student("Hasan", 7)
# ### 3 arguments because init has self and we're passing 2 all sums up to 3 arguments
class Student:
def __init__(self, name, rollNumber): #self - the object we're creating
self.name = name
self.rollNumber = rollNumber
s3 = Student("Hasan", 7)
s3.__dict__
# +
#Predict the output
class Student:
def __init__(self,name,age):
self.name = "Rohan"
self.age = 20
def print_student_details():
print(self.name, end= " ")
print(self.age)
s = Student()
s.print_student_details()
# -
| 03 OOPS-1/3.04 Init Method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="0KrR-Qd_wXf-"
# # Introduction
# + [markdown] id="I6e2m4XsuRoL"
# Since Jan. 1, 2015, [The Washington Post](https://www.washingtonpost.com/) has been compiling a database of every fatal shooting in the US by a police officer in the line of duty.
#
# <center><img src=https://i.imgur.com/sX3K62b.png></center>
#
# While there are many challenges regarding data collection and reporting, The Washington Post has been tracking more than a dozen details about each killing. This includes the race, age and gender of the deceased, whether the person was armed, and whether the victim was experiencing a mental-health crisis. The Washington Post has gathered this supplemental information from law enforcement websites, local new reports, social media, and by monitoring independent databases such as "Killed by police" and "Fatal Encounters". The Post has also conducted additional reporting in many cases.
#
# There are 4 additional datasets: US census data on poverty rate, high school graduation rate, median household income, and racial demographics. [Source of census data](https://factfinder.census.gov/faces/nav/jsf/pages/community_facts.xhtml).
# + [markdown] id="JwGJl3mnw2Af"
# ### Upgrade Plotly
#
# Run the cell below if you are working with Google Colab
# + id="KvA9PBoRwvQG" outputId="ca6ee2a1-8da5-4f2d-f2b2-ea019cd92cae" colab={"base_uri": "https://localhost:8080/"}
# %pip install --upgrade plotly
# + [markdown] id="jqXsMLNJxELp"
# ## Import Statements
# + id="TPhEXC2FxGSr"
import numpy as np
import pandas as pd
import plotly.express as px
import matplotlib.pyplot as plt
import seaborn as sns
# This might be helpful:
from collections import Counter
# + [markdown] id="SwKGmmaLxS2s"
# ## Notebook Presentation
# + id="ayQQqhKtxBxj"
pd.options.display.float_format = '{:,.2f}'.format
# + [markdown] id="nKxJMOiMyAUF"
# ## Load the Data
# + _cell_guid="abb0685f-f844-43f8-84c0-4cd54361c7bf" _uuid="9c06ed7c41db79ae9d940f2d52aeff23621b3037" id="8KmGeg6ouRoP"
df_hh_income = pd.read_csv('Median_Household_Income_2015.csv', encoding="windows-1252")
df_pct_poverty = pd.read_csv('Pct_People_Below_Poverty_Level.csv', encoding="windows-1252")
df_pct_completed_hs = pd.read_csv('Pct_Over_25_Completed_High_School.csv', encoding="windows-1252")
df_share_race_city = pd.read_csv('Share_of_Race_By_City.csv', encoding="windows-1252")
df_fatalities = pd.read_csv('Deaths_by_Police_US.csv', encoding="windows-1252")
# + [markdown] id="6RuA5SvUyEaB"
# # Preliminary Data Exploration
#
# * What is the shape of the DataFrames?
# * How many rows and columns do they have?
# * What are the column names?
# * Are there any NaN values or duplicates?
# + id="3U9RcS1WyrRJ"
# + id="DmIARadGyrgs"
# + id="OSgTJoX7yrcS"
# + [markdown] id="p72TfMvayr2C"
# ## Data Cleaning - Check for Missing Values and Duplicates
#
# Consider how to deal with the NaN values. Perhaps substituting 0 is appropriate.
# + id="WEZuPaXvyxxp"
# + id="550aSczryx01"
# + [markdown] _uuid="52b308c936f5835c5cd186497fe3f96e96372483" id="n3DrcXVQuRoZ"
# # Chart the Poverty Rate in each US State
#
# Create a bar chart that ranks the poverty rate from highest to lowest by US state. Which state has the highest poverty rate? Which state has the lowest poverty rate? Bar Plot
# + id="mGxEzMDqzqmn"
# + id="B3AM6oJQ1D_P"
# + [markdown] id="GhUibVz5z4sn"
# # Chart the High School Graduation Rate by US State
#
# Show the High School Graduation Rate in ascending order of US States. Which state has the lowest high school graduation rate? Which state has the highest?
# + id="laD6W9eG0JFR"
# + [markdown] id="ENGTEmPQ0Rvt"
# # Visualise the Relationship between Poverty Rates and High School Graduation Rates
#
# #### Create a line chart with two y-axes to show if the rations of poverty and high school graduation move together.
# + id="MkNk5U8v0JPB"
# + id="urRMqQG_0Nmo"
# + [markdown] id="Zavx_8PL1Jy9"
# #### Now use a Seaborn .jointplot() with a Kernel Density Estimate (KDE) and/or scatter plot to visualise the same relationship
# + id="u77CHfdX0pjW"
# + id="CYy89o-u1zu0"
# + [markdown] id="PXNFQmHl10Fx"
# #### Seaborn's `.lmplot()` or `.regplot()` to show a linear regression between the poverty ratio and the high school graduation ratio.
# + id="F1xrjFgT1znI"
# + [markdown] id="QJTbZZKg0p6i"
# # Create a Bar Chart with Subsections Showing the Racial Makeup of Each US State
#
# Visualise the share of the white, black, hispanic, asian and native american population in each US State using a bar chart with sub sections.
# + id="YQbXb6Ex09D9"
# + id="LqrimlLo09Na"
# + [markdown] _uuid="77493cfd86a031612be8fb8d60f27713e74e6d80" id="K7xvQXkjuRo7"
# # Create Donut Chart by of People Killed by Race
#
# Hint: Use `.value_counts()`
# + id="1nOafPdy1u9Z"
# + id="YN3hzJTe1vHJ"
# + [markdown] id="kr-8Rezn3Bep"
# # Create a Chart Comparing the Total Number of Deaths of Men and Women
#
# Use `df_fatalities` to illustrate how many more men are killed compared to women.
# + id="joH1L4303BsJ"
# + id="X4MrMkVa3MTk"
# + [markdown] _uuid="cd1531c8421bfb6967cebf7039056779f9b960a6" id="ZxbBdvM0uRpQ"
# # Create a Box Plot Showing the Age and Manner of Death
#
# Break out the data by gender using `df_fatalities`. Is there a difference between men and women in the manner of death?
# + id="WdPPZLJY2shA"
# + id="tpPq4uFc2sql"
# + id="EnFbXlRm3XK6"
# + [markdown] id="h1pAgkyb3Xb0"
# # Were People Armed?
#
# In what percentage of police killings were people armed? Create chart that show what kind of weapon (if any) the deceased was carrying. How many of the people killed by police were armed with guns versus unarmed?
# + id="0d9B3Pec4TdM"
# + id="vdGTO3pW4Tp5"
# + id="qnO_rckl4WuJ"
# + [markdown] id="IgD2a5-L4W_I"
# # How Old Were the People Killed?
# + [markdown] id="BNPrdT-W9lf3"
# Work out what percentage of people killed were under 25 years old.
# + id="zSyDgzb542O9"
# + [markdown] id="5XrAD_ey9qBB"
# Create a histogram and KDE plot that shows the distribution of ages of the people killed by police.
# + id="U6B1Qp7l42H8"
# + [markdown] id="zcqqPDo79rTx"
# Create a seperate KDE plot for each race. Is there a difference between the distributions?
# + id="ubUnMjok43Zo"
# + [markdown] id="JkcHvMUt43uE"
# # Race of People Killed
#
# Create a chart that shows the total number of people killed by race.
# + id="e8wZvBrt5BEb"
# + id="_aqN0wC45BOl"
# + [markdown] id="SJaNm4rC5nWm"
# # Mental Illness and Police Killings
#
# What percentage of people killed by police have been diagnosed with a mental illness?
# + id="dbuGvCnN5ytC"
# + id="BsKDVLiY5CP_"
# + [markdown] id="R0WSPeKD5Cir"
# # In Which Cities Do the Most Police Killings Take Place?
#
# Create a chart ranking the top 10 cities with the most police killings. Which cities are the most dangerous?
# + id="NYVwo3qy5UV3"
# + id="T_FU4GFk5Upa"
# + [markdown] id="AqcOW6ca8JMD"
# # Rate of Death by Race
#
# Find the share of each race in the top 10 cities. Contrast this with the top 10 cities of police killings to work out the rate at which people are killed by race for each city.
# + id="FkjAUlKy8I2g"
# + id="dA3Hs9b55V59"
# + [markdown] id="1HUJksrI5WFb"
# # Create a Choropleth Map of Police Killings by US State
#
# Which states are the most dangerous? Compare your map with your previous chart. Are these the same states with high degrees of poverty?
# + id="u3UuX4Yo5dBA"
# + id="Y6VLPDeN5dKO"
# + [markdown] id="37UsmVYF6hA2"
# # Number of Police Killings Over Time
#
# Analyse the Number of Police Killings over Time. Is there a trend in the data?
# + id="nITf1IhP6gpI"
# + id="Q_vxjD_A7JRh"
# + id="mOqtie707JU8"
# + id="5fjhL8Lk7JKa"
# + [markdown] id="9o6uLFpS61nw"
# # Epilogue
#
# Now that you have analysed the data yourself, read [The Washington Post's analysis here](https://www.washingtonpost.com/graphics/investigations/police-shootings-database/).
# + id="8PjgzuNa61I1"
| day99/deaths-by-police/Fatal_Force.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="dyAvAsfq5OVZ" outputId="0af81ac3-bc84-4223-a369-05ae6a7286ec"
#Mounting Google Drive Locally
from google.colab import drive
drive.mount('/content/drive')
# + id="gzAMmDNqbgLS"
#set google drive path from personal google drive
import os
os.chdir('/content/drive/MyDrive/Colab Notebooks')
# + [markdown] id="7fjAWh20pMlH"
# **Importing the libraries**
# + id="GaHTw0BJ5cGL"
import tensorflow as tf
from keras.preprocessing.image import ImageDataGenerator
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="EmhJ7ANq5cJa" outputId="887fcb56-9f25-44b5-be48-f593b9adc12e"
#Specifying Tensorflow version
tf.__version__
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="gOm_8Co65cMB" outputId="cc40c1d9-399c-44ea-babe-ca04be0c19e7"
#testing gpu
import tensorflow as tf
tf.test.gpu_device_name()
#note:Without turning on GPU on google colab, training CNN will take forever.
#If your personal computer doesn't have gpu, is best to use google colab
# + [markdown] id="Ot82hmrFcM1c"
#
# **Data Preprocessing**
# + colab={"base_uri": "https://localhost:8080/"} id="qS5nfODH5cOi" outputId="2cdfcdc6-45f0-4a8a-9510-af9d82bd5e08"
#Inserting Training data, a bunch of pictures of dogs.
train_datagen = ImageDataGenerator(rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
training_set = train_datagen.flow_from_directory('/content/drive/MyDrive/Colab Notebooks/training_set',
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
# + colab={"base_uri": "https://localhost:8080/"} id="uGiKuWY25cRj" outputId="4239a99e-a584-459d-98b9-9db376ca3b77"
#the pretesting data, more pictures of dogs
test_datagen = ImageDataGenerator(rescale = 1./255)
test_set = test_datagen.flow_from_directory('/content/drive/MyDrive/Colab Notebooks/test_set',
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
# + [markdown] id="_BDp1vxpc-k-"
# # **Building CNN**
# + id="n2ff_7MS5cUR"
#inititialising CNN
cnn = tf.keras.models.Sequential()
# + id="11xNH2H05cZQ"
#Convolution using 3x3 matrix and relu activation
cnn.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, activation='relu', input_shape=[64, 64, 3]))
#Pooling with 2x2 kernel using 2 step stride
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
#adding second layer of convolution
cnn.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, activation='relu'))
#adding a second layer of pooling
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
#finish of with flattening (vectorising matreces)
cnn.add(tf.keras.layers.Flatten())
# + colab={"base_uri": "https://localhost:8080/"} id="HfaQyBwFeFla" outputId="c88d45cd-ab47-435d-a412-1f5ff2df5198"
#Full Conection
cnn.add(tf.keras.layers.Dense(units=128, activation='relu'))
#Output layer using sigmoid as activation, could use softmax
cnn.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
#model summary
cnn.summary()
# + [markdown] id="XLJ1BouKg0oU"
# ## **Training CNN**
# + id="iY6l55Rpin4T"
#Compiling CNN
cnn.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="4Jsg5M9HeFor" outputId="4dc8d502-9f87-41dc-a457-8ae37b210a58"
## Training or optimizing weights and bias using chaine rule including gradient of activation \
#as a result of output accuracy up to 30 epochs from training set evalution of the test set
cnn.fit(x = training_set, validation_data = test_set, epochs = 30)
# + [markdown] id="0zWMwbl0lTKA"
#
# + id="G5jV7zGLeFrp"
#Will test a picture of a cat
import numpy as np
from keras.preprocessing import image
test_image = image.load_img('/content/drive/MyDrive/Copy of single_prediction/cat_or_dog_2.jpg', target_size = (64, 64))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = cnn.predict(test_image)
training_set.class_indices
if result[0][0] == 1:
prediction = 'dog'
else:
prediction = 'cat'
# + colab={"base_uri": "https://localhost:8080/"} id="60X-S0mreF6e" outputId="958c1183-c5ee-457a-a229-84f6677019bc"
#at 92% accuracy, it should be able to predict what a cat is
print(prediction)
| Projects/CNN_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py35]
# language: python
# name: conda-env-py35-py
# ---
# ## French Wikipedia Heading Frequency
# This notebook serves to sort German Wikipedia section headers by frequency as related to this [research project](https://meta.wikimedia.org/wiki/Research:Investigate_frequency_of_section_titles_in_5_large_Wikipedias).
#
import numpy as np
import pandas as pd
# read in headers file by chunks of 100000 to conserve memory
# https://stackoverflow.com/questions/25962114/how-to-read-a-6-gb-csv-file-with-pandas
tp = pd.read_csv('frwiki_20161101_headings_2.tsv', sep='\t', header=0, dtype={'page_id': np.int32, 'page_title': object, 'page_ns': np.int16, 'heading_level': np.int8, 'heading_text': object}, iterator=True, chunksize=100000)
# concatenate all rows into a pandas dataframe
fr_DF = pd.concat([chunk for chunk in tp])
fr_DF.head()
fr_DF.page_ns.unique()
# determine number of unique articles in this dataset
len(fr_DF['page_title'].unique())
# remove leading and trailing whitespace from heading_text column
fr_DF['heading_text'] = pd.core.strings.str_strip(fr_DF['heading_text'])
# groupby heading_text and count the number of unique page_titles each heading appears in
# sort in descending order
# this returns a pandas series object
article_count = fr_DF.groupby('heading_text')['page_title'].apply(lambda x: len(x.unique())).sort_values(ascending=False)
# turn pandas series object into pandas dataframe
fr_article_count_DF = pd.DataFrame({'section_title':article_count.index, 'number_of_articles':article_count.values})
# add a column for the percentage of articles that header appears in
fr_article_count_DF['article_percentage'] = (fr_article_count_DF['number_of_articles']/1809018)*100
# set pandas options to display 100 rows
# round percentage to 2 decimal places and show top 100 results
pd.options.display.max_rows = 100
fr_article_count_DF.round({'article_percentage': 2}).head(100)
| frwiki_heading_freq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import sympy as sym
import pydae.build as db
# https://ctms.engin.umich.edu/CTMS/index.php?aux=Extras_Leadlag
# +
params_dict = {'T_wo':5} # parameters with default values
u_ini_dict = {'u_wo':0.0} # input for the initialization problem
u_run_dict = {'u_wo':0.0} # input for the running problem, its value is updated
x_list = ['x_wo'] # dynamic states
y_ini_list = ['z_wo'] # algebraic states for the initialization problem
y_run_list = ['z_wo'] # algebraic for the running problem
sys_vars = {'params':params_dict,
'u_list':u_run_dict,
'x_list':x_list,
'y_list':y_run_list}
exec(db.sym_gen_str()) # exec to generate the required symbolic varables and constants
# -
dx_wo = (u_wo - x_wo)/T_wo
g_wo = (u_wo - x_wo) - z_wo
# +
sys = {'name':'washout',
'params_dict':params_dict,
'f_list':[dx_wo],
'g_list':[g_wo],
'x_list':[x_wo],
'y_ini_list':y_ini_list,
'y_run_list':y_run_list,
'u_run_dict':u_run_dict,
'u_ini_dict':u_ini_dict,
'h_dict':{'u_wo':u_wo}}
sys = db.system(sys)
db.sys2num(sys)
# -
| examples/ctrl/basic/washout_builder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8 - Tensorflow
# language: python
# name: azureml_py38_tensorflow
# ---
# # Run BERT-Large inference workload
# + language="bash"
#
# # Download datasets, checkpoints and pre-trained model
# rm -rf ~/TF/bert-large
# mkdir -p ~/TF/bert-large/SQuAD-1.1
# cd ~/TF/bert-large/SQuAD-1.1
# wget https://github.com/oap-project/oap-project.github.io/raw/master/resources/ai/bert/dev-v1.1.json
# wget https://github.com/oap-project/oap-project.github.io/raw/master/resources/ai/bert/evaluate-v1.1.py
# wget https://github.com/oap-project/oap-project.github.io/raw/master/resources/ai/bert/train-v1.1.json
#
# cd ~/TF/bert-large
# wget https://storage.googleapis.com/intel-optimized-tensorflow/models/v1_8/bert_large_checkpoints.zip
# unzip bert_large_checkpoints.zip
#
# cd ~/TF/bert-large
# wget https://storage.googleapis.com/bert_models/2019_05_30/wwm_uncased_L-24_H-1024_A-16.zip
# unzip wwm_uncased_L-24_H-1024_A-16.zip
# + language="bash"
#
# # BERT-Large Inference
# # Install necessary packages
# sudo apt-get install -y numactl
# # Create ckpt directory
# rm -rf ~/TF/bert-large/inference/*
# mkdir -p ~/TF/bert-large/inference/BERT-Large-output
# # Download IntelAI benchmark
# cd ~/TF/bert-large/inference
# wget https://github.com/IntelAI/models/archive/refs/tags/v1.8.1.zip
# unzip v1.8.1.zip
# wget https://github.com/oap-project/oap-tools/raw/master/integrations/ml/databricks/benchmark/IntelAI_models_bertlarge_inference_realtime_throughput.patch
# cd ./models-1.8.1/
# git apply ../IntelAI_models_bertlarge_inference_realtime_throughput.patch
# + language="bash"
#
#
# #Bert-Large Inference
# export BERT_LARGE_OUTPUT=~/TF/bert-large/inference/BERT-Large-output
# export SQUAD_DIR=~/TF/bert-large/SQuAD-1.1/
# export BERT_LARGE_DIR=~/TF/bert-large/
# export PYTHONPATH=~/TF/bert-large/inference/models-1.8.1/benchmarks/
# cd ~/TF/bert-large/inference/models-1.8.1/benchmarks/
#
# function run_inference_without_numabind() {
# /anaconda/envs/azureml_py38_tensorflow/bin/python launch_benchmark.py \
# --model-name=bert_large \
# --precision=fp32 \
# --mode=inference \
# --framework=tensorflow \
# --batch-size=32 \
# --data-location $BERT_LARGE_DIR/wwm_uncased_L-24_H-1024_A-16 \
# --checkpoint $BERT_LARGE_DIR/bert_large_checkpoints \
# --output-dir $BERT_LARGE_OUTPUT/bert-squad-output \
# --verbose \
# --infer_option=SQuAD \
# DEBIAN_FRONTEND=noninteractive \
# predict_file=$SQUAD_DIR/dev-v1.1.json \
# experimental-gelu=False \
# init_checkpoint=model.ckpt-3649
# }
#
#
# function run_inference_with_numabind() {
# nohup /anaconda/envs/azureml_py38_tensorflow/bin/python launch_benchmark.py \
# --model-name=bert_large \
# --precision=fp32 \
# --mode=inference \
# --framework=tensorflow \
# --batch-size=32 \
# --socket-id 0 \
# --data-location $BERT_LARGE_DIR/wwm_uncased_L-24_H-1024_A-16 \
# --checkpoint $BERT_LARGE_DIR/bert_large_checkpoints \
# --output-dir $BERT_LARGE_OUTPUT/bert-squad-output \
# --verbose \
# --infer_option=SQuAD \
# DEBIAN_FRONTEND=noninteractive \
# predict_file=$SQUAD_DIR/dev-v1.1.json \
# experimental-gelu=False \
# init_checkpoint=model.ckpt-3649 >> socket0-inference-log &
#
# nohup /anaconda/envs/azureml_py38_tensorflow/bin/python launch_benchmark.py \
# --model-name=bert_large \
# --precision=fp32 \
# --mode=inference \
# --framework=tensorflow \
# --batch-size=32 \
# --socket-id 1 \
# --data-location $BERT_LARGE_DIR/wwm_uncased_L-24_H-1024_A-16 \
# --checkpoint $BERT_LARGE_DIR/bert_large_checkpoints \
# --output-dir $BERT_LARGE_OUTPUT/bert-squad-output \
# --verbose \
# -- infer_option=SQuAD \
# DEBIAN_FRONTEND=noninteractive \
# predict_file=$SQUAD_DIR/dev-v1.1.json \
# experimental-gelu=False \
# init_checkpoint=model.ckpt-3649 >> socket1-inference-log &
# }
#
# numa_nodes=$(lscpu | awk '/^NUMA node\(s\)/{ print $3 }')
#
# if [ "$numa_nodes" = "1" ];then
# run_inference_without_numabind
# else
# run_inference_with_numabind
# fi
# + language="bash"
#
# # Get the inference result
# numa_nodes=$(lscpu | awk '/^NUMA node\(s\)/{ print $3 }')
# if [ "$numa_nodes" = "1" ];then
# cd ~/TF/bert-large/inference/BERT-Large-output/bert-squad-output/
# cat benchmark*.log | grep "throughput((num_processed_examples-threshod_examples)/Elapsedtime)"
# else
# cd ~/TF/bert-large/inference/models-1.8.1/benchmarks/
# cat socket*-log | grep "throughput((num_processed_examples-threshod_examples)/Elapsedtime)"
# fi
# +
# Print TensorFlow version, and check whether is intel-optimized
import tensorflow
print("tensorflow version: " + tensorflow.__version__)
from packaging import version
if (version.parse("2.5.0") <= version.parse(tensorflow.__version__)):
from tensorflow.python.util import _pywrap_util_port
print( _pywrap_util_port.IsMklEnabled())
else:
from tensorflow.python import _pywrap_util_port
print(_pywrap_util_port.IsMklEnabled())
| integrations/ml/azure/benchmark/benchmark_tensorflow_bertlarge_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="7SCDe05G1QSx" executionInfo={"status": "ok", "timestamp": 1620185578973, "user_tz": -330, "elapsed": 9152, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="79b0af8b-de7d-4b35-909c-66990c44fad4"
# !pip install vowpalwabbit
# + colab={"base_uri": "https://localhost:8080/"} id="njhsjoqcSq3_" executionInfo={"status": "ok", "timestamp": 1620231073879, "user_tz": -330, "elapsed": 2707, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="afcf7f99-6177-47d8-b259-e1a0cdf9675b"
# !zip -r app.zip /content/dash-sample-apps/apps/dash-clinical-analytics
# + id="U17IKmm51Sa_" executionInfo={"status": "ok", "timestamp": 1620186109593, "user_tz": -330, "elapsed": 1945, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
from vowpalwabbit import pyvw
import random
import matplotlib.pyplot as plt
import pandas as pd
from itertools import product
# + id="AFdEwthO1U6N" executionInfo={"status": "ok", "timestamp": 1620185581218, "user_tz": -330, "elapsed": 2191, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
USER_LIKED_ARTICLE = -1.0
USER_DISLIKED_ARTICLE = 0.0
# + id="Joz5em0y7kfi"
users = ['A','B','C']
items = ['Item1','Item2','Item3','Item4','Item5','Item6']
context1 = ['morning','evening']
context2 = ['summer','winter']
context = pd.DataFrame(list(product(users, context1, context2, items)), columns=['users', 'context1', 'context2', 'items'])
context['reward'] = 0
#user 1 likes Item 1 in morning, and Item 6 in summer
context.loc[(context.users=='A') & \
(context.context1=='morning') & \
(context['items']=='Item1'), \
'reward'] = 1
context.loc[(context.users=='A') & \
(context.context2=='summer') & \
(context['items']=='Item6'), \
'reward'] = 1
#user 2 likes Item 2 in winter, and Item 5 in summer morning
context.loc[(context.users=='B') & \
(context.context2=='winter') & \
(context['items']=='Item2'), \
'reward'] = 1
context.loc[(context.users=='B') & \
(context.context1=='morning') & \
(context.context2=='summer') & \
(context['items']=='Item5'), \
'reward'] = 1
#user 3 likes Item 2 in morning, Item 3 in evening, and item 4 in winter morning
context.loc[(context.users=='C') & \
(context.context1=='morning') & \
(context['items']=='Item2'), \
'reward'] = 1
context.loc[(context.users=='C') & \
(context.context1=='evening') & \
(context['items']=='Item3'), \
'reward'] = 1
context.loc[(context.users=='C') & \
(context.context1=='morning') & \
(context.context2=='winter') & \
(context['items']=='Item4'), \
'reward'] = 1
context['cost'] = context['reward']*-1
contextdf = context.copy()
# + id="_S9shNn-liOP" executionInfo={"status": "ok", "timestamp": 1620191384481, "user_tz": -330, "elapsed": 1733, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
users = ['A','B','C']
items = ['Item1','Item2','Item3','Item4','Item5','Item6']
context1 = ['morning','evening']
context2 = ['summer','winter']
context = pd.DataFrame(list(product(users, context1, context2, items)), columns=['users', 'context1', 'context2', 'items'])
context['reward'] = 0
#user 1 likes Item 1 in morning, and Item 6 in summer
context.loc[(context.users=='A') & \
(context.context1=='morning') & \
(context['items']=='Item1'), \
'reward'] = 1
context.loc[(context.users=='A') & \
(context.context2=='summer') & \
(context['items']=='Item6'), \
'reward'] = 1
#user 2 likes Item 2 in winter, and Item 5 in summer morning
context.loc[(context.users=='B') & \
(context.context2=='winter') & \
(context['items']=='Item2'), \
'reward'] = 1
context.loc[(context.users=='B') & \
(context.context1=='morning') & \
(context.context2=='summer') & \
(context['items']=='Item5'), \
'reward'] = 1
#user 3 likes Item 2 in morning, Item 3 in evening, and item 4 in winter morning
context.loc[(context.users=='C') & \
(context.context1=='morning') & \
(context['items']=='Item2'), \
'reward'] = 1
context.loc[(context.users=='C') & \
(context.context1=='evening') & \
(context['items']=='Item3'), \
'reward'] = 1
context.loc[(context.users=='C') & \
(context.context1=='morning') & \
(context.context2=='winter') & \
(context['items']=='Item4'), \
'reward'] = 1
context['cost'] = context['reward']*-1
contextdf = context.copy()
# + colab={"base_uri": "https://localhost:8080/"} id="rpgSzAFjv_Rh" executionInfo={"status": "ok", "timestamp": 1620191384483, "user_tz": -330, "elapsed": 938, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="29a5d4c3-92f6-4ee4-f56f-31752de3c4b0"
contextdf.cost.value_counts()
# + id="Z8Uuu0axyu-C"
# def get_cost(context,action):
# if context['user'] == "Tom":
# if context['time_of_day'] == "morning" and action == 'politics':
# return USER_LIKED_ARTICLE
# elif context['time_of_day'] == "afternoon" and action == 'music':
# return USER_LIKED_ARTICLE
# else:
# return USER_DISLIKED_ARTICLE
# elif context['user'] == "Anna":
# if context['time_of_day'] == "morning" and action == 'sports':
# return USER_LIKED_ARTICLE
# elif context['time_of_day'] == "afternoon" and action == 'politics':
# return USER_LIKED_ARTICLE
# else:
# return USER_DISLIKED_ARTICLE
# + id="QlOd83hd1Z3F" executionInfo={"status": "ok", "timestamp": 1620191846746, "user_tz": -330, "elapsed": 671, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def get_cost(context,action):
return contextdf.loc[(contextdf['users']==context['user']) & \
(contextdf.context1==context['context1']) & \
(contextdf.context2==context['context2']) & \
(contextdf['items']==action), \
'cost'].values[0]
# + colab={"base_uri": "https://localhost:8080/"} id="6P_HYhhW8UWO" executionInfo={"status": "ok", "timestamp": 1620191848461, "user_tz": -330, "elapsed": 1576, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="74299b34-3a40-4141-df03-76edf4b56edf"
get_cost({'user':'A','context1':'morning','context2':'summer'},'Item2')
# + id="ub-6-Y5P1bhd" executionInfo={"status": "ok", "timestamp": 1620156279124, "user_tz": -330, "elapsed": 1383, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
# # This function modifies (context, action, cost, probability) to VW friendly format
# def to_vw_example_format(context, actions, cb_label = None):
# if cb_label is not None:
# chosen_action, cost, prob = cb_label
# example_string = ""
# example_string += "shared |User user={} time_of_day={}\n".format(context["user"], context["time_of_day"])
# for action in actions:
# if cb_label is not None and action == chosen_action:
# example_string += "0:{}:{} ".format(cost, prob)
# example_string += "|Action article={} \n".format(action)
# #Strip the last newline
# return example_string[:-1]
# + colab={"base_uri": "https://localhost:8080/"} id="onZzvDqJ1uNW" executionInfo={"status": "ok", "timestamp": 1620156324184, "user_tz": -330, "elapsed": 863, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="ef1a60f4-5acc-4ef9-dfde-a1e9f1b64fde"
# context = {"user":"Tom","time_of_day":"morning"}
# actions = ["politics", "sports", "music", "food"]
# print(to_vw_example_format(context,actions))
# + id="pTCHTNFf3jRe" executionInfo={"status": "ok", "timestamp": 1620191461983, "user_tz": -330, "elapsed": 1313, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
# This function modifies (context, action, cost, probability) to VW friendly format
def to_vw_example_format(context, actions, cb_label = None):
if cb_label is not None:
chosen_action, cost, prob = cb_label
example_string = ""
example_string += "shared |User users={} context1={} context2={}\n".format(context["user"], context["context1"], context["context2"])
for action in actions:
if cb_label is not None and action == chosen_action:
example_string += "0:{}:{} ".format(cost, prob)
example_string += "|Action items={} \n".format(action)
#Strip the last newline
return example_string[:-1]
# + colab={"base_uri": "https://localhost:8080/"} id="k8dLifwF3eDJ" executionInfo={"status": "ok", "timestamp": 1620191461985, "user_tz": -330, "elapsed": 802, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="54172834-13fb-4e48-a228-daf9a5ec0680"
context = {"user":"A","context1":"morning","context2":"summer"}
print(to_vw_example_format(context,item))
# + id="dM_aaYyb5xe9" executionInfo={"status": "ok", "timestamp": 1620191478597, "user_tz": -330, "elapsed": 1590, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def sample_custom_pmf(pmf):
total = sum(pmf)
scale = 1 / total
pmf = [x * scale for x in pmf]
draw = random.random()
sum_prob = 0.0
for index, prob in enumerate(pmf):
sum_prob += prob
if(sum_prob > draw):
return index, prob
# + id="1xeAG23o2EGj" executionInfo={"status": "ok", "timestamp": 1620191478599, "user_tz": -330, "elapsed": 1413, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def get_action(vw, context, actions):
vw_text_example = to_vw_example_format(context,actions)
pmf = vw.predict(vw_text_example)
chosen_action_index, prob = sample_custom_pmf(pmf)
return actions[chosen_action_index], prob
# + id="VWsWZZWgjVZR"
# + id="az_BHiJ32EwX" executionInfo={"status": "ok", "timestamp": 1620191478600, "user_tz": -330, "elapsed": 1293, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def choose_user(users):
return random.choice(users)
def choose_context1(context1):
return random.choice(context1)
def choose_context2(context2):
return random.choice(context2)
# + id="XqWD3pqt2GTr" executionInfo={"status": "ok", "timestamp": 1620192002733, "user_tz": -330, "elapsed": 1076, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def run_simulation(vw, num_iterations, users, contexts1, contexts2, actions, cost_function, do_learn = True):
cost_sum = 0.
ctr = []
for i in range(1, num_iterations+1):
user = choose_user(users)
context1 = choose_context1(contexts1)
context2 = choose_context2(contexts2)
context = {'user': user, 'context1': context1, 'context2': context2}
# print(context)
action, prob = get_action(vw, context, actions)
# print(action, prob)
cost = cost_function(context, action)
# print(cost)
cost_sum += cost
if do_learn:
# 5. Inform VW of what happened so we can learn from it
vw_format = vw.parse(to_vw_example_format(context, actions, (action, cost, prob)),pyvw.vw.lContextualBandit)
# 6. Learn
vw.learn(vw_format)
# 7. Let VW know you're done with these objects
vw.finish_example(vw_format)
# We negate this so that on the plot instead of minimizing cost, we are maximizing reward
ctr.append(-1*cost_sum/i)
return ctr
# + id="fPDQ5H512KEj" executionInfo={"status": "ok", "timestamp": 1620192004222, "user_tz": -330, "elapsed": 813, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def plot_ctr(num_iterations, ctr):
plt.plot(range(1,num_iterations+1), ctr)
plt.xlabel('num_iterations', fontsize=14)
plt.ylabel('ctr', fontsize=14)
plt.ylim([0,1])
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="wPhRpLyo2MmF" executionInfo={"status": "ok", "timestamp": 1620192011595, "user_tz": -330, "elapsed": 8030, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="a5b04514-516a-4c4f-f05c-f6437d3b23de"
# Instantiate learner in VW
vw = pyvw.vw("--cb_explore_adf -q UA --quiet --epsilon 0.2")
num_iterations = 5000
ctr = run_simulation(vw, num_iterations, users, context1, context2, items, get_cost)
plot_ctr(num_iterations, ctr)
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="TL81VOqK2Tbq" executionInfo={"status": "ok", "timestamp": 1620192585543, "user_tz": -330, "elapsed": 7245, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="1171c8f1-11f3-41bc-ad97-a3e518693896"
# Instantiate learner in VW but without -q
vw = pyvw.vw("--cb_explore_adf --quiet --epsilon 0.2")
num_iterations = 5000
ctr = run_simulation(vw, num_iterations, users, context1, context2, items, get_cost)
plot_ctr(num_iterations, ctr)
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="2vU5Kozv2WDg" executionInfo={"status": "ok", "timestamp": 1620192610855, "user_tz": -330, "elapsed": 6551, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="3dae15ae-38e8-418b-dd1b-37e78ce6f828"
# Instantiate learner in VW
vw = pyvw.vw("--cb_explore_adf -q UA --quiet --epsilon 0.2")
num_iterations = 5000
ctr = run_simulation(vw, num_iterations, users, context1, context2, items, get_cost, do_learn=False)
plot_ctr(num_iterations, ctr)
# + id="UzavSLvC2ZHm" executionInfo={"status": "ok", "timestamp": 1620192940933, "user_tz": -330, "elapsed": 1344, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
users = ['A','B','C']
items = ['Item1','Item2','Item3','Item4','Item5','Item6']
context1 = ['morning','evening']
context2 = ['summer','winter']
context = pd.DataFrame(list(product(users, context1, context2, items)), columns=['users', 'context1', 'context2', 'items'])
context['reward'] = 0
#user 1 likes Item 2 in morning, and Item 5 in summer
context.loc[(context.users=='A') & \
(context.context1=='morning') & \
(context['items']=='Item2'), \
'reward'] = 1
context.loc[(context.users=='A') & \
(context.context2=='summer') & \
(context['items']=='Item5'), \
'reward'] = 1
#user 2 likes Item 2 in summer, and Item 5 in morning
context.loc[(context.users=='B') & \
(context.context2=='summer') & \
(context['items']=='Item2'), \
'reward'] = 1
context.loc[(context.users=='B') & \
(context.context1=='morning') & \
(context['items']=='Item5'), \
'reward'] = 1
#user 3 likes Item 4 in morning, Item 3 in evening, and item 4 in winter evening
context.loc[(context.users=='C') & \
(context.context1=='morning') & \
(context['items']=='Item4'), \
'reward'] = 1
context.loc[(context.users=='C') & \
(context.context1=='evening') & \
(context['items']=='Item3'), \
'reward'] = 1
context.loc[(context.users=='C') & \
(context.context1=='evening') & \
(context.context2=='winter') & \
(context['items']=='Item4'), \
'reward'] = 1
context['cost'] = context['reward']*-1
contextdf_new = context.copy()
def get_cost_new1(context,action):
return contextdf_new.loc[(contextdf_new['users']==context['user']) & \
(contextdf_new.context1==context['context1']) & \
(contextdf_new.context2==context['context2']) & \
(contextdf_new['items']==action), \
'cost'].values[0]
# + id="Mqvn5gLc2ik2" executionInfo={"status": "ok", "timestamp": 1620193132066, "user_tz": -330, "elapsed": 1346, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def run_simulation_multiple_cost_functions(vw, num_iterations, users, contexts1, contexts2, actions, cost_functions, do_learn = True):
cost_sum = 0.
ctr = []
start_counter = 1
end_counter = start_counter + num_iterations
for cost_function in cost_functions:
for i in range(start_counter, end_counter):
user = choose_user(users)
context1 = choose_context1(contexts1)
context2 = choose_context2(contexts2)
context = {'user': user, 'context1': context1, 'context2': context2}
action, prob = get_action(vw, context, actions)
cost = cost_function(context, action)
cost_sum += cost
if do_learn:
vw_format = vw.parse(to_vw_example_format(context, actions, (action, cost, prob)),pyvw.vw.lContextualBandit)
vw.learn(vw_format)
ctr.append(-1*cost_sum/i)
start_counter = end_counter
end_counter = start_counter + num_iterations
return ctr
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="ExchBfTr2lBS" executionInfo={"status": "ok", "timestamp": 1620193222170, "user_tz": -330, "elapsed": 13076, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="3c23545d-652c-4deb-971c-8da1b409fbd9"
# use first reward function initially and then switch to second reward function
# Instantiate learner in VW
vw = pyvw.vw("--cb_explore_adf -q UA --quiet --epsilon 0.2")
num_iterations_per_cost_func = 5000
cost_functions = [get_cost, get_cost_new1]
total_iterations = num_iterations_per_cost_func * len(cost_functions)
ctr = run_simulation_multiple_cost_functions(vw, num_iterations_per_cost_func, users, context1, context2, items, cost_functions)
plot_ctr(total_iterations, ctr)
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="mgaEviR42mlI" executionInfo={"status": "ok", "timestamp": 1620193274550, "user_tz": -330, "elapsed": 12387, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="00ccc48f-f222-4a58-fe14-d8675d4b13cc"
# Do not learn
# use first reward function initially and then switch to second reward function
# Instantiate learner in VW
vw = pyvw.vw("--cb_explore_adf -q UA --quiet --epsilon 0.2")
num_iterations_per_cost_func = 5000
cost_functions = [get_cost, get_cost_new1]
total_iterations = num_iterations_per_cost_func * len(cost_functions)
ctr = run_simulation_multiple_cost_functions(vw, num_iterations_per_cost_func, users, context1, context2, items, cost_functions, do_learn=False)
plot_ctr(total_iterations, ctr)
# + id="4EEJ473k2xwx"
| _docs/nbs/VW-Contextual-Mod1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
# %matplotlib inline
from __future__ import print_function
# Declaration
Nephropathy = ['A', 'B', 'C']
Neuropathy = ['1', '2', '3']
class Model(object):
"""A general model class"""
def __init__(self, states, parameters=None, submodels=None):
# States should be a dictionary for the different states
self.states = states
# Make to types of states: Discrete states and continuous states
self.parameters = parameters
self.time = []
self.submodels = submodels
def rhs(self, state, time):
P = self.parameters["P"]
return P
def update(self, state, time):
# Update all submodels
if self.submodels:
self.submodels.update(self.submodels.states, time)
# Make this somehow into a loop over all states going through a dictionary of functions
# A1c state
currentstate = self.states["A1c"][-1]
newstate = currentstate + 0.01*np.random.random_sample()
self.states["A1c"].append(newstate)
# Nephropathy
currentstate = self.states["Nephropathy"][-1]
newstate = np.random.choice(Nephropathy, None, p=self.rhs(currentstate, time)[Nephropathy.index(currentstate),:])
self.states["Nephropathy"].append(newstate)
# Time step
self.time.append(time)
# Simulate Model
def simulate(model, tspan):
for t in tspan:
model.update(model.states, t)
# +
parameters = {"P" : np.array([[0.9, 0.1, 0],[0, 0.8, 0.2],[0.01, 0, 0.99]])}
states = {
"Nephropathy": [np.random.choice(Nephropathy)],
"A1c" : [0.05]
}
substates = {
"Nephropathy": [np.random.choice(Nephropathy)],
"A1c" : [0.05]
}
submodel = Model(substates,parameters)
model = Model(states,parameters,submodel)
simulate(model, range(100))
print(model.submodels.states)
print(model.states)
# +
# Simulate Cohort
tspan = range(100)
cohort = []
for i in range(100):
# Initialize
states = {
"Nephropathy": [np.random.choice(Nephropathy)],
"A1c" : [0.05]
}
substates = {
"Nephropathy": [np.random.choice(Nephropathy)],
"A1c" : [0.05]
}
submodel = Model(substates,parameters)
cohort.append(Model(states,parameters,submodel))
# Simulate
simulate(cohort[-1],tspan)
# Output
print(len(cohort),cohort[1].states)
# -
# ## Inheritance
P = np.array([[0.9, 0.1, 0],[0, 0.8, 0.2],[0.01, 0, 0.99]])
x=[]
x.append(0)
for i in range(100):
x.append(np.random.choice([0, 1 ,2], 1, p=P[x[i],:])[0])
print(x)
plt.hist(x)
| ModelFramework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# ## Scaling estimates figure
# +
#import gsw as sw # Gibbs seawater package
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
import matplotlib.gridspec as gspec
# %matplotlib inline
from matplotlib.ticker import FormatStrFormatter
from netCDF4 import Dataset
import numpy as np
import pandas as pd
import seaborn as sns
import sys
import scipy.stats
import xarray as xr
import canyon_tools.readout_tools as rout
import canyon_tools.metrics_tools as mpt
# -
sns.set_context('paper')
sns.set_style('white')
# +
CanyonGrid='/data/kramosmu/results/TracerExperiments/CNTDIFF_STEP/run38/gridGlob.nc'
CanyonGridOut = Dataset(CanyonGrid)
CanyonGridNoC='/data/kramosmu/results/TracerExperiments/CNTDIFF_STEP/run42/gridGlob.nc'
CanyonGridOutNoC = Dataset(CanyonGridNoC)
CanyonState='/data/kramosmu/results/TracerExperiments/CNTDIFF_STEP/run38/stateGlob.nc'
CanyonStateOut = Dataset(CanyonState)
# Grid variables
nx = 616
ny = 360
nz = 90
nt = 19 # t dimension size
time = CanyonStateOut.variables['T']
# +
# Constants and scales
L = 6400.0 # canyon length
R = 5000.0 # Upstream radius of curvature
g = 9.81 # accel. gravity
Wsb = 13000 # Width at shelf break
Hs = 147.5 # Shelf break depth
s = 0.005 # shelf slope
W = 8300 # mid-length width
Hh=97.5 # head depth
Hr = 132.0 # rim depth at DnS
# NOTE: The default values of all functions correspond to the base case
def Dh(f=9.66E-4,L=6400.0,N=5.5E-3):
'''Vertical scale Dh'''
return((f*L)/(N))
def Ro(U=0.37,f=9.66E-4,R=5000.0):
'''Rossby number using radius of curvature as length scale'''
return(U/(f*R))
def F(Ro):
'''Function that estimates the ability of the flow to follow isobaths'''
return(Ro/(0.9+Ro))
def Bu(N=5.5E-3,f=9.66E-5,W=8300,Hs=150.0):
'''Burger number'''
return(N*Hs/(f*W))
def RossbyRad(N=5.5E-3,Hs=150.0,f=9.66E-4):
'''1st Rossby radius of deformation'''
return(N*Hs/f)
# -
# +
import canyon_records
records = canyon_records.main()
# +
# records2 has all the runs without the ones where K_bg changes. I don't want to use this ones for fitting the data
ind = [0,3,4,5,6,7,8,9,12,13,14,15,16,17,18,19,20,21]
records2 = []
for ii in ind:
records2.append(records[ii])
# -
# records3 has all the runs without the ones where K_bg changes and run with low U high N.
ind = [0,3,4,5,6,7,8,9,12,13,14,15,16,17,18,19,21]
records3 = []
for ii in ind:
records3.append(records[ii])
# +
select_rec = [0,1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28]
for ind in select_rec:
file = ('/data/kramosmu/results/TracerExperiments/%s/HCW_TrMass_%s%s.csv' %(records[ind].exp_code,
records[ind].exp_code,
records[ind].run_num))
#fileNoC = ('/data/kramosmu/results/TracerExperiments/%s/HCW_TrMass_%s%s.csv' %(recordsNoC[ind].exp_code,
# recordsNoC[ind].exp_code,
# recordsNoC[ind].run_num))
print(file)
dfcan = pd.read_csv(file)
#dfnoc = pd.read_csv(fileNoC)
records[ind].HCW = dfcan['HCW']
#recordsNoC[ind].HCW = dfnoc['HCW']
records[ind].HCWTr1 = dfcan['HCWTr1']
#recordsNoC[ind].HCWTr1 = dfnoc['HCWTr1']
records[ind].TrMass = dfcan['TrMassHCW']
#recordsNoC[ind].TrMass = dfnoc['TrMassHCW']
records[ind].TrMassTr1 = dfcan['TrMassHCWTr1']
#recordsNoC[ind].TrMassTr1 = dfnoc['TrMassHCWTr1']
records[ind].TrMassTr2 = dfcan['TrMassHCWTr2']
#recordsNoC[ind].TrMassTr2 = dfnoc['TrMassHCWTr2']
records[ind].TrMassTot = dfcan['TotTrMass']
#recordsNoC[ind].TrMassTot = dfnoc['TotTrMass']
records[ind].TrMassTotTr2 = dfcan['TotTrMassTr2']
#recordsNoC[ind].TrMassTotTr2 = dfnoc['TotTrMassTr2']
records[ind].TrMassTotTr1 = dfcan['TotTrMassTr1']
#recordsNoC[ind].TrMassTotTr1 = dfnoc['TotTrMassTr1']
# -
# +
# Save mean maximum N of days 3-6 and std for each run.
keys = ['N2_tt08','N2_tt10','N2_tt12','N2_tt14','N2_tt16',]
key0 = 'N2_tt00'
stname = 'DnC' # Station at downstream side of canyon
for record in records:
filename1 = ('/ocean/kramosmu/OutputAnalysis/outputanalysisnotebooks/results/metricsDataFrames/N2_%s_%s.csv' %
(record.name,stname))
df = pd.read_csv(filename1)
df_anom=(df.sub(df[key0].squeeze(),axis=0)).add(df[key0][0])
maxd3 = max(df_anom[keys[0]][26:]) # 22:26 for other scaling
maxd4 = max(df_anom[keys[1]][26:])
maxd5 = max(df_anom[keys[2]][26:])
maxd6 = max(df_anom[keys[3]][26:])
maxd7 = max(df_anom[keys[4]][26:])
record.maxN = np.mean(np.array([maxd3,maxd4,maxd5,maxd6,maxd7]))
record.stdN = np.std(np.array([maxd3,maxd4,maxd5,maxd6,maxd7]))
maxd3a = max(df_anom[keys[0]][22:26]) # 22:26 for other scaling
maxd4a = max(df_anom[keys[1]][22:26])
maxd5a = max(df_anom[keys[2]][22:26])
maxd6a = max(df_anom[keys[3]][22:26])
maxd7a = max(df_anom[keys[4]][22:26])
record.maxNabove = np.mean(np.array([maxd3a,maxd4a,maxd5a,maxd6a,maxd7a]))
record.stdNabove = np.std(np.array([maxd3a,maxd4a,maxd5a,maxd6a,maxd7a]))
print(filename1)
# -
# +
# Save mean maximum N of days 3-6 and std for each run.
keys = ['<KEY>','<KEY> <KEY>','<KEY>','<KEY>']
key0 = '<KEY>'
stname = 'DnC' # Station at downstream side of canyon
for record in records:
filename1 = ('/ocean/kramosmu/OutputAnalysis/outputanalysisnotebooks/results/metricsDataFrames/dTr1dz_%s_%s.csv' %
(record.name,stname))
df = pd.read_csv(filename1)
df_anom=(df.sub(df[key0].squeeze(),axis=0)).add(df[key0][0])
maxd3 = min(df_anom[keys[0]][22:26])
maxd4 = min(df_anom[keys[1]][22:26])
maxd5 = min(df_anom[keys[2]][22:26])
maxd6 = min(df_anom[keys[3]][22:26])
maxd7 = min(df_anom[keys[4]][22:26])
record.maxdTrdz = np.mean(np.array([maxd3,maxd4,maxd5,maxd6,maxd7]))
record.stddTrdz = np.std(np.array([maxd3,maxd4,maxd5,maxd6,maxd7]))
record.inidTrdz = df[key0][30]
print('The initial tracer gradient is %f ' %(-1*record.inidTrdz))
# +
keys = ['Tr_profile_tt08','Tr_profile_tt10','Tr_profile_tt12','Tr_profile_tt14','Tr_profile_tt16']
key0 = 'Tr_profile_tt00'
stname = 'DnC' # Station at downstream side of canyon
for record in records:
filename1 = ('/ocean/kramosmu/OutputAnalysis/outputanalysisnotebooks/results/metricsDataFrames/Tr1_profile_%s_%s.csv' % (record.name,stname))
df = pd.read_csv(filename1)
df_anom=(df.sub(df[key0].squeeze(),axis=0))
maxd4 = np.mean(df[keys[0]][25:32])
maxd5 = np.mean(df[keys[1]][25:32])
maxd6 = np.mean(df[keys[2]][25:32])
maxd7 = np.mean(df[keys[3]][25:32])
maxd8 = np.mean(df[keys[4]][25:32])
maxd0 = df[key0][29]
record.maxTr = np.mean(np.array([maxd4,maxd5,maxd6,maxd7,maxd8]))
record.stdTr = np.std(np.array([maxd4,maxd5,maxd6,maxd7,maxd8]))
record.TrSB = maxd0
Ctop = df[key0][0]
# +
plt.rcParams['font.size'] = 8.0
f = plt.figure(figsize = (7,8.5)) # 190mm = 7.48 in, 230cm = 9.05in
gs = gspec.GridSpec(3, 2)
ax0 = plt.subplot(gs[0,0])
ax1 = plt.subplot(gs[0,1])
ax2 = plt.subplot(gs[1,0])
ax3 = plt.subplot(gs[1,1])
ax4 = plt.subplot(gs[2,0])
ax5 = plt.subplot(gs[2,1])
t=6.5 # days
# Tr
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% N/No above %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax0.errorbar((4.10*(1+(Z+Zdif)/Hh))-3.00,rec.maxNabove/(rec.N**2),
yerr = rec.stdNabove/(rec.N**2), # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
ax0.set_ylabel('$N^2/N^2_0$',labelpad=0.5)
ax0.set_xlabel(r'$4.10(1+(Z+Z_{dif})/H_h)-3.00$',labelpad=0.5)
maxN_array_Kv = np.array([rec.maxNabove/(rec.N**2) for rec in records2])
tilt_array_Kv = np.array([1+(rec.Z+rec.Zdif)/Hh for rec in records2])
x_fit = np.linspace(1.0, 7, 50)
mean_sq_err = np.mean(((maxN_array_Kv)-(4.10*tilt_array_Kv-3.00))**2)
upper_bound = ax0.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax0.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for N/No above is %f' %mean_sq_err)
ax0.plot(np.linspace(1.0, 7, 50),np.linspace(1.0, 7, 50),'k-')
ax0.set_ylim(1,7)
ax0.set_xlim(1,7)
ax0.text(0.92,0.05,'(a)',transform=ax0.transAxes,fontsize=10)
ax0.text(0.05,0.9,'Eqn. 21',transform=ax0.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% N/No %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax1.errorbar((4.82*(1+(Z-Zdif)/Hh))-3.47,rec.maxN/(rec.N**2),
yerr = rec.stdN/(rec.N**2), # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
ax1.set_ylabel('$N^2/N^2_0$',labelpad=0.5)
ax1.set_xlabel(r'$4.82(1+(Z-Z_{dif})/H_h)-3.47$',labelpad=0.5)
maxN_array = np.array([rec.maxN/(rec.N**2) for rec in records2])
tilt_array = np.array([((rec.Z-rec.Zdif)/Hh) for rec in records2])
x_fit = np.linspace(0, 7, 50)
mean_sq_err = np.mean(((maxN_array_Kv)-(4.82*tilt_array_Kv-3.47))**2)
upper_bound = ax1.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax1.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for N/N0 below is %f' %mean_sq_err)
ax1.plot(np.linspace(0, 7, 50),np.linspace(0, 7, 50),'k-')
ax1.set_ylim(0,7)
ax1.set_xlim(0,7)
ax1.text(0.92,0.05,'(b)',transform=ax1.transAxes,fontsize=10)
ax1.text(0.05,0.9,'Eqn. 23',transform=ax1.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% dTr/dz %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
ax2.plot(np.linspace(1,6.5, 50),np.linspace(1,6.5, 50),'k-')
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax2.errorbar(4.17*(1+(Z+Zdif)/Hh)-3.25,rec.maxdTrdz/rec.inidTrdz,
yerr = rec.stddTrdz/rec.inidTrdz, # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
ax2.set_ylabel('$\partial_zC/\partial_zC_0$',labelpad=0.5)
ax2.set_xlabel(r'$4.27(1+(Z+Z_{dif})/H_h)-3.25$',labelpad=0.5)
maxN_array = np.array([rec.maxdTrdz/rec.inidTrdz for rec in records2])
tilt_array = np.array([1+(rec.Z+rec.Zdif)/Hh for rec in records2])
x_fit = np.linspace(1,7, 50)
mean_sq_err = np.mean(((maxN_array)-(4.17*tilt_array-3.25))**2)
upper_bound = ax2.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax2.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for dTr/dz above is %f' %mean_sq_err)
ax2.set_ylim(1,7)
ax2.set_xlim(1,7)
ax2.text(0.92,0.05,'(c)',transform=ax2.transAxes,fontsize=10)
ax2.text(0.05,0.9,'Eqn. 22',transform=ax2.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Tracer Conc %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
ax3.plot(np.linspace(0,1.4,50),np.linspace(0,1.4,50),'k-')
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
dTrdz = -rec.inidTrdz*(4.17*(1+(Z+Zdif)/Hh)-3.25)
plt1 = ax3.errorbar((0.01*(dTrdz*(Hh+132.0)/2.0)) + 1.00,rec.maxTr/rec.TrSB,
yerr = rec.stdTr/rec.TrSB,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
rec.dTr = dTrdz
ax3.set_xlim(0.95,1.4)
ax3.set_ylim(0.95,1.4)
#ax3.set_title('Max tracer concetration between head nad sh, Dn')
ax3.set_ylabel('$C_{max}$/$C_{0}$',labelpad=0.5)
ax3.set_xlabel(r'$\bar{C}/C_{0}=%1.2f\partial_zC(H_{sb}+H_r)/2+%1.1f$' %((0.01),(1.00)),labelpad=0.5)
fmt = FormatStrFormatter('%1.2f')
ax3.xaxis.set_major_formatter(fmt)
ax3.yaxis.set_major_formatter(fmt)
ax3.tick_params(axis='x', pad=1)
ax3.tick_params(axis='y', pad=1)
maxN_array = np.array([rec.maxTr/rec.TrSB for rec in records2])
tilt_array = np.array([(0.01*(rec.dTr*(Hh+132.0)/2.0))+1.00 for rec in records2])
x_fit = np.linspace(0,1.4, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax3.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax3.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for C/Co is %f' %mean_sq_err)
ax3.text(0.92,0.05,'(d)',transform=ax3.transAxes,fontsize=10)
ax3.text(0.05,0.9,'Eqn. 26',transform=ax3.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
t=4
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Se = (s*rec.N)/(rec.f*((F(Ro(rec.u_mod,rec.f,W))/Ro(rec.u_mod,rec.f,L))**(1/2)))
HA2013=((0.91*(F(Ro(rec.u_mod,rec.f,W))**(3/2))*(Ro(rec.u_mod,rec.f,L)**(1/2))*((1-1.21*Se)**3))+0.07)
can_eff = rec.HCW
Phi = np.mean(np.array([(can_eff[ii]-can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
Phi_std = np.std(np.array([(can_eff[ii]-can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
#Phi = np.mean(np.array([(can_eff[ii]) for ii in range (8,14)]))
plt1 = ax4.errorbar(HA2013*(rec.u_mod*W*Dh(rec.f,L,rec.N))/1E4,Phi/1E4,
yerr=Phi_std/1E4,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Phi = Phi/1E4
rec.HA2013 = HA2013*(rec.u_mod*W*Dh(rec.f,L,rec.N))/1E4
ax4.set_ylabel(r'Upwelling flux ($10^4$ m$^3$s$^{-1}$) ',labelpad=0.5)
ax4.set_xlabel(r'$\Phi$ ($10^4$ m$^3$s$^{-1}$)',labelpad=0.5)
maxN_array = np.array([rec.Phi for rec in records3])
tilt_array = np.array([rec.HA2013 for rec in records3])
x_fit = np.linspace(0,5, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax4.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax4.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for Phi is %f' %mean_sq_err)
ax4.plot(np.linspace(0,5, 50),np.linspace(0,5, 50),'k-')
ax4.set_ylim(0,5.0)
ax4.text(0.92,0.05,'(e)',transform=ax4.transAxes,fontsize=10)
ax4.text(0.05,0.9,'Eqn. 27',transform=ax4.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi_Tr %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
Capprox=rec.TrSB*((0.01*(rec.dTr*(Hh+132.0)/2.0))+1.0)
Se = (s*rec.N)/(rec.f*((F(Ro(rec.u_mod,rec.f,W))/Ro(rec.u_mod,rec.f,L))**(1/2)))
HA2013=(rec.u_mod*W*Dh(rec.f,L,rec.N))*((0.91*(F(Ro(rec.u_mod,rec.f,W))**(3/2))*(Ro(rec.u_mod,rec.f,L)**(1/2))*((1-1.21*Se)**3))+0.07)
mass_can_eff = rec.TrMass
can_eff = rec.HCW
phiTr = np.mean(np.array([(mass_can_eff[ii]-mass_can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
phiTr_std = np.std(np.array([(mass_can_eff[ii]-mass_can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
plt1 = ax5.errorbar(HA2013*(Capprox)/1E5, # 1000 is a conversion factor
(phiTr)/1E5,
yerr = phiTr_std/1E5,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.PhiTr = (phiTr)/1E5
rec.PhiTrScale = HA2013*(Capprox)/1E5
ax5.set_xlabel(r'$\Phi_{Tr}=\Phi \bar{C}$ ($10^5$ $\mu$Mm$^{3}$s$^{-1}$)',labelpad=0.5)
ax5.set_ylabel(r'Tracer upwelling flux ($10^5$ $\mu$Mm$^{3}$s$^{-1}$)',labelpad=0.5)
maxN_array = np.array([rec.PhiTr for rec in records3])
tilt_array = np.array([rec.PhiTrScale for rec in records3])
x_fit = np.linspace(0,4, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax5.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax5.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for PhiTr/1E5 above is %f' %mean_sq_err)
ax5.set_ylim(0,4)
ax5.plot(np.linspace(0,4, 50),np.linspace(0,4, 50),'k-')
ax5.text(0.93,0.05,'(f)',transform=ax5.transAxes,fontsize=10)
ax5.text(0.05,0.9,'Eqn. 28',transform=ax5.transAxes,fontsize=10)
ax5.legend(bbox_to_anchor=(1.0,-0.15),ncol=5, columnspacing=0.1)
ax0.tick_params(axis='x', pad=1)
ax1.tick_params(axis='x', pad=1)
ax2.tick_params(axis='x', pad=1)
ax4.tick_params(axis='x', pad=1)
ax5.tick_params(axis='x', pad=1)
ax0.tick_params(axis='y', pad=3)
ax1.tick_params(axis='y', pad=3)
ax2.tick_params(axis='y', pad=3)
ax4.tick_params(axis='y', pad=3)
ax5.tick_params(axis='y', pad=3)
plt.savefig('figure9.eps',format='eps',bbox_inches='tight')
print(rec.inidTrdz)
# +
plt.rcParams['font.size'] = 8.0
f = plt.figure(figsize = (7,8.5)) # 190mm = 7.48 in, 230cm = 9.05in
gs = gspec.GridSpec(3, 2)
ax0 = plt.subplot(gs[0,0])
ax1 = plt.subplot(gs[0,1])
ax2 = plt.subplot(gs[1,0])
ax3 = plt.subplot(gs[1,1])
ax4 = plt.subplot(gs[2,0])
ax5 = plt.subplot(gs[2,1])
t=6.5 # days
# Tr
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% N/No above %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax0.errorbar((4.10*(1+(Z+Zdif)/Hh))-3.00,rec.maxNabove/(rec.N**2),
yerr = rec.stdNabove/(rec.N**2), # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
markeredgewidth=1.0,
markeredgecolor = 'k',
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
ax0.set_ylabel('$N^2/N^2_0$',labelpad=0.5)
ax0.set_xlabel(r'$4.10(1+(Z+Z_{dif})/H_h)-3.00$',labelpad=0.5)
maxN_array_Kv = np.array([rec.maxNabove/(rec.N**2) for rec in records2])
tilt_array_Kv = np.array([1+(rec.Z+rec.Zdif)/Hh for rec in records2])
x_fit = np.linspace(1.0, 7, 50)
mean_sq_err = np.mean(((maxN_array_Kv)-(4.10*tilt_array_Kv-3.00))**2)
upper_bound = ax0.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax0.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for N/No above is %f' %mean_sq_err)
ax0.plot(np.linspace(1.0, 7, 50),np.linspace(1.0, 7, 50),'k-')
#ax0.set_ylim(1,7)
#ax0.set_xlim(1,7)
ax0.text(0.92,0.05,'(a)',transform=ax0.transAxes,fontsize=10)
ax0.text(0.05,0.9,'Eqn. 21',transform=ax0.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% N/No %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax1.errorbar((4.82*(1+(Z-Zdif)/Hh))-3.47,rec.maxN/(rec.N**2),
yerr = rec.stdN/(rec.N**2), # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
markeredgewidth=1.0,
markeredgecolor = 'k',
label=rec.label)
rec.Z = Z
rec.N_below = np.sqrt(((4.82*(1+(Z-Zdif)/Hh))-3.47)*rec.N**2)
ax1.set_ylabel('$N^2/N^2_0$',labelpad=0.5)
ax1.set_xlabel(r'$4.82(1+(Z-Z_{dif})/H_h)-3.47$',labelpad=0.5)
maxN_array = np.array([rec.maxN/(rec.N**2) for rec in records2])
tilt_array = np.array([((rec.Z-rec.Zdif)/Hh) for rec in records2])
x_fit = np.linspace(0, 7, 50)
mean_sq_err = np.mean(((maxN_array_Kv)-(4.82*tilt_array_Kv-3.47))**2)
upper_bound = ax1.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax1.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for N/N0 below is %f' %mean_sq_err)
ax1.plot(np.linspace(0, 7, 50),np.linspace(0, 7, 50),'k-')
#ax1.set_ylim(0,7)
#ax1.set_xlim(0,7)
ax1.text(0.92,0.05,'(b)',transform=ax1.transAxes,fontsize=10)
ax1.text(0.05,0.9,'Eqn. 23',transform=ax1.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% dTr/dz %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
ax2.plot(np.linspace(1,6.5, 50),np.linspace(1,6.5, 50),'k-')
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax2.errorbar(4.17*(1+(Z+Zdif)/Hh)-3.25,rec.maxdTrdz/rec.inidTrdz,
yerr = rec.stddTrdz/rec.inidTrdz, # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
markeredgewidth=1.0,
markeredgecolor = 'k',
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
ax2.set_ylabel('$\partial_zC/\partial_zC_0$',labelpad=0.5)
ax2.set_xlabel(r'$4.27(1+(Z+Z_{dif})/H_h)-3.25$',labelpad=0.5)
maxN_array = np.array([rec.maxdTrdz/rec.inidTrdz for rec in records2])
tilt_array = np.array([1+(rec.Z+rec.Zdif)/Hh for rec in records2])
x_fit = np.linspace(1,7, 50)
mean_sq_err = np.mean(((maxN_array)-(4.17*tilt_array-3.25))**2)
upper_bound = ax2.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax2.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for dTr/dz above is %f' %mean_sq_err)
#ax2.set_ylim(1,7)
#ax2.set_xlim(1,7)
ax2.text(0.92,0.05,'(c)',transform=ax2.transAxes,fontsize=10)
ax2.text(0.05,0.9,'Eqn. 22',transform=ax2.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Tracer Conc %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
ax3.plot(np.linspace(1,1.4,50),np.linspace(1,1.4,50),'k-')
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
dTrdz = -rec.inidTrdz*(4.17*(1+(Z+Zdif)/Hh)-3.25)
plt1 = ax3.errorbar((0.01*(dTrdz*(Hh+132.0)/2.0)) + 1.00,rec.maxTr/rec.TrSB,
yerr = rec.stdTr/rec.TrSB,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
markeredgewidth=1.0,
markeredgecolor = 'k',
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
rec.dTr = dTrdz
#ax3.set_xlim(0.95,1.4)
#ax3.set_ylim(0.95,1.4)
#ax3.set_title('Max tracer concetration between head nad sh, Dn')
ax3.set_ylabel('$C_{max}$/$C_{0}$',labelpad=0.5)
ax3.set_xlabel(r'$\bar{C}/C_{0}=%1.2f\partial_zC(H_{sb}+H_r)/2+%1.1f$' %((0.01),(1.00)),labelpad=0.5)
fmt = FormatStrFormatter('%1.2f')
ax3.xaxis.set_major_formatter(fmt)
ax3.yaxis.set_major_formatter(fmt)
ax3.tick_params(axis='x', pad=1)
ax3.tick_params(axis='y', pad=1)
maxN_array = np.array([rec.maxTr/rec.TrSB for rec in records2])
tilt_array = np.array([(0.01*(rec.dTr*(Hh+132.0)/2.0))+1.00 for rec in records2])
x_fit = np.linspace(1,1.4, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax3.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax3.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for C/Co is %f' %mean_sq_err)
ax3.text(0.92,0.05,'(d)',transform=ax3.transAxes,fontsize=10)
ax3.text(0.05,0.9,'Eqn. 26',transform=ax3.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
t=4
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Se = (s*rec.N)/(rec.f*((F(Ro(rec.u_mod,rec.f,W))/Ro(rec.u_mod,rec.f,L))**(1/2)))
HA2013=((0.91*(F(Ro(rec.u_mod,rec.f,W))**(3/2))*(Ro(rec.u_mod,rec.f,L)**(1/2))*((1-1.21*Se)**3))+0.07)
can_eff = rec.HCW
Phi = np.mean(np.array([(can_eff[ii]-can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
Phi_std = np.std(np.array([(can_eff[ii]-can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
#Phi = np.mean(np.array([(can_eff[ii]) for ii in range (8,14)]))
plt1 = ax4.errorbar(HA2013*(rec.u_mod*W*Dh(rec.f,L,rec.N))/1E4,Phi/1E4,
yerr=Phi_std/1E4,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
markeredgewidth=1.0,
markeredgecolor = 'k',
label=rec.label)
rec.Phi = Phi/1E4
rec.HA2013 = HA2013*(rec.u_mod*W*Dh(rec.f,L,rec.N))/1E4
ax4.set_ylabel(r'Upwelling flux ($10^4$ m$^3$s$^{-1}$) ',labelpad=0.5)
ax4.set_xlabel(r'$\Phi$ ($10^4$ m$^3$s$^{-1}$)',labelpad=0.5)
maxN_array = np.array([rec.Phi for rec in records3])
tilt_array = np.array([rec.HA2013 for rec in records3])
x_fit = np.linspace(0,5, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax4.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax4.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for Phi is %f' %mean_sq_err)
ax4.plot(np.linspace(0,5, 50),np.linspace(0,5, 50),'k-')
#ax4.set_ylim(0,5.0)
ax4.text(0.92,0.05,'(e)',transform=ax4.transAxes,fontsize=10)
ax4.text(0.05,0.9,'Eqn. 27',transform=ax4.transAxes,fontsize=10)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi_Tr %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
Capprox=rec.TrSB*((0.01*(rec.dTr*(Hh+132.0)/2.0))+1.0)
Se = (s*rec.N)/(rec.f*((F(Ro(rec.u_mod,rec.f,W))/Ro(rec.u_mod,rec.f,L))**(1/2)))
HA2013=(rec.u_mod*W*Dh(rec.f,L,rec.N))*((0.91*(F(Ro(rec.u_mod,rec.f,W))**(3/2))*(Ro(rec.u_mod,rec.f,L)**(1/2))*((1-1.21*Se)**3))+0.07)
mass_can_eff = rec.TrMass
can_eff = rec.HCW
phiTr = np.mean(np.array([(mass_can_eff[ii]-mass_can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
phiTr_std = np.std(np.array([(mass_can_eff[ii]-mass_can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
plt1 = ax5.errorbar(HA2013*(Capprox)/1E5, # 1000 is a conversion factor
(phiTr)/1E5,
yerr = phiTr_std/1E5,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
markeredgewidth=1.0,
markeredgecolor = 'k',
label=rec.label)
rec.PhiTr = (phiTr)/1E5
rec.PhiTrScale = HA2013*(Capprox)/1E5
ax5.set_xlabel(r'$\Phi_{Tr}=\Phi \bar{C}$ ($10^5$ $\mu$Mm$^{3}$s$^{-1}$)',labelpad=0.5)
ax5.set_ylabel(r'Tracer upwelling flux ($10^5$ $\mu$Mm$^{3}$s$^{-1}$)',labelpad=0.5)
maxN_array = np.array([rec.PhiTr for rec in records3])
tilt_array = np.array([rec.PhiTrScale for rec in records3])
x_fit = np.linspace(0,4, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax5.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax5.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for PhiTr/1E5 above is %f' %mean_sq_err)
#ax5.set_ylim(0,4)
ax5.plot(np.linspace(0,4, 50),np.linspace(0,4, 50),'k-')
ax5.text(0.93,0.05,'(f)',transform=ax5.transAxes,fontsize=10)
ax5.text(0.05,0.9,'Eqn. 28',transform=ax5.transAxes,fontsize=10)
ax5.legend(bbox_to_anchor=(1.0,-0.15),ncol=5, columnspacing=0.1)
ax0.tick_params(axis='x', pad=1)
ax1.tick_params(axis='x', pad=1)
ax2.tick_params(axis='x', pad=1)
ax4.tick_params(axis='x', pad=1)
ax5.tick_params(axis='x', pad=1)
ax0.tick_params(axis='y', pad=3)
ax1.tick_params(axis='y', pad=3)
ax2.tick_params(axis='y', pad=3)
ax4.tick_params(axis='y', pad=3)
ax5.tick_params(axis='y', pad=3)
#plt.savefig('figure9.eps',format='eps',bbox_inches='tight')
print(rec.inidTrdz)
# -
-
# +
print ("\t".join(['Experiment &','$\kappa_{bg}$ &','$\kappa_{can}$&', '$N$ (s$^{-1}$)$&',
'$f$ (s$^{-1}$)&', 'U (ms$^{-1}$)&', '$Bu$&', '$Ro$' ,
]))
for rec,recNoC in zip(records,recordsNoC):
print ("\t".join(['%s\t&$%0.2e$\t&$%0.2e$\t&$%.1e$\t&$%.2e$\t&$%.2f$\t&$%.2f$\t&$%.2f$\t '
% (rec.paperName,
rec.kv,
rec.kv,
rec.N,
rec.f,
rec.u_mod,
Bu(rec.N, rec.f,Wsb,Hs),
Ro(U=rec.u_mod, f=rec.f, R=L),
)
]))
# -
# ## Individual plots for talks
# +
labels_Bark = ['Lin','Sal','Oxy','Nit','Sil','Pho','NitA','Met']
colours_Bark = ['purple','blue','green','gold','orange','red','orchid','teal']
PhiTr_scaled_barkley = [981713.2 ,1021419.4,-5917704.4,1439242.2,2458652.0,67971.1,907062.9,3573535.6]
PhiTr_model_barkley = [1093887.3,1148391.8,3480067.7,1115451.9,1626847.6,76542.7,901950.0,777955.9]
# +
sns.set_context('talk')
f0,ax0 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
t=5 # days
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% N/No above %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
for rec,recNoC in zip(records,recordsNoC):
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax0.errorbar(4.10*(1+(Z+Zdif)/Hh)-3.00,rec.maxNabove/(rec.N**2),
yerr = rec.stdNabove/(rec.N**2), # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
ax0.set_ylabel('$N^2/N^2_0$',labelpad=0.5)
ax0.set_xlabel(r'$4.10(1+(Z+Z_{dif})/H_s)-3.0$',labelpad=0.5)
maxN_array_Kv = np.array([rec.maxNabove/(rec.N**2) for rec in records2])
tilt_array_Kv = np.array([1+(rec.Z+rec.Zdif)/Hh for rec in records2])
x_fit = np.linspace(1.0, 7, 50)
mean_sq_err = np.mean(((maxN_array_Kv)-(4.10*tilt_array_Kv-3.00))**2)
upper_bound = ax0.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax0.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for N/No above is %f' %mean_sq_err)
ax0.plot(np.linspace(1.0, 7, 50),np.linspace(1.0, 7, 50),'k-')
ax0.set_ylim(1,7)
ax0.set_xlim(1,7)
plt.savefig('scaling_topN.eps',format='eps',bbox_inches='tight')
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% N/No %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
f1,ax1 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
for rec,recNoC in zip(records,recordsNoC):
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax1.errorbar((4.82*(1+(Z-Zdif)/Hh))-3.47,rec.maxN/(rec.N**2),
yerr = rec.stdN/(rec.N**2), # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
ax1.set_ylabel('$N^2/N^2_0$',labelpad=0.5)
ax1.set_xlabel(r'$4.82(1+(Z-Z_{dif})/H_h)-3.47$',labelpad=0.5)
maxN_array = np.array([rec.maxN/(rec.N**2) for rec in records2])
tilt_array = np.array([((rec.Z-rec.Zdif)/Hh) for rec in records2])
x_fit = np.linspace(0, 7, 50)
mean_sq_err = np.mean(((maxN_array_Kv)-(4.82*tilt_array_Kv-3.47))**2)
upper_bound = ax1.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax1.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for N/N0 below is %f' %mean_sq_err)
ax1.plot(np.linspace(0, 7, 50),np.linspace(0, 7, 50),'k-')
ax1.set_ylim(0,7)
ax1.set_xlim(0,7)
plt.savefig('scaling_N.eps',format='eps',bbox_inches='tight')
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% dTr/dz %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
f2,ax2 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
ax2.plot(np.linspace(1,6.5, 50),np.linspace(1,6.5, 50),'k-')
for rec,recNoC in zip(records,recordsNoC):
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
plt1 = ax2.errorbar(4.17*(1+(Z+Zdif)/Hh)-3.25,rec.maxdTrdz/rec.inidTrdz,
yerr = rec.stddTrdz/rec.inidTrdz, # since rec.N is a constant
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
ax2.set_ylabel('$\partial_zC/\partial_zC_0$',labelpad=0.5)
ax2.set_xlabel(r'$4.27(1+(Z+Z_{dif})/H_h)-3.25$',labelpad=0.5)
maxN_array = np.array([rec.maxdTrdz/rec.inidTrdz for rec in records2])
tilt_array = np.array([1+(rec.Z+rec.Zdif)/Hh for rec in records2])
x_fit = np.linspace(1,7, 50)
mean_sq_err = np.mean(((maxN_array)-(4.17*tilt_array-3.25))**2)
upper_bound = ax2.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax2.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for dTr/dz above is %f' %mean_sq_err)
ax2.set_ylim(1,7)
ax2.set_xlim(1,7)
plt.savefig('scaling_dC.eps',format='eps',bbox_inches='tight')
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Tracer Conc %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
f3,ax3 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
for rec in records:
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
ax3.plot(np.linspace(0,1.4,50),np.linspace(0,1.4,50),'k-')
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
dTrdz = -rec.inidTrdz*(6.27*(((Z+Zdif)/Hs))+0.98)
plt1 = ax3.errorbar((0.08*(dTrdz*(Hh+132.0)/2.0))/rec.TrSB + 7.19/rec.TrSB,rec.maxTr/rec.TrSB,
yerr = rec.stdTr/rec.TrSB,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Z = Z
rec.Zdif = Zdif
rec.dTr = dTrdz
ax3.set_xlim(0.95,1.4)
ax3.set_ylim(0.95,1.4)
#ax3.set_title('Max tracer concetration between head nad sh, Dn')
ax3.set_ylabel('$C_{max}$/$C_{0}$',labelpad=0.5)
ax3.set_xlabel(r'$\bar{C}/C_{0}=%1.2f\partial_zC(H_{sb}+H_r)/2+%1.1f$' %((0.08/2),(7.19/rec.TrSB)),labelpad=0.5)
C = (0.08*(dTrdz*(Hh+132.0)/2.0)) + 7.19
err_C = abs(1-(C/rec.maxTr))*100
print('The sigma error for bar{C} %s is %f' %(rec.name,err_C))
fmt = FormatStrFormatter('%1.2f')
ax3.xaxis.set_major_formatter(fmt)
ax3.yaxis.set_major_formatter(fmt)
ax3.tick_params(axis='x', pad=1)
ax3.tick_params(axis='y', pad=1)
maxN_array = np.array([rec.maxTr/rec.TrSB for rec in records2])
tilt_array = np.array([(0.08*(rec.dTr*(Hh+132.0)/2.0))/rec.TrSB+7.19/rec.TrSB for rec in records2])
x_fit = np.linspace(0,1.4, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax3.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax3.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for C/Co is %f' %mean_sq_err)
plt.savefig('scaling_C.eps',format='eps',bbox_inches='tight')
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
f4,ax4 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
t=4
for rec,recNoC in zip(records,recordsNoC):
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Se = (s*rec.N)/(rec.f*((F(Ro(rec.u_mod,rec.f,W))/Ro(rec.u_mod,rec.f,L))**(1/2)))
HA2013=((0.91*(F(Ro(rec.u_mod,rec.f,W))**(3/2))*(Ro(rec.u_mod,rec.f,L)**(1/2))*((1-1.21*Se)**3))+0.07)
can_eff = rec.HCW
Phi = np.mean(np.array([(can_eff[ii]-can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
Phi_std = np.std(np.array([(can_eff[ii]-can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
#Phi = np.mean(np.array([(can_eff[ii]) for ii in range (8,14)]))
plt1 = ax4.errorbar(HA2013*(rec.u_mod*W*Dh(rec.f,L,rec.N))/1E4,Phi/1E4,
yerr=Phi_std/1E4,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.Phi = Phi/1E4
rec.HA2013 = HA2013*(rec.u_mod*W*Dh(rec.f,L,rec.N))/1E4
ax4.set_ylabel(r'Upwelling flux ($10^4$ m$^3$s$^{-1}$) ',labelpad=0.5)
ax4.set_xlabel(r'$\Phi$ ($10^4$ m$^3$s$^{-1}$)',labelpad=0.5)
maxN_array = np.array([rec.Phi for rec in records3])
tilt_array = np.array([rec.HA2013 for rec in records3])
x_fit = np.linspace(0,5, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax4.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax4.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for Phi is %f' %mean_sq_err)
ax4.plot(np.linspace(0,5, 50),np.linspace(0,5, 50),'k-')
ax4.set_ylim(0,5.0)
plt.savefig('scaling_Phi.eps',format='eps',bbox_inches='tight')
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi_Tr %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
f5,ax5 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
for rec,recNoC in zip(records,recordsNoC):
Z = ((rec.f*rec.u_mod*F(Ro(rec.u_mod,rec.f,R))*L)**(0.5))/rec.N
Zdif = np.sqrt(t*3600*24*(rec.kv-rec.kbg))
Capprox=rec.TrSB*((0.01*(rec.dTr*(Hh+132.0)/2.0))+1.0)
Se = (s*rec.N)/(rec.f*((F(Ro(rec.u_mod,rec.f,W))/Ro(rec.u_mod,rec.f,L))**(1/2)))
HA2013=(rec.u_mod*W*Dh(rec.f,L,rec.N))*((0.91*(F(Ro(rec.u_mod,rec.f,W))**(3/2))*(Ro(rec.u_mod,rec.f,L)**(1/2))*((1-1.21*Se)**3))+0.07)
mass_can_eff = rec.TrMass
can_eff = rec.HCW
phiTr = np.mean(np.array([(mass_can_eff[ii]-mass_can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
phiTr_std = np.std(np.array([(mass_can_eff[ii]-mass_can_eff[ii-1])/(time[ii]-time[ii-1]) for ii in range (8,18)]))
plt1 = ax5.errorbar(HA2013*(Capprox)/1E5, # 1000 is a conversion factor
(phiTr)/1E5,
yerr = phiTr_std/1E5,
marker = rec.mstyle,
markersize = 10,
color = sns.xkcd_rgb[rec.color2],
label=rec.label)
rec.PhiTr = (phiTr)/1E5
rec.PhiTrScale = HA2013*(Capprox)/1E5
ax5.set_xlabel(r'$\Phi_{Tr}=\Phi \bar{C}$ ($10^5$ $\mu$Mm$^{3}$s$^{-1}$)',labelpad=0.5)
ax5.set_ylabel(r'Tracer upwelling flux ($10^5$ $\mu$Mm$^{3}$s$^{-1}$)',labelpad=0.5)
maxN_array = np.array([rec.PhiTr for rec in records3])
tilt_array = np.array([rec.PhiTrScale for rec in records3])
x_fit = np.linspace(0,4, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax5.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax5.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
print('The rms error for PhiTr/1E5 above is %f' %mean_sq_err)
ax5.set_ylim(0,4)
ax5.plot(np.linspace(0,4, 50),np.linspace(0,4, 50),'k-')
plt.savefig('scaling_PhiTr.eps',format='eps',bbox_inches='tight')
ax5.legend(bbox_to_anchor=(1.2,-0.15),ncol=6, columnspacing=0.1)
# %%%%%%%%%%%%%%%%%%%%%%%%%%%%%% Phi_Tr with barkley profiles %%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%%
f6,ax6 = plt.subplots(1,1,figsize = (5,5)) # 190mm = 7.48 in, 230cm = 9.05in
ax6.set_xlabel(r'$\Phi_{Tr}/UWD_hC_{sb}$',labelpad=0.5)
ax6.set_ylabel(r'Tracer upwelling flux/UWD_hC_{sb}',labelpad=0.5)
maxN_array = np.array([rec.PhiTr for rec in records3])
tilt_array = np.array([rec.PhiTrScale for rec in records3])
x_fit = np.linspace(0,1, 50)
mean_sq_err = np.mean(((maxN_array)-(tilt_array))**2)
upper_bound = ax6.plot(x_fit,x_fit+(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
lower_bound = ax6.plot(x_fit,x_fit-(mean_sq_err)**(0.5),linestyle = '--',color='0.5')
ax6.plot(np.linspace(0,0.2, 50),np.linspace(0,0.2, 50),'k-')
#for TrMass,Trmodel, lab, col in zip(PhiTr_scaled_barkley, PhiTr_model_barkley,labels_Bark,colours_Bark):
# ax6.plot(TrMass, Trmodel, color=col, label=lab, marker='p')
ax6.set_ylim(0,0.2)
ax6.set_xlim(0,0.2)
ax6.legend(bbox_to_anchor=(1.2,-0.15),ncol=6, columnspacing=0.1)
plt.savefig('scaling_PhiTr_wBarkley.eps',format='eps',bbox_inches='tight')
# -
| PythonScripts/Paper1Figures/fig_scaling_estimates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/stmnk/qa/blob/master/data/dev/NQ_dataset_sample-local.ipynb)
# Import utilities.
import gzip
import shutil
# Extract a `.gz` file.
with gzip.open(f'nq-dev-00.jsonl.gz', 'rb') as f_in:
with open(f'nq-dev-00.jsonl', 'wb') as f_out:
shutil.copyfileobj(f_in, f_out)
# Read data from a `.jsonl` file.
# +
import json
from pandas.io.json import json_normalize
import pandas as pd
def read_jsonl(input_path) -> list:
"""
Read list of objects from a JSON lines file.
"""
data = []
with open(input_path, 'r', encoding='utf-8') as f:
for line in f:
data.append(json.loads(line.rstrip('\n|\r')))
print(f'Loaded {len(data)} records from {input_path}'
return data
# -
# Inspect the data frame.
# +
nq_sample_list = read_jsonl(f'nq-dev-00.jsonl')
df = pd.DataFrame(nq_sample_list, columns=[
'example_id',
'question_text', 'question_tokens',
'document_url', 'document_html', # 'document_tokens',
'long_answer_candidates',
'annotations',
])
df
# -
# Write data to a `.jsonl` file.
# +
def write_jsonl(data, output_path, append=False):
"""
Write list of objects to a JSON lines file.
"""
mode = 'a+' if append else 'w'
with open(output_path, mode, encoding='utf-8') as f:
for line in data:
json_record = json.dumps(line, ensure_ascii=False)
f.write(json_record + '\n')
print('Wrote {} records to {}'.format(len(data), output_path))
| data/dev/NQ_dataset_sample_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="MhoQ0WE77laV"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="_ckMIh7O7s6D"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" colab={} colab_type="code" id="vasWnqRgy1H4"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] colab_type="text" id="YenH_9hJbFk1"
# # Clasificacion Basica: Predecir una imagen de moda
# + [markdown] colab_type="text" id="S5Uhzt6vVIB2"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/es-419/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/es-419/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/es-419/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="Uo47Ynr8gNAU"
# Note: Nuestra comunidad de Tensorflow ha traducido estos documentos. Como las traducciones de la comunidad
# son basados en el "mejor esfuerzo", no hay ninguna garantia que esta sea un reflejo preciso y actual
# de la [Documentacion Oficial en Ingles](https://www.tensorflow.org/?hl=en).
# Si tienen sugerencias sobre como mejorar esta traduccion, por favor envian un "Pull request"
# al siguiente repositorio [tensorflow/docs](https://github.com/tensorflow/docs).
# Para ofrecerse como voluntario o hacer revision de las traducciones de la Comunidad
# por favor contacten al siguiente grupo [<EMAIL> list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs).
# + [markdown] colab_type="text" id="FbVhjPpzn6BM"
# Esta Guia entrena un modelo de red neuronal para clasificar imagenes de ropa como, tennis y camisetas. No hay problema sino entiende todos los detalles; es un repaso rapido de un programa completo de Tensorflow con los detalles explicados a medida que avanza.
#
# Esta Guia usa [tf.keras](https://www.tensorflow.org/guide/keras), un API de alto nivel para construir y entrenar modelos en Tensorflow.
# + colab={} colab_type="code" id="dzLKpmZICaWN"
# TensorFlow y tf.keras
import tensorflow as tf
from tensorflow import keras
# Librerias de ayuda
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# + [markdown] colab_type="text" id="yR0EdgrLCaWR"
# ## Importar el set de datos de moda de MNIST
# + [markdown] colab_type="text" id="DLdCchMdCaWQ"
# Esta guia usa el set de datos de [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist)
# que contiene mas de 70,000 imagenes en 10 categorias. Las imagenes muestran articulos individuales de ropa a una resolucion baja (28 por 28 pixeles) como se ve aca:
#
# <table>
# <tr><td>
# <img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
# alt="Fashion MNIST sprite" width="600">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST samples</a> (by Zalando, MIT License).<br/>
# </td></tr>
# </table>
#
# Moda MNIST esta construida como un reemplazo para el set de datos clasico [MNIST](http://yann.lecun.com/exdb/mnist/)
# casi siempre utilizado como el "Hola Mundo" de programas de aprendizaje automatico (ML) para computo de vision. El set de datos de MNIST contiene imagenes de digitos escrito a mano (0, 1, 2, etc.) en un formato identico al de los articulos de ropa que va a utilizar aca.
#
# Esta guia utiliza Moda MNIST para variedad y por que es un poco mas retador que la regular MNIST. Ambos set de datos son relativamente pequenos y son usados para verificar que el algoritmo funciona como debe.
#
# Aca, 60,000 imagenes son usadas para entrenar la red neuronal y 10,000 imagenes son usadas para evaluar que tan exacto aprendia la red a clasificar imagenes. Pueden acceder al set de moda de MNIST directamente desde TensorFlow. Para importar y cargar el set de datos de MNIST directamente de TensorFlow:
# + colab={} colab_type="code" id="7MqDQO0KCaWS"
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# + [markdown] colab_type="text" id="t9FDsUlxCaWW"
# [link text](https://)Al cargar el set de datos retorna cuatro arreglos en NumPy:
#
# * El arreglo `train_images` y `train_labels` son los arreglos que *training set*—el modelo de datos usa para aprender.
# * el modelo es probado contra los arreglos *test set*, el `test_images`, y `test_labels`.
#
# Las imagenes son 28x28 arreglos de NumPy, con valores de pixel que varian de 0 a 255. Los *labels* son un arreglo de integros, que van del 0 al 9. Estos corresponden a la *class* de ropa que la imagen representa.
#
# <table>
# <tr>
# <th>Label</th>
# <th>Class</th>
# </tr>
# <tr>
# <td>0</td>
# <td>T-shirt/top</td>
# </tr>
# <tr>
# <td>1</td>
# <td>Trouser</td>
# </tr>
# <tr>
# <td>2</td>
# <td>Pullover</td>
# </tr>
# <tr>
# <td>3</td>
# <td>Dress</td>
# </tr>
# <tr>
# <td>4</td>
# <td>Coat</td>
# </tr>
# <tr>
# <td>5</td>
# <td>Sandal</td>
# </tr>
# <tr>
# <td>6</td>
# <td>Shirt</td>
# </tr>
# <tr>
# <td>7</td>
# <td>Sneaker</td>
# </tr>
# <tr>
# <td>8</td>
# <td>Bag</td>
# </tr>
# <tr>
# <td>9</td>
# <td>Ankle boot</td>
# </tr>
# </table>
#
# Cada imagen es mapeada a una unica etiqueta. Ya que los *Class names* no estan incluidoen el dataset, almacenelo aca para usarlos luego cuando se visualicen las imagenes:
# + colab={} colab_type="code" id="IjnLH5S2CaWx"
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + [markdown] colab_type="text" id="Brm0b_KACaWX"
# ## Explore el set de datos
#
# Explore el formato de el set de datos antes de entrenar el modelo. Lo siguiente muestra que hay 60,000 imagenes en el set de entrenamiento, con cada imagen representada por pixeles de 28x28:
# + colab={} colab_type="code" id="zW5k_xz1CaWX"
train_images.shape
# + [markdown] colab_type="text" id="cIAcvQqMCaWf"
# Asimismo, hay 60,000 etiquetas en el set de entrenamiento:
# + colab={} colab_type="code" id="TRFYHB2mCaWb"
len(train_labels)
# + [markdown] colab_type="text" id="YSlYxFuRCaWk"
# Cada etiqueta es un integro entre 0 y 9:
# + colab={} colab_type="code" id="XKnCTHz4CaWg"
train_labels
# + [markdown] colab_type="text" id="TMPI88iZpO2T"
# Hay 10,000 imagenes en el set de pruebas. Otra vez, cada imagen es representada como pixeles de 28x28:
# + colab={} colab_type="code" id="2KFnYlcwCaWl"
test_images.shape
# + [markdown] colab_type="text" id="rd0A0Iu0CaWq"
# Y el set de pruebas contiene 10,000 etiquetas de imagen:
# + colab={} colab_type="code" id="iJmPr5-ACaWn"
len(test_labels)
# + [markdown] colab_type="text" id="ES6uQoLKCaWr"
# ## Pre-procese el set de datos
#
# El set de datos debe ser pre-procesada antes de entrenar la red. Si usted inspecciona la primera imagen en el set de entrenamiento, va a encontrar que los valores de los pixeles estan entre 0 y 255:
# + colab={} colab_type="code" id="m4VEw8Ud9Quh"
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
# + [markdown] colab_type="text" id="Wz7l27Lz9S1P"
# Escale estos valores en un rango de 0 a 1 antes de alimentarlos al modelo de la red neuronal. Para hacero, divida los valores por 255. Es importante que el *training set* y el *testing set* se pre-procesen de la misma forma:
# + colab={} colab_type="code" id="bW5WzIPlCaWv"
train_images = train_images / 255.0
test_images = test_images / 255.0
# + [markdown] colab_type="text" id="Ee638AlnCaWz"
# Para verificar que el set de datos esta en el formato adecuado y que estan listos para construir y entrenar la red, vamos a desplegar las primeras 25 imagenes de el *training set* y despleguemos el nombre de cada clase debajo de cada imagen.
# + colab={} colab_type="code" id="oZTImqg_CaW1"
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
# + [markdown] colab_type="text" id="59veuiEZCaW4"
# ## Construir el Modelo
#
# Construir la red neuronal requiere configurar las capas del modelo y luego compilar el modelo.
# + [markdown] colab_type="text" id="Gxg1XGm0eOBy"
# ### Configurar las Capas
#
# Los bloques de construccion basicos de una red neuronal son las *capas* o *layers*. Las capas extraen representaciones de el set de datos que se les alimentan. Con suerte, estas representaciones son considerables para el problema que estamos solucionando.
#
# La mayoria de aprendizaje profundo consiste de unir capas sencillas.
# La mayoria de las capas como `tf.keras.layers.Dense`, tienen parametros que son aprendidos durante el entrenamiento.
# + colab={} colab_type="code" id="9ODch-OFCaW4"
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation='relu'),
keras.layers.Dense(10, activation='softmax')
])
# + [markdown] colab_type="text" id="gut8A_7rCaW6"
# La primera capa de esta red, `tf.keras.layers.Flatten`,
# transforma el formato de las imagenes de un arreglo bi-dimensional (de 28 por 28 pixeles) a un arreglo uni dimensional (de 28*28 pixeles = 784 pixeles). Observe esta capa como una capa no apilada de filas de pixeles en la misma imagen y alineandolo. Esta capa no tiene parametros que aprender; solo reformatea el set de datos.
#
# Despues de que los pixeles estan "aplanados", la secuencia consiste de dos capas`tf.keras.layers.Dense`. Estas estan densamente conectadas, o completamente conectadas. La primera capa `Dense` tiene 128 nodos (o neuronas). La segunda (y ultima) capa es una capa de 10 nodos *softmax* que devuelve un arreglo de 10 probabilidades que suman a 1. Cada nodo contiene una calificacion que indica la probabilidad que la actual imagen pertenece a una de las 10 clases.
#
# ### Compile el modelo
#
# Antes de que el modelo este listo para entrenar , se necesitan algunas configuraciones mas. Estas son agregadas durante el paso de compilacion del modelo:
#
# * *Loss function* —Esto mide que tan exacto es el modelo durante el entrenamiento. Quiere minimizar esta funcion para dirigir el modelo en la direccion adecuada.
# * *Optimizer* — Esto es como el modelo se actualiza basado en el set de datos que ve y la funcion de perdida.
# * *Metrics* — Se usan para monitorear los pasos de entrenamiento y de pruebas.
# El siguiente ejemplo usa *accuracy* (exactitud) , la fraccion de la imagenes que son correctamente clasificadas.
# + colab={} colab_type="code" id="Lhan11blCaW7"
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="qKF6uW-BCaW-"
# ## Entrenar el Modelo
#
# Entrenar el modelo de red neuronal requiere de los siguientes pasos:
#
# 1. Entregue los datos de entrenamiento al modelo. En este ejemplo , el set de datos de entrenamiento estan en los arreglos `train_images` y `train_labels`.
# 2. el modelo aprende a asociar imagenes y etiquetas.
# 3. Usted le pregunta al modelo que haga predicciones sobre un set de datos que se encuentran en el ejemplo,incluido en el arreglo `test_images`. Verifique que las predicciones sean iguales a las etiquetas de el arreglo`test_labels`.
#
# Para comenzar a entrenar, llame el metodo `model.fit`, es llamado asi por que *fit* (ajusta) el modelo a el set de datos de entrenamiento:
# + colab={} colab_type="code" id="xvwvpA64CaW_"
model.fit(train_images, train_labels, epochs=10)
# + [markdown] colab_type="text" id="W3ZVOhugCaXA"
# A medida que el modelo entrena, la perdida y la exactitud son desplegadas. Este modelo alcanza una exactitud de 0.88 (o 88%) sobre el set de datos de entrenamiento.
# + [markdown] colab_type="text" id="oEw4bZgGCaXB"
# ## Evaluar Exactitud
#
# Siguente, compare como el rendimiento del modelo sobre el set de datos:
# + colab={} colab_type="code" id="VflXLEeECaXC"
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print('\nTest accuracy:', test_acc)
# + [markdown] colab_type="text" id="yWfgsmVXCaXG"
# Resulta que la exactitud sobre el set de datos es un poco menor que la exactitud sobre el set de entrenamiento. Esta diferencia entre el entrenamiento y el test se debe a *overfitting* (sobre ajuste). Sobre ajuste sucede cuando un modelo de aprendizaje de maquina (ML) tiene un rendimiento peor sobre un set de datos nuevo, que nunca antes ha visto comparado con el de entrenamiento.
# + [markdown] colab_type="text" id="xsoS7CPDCaXH"
# ## Hacer predicciones
#
# Con el modelo entrenado usted puede usarlo para hacer predicciones sobre imagenes.
# + colab={} colab_type="code" id="Gl91RPhdCaXI"
predictions = model.predict(test_images)
# + [markdown] colab_type="text" id="x9Kk1voUCaXJ"
# Aca, el modelo ha predecido la etiqueta para cada imagen en el set de datos de *test* (prueba). Miremos la primera prediccion:
# + colab={} colab_type="code" id="3DmJEUinCaXK"
predictions[0]
# + [markdown] colab_type="text" id="-hw1hgeSCaXN"
# *una* prediccion es un arreglo de 10 numeros. Estos representan el nivel de "confianza" del modelo sobre las imagenes de cada uno de los 10 articulos de moda/ropa. Ustedes pueden revisar cual tiene el nivel mas alto de confianza:
# + colab={} colab_type="code" id="qsqenuPnCaXO"
np.argmax(predictions[0])
# + [markdown] colab_type="text" id="E51yS7iCCaXO"
# Entonces,el modelo tiene mayor confianza que esta imagen es un bota de tobillo "ankle boot" o `class_names[9]`. Examinando las etiquetas de *test* o de pruebas muestra que esta clasificaion es correcta:
# + colab={} colab_type="code" id="Sd7Pgsu6CaXP"
test_labels[0]
# + [markdown] colab_type="text" id="ygh2yYC972ne"
# **Grafique** esto para poder ver todo el set de la prediccion de las 10 clases.
# + colab={} colab_type="code" id="DvYmmrpIy6Y1"
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array, true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array, true_label[i]
plt.grid(False)
plt.xticks(range(10))
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + [markdown] colab_type="text" id="d4Ov9OFDMmOD"
# Miremos la imagen [0], sus predicciones y el arreglo de predicciones. Las etiquetas de prediccion correctas estan en azul y las incorrectas estan en rojo. El numero entrega el porcentaje (sobre 100) para la etiqueta predecida.
# + colab={} colab_type="code" id="HV5jw-5HwSmO"
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], test_labels)
plt.show()
# + colab={} colab_type="code" id="Ko-uzOufSCSe"
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions[i], test_labels)
plt.show()
# + [markdown] colab_type="text" id="kgdvGD52CaXR"
# Vamos a graficar multiples imagenes con sus predicciones. Notese que el modelo puede estar equivocado aun cuando tiene mucha confianza.
# + colab={} colab_type="code" id="hQlnbqaw2Qu_"
# Plot the first X test images, their predicted labels, and the true labels.
# Color correct predictions in blue and incorrect predictions in red.
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions[i], test_labels)
plt.tight_layout()
plt.show()
# + [markdown] colab_type="text" id="R32zteKHCaXT"
# Finalmente, usamos el modelo entrenado para hacer una prediccion sobre una unica imagen.
# + colab={} colab_type="code" id="yRJ7JU7JCaXT"
# Grab an image from the test dataset.
img = test_images[1]
print(img.shape)
# + [markdown] colab_type="text" id="vz3bVp21CaXV"
# Los modelos de `tf.keras` son optimizados sobre *batch* o bloques,
# o coleciones de ejemplos por vez.
# De acuerdo a esto, aunque use una unica imagen toca agregarla a una lista:
# + colab={} colab_type="code" id="lDFh5yF_CaXW"
# Add the image to a batch where it's the only member.
img = (np.expand_dims(img,0))
print(img.shape)
# + [markdown] colab_type="text" id="EQ5wLTkcCaXY"
# Ahora prediga la etiqueta correcta para esta imagen:
# + colab={} colab_type="code" id="o_rzNSdrCaXY"
predictions_single = model.predict(img)
print(predictions_single)
# + colab={} colab_type="code" id="6Ai-cpLjO-3A"
plot_value_array(1, predictions_single[0], test_labels)
_ = plt.xticks(range(10), class_names, rotation=45)
# + [markdown] colab_type="text" id="cU1Y2OAMCaXb"
# `model.predict` retorna una lista de listas para cada imagen dentro del *batch* o bloque de datos. Tome la prediccion para nuestra unica imagen dentro del *batch* o bloque:
# + colab={} colab_type="code" id="2tRmdq_8CaXb"
np.argmax(predictions_single[0])
# + [markdown] colab_type="text" id="YFc2HbEVCaXd"
# Y el modelo predice una etiqueta de 2.
| site/es-419/tutorials/keras/classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datascience import *
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
# -
# # Lecture 11 #
# ## Lists
simple_list = ['hello', 7, 3.14, True]
simple_list
# +
my_array = make_array(1, 2, 3)
crowded_list = [my_array, 'what is going on', -10]
crowded_list
# -
Table().with_columns('Numbers', [1, 2, 3])
drinks = Table(['Drink', 'Cafe', 'Price'])
drinks
drinks = drinks.with_rows([
['Milk Tea', 'Asha', 5.5],
['Espresso', 'Strada', 1.75],
['Latte', 'Strada', 3.25],
['Espresso', "FSM", 2]
])
drinks
# ## Grouping by one column
survey = Table.read_table('welcome_survey_v4.csv')
survey.show(3)
survey.group('Sleep position').show()
survey.group('Sleep position', np.average)
(survey.select('Sleep position', 'Hours of sleep').group('Sleep position', np.average)
# ## Cross-classification: grouping by two columns
survey.group(['Handedness','Sleep position']).show()
survey.pivot('Sleep position', 'Handedness')
survey.pivot('Sleep position', 'Handedness', 'Hours of sleep', np.average)
(survey.select('Handedness', 'Sleep position', 'Hours of sleep')
.group(['Handedness','Sleep position'], np.average)).show()
# Here, pivot doesn't know how to combine all the hours of sleep
# for each subgroup of students
survey.pivot('Sleep position', 'Handedness', 'Hours of sleep')
# ## Challenge Question ##
sky = Table.read_table('skyscrapers_v2.csv')
sky = (sky.with_column('age', 2020 - sky.column('completed'))
.drop('completed'))
sky.show(3)
# +
# 1. For each city, what’s the tallest building for each material?
# +
# 2. For each city, what’s the height difference between the tallest
# steel building and the tallest concrete building?
# -
# Don't read ahead until you try the challenge questions yourself first!
sky.select('material', 'city', 'height').group(['city', 'material'], max)
sky_p = sky.pivot('material', 'city', 'height', max)
sky_p.show()
sky_p = sky_p.with_column(
'difference',
abs(sky_p.column('steel') - sky_p.column('concrete'))
)
sky_p
sky_p.sort('difference', True)
# ### Take-home question: try it here!
# +
# Generate a table of the names of the oldest buildings for each
# material for each city:
# -
# ## Joins ##
drinks
discounts = Table().with_columns(
'Coupon % off', make_array(10, 25, 5),
'Location', make_array('Asha', 'Strada', 'Asha')
)
discounts
combined = drinks.join('Cafe', discounts, 'Location')
combined
discounted_frac = 1 - combined.column('Coupon % off') / 100
combined.with_column(
'Discounted Price',
combined.column('Price') * discounted_frac
)
drinks.join('Cafe', drinks, 'Cafe')
| lec/lec11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import scipy as sp
import scipy.stats
import itertools
import logging
import matplotlib.pyplot as plt
import pandas as pd
import torch.utils.data as utils
import math
import time
import tqdm
import torch
import torch.optim as optim
import torch.nn.functional as F
from argparse import ArgumentParser
from torch.distributions import MultivariateNormal
import torch.nn as nn
import torch.nn.init as init
# -
# # use 3 prong loss and purebkg on (rnd2prong, rndbkg)
# +
bkgaeloss_3prong = np.load(f'../data_strings/bkgae_purebkg_loss_2prong.npy')
sigaeloss_3prong = np.load(f'../data_strings/sigae_3prong_loss_2prong.npy')
print(len(bkgloss_3prong),len(sigaeloss_3prong))
# +
bkgaeloss_purebkg = np.load(f'../data_strings/bkgae_purebkg_loss_purebkg.npy')
sigaeloss_purebkg = np.load(f'../data_strings/sigae_3prong_loss_purebkg.npy')
print(len(bkgloss_purebkg), len(sigaeloss_purebkg))
# -
correct = np.where( (sigaeloss_3prong<100) & (bkgaeloss_3prong<100))[0]
bkgaeloss_3prong = bkgaeloss_3prong[correct]
sigaeloss_3prong = sigaeloss_3prong[correct]
print(len(bkgloss_3prong),len(sigaeloss_3prong))
correct = np.where( (sigaeloss_purebkg<100) & (bkgaeloss_purebkg<100))[0]
correct.shape
bkgaeloss_purebkg = bkgaeloss_purebkg[correct]
sigaeloss_purebkg = sigaeloss_purebkg[correct]
print(len(bkgaeloss_purebkg),len(sigaeloss_purebkg))
plt.scatter(bkgloss_3prong[:300], sigaeloss_3prong[:300], marker='.', color='red',alpha=0.2)
plt.scatter(bkgloss_purebkg[:300], sigaeloss_purebkg[:300], marker='x', color='blue',alpha=0.2)
plt.xlim(0, 2)
plt.ylim(0, 10)
X_bkg = np.column_stack((bkgaeloss_purebkg,sigaeloss_purebkg))
X_sig = np.column_stack((bkgaeloss_3prong,sigaeloss_3prong))
X = np.vstack((X_bkg,X_sig))
X.shape
# +
y_bkg = np.zeros(908999)
y_sig = np.ones(58821)
Y = np.concatenate((y_bkg,y_sig),axis=0)
# -
Y.shape
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis, QuadraticDiscriminantAnalysis
clf = LinearDiscriminantAnalysis()
clf.fit(X, Y)
X_r2 = clf.transform(X)
X_r2.shape
plt.hist(X_r2[Y==1],bins = np.linspace(-5,5,100),alpha=0.3);
plt.hist(X_r2[Y==0],bins = np.linspace(-5,5,100),alpha=0.3);
plt.hist(sigaeloss_3prong,bins = np.linspace(0,5,100),alpha=0.3);
plt.hist(sigaeloss_purebkg,bins = np.linspace(0,5,100),alpha=0.3);
f_rnd = pd.read_hdf("/data/t3home000/spark/QUASAR/preprocessing/conventional_tau_rnd.h5")
f_3prong = pd.read_hdf("/data/t3home000/spark/QUASAR/preprocessing/conventional_tau_3prong_rnd.h5")
f_purebkg = pd.read_hdf("/data/t3home000/spark/QUASAR/preprocessing/conventional_tau_background.h5")
f_BB1 = pd.read_hdf("/data/t3home000/spark/QUASAR/preprocessing/conventional_tau_BB1_rnd.h5")
f_BB2 = pd.read_hdf("/data/t3home000/spark/QUASAR/preprocessing/conventional_tau_BB2.h5")
f_BB3 = pd.read_hdf("/data/t3home000/spark/QUASAR/preprocessing/conventional_tau_BB3.h5")
dt = f_BB1.values
# +
correct = (dt[:,3]>0) &(dt[:,19]>0)
dt = dt[correct]
for i in range(13,19):
dt[:,i] = dt[:,i]/dt[:,3]
for i in range(29,35):
dt[:,i] = dt[:,i]/(dt[:,19])
correct = (dt[:,29]>=0) &(dt[:,29]<=1)&(dt[:,30]>=0) &(dt[:,30]<=1)&(dt[:,31]>=0) &(dt[:,31]<=1)&(dt[:,32]>=0) &(dt[:,32]<=1)&(dt[:,33]>=0) &(dt[:,33]<=1)&(dt[:,34]>=-0.01) &(dt[:,34]<=1)
dt = dt[correct]
# -
bb_mass = dt[:,0]
dt_rnd = f_rnd.values
rnd_idx = dt_rnd[:,-1]
# +
#improve purity of trainig by only selecting properly reconstructed jets
# -
plt.hist(dt_rnd[rnd_idx==1,3])
# # APPLY PRESELECTION BEFORE DOING THE FIT TO THE LDA
np.finfo(np.float32).max
bkgloss_0 = np.load(f'../data_strings/bkgae_rndbkg_loss_bb1.npy')
sigaeloss_0 = np.load(f'../data_strings/sigae_2prong_loss_bb1.npy')
def generate_X(whichsample):
#bkgloss_0 = np.load(f'../data_strings/bkgae_rndbkg_loss_{whichsample}.npy')
bkgloss_1 = np.load(f'../data_strings/bkgae_purebkg_loss_{whichsample}.npy')
sigaeloss_0 = np.load(f'../data_strings/sigae_2prong_loss_{whichsample}.npy')
sigaeloss_1 = np.load(f'../data_strings/sigae_3prong_loss_{whichsample}.npy')
sigaeloss_0[sigaeloss_0 > 2000 ] = 2000
sigaeloss_1[sigaeloss_1 > 2000 ] = 2000
#print(np.max(bkgloss_0))
print(np.max(bkgloss_1))
print(np.max(sigaeloss_0))
print(np.max(sigaeloss_1))
return np.column_stack((bkgloss_1,sigaeloss_0,sigaeloss_1))
X = np.vstack((generate_X('rndbkg'),generate_X('purebkg'),generate_X('2prong'),generate_X('3prong')))
X.shape
def get_len(whichsample):
bkgloss_0 = np.load(f'../data_strings/bkgae_{whichsample}_loss_rndbkg.npy')
bkgloss_1 = np.load(f'../data_strings/bkgae_{whichsample}_loss_purebkg.npy')
sigaeloss_0 = np.load(f'../data_strings/sigae_2prong_loss_2prong.npy')
sigaeloss_1 = np.load(f'../data_strings/sigae_2prong_loss_3prong.npy')
return len(bkgloss_0), len(bkgloss_1), len(sigaeloss_0), len(sigaeloss_1)
get_len('rndbkg')
y_rndbkg = np.zeros(938331)
y_purebkg = np.zeros(930647)
y_2prong = np.ones(57042)
y_3prong = np.ones(58822)
Y = np.concatenate((y_rndbkg,y_purebkg,y_2prong,y_3prong),axis=0)
Y.shape
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis, QuadraticDiscriminantAnalysis
clf = QuadraticDiscriminantAnalysis()
clf.fit(X, Y)
BB1 = generate_X('bb1')
BB1.shape
BB1_prob = clf.predict_proba(BB1)
https://mit.zoom.us/j/7287742919?pwd=<PASSWORD>VGpLNFQ5R0ZKS0pvWlZQRzcxdz09
bb1_index = np.where(BB1_prob[:,1]>0.91)[0]
# +
#bb1_index = np.where((BB1[:,0]>1.5) & (BB1[:,1]>1.5) & (BB1[:,2]<3.)&(BB1[:,3]<4.))[0]
# -
len(bb1_index)
bb1_index
plt.hist(bb_mass[bb1_index],bins=np.linspace(2800,7000,33),alpha=0.3);
plt.axvline(x=3823)
from scipy.stats import percentileofscore
percentiles = [percentileofscore(BB1[:,2], i) for i in BB1[:,2]]
plt.scatter(BB1[:,0],BB1_prob[:,1])
plt.xscale('log')
def get_loss(X):
| training/LDA_test_on_BB1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data download and basic preparation
# ### <NAME>
# ---
# This script uses the modified Yahoo! Finance api (through the `yfinance` library) to access and download historical data from selected markets. It also saves the original data (which consists on daily observations) and a copy of yearly observations as `.csv` files.
# Import the required libraries
import glob
import os
import pandas as pd
import yfinance as yf
# The selected market data to download is passed to the script via the `tickers` dictionary. The key of the dictionary is a descriptive name for the data (for example, `sp500`), and the value of the dictionary is the corresponding symbol used at the Yahoo! Finance website (for example, `^GSPC`). Both are `string`s.
# Dictionary of data to download
tickers = {
'ftse100': '^FTSE',
'ni225': '^N225',
'sp500': '^GSPC'
}
# ## Download data
# The original (daily) data is saved in a `downloads` folder as `name.csv`, where each value of `name` is a key of the dictionary `tickers`.
# +
results = {ticker: None for ticker in tickers}
for ticker in tickers.keys():
# Download data
t = yf.Ticker(tickers[ticker]).history(period = 'max')
# Keep only the closing price
t = pd.DataFrame(t['Close'])
# Keep only the values up to 2020
t = t.query('Date <= "2020-12-31"')
# Save the Data Frame to a .csv file
t.to_csv(f'downloads/{ticker}.csv')
# Report number of results found
results[ticker] = len(t)
# -
# Show number of observations for each ticker
for ticker in results:
print(f'{ticker}: {str(results[ticker])} observations found.')
# ## Convert to yearly observations
# A copy of yearly data is saved in a `downloads` folder as `name_yearly.csv`, where each value of `name` is a key of the dictionary `tickers`.
# +
# Get list of files
files = glob.glob('downloads/*.csv')
# Show the list of files
for file in files:
print(file)
# -
for file in files:
# Get data name:
loc1 = file.find('/') + 1
loc2 = file.find('.')
name = file[loc1:loc2]
#Load data
data = pd.read_csv(file)
# Convert 'Date' column to Datetime type
data['Date'] = pd.to_datetime(data['Date'])
# Create a list 'years' with the years available in the data
years = set([date.year for date in data['Date']])
years = list(years)
years.sort()
# Set the column 'Date' as the index of the data frame
# This is required to use the .last() method
data = data.set_index('Date')
# Loop through the years and extract the last day info
# Each will be stored in a Data Frame inside a list named ' last_data'
last_data = []
for year in years:
last = data.query(f'Date >= "{year}-01-01" and Date <= "{year}-12-31"').last(offset = '1D')
last_data.append(pd.DataFrame(last))
# Join the extracted data in a single Data Frame called ' closing'
closing = pd.concat(last_data)
# Save the Data Frame in a .csv file
closing.to_csv(f'downloads/{name}_yearly.csv')
| script_get_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="R0AV1AZygDzp" colab_type="text"
# # Introduction
#
# This project is completed as part of the [Kaggle's Titanic Competition](https://www.kaggle.com/c/titanic).
#
#
# The goal of this project to predict what sorts of people were likely to survive.
#
# ---
#
# + id="MXb-rRXGgv1J" colab_type="code" colab={}
# Importing libraries
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
import matplotlib.pyplot as plt
import matplotlib.pylab as pylab
import seaborn as sns
from numpy import percentile
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
from xgboost import XGBClassifier
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_validate
from sklearn.model_selection import cross_val_score
# + id="LzN2L789g526" colab_type="code" colab={}
# Loading the datasets
train = pd.read_csv('train.csv')
test = pd.read_csv('test.csv')
# + [markdown] id="a3MGCSmQi_EY" colab_type="text"
# # 1. Exploratory Analysis with Pandas
# + [markdown] id="_fon9oeHhJiN" colab_type="text"
# Checking the data dimension.
# + id="HnFLQANvhMY0" colab_type="code" outputId="9dcca4e8-7492-4b72-9132-78ac73b73070" colab={"base_uri": "https://localhost:8080/", "height": 34}
train.shape
# + id="lPqlS8cshXX7" colab_type="code" outputId="ebd936f8-900b-45e9-cf40-64438f5c6eec" colab={"base_uri": "https://localhost:8080/", "height": 306}
# General information about the data
train.info()
# + [markdown] id="b8Gm9iXChatl" colab_type="text"
# **int64**, **object**, and **float64** are the types of the features.
# + id="z_Qg_bEXhhpN" colab_type="code" outputId="96db3ffa-5834-4e86-934e-d09453751be5" colab={"base_uri": "https://localhost:8080/", "height": 297}
# Statistical characteristics of numerical features
train.describe()
# + id="Hm2tQdnKhlwX" colab_type="code" outputId="a4d57451-6927-4741-ae7c-6a1147f6b0f1" colab={"base_uri": "https://localhost:8080/", "height": 173}
# Statistical characteristics of categorical features
train.describe(include=['object'])
# + [markdown] id="mPnmeWfkhpz_" colab_type="text"
# Let's see, how many male or female survived?
# + id="yjzetv8KhrDi" colab_type="code" outputId="dc368ad3-172b-4d4a-c035-55381182578f" colab={"base_uri": "https://localhost:8080/", "height": 142}
pd.crosstab(train['Survived'], train['Sex'])
# + [markdown] id="mNR3OBkAhv4w" colab_type="text"
# 233 females and 109 males were survived. It becomes 68% females and 32% males who survived.
# + [markdown] id="xKYCsSSZh33F" colab_type="text"
# Let's see, what age of people mostly survived?
# + id="1KxPNI2ph9Ma" colab_type="code" outputId="4cf74305-8965-4c3d-b022-67fc84dc079d" colab={"base_uri": "https://localhost:8080/", "height": 142}
pd.crosstab(train['Survived'], train['Age'] <= 15)
# + [markdown] id="oXmDXqG0iBDx" colab_type="text"
# There were 83 children on the ship with age less than or equal to 15 years. Among them 59% Survived
# + id="y4DkWtE1iHPj" colab_type="code" outputId="17282d8f-799b-47eb-e6cc-41b71862e810" colab={"base_uri": "https://localhost:8080/", "height": 142}
pd.crosstab(train['Survived'], (train['Age'] > 15) & (train['Age'] <= 35))
# + [markdown] id="OIpVDr7uiS49" colab_type="text"
# There were 414 people with age range 16 to 35 in which 38% Survived.
# + id="Xu-NR4WqiXrf" colab_type="code" outputId="c5807481-468d-460e-923f-682de0eea2ec" colab={"base_uri": "https://localhost:8080/", "height": 142}
pd.crosstab(train['Survived'], (train['Age'] > 35) & (train['Age'] <= 90))
# + [markdown] id="oF8znaa6iblg" colab_type="text"
# There were 217 people in middle or old ages in which 38% Survived.
#
# So, we come to the conclusion that, mostly children were survived in the incident.
#
#
#
# ---
# + [markdown] id="3ES8pI79izcu" colab_type="text"
# # 2. Visual Exploratory Analysis
#
# First, we'll check whether the numerical features are normally distributed or not.
# + id="x6QHvb0GjNz-" colab_type="code" outputId="b50b336f-2d45-4f47-fc27-e1b7c3c82ce9" colab={"base_uri": "https://localhost:8080/", "height": 361}
sns.distplot(train['Pclass']);
# + id="SkKYNxUQjSVD" colab_type="code" outputId="b66a471e-b7ac-4efc-f3f5-4545b385232d" colab={"base_uri": "https://localhost:8080/", "height": 361}
sns.distplot(train['SibSp']);
# + id="ohVbpgROjUxJ" colab_type="code" outputId="afb8f50f-d8ca-4bd1-c96d-95ef242e3739" colab={"base_uri": "https://localhost:8080/", "height": 361}
sns.distplot(train['Fare']);
# + [markdown] id="ZNtph0ZMjc7l" colab_type="text"
# From the plots, we can see that the dataset is not normally distributed.
# + id="u0M2pkI2jhYP" colab_type="code" outputId="ee089b26-fa66-4382-baff-a74ffd7cd205" colab={"base_uri": "https://localhost:8080/", "height": 279}
# Checking the frequency distribution of the target variable
train['Survived'].value_counts()
_, axes = plt.subplots(nrows=1, ncols=1, figsize=(12, 4))
sns.countplot(x='Survived', data=train, ax=axes);
# + [markdown] id="XYsc4GNTjo_V" colab_type="text"
# Let’s look at the correlations among the numerical variables in our dataset.
# + id="6hXw8ErAjsJj" colab_type="code" outputId="797e1efe-8dba-494b-8cea-1dc310377adc" colab={"base_uri": "https://localhost:8080/", "height": 347}
# Drop non-numerical variables
numerical = list(set(train.columns) - set(['Survived', 'Name', 'Sex', 'Ticket', 'Cabin', 'Embarked']))
# Calculate and plot
corr_matrix = train[numerical].corr()
sns.heatmap(corr_matrix);
# + [markdown] id="ecfBEulFk_FW" colab_type="text"
# we can see the correlation between **Parch** and **SibSp**.
#
# Sometimes you can analyze an ordinal variable as numerical one:
# + id="VsycO_hVlJTH" colab_type="code" outputId="5ef1f612-10e8-48c0-da2a-be7a51c42fe8" colab={"base_uri": "https://localhost:8080/", "height": 509}
fig, axes = plt.subplots(nrows=2, ncols=4, figsize=(10, 7))
for idx, feat in enumerate(numerical):
ax = axes[int(idx / 4), idx % 4]
sns.boxplot(x='Survived', y=feat, data=train, ax=ax)
ax.set_xlabel('')
ax.set_ylabel(feat)
fig.tight_layout();
# + [markdown] id="gJRVJ11ElUuF" colab_type="text"
# The above subplots shows some interesting facts.
#
# 1. People with 8 and 5 siblings on the ship didn't survive at all.
# 2. Passenger class (Pclass) Lower class (3) didn't survive at all.
#
# ---
#
# # 3. Data Preprocessing
# + [markdown] id="ZrutH3QsnOt_" colab_type="text"
# ## 3.1. Handling Missing Values
# + id="-dm9bOdfmavR" colab_type="code" outputId="5d87890c-8d73-4dc2-b67f-b4183da928d7" colab={"base_uri": "https://localhost:8080/", "height": 238}
# Checking for missing values
train.isnull().sum()
# + [markdown] id="2t-QSHdWmp_p" colab_type="text"
# Training dataset have missing values in Age (float64) and Embarked (object).
# + id="IM42ixbWmlVv" colab_type="code" outputId="45caa5bd-1702-47e0-95f7-456805a42abc" colab={"base_uri": "https://localhost:8080/", "height": 221}
test.isnull().sum()
# + [markdown] id="_JK556I7muV_" colab_type="text"
# Test set have missing values in Age (float64) and Fare (float64)
# + id="hgs7PtU6myi_" colab_type="code" colab={}
# Function to handle numerical features missing values
def handle_num_missing_values(dataset, columns):
for i in range(0, len(columns)):
dataset[columns[i]].fillna(dataset[columns[i]].median(), inplace=True)
# Function to handle categorical features missing values
def handle_obj_missing_values(dataset, columns):
for i in range(0, len(columns)):
dataset[columns[i]].fillna(method='bfill', inplace=True)
handle_num_missing_values(train, ['Age'])
handle_num_missing_values(test, ['Age', 'Fare'])
handle_obj_missing_values(train, ['Embarked'])
# + [markdown] id="PDmOlKI6nJu9" colab_type="text"
# ## 3.2. Feature Engineering
# + [markdown] id="AfuYfRNxncSk" colab_type="text"
# We'll create a feature from **Cabin**, if empty Cabin, assign feature value as *0*, else *1*.
# + id="uLfAk_Bdnvdz" colab_type="code" outputId="8bce3f28-c7ca-48e2-9fe3-78812d0a1a6c" colab={"base_uri": "https://localhost:8080/", "height": 179}
train['HasCabin'] = train['Cabin'].where(train['Cabin'].isnull(), 1).fillna(0).astype(int)
test['HasCabin'] = test['Cabin'].where(test['Cabin'].isnull(), 1).fillna(0).astype(int)
train.head(2)
# + id="ZZegECSQoB96" colab_type="code" colab={}
def feature_engineering(dataset):
# Create new feature 'Title' from 'Name'
dataset['Title'] = dataset['Name'].str.split(',', expand=True)[1].str.split('.', expand=True)[0]
# Create new feature 'FamilySize', by adding SibSp, Parch, 1 (The Passenger)
dataset['FamilySize'] = dataset['SibSp'] + dataset['Parch'] + 1
# Create new feature 'IsAlone'
dataset['IsAlone'] = 1
dataset['IsAlone'].loc[dataset['FamilySize'] > 1] = 0
# Create Age bins
dataset['AgeBin'] = pd.cut(dataset['Age'].astype(int), 4, labels=['1 -20', '20-40', '40-60', '60-80'])
# Create Fare Bin
dataset['FareBin'] = pd.qcut(dataset['Fare'], 4, labels=['1-200', '200-400', '400-500', '500-600'])
return dataset
train, test = feature_engineering(train), feature_engineering(test)
# + id="0edXw8GdoSbG" colab_type="code" outputId="9e97c844-a936-4452-9124-ceadd1ee6c09" colab={"base_uri": "https://localhost:8080/", "height": 267}
train.head(2)
# + [markdown] id="dlXGauzxoccj" colab_type="text"
# We have created 5 new features from the existing features.
#
# 1. Title: Splitting the titles from name e-g, Mr, Ms, Dr.
# 2. FamilySize: Adding SibSp (Siblings etc) plus Parch(Parents etc) + the pessengers.
# 3. IsAlone: Create a new column to check whether the passenger is alone or not on the ship.
# 4. AgeBin: Create a new feature containing age ranges that describes in which age range the passenger fall into.
# 5. FareBin: Similary, create a 'FareBin' feature.
#
# Let's check the unique title in the '**Title**' feature
# + id="DVg-IIRCooxt" colab_type="code" outputId="0f9a69ff-243a-4451-b330-87a02537b767" colab={"base_uri": "https://localhost:8080/", "height": 323}
train['Title'].value_counts()
# + [markdown] id="D6V7gY7-os2u" colab_type="text"
# There are 14 unique values. So we're going to give a unique name 'Misc' to the titles which are occurring less than 10 times.
# + id="UyRmyYR4oxo5" colab_type="code" outputId="dc08bf43-ed5a-43ab-83af-ea54d114bb63" colab={"base_uri": "https://localhost:8080/", "height": 315}
# Function to clean up rare titles
def clean_titles(dataset):
title_names = dataset['Title'].value_counts() < 10
dataset['Title'] = dataset['Title'].apply(lambda x: 'Misc' if title_names.loc[x] == True else x)
return dataset
train = clean_titles(train)
test = clean_titles(test)
train['Title'].value_counts()
train.head(3)
# + [markdown] id="xJS3wGHUo6j8" colab_type="text"
# We'll create 2 new features (**Last Name and Frequency of Last Name**) from the **Name** column.
# + id="apAt_m8hpIOO" colab_type="code" outputId="d5c55e09-a3fe-4d1c-98b9-0e6117b274e6" colab={"base_uri": "https://localhost:8080/", "height": 496}
def get_frequence_of_last_names(dataset):
dataset['LName'] = dataset['Name'].str.split(',', expand=True)[0]
dataset['LName_freq'] = dataset['LName'].map(dataset['LName'].value_counts())
return dataset
train, test = get_frequence_of_last_names(train), get_frequence_of_last_names(test)
train.head(5)
# + [markdown] id="p4SmFv3OpYYp" colab_type="text"
# Splitting the target varible from the training set.
# + id="Q7M4_nN7pagw" colab_type="code" colab={}
y_train = train.iloc[:, 1].values
# + [markdown] id="QrByD3tJpeXQ" colab_type="text"
# Removing unnecessary features.
# + id="YGY_GwZfphuR" colab_type="code" colab={}
train.drop(['Name', 'LName', 'Survived', 'PassengerId', 'Ticket', 'Cabin'], inplace=True, axis=1)
test.drop(['Name', 'LName', 'PassengerId', 'Ticket', 'Cabin'], inplace=True, axis=1)
# + id="838cqLnypjEO" colab_type="code" outputId="2e2a7ebe-77e1-42f0-ed2e-b2fc980f6de6" colab={"base_uri": "https://localhost:8080/", "height": 224}
train.head(5)
# + [markdown] id="1YWKQxNPrHzZ" colab_type="text"
# ## 3.3. Encoding
#
# We are going to label encode Sex, Embarked, Title, AgeBin, and FareBin features.
#
# **Note:** This is one of the most important data preprocessing part. So, please stick with me on this.
#
# **Problem:**
#
# When you apply encoding and scaling on training and test set. You apply *`fit_transform()`* to the training set and *`transform()`* to the test set.
#
# But, the problem is that using *`fit_transform()`*, model learns parameters using training set. So, when you apply *`transform()`*, you might face an error:
#
# *`ValueError: Unseen labels or values`*
#
# This error happens when test set have new unseen values that your training set don't have. So, you'll have to take care of the error.
# + id="ET9j9yn4s3LD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 162} outputId="b1544e4a-253e-4a15-d5f9-c3422ac7cab3"
# Label Encoding
le = LabelEncoder()
train['Sex'] = le.fit_transform(train['Sex'])
test['Sex'] = le.transform(test['Sex'])
train['Embarked'] = le.fit_transform(train['Embarked'])
test['Embarked'] = le.transform(test['Embarked'])
train['Title'] = le.fit_transform(train['Title'])
test['Title'] = le.transform(test['Title'])
train['AgeBin'] = le.fit_transform(train['AgeBin'])
test['AgeBin'] = le.transform(test['AgeBin'])
train['FareBin'] = le.fit_transform(train['FareBin'])
test['FareBin'] = le.transform(test['FareBin'])
train.head(3)
# + id="4oA2QB6ts9mK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 191} outputId="c5f0f83f-280f-4da9-fc23-0682266ea9f4"
# One Hot Encoding
def one_hot(df, cols):
for each in cols:
dummies = pd.get_dummies(df[each], prefix=each, drop_first=True)
#dummies = pd.get_dummies(df[each], prefix=each)
df = pd.concat([df, dummies], axis=1)
return df
train = one_hot(train, cols = ['Pclass', 'Embarked', 'Title', 'AgeBin', 'FareBin'])
test = one_hot(test, cols = ['Pclass', 'Embarked', 'Title', 'AgeBin', 'FareBin'])
train.head(3)
# + [markdown] id="aPFJXnrJtOH_" colab_type="text"
# **Note:** I have removed first variable for avoiding the dummy variable trap.
#
# ## 3.4. Feature Scaling
# + id="yZ5kM9khte6u" colab_type="code" colab={}
sc = StandardScaler()
X_train = sc.fit_transform(train.values)
X_test = sc.transform(test.values)
# + id="Lada9EGctllk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="9f308672-be66-4e5b-e035-ee667e027d02"
X_train[3, :]
# + [markdown] id="p2YogA-ot29G" colab_type="text"
# # 4. Modeling
#
# I'm going to implement 3 models to the dataset: XGBoost, Support Vector Machines, and Logistic Regression.
#
# **Note:** Parameter tuning has already been done using GridSearchCV.
#
# ## 4.1. XGBoost
# + id="h7NzW5xmuqXq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="506b8524-12f9-4bb9-9363-1da2d2c7a735"
clf_1 = XGBClassifier(learning_rate = 0.01, gamma = 0.01, max_depth = 4, n_estimators = 200, subsample = 1,
colsample_bytree = 1, objective = 'binary:logistic', random_state=0)
clf_1.fit(X_train, y_train)
y_pred_1 = clf_1.predict(X_test)
y_pred_1
# + [markdown] id="cca8H2wRx2Hz" colab_type="text"
# Above are the predictions I got from the XGBoost model.
#
# ***Kaggle Score: 0.77990***
# + [markdown] id="ya_q46_VyRVw" colab_type="text"
# ## 4.2. Logistic Regression
# + id="hPoqwxe0yWjZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="29923597-7739-4936-abaa-7bd26caee7e5"
clf_2 = LogisticRegression(C = 0.1, max_iter = 10, penalty = 'l2', solver = 'newton-cg')
clf_2.fit(X_train, y_train)
y_pred_2 = clf_2.predict(X_test)
y_pred_2
# + [markdown] id="bYLCD75WyyMm" colab_type="text"
# ***Kaggle Score: 0.77033***
# + [markdown] id="fDg-lmDMy_Nk" colab_type="text"
# ## 4.3. Support Vector Machines
# + id="oO8-GyzRzDiv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="736157e2-e102-4e8e-aab8-ac8e8fc65351"
clf_3 = SVC(C = 10, gamma = 0.01, kernel = 'rbf', random_state = 0)
clf_3.fit(X_train, y_train)
y_pred_3 = clf_3.predict(X_test)
y_pred_3
# + [markdown] id="bftAeYvnzRvr" colab_type="text"
# ***Kaggle Score: 0.77511***
#
# # Conclusion
#
# XGboost performed better than SVM and Logistic Regression. I could have make the results better by doing some more feature engineering and parameter tuning.
#
# **Note:** No cheating was done in the competition, because I know how to cheat and get the score 1. :)
| titanic/titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="pg7sF9FdJ96E"
import keras
import os
model=keras.models.load_model('/content/drive/MyDrive/All fruits/temp4/FruitsBestModel2.h5')
# + colab={"base_uri": "https://localhost:8080/"} id="qpzI96ULPo3Z" outputId="ae7ea638-e8a3-49d0-81f8-c58f04da5395"
label_fruit_map={}
for index,fruit in enumerate(fruit_lt):
label_fruit_map[index]=fruit
label_fruit_map
# + colab={"base_uri": "https://localhost:8080/"} id="dEuC6O6XMvaC" outputId="b350962a-8303-4a33-c99f-a603a0688033"
fruit_label_map={}
for key,value in label_fruit_map.items():
fruit_label_map[value]=key
fruit_label_map
# + colab={"base_uri": "https://localhost:8080/"} id="2U-9kLbmKBPb" outputId="34a0dc2a-cce2-4e2e-920d-789a0fd5f988"
new_imgs=os.listdir('/content/drive/MyDrive/All fruits/Evaluation fruit')
new_imgs,len(new_imgs)
# + id="Ngw19fbUKRzA"
import cv2
import numpy as np
def get_preprocessed_imgs(dir_path):
lt=[]
imgs=[]
imgs_lt=os.listdir(dir_path)
for img in imgs_lt:
image=cv2.imread(os.path.join(dir_path,img))
try:
resized_img=cv2.resize(image,(200,200))
lt.append(resized_img)
imgs.append(img)
except:
print(img)
return np.array(lt),imgs
# + colab={"base_uri": "https://localhost:8080/"} id="SM0XjP70LKsh" outputId="c949284f-ee61-49d7-d80a-64bf18d23279"
image_data,image_names=get_preprocessed_imgs('/content/drive/MyDrive/All fruits/Evaluation fruit')
image_data.shape,len(image_names)
# + colab={"base_uri": "https://localhost:8080/"} id="qZfeZtthLS3d" outputId="94d199c4-34b6-414e-84b3-fcfef0016155"
y_pred=model.predict(image_data)
y_pred
# + colab={"base_uri": "https://localhost:8080/"} id="Pktz5z22L2QC" outputId="e7367bb4-d105-4e00-c26f-646a67c7cb87"
pred_classes=np.argmax(y_pred,axis=1)
pred_classes
# + colab={"base_uri": "https://localhost:8080/"} id="WGHw5V1wPwI4" outputId="9299acf6-3c69-4655-b943-b3f9193c533c"
label_fruit_map.values()
# + colab={"base_uri": "https://localhost:8080/"} id="FiUKf1ZoMYaC" outputId="6fd14e1a-d449-4cca-c9bf-226210edac61"
count=0
wrong_predicted_imgs=[]
wrong_true_label=[]
wrong_predicted_label=[]
for index,val in enumerate(pred_classes):
img_name=image_names[index].split('.')[0][:-1].lower()
label=label_fruit_map[val].split('_')[0].lower()
if(img_name==label or label.startswith(img_name) or img_name.startswith(label)):
count+=1
else:
wrong_predicted_imgs.append(index)
wrong_predicted_label.append(label_fruit_map[val])
wrong_true_label.append(image_names[index])
print(f'Label: {label} img_name: {img_name} img_index: {index}')
print(count)
#print(f'{label_fruit_map[val]}-->{image_names[index]}')
#Cherry_fruit-->Acerola1.jpg,Pummelo_fruit-->Apple1.jpg,
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="C0UPqQCIMmN3" outputId="f0cfbcfb-1fde-4297-fd10-6a5e5be8b7f4"
image_names[0].split('.')[0][:-1].lower()
# + colab={"base_uri": "https://localhost:8080/"} id="qS2LSJIIRTIH" outputId="84e9c10b-fdcf-4f45-e932-317fef532762"
wrong_predicted_imgs[0],wrong_true_label[0],wrong_predicted_label[0]
# + id="ois-gpRaOKzM"
from google.colab.patches import cv2_imshow
dir_path='/content/drive/MyDrive/All fruits/Evaluation fruit'
images=[]
for img in wrong_true_label[5:]:
img_path=os.path.join(dir_path,img)
img=cv2.imread(img_path)
resized=cv2.resize(img,(200,200))
images.append(resized)
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="M7i75eUFSq_b" outputId="4bb4e905-fbf0-4388-91b4-f0257552db3d"
cv2_imshow(images[0]),wrong_predicted_label[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="6Wt2WaHlSuAl" outputId="16da73fc-917c-4828-e4a8-922c2987e08f"
cv2_imshow(images[1]),wrong_predicted_label[1]
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="gwhBaJjZS5Dg" outputId="9d92f856-606a-4e18-96a3-19a0b0b777a3"
cv2_imshow(images[2]),wrong_predicted_label[2]
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="LwxYHsjGS6CN" outputId="6d6fc6bf-0a00-44c5-d1c7-1d41e67431fe"
cv2_imshow(images[3]),wrong_predicted_label[3]
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="igaC-vWTS6g8" outputId="becf408a-f7b3-4692-d179-914f016bac27"
cv2_imshow(images[4]),wrong_predicted_label[4]
# + [markdown] id="e0Jq3u6HBYf1"
# Prepare Target
# + colab={"base_uri": "https://localhost:8080/"} id="q8IX9yU1_vkY" outputId="1ee5504b-9067-40c2-af8f-9da677bfe2a0"
folder_path='/content/drive/MyDrive/All fruits/temp1'
fruit_lt=os.listdir(folder_path)
label=[]
for i in range(len(fruit_lt)):
file_path=os.path.join(folder_path,fruit_lt[i])
fruit_data=np.load(open(file_path,'rb'))
print(fruit_data.shape)
label.extend([i]*fruit_data.shape[0])
# + colab={"base_uri": "https://localhost:8080/"} id="jplhoCmt_vkd" outputId="d3e94b9c-3732-4bc8-c880-7bf9a34b0920"
print(len(label))
# + colab={"base_uri": "https://localhost:8080/"} id="Tqf_TUJT_vkf" outputId="637d2928-be44-4178-9b3b-aaf66f4dcf25"
target=np.array(label)
target.shape
# + colab={"base_uri": "https://localhost:8080/"} id="I-9LaXkK_vkh" outputId="bde34f55-0d1d-4d77-88aa-5b9dc4242d4b"
from tensorflow.keras.utils import to_categorical
target_categorical=to_categorical(target)
target_categorical.shape
# + colab={"base_uri": "https://localhost:8080/"} id="59g6eLxv_vki" outputId="3e5e7624-f972-4eab-d52d-6662f9224952"
target_categorical
# + id="lAs99Fae_vkk"
np.save(open('/content/drive/MyDrive/All fruits/target_new.npy','wb'),target_categorical)
| 5_Evaluation_on_New_Images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # STANDALONE STUDY
# ## **SPATIAL FOURBAR**
# -----------------
# ## Environment Setup
# --------------------
# ### Colab Session
# *Run if in a Colab Session Only!*
# #### Setting-up Colab Virtual Machine
# This will clone the package repository and install the needed modules into the Virtual Machine dedicated for the running notebook session.
# !git clone https://github.com/khaledghobashy/smbd.git
# !cd smbd/smbd/numenv/python/numerics/core/math_funcs/_cython_definitions && python setup.py build_ext -if
# !pip install sympy==1.4
# Adding package path to the python system path and creating a new folder, **smbd_project**, where we can store the model files and results.
# +
import os
import sys
pkg_path = '/content/smbd'
sys.path.append(pkg_path)
os.mkdir('smbd_project')
os.chdir('smbd_project')
# -
# #### Rendering SymPy in Colab
# Rendering SymPy equations in Colab requires the MathJax javascript library to be available within each cell output. The following defines an execution hook that loads it automatically each time you execute a cell.
# +
from IPython.display import Math, HTML
def load_mathjax_in_cell_output():
display(HTML("<script src='https://www.gstatic.com/external_hosted/"
"mathjax/latest/MathJax.js?config=default'></script>"))
get_ipython().events.register('pre_run_cell', load_mathjax_in_cell_output)
# -
# ---------------------------------------------------------------
# ### Local Session
# *Run if on your Local Machine!*</br>
#
# The ```pkg_path``` variable is where you have the repository on your machine and should be changed to where you cloned the repository.</br>
# *Uncomment the code below and change the ```pkg_path``` variable!*
# +
#import sys
#pkg_path = '/path/to/smbd'
#sys.path.append(pkg_path)
# -
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# Creating Project Directory
from smbd.systems import standalone_project
project = standalone_project()
project.create()
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ## **SYSTEM DESCRIPTION**
# -------------------------
# ### **Important Note**
# The tool uses [euler-parameters](https://en.wikibooks.org/wiki/Multibody_Mechanics/Euler_Parameters) -which is a 4D unit quaternion- to represents bodies orientation in space. This makes the generalized coordinates used to fully define a body in space to be **7,** instead of **6**, it also adds an algebraic equation to the constraints that ensures the unity/normalization of the body quaternion. This is an important remark as the calculations of the degrees-of-freedom depends on it.
# ### **Summary**
# A four-bar linkage is the simplest movable closed chain linkage. It consists of four bodies, called bars or links, connected in a loop by four joints that gives the mechanism only 1 degree-of-freedom. More general information can be found on [wikipedia](https://en.wikipedia.org/wiki/Four-bar_linkage).
# ### **Topology Layout**
# The mechanism consists of 3 Bodies + 1 Ground. Therefore, total system coordinates -including the ground- is $$n=n_b\times7 = 4\times7 = 28$$
# where $n_b$ is the total number of bodies. The list of bodies is given below:
# - Crank $l_1$.
# - Connecting-Rod $l_2$.
# - Rocker $l_3$.
#
# The system connectivity is as follows:
# - Crank $l_1$ is connected to the ground by a revolute joint, resulting in constraint equations $n_{c,rev} = 5$
# - Connecting-Rod $l_2$ is connected to the Crank $l_1$ by a spherical joint, resulting in constraint equations $n_{c,sph} = 3$
# - Rocker $l_3$ is connected to Connecting-Rod $l_2$ by a universal joint, resulting in constraint equations $n_{c,uni} = 4$
# - Rocker $l_3$ is connected to the ground by a revolute joint, resulting in constraint equations $n_{c,tri} = 5$
# - Total free DOF is $$n-( n_{c,rev}+n_{c,sph}+n_{c,uni}+n_{c,P}+n_{c,g}) = 28-(10+3+4+(3\times1)+7) = 28-27 = 1$$, where the $n_{c,P}$ and $n_{c,g}$ represents the constraints due to euler-parameters normalization equations and the ground-constraints respectively.
# - The Crank revolute joint is actuated by a rotational actuator, resulting in constraint equations $n_{c,act} = 1$
#
# | Joint Name | Body i | Body j | Joint Type | $n_c$ |
# |:-----------:|:-------------- |:-------------- | ---------- | ----- |
# | a | Ground | Crank | Revolute | 5 |
# | b | Crank | Connecting-Rod | Spherical | 3 |
# | c | Connecting-Rod | Rocker | Universal | 4 |
# | d | Rocker | Ground | Revolute | 5 |
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ## **SYMBOLIC TOPOLOGY**
# ------------------------
# In this section, we create the symbolic topology that captures the topological layout that we just discussed.</br>
# Defining the topology is very simple. We start by importing the ```standalone_topology``` class and create a new instance that represents our symbolic model. Then we start adding the components we discussed earlier, starting by the bodies, then the joints, actuators and forces, and thats it.</br>
# These components will be represented symbolically, and therefore there is no need for any numerical inputs at this step.
#
# The system is stored in a form of a network graph that stores all the data needed for the assemblage of the system equations later. But even before the assemblage process, we can gain helpful insights about our system as well be shown.
#
from smbd.systems import standalone_topology, configuration
project_name = 'spatial_fourbar'
sym_model = standalone_topology(project_name)
# ### Adding Bodies
sym_model.add_body('l1')
sym_model.add_body('l2')
sym_model.add_body('l3')
# ### Adding Joints
sym_model.add_joint.revolute('a','ground','rbs_l1')
sym_model.add_joint.spherical('b','rbs_l1','rbs_l2')
sym_model.add_joint.universal('c','rbs_l2','rbs_l3')
sym_model.add_joint.revolute('d','rbs_l3','ground')
# ### Adding Actuators
sym_model.add_actuator.rotational_actuator('act', 'jcs_a')
# ### Adding Forces
# +
#model.add_force.internal_force('force_name', 'body_1', 'body_2', mirror=False)
# + [markdown] colab_type="text" id="goKtOBuLd1Kh"
# ### **SYMBOLIC CHARACTERISTICS**
# -
# #### Topology Graph
# Visualizing the connectivity of the system as a network graph, where the nodes represent the bodies, and the edges represent the joints, forces and/or actuators between the bodies.
# Checking the system's number of generalized coordinates $n$ and number of constraints $n_c$.
# ### **ASSEMBLING**
# This is the last step of the symbolic building process, where we make the system starts the assemblage process of the governing equations, which will be used then in the code generation for the numerical simulation, and also can be used for further symbolic manipulations.</br>
#
# *Note: The equations' notations will be discussed in the tool documentation files.*
sym_model.assemble()
# #### Checking the System Equations
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ## **SYSTEM CONFIGURATION**
# ---------------------------
# In this step we define a symbolic configuration of our symbolic topology. As you may have noticed in the symbolic topology building step, we only cared about the **_topology_**, thats is the system bodies and their connectivity, and we did not care explicitly with how these components are configured in space.</br>
#
# In order to create a valid numerical simulation session, we have to provide the system with its numerical configuration needed, for example, the joints' locations and orientations. The symbolic topology in its raw form will require you to manually enter all these numerical arguments, which can be cumbersome even for smaller systems. This can be checked by checking the configuration inputs of the symbolic configuration as ```sym_config.config.input_nodes```
#
# Here we start by stating the symbolic inputs we wish to use instead of the default inputs set, and then we define the relation between these newly defined arguments and the original ones.
#
# **_The details of this process will be provided in the documentation._**
# +
# Symbolic configuration name.
config_name = '%s_cfg'%project_name
# Symbolic configuration instance.
sym_config = configuration(config_name, sym_model)
# -
# ### CONFIGURATION INPUTS
# +
# Adding the desired set of UserInputs
# ====================================
sym_config.add_point.UserInput('a')
sym_config.add_point.UserInput('b')
sym_config.add_point.UserInput('c')
sym_config.add_point.UserInput('d')
sym_config.add_vector.UserInput('x')
sym_config.add_vector.UserInput('y')
sym_config.add_vector.UserInput('z')
# -
# ### CONFIGURATION RELATIONS
# +
# Defining Relations between original topology inputs
# and our desired UserInputs.
# ===================================================
# Revolute Joint (a) location and orientation
sym_config.add_relation.Equal_to('pt1_jcs_a', ('hps_a',))
sym_config.add_relation.Equal_to('ax1_jcs_a', ('vcs_x',))
# Spherical Joint (b) location and orientation
sym_config.add_relation.Equal_to('pt1_jcs_b', ('hps_b',))
sym_config.add_relation.Equal_to('ax1_jcs_b', ('vcs_z',))
# Universal Joint (c) location and orientation
sym_config.add_relation.Equal_to('pt1_jcs_c', ('hps_c',))
sym_config.add_relation.Oriented('ax1_jcs_c', ('hps_b', 'hps_c'))
sym_config.add_relation.Oriented('ax2_jcs_c', ('hps_c', 'hps_b'))
# Revolute Joint (d) location and orientation
sym_config.add_relation.Equal_to('pt1_jcs_d', ('hps_d',))
sym_config.add_relation.Equal_to('ax1_jcs_d', ('vcs_y',))
# -
# ### GEOMETRIES
# Here we start defining basic geometric shapes that can represents the shapes of the bodies in our system. This serves two points:
# - Visualization and Animation in blender.
# - Evaluating the bodies inertia properties from these basic geometries instead of explicit definition.
# +
# links radius
sym_config.add_scalar.UserInput('links_ro')
# Link 1 geometry
sym_config.add_geometry.Cylinder_Geometry('l1', ('hps_a','hps_b','s_links_ro'))
sym_config.assign_geometry_to_body('rbs_l1', 'gms_l1')
# Link 2 geometry
sym_config.add_geometry.Cylinder_Geometry('l2', ('hps_b','hps_c','s_links_ro'))
sym_config.assign_geometry_to_body('rbs_l2', 'gms_l2')
# Link 3 geometry
sym_config.add_geometry.Cylinder_Geometry('l3', ('hps_c','hps_d','s_links_ro'))
sym_config.assign_geometry_to_body('rbs_l3', 'gms_l3')
# -
# Now lets check the current ```input_nodes```.
# ### ASSEMBLING
sym_config.export_JSON_file()
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ## **CODE GENERATION**
# ----------------------
# This step aims to create a valid code that can be used for numerical simulation. We will use both the ***python code generator*** and the **C++ code generator** to create a valid numerical simulation environment in python and C++.</br>
#
# Theoretically, the symbolic environment is uncoupled from the simulation environment, which opens the door to create various simulation environments that can be in any language.
# + [markdown] colab_type="text" id="udnBb6lvi9fo"
# ### Generating Python Numerical Environment
# -
from smbd.numenv.python.codegen import projects as py_numenv
# Creating the needed directories structure for the source code.
py_project = py_numenv.standalone_project(parent_dir='')
py_project.create_dirs()
# Generating the source code.
py_project.write_topology_code(sym_model.topology)
py_project.write_configuration_code(sym_config.config)
# + [markdown] colab_type="text" id="udnBb6lvi9fo"
# ### Generating C++ Numerical Environment
# -
from smbd.numenv.cpp_eigen.codegen import projects as cpp_numenv
cpp_project = cpp_numenv.standalone_project(parent_dir='')
cpp_project.create_dirs()
# Generating the source code.
cpp_project.write_topology_code(sym_model.topology)
cpp_project.write_configuration_code(sym_config.config)
# Generating the makefile and main.cpp files.
cpp_project.write_mainfile()
cpp_project.write_makefile()
# ### Generating Blender Script
from smbd.utilities.blender import codegen
blender_code = codegen.script_generator(sym_config.config)
blender_code.write_code_file('numenv/')
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ## **PYTHON NUMERICAL SIMULATION**
# ---------------------------
# The **numerical simulation** step is also a straight forward one. We start by importing helper libraries like **numpy** and **pandas**, then we import the ```multibody_system``` class and the ```simulation``` class that will be used to create a numerical system and a simulation instance respectively. And finally we import the generated code files from the previous step.
#
# The steps then can be summarized as follows:
# 1. Create a numerical model instance by passing the generated topology code to the ```multibody_system``` class.
# 2. Make the configuration of that instance refers to the generated configuration code.
# 3. Setting the numerical data of the configuration.
# 4. Setting the **user-defined functions** that will be used for the actuators/forces.
# 5. Creating a **simulation instance**, set the desired time-sampling and run the simulation.
# 6. Plotting the results.
# + [markdown] colab_type="text" id="w-7fcrPBjUie"
# ### Imports
# -
import numpy as np
from smbd.numenv.python.numerics.core.systems import multibody_system, simulation, configuration
# Importing the generated modules/souce-code of the model.
from numenv.python.src import spatial_fourbar
# + [markdown] colab_type="text" id="vkHoR9CCjb6u"
# ### Creating Numerical Model
# +
num_model = multibody_system(spatial_fourbar)
num_config = configuration('base')
num_config.construct_from_json('spatial_fourbar_cfg.json')
num_model.topology.config = num_config
# + [markdown] colab_type="text" id="85ZRAWmRjitu"
# ### Setting Configuration Numerical Data
# -
# +
num_config.hps_a.flat[:] = 0, 0, 0
num_config.hps_b.flat[:] = 0, 0, 200
num_config.hps_c.flat[:] = -750, -850, 650
num_config.hps_d.flat[:] = -400, -850, 0
num_config.vcs_x.flat[:] = 1, 0, 0
num_config.vcs_y.flat[:] = 0, 1, 0
num_config.vcs_z.flat[:] = 0, 0, 1
num_config.s_links_ro = 20
# -
num_config.assemble()
# ### Setting User Functions.
# #### Actuators
# Here we set the actuation functions of the system actuators.
num_config.UF_mcs_act = lambda t : -np.deg2rad(360)*t
# ### Assembling the numerical configuration.
# Here, we pass the numerical inputs instance ```num_inputs``` to the ```assemble()``` method of the ```num_config``` instance of our model.
# + [markdown] colab_type="text" id="APxcuvdMjq4-"
# ### Creating Simulation Instance
# -
# A simulation instance takes in three arguments, the simulation ```name```, the numerical ```model``` and the simulation ```typ```. The simulation ```typ``` argument should be either *kds* for kinematically driven systems or *dds* for dynamically driven systems. For our case, the system is fully defined kinematically, and therefore we will set the ```typ``` as *kds*.
sim = simulation('sim', num_model, 'kds')
sim.set_time_array(5, 2e-2)
sim.solve()
sim.soln.pos_dataframe.plot(x='time', y='rbs_l3.x', grid=True, figsize=(10,4))
sim.soln.vel_dataframe.plot(x='time', y='rbs_l3.x', grid=True, figsize=(10,4))
sim.soln.acc_dataframe.plot(x='time', y='rbs_l3.x', grid=True, figsize=(10,4))
# ### Evaluating System Reactions
# Here we take the simulation a step further by making it an inverse dynamics problem and evaluating the reactions resulted due the imposed constraints, either by joints or actuators. This step is only valid if you have defined the numerical values of the inertia properties of the system.
sim.eval_reactions()
sim.soln.reactions_dataframe.plot(x='time', y=['T_ground_mcs_act.x', 'T_ground_jcs_a.x'],
grid=True, figsize=(10,4))
# ---------------------------------------------------------------
# ---------------------------------------------------------------
# ## **C++ NUMERICAL SIMULATION**
#
# _Only for Linux Machines_
#
#
# -------------------------------
#
# ***TO DO BRIEF***
# The C++ numerical environment provides samples of **Makefile** and **main.cpp** files that can be used to create, compile and run an executable of the created model.
#
# In order to create a valid environment, we have to edit the **Makefile** and the **main.cpp** files to match the model variables as shown below. These edits can be carried out on the text files using any text editor normally, but for completeness and compatibility with the **Jupyter Notebook** environment the files can be edited here locally using the *Jupyter magic functions*.
#
# First, we check the current working directory, it should be the parent directory that contains [numenv, config_inputs, results, ..], then we use the command```%cd numenv/cpp_eigen/``` to change the working directory.
# Changing the current working directory to 'numenv/cpp_eigen/'
# %cd numenv/cpp_eigen/
# ---------------------------------------------------
# ### **Editing the main.cpp file**
# #### Using the ```%load ``` magic function.
# -------------------------------------------
# Then we use the command ```%load main.cpp```, this will load the generated **main.cpp** content in the current cell where we can edit the content. The edits are as follows:
# - Remove the first line ```# %load main.cpp``` and insert the line ```%%writefile new_main.cpp```.
# - Change the variables topology and configuration in the include directive to match the names of the model generated source files.
# - Start specifying the configuration numerical data via the ```Config``` class.
#
# After these edits, just run the cell like any normal cell via Shift+Enter.
# #### Using the ```%cat ``` magic function.
# -------------------------------------------
# Then we use the command ```%cat main.cpp```, this will load the generated **main.cpp** content as the output of the current cell, we can then copy the content and past and edit it in another cell. The edits are as follows:
# - Insert the line ```%%writefile new_main.cpp``` at the top of the cell.
# - Change the variables topology and configuration in the include directive to match the names of the model generated source files.
# - Start specifying the configuration numerical data via the ```Config``` class.
#
# After these edits, just run the cell like any normal cell via Shift+Enter.
# %cat main.cpp
# +
# %%writefile new_main.cpp
#include <iostream>
#include "smbd/solvers.hpp"
#include "src/spatial_fourbar.hpp"
#include "src/spatial_fourbar_cfg.hpp"
int main()
{
Topology model("");
auto Config = ConfigurationInputs<Configuration>(model.config);
// assign the configuration inputs needed ...
Config.R_ground << 0, 0, 0 ;
Config.P_ground << 1, 0, 0, 0 ;
Config.hps_a << 0, 0, 0 ;
Config.hps_b << 0, 0, 200 ;
Config.hps_c << -750, -850, 650 ;
Config.hps_d << -400, -850, 0 ;
Config.vcs_x << 1, 0, 0 ;
Config.vcs_y << 0, 1, 0 ;
Config.vcs_z << 0, 0, 1 ;
Config.s_links_ro = 20 ;
Config.UF_mcs_act = [](double t)->double{return 2*(22/7)*t;};
Config.assemble();
Solver<Topology> Soln(model);
Soln.set_time_array(5, 250);
Soln.Solve();
Soln.ExportResultsCSV("../../results/", 0);
};
# -
# ---------------------------------------------------
# ### **Editing the Makefile file**
# Following the same steps done for the main.cpp file, then apply the following edits:
# - Change the variables MODEL, CONFIG and MAIN to match the generated source files of the topology, configuration and the edited new_main.cpp file.
#
# After these edits, just run the cell like any normal cell via Shift+Enter.
# %cat Makefile
# +
# %%writefile Makefile
# Change MODEL, CONFG and MAIN to match the source files you want to build
# ========================================================================
MODEL := spatial_fourbar
CONFG := spatial_fourbar_cfg
MAIN := new_main.cpp
# ========================================================================
M_BUILD := build/
M_SRC := src/
M_BIN := bin/
NUM_DIR := /home/khaledghobashy/Documents/smbd/smbd/numenv/cpp_eigen/numerics
SMBD_SRC := $(NUM_DIR)/src
SMBD_BUILD := $(NUM_DIR)/build
SMBD_OBJS = $(SMBD_BUILD)/*.o
DEPS := $(M_BUILD)$(MODEL).o $(MAIN) $(M_SRC)$(CONFG).hpp $(SMBD_SRC)/smbd/solvers.hpp
INC := -I $(SMBD_SRC)
CC := g++
$(M_BIN)$(MODEL): $(DEPS) $(SMBD_OBJS)
$(CC) $(INC) $(M_BUILD)$(MODEL).o $(MAIN) $(SMBD_OBJS) -o $@
$(M_BUILD)$(MODEL).o: $(M_SRC)$(MODEL).cpp $(M_SRC)$(MODEL).hpp
$(CC) $(INC) -c -o $@ $<
$(SMBD_BUILD)/%.o: $(SMBD_SRC)/smbd/%.cpp $(SMBD_SRC)/smbd/%.hpp
cd $(SMBD_SRC)/../ && make
clear:
rm $(M_BUILD)*.o $(M_BIN)$(MODEL)
# -
# ### Running ```make``` command
# !make
# ### Running the executable.
# !bin/spatial_fourbar
# ### Loading and Plotting Results.
import pandas as pd
import matplotlib.pyplot as plt
data = pd.read_csv('../../results/Positions.csv')
data.plot(x='time', y='rbs_l3.x', grid=True, figsize=(10,4))
data.plot(x='time', y='rbs_l1.y', grid=True, figsize=(10,4))
| uraeus/smbd/utilities/serialization/structural/test/spatial_fourbar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pyspark
from pyspark.sql import *
from pyspark.sql.types import *
from pyspark.sql.functions import *
from pyspark import SparkContext, SparkConf
from pyspark.sql import SQLContext # https://spark.apache.org/docs/1.6.1/sql-programming-guide.html
from os.path import join, abspath
import psycopg2
import os
# pip install psycopg2-binary
# # now create connection
connection = psycopg2.connect(
host = 'c451_db_1',
database = 'irs990',
user = 'postgres',
password = '<PASSWORD>',
port='5432'
)
cur = connection.cursor()
cur.execute("""SELECT datname from pg_database""")
rows = cur.fetchall()
print('\nShow me the databases:\n')
for row in rows:
print(' ', row[0])
pd.read_sql_query("SELECT * FROM address_table LIMIT 3", connection)
# +
# Create the session
conf = (SparkConf()
.set("spark.ui.port", "4041")
.set('spark.executor.memory', '4G')
.set('spark.driver.memory', '45G')
.set('spark.driver.maxResultSize', '10G')
.set('spark.jars', '/home/jovyan/scratch/postgresql-42.2.18.jar'))
# Create the context
# sc = pyspark.SparkContext(conf=conf)
# sqlContext = SQLContext(sc)
spark = SparkSession.builder \
.appName('test') \
.config(conf=conf) \
.getOrCreate()
# -
# spark.stop()
properties = {
'driver': 'org.postgresql.Driver',
'url': 'jdbc:postgresql://c451_db_1:5432/irs990',
'user': 'postgres',
'password': '<PASSWORD>',
'dbtable': 'address_table',
}
spark
os.listdir()
os.getcwd()
jdbcDF = spark.read \
.format('jdbc') \
.option('driver', properties['driver']) \
.option('url', properties['url']) \
.option('dbtable', properties['dbtable']) \
.option('user', properties['user']) \
.load()
jdbcDF.show(10)
| scripts/postgres_docker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 数学函数、字符串和对象
# ## 本章介绍Python函数来执行常见的数学运算
# - 函数是完成一个特殊任务的一组语句,可以理解为一个函数相当于一个小功能,但是在开发中,需要注意一个函数的长度最好不要超过一屏
# - Python中的内置函数是不需要Import导入的
# <img src="../Photo/15.png"></img>
min(-1,0,1)
pow(2,3)
abs(-10)
max('abc')
max(1,2,3)
import os
import random
a=eval(input('please input Number '))
b=random.randint(0,2)
pritn(b)
if max(a,b)==a
os.system('calc')
else:
os.system('calc')
# +
a,b,c=eval(input('输入三个数,a,b,c'))
# -
#
a = -10
print(abs(a))
b = -10.1
print(abs(b))
c = 0
print(abs(c))
max(1, 2, 3, 4, 5)
min(1, 2, 3, 4, 5)
min(1, 2, 3, -4, 5)
for i in range(10):
print(i)
pow(2, 4, 2) # 幂指数运算,第三个参数是取模运算
round(10.67, 1) # 一个参数就是四舍五入,保留小数位数
# ## 尝试练习Python内置函数
# ## Python中的math模块提供了许多数学函数
# <img src="../Photo/16.png"></img>
# <img src="../Photo/17.png"></img>
#
x,y=eval(input(x,y))
a=eval(input(a))
y=eval(input(y))
L=-[yloga+(1-y)log(1-a)]
print(L)
import math
math.fabs(-4.0)
import math
math.ceil(-3.9)
import time
start=time.time()
num=0
for i in range(1000000):
num +=i
end=time.time()
print(end - start)
# +
import math # 导入数学包
a1 = math.fabs(-2)
print(a1)
print(math.log(2.71828))
print(math.asin(1.0))
# -
b1 = math.cos(math.radians(90)) # cos代入的是弧度值,very important!
print(b1)
c1 = 3.1415926
print(math.degrees(c1))
math.sqrt(9)
math.sin(2 * math.pi)
math.cos(2 * math.pi)
min(2, 2, 1)
math.log(math.e ** 2)
math.exp(1)
max(2, 3, 4)
math.ceil(-2.5)
# +
# 验证码系统
first_num, second_num = 3, 4
print('验证码', first_num ,'+', second_num, '= ?')
answer = eval(input('写出结果: '))
if answer == first_num + second_num:
print('验证码正确')
else:
print('验证码错误')
# +
import random
import math
first_num, second_num = 3, 4
list = ['+', '-', '*', '/']
randl = random.randint(0, 3)
if list[randl]=='+':
print('验证码', first_num ,'+', second_num, '= ?')
right_answer = first_num + second_num
elif list[randl]=='-':
print('验证码', first_num ,'-', second_num, '= ?')
right_answer = first_num - second_num
elif list[randl]=='-':
print('验证码', first_num ,'*', second_num, '= ?')
right_answer = first_num * second_num
else:
print('验证码', first_num ,'/', second_num, '= ?')
right_answer = first_num / second_num
answer = eval(input('写出结果: '))
if answer == right_answer:
print('验证码正确')
else:
print('验证码错误')
# +
# 验证码系统
import random
first_num = random.randint(0, 9)
second_num = random.randint(0, 9)
fuhao = random.randint(0, 3)
if fuhao==0:
print('验证码', first_num ,'+', second_num, '= ?')
right_answer = first_num + second_num
elif fuhao==1:
print('验证码', first_num ,'-', second_num, '= ?')
right_answer = first_num - second_num
elif fuhao==2:
print('验证码', first_num ,'*', second_num, '= ?')
right_answer = first_num * second_num
else:
print('验证码', first_num ,'/', second_num, '= ?')
right_answer = first_num / second_num
answer = eval(input('写出结果: '))
if answer == right_answer:
print('验证码正确')
else:
print('验证码错误')
# -
import random
list = ['+', '-', '*', '/']
c = random.sample(list, 1)
print(c)
# +
import random
import math
first_num = random.randint(0, 9)
second_num = random.randint(0, 9)
list = ['+', '-', '*', '/']
fuhao = random.sample(list, 1)
if fuhao=='+':
print('验证码', first_num ,'+', second_num, '= ?')
right_answer = first_num + second_num
elif fuhao=='-':
print('验证码', first_num ,'-', second_num, '= ?')
right_answer = first_num - second_num
elif fuhao=='-':
print('验证码', first_num ,'*', second_num, '= ?')
right_answer = first_num * second_num
else:
print('验证码', first_num ,'/', second_num, '= ?')
right_answer = first_num / second_num
answer = eval(input('写出结果: '))
if answer == right_answer:
print('验证码正确')
else:
print('验证码错误')
# -
import PIL
# ## 两个数学常量PI和e,可以通过使用math.pi 和math.e调用
import math
print(math.pi)
print(math.e)
# ## EP:
# - 通过math库,写一个程序,使得用户输入三个顶点(x,y)返回三个角度
# - 注意:Python计算角度为弧度制,需要将其转换为角度
# <img src="../Photo/18.png">
import math
x1,x2=eval(input('x1,x2'))
y1,y2=eval(input('y1,y2'))
x3,y3=eval(input('x3,y3'))
c=math.sqrt((x1-x2)**2+(y1-y2)**2)
b=math.sqrt((x1-x3)**2+(y1-y3)**2)
a=math.sqrt((x2-x3)**2+(y2-y3)**2)
A=math.degrees(math.acos((a*a-b*b-c*c)/(-2*b*c)))
B=math.degrees(math.acos((b*b-a*a-c*c)/(-2*a*c)))
C=math.degrees(math.acos((c*c-b*b-a*a)/(-2*a*b)))
print(A,B,C)
# +
import math
x1, y1 = eval(input('输入A点坐标:'))
x2, y2 = eval(input('输入B点坐标:'))
x3, y3 = eval(input('输入C点坐标:'))
a = math.sqrt((x1 - x2) ** 2 + (y1 - y2) ** 2)
b = math.sqrt((x1 - x3) ** 2 + (y1 - y3) ** 2)
c = math.sqrt((x2 - x3) ** 2 + (y2 - y3) ** 2)
A = math.degrees(math.acos((a * a - b * b - c * c) / (-2 * b * c)))
B = math.degrees(math.acos((b * b - a * a - c * c) / (-2 * a * c)))
C = math.degrees(math.acos((c * c - b * b - a * a) / (-2 * a * b)))
print('三角形的三个角分别为', A, B, C)
# -
# ## 字符串和字符
# - 在Python中,字符串必须是在单引号或者双引号内,在多段换行的字符串中可以使用“”“
# - 在使用”“”时,给予其变量则变为字符串,否则当多行注释使用
# +
a = 'joker'
b = "Kate"
c = """在Python中,字符串必须是在单引号或者双引号内,在多段换行的字符串中可以使用“”“
在使用”“”时,给予其变量则变为字符串,否则当多行注释使用""" #字符串有多行时,添加三个单引号或者三个双引号
"""在Python中,字符串必须是在单引号或者双引号内,在多段换行的字符串中可以使用“”“
在使用”“”时,给予其变量则变为字符串,否则当多行注释使用""" #三引号可以表示多行注释
# 当6个引号没有赋值时,那么它是注释的作用
# 6个引号的作用,多行文本
print(type(a), type(b), type(c))
# -
# ## ASCII码与Unicode码
# - <img src="../Photo/19.png"></img>
# - <img src="../Photo/20.png"></img>
# - <img src="../Photo/21.png"></img>
# ## 函数ord、chr
# - ord 返回ASCII码值
# - chr 返回字符
ord('a')
joker = 'A'
ord(joker)
print(ord('q'), ord('Z'))
print(chr(65))
print(chr(90))
import numpy as np
np.nonzero(1)
# ## EP:
# - 利用ord与chr进行简单邮箱加密
email='759290425@<EMAIL>'
i=0
for i in email:
mail=ord(i)+1
mail1=chr(mail)
print(mail1,end=(''))
email = '<EMAIL>' # 邮箱加密过程
j = 0
for i in email:
text = ord(i) + 1
re_text = chr(text)
print(re_text,end=(''))
import hashlib
str1 = 'this is a test.'
h1 = hashlib.md5()
h1.update(str1.encode(encoding = 'utf-8'))
print('MD5加密之后为:', h1.hexdigest())
# ## 转义序列 \
# - a = "He said,"Johon's program is easy to read""
# - 转掉原来的意思
# - 一般情况下,只有当语句与默认语句相撞的时候,就需要转义
a = "He said,\"Johon's program is easy to read\"" #z正则表达式中常用转义字符\
print(a)
# ## 高级print
# - 参数 end: 以什么方式结束打印
# - 默认换行打印
email = '<EMAIL>' # 邮箱加密过程
j = 0
for i in email:
text = ord(i) + 1
re_text = chr(text)
print(re_text, end = '')
# ## 函数str
# - 将类型强制转换成字符串类型
# - 其他一些以后会学到(list,set,tuple...)
a = 100.12
type(str(a))
# ## 字符串连接操作
# - 直接使用 “+”
# - join() 函数
# %time ' '.join(('a','b'))
''.join(('a','b'))
for i in range(10):
url='https://www.so.com/s?q=%E5%9B%BE%E7%89%87&ie=utf-8&src=se7_newtab_new'
url2=''.join(('url','i'))
print(url2)
a1 = 'www.baidu.com/image.page='
a2 = '1'
for i in range(0, 10):
a2 = a1 + str(i)
print(a2)
joint = '^'
# %time joint.join(('a', 'b', 'c', 'd')) # join的参数需要在一个元组之中
# %time '*'.join(('a', 'b', 'c', 'd')) # join的参数需要在一个元组之中
# %time 'A' + 'B' + 'C'
# ## EP:
# - 将 “Welcome” “to” "Python" 拼接
# - 将int型 100 与 “joker is a bad man” 拼接
# - 从控制台读取字符串
# > 输入一个名字返回夸奖此人是一个帅哥
test=' '.join(('Welcome','to','Python'))
a=100
test2=str(a)
test3=''.join((test2,'Joker is a bad man'))
print(test,'\n','test2','\n',test3)
a=input('输入一个名字 ')
b=' '.join((a ,'是一个帅哥'))
print(b)
text1 = ' '.join(('Welcome', 'to', 'Python'))
i = 100
text2 = str(i)
text3 = ' '.join((text2, 'Joker is a bad man'))
print(text1, '\n', text2 ,'\n', text3)
name = input('输入名字:')
text = ' '.join((name, 'is a good boy.'))
print(text)
# ## 实例研究:最小数量硬币
# - 开发一个程序,让用户输入总金额,这是一个用美元和美分表示的浮点值,返回一个由美元、两角五分的硬币、一角的硬币、五分硬币、以及美分个数
# <img src="../Photo/22.png"></img>
amount = eval(input('Enter an amount, for example 11.56: '))
fenshuAmount = int(amount * 100)
dollorAmount = fenshuAmount // 100
remainDollorAmount = fenshuAmount % 100
jiaoAmount = remainDollorAmount // 25
remainJiaoAmount = remainDollorAmount % 25
fenAmount = remainJiaoAmount // 10
remainFenAmount = remainJiaoAmount % 10
fenAmount2 = remainFenAmount // 5
remainFenAmount2 = remainFenAmount % 5
fenFinalAmount = remainFenAmount2
print('美元个数为',dollorAmount,'\n', '两角五分硬币个数为',
jiaoAmount, '\n','一角个数为', fenAmount, '\n','五美分个数为', fenAmount2,'\n', '一美分个数为',fenFinalAmount)
amount = eval(input('Ennter an amount,for example 11.56:'))
remainingAmount = int(amount * 100)
print(remainingAmount)
numberOfOneDollars = remainingAmount //100
remainingAmount = remainingAmount % 100
numberOfQuarters = remainingAmount // 25
remainingAmount = remainingAmount % 25
numberOfDimes = remainingAmount // 10
remainingAmount = remainingAmount % 10
numberOfNickls = remainingAmount // 5
remainingAmount = remainingAmount % 5
numberOfPenies = remainingAmount
print(numberOfOneDollars,numberOfQuarters,numberOfDimes,numberOfNickls,numberOfPenies)
# - Python弱项,对于浮点型的处理并不是很好,但是处理数据的时候使用的是Numpy类型
# <img src="../Photo/23.png"></img>
remainingAmount = eval(input('Ennter an amount,for example 11.56:'))
print(remainingAmount)
numberOfOneDollars = remainingAmount //100
remainingAmount = remainingAmount % 100
numberOfQuarters = remainingAmount // 25
remainingAmount = remainingAmount % 25
numberOfDimes = remainingAmount // 10
remainingAmount = remainingAmount % 10
numberOfNickls = remainingAmount // 5
remainingAmount = remainingAmount % 5
numberOfPenies = remainingAmount
print(numberOfOneDollars,numberOfQuarters,numberOfDimes,numberOfNickls,numberOfPenies)
# ## id与type
# - id 查看内存地址,在判断语句中将会使用
# - type 查看元素类型
a = 100
id(a)
id(True)
100 == 100
112345678800000000 is '112345678800000000'
112345678800000000 is 112345678800000000
a = True
b = False
print(id(a), id(b))
a is b
# ## 其他格式化语句见书
# # Homework
# - 1
# <img src="../Photo/24.png"><img>
# <img src="../Photo/25.png"><img>
import math
r=eval(input('输入顶点到中心距离r '))
s=2*r*math.sin(math.pi/5)
A=(5*s*s)/(4*math.tan(math.pi/5))
print('边长为 %.4f 的五边形面积为 %.4f' %(s,A))
import math
radius = eval(input('输入五边形顶点到中心的距离:'))
s = 2 * radius * math.sin(math.pi / 5)
area = 5 * s * s / (4 * math.tan(math.pi / 5))
print('计算结果:边长为 %.4f 的五边形的面积为 %.4f' %(s, area))
# - 2
# <img src="../Photo/26.png"><img>
import math
x1,y1=eval(input('输入第一个点的经度和纬度'))
x2,y2=eval(input('输入第二个点的经度和纬度'))
r=6371.01
x1=math.radians(x1)
x2=math.radians(x2)
y1=math.radians(y1)
y2=math.radians(y2)
d=r*math.acos((math.sin(x1)*math.sin(x2))+(math.cos(x1)*math.cos(x2))*(math.cos(y1-y2)))
print('地球两点之间距离为:',d, 'km')
import math
x1, y1 = eval(input('输入第一个点的经度和纬度:'))
x2, y2 = eval(input('输入第二个点的经度和纬度:'))
x1 = math.radians(x1)
x2 = math.radians(x2)
y1 = math.radians(y1)
y2 = math.radians(y2)
radius = 6371.01
#print(x1, y1, x2, y2)
d = radius * math.acos(math.sin(x1) * math.sin(x2) + math.cos(x1) * math.cos(x2) * math.cos(y1 - y2))
print('地球上两个点之间的距离为:', d, 'km')
# - 3
# <img src="../Photo/27.png"><img>
import math
s=eval(input('输入五角形边长 '))
A=5*s**2/(4*math.tan(math.pi/5))
print('输出面积', A)
import math
s = eval(input('输入五边形的边长:'))
area = 5 * s * s / (4 * math.tan(math.pi / 5))
print('计算结果:边长为%.4f的五边形的面积为%.4f' %(s, area))
# - 4
# <img src="../Photo/28.png"><img>
import math
s=eval(input('输入正多边形边长 '))
n=eval(input('输入正多边形边数 '))
A=n*s**2/(4*math.tan(math.pi/n))
print('输出面积', A)
import math
num = eval(input('输入正多边形的边数:'))
s = eval(input('输入正多边形的边长:'))
area = num * s * s / (4 * math.tan(math.pi / num))
print('计算结果:边长为%.4f的%d边形的面积为%.4f' %(s, num,area))
# - 5
# <img src="../Photo/29.png"><img>
# <img src="../Photo/30.png"><img>
n=eval(input('输入一个ASCII值 '))
print('返回值为 ',chr(n))
num = eval(input('输入一个介于0到127之间的整数ASCII码值:'))
alp = chr(num)
print('ASCII码值为', num, '的字母为',alp)
# - 6
# <img src="../Photo/31.png"><img>
n=input('输入名字 ')
t=eval(input('输入工作时间 '))
h=eval(input('输入每小时报酬 '))
r=eval(input('输入联扣税率 '))
r1=eval(input('输入州扣税率 '))
g=t*h
#print('工作总时间所获得的报酬 ',G)
f=g*r
#print('所获得的报酬扣除联邦税',F)
t1=g*r1
#print('所获得的报酬扣除州税',T)
t2=f+t1
#print('总结算',T1)
print('姓名 ',n,'\n','工作时间',t,'\n','每小时报酬',h,'\n','联扣税率$',r,'\n','州扣税率$',r1,'\n',
'工作总时间所获得的报酬 ',g,'\n','所获得的报酬扣除联邦税$',f,'\n','所获得的报酬扣除州税$',t1,'\n','总结算',t2)
# +
name = input('输入雇员的姓名:')
hourTime = eval(input('输入雇员一周工作的时间:'))
payRate = eval(input('输入雇员每小时报酬:'))
federalTax = eval(input('输入联邦扣税率:'))
stateTax= eval(input('输入州扣税率:'))
grossPay = hourTime * payRate
federalWithholding = grossPay * federalTax
stateWithholding = grossPay * stateTax
totalDeduction = federalWithholding + stateWithholding
netPay = grossPay - totalDeduction
print('雇员姓名:',name, '\n','一周工作时间:', hourTime, '\n', '每小时报酬:$', payRate,'\n',
'总收入:$', grossPay,'\n', '扣税:','\n', '联邦扣税额:$',federalWithholding,'\n',
'州扣税额:$',stateWithholding,'\n', '总扣税额:$',totalDeduction ,'\n',
'剩余工资:$', netPay)
# -
# - 7
# <img src="../Photo/32.png"><img>
n=input('输入四位数')
r=''
for i in n:
r=i+r
print('反向数字为',r)
number = input('输入一个四位整数:')
reverNumber = ''
for i in number:
reverNumber = i + reverNumber
print('四位整数',number, '的反向数字为',reverNumber)
# - 8 进阶:
# > 加密一串文本,并将解密后的文件写入本地保存
# +
email = '<EMAIL>' # 邮箱加密过程
re_text = ''
for i in email:
text = ord(i) + 1
re_text = ''.join((re_text, chr(text)))
print('加密之后的信息为:', re_text)
reves_text = ''
for i in re_text:
text = ord(i) - 1
reves_text = ''.join((reves_text, chr(text)))
print('解密之后的信息为:', reves_text)
outputfile = open("day02.txt",'w')
outputfile.write(reves_text)
outputfile.close()
| 7.17.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Wikidata.org
#
# Wikidata.org allows you to get information from Wiki sites through an API.
#
# Using the API requires you to write the query in [SPARQL](https://en.wikipedia.org/wiki/SPARQL), but there is plenty of example queries to work from at https://query.wikidata.org allowing you to get started without previous knowledge about this SPARQL.
#
# ### Wikidata and Machinelearning
# As Wiki is an internatinal project in +300 languages and has a lot of articles, Wikidata is a good start for harvesting data to use in machinelearning
# * Example SPARQL queries
# * All [Swedish politicians in the Swedish Goverment](http://tinyurl.com/y4jo4lva)
# * just those with a [twitter account](http://tinyurl.com/y4qb32q9)
# * In may started also the [Lexicographical data project](https://www.wikidata.org/wiki/Wikidata:Lexicographical_data/en) with the goal to "describe precisely all words in all languages" in a machinereadable form
#
# More information:
# * [Wikidata](https://www.wikidata.org/wiki/Wikidata:Main_Page)
# * [The query editor](https://query.wikidata.org/)
from IPython.display import YouTubeVideo
YouTubeVideo("AR66WVBViBQ")
# ## Example: Swedish Prime ministers
# +
# Install a dependency using a terminal command
# !pip install --quiet sparqlwrapper
# Allows us to use IPython.display.JSON
import IPython.display
# +
from SPARQLWrapper import SPARQLWrapper, JSON
endpoint_url = "https://query.wikidata.org/sparql"
query = """# Query Find in WikiData people with position held P39 Swedish Prime minister Q687075
#
select ?person ?personLabel ?personDescription ?replacedbyLabel ?start ?pic ?end {
{
?person wdt:P39 wd:Q687075;
p:P39 [
ps:P39 wd:Q687075;
pq:P580 ?start;
pq:P582 ?end;
pq:P1365 ?replace; #Replace
pq:P1366 ?replacedby #Replaced by
].
OPTIONAL{ ?person wdt:P18 ?pic .} # If we have an illustration
}
SERVICE wikibase:label { bd:serviceParam wikibase:language "en"}
}
Order by ?start"""
def get_results(endpoint_url, query):
sparql = SPARQLWrapper(endpoint_url)
sparql.setQuery(query)
sparql.setReturnFormat(JSON)
return sparql.query().convert()
results = get_results(endpoint_url, query)
IPython.display.JSON(results)
# -
for result in results["results"]["bindings"]:
print(result["personLabel"]["value"], result["start"]["value"], " - ", result["end"]["value"])
# ## Example: Airports near Berlin
# +
from SPARQLWrapper import SPARQLWrapper, JSON
endpoint_url = "https://query.wikidata.org/sparql"
query = """SELECT ?place ?placeLabel ?location
WHERE
{
# Berlin coordinates
wd:Q64 wdt:P625 ?berlinLoc .
SERVICE wikibase:around {
?place wdt:P625 ?location .
bd:serviceParam wikibase:center ?berlinLoc .
bd:serviceParam wikibase:radius "100" .
} .
# Is an airport
FILTER EXISTS { ?place wdt:P31/wdt:P279* wd:Q1248784 } .
SERVICE wikibase:label {
bd:serviceParam wikibase:language "[AUTO_LANGUAGE],en" .
}
}"""
def get_results(endpoint_url, query):
sparql = SPARQLWrapper(endpoint_url)
sparql.setQuery(query)
sparql.setReturnFormat(JSON)
return sparql.query().convert()
results = get_results(endpoint_url, query)
IPython.display.JSON(results)
# -
for result in results["results"]["bindings"]:
print(result["placeLabel"]["value"])
| wikidata-org.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Now it's time to test new tree.
# There must be already some scripts for the testing.
# +
from tree import treemodule
from tree import treeutils
import numpy as np
alltrees = treemodule.CTree()
wdir = '/home/hoseung/Work/data/05427/'
alltrees.load(filename= wdir + 'rhalo/Trees/tree_0_0_0.dat')
# + active=""
# How to extract a full tree and plot it?
#
# Unlike TMtree, A full tree is given in continuous lines.
#
# 1) extract one tree
# 2) from the last snapshots, find progenitors
# 3) link progenitors with the descendant.
# +
def extract_a_tree(alltrees, idx_last):
i_last = np.where(alltrees['id'] == idx_last)
return alltrees[np.where(alltrees['tree_root_id'] == idx_last)]
def get_progenitors(treedata, idx):
iprgs = np.where(treedata['desc_id'] == idx)
idx_prgs = treedata['id'][iprgs]
return idx_prgs
# -
atree = extract_a_tree(alltrees.data, 110226)
get_progenitors(atree, 107277)
# +
nout_max = alltrees.data['nout'].max()
nout_fi = 187
alltrees.data['nout'] += nout_fi - nout_max
#max_nout = nout_fi
i_final = np.where(alltrees.data["nout"] == nout_fi)
ttt_sub = alltrees.data[i_final]
#ttt.data[0]
# +
def link_circle_up(x, y, r, ax, finish=0):
"""
Given two points, draw circle at the first point and link it to the second point
without drawing the second point by default (so that it can repeat to build a long thread of bids).
for the last point, pass the radius of the last circle to the argument 'finish'
For example,
fig = plt.figure()
ax = fig.add_subplot(111)
xpos = [1,1] & ypos = [2,4]
link_circle(xpos, ypos, 10, ax)
xpos = [1,2] & ypos = [4,6]
link_circle(xpos, ypos, 30, ax, finish=30)
fig.show()
"""
ax.plot(x[0], y[0], 'o', ms=r, lw=2, alpha=0.7, mfc='orange')
ax.plot(x, y, '-', c='black',alpha=0.7)
if finish > 0:
ax.plot(x[1], y[1], 'o', ms=20, lw=2, alpha=0.7, mfc='orange')
def recursive_tree(idx, tt, nstep, ax, x0, y0, dx, mass_unit=1e10):
import tree.draw_merger_tree as dmt
prgs = get_progenitors(tt, idx)
i_this_gal = np.where(tt['id'] == idx)
m = np.sqrt(tt[i_this_gal]["mvir"] / mass_unit)
#print("IDX:", idx, "prgs: ",prgs, "mass:", m, i_this_gal)
nprg = len(prgs)
if nstep == 0:
return
else:
if nprg == 0:
return
else:
if nprg > 1:
dx *= 0.9
# print("Branch!", nprg)
xarr = dmt.get_xarr(nprg) * dx + x0
for i, x in zip(prgs, xarr):
link_circle_up([x0, x], [y0, y0 + 1], m, ax)
recursive_tree(i, tt, nstep - 1, ax, x, y0 + 1, dx, mass_unit=mass_unit)
# + active=""
# It's working well.
# Now search use the tree to calculate construct catalog, and search for mergers.
# -
#iii = np.where(ttt_sub['Orig_halo_id'] == 7)[0]
iii = np.where(ttt_sub['id'] == 110226)[0]
ttt_sub[iii]
# +
import matplotlib.pyplot as plt
galid = 110478
nout_fi = 187
nout_ini = 30
final_gals = ttt_sub['id']
final_gals_org = ttt_sub['Orig_halo_id']
#for galid in final_gals[2:3]:
for galid in [110478]:
print(galid)
fig = plt.figure(figsize=[6,6])
plt.ioff()
ax = fig.add_subplot(111)
#galid = 6033
sidgal = str(galid).zfill(5)
nouts = np.unique(range(nout_fi - nout_ini + 2))
zreds = np.unique(alltrees.data["aexp"])[:len(nouts)]
zreds = ["%.2f" % i for i in zreds]
#print(zreds)
atree = extract_a_tree(alltrees.data, galid)
recursive_tree(galid, atree, 120, ax, 0, 0, 1, mass_unit=1e8)
# y axis label (redshift)
ax.set_ylabel("Redshift")
ax.set_xlim([-3,1])
ax.set_ylim([0,151])
plt.yticks(nouts[1:151:10], zreds[1:151:10])
ax.set_title(sidgal)
#fig.show()
plt.savefig(wdir + "mergertrees/" + sidgal + '.png')
plt.close()
# -
ttt.data.dtype
sum(trees.data['nprog'] > 1)
import matplotlib.pyplot as plt
plt.hist(trees.data['nprog'])
plt.show()
prgs, inds = treeutils.get_main_prg(trees, haloinds=[11])
import matplotlib.pyplot as plt
#%%
import tree.treeplots as trp
a = trees.data[inds]
trp.plot_all(a, a['id'][0], save=True, out_dir=wdir)
| scripts/notebooks/halo/Test_Trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import urllib
import numpy as np
import requests
import matplotlib.pyplot as plt
def main():
print(cv2.__version__)
print("Hello")
if __name__ == "__main__":
main()
# -
"""Get a image from website and display it"""
def get_image(self,url):
req = urllib.request.urlopen(url)
arr = np.asarray(bytearray(req.read()), dtype=np.uint8)
img = cv2.imdecode(arr, cv2.IMREAD_COLOR)
return img
img = cv2.imread('kitchen.jpg')
img_gray = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
# +
""""finding the contors in the image"""
_,contor,_ = cv2.findContours(img_gray,cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_SIMPLE)
print("no of shapes {0}".format(len(contor)))
for cnt in contor:
epsilon = 1*cv2.arcLength(cnt,True)
approx = cv2.approxPolyDP(cnt,epsilon,True)
cnt = approx
rect = cv2.minAreaRect(cnt)
box = cv2.boxPoints(rect)
box = np.int0(box)
cv2.drawContours(img,[box],0,(0,0,255),3)
# -
#cv2.imshow("image",img)
#cv2.imshow("gray",img_gray)
#cv2.waitKey(0) #this is to keep the image display otherwise it will exit immediately
plt.imshow(img_gray)
plt.figure("image")
plt.imshow(img)
plt.show()
#speed test
x = 5
# %timeit y=x*x
# +
import cv2
import numpy as np
import matplotlib.pyplot as plt
image = cv2.imread('kitchen.jpg',0)
#cv2.imshow("orginal",image)
#plt.imshow(image)
down_sized = cv2.resize(image,(64,60))
cv2.imwrite("64x60.png",down_sized)
#plt.imshow(down_sized)
# -
| openCV.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# -*- coding: UTF-8
# -
# # <p style="text-align: center;"> JSTOR Text Analysis Project:<br/> Refining Expert-Built Dictionaries with word2vec
# Authors: <NAME>, <NAME>, <NAME>, <NAME><br/>
# Institution: University of California, Berkeley<br/>
# Date created: July 20, 2018<br/>
# Date last modified: September 22, 2020
# ## Initialize Python
# Install missing packages
# !pip install gensim
# !pip install nltk
# !pip install tqdm
# +
# IMPORTING KEY PACKAGES
import gensim # for word embedding models
import _pickle as cPickle # Optimized version of pickle
import gc # For managing garbage collector
from collections import Counter # For counting terms across the corpus
import re # For filtering most_similar() output--remove surnames
import csv # For saving csv files
import sys; sys.path.insert(0, "../../../data_management/tools/") # To load functions from files in data_management/tools
from textlist_file import write_list, load_list # For saving and loading text lists to/from file
# +
# FOR VISUALIZATIONS
import matplotlib
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE # For visualizing word embeddings
from scipy.spatial import distance # To use cosine distances for tSNE metric
# Visualization parameters
# %pylab inline
# %matplotlib inline
#matplotlib.style.use('white')
import seaborn as sns # To make matplotlib prettier
sns.set(style='white')
#sns.despine()
# +
# Define model file paths (Your job to figure out how to load these in!)
wem_path = "../../../models_storage/word_embeddings_data/word2vec_phrased_filtered_300d_2020_sept5.bin" # old: dec11
#wem_path_npy = "../../../models_storage/word_embeddings_data/word2vec_phrased_filtered_300d_aug14.bin.wv.vectors.npy"
#wem_path_old = "../yoon/word2vec_phrased_filtered_300d_july18.bin"
# Define dictionary file paths:
culture_path = "../../Dictionary Mapping/Dictionaries/core/cultural_core.csv"
relational_path = "../../Dictionary Mapping/Dictionaries/core/relational_core.csv"
demographic_path = "../../Dictionary Mapping/Dictionaries/core/demographic_core.csv"
culture_orgs_path = "../../Dictionary Mapping/Dictionaries/core/cultural_core_orgs.csv"
relational_orgs_path = "../../Dictionary Mapping/Dictionaries/core/relational_core_orgs.csv"
demographic_orgs_path = "../../Dictionary Mapping/Dictionaries/core/demographic_core_orgs.csv"
culture_full_path = "../../Dictionary Mapping/Dictionaries/cultural_original.csv"
relational_full_path = "../../Dictionary Mapping/Dictionaries/relational_original.csv"
demographic_full_path = "../../Dictionary Mapping/Dictionaries/demographic_original.csv"
# -
# ## Define helper functions
def dict_cohere(thisdict, wem_model):
'''Computes the average cosine similarity score of terms within one dictionary with all other terms in that same dictionary,
effectively measuring the coherence of the dictionary.
...question for development: does it make sense to compare the average cosine similarity score between all terms
in thisdict and the average cosine similarity among the total model vocabulary? (Could that be, by definition, 0?)
NOTE: For an unknown reason, calling this function deletes terms from thisdict.
Inputs: List of key terms, word2vec model.
Output: Average cosine similarity score of each word with all other words in the list of key terms.'''
# Initialize average distance variables:
word_avg_dist = 0
word_avg_dists = 0
dict_avg_sim = 0
all_avg_dists = 0
model_avg_dists = 0
# Compute average cosine similarity score of each word with other dict words:
for word in thisdict:
word_avg_dist = (wem_model.wv.distances(word, other_words=thisdict).sum())/len(thisdict) # Total diffs of word with all other words, take average
word_avg_dists += word_avg_dist # Add up each average distance, incrementally
dict_avg_sim = 1 - word_avg_dists/len(thisdict) # Find average cosine similarity score by subtracting avg. distance from 1
#print("Dictionary coherence (avg. cosine similarity): " + str(dict_avg_sim))
return dict_avg_sim
def term_or_part_in_blacklist(term, blacklist):
if term in blacklist:
return True
for part in term.split("_"):
if part in blacklist:
return True
return False
# ## Load & check word2vec model
# +
# Load word2vec model using gensim:
model = gensim.models.KeyedVectors.load(wem_path)
# For reference, standard code looks like:
#model = gensim.models.KeyedVectors.load_word2vec_format(wem_path_old, binary=True)
# -
# ### Check similar terms to authors of foundational texts per perspective
model.most_similar("pfeffer_salancik", topn=50) # foundational relational author
model.most_similar(positive=["meyer_rowan", "dimaggio_powell"], topn=50) # foundational cultural authors
model.most_similar("hannan_freeman", topn=50) # foundational demographic author
# ## Load and refine black list
# +
# Load the text file of surnames for further processing.
with open('../../surnames.txt', 'r') as file:
data = file.read().replace('\n', '')
surname = data.lower().split('|')
with open('../../expanded_dict_blacklist.csv', 'r') as csv_file:
csv_reader = csv.reader(csv_file)
for row in csv_reader:
surname += row
# -
print(surname)
# Add garbage terms into blacklist.
for i in model.wv.vocab:
if ("valign" in i) or ("oasis" in i) or ("colwidth" in i):
surname.append(i)
len(surname)
# ## Remove blacklist terms from model
# +
# https://stackoverflow.com/questions/48941648/how-to-remove-a-word-completely-from-a-word2vec-model-in-gensim
from tqdm.notebook import tqdm
import json
import nltk
nltk.download("wordnet")
from nltk.corpus import wordnet
with open('../../Dictionary Mapping/Dictionaries/words_dictionary.json') as f:
whitelist = json.load(f)
blacklist = set(surname)
def remove_from_w2v(w2v, blacklist):
new_vectors = []
new_vocab = {}
new_index2entity = []
new_vectors_norm = []
for i in tqdm(range(len(w2v.wv.vocab))):
word = w2v.wv.index2entity[i]
vec = w2v.wv.vectors[i]
vocab = w2v.wv.vocab[word]
vec_norm = w2v.wv.vectors_norm[i]
if len(wordnet.synsets(word)) > 0 or all([len(wordnet.synsets(x)) > 0 for x in word.split("_")]):
# if word in whitelist or all([x in whitelist for x in word.split("_")]):
if not word in blacklist or any([w in blacklist for w in word.split("_")]):
vocab.index = len(new_index2entity)
new_index2entity.append(word)
new_vocab[word] = vocab
new_vectors.append(vec)
new_vectors_norm.append(vec_norm)
w2v.wv.vocab = new_vocab
w2v.wv.vectors = np.array(new_vectors)
w2v.wv.index2entity = np.array(new_index2entity)
w2v.wv.index2word = np.array(new_index2entity)
w2v.wv.vectors_norm = np.array(new_vectors_norm)
model.wv.init_sims() # needs to be called for remove_from_w2v to work
print("Vocab size before removal: " + str(len(model.wv.vocab)))
remove_from_w2v(model, surname)
print("Vocab size after: " + str(len(model.wv.vocab)))
# -
# ## Load and clean dictionaries
# ### Cultural dictionary
# +
# Load the raw culture dictionary (full and seed) and expanded version (50 terms + 22 orgs terms).
culture_full = [elem.strip('\n').replace(",", " ") for elem in load_list(culture_full_path)] # Load full culture dictionary
culture_full = list(set(culture_full)) # Remove duplicates
culture_orgs = [elem.strip('\n').replace(",", " ") for elem in load_list(culture_orgs_path)] # Load orgs-specific culture dictionary
culture_orgs = list(set(culture_orgs)) # Remove duplicates
culture = [elem.strip('\n').replace(",", " ") for elem in load_list(culture_path)] # Load seed culture dictionary
culture = list(set(culture)) # Remove duplicates
sorted(culture)
# +
# Remove any terms from culture dict NOT in current model (these will have to be replaced):
for i in range(5):
for word in culture:
if word not in list(model.wv.vocab):
culture.remove(word)
print("Removed " + str(word) + " from culture dictionary.")
# Repeat for quality (second pass sometimes catches more):
for word in culture:
if word not in list(model.wv.vocab):
culture.remove(word)
print("Removed " + str(word) + " from culture dictionary.")
# Remove surnames in culture
for word in culture:
if word in surname:
culture.remove(word)
print("Removed " + str(word) + " from culture dictionary.")
print("Length of culture dictionary filtered into vector space:", len(culture))
culture
# +
coherence, coherence_orgs = dict_cohere(culture, model), dict_cohere(culture_orgs, model)
print("Coherence of " + str(len(culture)) + "-term cultural dictionary: ", str(coherence))
print("Coherence of " + str(len(culture_orgs)) + "-term cultural dictionary: ", str(coherence_orgs))
# -
# Check out most similar words
model.wv.most_similar(culture_orgs, topn=20)
# ### Relational dictionary
# +
# Load the raw + orgs relational dictionaries.
relational = []
for item in load_list(relational_path):
relational.append(item.strip("\n").replace(",", " "))
relational_orgs = [elem.strip('\n').replace(",", " ") for elem in load_list(relational_orgs_path)] # Load orgs-specific culture dictionary
relational_orgs = list(set(relational_orgs)) # Remove duplicates
relational
# +
# Remove any terms from relational dict NOT in current model (these will have to be replaced):
for i in range(5):
for word in relational:
if word not in list(model.wv.vocab):
relational.remove(word)
print("Removed " + str(word) + " from relational dictionary.")
# Repeat for quality (second pass sometimes catches more):
for word in relational:
if word not in list(model.wv.vocab):
relational.remove(word)
print("Removed " + str(word) + " from relational dictionary.")
# Remove surnames in relational
for word in relational:
if word in surname:
relational.remove(word)
print("Removed " + str(word) + " from relational dictionary.")
print()
print("Length of relational dictionary filtered into vector space:", len(relational))
relational
# -
coherence, coherence_orgs = dict_cohere(relational, model), dict_cohere(relational_orgs, model)
print("Coherence of " + str(len(relational)) + "-term relational dictionary: ", str(coherence))
print("Coherence of " + str(len(relational_orgs)) + "-term relational dictionary: ", str(coherence_orgs))
# Check out most similar words
model.wv.most_similar(relational_orgs, topn=20)
# ### Demographic dictionary
# +
# Load the raw + orgs demographic dictionaries.
demographic = []
for item in load_list(demographic_path):
demographic.append(item.strip("\n").replace(",", " "))
demographic_orgs = [elem.strip('\n').replace(",", " ") for elem in load_list(demographic_orgs_path)] # Load orgs-specific culture dictionary
demographic_orgs = list(set(demographic_orgs)) # Remove duplicates
demographic
# +
# Remove any terms from demographic dict NOT in current model (these will have to be replaced):
for i in range(5):
for word in demographic:
if word not in list(model.wv.vocab):
demographic.remove(word)
print("Removed " + str(word) + " from demographic dictionary.")
# Repeat for quality (second pass sometimes catches more):
for word in demographic:
if word not in list(model.wv.vocab):
demographic.remove(word)
print("Removed " + str(word) + " from demographic dictionary.")
# Remove surnames in demographic
for word in demographic:
if word in surname:
demographic.remove(word)
print("Removed " + str(word) + " from demographic dictionary.")
print()
print("Length of demographic dictionary filtered into vector space:", len(demographic))
demographic
# -
coherence, coherence_orgs = dict_cohere(demographic, model), dict_cohere(demographic_orgs, model)
print("Coherence of " + str(len(demographic)) + "-term demographic dictionary: ", str(coherence))
print("Coherence of " + str(len(demographic_orgs)) + "-term demographic dictionary: ", str(coherence_orgs))
# Check out most similar words
model.wv.most_similar(demographic_orgs, topn=20)
# ## Expand dictionaries
# By using the model to look at similar words across terms, create a list of candidate terms for a bigger conceptual dictionary. Manually search all these candidate terms for those that are tightly conceptually related to the seed dictionary. This process blends data-driven search from the model with hand-driven search across the candidate terms.
#
# By searching through the above candidate terms/phrases, expand from the seed terms into a larger--but still conceptually coherent--list!
dictionary_lengths = list(range(30, 100, 10)) + list(range(100, 1001, 100))
expanded_dicts_path = '../../Dictionary Mapping/Dictionaries/Expanded/wordnet_english3/'
filename_template = 'closest_{}_{}.csv' # filename_template.format(perspective, length)
# +
expanded_dict = culture.copy()
for length in dictionary_lengths:
expanded_dict += [x for x, _ in model.wv.most_similar(expanded_dict, topn = length - len(expanded_dict))]
coherence = dict_cohere(expanded_dict, model)
print("Coherence of " + str(length) + "-term cultural dictionary: ", str(coherence))
wtr = csv.writer(open(expanded_dicts_path + filename_template.format('culture', str(length)), 'w'), delimiter=',', lineterminator='\n')
for x in expanded_dict:
wtr.writerow([x])
culture_expanded = expanded_dict.copy()
print()
expanded_dict = relational.copy()
for length in dictionary_lengths:
expanded_dict += [x for x, _ in model.wv.most_similar(expanded_dict, topn = length - len(expanded_dict))]
coherence = dict_cohere(expanded_dict, model)
print("Coherence of " + str(length) + "-term relational dictionary: ", str(coherence))
wtr = csv.writer(open(expanded_dicts_path + filename_template.format('relational', str(length)), 'w'), delimiter=',', lineterminator='\n')
for x in expanded_dict:
wtr.writerow([x])
relational_expanded = expanded_dict.copy()
print()
expanded_dict = demographic.copy()
for length in dictionary_lengths:
if length == 1000:
i = 0
expanded_dict += [x for x, _ in model.wv.most_similar(expanded_dict, topn = length - len(expanded_dict))]
coherence = dict_cohere(expanded_dict, model)
print("Coherence of " + str(length) + "-term demographic dictionary: ", str(coherence))
with open(expanded_dicts_path + filename_template.format('demographic', str(length)), 'w') as f:
wtr = csv.writer(f)
for x in expanded_dict:
wtr.writerow([x])
demographic_expanded = expanded_dict.copy()
# -
# ## Find more blacklist candidates
counter = 0
for w in tqdm(demographic_expanded):
if w not in whitelist and "".join(w.split("_")) not in whitelist:
for part in w.split("_"):
if part not in whitelist:
print(w)
counter += 1
break
print(counter)
counter = 0
for w in tqdm(culture_expanded):
if w not in whitelist and "".join(w.split("_")) not in whitelist:
for part in w.split("_"):
if part not in whitelist:
print(w)
counter += 1
break
print(counter)
counter = 0
for w in tqdm(relational_expanded):
if w not in whitelist and "".join(w.split("_")) not in whitelist:
for part in w.split("_"):
if part not in whitelist:
print(w)
counter += 1
break
print(counter)
| refine_dictionaries/refine_dict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 32-bit
# language: python
# name: python38332bit5e51badee2b94e4eb6b116e6ca3abae6
# ---
# # Background & Overview
# Online retail is a transnational data set which contains all the transactions occurring between January 12, 2010 - September 12, 2011 for a store in the United Kingdoms. The company mainly sells unique all-occasion gifts. I will be pretending to be a data scientist who was hired by a global retail marketing company to analyze consumer data of this UK store, so that the company can create enriching marketing campaigns to attract more customers into buying their gift items.
# # EDA (Exploratory Data Analysis) Part 1
# import required libraries for dataframe and visualization
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import datetime as dt
import altair as alt
from altair import datum
alt.data_transformers.disable_max_rows()
alt.data_transformers.enable('json')
# import required libraries for clustering
import sklearn
from sklearn.preprocessing import StandardScaler
from sklearn.cluster import KMeans
from yellowbrick.cluster import KElbowVisualizer
from sklearn.decomposition import PCA
from sklearn.cluster import SpectralClustering
from sklearn.metrics import silhouette_score
# #### Loading in the dataset into my dataframe variable
df = pd.read_csv(r'D:\OnlineRetail.csv',encoding= 'unicode_escape')
df.head(5)
# ### Creating a more comprehensive look into the datasets columns, data-types, and more.
#
# Instead of listing out the data-types, column names, and other specifices. I decided to create a list-style display to show all information regarding the dataset. You will see that this dataset contains the following below:
# - 8 columns
# - 540k rows
# - 1 int64 datatype
# - 2 float datatypes
# - 5 object datatypes
# - 64k Column distinctive values
# - 13k NULL/NAN values
def initial_eda(df):
if isinstance(df, pd.DataFrame):
total_na = df.isna().sum().sum()
print("Dimensions : %d rows, %d columns" % (df.shape[0], df.shape[1]))
print("Total NA Values : %d " % (total_na))
print("%38s %10s %10s %10s" % ("Column Name", "Data Type", "#Distinct", "NA Values"))
col_name = df.columns
dtyp = df.dtypes
uniq = df.nunique()
na_val = df.isna().sum()
for i in range(len(df.columns)):
print("%38s %10s %10s %10s" % (col_name[i], dtyp[i], uniq[i], na_val[i]))
else:
print("Expect a DataFrame but got a %15s" % (type(df)))
initial_eda(df)
# ### Descriptive Statistics
#
# Since the dataset has a mixture of all values I wanted to run a quick statistical description view on the dataset to gain more instance. From reviewing the output we can see that the columns 'Quantity' and 'Unitprice' are returned but not 'CustomerID'. This was down as CustomerID is more of an identifier, now with Quantity you can see that there is a strange minimum value of -80995.00 and maximum value 80995.00. This tells me that this column is very likely to contain outliers same with the column Unitprice having a minimum value of -11062.06. Later on in the notebook you see that I am removing the low and upper values that contains those such outliers.
df.drop(columns='CustomerID').describe().T
df= df[df.Quantity > 0]
df= df[df.UnitPrice > 0]
df.drop(columns='CustomerID').describe().T
# ### Data cleaning
#
# As shown above we have 132220 null values in total but when breaking it down to individual columns majority of the nulls come from the CustomerID. We can now see that after dropping the rows that contained any NULL values are total entries drop to 397884, this would be about a 10% decrease in the total size we originally had at the start.
df.isnull().sum()
df.dropna(inplace=True)
df.isnull().sum()
initial_eda(df)
# ### Data attribute information
# - InvoiceNo: Invoice number 6-digit integral number, Object.
# - StockCode: Product code 5-digit integral number, Object.
# - Description: Product name, Object.
# - Quantity: The quantities of each product per transaction, int64.
# - InvoiceDate: Invice date and time, int64.
# - UnitPrice: Unit price, int64.
# - CustomerID: Customer number, Object.
# - Country: Country name, Object.
# ### Altering the columns
#
# You will see below me altering the existing columns in the data set as well as adding new columns to strengthen my model further down in the notebook.
# change the column names
df.rename(index=str, columns={'InvoiceNo': 'invoice_num',
'StockCode' : 'stock_code',
'Description' : 'description',
'Quantity' : 'quantity',
'InvoiceDate' : 'invoice_date',
'UnitPrice' : 'unit_price',
'CustomerID' : 'cust_id',
'Country' : 'country'}, inplace=True)
# change the invoice_date format - String to Timestamp format
df['invoice_date'] = pd.to_datetime(df.invoice_date, format='%d-%m-%Y %H:%M')
df['cust_id'] = df['cust_id'].astype('int64')
# change description - UPPER case to LOWER case
df['description'] = df.description.str.lower()
# Creating a new column in the dataframe for total price of a unit and rearranging the dataframe/dataset
df['amount_spent'] = df['quantity'] * df['unit_price']
df= df[['invoice_num','invoice_date','stock_code','description','quantity','unit_price','amount_spent','cust_id','country']]
df.head()
df.insert(loc=2, column='year_month', value=df['invoice_date'].map(lambda x: 100*x.year + x.month))
df.insert(loc=3, column='month', value=df.invoice_date.dt.month)
# # +1 to make Monday=1.....until Sunday=7
df.insert(loc=4, column='day', value=(df.invoice_date.dt.dayofweek)+1)
df.insert(loc=5, column='hour', value=df.invoice_date.dt.hour)
df.head()
# # Data Vizualization
# #### Number of Orders for different Customers
# As we can see here that the data is fairly balanced with the number of orders made by customers except for the few outliers that we do have. These outliers can be seen with the bars that break out of the normal trend of not passing 2k-3k orders made. We can also see that The top 4 out of 5 customers who have the most orders are from the United Kingdoms.
# +
orders = df.groupby(by=['cust_id','country'], as_index=False)['invoice_num'].count()
plt.subplots(figsize=(15,6))
plt.plot(orders.cust_id, orders.invoice_num)
plt.xlabel('Customers ID')
plt.ylabel('Number of Orders')
plt.title('Number of Orders for different Customers')
plt.show()
# -
print('The TOP 5 customers with most number of orders...')
orders.sort_values(by='invoice_num', ascending=False).head()
# #### Money Spent for different Customers
# From what you can see below you have the most money spent per customer but with this being shown there are more outliers given an a lower amount of data that is in a normal range you can see that when the amount spent goes over 50k you can detemine this to be an outlier. Lastly, when comparing the top 5 orders to top 5 money spent there is only one match and that is.
# - 14911 EIRE with 5675 orders and $143,825.06 spent
# +
money_spent = df.groupby(by=['cust_id','country'], as_index=False)['amount_spent'].sum()
plt.subplots(figsize=(15,6))
plt.plot(money_spent.cust_id, money_spent.amount_spent)
plt.xlabel('Customers ID')
plt.ylabel('Money spent (Dollar)')
plt.title('Money Spent for different Customers')
plt.show()
# -
print('The TOP 5 customers with highest money spent...')
money_spent.sort_values(by='amount_spent', ascending=False).head()
# #### Number of orders for different Months (1st Dec 2010 - 9th Dec 2011)
# Below you will see the number of orders that have been place from the 1st Dec 2010 - 9th Dec 2011. In the bar graph you can see that there is a trend occurring every 2 to 3 months where the number of orders are rising and failing. If there is more to investigate would to see why there is a consistent rise and fall in throughout the year.
color = sns.color_palette()
ax = df.groupby('invoice_num')['year_month'].unique().value_counts().sort_index().plot(kind='bar',color=color[0],figsize=(15,6))
ax.set_xlabel('Month',fontsize=15)
ax.set_ylabel('Number of Orders',fontsize=15)
ax.set_title('Number of orders for different Months (1st Dec 2010 - 9th Dec 2011)',fontsize=15)
ax.set_xticklabels(('Dec_10','Jan_11','Feb_11','Mar_11','Apr_11','May_11','Jun_11','July_11','Aug_11','Sep_11','Oct_11','Nov_11','Dec_11'), rotation='horizontal', fontsize=13)
plt.show()
# #### Number of orders for different Days
#
# We can see here that the most popular day to order an item is on Thrusday. This can be useful to increase targeting customers before and during this day to increase it further or find out why that is so that we can bring up the weekend orders.
ax = df.groupby('invoice_num')['day'].unique().value_counts().sort_index().plot(kind='bar',color=color[0],figsize=(15,6))
ax.set_xlabel('Day',fontsize=15)
ax.set_ylabel('Number of Orders',fontsize=15)
ax.set_title('Number of orders for different Days',fontsize=15)
ax.set_xticklabels(('Mon','Tue','Wed','Thur','Fri','Sun'), rotation='horizontal', fontsize=15)
plt.show()
# #### Number of orders for different Hours
#
# From reviewing the chart below we can see that the optimal range of when customer buy items is between 10am-3pm. This will be very useful to try and push ads through out this timeframe.
ax = df.groupby('invoice_num')['hour'].unique().value_counts().iloc[:-1].sort_index().plot(kind='bar',color=color[0],figsize=(15,6))
ax.set_xlabel('Hour',fontsize=15)
ax.set_ylabel('Number of Orders',fontsize=15)
ax.set_title('Number of orders for different Hours',fontsize=15)
ax.set_xticklabels(range(6,21), rotation='horizontal', fontsize=15)
plt.show()
# #### Money Spent by different Countries
#
# We can see that that number one country that spent the most money is the United Kingdom, however that does not give the full picture as to what the other countries spent. So you can see here that I have a chart below that shows the total amount spent by country with and without the UK so that we can get a full look into the amount spent for every country.
# +
group_country_amount_spent = df.groupby('country')['amount_spent'].sum().sort_values()
# del group_country_orders['United Kingdom']
# plot total money spent by each country (with UK)
plt.subplots(figsize=(15,8))
group_country_amount_spent.plot(kind='barh', fontsize=12, color=color[0])
plt.xlabel('Money Spent (Dollar)', fontsize=12)
plt.ylabel('Country', fontsize=12)
plt.title('Money Spent by different Countries', fontsize=12)
plt.show()
# +
group_country_amount_spent = df.groupby('country')['amount_spent'].sum().sort_values()
del group_country_amount_spent['United Kingdom']
# plot total money spent by each country (without UK)
plt.subplots(figsize=(15,8))
group_country_amount_spent.plot(kind='barh', fontsize=12, color=color[0])
plt.xlabel('Money Spent (Dollar)', fontsize=12)
plt.ylabel('Country', fontsize=12)
plt.title('Money Spent by different Countries', fontsize=12)
plt.show()
# -
# # Feature Engineering
# Since we can see that the mass amounts of order buy and have are from the United Kingdom let us focus on this particular country as this is the biggest customer. To havee a better breakdown on to what the Unit Kingdom is ordering/buying I am going to create a couple of frames to display the information.
# - Number of days since last purchase
# - Number of transactions
# - Total amount of transactions (revenue contributed)
df.head()
# ### Creating three field that will be used to show clustering data.
#
# I am creating three new fields and assigning them a new dataframe in which I will be reviewing for the clustering of the dataset.
# - (Recency): Number of days since last purchase
# - (Frequency): Number of tracsactions
# - (Monetary): Total amount of transactions (revenue contributed)
# New Attribute : Monetary
df['Amount'] = df['quantity']*df['unit_price']
amount = df.groupby('cust_id')['Amount'].sum()
amount = amount.reset_index()
amount.head()
# New Attribute : Frequency
Frequency = df.groupby('cust_id')['invoice_num'].count()
Frequency = Frequency.reset_index()
Frequency.columns = ['cust_id', 'Frequency']
Frequency.head()
AF = pd.merge(amount, Frequency, on='cust_id', how='inner')
AF.head()
# Compute the maximum date to know the last transaction date
max_date = max(df['invoice_date'])
max_date
df['Diff'] = max_date - df['invoice_date']
df.head()
DaysSince = df.groupby('cust_id')['Diff'].min().dt.days
DaysSince = DaysSince.reset_index()
DaysSince.head()
AFD = pd.merge(AF, DaysSince, on='cust_id', how='inner')
AFD.columns = ['CustomerID', 'Amount', 'Frequency', 'Recency']
AFD.head()
# ### Outlier check.
#
# Since we created new fields with in the dataframe I would like to take away any outliers it may have. To better help me see this I am using a box and whisker plot to do so. Then I am taking the data I have found and removing it.
attributes = ['Amount','Frequency','Recency']
plt.rcParams['figure.figsize'] = [10,8]
sns.boxplot(data = AFD[attributes], orient="v", palette="Set2" ,whis=1.5,saturation=1, width=0.7)
plt.title("Outliers Variable Distribution", fontsize = 14, fontweight = 'bold')
plt.ylabel("Range", fontweight = 'bold')
plt.xlabel("Attributes", fontweight = 'bold')
# +
# Removing (statistical) outliers for Amount
Q1 = AFD.Amount.quantile(0.05)
Q3 = AFD.Amount.quantile(0.95)
IQR = Q3 - Q1
AFD = AFD[(AFD.Amount >= Q1 - 1.5*IQR) & (AFD.Amount <= Q3 + 1.5*IQR)]
# Removing (statistical) outliers for Recency
Q1 = AFD.Recency.quantile(0.05)
Q3 = AFD.Recency.quantile(0.95)
IQR = Q3 - Q1
AFD = AFD[(AFD.Recency >= Q1 - 1.5*IQR) & (AFD.Recency <= Q3 + 1.5*IQR)]
# Removing (statistical) outliers for Frequency
Q1 = AFD.Frequency.quantile(0.05)
Q3 = AFD.Frequency.quantile(0.95)
IQR = Q3 - Q1
AFD = AFD[(AFD.Frequency >= Q1 - 1.5*IQR) & (AFD.Frequency <= Q3 + 1.5*IQR)]
# -
# # Build the clustering models with comparison
# ### Rescaling the Attributes
# It is extremely important to rescale the variables so that they have a comparable scale.| There are two common ways of rescaling:
#
# Min-Max scaling
# Standardisation (mean-0, sigma-1)
# Here, we will use Standardisation Scaling.
AFD = AFD[['Amount', 'Frequency', 'Recency']]
# Instantiate
scaler = StandardScaler()
# fit_transform
AFD_scaled = scaler.fit_transform(AFD)
AFD_scaled.shape
AFD_scaled = pd.DataFrame(AFD_scaled)
AFD_scaled.columns = ['Amount', 'Frequency', 'Recency']
AFD_scaled.head()
# ### K-Means Clustering
# K-means clustering is one of the simplest and popular unsupervised machine learning algorithms.
#
# The algorithm works as follows:
#
# First we initialize k points, called means, randomly.
# We categorize each item to its closest mean and we update the mean’s coordinates, which are the averages of the items categorized in that mean so far.
# We repeat the process for a given number of iterations and at the end, we have our clusters.
kmeans = KMeans(n_clusters=4, max_iter=50)
kmeans.fit(AFD_scaled)
kmeans.labels_
# ### Finding the Optimal Number of Clusters
#
# Elbow Curve to get the right number of Clusters.
#
# A fundamental step for any unsupervised algorithm is to determine the optimal number of clusters into which the data may be clustered. The Elbow Method is one of the most popular methods to determine this optimal value of k.
# Elbow-curve/SSD
ssd = []
range_n_clusters = [2, 3, 4, 5, 6, 7, 8]
for num_clusters in range_n_clusters:
kmeans = KMeans(n_clusters=num_clusters, max_iter=50)
kmeans.fit(AFD_scaled)
ssd.append(kmeans.inertia_)
# plot the SSDs for each n_clusters
plt.plot(ssd)
k_means = KMeans(n_clusters= 3, init='k-means++', random_state=0).fit(AFD_scaled)
clusters = k_means.predict(AFD_scaled)
AFD['Cluster_Label'] = cl_labels_k = k_means.labels_
AFD.head()
# ### K-Means Clustering with 3 Cluster Ids Inference
#
# - Customers with Cluster Id 2 are the customers with high amount of transactions as compared to other customers.
# - Customers with Cluster Id 2 are frequent buyers.
# - Customers with Cluster Id 2 are not recent buyers and hence least of importance from business point of view.
# - Customer with Cluster Id 0,1 are very low compared to the second cluster and this will have to be targeted more
# - Customer with Cluster Id 0,1 are also mostly likely not to be frequent buyers. We should investigate the reason as to why they are low compared to Cluster 2.
sns.boxplot(x='Cluster_Label', y='Amount', data=AFD)
sns.boxplot(x='Cluster_Label', y='Frequency', data=AFD)
sns.boxplot(x='Cluster_Label', y='Recency', data=AFD)
# # Conclusion
#
# We can see here after looking at the cluster zero, one, and two that the customer that fail into the cluster 2 does not need to be focused on to grow the sales of the store. But what needs to happen is that the store will have to focus more on the customer who fall into the clusters of 0 and 1 becasue both clusters compared to cluster 2 did not report as well on the amount of spend and how often they bought an item. From this we will have to create some marketing strategies to target this specific customer to spend more and become more frequent when buy our items. More inmportantl customer from cluster 1 as they are the news customers to the store.
| Assignments&Projects/Clustering/Clustering ML.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tasks 2020
# Task submissions for October - December 2020
# ***
#
# ### Task 1
# Write a Python function called counts that takes a list as input and returns a dictionary of unique items in the list as keys and the number of times each item appears as values
# ***
# * For this task I write a function `counts` which takes a list as an argument
# * There are a number of possible methods which add lists to dictionaries, such as using list comrehensions as well and through the importation of modules (`collections` module) amongst others. This blog post at geesforgeeks.org details some of these [1]
# * I found the simplest and most useful solution on stackoverflow.com which I demonstrate below [2]. This method is also suggested at kite.com [3] <br>
# <br>
#
# ##### Function description
# * Inside the `counts` function I create an empty dictionary `d`
# * Next I use a `for` loop to iterate over the list, adding a list-item to the dictionary on each iteration
# * Inside the `for` loop I use conditional statements to determine whether a new key-value pair should be added to the dictionary. If the key exists already, its corresponding value increases by 1
# * When the `for` loop ends, the dictionary containing the list items is returned
# * Finally, I call the function and print the returned dictionary to the console
# <br>
#
#
#
#
# <br>
#
# ### References
#
# [1] geeksforgeeks.com; Python | Ways to create a dictionary of Lists; https://www.geeksforgeeks.org/python-ways-to-create-a-dictionary-of-lists/ <br>
# [2] stackoverflow.com; How to add list elements into dictionary https://stackoverflow.com/questions/30208044/how-to-add-list-elements-into-dictionary<br>
# [3] kite.com; How to append an element to a key in a dictionary with Python; https://www.kite.com/python/answers/how-to-append-an-element-to-a-key-in-a-dictionary-with-python <br>
# <br>
#
# ***
#
# **Function code**
# create a simple list of 5 elements
l = ['A', 'A', 'B', 'C', 'A']
# +
'''
This function takes a list as an argument and adds list items to a dictionary
'''
def counts(lst):
# empy dictionary d initialised
d = {}
# iterate through list items and add to dict d
for item in lst:
if item in d:
d[item] += 1
else:
d[item] = 1
return d
# -
# call function and print to console
print(counts(l))
# #### End task 1
# ***
# <br>
# <br>
#
# ### Task 2
# Write a Python function called `dicerolls()` that simulates rolling dice. Your function should take two parameters: the number of dice *k* and the number of times to roll the dice *n*. The function should simulate randomly rolling *k* dice *n* times, keeping track of each total face value. It should then return a dictionary with the number of times each possible total face value occurred.
# ***
# * As requested by the task, the function takes two parameters: k (no. of dice) and n (no. of rolls).
# * I have used nested for loops to carry out the simulation. I have based the algorithm used on useful information about nested loops at w3schools.com [1].
# * The outer loop simulates the number of times the dice are rolled while the inner loop simulates the number of dice.
# * To simulate a random roll of a single die, I use the `integers()` function from `numpy.random` package.
# * On every iteration of the outer loop (n dice rolls), the inner loop runs k times (no. of dice)
# * The results of each iteration of the inner loop are appended to list l (initialised as an empty list within the function)
# * I use the function from Task 1 (above) to add each list item to dictionary d (initialised as an empty dictionary within the function)
# * The function returns a sorted dictionary. I found the code for this on a discussion on stackoverflow.com [2]
# * I call the `dicerolls()` 4 times. First with 2 dice, then 3, 4 and 5. The dice are rolled 100,000 times on each functon call.
# <br>
#
# ##### Plotting the output
# * I plot the output of each function call on a separate bar charts. To plot a dictionary on a bar chart, I used code found on a stackoverflow.com discussion [3].
# * I used the matplotlib documentation to plot 4 subplots on a single figure [4].
# * From observation of the bar charts, it is clear that the results are normally distributed, with the curve becoming increasingly rounded the more dice are thrown.
#
#
#
#
# <br>
#
# ### References
# [1] w3schools; Python Nested Loops; https://www.w3schools.com/python/gloss_python_for_nested.asp <br>
# [2] https://stackoverflow.com/questions/9001509/how-can-i-sort-a-dictionary-by-key%22 <br>
# [3] https://stackoverflow.com/questions/21195179/plot-a-histogram-from-a-dictionary <br>
# [4] Pyplot tutorial; intro to pyplot; https://matplotlib.org/tutorials/introductory/pyplot.html<br>
# <br>
#
# #### Build function and run simulation
# import default_rng for random number generation, matplotlib.pyplot for visualisation
from numpy.random import default_rng
import matplotlib.pyplot as plt
# construct a new Generator object
rng = default_rng()
# +
'''
This function simulates the rolling of k=int dice, n=int times
'''
def dicerolls(k, n):
l = [] # initialise empty list
d = {} # initialise empty dict
for roll in range(n): # outer loop simulates no. of times dice are rolled
dice_sum = 0 # dice_sum counter set to 0
for die in range(k): # inner loop simulates each of k dice thrown at random
dice = rng.integers(1, 7)
dice_sum += dice # face value of each dice added together
l.append(dice_sum)
d = counts(l) # function from Task 1 called
return dict(sorted(d.items()))
# -
# 2 dice rolled 100,000 times and dictionary output
two_dice = dicerolls(2, 100000)
two_dice
# <br>
#
# #### Plot output: Different values for k (number of dice)
# +
# plot 4 subplots on a single figure for better visualisation comparison
plt.figure(figsize=(10, 7))
# plot bar chart for 2 dice rolled
plt.subplot(221)
plt.bar(list(two_dice.keys()), two_dice.values())
plt.title('2 dice')
# plot bar chart for 3 dice rolled
three_dice = dicerolls(3, 100000)
plt.subplot(222)
plt.bar(list(three_dice.keys()), three_dice.values())
plt.title('3 dice')
# plot bar chart for 4 dice rolled
four_dice = dicerolls(4, 100000)
plt.subplot(223)
plt.bar(list(four_dice.keys()), four_dice.values())
plt.title('4 dice')
# plot bar chart for 5 dice rolled
five_dice = dicerolls(5, 100000)
plt.subplot(224)
plt.bar(list(five_dice.keys()), five_dice.values())
plt.title('5 dice')
# -
# ##### Note the increased roundedness of the curve the more dice are thrown. All 4 have the appearance of a normal distribution.
#
# <br>
# #### End task 2
# ***
# <br>
#
#
# ### Task 3
#
# Write some python code that simulates flipping a coin 100 times. Then run this code 1,000 times, keeping track of the number of heads in each of the 1,000 simulations. Select an appropriate plot to depict the resulting list of 1,000 numbers, showing that it roughly follows a bell-shaped curve. Use the `numpy.random.binomial` function to generate the simulation.
#
# * To demonstrate the binomial distribution in practical terms, the coin toss example is often used [1], [2], [3].
# * The coin toss is also an example of a Bernoulli trial. This is a single trial from which there are exactly two possible outcomes, usually denoted as 'success' or 'failure'. [4]
# * When we toss a fair coin, there are only two possible outcomes - heads or tails - and each outcome has an equal probability (p=0.5) of arising.
# * If we say that 'heads' denotes a 'success', we can perform the coin toss n number of times, counting the number of successes we observe. This number will have a binomial distribution.
# * Using rng.binomial below, I simulate the coin toss and plot the resulting distribution on a histogram. The bell shaped curve is evident.
# * There are 100 trials (n=100), with a 50% probability of success (p=0.5). This is performed 1,000 times.
#
# <br>
#
# ### References
# [1] <NAME>; Python for Data 22: Probability Distributions; https://www.kaggle.com/hamelg/python-for-data-22-probability-distributions <br>
# [2] onlinestatsbook.com; Binomial Distribution; http://onlinestatbook.com/2/probability/binomial.html <br>
# [3] Wikipedia; Bernoulli Trial; https://en.wikipedia.org/wiki/Bernoulli_trial<br>
# [4] *Ibid* <br>
# construct a new Generator object
rng = default_rng()
n, p = 100, .5 # number of trials, probability of each trial
unbiased_coin = rng.binomial(n, p, 10000) # result of tossing a coin 10 times, tested 1000 times.
plt.hist(unbiased_coin, color='green')
plt.title("unbiased coin")
plt.show()
# This has the appearance of a normal distribution (explored below). If the probability of success is changed however (i.e. we add a bias to the coin), we observe a change in the shape of the distribution - it becomes asymmetrical:
n, p = 100, .9 # number of trials, probability of success
biased_coin = rng.binomial(n, p, 10000) # result of tossing a coin 10 times, tested 10000 times.
plt.hist(biased_coin, color='green')
plt.title("biased coin")
plt.show()
# #### End task 3
# ***
#
# <br>
#
#
# ### Task 4
#
# Use numpy to create four data sets, each with an `x` array and a corresponding `y` array, to demonstrate Simpson’s paradox. You might create your `x` arrays using `numpy.linspace` and create the `y` array for each `x` using notation like `y = a * x + b` where you choose the `a` and `b` for each `x`, `y` pair to demonstrate the paradox. You might see the Wikipedia page for Simpson’s paradox for inspiration.
# <br>
#
#
# #### Simpson's Paradox
#
# * Simpson's Paradox is a phenomenon in statistics whereby seemingly contradictory trends are observed within the same data set, depending on how the data set is analysed.
# * For example, the data may be split into four smaller groups and within each of these groups, a positive trend is observed. However when taken in aggregate, the overall trend is seen to be negative [1].
#
# <br>
#
# * In order to demonstrate this phenomenon I have generated four data sets below, using `numpy.linspace`. I then plot these data sets on a graph for visualisation.
#
# <br>
#
#
# #### Data set generation
# * First, I import `numpy` and `matplotlib.pylot` for data set generation and subsequent visualisation respectively:
import numpy as np
import matplotlib.pyplot as plt
# <br>
#
#
# * Next, I generate four data sets of equal sample size (20).
# * The coordinates for the x values and the corresponding y values have been intentionally chosen to so that each individual data set will show a positive trend.
# * These same data sets, when taken in aggregate however, show an overall negative trend.
# * When plotted on a graph (below), Simpson's paradox can be visualised and thus understood intuitively. There are many examples of such visualisations which guided the selection of values for my data sets [2], [3], [4].
# * The `linspace` function generates the x array while the corresponding y array is populated using the formula `a * x + b` as suggested in the task brief.
# * I have added the function `numpy.radnom.randn` to the formula in order to closer simulate the random nature of reality. I use code found on a discussion on stackoverflow.com for this [5].
#
# <br>
#
# +
# Assign values for a and b
a = 1
b = 20
# Generate an x array using the numpy.linspace function
x = np.linspace(3, 10, 20)
# Calculate corresponding y values
y = a * x + b + np.random.randn(*x.shape)
# +
b = 15
x2 = np.linspace(5, 12, 20)
y2 = a * x + b + np.random.randn(*x2.shape)
# +
b = 10
x3 = np.linspace(7, 17, 20)
y3 = a * x + b + np.random.randn(*x3.shape)
# +
b = 5
x4 = np.linspace(9, 22, 20)
y4 = a * x + b + np.random.randn(*x4.shape)
# -
# <br>
#
#
# #### Plotting and Visualtisation
#
# * When we plot and visualise the data on a scatter plot, we observe a positive trend in each of the four data sets individually, while there is a clearly observable negative trend in the overall relationship between the variables if we take the data in aggregate.
# * To generate this plot, I used code found in a discussion on stackoverflow.com [6].
#
# <br>
# <br>
#
#
# When we plot one individual data set, we can clearly see a positive trend in the relationship between the x and y variables.
# Plot single data set 'x'
plt.scatter (x, y)
plt.xlim(0, 15)
plt.ylim(20, 40)
# <br>
#
#
# Plotting each data set on the same plot however, reveals a different picture. On the plot below, we observer that each of the four individual data is associated with a positive trend, while the overall data reveals that there is a negative trend in the relationship between both variables.
# +
# Initialise a figure with a single axes
fig = plt.figure()
ax1 = fig.add_subplot(111)
# Plot each data set on a scatter plot
ax1.scatter(x, y, s=10, c='b', marker="s", label='x')
ax1.scatter(x2 ,y2 , s=10, c='r', marker="o", label='x2')
ax1.scatter(x3, y3, s=10, c='g', marker="x", label='x3')
ax1.scatter(x4 ,y4 , s=10, c='y', marker="^", label='x4')
plt.xlim(-5, 30)
plt.ylim(0, 40)
plt.legend(loc='upper right');
plt.xlabel('supply', fontsize=16)
plt.ylabel('price', fontsize=16)
plt.show()
# -
# * In a real world scenario, this data example might represent a price-supply relationship (units are arbitrary) [7].
# * Looking at the data in totality, when the supply of products increases, we see a reduction in cost of those products overall.
# * However, taken individually, each product displays a price-supply trend which seems to contradict the overall direction.
# * There may be any number of plausible reasons why this might be the case, depending on the context of the data.
# * Generally speaking, this is an important aspect to take into account when considering Simpson's paradox. The statistical evidence may not always be sufficient in explaining the results it yields.
# * One must always be aware of the real world context in which the data was collected and exists
#
# ### References
#
# [1] wikipedia.org; Simpson's Paradox; https://en.wikipedia.org/wiki/Simpson%27s_paradox <br>
# [2] <NAME>.; Simpson's Paradox in Psychological Science: A Practical Guide; https://www.researchgate.net/figure/Alcohol-use-and-intelligence-Simulated-data-illustrating-that-despite-a-positive_fig2_256074671 <br>
# [3] stackexchange.com; Examples of Simpson's Paradox being resolved by choosing the aggregate data; https://stats.stackexchange.com/questions/478463/examples-of-simpsons-paradox-being-resolved-by-choosing-the-aggregate-data <br>
# [4] wikipedia.org; Simpson's Paradox; https://en.wikipedia.org/wiki/Simpson%27s_paradox <br>
# [5] stackoverflow.com; Linear regression minimizing errors only above the linear; https://stackoverflow.com/questions/58090204/linear-regression-minimizing-errors-only-above-the-linear <br>
# [6] stackoverflow.com; MatPlotLib: Multiple datasets on the same scatter plot; https://stackoverflow.com/questions/4270301/matplotlib-multiple-datasets-on-the-same-scatter-plot <br>
# [7] icoachmath.com; Trend; http://www.icoachmath.com/math_dictionary/Trend <br>
# <br>
#
# #### End task 4
# ***
| FDA _Tasks2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning from Pre-Trained Models with Keras
#
# ## Introduction
#
# ImageNet, an image recognition benchmark dataset*, helped trigger the modern AI explosion. In 2012, the AlexNet architecture (a deep convolutional-neural-network) rocked the ImageNet benchmark competition, handily beating the next best entrant. By 2014, all the leading competitors were deep learning based. Since then, accuracy scores continued to improve, eventually surpassing human performance.
#
# In this hands-on tutorial we will build on this pioneering work to create our own neural-network architecture for image recognition. Participants will use the elegant Keras deep learning programming interface to build and train TensorFlow models for image classification tasks on the CIFAR-10 / MNIST datasets*. We will demonstrate the use of transfer learning* (to give our networks a head-start by building on top of existing, ImageNet pre-trained, network layers*), and explore how to improve model performance for standard deep learning pipelines. We will use cloud-based interactive Jupyter notebooks to work through our explorations step-by-step. Once participants have successfully trained their custom model we will show them how to submit their model's predictions to Kaggle for scoring*.
#
# This tutorial aims to prepare participants for the HPC Saudi 2020 Student AI Competition.
#
# Participants are expected to bring their own laptops and sign-up for free online cloud services (e.g., Google Colab, Kaggle). They may also need to download free, open-source software prior to arriving for the workshop.
#
# This tutorial assumes some basic knowledge of neural networks. If you’re not already familiar with neural networks, then you can learn the basics concepts behind neural networks at [course.fast.ai](https://course.fast.ai/).
#
# * Tutorial materials are derived from:
# * [PyTorch Tutorials](https://github.com/kaust-vislab/pytorch-tutorials) by <NAME>.
# * [What is torch.nn really?](https://pytorch.org/tutorials/beginner/nn_tutorial.html) by <NAME>, <NAME>, <NAME>.
# * [Machine Learning Notebooks](https://github.com/ageron/handson-ml2) (2nd Ed.) by <NAME>.
# * *Deep Learning with Python* by <NAME>.
# ### Jupyter Notebooks
#
# This is a Jupyter Notebook. It provides a simple, cell-based, IDE for developing and exploring complex ideas via code, visualizations, and documentation.
#
# A notebook has two primary types of cells: i) `markdown` cells for textual notes and documentation, such as the one you are reading now, and ii) `code` cells, which contain snippets of code (typically *Python*, but also *bash* scripts) that can be executed.
#
# The currently selected cell appears within a box. A green box indicates that the cell is editable. Clicking inside a *code* cell makes it selected and editable. Double-click inside *markdown* cells to edit.
#
# Use `Tab` for context-sensitive code-completion assistance when editing Python code in *code* cells. For example, use code assistance after a `.` seperator to find available object members. For help documentation, create a new *code* cell, and use commands like `dir(`*module*`)`, `help(`*topic*`)`, `?`*name*, or `??`*function* for user provided *module*, *topic*, variable *name*, or *function* name. The magic `?` and `??` commands show documentation / source code in a separate pane.
#
# Clicking on `[Run]` or pressing `Ctrl-Enter` will execute the contents of a cell. A *markdown* cell converts to its display version, and a *code* cell runs the code inside. To the left of a *code* cell is a small text bracket `In [ ]:`. If the bracket contains an asterix, e.g., `In [*]:`, that cell is currently executing. Only one cell executes at a time (if multiple cells are *Run*, they are queued up to execute in the order they were run). When a *code* cell finishes executing, the bracket shows an execution count in the bracket – each *code* cell execution increments the counter and provides a way to determine the order in which codes were executed – e.g., `In [7]` for the seventh cell to complete.
#
# The output produced by a *code* cell appears at the bottom of that cell after it executes. The output generated by a code cell includes anything printed to the output during execution (e.g., print statements, or thrown errors) and the final value generated by the cell (i.e., not the intermediate values). The final value is 'pretty printed' by Jupyter.
#
# Typically, notebooks are written to be executed in order, from top to bottom. Behind the scenes, however, each Notebook has a single Python state (the `kernel`), and each *code* cell that executes, modifies that state. It is possible to modify and re-run earlier cells; however, care must be taken to also re-run any other cells that depend upon the modified one. List the Python state global variables with the magic command `%wgets`. The *kernel* can be restarted to a known state, and cell output cleared, if the Python state becomes too confusing to fix manually (choose `Restart & Clear Output` from the Jupyter `Kernel` menu) – this requires running each *code* cell again.
#
# Complete user documentation is available at [jupyter-notebook.readthedocs.io](https://jupyter-notebook.readthedocs.io/en/stable/notebook.html#notebook-user-interface). <br/>
# Many helpful tips and techniques from [28 Jupyter Notebook Tips, Tricks, and Shortcuts](https://www.dataquest.io/blog/jupyter-notebook-tips-tricks-shortcuts/).
# ## Setup
# ### Create a Kaggle Account
#
# #### 1. Register for an account
#
# In order to download Kaggle competition data you will first need to create a [Kaggle](https://www.kaggle.com/) account.
#
# #### 2. Create an API key
#
# Once you have registered for a Kaggle account you will need to create [API credentials](https://github.com/Kaggle/kaggle-api#api-credentials) in order to be able to use the `kaggle` CLI to download data.
#
# * Go to the `Account` tab of your user profile,
# * and click `Create New API Token` from the API section.
#
# This generates a `kaggle.json` file (with 'username' and 'key' values) to download.
#
#
# ### Setup Colab
#
# In order to run this notebook in [Google Colab](https://colab.research.google.com) you will need a [Google Account](https://accounts.google.com/). Sign-in to your Google account, if necessary, and then start the notebook.
#
# Change Google Colab runtime to use GPU:
#
# * Click `Runtime` -> `Change runtime type` menu item
# * Specify `Runtime type` as `Python 3`
# * Specify `Hardware accelerator` as `GPU`
# * Click **[Save]** button
#
# The session indicator (toolbar / status ribbon under menu) should briefly appear as `Connecting...`. When the session restarts, continue with the next cell (specifying TensorFlow version v2.x):
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# ### Download Data
#
# There are two image datasets ([CIFAR10](https://www.cs.toronto.edu/~kriz/cifar.html) and [MNIST](http://yann.lecun.com/exdb/mnist/index.html)) which these tutorial / exercise notebooks use.
#
# These datasets are available from a variety of sources, including this repository – depending on how the notebook was launched (e.g., Git+LFS / Binder contains entire repository, Google Colab only contains the notebook).
#
# Because data is the fundamental fuel for deep learning, we need to ensure the required datasets for this tutorial are available to the current notebook session. The following steps will ensure the data is already available (or downloaded), and cached where Keras can find them.
#
# Follow instructions and run the cells below to acquire required datasets:
# +
import pathlib
import tensorflow.keras.utils as Kutils
def cache_mnist_data():
for n in ["mnist.npz", "kaggle/train.csv", "kaggle/test.csv"]:
path = pathlib.Path("../datasets/mnist/%s" % n).absolute()
DATA_URL = "file:///" + str(path)
data_file_path = Kutils.get_file(n.replace('/','-mnist-'), DATA_URL)
print("cached file: %s" % n)
def cache_cifar10_data():
for n in ["cifar-10.npz", "cifar-10-batches-py.tar.gz"]:
path = pathlib.Path("../datasets/cifar10/%s" % n).absolute()
DATA_URL = "file:///" + str(path)
if path.is_file():
data_file_path = Kutils.get_file(n, DATA_URL)
print("cached file: %s" % n)
else:
print("FAILED: First fetch file: %s" % n)
def cache_models():
for n in ["vgg16_weights_tf_dim_ordering_tf_kernels_notop.h5"]:
path = pathlib.Path("../models/%s" % n).absolute()
DATA_URL = "file:///" + str(path)
if path.is_file():
data_file_path = Kutils.get_file(n, DATA_URL, cache_subdir='models')
print("cached file: %s" % n)
# -
# #### Download MNIST Data
# If you are using Binder to run this notebook, then the data is already downloaded and available. Skip to the next step.
#
# If you are using Google Colab to run this notebook, then you will need to download the data before proceeding.
#
# ##### Download MNIST from Kaggle
#
# **Note:** Before attempting to download the competition data you will need to login to your [Kaggle](https://www.kaggle.com) account and accept the rules for this competition.
#
# Set your Kaggle username and API key (from the `kaggle.json` file) into the cell below, and execute the code to download the Kaggle [Digit Recognizer: Learn computer vision with the famous MNIST data](https://www.kaggle.com/c/digit-recognizer) competition data.
# + language="bash"
# # NOTE: Replace YOUR_USERNAME and YOUR_API_KEY with actual credentials
# export KAGGLE_USERNAME="YOUR_USERNAME"
# export KAGGLE_KEY="YOUR_API_KEY"
# kaggle competitions download -c digit-recognizer -p ../datasets/mnist/kaggle
# + language="bash"
# unzip -n ../datasets/mnist/kaggle/digit-recognizer.zip -d ../datasets/mnist/kaggle
# -
# ##### (Alternative) Download MNIST from GitHub
#
# If you are running this notebook using Google Colab, but did *not* create a Kaggle account and API key, then dowload the data from our GitHub repository by running the code in the following cells.
# +
import pathlib
import requests
def fetch_mnist_data():
RAW_URL = "https://github.com/holstgr-kaust/keras-tutorials/raw/master/datasets/mnist"
DEST_DIR = pathlib.Path('../datasets/mnist')
DEST_DIR.mkdir(parents=True, exist_ok=True)
for n in ["mnist.npz", "kaggle/train.csv", "kaggle/test.csv", "kaggle/sample_submission.csv"]:
path = DEST_DIR / n
if not path.is_file(): # Don't download if file exists
with path.open(mode = 'wb') as f:
response = requests.get(RAW_URL + "/" + n)
f.write(response.content)
# -
fetch_mnist_data()
cache_mnist_data()
# ##### (Alternative) Download MNIST with Keras
#
# If you are running this notebook using Google Colab, but did *not* create a Kaggle account and API key, then dowload the data using the Keras load_data() API by running the code in the following cells.
from tensorflow.keras.datasets import mnist
cache_mnist_data()
mnist.load_data();
# #### Download CIFAR10 Data
#
# If you are using Binder to run this notebook, then the data is already downloaded and available. Skip to the next step.
#
# If you are using Google Colab to run this notebook, then you will need to download the data before proceeding.
#
# ##### Download CIFAR10 from Kaggle
#
# **Note:** Before attempting to download the competition data you will need to login to your [Kaggle](https://www.kaggle.com) account.
#
# Set your Kaggle username and API key (from the `kaggle.json` file) into the cell below, and execute the code to download the Kaggle [Digit Recognizer: Learn computer vision with the famous MNIST data](https://www.kaggle.com/c/digit-recognizer) competition data.
# + language="bash"
# # NOTE: Replace YOUR_USERNAME and YOUR_API_KEY with actual credentials
# export KAGGLE_USERNAME="YOUR_USERNAME"
# export KAGGLE_KEY="YOUR_API_KEY"
# kaggle datasets download guesejustin/cifar10-keras-files-cifar10load-data -p ../datasets/cifar10/
# + language="bash"
# unzip -n ../datasets/cifar10/cifar10-keras-files-cifar10load-data.zip -d ../datasets/cifar10
# -
# ##### (Alternative) Download CIFAR10 from GitHub
#
# If you are running this notebook using Google Colab, but did *not* create a Kaggle account and API key, then dowload the data from our GitHub repository by running the code in the following cells.
# +
import os
import pathlib
import requests
def fetch_cifar10_data():
RAW_URL = "https://github.com/holstgr-kaust/keras-tutorials/raw/master/datasets/cifar10"
DEST_DIR = pathlib.Path('../datasets/cifar10')
DEST_DIR.mkdir(parents=True, exist_ok=True)
for n in ["cifar-10.npz", "cifar-10-batches-py.tar.gz"]:
path = DEST_DIR / n
if not path.is_file(): # Don't download if file exists
with path.open(mode = 'wb') as f:
response = requests.get(RAW_URL + "/" + n)
f.write(response.content)
print("downloaded file: %s" % n)
# -
fetch_cifar10_data()
cache_cifar10_data()
# + language="bash"
# DEST_DIR='../datasets/cifar10'
# tar xvpf "${DEST_DIR}/cifar-10-batches-py.tar.gz" --directory="${DEST_DIR}"
# -
# ##### (Alternative) Download CIFAR10 with Keras
#
# If you are running this notebook using Google Colab, but did *not* create a Kaggle account and API key, then dowload the data using the Keras load_data() API by running the code in the following cells.
from tensorflow.keras.datasets import cifar10
cache_cifar10_data()
cifar10.load_data();
# ## Tutorial
#
# ### Setup
#
# Initialize the Python environment by importing and verifying the modules we will use.
import os
import sys
import pathlib
import random
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow.keras as keras
# `%matplotlib inline` is a magic command that makes *matplotlib* charts and plots appear was outputs in the notebook.
#
# `%matplotlib notebook` enables semi-interactive plots that can be enlarged, zoomed, and cropped while the plot is active. One issue with this option is that new plots appear in the active plot widget, not in the cell where the data was produced.
# %matplotlib inline
# Now check the runtime environment to ensure it can run this notebook. If there is an `Exception`, or if there are no GPUs, you will need to run this notebook in a more capable environment (see `README.md`, or ask instructor for additional help).
# +
# Verify runtime environment
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
IS_COLAB = True
except Exception:
IS_COLAB = False
print("is_colab:", IS_COLAB)
assert tf.__version__ >= "2.0", "TensorFlow version >= 2.0 required."
print("tensorflow_version:", tf.__version__)
assert sys.version_info >= (3, 5), "Python >= 3.5 required."
print("python_version:", "%s.%s.%s-%s" % (sys.version_info.major,
sys.version_info.minor,
sys.version_info.micro,
sys.version_info.releaselevel
))
print("executing_eagerly:", tf.executing_eagerly())
__physical_devices = tf.config.experimental.list_physical_devices('GPU')
if len(__physical_devices) == 0:
print("No GPUs available. Expect training to be very slow.")
if IS_COLAB:
print("Go to `Runtime` > `Change runtime` and select a GPU hardware accelerator."
"Then `Save` to restart session.")
else:
print("is_built_with_cuda:", tf.test.is_built_with_cuda())
print("is_gpu_available:", tf.test.is_gpu_available(), [d.name for d in __physical_devices])
# -
# ### CIFAR10 - Dataset Processing
#
# The previously acquired CIFAR10 dataset is the essential input needed to train an image classification model. Before using the dataset, there are several preprocessing steps required to load the data, and create the correctly sized training, validation, and testing arrays used as input to the network.
#
# The following data preparation steps are needed before they can become inputs to the network:
#
# * Cache the downloaded dataset (to use Keras `load_data()` functionality).
# * Load the dataset (CIFAR10 is small, and fits into a `numpy` array).
# * Verify the shape and type of the data, and understand it...
# * Convert label indices into categorical vectors.
# * Convert image data from integer to float values, and normalize.
# * Verify converted input data.
# #### Cache Data
#
# Make downloaded data available to Keras (and check if it's really there). Provide dataset utility functions.
# Cache CIFAR10 Datasets
cache_cifar10_data()
# + language="bash"
# find ~/.keras -name "cifar-10*" -type f
# -
# These helper function assist with managing the three label representations we will encounter:
#
# * label index: a number representing a class
# * label names: a *human readable* text representation of a class
# * category vector: a vector space to represent the categories
#
# The label index `1` represents an `automobile`, and `2` represents a `bird`; but, `1.5` doesn't make a `bird-mobile`. We need a representation where each dimension is a continuum of that feature. There are 10 distinct categories, so we encode them as a 10-dimensional vector space, where the i-th dimension represents the i-th class. An `automobile` becomes `[0,1,0,0,0,0,0,0,0,0]`, a `bird` becomes `[0,0,1,0,0,0,0,0,0,0]` (these are called *one-hot encodings*), and a `bird-mobile` (which we couldn't represent previously) can be encoded as `[0,0.5,0.5,0,0,0,0,0,0,0]`.
#
# **Note:** We already know how our dataset is represented. Typically, one would load the data first, analyse the class representation, and then write the helper functions.
# +
# Helper functionality to provide human-readable labels
cifar10_label_names = ['airplane', 'automobile',
'bird', 'cat', 'deer', 'dog', 'frog', 'horse',
'ship', 'truck']
def cifar10_index_label(idx):
return cifar10_label_names[int(idx)]
def cifar10_category_label(cat):
return cifar10_index_label(cat.argmax())
def cifar10_label(v):
return cifar10_index_label(v) if np.isscalar(v) or np.size(v) == 1 else cifar10_category_label(v)
# -
# #### Load Data
#
# Datasets for classification require two parts: i) the input data (`x` in our nomenclature), and ii) the labels (`y`). Classifiction takes an `x` as input, and returns a `y` (the class) as output.
#
# When training a model from a dataset (called the `train`ing dataset), it is important to keep some of the data aside (called the `test` set). If we didn't, the model could just memorize the data without learning a generalization that would apply to novel related data. The `test` set is used to evaluate the typical real performance of the model.
# +
from tensorflow.keras.datasets import cifar10
# The data, split between train and test sets:
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# -
# **Note:** Backup plan: Run the following cell if the data didn't load via `cifar10.load_data` above.
# Try secondary data source if the first didn't work
try:
print("data loaded." if type((x_train, y_train, x_test, y_test)) else "load failed...")
except NameError:
with np.load('../datasets/cifar10/cifar-10.npz') as data:
x_train = data['x_train']
y_train = data['y_train']
x_test = data['x_test']
y_test = data['y_test']
print("alternate data load." if type((x_train, y_train, x_test, y_test)) else "failed...")
# #### Explore Data
#
# Explore data types, shape, and value ranges. Ensure they make sense, and you understand the data well.
print('x_train type:', type(x_train), ',', 'y_train type:', type(y_train))
print('x_train dtype:', x_train.dtype, ',', 'y_train dtype:', y_train.dtype)
print('x_train shape:', x_train.shape, ',', 'y_train shape:', y_train.shape)
print('x_test shape:', x_test.shape, ',', 'y_test shape:', y_test.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
print('x_train (min, max, mean): (%s, %s, %s)' % (x_train.min(), x_train.max(), x_train.mean()))
print('y_train (min, max): (%s, %s)' % (y_train.min(), y_train.max()))
# * The data is stored in Numpy arrays.
# * The datatype for both input data and labels is a small unsigned int. They represent different things though. The input data represents pixel value, the labels represent the category.
# * There are 50000 training data samples, and 10000 testing samples.
# * Each input sample is a colour images of 32x32 pixels, with 3 channels of colour (RGB), for a total size of 3072 bytes. Each label sample is a single byte.
# * A 32x32 pixel, 3-channel colour image (2-D) can be represented as a point in a 3072 dimensional vector space.
# * We can see that pixel values range between 0-255 (that is the range of `uint8`) and the mean value is close to the middle. The label values range between 0-9, which corresponds to the 10 categories the labels represent.
#
# Lets explore the dataset visually, looking at some actual images, and get a statistical overview of the data.
#
# Most of the code in the plotting function below is there to tweak the appearance of the output. The key functionality comes from `matplotlib` functions `imshow` and `hist`, and `numpy` function `histogram`.
def cifar10_imageset_plot(img_data=None):
(x_imgs, y_imgs) = img_data if img_data else (x_train, y_train)
fig = plt.figure(figsize=(16,8))
for i in range(40):
plt.subplot(4, 10, i + 1)
plt.xticks([])
plt.yticks([])
idx = int(random.uniform(0, x_imgs.shape[0]))
plt.title(cifar10_label(y_imgs[idx]))
plt.imshow(x_imgs[idx], cmap=plt.get_cmap('gray'))
plt.show()
# Show array of random labelled images with matplotlib (re-run cell to see new examples)
cifar10_imageset_plot((x_train, y_train))
def histogram_plot(img_data=None):
(x_data, y_data) = img_data if img_data else (x_train, y_train)
hist, bins = np.histogram(y_data, bins = range(int(y_data.min()), int(y_data.max() + 2)))
fig = plt.figure(figsize=(12,5))
plt.subplot(1,2,1)
plt.hist(y_data, bins = range(int(y_data.min()), int(y_data.max() + 2)))
plt.xticks(range(int(y_data.min()), int(y_data.max() + 2)))
plt.title("y histogram")
plt.subplot(1,2,2)
plt.hist(x_data.flat, bins = range(int(x_data.min()), int(x_data.max() + 2)))
plt.title("x histogram")
plt.tight_layout()
plt.show()
print('y histogram counts:', hist)
histogram_plot((x_train, y_train))
histogram_plot((x_test, y_test))
# The data looks reasonable: there are sufficient examples for each category (y_train) and a near-normal distribution of pixel values that appears similar in both the train and test datasets.
#
# The next aspect of the input data to grapple with is how the input vector space corresponds with the output category space. Is the correspondence simple, e.g., distances in the input space relate to distances in the output space; or, more complex.
#
# ##### Visualizing training samples using PCA
#
# [Principal Components Analysis (PCA)](https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) can be used as a visualization tool to see if there are any obvious patterns in the training samples.
#
# PCA re-represents the input data by changing the basis vectors that represent them. These new orthonormal basis vectors (eigen vectors) represent variance in the data (ordered from largest to smallest). Projecting the data samples onto the first few (2 or 3) dimensions will let us see the data with the biggest differences accounted for.
#
# The following cell uses `scikit-learn` to calculate PCA eigen vectors for a random subset of the data (10%).
# +
import sklearn
import sklearn.decomposition
_prng = np.random.RandomState(42)
pca = sklearn.decomposition.PCA(n_components=40, random_state=_prng)
x_train_flat = x_train.reshape(*x_train.shape[:1], -1)
y_train_flat = y_train.reshape(y_train.shape[0])
print("x_train:", x_train.shape, "y_train", y_train.shape)
print("x_train_flat:", x_train_flat.shape, "y_train_flat", y_train_flat.shape)
pca_train_features = pca.fit_transform(x_train_flat, y_train_flat)
print("pca_train_features:", pca_train_features.shape)
# Sample 10% of the PCA results
_idxs = _prng.randint(y_train_flat.shape[0], size=y_train_flat.shape[0] // 10)
pca_features = pca_train_features[_idxs]
pca_category = y_train_flat[_idxs]
print("pca_features:", pca_features.shape,
"pca_category", pca_category.shape,
"min,max category:", pca_category.min(), pca_category.max())
# -
def pca_components_plot(components_, shape_=(32, 32, 3)):
fig = plt.figure(figsize=(16,8))
for i in range(min(40, components_.shape[0])):
plt.subplot(4, 10, i + 1)
plt.xticks([])
plt.yticks([])
eigen_vect = (components_[i] - np.min(components_[i])) / np.ptp(pca.components_[i])
plt.title('component: %s' % i)
plt.imshow(eigen_vect.reshape(shape_), cmap=plt.get_cmap('gray'))
plt.show()
# This plot shows the new eigen vector basis functions suggested by the PCA analysis. Any image in our dataset can be created as a linear combination of these basis vectors. At a guess, the most prevalent feature of the dataset is that there is something at the centre of the image that is distinct from the background (components 0 & 2) and there is often a difference between 'land' & 'sky' (component 1) – compare with the sample images shown previously.
pca_components_plot(pca.components_)
# These are 2D and 3D scatter plot functions that colour the points by their labels (so we can see if any 'clumps' of points correspond to actual categories).
def category_scatter_plot(features, category, title='CIFAR10'):
num_category = 1 + category.max() - category.min()
fig, ax = plt.subplots(1, 1, figsize=(12, 10))
cm = plt.cm.get_cmap('tab10', num_category)
sc = ax.scatter(features[:,0], features[:,1], c=category, alpha=0.4, cmap=cm)
ax.set_xlabel("Component 1")
ax.set_ylabel("Component 2")
ax.set_title(title)
plt.colorbar(sc)
plt.show()
# +
from mpl_toolkits.mplot3d import Axes3D
def category_scatter3d_plot(features, category, title='CIFAR10'):
num_category = 1 + category.max() - category.min()
mean_feat = np.mean(features, axis=0)
std_feat = np.std(features, axis=0)
min_range = mean_feat - std_feat
max_range = mean_feat + std_feat
fig = plt.figure(figsize=(12, 10))
cm = plt.cm.get_cmap('tab10', num_category)
ax = fig.add_subplot(111, projection='3d')
sc = ax.scatter(features[:,0], features[:,1], features[:,2],
c=category, alpha=0.85, cmap=cm)
ax.set_xlabel("Component 1")
ax.set_ylabel("Component 2")
ax.set_zlabel("Component 3")
ax.set_title(title)
ax.set_xlim(2.0 * min_range[0], 2.0 * max_range[0])
ax.set_ylim(2.0 * min_range[1], 2.0 * max_range[1])
ax.set_zlim(2.0 * min_range[2], 2.0 * max_range[2])
plt.colorbar(sc)
plt.show()
# -
category_scatter_plot(pca_features, pca_category, title='CIFAR10 - PCA')
# **Note:** 3D PCA plot works best with `%matplotlib notebook` to enable interactive rotation (enabled at start of session).
category_scatter3d_plot(pca_features, pca_category, title='CIFAR10 - PCA')
# The data in its original image space does not appear to cluster into corresponding categories.
#
# ##### Visualizing training sample using t-SNE
#
# [t-distributed Stochastic Neighbor Embedding (t-SNE)](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html#sklearn.manifold.TSNE) is a tool to visualize high-dimensional data. It converts similarities between data points to joint probabilities and tries to minimize the Kullback-Leibler divergence between the joint probabilities of the low-dimensional embedding and the high-dimensional data. For more details on t-SNE including other use cases see this excellent *Toward Data Science* [blog post](https://towardsdatascience.com/an-introduction-to-t-sne-with-python-example-5a3a293108d1).
#
# Informally, t-SNE is preserving the local neighbourhood of data points to help uncover the manifold on which the data lies. For example, a flat piece of paper with two coloured (e.g., red and blue) regions would be a simple manifold to characterize in 3D space; but, if the paper is crumpled up, it becomes very hard to characterize in the original 3D space (blue and red regions could be very close in this representational space) – instead, by following the cumpled paper (manifold) we would recover the fact that blue and red regions are really very distant, and not nearby at all.
#
# It is highly recommended to use another dimensionality reduction method (e.g. PCA) to reduce the number of dimensions to a reasonable amount if the number of features is very high. This will suppress some noise and speed up the computation of pairwise distances between samples.
#
# * [An Introduction to t-SNE with Python Example](https://towardsdatascience.com/an-introduction-to-t-sne-with-python-example-5a3a293108d1)
# +
import sklearn
import sklearn.decomposition
import sklearn.pipeline
import sklearn.manifold
_prng = np.random.RandomState(42)
embedding2_pipeline = sklearn.pipeline.make_pipeline(
sklearn.decomposition.PCA(n_components=0.95, random_state=_prng),
sklearn.manifold.TSNE(n_components=2, random_state=_prng))
embedding3_pipeline = sklearn.pipeline.make_pipeline(
sklearn.decomposition.PCA(n_components=0.95, random_state=_prng),
sklearn.manifold.TSNE(n_components=3, random_state=_prng))
# +
# Sample 10% of the data
_prng = np.random.RandomState(42)
_idxs = _prng.randint(y_train_flat.shape[0], size=y_train_flat.shape[0] // 10)
tsne_features = x_train_flat[_idxs]
tsne_category = y_train_flat[_idxs]
print("tsne_features:", tsne_features.shape,
"tsne_category", tsne_category.shape,
"min,max category:", tsne_category.min(), tsne_category.max())
# +
# t-SNE is SLOW (but can be GPU accelerated!);
# lengthy operation, be prepared to wait...
transform2_tsne_features = embedding2_pipeline.fit_transform(tsne_features)
print("transform2_tsne_features:", transform2_tsne_features.shape)
for i in range(2):
print("min,max features[%s]:" % i,
transform2_tsne_features[:,i].min(),
transform2_tsne_features[:,i].max())
# -
category_scatter_plot(transform2_tsne_features, tsne_category, title='CIFAR10 - t-SNE')
# **Note:** Skip this step during the tutorial, it will take too long to complete.
# +
# t-SNE is SLOW (but can be GPU accelerated!);
# extremely lengthy operation, be prepared to wait... and wait...
transform3_tsne_features = embedding3_pipeline.fit_transform(tsne_features)
print("transform3_tsne_features:", transform3_tsne_features.shape)
for i in range(3):
print("min,max features[%s]:" % i,
transform3_tsne_features[:,i].min(),
transform3_tsne_features[:,i].max())
# -
category_scatter3d_plot(transform3_tsne_features, tsne_category, title='CIFAR10 - t-SNE')
# t-SNE relates the data points (images) according to their closest neighbours. Hints of underlying categories appear; but are not cleanly seperable into the original categories.
# #### Data Conversion
#
# The data type for the training data is `uint8`, while the input type for the network will be `float32` so the data must be converted. Also, the labels need to be categorical, or *one-hot encoded*, as discussed previously. Keras provides utility functions to convert labels to categories (`to_categorical`), and `numpy` makes it easy to perform operations over entire arrays.
#
# * https://keras.io/examples/cifar10_cnn/
# +
num_classes = (y_train.max() - y_train.min()) + 1
print('num_classes =', num_classes)
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# +
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
train_data = (x_train, y_train)
test_data = (x_test, y_test)
# -
# After the data conversion, notice that the datatypes are `float32`, the input `x` data shapes are the same; but, the `y` classification labels are now 10-dimensional, instead of scalar.
# +
print('x_train type:', type(x_train))
print('x_train dtype:', x_train.dtype)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print('y_train type:', type(y_train))
print('y_train dtype:', y_train.dtype)
print('y_train shape:', y_train.shape)
print('y_test shape:', y_test.shape)
# -
# ## Acquire Pre-Trained Network
#
# Download an *ImageNet* pretrained VGG16 network[<sup>1</sup>](#fn1), sans classification layer, shaped for 32x32px colour images<sup>[*](https://github.com/fchollet/deep-learning-models/releases/download/v0.1/vgg16_weights_tf_dim_ordering_tf_kernels_notop.h5)</sup> (the smallest supported size). This image-feature detection network is an example of a deep CNN (Convolutional Neural Network).
#
# **Note:** The network must be fixed – it was already trained on a very large dataset, so training it on our smaller dataset would result in it un-learning valuable generic features.
#
# <span id="fn1"><sup>[1]</sup> *Very Deep Convolutional Networks for Large-Scale Image Recognition** by <NAME> and <NAME>, [arXiv (2014)](https://arxiv.org/abs/1409.1556).</span>
cache_models()
# +
from tensorflow.keras.applications import VGG16
conv_base = VGG16(weights='imagenet', include_top=False, input_shape=(32, 32, 3))
conv_base.trainable = False
conv_base.summary()
# -
# The summary shows the layers, starting from the InputLayer and proceeding through Conv2D convolutional layers, which are then collected at MaxPooling2D layers.
#
# A convolutional kernel is a small matrix that looks for a specific, localized, pattern on its inputs. This pattern is called a `feature`. The kernel is applied at each location on the input image, and the output is another image – a feature image – that represent the strength of that feature at the given location.
#
# Because the inputs to convolution are images, and the outputs are also images – but transformed into a different feature space – it is possible to stack many convolutional layers on top of each other.
#
# A feature image can be reduced in size with a MaxPooling2D layer. This layer 'pools' an `MxN` region to a single value, taking the largest value from the 'pool'. The 'Max' in 'MaxPooling' is keeping the *best* evidence for that feature, found in the original region.
#
# The InputLayer shape and data type should match with the input data:
#
# *Note:* The first dimension of the shape will differ; the input layer has `None` to indicate it accepts a batch sized collection of arrays of the remaining shape. The input data shape will indicate, in that first axis, how many samples it contains.
print("input layer shape:", conv_base.layers[0].input.shape)
print("input layer dtype:", conv_base.layers[0].input.dtype)
print("input layer type:", type(conv_base.layers[0].input))
print("input data shape:", x_train.shape)
print("input data dtype:", x_train.dtype)
print("input data type:", type(x_train))
# ### Explore Convolutional Layers
#
# The following are visualization functions (and helpers) for understanding what the convolutional layers in a network have learned.
#
# We may ask questions about each convolutional kernal in a convolutional layer:
#
# * What local features is the kernel looking for: `visualize_conv_layer_weights`
# * For a given input image, what feature image will the kernal produce: `visualize_conv_layer_output`
# * What input image makes the kernel respond most strongly: `visualize_conv_layer_response`
# +
def cifar10_image_plot(img_data=None, image_index=None):
(x_imgs, y_imgs) = img_data if img_data else (x_train, y_train)
if not image_index:
image_index = int(random.uniform(0, x_imgs.shape[0]))
plt.imshow(x_imgs[image_index], cmap='gray')
plt.title("%s" % cifar10_label(y_imgs[image_index]))
plt.xlabel("#%s" % image_index)
plt.show()
return image_index
def get_model_layer(model, layer_name):
if type(layer_name) == str:
layer = model.get_layer(layer_name)
else:
m = model
for ln in layer_name:
model = m
m = m.get_layer(ln)
layer = m
return (model, layer)
# -
def visualize_conv_layer_weights(model, layer_name):
(model, layer) = get_model_layer(model, layer_name)
layer_weights = layer.weights[0]
max_size = layer_weights.shape[3]
col_size = 12
row_size = int(np.ceil(float(max_size) / float(col_size)))
print("conv layer: %s shape: %s size: (%s,%s) count: %s" %
(layer_name,
layer_weights.shape,
layer_weights.shape[0], layer_weights.shape[1],
max_size))
fig, ax = plt.subplots(row_size, col_size, figsize=(12, 1.2 * row_size))
idx = 0
for row in range(0,row_size):
for col in range(0,col_size):
ax[row][col].set_xticks([])
ax[row][col].set_yticks([])
if idx < max_size:
ax[row][col].imshow(layer_weights[:, :, 0, idx], cmap='gray')
else:
fig.delaxes(ax[row][col])
idx += 1
plt.tight_layout()
plt.show()
def visualize_conv_layer_output(model, layer_name, image_index=None):
(model, layer) = get_model_layer(model, layer_name)
layer_output = layer.output
if not image_index:
image_index = cifar10_image_plot()
intermediate_model = keras.models.Model(inputs = model.input, outputs=layer_output)
intermediate_prediction = intermediate_model.predict(x_train[image_index].reshape(1,32,32,3))
max_size = layer_output.shape[3]
col_size = 10
row_size = int(np.ceil(float(max_size) / float(col_size)))
print("conv layer: %s shape: %s size: (%s,%s) count: %s" %
(layer_name,
layer_output.shape,
layer_output.shape[1], layer_output.shape[2],
max_size))
fig, ax = plt.subplots(row_size, col_size, figsize=(12, 1.2 * row_size))
idx = 0
for row in range(0,row_size):
for col in range(0,col_size):
ax[row][col].set_xticks([])
ax[row][col].set_yticks([])
if idx < max_size:
ax[row][col].imshow(intermediate_prediction[0, :, :, idx], cmap='gray')
else:
fig.delaxes(ax[row][col])
idx += 1
plt.tight_layout()
plt.show()
# +
from tensorflow.keras import backend as K
def process_image(x):
epsilon = 1e-5
# Normalizes the tensor: centers on 0, ensures that std is 0.1 Clips to [0, 1]
x -= x.mean()
x /= (x.std() + epsilon)
x *= 0.1
x += 0.5
x = np.clip(x, 0, 1)
x *= 255
x = np.clip(x, 0, 255).astype('uint8')
return x
def generate_response_pattern(model, conv_layer_output, filter_index=0):
#step_size = 1.0
epsilon = 1e-5
img_tensor = tf.Variable(tf.random.uniform((1, 32, 32, 3)) * 20 + 128.0, trainable=True)
response_model = keras.models.Model([model.inputs], [conv_layer_output])
for i in range(40):
with tf.GradientTape() as gtape:
layer_output = response_model(img_tensor)
loss = K.mean(layer_output[0, :, :, filter_index])
grads = gtape.gradient(loss, img_tensor)
grads /= (K.sqrt(K.mean(K.square(grads))) + epsilon)
img_tensor = tf.Variable(tf.add(img_tensor, grads))
img = np.array(img_tensor[0])
return process_image(img)
# -
def visualize_conv_layer_response(model, layer_name):
(model, layer) = get_model_layer(model, layer_name)
layer_output = layer.output
max_size = layer_output.shape[3]
col_size = 12
row_size = int(np.ceil(float(max_size) / float(col_size)))
print("conv layer: %s shape: %s size: (%s,%s) count: %s" %
(layer_name,
layer_output.shape,
layer_output.shape[1], layer_output.shape[2],
max_size))
fig, ax = plt.subplots(row_size, col_size, figsize=(12, 1.2 * row_size))
idx = 0
for row in range(0,row_size):
for col in range(0,col_size):
ax[row][col].set_xticks([])
ax[row][col].set_yticks([])
if idx < max_size:
img = generate_response_pattern(model, layer_output, idx)
ax[row][col].imshow(img, cmap='gray')
ax[row][col].set_title("%s" % idx)
else:
fig.delaxes(ax[row][col])
idx += 1
plt.tight_layout()
plt.show()
# Looking at the the first 4 convolution layers, we see that:
#
# * All the kernels are 3x3 (i.e., 9 elements each)
# * Layers 1 & 2 have 64 kernels each (64 different possible features)
# * Layers 3 & 4 have 128 kernels each (128 different possible features)
# * Light pixels indicate preference for an activated pixel
# * Dark pixels indicate preference for an inactive pixel
# * The kernels seem to represent edges and lines at various angles
for n in [l.name for l in conv_base.layers if isinstance(l, keras.layers.Conv2D)][:4]:
visualize_conv_layer_weights(conv_base, n)
# For the given input image, show the corresponding feature image. At the lower level layers (e.g., first Conv2D layer), the feature images seem to capture concepts like 'edges' or maybe 'solid colour'?
#
# At higher layers, the size of the feature images decrease because of the MaxPooling. They also appear more abstract – harder to visually recognize than the original image – however, the features are spatially related to the original image (e.g., if there is a white/high value in the lower-left corner of the feature image, then somewhere on the lower-left corner of the original image, there exists pixels that the network is confident represent the feature in question).
image_index = cifar10_image_plot()
for n in [l.name for l in conv_base.layers if isinstance(l, keras.layers.Conv2D)][:7]:
visualize_conv_layer_output(conv_base, n, image_index)
# This plot shows which input images cause the greatest response from the convolution kernels. At lower layers, we see many simple 'wave' textures showing that these kernals like to see edges at particular angles. At lower-middle layers, the paterns show larger scale and more complexity (like dots and curves); but, still lots of angled edges.
for n in [l.name for l in conv_base.layers if isinstance(l, keras.layers.Conv2D)][:4]:
visualize_conv_layer_response(conv_base, n)
# The patterns in the higher levels can get even more complex; but, some of them don't seem to encode for anything but noise. Maybe these could be pruned to make a smaller network...
#
# **Note:** Skip this step during the tutorial, it will take too long to complete.
# NOTE: Visualize mid to higher level convolutional layers;
# lengthy operation, be prepared to wait...
for n in [l.name for l in conv_base.layers if isinstance(l, keras.layers.Conv2D)][4:]:
visualize_conv_layer_response(conv_base, n)
# ### CNN Base + Classifier Model
#
# Create a simple model that has the pre-trained CNN (Convolutional Neural Network) as a base, and adds a basic classifier on top.
#
# The new layer types are Flatten, Dense, Dropout, and Activation.
#
# The Flatten layer reshapes the input dimensions (2D + 1 channel) into a single dimension.
#
# The Dense(x) layer is a layer of (`x`) neurons (represented as a flat 1D array) connected to a flat input. The size of the input and outputs do not need to match.
#
# The Dropout(x) layer withholds a random fraction (`x`) of the input neurons from training during each batch of data. This limits the ability of the network to `overfit` on the training data (i.e., memorize training data, rather than learn generalizable rules).
#
# Activation is an essential part of (or addition to) each layer. Layers like Dense are simply linear functions (weighted sums + a bias). Without a non-linear component, the network could not learn a non-linear function. Activations like 'relu' (Rectified Linear Unit), 'tanh', or 'sigmoid' are functions to introduce a non-linearity. They also clamp output values within known ranges.
#
# The 'softmax' activation is used to produce probability distributions over multiple categories.
#
# This example uses the Sequential API to build the final network.
#
# * [Activation Functions in Neural Networks](https://towardsdatascience.com/activation-functions-neural-networks-1cbd9f8d91d6)
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Flatten, Dense, Activation, Dropout
from tensorflow.keras.applications import VGG16
def create_cnnbase_classifier_model(conv_base=None):
if not conv_base:
conv_base = VGG16(weights='imagenet', include_top=False, input_shape=(32, 32, 3))
conv_base.trainable = False
model = Sequential()
model.add(conv_base)
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
return model
# -
# Create our model *model_transfer_cnn* by calling the creation function *create_cnnbase_classifier_model* above.
#
# Notice the split of total parameters (\~15 million) between trainable (\~0.3 million for our classifier) and non-trainable (\~14.7 million for the pre-trained CNN).
#
# Note also that the final Dense layer squeezes the network down to the number of categories.
model_transfer_cnn = create_cnnbase_classifier_model(conv_base)
model_transfer_cnn.summary()
# ### Train Model
#
# Training a model typically involves setting relevant hyperparameters that control aspects of the training process. Common hyperparameters include:
#
# * `epochs`: The number of training passes through the entire dataset. The number of epochs depends upon the complexity of the dataset, and how effectively the network architecture of the model can learn it. If the value is too small, the model accuracy will be low. If the value is too big, then the training will take too long for no additional benefit, as the model accuracy will plateau.
# * `batch_size`: The number of samples to train during each step. The number should be set so that the GPU memory and compute are well utilized. The `learning_rate` needs to be set accordingly.
# * `learning_rate`: The step-size to update model weights during the training update phase (backpropagation). Too small, and learning takes too long. Too large, and we may step over the minima we are trying to find. The learning rate can be increased as the batch sizes increases (with some caveats), on the assumption that with more data in a larger batch, the error gradient will be more accurate, so therefore, we can take a larger step.
# * `decay`: Used by some optimizers to decrease the `learning_rate` over time, on the assumption that as we get closer to our goal, we should focus on smaller refinement steps.
batch_size = 128 #32
epochs = 25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
# The model needs to be compiled prior to use. This step enables the model to train efficiently on the GPU device.
#
# This step also specifies the loss functions, accuracy metrics, learning strategy (optimizers), and more.
#
# Our `loss` is *categorical_crossentropy* because we are doing multi-category classification.
#
# We use an RMSprop optimizer, which is a varient of standard gradient descent optimizers that also includes momentum. Momentum is used to speed up learning in directions where it has been making more progress.
#
# * [A Look at Gradient Descent and RMSprop Optimizers](https://towardsdatascience.com/a-look-at-gradient-descent-and-rmsprop-optimizers-f77d483ef08b)
# * [Understanding RMSprop — faster neural network learning](https://towardsdatascience.com/understanding-rmsprop-faster-neural-network-learning-62e116fcf29a)
# +
from tensorflow.keras.optimizers import RMSprop
model_transfer_cnn.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
# -
# The model `fit` function trains the network, and returns a history of training and testing accuracy.
#
# *Note:* Because we already have a test dataset, and we are not validating our hyperparameters, we will use the test dataset for validation. We could have also reserved a fraction of the training data to use for validation.
history = model_transfer_cnn.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# ### Evaluate Model
# Visualize accuracy and loss for training and validation.
#
# * https://keras.io/visualization/
def history_plot(history):
fig = plt.figure(figsize=(12,5))
plt.title('Model accuracy & loss')
# Plot training & validation accuracy values
ax1 = fig.add_subplot()
#ax1.set_ylim(0, 1.1 * max(history.history['loss']+history.history['val_loss']))
ax1.set_prop_cycle(color=['green', 'red'])
p1 = ax1.plot(history.history['loss'], label='Train Loss')
p2 = ax1.plot(history.history['val_loss'], label='Test Loss')
# Plot training & validation loss values
ax2 = ax1.twinx()
ax2.set_ylim(0, 1.1 * max(history.history['accuracy']+history.history['val_accuracy']))
ax2.set_prop_cycle(color=['blue', 'orange'])
p3 = ax2.plot(history.history['accuracy'], label='Train Acc')
p4 = ax2.plot(history.history['val_accuracy'], label='Test Acc')
ax1.set_ylabel('Loss')
ax1.set_xlabel('Epoch')
ax2.set_ylabel('Accuracy')
pz = p3 + p4 + p1 + p2
plt.legend(pz, [l.get_label() for l in pz], loc='center right')
plt.show()
# The history plot shows characteristic features of training performance over successive epochs. Accuracy and loss are related, in that a reduction in loss produces an increase in accuracy. The graph shows characteristic arcs for training and testing accuracy / loss over training time (epochs).
#
# The primary measure to improve is *testing accuracy*, because that indicates how well the model generalizes to data it must typically classify.
#
# The accuracy curves show that testing accuracy has plateaued (with some variability), while training accuracy increases (but at a slowing rate). The difference between training and testing accuracy shows overfitting of the model (i.e., the model can memorize what it has seen better than it can generalize the classification rules).
#
# We would like a model that *can* overfit (otherwise it might not be large enough to capture the complexity of the data domain), but doesn't. And then, it is only trained until *test accuracy* peaks.
#
# Could the model 100% overfit the data? The graph doesn't answer definitively yet, but training accuracy seems to be slowing, while training loss is still decreasing (with lots of room to improve – the loss axis does not start at zero).
#
# *Note:* The model contains Dropout layers to help prevent overfitting. What happens to training and testing accuracy when those layers are removed?
history_plot(history)
# Score trained model.
scores = model_transfer_cnn.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# The following prediction plot functions provide insight into aspects of model prediction.
def prediction_plot(model, test_data):
(x_test, y_test) = test_data
fig = plt.figure(figsize=(16,8))
correct = 0
total = 0
rSym = ''
for i in range(40):
plt.subplot(4, 10, i + 1)
plt.xticks([])
plt.yticks([])
idx = int(random.uniform(0, x_test.shape[0]))
result = model.predict(x_test[idx:idx+1])[0]
if y_test is not None:
rCorrect = True if cifar10_label(y_test[idx]) == cifar10_label(result) else False
rSym = '✔' if rCorrect else '✘'
correct += 1 if rCorrect else 0
total += 1
plt.title("%s %s" % (rSym, cifar10_label(result)))
plt.imshow(x_test[idx], cmap=plt.get_cmap('gray'))
plt.show()
if y_test is not None:
print("% 3.2f%% correct (%s/%s)" % (100.0 * float(correct) / float(total), correct, total))
def prediction_classes_plot(model, test_data):
(x_test, y_test) = test_data
fig = plt.figure(figsize=(16,8))
correct = 0
total = 0
rSym = ''
for i in range(40):
plt.subplot(4, 10, i + 1)
plt.xticks([])
plt.yticks([])
idx = int(random.uniform(0, x_test.shape[0]))
result = model.predict_classes(x_test[idx:idx+1])[0]
if y_test is not None:
rCorrect = True if cifar10_label(y_test[idx]) == cifar10_label(result) else False
rSym = '✔' if rCorrect else '✘'
correct += 1 if rCorrect else 0
total += 1
plt.title("%s %s" % (rSym, cifar10_label(result)))
plt.imshow(x_test[idx], cmap=plt.get_cmap('gray'))
plt.show()
if y_test is not None:
print("% 3.2f%% correct (%s/%s)" % (100.0 * float(correct) / float(total), correct, total))
def prediction_proba_plot(model, test_data):
(x_test, y_test) = test_data
fig = plt.figure(figsize=(15,15))
for i in range(10):
plt.subplot(10, 2, (2*i) + 1)
plt.xticks([])
plt.yticks([])
idx = int(random.uniform(0, x_test.shape[0]))
result = model.predict_proba(x_test[idx:idx+1])[0] * 100 # prob -> percent
if y_test is not None:
plt.title("%s" % cifar10_label(y_test[idx]))
plt.xlabel("#%s" % idx)
plt.imshow(x_test[idx], cmap=plt.get_cmap('gray'))
ax = plt.subplot(10, 2, (2*i) + 2)
plt.bar(np.arange(len(result)), result, label='%')
plt.xticks(range(0, len(result) + 1))
ax.set_xticklabels(cifar10_label_names)
plt.title("classifier probabilities")
plt.tight_layout()
plt.show()
# * *Grad-CAM: Visual Explanations from Deep Networks via Gradient-based Localization* by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, and <NAME> [arXiv (2016)](https://arxiv.org/abs/1610.02391)
# * https://jacobgil.github.io/deeplearning/class-activation-maps
# +
from tensorflow.keras import backend as K
def generate_activation_pattern(model, conv_layer_output, category_idx, image):
epsilon = 1e-10
activation_model = keras.models.Model([model.inputs], [conv_layer_output, model.output])
with tf.GradientTape() as gtape:
conv_output, prediction = activation_model(image)
category_output = prediction[:, category_idx]
grads = gtape.gradient(category_output, conv_output)
pooled_grads = K.mean(grads, axis=(0, 1, 2))
heatmap = tf.reduce_mean(tf.multiply(pooled_grads, conv_output), axis=-1) * -1.
heatmap = np.maximum(heatmap, 0)
heatmap /= np.max(heatmap) + epsilon
return(heatmap)
# -
def activation_plot(model, layer_name, image_data, image_index=None):
(layer_model, conv_layer) = get_model_layer(model, layer_name)
(x_imgs, y_cat) = image_data
if not image_index:
image_index = int(random.uniform(0, x_imgs.shape[0]))
image = x_imgs[image_index:image_index+1]
fig = plt.figure(figsize=(16,8))
plt.subplot(1, num_classes + 2, 1)
plt.xticks([])
plt.yticks([])
plt.title(cifar10_label(y_cat[image_index]))
plt.xlabel("#%s" % image_index)
plt.imshow(image.reshape(32, 32, 3))
result = model.predict(image)[0]
for i in range(num_classes):
activation = generate_activation_pattern(model, conv_layer.output, i, image)
activation = np.copy(activation)
plt.subplot(1, num_classes + 2, i + 2)
plt.xticks([])
plt.yticks([])
plt.title(cifar10_label(i))
plt.xlabel("(% 3.2f%%)" % (result[i] * 100.0))
plt.imshow(activation[0])
plt.show()
# This plot shows what the model thinks is the most likely class for each image.
prediction_classes_plot(model_transfer_cnn, (x_test, y_test))
# This plot shows the probabilities that the model assigns to each category class, and provides a sense of how confident the network is with its classifications.
prediction_proba_plot(model_transfer_cnn, (x_test, y_test))
# +
# TODO: Complete activation plot
#activation_plot(model_transfer_cnn, ('vgg16', 'block5_conv3'), (x_test, y_test), 1)
# -
# ### CNN Classifier Model
#
# Create a basic CNN (Convolutional Neural Network) based classifier from scratch.
#
# We have encountered Conv2D and MaxPooling2D layers previously, but here we see how they are declared. Conv2D layers specify the number of convolution kernels and their shape. MaxPooling2D layers specify the size of each pool (i.e., the scaling factors).
#
# Notice the total number of parameters (\~1.25 million) in this smaller network.
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Flatten, Dense, Activation, Dropout, Conv2D, MaxPooling2D
def create_cnn_classifier_model():
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same',
input_shape=x_train.shape[1:]))
model.add(Activation('relu'))
model.add(Conv2D(32, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(64, (3, 3), padding='same'))
model.add(Activation('relu'))
model.add(Conv2D(64, (3, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(512))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
return model
# -
model_simple_cnn = create_cnn_classifier_model()
model_simple_cnn.summary()
batch_size = 128 #32
epochs = 25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
# +
from tensorflow.keras.optimizers import RMSprop
model_simple_cnn.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
# -
# %%time
history = model_simple_cnn.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# The notable features of the history plot for this model are:
#
# * Training accuracy is ~10 percentage points better than the previous model,
# * test accuracy more closely tracks training accuracy, and
# * test accuracy shows more variability.
history_plot(history)
# Score trained model.
scores = model_simple_cnn.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
prediction_classes_plot(model_simple_cnn, (x_test, y_test))
prediction_proba_plot(model_simple_cnn, (x_test, y_test))
for n in [l.name for l in model_simple_cnn.layers if isinstance(l, keras.layers.Conv2D)][:4]:
visualize_conv_layer_weights(model_simple_cnn, n)
image_index = cifar10_image_plot()
for n in [l.name for l in model_simple_cnn.layers if isinstance(l, keras.layers.Conv2D)]:
visualize_conv_layer_output(model_simple_cnn, n, image_index)
# Interesting aspects of the convolutional layer response for our *model_simple_cnn* model:
#
# * There are fewer Conv2D layers in this simple model
# * Compared to the pre-trained VGG16 convolutional base network,
# * the latter levels are the first edge detection kernels, and
# * there are no layers with higher-level features.
for n in [l.name for l in model_simple_cnn.layers if isinstance(l, keras.layers.Conv2D)][:4]:
visualize_conv_layer_response(model_simple_cnn, n)
# This plot shows which pixels of the original image contributed the most 'confidence' to the classification categories.
#
# The technique is better applied to larger images where the object of interest might be anywhere inside the image.
n = [l.name for l in model_simple_cnn.layers if isinstance(l, keras.layers.Conv2D)][-1]
print(n)
for i in range(5):
activation_plot(model_simple_cnn, n, (x_test, y_test))
# ### Combined Models
#
# Keras supports a functional interface to take network architectures beyond simply sequential networks.
#
# The new layer types are Input and Concatenate; and, there is an explicit Model class.
#
# The Input layer is a special layer denoting sources of input from training batches.
#
# The Concatenate layer combines multiple inputs (along an axis with the same size) and creates a larger layer incorporating all the input values.
#
# Model construction is also different. Instead of using a `Sequential` model, and `add`ing layers to it:
#
# * An explicit Input layer is created,
# * we pass inputs into the layers explicity,
# * the output from a layer become input for arbitrary other layers, and finally,
# * A Model object is created with the source Input layer as inputs and outputs from the final layer.
#
# We'll demonstrate by creating a new network which combines the two CNN classifier networks we created previously.
#
# *Note:* Network models provided as an argument are changed to be non-trainable (the assumption is that they were already trained).
# +
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Input, Concatenate, Flatten, Dense, Activation, Dropout
from tensorflow.keras.optimizers import RMSprop
def create_combined_classifier_model(trained_model1=None, trained_model2=None):
if trained_model1:
network1 = trained_model1
network1.trainable = False
else:
network1 = create_cnnbase_classifier_model()
if trained_model2:
network2 = trained_model2
network2.trainable = False
else:
network2 = create_cnn_classifier_model()
inputs = Input(shape=(32,32,3), name='cifar10_image')
c1 = network1(inputs)
c2 = network2(inputs)
c = Concatenate()([c1, c2])
x = Dense(512)(c)
x = Activation('relu')(x)
x = Dropout(0.5)(x)
x = Dense(num_classes)(x)
outputs = Activation('softmax')(x)
model = Model(inputs=inputs, outputs=outputs, name='combined_cnn_classifier')
return model
# -
# #### Combining Pre-Trained Models
#
# This version of the combined classifier uses both of the trained networks we created previously.
#
# Notice the trainable parameters (~16,000) is very small. How will this affect training?
model_combined = create_combined_classifier_model(model_transfer_cnn, model_simple_cnn)
model_combined.summary()
# This plot shows a graph representation of the layer connections. Notice how a single input feeds the previously created Sequential networks, their output is combine via Concatenate, and then a classifier network is added on top.
keras.utils.plot_model(model_combined)
# Reduce number of `epochs` because this network is mostly trained (execpt for the final classifier), and there are few trainable parameters.
# +
batch_size = 128 #32
epochs = 5 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_combined.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
# -
history = model_combined.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# It looks like everything we needed to learn was learned in a single epoch.
history_plot(history)
# Here is an interesting, possibly counter-intuitive, result: combining two weaker networks can create a stronger one.
#
# The reason is that the weakness in one model, might be a strength in the other model (each has 'knowledge' that the other doesn't); we just need a layer to discriminate when to trust each model. At a larger scale (of layers and models) is what is happening at the lower level of the neurons themselves.
# Score trained model.
scores = model_combined.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# NOTE: Sequential Model provides `predict_classes` or `predict_proba`
# Functional API Model does not; because it may have multiple outputs
# Using simple `predict` plot instead
prediction_plot(model_combined, (x_test, y_test))
# The combine model improves accuracy by 2%, and takes 1/5<sup>th</sup> of the time to train.
#
# #### Training Combining Models
#
# This version of the combined classifier uses both network architectures seen previously; except, in this version, the models need to be trained from scratch. The following cells repeat the previous experiments with this combined classifier.
#
# *Spoiler:* The combined network doesn't perform any better than the partially trained one did, but takes much longer to train (more epochs).
#
# **Note:** Skip this step during the tutorial, it will cause unecessary delay.
# +
batch_size = 128 #32
epochs = 25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_combined = create_combined_classifier_model()
model_combined.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
history = model_combined.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# -
history_plot(history)
# Score trained model.
scores = model_combined.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# ### Skip Connections
#
# From previous comparisons of the `visualize_conv_layer_response` plots of the two basic CNN models, it becomes apparent that the pre-trained VGG16 network contains more complex *knowledge* about images: there were more convolutional layers with a greater variety of patterns and features they could represent.
#
# In the previous cnnbase_classifier model `model_transfer_cnn`, only the last Conv2D layer fed directly to the classifier, and the feature information contained in the middle layers wasn't directly available to the classifier.
#
# Skip Connections are a way to bring lower level feature encodings to higher levels of the network directly. They are also useful during training very deep networks to deal with the problem of *vanishing gradients*.
#
# In the following example, the original CNN base of the pre-trained VGG16 model is decomposed into layered groups, and a new network created that feeds these intermediate layers to the top of the network, where they are concatenated together to perform the final classification.
#
# * https://towardsdatascience.com/understanding-and-coding-a-resnet-in-keras-446d7ff84d33
# * https://arxiv.org/abs/1608.04117
# +
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Input, Concatenate, Flatten, Dense, Activation, Dropout
from tensorflow.keras.applications import VGG16
from tensorflow.keras.optimizers import RMSprop
def create_cnnbase_skipconnected_classifier_model(conv_base=None):
if not conv_base:
conv_base = VGG16(weights='imagenet', include_top=False, input_shape=(32, 32, 3))
conv_base.trainable = False
# Split conv_base into groups of CNN layers topped by a MaxPooling2D layer
cb_idxs = [i for (i,l) in enumerate(conv_base.layers) if isinstance(l, keras.layers.MaxPooling2D)]
all_idxs = [-1] + cb_idxs
idx_pairs = [l for l in zip(all_idxs, cb_idxs)]
cb_layers = [conv_base.layers[i+1:j+1] for (i,j) in idx_pairs]
# Dense Pre-Classifier Layers creation function - used repeatedly at multiple network locations
def dense_classes(l):
x = Dense(512)(l)
x = Activation('relu')(x)
x = Dropout(0.5)(x)
x = Dense(num_classes)(x)
return x
inputs = Input(shape=(32,32,3), name='cifar10_image')
# Join split groups into a sequence, but keep track of their outputs to create skip connections
skips = []
inz = inputs
for lz in cb_layers:
m = Sequential()
m.trainable = False
for ls in lz:
m.add(ls)
# inz is the output of model m, but the input for next layer group
inz = m(inz)
skips += [inz]
# Flatten all outputs (which had different dimensions) to Concatenate them on a common axis
flats = [dense_classes(Flatten()(l)) for l in skips]
c = Concatenate()(flats)
x = dense_classes(c)
outputs = Activation('softmax')(x)
model = Model(inputs=inputs, outputs=outputs)
return model
# -
model_skipconnected = create_cnnbase_skipconnected_classifier_model(conv_base)
model_skipconnected.summary()
keras.utils.plot_model(model_skipconnected)
# +
batch_size = 128 #32
epochs = 25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_skipconnected.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
history = model_skipconnected.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# -
history_plot(history)
# A significant improvement over the first pre-trained model.
# Score trained model.
scores = model_skipconnected.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# Using simple `predict` plot because model uses Functional API
prediction_plot(model_skipconnected, (x_test, y_test))
# ### Data Agumentation
#
# Data augmentation is a technique to expand the set of available training data and can significantly improve the performance of image processing networks.
#
# **Note:** Training examples in this section may take significant time. The approach does not improve accuracy results on this simple dataset, but is included here for illustration of the technique.
# +
from tensorflow.keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
featurewise_center=False, # set input mean to 0 over the dataset
samplewise_center=False, # set each sample mean to 0
featurewise_std_normalization=False, # divide inputs by std of the dataset
samplewise_std_normalization=False, # divide each input by its std
zca_whitening=False, # apply ZCA whitening
zca_epsilon=1e-06, # epsilon for ZCA whitening
rotation_range=10, # randomly rotate images in the range (degrees, 0 to 180)
# randomly shift images horizontally (fraction of total width)
width_shift_range=0.1,
# randomly shift images vertically (fraction of total height)
height_shift_range=0.1,
shear_range=0.1, # set range for random shear
zoom_range=0.1, # set range for random zoom
channel_shift_range=0.0, # set range for random channel shifts
# set mode for filling points outside the input boundaries
fill_mode='nearest',
cval=0.0, # value used for fill_mode = "constant"
horizontal_flip=True, # randomly flip images
vertical_flip=False, # randomly flip images
# set rescaling factor (applied before any other transformation)
rescale=None,
# set function that will be applied on each input
preprocessing_function=None,
# image data format, either "channels_first" or "channels_last"
data_format=None,
# fraction of images reserved for validation (strictly between 0 and 1)
validation_split=0.0
)
# Compute quantities required for feature-wise normalization
# (std, mean, and principal components if ZCA whitening is applied).
datagen.fit(x_train)
# +
exampledata = datagen.flow(x_train, y_train, batch_size=batch_size)
cifar10_imageset_plot((exampledata[0][0], exampledata[0][1]))
# -
# #### CNN Base + Classifier Model Agumented
# +
batch_size = 128 #32
epochs = 12 #25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_augmented = create_cnnbase_classifier_model(conv_base)
model_augmented.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
history = model_augmented.fit(datagen.flow(x_train, y_train, batch_size=batch_size),
validation_data=(x_test, y_test),
epochs=epochs,
shuffle=True,
use_multiprocessing=True, workers=4
)
# -
history_plot(history)
# Score trained model.
scores = model_augmented.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# #### CNN Classifier Model Augmented
# +
batch_size = 128 #32
epochs = 12 #25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_augmented = create_cnn_classifier_model()
model_augmented.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
history = model_augmented.fit(datagen.flow(x_train, y_train, batch_size=batch_size),
validation_data=(x_test, y_test),
epochs=epochs,
shuffle=True,
use_multiprocessing=True, workers=4
)
# -
history_plot(history)
# Score trained model.
scores = model_augmented.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# #### CNN Base + Skip Connected Classifier Model Agumented
# +
batch_size = 128 #32
epochs = 12 #25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_augmented = create_cnnbase_skipconnected_classifier_model(conv_base)
model_augmented.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
history = model_augmented.fit(datagen.flow(x_train, y_train, batch_size=batch_size),
validation_data=(x_test, y_test),
epochs=epochs,
shuffle=True,
use_multiprocessing=True, workers=4
)
# -
history_plot(history)
# Score trained model.
scores = model_augmented.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# ### Mixed Precision
#
# **TODO:** Fix performance issues
#
# **Note:** Mixed Precision is still experimental...
#
# * https://www.tensorflow.org/guide/keras/mixed_precision
# * https://www.tensorflow.org/api_docs/python/tf/keras/mixed_precision/experimental/Policy
# * https://developer.nvidia.com/automatic-mixed-precision
# * https://docs.nvidia.com/deeplearning/sdk/mixed-precision-training/index.html
#
# ```python
# opt = tf.train.experimental.enable_mixed_precision_graph_rewrite(opt)
# ```
# +
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Flatten, Dense, Activation, Dropout, Conv2D, MaxPooling2D
policy = tf.keras.mixed_precision.experimental.Policy('mixed_float16')
#tf.keras.mixed_precision.experimental.set_policy('mixed_float16')
def create_mixed_precision_cnn_classifier_model():
model = Sequential()
model.add(Conv2D(32, (3, 3), padding='same',
input_shape=x_train.shape[1:],
dtype=policy))
model.add(Activation('relu', dtype=policy))
model.add(Conv2D(32, (3, 3), dtype=policy))
model.add(Activation('relu', dtype=policy))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25, dtype=policy))
model.add(Conv2D(64, (3, 3), padding='same', dtype=policy))
model.add(Activation('relu', dtype=policy))
model.add(Conv2D(64, (3, 3), dtype=policy))
model.add(Activation('relu', dtype=policy))
model.add(MaxPooling2D(pool_size=(2, 2), dtype=policy))
model.add(Dropout(0.25, dtype=policy))
model.add(Flatten(dtype=policy))
# Dense layers use global policy of 'mixed_float16';
# does computations in float16, keeps variables in float32.
model.add(Dense(512, dtype=policy))
model.add(Activation('relu', dtype=policy))
model.add(Dropout(0.5, dtype=policy))
model.add(Dense(num_classes, dtype=policy))
# Softmax should be done in float32 for numeric stability. We pass
# dtype='float32' to use float32 instead of the global policy.
model.add(Activation('softmax', dtype='float32'))
return model
# +
from tensorflow.keras.optimizers import RMSprop
model_mixedprecision_cnn = create_mixed_precision_cnn_classifier_model()
model_mixedprecision_cnn.summary()
batch_size = 128 #32
epochs = 25 #100
learning_rate = 1e-3 #1e-4
decay = 1e-6
model_mixedprecision_cnn.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay),
metrics=['accuracy'])
# -
# %%time
history = model_mixedprecision_cnn.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True)
# Score trained model.
scores = model_mixedprecision_cnn.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# Reset Policy
tf.keras.mixed_precision.experimental.set_policy('float32')
# ### Multi-GPU Example
#
# Using multiple GPUs on a single node is a simple way to speed up deep learning. Keras / TensorFlow support this with a small modification to code.
#
# First, determine if multiple GPUs are available:
physical_devices = tf.config.experimental.list_physical_devices('GPU')
device_count = len(physical_devices)
print("GPU count:", device_count)
print("GPU devices:", physical_devices)
# When scaling to `n` GPUs, there is `n *` the available GPU memory, so we can increase the batch_size by `n`. A larger batch size means that there is more data evaluated by the batch step, which creates a more accurate and representative loss gradient – so we can take a larger corrective step by multiply the learning_rate by `n`. Because we are learning `n *` more each epoch, we only need `1/n`<sup>th</sup> the number of training epochs.
#
# There are additional subtleties and mitigating strategies to be aware of when scaling batch sizes larger. Some of these are discussed in [Deep Learning at scale: Accurate, Large Mini batch SGD](https://towardsdatascience.com/deep-learning-at-scale-accurate-large-mini-batch-sgd-8207d54bfe02).
# +
# Multi-GPU Example
assert device_count >= 2, "Two or more GPUs required to demonstrate multi-gpu functionality"
from tensorflow.keras.optimizers import Adam, RMSprop
from tensorflow.keras.callbacks import LearningRateScheduler, ReduceLROnPlateau
batch_size = device_count * 128 #32
epochs = 25 // device_count + 1 #100
learning_rate = device_count * 1e-3 #1e-4
decay = 1e-6
def lr_schedule(epoch):
initial_lr = device_count * 1e-3
warmup_epochs = 5
warmup_lr = (epoch + 1) * initial_lr / warmup_epochs
return warmup_lr if epoch <= warmup_epochs else initial_lr
lr_scheduler = LearningRateScheduler(lr_schedule, verbose=1)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1), cooldown=0, patience=5, min_lr=0.5e-6)
callbacks = [lr_reducer, lr_scheduler]
strategy = tf.distribute.MirroredStrategy()
with strategy.scope():
model_multigpu = create_cnnbase_classifier_model()
model_multigpu.compile(loss='categorical_crossentropy',
optimizer=RMSprop(learning_rate=learning_rate, decay=decay, momentum=0.5),
# TODO: Explore Adam without lr_scheduling
#optimizer=Adam(learning_rate=learning_rate),
metrics=['accuracy'])
history = model_multigpu.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
validation_data=(x_test, y_test),
shuffle=True,
callbacks=callbacks,
use_multiprocessing=True, workers=4
)
# -
history_plot(history)
# Score trained model.
scores = model_multigpu.evaluate(x_test, y_test, verbose=0)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
# ## Speaker Bios
#
# <NAME> is a Staff Scientist in the Visualization Core Lab at KAUST (King Abdullah University of Science and Technology) specializing in HPC workflow solutions for deep learning, image processing, and scientific visualization.
#
# <NAME> is a Computational Scientist in the Supercomputing Core Lab at KAUST (King Abdullah University of Science and Technology) specializing in large scale HPC applications and GPGPU support for users on Ibex (cluster) and Shaheen (supercomputer). Mohsin holds a PhD in Computational Bioengineering, and a Post Doc, from University of Canterbury, New Zealand.
# ## References
#
# * https://qz.com/1034972/the-data-that-changed-the-direction-of-ai-research-and-possibly-the-world/
# * https://www.cs.toronto.edu/~kriz/cifar.html
# * http://yann.lecun.com/exdb/mnist/index.html
# * https://towardsdatascience.com/transfer-learning-from-pre-trained-models-f2393f124751 <br/>
# https://towardsdatascience.com/keras-transfer-learning-for-beginners-6c9b8b7143e <br/>
# https://machinelearningmastery.com/how-to-improve-performance-with-transfer-learning-for-deep-learning-neural-networks/
# * https://towardsdatascience.com/deep-learning-at-scale-accurate-large-mini-batch-sgd-8207d54bfe02
# * https://arxiv.org/abs/1409.1556 <br/>
# https://arxiv.org/abs/1610.02391
# * https://www.kaggle.com/c/digit-recognizer
# * https://jupyter-notebook.readthedocs.io/en/stable/
# * https://github.com/kaust-vislab/handson-ml2
# * https://keras.io/examples/cifar10_cnn/ <br/>
# https://keras.io/examples/cifar10_resnet/
| notebooks/keras-transfer-learning-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: sbi
# language: python
# name: sbi
# ---
# # calculating the bispectrum for survey geometry
import os, time
import numpy as np
from simbig import halos as Halos
from simbig import galaxies as Galaxies
from simbig import forwardmodel as FM
from simbig import obs as CosmoObs
# --- plotting ---
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
# ## Read in `Quijote` halo catalog and populate with galaxies
# I'm using `i=1118`th cosmology in the LHC because that's the closest to the cosmology used in Manera+(2015)
# +
# read in halo catalog
halos = Halos.Quijote_LHC_HR(1118, z=0.5)
print('Om, Ob, h, ns, s8:')
print(Halos.Quijote_LHC_cosmo(1118))
# get LOWZ HOD parameters
theta_hod = Galaxies.thetahod_lowz_sgc()
# populate halos
hod = Galaxies.hodGalaxies(halos, theta_hod, seed=0)
# -
# ## Forward model survey geometry and generate matching randoms
gals = FM.BOSS(hod, sample='lowz-south', seed=0, veto=False, fiber_collision=False, silent=False)
rand = FM.BOSS_randoms(gals, sample='lowz-south', veto=False) # random without veto mask
# ## Calculate bispectrum using `simbig.obs.B0k_survey`
t0 = time.time()
b123 = CosmoObs.B0k_survey(gals, rand,
P0=1e4,
Ngrid=360,
Lbox=1400,
silent=False)
print('B0 take %f sec' % ((time.time() - t0)))
klim = (b123[0] < 0.5) & (b123[1] < 0.5) & (b123[2] < 0.5)
fig = plt.figure(figsize=(20,5))
sub = fig.add_subplot(111)
sub.plot(range(np.sum(klim)), b123[3][klim])
sub.set_xlabel('triangle configurations', fontsize=25)
sub.set_xlim(0, np.sum(klim))
sub.set_ylabel('$B_0(k_1, k_2, k_3)$', fontsize=25)
sub.set_yscale('log')
| nb/demo_bk_survey.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %run ../Python_files/util.py
road_seg_inr_capac = zload('../temp_files/road_seg_inr_capac_ext.pkz')
raw_data_folder = '/home/jzh/INRIX/All_INRIX_2012/'
import csv
import gzip
import glob
for input_file in glob.glob(raw_data_folder + '*.csv.gz'):
with gzip.open(input_file, 'rb') as inp, \
open(input_file + '.filtered.csv', 'wb') as out:
writer = csv.writer(out)
for row in csv.reader(inp):
if row[0] in road_seg_inr_capac.tmc:
writer.writerow(row)
# -
| 01_INRIX_data_preprocessing_ifac17/INRIX_data_preprocessing_03_filter_speed_data_ext.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 6 : Visualization
# # Matplotlib version and modules
# +
import pkgutil as pu
import numpy as np
import matplotlib as mpl
import scipy as sp
import pandas as pd
import pydoc
print("Matplotlib version", mpl.__version__)
def clean(astr):
s = astr
# remove multiple spaces
s = ' '.join(s.split())
s = s.replace('=','')
return s
def print_desc(prefix, pkg_path):
for pkg in pu.iter_modules(path=pkg_path):
name = prefix + "." + pkg[1]
if pkg[2] == True:
try:
docstr = pydoc.plain(pydoc.render_doc(name))
docstr = clean(docstr)
start = docstr.find("DESCRIPTION")
docstr = docstr[start: start + 140]
print(name, docstr)
except:
continue
print("\n")
print_desc("matplotlib", mpl.__path__)
# -
# # Basic matplotlib plot
# +
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
x = np.linspace(0, 20)
plt.plot(x, .5 + x)
plt.plot(x, 1 + 2 * x, '--')
plt.show()
# -
# # Logarithmic plot
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
years = df.index.values
counts = df['trans_count'].values
poly = np.polyfit(years, np.log(counts), deg=1)
print("Poly", poly)
plt.semilogy(years, counts, 'o')
plt.semilogy(years, np.exp(np.polyval(poly, years)))
plt.show()
# -
# ## Scatter Plots
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True, right_index=True)
df = df.replace(np.nan, 0)
years = df.index.values
counts = df['trans_count'].values
gpu_counts = df['gpu_trans_count'].values
cnt_log = np.log(counts)
plt.scatter(years, cnt_log, c= 200 * years, s=20 + 200 * gpu_counts/gpu_counts.max(), alpha=0.5)
plt.show()
# -
# # Legends and annotaions
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True, right_index=True)
df = df.replace(np.nan, 0)
years = df.index.values
counts = df['trans_count'].values
gpu_counts = df['gpu_trans_count'].values
poly = np.polyfit(years, np.log(counts), deg=1)
plt.plot(years, np.polyval(poly, years), label='Fit')
gpu_start = gpu.index.values.min()
y_ann = np.log(df.at[gpu_start, 'trans_count'])
ann_str = "First GPU\n %d" % gpu_start
plt.annotate(ann_str, xy=(gpu_start, y_ann), arrowprops=dict(arrowstyle="->"), xytext=(-30, +70), textcoords='offset points')
cnt_log = np.log(counts)
plt.scatter(years, cnt_log, c= 200 * years, s=20 + 200 * gpu_counts/gpu_counts.max(), alpha=0.5, label="Scatter Plot")
plt.legend(loc='upper left')
plt.grid()
plt.xlabel("Year")
plt.ylabel("Log Transistor Counts", fontsize=16)
plt.title("Moore's Law & Transistor Counts")
plt.show()
# -
# # Three-dimensional plots
# +
from mpl_toolkits.mplot3d.axes3d import Axes3D
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True, right_index=True)
df = df.replace(np.nan, 0)
fig = plt.figure()
ax = Axes3D(fig)
X = df.index.values
Y = np.log(df['trans_count'].values)
Y = np.where(df['trans_count'].values>0, np.log(df['trans_count'].values), 0)
X, Y = np.meshgrid(X, Y)
#Z = np.log(df['gpu_trans_count'].values)
Z = np.where(df['gpu_trans_count'].values>0, np.log(df['gpu_trans_count'].values), 0)
print(Y.shape, Z.shape)
ax.plot_surface(X, Y, Z)
ax.set_xlabel('Year')
ax.set_ylabel('Log CPU transistor counts')
ax.set_zlabel('Log GPU transistor counts')
ax.set_title("Moore's Law & Transistor Counts")
plt.show()
# -
# # plotting in pandas
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True,
right_index=True)
df = df.replace(np.nan, 0)
df.plot()
df.plot(logy=True)
df[df['gpu_trans_count'] > 0].plot(kind='scatter',
x='trans_count', y='gpu_trans_count', loglog=True)
plt.show()
# -
# # Lag plots
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pandas.tools.plotting import lag_plot
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True, right_index=True)
df = df.replace(np.nan, 0)
lag_plot(np.log(df['trans_count']))
plt.show()
# -
# # autocorrelation plots
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pandas.tools.plotting import autocorrelation_plot
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True, right_index=True)
df = df.replace(np.nan, 0)
autocorrelation_plot(np.log(df['trans_count']))
plt.show()
# -
# # plot.ly
# +
import plotly.plotly as py
from plotly.graph_objs import *
import numpy as np
import pandas as pd
df = pd.read_csv('transcount.csv')
df = df.groupby('year').aggregate(np.mean)
gpu = pd.read_csv('gpu_transcount.csv')
gpu = gpu.groupby('year').aggregate(np.mean)
df = pd.merge(df, gpu, how='outer', left_index=True, right_index=True)
df = df.replace(np.nan, 0)
# Change the user and api_key to your own username and api_key
py.sign_in('zhanghaogithub123', '<KEY>')
counts = np.log(df['trans_count'].values)
gpu_counts = np.log(df['gpu_trans_count'].values)
data = Data([Box(y=counts), Box(y=gpu_counts)])
plot_url = py.plot(data, filename='moore-law-scatter')
print(plot_url)
# -
| Chapter06/ch-06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Python - základy na prežitie
#
# __Rada: nedávajte medzery a diakritiku v názvoch NB. Tu, v textových bunkách, si môžete robiť, čo srdco ráči.__
# + [markdown] slideshow={"slide_type": "slide"}
# ### Premenné v Pythone a objekty.
# **Nie je treba deklarovať ich typ a aj práca s pamäťou (alokácia a uvoľňovanie) je automatická.**
#
# **Python je od prírody objektovo orientovaný, ale netreba to obkecávať.**
# + [markdown] slideshow={"slide_type": "slide"}
# ### Čísla, textové reťazce
# + slideshow={"slide_type": "fragment"}
# Toto je komentar a aj nizsie budu. Kvoli jednoduchosti nepouzivame diakritiku, hoci by sme mohli...
n = 1 # V premennej n bude cele cislo, jeho typ je trieda (class) int
print("Premenna n ma hodnotu",n, "jej typ je", type(n))
# + slideshow={"slide_type": "fragment"}
f = .7e13 # Do premennej f priradime desatinne cislo (float) - pozname podla desatinnej bodky
print("Premenna f ma hodnotu",f, "jej typ je", type(f))
# + slideshow={"slide_type": "slide"}
c = 'Ahoj, svet a "vesmír" ' # Textovy retazec (str), bud v uvodzovkach alebo v apostrofoch sa dava
# ak je v apostrofoch, vnutri mozeme beztrestne doslova pouzit uvodzovky apod...
print("Premenna c hovori, ze: ",c, "jej typ je", type(c))
# + slideshow={"slide_type": "fragment"}
# a este jeden zaujimavy typ
nic = None
print(nic)
print(type(nic))
# + slideshow={"slide_type": "slide"}
# v jednoduchosti je krasa
x = 2**3 # umocnovanie
y = -.9
x,y = y,x # vymenenie hodnot bez pomocnej prememnnej
print("x =",x, " y =",y)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Zoznamy, množiny, slovníky
# + slideshow={"slide_type": "fragment"}
# Z fleku - priklady "zlozenych" datovych typov:
L = [3,c,-7.5,'abc'] # Zoznam (list) - moze obsahovat LUBOVOLNE Python objekty. Nie je to "pole"!
# Zapisujeme ho do hranatych zatvoriek.
print("\nZoznam L sa vypise ako",L,"jeho typ je", type(L))
print("Zoznam L obsahuje ako prvok retazec c?", 'fff' in L) # veru obsahuje, True (pravda) je to
# + slideshow={"slide_type": "fragment"}
# Usporiadana n-tica
N = (L,33,'a')
print(N, type(N))
# + slideshow={"slide_type": "fragment"}
# Mnozina (set). Toto nema kazdy jazyk - a tak jednoducho.
M = {1,2,3,2,5,1}
print("Mnozina M sa vypise ako",M,"jej typ je", type(M)) #vsimnite si, ze kazdy prvok sa v nej nachadza iba raz
print("Prvok 3 je v mnozine M?", 3 in M) # True je booleovska premenna - pravda je
print("Prvok 0 je v mnozine M?", 0 in M) # tak toto pravda nie je
# + slideshow={"slide_type": "slide"}
# Nakoniec, slovnik (dict). Mnozina dvojic kluc - hodnota.
D = {'a':'and',1:'one',(1,2): 4.5} # kluce musia byt text. retazce, cisla, alebo usporiadane n-tice
print("Slovnik D sa vypise ako",D,"jeho typ je", type(D))
#Vyskusajte si aj prikazy D.keys() a D.values()
# -
del D['a']
D
# + [markdown] slideshow={"slide_type": "slide"}
# ## List comprehension
# Využíva sa pri vytváraní premennej typu list pomocou spracovania "väčšieho" zoznamu.
#
# Je to takáto konštrukcia
# ## `Z = [f(p) for p in X if V(p)]`
#
# kde `f(p)` je funkcia (vstavaná, alebo nami vytvorená),
#
# `X` je zoznam, ktorý spracúvame,
#
# `V(p)` je podmienka na prvky zoznamu `X`, po ktorom prechádzame
# -
list(range(11))
# + slideshow={"slide_type": "slide"}
Z = [p for p in range(0,11) if p%2 == 0]
Z2 = [p**2 for p in range(0,11) if p%2 == 0] # druha mocnina p v rozsahu od 0 po 9, ak je p parne
print(Z,Z2,sep='\n')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Cykly for a while
# __Veľký pozor - bloky kódu v Pythone sú určené odsadením.__
# + slideshow={"slide_type": "fragment"}
# Cyklus, jednoduchy. Bloky kodu v Pythone su urcene ODSADENIM, ziadne zatvorky a pod.
for p in L:
print(p + p, ",", 3 * p)
print("\nA toto už nie je v cykle") # \n je prechod na novy riadok, ako v C
# + slideshow={"slide_type": "slide"}
# Typ range
for p in range(1,11): # rozsah od 1 po 10 (11-1)
print(p)
print(list(range(0,10))) # pretypovanie, co by bolo bez neho?
# + [markdown] slideshow={"slide_type": "slide"}
# **Dvojbodka ja súčasťou syntaxe a používa sa v 3 prípadoch:**
# - **cykly**
# - **podmienky**
# - **definície funkcií**
# + slideshow={"slide_type": "slide"}
# Cyklus, pokial plati podmienka
k = 1
while (k < 7):
print("k = ",k)
k = k + 1 # mozete tiez ako v C: k += 1
print("Toto uz nie je v cykle, bo nie je odsadene...")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Definícia funkcie a jej volanie
# + slideshow={"slide_type": "fragment"}
# Jednoducha definicia funkcie a ukazka podmienok (vetvenie)
def sgn(x):
if x != 0:
if x > 0:
return 1
else: # namiesto else if mozeme pouzit skratene elif
return -1
return 0 # to nastane, ak x == 0 (porovnavanie, ako v C)
print("sgn(-5.4) = ",sgn(-5.4),", sgn(0) = ",sgn(0), ", sgn(1.7e10) = ", sgn(1.7e10))
# + slideshow={"slide_type": "slide"}
# Magic - ake mame premenne a ine objekty
# %whos
# + slideshow={"slide_type": "slide"}
# Skratena definicia funkcie
sqr = lambda x : x**2
print(type(sqr))
print(sqr(3))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Indexovanie premenných sekvenčného typu a dvojbodková notácia
# + slideshow={"slide_type": "fragment"}
# Indexuje sa pomocou hranatych zatvoriek. Indexy su od nuly, ako v C.
print(c,"\nVybrane: ",c[0],c[3],c[4])
print(L,"\nVybrane: ",L[1])
# + slideshow={"slide_type": "fragment"}
# Dvojbodkova notacia (slices), vseobecne od:do:krok
# ked chyba od, do berie sa cely rozsah. Ked chyba krok, berie sa 1.
print(c[:2]) # prve dva prvky
print(c[2:5]) # prvky s indexami 2,3,4
# Pre pripomenutie
print("L je zoznam: ",L)
# Dvojite indexovanie
print(L[1][:5])
# + slideshow={"slide_type": "slide"}
# Zaporne indexovanie - od konca
print(L[-1]) # posledny prvok L
print(c[-5:]) # poslednych 5 znakov c (je tam aj medzera, tu nevidno)
# + slideshow={"slide_type": "fragment"}
print(c[::2]) # kazde druhe pismeno
print(c.find('s')) # index prveho vyskytu hladaneho podretazca
print(c.count('s')) # pocet vyskytov hladaneho podretazca
print(c.lower()) # vsetky pismena zmeni na male
# + [markdown] slideshow={"slide_type": "slide"}
# ### Slovník - kľúče ako indexy
# + slideshow={"slide_type": "fragment"}
# Mnozina sa indexovat neda, nie je v nej ziadne poradie urcene. Iba ci do nej nieco patri, alebo nie.
# Slovnik sa indexuje klucmi namiesto indexov 0,1,...
print("povodny D", D) # pre pripomenutie
# print("D['a'] = " ,D['a'], "\nD[(1,2)] = ", D[(1,2)])
# dalsi kluc a hodnota sa jednoducho prida pomocou indexovania
D['Janko'] = 'Marienka'
print("zmeneny D", D)
| Uvod/Python_zaklady_na_prezitie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["meta", "draft"]
# # GPS tracks
# + [markdown] tags=["hide"]
# http://geopandas.org/gallery/plotting_basemap_background.html#adding-a-background-map-to-plots
# -
# https://ocefpaf.github.io/python4oceanographers/blog/2015/08/03/fiona_gpx/
# + tags=["hide"]
import pandas as pd
import geopandas as gpd
# -
# - **TODO**: put the following lists in a JSON dict and make it avaliable in a public Git repository (it can be usefull for other uses)
# - **TODO**: put the generated GeoJSON files in a public Git repository
df = gpd.read_file("communes-20181110.shp")
# !head test.gpx
# !head test.csv
# https://gis.stackexchange.com/questions/114066/handling-kml-csv-with-geopandas-drivererror-unsupported-driver-ucsv
df_tracks = pd.read_csv("test.csv", skiprows=3)
df_tracks.head()
df_tracks.columns
# +
from shapely.geometry import LineString
# https://shapely.readthedocs.io/en/stable/manual.html
positions = df_tracks.loc[:, ["Longitude (deg)", "Latitude (deg)"]]
positions
# -
LineString(positions.values)
# https://stackoverflow.com/questions/38961816/geopandas-set-crs-on-points
df_tracks = gpd.GeoDataFrame(geometry=[LineString(positions.values)], crs = {'init' :'epsg:4326'})
df_tracks.head()
df_tracks.plot()
# +
communes_list = [
"78160", # Chevreuse
"78575", # Saint-Rémy-lès-Chevreuse
]
df = df.loc[df.insee.isin(communes_list)]
# -
df
ax = df_tracks.plot(figsize=(10, 10), alpha=0.5, edgecolor='k')
ax = df.plot(ax=ax, alpha=0.5, edgecolor='k')
#df.plot(ax=ax)
# ## Convert the data to Web Mercator
df_tracks_wm = df_tracks.to_crs(epsg=3857)
df_wm = df.to_crs(epsg=3857)
df_tracks_wm
ax = df_tracks_wm.plot(figsize=(10, 10), alpha=0.5, edgecolor='k')
# ## Contextily helper function
# +
import contextily as ctx
def add_basemap(ax, zoom, url='http://tile.stamen.com/terrain/tileZ/tileX/tileY.png'):
xmin, xmax, ymin, ymax = ax.axis()
basemap, extent = ctx.bounds2img(xmin, ymin, xmax, ymax, zoom=zoom, url=url)
ax.imshow(basemap, extent=extent, interpolation='bilinear')
# restore original x/y limits
ax.axis((xmin, xmax, ymin, ymax))
# -
# ## Add background tiles to plot
# +
ax = df_tracks_wm.plot(figsize=(16, 16), alpha=0.5, edgecolor='k')
ax = df_wm.plot(ax=ax, alpha=0.5, edgecolor='k')
#add_basemap(ax, zoom=13, url=ctx.sources.ST_TONER_LITE)
add_basemap(ax, zoom=14)
ax.set_axis_off()
# -
# ## Save selected departments into a GeoJSON file
import fiona
fiona.supported_drivers
# !rm tracks.geojson
df_tracks.to_file("tracks.geojson", driver="GeoJSON")
# !ls -lh tracks.geojson
df = gpd.read_file("tracks.geojson")
df
ax = df.plot(figsize=(10, 10), alpha=0.5, edgecolor='k')
| nb_dev_python/python_geopandas_gps_tracks_lines_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 03 - Conjoint analysis
# The exercises in this notebook are inspired by [Traditional Conjoint Analysis with Excel](https://sawtoothsoftware.com/resources/technical-papers/analysis-of-traditional-conjoint-using-excel-an-introductory-example). We use the same data and run the same analysis, but we perform the analysis in Python - which is more extensible.
#
# ## Example Problem
# Consider a product range where each product has three attributes $(\text{Brand}, \text{Colour}, \text{Price})$ which can each take on the following values:
#
# | Brand | Colour | Price |
# | -------- | -------- | -------- |
# | A | Red | £50 |
# | B | Blue | £100 |
# | C | | £150 |
#
# For example, a particular product instance might have attribute-values $(\text{B}, \text{Red}, \text{£150})$. In total, there are $18$ possible products that can be created from these attribute values.
#
# $$\text{3 brands} × \text{2 colors} × \text{3 prices} = \text{18 products}$$
#
# Assume that each of these products are tested is a trial where partipants rate each of the products on a scale from $0$ to $10$, where $10$ represents the highest degree of preference.
# ## Example data
# Let's assume we have data from one participant, who rated every product. Run the code below to load their data into a `DataFrame`.
# +
import pandas as pd
columns = ['Product', 'Brand', 'Colour', 'Price', 'Preference']
data = [
[ 1, 'A', 'Red', '£50', 5],
[ 2, 'A', 'Red', '£100', 5],
[ 3, 'A', 'Red', '£150', 0],
[ 4, 'A', 'Blue', '£50', 8],
[ 5, 'A', 'Blue', '£100', 5],
[ 6, 'A', 'Blue', '£150', 2],
[ 7, 'B', 'Red', '£50', 7],
[ 8, 'B', 'Red', '£100', 5],
[ 9, 'B', 'Red', '£150', 3],
[10, 'B', 'Blue', '£50', 9],
[11, 'B', 'Blue', '£100', 6],
[12, 'B', 'Blue', '£150', 5],
[13, 'C', 'Red', '£50', 10],
[14, 'C', 'Red', '£100', 7],
[15, 'C', 'Red', '£150', 5],
[16, 'C', 'Blue', '£50', 9],
[17, 'C', 'Blue', '£100', 7],
[18, 'C', 'Blue', '£150', 6]
]
df_responses = pd.DataFrame(data=data,columns=columns)
df_responses.head(20)
# -
# ## Coding attribute levels
# To perform a conjoint analysis on the above data, we first need to format (/code) the attribute values/levels. We can do this using the `get_dummies` method.
attributes = ['Brand', 'Colour', 'Price']
df_dummies = pd.get_dummies(df_responses, columns=attributes)
df_dummies.head(20)
# ## Resolving linear dependencies
# The problem with the above coding, is the linear dependency between input features. To fix this problem, we can use the `drop_first=True` argument to produce a better set of input features.
df_coded = pd.get_dummies(df_responses, columns=attributes, drop_first=True)
df_coded.head(20)
# ## Multiple regression analysis
# We can now use the `sklearn` package to perform regression analysis.
# +
from sklearn import linear_model
regr = linear_model.LinearRegression()
dependent_var = 'Preference'
independent_vars = ['Brand_B', 'Brand_C', 'Colour_Red', 'Price_£150', 'Price_£50']
y = df_coded[dependent_var]
X = df_coded[independent_vars]
regr.fit(X, y)
print(regr.coef_)
# -
# These regression coefficients, show the effect of each attribute level relative to the one missing. To see this, we can print the final results in a more human-readible format.
# +
for attribute in attributes:
attribute_levels = [level for level in df_dummies.columns if level.startswith(attribute)]
print(attribute)
for level in attribute_levels:
value = regr.coef_[independent_vars.index(level)] if (level in independent_vars) else 0.00
level = level.split('_')[-1]
print(f' {level}={value:.2f}')
# -
# Look at results compared to the original [article](https://sawtoothsoftware.com/resources/technical-papers/analysis-of-traditional-conjoint-using-excel-an-introductory-example). Why are they slightly different?
#
# Also, read the original [article](https://sawtoothsoftware.com/resources/technical-papers/analysis-of-traditional-conjoint-using-excel-an-introductory-example) for more details.
| solutions/03 - Conjoint analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
import keras
from keras.layers import Dense, Input, Conv2D, BatchNormalization, MaxPool2D, Flatten
from keras.models import Model
from keras.utils import to_categorical, multi_gpu_model
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
train = pd.read_csv('train.csv')
train.head()
labels = train.label.to_numpy().reshape((42000,1))
images = train.drop('label', axis=1).to_numpy().reshape((42000,28,28,1))
images = images / 255
#plt.imshow(images[0])
labelsEncoding = to_categorical(labels)
def convModel():
inputs = Input(shape=(28,28,1))
X = Conv2D(32, 3, padding='same')(inputs)
X = MaxPool2D(pool_size=(2,2))(X)
X = Conv2D(64, 3, padding='same')(X)
X = MaxPool2D(pool_size=(2,2))(X)
X = Flatten()(X)
X = Dense(32, kernel_regularizer='l2', activation='relu')(X)
X = Dense(32, kernel_regularizer='l2', activation='relu')(X)
outputs = Dense(10, kernel_regularizer='l2', activation='softmax')(X)
model = Model(inputs = inputs, outputs = outputs)
return model
model = convModel()
model = multi_gpu_model(model, gpus=2)
model.compile(optimizer='adam', loss = 'categorical_crossentropy', metrics=['accuracy'])
model.fit(images, labelsEncoding, batch_size=128, epochs=10)
model.save_weights('conv_nobatchnorm.h5')
test = pd.read_csv('test.csv')
test.head()
testimages = test.to_numpy().reshape((28000,28,28,1))
testimages = testimages/255
predictions = model.predict(testimages, batch_size=128, verbose=1)
predictionLabels = pd.DataFrame({'Label':predictions.argmax(axis=-1).reshape((28000,1))[:,0]})
ImageId = pd.DataFrame({'ImageId': [i+1 for i in range(28000)]})
submission = pd.concat([ImageId, predictionLabels], axis = 1)
submission.to_csv('submission_conv_nobatchnorm.csv', index=False)
def convModelBatchNorm():
inputs = Input(shape=(28,28,1))
X = Conv2D(32, 3, padding='same', activation='relu')(inputs)
X = BatchNormalization()(X)
X = MaxPool2D(pool_size=(2,2))(X)
X = Conv2D(64, 3, padding='same', activation='relu')(X)
X = BatchNormalization()(X)
X = MaxPool2D(pool_size=(2,2))(X)
X = Flatten()(X)
X = Dense(128, kernel_regularizer='l2', activation='relu')(X)
X = Dense(128, kernel_regularizer='l2', activation='relu')(X)
outputs = Dense(10, kernel_regularizer='l2', activation='softmax')(X)
model = Model(inputs = inputs, outputs = outputs)
return model
model = convModelBatchNorm()
model = multi_gpu_model(model, gpus=2)
model.compile(optimizer='adam', loss = 'categorical_crossentropy', metrics=['accuracy'])
model.fit(images, labelsEncoding, batch_size=128, epochs=10)
model.save_weights('conv_batchnorm_relu.h5')
predictions_batchnorm = model.predict(testimages, batch_size=128, verbose=1)
predictionLabels_batchnorm = pd.DataFrame({'Label':predictions_batchnorm.argmax(axis=-1).reshape((28000,1))[:,0]})
ImageId_batchnorm = pd.DataFrame({'ImageId': [i+1 for i in range(28000)]})
submission_batchnorm = pd.concat([ImageId_batchnorm, predictionLabels_batchnorm], axis = 1)
submission.to_csv('submission_conv_batchnorm_relu2.csv', index=False)
predictionLabels.equals(predictionLabels_batchnorm)
| MNIST_conv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Example #1
#
# Author : <NAME> (<EMAIL>)
#
# We will calculate cross power, coherence, and cross phase between two data sets.
# #### Step #0
# Import modules.
# %matplotlib inline
import sys, os
sys.path.append(os.pardir)
from fluctana import *
# #### Step #1
# Make two data sets to be compared.
#
# One data set has four channels ('ECEI_G1401', 'ECEI_G1402', 'ECEI_G1403', 'ECEI_G1404').
#
# The other has four channels ('ECEI_G1501', 'ECEI_G1502', 'ECEI_G1503', 'ECEI_G1504').
A = FluctAna()
A.add_data(KstarEcei(shot=19348, clist=['ECEI_G1401-1404']), trange=[8.2, 8.25])
A.add_data(KstarEcei(shot=19348, clist=['ECEI_G1501-1504']), trange=[8.2, 8.25])
# #### Step #2
# Do fft transform with proper option parameters
A.fftbins(nfft=512,window='hann',overlap=0.5,detrend=1)
# The message means that
#
# data set number 0 has three channel and each data is divided into 95 sections (or bins) with overlap ratio = 0.5. Each section (bin) has 512 data points.
#
# data set number 1 has three channels and each data is divided into 95 sections (or bins) with overlap ratio = 0.5. Each section (bin) has 512 data points.
# #### Step #3
# Calculate cross power between two data sets.
#
# Set the reference data set (done) as data set number 0. If the reference data set has only one channel, calculations will be done between the one reference channel and all channels in dtwo. Otherwise, done can have same number of channels with dtwo, and calculations between pairs of same channel numbers will be done.
#
# Set dtwo as data set number 1
A.cross_power(done=0,dtwo=1)
# #### Step #4
# Plot the result.
#
# The result is saved for data set number 1 (dtwo)
# Plot the result of the channel number 0, 1, 2 (in dtwo=1) in multiple windows
A.mplot(dnum=1,cnum=[0,1,2,3],type='val')
# Plot the result of the channel number 0, 1, 2 (in dtwo=1) in single window (overlapping)
A.oplot(dnum=1,cnum=[0,1,2,3],type='val')
# Plot the rms amplitude image (3 points in this example) by adding up the cross power over a frequency range
#
# snum : sample channel number
#
# frange : integration frequency range
#
# vlimits : color scale limits
A.cplot(dnum=1,snum=2,frange=[0,100],vlimits=[0,0.05])
# You can calculate and plot coherence similarily.
A.coherence(done=0,dtwo=1)
A.mplot(dnum=1,cnum=[0,1,2,3],type='val',ylimits=[0,1])
A.oplot(dnum=1,cnum=[0,1,2,3],type='val',ylimits=[0,1])
A.cplot(dnum=1,snum=2,frange=[0,100],vlimits=[0,0.3])
# You can calculate and plot cross phase similarily.
A.cross_phase(done=0,dtwo=1)
A.mplot(dnum=1,cnum=range(4),type='val')
A.oplot(dnum=1,cnum=range(4),type='val')
A.cplot(dnum=1,snum=2,frange=[0,50],vlimits=[-10,10])
A.Dlist[1].pdata
| examples/example1_19348.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from eigenwell.src.eigen_guide import *
from eigenwell.src.constants import *
from eigenwell.src.structure import *
import scipy.sparse.linalg as la
import os
L0 = 1e-6;
omega_p = 0.72*np.pi*1e15;
gamma = 5.5e12;
# ## Example of a Dispersive Eigensolver
# We're using a material which changes in dielectric with frequency $\epsilon(\omega)$
# !pwd
# !ls ../
# +
omega_cutoff = 0.83020*omega_p;
wvlen_cutoff = 2*np.pi*C0/omega_cutoff/1e-6;
wvlen_cutoff2 = 2*np.pi*C0/(0.92*omega_p)/1e-6;
lambda_p = 2*np.pi*C0/omega_p/1e-6;
print(wvlen_cutoff, lambda_p, omega_p)
wvlen_scan = np.linspace(0.7,20, 1000);
epsilon_diel = 16;
a = 0.2*L0; #lattice constant
Nx = 500
eps_r = epsilon_diel*np.ones((Nx, 1))
eps_r = eps_r.astype('complex')
print(eps_r.shape)
fill_factor = 0.2;
dx= a/Nx;
dL = [dx, 1];
N = [Nx, 1];
L = [a, 1]
print(dL)
fd = FiniteDifferenceGrid(dL,N)
# +
struct = Structure(eps_r, L)
eigen = EigenGuide2D(struct, polarization = 'TM');
# +
kspectra = list();
for i,wvlen in enumerate(wvlen_scan):
omega = 2*np.pi*C0/wvlen/L0;
epsilon_metal = 1-omega_p**2/(omega**2 - 1j*(gamma*omega))
eps_r[int(Nx/2-fill_factor*Nx/2): int(Nx/2+fill_factor*Nx/2)] = epsilon_metal;
struct = Structure(eps_r, L)
eigen.update_structure(struct);
eigen.make_operator_components(omega);
neff = np.sqrt(np.max(np.real(eps_r)));
beta_est = abs(2*np.pi*neff / (wvlen*L0));
sigma = beta_est**2;
Aop = eigen.A;
kvals, modes = la.eigs(Aop, sigma = sigma, k = 10);
kspectra.append(np.sqrt(kvals));
kspectra = np.array(kspectra);
# +
omega_scan = 2*np.pi*C0/wvlen_scan/1e-6
plt.figure(figsize = (5,5));
plt.plot(np.real(kspectra)*1e-6, omega_scan/omega_p, '.b', markersize = 2);
plt.plot(np.imag(kspectra)*1e-6, omega_scan/omega_p, '.r', markersize = 2);
plt.xlim((-30, 50))
plt.xlabel('k (1/($\mu$m))')
plt.ylabel('$\omega/\omega_p$')
#plt.savefig("../img/IMI_band_structure.png",dpi = 300)
plt.show();
#plt.plot(np.imag(kspectra), omega_scan/omega_p, '.r', markersize = 2);
print(os.listdir("../img"))
# -
# ## eigenTE
eigen_te = EigenGuide2D(struct, polarization = 'TE');
kspectra_te = list();
for i,wvlen in enumerate(wvlen_scan):
omega = 2*np.pi*C0/wvlen/L0;
epsilon_metal = 1-omega_p**2/(omega**2 - 1j*(gamma*omega))
eps_r[int(Nx/2-fill_factor*Nx/2): int(Nx/2+fill_factor*Nx/2)] = epsilon_metal;
eigen.update_structure(struct);
eigen.make_operator_components(omega);
neff = np.sqrt(np.max(np.real(eps_r)));
beta_est = abs(2*np.pi*neff / (wvlen*L0));
sigma = beta_est**2;
Aop = eigen.A;
kvals, modes = la.eigs(Aop, sigma = sigma, k = 10);
kspectra_te.append(np.sqrt(kvals));
plt.figure(figsize = (5,5));
plt.plot(np.real(kspectra), omega_scan/omega_p, '.b', markersize = 2);
plt.plot(np.imag(kspectra), omega_scan/omega_p, '.r', markersize = 2);
plt.xlim((-30e6, 50e6))
plt.show();
| notebooks/.ipynb_checkpoints/IMI-MIM k-eigensolver-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Download** (right-click, save target as ...) this page as a jupyterlab notebook from: [Exam 2](http://192.168.3.11/engr-1330-webroot/5-ExamProblems/Exam2/Exam2/.fall2021/Exam2-Fall2021-Deploy.ipynb)
# + language="html"
# <!--Script block to left align Markdown Tables-->
# <style>
# table {margin-left: 0 !important;}
# </style>
# -
# # ENGR 1330 Exam 2 Sec 001 Fall 2021
# ---
# Instructions:
# - Work directly in this notebook, upon completion render the notebook as a PDF file.
# - Upload your solution PDF **AND** the .ipynb (2 files) to the Blackboard repository for grading!
# - Multiple attempts are allowed in the 48-hour time window
# - Internet resources are allowed, remember to cite what you copy from elsewhere.
# - Consulting with each other is **not** permitted.
# - Sharing work with each other is **not** permitted.
# - Working together is **not** permitted.
#
# ---
#
# **Castillo,Andres**
#
# **R11614756**
#
# ENGR 1330 Exam 2 - Take-Home
# ---
#
# ## Problem 0 (1 pts):
# Run the cell below, and leave the results in your notebook.
# If you get an ERROR message, leave it and continue.
#### RUN! this Cell ####
import sys
# ! date
# ! hostname
# ! whoami
print(sys.executable) # OK if generates an exception message on Windows machines
# tested ok MacOS, arm linux, x86-64 linux, Windows 10VM, Windows Server
# ---
# ## Problem 1. Do you want to play a game? (14 pts)
#
# **Problem Statement**
#
# Create a script that will play the Hi-Lo guessing game. The user picks a secret number between 1 and 100. The computer tries to guess the number in less than five turns. When the computer makes a guess, the user enters a hint; the letter 'H' indicates the computer's guess was too high, the letter 'L' means the computer guessed too low. The program should use the hint to revise its guess. If the user enters the letter 'W' that means the computer guessed the number correctly and the computer should reply "I won. As agreed you should now self-terminate". If the computer takes 5 guesses without winning it should print a message "I lost. However I cannot self-terminate, you will have to end me!"
#
# **Deliverables**
# 1. Script to prompt for user input and play the Hi-Lo game
# - Echo each input
# - Report total computer guesses, and outcome (winner or loser)
# 2. Demonstration run
# 3. Play your game 10 times, how many times does the computer win? (you don't need to show each play, just answer the question)
# +
# code here
low = 1
high = 100
print("Enter H if guess is too high \n Enter L if guess is too low \n Enter W if the computer guessed correctly")
starting = (low + high)//2
count = 1
print(starting)
letter = input()
while (letter != 'W') and (count<5):
if letter=='H':
high = starting-1
else:
low = starting+1
starting = (low + high) // 2
count += 1
print(starting)
print('current count:',count)
letter = input()
if letter=='W':
print("I won. As agreed you should now self-terminate")
else:
print("I lost. However I cannot self-terminate, you will have to end me!")
# -
# ---
#
# ## Problem 2 Data interpolation, and plotting (15 pts.)
#
# **Problem Statement**
#
# Plot the following data with time on the horizontal axis, and pressure on the vertical axis.
# Use red circles as the marker.
# Create and plot a data model using a blue curve for the model.
# Use the data model to estimate the pressure at 3 minutes.
#
#
# |Time/min |Pressure/mmHg|
# |---|---|
# |0.0 |7.5 |
# |2.5 |10.5 |
# |5.0 |12.5 |
# |10 |15.8 |
# |15.0 |17.9|
# |20.0 |19.4|
#
# **Deliverables**
#
# 1. Script to generate observed data and plots
# 2. Plot of pressure vs. time with
# - Axis labels and units
# - Plot title
# - Correct marker type and color as per specifications
# 3. Script to generate data model (interpolating polynomial, or power-law,logarithmic,exponential, or polynomial functional form)
# 4. Plot of the data model of pressure vs. time on same chart as 1. above with
# - Correct line type and color
# - Legend to identify the two plotted series.
# 5. Estimated pressure at 3 minutes using the data model.
#
#
# +
# code here
def lagint(xlist,ylist,xpred):
lagint = 0.0
norder = len(xlist)
for i in range(norder):
term = ylist[i]
for j in range(norder):
if (i != j):
term = term * (xpred-xlist[j])/(xlist[i]-xlist[j])
lagint = lagint + term
return(lagint)
xtable = [0.0,2.5,5.0,10,15.0,20.0]
ytable = [7.5,10.5,12.5,15.8,17.9,19.4]
xpred = []
ypred = []
step_size = 0.1
how_many = int(1.0/step_size)
for i in range(how_many+1):
xpred.append(float(i)*step_size)
ypred.append(lagint(xtable,ytable,float(i)*step_size))
import matplotlib.pyplot
myfigure = matplotlib.pyplot.figure(figsize = (9,6))
matplotlib.pyplot.scatter(xtable, ytable ,color ='red')
matplotlib.pyplot.plot(xpred, ypred, color ='blue')
matplotlib.pyplot.xlabel("Time/Min")
matplotlib.pyplot.ylabel("Pressure/mmHg")
matplotlib.pyplot.legend(['Blue Curve is Fitted Polynomial','Red Markers are Observations'])
mytitle = "Interpolating Polynomial Fit to Observations\n "
matplotlib.pyplot.title(mytitle)
matplotlib.pyplot.show()
# -
# ---
#
# ## Problem 3. Data modeling to set production goals (15 pts.)
#
# **Problem Statement**
#
# The revenue and cost of operating from a small Beverly Hills auto bumper gold-plating company was collected over some time. Plot the cost, revenue, and profit( = revenue - cost) vs. number of bumpers plated. Plot the cost data using circular markers, revenue using square markers, and profit using triangular markers.
#
# Build a data model of the cost, revenue, and profit.
# Use the data model to predict the number of units to plate to:
#
# - Minimize cost
# - Maximize profit
#
#
# |Bumpers Plated|Cost(Dollars)|Revenue(Dollars)|
# |---|---|---|
# | 0 |432 |0|
# | 1 |426 |500|
# | 3 |396 |1500|
# | 4 |384 |2000|
# | 9 |594 |4500|
# |13 |1446|6500|
#
#
#
# **Deliverables**
#
# 1. Script to generate observed data and plots of cost, revenue, and profit
# 2. Plot of cost,revenue, and profit vs. units plated with
# - Axis labels and units
# - Plot title
# - Correct marker type as per specifications
# 3. Script to generate data model(s) (power-law, logarithmic, exponential, or polynomial functional form)
# 4. Analysis using the data model you choose
# - Best "fit" using trial-and-error
# - Report sum of squared residuals
# 5. Plot of the data model of cost vs. units plated, revenue vs. units plated, and profit vs. units plated on same chart as 2. above with
# - Correct line type and color (cost = red, revenue = blue, profit = green)
# - Legend to identify the plotted series.
# 6. Recomended production rate to maximize profit.
# +
# code here
import pandas as pd
import matplotlib.pyplot as plt
# Create the data
data = [[0, 432, 0], [1,426, 500], [3, 396, 1500], [4, 384, 2000], [9,594,4500], [13,1446,6500]]
df = pd.DataFrame(data, columns =['Bumpers Plated', 'Cost (Dollars)', 'Revenue (Dollars)'], dtype = float)
print(df)
#Code to Find profit
profitcode = df['Revenue (Dollars)'] - df['Cost (Dollars)']
print(profitcode)
#Add Profit to data frame
dfprofit = df.assign(Profit = profitcode)
Bumpers Plated Cost (Dollars) Revenue (Dollars)
print(dfprofit)
#Plot the data
profitplot = dfprofit.plot('Bumpers Plated',y=['Cost (Dollars)','Revenue (Dollars)', 'Profit'], style = ['o-','s-'
plt.xlabel('Bumpers Plated')
plt.ylabel('Dollar Amount')
plt.show()
# Plot for trail and error fit
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
def maxprofitplot(listx1,listy1,listx2,listy2,strlablx,strlably,strtitle):
profitplot = plt.figure(figsize = (10,5))
plt.plot(listx1,listy1, c='red', marker='o',linewidth=0)
plt.plot(listx2,listy2, c='blue',linewidth=1)
plt.xlabel(strlablx)
plt.ylabel(strlably)
plt.legend(['Bumpers Plated','Profit'])
plt.title(strtitle)
plt.show()
return()
b0 = float(input('Enter a b0 value'))
b1 = float(input('Enter a b1 value:'))
model = b0 + b1*dfprofit['Profit']
maxprofitplot(dfprofit['Profit'],dfprofit['Revenue (Dollars)'],dfprofit['Profit'],model,"Bumpers Plated","Profit"
#Sum of Sqaured Residuals
def sumofsquares(res1,res2,res3):
if len(res1)!=len(res2) or len(res1)!=len(res3):
return
for i in range(len(res1)):
res3[i]=res1[i]-res2[i]
return(res3)
residuals = [0 for i in range(len(dfprofit['Profit']))] # empty list
sumofsquares(dfprofit['Profit'],model,residuals)
print(sum(residuals))
for i in [range(len(residuals))]:
residuals[i]=residuals[i]**2
print(sum(residuals))
#Cost vs Bumpers Plated
profitplot = dfprofit.plot('Bumpers Plated',y=['Cost (Dollars)'], style = ['o-r'])
plt.xlabel('Bumpers Plated')
plt.ylabel('Dollar Amount')
plt.show()
#Revenue vs Bumpers Plated
profitplot = dfprofit.plot('Bumpers Plated',y=['Revenue (Dollars)'], style = ['s-b'])
plt.xlabel('Bumpers Plated')
plt.ylabel('Dollar Amount')
plt.show()
#Bumpers Plated Vs Profit
profitplot = dfprofit.plot('Bumpers Plated',y=['Cost (Dollars)','Revenue (Dollars)', 'Profit'], style = ['o-r','s-b'
plt.xlabel('Bumpers Plated')
plt.ylabel('Dollar Amount')
plt.show()
#Bumpers Plated Vs Profit
profitplot = dfprofit.plot('Bumpers Plated',y=['Profit'], style = ['v-g'])
plt.xlabel('Bumpers Plated')
plt.ylabel('Dollar Amount')
plt.show()
# -
# ___
# ## Problem 4. (40 pts): On "Bottled Poetry" ...
#
# <img src="https://w7w5t4b3.rocketcdn.me/wp-content/uploads/2016/04/game-of-thonres-tyrion-lannister-i-drink-and-i-know-things.jpg" width="500">
#
#
# The "winequality-red.csv" and "winequality-white.csv" datasets provide information related to red and white variants of vinho verde wine samples, from the north of Portugal. The goal is to model wine quality based on physicochemical tests. Follow the steps and answer the question. *Due to privacy and logistic issues, only physicochemical (inputs) and sensory (the output) variables are available (e.g. there is no data about grape types, wine brand, wine selling price, etc.).*
#
#
# #### The datasets consists of several Input variables (based on physicochemical tests).
#
# |Columns|Info.|
# |---:|---:|
# |fixed acidity |most acids involved with wine or fixed or nonvolatile (do not evaporate readily)|
# |volatile acidity |the amount of acetic acid in wine, which at too high of levels can lead to an unpleasant, vinegar taste|
# |citric acid |found in small quantities, citric acid can add 'freshness' and flavor to wines|
# |residual sugar |the amount of sugar remaining after fermentation stops, it's rare to find wines with less than 1 gram/liter|
# |chlorides |the amount of salt in the wine|
# |free sulfur dioxide |the free form of SO2 exists in equilibrium between molecular SO2 (as a dissolved gas) and bisulfite ion|
# |total sulfur dioxide |amount of free and bound forms of S02; in low concentrations, SO2 is mostly undetectable in wine|
# |density |the density of water is close to that of water depending on the percent alcohol and sugar content|
# |pH |describes how acidic or basic a wine is on a scale from 0 (very acidic) to 14 (very basic); most wines are between 3-4|
# |sulphates |a wine additive which can contribute to sulfur dioxide gas (S02) levels, wich acts as an antimicrobial|
# |alcohol |the percent alcohol content of the wine|
# |quality (score between 0 and 10) |output variable (based on sensory data, score between 0 and 10)|
#
#
#
# **References:**
#
# <NAME>, <NAME>, <NAME>, <NAME> and <NAME>. Modeling wine preferences by data mining from physicochemical properties. In Decision Support Systems, Elsevier, 47(4):547-553, 2009. [https://www.sciencedirect.com/science/article/pii/S0167923609001377?via%3Dihub](https://www.sciencedirect.com/science/article/pii/S0167923609001377?via%3Dihub)
#
# <!--
# https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv
#
# https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv
#
# https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv
# -->
#
# Download the red-variety dataset from [https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv](https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv) using the script below:
#
# **If you get an error, then download using your browser and mouse**
######### CODE TO AUTOMATICALLY DOWNLOAD THE DATABASE ################
# #! pip install requests #install packages into local environment
import requests # import needed modules to interact with the internet
# make the connection to the remote file (actually its implementing "bash curl -O http://fqdn/path ...")
remote_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv' # a csv file
response = requests.get(remote_url) # Gets the file contents puts into an object
output = open('winequality-red.csv', 'wb') # Prepare a destination, local
output.write(response.content) # write contents of object to named local file
output.close() # close the connection
# 1. Read the data as a data frame and print the first few rows. In a few lines explain what can be understood about the data from this.
# 2. Use the appropriate function and get a summary of information on the data frame. Explain what you can learn from this summary report.
# 3. Are there any missing values in the data? Justify your answer.
# 4. Use the appropriate function and get the 5-number summary for the data frame. Explain what you can learn from this summary report for each column.
# 5. Rename the "quality (score _0to10)" column heading to "quality"
# 6. Make a subset of all the wines with a quality above 7. Name this subset "TopQ".
# 7. What percentage of wines in "TopQ" has an alcohol content less than 10%? What is this percentage out of the entire set of wine (the original data)?
# 8. Print the above subset of the dataframe, sorted by wine quality.
# 9. Define a function that labels the wines based on their quality according to the table below: <br>
#
# |Quality Score|Label.|
# |---:|---:|
# |q>= 7|Top|
# |5<q<7|Average|
# |q<=5|Low|
#
# 10. Apply the function on the data frame and store the result in a new column "Qlabel".
# 11. Report the share of each quality label in percentage.
# 12. Plot a histogram of pH for all the Low quality wines. Explain what you can infer from this plot.
# 13. Make a similar histogram for pH for all the Top quality wines. Put the new histogram and the previous one next to each other and explain what you can infer by comparing them.
# +
# Put your code here - start with scaffolding and back-fill code
# Read the data as a data frame and print the first few rows.
# Summary of information on the data frame.
# Are there any missing values in the data?
# ... put rest of scaffolding here
# ...
# ...
# Make a similar histogram for pH for all the Top quality wines.
# ...
# -
import pandas as pd
# Read the data as a data frame and print the first few rows.
winedata = pd.read_csv('winequality-red.csv', sep=';', engine='python')
winedata.head(4)
# Summary of information on the data frame.
# Are there any missing values in the data?
# ... put rest of scaffolding here
# ...
# ...
# Make a similar histogram for pH for all the Top quality wines.
winedata.info()
winedata.describe()
# +
TopQ = winedata.loc[(winedata['quality'] > 7)]
display(TopQ)
#Find all wines with alcohol content below 10.0
winedatabelow10 = winedata.loc[(winedata['alcohol'] < 10.0)]
display(winedatabelow10)
#Step 7
print('Number of top quality wines')
len(TopQ)
# -
print('Total Number of Wines with Alcohol Content Below 10')
len(winedatabelow10)
def assignNewLabels(label):
if label >= 7:
return "Top"
elif 5 < label < 7 :
return "Average"
elif label <= 5 :
return "Low"
else:
return 'NA'
winedata['QLabel'] = winedata['quality'].apply(assignNewLabels)
display(winedata)
# +
winedata['QLabel'].value_counts(normalize=True) * 100
lowqualitywine = winedata.loc[(winedata['QLabel'] == 'Low')]
display(lowqualitywine)
topqualitywine = winedata.loc[(winedata['QLabel'] == "Top")]
display(topqualitywine)
#Step 12
import matplotlib.pyplot as plt
lowqualitywine.plot(kind='hist',x='quality',y='pH',color='purple', bins = 20)
plt.xlabel('pH Quality')
plt.ylabel('Samples')
plt.title('Low Quality Wine Samples and Its pH')
plt.show()
#Step 13
topqualitywine.plot(kind='hist',x='quality',y='pH',color='pink', bins = 20)
plt.xlabel('pH Quality')
plt.ylabel('Samples')
plt.title('Top Quality Wine Samples and Its pH')
plt.show()
a = lowqualitywine['pH']
b = topqualitywine['pH']
plt.figure(figsize=(8,6))
plt.hist(a, bins=20, alpha=0.5, label="Low Quality", color = 'purple')
plt.hist(b, bins=20, alpha=0.5, label="Top Quality", color = 'pink')
plt.xlabel("pH Level", size=14)
plt.ylabel("Sample Frequency", size=10)
plt.title("Top Quality and Low Q")
# -
# **Deliverables**
# 1. Script to read datafile and process as directed above
# 2. Function to generate wine quality label classifications as directed above.
# 3. Analysis using the function
# 4. Histograms (side-by-side) of low quality and high quality wines, as classified above.
# - Axis labels and units
# - Plot title
# - Suitable bin count
# winedata.info()
# ___
# # Problem 5 (15 pts): To cause or to correlate; that is the question!
#
# <img src="https://aspectmr.com/wp-content/uploads/2019/11/chicken_makes_you_go_to_the_moon.jpg" align="right" width="300">
#
#
# 1. What is the message behind the above image in relation to causation and correlation?
# 2. Why is it important to understand the difference between causation and correlation?
# 3. Using your wine database, assume that the quality score can be treated as a continuous $~~~$ <br> variable (a *float*). Determine the Pearson's correlation coefficient between pH and the quality score.
# 4. What do you conclude from the correlation?
# code here - correlation analysis
qualityscore = winedata['quality']
pHscore = winedata['pH']
pearsoncorrelation = qualityscore.corr(pHscore)
print(pearsoncorrelation)
winedata.plot.scatter(x='pH', y='quality', s=22, alpha=0.25, c='purple', figsize=(8, 6))
plt.xlabel("pH")
plt.ylabel("Quality")
plt.title("Plot of Quality and its pH")
plt.legend(loc='upper right')
# **Deliverables**
# 1. Answers to questions 1 and 2 above.
# 2. Working correlation analysis script on appropriate dataframe elements
# 3. Scatterplot of quality score (y-axis) versus pH (x-axis)
# - Axis labels and units
# - Plot title
# 4. Conclusion (answer to last question above)
# ---
# ___
#
# ## <font color=green>Bonus Problem (10 points)</font>
#
# Repeat Problem 4 using the white wine database. Download the white-variety dataset from [https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv](https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv)
#
# Is there evidence that pH is a predictor of wine quality?
# +
# code here
# -
# ---
# ___
#
# ## <font color=blue>Bonus Problem (20 points)</font>
#
# Consider the polynomial data model below:
#
# > Polynomial Model: $y_{model} = \beta_0 + \beta_1 x_{obs} + \beta_2 x_{obs}^2 + ... + \beta_n x_{obs}^n$ <br>
# <!--Logarithmic Model: $y_{model} = \beta_0 + \beta_1 log(x_{obs}) $<br>
# Power-Law Model: $y_{model} = \beta_0 x_{obs}^{\beta_1} $ <br>
# Exponential Model: $y_{model} = \beta_0 e^{{\beta_1}x_{obs}} $ -->
#
# One way to "fit" this model to data is to construct a design matrix $X$ comprised of $x_{obs}$ and ones (1). Then construct a linear system related to this design matrix.
#
# The data model as a linear system is:
#
# $$\begin{gather}
# \mathbf{X} \cdot \mathbf{\beta} = \mathbf{Y}
# \end{gather}$$
#
# For example using the Polynomial Model (order 2 for brevity, but extendable as justified)
#
# \begin{gather}
# \mathbf{X}=
# \begin{pmatrix}
# 1 & x_1 & x_1^2\\
# ~\\
# 1 & x_2 & x_2^2\\
# ~ \\
# 1 & x_3 & x_3^2\\
# \dots & \dots & \dots \\
# 1 & x_n & x_n^2\\
# \end{pmatrix}
# \end{gather}
#
# \begin{gather}
# \mathbf{\beta}=
# \begin{pmatrix}
# \beta_0 \\
# ~\\
# \beta_1 \\
# ~ \\
# \beta_2 \\
# \end{pmatrix}
# \end{gather}
#
# \begin{gather}
# \mathbf{Y}=
# \begin{pmatrix}
# y_1 \\
# ~\\
# y_2 \\
# ~ \\
# y_3 \\
# \dots \\
# y_n \\
# \end{pmatrix}
# \end{gather}
#
# To find the unknown $\beta$ values the solution of the linear system below provides a "best linear unbiased estimator (BLUE)" fit
#
# $$\begin{gather}
# [\mathbf{X^T}\mathbf{X}] \cdot \mathbf{\beta} = [\mathbf{X^T}]\mathbf{Y}
# \end{gather}$$
#
# or an alternative expression is
#
# $$\begin{gather}
# \mathbf{\beta} = [\mathbf{X^T}\mathbf{X}]^{-1}[\mathbf{X^T}]\mathbf{Y}
# \end{gather}$$
#
# Once the values for $\beta$ are obtained then we can apply our plotting tools and use the model to extrapolate and interpolate. Most of the arithmetic can be accomplished using numpy as in Lesson 9.
#
# Consider the data collected during the boost-phase of a ballistic missle. The maximum speed of a solid-fueled missle at burn-out (when the boost-phase ends) is about 7km/s. Using this knowledge and the early-time telemetry below; fit a data model using the linear system approach above.
#
# |Elapsed Time (s)|Speed (m/s)|
# |---:|---:|
# |0 |0|
# |1.0 |3|
# |2.0 |7.4|
# |3.0 |16.2|
# |4.0 |23.5|
# |5.0 |32.2|
# |6.0 | 42.2|
# |7.0 | 65.1 |
# |8.0 | 73.5 |
# |9.0 | 99.3 |
# |10.0| 123.4|
#
# Use the model to estimate boost phase burn-out time. Plot the model and data on the same axis to demonstrate the quality of the fit.
# +
# code here
import numpy as np
import matplotlib.pyplot as plt
Elapsed_Time = np.array([0,1.0,2.0,3.0,4.0,5.0,6.0,7.0,8.0,9.0,10.0])
Speed = np.array([0,3,7.4,16.2,23.5,32.2,42.2,65.1,73.5,99.3,123.4])
plt.rcParams['figure.figsize'] = (10,7)
plt.scatter(Elapsed_Time,Speed,color="blue")
plt.xlabel("time")
plt.ylabel("speed")
m,b = np.polyfit(Elapsed_Time,Speed,1)
plt.plot(Elapsed_Time, m*Elapsed_Time+b, color="red")
plt.show()
# -
| 7-SpecialNotebooks/.ipynb_checkpoints/Exam2-Fall2021-Deploy (1)-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # On Benford's Law and NYC Taxi Trip Durations
# In this notebook, we will demonstrate how [Benford's Law](https://en.wikipedia.org/wiki/Benford%27s_law) is reflected in the first digit of time durations of [NYC taxi trips](https://www.kaggle.com/c/nyc-taxi-trip-duration/data).
#
# Last night, I woke up at 2AM and could not get back to sleep and started TV surfing and stublled upon Netflix [Connected, the Digits eposide](https://www.netflix.com/title/81031737). The reporter was talking about the usage of Benford's law. As I was watching it, I was constantly saying "No Way!". At the end of the episode, I looked up the [law](https://en.wikipedia.org/wiki/Benford%27s_lawhttps://en.wikipedia.org/wiki/Benford%27s_law) (oh...and never went back to sleep that night due to the excitement :-) and went about replicating the curve with some data to "see" the curve.
#
# Though the law can be applied to a "small" set, the more data you have, the better. That is why I used my fav dataset (NYC Taxi Data) for BigData analysis. And, a 1 through 9, as a first digit in the data, has to have a very good likelihood of occurance. For example, latitude and longitude values in a restricted area like NYC are NOT good candicate, as 7 and 4 are the only values (see below the extent values). However, a trip duration value is a GREAT candidate.
#
# So...this notebook uses [Apache Spark](https://spark.apache.org/) to prep the data, in such that we can "see" a Benford curve in the processed output.
import os
import numpy as np
import pandas as pd
import seaborn as sms
import matplotlib.pyplot as plt
plt.style.use('dark_background')
# Download the [data](https://www.kaggle.com/c/nyc-taxi-trip-duration/datahttps://www.kaggle.com/c/nyc-taxi-trip-duration/data) into a folder named `kaggle` in your home directory.
load_path = os.path.expanduser(os.path.join("~","kaggle","nyc-taxi-trip-duration","train.csv"))
# Load the data into a Spark dataframe and map it to a SQL view named `v0`.
# +
schema = ",".join([
"id string",
"vendor_id string",
"pickup_datetime timestamp",
"dropoff_datetime timestamp",
"passenger_count integer",
"pickup_longitude double",
"pickup_latitude double",
"dropoff_longitude double",
"dropoff_latitude double",
"store_and_fwd_flag string",
"trip_duration integer"
])
df = spark\
.read\
.format("csv")\
.option("header",True)\
.option("parserLib", "univocity")\
.option("mode", "permissive")\
.schema(schema)\
.load(load_path)\
.drop("id","vendor_id","passenger_count","store_and_fwd_flag")\
.selectExpr("*","hour(pickup_datetime) pickup_hour")\
.cache()
df.createOrReplaceTempView("v0")
# -
# Restrict the pickups and dropoffs to an area around Manhattan.
#
# 
lon_min,lat_min,lon_max,lat_max = (-75.08761590197491, 40.122033125848525, -72.33354542760787, 41.68167992377412)
# Accumulate the first digit of the trip duration field statistics and collect the result as a Pandas dataframe.
#
# Note here we are convering the `trip_duration` numerical value to a string value, in such that we can pick up the first digit as a character using `substr` function.
pdf = spark.sql(f"""
select b,count(1) pop
from(
select substr(cast(trip_duration as string),0,1) b
from v0
where pickup_longitude between {lon_min} and {lon_max}
and pickup_latitude between {lat_min} and {lat_max}
and dropoff_longitude between {lon_min} and {lon_max}
and dropoff_latitude between {lat_min} and {lat_max}
and trip_duration > 0
)
group by b
order by b
""")\
.toPandas()
# Add a percent pandas series.
pdf['pct'] = 100.0 * pdf['pop'] / pdf['pop'].sum()
# Compute the Benford values.
bf = 100.0 * np.log10(1.0+1.0/np.arange(1,10,1))
# Plot the occurence and the Benford curve.
plt.figure(figsize=(16,6))
bar = plt.bar(pdf['b'],pdf['pct'], label='Percent of Occurence')
plt.plot(pdf['b'], bf, 'r', label='Benford')
plt.xlabel('First Digit of Trip Duration Field')
# plt.xticks(rotation=90)
plt.legend()
plt.show()
# # MIND BLOWN !!!
# + [markdown] pycharm={"name": "#%% md\n"}
# **PS**: Could not sleep last nite is because my mind was rattled by what happened to my beloved Beirut. "Paris of the Middle East" did not need this in the middle of corona and an economic crisis.
#
# This article is dedicated to all that passed away and injured, specially my friend Dr. Georgie. Speedy recovery.
#
# "I Love you ya Lubnan, Ya watani" - Fairuz
| taxi_trips_benford.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Class Coding Lab: Walkthrough
#
# For this class coding lab paste the code provided by the walk through into the cell below. Execute the cell to verfiy the program runs correctly and also saves the notebook output.
print("Testing to ensure all modudles are installed...")
import requests
import json
import pandas as pd
import numpy as np
import matplotlib
import plotly
import cufflinks as cf
import folium
from IPython.display import display, HTML
display(HTML("This is <span style='color: blue;'>Working</span>"))
| content/lessons/01/Class-Coding-Lab/CCL-walkthrough.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="qt7QZeHklDbq" colab={"base_uri": "https://localhost:8080/"} outputId="09368bb7-15cd-44e3-84ca-879683ba4f6e"
# !pip install -q facenet_pytorch
from facenet_pytorch import MTCNN
from google.colab import files
from PIL import Image
import os
from pathlib import Path
import matplotlib.pyplot as plt
import torch
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# + id="wpnLfebgkUCq"
mtcnn = MTCNN(keep_all=False, image_size=256, margin=50,post_process=True,min_face_size=100,device=device)
# + id="gjQNJDjXkX2h"
def getFaces(images,path,out_path):
Path(out_path).mkdir(parents=True, exist_ok=True)
for image in images:
filename = path+image
new_name = out_path+image
im = Image.open(filename)
save_paths = new_name
mtcnn(im, save_path=save_paths)
print("All Faces Extracted")
# + id="RJ0EQQz_kxiY"
game_path = "/content/drive/MyDrive/Game-Movie/images/game/" #path for games and images
movies_path = "/content/drive/MyDrive/Game-Movie/images/movies/"
game_images = next(os.walk(game_path))[2]
movie_images = next(os.walk(movies_path))[2]
# + colab={"base_uri": "https://localhost:8080/"} id="4iQGNYlHl71f" outputId="f2e2a9bc-a06f-4582-c788-ca70ff69093a"
getFaces(game_images,game_path,"/content/drive/MyDrive/Game-Movie/face_images/game/")
getFaces(movie_images,movies_path,"/content/drive/MyDrive/Game-Movie/face_images/movies/")
# + colab={"base_uri": "https://localhost:8080/"} id="1JywWZEwtHNA" outputId="44376ae9-e236-4069-a5f7-5178900f4f11"
import h5py
from PIL import Image
import os, os.path
import random
import numpy as np
game_path = "/content/drive/MyDrive/Game-Movie/face_images/game/"
movie_path = "/content/drive/MyDrive/Game-Movie/face_images/movies/"
game_fileName = "/content/drive/MyDrive/Game-Movie/face_images/h5/game_faces.h5"
movie_fileName = "/content/drive/MyDrive/Game-Movie/face_images/h5/movie_faces.h5"
def converth5(path,output_path):
#get names of the images and randomize
# Path(output_path).mkdir(parents=True, exist_ok=True)
files = [name for name in os.listdir(path)]
random.shuffle(files)#inplace operation
#define some variable according to the size
size = 256
numTrain = int(0.15*len(files))
numTest = len(files)-numTrain
#create the h5 files
f = h5py.File(output_path, "w")
with f as out:
out.create_dataset("X_train",(numTrain,size,size,3),dtype='u1')
out.create_dataset("X_test",(numTest,size,size,3),dtype='u1')
#populate the h5 file
f = h5py.File(output_path, "a")
with f as out:
for index,img_name in enumerate(files[:numTrain]):
img = Image.open(path+img_name)
out['X_train'][index] = np.asarray(img)
print("Conversion Complete")
converth5(game_path,game_fileName)
# + colab={"base_uri": "https://localhost:8080/"} id="13KtuGHEmt6C" outputId="6d50ecd1-a829-4148-fa00-9f2b51d418e1"
# converth5(game_path,game_fileName)
converth5(movie_path,movie_fileName)
| Face_Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Cartilha criada com auxílio do canal 'Hashtag Programação'
# ### Importando o pandas
import pandas as pd
# ### Criando um dataframe a partir de um dicionário
# dataframe = pd.DataFrame()
venda = {'data': ['15/02/2021', '16/02/2021'],
'valor': [500, 300],
'produto': ['feijao', 'arroz'],
'qtde': [50, 70],
}
venda_df = pd.DataFrame(venda)
# ### Visualização dos Dados
# - print
# - display
print(venda_df)
display(venda_df)
# ### Importando arquivos e bases de dados
planilha_vendas = pd.read_excel('Vendas.xlsx')
display(planilha_vendas)
# ### Resumos de Visualização de Dados simples e úteis
# - head
# - shape
# - describe
planilha_vendas.head()
planilha_vendas.shape
planilha_vendas.describe()
# ### Pegar 1 coluna (e os pd.Series)
#Series = colunas do Dataframe
produtos = planilha_vendas['Produto']
display(produtos)
# # + DE UMA COLUNA
produtos = planilha_vendas[['Produto', 'ID Loja']]
display(produtos)
# ### .loc, um método muito importante
# - Pegar 1 linha
# - Pegar linhas de acordo com alguma condição
# - Pegar linhas e colunas específicas
# - Pegar 1 valor específico
# -> O metodo loc trabalha com os indices(nº ao lado direito da tabela)
#
# -> dataframe.loc['linhas','colunas']
#Pegar uma linha
display(planilha_vendas.loc[1])
#Pegar de uma até outra
display(planilha_vendas.loc[1:5])
#Pegar linhas que correspondem a uma condição
vendas_iguatemi_esp = planilha_vendas.loc[planilha_vendas['ID Loja'] == 'Iguatemi Esplanada']
display(vendas_iguatemi_esp)
#Pegar várias linhas e colunas usando loc
vendas_norte_shop = planilha_vendas.loc[planilha_vendas['ID Loja'] == 'Norte Shopping',['ID Loja','Produto','Quantidade']]
display(vendas_norte_shop)
#Pegar valor especifico
print(planilha_vendas.loc[1, 'Produto'])
# ### Adicionar 1 coluna
#A partir de uma coluna que exite
planilha_vendas['Comissão'] = planilha_vendas['Valor Final'] * 0.05
display(planilha_vendas)
#A partir de um valor padrão
planilha_vendas.loc[:,'Imposto'] = 0
display(planilha_vendas)
# ### Adicionar 1 linha
# - Linhas de um complemento da base de dados
vendas_dez = pd.read_excel('Vendas - Dez.xlsx')
planilha_vendas = planilha_vendas.append(vendas_dez)
display(planilha_vendas)
# ### Excluir linhas e colunas
planilha_vendas = planilha_vendas.drop('Imposto',axis=1) #1 é o eixo da coluna, 0 é o eixo da linha
planilha_vendas = planilha_vendas.drop(0,axis=0)
display(planilha_vendas)
# ### Valores Vazios
# - Deletar linhas/colunas vazias (NaN)
# - Deletar linhas que possuem valores vazios
# - Preencher valores vazios (média e último valor)
# +
#Deletar linhas e colunas completamente vazias
planilha_vendas = planilha_vendas.dropna(how = 'all', axis =1)
#deletar se tiver pelo menos um valor vazio
planilha_vendas = planilha_vendas.dropna()
#Preencher valores vazios
#Preencher com valor fixo
planilha_vendas['Comissão'] = planilha_vendas['Comissão'].fillna(1)
#Preencher com média da coluna
planilha_vendas['Comissão'] = planilha_vendas['Comissão'].fillna(planilha_vendas['Comissão'].mean())
display(planilha_vendas)
#Prencher com ultimo valor (Pegar o valor de cima e coloca embaixo)
planilha_vendas = planilha_vendas.ffill()
# -
# ### Calcular Indicadores
# - Groupby
# - Value Counts = Conta quantas vezes um determinado item aparece
#Value counts
transacoes_loja = planilha_vendas['ID Loja'].value_counts()
display(transacoes_loja)
#Group By (Funciona melhor com coluna especificada para agrupar e colunas numericas para fazer o metodo desejado)
faturamento_produto = planilha_vendas[['Produto','Valor Final']].groupby('Produto').sum()
display(faturamento_produto)
# ### Mesclar 2 dataframes (Procurar informações de um dataframe em outro)
#é necessário ambos dataframes terem uma coluna em comum
gerentes_df = pd.read_excel('Gerentes.xlsx')
planilha_vendas = planilha_vendas.merge(gerentes_df)
display(planilha_vendas)
| Python_Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Zemberek: Word Analysis Example
Documentation: https://bit.ly/2MTmfr1
Java Code Example: https://bit.ly/2MV2Hmj
"""
from os.path import join
from jpype import JClass, JString, getDefaultJVMPath, shutdownJVM, startJVM
if __name__ == '__main__':
ZEMBEREK_PATH: str = join( 'bin', 'zemberek-full.jar')
startJVM(
getDefaultJVMPath(),
'-ea',
f'-Djava.class.path={ZEMBEREK_PATH}',
convertStrings=False
)
# -
TurkishMorphology: JClass = JClass('zemberek.morphology.TurkishMorphology')
AnalysisFormatters: JClass = JClass('zemberek.morphology.analysis.AnalysisFormatters')
WordAnalysis: JClass = JClass('zemberek.morphology.analysis.WordAnalysis')
morphology: TurkishMorphology = TurkishMorphology.createWithDefaults()
word: str = 'okumak'
print(f'\nWord: {word}')
results: morphology = morphology.analyze(JString(word))
for result in results:
print(result.getStem())
shutdownJVM()
| Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. test data
# ## 1.1 init
import pandas as pd
import sys
sys.path.append('C:/ProgramData/Anaconda3/Lib/site-packages')
import ERgene
import matplotlib.pyplot as plt
data=pd.read_csv('C:/Users/FernandoZeng/Desktop/GSE103931_series_matrix.txt',sep='\t')
data=data.set_index(data.columns[0])
font1 = {
'weight' : 'bold',
'size' : 18,
}
data.plot(kind = 'density')
plt.xlim(-5,10)
import matplotlib.pyplot as plt
data=pd.read_csv('test.csv')
data=data.set_index(data.columns[0])
font1 = {
'weight' : 'bold',
'size' : 18,
}
data.plot(kind = 'density')
plt.xlim(-1000,1000)
data.plot(kind = 'box')
#plt.ylim(-200,800)
# ## 1.2 Find ERgene
ERgene.FindERG(data,10)
# ## 1.3 normalizationdata
data2=ERgene.normalizationdata(data,'miR-92a')
data2.plot(kind = 'density')
plt.xlim(-5,10)
data2.plot(kind = 'box')
#plt.ylim(-200,800)
data2[data2['ID_REF']=='miR-486-3p'].to_csv('tttt.csv')
| example/origin_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
#
#
# This notebook shows how **CKG** can be used to download data from the Proteomics Identifications Database - PRIDE - (https://www.ebi.ac.uk/pride/) and quickly formated to start analyzing them with the functionality in the analytics core.
#
#
#
#
# +
import os
import ckg_utils
from graphdb_builder import builder_utils
from graphdb_builder.experiments.parsers import proteomicsParser
from analytics_core.analytics import analytics
# -
# ##### CKG path
ckg_location = '/Users/albertosantos/Development/Clinical_Proteomics_Department/ClinicalKnowledgeGraph(CKG)/code'
# #### Define where the data should be downloaded
analysis_dir = os.path.join(ckg_location, '/data/tmp/Deshmukh2019')
ckg_utils.checkDirectory(analysis_dir)
# ##### Specify the PRIDE identifier and file to be downloaded
pxd_id = 'PXD008541'
file_name='SearchEngineResults_secretome.zip.rar'
# ##### Download data
#
# We can use functionality in graphdb_builder to directly download data files from EBI's PRIDE database (https://www.ebi.ac.uk/pride/). For that you just need to specify the PRIDE identifier for the project (PXD...) and the name of the file to download. In this case, the project identifier is **PXD008541** and the file we will use is **SearchEngineResults_secretome.zip.rar**, a RAR compressed file with the output files from MaxQuant.
builder_utils.download_PRIDE_data(pxd_id=pxd_id,
file_name=file_name,
to=analysis_dir)
# ## Read Data In
# ### Decompress File
builder_utils.unrar(filepath=os.path.join(analysis_dir, file_name), to=analysis_dir)
# The list of files within the compressed folder can be listed using the listDirectoryFiles functionality in gaphdb_builder.
builder_utils.listDirectoryFiles(analysis_dir)
# We use the proteinGroups file that contains the proteomics data processed using MaxQuant software.
proteinGroups_file = os.path.join(analysis_dir, 'proteinGroups.txt')
# ### Parse Contents
#
# CKG has parsers for MaxQuant and Spectronaut output files. The default configuration needed to parse these files needs to be updated with the name of the columns containing the protein quantifications for each sample. Also, the default configuration can be adapted to the experiment by selected specific filters or removing non-used columns. For example, in this study the output file did not have columns: Score, Q-value, so we removed them from the configuration and the column 'Potential contaminant' was renamed to 'Contaminant' so we changed the name in the filters.
# +
#d = pd.read_csv(proteinGroups_file, sep='\t')
#d.columns.tolist()
# -
columns = ['LFQ intensity BAT_NE1',
'LFQ intensity BAT_NE2',
'LFQ intensity BAT_NE3',
'LFQ intensity BAT_NE4',
'LFQ intensity BAT_NE5',
'LFQ intensity BAT_woNE1',
'LFQ intensity BAT_woNE2',
'LFQ intensity BAT_woNE3',
'LFQ intensity BAT_woNE4',
'LFQ intensity BAT_woNE5',
'LFQ intensity WAT_NE1',
'LFQ intensity WAT_NE2',
'LFQ intensity WAT_NE3',
'LFQ intensity WAT_NE4',
'LFQ intensity WAT_NE5',
'LFQ intensity WAT_woNE1',
'LFQ intensity WAT_woNE2',
'LFQ intensity WAT_woNE3',
'LFQ intensity WAT_woNE4',
'LFQ intensity WAT_woNE5',
'Contaminant']
configuration = proteomicsParser.update_configuration(data_type='proteins',
processing_tool='maxquant',
value_col='LFQ intensity',
columns=columns,
drop_cols=['Score', 'Q-value', 'Potential contaminant'],
filters=['Reverse', 'Only identified by site', 'Contaminant'])
configuration
# When we parse the data, we obtain a matrix in an edge list following CKG's graph format: sample, protein, realtionship_type, value, protein_group_id, is_razor
data = proteomicsParser.parser_from_file(proteinGroups_file, configuration=configuration, data_type='proteins', is_standard=False)[('proteins', 'w')]
data.head()
data.columns = ['sample', 'identifier', 'relationship', 'LFQ intensity', 'id', 'is_razor']
data.head()
data.shape
data = data[data.is_razor]
data.shape
# We can use the sample names to extract the group information: BAT_NE, WAT_NE, BAT_woNE, WAT_woNE
#
# With this last column, we obtain the **original dataframe** used as starting point in CKG' analysis pipelines.
data['group'] = data['sample'].apply(lambda x: re.sub('\d', '', x))
data.head()
original = data[['group', 'sample', 'identifier', 'LFQ intensity']]
# ##### --> the original dataframe is the starting point in CKG's proteomics analysis.
# ## Data Preparation
#
# In order to prepare the data we follow the steps:
#
# 1) Filtering based on missing values
#
# 2) Imputation of missing values using a mixed model estrategy: KNN and MinProb
#
# These steps will generate the **processed dataframe**, a complete matrix that can be used in the exploratory and statistical analysis.
processed_data = analytics.get_proteomics_measurements_ready(original,
index_cols=['group', 'sample'],
drop_cols=['sample'],
group='group',
identifier='identifier',
extra_identifier=None,
imputation=True,
method='mixed',
knn_cutoff=0.4,
missing_method='at_least_x',
missing_per_group=True,
min_valid=3,
value_col='LFQ intensity',
shift=1.8,
nstd=0.3)
processed_data.head()
| src/notebooks/recipes/Download_PRIDE_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# Read dataset file ckd.csv
dataset = pd.read_csv("kidney_disease_no_id.csv",header=0, na_values="?")
# Replace null values "?" by numpy.NaN
dataset.replace("?", np.NaN)
# Convert nominal values to binary values
cleanup = {"rbc": {"normal": 1, "abnormal": 0},
"pc": {"normal": 1, "abnormal": 0},
"pcc": {"present": 1, "notpresent": 0},
"ba": {"present": 1, "notpresent": 0},
"htn": {"yes": 1, "no": 0},
"dm": {"yes": 1, "no": 0},
"cad": {"yes": 1, "no": 0},
"appet": {"good": 1, "poor": 0},
"pe": {"yes": 1, "no": 0},
"ane": {"yes": 1, "no": 0}}
# Replace binary values into dataset
dataset.replace(cleanup, inplace=True)
# Fill null values with mean value of the respective column
dataset.fillna(round(dataset.mean(),2), inplace=True)
dataset.to_csv("Preprocessed_ckd.csv", sep=',', index=False)
data =pd.read_csv('Preprocessed_ckd.csv')
data
data.corr()
columns_to_retain = ["Sg", "Al", "Hemo", "Pcv", "Rbcc", "Htn", "Dm", "Class"]
modified_data = data.drop([col for col in data.columns if not col in columns_to_retain], axis=1)
modified_data
modified_data.to_csv('modified_ckd_with_1_and_0.csv', sep=',', index=False)
cleanup = {"Htn": {1: "yes", 0: "no"},
"Dm": {1: "yes", 0: "no"},
"Class": {1: "yes", 0: "no"}}
modified_data.replace(cleanup, inplace=True)
modified_data
modified_data.to_csv('final_modified_ckd_for_prediction.csv', sep=',', index=False)
| Dataset Preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hands-on Federated Learning: Image Classification
#
# In their recent (and exteremly thorough!) review of the federated learning literature [*Kairouz, et al (2019)*](https://arxiv.org/pdf/1912.04977.pdf) define federated learning as a machine learning setting where multiple entities (clients) collaborate in solving a machine learning problem, under the coordination of a central server or service provider. Each client’s raw data is stored locally and not exchanged or transferred; instead, focused updates intended for immediate aggregation are used to achieve the learning objective.
#
# In this tutorial we will use a federated version of the classic MNIST dataset to introduce the Federated Learning (FL) API layer of TensorFlow Federated (TFF), [`tff.learning`](https://www.tensorflow.org/federated/api_docs/python/tff/learning) - a set of high-level interfaces that can be used to perform common types of federated learning tasks, such as federated training, against user-supplied models implemented in TensorFlow or Keras.
# # Preliminaries
# +
import collections
import os
import typing
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow.keras as keras
import tensorflow_federated as tff
# -
# required to run TFF inside Jupyter notebooks
import nest_asyncio
nest_asyncio.apply()
tff.federated_computation(lambda: 'Hello, World!')()
# # Preparing the data
#
# In the IID setting the local data on each "client" is assumed to be a representative sample of the global data distribution. This is typically the case by construction when performing data parallel training of deep learning models across multiple CPU/GPU "clients".
#
# The non-IID case is significantly more complicated as there are many ways in which data can be non-IID and different degress of "non-IIDness". Consider a supervised task with features $X$ and labels $y$. A statistical model of federated learning involves two levels of sampling:
#
# 1. Sampling a client $i$ from the distribution over available clients $Q$
# 2. Sampling an example $(X,y)$ from that client’s local data distribution $P_i(X,y)$.
#
# Non-IID data in federated learning typically refers to differences between $P_i$ and $P_j$ for different clients $i$ and $j$. However, it is worth remembering that both the distribution of available clients, $Q$, and the distribution of local data for client $i$, $P_i$, may change over time which introduces another dimension of “non-IIDness”. Finally, if the local data on a client's device is insufficiently randomized, perhaps ordered by time, then independence is violated locally as well.
#
# In order to facilitate experimentation TFF includes federated versions of several popular datasets that exhibit different forms and degrees of non-IIDness.
# What datasets are available?
tff.simulation.datasets.
# This tutorial uses a version of MNIST that contains a version of the original NIST dataset that has been re-processed using [LEAF](https://leaf.cmu.edu/) so that the data is keyed by the original writer of the digits.
#
# The federated MNIST dataset displays a particular type of non-IIDness: feature distribution skew (covariate shift). Whith feature distribution skew the marginal distributions $P_i(X)$ vary across clients, even though $P(y|X)$ is shared. In the federated MNIST dataset users are writing the same numbers but each user has a different writing style characterized but different stroke width, slant, etc.
# +
# tff.simulation.datasets.emnist.load_data?
# -
emnist_train, emnist_test = (tff.simulation
.datasets
.emnist
.load_data(only_digits=True, cache_dir="../data"))
NUMBER_CLIENTS = len(emnist_train.client_ids)
NUMBER_CLIENTS
def sample_client_ids(client_ids: typing.List[str],
sample_size: typing.Union[float, int],
random_state: np.random.RandomState) -> typing.List[str]:
"""Randomly selects a subset of clients ids."""
number_clients = len(client_ids)
error_msg = "'client_ids' must be non-emtpy."
assert number_clients > 0, error_msg
if isinstance(sample_size, float):
error_msg = "Sample size must be between 0 and 1."
assert 0 <= sample_size <= 1, error_msg
size = int(sample_size * number_clients)
elif isinstance(sample_size, int):
error_msg = f"Sample size must be between 0 and {number_clients}."
assert 0 <= sample_size <= number_clients, error_msg
size = sample_size
else:
error_msg = "Type of 'sample_size' must be 'float' or 'int'."
raise TypeError(error_msg)
random_idxs = random_state.randint(number_clients, size=size)
return [client_ids[i] for i in random_idxs]
# these are what the client ids look like
_random_state = np.random.RandomState(42)
sample_client_ids(emnist_train.client_ids, 10, _random_state)
# +
def create_tf_datasets(source: tff.simulation.ClientData,
client_ids: typing.Union[None, typing.List[str]]) -> typing.Dict[str, tf.data.Dataset]:
"""Create tf.data.Dataset instances for clients using their client_id."""
if client_ids is None:
client_ids = source.client_ids
datasets = {client_id: source.create_tf_dataset_for_client(client_id) for client_id in client_ids}
return datasets
def sample_client_datasets(source: tff.simulation.ClientData,
sample_size: typing.Union[float, int],
random_state: np.random.RandomState) -> typing.Dict[str, tf.data.Dataset]:
"""Randomly selects a subset of client datasets."""
client_ids = sample_client_ids(source.client_ids, sample_size, random_state)
client_datasets = create_tf_datasets(source, client_ids)
return client_datasets
# +
_random_state = np.random.RandomState()
client_datasets = sample_client_datasets(emnist_train, sample_size=1, random_state=_random_state)
(client_id, client_dataset), *_ = client_datasets.items()
fig, axes = plt.subplots(1, 5, figsize=(12,6), sharex=True, sharey=True)
for i, example in enumerate(client_dataset.take(5)):
axes[i].imshow(example["pixels"].numpy(), cmap="gray")
axes[i].set_title(example["label"].numpy())
_ = fig.suptitle(x= 0.5, y=0.75, t=f"Training examples for a client {client_id}", fontsize=15)
# -
# ## Data preprocessing
#
# Since each client dataset is already a [`tf.data.Dataset`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset), preprocessing can be accomplished using Dataset transformations. Another option would be to use preprocessing operations from [`sklearn.preprocessing`](https://scikit-learn.org/stable/modules/preprocessing.html).
#
# Preprocessing consists of the following steps:
#
# 1. `map` a function that flattens the 28 x 28 images into 784-element tensors
# 2. `map` a function that rename the features from pixels and label to X and y for use with Keras
# 3. `shuffle` the individual examples
# 4. `batch` the into training batches
#
# We also throw in a `repeat` over the data set to run several epochs on each client device before sending parameters to the server for averaging.
# +
AUTOTUNE = (tf.data
.experimental
.AUTOTUNE)
SHUFFLE_BUFFER_SIZE = 1000
NUMBER_TRAINING_EPOCHS = 5 # number of local updates!
TRAINING_BATCH_SIZE = 32
TESTING_BATCH_SIZE = 32
NUMBER_FEATURES = 28 * 28
NUMBER_TARGETS = 10
# +
def _reshape(training_batch):
"""Extracts and reshapes data from a training sample """
pixels = training_batch["pixels"]
label = training_batch["label"]
X = tf.reshape(pixels, shape=[-1]) # flattens 2D pixels to 1D
y = tf.reshape(label, shape=[1])
return X, y
def create_training_dataset(client_dataset: tf.data.Dataset) -> tf.data.Dataset:
"""Create a training dataset for a client from a raw client dataset."""
training_dataset = (client_dataset.map(_reshape, num_parallel_calls=AUTOTUNE)
.shuffle(SHUFFLE_BUFFER_SIZE, seed=None, reshuffle_each_iteration=True)
.repeat(NUMBER_TRAINING_EPOCHS)
.batch(TRAINING_BATCH_SIZE)
.prefetch(buffer_size=AUTOTUNE))
return training_dataset
def create_testing_dataset(client_dataset: tf.data.Dataset) -> tf.data.Dataset:
"""Create a testing dataset for a client from a raw client dataset."""
testing_dataset = (client_dataset.map(_reshape, num_parallel_calls=AUTOTUNE)
.batch(TESTING_BATCH_SIZE))
return testing_dataset
# -
# ## How to choose the clients included in each training round
#
# In a typical federated training scenario there will be a very large population of user devices however only a fraction of these devices are likely to be available for training at a given point in time. For example, if the client devices are mobile phones then they might only participate in training when plugged into a power source, off a metered network, and otherwise idle.
#
# In a simulated environment, where all data is locally available, an approach is to simply sample a random subset of the clients to be involved in each round of training so that the subset of clients involved will vary from round to round.
#
# ### How many clients to include in each round?
#
# Updating and averaging a larger number of client models per training round yields better convergence and in a simulated training environment probably makes sense to include as many clients as is computationally feasible. However in real-world training scenario while averaging a larger number of clients improve convergence, it also makes training vulnerable to slowdown due to unpredictable tail delays in computation/communication at/with the clients.
def create_federated_data(training_source: tff.simulation.ClientData,
testing_source: tff.simulation.ClientData,
sample_size: typing.Union[float, int],
random_state: np.random.RandomState) -> typing.Dict[str, typing.Tuple[tf.data.Dataset, tf.data.Dataset]]:
# sample clients ids from the training dataset
client_ids = sample_client_ids(training_source.client_ids, sample_size, random_state)
federated_data = {}
for client_id in client_ids:
# create training dataset for the client
_tf_dataset = training_source.create_tf_dataset_for_client(client_id)
training_dataset = create_training_dataset(_tf_dataset)
# create the testing dataset for the client
_tf_dataset = testing_source.create_tf_dataset_for_client(client_id)
testing_dataset = create_testing_dataset(_tf_dataset)
federated_data[client_id] = (training_dataset, testing_dataset)
return federated_data
_random_state = np.random.RandomState(42)
federated_data = create_federated_data(emnist_train,
emnist_test,
sample_size=0.01,
random_state=_random_state)
# keys are client ids, values are (training_dataset, testing_dataset) pairs
len(federated_data)
# # Creating a model with Keras
#
# If you are using Keras, you likely already have code that constructs a Keras model. Since the model will need to be replicated on each of the client devices we wrap the model in a no-argument Python function, a representation of which, will eventually be invoked on each client to create the model on that client.
def create_keras_model_fn() -> keras.Model:
model_fn = keras.models.Sequential([
keras.layers.Input(shape=(NUMBER_FEATURES,)),
keras.layers.Dense(units=NUMBER_TARGETS),
keras.layers.Softmax(),
])
return model_fn
# In order to use any model with TFF, it needs to be wrapped in an instance of the [`tff.learning.Model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/Model) interface, which exposes methods to stamp the model's forward pass, metadata properties, etc, and also introduces additional elements such as ways to control the process of computing federated metrics.
#
# Once you have a Keras model like the one we've just defined above, you can have TFF wrap it for you by invoking [`tff.learning.from_keras_model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/from_keras_model), passing the model and a sample data batch as arguments, as shown below.
# +
# tff.learning.from_keras_model?
# -
def create_tff_model_fn() -> tff.learning.Model:
keras_model = create_keras_model_fn()
dummy_batch = (tf.constant(0.0, shape=(TRAINING_BATCH_SIZE, NUMBER_FEATURES), dtype=tf.float32),
tf.constant(0, shape=(TRAINING_BATCH_SIZE, 1), dtype=tf.int32))
loss_fn = (keras.losses
.SparseCategoricalCrossentropy())
metrics = [
keras.metrics.SparseCategoricalAccuracy()
]
tff_model_fn = (tff.learning
.from_keras_model(keras_model, dummy_batch, loss_fn, None, metrics))
return tff_model_fn
# Again, since our model will need to be replicated on each of the client devices we wrap the model in a no-argument Python function, a representation of which, will eventually be invoked on each client to create the model on that client.
# # Training the model on federated data
#
# Now that we have a model wrapped as `tff.learning.Model` for use with TFF, we can let TFF construct a Federated Averaging algorithm by invoking the helper function `tff.learning.build_federated_averaging_process` as follows.
#
# Keep in mind that the argument needs to be a constructor (such as `create_tff_model_fn` above), not an already-constructed instance, so that the construction of your model can happen in a context controlled by TFF.
#
# One critical note on the Federated Averaging algorithm below, there are 2 optimizers: a
#
# 1. `client_optimizer_fn` which is only used to compute local model updates on each client.
# 2. `server_optimizer_fn` applies the averaged update to the global model on the server.
#
# N.B. the choice of optimizer and learning rate may need to be different than those you would use to train the model on a standard i.i.d. dataset. Start with stochastic gradient descent with a smaller (than normal) learning rate.
# +
# tff.learning.build_federated_averaging_process?
# +
CLIENT_LEARNING_RATE = 1e-2
SERVER_LEARNING_RATE = 1e0
def create_client_optimizer(learning_rate: float = CLIENT_LEARNING_RATE,
momentum: float = 0.0,
nesterov: bool = False) -> keras.optimizers.Optimizer:
client_optimizer = (keras.optimizers
.SGD(learning_rate, momentum, nesterov))
return client_optimizer
def create_server_optimizer(learning_rate: float = SERVER_LEARNING_RATE,
momentum: float = 0.0,
nesterov: bool = False) -> keras.optimizers.Optimizer:
server_optimizer = (keras.optimizers
.SGD(learning_rate, momentum, nesterov))
return server_optimizer
federated_averaging_process = (tff.learning
.build_federated_averaging_process(create_tff_model_fn,
create_client_optimizer,
create_server_optimizer,
client_weight_fn=None,
stateful_delta_aggregate_fn=None,
stateful_model_broadcast_fn=None))
# -
# What just happened? TFF has constructed a pair of *federated computations* (i.e., programs in TFF's internal glue language) and packaged them into a [`tff.utils.IterativeProcess`](https://www.tensorflow.org/federated/api_docs/python/tff/utils/IterativeProcess) in which these computations are available as a pair of properties `initialize` and `next`.
#
# It is a goal of TFF to define computations in a way that they could be executed in real federated learning settings, but currently only local execution simulation runtime is implemented. To execute a computation in a simulator, you simply invoke it like a Python function. This default interpreted environment is not designed for high performance, but it will suffice for this tutorial.
#
# ## `initialize`
#
# A function that takes no arguments and returns the state of the federated averaging process on the server. This function is only called to initialize a federated averaging process after it has been created.
# () -> SERVER_STATE
print(federated_averaging_process.initialize.type_signature)
state = federated_averaging_process.initialize()
# ## `next`
#
# A function that takes current server state and federated data as arguments and returns the updated server state as well as any training metrics. Calling `next` performs a single round of federated averaging consisting of the following steps.
#
# 1. pushing the server state (including the model parameters) to the clients
# 2. on-device training on their local data
# 3. collecting and averaging model updates
# 4. producing a new updated model at the server.
# +
# extract the training datasets from the federated data
federated_training_data = [training_dataset for _, (training_dataset, _) in federated_data.items()]
# SERVER_STATE, FEDERATED_DATA -> SERVER_STATE, TRAINING_METRICS
state, metrics = federated_averaging_process.next(state, federated_training_data)
print(f"round: 0, metrics: {metrics}")
# -
# Let's run a few more rounds on the same training data (which will over-fit to a particular set of clients but will converge faster).
number_training_rounds = 15
for n in range(1, number_training_rounds):
state, metrics = federated_averaging_process.next(state, federated_training_data)
print(f"round:{n}, metrics:{metrics}")
# # First attempt at simulating federated averaging
#
# A proper federated averaging simulation would randomly sample new clients for each training round, allow for evaluation of training progress on training and testing data, and log training and testing metrics to TensorBoard for reference.
#
# Here we define a function that randomly sample new clients prior to each training round and logs training metrics TensorBoard. We defer handling testing data until we discuss federated evaluation towards the end of the tutorial.
def simulate_federated_averaging(federated_averaging_process: tff.utils.IterativeProcess,
training_source: tff.simulation.ClientData,
testing_source: tff.simulation.ClientData,
sample_size: typing.Union[float, int],
random_state: np.random.RandomState,
number_rounds: int,
initial_state: None = None,
tensorboard_logging_dir: str = None):
state = federated_averaging_process.initialize() if initial_state is None else initial_state
if tensorboard_logging_dir is not None:
if not os.path.isdir(tensorboard_logging_dir):
os.makedirs(tensorboard_logging_dir)
summary_writer = (tf.summary
.create_file_writer(tensorboard_logging_dir))
with summary_writer.as_default():
for n in range(number_rounds):
federated_data = create_federated_data(training_source,
testing_source,
sample_size,
random_state)
anonymized_training_data = [dataset for _, (dataset, _) in federated_data.items()]
state, metrics = federated_averaging_process.next(state, anonymized_training_data)
print(f"Round: {n}, Training metrics: {metrics}")
for name, value in metrics._asdict().items():
tf.summary.scalar(name, value, step=n)
else:
for n in range(number_rounds):
federated_data = create_federated_data(training_source,
testing_source,
sample_size,
random_state)
anonymized_training_data = [dataset for _, (dataset, _) in federated_data.items()]
state, metrics = federated_averaging_process.next(state, anonymized_training_data)
print(f"Round: {n}, Training metrics: {metrics}")
return state, metrics
federated_averaging_process = (tff.learning
.build_federated_averaging_process(create_tff_model_fn,
create_client_optimizer,
create_server_optimizer,
client_weight_fn=None,
stateful_delta_aggregate_fn=None,
stateful_model_broadcast_fn=None))
_random_state = np.random.RandomState(42)
_tensorboard_logging_dir = "../results/logs/tensorboard"
updated_state, current_metrics = simulate_federated_averaging(federated_averaging_process,
training_source=emnist_train,
testing_source=emnist_test,
sample_size=0.01,
random_state=_random_state,
number_rounds=5,
tensorboard_logging_dir=_tensorboard_logging_dir)
updated_state
current_metrics
# # Customizing the model implementation
#
# Keras is the recommended high-level model API for TensorFlow and you should be using Keras models and creating TFF models using [`tff.learning.from_keras_model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/from_keras_model) whenever possible.
#
# However, [`tff.learning`](https://www.tensorflow.org/federated/api_docs/python/tff/learning) provides a lower-level model interface, [`tff.learning.Model`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/Model), that exposes the minimal functionality necessary for using a model for federated learning. Directly implementing this interface (possibly still using building blocks from [`keras`](https://www.tensorflow.org/guide/keras)) allows for maximum customization without modifying the internals of the federated learning algorithms.
#
# Now we are going to repeat the above from scratch!
# ## Defining model variables
#
# We start by defining a new Python class that inherits from `tff.learning.Model`. In the class constructor (i.e., the `__init__` method) we will initialize all relevant variables using TF primatives as well as define the our "input spec" which defines the shape and types of the tensors that will hold input data.
# +
class MNISTModel(tff.learning.Model):
def __init__(self):
# initialize some trainable variables
self._weights = tf.Variable(
initial_value=lambda: tf.zeros(dtype=tf.float32, shape=(NUMBER_FEATURES, NUMBER_TARGETS)),
name="weights",
trainable=True
)
self._bias = tf.Variable(
initial_value=lambda: tf.zeros(dtype=tf.float32, shape=(NUMBER_TARGETS,)),
name="bias",
trainable=True
)
# initialize some variables used in computing metrics
self._number_examples = tf.Variable(0.0, name='number_examples', trainable=False)
self._total_loss = tf.Variable(0.0, name='total_loss', trainable=False)
self._number_true_positives = tf.Variable(0.0, name='number_true_positives', trainable=False)
# define the input spec
self._input_spec = collections.OrderedDict([
('X', tf.TensorSpec([None, NUMBER_FEATURES], tf.float32)),
('y', tf.TensorSpec([None, 1], tf.int32))
])
@property
def input_spec(self):
return self._input_spec
@property
def local_variables(self):
return [self._number_examples, self._total_loss, self._number_true_positives]
@property
def non_trainable_variables(self):
return []
@property
def trainable_variables(self):
return [self._weights, self._bias]
# -
# ## Defining the forward pass
#
# With the variables for model parameters and cumulative statistics in place we can now define the `forward_pass` method that computes loss, makes predictions, and updates the cumulative statistics for a single batch of input data.
class MNISTModel(tff.learning.Model):
def __init__(self):
# initialize some trainable variables
self._weights = tf.Variable(
initial_value=lambda: tf.zeros(dtype=tf.float32, shape=(NUMBER_FEATURES, NUMBER_TARGETS)),
name="weights",
trainable=True
)
self._bias = tf.Variable(
initial_value=lambda: tf.zeros(dtype=tf.float32, shape=(NUMBER_TARGETS,)),
name="bias",
trainable=True
)
# initialize some variables used in computing metrics
self._number_examples = tf.Variable(0.0, name='number_examples', trainable=False)
self._total_loss = tf.Variable(0.0, name='total_loss', trainable=False)
self._number_true_positives = tf.Variable(0.0, name='number_true_positives', trainable=False)
# define the input spec
self._input_spec = collections.OrderedDict([
('X', tf.TensorSpec([None, NUMBER_FEATURES], tf.float32)),
('y', tf.TensorSpec([None, 1], tf.int32))
])
@property
def input_spec(self):
return self._input_spec
@property
def local_variables(self):
return [self._number_examples, self._total_loss, self._number_true_positives]
@property
def non_trainable_variables(self):
return []
@property
def trainable_variables(self):
return [self._weights, self._bias]
@tf.function
def _count_true_positives(self, y_true, y_pred):
return tf.reduce_sum(tf.cast(tf.equal(y_true, y_pred), tf.float32))
@tf.function
def _linear_transformation(self, batch):
X = batch['X']
W, b = self.trainable_variables
Z = tf.matmul(X, W) + b
return Z
@tf.function
def _loss_fn(self, y_true, probabilities):
return -tf.reduce_mean(tf.reduce_sum(tf.one_hot(y_true, NUMBER_TARGETS) * tf.math.log(probabilities), axis=1))
@tf.function
def _model_fn(self, batch):
Z = self._linear_transformation(batch)
probabilities = tf.nn.softmax(Z)
return probabilities
@tf.function
def forward_pass(self, batch, training=True):
probabilities = self._model_fn(batch)
y_pred = tf.argmax(probabilities, axis=1, output_type=tf.int32)
y_true = tf.reshape(batch['y'], shape=[-1])
# compute local variables
loss = self._loss_fn(y_true, probabilities)
true_positives = self._count_true_positives(y_true, y_pred)
number_examples = tf.size(y_true, out_type=tf.float32)
# update local variables
self._total_loss.assign_add(loss)
self._number_true_positives.assign_add(true_positives)
self._number_examples.assign_add(number_examples)
batch_output = tff.learning.BatchOutput(
loss=loss,
predictions=y_pred,
num_examples=tf.cast(number_examples, tf.int32)
)
return batch_output
# ## Defining the local metrics
#
# Next, we define a method `report_local_outputs` that returns a set of local metrics. These are the values, in addition to model updates (which are handled automatically), that are eligible to be aggregated to the server in a federated learning or evaluation process.
#
# Finally, we need to determine how to aggregate the local metrics emitted by each device by defining `federated_output_computation`. This is the only part of the code that isn't written in TensorFlow - it's a federated computation expressed in TFF.
class MNISTModel(tff.learning.Model):
def __init__(self):
# initialize some trainable variables
self._weights = tf.Variable(
initial_value=lambda: tf.zeros(dtype=tf.float32, shape=(NUMBER_FEATURES, NUMBER_TARGETS)),
name="weights",
trainable=True
)
self._bias = tf.Variable(
initial_value=lambda: tf.zeros(dtype=tf.float32, shape=(NUMBER_TARGETS,)),
name="bias",
trainable=True
)
# initialize some variables used in computing metrics
self._number_examples = tf.Variable(0.0, name='number_examples', trainable=False)
self._total_loss = tf.Variable(0.0, name='total_loss', trainable=False)
self._number_true_positives = tf.Variable(0.0, name='number_true_positives', trainable=False)
# define the input spec
self._input_spec = collections.OrderedDict([
('X', tf.TensorSpec([None, NUMBER_FEATURES], tf.float32)),
('y', tf.TensorSpec([None, 1], tf.int32))
])
@property
def federated_output_computation(self):
return self._aggregate_metrics_across_clients
@property
def input_spec(self):
return self._input_spec
@property
def local_variables(self):
return [self._number_examples, self._total_loss, self._number_true_positives]
@property
def non_trainable_variables(self):
return []
@property
def trainable_variables(self):
return [self._weights, self._bias]
@tff.federated_computation
def _aggregate_metrics_across_clients(metrics):
aggregated_metrics = {
'number_examples': tff.federated_sum(metrics.number_examples),
'average_loss': tff.federated_mean(metrics.average_loss, metrics.number_examples),
'accuracy': tff.federated_mean(metrics.accuracy, metrics.number_examples)
}
return aggregated_metrics
@tf.function
def _count_true_positives(self, y_true, y_pred):
return tf.reduce_sum(tf.cast(tf.equal(y_true, y_pred), tf.float32))
@tf.function
def _linear_transformation(self, batch):
X = batch['X']
W, b = self.trainable_variables
Z = tf.matmul(X, W) + b
return Z
@tf.function
def _loss_fn(self, y_true, probabilities):
return -tf.reduce_mean(tf.reduce_sum(tf.one_hot(y_true, NUMBER_TARGETS) * tf.math.log(probabilities), axis=1))
@tf.function
def _model_fn(self, batch):
Z = self._linear_transformation(batch)
probabilities = tf.nn.softmax(Z)
return probabilities
@tf.function
def forward_pass(self, batch, training=True):
probabilities = self._model_fn(batch)
y_pred = tf.argmax(probabilities, axis=1, output_type=tf.int32)
y_true = tf.reshape(batch['y'], shape=[-1])
# compute local variables
loss = self._loss_fn(y_true, probabilities)
true_positives = self._count_true_positives(y_true, y_pred)
number_examples = tf.cast(tf.size(y_true), tf.float32)
# update local variables
self._total_loss.assign_add(loss)
self._number_true_positives.assign_add(true_positives)
self._number_examples.assign_add(number_examples)
batch_output = tff.learning.BatchOutput(
loss=loss,
predictions=y_pred,
num_examples=tf.cast(number_examples, tf.int32)
)
return batch_output
@tf.function
def report_local_outputs(self):
local_metrics = collections.OrderedDict([
('number_examples', self._number_examples),
('average_loss', self._total_loss / self._number_examples),
('accuracy', self._number_true_positives / self._number_examples)
])
return local_metrics
# Here are a few points worth highlighting:
#
# * All state that your model will use must be captured as TensorFlow variables, as TFF does not use Python at runtime (remember your code should be written such that it can be deployed to mobile devices).
# * Your model should describe what form of data it accepts (input_spec), as in general, TFF is a strongly-typed environment and wants to determine type signatures for all components. Declaring the format of your model's input is an essential part of it.
# * Although technically not required, we recommend wrapping all TensorFlow logic (forward pass, metric calculations, etc.) as tf.functions, as this helps ensure the TensorFlow can be serialized, and removes the need for explicit control dependencies.
#
# The above is sufficient for evaluation and algorithms like Federated SGD. However, for Federated Averaging, we need to specify how the model should train locally on each batch.
class MNISTrainableModel(MNISTModel, tff.learning.TrainableModel):
def __init__(self, optimizer):
super().__init__()
self._optimizer = optimizer
@tf.function
def train_on_batch(self, batch):
with tf.GradientTape() as tape:
output = self.forward_pass(batch)
gradients = tape.gradient(output.loss, self.trainable_variables)
self._optimizer.apply_gradients(zip(tf.nest.flatten(gradients), tf.nest.flatten(self.trainable_variables)))
return output
# # Simulating federated training with the new model
#
# With all the above in place, the remainder of the process looks like what we've seen already - just replace the model constructor with the constructor of our new model class, and use the two federated computations in the iterative process you created to cycle through training rounds.
# +
def create_custom_tff_model_fn():
optimizer = keras.optimizers.SGD(learning_rate=0.02)
return MNISTrainableModel(optimizer)
federated_averaging_process = (tff.learning
.build_federated_averaging_process(create_custom_tff_model_fn))
_random_state = np.random.RandomState(42)
updated_state, current_metrics = simulate_federated_averaging(federated_averaging_process,
training_source=emnist_train,
testing_source=emnist_test,
sample_size=0.01,
random_state=_random_state,
number_rounds=10)
# -
updated_state
current_metrics
# # Evaluation
#
# All of our experiments so far presented only federated training metrics - the average metrics over all batches of data trained across all clients in the round. Should we be concerened about overfitting? Yes! In federated averaging algorithms there are two different ways to over-fit.
#
# 1. Overfitting the shared model (especially if we use the same set of clients on each round).
# 2. Over-ftting local models on the clients.
#
# ## Federated evaluation
#
# To perform evaluation on federated data, you can construct another federated computation designed for just this purpose, using the [`tff.learning.build_federated_evaluation`](https://www.tensorflow.org/federated/api_docs/python/tff/learning/build_federated_evaluation) function, and passing in your model constructor as an argument. Note that evaluation doesn't perform gradient descent and there's no need to construct optimizers.
#
# +
# tff.learning.build_federated_evaluation?
# -
federated_evaluation = (tff.learning
.build_federated_evaluation(create_custom_tff_model_fn))
# function type signature: SERVER_MODEL, FEDERATED_DATA -> METRICS
print(federate_evaluation.type_signature)
# The `federated_evaluation` function is similar to `tff.utils.IterativeProcess.next` but with two important differences.
#
# 1. Function does not return the server state; since evaluation doesn't modify the model or any other aspect of state - you can think of it as stateless.
# 2. Function only needs the model and doesn't require any other part of server state that might be associated with training, such as optimizer variables.
training_metrics = federated_evaluation(updated_state.model, federated_training_data)
training_metrics
# Note the numbers may look marginally better than what was reported by the last round of training. By convention, the training metrics reported by the iterative training process generally reflect the performance of the model at the beginning of the training round, so the evaluation metrics will always be one step ahead.
# ## Evaluating on client data not used in training
#
# Since we are training a shared model for digit classication we might also want to evaluate the performance of the model on client test datasets where the corresponding training dataset was not used in training.
_random_state = np.random.RandomState(42)
client_datasets = sample_client_datasets(emnist_test, sample_size=0.01, random_state=_random_state)
federated_testing_data = [create_testing_dataset(client_dataset) for _, client_dataset in client_datasets.items()]
testing_metrics = federated_evaluation(updated_state.model, federated_testing_data)
testing_metrics
# # Adding evaluation to our federated averaging simulation
def simulate_federated_averaging(federated_averaging_process: tff.utils.IterativeProcess,
federated_evaluation,
training_source: tff.simulation.ClientData,
testing_source: tff.simulation.ClientData,
sample_size: typing.Union[float, int],
random_state: np.random.RandomState,
number_rounds: int,
tensorboard_logging_dir: str = None):
state = federated_averaging_process.initialize()
if tensorboard_logging_dir is not None:
if not os.path.isdir(tensorboard_logging_dir):
os.makedirs(tensorboard_logging_dir)
summary_writer = (tf.summary
.create_file_writer(tensorboard_logging_dir))
with summary_writer.as_default():
for n in range(number_rounds):
federated_data = create_federated_data(training_source,
testing_source,
sample_size,
random_state)
# extract the training and testing datasets
anonymized_training_data = []
anonymized_testing_data = []
for training_dataset, testing_dataset in federated_data.values():
anonymized_training_data.append(training_dataset)
anonymized_testing_data.append(testing_dataset)
state, _ = federated_averaging_process.next(state, anonymized_training_data)
training_metrics = federated_evaluation(state.model, anonymized_training_data)
testing_metrics = federated_evaluation(state.model, anonymized_testing_data)
print(f"Round: {n}, Training metrics: {training_metrics}, Testing metrics: {testing_metrics}")
# tensorboard logging
for name, value in training_metrics._asdict().items():
tf.summary.scalar(name, value, step=n)
for name, value in testing_metrics._asdict().items():
tf.summary.scalar(name, value, step=n)
else:
for n in range(number_rounds):
federated_data = create_federated_data(training_source,
testing_source,
sample_size,
random_state)
# extract the training and testing datasets
anonymized_training_data = []
anonymized_testing_data = []
for training_dataset, testing_dataset in federated_data.values():
anonymized_training_data.append(training_dataset)
anonymized_testing_data.append(testing_dataset)
state, _ = federated_averaging_process.next(state, anonymized_training_data)
training_metrics = federated_evaluation(state.model, anonymized_training_data)
testing_metrics = federated_evaluation(state.model, anonymized_testing_data)
print(f"Round: {n}, Training metrics: {training_metrics}, Testing metrics: {testing_metrics}")
return state, (training_metrics, testing_metrics)
# +
federated_averaging_process = (tff.learning
.build_federated_averaging_process(create_tff_model_fn,
create_client_optimizer,
create_server_optimizer,
client_weight_fn=None,
stateful_delta_aggregate_fn=None,
stateful_model_broadcast_fn=None))
federated_evaluation = (tff.learning
.build_federated_evaluation(create_tff_model_fn))
_random_state = np.random.RandomState(42)
updated_state, current_metrics = simulate_federated_averaging(federated_averaging_process,
federated_evaluation,
training_source=emnist_train,
testing_source=emnist_test,
sample_size=0.01,
random_state=_random_state,
number_rounds=15)
# -
# # Wrapping up
# ## Interesting resources
#
# [PySyft](https://github.com/OpenMined/PySyft) is a Python library for secure and private Deep Learning created by [OpenMined](https://www.openmined.org/). PySyft decouples private data from model training, using
# [Federated Learning](https://ai.googleblog.com/2017/04/federated-learning-collaborative.html),
# [Differential Privacy](https://en.wikipedia.org/wiki/Differential_privacy),
# and [Multi-Party Computation (MPC)](https://en.wikipedia.org/wiki/Secure_multi-party_computation) within the main Deep Learning frameworks like PyTorch and TensorFlow.
#
| notebooks/image-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ae (local)
# language: python
# metadata:
# is_prebuilt: false
# kernel_name: ae_local
# name: ae_local
# ---
# + [markdown] originalKey="480abf47-77be-4c72-819e-647600340428" showInput=true customInput code_folding=[] hidden_ranges=[]
# # Risk averse Bayesian optimization with environmental variables
#
# This notebook considers risk averse Bayesian optimization of objectives $f(x, w)$, where $x$ denotes the design variable and $w$ denotes the environmental variable.
# The design variable $x$ is fully controlled by the practitioner, however, the environmental variable $w$ is only controllable at the experimentation phase and is determined by the environment once the decision $x$ is implemented, according to some probability distribution.
# In this setting, with the $W$ denoting the random environmental variable, the objective we want to optimize becomes a random function, written as $f(x, W)$, whose value is determined only once the environmental variable $W$ is realized.
# This formulation is relevant whenever we need to make a decision to be implemented in an unknown future environment, and we can simulate the environment during the optimization phase.
#
# For this problem setting, [1] proposes to optimize a risk measure of the random function, written as $\rho[f(x, W)]$, where $\rho$ denotes a risk measure, which is a functional that maps a random variable (in this case $f(x, W)$ induced by $W$) to a real number.
# They propose the $\rho$KG acquisition function, which extends the well-known knowledge-gradient acquisition function, and requires access to posterior mean of the objective, i.e., $\mathbb{E}_n[\rho[f(x, W)]]$, where the expectation is taken over the sample paths of the GP model.
# Unlike the posterior mean of the function $f(x, w)$, the posterior mean of the risk measure is not available in closed-form and needs to be estimated via sampling.
# The procedure for estimating $\mathbb{E}_n[\rho[f(x, W)]]$ for a given $x$ is as follows:
# - Draw a set of `n_w` samples of $W$ according to the probability distribution. Let's call this `w_set`.
# - Append each $w$ in `w_set` to the given $x$ to get $(x, w)$ pairs. Note that for a single $x$, we now have `n_w` pairs of $(x, w)$.
# - Draw samples from the joint posterior distribution of these `n_w` pairs of $(x, w)$. Note that the joint distribution here is an `n_w`-dimensional Gaussian distribution.
# - Calculate the empirical risk measure corresponding to each sample, converting each `n_w`-dimensional posterior sample to a scalar sample of the risk measure.
# - Take the average of these risk measure samples to get the Monte-Carlo estimate of the posterior mean of the risk measure.
#
# Now that the background is established, we are ready to implement a one-shot version of the $\rho$KG acquisition function proposed in [1], in native BoTorch. We will:
# - Use `AppendFeatures` input transform to add the set of $W$ samples to each given $x$;
# - Calculate the joint posterior over these samples;
# - Use `RiskMeasureMCObjective` to convert these joint samples into samples of the risk measure;
# - And use the samples of the risk measure in `qMultiFidelityKnowledgeGradient` to define the $\rho$KG acquisition function.
#
# We will use the (negated) Branin function as $f(x, w)$ with the first input dimension denoting $x$ and the second input dimension denoting $w$, and find the $x$ maximizing the CVaR risk measure at risk level $\alpha=0.7$. We will assume that $W$ has a uniform distribution over $[0, 1]$ and approximate the risk measure using $16$ (qMC) samples of $W$ at a given time.
#
# CVaR, the Conditional Value-at-Risk, is a risk measure that measures the expectation of the worst outcomes (small rewards or large losses) with a total probability of $1 - \alpha$.
# It is commonly defined as the conditional expectation of the reward function, with the condition that the reward is smaller than the corresponding $1 - \alpha$ quantile.
#
# Note: Risk measures are typically studied in the context of a minimization problem (including in [1]), since it makes more sense to minimize "risk", and treat the larger values as being undesirable. Since the default behavior in BoTorch is to maximize the objective, the `RiskMeasureMCObjective` (and its subclasses) is defined w.r.t. the lower tail of the random variable, i.e., by treating the smaller values as undesirable. With this implementation, all that is needed to minimize a risk measure (of the original objective) is to negate the objective, as is done in this notebook.
#
# [1] [<NAME>, <NAME>, <NAME>, and <NAME>. Bayesian Optimization of Risk Measures. Advances in Neural Information Processing Systems 33, 2020.](https://arxiv.org/abs/2007.05554)
# + originalKey="3883d166-cec1-4027-8678-fac809a02e21" code_folding=[] hidden_ranges=[] requestMsgId="a44e7db8-06a5-405b-b2bd-0e73a3c2bf0b"
import os
import warnings
from time import time
import matplotlib.pyplot as plt
import torch
from botorch import fit_gpytorch_model
from botorch.acquisition import qMultiFidelityKnowledgeGradient, qSimpleRegret
from botorch.acquisition.risk_measures import CVaR
from botorch.models import SingleTaskGP
from botorch.models.transforms import Standardize
from botorch.models.transforms.input import AppendFeatures
from botorch.optim import optimize_acqf
from botorch.utils.sampling import draw_sobol_samples
from botorch.utils.transforms import unnormalize
from botorch.test_functions import Branin
from gpytorch import ExactMarginalLogLikelihood
from torch import Tensor
warnings.filterwarnings("ignore")
SMOKE_TEST = os.environ.get("SMOKE_TEST")
BATCH_SIZE = 2 if not SMOKE_TEST else 1
NUM_RESTARTS = 10 if not SMOKE_TEST else 2
RAW_SAMPLES = 128 if not SMOKE_TEST else 4
N_W = 16 if not SMOKE_TEST else 2
NUM_ITERATIONS = 20 if not SMOKE_TEST else 2
NUM_FANTASIES = 16 if not SMOKE_TEST else 2
tkwargs = {"device": "cpu", "dtype": torch.double}
# + [markdown] originalKey="<KEY>" showInput=true customInput code_folding=[] hidden_ranges=[]
# ## Problem setup
# We will initialize the `Branin` test function and define a wrapper around it to normalize the domain to $[0, 1]^2$.
# + originalKey="<KEY>" showInput=true customInput code_folding=[] hidden_ranges=[] requestMsgId="91d909e4-016a-4f3b-9efb-00a92cc9c580"
test_function = Branin(negate=True)
dim = test_function.dim
def evaluate_function(X: Tensor) -> Tensor:
return test_function(unnormalize(X, test_function.bounds)).view(*X.shape[:-1], 1)
# + [markdown] originalKey="3742d3d6-f77a-4013-9823-f797480c5d44" showInput=true customInput code_folding=[] hidden_ranges=[]
# ### Model initialization
# We will initialize the `SingleTaskGP` model on $8$ Sobol points drawn from the $(x, w)$ space.
# In doing so, we will also pass in the `AppendFeatures`. We will re-initialize `AppendFeatures` with a new `w_set` at every model training to ensure adequate coverage of the $W$ space.
# + originalKey="5a461471-35be-4c7e-9f1f-3c843288592e" showInput=true customInput code_folding=[] hidden_ranges=[] requestMsgId="e6435a8b-d26d-4949-96e5-b69661979e50"
bounds = torch.stack([torch.zeros(dim), torch.ones(dim)]).to(**tkwargs)
train_X = draw_sobol_samples(bounds=bounds, n=8, q=1).squeeze(-2).to(**tkwargs)
train_Y = evaluate_function(train_X)
def train_model(train_X: Tensor, train_Y: Tensor) -> SingleTaskGP:
r"""Returns a `SingleTaskGP` model trained on the inputs"""
w_set = draw_sobol_samples(n=N_W, q=1, bounds=bounds[:, -1:]).squeeze(-2).to(**tkwargs)
model = SingleTaskGP(
train_X, train_Y, input_transform=AppendFeatures(feature_set=w_set), outcome_transform=Standardize(m=1)
)
mll = ExactMarginalLogLikelihood(model.likelihood, model)
fit_gpytorch_model(mll)
return model
model = train_model(train_X, train_Y)
# + [markdown] originalKey="ffd3d6f1-32ed-496a-934b-e24868f41eae" showInput=true customInput code_folding=[] hidden_ranges=[]
# ### Define a helper function that performs the BO step
# The helper function will initialize the `qMultiFidelityKnowledgeGradient` acquisition function with the risk measure objective, and optimize it to find the candidate to evaluate.
# We use `qMultiFidelityKnowledgeGradient` instead of `qKnowledgeGraient` since it accepts a `project` callable, which we will use to ignore the $w$ present in the fantasy solutions before adding the `w_set` via the `AppendFeatures` input transform.
# + originalKey="<KEY>" showInput=true customInput code_folding=[] hidden_ranges=[] requestMsgId="b5910ef1-4634-4bb4-b72e-af5713d72f45"
risk_measure = CVaR(alpha=0.7, n_w=N_W)
def ignore_w(X: Tensor) -> Tensor:
r"""Remove `w` from the input."""
return X[..., :-1]
def optimize_rho_kg_and_get_observation():
r"""Optimizes the rhoKG acquisition function, and returns a new candidate and observation."""
acqf = qMultiFidelityKnowledgeGradient(
model=model,
num_fantasies=NUM_FANTASIES,
objective=risk_measure,
project=ignore_w,
)
candidate, _ = optimize_acqf(
acq_function=acqf,
bounds=bounds,
q=BATCH_SIZE,
num_restarts=NUM_RESTARTS,
raw_samples=RAW_SAMPLES,
)
new_observations = evaluate_function(candidate)
return candidate, new_observations
# + [markdown] originalKey="<KEY>" showInput=true customInput code_folding=[] hidden_ranges=[]
# ## Perform the Bayesian optimization loop with $\rho$KG
# The BO loop iterates the following steps:
# - Given the surrogate model, maximize the acquisition function to find the candidate(s) $(x, w)$ to evaluate;
# - Observe $f(x, w)$ for each candidate;
# - Update the surrogate model with the new observation.
#
# Note: Running this may take a while.
# + originalKey="e7151464-ac1d-48a6-8ffe-638d8c0f8a8c" showInput=true customInput code_folding=[] hidden_ranges=[] requestMsgId="372f60a2-efee-42ac-b2a7-1a08c23b5853"
start_time = time()
for i in range(NUM_ITERATIONS):
print(f"Starting iteration {i}, total time: {time() - start_time:.3f} seconds.")
# optimize the acquisition function and get the observations
candidate, observations = optimize_rho_kg_and_get_observation()
# update the model with new observations
train_X = torch.cat([train_X, candidate], dim=0)
train_Y = torch.cat([train_Y, observations], dim=0)
model = train_model(train_X, train_Y)
# + [markdown] originalKey="a3c58456-524d-4cfd-8619-3dab53e7d8ac" showInput=true customInput code_folding=[] hidden_ranges=[]
# ### Find the solution to implement
# We will choose the solution to implement as the point maximizing the posterior expectation of the risk measure. Since this expectation is not available in closed form, we will maximize its qMC estimate as a surrogate. We will use a larger `w_set` here to get a more precise estimate.
# + originalKey="09fa7c2b-6369-404e-928e-fb1913fe5215" showInput=true customInput code_folding=[] hidden_ranges=[] requestMsgId="f3b697a7-0ccf-4ba2-8195-cf00dbdb9cb6"
# update the input transform of the already trained model
w_set = draw_sobol_samples(n=128, q=1, bounds=bounds[:, -1:]).squeeze(-2).to(**tkwargs)
new_transform = AppendFeatures(feature_set=w_set).eval()
model.input_transform = new_transform
risk_measure = CVaR(alpha=0.7, n_w=128)
expected_risk_measure = qSimpleRegret(model=model, objective=risk_measure)
final_candidate, expected_objective = optimize_acqf(
acq_function=expected_risk_measure,
bounds=bounds[:, :1],
q=1,
num_restarts=NUM_RESTARTS,
raw_samples=RAW_SAMPLES,
)
# + [markdown] originalKey="649ac54a-bfe6-48f8-b33b-1dcad0609a26" showInput=true customInput code_folding=[] hidden_ranges=[]
# ### Let's plot the true risk measure and see how we did
# We can use the input transform and the risk measure we previously defined to make this part easier!
#
# The plot shows that we found the global optimal solution and that our estimate of the risk measure at the optimal point is quite accurate.
# + originalKey="<KEY>" showInput=true customInput code_folding=[] hidden_ranges=[] requestMsgId="29cef7c5-cb54-48c8-ba40-b07be775d672"
plot_x = torch.linspace(0, 1, 100, **tkwargs).view(-1, 1)
eval_X = new_transform(plot_x)
eval_Y = evaluate_function(eval_X)
plot_risk_measure = risk_measure(eval_Y)
plt.figure(figsize=(12, 8))
plt.title("True Risk Measure Objective and Solution Found")
plt.plot(plot_x, plot_risk_measure)
plt.scatter(final_candidate, expected_objective, marker="*", color="red", s=500)
plt.xlabel("x")
plt.ylabel("$\\rho[f(x, w)]$")
plt.show()
| tutorials/risk_averse_bo_with_environmental_variables.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.2
# language: julia
# name: julia-0.5
# ---
# ## Starter Problems
# #### Note: These 4 problems are from https://lectures.quantecon.org/jl/julia_by_example.html
# #### Strang Matrix Problem
#
# Use Julia's array and control flow syntax in order to define the NxN Strang matrix:
#
# $$ \left[\begin{array}{ccccc}
# -2 & 1\\
# 1 & -2 & 1\\
# & \ddots & \ddots & \ddots\\
# & & \ddots & \ddots & 1\\
# & & & 1 & -2
# \end{array}\right] $$
#
# i.e. a matrix with `-2` on the diagonal, 1 on the off-diagonals, and 0 elsewere.
# #### Factorial Problem*
#
# Using a `for` loop, write a function `my_factorial(n)` that computes the `n`th factorial. Try your function on integers like `15`, and then use `BigInt` inputs like `big(100)`. Make your function's output type match the input type for `n`.
# #### Binomial Problem*
#
# A random variable `X~Bin(n,p)` is defined the number of successes in `n` trials where each trial has a success probability `p`. For example, if `Bin(10,0.5)`, then `X` is the number of coin flips that turn up heads in `10` flips.
#
# Using only `rand()` (uniform random numbers), write a function `binomial_rv(n,p)` that produces one draw of `Bin(n,p)`.
# #### Monte Carlo $\pi$ Problem*
#
# Use random number generation to estimate $\pi$. To do so, mentally draw the unit circle. It is encompassed in the square $[-1,1]\times[-1,1]$. The area of the circle is $\pi r^2 = \pi$. The area of the square is $4$. Thus if points are randomly taken evenly from $[-1,1]\times[-1,1]$, then the probability they land in the circle ($x^2 + y^2\leq 1$) is $\frac{\pi}{4}$. Use this to estimate $\pi$.
# ## Integration Problems
#
# These problems integrate basic workflow tools to solve some standard data science and scientific computing problems.
# #### Timeseries Generation Problem*
#
# An AR1 timeseries is defined by
#
# $$ x_{t+1} = \alpha x_i + \epsilon_{t+1} $$
#
# where $x_0 = 0$ and $t=0,\ldots,T$. The shocks ${\epsilon_t}$ are i.i.d. standard normal (`N(0,1)`, given by `randn()`). Using $T=200$
#
# 1) $\alpha = 0$
# 2) $\alpha = 0.5$
# 3) $\alpha = 0.9$
#
# use Plots.jl to plot a timecourse for each of the parameters. Label the lines for the values of $\alpha$ that generate them using the `label` argument in `plot`.
# #### Regression Problem
# +
#### Prepare Data For Regression Problem
X = rand(1000, 3) # feature matrix
a0 = rand(3) # ground truths
y = X * a0 + 0.1 * randn(1000); # generate response
# Data For Regression Problem Part 2
X = rand(100);
y = 2X + 0.1 * randn(100);
# -
# Given an Nx3 array of data (`randn(N,3)`) and a Nx1 array of outcomes, produce the data matrix `X` which appends a column of 1's to the front of the data matrix, and solve for the 4x1 array `β` via `βX = b` using `qrfact`, or `\`, or [the definition of the OLS estimator](https://en.wikipedia.org/wiki/Ordinary_least_squares#Estimation). (Note: This is linear regression).
#
# Compare your results to that of using `llsq` from `MultivariateStats.jl` (note: you need to go find the documentation to find out how to use this!). Compare your results to that of using ordinary least squares regression from `GLM.jl`.
#
# #### Regression Problem Part 2
#
# Using your OLS estimator or one of the aforementioned packages, solve for the regression line using the (X,y) data above. Plot the (X,y) scatter plot using `scatter!` from Plots.jl. Add the regression line using `abline!`. Add a title saying "Regression Plot on Fake Data", and label the x and y axis.
# #### Logistic Map Problem
#
# The logistic difference equation is defined by the recursion
#
# $$ b_{n+1}=r*b_{n}(1-b_{n}) $$
#
# where $b_{n}$ is the number of bunnies at time $n$. Starting with $b_{0}=.25$, by around $400$ iterations this will reach a steady state. This steady state (or steady periodic state) is dependent on $r$. Write a function which plots the steady state attractor. This is done as follows:
#
# 1) Solve for the steady state(s) for each given $r$ (i.e. iterate the relation 400 times).
#
# 2) Calculate "every state" in the steady state attractor. This means, at steady state (after the first 400 iterations), save the next 150 values. Call this set of values $y_s(r)$.
#
# 3) Do steps (1) and (2) with $r\in\left(2.9,4\right)$, `dr=.001`. Plot $r$ x-axis vs $y_s(r)$=value seen in the attractor) using Plots.jl. Your result should be the [Logistic equation bifurcation diagram](https://upload.wikimedia.org/wikipedia/commons/7/7d/LogisticMap_BifurcationDiagram.png).
| Notebooks/BasicProblems.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''api_book'': venv)'
# name: python3
# ---
# # Building blocks of a computer
# All the creation of the ML models and model serving could not be possible without a computer. Ussualy, when a project starts and the initial meetings take place, a large part of the conversation is about what hardware to use and how much will it cost. In order to estimate the potential costs and having in mind the scope of the project, it is imperative to know at least on the high level what are the main components of the computer from the standpoint of an Machine Learning engineer. The main parts, along with the popular acronyms, are:
#
# * Disk space - SSD and HDD
# * Processors - CPU units
# * Random access memory - RAM
# * Operating systems
# # What is a computer?
# As per wiki {cite}`wiki:computers`:
#
# ***"A computer is a machine that can be programmed to carry out sequences of arithmetic or logical operations automatically."***
#
# The author <NAME> augments this definition with some of this thoughts:
#
# ***"Sending email and playing games are made possible by programs running on computers. The computer itself is like a newborn baby. It doesn’t really know how to do much. We hardly ever think about the basic machinery of human beings, because we mostly interact with the personalities that are running on that basic machinery, just like programs running on computers. For example, when you’re on a web page, you’re not reading it using just the computer itself; you’re reading it using programs someone else wrote that are running on your computer, the computer hosting the web page, and all of the computers in between that make the internet function."***
#
# Putting these definitions and thoughts in simpler terms, a computer is a machine with various hardware parts working together in order to run software programs.
#
# From a machine learning practitioner point of view, **a computer is a machine which uses disk space, processors, computer memory, graphical units and operating systems in order to serve machine learning models to the client**.
# # Main component of modern computers
# Computers would not be that usefull if we could not store some data into it and the access it later. The building blocks of computer memory are **bits**. The bit represents a logical state with one of two possible values most commonly denoted as **1 or 0**, **True or False**. How does a computer know when a bit has a value of 1 or 0? This is where transistors{cite}`transistors` come into play. A transistor is one of the most influential inventions in the modern history, enabling anyone to have a powerfull and cheap computers in their households, communications between computers and overall transistors enable the internal calculation logic inside the computers.
#
# **"You don’t need to know everything about the guts of transistors. The important thing to know is that a transistor is made on a substrate, or slab, of some semiconducting material, usually silicon. Unlike other technologies such as gears, valves, relays, and vacuum tubes, transistors aren’t individually manufactured objects. They’re made through a process called photolithography, which involves projecting a picture of a transistor onto a silicon wafer and developing it. This process is suitable for mass production because large numbers of transistors can be projected onto a single silicon wafer substrate, developed, and then sliced up into individual components."**{cite}`secret_life_computers`
#
# 
# One of the main uses of a transistor is that it can "hold" voltage. To simplify things and to build intuition, we can say that a transistor is "on" or has a state of "1" when there is voltage in it, and the transistor is "off" or it's state is "0" if there is no voltage in it.
#
# A single transistor would not be much usefull, because we could only store 1 bit of information in it. Modern computers have integrated circuits that have millions and more transistors inside of them.
#
# 
#
# This means that a single circuit can hold millions, billions and even more bits. These bits can be accessed by a computer in a billionths of a second. If you open up your desktop computer you would see alot of integrated circuits bunched up together which forms a certain architecture. But no matter how many circuits and transistors there are in a computer, no matter the marketing messages, all the modern computer computation and data storage logic depends on a very simple fact that a single transistor can be either "1" or "0".
# # Disk space
# In order to store files, data, models and to execute **scripts.py** files we need storage space inside a computer to put them in. Nowadays, there are two main types of hardware used for storage: **hard disk drives (HDD)** and **solid state drives (SSD)**.
# ## HDDs
# A typical hard disk drive looks like this:
#
# 
#
# The most important parts are the disk where the data is stored and the "needle" on top of the round disk which is used to extract the data. That needle is refered to as the **head**.
#
# **"It takes time to talk to memory. Imagine you had to go to the store every time you needed a cup of flour. It’s much more practical to go to the store once and bring home a whole sack of flour. Larger memory devices use this principle. Think warehouse shopping for bits. Disk drives, also known as mass storage, are great for storing immense amounts of data. \<...>\. They’re often referred to as mass storage. Some religious institutions use mass storage for their ceremonies in between use. Disk drives store bits on rotating magnetic platters, sort of like a lazy Susan. Bits periodically come around to where you’re sitting, and you use your hand to pluck them off or put them on.**
#
# **In a disk drive, your hand is replaced by the disk head. Disk drives are relatively slow compared to other types of memory. If you want something that just passed by the head, you have to wait almost an entire rotation for it to come around again. Modern disks spin at 7,200 rotations per minute (RPM), which means a rotation takes slightly longer than 8 milliseconds. The big problem with disk drives is that they’re mechanical and wear out. Bearing wear is one of the big causes of disk failure The difference between commercial and consumer-grade devices is primarily the amount of grease in the bearing—manufacturers are able to charge hundreds of dollars for something that costs less than a penny Disk drives store data by magnetizing areas on the disk, which makes them nonvolatile just like core memory. Disk drives are a trade-off between speed and density. They’re slow because of the time it takes for the bits you want to show up under the head, but because the data is being brought to the head, no space is required for address and data connections, unlike, for example, in a DRAM. They’re built in sealed containers because dust and dirt would cause them to fail."**{cite}`secret_life_computers`
# ## SSDs
# A solid state drive is a type of memory storing hardware that uses transistors and bites to store information. A typical SSD drive looks like this:
#
# 
#
# As one can see, an SSD has electronic circuits in it which are used to store data. Because there are no spinning parts and the memory can be accessed almost instantly, the SSD is becoming the more dominant technology in storing data.
#
# **"It works like RAM for reading and also for writing a blank device filled with 0s. But although 0s can be turned into 1s, they can’t be turned back without being erased first. Flash memory is internally divided into blocks, and only blocks can be erased, not individual locations. Flash memory devices are random-access for reads, and block-access for writes. Disk drives are slowly being replaced by solid-state disk drives, which are pretty much just flash memory packaged up to look like a disk drive. Right now their price per bit is much higher than spinning disks, but that’s expected to change. Because flash memory wears out, solid-state drives include a processor that keeps track of the usages in different blocks and tries to even it out so that all blocks wear out at the same rate."**{cite}`secret_life_computers`
#
# Acording to a research by <NAME>{cite}`wikibon` the SSDs will become more cheap in terms of cost per gigabyte by 2026:
#
# 
# # CPU
# A **Central Procesing Unit** in a computer gives out commands to all the other hardware inside of the computer. It also serves as an interaction point between a program (piece of software code) and the internal hardware. A typical CPU:
#
# 
#
# A CPU is made of billions of transistors which are bundled together into electronic circuits. Circuits bundled up with a processor form a **core**.
#
# A processor is a circuit that executes instructions from the programs to the hardware components and back.
#
# In the past, all the CPUs were single core units. Nowadays, the most common form of a CPU is a multi-core CPU:
#
# 
#
# All individual cores have an environment dedicated to them (memory, instructions, objects, etc.) and there is a shared memory space.
#
# To list the information about a CPU in an Ubuntu machine use the command:
#
# ```
# lscpu
# ```
#
# It should print out something similar:
#
# ```
# Architecture: x86_64
# CPU op-mode(s): 32-bit, 64-bit
# Byte Order: Little Endian
# Address sizes: 48 bits physical, 48 bits virtual
# CPU(s): 16
# On-line CPU(s) list: 0-15
# Thread(s) per core: 2
# Core(s) per socket: 8
# Socket(s): 1
# NUMA node(s): 1
# Vendor ID: AuthenticAMD
# CPU family: 25
# Model: 33
# Model name: AMD Ryzen 7 5800X 8-Core Processor
# ...
# ```
#
# The CPU with the above information has 16 cores and 32 **threads** in total (**2 Thread(s) per core** * **CPU(s)** = 16 * 2 = 32).
# ## Threads
# There are two types of threads in regards to a CPU: **hardware threads** and **software threads**.
#
# A hardware thred refers to the highest level of code (set of instructions) executed by a processor. Each CPU core has at least one thread, but nowadays, in terms of hardware, there are two threads per CPU. So, in the case of the example above, while having 16 physical cores on a chip, there are 16 additional "virtual" cores who can be used to give out instructions to programs. In practise, from a program point of view, a computer with 16 cores and 2 threads per core has 32 cores.
#
# Software threads are the smallest sequence of instructons that can be managed independently. Even if you have 32 hardware threads, you can have thousands of software threads. Software threads form units of **processes**. Each process can have many threads. For example, a command in Linux to show the CPU usage is **htop**:
#
# ```
# # Install first if not present with
# # apt-get install htop
#
# # Run the command
# htop
# ```
#
# A snapshot of a typical output:
#
# 
#
# The above output shows the physical cores available (16) and, along other things, it shows that the total number of active processes running (tasks) are **281** and those proceses have **1561** threads running. All the individual processes are listed in real time as well. Each process has a PID (process identification number) assigned to it.
# # RAM
# RAM is short for **Random Access Memory**. A typical hardware piece of RAM looks like this:
#
# 
#
# A RAM piece has transistors and circuits on it and the main usage for RAM is to almost instantaneously transfer information from CPU back and forth (hence the term access memory). The random part is due to the fact that the information is assigned to random **addresses** of memory storage.
#
# **"Memory is like a long street full of houses. Each house is exactly the same size and has room for a certain number of bits. Building codes have pretty much settled on 1 byte per house. And just like on a real street, each house has an address, which is just a number. If you have 64 MiB of memory in your computer, that’s 64 × 1,024 × 1,024 = 67,108,864 bytes (or 536,870,912 bits). The bytes have addresses from 0 to 67,108,863. This numbering makes sense, unlike the numbering on many real streets."**{cite}`secret_life_computers`
#
# Thus each byte of information has it's own address on a RAM board. If a place is already allocated, it means that the voltage in a certain transistors is not 0 and the place in RAM cannot be occupied by another piece of data. Bytes can only be stored to a place in RAM where the space is currently free (meaning - no voltage). If a computer is turned off, then all the information in RAM is lost.
#
# To check out the information about RAMs on a Linux machine use the command:
#
# ```
# free -h
# ```
#
# A general output of the command:
#
# ```
# total used free shared buff/cache available
# Mem: 62Gi 5,7Gi 53Gi 265Mi 3,2Gi 56Gi
# Swap: 2,0Gi 0B 2,0Gi
# ```
#
# The **Mem:** row indicates the memory available in the RAM hardware.
#
# The **Swap:** row indicates the available SSD space which is dedicated for random access. If all the RAM hardware is depleted, then CPU will write data to a part in SSD.
#
# The **total** collumn indicates the total amount of RAM available on a machine. This is the total amount of unique addresses which data can be stored for an instant access for any CPU proccesses.
#
# The **used** collumn indicates the currently used up memory in random access hardware.
#
# The **free** collumn indicates the total available memory which can be populated by processes running in the computer.
# # Putting all the pieces together
#
# The three major parts of a computer which strongly influence costs and are the main resources for running programs are:
#
# **Disk space** - either HHD or an SSD.
#
# **CPU** - amount of processes managed and their speed of execution.
#
# **RAM** - memory space for storing data created by various processes.
#
# 
#
# When a user initiates a python script with the command:
#
# ```
# python script.py
# ```
#
# There are many threads and processes that are sprung but the main parts are the following:
#
# 1) The process gets assigned a PID.
# 2) The CPU then finds the script in the dedicated space the script is in the computer file storage.
# 3) The python language deciphers the text in the script from human readable code to compiler readable instructions.
# 4) One of the cores and the CPU is then assigned to calculate all the instructions from the python script.
# 5) All while calculating, a portion of RAM is occupied by the objects created in the script and can be accessed very quickly.
# 6) After all the calculations, an output is presented to the user.
#
# All the intermediate steps (which can be hundreds or thousands) are separate threads that are assigned to the main PID.
# +
# The execution of this cell is a separate process and gets a unique PID. Every intermediate step is a software thread.
# Storing the data in RAM
a = 5
b = 10
# Creating another object in memory;
c = a + b
# Printing out the result to the user
print(f"The sum of {a} and {b} is: {c}")
# -
# Altough the above script in the cell is straightforward, it initiates hundreds of subprocesses in the computer hardware.
# # Contributions
#
# If you enjoyed the book so far and feel like donating, feel free to do so. The link to do a one time donation is [via Stripe](https://buy.stripe.com/14k17A6lQ8lAat2aEI).
#
# Additionaly, if you want me to add another chapter or to expand an existing one, please create an issue on [Github](https://github.com/Eligijus112/api-book).
| api-book/_build/html/_sources/chapter-1-computer/computer-specs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
# %matplotlib inline
import pprint
pp = pprint.PrettyPrinter(indent=4)
# +
import nltk
from nltk.sentiment.vader import SentimentIntensityAnalyzer
nltk.download('vader_lexicon')
analyzer = SentimentIntensityAnalyzer()
# -
# # News Headlines Sentiment
#
# Use the news api to pull the latest news articles for bitcoin and ethereum and create a DataFrame of sentiment scores for each coin.
#
# Use descriptive statistics to answer the following questions:
# 1. Which coin had the highest mean positive score?
# 2. Which coin had the highest negative score?
# 3. Which coin had the highest positive score?
# Read your api key environment variable
from newsapi import NewsApiClient
newsapi = NewsApiClient(api_key=os.getenv('NEWS_API'))
print(newsapi)
# + [markdown] jupyter={"outputs_hidden": true}
# # Lotsa Functions
# 1. get_sentiment_scores
# 2. get_sentiments_on_topic
# 3. sentiment_to_df
# -
# Fetch the Bitcoin news articles
def get_sentiment_scores(text, date, source, url):
sentiment_scores = {}
# Sentiment scoring with VADER
text_sentiment = analyzer.polarity_scores(text)
sentiment_scores["date"] = date
sentiment_scores["text"] = text
sentiment_scores["source"] = source
sentiment_scores["url"] = url
sentiment_scores["compound"] = text_sentiment["compound"]
sentiment_scores["positive"] = text_sentiment["pos"]
sentiment_scores["neutral"] = text_sentiment["neu"]
sentiment_scores["negative"] = text_sentiment["neg"]
if text_sentiment["compound"] >= 0.05: # Positive
sentiment_scores["normalized"] = 1
elif text_sentiment["compound"] <= -0.05: # Negative
sentiment_scores["normalized"] = -1
else:
sentiment_scores["normalized"] = 0 # Neutral
return sentiment_scores
def get_sentiments_on_topic(topic):
""" We loke documentation"""
sentiments_data = []
# Loop through all the news articles
for article in newsapi.get_everything(q=topic, language="en", page_size=100)["articles"]:
try:
# Get sentiment scoring using the get_sentiment_score() function
sentiments_data.append(
get_sentiment_scores(
article["content"],
article["publishedAt"][:10],
article["source"]["name"],
article["url"],
)
)
except AttributeError:
pass
return sentiments_data
def sentiment_to_df(sentiments):
# Create a DataFrame with the news articles' data and their sentiment scoring results
news_df = pd.DataFrame(sentiments)
# Sort the DataFrame rows by date
news_df = news_df.sort_values(by="date")
# Define the date column as the DataFrame's index
news_df.set_index("date", inplace=True)
return news_df
# # Use the functions
# +
topics = ['Bitcoin', 'Ethereum']
btc_sentiment = get_sentiments_on_topic(topics[0])
btc_df = sentiment_to_df(btc_sentiment)
display(btc_df.head())
display("btc_df.describe()")
display(btc_df.describe())
# -
eth_sentiment = get_sentiments_on_topic(topics[1])
eth_df = sentiment_to_df(eth_sentiment)
display(eth_df.head())
display(eth_df.describe())
# ### Questions:
#
# Q: Which coin had the highest mean positive score?
#
# A: **BTC**
#
# Q: Which coin had the highest compound score?
#
# A: **BTC**
#
# Q. Which coin had the highest positive score?
#
# A: **BTC**
# ---
# # Tokenizer
#
# In this section, you will use NLTK and Python to tokenize the text for each coin. Be sure to:
# 1. Lowercase each word
# 2. Remove Punctuation
# 3. Remove Stopwords
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer, PorterStemmer
from string import punctuation
import re
# Expand the default stop words list if necessary
nltk.download('wordnet')
# +
# Complete the tokenizer function
def tokenizer(text):
"""returns a list of words that is lemmatized, stopworded, tokenized, and free of any non-letter characters. """
# Create a list of the words
# Convert the words to lowercase
# Remove the punctuation
# Remove the stop words
# Lemmatize Words into root words
lemmatizer = WordNetLemmatizer()
sw = set(stopwords.words('english'))
regex = re.compile("[^a-zA-Z ]")
re_clean = regex.sub('', text)
words = word_tokenize(re_clean)
return [lemmatizer.lemmatize(word.lower()) for word in words if word.lower() not in set(stopwords.words('english'))]
# -
# Create a new tokens column for bitcoin
btc_df["tokens"] = btc_df["text"].apply(tokenizer)
btc_df.head()
# Create a new tokens column for ethereum
eth_df["tokens"] = eth_df["text"].apply(tokenizer)
eth_df.head()
# ---
# # NGrams and Frequency Analysis
#
# In this section you will look at the ngrams and word frequency for each coin.
#
# 1. Use NLTK to produce the n-grams for N = 2.
# 2. List the top 10 words for each coin.
from collections import Counter
from nltk import ngrams
# Generate the Bitcoin N-grams where N=2
flat_btc_tokens = [item for sublist in btc_df.tokens.to_list() for item in sublist]
bigram_counts = Counter(ngrams(flat_btc_tokens, n=2))
bigram_counts.most_common(20)
# Generate the Ethereum N-grams where N=2
flat_eth_tokens = [item for sublist in eth_df.tokens.to_list() for item in sublist]
eth_bigram_counts = Counter(ngrams(flat_eth_tokens, n=2))
eth_bigram_counts.most_common(20)
# Use the token_count function to generate the top 10 words from each coin
def token_count(tokens, N=10):
"""Returns the top N tokens from the frequency count"""
return Counter(tokens).most_common(N)
# Get the top 10 words for Bitcoin
token_count(flat_btc_tokens)
# Get the top 10 words for Ethereum
token_count(flat_eth_tokens)
# # Word Clouds
#
# In this section, you will generate word clouds for each coin to summarize the news for each coin
from wordcloud import WordCloud
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = [20.0, 10.0]
# Generate the Bitcoin word cloud
wordcloud = WordCloud(colormap="RdYlBu").generate(" ".join(flat_btc_tokens))
plt.imshow(wordcloud)
plt.axis("off")
fontdict = {"fontsize": 50, "fontweight": "bold"}
plt.title("Bitcoin Word Cloud", fontdict=fontdict)
plt.show()
# Generate the Ethereum word cloud
wordcloud = WordCloud(colormap="RdYlBu").generate(" ".join(flat_eth_tokens))
plt.imshow(wordcloud)
plt.axis("off")
fontdict = {"fontsize": 50, "fontweight": "bold"}
plt.title("Bitcoin Word Cloud", fontdict=fontdict)
plt.show()
# # Named Entity Recognition
#
# In this section, you will build a named entity recognition model for both coins and visualize the tags using SpaCy.
import spacy
from spacy import displacy
# +
# Optional - download a language model for SpaCy
# # !python -m spacy download en_core_web_sm
# -
# Load the spaCy model
nlp = spacy.load('en_core_web_sm')
# ## Bitcoin NER
# Concatenate all of the bitcoin text together
flat_btc_text_str = " ".join(btc_df.text.to_list())
print(flat_btc_text_str)
# Run the NER processor on all of the text
btc_doc = nlp(flat_btc_text_str)
btc_doc.user_data["title"] = "Bitcoin NER"
# Add a title to the document
# Render the visualization
displacy.render(btc_doc,style='ent')
# List all Entities
for ent in btc_doc.ents:
print(ent.text, ent.label_)
# ---
# ## Ethereum NER
# Concatenate all of the bitcoin text together
#
# https://spacy.io/usage/visualizers
# colors = {"ORG": "linear-gradient(90deg, #aa9cfc, #fc9ce7)"}
# options = {"ents": ["ORG"], "colors": colors}
# displacy.serve(doc, style="ent", options=options)
#
flat_eth_text_str = " ".join(eth_df.text.to_list())
#print(flat_eth_text_str)
eth_doc = nlp(flat_eth_text_str)
eth_doc.user_data["title"] = "Ethereum NER"
colors = {"ORG": "linear-gradient(90deg, #aa9cfc, #fc9ce7)"}
options = {"ents": ["ORG"], "colors": colors}
displacy.render(eth_doc,style='ent', options=options)
# +
# Run the NER processor on all of the text
# Add a title to the document
# List all Entities
for ent in eth_doc.ents:
print(ent.text, ent.label_)
# -
| Starter_Code/crypto_sentiment.ipynb |