code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="t8dgI_OxP9OE" colab_type="text"
# #`This notebook is an Illustration to implement Linear Discriminant Algorithm and Quadratic Discriminant Algorithm with Sklearn using Python🐍📓`
#
# For more information, check out the [LDA](https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.LinearDiscriminantAnalysis.html#sklearn.discriminant_analysis.LinearDiscriminantAnalysis) & [QDA](https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.QuadraticDiscriminantAnalysis.html#sklearn.discriminant_analysis.QuadraticDiscriminantAnalysis)
# + [markdown] id="EMJ-GphnQj16" colab_type="text"
# ## **Setup**
# Let's setup the imports:
# + id="4Kjc9AriD6vp" colab_type="code" colab={}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
sns.set_style()
from matplotlib.colors import ListedColormap
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
# custom color map
cm_dark = ListedColormap(['#ff6060', '#8282ff','#ffaa00','#fff244','#4df9b9','#76e8fc','#3ad628'])
# + [markdown] id="rKNqK6ANQlCp" colab_type="text"
# Let's load the Iris dataset which is in-build in Seaborn and do some preprocessing:
# + id="9TKbw_UsD6tI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="b6e74bbf-847e-4da7-926b-cffd7988574e"
df = sns.load_dataset('iris')
df = df[df['species'] != 'setosa']
col = ['petal_length', 'petal_width']
X = df.loc[:, col]
species_to_num = {'versicolor': 0, 'virginica': 1}
df['tmp'] = df['species'].map(species_to_num)
y = df['tmp']
X.head()
# + [markdown] id="fpEt-hNtQwTe" colab_type="text"
# ## LINEAR DISCRIMINANT ALGORITHM
# + id="4EyFj6XVD6oI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="c9e6a7fd-053d-484e-85a2-88b347643882"
clf = LinearDiscriminantAnalysis(shrinkage=0.2, solver='eigen')
clf.fit(X, y)
# + id="3esz1v2dD6lv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="8e3cad6c-bee9-4b0e-a3ec-b0d539186253"
Xv = X.values.reshape(-1,1)
h = 0.02
x_min, x_max = Xv.min(), Xv.max() + 1
y_min, y_max = y.min(), y.max() + 2
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
z = z.reshape(xx.shape)
fig = plt.figure(figsize=(7,6))
ax = plt.pcolormesh(xx, yy, z, cmap = cm_dark, alpha=0.5);
plt.scatter(X.values[:, 0], X.values[:, 1], c=y, s=100,
alpha=0.9, edgecolors='k');
plt.title("Linear Discriminant Analysis Of Iris Dataset", fontweight='bold');
# + [markdown] id="AvTvNFJdQ1-b" colab_type="text"
# ## QUADRATIC DISCRIMINANT ALGORITHM
# + id="VbnSsEU3H8cA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="e314d81e-2aa0-4976-b160-f7f41f155389"
clf1 = QuadraticDiscriminantAnalysis(store_covariance=False)
clf1.fit(X, y)
# + id="BUhBfxU1IGf8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="6032bd69-8a2c-479f-ada8-5326737163e9"
Xv = X.values.reshape(-1,1)
h = 0.02
x_min, x_max = Xv.min(), Xv.max() + 1
y_min, y_max = y.min(), y.max() + 2
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
z = clf1.predict(np.c_[xx.ravel(), yy.ravel()])
z = z.reshape(xx.shape)
fig = plt.figure(figsize=(7,6))
ax = plt.pcolormesh(xx, yy, z, cmap = cm_dark, alpha=0.3);
plt.scatter(X.values[:, 0], X.values[:, 1], c=y, s=70,
alpha=0.9, edgecolors='k');
plt.title("Quadratic Discriminant Analysis Of Iris Dataset", fontweight='bold');
# + [markdown] id="2DzIPs-fap4b" colab_type="text"
# ## Summary
#
# - You learned how to implement LDA and QDA
# - Visualize the Gaussian Distribution for each class
#
# + [markdown] id="3l-7J41LaqFh" colab_type="text"
# ## Reference
# - [LDA](https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.LinearDiscriminantAnalysis.html#sklearn.discriminant_analysis.LinearDiscriminantAnalysis)
# - [QDA](https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.QuadraticDiscriminantAnalysis.html#sklearn.discriminant_analysis.QuadraticDiscriminantAnalysis)
| LDA_QDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn import linear_model
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
matplotlib.style.use('ggplot')
# +
data = {'x1' : [158,158,158,158,158,158,158,160,160,160,160,160,160,160,162,162,162,162,162,162,162,162,164,164,164,164,164,164,164,164,166,166,166,166,166,166,166,166,166,166,166,168,168,168,168,168,168,168,168,168,168,168,170,170,170,170,170,170,170,170,170,170,170,170,170,170,172,172,172,172,172,172,172,172,172,172,172,172,172,174,174,174,174,174,174,174,174,174,174,174,174,174,176,176,176,176,176,176,176,176,176,176,176,176,176,176,178,178,178,178,178,178,178,178,178,178,178,178,178,178,180,180,180,180,180,180,180,180,180,180,180,180,182,182,182,182,182,182,182,182,182,184,184,184,184,184,184,184,184,184],
'x2' : [56,58,60,62,64,66,68,56,58,60,62,64,66,68,56,58,60,62,64,66,68,70,56,58,60,62,64,66,68,70,56,58,60,62,64,66,68,70,72,74,76,56,58,60,62,64,66,68,70,72,74,76,56,58,60,62,64,66,68,70,72,74,76,78,80,82,56,58,60,62,64,66,68,70,72,74,76,78,80,56,58,60,62,64,66,68,70,72,74,76,78,80,56,58,60,62,64,66,68,70,72,74,76,78,80,82,56,58,60,62,64,66,68,70,72,74,76,78,80,82,60,62,64,66,68,70,72,74,76,78,80,82,66,68,70,72,74,76,78,80,82,66,68,70,72,74,76,78,80,82],
'y' : [90,90,95,95,95,95,100,95,95,95,95,95,95,100,95,95,95,95,95,95,100,100,95,95,95,95,95,95,95,95,95,95,95,95,95,95,95,100,100,100,100,95,95,95,95,100,100,100,100,100,100,100,95,95,95,100,100,100,100,100,100,100,100,105,105,105,100,100,100,100,100,100,100,100,100,100,105,105,105,100,100,100,100,100,100,100,100,100,100,105,105,105,100,100,100,100,100,100,100,100,100,100,105,105,105,110,100,100,100,105,105,105,105,105,105,105,105,105,110,110,105,105,105,105,105,105,105,105,105,105,110,110,105,105,105,105,105,105,110,110,110,105,105,105,105,105,105,110,110,110]}
# data 변수 안에 dictionary형태의 x1, x2, y key 이름을 가진 데이터를 생성한다.
data = pd.DataFrame(data)
# data변수 안에 저장된 데이터를 pandas.DataFrame 함수를 통해 2차원의 수정 가능한 테이블 형태의 구조로 변경 후 data 변수에 저장
X = data[['x1', 'x2']]
# 독립변수들을 따로 변수에 저장. "data"데이터 프레임 안에 독립변수 "x1"과 "x2"를 "X"라는 변수에 저장.
y = data['y']
data
# -
linear_regression = linear_model.LinearRegression()
linear_regression.fit(X = pd.DataFrame(X), y = y)
prediction = linear_regression.predict(X = pd.DataFrame(X))
a = linear_regression.intercept_
b = linear_regression.coef_
print('a value = ', linear_regression.intercept_)
print('b value = ', linear_regression.coef_)
value = a+(166.13*b[0] + 56.8*b[1])
print(value)
residuals = y-prediction
residuals.describe()
from sklearn.metrics import mean_squared_error
print('score = ', linear_regression.score(X =pd.DataFrame(X), y=y))
print('Mean_Squared_Error = ', mean_squared_error(prediction, y))
print('RMSE = ', mean_squared_error(prediction, y)**0.5)
| top size.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# HIDDEN
from datascience import *
import matplotlib
matplotlib.use('Agg', warn=False)
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import numpy as np
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import nbinteract as nbi
# -
# ### Sampling from a Population ###
#
# The law of averages also holds when the random sample is drawn from individuals in a large population.
#
# As an example, we will study a population of flight delay times. The table `united` contains data for United Airlines domestic flights departing from San Francisco in the summer of 2015. The data are made publicly available by the [Bureau of Transportation Statistics](http://www.transtats.bts.gov/Fields.asp?Table_ID=293) in the United States Department of Transportation.
#
# There are 13,825 rows, each corresponding to a flight. The columns are the date of the flight, the flight number, the destination airport code, and the departure delay time in minutes. Some delay times are negative; those flights left early.
united = Table.read_table('http://inferentialthinking.com/notebooks/united_summer2015.csv')
united
# One flight departed 16 minutes early, and one was 580 minutes late. The other delay times were almost all between -10 minutes and 200 minutes, as the histogram below shows.
united.column('Delay').min()
united.column('Delay').max()
# +
delay_opts = {
'xlabel': 'Delay (minute)',
'ylabel': 'Percent per minute',
'xlim': (-20, 600),
'ylim': (0, 0.045),
'bins': 62,
}
nbi.hist(united.column('Delay'), options=delay_opts)
# -
# For the purposes of this section, it is enough to zoom in on the bulk of the data and ignore the 0.8% of flights that had delays of more than 200 minutes. This restriction is just for visual convenience; the table still retains all the data.
united.where('Delay', are.above(200)).num_rows/united.num_rows
# +
delay_opts = {
'xlabel': 'Delay (minute)',
'ylabel': 'Percent per minute',
'xlim': (-20, 200),
'ylim': (0, 0.045),
'bins': 22,
}
nbi.hist(united.column('Delay'), options=delay_opts)
# -
# The height of the [0, 10) bar is just under 3% per minute, which means that just under 30% of the flights had delays between 0 and 10 minutes. That is confirmed by counting rows:
united.where('Delay', are.between(0, 10)).num_rows/united.num_rows
# ### Empirical Distribution of the Sample ###
#
# Let us now think of the 13,825 flights as a population, and draw random samples from it with replacement. It is helpful to package our analysis code into a function. The function `empirical_delays` takes the sample size as its argument and returns the array of sampled flight delays.
def empirical_hist_delay(sample_size):
return united.sample(sample_size).column('Delay')
# As we saw with the dice, as the sample size increases, the empirical histogram of the sample more closely resembles the histogram of the population. Compare these histograms to the population histogram above.
nbi.hist(empirical_hist_delay,
options=delay_opts,
sample_size=widgets.ToggleButtons(options=[10, 100, 1000, 10000], description='Sample Size: '))
# The most consistently visible discrepancies are among the values that are rare in the population. In our example, those values are in the the right hand tail of the distribution. But as the sample size increases, even those values begin to appear in the sample in roughly the correct proportions.
# ### Convergence of the Empirical Histogram of the Sample ###
# What we have observed in this section can be summarized as follows:
#
# For a large random sample, the empirical histogram of the sample resembles the histogram of the population, with high probability.
#
# This justifies the use of large random samples in statistical inference. The idea is that since a large random sample is likely to resemble the population from which it is drawn, quantities computed from the values in the sample are likely to be close to the corresponding quantities in the population.
| packages/nbinteract-core/example-notebooks/examples_sampling_from_a_population.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 7: 48 Crayons
# Let's review where we've been so far:
#
# * We got a basic neural network up and running and trained it to predict the name of a few colors.
# * We added more colors until we ran into problems with the training taking too many repetitions, *epochs*, and not getting much better after awhile.
# * So we *perturbed* our data, wiggling the color values around a little bit and calling the new values the same color as the original. This gave the network lots more examples - which networks typically like a lot! - and helped us converge to a lower loss in only a few epochs, but at the cost of each epoch taking a lot of time.
#
# We've not discussed exactly what the network does inside. That's a lot of detail that we will start to get into soon.
#
# In the meantime, as promised last time, it's time to try doubling our crayon colors from 24 to 48. Let's give it a go and see where things end up.
#
# Hint: There are two new changes in this code from last time. See if you can spot them; we'll use them below.
# +
from keras.layers import Activation, Dense, Dropout
from keras.models import Sequential
import keras.optimizers, keras.utils, numpy
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelBinarizer
def train(rgbValues, colorNames, epochs = 16, perceptronsPerColorName = 8, batchSize = 1):
"""
Trains a neural network to understand how to map color names to RGB triples.
The provided lists of RGB triples must be floating point triples with each
value in the range [0.0, 1.0], and the number of color names must be the same length.
Different names are allowed to map to the same RGB triple.
Returns a trained model that can be used for recognize().
"""
# Convert the Python map RGB values into a numpy array needed for training.
rgbNumpyArray = numpy.array(rgbValues, numpy.float64)
# Convert the color labels into a one-hot feature array.
# Text labels for each array position are in the classes_ list on the binarizer.
labelBinarizer = LabelBinarizer()
oneHotLabels = labelBinarizer.fit_transform(colorNames)
numColors = len(labelBinarizer.classes_)
colorLabels = labelBinarizer.classes_
# Hyperparameters to define the network shape.
numFullyConnectedPerceptrons = numColors * perceptronsPerColorName
model = Sequential([
# Layer 1: Fully connected layer with ReLU activation.
Dense(numFullyConnectedPerceptrons, activation='relu', kernel_initializer='TruncatedNormal', input_shape=(3,)),
# Outputs: SoftMax activation to get probabilities by color.
Dense(numColors, activation='softmax')
])
print(model.summary())
# Compile for categorization.
model.compile(
optimizer = keras.optimizers.SGD(lr = 0.01, momentum = 0.9, decay = 1e-6, nesterov = True),
loss = 'categorical_crossentropy',
metrics = [ 'accuracy' ])
history = model.fit(rgbNumpyArray, oneHotLabels, epochs=epochs, batch_size=batchSize)
return (model, colorLabels)
def createMoreTrainingData(colorNameToRGBMap):
# The incoming color map is not typically going to be oversubscribed with e.g.
# extra 'red' samples pointing to slightly different colors. We generate a
# training dataset by perturbing each color by a small amount positive and
# negative. We do this for each color individually, by pairs, and for all three
# at once, for each positive and negative value, resulting in dataset that is
# many times as large.
perturbValues = [ 0.0, 0.01, 0.02, 0.03 ] # TODO: Experiment with adding 0.04, 0.05
rgbValues = []
labels = []
for colorName, rgb in colorNameToRGBMap.items():
reds = []
greens = []
blues = []
for perturb in perturbValues:
if rgb[0] + perturb <= 1.0:
reds.append(rgb[0] + perturb)
if perturb != 0.0 and rgb[0] - perturb >= 0.0:
reds.append(rgb[0] - perturb)
if rgb[1] + perturb <= 1.0:
greens.append(rgb[1] + perturb)
if perturb != 0.0 and rgb[1] - perturb >= 0.0:
greens.append(rgb[1] - perturb)
if rgb[2] + perturb <= 1.0:
blues.append(rgb[2] + perturb)
if perturb != 0.0 and rgb[2] - perturb >= 0.0:
blues.append(rgb[2] - perturb)
for red in reds:
for green in greens:
for blue in blues:
rgbValues.append((red, green, blue))
labels.append(colorName)
return (rgbValues, labels)
# -
# And then our newly expanded color list, and training a small number of epochs:
# +
def rgbToFloat(r, g, b): # r, g, b in 0-255 range
return (float(r) / 255.0, float(g) / 255.0, float(b) / 255.0)
# http://www.jennyscrayoncollection.com/2017/10/complete-list-of-current-crayola-crayon.html
colorMap = {
# 8-crayon box colors
'red': rgbToFloat(238, 32, 77),
'yellow': rgbToFloat(252, 232, 131),
'blue': rgbToFloat(31, 117, 254),
'brown': rgbToFloat(180, 103, 77),
'orange': rgbToFloat(255, 117, 56),
'green': rgbToFloat(28, 172, 20),
'violet': rgbToFloat(146, 110, 174),
'black': rgbToFloat(35, 35, 35),
# Additional for 16-count box
'red-violet': rgbToFloat(192, 68, 143),
'red-orange': rgbToFloat(255, 117, 56),
'yellow-green': rgbToFloat(197, 227, 132),
'blue-violet': rgbToFloat(115, 102, 189),
'carnation-pink': rgbToFloat(255, 170, 204),
'yellow-orange': rgbToFloat(255, 182, 83),
'blue-green': rgbToFloat(25, 158, 189),
'white': rgbToFloat(237, 237, 237),
# Additional for 24-count box
'violet-red': rgbToFloat(247, 83 ,148),
'apricot': rgbToFloat(253, 217, 181),
'cerulean': rgbToFloat(29, 172, 214),
'indigo': rgbToFloat(93, 118, 203),
'scarlet': rgbToFloat(242, 40, 71),
'green-yellow': rgbToFloat(240, 232, 145),
'bluetiful': rgbToFloat(46, 80, 144),
'gray': rgbToFloat(149, 145, 140),
# Additional for 32-count box
'chestnut': rgbToFloat(188, 93, 88),
'peach': rgbToFloat(255, 207, 171),
'sky-blue': rgbToFloat(128, 215, 235),
'cadet-blue': rgbToFloat(176, 183, 198),
'melon': rgbToFloat(253, 188, 180),
'tan': rgbToFloat(250, 167, 108),
'wisteria': rgbToFloat(205, 164, 222),
'timberwolf': rgbToFloat(219, 215, 210),
# Additional for 48-count box
'lavender': rgbToFloat(252, 180, 213),
'burnt-sienna': rgbToFloat(234, 126, 93),
'olive-green': rgbToFloat(186, 184, 108),
'purple-mountains-majesty': rgbToFloat(157, 129, 186),
'salmon': rgbToFloat(255, 155, 170),
'macaroni-and-cheese': rgbToFloat(255, 189, 136),
'granny-smith-apple': rgbToFloat(168, 228, 160),
'sepia': rgbToFloat(165, 105, 79),
'mauvelous': rgbToFloat(239, 152, 170),
'goldenrod': rgbToFloat(255, 217, 117),
'sea-green': rgbToFloat(159, 226, 191),
'raw-sienna': rgbToFloat(214, 138, 89),
'mahogany': rgbToFloat(205, 74, 74),
'spring-green': rgbToFloat(236, 234, 190),
'cornflower': rgbToFloat(154, 206, 235),
'tumbleweed': rgbToFloat(222, 170, 136),
}
(rgbValues, colorNames) = createMoreTrainingData(colorMap)
(colorModel, colorLabels) = train(rgbValues, colorNames, 5)
# -
# In 3 epochs my machine went from 1.2 down to 0.43, then by 5 epochs it got down to 0.37. Plus note it takes longer since we have more colors, more added perturbed colors, and the network itself needs to be bigger to handle more colors.
#
# Let's see how it performs by testing with the sliders:
# +
from ipywidgets import interact
from IPython.core.display import display, HTML
def displayColor(r, g, b):
rInt = min(255, max(0, int(r * 255.0)))
gInt = min(255, max(0, int(g * 255.0)))
bInt = min(255, max(0, int(b * 255.0)))
hexColor = "#%02X%02X%02X" % (rInt, gInt, bInt)
display(HTML('<div style="width: 50%; height: 50px; background: ' + hexColor + ';"></div>'))
numPredictionsToShow = 5
@interact(r = (0.0, 1.0, 0.01), g = (0.0, 1.0, 0.01), b = (0.0, 1.0, 0.01))
def getTopPredictionsFromModel(r, g, b):
testColor = numpy.array([ (r, g, b) ])
predictions = colorModel.predict(testColor, verbose=0) # Predictions shape (1, numColors)
predictions *= 100.0
predColorTuples = []
for i in range(0, len(colorLabels)):
predColorTuples.append((predictions[0][i], colorLabels[i]))
predAndNames = numpy.array(predColorTuples, dtype=[('pred', float), ('colorName', 'U50')])
sorted = numpy.sort(predAndNames, order=['pred', 'colorName'])
sorted = sorted[::-1] # reverse rows to get highest on top
for i in range(0, numPredictionsToShow):
print("%2.1f" % sorted[i][0] + "%", sorted[i][1])
displayColor(r, g, b)
# -
# Rather good results! With so many more colors to choose from, the network is predicting rather well the in-between colors.
#
# And of course, next is the slider that lets you play around with the epochs. But this time there are new sliders you can play with:
#
# * How many perceptrons - the elements that make up our network, which we'll describe more later on - are added for each color name. More perceptrons do not always mean more accuracy. And each perceptron we add increases the amount of math needed to train and use the network.
# * The batch size. This is how many color examples are given at a time before the network is told to adjust itself to the new data (which is called *backpropagation*, something we'll be learning more about later). We've been running with a batch size of 1 so far, which is slow but best for accuracy, whereas bigger numbers train faster but tend to be less accurate, meaning you might have to increase the epochs as well. Try running with batch sizes 3 and 5 and see if your results with the predicted colors are very different.
#
# Play with the numbers and see the results to get a feel for things.
@interact(epochs = (1, 10), perceptronsPerColorName = (1, 32), batchSize = (1, 10))
def trainModel(epochs=3, perceptronsPerColorName=8, batchSize=1):
global colorModel
global colorLabels
(colorModel, colorLabels) = train(rgbValues, colorNames, epochs=epochs, perceptronsPerColorName=perceptronsPerColorName, batchSize=batchSize)
interact(getTopPredictionsFromModel, r = (0.0, 1.0, 0.01), g = (0.0, 1.0, 0.01), b = (0.0, 1.0, 0.01))
# + [markdown] slideshow={"slide_type": "subslide"}
# What I found was that 2 and 3 perceptrons per color name were too low, the predictions were pretty bad at 3 epochs. At 4 it gets about as good as when it was at 8, and setting it to 28 did not much better at all.
#
# Similarly, batch size 2 seemed about as good as 1, but trained twice as fast, 3 a little bit less accurate but faster, but by 10 or so it was relatively inaccurate - loss of 0.86 instead of 0.4 - but had a lot faster training.
#
# Numbers like epochs, perceptron counts, and batch sizes are called *hyperparameters* to the neural network. Adjusting them can make the network better or worse at its job, and in fact for a given problem and set of training data there is not always an exact right set of numbers, instead you often have to run many experiments with different hyperparameters to try to find the best results.
#
# And if you have a lot of experiments, a lot of data, and a lot of perceptrons, you get even more math...
#
# ### Coming up...
# We'll start using the hyperparameters to save a bit of training time at the cost of some accuracy. And we'll learn why perceptrons get *triggered*...
# -
| Part07_48_Crayons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #Cahn-Hilliard with Primtive and Legendre Bases
#
# This example uses a Cahn-Hilliard model to compare two different bases representations to discretize the microstructure. One basis representaion uses the primitive (or hat) basis and the other uses Legendre polynomials. The example includes the background theory about using Legendre polynomials as a basis in MKS. The MKS with two different bases are compared with the standard spectral solution for the Cahn-Hilliard solution at both the calibration domain size and a scaled domain size.
# ###Cahn-Hilliard Equation
#
# The Cahn-Hilliard equation is used to simulate microstructure evolution during spinodial decomposition and has the following form,
#
# $$ \dot{\phi} = \nabla^2 \left( \phi^3 - \phi \right) - \gamma \nabla^4 \phi $$
#
# where $\phi$ is a conserved ordered parameter and $\sqrt{\gamma}$ represents the width of the interface. In this example, the Cahn-Hilliard equation is solved using a semi-implicit spectral scheme with periodic boundary conditions, see [Chang and Rutenberg](http://dx.doi.org/10.1103/PhysRevE.72.055701) for more details.
# ### Basis Functions for the Microstructure Function and Influence Function
#
# In this example, we will explore the differences when using the
# Legendre polynomials as the basis function compared to the primitive
# (or hat) basis for the microstructure function and the influence coefficients.
#
# For more information about both of these basis please see the [theory section](THEORY.html).
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
# -
# ##Modeling with MKS
#
# ###Generating Calibration Datasets
#
# Because the microstructure is a continuous field that can have a range of values and changes over time, the first order influence coefficients cannot be calibrated with delta microstructures. Instead, a large number of simulations with random initial conditions will be used to calibrate the first order influence coefficients using linear regression. Let's show how this is done.
#
# The function `make_cahnHilliard` from `pymks.datasets` provides a nice interface to generate calibration datasets for the influence coefficients. The function `make_cahnHilliard` requires the number of calibration samples, given by `n_samples`, and the size and shape of the domain, given by `size`.
# +
import pymks
from pymks.datasets import make_cahn_hilliard
length = 41
n_samples = 400
dt = 1e-2
np.random.seed(101)
size=(length, length)
X, y = make_cahn_hilliard(n_samples=n_samples, size=size, dt=dt)
# -
# The function `make_cahnHilliard` has generated `n_samples` number of random microstructures, `X`, and returned the same microstructures after they have evolved for one time step, given by `y`. Let's take a look at one of them.
# +
from pymks.tools import draw_concentrations
draw_concentrations((X[0], y[0]),('Calibration Input', 'Calibration Output'))
# -
# ### Calibrate Influence Coefficients
#
# In this example, we compare the difference between using the primitive (or hat) basis and the Legendre polynomial basis to represent the microstructure function. As mentioned above, the microstructures (concentration fields) are not discrete phases. This leaves the number of local states in local state space `n_states` as a free hyperparameter. In the next section, we look to see what a practical number of local states for bases would be.
#
# ### Optimizing the Number of Local States
#
# Below, we compare the difference in performance, as we vary the local state, when we choose the primitive basis and the Legendre polynomial basis.
#
# The `(X, y)` sample data is split into training and test data. The code then optimizes `n_states` between `2` and `11` and the two `basis` with the `parameters_to_tune` variable. The `GridSearchCV` takes an `MKSLocalizationModel` instance, a `scoring` function (figure of merit) and the `parameters_to_tune` and then finds the optimal parameters with a grid search.
# +
from pymks.bases import PrimitiveBasis
from sklearn.grid_search import GridSearchCV
from sklearn import metrics
mse = metrics.mean_squared_error
from pymks.bases import LegendreBasis
from pymks import MKSLocalizationModel
from sklearn.cross_validation import train_test_split
train_split_shape = (X.shape[0],) + (np.prod(X.shape[1:]),)
X_train, X_test, y_train, y_test = train_test_split(X.reshape(train_split_shape),
y.reshape(train_split_shape),
test_size=0.5, random_state=3)
prim_basis = PrimitiveBasis(2, [-1, 1])
leg_basis = LegendreBasis(2, [-1, 1])
params_to_tune = {'n_states': np.arange(2, 11),
'basis': [prim_basis, leg_basis]}
Model = MKSLocalizationModel(prim_basis)
scoring = metrics.make_scorer(lambda a, b: -mse(a, b))
fit_params = {'size': size}
gs = GridSearchCV(Model, params_to_tune, cv=5,
fit_params=fit_params, n_jobs=3).fit(X_train, y_train)
# -
# The optimal parameters are the `LegendreBasis` with only 4 local states. More terms don't improve the R-squared value.
print(gs.best_estimator_)
print(gs.score(X_test, y_test))
# +
from pymks.tools import draw_gridscores
lgs = [x for x in gs.grid_scores_ \
if type(x.parameters['basis']) is type(leg_basis)]
pgs = [x for x in gs.grid_scores_ \
if type(x.parameters['basis']) is type(prim_basis)]
draw_gridscores([lgs, pgs], 'n_states', data_labels=['Legendre', 'Primitve'],
colors=['#f46d43', '#1a9641'], score_label='R-Squared',
param_label = 'L - Total Number of Local States')
# -
# As you can see the `LegendreBasis` converges faster than the `PrimitiveBasis`. In order to further compare performance between the two models, lets select 4 local states for both bases.
# ### Comparing the Bases for `n_states=4`
# +
prim_basis = PrimitiveBasis(n_states=4, domain=[-1, 1])
prim_model = MKSLocalizationModel(basis=prim_basis)
prim_model.fit(X, y)
leg_basis = LegendreBasis(4, [-1, 1])
leg_model = MKSLocalizationModel(basis=leg_basis)
leg_model.fit(X, y)
# -
# Now let's look at the influence coefficients for both bases.
#
# First, the `PrimitiveBasis` influence coefficients:
# +
from pymks.tools import draw_coeff
draw_coeff(prim_model.coef_)
# -
# Now, the `LegendreBasis` influence coefficients:
draw_coeff(leg_model.coef_)
# Now, let's do some simulations with both sets of coefficients and compare the results.
# ###Predict Microstructure Evolution
#
# In order to compare the difference between the two bases, we need to have the Cahn-Hilliard simulation and the two MKS models start with the same initial concentration `phi0` and evolve in time. In order to do the Cahn-Hilliard simulation, we need an instance of the class `CahnHilliardSimulation`.
# +
from pymks.datasets.cahn_hilliard_simulation import CahnHilliardSimulation
np.random.seed(66)
phi0 = np.random.normal(0, 1e-9, ((1,) + size))
ch_sim = CahnHilliardSimulation(dt=dt)
phi_sim = phi0.copy()
phi_prim = phi0.copy()
phi_legendre = phi0.copy()
# -
# Let's look at the inital concentration field.
draw_concentrations([phi0[0]], ['Initial Concentration'])
# In order to move forward in time, we need to feed the concentration back into the Cahn-Hilliard simulation and the MKS models.
# +
time_steps = 50
for steps in range(time_steps):
ch_sim.run(phi_sim)
phi_sim = ch_sim.response
phi_prim = prim_model.predict(phi_prim)
phi_legendre = leg_model.predict(phi_legendre)
# -
# Let's take a look at the concentration fields.
# +
from pymks.tools import draw_concentrations
draw_concentrations((phi_sim[0], phi_prim[0], phi_legendre[0]),
('Simulation', 'Primative', 'Legendre'))
# -
# By just looking at the three microstructures is it difficult to see any differences. Below, we plot the difference between the two MKS models and the simulation.
# +
from sklearn import metrics
from pymks.tools import draw_differences
mse = metrics.mean_squared_error
draw_differences([(phi_sim[0] - phi_prim[0]), (phi_sim[0] - phi_legendre[0])],
['Simulaiton - Prmitive', 'Simulation - Legendre'])
print 'Primative mse =', mse(phi_sim[0], phi_prim[0])
print 'Legendre mse =', mse(phi_sim[0], phi_legendre[0])
# -
# The `LegendreBasis` basis clearly outperforms the `PrimitiveBasis` for the same value of `n_states`.
# ##Resizing the Coefficients to use on Larger Systems
#
# Below we compare the bases after the coefficients are resized.
# +
big_length = 3 * length
big_size = (big_length, big_length)
prim_model.resize_coeff(big_size)
leg_model.resize_coeff(big_size)
phi0 = np.random.normal(0, 1e-9, (1,) + big_size)
phi_sim = phi0.copy()
phi_prim = phi0.copy()
phi_legendre = phi0.copy()
# -
# Let's take a look at the initial large concentration field.
draw_concentrations([phi0[0]], ['Initial Concentration'])
# Let's look at the resized coefficients.
#
# First, the influence coefficients from the `PrimitiveBasis`.
draw_coeff(prim_model.coef_)
# Now, the influence coefficients from the `LegendreBases`.
draw_coeff(leg_model.coef_)
# Once again, we are going to march forward in time by feeding the concentration fields back into the Cahn-Hilliard simulation and the MKS models.
for steps in range(time_steps):
ch_sim.run(phi_sim)
phi_sim = ch_sim.response
phi_prim = prim_model.predict(phi_prim)
phi_legendre = leg_model.predict(phi_legendre)
draw_concentrations((phi_sim[0], phi_prim[0], phi_legendre[0]), ('Simulation', 'Primiative', 'Legendre'))
# Both MKS models seem to predict the concentration faily well. However, the Legendre polynomial basis looks to be better. Again, let's look at the difference between the simulation and the MKS models.
# +
draw_differences([(phi_sim[0] - phi_prim[0]), (phi_sim[0] - phi_legendre[0])],
['Simulaiton - Primiative','Simulation - Legendre'])
print 'Primative mse =', mse(phi_sim[0], phi_prim[0])
print 'Legendre mse =', mse(phi_sim[0], phi_legendre[0])
# -
# With the resized influence coefficients, the `LegendreBasis` outperforms the `PrimitiveBasis` for the same value of `n_states`. The value of `n_states` does not necessarily guarantee a fair comparison between the two basis in terms of floating point calculations and memory used.
| notebooks/localization_cahn_hilliard_Legendre_2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0-rc4
# language: julia
# name: julia-0.5
# ---
workspace()
#addprocs(3)
include("../src/UNSflow.jl")
include("../src/UNSflow.jl")
using UNSflow
# +
#Construct dimensionless quantities from given values
c_d = 0.2
b_d = 6
AR = b_d/c_d
u_d = 10
f_d = 0.8889
k = 2*pi*f_d*c_d/(2*u_d)
#_dAll values below are nondimensional
c = 1
u = 1
w = 2*k
T = (2*pi/w)
ncyc = 8
t_tot = ncyc*T
#modedata = readdlm("../test/anto_mode1.dat");
#mode_spl = Spline1D(modedata[:,3],modedata[:,4])
# +
Tn = T*c_d/u_d
#dlt = readcsv("../test/anto_cl.csv")
#dlt = dlt[2:end,:]
#t_ant = dlt[:,1]/Tn
#len = indmin(abs(t_ant-1))
ant_steady = 0.4946
#ant_ocl = dlt[:,2] - ant_steady
# -
# # 2% of the bending mode
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
#scale = 0.02
# +
#Run LDVM at all these locations:
alpha_amp = 1*pi/180
alpha_mean = 5*pi/180
h_amp = 0.
#for i = 1:n_span
# y_d = -b_d*cos(psi[i])/2.
# h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
#end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
hdef = ConstDef(h_amp)
alphadef = CosDef(alpha_mean, alpha_amp, w, 0.)
udef = ConstDef(u)
pvt = 0.25 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
#surf_in = TwoDSurf[]
#field_in = TwoDFlowField[]
#n_in = Int64[]
#dt_in = Float64[]
#del_in = DelVortDef[]
#for i = 1:Int(n_span/2)
#hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
#push!(surf_in,surf)
#push!(field_in, curfield)
#push!(n_in,nsteps,)
#push!(dt_in,dtstar)
#push!(del_in,del)
#end
#@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
tmat, tsurf, tcur = ldvm(surf, curfield, nsteps, dtstar, del)
for i = 1:Int(n_span/2)
push!(W_mat,tmat)
push!(W_surf, tsurf)
push!(W_curfield, tcur)
end
# -
nsteps
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
# +
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
# ### 10% scale
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
scale = 0.1
# +
#Run LDVM at all these locations:
alpha_amp = 5*pi/180
h_amp = zeros(n_span)
for i = 1:n_span
y_d = -b_d*cos(psi[i])/2.
h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
alphadef = ConstDef(alpha_amp)
udef = ConstDef(u)
pvt = 0.0 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
surf_in = TwoDSurf[]
field_in = TwoDFlowField[]
n_in = Int64[]
dt_in = Float64[]
del_in = DelVortDef[]
for i = 1:Int(n_span/2)
hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
push!(surf_in,surf)
push!(field_in, curfield)
push!(n_in,nsteps,)
push!(dt_in,dtstar)
push!(del_in,del)
end
@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
for i = 1:Int(n_span/2)
push!(W_mat,fullsol[i][1])
push!(W_surf, fullsol[i][2])
push!(W_curfield, fullsol[i][3])
end
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
# ### 25% scale
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
scale = 0.25
# +
#Run LDVM at all these locations:
alpha_amp = 5*pi/180
h_amp = zeros(n_span)
for i = 1:n_span
y_d = -b_d*cos(psi[i])/2.
h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
alphadef = ConstDef(alpha_amp)
udef = ConstDef(u)
pvt = 0.0 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
surf_in = TwoDSurf[]
field_in = TwoDFlowField[]
n_in = Int64[]
dt_in = Float64[]
del_in = DelVortDef[]
for i = 1:Int(n_span/2)
hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
push!(surf_in,surf)
push!(field_in, curfield)
push!(n_in,nsteps,)
push!(dt_in,dtstar)
push!(del_in,del)
end
@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
for i = 1:Int(n_span/2)
push!(W_mat,fullsol[i][1])
push!(W_surf, fullsol[i][2])
push!(W_curfield, fullsol[i][3])
end
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
# ### 50% scale
#
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
scale = 0.5
# +
#Run LDVM at all these locations:
alpha_amp = 5*pi/180
h_amp = zeros(n_span)
for i = 1:n_span
y_d = -b_d*cos(psi[i])/2.
h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
alphadef = ConstDef(alpha_amp)
udef = ConstDef(u)
pvt = 0.0 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
surf_in = TwoDSurf[]
field_in = TwoDFlowField[]
n_in = Int64[]
dt_in = Float64[]
del_in = DelVortDef[]
for i = 1:Int(n_span/2)
hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
push!(surf_in,surf)
push!(field_in, curfield)
push!(n_in,nsteps,)
push!(dt_in,dtstar)
push!(del_in,del)
end
@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
for i = 1:Int(n_span/2)
push!(W_mat,fullsol[i][1])
push!(W_surf, fullsol[i][2])
push!(W_curfield, fullsol[i][3])
end
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
# ### 75% scale
#
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
scale = 0.75
# +
#Run LDVM at all these locations:
alpha_amp = 5*pi/180
h_amp = zeros(n_span)
for i = 1:n_span
y_d = -b_d*cos(psi[i])/2.
h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
alphadef = ConstDef(alpha_amp)
udef = ConstDef(u)
pvt = 0.0 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
surf_in = TwoDSurf[]
field_in = TwoDFlowField[]
n_in = Int64[]
dt_in = Float64[]
del_in = DelVortDef[]
for i = 1:Int(n_span/2)
hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
push!(surf_in,surf)
push!(field_in, curfield)
push!(n_in,nsteps,)
push!(dt_in,dtstar)
push!(del_in,del)
end
@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
for i = 1:Int(n_span/2)
push!(W_mat,fullsol[i][1])
push!(W_surf, fullsol[i][2])
push!(W_curfield, fullsol[i][3])
end
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
# ### 100% scale
#
#
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
scale = 1.0
# +
#Run LDVM at all these locations:
alpha_amp = 5*pi/180
h_amp = zeros(n_span)
for i = 1:n_span
y_d = -b_d*cos(psi[i])/2.
h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
alphadef = ConstDef(alpha_amp)
udef = ConstDef(u)
pvt = 0.0 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
surf_in = TwoDSurf[]
field_in = TwoDFlowField[]
n_in = Int64[]
dt_in = Float64[]
del_in = DelVortDef[]
for i = 1:Int(n_span/2)
hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
push!(surf_in,surf)
push!(field_in, curfield)
push!(n_in,nsteps,)
push!(dt_in,dtstar)
push!(del_in,del)
end
@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
for i = 1:Int(n_span/2)
push!(W_mat,fullsol[i][1])
push!(W_surf, fullsol[i][2])
push!(W_curfield, fullsol[i][3])
end
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
# ### 200% scale
#
# +
n_span = 12
n_bterm = 40
psi = zeros(n_span)
dpsi = pi/n_span
for i = 1:n_span
psi[i] = (real(i)-0.5)*dpsi
end
scale = 2.0
# +
#Run LDVM at all these locations:
alpha_amp = 5*pi/180
h_amp = zeros(n_span)
for i = 1:n_span
y_d = -b_d*cos(psi[i])/2.
h_amp[i] = evaluate(mode_spl,y_d)*scale/c_d
end
dtstar = min(0.015*8,0.015*0.2*2/(k*maximum(h_amp)))
nsteps = round(Int,t_tot/dtstar)+1
alphadef = ConstDef(alpha_amp)
udef = ConstDef(u)
pvt = 0.0 #Doesnt matter, no pitch
lespcrit = [21;] #high value to turn off LEV shedding
W_mat = Array{Float64,2}[]
W_surf = TwoDSurf[]
W_curfield = TwoDFlowField[]
surf_in = TwoDSurf[]
field_in = TwoDFlowField[]
n_in = Int64[]
dt_in = Float64[]
del_in = DelVortDef[]
for i = 1:Int(n_span/2)
hdef = CosDef(0., h_amp[i], w, 0.)
full_kinem = KinemDef(alphadef, hdef, udef)
surf = TwoDSurf("FlatPlate", pvt, full_kinem, lespcrit)
curfield = TwoDFlowField()
del = DelVortDef(1, 500, 10)
push!(surf_in,surf)
push!(field_in, curfield)
push!(n_in,nsteps,)
push!(dt_in,dtstar)
push!(del_in,del)
end
@time fullsol = pmap((a1,a2,a3,a4,a5)->ldvm(a1,a2,a3,a4),surf_in,field_in,n_in,dt_in,del_in)
for i = 1:Int(n_span/2)
push!(W_mat,fullsol[i][1])
push!(W_surf, fullsol[i][2])
push!(W_curfield, fullsol[i][3])
end
# +
#Mirror image for the rest of the span
for i = Int(n_span/2)+1:n_span
mt = W_mat[n_span - i + 1]
st = W_surf[n_span - i + 1]
ct = W_curfield[n_span - i + 1]
push!(W_mat, mt)
push!(W_surf, st)
push!(W_curfield, ct)
end
lhs = zeros(n_span,n_bterm)
rhs = zeros(n_span)
b_coeff = zeros(nsteps,n_bterm)
dt = W_mat[1][2,1] - W_mat[1][1,1]
cnc_f = zeros(nsteps)
cnnc_f = zeros(nsteps)
bdot = zeros(nsteps,n_bterm)
for i = 1:nsteps
for j = 1:n_span
for n = 1:n_bterm
lhs[j,n] = sin(n*psi[j])*(sin(psi[j]) + (n*pi/(2*AR)))
end
rhs[j] = pi*sin(psi[j])*W_mat[j][i,9]/(2*AR)
end
b_coeff[i,:] = \(lhs, rhs)
if i >= 2
bdot[i,:] = (b_coeff[i,:] - b_coeff[i-1,:])/dt
end
end
a03d = zeros(nsteps,n_span)
cd_ind = zeros(nsteps)
a0dot3d = zeros(nsteps,n_span)
for i = 1:nsteps
cd_ind[i] = 0
for n = 1:n_bterm
cd_ind[i] = cd_ind[i] + real(n)*b_coeff[i,n]^2
end
cd_ind[i] = cd_ind[i]*pi*AR
for j = 1:n_span
a03d[i,j] = 0
for n = 1:n_bterm
a03d[i,j] = a03d[i,j] - real(n)*b_coeff[i,n]*sin(n*psi[j])/sin(psi[j])
a0dot3d[i,j] = a0dot3d[i,j] - real(n)*bdot[i,n]*sin(n*psi[j])/sin(psi[j])
end
end
end
W_alpha = zeros(nsteps,n_span)
W_h = zeros(nsteps,n_span)
W_hdot = zeros(nsteps,n_span)
W_u = zeros(nsteps,n_span)
W_u[:,:] = 1
W_alpha[:,:] = 5*pi/180
for i = 1:nsteps
for j = 1:n_span
hdef = CosDef(0., h_amp[j], w, 0.)
tt = W_mat[1][i,1]
W_h[i,j] = hdef(tt)*c
W_hdot[i,j] = ForwardDiff.derivative(hdef,tt)*u
end
end
W_cn = zeros(nsteps)
W_cs = zeros(nsteps)
W_cl = zeros(nsteps)
W_cd = zeros(nsteps)
W_cdi = zeros(nsteps)
cn3d = zeros(nsteps, n_span)
cs3d = zeros(nsteps, n_span)
cl3d = zeros(nsteps, n_span)
cd3d = zeros(nsteps, n_span)
for i = 1:nsteps
W_cn[i] = 0
W_cs[i] = 0
for j = 1:n_span
cn3d[i,j] = W_mat[j][i,10] + (2*pi/u)*(W_u[i,j]*cos(W_alpha[i,j]) + W_hdot[i,j]*sin(W_alpha[i,j]))*a03d[1,j] + (2*pi*c/u)*(3*a0dot3d[i,j]/4)
cs3d[i,j] = W_mat[j][i,11] + 2*pi*a03d[i,j]^2
cl3d[i,j] = cn3d[i,j]*cos(W_alpha[i,j]) + cs3d[i,j]*sin(W_alpha[i,j])
cd3d[i,j] = cn3d[i,j]*sin(W_alpha[i,j]) - cs3d[i,j]*cos(W_alpha[i,j])
W_cn[i] = W_cn[i] + cn3d[i,j]*sin(psi[j])*dpsi/2
W_cs[i] = W_cs[i] + cs3d[i,j]*sin(psi[j])*dpsi/2
W_cl[i] = W_cl[i] + cl3d[i,j]*sin(psi[j])*dpsi/2
W_cd[i] = W_cd[i] + cd3d[i,j]*sin(psi[j])*dpsi/2
end
end
t_th = W_mat[1][:,1]*c_d/u_d
range = round(Int,(ncyc-1)*nsteps/ncyc)+1:nsteps
tbyT = (t_th[range]-t_th[range[1]])/Tn
kr_spl = Spline1D(tbyT,W_cl[range])
kr_eval = evaluate(kr_spl,t_ant)
ant_cl = ant_steady + ant_ocl*scale
len = indmin(abs(t_ant-1))
err = Array(Float64,len)
for i = 1:len
err[i] = (ant_cl[i] - kr_eval[i])/ant_cl[i]
end
rms_err = sqrt(mean(err.^2))*100
# -
plot(t_ant,kr_eval)
plot(t_ant,ant_cl)
| Notebooks/IFASD testcases.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Jupyter-flex allows you to create interactive dashboards based on Jupyter Notebooks based on two simple concepts:
#
# 1. Control the layout of the dashboard using markdown headers
# 2. Define the dashboard components using Jupyter Notebook cell tags
#
# ## Your first dashboard
#
# Let's take a very simple Jupyter Notebook with 3 cells and one plot and convert it to a dashboard.
import plotly.express as px
df = px.data.iris()
fig = px.scatter(df, x="sepal_width", y="sepal_length")
fig.show()
# All you need to do to convert this to a dashboard is to add tag with the value `body` to the that outs the plot.
#
# <div class="admonition tip">
# <p class="admonition-title">How to view and add tags to cells in Jupyter Notebook</p>
# <ol>
# <li>In the top navigation go to View > Cell Toolbar > Tags</li>
# <li>Then type "body" in the new input of target cell and click on "Add tag" or press enter</li>
# </ol>
# </div>
# + tags=["body"]
fig = px.scatter(df, x="sepal_width", y="sepal_length")
fig.show()
# -
# ### Converting the Notebook to a dashboard
#
# From here there is a couple of options to convert the notebook to a html dashboard.
#
# 1. You can execute the notebook as you normaly do in the Jupyter Notebook UI and then select: `File > Download as > Flex Dashboard (.html)`:
# 
# 2. You can go in a terminal and run `nbconvert`:
#
# <p class="code-header">Terminal</p>
#
# ```
# $ jupyter nbconvert --to flex notebook.ipynb
# ```
#
# Optionally add the `--execute` flag to execute the notebook before converting it so the outputs are shown in the dashboard.
#
# <p class="code-header">Terminal</p>
#
# ```
# $ jupyter nbconvert --to flex notebook.ipynb --execute
# ```
#
# Open the resulting `.html` file in a browser and the result will be:
#
# [](/examples/one-plot.html)
#
# <p class="img-caption">Click on the image to open the rendered dashboard</p>
#
# You might notice that the default title of the dashboard is the name of the notebook file, you can customize these using [parameters](#parameters-orientation-and-title).
#
# This is a very simple example, now let's look at the card concept of Jupyter-flex.
# ## Cards: Multiple outputs
#
# A Card is an object that holds one or more components of the dashboard such as markdown or any output generated from the execution of a Cell such as plots, text and widgets.
#
# To learn more about cards and its options go to [Layout > Cards](/layouts/#cards).
#
# You define a new Card by adding a level-3 markdown header (`###`).
#
# Any output from a tagged Cell will be added to the current Card until a new Card, Section or Page is defined.
#
# Going back to the notebook example we can add a new plot to the by adding two new cells:
#
# 1. One markdown cell with a level-3 markdown header (`###`)
# 2. One code cell with the `body` tag
# +
### Second plot
# + tags=["body"]
fig = px.scatter(df, x="petal_width", y="petal_length")
fig.show()
# -
# [](/examples/two-plots.html)
#
# <p class="img-caption">Click on the image to open the rendered dashboard</p>
#
# You will notice two things:
#
# 1. The default layout is a single column with cards stacked vertically and sized to fill available browser height.
# 2. The value of the level-3 markdown header is added to the Card header
# ## Sections: Multiple columns
#
# To add another column to the dashboard define a new Section using a level 2 markdown header (`##`)
#
# In this case, the value of the header is irrelevant (since the default theme doesn't show it), it acts as an indicator to create a new Section.
# +
## Column
# + tags=["body"]
fig = px.scatter(df, x="sepal_length", y="petal_length")
fig.show()
# -
# In this case the result would be:
#
# [](/examples/two-columns.html)
#
# <p class="img-caption">Click on the image to open the rendered dashboard</p>
#
# You will notice another default orientation: to have multiple sections as columns.
# ## Parameters: Orientation and title
#
# You can control the parameters of the dashboard such as title and orientation to be based of rows instead on columns
# by tagging a code cell with `parameters`.
#
# Let's change the orientation of the plot to `rows` and add a title of `A Flex dashboard`.
# + tags=["parameters"]
flex_title = "A flex dashboard"
flex_orientation = "rows"
# -
# [](/examples/two-rows.html)
#
# <p class="img-caption">Click on the image to open the rendered dashboard</p>
#
# ## Learning more
#
# Well done! You have created you first Flex dashboard.
#
# The [Layouts](/layouts) page goes in depth about all the options to control the content of Jupyter-flex dashboards.
#
# The [Plotting](/plotting) page goes through some considerations around different plotting libraries in Jupyter-flex dashboards.
#
# The [Voila and Jupyter Widgets](/voila-widgets/) page describes how to leverage Voila to create dashboards that use a live Jupyter kernel that enable viewers to change underlying parameters and see the results immediately using [Jupyter widgets](https://ipywidgets.readthedocs.io/).
| docs/getting-started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: uncertify
# language: python
# name: uncertify
# ---
# +
# %load_ext autoreload
# %autoreload 2
from context import uncertify
# +
import logging
from uncertify.log import setup_logging
setup_logging()
LOG = logging.getLogger(__name__)
# Matplotlib DEBUG logging spits out a whole bunch of crap
mpl_logger = logging.getLogger('matplotlib')
mpl_logger.setLevel(logging.WARNING)
# + pycharm={"name": "#%%\n"}
LOG.info(f'Your code goes here... "{uncertify.__package__}" loaded successfully from context.py')
| notebooks/.ipynb_checkpoints/template_notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pickle
import numpy
# Functions from other notebook file.
from ipynb.fs.full.shared_functions_server import *
# -
# Move one directory back to the project root.
os.chdir("..")
# Suppress tensorflow log messages.
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# ---
# GLOBALS
STORED_MODEL = 'model-info/t-net'
SERIALIZED_TRAINING = r'model-info/t-net-training'
# Open serialized training history data.
with open(SERIALIZED_TRAINING, 'rb') as input_file:
history = pickle.load(input_file, encoding='bytes')
# +
# GLOBALS
PAD_SIZE = 20
TICK_FONT = 15
LABEL_FONT = 20
TRAINING_ACCURACY = history['acc']
VALIDATION_ACCURACY = history['val_acc']
TRAINING_LOSS = history['loss']
VALIDATION_LOSS = history['val_loss']
EPOCHS = range(0, 50)
ACCURACY_FIGURE = 'T-NET-accuracy.png'
LOSS_FIGURE = 'T-NET-loss.png'
# -
# ---
# ## Evaluate Model History
# +
validation_score = VALIDATION_ACCURACY[-1] # get the last entry
validation_score = round(validation_score * 100, 2)
print('Validation accuracy (latest): {}%'.format(validation_score))
training_score = TRAINING_ACCURACY[-1] # get the last entry
training_score = round(training_score * 100, 2)
print('Training accuracy (latest): {}%'.format(training_score))
# -
# ---
# +
validation_score = VALIDATION_LOSS[-1] # get the last entry
validation_score = round(validation_score, 2)
print('Validation loss (latest): {}'.format(validation_score))
training_score = TRAINING_LOSS[-1] # get the last entry
training_score = round(training_score, 2)
print('Training loss (latest): {}'.format(training_score))
# -
# ---
# ## Accuracy Figure
# +
pyplot.plot(EPOCHS, TRAINING_ACCURACY, 'm--', label='Training Accuracy')
pyplot.plot(EPOCHS, VALIDATION_ACCURACY, 'co-', label='Validation Accuracy')
pyplot.xlabel('Epochs', fontsize=LABEL_FONT, labelpad=PAD_SIZE)
pyplot.xticks(fontsize=TICK_FONT)
pyplot.ylabel('Accuracy', fontsize=LABEL_FONT, labelpad=PAD_SIZE)
pyplot.yticks(numpy.arange(0, 1.1, step=0.1), fontsize=TICK_FONT)
pyplot.legend(loc=4, prop={'size': 15})
acc_fig = pyplot.gcf()
acc_fig.savefig(ACCURACY_FIGURE, dpi=400, bbox_inches='tight')
# -
# ---
# ## Loss Figure
# +
pyplot.plot(EPOCHS, TRAINING_LOSS, 'r--', label='Training Loss')
pyplot.plot(EPOCHS, VALIDATION_LOSS, 'bo-', label='Validation Loss')
pyplot.xlabel('Epochs', fontsize=LABEL_FONT, labelpad=PAD_SIZE)
pyplot.xticks(fontsize=TICK_FONT)
pyplot.ylabel('Loss', fontsize=LABEL_FONT, labelpad=PAD_SIZE)
pyplot.yticks(fontsize=TICK_FONT)
pyplot.legend(loc=1, prop={'size': 15})
loss_fig = pyplot.gcf()
loss_fig.savefig(LOSS_FIGURE, dpi=400, bbox_inches='tight')
# -
# ---
# ## Model Architecture
model = models.load_model(STORED_MODEL)
model.summary()
| notebooks/t-net-evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a name="pagetop"></a>
# <div style="width:1000 px">
#
# <div style="float:right; width:98 px; height:98px;">
# <img src="https://raw.githubusercontent.com/Unidata/MetPy/master/src/metpy/plots/_static/unidata_150x150.png" alt="Unidata Logo" style="height: 98px;">
# </div>
#
# <h1>NumPy Basics</h1>
# <h3>Unidata Python Workshop</h3>
#
# <div style="clear:both"></div>
# </div>
#
# <hr style="height:2px;">
#
# <div style="float:right; width:250 px"><img src="http://www.contribute.geeksforgeeks.org/wp-content/uploads/numpy-logo1.jpg" alt="NumPy Logo" style="height: 250px;"></div>
#
# ### Questions
# 1. What are arrays?
# 2. How can arrays be manipulated effectively in Python?
#
# ### Objectives
# 1. Create an array of ‘data’.
# 2. Perform basic calculations on this data using python math functions.
# 3. Slice and index the array
# NumPy is the fundamental package for scientific computing with Python. It contains among other things:
# - a powerful N-dimensional array object
# - sophisticated (broadcasting) functions
# - useful linear algebra, Fourier transform, and random number capabilities
#
# The NumPy array object is the common interface for working with typed arrays of data across a wide-variety of scientific Python packages. NumPy also features a C-API, which enables interfacing existing Fortran/C/C++ libraries with Python and NumPy.
# ## Create an array of 'data'
#
# The NumPy array represents a *contiguous* block of memory, holding entries of a given type (and hence fixed size). The entries are laid out in memory according to the shape, or list of dimension sizes.
# + slideshow={"slide_type": "subslide"}
# Convention for import to get shortened namespace
import numpy as np
# + slideshow={"slide_type": "fragment"}
# Create a simple array from a list of integers
a = np.array([1, 2, 3])
a
# -
# See how many dimensions the array has
a.ndim
# + slideshow={"slide_type": "fragment"}
# Print out the shape attribute
a.shape
# + slideshow={"slide_type": "fragment"}
# Print out the data type attribute
a.dtype
# + slideshow={"slide_type": "subslide"}
# This time use a nested list of floats
a = np.array([[1., 2., 3., 4., 5.]])
a
# -
# See how many dimensions the array has
a.ndim
# + slideshow={"slide_type": "fragment"}
# Print out the shape attribute
a.shape
# + slideshow={"slide_type": "fragment"}
# Print out the data type attribute
a.dtype
# -
# <div class="alert alert-warning">
# <h2>Poll</h2>
# Please go to <a href="http://www.PollEv.com/johnleeman205">http://www.PollEv.com/johnleeman205</a> to take a quick poll.
# </div>
# + [markdown] slideshow={"slide_type": "subslide"}
# NumPy also provides helper functions for generating arrays of data to save you typing for regularly spaced data.
#
# * `arange(start, stop, interval)` creates a range of values in the interval `[start,stop)` with `step` spacing.
# * `linspace(start, stop, num)` creates a range of `num` evenly spaced values over the range `[start,stop]`.
# -
# ### arange
# + slideshow={"slide_type": "fragment"}
a = np.arange(5)
print(a)
# -
a = np.arange(3, 11)
print(a)
# + slideshow={"slide_type": "fragment"}
a = np.arange(1, 10, 2)
print(a)
# -
# <div class="alert alert-warning">
# <h2>Poll</h2>
# Please go to <a href="http://www.PollEv.com/johnleeman205">http://www.PollEv.com/johnleeman205</a> to take a quick poll.
# </div>
# ### linspace
# + slideshow={"slide_type": "fragment"}
b = np.linspace(5, 15, 5)
print(b)
# -
b = np.linspace(2.5, 10.25, 11)
print(b)
# <div class="alert alert-warning">
# <h2>Poll</h2>
# Please go to <a href="http://www.PollEv.com/johnleeman205">http://www.PollEv.com/johnleeman205</a> to take a quick poll.
# </div>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Perform basic calculations with Python
#
# ### Basic math
#
# In core Python, that is *without* NumPy, creating sequences of values and adding them together requires writing a lot of manual loops, just like one would do in C/C++:
# -
a = range(5, 10)
b = [3 + i * 1.5/4 for i in range(5)]
# + slideshow={"slide_type": "fragment"}
result = []
for x, y in zip(a, b):
result.append(x + y)
print(result)
# + [markdown] slideshow={"slide_type": "subslide"}
# That is very verbose and not very intuitive. Using NumPy this becomes:
# -
a = np.arange(5, 10)
b = np.linspace(3, 4.5, 5)
# + slideshow={"slide_type": "fragment"}
a + b
# -
# The four major mathematical operations operate in the same way. They perform an element-by-element calculation of the two arrays. The two must be the same shape though!
# + slideshow={"slide_type": "fragment"}
a * b
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Constants
#
# NumPy proves us access to some useful constants as well - remember you should never be typing these in manually! Other libraries such as SciPy and MetPy have their own set of constants that are more domain specific.
# -
np.pi
np.e
# + slideshow={"slide_type": "fragment"}
# This makes working with radians effortless!
t = np.arange(0, 2 * np.pi + np.pi / 4, np.pi / 4)
t
# -
# ### Array math functions
#
# NumPy also has math functions that can operate on arrays. Similar to the math operations, these greatly simplify and speed up these operations. Be sure to checkout the [listing](https://docs.scipy.org/doc/numpy/reference/routines.math.html) of mathematical functions in the NumPy documentation.
# + slideshow={"slide_type": "fragment"}
# Calculate the sine function
sin_t = np.sin(t)
print(sin_t)
# -
# Round to three decimal places
print(np.round(sin_t, 3))
# Calculate the cosine function
cos_t = np.cos(t)
print(cos_t)
# Convert radians to degrees
degrees = np.rad2deg(t)
print(degrees)
# Integrate the sine function with the trapezoidal rule
sine_integral = np.trapz(sin_t, t)
print(np.round(sine_integral, 3))
# Sum the values of the cosine
cos_sum = np.sum(cos_t)
print(cos_sum)
# Calculate the cumulative sum of the cosine
cos_csum = np.cumsum(cos_t)
print(cos_csum)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Index and slice arrays
#
# Indexing is how we pull individual data items out of an array. Slicing extends this process to pulling out a regular set of the items.
# + slideshow={"slide_type": "subslide"}
# Create an array for testing
a = np.arange(12).reshape(3, 4)
# + slideshow={"slide_type": "fragment"}
a
# + [markdown] slideshow={"slide_type": "subslide"}
# Indexing in Python is 0-based, so the command below looks for the 2nd item along the first dimension (row) and the 3rd along the second dimension (column).
#
# 
# + slideshow={"slide_type": "fragment"}
a[1, 2]
# + [markdown] slideshow={"slide_type": "subslide"}
# Can also just index on one dimension
# + slideshow={"slide_type": "fragment"}
a[2]
# + [markdown] slideshow={"slide_type": "subslide"}
# Negative indices are also allowed, which permit indexing relative to the end of the array.
# + slideshow={"slide_type": "fragment"}
a[0, -1]
# -
# <div class="alert alert-warning">
# <h2>Poll</h2>
# Please go to <a href="http://www.PollEv.com/johnleeman205">http://www.PollEv.com/johnleeman205</a> to take a quick poll.
# </div>
# + [markdown] slideshow={"slide_type": "subslide"}
# Slicing syntax is written as `start:stop[:step]`, where all numbers are optional.
# - defaults:
# - start = 0
# - stop = len(dim)
# - step = 1
# - The second colon is also optional if no step is used.
#
# It should be noted that end represents one past the last item; one can also think of it as a half open interval: `[start, end)`
# + slideshow={"slide_type": "subslide"}
# Get the 2nd and 3rd rows
a[1:3]
# + slideshow={"slide_type": "fragment"}
# All rows and 3rd column
a[:, 2]
# + slideshow={"slide_type": "fragment"}
# ... can be used to replace one or more full slices
a[..., 2]
# -
# Slice every other row
a[::2]
# + [markdown] slideshow={"slide_type": "fragment"}
# <div class="alert alert-warning">
# <h2>Poll</h2>
# Please go to <a href="http://www.PollEv.com/johnleeman205">http://www.PollEv.com/johnleeman205</a> to take a quick poll.
# </div>
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>The code below calculates a two point average using a Python list and loop. Convert it do obtain the same results using NumPy slicing</li>
# <li>Bonus points: Can you extend the NumPy version to do a 3 point (running) average?</li>
# </ul>
# </div>
# +
data = [1, 3, 5, 7, 9, 11]
out = []
# Look carefully at the loop. Think carefully about the sequence of values
# that data[i] takes--is there some way to get those values as a numpy slice?
# What about for data[i + 1]?
for i in range(len(data) - 1):
out.append((data[i] + data[i + 1]) / 2)
print(out)
# +
# YOUR CODE GOES HERE
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/slice.py
# +
# YOUR BONUS CODE GOES HERE
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/slice_bonus.py
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>Given the array of data below, calculate the total of each of the columns (i.e. add each of the three rows together):</li>
# </ul>
# </div>
# +
data = np.arange(12).reshape(3, 4)
# YOUR CODE GOES HERE
# total = ?
# -
# <div class="alert alert-info">
# <b>SOLUTION</b>
# </div>
# +
# # %load solutions/sum_row.py
# -
# ## Resources
#
# The goal of this tutorial is to provide an overview of the use of the NumPy library. It tries to hit all of the important parts, but it is by no means comprehensive. For more information, try looking at the:
# - [Tentative NumPy Tutorial](http://wiki.scipy.org/Tentative_NumPy_Tutorial)
# - [NumPy User Guide](http://docs.scipy.org/doc/numpy/user/)
# - [SciPy Lecture Notes](https://scipy-lectures.org/)
# <a href="#pagetop">Top</a>
# <hr style="height:2px;">
| pages/workshop/NumPy/Numpy Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# +
import dipy.reconst.dti as dti
import os
import numpy as np
import math
import SimpleITK as sitk
from scipy import ndimage
import nibabel as nib
from PIL import Image
import scipy.misc
from scipy import signal
import warnings
import SimpleITK as sitk
from dipy.reconst.dti import *
# -
from ndreg import *
from dipy.tracking.eudx import EuDX
# +
def plot_rgb(im):
plt.rcParams.update({'axes.labelsize': 'x-large',
'axes.titlesize': 'x-large'})
if im.shape == (182, 218, 182):
x = [78, 90, 100]
y = [82, 107, 142]
z = [88, 103, 107]
else:
shap = im.shape
x = [int(shap[0]*0.35), int(shap[0]*0.51), int(shap[0]*0.65)]
y = [int(shap[1]*0.35), int(shap[1]*0.51), int(shap[1]*0.65)]
z = [int(shap[2]*0.35), int(shap[2]*0.51), int(shap[2]*0.65)]
coords = (x, y, z)
labs = ['Sagittal Slice (YZ fixed)',
'Coronal Slice (XZ fixed)',
'Axial Slice (XY fixed)']
var = ['X', 'Y', 'Z']
idx = 0
for i, coord in enumerate(coords):
for pos in coord:
idx += 1
ax = plt.subplot(3, 3, idx)
ax.set_title(var[i] + " = " + str(pos))
if i == 0:
image = ndimage.rotate(im[pos, :, :], 90)
elif i == 1:
image = ndimage.rotate(im[:, pos, :], 90)
else:
image = im[:, :, pos]
if idx % 3 == 1:
ax.set_ylabel(labs[i])
ax.yaxis.set_ticks([0, image.shape[0]/2, image.shape[0] - 1])
ax.xaxis.set_ticks([0, image.shape[1]/2, image.shape[1] - 1])
plt.imshow(image)
fig = plt.gcf()
fig.set_size_inches(12.5, 10.5, forward=True)
return fig
def tiff_to_array(folder_path, input_path):
"""
Function takes a single image (TIFF, or other also works), and returns
the single image as a numpy array. Called by tiff_stack_to_array.
:param input_path: Single image file to open.
:return: Numpy representation of image.
"""
# The convert tag makes sure that we're dealing with floats, not uint8
# This prevents underflow.
im = Image.open(folder_path + input_path).convert("F")
# im.show()
imarray = np.array(im)
# print(imarray)
# print(imarray.dtype)
return imarray
def tiff_stack_to_array(input_path):
"""
Function takes input_path, which should should lead to a directory.
Loads all TIFFs in input_path, then generates numpy arrays from the
TIFF stack by calling tiff_to_array helper function. Make sure TIFF
images are ordered in numerical order.
:param input_path: Folder or directory containing .tiff stack.
:return: Numpy array of tiff stack.
"""
im_list = [];
for filename in os.listdir(input_path):
if filename.endswith(".tif"):
# print(os.path.join(directory, filename))
im_arr = tiff_to_array(input_path, filename)
im_list.append(im_arr)
s = np.stack(im_list, axis=2)
print s.shape
return s
# +
# A Python implementation of Ailey's matlab tensor code.
import os
import numpy as np
import math
import SimpleITK as sitk
from scipy import ndimage
import nibabel as nib
from PIL import Image
import scipy.misc
from scipy import signal
import warnings
#warnings.filterwarnings("ignore")
def doggen(sigma):
"""
Helper function to generate derivatives of Gaussian kernels, in either 1D, 2D, or 3D.
Source code in MATLAB obtained from <NAME>, Stanford University, September 2015
:param sigma: Sigma for use (see defaults in generate_FSL_structure_tensor)
:return: Derivative of Gaussian kernel with dimensions of sigma.
"""
halfsize = np.ceil(3 * np.max(sigma))
x = range(np.single(-halfsize), np.single(halfsize + 1)); # Python colon is not inclusive at end, while MATLAB is.
dim = len(sigma);
if dim == 1:
X = np.array(x); # Remember that, by default, numpy arrays are elementwise multiplicative
X = X.astype(float);
k = -X * np.exp(-X**2/(2 * sigma**2));
elif dim == 2:
[X, Y] = np.meshgrid(x, x);
X = X.astype(float);
Y = Y.astype(float);
k = -X * np.exp(-X**2/(2*sigma[0]^2) * np.exp(-Y**2))
elif dim == 3:
[X, Y, Z] = np.meshgrid(x, x, x);
X = X.transpose(0, 2, 1); # Obtained through vigorous testing (see below...)
Y = Y.transpose(2, 0, 1);
Z = Z.transpose(2, 1, 0);
X = X.astype(float);
Y = Y.astype(float);
Z = Z.astype(float);
k = -X * np.exp(np.divide(-np.power(X, 2), 2 * np.power(sigma[0], 2))) * np.exp(np.divide(-np.power(Y,2), 2 * np.power(sigma[1],2))) * np.exp(np.divide(-np.power(Z,2), 2 * np.power(sigma[2],2)))
else:
print 'Only supports up to 3 dimensions'
return np.divide(k, np.sum(np.abs(k[:])));
def gaussgen(sigma):
"""
Function to generate Gaussian kernels, in 1D, 2D and 3D.
Source code in MATLAB obtained from <NAME>, Stanford University, September 2015
:param sigma: Sigma for use in generating Gaussian kernel (see defaults in generate_FSL_structure_tensor)
:return: Gaussian kernel with dimensions of sigma.
"""
halfsize = np.ceil(3 * max(sigma));
x = range(np.single(-halfsize), np.single(halfsize + 1));
dim = len(sigma);
if dim == 1:
x = x.astype(float);
k = np.exp(-x**2 / (2 * sigma^2));
elif dim == 2:
[X, Y] = np.meshgrid(x, x);
X = X.astype(float);
Y = Y.astype(float);
k = np.exp(-X**2 / (2 * sigma[0]**2)) * np.exp(-Y**2 / (2 * sigma[1]**2));
elif dim == 3:
[X, Y, Z] = np.meshgrid(x, x, x);
X = X.transpose(0, 2, 1); # Obtained through vigorous testing (see below...)
Y = Y.transpose(2, 0, 1);
Z = Z.transpose(2, 1, 0);
X = X.astype(float);
Y = Y.astype(float);
Z = Z.astype(float);
k = np.exp(-X**2 / (2 * sigma[0]**2)) * np.exp(-Y**2 / (2 * sigma[1]**2)) * np.exp(-Z**2 / (2 * sigma[2]**2));
else:
print 'Only supports up to dimension 3'
return np.divide(k, np.sum(np.abs(k)));
def tiff_to_array(folder_path, input_path):
"""
Function takes a single image (TIFF, or other also works), and returns
the single image as a numpy array. Called by tiff_stack_to_array.
:param input_path: Single image file to open.
:return: Numpy representation of image.
"""
# The convert tag makes sure that we're dealing with floats, not uint8
# This prevents underflow.
im = Image.open(folder_path + input_path).convert("F")
# im.show()
imarray = np.array(im)
# print(imarray)
# print(imarray.dtype)
return imarray
def tiff_stack_to_array(input_path):
"""
Function takes input_path, which should should lead to a directory.
Loads all TIFFs in input_path, then generates numpy arrays from the
TIFF stack by calling tiff_to_array helper function. Make sure TIFF
images are ordered in numerical order.
:param input_path: Folder or directory containing .tiff stack.
:return: Numpy array of tiff stack.
"""
im_list = [];
for filename in os.listdir(input_path):
if filename.endswith(".tiff"):
# print(os.path.join(directory, filename))
im_arr = tiff_to_array(input_path, filename)
im_list.append(im_arr)
s = np.stack(im_list, axis=2)
print s.shape
return s
def nii_to_tiff_stack(input_path, token):
"""
Function loads an nii using SITK, then converts the nii into a folder containing a TIFF stack.
This function is useful later on for generating the structure tensor.
:param input_path: Path to .nii file.
:param token: Name of token.
"""
image = sitk.ReadImage(input_path);
planes_number = image.GetSize();
data = sitk.GetArrayFromImage(image)
z_dimension = planes_number[2];
## if we have (i, j, k), we want (k, j, i) (converts nibabel format to sitk format)
##new_im = aut_1367.swapaxes(0,2) # just swap i and k
if not os.path.exists(token + "_TIFFs"):
os.makedirs(token + "_TIFFs");
plane = 0;
for plane in range(0, z_dimension):
output = data[plane, :, :]
scipy.misc.toimage(output).save(token + "_TIFFs/" + token + "_" + str(plane) + '.tiff')
def generate_FSL_structure_tensor(img_data, filename, dogsigmaArr=[1], gausigmaArr=[2.3], angleArr=[25]):
"""
Function takes a numpy array (from TIFF_stack_to_array) and saves output
FSL structure tensor as filename string. Allows inputting alternate dogsigmaArr,
gausigmaArr, angleArr, although defaults to currently to parameters from MATLAB script.
Also returns tensorfsl (the tensor fsl structure) image numpy array.
## Parameters (the script loops through all parameters and saves each result automatically)
# dogsigmaArr = [1]; Sigma values for derivative of gaussian filter, recommended value: 0.6 - 1.3 (based on actual data)
# gausigmaArr = [2.3]; Sigma values for gaussian filter, recommended value: 1.3 - 2.3 (based on actual data)
# angleArr = [25]; Angle thresholds for fiber tracking, recommended value: 20 - 30.
Follows code from MATLAB CAPTURE scripts.
:param img_data: Numpy array of image, typically from tiff_stack_to_array called on a directory of TIFFs.
:param filename: Name to save the FSL structure tensor as.
:param dogsigmaArr: Sigma values for derivative of Gaussian filter, with recommended values between 0.6 - 1.3.
:param gausigmaArr: Sigma values for Gaussian filter, with recommended values between 1.3 - 2.3.
:param angleArr: Angle threshold for fiber tracking, with recommended values between 20 - 30.
:return tensorfsl: TensorFSL format of structure tensor (upper triangular matrix)
"""
for jj in range(len(dogsigmaArr)):
dogsigma = dogsigmaArr[jj];
print "Start DoG Sigma on " + str(dogsigma);
# Generate dog kernels
dogkercc = doggen([dogsigma, dogsigma, dogsigma]);
dogkercc = np.transpose(dogkercc, (0, 2, 1)); # annoying
#print dogkercc.shape;
#print dogkercc[:, :, 0];
dogkerrr = np.transpose(dogkercc, (1, 0, 2));
#print dogkerrr[:, :, 0];
dogkerzz = np.transpose(dogkercc, (0, 2, 1));
#print dogkerzz[:, :, 0];
# Compute gradients
grr = signal.convolve(img_data, dogkerrr, 'same');
#print grr[:, :, 0];
gcc = signal.convolve(img_data, dogkercc, 'same');
#print gcc[:, :, 0];
gzz = signal.convolve(img_data, dogkerzz, 'same');
#print gzz[:, :, 0];
# Compute gradient products
gprrrr = np.multiply(grr, grr);
#print gprrrr[:, :, 0];
gprrcc = np.multiply(grr, gcc);
#print gprrcc[:, :, 0];
gprrzz = np.multiply(grr, gzz);
#print gprrzz[:, :, 0]
gpcccc = np.multiply(gcc, gcc);
gpcczz = np.multiply(gcc, gzz);
gpzzzz = np.multiply(gzz, gzz);
# Compute gradient amplitudes
# print ga.dtype;
ga = np.sqrt(gprrrr + gpcccc + gpzzzz);
#print ga[:, :, 0];
#print "GA SHAPE:"
#print ga.shape;
# Convert numpy ndarray object to Nifti data type
gradient_amplitudes_data = nib.Nifti1Image(ga, affine=np.eye(4));
# Save gradient amplitudes image
nib.save(gradient_amplitudes_data, 'gradient_amplitudes.nii');
# Compute gradient vectors
gv = np.concatenate((grr[..., np.newaxis], gcc[..., np.newaxis], gzz[..., np.newaxis]), axis = 3);
#print gv[:, :, 0, 0];
gv = np.divide(gv, np.tile(ga[..., None], [1, 1, 1, 3]));
#print gv[:, :, 0, 1];
#print "GV SHAPE:"
#print gv.shape;
# Convert numpy ndarray object to Nifti data type
gradient_vectors_data = nib.Nifti1Image(gv, affine=np.eye(4));
# Save gradient vectors
nib.save(gradient_vectors_data, 'gradient_vectors.nii');
# Compute structure tensor
for kk in range(len(gausigmaArr)):
gausigma = gausigmaArr[kk];
print "Start Gauss Sigma with gausigma = " + str(gausigma);
print "Generating Gaussian kernel..."
gaussker = np.single(gaussgen([gausigma, gausigma, gausigma]));
#print gaussker[:, :, 0];
print "Blurring gradient products..."
gprrrrgauss = signal.convolve(gprrrr, gaussker, "same");
#print gprrrrgauss[:, :, 0];
gprrccgauss = signal.convolve(gprrcc, gaussker, "same");
#print gprrccgauss[:, :, 0];
gprrzzgauss = signal.convolve(gprrzz, gaussker, "same");
gpccccgauss = signal.convolve(gpcccc, gaussker, "same");
gpcczzgauss = signal.convolve(gpcczz, gaussker, "same");
gpzzzzgauss = signal.convolve(gpzzzz, gaussker, "same");
print "Saving a copy for this Gaussian sigma..."
tensorfsl = np.concatenate((gprrrrgauss[..., np.newaxis], gprrccgauss[..., np.newaxis], gprrzzgauss[..., np.newaxis], gpccccgauss[..., np.newaxis], gpcczzgauss[..., np.newaxis], gpzzzzgauss[..., np.newaxis]), axis = 3);
tmp = np.copy(tensorfsl[:,:,:,3])
tensorfsl[:,:,:,3] = tensorfsl[:,:,:,2]
tensorfsl[:,:,:,2] = tmp
# Convert numpy ndarray object to Nifti data type
tensor_fsl_data = nib.Nifti1Image(tensorfsl, affine=np.eye(4));
nib.save(tensor_fsl_data, str(filename) + "dogsigma_" + str(jj) + "gausigma_" + str(kk) + 'tensorfsl.nii');
print 'Completed computing structure tensor on ' + str(filename) + '!'
return tensorfsl
def plot_rgb(im):
plt.rcParams.update({'axes.labelsize': 'x-large',
'axes.titlesize': 'x-large'})
if im.shape == (182, 218, 182):
x = [78, 90, 100]
y = [82, 107, 142]
z = [88, 103, 107]
else:
shap = im.shape
x = [int(shap[0]*0.35), int(shap[0]*0.51), int(shap[0]*0.65)]
y = [int(shap[1]*0.35), int(shap[1]*0.51), int(shap[1]*0.65)]
z = [int(shap[2]*0.35), int(shap[2]*0.51), int(shap[2]*0.65)]
coords = (x, y, z)
labs = ['Sagittal Slice (YZ fixed)',
'Coronal Slice (XZ fixed)',
'Axial Slice (XY fixed)']
var = ['X', 'Y', 'Z']
idx = 0
for i, coord in enumerate(coords):
for pos in coord:
idx += 1
ax = plt.subplot(3, 3, idx)
ax.set_title(var[i] + " = " + str(pos))
if i == 0:
image = ndimage.rotate(im[pos, :, :,0:3], 90)
elif i == 1:
image = ndimage.rotate(im[:, pos, :,0:3], 90)
else:
image = im[:, :, pos,0:3]
print image.shape
if idx % 3 == 1:
ax.set_ylabel(labs[i])
ax.yaxis.set_ticks([0, image.shape[0]/2, image.shape[0] - 1])
ax.xaxis.set_ticks([0, image.shape[1]/2, image.shape[1] - 1])
plt.imshow(image)
fig = plt.gcf()
fig.set_size_inches(12.5, 10.5, forward=True)
return fig
def fiber_stream(f):
test = f
print len(test)
fig = plt.figure(1)
plt.subplots(figsize=(10, 10))
plt.subplot(311)
plt.title("Y-axis vs X-axis (" + str(len(test)) + " fibers)")
for i in range(len(test)):
plt.plot(test[i][:,0], test[i][:,1])
plt.subplot(312)
plt.title("Z-axis vs X-axis (" + str(len(test)) + " fibers)")
for i in range(len(test)):
plt.plot(test[i][:,0], test[i][:,2])
plt.subplot(313)
plt.title("Z-axis vs Y-axis (" + str(len(test)) + " fibers)")
for i in range(len(test)):
plt.plot(test[i][:,1], test[i][:,2])
plt.tight_layout()
#fig = plt.show()
fig.savefig('tensor_streamlines.png')
def tensor2tract(struct_tensor, is_fsl):
if is_fsl:
tmp = np.copy(struct_tensor[:,:,:,3])
struct_tensor[:,:,:,3] = struct_tensor[:,:,:,2]
struct_tensor[:,:,:,2] = tmp
output = from_lower_triangular(struct_tensor)
evals, evecs = decompose_tensor(output)
FA = fractional_anisotropy(evals)
RGB = color_fa(FA, evecs)
# nb.save(nb.Nifti1Image(np.array(255 * RGB, 'uint8'), result.get_affine()), 'fsl_tensor_rgb_upper.nii.gz')
affine = img.get_affine()
fa = nib.Nifti1Image(np.array(255 * RGB, 'uint8'), affine)
im = fa.get_data()
fig = plot_rgb(im)
plt.savefig('tensor_field_brain.png')
sphere = get_sphere('symmetric724')
peak_indices = quantize_evecs(evecs, sphere.vertices)
eu = EuDX(FA.astype('f8'), peak_indices, seeds=50000, odf_vertices = sphere.vertices, a_low=0.2)
tensor_streamlines = [streamline for streamline in eu]
return tensor_streamlines
# -
def tensor2tract(struct_tensor, is_fsl):
if is_fsl:
tmp = np.copy(struct_tensor[:,:,:,3])
struct_tensor[:,:,:,3] = struct_tensor[:,:,:,2]
struct_tensor[:,:,:,2] = tmp
output = from_lower_triangular(struct_tensor)
evals, evecs = decompose_tensor(output)
FA = fractional_anisotropy(evals)
RGB = color_fa(FA, evecs)
# nb.save(nb.Nifti1Image(np.array(255 * RGB, 'uint8'), result.get_affine()), 'fsl_tensor_rgb_upper.nii.gz')
#affine = struct_tensor.get_affine()
fa = nib.Nifti1Image(np.array(255 * RGB, 'uint8'), affine)
im = fa.get_data()
fig = plot_rgb(im)
plt.savefig('tensor_field_brain.png')
sphere = get_sphere('symmetric724')
peak_indices = quantize_evecs(evecs, sphere.vertices)
eu = EuDX(FA.astype('f8'), peak_indices, seeds=50000, odf_vertices = sphere.vertices, a_low=0.2)
tensor_streamlines = [streamline for streamline in eu]
return tensor_streamlines
import tractography_latest as tract
reload(tract)
formatimg = tract.tiff_stack_to_array('CTT/demo/data/')
fsl, dtk = tract.generate_FSL_and_DTK_structure_tensor(formatimg, 'jovo', dogsigmaArr=[1], gausigmaArr=[2.3]);
mask = tract.tiff_stack_to_nii('CTT/demo/mask-brain/', 'brainmask')
mask = tract.tiff_stack_to_array('CTT/demo/mask-brain/')
print mask.shape
newmask = nib.load('CTT/demo/result/mask-brain.nii.gz')
print newmask.shape
print newmask.affine
affine = newmask.affine
streamlines = tract.tensor2tract(dtk, False)
affine = newmask.affine
streamlines = tensor2tract(dtk, False)
np.savez('neweststreams', streamlines)
# +
# #!python multigraphs.py neweststreams.npz hello/ CTT/demo/result/dog1gau0.5/gv.nii.gz
# -
# !python multigraphs.py neweststreams.npz hello/ CTT/demo/result/data.nii.gz
# !python multigraphs.py neweststreams.npz ara/ CTT/demo/result/mask-roi1.nii.gz
import networkx as nx
#'/home/graph.graphml'
path = 'hello/graphs/ga/neweststreams_ga.gpickle'
import networkx as nx
#'/home/graph.graphml'
path = 'ara/graphs/mask-roi1/neweststreams_mask-roi1.gpickle'
g = nx.read_gpickle(path)
g = nx.adj_matrix(g).todense()
fig = plt.figure(figsize=(7,7))
p = plt.imshow(g, interpolation='None')
affine = newmask.affine
streamlines = tract.tensor2tract(dtk, False)
| Albert_Jupyter/connectomes+final.ipynb |
# %matplotlib inline
from fenics import *
parameters["plotting_backend"] = 'matplotlib'
import pylab
# +
# Define discrete Functionspace
mesh = UnitSquareMesh(100, 100)
U = FiniteElement("Lagrange", triangle, 1) # Finite element for forward PDE space
V = FiniteElement("Lagrange", triangle, 1) # Finite element for adjoint PDE space
M = FiniteElement("DG", triangle, 0) # Finite lement for control space
W = FunctionSpace(mesh, MixedElement([U, V, M]))
# Define Functions
w = Function(W)
u, lmbd, f = split(w) # Solution functions
x = TestFunction(W) # Test functions
# Define variational problem
a = inner(grad(u), grad(lmbd))*dx
L = f*lmbd*dx
# Define functional
ud = Expression("sin(pi*x[0])*sin(pi*x[1])", degree=4) # Desired temperature profile
alpha = Constant(1e-6) # Regularization parameter
J = (u-ud)**2*dx + alpha*f**2*dx
# Define boundary conditions
bc_u = DirichletBC(W.sub(0), 0.0, "on_boundary")
bc_lmbd = DirichletBC(W.sub(1), 0.0, "on_boundary")
bcs = [bc_u, bc_lmbd]
# Derive optimality conditions
lagrang = J + a + L
kkt = derivative(lagrang, w, x)
# Solve Poisson problem
solve(kkt == 0, w, bcs)
plot(w[0], title="Temperature")
pylab.show()
plot(w[2], title="Control")
pylab.show()
plot(ud-w[0], title="Temperature difference")
interactive()
| notebooks/15_one_shot_optimisation/poisson_one_shot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Scrapping diamonds data from 'James Allen' website for our project
# ### A view of the website:
# 
# 
# ### A view at diamond's information table:
# 
# ### Import libraries
import pandas as pd
import requests
from bs4 import BeautifulSoup as bs
import random
# +
# Creating a function to retrieve data from a diamond's page.
def retrieve_data(page, listing_divs):
listing_list = []
#itterate each diamond in the page (there are maximum 23 diamonds in each page)
for index in range(23):
each_listing = []
#try to get a specific diamond's data in html
try:
current_diamond = listing_divs.select('div[id*=_{page}_{index}]'.format(page=page, index=index))[0]
except:
break
#scraping diamond features
#url
try:
url = current_diamond.select('a[class*=image-container]')[0]['href']
url = 'https://www.jamesallen.com/'+url
#scrapping the diamond's page
r = requests.get(url)
soup = bs(r.content, "html5lib")
#get the diamond's information table
try:
data = soup.select('div[class*=tablesContainer]')[0]
except:
break
#get the diamond's attributes:
try:
carat = data.select('div[data-qa*=stone_carat_value]')[0].string
except:
carat = None
try:
color = data.select('div[data-qa*=stone_color_value]')[0].string
except:
color = None
try:
clarity = data.select('div[data-qa*=stone_clarity_value]')[0].string
except:
clarity = None
try:
cut = data.select('div[data-qa*=stone_cut_value]')[0].string
except:
cut = None
try:
polish = data.select('div[data-qa*=stone_polish_value]')[0].string
except:
polish = None
try:
shape = data.select('div[data-qa*=stone_shape_value]')[0].string
except:
shape = None
try:
symmetry = data.select('div[data-qa*=stone_symmetry_value]')[0].string
except:
symmetry = None
try:
fluorescence = data.select('div[data-qa*=stone_fluorescence_value]')[0].string
except:
fluorescence = None
try:
lw = data.select('div[data-qa*=stone_lw_value]')[0].string
except:
lw = None
try:
lw_ratio = data.select('div[data-qa*=stone_lw_ratio_value]')[0].string
except:
lw_ratio = None
try:
certificate = data.select('div[data-qa*=stone_certificate_value]')[0].string
except:
certificate = None
#we scrapped the price from the original page
price = current_diamond.select('div[class*=salePrice]')
if len(price)==0:
price = current_diamond.select('div[class*=price]')
price = price[0].string
#appending the diamond's attributes to the list
each_listing.append(url)
each_listing.append(carat)
each_listing.append(color)
each_listing.append(clarity)
each_listing.append(cut)
each_listing.append(polish)
each_listing.append(shape)
each_listing.append(symmetry)
each_listing.append(fluorescence)
each_listing.append(lw)
each_listing.append(lw_ratio)
each_listing.append(certificate)
each_listing.append(price)
listing_list.append(each_listing)
except:
break
#return the list of all the diamonds in this page
return listing_list
# +
#scraping random pages from price range.
# this function goes through the range of the prices (from 200-100000) and genarate random pages for scraping
def scrape(prices):
#the beginning of the url
url_prefix = 'http://www.jamesallen.com/loose-diamonds/all-diamonds/page-'
#a loop which generate a random page in a range of 200 dollar price
for start_price in prices:
print('scraping page number' + str(start_price//200) + '...(price range : $' + str(start_price) + '-$' + str(start_price+200) + ')')
#generate a random number
random_page = random.randint(1,20) ## there are maximum 20 pages in each search.
#generate the url:
#special case for the first page(the url is a little different)
if random_page==1:
target_page ='http://www.jamesallen.com/loose-diamonds/all-diamonds/?Color=M,L,K,J,I,H,G,F,E,D&Cut=Good,Very+Good,Ideal,TrueHearts&Shape=all-diamonds&Clarity=I1,SI2,SI1,VS2,VS1,VVS2,VVS1,IF,FL&PriceFrom=' + str(start_price)+ '&PriceTo=' + str(start_price+200)+ '&CaratFrom=0.05'
#other pages
else:
target_page = url_prefix + str(random_page) + '/?Color=M,L,K,J,I,H,G,F,E,D&Cut=Good,Very+Good,Ideal,TrueHearts&Shape=all-diamonds&Clarity=I1,SI2,SI1,VS2,VS1,VVS2,VVS1,IF,FL&PriceFrom=' + str(start_price)+ '&PriceTo=' + str(start_price+200)+ '&CaratFrom=0.05'
#scrapping the page html
r = requests.get(target_page)
# Getting a BeautifulSoup instance to be able to retrieve data
soup = bs(r.content, "html5lib")
#get the diamonds in this page
listing_divs = soup.find('div', attrs={'id':'data-page-container'})
#in case this page doent exit/empty try the first page in this search
if listing_divs == None:
print('This random generated page did not found, lets try to scrape the first page...')
random_page=1
r = requests.get('http://www.jamesallen.com/loose-diamonds/all-diamonds/?Color=M,L,K,J,I,H,G,F,E,D&Cut=Good,Very+Good,Ideal,TrueHearts&Shape=all-diamonds&Clarity=I1,SI2,SI1,VS2,VS1,VVS2,VVS1,IF,FL&PriceFrom=' + str(start_price)+ '&PriceTo=' + str(start_price+200)+ '&CaratFrom=0.05')
# Getting a BeautifulSoup instance to be able to retrieve data
soup = bs(r.content, "html5lib")
listing_divs = soup.find('div', attrs={'id':'data-page-container'})
#in case the 1st page doent exit/empty continue to the next price range
if listing_divs == None:
continue
#now we have a page with data, let's send it to retrieve_data fun to get the infos of this page's diamonds
one_page_parsed = retrieve_data(random_page, listing_divs)
#create a dataframe from the data and add it to our local csv file
df = pd.DataFrame(one_page_parsed, columns=['url','carat','color','clarity','cut', 'polish', 'shape', 'symmetry', 'fluorescence', 'lw', 'lw_ratio','certificate', 'price'])
df.to_csv('scraped_Diamonds', mode='a', header=False)
# -
#creating a local csv file in our computer with the column labels
file = pd.DataFrame(columns=['url','carat','color','clarity','cut', 'polish', 'shape', 'symmetry', 'fluorescence', 'lw', 'lw_ratio','certificate', 'price'])
file.to_csv('scraped_Diamonds')
#creating a list of prices with skips of 200 dollars
prices = [i for i in range(200, 100001, 200)] ##diamonds till 100,000 dollars.
#start scraping
scrape(prices)
| Diamonds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from expresso.pycas import *
# +
f = Function('f',argc = 1)
g = Function('g',argc = 2)
x,y = symbols("x, y",type=Types.Real)
z = Symbol('z',type=Types.Complex)
# +
import numpy as np
def my_function_python(x,y):
return (abs(x)/(1+abs(y)))%1
my_function_ccode = '''
double myfunction(double x,double y){
return fmod(abs(x)/(1+abs(y)),1);
}
'''
pycas_custom_function = custom_function("myfunction",
argc = 2,
python_function=my_function_python,
ccode = my_function_ccode,
return_type=Types.Real)
# -
import numpy as np
npx,npy = np.meshgrid(np.linspace(-10, 10, 1001),np.linspace(-10, 10, 950))
snpx = array('np_x',npx+2*np.random.rand(*npy.shape))
p = parameter('p',1)
expr = piecewise((pi*snpx(sin(x)*cos(y)*1000,y*50),y>0),(e*pycas_custom_function(y-p*x,x+p*y)*10,True))
expr
fs_def = FunctionDefinition('f_single',(x,y),expr,return_type=Types.Real,parallel=False)
fp_def = FunctionDefinition('f_parallel',(x,y),expr,return_type=Types.Real,parallel=True)
clib = ccompile(fp_def,fs_def)
# +
p.set_value(2)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.imshow(clib.f_parallel(npx,npy))
plt.colorbar();
# -
nlib = ncompile(fp_def,fs_def)
plt.imshow(nlib.f_parallel(npx,npy))
plt.colorbar();
#plt.imshow(nlib.f_single(npx,npy) - clib.f_parallel(npx,npy))
#plt.colorbar();
npx[ np.where(abs(nlib.f_parallel(npx,npy) - clib.f_parallel(npx,npy)) != 0) ],npy[ np.where(abs(nlib.f_parallel(npx,npy) - clib.f_parallel(npx,npy)) != 0) ]
# +
lf = lambdify(expr)
N = mpmathify(expr)
for (vx,vy) in zip(10*(np.random.rand(1000)-0.5),10*(np.random.rand(1000)-0.5)):
assert np.isclose(nlib.f_single(vx,vy),lf(x=vx,y=vy))
assert np.isclose(lf(x=vx,y=vy),float(N(x=vx,y=vy)))
# +
lib_f_single_t = clib.f_single(npx,npy)
lib_f_parallel_t = clib.f_parallel(npx,npy)
cf_single_t = nlib.f_single(npx,npy)
cf_parallel_t = nlib.f_parallel(npx,npy)
assert np.allclose(lib_f_single_t, lib_f_parallel_t)
assert np.allclose(cf_single_t, cf_parallel_t)
assert np.allclose(cf_parallel_t, lib_f_single_t)
# -
# %timeit nlib.f_single(npx,npy)
# %timeit nlib.f_parallel(npx,npy)
# %timeit clib.f_single(npx,npy)
# %timeit clib.f_parallel(npx,npy)
| examples/PyCAS Compiler Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Intel® Extension for Scikit-learn KNN for MNIST dataset
from time import time
from sklearn import metrics
from sklearn.model_selection import train_test_split
from sklearn.datasets import fetch_openml
x, y = fetch_openml(name='mnist_784', return_X_y=True)
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2, random_state=72)
# Intel Extension for Scikit-learn (previously known as daal4py) contains drop-in replacement functionality for the stock scikit-learn package. You can take advantage of the performance optimizations of Intel Extension for Scikit-learn by adding just two lines of code before the usual scikit-learn imports:
from sklearnex import patch_sklearn
patch_sklearn()
# Intel(R) Extension for Scikit-learn patching affects performance of specific Scikit-learn functionality. Refer to the [list of supported algorithms and parameters](https://intel.github.io/scikit-learn-intelex/algorithms.html) for details. In cases when unsupported parameters are used, the package fallbacks into original Scikit-learn. If the patching does not cover your scenarios, [submit an issue on GitHub](https://github.com/intel/scikit-learn-intelex/issues).
params = {
'n_neighbors': 40,
'weights': 'distance',
'n_jobs': -1
}
# Training and predict KNN algorithm with Intel(R) Extension for Scikit-learn for MNIST dataset
start = time()
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(**params).fit(x_train, y_train)
predicted = knn.predict(x_test)
f"Intel(R) extension for Scikit-learn time: {(time() - start):.2f} s"
report = metrics.classification_report(y_test, predicted)
print(f"Classification report for KNN:\n{report}\n")
# *The first column of the classification report above is the class labels.*
#
# In order to cancel optimizations, we use *unpatch_sklearn* and reimport the class KNeighborsClassifier.
from sklearnex import unpatch_sklearn
unpatch_sklearn()
# Training and predict KNN algorithm with original scikit-learn library for MNSIT dataset
start = time()
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(**params).fit(x_train, y_train)
predicted = knn.predict(x_test)
f"Original Scikit-learn time: {(time() - start):.2f} s"
report = metrics.classification_report(y_test, predicted)
print(f"Classification report for KNN:\n{report}\n")
# With scikit-learn-intelex patching you can:
#
# - Use your scikit-learn code for training and prediction with minimal changes (a couple of lines of code);
# - Fast execution training and prediction of scikit-learn models;
# - Get the same quality;
# - Get speedup more than **24** times.
| examples/notebooks/knn_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Questionário 43 (Q43)
#
#
# Orientações:
#
# - Registre suas respostas no questionário de mesmo nome no SIGAA.
# - O tempo de registro das respostas no questionário será de 10 minutos. Portanto, resolva primeiro as questões e depois registre-as.
# - Haverá apenas 1 (uma) tentativa de resposta.
# - Submeta seu arquivo-fonte (utilizado para resolver as questões) em formato _.ipynb_ pelo SIGAA anexando-o à Tarefa denominada "Envio de arquivo" correspondente ao questionário.
#
# *Nota:* o arquivo-fonte será utilizado apenas como prova de execução da tarefa. Nenhuma avaliação será feita quanto ao estilo de programação.
#
# <hr>
# Para responder este questionário, utilize o banco de dados [brasileirao2021.csv](https://github.com/gcpeixoto/ICD/blob/main/database/brasileirao2021.csv). Fonte: [[CBF]](https://www.cbf.com.br/futebol-brasileiro/competicoes/campeonato-brasileiro-serie-a).
#
# **Obs.:** use o _dataset_ do repositório Git e não o do site da CBF, visto que este é atualizado após cada jogo.
#
# <br>
#
# **Questão 1.** Utilizando o método de _z-scores_ e o _dataset_, identifique todos os times cuja pontuação superou a média do campeonato e assinale a alternativa correta quanto às posições que ocupavam no ranque do Brasileirão 2021 no momento em que o _dataset_ havia sido gerado.
#
# A. 1a. a 6a.
#
# B. 3a. a 5a.
#
# C. 1a. a 9a.
#
# D. 2a. a 8a.
# **Questão 2.** O _dataset_ descreve o desempenho de cada time através de marcadores clássicos do futebol, a saber: Pontos (_PTS_), Jogos (_J_), Vitórias (_V_), Empates (_E_), Derrotas (_D_), Gols Marcados (Pró) (_GP_), Gol Sofridos (Contra) (_GC_), Saldo de Gols (_SG_), Cartões Amarelos (_CA_) e Cartões Vermelhos (_CV_).
#
# Considerando $X$ a série correspondente a _PTS_, determine as variáveis correspondentes às séries $Y_1$ e $Y_2$, tais que, $\text{cov}(X,Y_1)$ seja a maior covariância positiva e $\text{cov}(X,Y_2)$ seja a maior covariância negativa.
#
#
# A. _J_ e _V_
#
# B. _GP_ e _GC_
#
# C. _SG_ e _GP_
#
# D. _SG_ e _GC_
#
# **Questão 3.** Considere o _DataFrame_ processado para a resolução da Questão 2. Entre as
# séries Gols Marcados (Pró) (_GP_), Gol Sofridos (Contra) (_GC_), Saldo de Gols (_SG_), Cartões Amarelos (_CA_) e Cartões Vermelhos (_CV_), identifique a que possui a mais forte correlação positiva com _E_ e a que possui a mais forte correlação negativa com _V_, respectivamente. Assinale a alternativa correta.
#
# A. _GP_ e _GC_
#
# B. _CV_ e _GP_
#
# C. _CA_ e _GC_
#
# D. _SG_ e _GC_
| _build/jupyter_execute/ipynb/Q43.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
headers = {
'User-Agent': 'Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/92.0.4515.159 Safari/537.36',
}
url = "https://www.vmware.com/bin/vmware/getadvisorieslist?resourcePath=/content/vmware/vmware-published-sites/us/security/advisories"
json_data = requests.get(url, headers=headers).json()
json_data["data"]
| VMware-Security-Advisories-WebCrawler.ipynb |
# ---
# jupyter:
# jupytext:
# formats: notebooks//ipynb,rmd//Rmd
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Use RMarkdown...
# ...when you want to weave together code (it doesn't have to be R!), and narrative (efficiently written in Markdown).
# ## With Python
# Let's demonstrate with the classic entrypoint to Python:
print('hello world')
# And now we'll do something a tiny bit more complicated: use numpy to generate an array of twenty random numbers, which we'll then use matplotlit to plot.
import numpy as np
import matplotlib.pyplot as plt
numbers = np.random.rand(20,1)
plt.plot(numbers)
plt.show()
# Now let's add a citation (I'm using Zotero, with the BetterBibTex plugin, and citation keys in the format `[authForeIni][authEtAl][year]`, and then exporting the bibliography as `refs.bib`, which needs to be saved in our `bits` folder) -- maybe something about Jupyter notebooks [@TKluyverEtAl2016] -- and we can run our bash script to turn this into a publishable PDF...
# +
# to run this from within the notebook, first comment out this line and save...
# ...and then uncomment the line to run it -- otherwise, Pweave will get stuck...
# ...in an infinite loop and be unable to finish processing the notebook
# # ! ../bits/publi.sh RMarkdown
# -
# ## References
| notebooks/RMarkdown.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vCvuDQIZVrg0"
# # Data Science Championship - South Zone
#
# > "Predicting the house rent of the given house details"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [predict, machine, hack, the, house, rent, price, data, science, championship, south, zone, learning]
# - hide: false
# + colab={"base_uri": "https://localhost:8080/"} id="7UeCd7W-AfMG" outputId="b5f3d7a9-672f-42c1-fd57-d89b3d97d873"
# Installing the modules
# !pip3 install category_encoders
# + id="QsV1vsJu5oWr" colab={"base_uri": "https://localhost:8080/"} outputId="289cab47-e541-46f9-f27b-d5dce9bdc83e"
# Required modules
import shutil
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import category_encoders as ce
from google.colab import drive
from matplotlib import pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
# + id="dAzDbAv050r5"
# Config
# %matplotlib inline
plt.rcParams['figure.figsize'] = (12, 12)
pd.set_option('display.max_columns', None)
# + colab={"base_uri": "https://localhost:8080/"} id="k7fUjICd57Q5" outputId="5ea56c5c-eebd-445a-c347-00a23e79265c"
# Mounting the drive
drive.mount('./mydrive')
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="9Zwv2tqP8Pdf" outputId="2864619e-748b-4a67-8031-ffe9ae59ae56"
# Moving files to workspace
shutil.copy('/content/mydrive/MyDrive/Machine Hack/Data Science Students Championship/train.csv', './train.csv')
shutil.copy('/content/mydrive/MyDrive/Machine Hack/Data Science Students Championship/test.csv', './test.csv')
shutil.copy('/content/mydrive/MyDrive/Machine Hack/Data Science Students Championship/submission.csv', './submission.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="_9nvPtYJ8pLT" outputId="2e0d8f55-01e4-4e87-f58e-eae1d84d03f2"
# Load the data
train = pd.read_csv('train.csv')
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 850} id="AMHGVz128ykU" outputId="06b05aff-9598-4ef7-a4bd-d8c8669c19b7"
# Inspect the data
train.info()
train.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 322} id="4gXgDrxl84qZ" outputId="dfbfad90-1ce1-4f5c-ac6c-bb5da2922431"
# Load the test data
test = pd.read_csv('./test.csv')
test.head()
# + colab={"base_uri": "https://localhost:8080/"} id="1eEAPF4vB7QR" outputId="0110778d-cad0-4a77-ce61-623c84a18daf"
# Checking for the missing values
if train.isna().any().any():
print(train.isna().any())
else:
print("No Missing Values")
# + [markdown] id="WWv1P63aCMpo"
# ## Feature Engineering
# + id="6l-1ybiF9ke7"
# Converting the categorical variables
train['layout_type'] = np.where(train['layout_type'] == 'BHK', 1, 0)
test['layout_type'] = np.where(test['layout_type'] == 'BHK', 1, 0)
for col in train.columns:
if train[col].dtype == 'object':
encoder = ce.cat_boost.CatBoostEncoder()
train[col] = encoder.fit_transform(train[col], train['price'])
test[col] = encoder.transform(test[col])
# + id="zlpri2CuCdwW"
# Seperating out features and labels
X = train.drop(['Property_ID', 'price'], axis=1)
y = train['price']
X_test = test.drop(['Property_ID', 'price'], axis=1)
# + id="4pPdOFBkMTp_"
# Scaling the train and test values
scaler = MinMaxScaler(feature_range=(0, 1))
X_scaled = scaler.fit_transform(X)
X_test_sclaed = scaler.transform(X_test)
scaler_label = MinMaxScaler(feature_range=(0, 1))
scaler_label.fit(y.values.reshape(-1, 1))
y_scaled = scaler_label.transform(y.values.reshape(-1, 1))
# + id="Bb_0V9-JCr6C"
# Train and test split
X_train, X_valid, y_train, y_valid = train_test_split(X_scaled, y_scaled, test_size=0.2, random_state=88)
# + [markdown] id="Gp4JLIKwCQLp"
# ## Model Building
# + colab={"base_uri": "https://localhost:8080/"} id="wy-crM5JWJqc" outputId="d9eaa2ac-14e8-45c6-f0eb-60a814980eb2"
# Model Definition
input_len = len(train.columns) - 2
model = tf.keras.models.Sequential([
tf.keras.layers.Input(shape=(input_len,)),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(32, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(8, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(1, activation='linear'),
])
model.summary()
# + id="s0gKrifkXrA0"
# Custom loss
def rmse(y_true, y_pred):
return tf.math.sqrt(tf.keras.losses.mean_squared_error(y_true, y_pred))
# + id="YNMQUAD9aeRC"
# Compiling the model
loss = rmse
optim = tf.keras.optimizers.Adam(learning_rate=0.0001)
checkpoint_callback = tf.keras.callbacks.ModelCheckpoint('custom_model_checkpoint.hdf5', save_best_only=True, custom_objects=[rmse])
model.compile(optimizer=optim, loss=loss, metrics=[tf.keras.losses.mean_squared_error])
# + colab={"base_uri": "https://localhost:8080/"} id="kFGADtutBsqx" outputId="66ee4526-f74d-4e1e-8afc-f8edc190fead"
# Fitting the model
epochs = 20
batch_size = 64
model.fit(X_train, y_train, validation_data=(X_valid, y_valid), epochs=epochs, batch_size=batch_size, shuffle=True)
# + colab={"base_uri": "https://localhost:8080/"} id="Uo2yMRfFFtBS" outputId="d8bf1fb1-f745-47d1-aa7b-ae4bb7aba787"
# Scoring the partitions
print(f"RMSE of Train is {mean_squared_error(scaler_label.inverse_transform(model.predict(X_train)), scaler_label.inverse_transform(y_train), squared=False)}")
print(f"RMSE of Valid is {mean_squared_error(scaler_label.inverse_transform(model.predict(X_valid)), scaler_label.inverse_transform(y_valid), squared=False)}")
# + id="ey2kvsYoG1Oq"
# Generate the submission file
test = pd.DataFrame(index=range(X_test.shape[0]))
test['price'] = model.predict(X_test)
test.to_csv('submission.csv', index=False)
| _notebooks/2022-05-14-Data-Science-Championship.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import import seaborn as sns
# %matplotlib inline
import numpy as np
employee = pd.read_csv("WA_Fn-UseC_-HR-Employee-Attrition.csv")
print(employee)
#evaluate first five rows
employee.head()
#describing dataset
employee.describe()
#job role
display(employee.isnull().any())
pd.crosstab(employee['JobRole'],employee['Attrition']).plot(kind='bar',stacked=False)
plt.title('Attrition with respect to JobRole')
plt.xlabel('JobRole')
plt.ylabel('Frequency of Attrition')
pd.crosstab(employee['DistanceFromHome'],employee['Attrition']).plot(kind='bar',stacked=False)
plt.title('Attrition with respect to DistanceFromHome')
plt.xlabel('EducationField')
plt.ylabel('Frequency of Attrition')
pd.crosstab(employee['AverageMonthlyIncome'],employee['Attrition']).plot(kind='bar',stacked=False)
plt.title('Attrition with respect to DistanceFromHome')
plt.xlabel('EducationField')
plt.ylabel('Frequency of Attrition')
| Assignment 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pylab as plt
plt.style.use('fivethirtyeight')
from animals import Island, Rabbit
import random
# # Testing
R = Rabbit(10)
R.age
R._age()
R.age
random.seed(123)
R2 = R.breed()
R2.survival_skill
I = Island(init_rabbits=10, max_pop=100)
I.rabbits
stats = I.compute_epoches(15)
stats[14]
# # Thousand Islands
# +
params = {'init_rabbits':10, 'max_pop':40}
years, N_islands = 15, 1000
islands = [Island(**params) for _ in range(N_islands)]
stats = [ island.compute_epoches(years) for island in islands]
# -
# # Harsh Islands
from animals import HarshIsland
# +
params = {'init_rabbits':10, 'max_pop':40, 'env_range':[10,90]}
years, N_islands = 15, 1000
h_islands = [HarshIsland(**params) for _ in range(N_islands)]
h_stats = [ island.compute_epoches(years) for island in h_islands]
# -
# ## Visualisation
# +
fig, axes = plt.subplots(4,2, figsize=(10,10), sharex=True)
for i, title in enumerate(('Population', 'Average age', 'Average Survival Skill', '% of rabbits with SSK > 75')):
axes[i][0].set_ylabel(title)
for i, (k, v) in enumerate({"Heaven Islands":stats,
'Harsh Islands':h_stats}.items()):
axes[0][i].set_title(k)
for s in v: # for each island
years = list(s.keys())
axes[0][i].plot(years, [v['pop'] for v in s.values()], c='red', alpha=.005)
axes[1][i].plot(years, [v.get('mean_age', None) for v in s.values()], c='blue', alpha=.005)
axes[2][i].plot(years, [v.get('mean_skill', None) for v in s.values()], c='green', alpha=.005)
axes[3][i].plot(years, [v.get('75_skill', None) for v in s.values()], c='purple', alpha=.005)
# -
| Chapter08/8_2_Simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # part 4: descriptive statistics and bariclot calculation
#
# * create summary statistics of the entire dataset using
# * compare training, validation, and testing sets for both leak and clot
# * calculate bariclot score for each patient in the clot training set
#
# this will generate
#
# * a `results` directory
# * a `results/descriptive_stats` subdirectory - holds all descriptive data generated by tableone
# * a `results/bariclot` subdirectory - holds bariclot results for the test cohort. we also produce a sanity check for bariclot on 2015 data to make sure it performs as expected
# preliminaries
# +
import pandas as pd
import numpy as np
import python_modules.constants as constants
import os
from tableone import TableOne
# +
np.random.seed(seed=1872)
# +
# Set ipython's max row display
pd.set_option('display.max_row', 100)
# Set iPython's max column display
pd.set_option('display.max_columns', 50)
# +
PATH_IMPORT = 'study_data/'
PATH_RESULTS = 'results/'
PATH_DESCRIPTIVE = 'results/descriptive_stats/'
PATH_BARICLOT = 'results/bariclot/'
# make dirs to hold outputs
# if folders already exist, this will throw errors
os.mkdir(f'{PATH_RESULTS}')
os.mkdir(f'{PATH_DESCRIPTIVE}')
os.mkdir(f'{PATH_BARICLOT}')
# -
# import data, specify variables to be included in table 1
# +
df_main = pd.read_csv(f'{PATH_IMPORT}/study_data_split.csv', low_memory=False, index_col=0)
# +
#labs_type = 'continuous'
#lim_intra = False
#
#table_one_cats = constants.categorical(labs = labs_type, lim_intra = lim_intra) + ['LEAK', 'CLOT']
#table_one_cons = constants.continuous(labs = labs_type, lim_intra = lim_intra)
#table_one_include = table_one_cats + table_one_cons
table_one_cats = constants.CATEGORICAL_PRE + constants.OUTCOME
table_one_cons = constants.CONTINUOUS_PRE + constants.CONTINUOUS_POST
table_one_include = table_one_cats + table_one_cons
# -
# ## descriptive statistics
# we split the data into quarters in the last notebook and somewhat confusingly named part of the training set `val_1`. therefore create another row to map the actual training, validation, and testing sets, ultimately creating the final training set by adding `val_1` patients to `train` patients.
# +
final_analysis_pop_dict = {'train':0, 'val_1':0, 'val_2':1, 'test': 2}
df_main['consolidated_clot_groups'] = df_main.CLOT_SET.map(final_analysis_pop_dict)
df_main['consolidated_leak_groups'] = df_main.LEAK_SET.map(final_analysis_pop_dict)
# -
# build summary tables
mytable = TableOne(df_main, table_one_include, table_one_cats)
mytable_clot = TableOne(df_main, table_one_include, table_one_cats, 'consolidated_clot_groups', pval=True)
mytable_leak = TableOne(df_main, table_one_include, table_one_cats, 'consolidated_leak_groups', pval=True)
# save them
#
#
mytable.to_csv(f'{PATH_DESCRIPTIVE}dataset_summary.csv')
mytable_clot.to_csv(f'{PATH_DESCRIPTIVE}clot_set_summary.csv')
mytable_leak.to_csv(f'{PATH_DESCRIPTIVE}leak_set_summary.csv')
# ## bariclot calculation
# dicts and functions to parse data into format needed for bariclot calculation
# +
d_funcstat = {'Independent':0, 'Partially Dependent':1, 'Totally Dependent':2}
d_clothist = {'No':0, 'Yes':1}
def d_race(race):
if race == 'Black or African American':
return 1
else:
return 0
# -
# bariclot calculation
# * <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Predicting venous thromboembolism following laparoscopic bariatric surgery: development of the BariClot tool using the MBSAQIP database. Surg Endosc. 2018; PMID:30003351; http://dx.doi.org/10.1007/s00464-018-6348-0
#
# +
def calculate_bariclot(row):
w = d_funcstat[row.FUNSTATPRESURG] * 3
x = d_clothist[row.HISTORY_DVT] * 9
y = d_race(row.race_PUF) * 3
z = row.OPLENGTH / 60
return np.sum([w,x,y,z])
def calculate_bariclot_groups(row):
bc = calculate_bariclot(row)
if bc < 1:
return 0
elif bc < 7:
return 1
elif bc < 10:
return 2
else:
return 3
# -
# ### test population
# some patients (333 in the total study cohort) have missing data for operative duration; replace these with the mean operative duration.
df_main['OPLENGTH'].fillna(df_main['OPLENGTH'].mean(), inplace = True)
# select down to clot test population
# +
df_test_clot = df_main[df_main['CLOT_SET'] == 'test']
# -
len(df_test_clot)
# ### run the calculator
bariclot_scores = df_test_clot.apply(calculate_bariclot, axis=1)
bariclot_score_groups = df_test_clot.apply(calculate_bariclot_groups, axis=1)
# save targets and values for post-processing in R
# +
#dataframe to hold results
df_bariclot = pd.DataFrame()
#populate dataframe with bariclot scores and target outcomes
df_bariclot['scores'] = bariclot_scores
df_bariclot['targs'] = df_test_clot['CLOT']
# +
df_bariclot = df_bariclot.reset_index(drop=True)
# -
df_bariclot.to_csv(f'{PATH_BARICLOT}bariclot_test.csv')
# ### look at auc for 2015 population just as a sanity check
#
# (it checks out when we run stats in R)
# +
df_2015_clot = df_main[df_main['OPYEAR'] == 2015]
# +
bariclot_scores_2015 = df_2015_clot.apply(calculate_bariclot, axis=1)
df_bariclot_2015 = pd.DataFrame()
df_bariclot_2015['scores'] = bariclot_scores_2015
df_bariclot_2015['targs'] = df_2015_clot['CLOT']
df_bariclot_2015 = df_bariclot_2015.reset_index(drop=True)
df_bariclot_2015.to_csv(f'{PATH_BARICLOT}bariclot_2015.csv')
| part4_descriptive-statistics-bariclot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GAN review
# ## Problem Setup
# Let $x$ be element from the data space and let $z$ be an element from the latent space.
# In GAN, we have a discriminator $D(x;\theta_d)$ and generator $G(z;\theta_g)$.
# The generator $G(z;\theta_g)$ generates fake data, which depends on the latent variable $z$ that is sampled from noise prior $p_z(z)$.
# The discriminator $D(x;\theta_d)$ examines an element from the data space and returns a probability that it is real or fake.
# The goal is to train the $G$ and the $D$ such that each becomes good at its task.
# ## Algorithm
# repeat k times:
# > **For discriminator:** \
# sample m noise values $\{ z_1, ..., z_m \}$ from noise prior $p_g(z)$ \
# sample m examples $\{ x_1, ..., x_m \}$ from data set. \
# update discriminator parameters $\theta_d$ using loss function $-\frac{1}{m} \sum_{i=1}^{m} [\log D(x_i) + \log(1-D(G(z_i)))]$ \
# **For generator:** \
# sample m noise values $\{ z_1, ..., z_m \}$ from noise prior $p_g(z)$ \
# update generator parameters $\theta_g$ using loss function $\frac{1}{m} \sum_{i=1}^m \log(1-D(G(z_i)))$
# In practice, we use $\frac{1}{m} \sum_{i=1}^m \log(D(G(z_i)))$, which provides larger gradient to the generator.
# This is because during the initial phase of the training, the discriminator has easier time distinguishing the fake data from real data,
# Therefore, D(G) tends to be small, and $\log(D(G))$ provides larger gradient than $\log (1-D(G))$ during the initial phase of training (see loss graph)
# >
# ## Notes
# Let $x=g(z)$, $p_g(x)dx=p_z(z)dz$ (as an integration measure): $p_g(x)$ is the probability distribution of fake data generated by $G$. Ideally, we have $p_g=p_{data}$
# Setup environment
from matplotlib import pyplot as plt
import numpy as np
import math
import torch
import torch.nn as nn
import torch.nn.functional as F
from torchvision import datasets, transforms
import torch.optim as optim
import datetime
from tqdm import tqdm
from maxout import Maxout
device = torch.device('cuda' if torch.cuda.is_available()
else torch.device('cpu'))
print(f"Training on {device}.")
# +
# load data
batch_size=128
# normalize image such that the pixels lie between -1 and 1 (generator outputs lie between -1 and 1)
train_loader = torch.utils.data.DataLoader(
datasets.MNIST('../MNIST_data', train=True, download=False,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))
])),
batch_size=batch_size, shuffle=True,pin_memory=True)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST('../MNIST_data', train=False, download=False,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))
])),
batch_size=batch_size, shuffle=True, pin_memory=True)
# -
# ### Notes
# We provide fully connected network and maxout network for discriminator. The original paper uses maxout network (with a slightly different design).
# +
class generator(nn.Module):
""" The generator network in GAN
Args:
z_dim: dimension of latent space
G_h1_dim: dimension of first hidden layer
G_h2_dim: dimension of second hidden layer
G_h3_dim: dimension of third hidden layer
G_out_dim: dimension of output space (for MNIST, this should be 28*28)
Returns:
subclass of nn.Module: generator network"""
def __init__(self,z_dim,G_h1_dim, G_h2_dim, G_h3_dim, G_out_dim):
super().__init__()
self.G_FC1 = nn.Linear(z_dim, G_h1_dim)
self.G_FC2 = nn.Linear(G_h1_dim, G_h2_dim)
self.G_FC3 = nn.Linear(G_h2_dim,G_h3_dim)
self.G_FC4 = nn.Linear(G_h3_dim,G_out_dim)
self._init_weights(irange=0.05)
def _init_weights(self,irange):
"""Overrides built-in initialization. The original paper uses irange of 0.05"""
for m in self.modules():
if type(m) in {nn.Linear}:
nn.init.uniform_(m.weight.data,-irange,irange)
if m.bias is not None:
torch.zero_(m.bias.data)
def forward(self,z_in):
out = F.relu(self.G_FC1(z_in))
out = F.relu(self.G_FC2(out))
out = F.relu(self.G_FC3(out))
out = torch.tanh(self.G_FC4(out))
return out
class discriminator(nn.Module):
""" The discriminator network in GAN
Args:
mode: for fully connected, use 'FC' and for maxout, use 'MO'
x_dim: dimension of input (for MNIST, this should be 28*28)
D_h1_dim: dimension of first hidden layer
D_h2_dim: dimension of second hidden layer
D_h3_dim: dimension of third hidden layer
Returns:
subclass of nn.Module: discriminator network"""
def __init__(self,mode,x_dim,D_h1_dim,D_h2_dim,D_h3_dim):
super().__init__()
self.mode = mode
self.x_dim = x_dim
if self.mode == 'FC':
self.D_FC1 = nn.Linear(x_dim,D_h1_dim)
self.D_FC2 = nn.Linear(D_h1_dim,D_h2_dim)
self.D_FC3 = nn.Linear(D_h2_dim,D_h3_dim)
self.D_FC4 = nn.Linear(D_h3_dim,1)
elif self.mode == 'MO':
self.D_MO1 = Maxout(x_dim,D_h1_dim,5)
self.D_MO2 = Maxout(D_h1_dim,D_h2_dim,5)
self.D_MO3 = Maxout(D_h2_dim,D_h3_dim,5)
self.D_FC1 = nn.Linear(D_h3_dim,1)
else:
raise NotImplementedError
self._init_weights(irange=0.005)
def _init_weights(self,irange):
"""Overrides built-in initialization. The original paper uses irange of 0.005"""
for m in self.modules():
if type(m) in {nn.Linear}:
nn.init.uniform_(m.weight.data,-irange,irange)
if m.bias is not None:
torch.zero_(m.bias.data)
def forward(self,x_in):
if self.mode == 'FC':
out = F.relu(self.D_FC1(x_in))
out = F.dropout(out,p=0.3)
out = F.relu(self.D_FC2(out))
out = F.dropout(out,p=0.3)
out = F.relu(self.D_FC3(out))
out = F.dropout(out,p=0.3)
out = self.D_FC4(out)
out = torch.sigmoid(out)
elif self.mode == 'MO':
out = F.dropout(self.D_MO1(x_in.view(-1,self.x_dim)),p=0.3)
out = F.dropout(self.D_MO2(out),p=0.3)
out = F.dropout(self.D_MO3(out),p=0.3)
out = self.D_FC1(out)
out = torch.sigmoid(out)
else:
raise NotImplementedError
return out
def noise(batch_size,z_dim,device):
return torch.randn(batch_size,z_dim,device=device)
z_dim=100
gen = generator(z_dim=z_dim,G_h1_dim=256,G_h2_dim=512,G_h3_dim=1024,G_out_dim=28*28).to(device)
dis = discriminator(mode='MO',x_dim=28*28,D_h1_dim=512,D_h2_dim=256,D_h3_dim=128).to(device)
# dis = discriminator(mode='FC',x_dim=28*28,D_h1_dim=1048,D_h2_dim=512,D_h3_dim=256).to(device)
# +
optimizer_D = optim.Adam(dis.parameters(),lr = 0.0001)
optimizer_G = optim.Adam(gen.parameters(),lr = 0.0001)
def train_discriminator(imgs,train=True):
"""Train the discriminator network
Args:
imgs: minibatch of images
train: when set to False, used for validation
Returns:
mean loss
"""
if train:
gen.eval(), dis.train()
noise_d = noise(batch_size,z_dim,device)
loss_real = -torch.log(dis(imgs.view(-1,28*28))).mean()
loss_fake = -torch.log(1.-dis(gen(noise_d))).mean()
loss_d = loss_real+loss_fake
optimizer_D.zero_grad()
loss_d.backward()
optimizer_D.step()
else:
gen.eval(), dis.eval()
noise_d = noise(batch_size,z_dim,device)
loss_real = -torch.log(dis(imgs.view(-1,28*28))).mean()
loss_fake = -torch.log(1.-dis(gen(noise_d))).mean()
loss_d = loss_real+loss_fake
return loss_d.detach()
def train_generator(imgs,train=True):
"""Train the generator network
Args:
imgs: minibatch of images
train: when set to false, used for validation
Returns:
mean loss
"""
if train:
gen.train(), dis.eval()
noise_g = noise(batch_size,z_dim,device)
loss_g = -torch.log(dis(gen(noise_g))).mean()
optimizer_G.zero_grad()
loss_g.backward()
optimizer_G.step()
else:
gen.eval(), dis.eval()
noise_g = noise(batch_size,z_dim,device)
loss_g = -torch.log(dis(gen(noise_g))).mean()
return loss_g.detach()
def training_loop(epochs,test = False):
"""Train generator and discriminator
Args:
epochs: number of epochs
test: if True, evaluate on validation set
Returns:
List of average loss per epoch.
If test is True, also returns list of average loss for test set.
The first element in the list contains the discriminator loss,
the second element in the list contains the generator loss.
"""
loss_list_trn = [[],[]]
loss_list_val = [[],[]]
for epoch in tqdm(range(epochs),desc = f'percent of epochs completed'):
for imgs, label in train_loader:
loss_d_batch_list_trn = []
loss_g_batch_list_trn = []
imgs = imgs.to(device)
loss_d_batch_trn=train_discriminator(imgs)
loss_g_batch_trn=train_generator(imgs)
loss_d_batch_list_trn.append(loss_d_batch_trn.item())
loss_g_batch_list_trn.append(loss_g_batch_trn.item())
loss_list_trn[0].append(np.mean(loss_d_batch_list_trn))
loss_list_trn[1].append(np.mean(loss_g_batch_list_trn))
if test:
with torch.no_grad():
for imgs, label in test_loader:
loss_d_batch_list_val = []
loss_g_batch_list_val = []
imgs = imgs.to(device)
loss_d_batch_val = train_discriminator(imgs,train=False)
loss_g_batch_val = train_generator(imgs,train=False)
loss_d_batch_list_val.append(loss_d_batch_val.item())
loss_g_batch_list_val.append(loss_g_batch_val.item())
loss_list_val[0].append(np.mean(loss_d_batch_list_val))
loss_list_val[1].append(np.mean(loss_g_batch_list_val))
return loss_list_trn, loss_list_val
# -
# print("training loop starting at",datetime.datetime.now())
loss_list = training_loop(epochs=100,test=True)
# print("finished training loop at",datetime.datetime.now())
# how well does discriminator distinguish real from fake and how well does generator fool the discriminator?
dis.eval()
gen.eval()
for imgs,_ in train_loader:
imgs=imgs.to(device)
print(dis(imgs.view(-1,28*28)).mean(),dis(gen(noise(100,z_dim,device))).mean())
break
for imgs,_ in test_loader:
imgs=imgs.to(device)
print(dis(imgs.view(-1,28*28)).mean(),dis(gen(noise(100,z_dim,device))).mean())
break
# ## Notes
# Notice that the discriminator loss increase slightly.
# This is because initially, the discriminator has an easy time distinguishing the randomly generated fake image from real ones.
# However, the generator soon learns to fool the discriminator, and we reach an equilibrium.
fig, ax = plt.subplots(nrows=1, ncols=2,figsize=(11,5))
ax[0].plot(range(1,len(loss_list[0][0])+1),loss_list[0][0],'.r',label = 'discriminator',markersize=12)
ax[0].plot(range(1,len(loss_list[0][1])+1),loss_list[0][1],'.b',label = 'generator',markersize=12)
ax[0].set_xlabel('epoch')
ax[0].set_ylabel('train loss')
ax[0].legend()
ax[1].plot(range(1,len(loss_list[1][0])+1),loss_list[0][0],'.r',label = 'discriminator',markersize=12)
ax[1].plot(range(1,len(loss_list[1][1])+1),loss_list[1][1],'.b',label = 'generator',markersize=12)
ax[1].set_xlabel('epoch')
ax[1].set_ylabel('test loss')
ax[1].legend()
plt.tight_layout()
# save model
torch.save(gen.state_dict(),'gen_MO.pth')
torch.save(dis.state_dict(),'dis_MO.pth')
def generate_img(gen,n_img):
from torchvision.utils import save_image
gen.eval()
generated = gen(noise(n_img,z_dim,device))
generated=generated.detach().view(n_img,1,28,28).to('cpu')
print(generated.shape)
with torch.no_grad():
save_image(generated,'gen_MO_img.pdf')
# for i in range(10):
# plt.imshow(np.array(a[i].view(28,28).detach().to('cpu')),cmap='gray')
# plt.show()
generate_img(gen,10)
| GAN with maxout/GAN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/janchorowski/ml_uwr/blob/fall2019/assignment2/Assignment2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="6JGmPStV4yiw"
# # Lab Assignment 2
# + [markdown] colab_type="text" id="aUenJ9L141My"
# **Submission deadline:**
# * **Regular problems: last lab session before or on Monday, 18.11.19**
# * **Bonus problems: deadline for Lab Assignment 3**
#
# **Points: 12 + 7 bonus points**
#
# Please note: some of the assignments are tedious or boring if you are already a NumPy ninja. The bonus problems were designed to give you a more satisfying alternative.
#
# The assignment is in the form of a Jupyter notebook. We will be using [Google Colab](https://colab.research.google.com) to solve it. Below you will find a "Setup" section. Follow instructions from this paragraph to download the notebook and open it using [Google Colab](https://colab.research.google.com).
#
# Your goal is to solve problems posted below. Whenever possible, add your solutions to the notebook.
#
# Please email us about any problems with it - we will try to correct them quickly. Also, please do not hesitate to use GitHub’s pull requests to send us corrections!
# + colab={} colab_type="code" id="NsnbuW1uzVcC"
# Please note that this code needs only to be run in a fresh runtime.
# However, it can be rerun afterwards too.
# !pip install -q gdown httpimport
# + colab={} colab_type="code" id="a4TIgG0bwlpS"
# Standard IPython notebook imports
# %matplotlib inline
import os
import string
import random
from collections import OrderedDict
from io import StringIO
import httpimport
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from tqdm import tqdm_notebook
import scipy.stats as sstats
import seaborn as sns
import sklearn.tree
import sklearn.ensemble
import graphviz
# In this way we can import functions straight from github
with httpimport.github_repo('janchorowski', 'nn_assignments',
module='common', branch='nn18'):
from common.plotting import plot_mat
sns.set_style('whitegrid')
# + [markdown] colab_type="text" id="a7qCaa3LRuzJ"
# # Problem 1 [2p] Naive Bayes Classifier
#
# The Bayes' theorem allows us to construct a classifier in which we
# model how the data is generated. Here we will describe a
# simple and popular example of such a classifier called the naive
# Bayes classifier. Despite its simplicity It is quite effective for
# classification of text documents (e.g. as spam and non-spam).
#
# Let a document be a sequence of words $D=W_1,W_2,\ldots,W_n$
# We will model generation of text documents as a two-stage process.
# First, document category $C_j$ is drawn at random with probability
# $p(C_j)$, also called the *a priori* probability.
# To define the class-conditional probability
# $p(D|C_j)$, we will make a simplifying (naive)
# assumption, that every word in the document is drawn independently at
# random with probability $p(W_i|C)$:
#
# \begin{equation*}
# p(D|C_j) = p(W_1,W_2,\ldots,W_n | C_j) \approx p(W_1|C_j)p(W_2|C_j)\ldots p(W_n|C_j).
# \end{equation*}
#
# To infer the class of a document we apply the Bayes theorem:
# \begin{equation*} p(C_j|D) = \frac{p(D|C_j)p(C_j)}{p(D)} = \frac{p(C_j)p(W_1|C_j)p(W_2|C_j)\ldots p(W_n|C_j)}{p(D)}.
# \end{equation*}
# Please note that since we assumed only a finite number of classes,
# we can compute the term $p(D)$ by making sure that the *a
# posteriori probabilities* $p(C_j|D)$ sum to $1$ over all classes.
#
# In this exercise we will try to mimic the language-guessing feature
# of [Google Translate](https://translate.google.com/), although
# on a much smaller scale. We are given an input which is a
# lower-case sequence of characters (such as *"some people like
# pineapple on their pizza"*), and we determine whether the
# sequence's language is English, Polish or Spanish.
# We will treat each character as a separate observation.
# The numbers are taken from [Wikipedia article on letter frequency](https://en.wikipedia.org/wiki/Letter_frequency#Relative_frequencies_of_letters_in_other_languages). We display the first few rows:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="N-8g1QuNbIfs" outputId="62e46485-55ae-4bdb-a2ee-154e49f802c9"
wiki_table = u"""English|French|German|Spanish|Portuguese|Esperanto|Italian|Turkish|Swedish|Polish|Dutch|Danish|Icelandic|Finnish|Czech
a|8.167|7.636|6.516|11.525|14.634|12.117|11.745|12.920|9.383|10.503|7.486|6.025|10.110|12.217|8.421
b|1.492|0.901|1.886|2.215|1.043|0.980|0.927|2.844|1.535|1.740|1.584|2.000|1.043|0.281|0.822
c|2.782|3.260|2.732|4.019|3.882|0.776|4.501|1.463|1.486|3.895|1.242|0.565|0|0.281|0.740
d|4.253|3.669|5.076|5.010|4.992|3.044|3.736|5.206|4.702|3.725|5.933|5.858|1.575|1.043|3.475
e|12.702|14.715|16.396|12.181|12.570|8.995|11.792|9.912|10.149|7.352|18.91|15.453|6.418|7.968|7.562
f|2.228|1.066|1.656|0.692|1.023|1.037|1.153|0.461|2.027|0.143|0.805|2.406|3.013|0.194|0.084
g|2.015|0.866|3.009|1.768|1.303|1.171|1.644|1.253|2.862|1.731|3.403|4.077|4.241|0.392|0.092
h|6.094|0.737|4.577|0.703|0.781|0.384|0.636|1.212|2.090|1.015|2.380|1.621|1.871|1.851|1.356
i|6.966|7.529|6.550|6.247|6.186|10.012|10.143|9.600|5.817|8.328|6.499|6.000|7.578|10.817|6.073
j|0.153|0.613|0.268|0.493|0.397|3.501|0.011|0.034|0.614|1.836|1.46|0.730|1.144|2.042|1.433
k|0.772|0.049|1.417|0.011|0.015|4.163|0.009|5.683|3.140|2.753|2.248|3.395|3.314|4.973|2.894
l|4.025|5.456|3.437|4.967|2.779|6.104|6.510|5.922|5.275|2.564|3.568|5.229|4.532|5.761|3.802
m|2.406|2.968|2.534|3.157|4.738|2.994|2.512|3.752|3.471|2.515|2.213|3.237|4.041|3.202|2.446
n|6.749|7.095|9.776|6.712|4.446|7.955|6.883|7.987|8.542|6.237|10.032|7.240|7.711|8.826|6.468
o|7.507|5.796|2.594|8.683|9.735|8.779|9.832|2.976|4.482|6.667|6.063|4.636|2.166|5.614|6.695
p|1.929|2.521|0.670|2.510|2.523|2.755|3.056|0.886|1.839|2.445|1.57|1.756|0.789|1.842|1.906
q|0.095|1.362|0.018|0.877|1.204|0|0.505|0|0.020|0|0.009|0.007|0|0.013|0.001
r|5.987|6.693|7.003|6.871|6.530|5.914|6.367|7.722|8.431|5.243|6.411|8.956|8.581|2.872|4.799
s|6.327|7.948|7.270|7.977|6.805|6.092|4.981|3.014|6.590|5.224|3.73|5.805|5.630|7.862|5.212
t|9.056|7.244|6.154|4.632|4.336|5.276|5.623|3.314|7.691|2.475|6.79|6.862|4.953|8.750|5.727
u|2.758|6.311|4.166|2.927|3.639|3.183|3.011|3.235|1.919|2.062|1.99|1.979|4.562|5.008|2.160
v|0.978|1.838|0.846|1.138|1.575|1.904|2.097|0.959|2.415|0.012|2.85|2.332|2.437|2.250|5.344
w|2.360|0.074|1.921|0.017|0.037|0|0.033|0|0.142|5.813|1.52|0.069|0|0.094|0.016
x|0.150|0.427|0.034|0.215|0.253|0|0.003|0|0.159|0.004|0.036|0.028|0.046|0.031|0.027
y|1.974|0.128|0.039|1.008|0.006|0|0.020|3.336|0.708|3.206|0.035|0.698|0.900|1.745|1.043
z|0.074|0.326|1.134|0.467|0.470|0.494|1.181|1.500|0.070|4.852|1.39|0.034|0|0.051|1.503
à|0|0.486|0|0|0.072|0|0.635|0|0|0|0|0|0|0|0
â|0|0.051|0|0|0.562|0|0|0|0|0|0|0|0|0|0
á|0|0|0|0.502|0.118|0|0|0|0|0|0|0|1.799|0|0.867
å|0|0|0|0|0|0|0|0|1.338|0|0|1.190|0|0.003|0
ä|0|0|0.578|0|0|0|0|0|1.797|0|0|0|0|3.577|0
ã|0|0|0|0|0.733|0|0|0|0|0|0|0|0|0|0
ą|0|0|0|0|0|0|0|0|0|0.699|0|0|0|0|0
æ|0|0|0|0|0|0|0|0|0|0|0|0.872|0.867|0|0
œ|0|0.018|0|0|0|0|0|0|0|0|0|0|0|0|0
ç|0|0.085|0|0|0.530|0|0|1.156|0|0|0|0|0|0|0
ĉ|0|0|0|0|0|0.657|0|0|0|0|0|0|0|0|0
ć|0|0|0|0|0|0|0|0|0|0.743|0|0|0|0|0
č|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.462
ď|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.015
ð|0|0|0|0|0|0|0|0|0|0|0|0|4.393|0|0
è|0|0.271|0|0|0|0|0.263|0|0|0|0|0|0|0|0
é|0|1.504|0|0.433|0.337|0|0|0|0|0|0|0|0.647|0|0.633
ê|0|0.218|0|0|0.450|0|0|0|0|0|0|0|0|0|0
ë|0|0.008|0|0|0|0|0|0|0|0|0|0|0|0|0
ę|0|0|0|0|0|0|0|0|0|1.035|0|0|0|0|0
ě|0|0|0|0|0|0|0|0|0|0|0|0|0|0|1.222
ĝ|0|0|0|0|0|0.691|0|0|0|0|0|0|0|0|0
ğ|0|0|0|0|0|0|0|1.125|0|0|0|0|0|0|0
ĥ|0|0|0|0|0|0.022|0|0|0|0|0|0|0|0|0
î|0|0.045|0|0|0|0|0|0|0|0|0|0|0|0|0
ì|0|0|0|0|0|0|0.030|0|0|0|0|0|0|0|0
í|0|0|0|0.725|0.132|0|0|0|0|0|0|0|1.570|0|1.643
ï|0|0.005|0|0|0|0|0|0|0|0|0|0|0|0|0
ı|0|0|0|0|0|0|0|5.114|0|0|0|0|0|0|0
ĵ|0|0|0|0|0|0.055|0|0|0|0|0|0|0|0|0
ł|0|0|0|0|0|0|0|0|0|2.109|0|0|0|0|0
ñ|0|0|0|0.311|0|0|0|0|0|0|0|0|0|0|0
ń|0|0|0|0|0|0|0|0|0|0.362|0|0|0|0|0
ň|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.007
ò|0|0|0|0|0|0|0.002|0|0|0|0|0|0|0|0
ö|0|0|0.443|0|0|0|0|0.777|1.305|0|0|0|0.777|0.444|0
ô|0|0.023|0|0|0.635|0|0|0|0|0|0|0|0|0|0
ó|0|0|0|0.827|0.296|0|0|0|0|1.141|0|0|0.994|0|0.024
õ|0|0|0|0|0.040|0|0|0|0|0|0|0|0|0|0
ø|0|0|0|0|0|0|0|0|0|0|0|0.939|0|0|0
ř|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.380
ŝ|0|0|0|0|0|0.385|0|0|0|0|0|0|0|0|0
ş|0|0|0|0|0|0|0|1.780|0|0|0|0|0|0|0
ś|0|0|0|0|0|0|0|0|0|0.814|0|0|0|0|0
š|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.688
ß|0|0|0.307|0|0|0|0|0|0|0|0|0|0|0|0
ť|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.006
þ|0|0|0|0|0|0|0|0|0|0|0|0|1.455|0|0
ù|0|0.058|0|0|0|0|0.166|0|0|0|0|0|0|0|0
ú|0|0|0|0.168|0.207|0|0|0|0|0|0|0|0.613|0|0.045
û|0|0.060|0|0|0|0|0|0|0|0|0|0|0|0|0
ŭ|0|0|0|0|0|0.520|0|0|0|0|0|0|0|0|0
ü|0|0|0.995|0.012|0.026|0|0|1.854|0|0|0|0|0|0|0
ů|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.204
ý|0|0|0|0|0|0|0|0|0|0|0|0|0.228|0|0.995
ź|0|0|0|0|0|0|0|0|0|0.078|0|0|0|0|0
ż|0|0|0|0|0|0|0|0|0|0.706|0|0|0|0|0
ž|0|0|0|0|0|0|0|0|0|0|0|0|0|0|0.721"""
df = pd.read_table(StringIO(wiki_table), sep='|', index_col=0)
df.head()
# + [markdown] colab_type="text" id="3Av5tmHDbKOn"
# Implement the language classifier and answer the following:
#
# 1. **[0.5p]** Naive Bayes can be implemented
# either by multiplying probabilities or by adding
# log-probabilities. Which one is better and why?
#
# Please type a short answer below.
# 2. **[1.5p]** What is the language of the following phrases, according to the classifier (below in a code cell)? Assume equal prior language probabilities $P(C)$.
# 3. **[bonus]** What happens when a Naive Bayes classifier
# is applied to a document with out-of-vocabulary words? Propose
# some solutions. Relate them to the concept of Bayesian
# priors discussed during the lecture.
#
# This question will be discussed during Class Assignment 2.
# + [markdown] colab_type="text" id="6Qpm3aaICM-7"
# Using log-probabilities is better because it doesn't go so fast to zero.
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" id="6qqxHPSF6pQ0" outputId="b63cf3be-eac1-412e-9b3a-1fbbdd5375ab"
# We can easiily manipulate the letter frequency table using Pandas
langs = list(df)
letters = list(df.index)
print('Languages:', ','.join(langs))
print('Letters:', ', '.join(letters))
print('P(ę|Polish) =', df.loc['ę', 'Polish'])
# + colab={"base_uri": "https://localhost:8080/", "height": 629} colab_type="code" id="kW6DtwD48nzD" outputId="c3f48d57-722f-4f53-d6f6-8bc8690266d5"
# The values are recentages of letter appearance, but curiously enough they don't
# sum to 100%.
print(f'\nTotal letter count by language:\n{df.sum(0)}')
# Normalize the data such that the letter frequencies add up to 1 for each language
df_norm = df / df.sum(0)
print(f'\nAfter normalization:\n{df_norm.sum(0)}')
# + colab={} colab_type="code" id="SZI1U6VgcfxL"
norm_sent = lambda sent: sent.lower().translate(
str.maketrans(
'',
'',
string.punctuation
+ string.digits
+ string.whitespace)
)
def naive_bayes(sent, langs, df):
"""Returns the most probable language of a sentence"""
# Try working with log-probabilities.
# to prevent taking log(0) you can e.g. add a very small amount (1e-100)
# to each tabulated frequency.
df_log = df.replace(0, 1e-100).apply(np.log)
# normalize the sentence: remove spaces and punctuations, take lower case
sent = norm_sent(sent)
log_probs = {}
for lang in langs:
log_probs[lang] = np.sum(
[
df_log.loc[l, lang]
for l in sent
]
)
log_probs = {k: np.exp(log_probs[k]) for k in log_probs}
prob_sum = sum(log_probs.values())
# TODO compute language probabilitie and order from most to least probable
probs = OrderedDict(
[
(k, v / prob_sum)
for k,v in sorted(log_probs.items(), key=lambda kv: kv[1])[::-1]
]
)
return probs
# + colab={"base_uri": "https://localhost:8080/", "height": 323} colab_type="code" id="H2PK1SnFBZVc" outputId="1bd607b7-e144-4419-8b12-ed543e761c4d"
sentences = [
"No dejes para mañana lo que puedas hacer hoy.",
"Przed wyruszeniem w drogę należy zebrać drużynę.",
"Żeby zrozumieć rekurencję, należy najpierw zrozumieć rekurencję.",
"Si vale la pena hacerlo vale la pena hacerlo bien.",
"Experience is what you get when you didn't get what you wanted.",
"Należy prowokować intelekt, nie intelektualistów.",
u"<NAME> kaj feliĉan no<NAME>.",
u"Vos enfants sont très beaux. Ils sont adoptes?",
u"Is het de rode of de groene knoop die de bom ontmantelen.",
u"Se tem mesmo que cortar as unhas dos pés, faça o favor de deixar a cozinha.",
u"Keine Elephanten in der Bar nach 8 Uhr!",
u"Gostarias de ir a minha casa e fazer as coisas que de qualquer forma direi às pessoas que fizemos?",
u"Vandaag heb ik 10 eetlepels soep gegeten",
u"cuando tengo sed",
u"hej boysy, looknijcie sobie przez windowsa",
]
for sent in sentences:
print(f'{sent}:')
for k, v in naive_bayes(sent, langs, df_norm).items():
if v<1e-3:
break
print(f'{k}: {v:.3f}\t', end='')
print('\n')
# -
# ##### 3
# It depend to frequency of this words. If they are not very common- it doesn't change classification. But if they are very common it's hard even to native speakers to recognized language of text.
# To solve this we can use spell-checker which can cut off all non-vocabulary words. it shouldn't change 'value' of text much.
# + [markdown] colab_type="text" id="J1kDmmrYcg2b"
# # Problem 2: Simple Kalman filtering [2p + 2b]
#
# Oh no, someone has kidnapped you! You feel that you are in the trunk of a moving car. Luckily, you have your phone with GPS. Unfortunately, the GPS is noisy. You want to combine your estimate of your location by combining your prior belief about where you can be with the noisy GPS. You set out to implement a [1D Kalman filter](https://en.wikipedia.org/wiki/Kalman_filter).
#
# Problem setup:
# - your prior belief about the location is a Gaussian with mean 0 and some initial standard deviation $\mathcal{N}(0, \sigma_i)$
# - the car moves in a brownian motion - each time step, it changes location by a normally distributed random amound sampled from $\mathcal{N}(0, \sigma_m)$
# - each time step, you get a GPS reading which is sampled around your true (and sadly unknown to you ) location from $\mathcal{N}(\text{true loc}, \sigma_g)$
#
# You want to use the following algorithm to track your location:
#
# 1. Initially, the PDF of your location is $p(x) = \mathcal{N}(x; \mu_l=0, \sigma_l=\sigma_i)$
# 2. For each time step, you update your belief about your location:
# 1. $p(x)$ is updated due to according to the car movement distribution
# 2. you use the Bayes formula to incorporate the GPS readout:
# $$
# p(x|\text{GPS readout}) = \frac{p(\text{GPS readout}|x)p(x)}
# {p(\text{GPS readout})}
# $$
# 3. you set $p(x) \gets p(x|\text{GPS readout})$ to be your prior belief about your locatin used during the next iteration.
#
#
# NB: the GPS is actually very noisy, and Kalman filters are routinely used to fuse information from the GPS, accelerometers and odometry in practical applications, such as GPS navigation.
#
# Hint: during the class assignments we have computed the pdf of
# $$
# p(x) = \mathcal{N}(x;\mu_1, \sigma_1)\mathcal{N}(x;\mu_2, \sigma_2)
# $$
# What disrtibution will the PDF belong to? Maybe you can simply compute the new mean and standard deviation?
#
# #### Problem [.5p]
#
# Implement below a simulator for your kidnapping, then fill in the code for plotting the true location and GPS readouts over time.
#
# #### Problem [1.5p]
#
# Implement a 1D Kalman filer using the algorithm stated above: maintian a probability distribution over your location, then at each timestep update it to account for car movement and GPS readouts.
#
# Plot the estimated location along with its standard deviation against the true location from the simluator.
#
# Experiemt with different setting for the standard deviations of the car's motion and the GPS. What happens if the simulator and the Kalman filter use different probability distributions?
#
# #### Problem [2p bonus]
#
# Suppose the car has a velocity, which is updated at each time step:
# $$
# \begin{split}
# v &\gets v + \mathcal{N}(0, \sigma_v) \\
# x &\gets x + v \\
# \text{GPS readout} &= \mathcal{N}(x, \sigma_g)
# \end{split}
# $$
#
# Update the Kalman filter code to track both the car's location and velocity. You can assume that the initial velocity is exactly 0.
# -
np.random.normal(0, 1, 1)
# + colab={} colab_type="code" id="8RXfUadZcl6s"
def simulate(initial_sigma, motion_sigma, gps_sigma, n_steps):
"""Simulate a sequence of locations and noisy GPS measurements
Args:
initial_sigma, motion_sigma, gps_sigma: parameters of the simulation
n_steps: number of timesteps
Returns:
a DataFrame with columns 'x' and 'gps' giving the true location and
gps readouts.
"""
# Sample an initial location from the distribution ovetr the initial loc
x = np.random.normal(0, initial_sigma, 1)[0]
loc_hist = []
for s in range(n_steps):
x = np.random.normal(x, motion_sigma, 1)[0]
gps_readout = np.random.normal(x, gps_sigma, 1)[0]
loc_hist.append((x, gps_readout))
loc_df = pd.DataFrame(loc_hist, columns=['x', 'gps'])
return loc_df
def kalman_predict(loc_df, initial_sigma, motion_sigma, gps_sigma):
# Set our initial belief about our location
prior_mu = 0#TODO
prior_sigma = 1#TODO
posterior_mu = prior_mu
posterior_sigma = prior_sigma
predictions = []
for gps_readout in loc_df.gps:
# expand the prior by the movement
prior_sigma = 15#TODO
# now do the bayes update
posterior_mu = (
posterior_mu * gps_sigma + motion_sigma * gps_readout
) / (motion_sigma + gps_sigma)
posterior_sigma = 1 / (1/motion_sigma + 1/gps_sigma)
predictions.append((posterior_mu, posterior_sigma))
predictions_df = pd.DataFrame(predictions, columns=['mu', 'sigma'])
return predictions_df
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="rMdxz44RcxXK" outputId="a5b5c506-2146-40c2-b42a-60422ed26735"
#@title Kalman Simulationb
initial_sigma = 10 #@param
motion_sigma = 5 #@param
gps_sigma = 20 #@param
n_steps = 50 #@param
loc_df = simulate(initial_sigma, motion_sigma, gps_sigma, n_steps)
predictions_df = kalman_predict(loc_df, initial_sigma, motion_sigma, gps_sigma)
plt.plot(loc_df.x, 'r', label='true position')
plt.plot(loc_df.gps, 'go', label='gps readout')
plt.plot(predictions_df.mu, 'b', label='kalman position')
plt.fill_between(range(len(predictions_df)),
predictions_df.mu + predictions_df.sigma,
predictions_df.mu - predictions_df.sigma, color='b', alpha=0.2)
plt.fill_between(range(len(predictions_df)),
predictions_df.mu + 3 * predictions_df.sigma,
predictions_df.mu - 3 * predictions_df.sigma, color='b', alpha=0.1)
plt.legend(loc='upper center', ncol=3, bbox_to_anchor=(0.5, 1.0), frameon=True)
plt.xlabel('time')
plt.ylabel('position')
plt.title('Kalman filtering of location data')
None
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="rMdxz44RcxXK" outputId="a5b5c506-2146-40c2-b42a-60422ed26735"
#@title Kalman Simulationb
initial_sigma = 10 #@param
motion_sigma = 45 #@param
gps_sigma = 20 #@param
n_steps = 50 #@param
loc_df = simulate(initial_sigma, motion_sigma, gps_sigma, n_steps)
predictions_df = kalman_predict(loc_df, initial_sigma, motion_sigma, gps_sigma)
plt.plot(loc_df.x, 'r', label='true position')
plt.plot(loc_df.gps, 'go', label='gps readout')
plt.plot(predictions_df.mu, 'b', label='kalman position')
plt.fill_between(range(len(predictions_df)),
predictions_df.mu + predictions_df.sigma,
predictions_df.mu - predictions_df.sigma, color='b', alpha=0.2)
plt.fill_between(range(len(predictions_df)),
predictions_df.mu + 3 * predictions_df.sigma,
predictions_df.mu - 3 * predictions_df.sigma, color='b', alpha=0.1)
plt.legend(loc='upper center', ncol=3, bbox_to_anchor=(0.5, 1.0), frameon=True)
plt.xlabel('time')
plt.ylabel('position')
plt.title('Kalman filtering of location data')
None
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="rMdxz44RcxXK" outputId="a5b5c506-2146-40c2-b42a-60422ed26735"
#@title Kalman Simulationb
initial_sigma = 10 #@param
motion_sigma = 5 #@param
gps_sigma = 80 #@param
n_steps = 50 #@param
loc_df = simulate(initial_sigma, motion_sigma, gps_sigma, n_steps)
predictions_df = kalman_predict(loc_df, initial_sigma, motion_sigma, gps_sigma)
plt.plot(loc_df.x, 'r', label='true position')
plt.plot(loc_df.gps, 'go', label='gps readout')
plt.plot(predictions_df.mu, 'b', label='kalman position')
plt.fill_between(range(len(predictions_df)),
predictions_df.mu + predictions_df.sigma,
predictions_df.mu - predictions_df.sigma, color='b', alpha=0.2)
plt.fill_between(range(len(predictions_df)),
predictions_df.mu + 3 * predictions_df.sigma,
predictions_df.mu - 3 * predictions_df.sigma, color='b', alpha=0.1)
plt.legend(loc='upper center', ncol=3, bbox_to_anchor=(0.5, 1.0), frameon=True)
plt.xlabel('time')
plt.ylabel('position')
plt.title('Kalman filtering of location data')
None
# + [markdown] colab_type="text" id="yQfI6oJHczUa"
# # Problem 3: Decision Tree Implementation
#
# Currently, there are no good implementations of Decision Trees in Python.
#
# Sadly, the machine leading toolkit [sklearn](https://scikit-learn.org/stable/index.html) doesn't handle categorical attributes. Let's use this as an excuse to implement Decision Tress ourselves.
#
#
# + colab={} colab_type="code" id="VtJg1W1MBjgr"
#@title Data Loading
# We will load a few commonly used datasets:
# - mushroom
# - iris
# - adult
# - congressional voting
# - german credit
# 1. Mushroom dataset
# https://archive.ics.uci.edu/ml/datasets/mushroom
# only categorical attributes with missing values
columns = [
"target", "cap-shape", "cap-surface", "cap-color", "bruises?", "odor",
"gill-attachment", "gill-spacing", "gill-size", "gill-color", "stalk-shape",
"stalk-root", "stalk-surface-above-ring", "stalk-surface-below-ring",
"stalk-color-above-ring", "stalk-color-below-ring", "veil-type", "veil-color",
"ring-number", "ring-type", "spore-print-color", "population", "habitat", ]
# Use read_csv to load the data.
url = 'http://archive.ics.uci.edu/ml/machine-learning-databases/mushroom/agaricus-lepiota.data'
mushroom_df = pd.read_csv(url, header=None, names=columns)
mushroom_idx_df = mushroom_df.reset_index()
# 2. Iris
iris_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data'
iris_df = pd.read_csv(
iris_url, header=None,
names=['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'target'])
# 3. Congressoinal Voting
# Binary attributes, binary class, missing data
vote_df = pd.read_csv(
'https://pkgstore.datahub.io/machine-learning/vote/vote_csv/data/65f1736301dee4a2ad032abfe2a61acb/vote_csv.csv'
).rename({'Class':'target'}, axis=1).fillna('na')
# 4. Adult
# census records, continuous and categorical attributes (some ordered), missing values
adult_names = [
"Age", "Workclass", "fnlwgt", "Education", "Education-Num", "Martial Status",
"Occupation", "Relationship", "Race", "Sex", "Capital Gain", "Capital Loss",
"Hours per week", "Country", "target"]
adult_df = pd.read_csv(
'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data',
names=adult_names, header=None, na_values="?")
adult_test_df = pd.read_csv(
'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test',
names=adult_names, header=None, na_values="?", skiprows=1)
# 5. German Credit
german_df = pd.read_csv(
'https://archive.ics.uci.edu/ml/machine-learning-databases/statlog/german/german.data',
names=[f'A{d}' for d in range(1,21)] + ['target'], header=None, sep=' ')
# + [markdown] colab_type="text" id="MHSSMQuWJ3nv"
# ## Decision Tree Task 1: Purity Measures [1p]
#
# Please fill the purity measures below.
#
# Verify the correctness by plotting the purity values if a two-class set with given class probabilities
# + colab={"base_uri": "https://localhost:8080/", "height": 275} colab_type="code" id="OJyyb5YY9o_H" outputId="e572c4cd-6748-4798-88f0-08a8fe500a49"
def entropy(counts):
_sum = np.sum(counts.values, axis=0)
return -np.sum(counts/_sum * np.log2(counts/_sum), axis=0)
def gini(counts):
# print(type(counts))
# print(counts)
_sum = np.sum(counts.values, axis=0)
return np.sum(counts/_sum * (1 - counts/_sum), axis=0)
def mean_err_rate(counts):
_sum = np.sum(counts.values, axis=0)
return np.min(counts/_sum, axis=0)
# Make a plot of the purity functions
x = np.linspace(0, 100, 500)
data = pd.DataFrame.from_dict({"c1": x, "c2": x.max(axis=0)-x}).transpose()
plt.axis('equal')
plt.plot(x/100, entropy(data))
plt.plot(x/100, gini(data))
plt.plot(x/100, mean_err_rate(data))
# + [markdown] colab_type="text" id="sYD_oPNBuuWk"
# ## Decision Tree Task 2: Categorical Splits [1p]
#
# ### The anatomy of a Decision Tree
#
#
# All internal (non-leaf) nodes of a split training examples according to a test implemented by the node. We capture this behavior using a generic `AbstractSplit` class which implements a split of data contained in the dataframe `df` using the attribute `attr`.
#
# The class features a lightweight constructor, `__init__` which only saves the information required to later split a training dataframe and recursively build the subtrees using the `build_subtrees` method.
#
# Fill in the blanks in the code below.
# + colab={} colab_type="code" id="IgLVlZhvy5hl"
class AbstractSplit:
"""Split the examples in a tree node according to a criterion.
"""
def __init__(self, attr):
self.attr = attr
def __call__(self, x):
"""Return the subtree corresponding to x."""
raise NotImplementedError
def build_subtrees(self, df, subtree_kwargs):
"""Recuisively build the subtrees."""
raise NotImplementedError
def iter_subtrees(self):
"""Return an iterator over subtrees."""
raise NotImplementedError
def add_to_graphviz(self, dot):
"""Add the split to the graphviz vizluzation."""
raise NotImplementedError
def __str__(self):
return f"{self.__class__.__name__}: {self.attr}"
# + [markdown] colab_type="text" id="IU6lhc_z9cx6"
# We will first implement a Multivariate Categorical split which has a subtree for each value that an attribute may take.
# + colab={} colab_type="code" id="CdUeZJTGwwHc"
class CategoricalMultivalueSplit(AbstractSplit):
def build_subtrees(self, df, subtree_kwargs):
self.subtrees = {}
for group_name, group_df in df.groupby(self.attr):
child = Tree(group_df, **subtree_kwargs)
self.subtrees[group_name] = child
def __call__(self, x):
# Return the subtree for the given example
return self.subtrees[x[self.attr]] #TODO
def iter_subtrees(self):
return self.subtrees.values()
def add_to_graphviz(self, dot, parent, print_info):
for split_name, child in self.subtrees.items():
child.add_to_graphviz(dot, print_info)
dot.edge(f'{id(parent)}', f'{id(child)}',
label=f'{split_name}')
# + colab={} colab_type="code" id="XUWaldXZ96Ha"
def get_categorical_split_and_purity(df, parent_purity, purity_fun, attr,
normalize_by_split_entropy=False):
"""Return a multivariate split and its purity.
Args:
df: a dataframe
parent_purity: purity of the parent node
purity_fun: function to compute the purity
attr: attribute over whihc to split the dataframe
normalize_by_split_entropy: if True, divide the purity gain by the split
entropy (to compute https://en.wikipedia.org/wiki/Information_gain_ratio)
Returns:
pair of (split, purity_gain)
"""
split = CategoricalMultivalueSplit(attr)
# Compute the purity after the split
x = df.groupby(attr).apply(lambda x: purity_fun(x['target'].value_counts()))
purity = np.sum(x * df[attr].value_counts() / len(df))
purity_gain = parent_purity - purity
if normalize_by_split_entropy:
purity_gain /= entropy(df[attr].value_counts())
return split, purity_gain
# + colab={} colab_type="code" id="2e_C9VVl6omi"
def get_split(df, criterion='infogain', nattrs=None):
# Implement termination criteria:
# 1. Node is pure
target_value_counts = df['target'].value_counts()
if len(target_value_counts) == 1:
return None
# 2. No split is possible
# First get alist of attributes that can be split
possible_splits = [x for x in df if x != 'target' and len(df[x].unique()) > 1] #TODO
# Terminate early if none are possivle
if not possible_splits:
return None
# Get the base purity measure and the purity function
if criterion in ['infogain', 'infogain_ratio']:
purity_fun = entropy
elif criterion in ['mean_err_rate']:
purity_fun = mean_err_rate
elif criterion in ['gini']:
purity_fun = gini
else:
raise Exception("Unknown criterion: " + criterion)
base_purity = purity_fun(target_value_counts)
best_purity_gain = -1
best_split = None
# Random Forest support
# Randomize the split by restricting the number of attributes
possible_splits = random.sample(possible_splits, min(nattrs, len(possible_splits)))
for attr in possible_splits:
if np.issubdtype(df[attr].dtype, np.number):
# Handling of numerical attributes will be defined later, in a manner
# similar to categorical ones
split_sel_fun = get_numrical_split_and_purity
else:
split_sel_fun = get_categorical_split_and_purity
split, purity_gain = split_sel_fun(
df, base_purity, purity_fun, attr,
normalize_by_split_entropy=criterion.endswith('ratio'))
if purity_gain > best_purity_gain:
best_purity_gain = purity_gain
best_split = split
return best_split
# + [markdown] colab_type="text" id="latO4p-WAHiG"
# We can now define a Tree class, which represents both a Decision Tree and its Nodes.
#
# Each node saves its class distribution in the `counts` attribute and debug/visualization information in the `info` field.
#
# Leaf nodes have `split == None`, while internal nodes have a split which points to subtrees.
#
# + colab={} colab_type="code" id="7-CMCry3AK7n"
class Tree:
def __init__(self, df, **kwargs):
super().__init__()
# Assert that threre are no missing values,
# TODO: remove this for bonus problem #XXX
assert not df.isnull().values.any()
# We need to let subrees know about all targets to properly color nodes
if 'all_targets' not in kwargs:
kwargs['all_targets'] = sorted(df['target'].unique())
# Save keyword arguments to build subtrees
kwargs_orig = dict(kwargs)
# Get kwargs we know about, remaning ones are for splitting
self.all_targets = kwargs.pop('all_targets')
# Save debug info for visualization
self.counts = df['target'].value_counts()
self.info = {
'num_samples': len(df),
'entropy': entropy(self.counts),
'gini': gini(self.counts),
'correct': 0,
'wrong': 0
}
# print("self info", self.info)
self.split = get_split(df, **kwargs)
if 'nattrs' in kwargs_orig:
if kwargs_orig['nattrs'] > 0:
kwargs_orig['nattrs'] -= 1
else:
self.split = None
if self.split:
#print('!!S', self.split)
self.split.build_subtrees(df, kwargs_orig)
def get_target_distribution(self, sample):
print("NotImplemented", sample)
# TODO: descend into subtrees and return the leaf target distribution
def create_stats(self, sample):
if sample['target'] == self.counts.idxmax():
self.info['correct'] += 1
else:
self.info['wrong'] += 1
if self.split:
try:
self.split(sample).create_stats(sample)
except:
return None
def prune(self):
if self.split:
wrongs = sum([x.info['wrong'] for x in self.split.iter_subtrees()])
if wrongs >= self.info['wrong']:
self.split = None
else:
for c in self.split.iter_subtrees():
c.prune()
def classify(self, sample):
if not self.split:
return self.counts.idxmax()
else:
try:
return self.split(sample).classify(sample)
except:
return self.counts.idxmax()
# TODO: classify the sample by descending into the appropriate subtrees.
def draw(self, print_info=True):
dot = graphviz.Digraph()
self.add_to_graphviz(dot, print_info)
return dot
def add_to_graphviz(self, dot, print_info):
freqs = self.counts / self.counts.sum()
freqs = dict(freqs)
colors = []
freqs_info = []
for i, c in enumerate(self.all_targets):
freq = freqs.get(c, 0.0)
if freq > 0:
colors.append(f"{i%9 + 1};{freq}")
freqs_info.append(f'{c}:{freq:.2f}')
colors = ':'.join(colors)
labels = [' '.join(freqs_info)]
if print_info:
for k,v in self.info.items():
labels.append(f'{k} = {v}')
if self.split:
labels.append(f'split by: {self.split.attr}')
dot.node(f'{id(self)}',
label='\n'.join(labels),
shape='box',
style='striped',
fillcolor=colors,
colorscheme='set19')
if self.split:
self.split.add_to_graphviz(dot, self, print_info)
# + colab={"base_uri": "https://localhost:8080/", "height": 857} colab_type="code" id="xpNExVwICWJL" outputId="30e3e1b8-ff9c-4db0-b4af-d6105b7a1e56"
# TODO: train a Decision Tree on the mushroom data.
# Plot the tree using the `.draw()` method.
# How many samples are classified correctly by a tree with only one split?
# Is the tree different when different purity functions are used?
mushroom_tree = Tree(mushroom_df, criterion='infogain')
mushroom_tree.draw()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="Q0Kv-kB99PkQ" outputId="9d58a25b-c993-4dac-e1cf-ee85417d3a80"
mushroom_tree = Tree(mushroom_df, criterion='infogain_ratio')
mushroom_tree.draw()
# + [markdown] colab_type="text" id="GABxeo7x2agz"
# ## Decision Tree Task 3: Numerical Splits [1p]
# A numerical split requires to search for the best threshold. Implement the selection of splits for numerical attributes below.
# + colab={} colab_type="code" id="4dqmM69UE64U"
class NumericalSplit(AbstractSplit):
def __init__(self, attr, th):
super(NumericalSplit, self).__init__(attr)
self.th = th
def build_subtrees(self, df, subtree_kwargs):
self.subtrees = (
Tree(df[df[self.attr] <= self.th], **subtree_kwargs),
Tree(df[df[self.attr] > self.th], **subtree_kwargs))
def __call__(self, x):
if x[self.attr] <= self.th:
return self.subtrees[0]
return self.subtrees[1]
def __str__(self):
return f"NumericalSplit: {self.attr} <= {self.th}"
def iter_subtrees(self):
return self.subtrees
def add_to_graphviz(self, dot, parent, print_info):
self.subtrees[0].add_to_graphviz(dot, print_info)
dot.edge(f'{id(parent)}', f'{id(self.subtrees[0])}',
label=f'<= {self.th:.2f}')
self.subtrees[1].add_to_graphviz(dot, print_info)
dot.edge(f'{id(parent)}', f'{id(self.subtrees[1])}',
label=f'> {self.th:.2f}')
def get_numrical_split_and_purity(df, parent_purity, purity_fun, attr,
normalize_by_split_entropy=False):
"""Find best split thereshold and compute the average purity after a split.
Args:
df: a dataframe
parent_purity: purity of the parent node
purity_fun: function to compute the purity
attr: attribute over whihc to split the dataframe
normalize_by_split_entropy: if True, divide the purity gain by the split
entropy (to compute https://en.wikipedia.org/wiki/Information_gain_ratio)
Returns:
pair of (split, purity_gain)
"""
attr_df = df[[attr, 'target']].sort_values(attr)
targets = attr_df['target']
values = attr_df[attr]
# Start with a split that puts all the samples into the right subtree
right_counts = targets.value_counts()
left_counts = right_counts * 0
best_split = None
best_purity_gain = -1
N = len(attr_df)
for row_i in range(N - 1):
# Update the counts of targets in the left and right subtree and compute
# the purity of the slipt for all possible thresholds!
# Return the best split found.
# Remember that the attribute may have duplicate values and all samples
# with the same attribute value must end in the same subtree!
threshold = values.iloc[row_i]
lower_equal = targets[values <= threshold]
greater = targets[values > threshold]
left_purity = purity_fun(lower_equal.value_counts())
right_purity = purity_fun(greater.value_counts())
purity = (
len(lower_equal) * left_purity
+ len(greater) * right_purity
) / len(values)
purity_gain = parent_purity - purity
if normalize_by_split_entropy:
purity_gain /= entropy(values.value_counts())
if best_purity_gain < purity_gain:
best_purity_gain = purity_gain
best_split = NumericalSplit(attr, threshold)
return best_split, best_purity_gain
# + colab={"base_uri": "https://localhost:8080/", "height": 693} colab_type="code" id="bKBwWQiABhID" outputId="70ca26d9-21e8-4366-bf63-5648f1cd5dd5"
# TODO: apply the tree to Iris with petal_length and petal_width attributes
iris2d = iris_df[['petal_length', 'petal_width', 'target']]
iris_tree = Tree(iris2d, criterion='infogain_ratio')
iris_tree.draw()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="LdatEZNFcBdu" outputId="2b2337a1-4691-41fc-c805-aa977ba12313"
def temp():
# TODO: to verify the tree building algorithm draw Iris2D decision boundary
# for different splitting criteria.
mesh_x, mesh_y = np.meshgrid(
np.linspace(iris2d.petal_length.min(), iris2d.petal_length.max(), 100),
np.linspace(iris2d.petal_width.min(), iris2d.petal_width.max(), 100),
)
mesh_data = np.hstack([mesh_x.reshape(-1, 1), mesh_y.reshape(-1, 1)])
mesh_data = pd.DataFrame(mesh_data, columns=iris2d.columns[:-1])
preds = np.empty( (len(mesh_data),))
for criterion in ['infogain', 'infogain_ratio', 'gini', 'mean_err_rate']:
iris2d_tree = Tree(iris2d, criterion=criterion)
for i, (_, r) in enumerate(mesh_data.iterrows()):
preds[i] = iris2d_tree.all_targets.index(iris2d_tree.classify(r))
plt.figure()
plt.title(f"Iris2D decision boundary for {criterion}.")
plt.contourf(mesh_x, mesh_y, preds.reshape(mesh_x.shape), cmap='Set1', vmin=0, vmax=7)
sns.scatterplot(x='petal_length', y='petal_width', hue='target', data=iris_df, palette='Set1', )
temp()
# + [markdown] colab_type="text" id="b2Q2ltNeSZGn"
# ## Decision Tree Task 4: Pruning [2p + 2bp]
#
# Tree pruning tries to remove splits that don't result in a decrease of the error rate.
#
# There are two possible strategies:
#
# ### 1. Reduced Error Rate Pruning
# Build a tree using all the data. Then split the training set into 10 crossvalidation subsets. Then in a loop over the tesintg crossvalidation subset:
# - put the data from the remaining 9 subsets through the tree, remember distributions at each node (leaf and internal nodes)
# - classify the samples in the testing subset, record the error rate for all nodes
# - remove leaf nodes that have a higher error rate than their parents.
#
# ### 2. Confidence-interval Pruning
# Build the deicision tree and record the class distribution in each node. For each node, estimate the upper confidence interval on the error rate. Remove nodes that have a higher upper bound on the error rate than their parents.
#
# As you can see, the two strategies are quite similar: both estimate the error rate for all nodes in the tree and remove subtrees that do not improve it. The difference stems from the way in which the error rates are computed.
#
# ### Task:
#
# Split the voting dataset into a training and testing set using a 70%-30% ratio.
#
# Train a decision tree and prune it using either method 1. or 2.
#
# Compare the error rates on the test set of the original and pruned tree.
#
# For bonus points: implement the other pruning algorithm.
#
# **Implementation hint**: you can store the information related to pruning in the `Tree.info` field. In this way, it will be printed by `Tree.draw` method.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="x28vJNM1SpHy" outputId="0456ff12-1b05-4d51-8192-efc80c81195e"
vote_tree = Tree(vote_df, criterion='infogain_ratio')
vote_tree.draw()
# +
def error_rate(tree, df):
err = 0
for _, row in df.iterrows():
ans = tree.classify(row)
err += ans != row['target']
return err/len(df)
def prune_tree(tree, df):
for _, row in df.iterrows():
tree.create_stats(row)
tree.prune()
# + colab={"base_uri": "https://localhost:8080/", "height": 992} colab_type="code" id="7hi_eo5XToE_" outputId="4a0d58b3-b698-4ce0-a683-252748f95b98"
N = len(vote_df)
d = 0.7
rand_perm = np.random.permutation(N)
train_idx = rand_perm[:int(d * N)]
test_idx = rand_perm[int(d * N):]
train_df = vote_df.iloc[train_idx]
test_df = vote_df.iloc[test_idx]
tree = Tree(train_df)
print("Unpruned err rate:", error_rate(tree, test_df))
prune_tree(tree, test_df)
print("Pruned err rate:", error_rate(tree, test_df))
tree.draw()
# + [markdown] colab_type="text" id="mBdvPmXKGon3"
# # Problem 4: Random Forest [3p]
#
# We will use the german credit dataeset. Please split it into a traiing and testing set using a 70%-30% ratio.
#
# Then train and test a regular decision tree on it.
#
# Then:
# 1. Implement selecting the split from a small random selection of attriutes.
# 2. Build a forest of at least 20 Random Trees, each selecting splits out of 1-3 attributes on the German Credit data. After adding each random tree:
# - Compute its test error rate and its OOB error rate
# - Record the accurracy of the RF after adding the tree to it.
#
# At the end ot training record the forest's OOB error rate.
#
#
# What is the mean accurracy of individual trees in the forest? What is the final forest accurracy?
#
# Define the agreement between two trees to be the fraction of test samples on which the answer of the two trees is identical. What is the mean aggreement of trees in the forest? How does it change with the number of attributes considered for each split? What is the impact of training each tree in the forest on a bootstrap sample, rather than on the train set?
# + colab={"base_uri": "https://localhost:8080/", "height": 236} colab_type="code" id="f-w0gDDTNhr6" outputId="fe7758c6-63f1-4043-f949-ca666897eba9"
N = len(german_df)
d = 0.7
rand_perm = np.random.permutation(N)
train_idx = rand_perm[:int(d * N)]
test_idx = rand_perm[int(d * N):]
train_df = german_df.iloc[train_idx]
test_df = german_df.iloc[test_idx]
tree = Tree(train_df)
print("Unpruned err rate:", error_rate(tree, test_df))
prune_tree(tree, test_df)
print("Pruned err rate:", error_rate(tree, test_df))
tree.draw()
# +
def error_rate2(tree, df, res, num):
err = 0
res[num] = np.zeros(len(res))
for i, (_, row) in enumerate(df.iterrows()):
ans = tree.classify(row)
err += ans != row['target']
res.iloc[i][num] = ans
tree.create_stats(row)
return err/len(df)
train_res = pd.DataFrame(index=range(len(train_df)))
test_res = pd.DataFrame(index=range(len(test_df)))
trees = []
for nattrs in range(1,4):
print("nattr: ", nattrs)
for i in range(3):
trees.append(Tree(train_df, nattrs=nattrs))
train_err = error_rate2(trees[i], train_df, train_res, i)
test_err = error_rate2(trees[i], test_df, test_res, i)
forrest_err = np.sum(np.array(train_res.mode(axis=1)).T == np.array(train_df['target']))/len(train_df)
print(f"Tree {i}: RF Err rate {forrest_err} Err rate {train_err} OOB err rate {test_err}")
# + [markdown] colab_type="text" id="nHW2hwFqmIdg"
# # Problem 5 [3bp]
#
# Implement the following extra analysis using a Random Forest:
# - variable importance
# - data clustering
# - data visualizatoin using MultiDimensional Sclaing (https://en.wikipedia.org/wiki/Multidimensional_scaling, https://scikit-learn.org/stable/modules/generated/sklearn.manifold.MDS.html).
#
# For details see https://www.stat.berkeley.edu/~breiman/Using_random_forests_v4.0.pdf.
| assignment2/Assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import PyPDF2
import pandas as pd
import tika
import re
import matplotlib.pyplot as plt
pdfFileObj = open('FICT.pdf', 'rb')
pdfReader = PyPDF2.PdfFileReader(pdfFileObj)
pageObj = pdfReader.getPage(15)
page1 = pageObj.
page1
pdfReader.numPages
# +
from tika import parser
file = 'FICT.pdf'
raw = parser.from_file(file)
# -
text = raw['content']
text = '\n'.join(text.split('ПІБ Група Код спец. Сер. бал№'))
text = re.sub('^\n*', '', text)
text = re.sub('(Курс Спеціальність \d{3} Група \w{2}-\d{2}(мп)?)', '', text)
text = re.sub('Курс Спеціальність', '', text)
text = '\n\n' + text[71:]
text = text.split('\n\n')
text = list(filter(None, text))
text
item = text[1]
item = re.sub(' ', ' ', item)
item = item.split(' ')
name = ' '.join(item[:3])
text[0]
item = item[3:]
item.insert(0, name)
# +
text = raw['content']
text = '\n'.join(text.split('ПІБ Група Код спец. Сер. бал№'))
text = re.sub('^\n*', '', text)
text = re.sub('(Курс Спеціальність \d{3} Група \w{2}-\d{2}(мп)?)', '', text)
text = re.sub('Курс Спеціальність', '', text)
text = '\n\n' + text[71:]
text = text.split('\n\n')
text = list(filter(None, text))
del_list = []
for index in range(len(text)-1):
item = text[index]
if len(item) > 4:
item = re.sub(' ', ' ', item)
item = item.split(' ')
name = ' '.join(item[:3])
item = item[3:]
item.insert(0, name)
text[index] = item
else:
del_list.append(index)
for i in range(len(text)-1):
if len(text[i]) > 4:
del(text[i][1])
for i in range(len(text)-1):
if type(text[i]) != type([]):
del_list.append(i)
for index in sorted(del_list, reverse=True):
del (text[index])
text[40] = list(filter(None, text[40]))
text[163][1] = None
text[163] = list(filter(None, text[163]))
text[611][1] = 'ІТ-72'
text[628][1] = 'ІТ-74'
del(text[-1])
# +
name_surname = []
group = []
spec_index = []
score = []
for item in text:
name_surname.append(item[0])
group.append(item[1])
spec_index.append(item[2])
score.append(float(item[3]))
# -
fict_scores = pd.DataFrame(
{'ПІБ': name_surname,
'Група': group,
'Індекс Спеціальності': spec_index,
'Бал': score
})
fict_scores['Група'].unique()
our = ['ІО-71', 'ІО-72', 'ІО-73', 'ІВ-71', 'ІВ-72', 'ІВ-73']
average_minus5 = fict_scores['Бал']*0.95
fict_scores['Бал -5%'] = average_minus5
fict_scores.where(fict_scores['Група'] == our)
our_boys = fict_scores[fict_scores['Група'].isin(our)]
sorted_boys = our_boys.sort_values(by ='Бал')[::-1]
import pdfkit as pdf
sorted_boys.to_html('test.html')
PdfFilename='pdfPrintOut.pdf'
pdf.from_file('test.html', PdfFilename)
our_boys
87*0.25
sorted_boys = sorted_boys.iloc[:40,]
sorted_boys.index = range(1, 41)
sorted_boys
| AverageForFICT(done).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Automated ML
#
# TODO: Import Dependencies. In the cell below, import all the dependencies that you will need to complete the project.
# ### Table of Contents
# 1. [Introduction]()
# 2. [Import all Dependencies](1.0.1)
# 3. [Setup Workspace and Experiment](1.1)
# 4. [Dataset](1.2)
# 5. [Hyperdrive Configuration](1.3)
# 6. [Model Deployment](1.6)
# 7. [Test](1.7)
# 8. [Get Logs](1.8)
# 9. [Delete Resources](1.9)
# ## Introduction
#
# This is a Capstone project in fulfilment of the Udacity Azure ML Nanodegree.
#
#
# This project is aimed at demonstrating the capabilities of the Azure ML studio in training a model and deploying it. There are two ways Azure ML studio achieves this: one is through AUTOML, a codeless configuration that automates machine learning. Another, is the HYPERDRIVE, a custom hyperparameter tuning functionality for optimizing a ML model's performance. Then, from any of these two functionalities of the Azure ML studio, a production model will emerge to enable us explore the Azure ML End-to-End production pipeline solution for enabling interaction between a deployed model and other web services.
#
#
# In this demo, the AUTOML outputed the VotingEnsemble model which is an ensemble of LightGBMClassifier and LogisticRegression classifier. The AUTOML best performing model, the VotingEnsemble, achieved a 99.978% accuracy.
# ### Import All Required Dependencies
# + gather={"logged": 1598423888013} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# System libraries
import os
import csv
import shutil
import logging
# Conda libraries
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import pkg_resources
# Azure core libraries
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
from azureml.core.dataset import Dataset
# Computer target core libraries
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Libraries for Visualizing run
from azureml.widgets import RunDetails
#library for Saving model
import joblib
# Clean data function defined in train.py script
from train import clean_data
# ONNX libraries
import sys
import json
from azureml.automl.core.onnx_convert import OnnxConvertConstants
from azureml.train.automl import constants
# Check core SDK version number
print("SDK version:", azureml.core.VERSION)
# -
# ### Initialize Workspace
# Initialize a workspace object from persisted configuration. Make sure the config file is present at .\config.json
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# ### Setup Experiment
# Here we will be creating an experiment named "heart-disease-automl".
# +
# choose a name for experiment
experiment_name = 'liver-disease-automl'
experiment=Experiment(ws, experiment_name)
# -
# ### Create or Attach an AmlCompute cluster
# You will need to create a compute target for your AutoML run. In this demo, you get the default AmlCompute as your training compute resource.
# +
#Create compute cluster
# Choose a name for your CPU cluster
cpu_cluster_name = "notebook137067"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS3_V2',
max_nodes=4)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# ## Dataset
#
# TODO: In this markdown cell, give an overview of the dataset you are using. Also mention the task you will be performing.
# TODO: Get data. In the cell below, write code to access the data you will be using in this project. Remember that the dataset needs to be external.
#
# ### Overview
# There is an increasing number of patients with liver disease in recent time due to life style and living habits such as excessive alcohol consumption, inhale of harmful gases, excessive weight gain, intake of contaminated food, abuse of drugs. This dataset is aimed at helping doctors during clinical diagnosis of liver disease to elevate burden and the stress involved in analyzing every single patients’ information. Therefore, the goal is to create a classifier that predicts whether a subject is healthy (non-liver patient) or ill (liver patient) based on some clinical and demographic features which are: age, gender, total Bilirubin, direct Bilirubin, total proteins, albumin, A/G ratio, SGPT, SGOT and Alkphos.
#load the liver dataset to datastore
data_path = "https://raw.githubusercontent.com/chollette/Liver-Disease-Classification-Azure-ML-Capstone-Project/master/starter_file/data/Liver%20Patient%20Dataset%20(LPD)_train.csv"
dataset = pd.read_csv(data_path)
# +
# Use the clean_data function to clean your data.
x, y = clean_data(dataset)
train_data = pd.concat([x, y], axis=1, sort=False)
#upload the cleaned marketing data to the default datastore (blob) of my workspace.
#first convert data to .csv
train_data.to_csv('train_data.csv',header=True)
#Then upload to datastore
datastore = ws.get_default_datastore()
datastore.upload_files(['train_data.csv'], target_path='', overwrite=True)
# -
#convert back to tabular dataset for running in AutoML
train_data = Dataset.Tabular.from_delimited_files(path = [(datastore, 'train_data.csv')])
label = "Result"
# ## AutoML Configuration
#
# TODO: Explain why you chose the automl settings and cofiguration you used below.
# + gather={"logged": 1598429217746} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# TODO: Put your automl settings here
automl_settings = {
"experiment_timeout_hours": 1,
#"experiment_timeout_minutes": 30,
"enable_early_stopping" : True,
"model_explainability" : True,
"iteration_timeout_minutes": 5,
"max_concurrent_iterations": 5,
"max_cores_per_iteration": -1,
"n_cross_validations": 10,
"primary_metric": 'accuracy',
"featurization": 'auto',
"verbosity": logging.INFO,
}
# TODO: Put your automl config here
automl_config = AutoMLConfig(task = 'classification',
compute_target=compute_target,
training_data = train_data,
label_column_name = label,
debug_log = "automl_errors.log",
**automl_settings
)
# + gather={"logged": 1598431107951} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# TODO: Submit your experiment
remote_run = experiment.submit(automl_config)
# -
# ## Run Details
#
# OPTIONAL: Write about the different models trained and their performance. Why do you think some models did better than others?
#
# TODO: In the cell below, use the `RunDetails` widget to show the different experiments.
# + gather={"logged": 1598431121770} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#Visualize experiment
RunDetails(remote_run).show()
# -
remote_run.wait_for_completion()
# ## Best Model
#
# TODO: In the cell below, get the best model from the automl experiments and display all the properties of the model.
#
#
# + gather={"logged": 1598431425670} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Retrieve and save your best automl model.
best_automl_run_metrics = remote_run.get_metrics()
print(best_automl_run_metrics)
# -
print("Best AutoML model Accuracy: ", best_automl_run_metrics['accuracy'])
# + gather={"logged": 1598431426111} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#Retrieve model details
best_run, fitted_model = remote_run.get_output()
print(best_run)
print(fitted_model.steps)
# -
#TODO: Save the best model
joblib.dump(fitted_model, 'automl-votingEnsemble_model.joblib')
# ### Register the Fitted Model for Deployment
#Register model
model = best_run.register_model(model_name='model',
model_path='outputs/model.pkl',
tags=best_run.get_metrics())
print(model.name, model.id, model.version, sep='\t')
compute_target.delete()
| Liver-Disease-Classification-Azure-ML-Capstone-Project-master/starter_file/automl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# remove warning message
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
# required library
import cv2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from local_utils import detect_lp
from os.path import splitext,basename
from keras.models import model_from_json
from keras.preprocessing.image import load_img, img_to_array
from keras.applications.mobilenet_v2 import preprocess_input
from sklearn.preprocessing import LabelEncoder
import glob
# -
# ### Part 1: Extract license plate from sample image
def load_model(path):
try:
path = splitext(path)[0]
with open('%s.json' % path, 'r') as json_file:
model_json = json_file.read()
model = model_from_json(model_json, custom_objects={})
model.load_weights('%s.h5' % path)
print("Loading model successfully...")
return model
except Exception as e:
print(e)
wpod_net_path = "wpod-net.json"
wpod_net = load_model(wpod_net_path)
# +
def preprocess_image(image_path,resize=False):
img = cv2.imread(image_path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
img = img / 255
if resize:
img = cv2.resize(img, (224,224))
return img
def get_plate(image_path, Dmax=608, Dmin = 608):
vehicle = preprocess_image(image_path)
ratio = float(max(vehicle.shape[:2])) / min(vehicle.shape[:2])
side = int(ratio * Dmin)
bound_dim = min(side, Dmax)
_ , LpImg, _, cor = detect_lp(wpod_net, vehicle, bound_dim, lp_threshold=0.5)
return vehicle, LpImg, cor
test_image_path = "Plate_examples/germany_car_plate.jpg"
vehicle, LpImg,cor = get_plate(test_image_path)
fig = plt.figure(figsize=(12,6))
grid = gridspec.GridSpec(ncols=2,nrows=1,figure=fig)
fig.add_subplot(grid[0])
plt.axis(False)
plt.imshow(vehicle)
grid = gridspec.GridSpec(ncols=2,nrows=1,figure=fig)
fig.add_subplot(grid[1])
plt.axis(False)
plt.imshow(LpImg[0])
# -
# ## Part 2: Segementing license characters
# +
if (len(LpImg)): #check if there is at least one license image
# Scales, calculates absolute values, and converts the result to 8-bit.
plate_image = cv2.convertScaleAbs(LpImg[0], alpha=(255.0))
# convert to grayscale and blur the image
gray = cv2.cvtColor(plate_image, cv2.COLOR_BGR2GRAY)
blur = cv2.GaussianBlur(gray,(7,7),0)
# Applied inversed thresh_binary
binary = cv2.threshold(blur, 180, 255,
cv2.THRESH_BINARY_INV + cv2.THRESH_OTSU)[1]
kernel3 = cv2.getStructuringElement(cv2.MORPH_RECT, (3, 3))
thre_mor = cv2.morphologyEx(binary, cv2.MORPH_DILATE, kernel3)
# visualize results
fig = plt.figure(figsize=(12,7))
plt.rcParams.update({"font.size":18})
grid = gridspec.GridSpec(ncols=2,nrows=3,figure = fig)
plot_image = [plate_image, gray, blur, binary,thre_mor]
plot_name = ["plate_image","gray","blur","binary","dilation"]
for i in range(len(plot_image)):
fig.add_subplot(grid[i])
plt.axis(False)
plt.title(plot_name[i])
if i ==0:
plt.imshow(plot_image[i])
else:
plt.imshow(plot_image[i],cmap="gray")
# plt.savefig("threshding.png", dpi=300)
# +
# Create sort_contours() function to grab the contour of each digit from left to right
def sort_contours(cnts,reverse = False):
i = 0
boundingBoxes = [cv2.boundingRect(c) for c in cnts]
(cnts, boundingBoxes) = zip(*sorted(zip(cnts, boundingBoxes),
key=lambda b: b[1][i], reverse=reverse))
return cnts
cont, _ = cv2.findContours(binary, cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
# creat a copy version "test_roi" of plat_image to draw bounding box
test_roi = plate_image.copy()
# Initialize a list which will be used to append charater image
crop_characters = []
# define standard width and height of character
digit_w, digit_h = 30, 60
for c in sort_contours(cont):
(x, y, w, h) = cv2.boundingRect(c)
ratio = h/w
if 1<=ratio<=3.5: # Only select contour with defined ratio
if h/plate_image.shape[0]>=0.5: # Select contour which has the height larger than 50% of the plate
# Draw bounding box arroung digit number
cv2.rectangle(test_roi, (x, y), (x + w, y + h), (0, 255,0), 2)
# Sperate number and gibe prediction
curr_num = thre_mor[y:y+h,x:x+w]
curr_num = cv2.resize(curr_num, dsize=(digit_w, digit_h))
_, curr_num = cv2.threshold(curr_num, 220, 255, cv2.THRESH_BINARY + cv2.THRESH_OTSU)
crop_characters.append(curr_num)
print("Detect {} letters...".format(len(crop_characters)))
fig = plt.figure(figsize=(10,6))
plt.axis(False)
plt.imshow(test_roi)
#plt.savefig('grab_digit_contour.png',dpi=300)
# +
fig = plt.figure(figsize=(14,4))
grid = gridspec.GridSpec(ncols=len(crop_characters),nrows=1,figure=fig)
for i in range(len(crop_characters)):
fig.add_subplot(grid[i])
plt.axis(False)
plt.imshow(crop_characters[i],cmap="gray")
#plt.savefig("segmented_leter.png",dpi=300)
# -
# ## Load pre-trained MobileNets model and predict
# +
# Load model architecture, weight and labels
json_file = open('MobileNets_character_recognition.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
model = model_from_json(loaded_model_json)
model.load_weights("License_character_recognition_weight.h5")
print("[INFO] Model loaded successfully...")
labels = LabelEncoder()
labels.classes_ = np.load('license_character_classes.npy')
print("[INFO] Labels loaded successfully...")
# -
# pre-processing input images and pedict with model
def predict_from_model(image,model,labels):
image = cv2.resize(image,(80,80))
image = np.stack((image,)*3, axis=-1)
prediction = labels.inverse_transform([np.argmax(model.predict(image[np.newaxis,:]))])
return prediction
# +
fig = plt.figure(figsize=(15,3))
cols = len(crop_characters)
grid = gridspec.GridSpec(ncols=cols,nrows=1,figure=fig)
final_string = ''
for i,character in enumerate(crop_characters):
fig.add_subplot(grid[i])
title = np.array2string(predict_from_model(character,model,labels))
plt.title('{}'.format(title.strip("'[]"),fontsize=20))
final_string+=title.strip("'[]")
plt.axis(False)
plt.imshow(character,cmap='gray')
print(final_string)
#plt.savefig('final_result.png', dpi=300)
# -
# # The end!
| [Part 3]End-to-end.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reciprocal cycles
# A unit fraction contains 1 in the numerator. The decimal representation of the unit fractions with denominators 2 to 10 are given:
#
# 1/2 = 0.5
# 1/3 = 0.(3)
# 1/4 = 0.25
# 1/5 = 0.2
# 1/6 = 0.1(6)
# 1/7 = 0.(142857)
# 1/8 = 0.125
# 1/9 = 0.(1)
# 1/10 = 0.1
# Where 0.1(6) means 0.166666..., and has a 1-digit recurring cycle. It can be seen that 1/7 has a 6-digit recurring cycle.
#
# Find the value of d < 1000 for which 1/d contains the longest recurring cycle in its decimal fraction part.
# ---
# ### Idea
# A problem of primary math calculation: 'calculate decimal for fraction'
#
# Record remainder of each division, and check if there is a repeatted pattern.
# ---
def get_remainders(d):
remainders = [1]
new_remiander = (remainders[-1] * 10) % d
while new_remiander:
remainders.append(new_remiander)
if new_remiander in remainders[:-1]:
break
new_remiander = (remainders[-1] * 10) % d
return remainders
get_remainders(2)
get_remainders(3)
get_remainders(4)
get_remainders(6)
get_remainders(7)
def get_recurring_interval(remainders):
r = remainders[-1]
for i, rr in enumerate(reversed(remainders[:-1]), 1):
if rr == r:
return i
return 0
get_recurring_interval(get_remainders(3))
get_recurring_interval(get_remainders(4))
get_recurring_interval(get_remainders(7))
def solve(bound):
return max(range(1, bound), key=lambda n: get_recurring_interval(get_remainders(n)))
solve(10)
solve(1000)
| 26-50/p26.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Inteligência Artificial - LAB02
#Execute esta celula
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# +
# import as bibliotecas necessárias
# %matplotlib inline
import numpy as np
import scipy as sp
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import pandas as pd
import time
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 200)
pd.set_option('display.notebook_repr_html', True)
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
import warnings
warnings.filterwarnings('ignore')
# %config InlineBackend.figure_format ='retina'
# -
dataset = sns.load_dataset('breast-cancer');
dataset.info();
dataset.columns
# +
# Qual foi a distribuição de nivel de malignidade dos tumores?
import seaborn as sns
sns.set(color_codes=True)
f, ax = plt.subplots(1,1, figsize=(8, 3));
ax = sns.distplot(dataset.deg_maling, kde=False, bins=20)
# bug
#ax = sns.distplot(titanic.age, kde=False, bins=20).set(xlim=(0, 90));
ax.set(xlim=(0, 4));
ax.set_ylabel('counts');
# -
f, ax = plt.subplots(1,1, figsize=(8, 3))
ax.hist(dataset.deg_maling, bins=20);
ax.set_xlim(0,4);
# +
# configure as cores
cmap = plt.get_cmap('Pastel1')
young = cmap(0.5)
middle = cmap(0.2)
older = cmap(0.8)
# obter o objeto que iremos alterar - patches é uma matriz com len: número de caixas
fig, ax = plt.subplots()
y_values, bins, patches = ax.hist(dataset.deg_maling, 10)
[patches[i].set_facecolor(young) for i in range(0,1)] # bin 0
[patches[i].set_facecolor(middle) for i in range(1,3)] # bins 1 and 2
[patches[i].set_facecolor(older) for i in range(3,10)] # 7 remaining bins
ax.grid(True)
fig.show()
# -
sns.kdeplot(dataset.deg_maling, bw=0.3, label="bw: 0.3", shade=True, color="r");
sns.kdeplot(dataset.deg_maling, bw=2, label="bw: 2", shade=True);
sns.kdeplot(dataset.deg_maling, bw=0.3, label="bw: 0.3", shade=True);
# seaborn
ax = sns.boxplot(x='deg_maling', data = dataset)
#ax = sns.boxplot(x=titanic['age']) # outra forma de codificar
ax.set_ylabel(None);
ax.set_xlabel('deg_maling', fontsize=14);
ax.set_title('Distribuição de idade no Titanic', fontsize=14);
ax = sns.boxplot(x='menopause', y='deg_maling', data=dataset)
print(dataset['deg-maling'].mean())
print(dataset['deg-maling'].median())
dataset.describe()
# +
plt.figure(figsize=(8, 6))
plt.bar(tumorsize['tumor-size'],idade['idade'])
plt.xlabel('Tamanho do tumor')
plt.ylabel('idade dos pacientes')
plt.title('Relação entre o tamanho de tumor com a idade do paciente')
plt.savefig('annual-real-gnp-us-1909-to-1970.png')
plt.show()
# +
plt.figure(figsize=(8, 6))
plt.bar(degmaling['deg-maling'],idade['idade'])
plt.xlabel('Tamanho do tumor')
plt.ylabel('idade dos pacientes')
plt.title('Relação entre o tamanho de tumor com a idade do paciente')
plt.savefig('annual-real-gnp-us-1909-to-1970.png')
plt.show()
| LAB02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Science Unit 4 Sprint Challenge 1 — Tree Ensembles
# ### Chicago Food Inspections
#
# For this Sprint Challenge, you'll use a dataset with information from inspections of restaurants and other food establishments in Chicago from January 1, 2010 to the present.
#
# [See this PDF](https://data.cityofchicago.org/api/assets/BAD5301B-681A-4202-9D25-51B2CAE672FF) for descriptions of the data elements included in this dataset.
#
# According to [Chicago Department of Public Health — Food Protection Services](https://www.chicago.gov/city/en/depts/cdph/provdrs/healthy_restaurants/svcs/food-protection-services.html), "Chicago is home to 16,000 food establishments like restaurants, grocery stores, bakeries, wholesalers, lunchrooms, mobile food vendors and more. Our business is food safety and sanitation with one goal, to prevent the spread of food-borne disease. We do this by inspecting food businesses, responding to complaints and food recalls."
# #### Your challenge: Predict whether inspections failed
#
# The target is the `Fail` column.
#
# - When the food establishment failed the inspection, the target is `1`.
# - When the establishment passed, the target is `0`.
# #### Run this cell to load the data:
# +
import pandas as pd
pd.options.display.max_columns = 200
pd.options.display.max_rows = 200
import category_encoders as ce
import pandas as pd
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from sklearn.tree import DecisionTreeClassifier
import seaborn as sns
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
from sklearn.metrics import roc_auc_score
from sklearn.datasets import make_classification
from sklearn.ensemble import ExtraTreesClassifier
# +
train_url = 'https://drive.google.com/uc?export=download&id=13_tP9JpLcZHSPVpWcua4t2rY44K_s4H5'
test_url = 'https://drive.google.com/uc?export=download&id=1GkDHjsiGrzOXoF_xcYjdzBTSjOIi3g5a'
train = pd.read_csv(train_url)
test = pd.read_csv(test_url)
assert train.shape == (51916, 17)
assert test.shape == (17306, 17)
# -
test.head(1000)
train.isnull().sum(axis = 0)
X_train = train.drop('Fail', axis=1)
X_test = test.drop('Fail', axis=1)
y_train = train['Fail']
y_test = test['Fail']
X_train['Violations'] = X_train['Violations'].fillna('No Violations')
X_train
# +
def wrangle(X):
X = X.copy()
# Drop irrelevant columns
X = X.drop(columns = ['Inspection ID', 'AKA Name', 'License #', 'Address', 'City', 'State', 'Latitude', 'Longitude', 'Location'])
# Change NaN in violations to no violations
X['Violations'] = X['Violations'].fillna('No Violations')
# Remaining NaN....
#Facility Type 224
#Risk 12
#Zip 26
#Inspection Type 1
X = X.fillna('Unknown')
# return wrangled data frame
return X
X_train = wrangle(X_train)
X_test = wrangle(X_test)
X_train.shape, X_test.shape
# +
# Random Forest Improves ROC AUC compared to Decision Tree
from sklearn.ensemble import RandomForestClassifier
pipe = make_pipeline(
ce.OrdinalEncoder(),
RandomForestClassifier(
n_estimators = 100,
class_weight = 'balanced',
min_samples_leaf = .005,
oob_score = True,
n_jobs = -1))
cross_val_score(pipe, X_train, y_train, cv=5, scoring='roc_auc', verbose = 10)
# -
m = pipe.fit(X_train, y_train)
m
from sklearn.metrics import accuracy_score, classification_report, recall_score
#m = RandomForestClassifier(n_estimators=100,min_samples_leaf=3 ,n_jobs=-1,max_features=0.25)
#time m.fit(X1_train, y1_train.values.ravel())
y_pred= m.predict(X_test)
accuracy_score(y_test, y_pred)
encoder = ce.OrdinalEncoder()
X_train = encoder.fit_transform(X_train)
X_test = encoder.transform(X_test)
gb = GradientBoostingClassifier()
gb.fit(X_train, y_train)
y_pred_proba = gb.predict_proba(X_test)[:,1]
print(roc_auc_score(y_test, y_pred_proba))
# +
from pdpbox.pdp import pdp_isolate, pdp_plot
feature='Risk'
pdp_isolated = pdp_isolate(model=gb, dataset=X_test,
model_features=X_test.columns, feature=feature)
pdp_plot(pdp_isolated, feature);
# +
import matplotlib.pyplot as plt
rf = RandomForestClassifier()
rf.fit(X_train, y_train)
rf.score(X_test, y_test)
feature_importances = pd.DataFrame(rf.feature_importances_,
index = X_train.columns,
columns=['importance']).sort_values('importance',ascending=True)
from pylab import rcParams
rcParams['figure.figsize'] = 10, 10
plt.plot(feature_importances)
# -
# ### Part 1: Preprocessing
#
# You may choose which features you want to use, and whether/how you will preprocess them. You may use any tools and techniques for categorical encoding. (Pandas, category_encoders, sklearn.preprocessing, or any other library.)
#
# _To earn a score of 3 for this part, engineer new features, and use any alternative categorical encoding instead of One-Hot or Ordinal/Label encoding._
#
# ### Part 2: Modeling
#
# Fit a Random Forest or Gradient Boosting model with the train set. (You may use scikit-learn, xgboost, or any other library.) Use cross-validation to estimate an ROC AUC validation score.
#
# Use your model to predict probabilities for the test set. Get an ROC AUC test score >= 0.60.
#
# _To earn a score of 3 for this part, get an ROC AUC test score >= 0.70._
#
#
# ### Part 3: Visualization
#
# Make one visualization for model interpretation. (You may use any libraries.) Choose one of these types:
# - Feature Importances
# - Permutation Importances
# - Partial Dependence Plot
#
# _To earn a score of 3 for this part, make at least two of these visualization types._
| SC_4_1/DS41SC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Custom TF-Hub Word Embedding with text2hub
#
# **Learning Objectives:**
# 1. Learn how to deploy AI Hub Kubeflow pipeline
# 1. Learn how to configure the run parameters for text2hub
# 1. Learn how to inspect text2hub generated artifacts and word embeddings in TensorBoard
# 1. Learn how to run TF 1.x generated hub module in TF 2.0
#
#
# ## Introduction
#
#
# Pre-trained text embeddings such as TF-Hub modules are a great tool for building machine learning models for text features, since they capture relationships between words. These embeddings are generally trained on vast but generic text corpora like Wikipedia or Google News, which means that they are usually very good at representing generic text, but not so much when the text comes from a very specialized domain with unique vocabulary, such as in the medical field.
#
#
# One problem in particular that arises when applying a TF-Hub text module which was pre-trained on a generic corpus to specialized text is that all of the unique, domain-specific words will be mapped to the same “out-of-vocabulary” (OOV) vector. By doing so we lose a very valuable part of the text information, because for specialized texts the most informative words are often the words that are very specific to that special domain. Another issue is that of commonly misspelled words from text gathered from say, customer feedback. Applying a generic pre-trained embedding will send the misspelled word to the OOV vectors, losing precious information. However, by creating a TF-Hub module tailored to the texts coming from that customer feedback means that common misspellings present in your real customer data will be part of the embedding vocabulary and should be close by closeby to the original word in the embedding space.
#
#
# In this notebook, we will learn how to generate a text TF-hub module specific to a particular domain using the text2hub Kubeflow pipeline available on Google AI Hub. This pipeline takes as input a corpus of text stored in a GCS bucket and outputs a TF-Hub module to a GCS bucket. The generated TF-Hub module can then be reused both in TF 1.x or in TF 2.0 code by referencing the output GCS bucket path when loading the module.
#
# Our first order of business will be to learn how to deploy a Kubeflow pipeline, namely text2hub, stored in AI Hub to a Kubeflow cluster. Then we will dig into the pipeline run parameter configuration and review the artifacts produced by the pipeline during its run. These artifacts are meant to help you assess how good the domain specific TF-hub module you generated is. In particular, we will explore the embedding space visually using TensorBoard projector, which provides a tool to list the nearest neighbors to a given word in the embedding space.
#
#
# At last, we will explain how to run the generated module both in TF 1.x and TF 2.0. Running the module in TF 2.0 will necessite a small trick that’s useful to know in itself because it allows you to use all the TF 1.x modules in TF hub in TF 2.0 as a Keras layer.
#
#
# !pip freeze | grep tensorflow-hub==0.7.0 || pip install tensorflow-hub==0.7.0
# +
import os
import tensorflow as tf
import tensorflow_hub as hub
# -
# Replace by your GCP project and bucket:
# +
PROJECT = "qwiklabs-gcp-03-3247cf88ddb1" # "your-gcp-project-here" # REPLACE WITH YOUR PROJECT NAME
BUCKET = "qwiklabs-gcp-03-3247cf88ddb1" #"your-gcp-bucket-here" # REPLACE WITH YOUR BUCKET NAME
os.environ["PROJECT"] = PROJECT
os.environ["BUCKET"] = BUCKET
# -
# ## Loading the dataset in GCS
# The corpus we chose is one of [Project Gutenberg medical texts](http://www.gutenberg.org/ebooks/bookshelf/48): [A Manual of the Operations of Surgery](http://www.gutenberg.org/ebooks/24564) by <NAME>, containing very specialized language.
#
# The first thing to do is to upload the text into a GCS bucket:
# + language="bash"
#
# URL=https://www.gutenberg.org/cache/epub/24564/pg24564.txt
# OUTDIR=gs://$BUCKET/custom_embedding
# CORPUS=surgery_manual.txt
#
# curl $URL > $CORPUS
# gsutil cp $CORPUS $OUTDIR/$CORPUS
# -
# It has very specialized language such as
#
# ```
# On the surface of the abdomen the position of this vessel would be
# indicated by a line drawn from about an inch on either side of the
# umbilicus to the middle of the space between the symphysis pubis
# and the crest of the ilium.
# ```
#
# Now let's go over the steps involved in creating your own embedding from that corpus.
# ## Step 1: Download the `text2hub` pipeline from AI Hub (TODO 1)
# Go on [AI Hub](https://aihub.cloud.google.com/u/0/) and search for the `text2hub` pipeline, or just follow [this link](https://aihub.cloud.google.com/u/0/p/products%2F4a91d2d0-1fb8-4e79-adf7-a35707071195).
# You'll land onto a page describing `text2hub`. Click on the "Download" button on that page to download the Kubeflow pipeline and click `Accept`.
# 
# The text2hub pipeline is a KubeFlow pipeline that comprises three components; namely:
#
#
# * The **text2cooc** component that computes a word co-occurrence matrix
# from a corpus of text
#
# * The **cooc2emb** component that factorizes the
# co-occurrence matrix using [Swivel](https://arxiv.org/pdf/1602.02215.pdf) into
# the word embeddings exported as a tsv file
#
# * The **emb2hub** component that takes the word
# embedding file and generates a TF Hub module from it
#
#
# Each component is implemented as a Docker container image that's stored into Google Cloud Docker registry, [gcr.io](https://cloud.google.com/container-registry/). The `pipeline.tar.gz` file that you downloaded is a yaml description of how these containers need to be composed as well as where to find the corresponding images.
#
# **Remark:** Each component can be run individually as a single component pipeline in exactly the same manner as the `text2hub` pipeline. On AI Hub, each component has a pipeline page describing it and from where you can download the associated single component pipeline:
#
# * [text2cooc](https://aihub.cloud.google.com/u/0/p/products%2F6d998d56-741e-4154-8400-0b3103f2a9bc)
# * [cooc2emb](https://aihub.cloud.google.com/u/0/p/products%2Fda367ed9-3d70-4ca6-ad14-fd6bf4a913d9)
# * [emb2hub](https://aihub.cloud.google.com/u/0/p/products%2F1ef7e52c-5da5-437b-a061-31111ab55312)
# ## Step 2: Upload the pipeline to the Kubeflow cluster (TODO 1)
# Go to your [Kubeflow cluster dashboard](https://console.cloud.google.com/ai-platform/pipelines/clusters) or navigate to `Navigation menu > AI Platform > Pipelines` and click `Open Pipelines Dashboard` then click on the `Pipelines` tab to create a new pipeline. You'll be prompted to upload the pipeline file you have just downloaded, click `Upload Pipeline`. Rename the generated pipeline name to be `text2hub` to keep things nice and clean.
# 
# ## Step 3: Create a pipeline run (TODO 1)
# After uploading the pipeline, you should see `text2hub` appear on the pipeline list. Click on it. This will bring you to a page describing the pipeline (explore!) and allowing you to create a run. You can inspect the input and output parameters of each of the pipeline components by clicking on the component node in the graph representing the pipeline. Click `Create Run`.
# 
# ## Step 4: Enter the run parameters (TODO 2)
# `text2hub` has the following run parameters you can configure:
#
# Argument | Description | Optional | Data Type | Accepted values | Default
# ------------------------------------------------ | ------------------------------------------------------------------------------------- | -------- | --------- | --------------- | -------
# gcs-path-to-the-text-corpus | A Cloud Storage location pattern (i.e., glob) where the text corpus will be read from | False | String | gs://... | -
# gcs-directory-path-for-pipeline-output | A Cloud Storage directory path where the pipeline output will be exported | False | String | gs://... | -
# number-of-epochs | Number of epochs to train the embedding algorithm (Swivel) on | True | Integer | - | 40
# embedding-dimension | Number of components of the generated embedding vectors | True | Integer | - | 128
# co-occurrence-word-window-size | Size of the sliding word window where co-occurrences are extracted from | True | Integer | - | 10
# number-of-out-of-vocabulary-buckets | Number of out-of-vocabulary buckets | True | Integer | - | 1
# minimum-occurrences-for-a-token-to-be-considered | Minimum number of occurrences for a token to be included in the vocabulary | True | Integer | - | 5
# You can leave most parameters with their default values except for
# `gcs-path-to-the-test-corpus` whose value should be set to
# !echo gs://$BUCKET/custom_embedding/surgery_manual.txt
# and for `gcs-directory-path-for-pipeline-output` which we will set to
# !echo gs://$BUCKET/custom_embedding
# **Remark**: `gcs-path-to-the-test-corpus` will accept a GCS pattern like `gs://BUCKET/data/*.txt` or simply a path like `gs://BUCKET/data/` to a GCS directory. All the files that match the pattern or that are in that directory will be parsed to create the word embedding TF-Hub module.
# 
# Make sure to choose experiment `default`. Once these values have been set, you can start the run by clicking on `Start`.
# ## Step 5: Inspect the run artifacts (TODO 3)
# Once the run has started you can see its state by going to the `Experiments` tab and clicking on the name of the run (here "text2hub-1").
# 
# It will show you the pipeline graph. The components in green have successfuly completed. You can then click on them and look at the artifacts that these components have produced.
#
# The `text2cooc` components has "co-occurrence extraction summary" showing you the GCS path where the co-occurrence data has been saved. Their is a corresponding link that you can paste into your browser to inspect the co-occurrence data from the GCS browser. Some statistics about the vocabulary are given to you such as the most and least frequent tokens. You can also download the vocabulary file containing the token to be embedded.
# 
# The `cooc2emb` has three artifacts
# * An "Embedding Extraction Summary" providing the information as where the model chekpoints and the embedding tables are exported to on GCP
# * A similarity matrix from a random sample of words giving you an indication whether the model associates close-by vectors to similar words
# * An button to start TensorBoard from the UI to inspect the model and visualize the word embeddings
# 
# We can have a look at the word embedding visualization provided by TensorBoard. Select the TF version: `TensorFlow 1.14.0`. Start TensorBoard by clicking on `Start Tensorboard` and then `Open Tensorboard` buttons, and then select "Projector".
#
# **Remark:** The projector tab may take some time to appear. If it takes too long it may be that your Kubeflow cluster is running an incompatible version of TensorBoard (your TB version should be between 1.13 and 1.15). If that's the case, just run Tensorboard from CloudShell or locally by issuing the following command.
# !echo tensorboard --port 8080 --logdir gs://$BUCKET/custom_embedding/embeddings
# The projector view will present you with a representation of the word vectors in a 3 dimensional space (the dim is reduced through PCA) that you can interact with. Enter in the search tool a few words like "ilium" and points in the 3D space will light up.
# 
# If you click on a word vector, you'll see appear the n nearest neighbors of that word in the embedding space. The nearset neighbors are both visualized in the center panel and presented as a flat list on the right.
#
# Explore the nearest neighbors of a given word and see if they kind of make sense. This will give you a rough understanding of the embedding quality. If it nearest neighbors do not make sense after trying for a few key words, you may need rerun `text2hub`, but this time with either more epochs or more data. Reducing the embedding dimension may help as well as modifying the co-occurence window size (choose a size that make sense given how your corpus is split into lines.)
#
# 
# The `emb2hub` artifacts give you a snippet of TensorFlow 1.x code showing you how to re-use the generated TF-Hub module in your code. We will demonstrate how to use the TF-Hub module in TF 2.0 in the next section.
# 
# # Step 7: Using the generated TF-Hub module (TODO)
# Let's see now how to load the TF-Hub module generated by `text2hub` in TF 2.0.
#
# We first store the GCS bucket path where the TF-Hub module has been exported into a variable:
MODULE = "gs://{bucket}/custom_embedding/hub-module".format(bucket=BUCKET)
MODULE
# Now we are ready to create a `KerasLayer` out of our custom text embedding.
med_embed = hub.KerasLayer(MODULE)
# That layer when called with a list of sentences will create a sentence vector for each sentence by averaging the word vectors of the sentence.
outputs = med_embed(tf.constant(['ilium', 'I have a fracture', 'aneurism']))
outputs
# If you use a version of TensorFlow Hub smaller than `tensorflow-hub==0.7.0`, then you'll need to use the following wrapper to instanciate the `KerasLayer`:
# ```python
# class Wrapper(tf.train.Checkpoint):
# def __init__(self, spec):
# super(Wrapper, self).__init__()
# self.module = hub.load(spec)
# self.variables = self.module.variables
# self.trainable_variables = []
# def __call__(self, x):
# return self.module.signatures["default"](x)["default"]
#
# med_embed = hub.KerasLayer(Wrapper(MODULE))
# ```
# Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive2/text_classification/solutions/custom_tf_hub_word_embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Ezu9OhdMDaES" colab_type="text"
# # Testing Altair-Saver
#
# <a href="https://colab.research.google.com/github/altair-viz/altair_saver/blob/master/AltairSaver.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# This notebook demonstrates the installation and use of [altair-saver](http://github.com/altair-viz/altair_saver). The following was tested in Colab.
#
#
# + id="ZiTDBCAM_Ni8" colab_type="code" outputId="547c4b1f-6d58-4356-9a41-6aed648dcc7b" colab={"base_uri": "https://localhost:8080/", "height": 68}
# !pip install -q altair_saver
# + [markdown] id="KJh7JMMzDjae" colab_type="text"
# ## Setup Selenium + Chromedriver
# + colab_type="code" id="lIYdn1woOS1n" outputId="ca17198b-8895-4d2e-c7be-c5bf033a2c9a" colab={"base_uri": "https://localhost:8080/", "height": 391}
# !apt-get -qq install chromium-chromedriver
# + [markdown] id="4pjnxI1bDosm" colab_type="text"
# ## Setup npm and the Vega CLI
# + id="if6t54-6_DvU" colab_type="code" outputId="6e247009-9fe0-44bc-ee1e-738d9d8cad65" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !npm install --silent vega-lite vega-cli canvas
# + [markdown] id="R5126EyEN4hT" colab_type="text"
# ## Create and save a chart
# + id="qd-Tcr8LBSqM" colab_type="code" outputId="05b179f7-abd4-4a64-ad2b-f943be0302cc" colab={"base_uri": "https://localhost:8080/", "height": 368}
import altair as alt
from vega_datasets import data
cars = data.cars.url
chart = alt.Chart(cars).mark_bar().encode(
x=alt.X('Miles_per_Gallon:Q', bin=True),
y='count()',
)
chart.display()
# + id="fAQLIM7qBX_u" colab_type="code" outputId="c40e4c70-3f72-4554-c071-07d3437c00ef" colab={"base_uri": "https://localhost:8080/", "height": 119}
from altair_saver import save
for fmt in ['json', 'vg.json', 'html', 'png', 'svg', 'pdf']:
save(chart, f'chart.{fmt}')
# !ls -lh chart.*
# + [markdown] id="x5v8HdYJGO3B" colab_type="text"
# ## View saved charts
#
# Here we use a variety of IPython display mechanisms to load and display the saved charts.
# + id="QVO2E2Y7Bhxx" colab_type="code" outputId="33c18996-d86b-4607-d6ad-b9f6ef8bba8b" colab={"base_uri": "https://localhost:8080/", "height": 364}
from PIL import Image
Image.open("chart.png")
# + id="ievDaZFaB6A1" colab_type="code" outputId="00d3ae91-1693-4332-d633-ad85e6ddf349" colab={"base_uri": "https://localhost:8080/", "height": 368}
from IPython.display import display, SVG
with open("chart.svg") as f:
display(SVG(f.read()))
# + id="zUMlMKIKCQIW" colab_type="code" outputId="f6c3630c-498a-4f44-92a4-41321eb70112" colab={"base_uri": "https://localhost:8080/", "height": 368}
import json
with open('chart.json') as f:
display(alt.VegaLite(json.load(f)))
# + id="ouREXcHvCkV0" colab_type="code" outputId="2dba7fbb-7b39-4402-dee5-f2f3bc2dc694" colab={"base_uri": "https://localhost:8080/", "height": 368}
import json
from altair import vega
vega.renderers.enable('colab')
with open('chart.vg.json') as f:
display(vega.Vega(json.load(f)))
# + id="JQU9wGCa21K-" colab_type="code" outputId="23317414-f3c2-4049-83e0-c3df3c432cc6" colab={"base_uri": "https://localhost:8080/", "height": 368}
from IPython.display import HTML
with open("chart.html") as f:
html = f.read()
HTML(html)
# + id="LTPTM_ZMJekM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1293f7cb-fe38-4ab7-ce37-c60a1192369e"
from IPython.display import HTML
import base64
with open("chart.pdf", 'rb') as f:
pdf_base64 = base64.b64encode(f.read()).decode()
HTML(f'Right-click and choose "Open In New Tab": <a download="chart.pdf" href="data:application/pdf;base64,{pdf_base64}">chart.pdf</a>')
# + [markdown] id="RT6t8A_4ie2Y" colab_type="text"
# ## Renderers
#
# Alternatively, you can enable an altair-viewer renderer and display the chart within a notebook as a specific type or set of types.
#
# For example, the following encodes both the vega-lite custom mimetype (which is supported by JupyterLab and other frontends via custom frontend extensions), as well as a PNG fallback that will be displayed in other environments:
# + id="IlgGMKKUixmL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 364} outputId="0d70e8b8-0014-4522-c64b-4b4bce9ed362"
alt.renderers.enable('altair_saver', fmts=['vega-lite', 'png'])
chart.display()
# + [markdown] id="nyZ_H5jZmrNx" colab_type="text"
# The content of the output can be confirmed by looking at the ``_repr_mimebundle_`` method, which is the special method used by IPython/Jupyter for rich rendering:
# + id="2uYnHFManE7t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 190} outputId="ef214440-0dcc-4314-adfb-47dc0c554785"
chart._repr_mimebundle_(include=None, exclude=None)
| AltairSaver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Experiment 1 - Xgboost
#
# Aim is to get a baseline model running and a prediction output for first kaggle submission.
#
# Need to preprocess the data and save to relevant folder, as well as save model and output file.
#
# Perform some light EDA to get insights about the dataset.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ## EDA
# import dataset - train
training_data = pd.read_csv('../data/raw/train (1).csv')
# inspect the data
training_data.head()
training_data.info()
training_data.describe()
training_data.shape
# how many distinct id's are there?
len(training_data.Id.unique())
# check distribution of the target
training_data.TARGET_5Yrs.value_counts()
# All variables are numeric, no missing values. Target is the last column, and use Id at column 2, remove column 1.
#
# 0 class is underrepresented, ratio about 16% 0, 84% 1
#
# This could affect the power of the model
# ## Prepare data for model
df_cleaned = training_data.copy()
# remove id_old
df_cleaned.drop('Id_old', axis=1, inplace=True)
df_cleaned.shape
# create Y
target = df_cleaned.pop('TARGET_5Yrs')
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(df_cleaned, target, test_size=0.2, random_state=8)
# ### Create the worst base model
# Worst case "dumb" model which predicts all 1's. Establish a baseline from which to improve.
# create a vector of all 1's
y_dumb = np.ones(len(y_train), dtype=np.int64)
print(y_dumb.size)
print(y_dumb)
# +
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
print(mse(y_train, y_dumb, squared=False))
print(mae(y_train, y_dumb))
# +
from sklearn.metrics import roc_auc_score
roc=roc_auc_score(y_train, y_dumb)
print("AUC: %.2f%% " % (roc *100))
# -
np.save('../data/processed/X_train', X_train)
np.save('../data/processed/X_val', X_val)
np.save('../data/processed/y_train', y_train)
np.save('../data/processed/y_val', y_val)
# ## Train model
# Following base model is based off this example https://machinelearningmastery.com/develop-first-xgboost-model-python-scikit-learn/
pip install xgboost
from xgboost import XGBClassifier
# instatiate model
model = XGBClassifier()
# fit on train set
model.fit(X_train, y_train)
# Save output to models folder
from joblib import dump
dump(model, '../models/xgboost.joblib')
# ## Predict
# predict class
y_train_preds = model.predict(X_train)
y_val_preds = model.predict(X_val)
# predict proabilities
y_train_preds_prob = model.predict_proba(X_train)
y_val_preds_prob = model.predict_proba(X_val)
print(y_train_preds_prob)
print(y_val_preds_prob)
from sklearn.metrics import mean_squared_error as mse
from sklearn.metrics import mean_absolute_error as mae
# +
print(mse(y_train, y_train_preds, squared=False))
print(mae(y_train, y_train_preds))
print(mse(y_val, y_val_preds, squared=False))
print(mae(y_val, y_val_preds))
# +
from sklearn.metrics import accuracy_score
accuracy = accuracy_score(y_val, y_val_preds)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# -
from sklearn.metrics import roc_auc_score ,recall_score, precision_score
precision=precision_score(y_val, y_val_preds)
recall=recall_score(y_val, y_val_preds)
roc=roc_auc_score(y_val, y_val_preds)
print("Precision: %.2f%% " % (precision *100))
print("Recall: %.2f%% " % (recall * 100))
print("AUC: %.2f%% " % (roc *100))
roc_auc_score(y_train, y_train_preds_prob[:,1])
roc_auc_score(y_val, y_val_preds_prob[:,1])
# Plot roc curve using code from here https://www.analyticsvidhya.com/blog/2020/06/auc-roc-curve-machine-learning/
# +
from sklearn.metrics import roc_curve
# roc curve for models
fpr, tpr, thresh = roc_curve(y_val, y_val_preds_prob[:,1], pos_label=1)
# roc curve for tpr = fpr
random_probs = [0 for i in range(len(y_val))]
p_fpr, p_tpr, _ = roc_curve(y_val, random_probs, pos_label=1)
# +
# use matplotlib to plot ROC curve
plt.style.use('seaborn')
# plot roc curves
plt.plot(fpr, tpr, linestyle='--',color='orange', label='XgBoost base')
plt.plot(p_fpr, p_tpr, linestyle='--', color='blue')
# title
plt.title('ROC curve')
# x label
plt.xlabel('False Positive Rate')
# y label
plt.ylabel('True Positive rate')
plt.legend(loc='best')
#plt.savefig('ROC',dpi=300)
plt.show()
# -
# predict on test set
test_data = pd.read_csv('../data/raw/test (1).csv')
test_data.info()
# remove old Id
test_cleaned = test_data.copy()
test_cleaned = test_cleaned.drop('Id_old', axis=1, inplace=True)
test_cleaned
y_test_preds = model.predict(test_cleaned)
y_test_preds_prob = model.predict_proba(test_cleaned)
print(y_test_preds)
print(y_test_preds_prob)
y_test_preds.sum()
# They are not all 1's, hooray!
# output predictions
id_col = test_cleaned.loc[:,['Id']]
print(id_col)
print(id_col.dtype)
probabilities = pd.DataFrame(y_test_preds_prob[:,1], columns = ['TARGET_5Yrs'])
print(probabilities)
# +
# concat columns
output = pd.concat([id_col,probabilities], axis=1)
#df_output = pd.DataFrame(output_list, columns = ['Id','TARGET_5Yrs'])
print(output)
# -
# save to csv
output.to_csv('../data/processed/output_xgboostbase.csv',index=False)
# # Kaggle AUC = 0.64325
| notebooks/tith_reasmey-10845345-week1_basemodel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.1
# language: julia
# name: julia-1.1
# ---
include("setup.jl")
# # Facility location problem
#
# **Originally Contributed by**: <NAME> and <NAME>
#
# Benchmark instances: http://resources.mpi-inf.mpg.de/departments/d1/projects/benchmarks/UflLib/
#
# ## Uncapacitated facility location
#
# ### Problem description
#
# * $M=\{1, \dots, m\}$ clients, $N=\{ 1, \dots, n\}$ sites where a facility can be built.
n = 2 # number of facility locations
m = 5 # number of clients
# * $f_j$: fixed cost of building a facility at site $j$
# * $c_{i, j}$: cost for serving customer $i$ from facility $j$
# Draw costs
Random.seed!(0)
f = rand(1000:10000, n);
c = rand(0:1000, m, n);
# ### MILP formulation
#
# $$
# \begin{array}{cl}
# \min_{x, y} \ \ \ &
# \sum_{i, j} c_{i, j} x_{i, j} +
# \sum_{j} f_{j} y_{j}\\
# s.t. &
# \sum_{j} x_{i, j} = 1, \ \ \ \forall i \in M\\
# & x_{i, j} \leq y_{j}, \ \ \ \forall i \in M, j \in N\\
# & x_{i, j}, y_{j} \in \{0, 1\}, \ \ \ \forall i \in M, j \in N
# \end{array}
# $$
model = Model();
# Create y variables
@variable(model, y[1:m], Bin);
@variable(model, x[1:m, 1:n], Bin);
# +
# set objective
F = sum([f[j]*y[j] for j in 1:n])
C = sum([c[i, j]*x[i, j] for i in 1:m for j in 1:n])
@objective(model, Min, F + C);
# +
# Add constraints
# Each client is serve exactly once
@constraint(
model,
[i in 1:m],
sum(x[i, j] for j in 1:n) == 1
)
# Fixed cost of opening facilities
@constraint(
model,
[i in 1:m, j in 1:n],
x[i, j] <= y[j]
);
# -
# Set optimizer
set_optimizer(model, with_optimizer(GLPK.Optimizer))
optimize!(model)
println("Optimal value: ", objective_value(model))
# Get y and x solutions
ysol = value.(y);
println("Optimal solution y: ", value.(y))
xsol = value.(x);
println("Optimal solution x: ", value.(x))
# +
# relax all binary variables
for var in x
is_binary(var) && unset_binary(var)
set_lower_bound(var, 0.0)
set_upper_bound(var, 1.0)
end
for var in y
is_binary(var) && unset_binary(var)
set_lower_bound(var, 0.0)
set_upper_bound(var, 1.0)
end
# Solve the LP relaxation
optimize!(model)
lp_val = objective_value(model)
println("Optimal value of relaxed model: ", lp_val)
# -
# Get y and x solutions
lp_ysol = value.(y);
println("Optimal solution y: ", value.(y))
lp_xsol = value.(x);
println("Optimal solution x: ", value.(x))
# +
# set all variables to be binary
for var in x
set_binary(var)
end
for var in y
set_binary(var)
end
optimize!(model)
mip_val = objective_value(model)
println("Optimal value of integer model: ", mip_val)
# -
(mip_val - lp_val) / mip_val # integrality gap
# ## Capacitated Facility location
#
# * Each client $i$ has a demand $a_{i}$, and each facility has a capacity $q_{j}$
# $$
# \begin{array}{cl}
# \min_{x, y} \ \ \ &
# \sum_{i, j} c_{i, j} x_{i, j} +
# \sum_{j} f_{j} y_{j}\\
# s.t. &
# \sum_{j} x_{i, j} = 1, \ \ \ \forall i \in M\\
# & \sum_{i} a_{i} x_{i, j} \leq q_{j} y_{j}, \ \ \ \forall j \in N\\
# & x_{i, j}, y_{j} \in \{0, 1\}, \ \ \ \forall i \in M, j \in N
# \end{array}
# $$
# +
n = 10 # number of facility locations
m = 30 # number of clients
# Draw costs
Random.seed!(0)
f = rand(1000:10000, n);
c = rand(0:1000, m, n);
# Clients' demands
a = rand(1:10, m);
# Capacities
q = rand(30:40, n);
# -
# Instantiate an empty model
model_cap = Model();
# Create variables
y = @variable(model_cap, y[1:n], Bin);
x = @variable(model_cap, x[1:m, 1:n], Bin);
# +
# set objective
C = sum([c[i, j]*x[i, j] for i in 1:m for j in 1:n]) # demand serving cost
F = sum([f[j]*y[j] for j in 1:n]) # fixed cost
@objective(model_cap, Min, C + F);
# +
# Add constraints
# Each client is serve exactly once
ctr_ = @constraint(
model_cap, # add constraints to model
[i in 1:m], # there are `m` constraints, indexed by `i`
sum(x[i, j] for j in 1:n) == 1 # the actual constraint
)
# Capacity constraints
ctr_capacity = @constraint(
model_cap,
[j in 1:n],
sum(a[i] * x[i, j] for i in 1:m) <= q[j]*y[j]
);
# Tighten LP relaxation
ctr_opening = @constraint(
model_cap,
[i in 1:m, j in 1:n],
x[i, j] <= y[j]
);
# -
# Set optimizer
set_optimizer(
model_cap,
with_optimizer(
GLPK.Optimizer,
msg_lev=3, # verbosity level
tm_lim=10000 # time limit, in ms
)
)
# Best solution found so far
println("Optimal value: ", objective_value(model_cap))
# try with Cbc
set_optimizer(model_cap, with_optimizer(Cbc.Optimizer))
optimize!(model_cap)
| FacilityLocation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: nft_analytics
# language: python
# name: nft_analytics
# ---
# # Find well priced NFTs on rising collections
#
# **Old calculation**
# ### ThorGuards #2716
# - Date: 15.11.2021
# - Price: 1.75 ETH
# - URL: https://opensea.io/assets/0xa98b29a8f5a247802149c268ecf860b8308b7291/390
#
# **Using Rarity tools calculation**
# ### Boss Beauties #2380
# - Date: 15.11.2021
# - Price: 4.9 ETH
# - URL: https://opensea.io/assets/0xb5c747561a185a146f83cfff25bdfd2455b31ff4/2380
#
# ### Gutter Dog #1290
# - Date: 15.11.2021
# - Price: 1.65 ETH
# - URL: https://opensea.io/assets/0x6e9da81ce622fb65abf6a8d8040e460ff2543add/1290
#
# ### DeadFellaz #8384
# - Date: 15.11.2021
# - Price: 1.29 ETH
# - URL: https://opensea.io/assets/0x2acab3dea77832c09420663b0e1cb386031ba17b/8545
#
# ### Kong #7981
# - Date: 15.11.2021
# - Price: 69.42 ETH
# - URL: https://opensea.io/assets/0xef0182dc0574cd5874494a120750fd222fdb909a/7981
#
# ### Cryptovoxels 1 Rack Pass
# - Date: 15.11.2021
# - Price: 3.5 ETH
# - URL: https://opensea.io/assets/0x79986af15539de2db9a5086382daeda917a9cf0c/2089
#
# ### The Shiboshis
# - Date: 15.11.2021
# - Price: ETH
# - URL:
# +
import os
from datetime import datetime
from dateutil.parser import parse
import time
import numpy as np
import pandas as pd
from scipy import interpolate
from scipy.optimize import curve_fit
import scipy.stats as scs
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
from pprint import pprint
from tqdm import tqdm
import seaborn as sns
sns.set_theme(style="whitegrid")
from src.nft_analytics import NFTAnalytics
from src.infura_api import InfuraAPI
# -
items_in_collection = 10000
nft = NFTAnalytics("0x11450058d796b02eb53e65374be59cff65d3fe7f")
asset_data = nft.fetch_data(max_offset=items_in_collection)
asset_data2 = nft.fetch_data(max_offset=1)
asset_data2
df = nft.calculate_rarity_df(asset_data, items_in_collection)
# +
df = pd.DataFrame(columns=["Name", "Price", "Rarity", "RarityPriceRatio"])
for idx, asset in enumerate(asset_data):
if asset["sell_orders"]:
if asset["sell_orders"][0]["payment_token_contract"]["symbol"] == "ETH":
price = float(asset["sell_orders"][0]["current_price"]) / 1e18
if price != 0:
rarity = 0
for trait in asset["traits"]:
trait_count = int(trait["trait_count"])
if trait_count != 0:
rarity += 1 / (trait_count / items_in_collection)
name = asset["name"]
df.loc[idx] = [name, price, rarity, rarity / price]
# -
# Filter out top 20% in rarity
df[df["Rarity"] > df["Rarity"].min() + 0.1 * (df["Rarity"].max() - df["Rarity"].min())].sort_values("RarityPriceRatio", ascending=False)
df.sort_values("RarityPriceRatio", ascending=False)
| notebooks/UndervaluedNFTs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 5.01: Ordinary Least Squares (OLS) as a Classifier
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
df = pd.read_csv('../Datasets/linear_classifier.csv')
df.head()
plt.figure(figsize=(10, 7))
for label, label_class in df.groupby('labels'):
plt.scatter(label_class.values[:,0], label_class.values[:,1],
label=f'Class {label}', marker=label, c='k')
plt.legend()
plt.title("Linear Classifier");
df_train, df_test = train_test_split(df.copy(), test_size=0.4, random_state=12)
# +
# Fit a linear regression model
model = LinearRegression()
model.fit(df_train.x.values.reshape((-1, 1)), df_train.y.values.reshape((-1, 1)))
# Print out the parameters
print(f'y = {model.coef_[0][0]}x + {model.intercept_[0]}')
# +
# Plot the trendline
trend = model.predict(np.linspace(0, 10).reshape((-1, 1)))
plt.figure(figsize=(10, 7))
for label, label_class in df_test.groupby('labels'):
plt.scatter(label_class.values[:,0], label_class.values[:,1],
label=f'Class {label}', marker=label, c='k')
plt.plot(np.linspace(0, 10), trend, c='k', label='Trendline')
plt.legend()
plt.title("Linear Classifier");
# +
# Make predictions
y_pred = model.predict(df_test.x.values.reshape((-1, 1)))
pred_labels = []
for _y, _y_pred in zip(df_test.y, y_pred):
if _y < _y_pred:
pred_labels.append('o')
else:
pred_labels.append('x')
df_test['Pred Labels'] = pred_labels
df_test.head()
# +
plt.figure(figsize=(10, 7))
for idx, label_class in df_test.iterrows():
if label_class.labels != label_class['Pred Labels']:
label = 'D'
s=70
else:
label = label_class.labels
s=50
plt.scatter(label_class.values[0], label_class.values[1],
label=f'Class {label}', marker=label, c='k', s=s)
plt.plot(np.linspace(0, 10), trend, c='k', label='Trendline')
plt.title("Linear Classifier");
incorrect_class = mlines.Line2D([], [], color='k', marker='D',
markersize=10, label='Incorrect Classification');
plt.legend(handles=[incorrect_class]);
# -
| Chapter05/Exercise5.01/Exercise5.01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploration Notebook to scrape several websites for job openings, store them in a database and build an own recommender for interesting jobs
#
# #### Use chrome inspector to get url, parameter values and expectet request method of website/API
# #### If data is received in JSON, use a JSON viewer or the preview window in chrome inspector to find where exactly in JSON file the intersting data is stored
# #### I search for thesis offers in the area of Data Science/Machine Learning/AI near Stuttgart, Germany at the moment . Therefore I scrape the larger and well-known companies in this area, which always have a few thesis offers on their websites
#
import undercover_request # custom module, install: pip install git+https://github.com/TimSchopf/undercover_request
import pandas as pd
import sqlite3
import os
# construct empty DataFrame, where scraped data gets temporary stored
data = pd.DataFrame(columns=['Title','Company','Location','Business_Unit','Description','Qualifications','ApplyURL','Release_Date','Valid','Interesting','New'])
# ## Scrape Daimler Carrer Website
# Daimler uses an API to send the data to the client.
# The Daimler Carrer Website itself is just a HTML template without data.
# Data and template render client side and are sent seperately to the client.
# In this case we must find the API URL that returns the raw data (the Daimler API sends JSON).
# The API URL already conatins the searched keywords, we don't need to append the daimler_keywords additionally
# searched keywords: thesis
daimler_keywords = {'_': 1555512030476}
daimler_api_url = 'https://global-jobboard-api.daimler.com/v3/search/%7B%22LanguageCode%22%3A%22DE%22%2C%22ScoreThreshold%22%3A80%2C%22SearchParameters%22%3A%7B%22MatchedObjectDescriptor%22%3A%5B%22PositionID%22%2C%22PositionTitle%22%2C%22PositionURI%22%2C%22LogoURI%22%2C%22OrganizationName%22%2C%22Organization%22%2C%22Organization.MemberCode%22%2C%22ParentOrganization%22%2C%22PositionLocation.CityName%22%2C%22PositionLocation.Longitude%22%2C%22PositionLocation.Latitude%22%2C%22PositionIndustry.Name%22%2C%22JobCategory.Name%22%2C%22JobCategory.Code%22%2C%22CareerLevel.Name%22%2C%22CareerLevel.Code%22%2C%22Facet%3AParentOrganization%22%2C%22Facet%3APositionLocation.CityName%22%2C%22Facet%3APositionLocation.CountryName%22%2C%22PublicationStartDate%22%2C%22ParentOrganizationGenesisID%22%5D%2C%22FirstItem%22%3A0%2C%22CountItem%22%3A1000000%7D%2C%22SearchCriteria%22%3A%5B%7B%22CriterionName%22%3A%22PublicationLanguage.Code%22%2C%22CriterionValue%22%3A%5B%22DE%22%5D%7D%2C%7B%22CriterionName%22%3A%22PublicationChannel.Code%22%2C%22CriterionValue%22%3A%5B%2212%22%5D%7D%2C%7B%22CriterionName%22%3A%22CareerLevel.Code%22%2C%22CriterionValue%22%3A%5B%2240%22%5D%7D%5D%7D?_=1555512030476'
daimler_req = undercover_request.request(daimler_api_url, request_type='get')
# +
# parse data
daimler_data = daimler_req.json()
# number of total hits for Daimler API
daimler_hits = daimler_data['SearchResult']['SearchResultCountAll']
# extract data from JSON and add it to DataFrame
for i in range(daimler_hits):
# job title
daimler_title = daimler_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionTitle']
# company name
daimler_company_logo_str = daimler_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['LogoURI'][0]
daimler_company = 'Daimler AG: ' + daimler_company_logo_str[25:len(daimler_company_logo_str)-14]
# location
daimler_location = daimler_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionLocation'][0]['CityName']
# business unit
daimler_business_unit = daimler_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['JobCategory'][0]['Name']
# no detailed job description and qualifications in Daimler JSON file
daimler_description = float('NaN')
daimler_qualifications = float('NaN')
# url to detailed job descrition and application possibility
daimler_applyURL = daimler_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionURI']
# release date of job offer
daimler_releaseDate = daimler_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PublicationStartDate']
# append to 'data' DataFrame
df = pd.DataFrame(data={'Title':[daimler_title],'Company':[daimler_company],'Location':[daimler_location],'Business_Unit':[daimler_business_unit],'Description':[daimler_description],'Qualifications':[daimler_qualifications],'ApplyURL':[daimler_applyURL],'Release_Date':[daimler_releaseDate],'Valid':[1],'Interesting':[float('NaN')],'New':[1]})
data = data.append(df, ignore_index=True)
# -
# ## Scrape Bosch Career Website
# +
# Bosch also uses an API, returns JSON and expects JSON post parameter
# The Bosch API only delivers 9 results at a time, but more results can be loaded on the website via load more button
# searched keywords: thesis
bosch_keywords = {"from":0,"size":9,"sort":[{"releasedDate":{"order":"desc"}}],"query":{"query_string":{"query":"thesis*","default_operator":"and"}}}
bosch_req = undercover_request.request('https://ro-dyn-backend.bosch.com/c-corpweb-tc/es/bosch-ro-backend*/ro_bosch_backend__job_posting/_search', request_type='post', json=bosch_keywords)
# +
# parse data
bosch_data = bosch_req.json()
# number of total hits for Bosch API
bosch_hits = bosch_data['hits']['total']
# -
# set size of post request parameter to number of total hits and get all hits at once, not only 9
bosch_keywords_test = {"from":0,"size":bosch_hits,"sort":[{"releasedDate":{"order":"desc"}}],"query":{"query_string":{"query":"thesis*","default_operator":"and"}}}
bosch_req = undercover_request.request('https://ro-dyn-backend.bosch.com/c-corpweb-tc/es/bosch-ro-backend*/ro_bosch_backend__job_posting/_search', request_type='post', json=bosch_keywords_test)
# +
# parse data
bosch_data = bosch_req.json()
# number of total hits for Bosch API
bosch_hits = bosch_data['hits']['total']
# extract data from JSON and add it to DataFrame
for i in range(bosch_hits):
# job title
bosch_title = bosch_data['hits']['hits'][i]['_source']['name']
# company name
bosch_company = bosch_data['hits']['hits'][i]['_source']['company']['name']
# loaction
bosch_city = bosch_data['hits']['hits'][i]['_source']['location']['city']
bosch_country = bosch_data['hits']['hits'][i]['_source']['location']['country']
bosch_postalCode = bosch_data['hits']['hits'][i]['_source']['location']['postalCode']
bosch_region = bosch_data['hits']['hits'][i]['_source']['location']['region']
bosch_location = str(bosch_postalCode) + ', ' + bosch_city + ', ' + bosch_country + ', ' + bosch_region
# business unit
bosch_business_unit = bosch_data['hits']['hits'][i]['_source']['function']['label']
# detailed job description
bosch_description = bosch_data['hits']['hits'][i]['_source']['jobAd']['sections']['jobDescription']['text']
# job qualifications
bosch_qualifications = bosch_data['hits']['hits'][i]['_source']['jobAd']['sections']['qualifications']['text']
# url to detailed job descrition and application possibility
bosch_applyURL = bosch_data['hits']['hits'][i]['_source']['applyUrl']
# release date of job offer
bosch_releaseDate = bosch_data['hits']['hits'][i]['_source']['releasedDate']
# append to 'data' DataFrame
df = pd.DataFrame(data={'Title':[bosch_title],'Company':[bosch_company],'Location':[bosch_location],'Business_Unit':[bosch_business_unit],'Description':[bosch_description],'Qualifications':[bosch_qualifications],'ApplyURL':[bosch_applyURL],'Release_Date':[bosch_releaseDate],'Valid':[1],'Interesting':[float('NaN')],'New':[1]})
data = data.append(df, ignore_index=True)
# -
# ## Scrape Porsche Career Website
# Posrsche also uses an API
# The API URL already conatins the searched keywords, we don't need to append the porsche_keywords additionally
# The API sends 10 results at a time, we need to modify the 'CountItem' Parameter or the url string to get all results/hits
porsche_keywords = '{"LanguageCode":"DE","SearchParameters":{"FirstItem":1,"CountItem":10,"Sort":[{"Criterion":"PublicationStartDate","Direction":"DESC"}],"MatchedObjectDescriptor":["ID","PositionTitle","PositionURI","PositionLocation.CountryName","PositionLocation.CityName","PositionLocation.Longitude","PositionLocation.Latitude","PositionLocation.PostalCode","PositionLocation.StreetName","PositionLocation.BuildingNumber","PositionLocation.Distance","JobCategory.Name","PublicationStartDate","ParentOrganizationName","ParentOrganization","OrganizationShortName","CareerLevel.Name","JobSector.Name","PositionIndustry.Name","PublicationCode","PublicationChannel.Id"]},"SearchCriteria":[{"CriterionName":"PublicationChannel.Code","CriterionValue":["12"]},{"CriterionName":"CareerLevel.Code","CriterionValue":["6"]}]}'
porsche_api_url = 'https://api-jobs.porsche.com/search/?data=%7B%22LanguageCode%22%3A%22DE%22%2C%22SearchParameters%22%3A%7B%22FirstItem%22%3A1%2C%22CountItem%22%3A10%2C%22Sort%22%3A%5B%7B%22Criterion%22%3A%22PublicationStartDate%22%2C%22Direction%22%3A%22DESC%22%7D%5D%2C%22MatchedObjectDescriptor%22%3A%5B%22ID%22%2C%22PositionTitle%22%2C%22PositionURI%22%2C%22PositionLocation.CountryName%22%2C%22PositionLocation.CityName%22%2C%22PositionLocation.Longitude%22%2C%22PositionLocation.Latitude%22%2C%22PositionLocation.PostalCode%22%2C%22PositionLocation.StreetName%22%2C%22PositionLocation.BuildingNumber%22%2C%22PositionLocation.Distance%22%2C%22JobCategory.Name%22%2C%22PublicationStartDate%22%2C%22ParentOrganizationName%22%2C%22ParentOrganization%22%2C%22OrganizationShortName%22%2C%22CareerLevel.Name%22%2C%22JobSector.Name%22%2C%22PositionIndustry.Name%22%2C%22PublicationCode%22%2C%22PublicationChannel.Id%22%5D%7D%2C%22SearchCriteria%22%3A%5B%7B%22CriterionName%22%3A%22PublicationChannel.Code%22%2C%22CriterionValue%22%3A%5B%2212%22%5D%7D%2C%7B%22CriterionName%22%3A%22CareerLevel.Code%22%2C%22CriterionValue%22%3A%5B%226%22%5D%7D%5D%7D'
porsche_req = undercover_request.request(porsche_api_url, request_type='get')
# +
# parse data
porsche_data = porsche_req.json()
# total number of hits from Porsche API
porsche_hits = porsche_data['SearchResult']['SearchResultCountAll']
# -
# modify API url string to get all search results at once
porsche_api_url = porsche_api_url[0:145] + str(porsche_hits) + porsche_api_url[147:len(porsche_api_url)]
porsche_req = undercover_request.request(url=porsche_api_url, request_type='get')
# +
# parse data
porsche_data = porsche_req.json()
# total number of hits from Porsche API
porsche_hits = porsche_data['SearchResult']['SearchResultCountAll']
# extract data from JSON and add it to DataFrame
for i in range(porsche_hits):
# job title
porsche_title = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionTitle']
# company name
porsche_company = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['ParentOrganizationName']
# location
porsche_city = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionLocation'][0]['CityName']
porsche_country = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionLocation'][0]['CountryName']
porsche_location = porsche_city + ', ' + porsche_country
#business unit
porsche_business_unit = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['JobCategory'][0]['Name']
# no detailed job description and qualifications in Porsche JSON file
porsche_description = float('NaN')
porsche_qualifications = float('NaN')
# url to detailed job descrition and application possibility
porsche_applyURL = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PositionURI']
# release date of job offer
porsche_releaseDate = porsche_data['SearchResult']['SearchResultItems'][i]['MatchedObjectDescriptor']['PublicationStartDate']
# append to 'data' DataFrame
df = pd.DataFrame(data={'Title':[porsche_title],'Company':[porsche_company],'Location':[porsche_location],'Business_Unit':[porsche_business_unit],'Description':[porsche_description],'Qualifications':[porsche_qualifications],'ApplyURL':[porsche_applyURL],'Release_Date':[porsche_releaseDate],'Valid':[1],'Interesting':[float('NaN')],'New':[1]})
data = data.append(df, ignore_index=True)
# -
# # Store scraped results in SQLite database
# +
# save 'data' DataFrame to persistent sqlite db
db_name = 'thesis_offers.db'
db_table_name = 'thesis_offers'
# if db does not exist, create new db
if not os.path.exists('./' + db_name):
conn = sqlite3.connect(db_name)
data.to_sql(name=db_table_name, con=conn, index=False)
# if db exists, write only new elements in db (which are not already in db) and check which job offer elements are currently valid
else:
conn = sqlite3.connect(db_name)
c = conn.cursor()
# set all Valids in db to 0
notValid = 0
Valid = 1
c.execute('UPDATE '+db_table_name+' SET Valid = ? WHERE Valid != ?', (notValid, notValid))
# set all New values in db to 0
c.execute('UPDATE '+db_table_name+' SET New = ? WHERE New != ?', (0, 0))
for i in range (data.shape[0]):
# use ApplyURL as PRIMARY KEY
t = (data['ApplyURL'][i],)
c.execute('SELECT * FROM thesis_offers WHERE ApplyURL=?', t)
# insert row of 'data' DataFrame if it is not in db
if len(c.fetchall()) == 0:
insert = [(data['ApplyURL'][i],data['Business_Unit'][i],data['Company'][i],data['Description'][i],data['Interesting'][i],data['Location'][i],data['New'][i],data['Qualifications'][i],data['Release_Date'][i],data['Title'][i],data['Valid'][i]),]
c.executemany('INSERT INTO '+db_table_name+' VALUES (?,?,?,?,?,?,?,?,?,?,?)', insert)
# set db Valid value to 1 if element is currently in 'data' DataFrame
else:
c.execute('UPDATE '+db_table_name+' SET Valid = ? WHERE ApplyURL=?', (Valid, data['ApplyURL'][i]))
# save db changes
conn.commit()
# close db connection
conn.close()
print('database updates finished')
| job_locator_web_scraper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning Implementation
# + [markdown] heading_collapsed=true
# ## Imports
# + hidden=true
import itertools
import json
import logging
import graphviz
import numpy as np
import pandas as pd
import plotly.offline as py
from graphviz import Digraph
from IPython.display import display
from plotly import graph_objects as go
from sklearn.datasets import load_boston, load_iris
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier, export_graphviz
# + [markdown] heading_collapsed=true
# ## Decision tree
# + [markdown] heading_collapsed=true hidden=true
# ### The maths
# + [markdown] hidden=true
# The decision tree is made by continuously splitting the data based on a certain feature and feature value. The feature and feature value used to split the data are chosen to increase the purity the most (or equivalently decrease the impurity the most).
# + [markdown] hidden=true
# Let the data at node $m$ be represented by $Q$.
#
# For a split $\theta = (j,t_m)$ consiting of feature $j$ and threshold value $t_m$ the imputiry $G$ of a split is given by
#
# $$
# G(Q,\theta) =
# \frac{n_{left}}{N_m}G(Q_{left}(\theta)) +
# \frac{n_{right}}{N_m}G(Q_{right}(\theta))
# $$
#
# Where the data $(x_i,y_i)$ is in $Q_{left}$ if $x_{i,j} <= t_m$ else $(x_i,y_i)$ is in $Q_{right}$
# + [markdown] hidden=true
# #### Classification
# + [markdown] hidden=true
# In the case of classification the Gini impurity is one the most common methods of measuring the impurity of the sample.
#
# If there are a set of classes $C$ often $C=\{0,1\}$ then for a given data set Q the impurity is defined as
#
# $$
# G(Q) = \sum_{c\in{C}} p_c(1-p_c)
# $$
# where $p_c$ is the probability of class $c$ in $Q$
# $$
# p_c = \frac{1}{N_Q}\sum_{x\in{Q}}\mathbb{1}(y_{class} = c)
# $$
# Where $N_Q = |Q|$
# + [markdown] hidden=true
# #### Regression
# + [markdown] hidden=true
# In regression, with a continuous target variable (y), the mean square error is often used as the impurity.
#
# $$
# G(Q) = \frac{1}{N_Q}\sum_{y_i\in Q}(y_i - \bar{y})^{2}
# $$
# where $\bar{y}$ is the mean value of $y$ in the node
# $$
# \bar{y} = \frac{1}{N_Q}\sum_{y_i\in Q}y_i
# $$
# + [markdown] heading_collapsed=true hidden=true
# ### Define the decision tree
# + hidden=true
logging.basicConfig()
logger = logging.getLogger('decision_tree')
logger.setLevel(logging.INFO)
logger.info(f'New logger with name {logger.name}')
# + hidden=true
class TreeNode():
count = itertools.count()
def __init__(self,
data,
max_depth,
min_samples_split,
min_samples_leaf,
n_classes=2,
max_features=None,
depth=0,
impurity='gini',
is_classifier=True):
"""
A single node in a decision tree
After recursive splitting of the input data, a given node
represents one split of the tree if it is not a leaf node. The
leaf node stores the training samples in that leaf to be used
for prediction.
The splitting nodes record the feature to split on as attribute
self.best_feature_index and the splitting value as attribute
self.best_feature_split_val
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features + 1 target)
Note the last column of the data are the target values
max_depth: int
The maximum depth allowed when "growing" a tree
min_samples_split: int
The minimum number of samples required to allow a split at
a the node
min_samples_leaf: int
The minimum number of samples allowed in a leaf. A split
candidate leading to less samples in a node than the
min_samples_leaf will be rejected
n_classes: int, optional, default 2
Number of classes in a classification setting. Ignored when
self.is_classifier = False
max_features: int, optional, default None
If set to 'sqrt' then only a random subset of features are
used to split at the node, the number of features used in
this case is sqrt(n_features).
Else all the features are considered when splitting at this
node
depth: int, optional, default 0
The depth of the node in the tree
impurity: str, optional, default 'gini'
The impurity measure to use when splitting at the node.
I have currently only implemented two
'gini' - Uses the gini impurity (for classification)
'mse' - Uses the mean square error - equal to variance (for
regression)
is_classifier: bool, optional, default True
Is the tree node used as part of a classification problem
or a regression problem. Should be set to True if
classification, False if regression
"""
self.data = data
self.max_depth = max_depth
self.min_samples_split = min_samples_split
self.min_samples_leaf = min_samples_leaf
self.n_classes = n_classes
self.max_features = max_features
self.depth = depth
self.impurity = impurity
self.is_classifier = is_classifier
self.data_shape = data.shape
self.split_attempted = False
self.best_split_impurity = None
self.best_feature_index = None
self.best_feature_split_val = None
self.is_leaf = False
self.node_impurity = self.calculate_impurity([data[:, -1]])
self.value = self._init_value(data)
self.id = str(next(self.count))
def __repr__(self):
return (
f'<TreeNode '
f'depth:{self.depth} '
f'node_impurity:{self.node_impurity:.2f} '
f'samples:{self.data_shape[0]} '
f'{"🌳" if self.is_root else ""}'
f'{"🍁" if self.is_leaf else ""}'
f'>')
@property
def is_root(self):
return self.depth == 0
def info(self):
return dict(
data_shape=self.data_shape,
n_classes=self.n_classes,
depth=self.depth,
min_samples_split=self.min_samples_split,
min_samples_leaf=self.min_samples_leaf,
node_impurity=self.node_impurity,
split_attempted=self.split_attempted,
best_split_impurity=self.best_split_impurity,
best_feature_index=self.best_feature_index,
best_feature_split_val=self.best_feature_split_val,
is_root=self.is_root)
def _init_value(self, data):
"""
Returns the terminal node value based on the input data
For a classifier this is the class_counts.
For a regressor this is the average y value.
Note this value can be access at a splitting node to see what
the prediction would have been at that level of the tree
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features + 1 target)
Note the last column of the data are the target values
Returns:
-------
numpy.ndarray or float:
Class counts if classifier, else mean of target values
"""
if self.is_classifier:
return np.bincount(
data[:, -1].astype(int),
minlength=self.n_classes)
else:
return np.mean(data[:, -1])
def split(self, feature_index, feature_split_val, only_y=True):
"""
Splits self.data on feature with index feature_index using
feature_split_val.
Each sample is included in left output if the feature value for
the sample is less than or equal to the feature_split_val else
it is included in the right output
Parameters:
----------
feature_index: int
Index of the feature (column) in self.data
feature_split_val: float
Feature value to use when splitting data
only_y: bool, optional, default True
Return only the y values in left and right - this is used
when checking candidate split purity increase
Returns:
-------
(numpy.ndarray, numpy.ndarray):
left and right splits of self.data
"""
assert feature_index in range(self.data.shape[1])
if only_y:
select = -1
else:
select = slice(None)
left_mask = self.data[:, feature_index] <= feature_split_val
right_mask = ~ left_mask
left = self.data[left_mask, select]
right = self.data[right_mask, select]
logger.debug(
f'Splitting on feature_index {feature_index} with '
f'feature_split_val = {feature_split_val} creates left '
f'with shape {left.shape} and right with '
f'shape {right.shape}')
return left, right
def gini_impurity(self, groups):
"""
Calculate the Gini impurity for groups of values
The impurity returned is the weighted average of the impurity
of the groups.
You can think of gini impurity as the probability of incorrectly
predicting a random sample from a group if the prediction was
made based purely on the distribution of class labels in the
group
Parameters:
----------
groups: tuple
The groups tuple is made up of arrays of values. It is
often called with groups = (left, right) to find the purity
of the candidate split
Returns:
-------
float:
Gini impurity
"""
gini = 0
total_samples = sum(group.shape[0] for group in groups)
for i, group in enumerate(groups):
group = group.astype(int)
class_counts = np.bincount(group, minlength=self.n_classes)
group_size = class_counts.sum()
class_probs = class_counts / group_size
unique_classes = np.count_nonzero(class_counts)
group_gini = (class_probs * (1 - class_probs)).sum()
gini += group_gini * (group_size / total_samples)
logger.debug(
f'Group {i} has size {group.shape[0]} with '
f'{unique_classes} unique classes '
f'with Gini index {group_gini:.3}')
return gini
def mean_square_impurity(self, groups):
"""
Calculates the mean square error impurity
The mse impurity is the weighted average of the group variances
Parameters:
----------
groups: tuple
The groups tuple is made up of arrays of values. It is
often called with groups = (left, right) to find the purity
of the candidate split
Returns:
-------
float:
Mean square error impurity
"""
mean_square_error = 0
total_samples = sum(group.shape[0] for group in groups)
for i, group in enumerate(groups):
group_size = group.shape[0]
group_mean = np.mean(group)
group_mean_square_error = np.mean((group - group_mean) ** 2)
mean_square_error += group_mean_square_error * \
(group_size / total_samples)
logger.debug(
f'Group {i} has size {group.shape[0]} with '
f'with MSE impurity {group_mean_square_error:.3}')
logger.debug(f'MSE candidate {mean_square_error}')
return mean_square_error
def calculate_impurity(self, groups):
"""
Calculates impurity based on self.impurity setting
Parameters:
----------
groups: tuple
The groups tuple is made up of arrays of values. It is
often called with groups = (left, right) to find the purity
of the candidate split
Returns:
-------
float:
Mean square error of groups if self.impurity = 'mse'
Gini impurity of groups if self.impurity = 'mse'
"""
if self.impurity == 'gini':
return self.gini_impurity(groups)
elif self.impurity == 'mse':
return self.mean_square_impurity(groups)
def check_split(self, feature_index, feature_split_val):
"""
Updates best split if candidate split is better
Splits the data in groups using self.split. Checks min samples
leaf condition after split. Calculates impurity of the split
then if impurity is less than best split already found and less
than the current node impurity the best_feature_index, the
best_feature_split_val and the best_split_impurity values are
updated.
Parameters:
----------
feature_index: int
Index of the feature (column) in self.data
feature_split_val: float
Feature value to use when splitting data
"""
groups = self.split(feature_index, feature_split_val)
if any(len(group) < self.min_samples_leaf for group in groups):
logger.debug(
f"Can't split node on feature {feature_index} with split "
f"val {feature_split_val} due to min_samples_leaf condition")
return None
split_impurity = self.calculate_impurity(groups)
best_current_impurity = (
10**10 if self.best_split_impurity is None
else self.best_split_impurity)
if ((split_impurity < best_current_impurity) and
(split_impurity < self.node_impurity)):
logger.debug(
f'Found new best split with feature_split_val='
f'{feature_split_val} for feature_index = {feature_index} '
f'and split_impurity = {split_impurity:.2f}')
self.best_feature_index = feature_index
self.best_feature_split_val = feature_split_val
self.best_split_impurity = split_impurity
def find_best_split(self):
"""
Finds best split at the node
Loops through each feature and each unique value of that feature
checking for the best candidate split (i.e. the split that
reduces the impurity the most)
The function first checks if we have reached the max depth or if
self.data < self.min_samples_split. In either case no further
split is allowed and the function returns
All features are considered unless self.max_features == 'sqrt'
in which case a random subset of features are used of size
sqrt(n_features)
"""
if self.depth == self.max_depth:
return
if self.data.shape[0] < self.min_samples_split:
logger.info(f"{self} can't split as samples < min_samples_split")
return None
if self.node_impurity == 0:
logger.info(f"Can't improve as node pure")
return None
n_features = self.data.shape[1] - 1
all_feature_indices = np.arange(n_features)
if self.max_features == 'sqrt':
features_to_check = np.random.choice(
all_feature_indices,
size=np.sqrt(n_features).astype(int))
else:
features_to_check = all_feature_indices
logger.info(f'Checking features {features_to_check}')
for feature_index in features_to_check:
for feature_split_val in np.unique(self.data[:, feature_index]):
self.check_split(feature_index, feature_split_val)
self.split_attempted = True
def recursive_split(self):
"""
Recursively grows tree by splitting to reduce impurity the most
The function finds the best split using the find_best_split
method. If there was a split found two nodes are created - left
and right. Finally the recursive_split method is called on each
of the new nodes.
Note the depth of the children node is incremented, otherwise
the node settings such as min_samples_split are passed to the
children nodes
"""
self.find_best_split()
if self.best_feature_index is not None:
logger.info(f'Splitting tree on feature_index '
f'{self.best_feature_index} and feature_split_val '
f'{self.best_feature_split_val:.2f}')
left, right = self.split(
feature_index=self.best_feature_index,
feature_split_val=self.best_feature_split_val,
only_y=False)
del self.data
self.left = TreeNode(
data=left,
max_depth=self.max_depth,
min_samples_split=self.min_samples_split,
min_samples_leaf=self.min_samples_leaf,
n_classes=self.n_classes,
max_features=self.max_features,
depth=self.depth + 1,
impurity=self.impurity,
is_classifier=self.is_classifier)
self.right = TreeNode(
data=right,
max_depth=self.max_depth,
min_samples_split=self.min_samples_split,
min_samples_leaf=self.min_samples_leaf,
n_classes=self.n_classes,
max_features=self.max_features,
depth=self.depth + 1,
impurity=self.impurity,
is_classifier=self.is_classifier)
self.left.recursive_split()
self.right.recursive_split()
else:
logger.info('Reached max depth or no splits reduce impurity')
self.is_leaf = True
def walk_depth_first(self, only_leaves=True):
"""
Generator traversing of all nodes below and including this node
Depth first so visiting children before siblings
Parameters:
----------
only_leaves: bool, optional, default True
Only return leaf nodes
Yields:
TreeNode: each node in tree
"""
if self.is_leaf:
yield self
else:
if not only_leaves:
yield self
for node in (self.left, self.right):
yield from node.walk_depth_first(only_leaves)
def walk_breadth_first(self, layer=None):
"""
Generator traversing of all nodes below and including this node
Breadth first so visiting siblings before children
Parameters:
----------
only_leaves: bool, optional, default True
Only return leaf nodes
Yields:
TreeNode: each node in tree
"""
if layer is None:
layer = [self]
for node in layer:
yield node
new_layer = [
child
for node_children in [[node.left, node.right]
for node in layer if not node.is_leaf]
for child in node_children]
if new_layer:
yield from self.walk_breadth_first(new_layer)
def print_tree(self):
"""
prints ascii representation of tree below this node
"""
for node in self.walk_depth_first(only_leaves=False):
print('--' * node.depth + str(node))
def predict_row_proba(self, row):
"""
Predicts class probabilities for input row by walking the tree
and returning the leaf node class probabilities
Parameters:
----------
row: numpy.ndarray
Input row, shape (n features,)
Returns:
-------
numpy.ndarray:
Class probabilities, shape (n classes, )
"""
if self.is_leaf:
group_size = self.value.sum()
class_probs = self.value / group_size
return class_probs
elif row[self.best_feature_index] <= self.best_feature_split_val:
return self.left.predict_row_proba(row)
else:
return self.right.predict_row_proba(row)
def predict_proba(self, data):
"""Predicts class probabilities for input data
Predicts class probabilities for each row in data by walking the
tree and returning the leaf node class probabilities
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features)
Returns:
-------
numpy.ndarray:
Predicted sample class probabilities,
shape (m samples, n classes)
"""
if not self.is_classifier:
raise Exception('Not a classifier')
if len(data.shape) == 2:
return np.stack([self.predict_row_proba(row)
for row in data])
else:
return self.predict_row_proba(data)
def predict_regressor_row(self, row):
"""
Predicts target value for input row by walking the tree
and returning the leaf node value
Parameters:
----------
row: numpy.ndarray
Input row, shape (n features,)
Returns:
-------
float:
Predicted target value
"""
if self.is_leaf:
return self.value
elif row[self.best_feature_index] <= self.best_feature_split_val:
return self.left.predict_regressor_row(row)
else:
return self.right.predict_regressor_row(row)
def predict_regressor(self, data):
"""
Predicts target values for each row in data by walking the
tree and returning the leaf node values
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features)
Returns:
-------
numpy.ndarray:
Predicted target values, shape (m samples, 1)
"""
if len(data.shape) == 2:
return np.stack([self.predict_regressor_row(row)
for row in data])
else:
return self.predict_regressor_row(data)
def predict(self, data):
"""Predicts target values or class labels for classification
Predicts target values/class for each row in data by walking the
tree and returning the leaf node value for regression or the
class with the largest predicted probability for classification
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features)
Returns:
-------
numpy.ndarray:
Predicted target values or class labels for classification
"""
if self.is_classifier:
return np.argmax(self.predict_proba(data), axis=-1)
else:
return self.predict_regressor(data)
def dot(self,
feature_names,
samples=True,
impurity=True,
value=True):
"""
Returns Digraph visualizing the tree below this node
Parameters:
----------
feature_names: list[str]
List of feature names
samples: bool, optional, default True
Whether to display the number of samples on this node
impurity: bool, optional, default True
Whether to display the impurity value on this node
value: bool, optional, default True
Whether to dispaly the value on this node
Returns:
-------
graphviz.Digraph:
dot for tree diagram visual
"""
dot = Digraph(
comment='Decsion Tree',
node_attr=dict(shape="rectangle",
style="rounded",
fillcolor="#028d35"))
for i, node in enumerate(self.walk_breadth_first()):
label = ""
if not node.is_leaf:
label += (
f'{feature_names[node.best_feature_index]} <= '
f'{node.best_feature_split_val}\n')
dot.edge(node.id, node.left.id)
dot.edge(node.id, node.right.id)
if samples:
label += f'Samples = {node.data_shape[0]}\n'
if impurity:
label += f'Impurity = {node.node_impurity:.2f}\n'
if value:
if self.is_classifier:
label += f'Class counts = {str(node.value)}\n'
else:
label += f'Average y = {node.value:.2f}\n'
dot.node(name=node.id, label=label)
return dot
# + hidden=true
class DecisionTree():
def __init__(self,
max_depth=2,
min_samples_split=2,
min_samples_leaf=1,
n_classes=2,
max_features=None,
impurity='gini',
is_classifier=True):
"""Decision tree model
Parameters:
----------
max_depth: int
The maximum depth allowed when "growing" a tree
min_samples_split: int
The minimum number of samples required to allow a split at
a the node
min_samples_leaf: int
The minimum number of samples allowed in a leaf. A split
candidate leading to less samples in a node than the
min_samples_leaf will be rejected
n_classes: int, optional, default 2
Number of classes in a classification setting. Ignored when
self.is_classifier = False
max_features: int, optional, default None
If set to 'sqrt' then only a random subset of features are
used to split at each node, the number of features used in
this case is sqrt(n_features).
Else all the features are considered when splitting at each
node
impurity: str, optional, default 'gini'
The impurity measure to use when splitting at each node.
I have currently only implemented two
'gini' - Uses the gini impurity (for classification)
'mse' - Uses the mean square error - equal to variance (for
regression)
is_classifier: bool, optional, default True
Is the model used as part of a classification problem
or a regression problem. Should be set to True if
classification, False if regression
"""
self.max_depth = max_depth
self.min_samples_split = min_samples_split
self.min_samples_leaf = min_samples_leaf
self.n_classes = n_classes
self.max_features = max_features
self.impurity = impurity
self.is_classifier = is_classifier
self.is_fitted = False
self.tree = None
def fit(self, X, y):
"""Fits the decision tree model
The tree is fitted by instantiaing a root TreeNode instance and
then calling the recursive_split method. This iteratively grows
the tree by finding the best split to reduce the impurity the
most.
Parameters:
----------
X: numpy.ndarray
Training data, shape (m samples, n features)
y: numpy.ndarray
Target values, shape (m samples, n features)
If classifier with n_classes the values are assumed to be in
0, ..., n-1
"""
y_shape = (X.shape[0], 1)
data = np.concatenate((X, y.reshape(y_shape)), axis=1)
self.tree = TreeNode(
data=data,
max_depth=self.max_depth,
min_samples_split=self.min_samples_split,
min_samples_leaf=self.min_samples_leaf,
n_classes=self.n_classes,
max_features=self.max_features,
impurity=self.impurity,
is_classifier=self.is_classifier)
self.tree.recursive_split()
self.is_fitted = True
def predict(self, data):
"""Predicts target values or class labels for classification
Predicts target values/class for each row in data by walking the
tree and returning the leaf node value for regression or the
class with the largest predicted probability for classification
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features)
Returns:
-------
numpy.ndarray:
Predicted target values or class labels for classification
"""
if not self.is_fitted:
raise Exception('Decision tree not fitted')
return self.tree.predict(data)
def predict_proba(self, data):
"""Predicts class probabilities for input data
Predicts class probabilities for each row in data by walking the
tree and returning the leaf node class probabilities
Parameters:
----------
data: numpy.ndarray
The input data with shape (m samples, n features)
Returns:
-------
numpy.ndarray:
Predicted sample class probabilities,
shape (m samples, n classes)
"""
if not self.is_fitted:
raise Exception('Decision tree not fitted')
return self.tree.predict_proba(data)
def render(self, feature_names):
"""Returns Digraph visualizing the decision tree (if fitted)
Parameters:
----------
feature_names: list[str]
List of feature names
Returns:
-------
graphviz.Digraph:
dot for tree diagram visual
"""
if not self.is_fitted:
print('Decision tree not fitted')
else:
return self.tree.dot(feature_names=feature_names)
# + [markdown] heading_collapsed=true
# ## Decision tree classifier - Iris data
# + [markdown] hidden=true
# ### Load iris data set
# + hidden=true
iris_data = load_iris()
iris_df = pd.DataFrame(iris_data['data'], columns=iris_data['feature_names'])
iris_df['y'] = iris_data['target']
iris_df = iris_df.sample(frac=1, random_state=42).reset_index(drop=True)
iris_sample = iris_df.head(5)
iris_sample
# + [markdown] hidden=true
# ### Fit tree decision tree classifier and visualise
# + hidden=true
# # for small sample
# iris_sample_vals = iris_sample.values
# X = iris_sample_vals[:,:-1]
# y = iris_sample_vals[:,-1]
X = iris_df.values[:,:-1]
y = iris_df.values[:,-1]
logger.setLevel(logging.INFO)
decision_tree = DecisionTree(n_classes=3, impurity='gini', is_classifier=True)
decision_tree.fit(X, y)
# For info or ansii tree
# decision_tree.tree.info()
# decision_tree.tree.print_tree()
feature_names = iris_data['feature_names']
decision_tree.render(feature_names=feature_names)
# + [markdown] hidden=true
# ### Example prediction on Iris data
# + hidden=true
decision_tree.predict(X[0,:])
# + hidden=true
decision_tree.predict(X)
# + hidden=true
# real y values
y.astype(int)
# + hidden=true
(decision_tree.predict(X) == y.astype(int)).sum() / len(y)
# + [markdown] heading_collapsed=true hidden=true
# ### Compare with Sklearn on Iris data
# + hidden=true
sk_decision_tree = DecisionTreeClassifier(
max_depth=decision_tree.max_depth,
min_samples_leaf=decision_tree.min_samples_leaf,
min_samples_split=decision_tree.min_samples_split)
sk_decision_tree.fit(X, y)
# Visualize the sklearn tree
# Note - same as ours except not using midpoints between split_vals
graphviz.Source(export_graphviz(sk_decision_tree))
# + [markdown] heading_collapsed=true
# ## Decision tree classifier - Titanic data
# + [markdown] heading_collapsed=true hidden=true
# ### Load titanic data
# + hidden=true
X_train = pd.read_feather('../data/titanic/processed/X_train.feather')
y_train = pd.read_feather('../data/titanic/processed/y_train.feather')
X_test = pd.read_feather('../data/titanic/processed/X_test.feather')
y_test = pd.read_feather('../data/titanic/processed/y_test.feather')
# + [markdown] heading_collapsed=true hidden=true
# ### Decision tree accuracy
# + hidden=true
# for best result use max depth = 4
titanic_decision_tree = DecisionTree(max_depth=2)
titanic_decision_tree.fit(X_train.values, y_train.values)
y_pred = titanic_decision_tree.predict(X_test.values)
test_acc = (y_pred == y_test.values.flatten()).sum() / len(y_test)
print(f'Test accuracy = {test_acc:.2%}')
# + [markdown] heading_collapsed=true hidden=true
# ### Visulaise titanic decision tree
# + hidden=true
titanic_features = list(X_train.columns)
titanic_decision_tree.render(titanic_features)
# + [markdown] heading_collapsed=true
# ## Decision tree regressor - Boston housing data
# + [markdown] heading_collapsed=true hidden=true
# ### Load boston data
# + hidden=true
boston_data = load_boston()
boston_df = pd.DataFrame(boston_data['data'], columns=boston_data['feature_names'])
boston_df['y'] = boston_data['target']
boston_df = boston_df.sample(frac=1, random_state=42).reset_index(drop=True)
boston_sample = boston_df.head(5)
boston_sample
# + [markdown] heading_collapsed=true hidden=true
# ### Fit decision tree on Boston data
# + hidden=true
logger.setLevel(logging.INFO)
X = boston_df.values[:,:-1]
y = boston_df.values[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
boston_decision_tree = DecisionTree(
impurity='mse',
is_classifier=False,
max_depth=2)
boston_decision_tree.fit(X_train, y_train)
# For info or ansii tree
# boston_decision_tree.tree.info()
# boston_decision_tree.tree.print_tree()
boston_feature_names = boston_data['feature_names']
boston_decision_tree.render(feature_names=boston_feature_names)
# + [markdown] heading_collapsed=true hidden=true
# ### Decision tree accuracy on Boston data
# + hidden=true
y_pred = boston_decision_tree.predict(X_test)
test_acc = r2_score(y_test, y_pred)
print(f'Test accuracy (R2 score) = {test_acc:.2%}')
# + [markdown] heading_collapsed=true
# ## end
| notebooks/decision_tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="VgPL0EJ69itt" executionInfo={"status": "ok", "timestamp": 1629147890965, "user_tz": 240, "elapsed": 7382, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="8af5ee27-af58-4911-dd48-5e8651186377"
# !pip install transformers
# + colab={"base_uri": "https://localhost:8080/"} id="1XxmkUAc8DAZ" executionInfo={"status": "ok", "timestamp": 1629147908603, "user_tz": 240, "elapsed": 17642, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="f87fbc86-dab5-4e30-ed84-81b97690c1b6"
from google.colab import drive
drive.mount('/content/drive')
# + id="TIOdl7669iT3"
from torch.utils.data import Dataset
import yaml
import torch
import pandas as pd
import torch
from torch.utils.data import random_split
from transformers import TrainingArguments, Trainer, GPTNeoForCausalLM
from transformers import GPT2TokenizerFast as GPT2Tokenizer
import yaml
import random
import numpy as np
from sklearn.model_selection import train_test_split
from tqdm import tqdm
import re
from os import listdir
# + id="hXIAT6c49obC" colab={"base_uri": "https://localhost:8080/", "height": 243, "referenced_widgets": ["6fc4033b49c6459d8eeacff377161c7d", "<KEY>", "e89f0c8f1e1d42bc863dfd2dfb739073", "9deedd74feaf4ed7bf6a7d786f226a35", "17b8e14e64b94a5cb7d8a7c376add37d", "c810c5e1dfd34fc082f7918d7722fd88", "0621d250263f408281d8762638a85f30", "4a02d54e2fb44e5c8b3d4d2fd45486e6", "10860fdccca5434392820d0d49d33258", "bfcef6145cf4495dbda0f136b0e9312f", "c779e09fd31648ca850b28696aaa913c", "7e52dcf2713740b59c7fc227c9bbdcbf", "682da4d36a9b4005843db359b733773a", "<KEY>", "<KEY>", "<KEY>", "0ce771a242e8404e83ef663626810c5c", "<KEY>", "ce9dce41f4024dc5aba1758402a31df5", "1ac753611acf4d4a924a5244671259d6", "aa83f10fe66246c9959c32ca09ae3844", "b5357a67e5ba425aa42235fff0ed46aa", "<KEY>", "<KEY>", "6c217a80584c4ecabef3565079ff2d7f", "6403e1968bd7475f85a47fa46d2d75dd", "517b0a854eb34af68189e8640510884d", "<KEY>", "54c840e0ea164908a9fb5fe5d14518f6", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "2de5b67e89b640abaf40a7021487bb7d", "<KEY>", "9e9f0f3c9eb14d0bba394be40f9d9add", "<KEY>", "80f88d4e905a4e78989c7d27c2ea00da", "<KEY>", "f7575eedbbd74d119773b688c49ea60c", "6694d333cc3d4a9b9e2dd890157a3005", "<KEY>", "7d7195039f764443a3df3e448bedfc84", "<KEY>", "a0abced697794606984aaa3ffdc3e510", "3894d76161124ca0b232698e38ad1a95", "d5e72a5bff7441daa46d6977a900cc1b", "7e6a3f1ee1864da0b05f07544434f2ac", "<KEY>", "5ecd3edcaf6048b58e67236d1e152702", "<KEY>", "bb0ffa735be14ce59e22b2b61605b991", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "f7e70f04d3a642ed9ac5273b698462b8", "87542461a408480c938d3442a3a3d66e", "<KEY>", "1cc6014837894b0ba47b2e596a51c658", "b7ca394f0d96422da75cda4bebb2d279", "c6964a17a29d4fc5b1e7d3ab32ca9709", "179407e88ea6440d80fecdf7e97412f7"]} executionInfo={"status": "ok", "timestamp": 1629147930454, "user_tz": 240, "elapsed": 17470, "user": {"displayName": "<NAME>00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="ab8dbe07-0440-4e7d-b5ac-860624d35318"
with open('/content/drive/MyDrive/data/config.yml', 'r') as f:
config = yaml.safe_load(f)
torch.cuda.empty_cache()
tokenizer = GPT2Tokenizer.from_pretrained(config['model']['model_name_internacional'], eos_token='<|endoftext|>', pad_token='<|pad|>')
model = GPTNeoForCausalLM.from_pretrained(config['model']['model_name_internacional'])
model.resize_token_embeddings(len(tokenizer))
"""Limpando Cache e Designando Variáveis Aleatórias"""
torch.cuda.empty_cache()
SEED = 42
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed_all(SEED)
# + id="MjTWRiSj_3Jm" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629147930457, "user_tz": 240, "elapsed": 11, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="1b750260-4cbc-4386-a31c-d51244e00770"
if torch.cuda.is_available():
device = torch.device('cuda')
else:
device = torch.device('cpu')
print(device)
# + id="A99j_26C815t" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629147930458, "user_tz": 240, "elapsed": 9, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="e23b9c24-7b10-4629-b4c4-cae09b528286"
# !nvidia-smi
# + id="RQ49K1JEmc2H"
path = config['data']['path_internacional']
# + colab={"base_uri": "https://localhost:8080/"} id="6qry0A3XnXiX" executionInfo={"status": "ok", "timestamp": 1629147930917, "user_tz": 240, "elapsed": 464, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="0bd0581c-01e4-44a5-e2b5-f48aebd9a25e"
for file in listdir(path):
print(file)
# + id="Kel_kpIxmlKY"
df = pd.DataFrame()
for file in listdir(path):
temp = pd.read_json(path + r'/' + file)
df = df.append(temp, ignore_index=True)
# + id="6uUFW3QKDurY"
def removendo_frases_repetidas(texto):
seen = set()
return [x for x in texto if x not in seen and not seen.add(x)]
# + id="Hxc-xbv5FYtp"
def removendo_sentencas_em_paratenses(x):
return [re.sub(r'\([^)]*\)', '', y) for y in x]
# + id="1k7Uwm-lGJOL"
def removendo_sentencas_em_cerquilhas(x):
return [re.sub(r'\{[^)]*\}', '', y) for y in x]
# + id="UzptAKrAnhP1"
def removendo_sentencas_em_colchetes(x):
return [re.sub(r'\[[^)]*\]', '', y) for y in x]
# + id="3egBO8GCG907"
def removendo_linha_vazia(x):
return list(filter(lambda x: x != '', x))
# + id="vpvViovJlCTU"
def removendo_refrao(x):
for y in x:
return re.sub('[Cc]horus', '', y)
# + id="N5QIgfTXnq7b"
def removendo_musica_sem_letra(x):
if len(x) <= 3:
return float('NaN')
else:
return x
# + id="ivo2G0A4D3nC"
df['letra'] = df['letra'].apply(lambda x: removendo_frases_repetidas(x))
df['letra'] = df['letra'].apply(lambda x: removendo_sentencas_em_paratenses(x))
df['letra'] = df['letra'].apply(lambda x: removendo_sentencas_em_colchetes(x))
df['letra'] = df['letra'].apply(lambda x: removendo_sentencas_em_cerquilhas(x))
df['letra'] = df['letra'].apply(lambda x: removendo_linha_vazia(x))
df['letra'] = df['letra'].apply(lambda x: removendo_musica_sem_letra(x))
df.dropna(inplace=True)
df.reset_index(drop=True, inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="w6CGtEtLruAS" executionInfo={"status": "ok", "timestamp": 1629147936937, "user_tz": 240, "elapsed": 16, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="44a05a24-1da7-47a0-bd3e-24faefb43be7"
len(df)
# + id="n7OIXz0C-Mb3"
#tornando letras em string
string = ''
for i, x in enumerate(df['letra']):
for y in x:
string += y + ' ' + '\n' + ' '
x = string
df['letra'][i] = x
string = ''
# + id="Joj1o01WHR3m"
train, test = train_test_split(df['letra'], test_size=0.15, shuffle=True, random_state=SEED)
# + id="hkP7MuCCMmzk"
class ShapingDataset(Dataset):
def __init__(self, texts):
self.texts = texts
self.tokenizer = tokenizer
self.labels = []
self.input_ids = []
self.attention_masks = []
self.max_length = max([len(re.findall('\w+', x)) for x in df['letra']]) #letra com maior número de palavras
for txt in texts:
dici = self.tokenizer(txt, padding='max_length', truncation=True, max_length=self.max_length, return_tensors='pt')
self.input_ids.append(dici['input_ids'][0])
self.attention_masks.append(dici['attention_mask'][0])
def __len__(self):
return len(self.texts)
def __getitem__(self, idx):
return self.input_ids[idx], self.attention_masks[idx]
# + id="M6LE9R81-Ku7"
train = ShapingDataset(train)
test = ShapingDataset(test)
# + id="qUSHaOsfR0-m"
training_args = TrainingArguments(output_dir=config['data']['output_dir'],
logging_dir=config['data']['logging_dir_internacional'],
num_train_epochs=config['model']['num_epochs'],
per_device_train_batch_size=config['model']['batch_size'],
per_device_eval_batch_size=config['model']['batch_size'],
logging_steps=config['data']['steps'],
save_steps = config['data']['steps'],
learning_rate=float(config['model']['learning_rate']),
warmup_steps=config['model']['num_epochs'])
# + id="KvBkUXDzlZji"
model.to(device)
trainer = Trainer(model=model, args=training_args, train_dataset=train, eval_dataset=test,
data_collator=lambda data:
{'input_ids': torch.stack([f[0] for f in data]),
'attention_mask': torch.stack([f[1] for f in data]),
'labels': torch.stack([f[0] for f in data])})
# + id="lY1nGuz-kp6G" colab={"base_uri": "https://localhost:8080/", "height": 972} executionInfo={"status": "ok", "timestamp": 1629159976337, "user_tz": 240, "elapsed": 12030902, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="b027dfc0-11c2-4309-ba0e-a6f9543104d3"
trainer.train()
# + id="M8rD9yqBuRzh"
saved_root = config['data']['saved_root_internacional']
# + id="Xd37efm2LUy5" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629159978377, "user_tz": 240, "elapsed": 2046, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="a5af6763-a60a-4b47-e6a8-86f9562a3bc0"
trainer.save_model(saved_root)
# + id="7H_Zatyksi3S" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629159978377, "user_tz": 240, "elapsed": 33, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="6c393f80-2fc2-427d-9fcc-91122d344ba1"
50000
# + id="rpfrX1tlPYQp" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1629159980310, "user_tz": 240, "elapsed": 1963, "user": {"displayName": "<NAME>\u00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="c90364b6-c22f-4759-81c6-90253f901f9a"
model = GPTNeoForCausalLM.from_pretrained(saved_root)
# + id="0njq13pJPc7b"
model.to('cuda')
generated = tokenizer('I miss you so much',return_tensors='pt').input_ids.cuda()
# + colab={"base_uri": "https://localhost:8080/"} id="ssgH28_BPlt5" executionInfo={"status": "ok", "timestamp": 1629159983661, "user_tz": 240, "elapsed": 3356, "user": {"displayName": "<NAME>00e3o", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj1j4nQ7Q_ayV5RNKMuyoCOHEsxIXlBdaTp36qA5g=s64", "userId": "15460989607803425555"}} outputId="c9143c2a-fbb2-4bb5-ac5a-9f91d1e4c3b9"
#Generating texts
sample_outputs = model.generate(generated,
# Use sampling instead of greedy decoding
do_sample=True,
# Keep only top 3 token with the highest probability
top_k=10,
# Maximum sequence length
max_length=200,
# Keep only the most probable tokens with cumulative probability of 95%
top_p=0.95,
# Changes randomness of generated sequences
temperature=2.,
# Number of sequences to generate
num_return_sequences=3)
# Decoding and printing sequences
for i, sample_output in enumerate(sample_outputs):
texto = tokenizer.decode(sample_output.tolist())
regex_padding = re.sub('<|pad|>', '', texto)
regex_barra = re.sub('[|+]', '', regex_padding)
espaço = re.sub('[ +]', ' ', regex_barra)
resultado = re.sub('[\n](2, )', '\n', espaço)
print(">> Texto {}: {}".format(i+1, resultado))
# + id="E-Ir3Kb1ufho"
| Finetuning Emo Bot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Organization: Matrix Structure
#
# >**Reference**: <NAME>, <NAME>, <NAME>, 2016. [*Temporal regularized matrix factorization for high-dimensional time series prediction*](http://www.cs.utexas.edu/~rofuyu/papers/tr-mf-nips.pdf). 30th Conference on Neural Information Processing Systems (*NIPS 2016*), Barcelona, Spain.
#
# We consider a dataset of $m$ discrete time series $\boldsymbol{y}_{i}\in\mathbb{R}^{f},i\in\left\{1,2,...,m\right\}$. The time series may have missing elements. We express spatio-temporal dataset as a matrix $Y\in\mathbb{R}^{m\times f}$ with $m$ rows (e.g., locations) and $f$ columns (e.g., discrete time intervals),
#
# $$Y=\left[ \begin{array}{cccc} y_{11} & y_{12} & \cdots & y_{1f} \\ y_{21} & y_{22} & \cdots & y_{2f} \\ \vdots & \vdots & \ddots & \vdots \\ y_{m1} & y_{m2} & \cdots & y_{mf} \\ \end{array} \right]\in\mathbb{R}^{m\times f}.$$
#
# # Temporal Regularized Matrix Factorization(TRMF)
# Temporal Regularized Matrix Factorization (TRMF) framework is an approach to incorporate temporal dependencies into matrix factorization models which use well-studied time series models to describe temporal dependencies
# among ${\boldsymbol{x}_t}$ explicitly.Such models take the form:
#
# $$\boldsymbol{x}_{t}\approx\sum_{l\in\mathcal{L}}\boldsymbol{\theta}_{l}\circledast\boldsymbol{x}_{t-l}$$
#
# where this autoregressive (AR) is specialized by a lag set $\mathcal{L}=\left\{l_1,l_2,...,l_d\right\}$ (e.g., $\mathcal{L}=\left\{1,2,144\right\}$) and weights $\boldsymbol{\theta}_{l}\in\mathbb{R}^{r},\forall l$, and we further define
#
# $$\mathcal{R}_{AR}\left(X\mid \mathcal{L},\Theta,\eta\right)=\frac{1}{2}\sum_{t=l_d+1}^{f}\left(\boldsymbol{x}_{t}-\sum_{l\in\mathcal{L}}\boldsymbol{\theta}_{l}\circledast\boldsymbol{x}_{t-l}\right)^T\left(\boldsymbol{x}_{t}-\sum_{l\in\mathcal{L}}\boldsymbol{\theta}_{l}\circledast\boldsymbol{x}_{t-l}\right)+\frac{\eta}{2}\sum_{t=1}^{f}\boldsymbol{x}_{t}^T\boldsymbol{x}_{t}.$$
#
# Thus, TRMF-AR is given by solving
#
# $$\min_{W,X,\Theta}\frac{1}{2}\underbrace{\sum_{(i,t)\in\Omega}\left(y_{it}-\boldsymbol{w}_{i}^T\boldsymbol{x}_{t}\right)^2}_{\text{sum of squared residual errors}}+\lambda_{w}\underbrace{\mathcal{R}_{w}\left(W\right)}_{W-\text{regularizer}}+\lambda_{x}\underbrace{\mathcal{R}_{AR}\left(X\mid \mathcal{L},\Theta,\eta\right)}_{\text{AR-regularizer}}+\lambda_{\theta}\underbrace{\mathcal{R}_{\theta}\left(\Theta\right)}_{\Theta-\text{regularizer}}$$
#
# where $\mathcal{R}_{w}\left(W\right)=\frac{1}{2}\sum_{i=1}^{m}\boldsymbol{w}_{i}^T\boldsymbol{w}_{i}$ and $\mathcal{R}_{\theta}\left(\Theta\right)=\frac{1}{2}\sum_{l\in\mathcal{L}}\boldsymbol{\theta}_{l}^T\boldsymbol{\theta}_{l}$ are regularization terms.
import numpy as np
from numpy.linalg import inv as inv
# # Matrix Computation Concepts
#
# ## Kronecker product
#
# - **Definition**:
#
# Given two matrices $A\in\mathbb{R}^{m_1\times n_1}$ and $B\in\mathbb{R}^{m_2\times n_2}$, then, the **Kronecker product** between these two matrices is defined as
#
# $$A\otimes B=\left[ \begin{array}{cccc} a_{11}B & a_{12}B & \cdots & a_{1m_2}B \\ a_{21}B & a_{22}B & \cdots & a_{2m_2}B \\ \vdots & \vdots & \ddots & \vdots \\ a_{m_11}B & a_{m_12}B & \cdots & a_{m_1m_2}B \\ \end{array} \right]$$
# where the symbol $\otimes$ denotes Kronecker product, and the size of resulted $A\otimes B$ is $(m_1m_2)\times (n_1n_2)$ (i.e., $m_1\times m_2$ columns and $n_1\times n_2$ rows).
#
# - **Example**:
#
# If $A=\left[ \begin{array}{cc} 1 & 2 \\ 3 & 4 \\ \end{array} \right]$ and $B=\left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10 \\ \end{array} \right]$, then, we have
#
# $$A\otimes B=\left[ \begin{array}{cc} 1\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] & 2\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] \\ 3\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] & 4\times \left[ \begin{array}{ccc} 5 & 6 & 7\\ 8 & 9 & 10\\ \end{array} \right] \\ \end{array} \right]$$
#
# $$=\left[ \begin{array}{cccccc} 5 & 6 & 7 & 10 & 12 & 14 \\ 8 & 9 & 10 & 16 & 18 & 20 \\ 15 & 18 & 21 & 20 & 24 & 28 \\ 24 & 27 & 30 & 32 & 36 & 40 \\ \end{array} \right]\in\mathbb{R}^{4\times 6}.$$
#
# ## Khatri-Rao product (`kr_prod`)
#
# - **Definition**:
#
# Given two matrices $A=\left( \boldsymbol{a}_1,\boldsymbol{a}_2,...,\boldsymbol{a}_r \right)\in\mathbb{R}^{m\times r}$ and $B=\left( \boldsymbol{b}_1,\boldsymbol{b}_2,...,\boldsymbol{b}_r \right)\in\mathbb{R}^{n\times r}$ with same number of columns, then, the **Khatri-Rao product** (or **column-wise Kronecker product**) between $A$ and $B$ is given as follows,
#
# $$A\odot B=\left( \boldsymbol{a}_1\otimes \boldsymbol{b}_1,\boldsymbol{a}_2\otimes \boldsymbol{b}_2,...,\boldsymbol{a}_r\otimes \boldsymbol{b}_r \right)\in\mathbb{R}^{(mn)\times r}$$
# where the symbol $\odot$ denotes Khatri-Rao product, and $\otimes$ denotes Kronecker product.
#
# - **Example**:
#
# If $A=\left[ \begin{array}{cc} 1 & 2 \\ 3 & 4 \\ \end{array} \right]=\left( \boldsymbol{a}_1,\boldsymbol{a}_2 \right) $ and $B=\left[ \begin{array}{cc} 5 & 6 \\ 7 & 8 \\ 9 & 10 \\ \end{array} \right]=\left( \boldsymbol{b}_1,\boldsymbol{b}_2 \right) $, then, we have
#
# $$A\odot B=\left( \boldsymbol{a}_1\otimes \boldsymbol{b}_1,\boldsymbol{a}_2\otimes \boldsymbol{b}_2 \right) $$
#
# $$=\left[ \begin{array}{cc} \left[ \begin{array}{c} 1 \\ 3 \\ \end{array} \right]\otimes \left[ \begin{array}{c} 5 \\ 7 \\ 9 \\ \end{array} \right] & \left[ \begin{array}{c} 2 \\ 4 \\ \end{array} \right]\otimes \left[ \begin{array}{c} 6 \\ 8 \\ 10 \\ \end{array} \right] \\ \end{array} \right]$$
#
# $$=\left[ \begin{array}{cc} 5 & 12 \\ 7 & 16 \\ 9 & 20 \\ 15 & 24 \\ 21 & 32 \\ 27 & 40 \\ \end{array} \right]\in\mathbb{R}^{6\times 2}.$$
def kr_prod(a, b):
return np.einsum('ir, jr -> ijr', a, b).reshape(a.shape[0] * b.shape[0], -1)
import numpy as np
A = np.array([[1, 2], [3, 4]])
B = np.array([[5, 6], [7, 8], [9, 10]])
print(kr_prod(A, B))
def TRMF(dense_mat, sparse_mat, init, time_lags, lambda_w, lambda_x, lambda_theta, eta, maxiter):
W = init["W"]
X = init["X"]
theta = init["theta"]
dim1, dim2 = sparse_mat.shape
binary_mat = np.zeros((dim1,dim2))
position = np.where((sparse_mat != 0))
binary_mat[position] = 1
pos = np.where((dense_mat != 0) & (sparse_mat == 0))
d = len(time_lags)
r = theta.shape[1]
mape = np.zeros(maxiter)
rmse = np.zeros(maxiter)
for iter in range(maxiter):
var1 = X.T
var2 = kr_prod(var1,var1)
var3 = np.matmul(var2,binary_mat.T)
var4 = np.matmul(var1,sparse_mat.T)
for i in range(dim1):
W[i,:] = np.matmul(inv((var3[:,i].reshape([r,r]))+lambda_w * np.eye(r)), var4[:,i])
var1 = W.T
var2 = kr_prod(var1,var1)
var3 = np.matmul(var2, binary_mat)
var4 = np.matmul(var1, sparse_mat)
for t in range(dim2):
Mt = np.zeros((r,r))
Nt = np.zeros(r)
if t < max(time_lags):
Pt = np.zeros((r,r))
Qt = np.zeros(r)
else:
Pt = np.eye(r)
Qt = np.einsum('ij, ij -> j', theta, X[t - time_lags, :])
if t < dim2 - np.min(time_lags):
if t >= np.max(time_lags) and t < dim2 - np.max(time_lags):
index = list(range(0, d))
else:
index = list(np.where((t + time_lags >= np.max(time_lags)) & (t + time_lags < dim2)))[0]
for k in index:
theta0 = theta.copy()
theta0[k, :] = 0
Mt = Mt + np.diag(theta[k, :]**2);
Nt = Nt + np.multiply(theta[k,:],(X[t+time_lags[k], :]
- np.einsum('ij, ij -> j', theta0,
X[t + time_lags[k] - time_lags, :])))
X[t,:] = np.matmul(inv(var3[:, t].reshape([r,r])
+ lambda_x * Pt + lambda_x * Mt + lambda_x * eta * np.eye(r)),
(var4[:, t] + lambda_x * Qt + lambda_x * Nt))
elif t >= dim2 - np.min(time_lags):
X[t, :] = np.matmul(inv(var3[:, t].reshape([r, r]) + lambda_x * Pt
+ lambda_x * eta * np.eye(r)), (var4[:, t] + Qt))
for k in range(d):
var1 = X[np.max(time_lags) - time_lags[k] : dim2 - time_lags[k], :]
var2 = inv(np.diag(np.einsum('ij, ij -> j', var1, var1)) + (lambda_theta / lambda_x) * np.eye(r))
var3 = np.zeros(r)
for t in range(np.max(time_lags) - time_lags[k], dim2 - time_lags[k]):
var3 = var3 + np.multiply(X[t, :],
(X[t + time_lags[k], :]
- np.einsum('ij, ij -> j', theta, X[t + time_lags[k] - time_lags, :])
+np.multiply(theta[k, :], X[t,:])))
theta[k, :] = np.matmul(var2,var3)
mat_hat = np.matmul(W, X.T)
mape[iter] = np.sum(np.abs(dense_mat[pos] - mat_hat[pos]) / dense_mat[pos]) / dense_mat[pos].shape[0]
rmse[iter] = np.sqrt(np.sum((dense_mat[pos] - mat_hat[pos])**2)/dense_mat[pos].shape[0])
return W, X, theta
def OnlineTRMF(sparse_vec, init, lambda_x, time_lags):
W = init["W"]
X = init["X"]
theta = init["theta"]
dim = sparse_vec.shape[0]
t, rank = X.shape
position = np.where(sparse_vec != 0)
binary_vec = np.zeros(dim)
binary_vec[position] = 1
xt_tilde = np.einsum('ij, ij -> j', theta, X[t - 1 - time_lags, :])
var1 = W.T
var2 = kr_prod(var1, var1)
var_mu = np.matmul(var1, sparse_vec) + lambda_x * xt_tilde
inv_var_Lambda = inv(np.matmul(var2, binary_vec).reshape([rank, rank]) + lambda_x * np.eye(rank))
X[t - 1, :] = np.matmul(inv_var_Lambda, var_mu)
mat_hat = np.matmul(W, X.T)
return X
def st_prediction(dense_mat, sparse_mat, time_lags, lambda_w, lambda_x, lambda_theta, eta,
rank, pred_time_steps, maxiter):
start_time = dense_mat.shape[1] - pred_time_steps
dense_mat0 = dense_mat[:, 0 : start_time]
sparse_mat0 = sparse_mat[:, 0 : start_time]
dim1 = sparse_mat0.shape[0]
dim2 = sparse_mat0.shape[1]
mat_hat = np.zeros((dim1, pred_time_steps))
for t in range(pred_time_steps):
if t == 0:
init = {"W": 0.1 * np.random.rand(dim1, rank), "X": 0.1 * np.random.rand(dim2, rank),
"theta": 0.1 * np.random.rand(time_lags.shape[0], rank)}
W, X, theta = TRMF(dense_mat0, sparse_mat0, init, time_lags,
lambda_w, lambda_x, lambda_theta, eta, maxiter)
X0 = np.zeros((dim2 + t + 1, rank))
X0[0 : dim2 + t, :] = X.copy()
X0[dim2 + t, :] = np.einsum('ij, ij -> j', theta, X0[dim2 + t - time_lags, :])
else:
sparse_vec = sparse_mat[:, start_time + t - 1]
if np.where(sparse_vec > 0)[0].shape[0] > rank:
init = {"W": W, "X": X0[- np.max(time_lags) - 1 :, :], "theta": theta}
X = OnlineTRMF(sparse_vec, init, lambda_x/dim2, time_lags)
X0 = np.zeros((np.max(time_lags) + 1, rank))
X0[0 : np.max(time_lags), :] = X[1 :, :].copy()
X0[np.max(time_lags), :] = np.einsum('ij, ij -> j', theta, X0[np.max(time_lags) - time_lags, :])
else:
X0 = np.zeros((np.max(time_lags) + 1, rank))
X0[0 : np.max(time_lags), :] = X[1 :, :]
X0[np.max(time_lags), :] = np.einsum('ij, ij -> j', theta, X0[np.max(time_lags) - time_lags, :])
mat_hat[:, t] = np.matmul(W, X0[-1, :])
if (t + 1) % 40 == 0:
print('Time step: {}'.format(t + 1))
small_dense_mat = dense_mat[:, start_time : dense_mat.shape[1]]
pos = np.where(small_dense_mat != 0)
final_mape = np.sum(np.abs(small_dense_mat[pos] -
mat_hat[pos])/small_dense_mat[pos])/small_dense_mat[pos].shape[0]
final_rmse = np.sqrt(np.sum((small_dense_mat[pos] -
mat_hat[pos]) ** 2)/small_dense_mat[pos].shape[0])
print('Final MAPE: {:.6}'.format(final_mape))
print('Final RMSE: {:.6}'.format(final_rmse))
print()
return mat_hat
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.0
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 144 * 5
time_lags = np.array([1, 2, 144])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 144 * 5
time_lags = np.array([1, 2, 144])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 144 * 5
time_lags = np.array([1, 2, 144])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 144 * 5
time_lags = np.array([1, 2, 144])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Guangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 144 * 5
time_lags = np.array([1, 2, 144])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# -
# **Experiment results** of spatial-temporal data prediction using TRMF:
#
# | scenario |`rank`|`Lambda_w`|`Lambda_x`|`Lambda_theta`|`eta`|`maxiter`| mape | rmse |
# |:----------|-----:|---------:|---------:|-------------:|----:|----------:|-----:|-----:|
# |**Original data**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.1065**| **4.30**|
# |**20%, RM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.1062**| **4.31**|
# |**40%, RM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.1062**| **4.30**|
# |**20%, NM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.1064**| **4.29**|
# |**40%, NM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.1071**| **4.32**|
#
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.0
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 18 * 7
time_lags = np.array([1, 2, 18])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 100
lambda_x = 100
lambda_theta = 100
eta = 0.01
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.1
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 18 * 7
time_lags = np.array([1, 2, 18])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 100
lambda_x = 100
lambda_theta = 100
eta = 0.01
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.3
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 18 * 7
time_lags = np.array([1, 2, 18])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 100
lambda_x = 100
lambda_theta = 100
eta = 0.01
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.1
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 18 * 7
time_lags = np.array([1, 2, 18])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 100
lambda_x = 100
lambda_theta = 100
eta = 0.01
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Birmingham-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Birmingham-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.3
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 18 * 7
time_lags = np.array([1, 2, 18])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 100
lambda_x = 100
lambda_theta = 100
eta = 0.01
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# -
# **Experiment results** of spatial-temporal data prediction using TRMF:
#
# | scenario |`rank`|`Lambda_w`|`Lambda_x`|`Lambda_theta`|`eta`|`back step`| mape | rmse |
# |:----------|-----:|---------:|---------:|-------------:|----:|----------:|-----:|-----:|
# |**Original data**| 10 | 100 | 100 | 100 | 0.01 | 200 | **0.3263**| **174.25**|
# |**20%, RM**| 10 | 100 | 100 | 100 | 0.01 | 200 | **0.3267**| **171.69**|
# |**40%, RM**| 10 | 100 | 100 | 100 | 0.01 | 200 | **0.3442**| **181.17**|
# |**20%, NM**| 10 | 100 | 100 | 100 | 0.01 | 200 | **0.3195**| **169.30**|
# |**40%, NM**| 10 | 100 | 100 | 100 | 0.01 | 200 | **0.3309**| **175.64**|
#
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.0
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 108 * 5
time_lags = np.array([1, 2, 108])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 1000
lambda_x = 1000
lambda_theta = 1000
eta = 0.05
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 108 * 5
time_lags = np.array([1, 2, 108])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 1000
lambda_x = 1000
lambda_theta = 1000
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
random_tensor.shape[1]
* random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
# binary_tensor = np.zeros(tensor.shape)
# for i1 in range(tensor.shape[0]):
# for i2 in range(tensor.shape[1]):
# binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
# binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
# * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 108 * 5
time_lags = np.array([1, 2, 108])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 1000
lambda_x = 1000
lambda_theta = 1000
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 108 * 5
time_lags = np.array([1, 2, 108])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 1000
lambda_x = 1000
lambda_theta = 1000
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import scipy.io
tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/tensor.mat')
tensor = tensor['tensor']
random_matrix = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_matrix.mat')
random_matrix = random_matrix['random_matrix']
random_tensor = scipy.io.loadmat('../datasets/Hangzhou-data-set/random_tensor.mat')
random_tensor = random_tensor['random_tensor']
dense_mat = tensor.reshape([tensor.shape[0], tensor.shape[1] * tensor.shape[2]])
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
# binary_mat = np.round(random_tensor + 0.5 - missing_rate).reshape([random_tensor.shape[0],
# random_tensor.shape[1]
# * random_tensor.shape[2]])
# =============================================================================
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros(tensor.shape)
for i1 in range(tensor.shape[0]):
for i2 in range(tensor.shape[1]):
binary_tensor[i1,i2,:] = np.round(random_matrix[i1,i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1]
* binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 108 * 5
time_lags = np.array([1, 2, 108])
dim1, dim2 = sparse_mat.shape
rank = 10
lambda_w = 1000
lambda_x = 1000
lambda_theta = 1000
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# -
# **Experiment results** of spatial-temporal data prediction using TRMF:
#
# | scenario |`rank`|`Lambda_w`|`Lambda_x`|`Lambda_theta`|`eta`|`maxiter`| mape | rmse |
# |:----------|-----:|---------:|---------:|-------------:|----:|--------:|-----:|-----:|
# |**Original data**| 10 | 1000 | 1000 | 1000 | 0.03 | 200 | **0.2777**| **39.99**|
# |**20%, RM**| 10 | 1000 | 1000 | 1000 | 0.03 | 200 | **0.2759**| **40.73**|
# |**40%, RM**| 10 | 1000 | 1000 | 1000 | 0.03 | 200 | **0.2668**| **47.80**|
# |**20%, NM**| 10 | 1000 | 1000 | 1000 | 0.03 | 200 | **0.2658**| **45.23**|
# |**40%, NM**| 10 | 1000 | 1000 | 1000 | 0.03 | 200 | **0.2878**| **41.02**|
#
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
RM_mat = pd.read_csv('../datasets/Seattle-data-set/RM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
RM_mat = RM_mat.values
missing_rate = 0.2
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(RM_mat + 0.5 - missing_rate)
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 288 * 5
time_lags = np.array([1, 2, 288])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
RM_mat = pd.read_csv('../datasets/Seattle-data-set/RM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
RM_mat = RM_mat.values
missing_rate = 0.4
# =============================================================================
### Random missing (RM) scenario
### Set the RM scenario by:
binary_mat = np.round(RM_mat + 0.5 - missing_rate)
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 288 * 5
time_lags = np.array([1, 2, 288])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
NM_mat = pd.read_csv('../datasets/Seattle-data-set/NM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
NM_mat = NM_mat.values
missing_rate = 0.2
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros((dense_mat.shape[0], 28, 288))
for i1 in range(binary_tensor.shape[0]):
for i2 in range(binary_tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(NM_mat[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 288 * 5
time_lags = np.array([1, 2, 288])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# +
import pandas as pd
dense_mat = pd.read_csv('../datasets/Seattle-data-set/mat.csv', index_col = 0)
NM_mat = pd.read_csv('../datasets/Seattle-data-set/NM_mat.csv', index_col = 0)
dense_mat = dense_mat.values
NM_mat = NM_mat.values
missing_rate = 0.4
# =============================================================================
### Non-random missing (NM) scenario
### Set the NM scenario by:
binary_tensor = np.zeros((dense_mat.shape[0], 28, 288))
for i1 in range(binary_tensor.shape[0]):
for i2 in range(binary_tensor.shape[1]):
binary_tensor[i1, i2, :] = np.round(NM_mat[i1, i2] + 0.5 - missing_rate)
binary_mat = binary_tensor.reshape([binary_tensor.shape[0], binary_tensor.shape[1] * binary_tensor.shape[2]])
# =============================================================================
sparse_mat = np.multiply(dense_mat, binary_mat)
# +
import time
start = time.time()
pred_time_steps = 288 * 5
time_lags = np.array([1, 2, 288])
dim1, dim2 = sparse_mat.shape
rank = 30
lambda_w = 500
lambda_x = 500
lambda_theta = 500
eta = 0.03
d = time_lags.shape[0]
maxiter = 200
mat_hat = st_prediction(dense_mat, dense_mat, time_lags, lambda_w, lambda_x, lambda_theta,
eta, rank, pred_time_steps, maxiter)
end = time.time()
print('Running time: %d seconds'%(end - start))
# -
# **Experiment results** of spatial-temporal data prediction using TRMF:
#
# | scenario |`rank`|`Lambda_w`|`Lambda_x`|`Lambda_theta`|`eta`|`maxiter`| mape | rmse |
# |:----------|-----:|---------:|---------:|-------------:|----:|----------:|-----:|-----:|
# |**Original data**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.0796** | **4.90**|
# |**20%, RM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.0795** | **4.90**|
# |**40%, RM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.0795** | **4.90**|
# |**20%, NM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.0794** | **4.89**|
# |**40%, NM**| 30 | 500 | 500 | 500 | 0.03 | 200 | **0.0796** | **4.90**|
#
| experiments/Prediction-ST-Online-TRMF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import qiskit
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from qiskit import BasicAer, IBMQ
from qiskit import QuantumCircuit, ClassicalRegister, QuantumRegister, execute
from qiskit.compiler import transpile
from qiskit.tools.visualization import plot_histogram
# -
# ### Initialization
# +
def init(circuit, f_in, f_out, n): #initialize the input states
for j in range(n):
circuit.h(f_in[j])
circuit.x(f_out)
circuit.h(f_out)
circuit.barrier()
clause_list = [[1,2,-3],[-1,-2,-3],[-1,2,3]]
f_in = QuantumRegister(3)
f_out = QuantumRegister(1)
auxx = QuantumRegister(1)
qc_aux = QuantumCircuit(auxx)
qc = QuantumCircuit(f_in, f_out)
init(qc, f_in, f_out, 3)
qc.draw(output = 'mpl')
# -
# ### Triple-controlled NOT gate
def tri_cx(f_in, aux, k):
tri_aux = QuantumRegister(1)
qc1 = QuantumCircuit(f_in, tri_aux, aux)
qc1.ccx(f_in[0], f_in[1], aux[k])
qc1.ccx(f_in[2], tri_aux, aux[k])
qc1.ccx(f_in[0], f_in[1], aux[k])
qc1.barrier()
return(qc1)
# circuit = circuit + qc1
# ### Bit-flipping clause
def flip_clause(f_in, clause, flip_aux, k):
qc = QuantumCircuit(f_in, flip_aux)
tri = tri_cx(f_in, flip_aux, k)
for (j,literal) in enumerate(clause):
if (literal < 0):
qc.x(f_in[j])
qc.cx(f_in,flip_aux[k])
qc.barrier()
qc = qc + tri
for (j,literal) in enumerate(clause):
if (literal < 0):
qc.x(f_in[j])
qc.barrier()
return(qc)
# ### Black-box function $U_f$
# +
def uf(f_in, f_out, clause_list):
aux = QuantumRegister(len(clause_list))
quf = QuantumCircuit(f_in, f_out, aux)
for(k, clause) in enumerate(clause_list):
quf = quf + flip_clause(f_in, clause, aux, k) #for(k, clause) in enumerate(clause_list):
quf = quf + tri_cx(aux, f_out, 0)
quf.barrier()
return(quf)
quff = uf(f_in, f_out, clause_list)
quff.draw(output = 'mpl', scale = 0.5)
# -
# ### Inversion about the average
def inversion(f_in, f_out)
q_in = QuantumRegister(f_in)
q_out = QuantumRegister(f_out)
qc = QuantumCircuit(q_in, q_out)
# ### $C^{n-1}Z$ gate
def cnz()
| Grover's Search Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### _Speech Processing Labs 2020: TTS: Module 4_
# run this first
import matplotlib.pyplot as plt
import numpy as np
import math
import IPython
# # 2 Building a Decision Tree (using pencil and paper)
#
# ### Learning Outcomes
# * Understand in detail how to choose the best partition of some data, to reduce entropy
# * Be able to calculate entropy for distributions of real data
# * Be able to grow a small decision tree without running any code
#
# ### Need to know
# * what entropy is (from the previous notebook)
# * Topic Videos: Prosody, Decision tree, Learning decision trees
#
# **This exercise requires pencil and paper, so go get some! I mean it!**
#
# We are going to do a few iterations of growing a Decision Tree by hand. This is not a handwritten tree though: we're running the normal algorithm, but we're going to perform it manually so we really understand every step. (Later, we'll run it in code.)
#
# You should do this notebook in groups of 2-3 students.
#
# Before you start, here's a short video to remind you how a decision tree is learned, to complement the topic videos.
from IPython.display import HTML
IPython.display.IFrame(width="640",height="428",src="https://fast.wistia.net/embed/iframe/9l5iq63ezi")
# ## 2.1 Raw data
#
# You are going to grow a small decision tree to perform the task of predicting where to place phrase breaks in a sentence, based only on the text. Fortunately, someone has already recorded some sentences, listened for where the speaker made phrase breaks, and labelled the data with those, so you don't need to. That's a relief! The raw data look like this, where "|" marks a phrase break:
# ```
# You know a little too much, | <NAME>. |
# It was obvious | that he was badly puzzled. |
# In half an hour | I was reading. |
# Then something awoke me. |
# He stammered a little, | like a man picking his words. |
# I pulled up a chair | and sat down on it. |
# I tore open the Tide Tables | and found Bradgate. |
# I took my head in my hands | and thought. |
# He came back | in ten minutes | with a long face. |
# He will be in London | at five. |
# Then the dark fell, | and silence. |
# But it was a chance, | the only possible chance. |
# It was obvious | that he was badly puzzled. |
# Then with some difficulty | I turned the car. |
# ```
# ## 2.2 Data preparation (feature extraction and feature engineering)
#
# We rarely use the raw data directly in machine learning. Usually, we have some choices to make, using our linguistic knowledge and our skills as machine learning engineers.
#
# ### 2.2.1 Dealing with variable length sequences
# As in so many problems involving a variable length sequence (here, the sentence), we reduce the problem to a fixed length sequence by using a sliding window. We need to do this because Decision Trees work with a fixed set of predictors, not a variable number.
#
# The sliding window will be placed around each location where a break might occur: each word juncture.
#
# ### 2.2.2 Choosing the predictors
# We also have to choose whether to use the raw predictor values (the orthographic words), or to process them in some way. Words are an open set with a very large numer of possible values, so the data will be very sparse. We will replace each word with a super-simple Part-Of-Speech (POS) tag:
#
# PUNC = punctuation
# CONT = content words
# FUNC = function words
#
# That makes the data look like this, stored as list of sentences:
corpus=[
"FUNC CONT FUNC CONT CONT CONT PUNC | CONT CONT PUNC |",
"FUNC CONT CONT | FUNC FUNC CONT CONT CONT PUNC |",
"FUNC CONT FUNC CONT | FUNC CONT CONT PUNC |",
"FUNC CONT CONT FUNC PUNC |",
"FUNC CONT FUNC CONT PUNC | CONT FUNC CONT CONT FUNC CONT PUNC |",
"FUNC CONT FUNC FUNC CONT | FUNC CONT CONT FUNC FUNC PUNC |",
"FUNC CONT CONT FUNC CONT CONT | FUNC CONT CONT PUNC |",
"FUNC CONT FUNC CONT FUNC FUNC CONT | FUNC CONT PUNC |",
"FUNC CONT CONT | FUNC CONT CONT | FUNC FUNC CONT CONT PUNC |",
"FUNC CONT FUNC FUNC CONT | FUNC CONT PUNC |",
"FUNC FUNC CONT CONT PUNC | FUNC CONT PUNC |",
"FUNC CONT CONT FUNC CONT PUNC | FUNC CONT CONT CONT PUNC |",
"FUNC CONT CONT | FUNC FUNC CONT CONT CONT PUNC |",
"FUNC FUNC FUNC CONT | FUNC CONT FUNC CONT PUNC |"]
# ### 2.2.3 What is the predictee?
# The predictee always has to have a value. In the raw data, it is only defined at word junctures where there was a break. We will rewrite the predictee so it is either "NB" (no break) or "B" (break) at *every word juncture*.
# ## 2.3 Data prepared for machine learning
#
# ### 2.3.1 Nicely formatted data
#
# From the raw data, we need to extract a set of data points for learning a decision tree. Each data point comprises the values of the predictors and the corresponding correct value of the predictee. This is the training data.
# +
data = []
for line in corpus:
# pad, ready for sliding window
line="PAD "+line+" PAD"
line=line.replace(" | ","-B-").replace(" "," NB ").replace("-B-"," B ")
words=line.split()
for i in range(0,len(words)):
if words[i] == "B" or words[i] == "NB":
data.append((words[i-1],words[i+1],words[i]))
print("Created",len(data),"data points. Each data point has the form (PREV, NEXT, PREDICTEE)")
for d in data:
print("{}".format(d))
# -
# ### 2.3.2 List all possible questions we can ask about the predictors
#
# There are two predictors - the POS of the previous and next word surrounding a word juncture. We've called them PREV and NEXT. Each can take one of a fixed set of values. Write down all possible questions you can ask, in the box below. Don't worry about whether they are *actually useful* - that's not your job! (The data will tell us that.) I've given you one question as a starting point:
# ```
# PREV=="FUNC"
# ```
# ...now edit this part to write down all other possible questions
# ## 2.4 Tree growing
#
# ### 2.4.1 Measure the entropy of the data before it is split
#
# Do this by hand. You need to find the distribution of predictee values in the above data set, then apply the entropy equation. Do the calculation on paper and not in Python! The only part you can't easily do by hand is to compute logarithms, so here's a function for you to do that:
p=0.5 # adjust this as required
print("{:.3}".format(math.log2(p)))
# ### 2.4.2 Pick a question and split the data, then measure the entropy of the two partitions
#
# I'll start you off, and give you some code that will split the data for you. If you prefer (and you might learn more by doing this), you could print out the data on paper and cut it into 145 pieces, then do all of this without any code.
# +
yes=[]
no=[]
for d in data:
# d[0] contains PREV and d[1] contains NEXT, with the predictee in d[2]
if d[0] == "PUNC": # <- change this for each of your questions in turn
yes.append(d)
else:
no.append(d)
yes_predictees = [d[2] for d in yes]
no_predictees = [d[2] for d in no]
print("Data for 'yes' is",yes,"\n")
print("Data for 'no' is",no,"\n\n")
yes_predictees.sort()
no_predictees.sort()
print("Predictee distribution for 'yes' is",yes_predictees,"\n")
print("Predictee distribution for 'no' is",no_predictees,"\n\n")
print("From a total of",len(data),"data points: 'yes' partition",len(yes),"/ 'no' partition",len(no))
# -
# Now we need to calculate the entropy of each of those two distributions, and compute the weighted sum. This will give the total entropy of the partitioned data.
#
# The 'yes' one is easy - they are all 'NB' so the entropy is 0. You do the one for 'no'.
#
# #### Repeat for all questions
#
# Repeat step 2.4.2 for every one of of the questions you came up with in 2.3.2 (divide the labour amongst your group), recording the entropy for each one.
#
# ### 2.4.3 Place the best question into the tree
#
# Once you have decided on the best question to split the whole data, write that as the root of the tree and draw a 'yes' and 'no' branch (really, do it on a piece of paper!).
#
# ### 2.4.4 Recurse
#
# Now recurse down one or both of those branches, treating each partition in turn as the training data. Your goal is to grow a *very* small tree. Keep going until you think you properly understand the algorithm.
#
# ## 2.5 Test
#
# When you are finished, make up a new sentence and use your tree to predict where the phrase breaks should be placed.
| tts/tts-m4-2-decision-tree-pencil-and-paper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import ssl
ssl._create_default_https_context = ssl._create_unverified_context
def printCounter(counter, min_threshold = 10):
print(newdict = {x: count for x, count in sorted(getNgrams(content, 2).items(), key=lambda item: -item[1]) if count >= min_threshold})
# +
from urllib.request import urlopen
from bs4 import BeautifulSoup
import re
import string
from collections import Counter
def cleanSentence(sentence):
sentence = sentence.split(' ')
sentence = [word.strip(string.punctuation+string.whitespace) for word in sentence]
sentence = [word for word in sentence if len(word) > 1 or (word.lower() == 'a' or word.lower() == 'i')]
return sentence
def cleanInput(content):
content = content.upper()
content = re.sub('\n', ' ', content)
content = bytes(content, 'UTF-8')
content = content.decode('ascii', 'ignore')
sentences = content.split('. ')
return [cleanSentence(sentence) for sentence in sentences]
def getNgramsFromSentence(content, n):
output = []
for i in range(len(content)-n+1):
output.append(content[i:i+n])
return output
def getNgrams(content, n):
content = cleanInput(content)
ngrams = Counter()
ngrams_list = []
for sentence in content:
newNgrams = [' '.join(ngram) for ngram in getNgramsFromSentence(sentence, n)]
ngrams_list.extend(newNgrams)
ngrams.update(newNgrams)
return(ngrams)
content = str(
urlopen('http://pythonscraping.com/files/inaugurationSpeech.txt').read(),
'utf-8')
ngrams = getNgrams(content, 3)
printCounter(ngrams)
# -
# +
def isCommon(ngram):
commonWords = ['THE', 'BE', 'AND', 'OF', 'A', 'IN', 'TO', 'HAVE', 'IT', 'I', 'THAT', 'FOR', 'YOU', 'HE', 'WITH', 'ON', 'DO', 'SAY', 'THIS', 'THEY', 'IS', 'AN', 'AT', 'BUT', 'WE', 'HIS', 'FROM', 'THAT', 'NOT', 'BY', 'SHE', 'OR', 'AS', 'WHAT', 'GO', 'THEIR', 'CAN', 'WHO', 'GET', 'IF', 'WOULD', 'HER', 'ALL', 'MY', 'MAKE', 'ABOUT', 'KNOW', 'WILL', 'AS', 'UP', 'ONE', 'TIME', 'HAS', 'BEEN', 'THERE', 'YEAR', 'SO', 'THINK', 'WHEN', 'WHICH', 'THEM', 'SOME', 'ME', 'PEOPLE', 'TAKE', 'OUT', 'INTO', 'JUST', 'SEE', 'HIM', 'YOUR', 'COME', 'COULD', 'NOW', 'THAN', 'LIKE', 'OTHER', 'HOW', 'THEN', 'ITS', 'OUR', 'TWO', 'MORE', 'THESE', 'WANT', 'WAY', 'LOOK', 'FIRST', 'ALSO', 'NEW', 'BECAUSE', 'DAY', 'MORE', 'USE', 'NO', 'MAN', 'FIND', 'HERE', 'THING', 'GIVE', 'MANY', 'WELL']
for word in ngram:
if word in commonWords:
return True
return False
def getNgramsFromSentence(content, n):
output = []
for i in range(len(content)-n+1):
if not isCommon(content[i:i+n]):
output.append(content[i:i+n])
return output
ngrams = getNgrams(content, 3)
print(ngrams)
# +
def getFirstSentenceContaining(ngram, content):
#print(ngram)
sentences = content.upper().split(". ")
for sentence in sentences:
if ngram in sentence:
return sentence+'\n'
return ""
print(getFirstSentenceContaining('EXCLUSIVE METALLIC CURRENCY', content))
print(getFirstSentenceContaining('EXECUTIVE DEPARTMENT', content))
print(getFirstSentenceContaining('GENERAL GOVERNMENT', content))
print(getFirstSentenceContaining('CALLED UPON', content))
print(getFirstSentenceContaining('CHIEF MAGISTRATE', content))
# +
from urllib.request import urlopen
from random import randint
def wordListSum(wordList):
sum = 0
for word, value in wordList.items():
sum += value
return sum
def retrieveRandomWord(wordList):
randIndex = randint(1, wordListSum(wordList))
for word, value in wordList.items():
randIndex -= value
if randIndex <= 0:
return word
def buildWordDict(text):
# Remove newlines and quotes
text = text.replace('\n', ' ');
text = text.replace('"', '');
# Make sure punctuation marks are treated as their own "words,"
# so that they will be included in the Markov chain
punctuation = [',','.',';',':']
for symbol in punctuation:
text = text.replace(symbol, ' {} '.format(symbol));
words = text.split(' ')
# Filter out empty words
words = [word for word in words if word != '']
wordDict = {}
for i in range(1, len(words)):
if words[i-1] not in wordDict:
# Create a new dictionary for this word
wordDict[words[i-1]] = {}
if words[i] not in wordDict[words[i-1]]:
wordDict[words[i-1]][words[i]] = 0
wordDict[words[i-1]][words[i]] += 1
return wordDict
text = str(urlopen('http://pythonscraping.com/files/inaugurationSpeech.txt')
.read(), 'utf-8')
wordDict = buildWordDict(text)
#Generate a Markov chain of length 100
length = 100
chain = ['I']
for i in range(0, length):
newWord = retrieveRandomWord(wordDict[chain[-1]])
chain.append(newWord)
print(' '.join(chain))
# -
# +
import pymysql
conn = pymysql.connect(host='127.0.0.1', unix_socket='/tmp/mysql.sock', user='root', passwd='<PASSWORD>', db='mysql', charset='utf8')
cur = conn.cursor()
cur.execute('USE wikipedia')
def getUrl(pageId):
cur.execute('SELECT url FROM pages WHERE id = %s', (int(pageId)))
return cur.fetchone()[0]
def getLinks(fromPageId):
cur.execute('SELECT toPageId FROM links WHERE fromPageId = %s', (int(fromPageId)))
if cur.rowcount == 0:
return []
return [x[0] for x in cur.fetchall()]
def searchBreadth(targetPageId, paths=[[1]]):
newPaths = []
for path in paths:
links = getLinks(path[-1])
for link in links:
if link == targetPageId:
return path + [link]
else:
newPaths.append(path+[link])
return searchBreadth(targetPageId, newPaths)
nodes = getLinks(1)
targetPageId = 28624
pageIds = searchBreadth(targetPageId)
for pageId in pageIds:
print(getUrl(pageId))
# -
'
| Chapter09_NaturalLanguages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Devanagri Character Dataset
#
# The dataset can be found [here](https://www.kaggle.com/ashokpant/devanagari-character-dataset)
#
# Let's start by importing the stuff we need. Note that this is not intended for any kind of production use. Using * for import is never a good idea except for prototyping
from fastai.vision import *
from fastai.metrics import *
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# The following lines of code expect that the data has been downloaded, extracted and laid out in the required directory structure
path = Path('DevanagariHandwrittenCharacterDataset')
path.ls()
train = path/'Train'
valid = path/'Valid'
valid.mkdir(parents=True, exist_ok=True)
# # Initialize directories for all categories in Train
#
# Make sure to run the below code only once.
#
# Substitute shutil.copy for os.rename if you want to run this more than once
for f in train.iterdir():
path = f.relative_to(train)
os.mkdir(valid/path)
# # Split files into train and validation set
split = 0.2
counter = 1
for f in train.iterdir():
n = os.listdir(f)
len_dir = len(n)
num_valid = round(split*len_dir)
valid_img = random.sample(n, num_valid)
for img in valid_img:
file = f/img
path = f.relative_to(train)
valid_path = valid/path/img
os.rename(file, valid_path)
data = ImageDataBunch.from_folder(path, train='Train', valid='Valid', ds_tfms = get_transforms(do_flip=False), size=28, bs=32)
data.show_batch(rows=4, figsize=(12, 12))
print(data.c)
print(data.classes)
learn = create_cnn(data, models.resnet34, metrics = error_rate)
learn.fit_one_cycle(4)
learn.save('iter1')
learn.unfreeze()
learn.fit_one_cycle(1)
learn.lr_find()
learn.recorder.plot()
learn.load('iter1')
interpret = ClassificationInterpretation.from_learner(learn)
interpret.plot_top_losses(4, figsize=(9, 16))
interpret.most_confused(min_val=10)
learn.fit_one_cycle(2)
learn.fit_one_cycle(4, max_lr=slice(1e-6, 1e-5))
learn.fit_one_cycle(4, max_lr=slice(1e-6, 1e-5))
learn.lr_find()
learn.recorder.plot()
learn.save('iter2')
learn.fit_one_cycle(4, max_lr=slice(1e-5, 1e-4))
data = ImageDataBunch.from_folder(path, train='Train', valid='Valid', ds_tfms = get_transforms(do_flip=False), bs=64, size=32)
learn1 = create_cnn(data, models.resnet50, metrics=error_rate)
learn1.fit_one_cycle(4)
learn1.lr_find()
learn1.recorder.plot()
learn1.fit_one_cycle(4)
learn1.lr_find()
learn1.recorder.plot()
learn.fit_one_cycle(4, max_lr=slice(1e-6, 1e-4))
learn1.lr_find()
learn1.recorder.plot()
learn1.fit_one_cycle(2, max_lr=slice(1e-6, 1e-5))
learn1.lr_find()
learn1.recorder.plot()
learn1.save('stage1')
learn1.fit_one_cycle(1, max_lr=slice(1e-6, 1e-5))
learn1.lr_find()
learn1.recorder.plot()
learn1.load('stage1')
learn1.fit_one_cycle(1)
# That is a pretty good result, considering we've only spent a couple of hours on this. The result above, as you can see is underfitting but in the interest of learning, I'll leave that as an exercise. Good luck!!
| Devanagri Character Dataset/devnagri-character-dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myenv
# language: python
# name: myenv
# ---
# # Import Libraries
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# %matplotlib notebook
from argparse import ArgumentParser
import yaml
import os
import math
import torch
# from torch import vmap
from functorch import vmap, grad
from models import FNN2d
from train_utils import Adam
# from train_utils.datasets import BurgersLoader
# from train_utils.train_2d import train_2d_burger
# from train_utils.eval_2d import eval_burgers
from solver.WaveEq import WaveEq1D
import traceback
import scipy.io
import torch.nn.functional as F
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
from tqdm import tqdm
from train_utils.utils import save_checkpoint
from train_utils.losses import LpLoss
from solver.my_random_fields import GRF_Mattern
from importlib import reload
try:
import wandb
except ImportError:
wandb = None
# -
# # Load/Update Config Functions:
# +
def update_config(config, file):
with open(file, 'w') as f:
config_updated = yaml.dump(config, f)
def load_config(file):
with open(file, 'r') as f:
config = yaml.load(f, yaml.FullLoader)
return config
# -
# # Define Data Loader:
class DataLoader(object):
def __init__(self, x_data, y_data, nx=128, nt=100, sub=1, sub_t=1, new=True):
# dataloader = MatReader(datapath)
self.sub = sub
self.sub_t = sub_t
s = nx
# if nx is odd
if (s % 2) == 1:
s = s - 1
self.s = s // sub
self.T = nt // sub_t
self.new = new
if new:
self.T += 1
self.x_data = x_data[:, 0:s:sub]
self.y_data = y_data[:, 0:self.T:sub_t, 0:s:sub]
def make_loader(self, n_sample, batch_size, start=0, train=True):
Xs = self.x_data[start:start + n_sample]
ys = self.y_data[start:start + n_sample]
if self.new:
gridx = torch.tensor(np.linspace(0, 1, self.s + 1)[:-1], dtype=torch.float)
gridt = torch.tensor(np.linspace(0, 1, self.T), dtype=torch.float)
else:
gridx = torch.tensor(np.linspace(0, 1, self.s), dtype=torch.float)
gridt = torch.tensor(np.linspace(0, 1, self.T + 1)[1:], dtype=torch.float)
gridx = gridx.reshape(1, 1, self.s)
gridt = gridt.reshape(1, self.T, 1)
Xs = Xs.reshape(n_sample, 1, self.s).repeat([1, self.T, 1])
Xs = torch.stack([Xs, gridx.repeat([n_sample, self.T, 1]), gridt.repeat([n_sample, 1, self.s])], dim=3)
dataset = torch.utils.data.TensorDataset(Xs, ys)
if train:
loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, shuffle=True)
else:
loader = torch.utils.data.DataLoader(dataset, batch_size=batch_size, shuffle=False)
return loader
# # Define Loss Functions
# ## Automatic Differentiation
# +
def Autograd_Wave(u, grid, c=1.0):
from torch.autograd import grad
gridt, gridx = grid
ut = grad(u.sum(), gridt, create_graph=True)[0]
utt = grad(ut.sum(), gridt, create_graph=True)[0]
ux = grad(u.sum(), gridx, create_graph=True)[0]
uxx = grad(ux.sum(), gridx, create_graph=True)[0]
Du = utt - c**2*uxx
return Du, uxx, utt
def AD_loss_Wave(u, u0, grid, index_ic=None, p=None, q=None, c=1.0):
batchsize = u.size(0)
# lploss = LpLoss(size_average=True)
Du, uxx, utt = Autograd_Wave(u, grid, c=c)
if index_ic is None:
# u in on a uniform grid
nt = u.size(1)
nx = u.size(2)
u = u.reshape(batchsize, nt, nx)
index_t = torch.zeros(nx,).long()
index_x = torch.tensor(range(nx)).long()
boundary_u = u[:, index_t, index_x]
# loss_bc0 = F.mse_loss(u[:, :, 0], u[:, :, -1])
# loss_bc1 = F.mse_loss(ux[:, :, 0], ux[:, :, -1])
else:
# u is randomly sampled, 0:p are BC, p:2p are ic, 2p:2p+q are interior
boundary_u = u[:, :p]
batch_index = torch.tensor(range(batchsize)).reshape(batchsize, 1).repeat(1, p)
u0 = u0[batch_index, index_ic]
# loss_bc0 = F.mse_loss(u[:, p:p+p//2], u[:, p+p//2:2*p])
# loss_bc1 = F.mse_loss(ux[:, p:p+p//2], ux[:, p+p//2:2*p])
loss_ic = F.mse_loss(boundary_u, u0)
f = torch.zeros(Du.shape, device=u.device)
loss_f = F.mse_loss(Du, f)
return loss_ic, loss_f
# -
# ## Spectral Derivatives
# +
def FDM_Wave(u, D=1, c=1.0):
batchsize = u.size(0)
nt = u.size(1)
nx = u.size(2)
u = u.reshape(batchsize, nt, nx)
dt = D / (nt-1)
dx = D / (nx)
u_h = torch.fft.fft(u, dim=2)
# Wavenumbers in y-direction
k_max = nx//2
k_x = torch.cat((torch.arange(start=0, end=k_max, step=1, device=u.device),
torch.arange(start=-k_max, end=0, step=1, device=u.device)), 0).reshape(1,1,nx)
ux_h = 2j *np.pi*k_x*u_h
uxx_h = 2j *np.pi*k_x*ux_h
ux = torch.fft.irfft(ux_h[:, :, :k_max+1], dim=2, n=nx)
uxx = torch.fft.irfft(uxx_h[:, :, :k_max+1], dim=2, n=nx)
ut = (u[:, 2:, :] - u[:, :-2, :]) / (2 * dt)
utt = (u[:, 2:, :] - 2.0*u[:, 1:-1, :] + u[:, :-2, :]) / (dt**2)
Du = utt - c**2 * uxx[:,1:-1,:]
return Du
def PINO_loss_wave(u, u0, c=1.0):
batchsize = u.size(0)
nt = u.size(1)
nx = u.size(2)
u = u.reshape(batchsize, nt, nx)
# lploss = LpLoss(size_average=True)
index_t = torch.zeros(nx,).long()
index_x = torch.tensor(range(nx)).long()
boundary_u = u[:, index_t, index_x]
loss_u = F.mse_loss(boundary_u, u0)
Du = FDM_Wave(u, c=c)[:, :, :]
f = torch.zeros(Du.shape, device=u.device)
loss_f = F.mse_loss(Du, f)
# loss_bc0 = F.mse_loss(u[:, :, 0], u[:, :, -1])
# loss_bc1 = F.mse_loss((u[:, :, 1] - u[:, :, -1]) /
# (2/(nx)), (u[:, :, 0] - u[:, :, -2])/(2/(nx)))
return loss_u, loss_f
# -
# # Define Training Function
def train_wave(model,
train_loader,
optimizer,
scheduler,
config,
rank=0,
log=False,
project='PINO-2d-default',
group='default',
tags=['default'],
use_tqdm=True):
if rank == 0 and wandb and log:
run = wandb.init(project=project,
entity='shawngr2',
group=group,
config=config,
tags=tags, reinit=True,
settings=wandb.Settings(start_method="fork"))
data_weight = config['train']['xy_loss']
f_weight = config['train']['f_loss']
ic_weight = config['train']['ic_loss']
c = config['data']['c']
ckpt_freq = config['train']['ckpt_freq']
model.train()
myloss = LpLoss(size_average=True)
pbar = range(config['train']['epochs'])
if use_tqdm:
pbar = tqdm(pbar, dynamic_ncols=True, smoothing=0.1)
for e in pbar:
model.train()
train_pino = 0.0
data_l2 = 0.0
train_ic = 0.0
train_loss = 0.0
for x, y in train_loader:
x, y = x.to(rank), y.to(rank)
# display(x.shape, y.shape)
out = model(x).reshape(y.shape)
data_loss = myloss(out, y)
loss_ic, loss_f = PINO_loss_wave(out, x[:, 0, :, 0], c=c)
total_loss = loss_ic * ic_weight + loss_f * f_weight + data_loss * data_weight
optimizer.zero_grad()
total_loss.backward()
optimizer.step()
data_l2 += data_loss.item()
train_pino += loss_f.item()
train_loss += total_loss.item()
train_ic += loss_ic.item()
scheduler.step()
data_l2 /= len(train_loader)
train_pino /= len(train_loader)
train_loss /= len(train_loader)
if use_tqdm:
pbar.set_description(
(
f'Epoch {e}, train loss: {train_loss:.5f} '
f'train f error: {train_pino:.5f}; '
f'data l2 error: {data_l2:.5f}; '
f'train ic error: {train_ic:.5f}'
)
)
if wandb and log:
wandb.log(
{
'Train f error': train_pino,
'Train L2 error': data_l2,
'Train ic error': loss_ic,
'Train loss': train_loss,
}
)
if e % ckpt_freq == 0:
save_checkpoint(config['train']['save_dir'],
config['train']['save_name'].replace('.pt', f'_{e}.pt'),
model, optimizer)
save_checkpoint(config['train']['save_dir'],
config['train']['save_name'],
model, optimizer)
print('Done!')
# # Evaluation Function
# +
def eval_wave(model,
dataloader,
config,
device,
use_tqdm=True):
model.eval()
myloss = LpLoss(size_average=True)
c = config['data']['c']
if use_tqdm:
pbar = tqdm(dataloader, dynamic_ncols=True, smoothing=0.05)
else:
pbar = dataloader
test_err = []
f_err = []
for x, y in pbar:
x, y = x.to(device), y.to(device)
out = model(x).reshape(y.shape)
data_loss = myloss(out, y)
loss_u, f_loss = PINO_loss_wave(out, x[:, 0, :, 0], c=c)
test_err.append(data_loss.item())
f_err.append(f_loss.item())
mean_f_err = np.mean(f_err)
std_f_err = np.std(f_err, ddof=1) / np.sqrt(len(f_err))
mean_err = np.mean(test_err)
std_err = np.std(test_err, ddof=1) / np.sqrt(len(test_err))
print(f'==Averaged relative L2 error mean: {mean_err}, std error: {std_err}==\n'
f'==Averaged equation error mean: {mean_f_err}, std error: {std_f_err}==')
# -
# # Checkpoint Loading
def load_checkpoint(model, ckpt_path, optimizer=None):
try:
ckpt = torch.load(ckpt_path)
model.load_state_dict(ckpt['model'])
print('Weights loaded from %s' % ckpt_path)
if optimizer is not None:
try:
optimizer.load_state_dict(ckpt['optim'])
print('Optimizer loaded from %s' % ckpt_path)
except: traceback.print_exc()
except:
traceback.print_exc()
# # Load Config File
config_file = 'configs/custom/wave-0000.yaml'
config = load_config(config_file)
display(config)
# # Parameters
# +
# dim = 1
# N = 4096
# Nx = 4096
# l = 0.1
# Nk = None
# Nsamples = 1000
# # jitter = 1e-12
# dt = 1.0e-4
# save_int = int(1e-2/dt)
# device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# grf = GaussianRF(dim, N, length=1.0, alpha=2.5, tau=5.0, device=device)
# U0 = grf.sample(Nsamples)
# +
Nsamples = config['data']['total_num']
N = config['data']['nx']
Nt0 = config['data']['nt']
c = config['data']['c']
sub_x = config['data']['sub']
sub_t = config['data']['sub_t']
Nx = N // sub_x
Nt = Nt0 // sub_t + 1
dim = 1
l = 0.1
L = 1.0
sigma = 0.2 #2.0
Nu = None # 2.0
dt = 1.0e-4
tend = 1.0
save_int = int(tend/dt/Nt)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# -
sub_x
4096//sub_x
# # Generate Random Fields
# +
# grf = GaussianRF(dim, N, length=1.0, alpha=2.5, tau=5.0, device=device)
# U0 = grf.sample(Nsamples)
# -
grf = GRF_Mattern(dim, N, length=L, nu=Nu, l=l, sigma=sigma, boundary="periodic", device=device)
U0 = grf.sample(Nsamples)
U0.shape
U0.shape
# +
# if dim == 1:
# U = np.array([GRF.plot_sample(X, sample, dim, shape) for sample in samples])
# if dim == 2:
# U = np.array([plot_surf(X, sample, shape) for sample in samples])
# -
wave_eq = WaveEq1D(Nx=N, c=c, dt=dt, device=device)
U = vmap(wave_eq.wave_driver, in_dims=(0, None))(U0, save_int)
# +
a = U0.cpu().float()
u = U.cpu().float()
display(u.shape,a.shape)
# +
# config_file_train = 'configs/custom/wave-train-0000.yaml'
# config_file_test = 'configs/custom/wave-test-0000.yaml'
# with open(config_file_train, 'r') as stream:
# config_train = yaml.load(stream, yaml.FullLoader)
# with open(config_file_test, 'r') as stream:
# config_test = yaml.load(stream, yaml.FullLoader)
# -
dataset = DataLoader(a, u, config['data']['nx'], config['data']['nt'], config['data']['sub'], config['data']['sub_t'])
train_loader = dataset.make_loader(config['data']['n_train'], config['train']['batchsize'], start=0, train=True)
test_loader = dataset.make_loader(config['data']['n_test'], config['test']['batchsize'], start=config['data']['n_train'], train=False)
# +
log = False
model = FNN2d(modes1=config['model']['modes1'],
modes2=config['model']['modes2'],
fc_dim=config['model']['fc_dim'],
layers=config['model']['layers'],
activation=config['model']['activation']).to(device)
optimizer = Adam(model.parameters(), betas=(0.9, 0.999),lr=config['train']['base_lr'])
scheduler = torch.optim.lr_scheduler.MultiStepLR(optimizer,
milestones=config['train']['milestones'],
gamma=config['train']['scheduler_gamma'])
# -
# # Load from checkpoint
load_checkpoint(model, ckpt_path=config['train']['ckpt'], optimizer=None)
# # Train the Model
# +
train_wave(model,
train_loader,
optimizer,
scheduler,
config,
rank=0,
log=log,
project=config['log']['project'],
group=config['log']['group'])
# -
# # Evaluate on Test Data
eval_wave(model, test_loader, config, device)
eval_wave(model, test_loader, config, device)
Nx = config['data']['nx'] // config['data']['sub']
Nt = config['data']['nt'] // config['data']['sub_t'] + 1
Ntest = config['data']['n_test']
model.eval()
test_x = np.zeros((Ntest,Nt,Nx,3))
preds_y = np.zeros((Ntest,Nt,Nx))
test_y = np.zeros((Ntest,Nt,Nx))
with torch.no_grad():
for i, data in enumerate(test_loader):
data_x, data_y = data
data_x, data_y = data_x.to(device), data_y.to(device)
pred_y = model(data_x).reshape(data_y.shape)
test_x[i] = data_x.cpu().numpy()
test_y[i] = data_y.cpu().numpy()
preds_y[i] = pred_y.cpu().numpy()
# data_loss = myloss(out, y)
# +
key = 1
pred = preds_y[key]
true = test_y[key]
a = test_x[key]
Nt, Nx, _ = a.shape
u0 = a[0,:,0]
T = a[:,:,2]
X = a[:,:,1]
x = X[0]
# -
plt.rcParams.update({'font.size': 11})
# +
fig = plt.figure(figsize=(24,5))
plt.subplot(1,4,1)
plt.plot(x, u0)
plt.xlabel('$x$')
plt.ylabel('$u$')
plt.title('Intial Condition $u(x)$')
plt.xlim([0,1])
plt.tight_layout()
plt.subplot(1,4,2)
# plt.pcolor(XX,TT, S_test, cmap='jet')
plt.pcolormesh(X, T, true, cmap='jet', shading='gouraud')
plt.colorbar()
plt.xlabel('$x$')
plt.ylabel('$t$')
plt.title(f'Exact $u(x,t)$')
plt.tight_layout()
plt.axis('square')
plt.subplot(1,4,3)
# plt.pcolor(XX,TT, S_pred, cmap='jet')
plt.pcolormesh(X, T, pred, cmap='jet', shading='gouraud')
plt.colorbar()
plt.xlabel('$x$')
plt.ylabel('$t$')
plt.title(f'Predict $u(x,t)$')
plt.axis('square')
plt.tight_layout()
plt.subplot(1,4,4)
# plt.pcolor(XX,TT, S_pred - S_test, cmap='jet')
plt.pcolormesh(X, T, pred - true, cmap='jet', shading='gouraud')
plt.colorbar()
plt.xlabel('$x$')
plt.ylabel('$t$')
plt.title('Absolute error')
plt.tight_layout()
plt.axis('square')
# plt.show()
# -
# +
# %matplotlib notebook
fig = plt.figure(figsize=(6,5))
ax = fig.add_subplot(111)
plt.ion()
fig.show()
fig.canvas.draw()
ax.plot(x, true[0], 'b-', label='Exact')
ax.plot(x, pred[0], 'r--', label='PINO Prediction')
ylim = plt.ylim()
xlim = [0, 1]
plt.tight_layout()
for i in range(Nt):
ax.clear()
ax.plot(x, true[i], 'b-', label='Exact')
ax.plot(x, pred[i], 'r--', label='PINO Prediction')
plt.ylim(ylim)
plt.xlim(xlim)
plt.xlabel(f'$x$')
plt.ylabel(f'$u$')
plt.title(f'Wave Equation')
plt.legend(loc='lower right')
plt.tight_layout()
fig.canvas.draw()
# -
# # Save and Load Data
# +
def save_data(data_path, test_x, test_y, preds_y):
data_dir, data_filename = os.path.split(data_path)
os.makedirs(data_dir, exist_ok=True)
np.savez(data_path, test_x=test_x, test_y=test_y, preds_y=preds_y)
def load_data(data_path):
data = np.load(data_path)
test_x = data['test_x']
test_y = data['test_y']
preds_y = data['preds_y']
return test_x, test_y, preds_y
# -
data_dir = 'data/Wave1D'
data_filename = 'data.npz'
data_path = os.path.join(data_dir, data_filename)
# os.makedirs(data_dir, exist_ok=True)
save_data(data_path, test_x, test_y, preds_y)
test_x, test_y, preds_y = load_data(data_path)
# +
def plot_predictions(key, test_x, test_y, preds_y, print_index=False, save_path=None, font_size=None):
if font_size is not None:
plt.rcParams.update({'font.size': font_size})
pred = preds_y[key]
true = test_y[key]
a = test_x[key]
Nt, Nx, _ = a.shape
u0 = a[0,:,0]
T = a[:,:,2]
X = a[:,:,1]
x = X[0]
# Plot
fig = plt.figure(figsize=(23,5))
plt.subplot(1,4,1)
plt.plot(x, u0)
plt.xlabel('$x$')
plt.ylabel('$u$')
plt.title('Intial Condition $u(x)$')
plt.xlim([0,1])
plt.tight_layout()
plt.subplot(1,4,2)
# plt.pcolor(XX,TT, S_test, cmap='jet')
plt.pcolormesh(X, T, true, cmap='jet', shading='gouraud')
plt.colorbar()
plt.xlabel('$x$')
plt.ylabel('$t$')
plt.title(f'Exact $u(x,t)$')
plt.tight_layout()
plt.axis('square')
plt.subplot(1,4,3)
# plt.pcolor(XX,TT, S_pred, cmap='jet')
plt.pcolormesh(X, T, pred, cmap='jet', shading='gouraud')
plt.colorbar()
plt.xlabel('$x$')
plt.ylabel('$t$')
plt.title(f'Predict $u(x,t)$')
plt.axis('square')
plt.tight_layout()
plt.subplot(1,4,4)
# plt.pcolor(XX,TT, S_pred - S_test, cmap='jet')
plt.pcolormesh(X, T, pred - true, cmap='jet', shading='gouraud')
plt.colorbar()
plt.xlabel('$x$')
plt.ylabel('$t$')
plt.title('Absolute Error')
plt.tight_layout()
plt.axis('square')
if save_path is not None:
plt.savefig(f'{save_path}.png', bbox_inches='tight')
plt.show()
# -
# +
# %matplotlib inline
figures_dir = 'Wave1D/figures/'
os.makedirs(figures_dir, exist_ok=True)
font_size = 12
for key in range(len(preds_y)):
# for key in range(10):
save_path = os.path.join(figures_dir, f'Wave1D_{key}')
# plot_predictions(key, test_x, test_y, preds_y, print_index=True, save_path=None)
plot_predictions(key, test_x, test_y, preds_y, print_index=True, save_path=save_path, font_size=font_size)
# -
# +
# %matplotlib notebook
fig = plt.figure(figsize=(6,5))
ax = fig.add_subplot(111)
plt.ion()
fig.show()
fig.canvas.draw()
diff = pred - true
ax.plot(x, pred[0] - true[0], 'b-', label='Difference')
# ax.plot(x, pred[0], 'r--', label='PINO Prediction')
ylim = [diff.min(), diff.max()]
xlim = [0, 1]
plt.xlim(xlim)
plt.ylim(ylim)
plt.tight_layout()
for i in range(Nt):
ax.clear()
# ax.plot(x, true[i], 'b-', label='Exact')
# ax.plot(x, pred[i], 'r--', label='PINO Prediction')
ax.plot(x, diff[i], 'b-', label='Difference')
plt.ylim(ylim)
plt.xlim(xlim)
plt.xlabel(f'$x$')
plt.ylabel(f'$u$')
plt.title(f'Wave Equation')
plt.legend(loc='lower right')
plt.tight_layout()
fig.canvas.draw()
| Wave1D_PINO.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting started with Variational Quantum Eigensolving
#
# Here, we show how to find a variational ground state to the following Hamiltonian:
#
# $$ H = \vec{\sigma}_1\cdot \vec{\sigma}_2 = \sigma^X_1 \sigma^X_2 + \sigma^Y_1 \sigma^Y_2 + \sigma^Z_1 \sigma^Z_2 $$
#
# We start by defining the Hamiltonian (in the form of an observable), and choose a variational circuit with two parameters:
# +
from qat.core import Observable, Term
from qat.lang.AQASM import Program, CNOT, H, RX, RY, RZ
H_XY = Observable(2,
pauli_terms=[Term(1., "XX", [0, 1]),
Term(1., "YY", [0, 1]),
Term(1., "ZZ", [0, 1])]
)
print("Hamiltonian:", H_XY)
prog = Program()
qbits = prog.qalloc(2)
alpha = prog.new_var(float, "\\alpha")
beta = prog.new_var(float, "\\beta")
gamma = prog.new_var(float, "\\gamma")
prog.apply(H, qbits[0])
prog.apply(RY(alpha), qbits[1])
prog.apply(CNOT, qbits)
prog.apply(RX(beta), qbits[0])
prog.apply(RY(gamma), qbits[1])
circuit = prog.to_circ()
# %qatdisplay circuit
# -
# ## Optimization
#
# We now perform the optimization, using a Variational Plugin:
# +
from qat.plugins import ScipyMinimizePlugin
from qat.qpus import LinAlg
linalg_qpu = LinAlg()
theta0 = [0.4, -0.3, 0.6]
optimizer_scipy = ScipyMinimizePlugin(method="COBYLA",
x0=theta0,
tol=1e-3,
options={"maxiter": 2000})
qpu = optimizer_scipy | linalg_qpu
job = circuit.to_job(job_type="OBS", observable=H_XY)
result = qpu.submit(job)
print("Minimum energy =", result.value)
print("Optimal angles =", result.meta_data["parameters"])
#print("==========Optimization data=============\n", result.meta_data['optimization_trace'])
# %matplotlib inline
import matplotlib.pyplot as plt
plt.plot(eval(result.meta_data['optimization_trace']))
plt.xlabel("Steps")
plt.ylabel("Energy");
# -
# ## True ground state
#
# In this simple case, we can compute the ground state energy by diagonalizing the matrix corresponding to the Hamiltonian:
# +
import numpy as np
from qat.dqs.hamiltonians import SpinHamiltonian
H_XY_spin = SpinHamiltonian(nqbits=H_XY.nbqbits, pauli_terms=H_XY.terms)
H_XY_matrix = H_XY_spin.get_matrix()
eigvals, eigvecs = np.linalg.eigh(H_XY_matrix)
print("Exact ground state energy: ", min(eigvals))
# -
# ## Going further:
#
# - play with the Hamiltonian
# - play with the ansatz
# - play with the QPU: try with a noisy QPU
# - play with the optimizer (change the method)
| misc/notebooks/tutorials/variational_algorithms/vqe_getting_started_heisenberg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Jupyter Intro
#
# ### First cell
#
# First we are going to run a couple of commands in the first cell, to avoid reloading libraries that we may be modifying and to get the ouput of the graphs in jupyter. (For IPython version 3.1, 4.x, and 5.x)
#
# `autoreload` reloads modules automatically before entering the execution of code typed at the IPython prompt.
#
# `%autoreload` Reload all modules (except those excluded by %aimport) automatically now. <br>
# `%autoreload 0` Disable automatic reloading. <br>
# `%autoreload 1` Reload all modules imported with %aimport every time before executing the Python code typed. <br>
# `%autoreload 2` Reload all modules (except those excluded by %aimport) every time before executing the Python code typed. <br>
# `%aimport` List modules which are to be automatically imported or not to be imported. <br>
# `%aimport foo` Import module ‘foo’ and mark it to be autoreloaded for %autoreload 1 <br>
#
#
# `%matplotlib inline` The output of plotting commands is displayed inline, directly below the code cell that produced it. The resulting plots will then also be stored in the notebook document.
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %aimport
# ### Other useful information:
# - Magic functions:
#
# <table style="margin: 0 auto">
# <tr>
# <td style="text-align:left">%quickref</td>
# <td style="text-align:left">IPython quick reference</td>
# </tr>
# <tr>
# <td style="text-align:left">%qtconsole</td>
# <td style="text-align:left">Open a Qt console aware of the current workspace</td>
# </tr>
# <tr>
# <td style="text-align:left">%debug</td>
# <td style="text-align:left">Enters the interactive debugger.</td>
# </tr>
# <tr>
# <td style="text-align:left">%hist</td>
# <td style="text-align:left">Print command input (and output) history.</td>
# </tr>
# <tr>
# <td style="text-align:left">%reset</td>
# <td style="text-align:left">Delete all variables and names defined in the current namespace.</td>
# </tr>
# <tr>
# <td style="text-align:left">%run</td>
# <td style="text-align:left">Run a python script inside a notebook. Ex: %run script.py</td>
# </tr>
# <tr>
# <td style="text-align:left">%prun my_function()</td>
# <td style="text-align:left">Runs a function (or code) within the python profiler.</td>
# </tr>
# <tr>
# <td style="text-align:left">%run -p my_script.py</td>
# <td style="text-align:left">Runs a script under the profiler.</td>
# </tr>
# <tr>
# <td style="text-align:left">%env</td>
# <td style="text-align:left">Set environment variables.</td>
# </tr>
# <tr>
# <td style="text-align:left">%load ./hello_world.py</td>
# <td style="text-align:left">Insert the code from an external script</td>
# </tr>
# <tr>
# <td style="text-align:left">%store</td>
# <td style="text-align:left">IPass variables between notebooks.</td>
# </tr>
# <tr>
# <td style="text-align:left">!ls</td>
# <td style="text-align:left">Executing Shell Commands (ex: ls)</td>
# </tr>
# <tr>
# <td style="text-align:left">%%py37</td>
# <td style="text-align:left">Run code from a different kernel (ex: py37)</td>
# </tr>
#
#
# </table><br>
# - Profiling:
#
# <table style="margin: 0 auto">
# <tr>
# <td style="text-align:left">%time</td>
# <td style="text-align:left">Time a single statement.</td>
# </tr>
# <tr>
# <td style="text-align:left">%timeit</td>
# <td style="text-align:left">Run a statement multiple times to get an average runtime.</td>
# </tr>
# <tr>
# <td style="text-align:left">%%timeit </td>
# <td style="text-align:left">Apply timeit to the full cell.</td>
# </tr>
# <tr>
# <td style="text-align:left">%lprun -f</td>
# <td style="text-align:left">for df.apply(lambda row: haversine(40.671, -73.985, row['latitude'], row['longitude']), axis=1) <br> %lprun -f haversine df.apply(lambda row: haversine(40.671, -73.985, row['latitude'], row['longitude']), axis=1)</td>
# </tr>
# </table> <br>
# - Jupyter (IPYTHON)
#
# <br>
# <table style="margin: 0 auto">
# <tr>
# <th colspan="2" align="centre"> Command Mode (press Esc to enable) <th>
# <th colspan="2" align="centre"> Edit Mode (press Enter to enable) <th>
# </tr>
# <tr>
# <td style="text-align:left">Enter</td>
# <td style="text-align:left">enter edit mode</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Tab</td>
# <td style="text-align:left">code completion or indent</td>
# </tr>
# <tr>
# <td style="text-align:left">Ctrl-Enter</td>
# <td style="text-align:left">run cell, select below</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Shift-Tab</td>
# <td style="text-align:left">tooltip</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Ctrl-Enter</td>
# <td style="text-align:left">run cell</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-[</td>
# <td style="text-align:left">Indent</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Alt-Enter</td>
# <td style="text-align:left">run cell, insert below</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-]</td>
# <td style="text-align:left">Dedent</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Y</td>
# <td style="text-align:left">to code</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-A</td>
# <td style="text-align:left">select all</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">M</td>
# <td style="text-align:left">to markdown</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-Z</td>
# <td style="text-align:left">undo</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">A/B</td>
# <td style="text-align:left">insert cell above/below</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-Y</td>
# <td style="text-align:left">redo</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">X</td>
# <td style="text-align:left">cut selected cell</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-Home</td>
# <td style="text-align:left">go to cell start</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">C</td>
# <td style="text-align:left">copy selected cell</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-Home / Ctrl-Up</td>
# <td style="text-align:left">go to cell start</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Shift-V / V</td>
# <td style="text-align:left">paste cell above/below</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-End / Ctrl-Down</td>
# <td style="text-align:left">go to cell end</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Z</td>
# <td style="text-align:left">undo last cell deletion</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">ESC</td>
# <td style="text-align:left">command mode</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Ctrl-S</td>
# <td style="text-align:left">Save and Checkpoint</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-M</td>
# <td style="text-align:left">command mode</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">L</td>
# <td style="text-align:left">toggle line numbers</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-Shift-Subtract</td>
# <td style="text-align:left">split cell</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">O</td>
# <td style="text-align:left">toggle output</td>
# <td style="text-align:left"></td>
# <td style="text-align:left">Ctrl-Shift-- </td>
# <td style="text-align:left">split cell</td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Shift-O</td>
# <td style="text-align:left">toggle output scrolling</td>
# <td style="text-align:left"></td>
# <td style="text-align:left"></td>
# <td style="text-align:left"></td>
# </tr>
# <tr align="left">
# <td style="text-align:left">Space / Shift-space</td>
# <td style="text-align:left">scroll down/up</td>
# <td style="text-align:left"></td>
# <td style="text-align:left"></td>
# <td style="text-align:left"></td>
# </tr>
# <tr align="left">
# <td style="text-align:left">1/2/3/...</td>
# <td style="text-align:left">to heading 1/2/3...</td>
# <td style="text-align:left"></td>
# <td style="text-align:left"></td>
# <td style="text-align:left"></td>
# </tr>
# </table><br>
# - Access the Docstring: command?<br>
# `str.replace?` <br><br>
#
# - Access the code: command??<br>
# `import numpy` <br>
# `numpy??` <br>
#
# - Links a terminal to the current workspace of a notebook: <br>
# `jupyter console --existing`
# - Markdown: [jupyter markdown](https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html), [markdown guide](https://www.markdownguide.org/basic-syntax/) or [cheatsheet](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet)<br>
# +
# To display the output of multiple variables at once:
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# Or modify ~/.ipython/profile_default/ipython_config.py with the lines:
# c = get_config()
# # Run all nodes interactively
# c.InteractiveShell.ast_node_interactivity = "all"
# -
| Notebooks/01_Jupyter_Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="KimMZUVqcJ8_"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="BRQ6HQ8zcV5v"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="BlWzg1D9_EhW"
# # Inspecting Quantization Errors with Quantization Debugger
# + [markdown] id="XLoHL19yb-a0"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/lite/performance/quantization_debugger"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/performance/quantization_debugger.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/performance/quantization_debugger.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/tensorflow/tensorflow/lite/g3doc/performance/quantization_debugger.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# <td>
# <a href="https://tfhub.dev/google/imagenet/mobilenet_v3_small_100_224/classification/5"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
# </td>
# </table>
# + [markdown] id="MWO_yYDGcGWY"
# Although full-integer quantization provides improved model size and latency, the
# quantized model won't always work as expected. It's usually expected for the
# model quality (e.g. accuracy, mAP, WER) to be slightly lower than the original
# float model. However, there are cases where the model quality can go below your
# expectation or generated completely wrong results.
#
# When this problem happens, it's tricky and painful to spot the root cause of the
# quantization error, and it's even more difficult to fix the quantization error.
# To assist this model inspection process, **quantization debugger** can be used
# to identify problematic layers, and **selective quantization** can leave those
# problematic layers in float so that the model accuracy can be recovered at the
# cost of reduced benefit from quantization.
#
# Note: This API is experimental, and there might be breaking changes in the API
# in the course of improvements.
# + [markdown] id="9kD29R1I_Mn6"
# ## Quantization Debugger
#
# Quantization debugger makes it possible to do quantization quality metric
# analysis in the existing model. Quantization debugger can automate processes for
# running model with a debug dataset, and collecting quantization quality metrics
# for each tensors.
#
# Note: Quantization debugger and selective quantization currently only works for
# full-integer quantization with int8 activations.
# + [markdown] id="221Qon7G_PmZ"
# ### Prerequisites
#
# If you already have a pipeline to quantize a model, you have all necessary
# pieces to run quantization debugger!
#
# * Model to quantize
# * Representative dataset
#
# In addition to model and data, you will need to use a data processing framework
# (e.g. pandas, Google Sheets) to analyze the exported results.
# + [markdown] id="qTEEzJWo_iZ_"
# ### Setup
#
# This section prepares libraries, MobileNet v3 model, and test dataset of 100
# images.
# + id="l7epUDUP_6qo"
# Quantization debugger is available from TensorFlow 2.7.0
# !pip uninstall -y tensorflow
# !pip install tf-nightly
# + id="LLsgiUZe_hIa"
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_hub as hub
# + cellView="form" id="veWjO3u32vzz"
#@title Boilerplates and helpers
MODEL_URI = 'https://tfhub.dev/google/imagenet/mobilenet_v3_small_100_224/classification/5'
def process_image(data):
data['image'] = tf.image.resize(data['image'], (224, 224)) / 255.0
return data
# Representative dataset
def representative_dataset(dataset):
def _data_gen():
for data in dataset.batch(1):
yield [data['image']]
return _data_gen
def eval_tflite(tflite_model, dataset):
"""Evaluates tensorflow lite classification model with the given dataset."""
interpreter = tf.lite.Interpreter(model_content=tflite_model)
interpreter.allocate_tensors()
input_idx = interpreter.get_input_details()[0]['index']
output_idx = interpreter.get_output_details()[0]['index']
results = []
for data in representative_dataset(dataset)():
interpreter.set_tensor(input_idx, data[0])
interpreter.invoke()
results.append(interpreter.get_tensor(output_idx).flatten())
results = np.array(results)
gt_labels = np.array(list(dataset.map(lambda data: data['label'] + 1)))
accuracy = (
np.sum(np.argsort(results, axis=1)[:, -5:] == gt_labels.reshape(-1, 1)) /
gt_labels.size)
print(f'Top-5 accuracy (quantized): {accuracy * 100:.2f}%')
model = tf.keras.Sequential([hub.KerasLayer(MODEL_URI)])
model.compile(
loss='sparse_categorical_crossentropy',
metrics='sparse_top_k_categorical_accuracy')
model.build([1, 224, 224, 3])
# Prepare dataset with 100 examples
ds = tfds.load('imagenet_v2', split='test[:1%]')
ds = ds.map(process_image)
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.representative_dataset = representative_dataset(ds)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
quantized_model = converter.convert()
# + id="7mX-R-xK4ADB"
test_ds = ds.map(lambda data: (data['image'], data['label'] + 1)).batch(16)
loss, acc = model.evaluate(test_ds)
print(f'Top-5 accuracy (float): {acc * 100:.2f}%')
# + id="Mnp6yBnJSCoh"
eval_tflite(quantized_model, ds)
# + [markdown] id="Tblkk3cxxpuw"
# We can see that the original model has 82% top-5 accuracy for our small dataset,
# while the quantized model shows 54% top-5 accuracy, which indicates a
# significant accuracy loss.
# + [markdown] id="dBBcfCQw_Wqd"
# ### Step 1. Debugger preparation
#
# Easiest way to use the quantization debugger is to provide
# `tf.lite.TFLiteConverter` that you have been using to quantize the model.
# + id="NOByihbD_NZZ"
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter.representative_dataset = representative_dataset(ds)
# my_debug_dataset should have the same format as my_representative_dataset
debugger = tf.lite.experimental.QuantizationDebugger(
converter=converter, debug_dataset=representative_dataset(ds))
# + [markdown] id="9vR1IIrmQS9W"
# ### Step 2. Running the debugger and getting the results
#
# When you call `QuantizationDebugger.run()`, the debugger will log differences
# between float tensors and quantized tensors for the same op location, and
# process them with given metrics.
# + id="HsUM54g-_E52"
debugger.run()
# + [markdown] id="yQpX_SBUQXvr"
# The processed metrics can be accessed with
# `QuantizationDebugger.layer_statistics`, or can be dumped to a text file in CSV
# format with `QuantizationDebugger.layer_statistics_dump()`.
# + id="U-AGYUAbQUmx"
RESULTS_FILE = '/tmp/debugger_results.csv'
with open(RESULTS_FILE, 'w') as f:
debugger.layer_statistics_dump(f)
# + id="LQzEi6VnQaen"
# !head /tmp/debugger_results.csv
# + [markdown] id="4np7VqU-Qfke"
# For each row in the dump, the op name and index comes first, followed by
# quantization parameters and error metrics (including
# [user-defined error metrics](#custom-metrics), if any). The resulting CSV file
# can be used to pick problematic layers with large quantization error metrics.
#
# With pandas or other data processing libraries, we can inspect detailed
# per-layer error metrics.
# + id="XUcSqYFGQb-f"
layer_stats = pd.read_csv(RESULTS_FILE)
layer_stats.head()
# + [markdown] id="7C_oHxWFOV6M"
# ### Step 3. Data analysis
#
# There are various ways to analyze the resulting. First, let's add some useful
# metrics derived from the debugger's outputs. (`scale` means the quantization
# scale factor for each tensor.)
#
# * Range (`256 / scale`)
# * RMSE / scale (`sqrt(mean_squared_error) / scale`)
#
# The `RMSE / scale` is close to `1 / sqrt(12)` (~ 0.289) when quantized
# distribution is similar to the original float distribution, indicating a good
# quantized model. The larger the value is, it's more likely for the layer not
# being quantized well.
# + id="mwviORyJN6e5"
layer_stats['range'] = 255.0 * layer_stats['scale']
layer_stats['rmse/scale'] = layer_stats.apply(
lambda row: np.sqrt(row['mean_squared_error']) / row['scale'], axis=1)
layer_stats[['op_name', 'range', 'rmse/scale']].head()
# + id="oAAv35CdPvc4"
plt.figure(figsize=(15, 5))
ax1 = plt.subplot(121)
ax1.bar(np.arange(len(layer_stats)), layer_stats['range'])
ax1.set_ylabel('range')
ax2 = plt.subplot(122)
ax2.bar(np.arange(len(layer_stats)), layer_stats['rmse/scale'])
ax2.set_ylabel('rmse/scale')
plt.show()
# + [markdown] id="8pqUQvRUWB3Q"
# There are many layers with wide ranges, and some layers that have high
# `RMSE/scale` values. Let's get the layers with high error metrics.
# + id="UqFsUX4_Q-cE"
layer_stats[layer_stats['rmse/scale'] > 0.7][[
'op_name', 'range', 'rmse/scale', 'tensor_name'
]]
# + [markdown] id="DHeALFTGWl_e"
# With these layers, you can try selective quantization to see if not quantizing
# those layers improves model quality.
# + id="cvdkjsbwYC6e"
suspected_layers = list(
layer_stats[layer_stats['rmse/scale'] > 0.7]['tensor_name'])
# + [markdown] id="W6RQw9JobOTR"
# In addition to these, skipping quantization for the first few layers also helps
# improving quantized model's quality.
# + id="ikF2bp6NZcXN"
suspected_layers.extend(list(layer_stats[:5]['tensor_name']))
# + [markdown] id="1DfT78w6W6Li"
# ## Selective Quantization
# + [markdown] id="-pubC-01cGEH"
# Selective quantization skips quantization for some nodes, so that the
# calculation can happen in the original floating-point domain. When correct
# layers are skipped, we can expect some model quality recovery at the cost of
# increased latency and model size.
#
# However, if you're planning to run quantized models on integer-only accelerators
# (e.g. Hexagon DSP, EdgeTPU), selective quantization would cause fragmentation of
# the model and would result in slower inference latency mainly caused by data
# transfer cost between CPU and those accelerators. To prevent this, you can
# consider running
# [quantization aware training](https://www.tensorflow.org/model_optimization/guide/quantization/training)
# to keep all the layers in integer while preserving the model accuracy.
# + [markdown] id="EQFBfR7YW-oh"
# Quantization debugger's option accepts `denylisted_nodes` and `denylisted_ops`
# options for skipping quantization for specific layers, or all instances of
# specific ops. Using `suspected_layers` we prepared from the previous step, we
# can use quantization debugger to get a selectively quantized model.
# + id="K5KD0JAEbpsv"
debug_options = tf.lite.experimental.QuantizationDebugOptions(
denylisted_nodes=suspected_layers)
debugger = tf.lite.experimental.QuantizationDebugger(
converter=converter,
debug_dataset=representative_dataset(ds),
debug_options=debug_options)
# + id="pfj9gzv4b7h4"
selective_quantized_model = debugger.get_nondebug_quantized_model()
eval_tflite(selective_quantized_model, ds)
# + [markdown] id="1RkfMYSHdtZy"
# The accuracy is still lower compared to 84% of the original float model, but we
# have 12pp improvement from the whole quantized model by skipping quantization
# for ~10 layers out of 111 layers.
#
# You can also try to not quantized all ops in the same class. For example, to
# skip quantization for all mean ops, you can pass `MEAN` to `denylisted_ops`.
# + id="ruUoP7SgcLpO"
debug_options = tf.lite.experimental.QuantizationDebugOptions(
denylisted_ops=['MEAN'])
debugger = tf.lite.experimental.QuantizationDebugger(
converter=converter,
debug_dataset=representative_dataset(ds),
debug_options=debug_options)
# + id="oY6kb5g_cO4H"
selective_quantized_model = debugger.get_nondebug_quantized_model()
eval_tflite(selective_quantized_model, ds)
# + [markdown] id="xa8488TeAyx-"
# With these techniques, we were able to improve the quantized MobileNet V3 model
# accuracy easily by 12pp. Next, we'll explore the advanced techniques to further
# debug and improve the model accuracy
# + [markdown] id="ZD75cY9PUb2u"
# ## Advanced usages
#
# Whith following features, you can futher customize your debugging pipeline.
# + [markdown] id="aVj9yrQoUfGo"
# ### Custom metrics
#
# By default, the quantization debugger emits five metrics for each float-quant
# difference: tensor size, standard deviation, mean error, max absolute error, and
# mean squared error. You can add more custom metrics by passing them to options.
# For each metrics, the result should be a single float value and the resulting
# metric will be an average of metrics from all examples.
#
# * `layer_debug_metrics`: calculate metric based on diff for each op outputs
# from float and quantized op outputs.
# * `layer_direct_compare_metrics`: rather than getting diff only, this will
# calculate metric based on raw float and quantized tensors, and its
# quantization parameters (scale, zero point)
# * `model_debug_metrics`: **only used when `float_model_(path|content)` is
# passed** to the debugger. In addition to the op-level metrics, final layer
# output is compared to the reference output from the original float model.
# + id="WqmRQSxoVVwu"
debug_options = tf.lite.experimental.QuantizationDebugOptions(
layer_debug_metrics={
'mean_abs_error': (lambda diff: np.mean(np.abs(diff)))
},
layer_direct_compare_metrics={
'correlation':
lambda f, q, s, zp: (np.corrcoef(f.flatten(),
(q.flatten() - zp) / s)[0, 1])
},
model_debug_metrics={
'argmax_accuracy': (lambda f, q: np.mean(np.argmax(f) == np.argmax(q)))
})
debugger = tf.lite.experimental.QuantizationDebugger(
converter=converter,
debug_dataset=representative_dataset(ds),
debug_options=debug_options)
# + id="PVQ4nEicXz2l"
debugger.run()
# + id="dfKA90csX9UL"
CUSTOM_RESULTS_FILE = '/tmp/debugger_results.csv'
with open(CUSTOM_RESULTS_FILE, 'w') as f:
debugger.layer_statistics_dump(f)
custom_layer_stats = pd.read_csv(CUSTOM_RESULTS_FILE)
custom_layer_stats[['op_name', 'mean_abs_error', 'correlation']].tail()
# + [markdown] id="Qqq30oWsZF5b"
# The result of `model_debug_metrics` can be separately seen from
# `debugger.model_statistics`.
# + id="wrXlmzEHYhQ5"
debugger.model_statistics
# + [markdown] id="DqJBLIsoUyIg"
# ### Using (internal) mlir_quantize API to access in-depth features
#
# Note: Some features in the folowing section,
# `TFLiteConverter._experimental_calibrate_only` and `converter.mlir_quantize` are
# experimental internal APIs, and subject to change in a non-backward compatible
# way.
# + id="VJm66Cz-XpeF"
from tensorflow.lite.python import convert
# + [markdown] id="2krUVzpiUp3u"
# #### Whole model verify mode
#
# The default behavior for the debug model generation is per-layer verify. In this
# mode, the input for float and quantize op pair is from the same source (previous
# quantized op). Another mode is whole-model verify, where the float and quantize
# models are separated. This mode would be useful to observe how the error is
# being propagated down the model. To enable, `enable_whole_model_verify=True` to
# `convert.mlir_quantize` while generating the debug model manually.
# + id="5zykINDlVLSg"
converter = tf.lite.TFLiteConverter.from_keras_model(model)
converter.representative_dataset = representative_dataset(ds)
converter.optimizations = [tf.lite.Optimize.DEFAULT]
converter._experimental_calibrate_only = True
calibrated_model = converter.convert()
# + id="eqvXlEiFXfSu"
# Note that enable_numeric_verify and enable_whole_model_verify are set.
quantized_model = convert.mlir_quantize(
calibrated_model,
enable_numeric_verify=True,
enable_whole_model_verify=True)
debugger = tf.lite.experimental.QuantizationDebugger(
quant_debug_model_content=quantized_model,
debug_dataset=representative_dataset(ds))
# + [markdown] id="xQ6TFsXQVHMe"
# #### Selective quantization from an already calibrated model
#
# You can directly call `convert.mlir_quantize` to get the selective quantized
# model from already calibrated model. This would be particularly useful when you
# want to calibrate the model once, and experiment with various denylist
# combinations.
# + id="ZCS-Fa9lbdc0"
selective_quantized_model = convert.mlir_quantize(
calibrated_model, denylisted_nodes=suspected_layers)
eval_tflite(selective_quantized_model, ds)
| tensorflow/lite/g3doc/performance/quantization_debugger.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep learning 2. Convolutional Neural Networks
# - The dense FFN used before contained 600000 parameters. These are expensive to train!
# - A picture is not a flat array of numbers, it is a 2D matrix with multiple color channels. Convolution kernels are able to find 2D hidden features.
# - A CNN's layers are operating on a 3D tensor of shape (height, width, channels). A colored imaged usually has three channels (RGB) but more are possible. We have grayscale thus only one channel.
# - The width and height dimensions tend to shrink as we go deeper in the network. Why? NNs are efective information filters.
#
# Why is this important for a biologist?
# - Sight is our main sense. Labelling pictures is much easier than other types of data!
# - Most biological data can be converted to image format. (including genomics, transcriptomics, etc)
# - Spatial transcriptomics, as well as some single cell data have multi-channel and spatial features.
# - Microscopy is biology too!
# +
from tensorflow.keras.datasets import mnist
from tensorflow.keras.utils import to_categorical
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images.reshape((60000, 28, 28, 1))
train_images = train_images.astype('float32') / 255
test_images = test_images.reshape((10000, 28, 28, 1))
test_images = test_images.astype('float32') / 255
train_labels = to_categorical(train_labels)
test_labels = to_categorical(test_labels)
# -
print(train_images.shape)
# ### Method:
#
# - The convolutional network will filter the image in a sequence, gradually expanding the complexity of hidden features and eliminating the noise via the "downsampling bottleneck".
# - A CNN's filtering principle is based on the idea of functional convolution, this is a mathematical way of comparing two functions in a temporal manner by sliding one over the other.
# - Parts: convolution, pooling and classification
from IPython.display import Image
Image(url= "../img/cnn.png", width=400, height=400)
Image(url= "../img/convolution.png", width=400, height=400)
Image(url= "../img/pooling.png", width=400, height=400)
# The layers:
# - The first block: 32 number of kernels (convolutional filters) each of 3 x 3 size followed by a max pooling operation with pool size of 2 x 2.
# - The second block: 64 number of kernels each of 3 x 3 size followed by a max pooling operation with pool size of 2 x 2 and a dropout of 20% to ensure the regularization and thus avoiding overfitting of the model.
# - classification block: flattening operation which transforms the data to 1 dimensional so as to feed it to fully connected or dense layer. The first dense layer consists of 128 neurons with relu activation while the final output layer consist of 10 neurons with softmax activation which will output the probability for each of the 10 classes.
#
# +
#from tensorflow.keras import layers
from tensorflow.keras.layers import Conv2D, MaxPooling2D, Dropout, Flatten, Dense
from tensorflow.keras import models
model = models.Sequential()
# first block
model.add(Conv2D(32, kernel_size=(3, 3), input_shape=(28, 28, 1), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
# second block
model.add(Conv2D(64, kernel_size=(3,3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.2))
# flattening followed by dense layer and final output layer
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dense(10,activation='softmax'))
model.summary()
# -
# - Loss:
# - Many functions are possible (mean square error, maximum likelihood)
# - cross-entropy loss (or log loss): sum for all predicted classes $- \sum_c y_c log(p_c)$, where $y_c$ is a binary inndication of classification success and $p_c$ is the probability value of the model prediction
#
#
# - Optimizers:
# - SGD: slower, classic, can get stuck in local minima, uses momentum to avoid small valleys
# - rmsprop (root mean square propagation): batches contain bias noise, weights are adjusted ortogonal to bias, leading to faster convergence.
# - Adam: combines the above
# - Read more at: https://medium.com/analytics-vidhya/momentum-rmsprop-and-adam-optimizer-5769721b4b19
#
#
# - Learning rate ($\alpha$): Gradient descent algorithms multiply the magnitude of the gradient (the rate of error change with respect to each weight) by a scalar known as learning rate (also sometimes called step size) to determine the next point. $w_{ij} = w_{ij} + \alpha \frac{dE}{dw_{ij}}$
#
# +
from tensorflow.keras.optimizers import Adam
# compiling the model
model.compile(loss='categorical_crossentropy', optimizer=Adam(lr=0.01), metrics=['accuracy'])
# train the model training dataset
history = model.fit(train_images, train_labels, epochs=5, validation_data=(test_images, test_labels), batch_size=128)
# save the model
model.save('cnn.h5')
# -
test_loss, test_acc = model.evaluate(test_images, test_labels)
print(test_loss, test_acc)
# %matplotlib inline
import matplotlib.pyplot as plt
print(history.history.keys())
# Ploting the accuracy graph
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='best')
plt.show()
# Ploting the loss graph
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'val'], loc='best')
plt.show()
# ### Good fit vs overfitting and underfitting
#
# - Good fit:
# - Training loss and Validation loss are close to each other with validation loss being slightly greater than the training loss.
# - Initially decreasing training and validation loss and a pretty flat training and validation loss after some point till the end.
#
#
# - Overfitting: Overfit models are similar to school kids that spend so much time learning that they are not able to generalize their knowledge, or hyperspecialists that cannot think outside their fields. If a model is over trained on the data it typically shows a perfect training set score and a gradually worsening test score. This is because it learned so much the training data noise that it fails to spot the signal in the test data. Can you spot above when the model overfits?
# - How to avoid: Don't use complex models for small simple datasets. Regularization.
# - How to spot:
# - Training loss and Validation loss are far away from each other.
# - Gradually decreasing validation loss (without flattening) upon adding training examples.
# - Very low training loss that’s very slightly increasing upon adding training examples.
#
#
# - Underfitting: The model doesn't learn from the data, giving a low score on both the training set and test/validation set.
# - How to avoid: Don't use simple models for a complex problem. Clean the noise from the dataset so that the signal is easier to capture by the model. (It may also be that the signal is missing or is misslabeled)
# - How to spot:
# - Increasing training loss upon adding training examples.
# - Training loss and validation loss are close to each other at the end.
# - Sudden dip in the training loss and validation loss at the end (not always).
#
#
# - This is a general ML problem, read here how you can use scikit learn to evaluare the loss/acc curves for any iterative learning algorithm:
# https://towardsdatascience.com/learning-curve-to-identify-overfitting-underfitting-problems-133177f38df5
#
#
# ### How do we improve model convergence?
#
# - Epochs: increase the number until the validation accuracy starts to decrease, even when the accuracy of the training data continues to increase (this is when we detect a potential overfitting).
# - Batches: A small number of samples per batch will avoid RAM issues.
# - Learning rate: if too big, the algorithm might converge fast to a local minimum, or bounce on the gradient well. If too small the convergence will be slower but better.
# - Choice of optimization methods and their parameters. (there are drawbacks to all)
# - Hyperparametrization. Note that there are also model specific hyper-parameters. Can you find them in this case (CNN)?
#
# Further read:
# - https://towardsdatascience.com/learning-process-of-a-deep-neural-network-5a9768d7a651
#
# **Task:** now retrain the model with a batch size of 32, 10 epochs, and no learning rate constraints (use defaults). Notice improvements in model fitting?
#
# ### Regularization
#
# - Large datasets cause overfitting! A common reason is having large weights in the network. They can result in small changes in the input having drastic effects in the output. More generally, you want to insure that your model is able to keep up with a certain level of input noise.
# - Regularization is a general ML problem, but in the NN context the main approaches are batch normalization and dropout layers.
#
# **Batch normalization**: normalize the inputs of each layer in such a way that, they have a mean activation output zero and a unit standard deviation.
# - This reduces the eliptical curvature of the error surface, helping gradient descent.
# - Since the normalization is only done batch wise, the normalization layers will still feed regular noise in the model!
#
# **Dropout**
# - By popular vote: A healthy mind is a mind that can forget.
# - A single model can be used to simulate having a large number of different network architectures by randomly dropping out nodes during training.
# - This makes the neurons less reliant on small perturbations from other neurons.
#
# **Task:**
# - apply a batch normalization layer
# - parametrise the dropout rate and measure loss.
#
#
# Further read:
# - https://medium.com/analytics-vidhya/everything-you-need-to-know-about-regularizer-eb477b0c82ba
# - https://www.analyticsvidhya.com/blog/2021/03/introduction-to-batch-normalization/
| day3/DL2_CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stochastic Volatility model
# +
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
np.random.seed(0)
# -
plt.rcParams.update({
"axes.prop_cycle": plt.cycler("color", ['#000000', '#1b6989', '#e69f00', '#009e73', '#f0e442', '#50b4e9', '#d55e00', '#cc79a7']),
"font.serif": ['Palatino',
'Palatino Linotype',
'Palatino LT STD',
'Book Antiqua',
'Georgia',
'DejaVu Serif'],
'font.family': 'serif',
'figure.facecolor': '#fffff8',
'axes.facecolor': '#fffff8',
'figure.constrained_layout.use': True,
'font.size': 14.0,
'hist.bins': 'auto',
'lines.linewidth': 1.0,
})
# Asset prices have time-varying volatility (variance of day over day `returns`). In some periods, returns are highly variable, while in others very stable. Stochastic volatility models model this with a latent volatility variable, modeled as a stochastic process. The following model is similar to the one described in the No-U-Turn Sampler paper, Hoffman (2011) p21.
#
# $$ \sigma \sim Exponential(50) $$
#
# $$ \nu \sim Exponential(.1) $$
#
# $$ s_i \sim Normal(s_{i-1}, \sigma^{-2}) $$
#
# $$ log(r_i) \sim t(\nu, 0, exp(-2 s_i)) $$
#
# Here, $r$ is the daily return series and $s$ is the latent log volatility process.
# ## Build Model
# First we load daily returns of the S&P 500, and calculate the daily log returns. This data is from May 2008 to November 2019.
returns = pd.read_csv(pm.get_data("SP500.csv"), index_col='Date')
returns["change"] = np.log(returns["Close"]).diff()
returns = returns.dropna()
returns.head()
# As you can see, the volatility seems to change over time quite a bit but cluster around certain time-periods. For example, the 2008 financial crisis is easy to pick out.
fig, ax = plt.subplots(figsize=(14, 4))
returns.plot(y="change", label='S&P 500', ax=ax)
ax.set(xlabel='time', ylabel='returns')
ax.legend();
# Specifying the model in `PyMC3` mirrors its statistical specification.
# +
def make_stochastic_volatility_model(data):
with pm.Model() as model:
step_size = pm.Exponential('step_size', 10)
volatility = pm.GaussianRandomWalk('volatility', sigma=step_size, shape=len(data))
nu = pm.Exponential('nu', 0.1)
returns = pm.StudentT('returns',
nu=nu,
lam=np.exp(-2*volatility),
observed=data["change"])
return model
stochastic_vol_model = make_stochastic_volatility_model(returns)
# -
# ## Checking the model
#
# Two good things to do to make sure our model is what we expect is to
# 1. Take a look at the model structure. This lets us know we specified the priors we wanted and the connections we wanted. It is also handy to remind ourselves of the size of the random variables.
# 2. Take a look at the prior predictive samples. This helps us interpret what our priors imply about the data.
#
pm.model_to_graphviz(stochastic_vol_model)
with stochastic_vol_model:
prior = pm.sample_prior_predictive(500)
# We plot and inspect the prior predictive. This is *many* orders of magnitude larger than the actual returns we observed. In fact, I cherry-picked a few draws to keep the plot from looking silly. This may suggest changing our priors: a return that our model considers plausible would violate all sorts of constraints by a huge margin: the total value of all goods and services the world produces is ~$\$10^9$, so we might reasonably *not* expect any returns above that magnitude.
#
# That said, we get somewhat reasonable results fitting this model anyways, and it is standard, so we leave it as is.
# +
fig, ax = plt.subplots(figsize=(14, 4))
returns['change'].plot(ax=ax, lw=1, color='black')
ax.plot(prior['returns'][4:6].T, 'g', alpha=0.5, lw=1, zorder=-10)
max_observed, max_simulated = np.max(np.abs(returns['change'])), np.max(np.abs(prior['returns']))
ax.set_title(f"Maximum observed: {max_observed:.2g}\nMaximum simulated: {max_simulated:.2g}(!)");
# -
# ## Fit Model
# Once we are happy with our model, we can sample from the posterior. This is a somewhat tricky model to fit even with NUTS, so we sample and tune a little longer than default.
with stochastic_vol_model:
trace = pm.sample(2000, tune=2000)
with stochastic_vol_model:
posterior_predictive = pm.sample_posterior_predictive(trace)
# Note that the `step_size` parameter does not look perfect: the different chains look somewhat different. This again indicates some weakness in our model: it may make sense to allow the step_size to change over time, especially over this 11 year time span.
pm.traceplot(trace, var_names=['step_size', 'nu']);
# Now we can look at our posterior estimates of the volatility in S&P 500 returns over time.
# +
fig, ax = plt.subplots(figsize=(14, 4))
y_vals = np.exp(trace['volatility'])[::5].T
x_vals = np.vstack([returns.index for _ in y_vals.T]).T.astype(np.datetime64)
plt.plot(x_vals, y_vals, 'k', alpha=0.002)
ax.set_xlim(x_vals.min(), x_vals.max())
ax.set_ylim(bottom=0)
ax.set(title='Estimated volatility over time', xlabel='Date', ylabel='Volatility');
# -
# Finally, we can use the posterior predictive distribution to see the how the learned volatility could have effected returns.
# +
fig, axes = plt.subplots(nrows=2, figsize=(14, 7), sharex=True)
returns['change'].plot(ax=axes[0], color='black')
axes[1].plot(np.exp(trace['volatility'][::100].T), 'r', alpha=0.5)
axes[0].plot(posterior_predictive['returns'][::100].T, 'g', alpha=0.5, zorder=-10)
axes[0].set_title("True log returns (black) and posterior predictive log returns (green)")
axes[1].set_title("Posterior volatility");
# -
# ## References
#
# 1. Hoffman & Gelman. (2011). [The No-U-Turn Sampler: Adaptively Setting Path Lengths in Hamiltonian Monte Carlo](http://arxiv.org/abs/1111.4246).
| docs/source/notebooks/stochastic_volatility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 2.3节 交叉验证
import numpy as np
from sklearn import datasets # 导入数据集
from sklearn.neighbors import KNeighborsClassifier # KNN分类算法
from sklearn.model_selection import KFold # 用于交叉验证
# 导入iris数据集
iris = datasets.load_iris()
X = iris.data
y = iris.target
print(X.shape, y.shape)
# +
# 定义8个不同的k值
ks = [1, 3, 5, 7, 9, 11, 13, 15]
# 5折交叉验证
kf = KFold(n_splits=5, random_state=2001, shuffle=True)
print(kf)
# -
| 02_knn/knn_check.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:darknet_env]
# language: python
# name: conda-env-darknet_env-py
# ---
# # XML to YOLOv3 label Pharser
# ## Created by <NAME> on 13/8/19
#
# Load the xml file
import xml.etree.ElementTree as ET
root = ET.parse('gopro_008.xml').getroot()
# get BBox info (example)
print(root[1][0][1][0][0].attrib)
print(root[1][0][1][0][0].get('x'))
# +
# BONUS: Count video frames
import cv2
video = cv2.VideoCapture('gopro_008.mp4')
total = 0
def count_frames_manual(video):
# initialize the total number of frames read
total = 0
# loop over the frames of the video
while True:
# grab the current frame
(grabbed, frame) = video.read()
# check to see if we have reached the end of the
# video
if not grabbed:
break
# increment the total number of frames read
total += 1
# return the total number of frames in the video file
return total
count_frames_manual(video)
# -
# creo i txt con le label (1 riga sola per ora) YOLO v3 format
import os
h_img = 1080
w_img = 1920
i = 0
for filename in os.listdir("output"):
if filename.endswith(".jpg"):
filePath = "output/" + filename[:-3] + "txt"
# creo il file del frame
f = open(filePath,"w+")
# le label sono dall angolo in alto a sx
# class x_center y_center W H
h = int(root[1][0][1][0][i].get('height'))
w = int(root[1][0][1][0][i].get('width'))
x = round(((int(root[1][0][1][0][i].get('x'))+(w/2))/1920),6)
y = round(((int(root[1][0][1][0][i].get('y'))+(h/2))/1080),6)
riga = "0 " + str(x) + " " + str(y) + " " +str(round(w/w_img,6)) + " " + str(round(h/h_img,6))
#print(riga)
f.write(riga) # scrivo la label nel file
f.close() # chiudiamo il file
i += 1
continue
else:
continue
print("done")
| XML_label_pharser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Z57Y5-H3uuOV" colab_type="text"
# #Combinatorial Optimization Problems on Variational Quantum Eigensolver(VQE) and QAOA
# + [markdown] id="23QCMzt_u3KT" colab_type="text"
# Combinatorial optimization problem is one of the usage of variational algorithm. Who want to start learning quantum-classical hybrid algorithm on quantum computing, it may be a good subject to strat.
#
# Combinatorial optimization problem is a problem to find the best solution by solving a minimized optimization problem. If you want to solve a social problem, to formulate this into a combination of binary number 0 and 1 and give some constraint on it.
# + [markdown] id="9Jbvpe9mu6qB" colab_type="text"
# ##Formulation of combinatorial optimization problem and VQE
# The formulation think the hamiltonian as a cost function for social problems and try to find the best answer from the qubits. The main step is,
#
# 1. Use ising model which is a basic physics model
# 2. Use combination of Z as a hamiltonian
# 3. More practically we use QUBO for the formulation which is 0 and 1 instead of -1 and +1 of ising model
#
# Usually we rarely use VQE without qaoa ansatz for the combinatorial optimization problem, but this time we just set a simple 1qubit problem.
#
# ```python
# h = -Z(0) - Z(0)*Z(1)
# ```
#
# The number after Z is the number of qubits. Here we are using 0th qubit and 1st qubit in the quantum circuit. and the coefficients in front of Z is important.
#
# The coefficient of Z(0) is a bias.
# The coefficient of Z(0)*Z(1) is a weight.
#
# Now we have both value of -1 as each coefficient.
#
# Z takes -1 or +1 as a expectation value. If the hamiltonian h takes smaller value it will be the answer.
#
# Here we just check the whole answer in the table.
#
# Z(0) | Z(1) | h
# --:|:----:|:--
# -1|-1|0
# -1|1|2
# 1|-1|0
# 1|1|-2
#
# The hamiltonian takes the minimu value of -2 when Z(0)=1 and Z(1)=1. VQE solve this automatically. This time as an ansatz we used 4 parameters of a,b,c,d. a, b for the 0th qubit, and c, d for the 1st qubit.
#
# Now we check it on blueqat, first we need to install blueqat and give this code.
# + id="opzc-Mvfu_gd" colab_type="code" outputId="79113e2c-7380-470a-e7b0-d7a06f95da3d" colab={"base_uri": "https://localhost:8080/", "height": 136}
# !pip3 install blueqat
# + id="qIc5WG5dumej" colab_type="code" outputId="69061199-2601-4f8d-ca3a-afcc65a354e6" colab={"base_uri": "https://localhost:8080/", "height": 51}
import numpy as np
from blueqat import Circuit
from blueqat.pauli import X, Y, Z, I
from blueqat.vqe import AnsatzBase, Vqe
class OneQubitAnsatz(AnsatzBase):
def __init__(self, hamiltonian):
super().__init__(hamiltonian.to_expr(), 4)
self.step = 1
def get_circuit(self, params):
a, b, c, d = params
return Circuit().ry(a)[0].rz(b)[0].ry(c)[1].rz(d)[1]
# hamiltonian is important
h = -Z(0) - Z(0)*Z(1)
runner = Vqe(OneQubitAnsatz(h))
result = runner.run()
print('Result by VQE')
print(runner.ansatz.get_energy(result.circuit, runner.sampler))
# + [markdown] id="6xNoXOE4vHUZ" colab_type="text"
# Now we get -2 as the expectation value of minimum value.
#
# For practical use we usually solve much bigger problems. Now we can see this problem as a kind of classification problem which bias of Z(0) shows that Alice belongs to the group1 and weight of Z(0)*Z(1) shows that Alice and Bob belong to the same group as a kind of max-cut problem.
#
# Here we used Z for operator, now we use a technique to transform it into 01 binary number.
#
# + [markdown] id="3DOTeXadvJzG" colab_type="text"
# ##Formulation with QUBO
# If use Z in the hamiltoinan Z takes -1 or +1 as an expectaion value. But it is little bit uncomfortable for the social problems. Usually we use 01 binary number so, we just transform hamiltonian into binary number using,
#
# $$
# q = \frac{Z + 1}{2}
# $$
#
# This is the transform of -1 to 0 and we now can use QUBO as a formulation.
# + [markdown] id="cvKkwYayvOga" colab_type="text"
# ##QUBO programming
# Let's use qubo on blueqat. Blueqat has a function for qubo. Now we can write the qubo form as,
#
# ```python
# h = -3*q(0)-3*q(1)-2*q(0)*q(1)
# ```
#
# This hamiltonian obviously takes the minimum value of -8 when q(0)=1 and q(1)=1. By solving on VQE we may get the similar result.
# + id="rkYdYTbWu-F4" colab_type="code" outputId="b3644963-3ce1-4900-e7a9-a60c44a3cb55" colab={"base_uri": "https://localhost:8080/", "height": 85}
import numpy as np
from blueqat import Circuit
from blueqat.pauli import X, Y, Z, I
from blueqat.pauli import qubo_bit as q
from blueqat.vqe import AnsatzBase, Vqe
class QubitAnsatz(AnsatzBase):
def __init__(self, hamiltonian):
super().__init__(hamiltonian, 4)
self.step = 1
def get_circuit(self, params):
a, b, c, d = params
return Circuit().ry(a)[0].rz(b)[0].ry(c)[1].rz(d)[1]
h = -3*q(0)-3*q(1)-2*q(0)*q(1)
h = h.to_expr().simplify()
runner = Vqe(QubitAnsatz(h))
result = runner.run()
print('Result by VQE')
print(runner.ansatz.get_energy(result.circuit, runner.sampler))
# Hamiltonian to matrix
mat = h.to_matrix()
# Calculate by numpy
print('Result by numpy')
print(np.linalg.eigh(mat)[0][0])
# + [markdown] id="DYsNMr6QvUnh" colab_type="text"
# We correctly the result. This time we just used 2qubits. But actually it is very difficult to get the right answer on bigger number of qubits. To compute much more efficiently we are using a new ansatz called QAOA ansatz for combinatorial optimization problem.
# + [markdown] id="etA_TEnsvWzL" colab_type="text"
# ##QAOA
# QAOA has a special form of ansatz especially to solve combinatorial optimization problem using the similar step of VQE. Now we try to use QAOA(Quantum Approximate Opitmization Alogirthm) in this chapter.
# + [markdown] id="9e4vdEK9vZq8" colab_type="text"
# ##2-5-1 Quantum Adiabatic Algorithm
# QAA is an algorithm to change the state vector adiabatically to keep the ground state of the hamiltonian through the time evolution.
#
# We set the initial hamiltonian as $H_{start}$ and the final Hamiltonian as $H_{fin}$. t is time and T is the whole step of schedule.
#
# $$
# H_{temp} = (1-\frac{t}{T})H_{start} + \frac{t}{T}H_{fin}
# $$
#
# The hamiltoinan adiabatically changed according to the time t and the eigenstate of state vector keep the ground state if T is enough big.
#
# $$
# H_{temp}\mid \psi \rangle = E_{0temp}\mid \psi \rangle
# $$
#
# Time evolution is,
#
# $$
# \mid \psi_{t+1} \rangle = U \mid \psi_t \rangle = e^{-iHt} \mid \psi_t \rangle
# $$
#
# + [markdown] id="FA9B9idIvgTH" colab_type="text"
# ##QAOA
# QAOA is basically using the idea of adiabatic process and transform it into a variational algorithm as ansatz of hamiltonian.
#
# 
# https://app.quantumcomputing.com/
#
# The first H on the circuit shows the initial eigen state of hamiltonian X. CX-Rz-CX shows the weight of hamiltonian and Rz shows the bias of the hamiltonian. Rx is a time evolution of initial hamiltonian X.
#
# Let's just see the inside of QaoaAnsatz of blueqat. Hamiltonian consists of Z operator and automatically make a variational transformation of time evolution ansatz.
#
#
# ```python
# class QaoaAnsatz(AnsatzBase):
# def __init__(self, hamiltonian, step=1, init_circuit=None):
# super().__init__(hamiltonian, step * 2)
# self.hamiltonian = hamiltonian.to_expr().simplify()
# if not self.check_hamiltonian():
# raise ValueError("Hamiltonian terms are not commutable")
#
# self.step = step
# self.n_qubits = self.hamiltonian.max_n() + 1
# if init_circuit:
# self.init_circuit = init_circuit
# if init_circuit.n_qubits > self.n_qubits:
# self.n_qubits = init_circuit.n_qubits
# else:
# self.init_circuit = Circuit(self.n_qubits).h[:]
# self.init_circuit.make_cache()
# self.time_evolutions = [term.get_time_evolution() for term in self.hamiltonian]
#
# def check_hamiltonian(self):
# """Check hamiltonian is commutable. This condition is required for QaoaAnsatz"""
# return self.hamiltonian.is_all_terms_commutable()
#
# def get_circuit(self, params):
# c = self.init_circuit.copy()
# betas = params[:self.step]
# gammas = params[self.step:]
# for beta, gamma in zip(betas, gammas):
# beta *= np.pi
# gamma *= 2 * np.pi
# for evo in self.time_evolutions:
# evo(c, gamma)
# c.rx(beta)[:]
# return c
# ```
#
# On actual use, the library automatically do the calculation and you don't to implement a complicated formulation or time evolution.
#
# Blueqat do most of the process and what you just need to do is to formulate the hamiltonian as a combinatorial optimization with binary number of 0 and 1.
#
# Now we see an example of simple problem on qubo.
#
# ```python
# cost = -3*q(0)-3*q(1)-2*q(0)*q(1)
# ```
#
# This take obviously -8 as a result. It is very easy, now we can think about the social problem as binary number. q(0) and q(1) has -3 of bias and q(0)*q(1) has -2 as a weight.
#
# + id="mcfRj-T-vSfz" colab_type="code" outputId="73f0d2c4-2d2a-4fd2-f6b5-21822297b472" colab={"base_uri": "https://localhost:8080/", "height": 34}
from blueqat import vqe
from blueqat.pauli import qubo_bit as q
h = -3*q(0)-3*q(1)-2*q(0)*q(1)
step = 2
result = vqe.Vqe(vqe.QaoaAnsatz(h, step)).run()
print(result.most_common(12))
# + [markdown] id="pgvdZYqGvtHP" colab_type="text"
# We get combination of (1,1) as the probability of 96%.
| tutorial/300_cop_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # DE Africa Coastlines useful tools <img align="right" src="https://github.com/digitalearthafrica/deafrica-sandbox-notebooks/raw/main/Supplementary_data/DE_Africa_Logo_Stacked_RGB_small.jpg">
#
# This notebook contains useful code snippets for processing and manipulating DE Africa Coastlines data.
#
#
# ---
# -
# ## Getting started
# Set working directory to top level of repo to ensure links work correctly:
# cd ..
# ### Load packages
#
# First we import the required Python packages, then we connect to the database, and load the catalog of virtual products.
# +
# %matplotlib inline
# %load_ext line_profiler
# %load_ext autoreload
# %autoreload 2
import os
import sys
import numpy as np
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
# -
# ## Extract style table from GeoPackage
import zipfile
with zipfile.ZipFile('../coastlines_v0.2.2.zip', 'r') as zip_ref:
zip_ref.extractall()
# Load 'layer_styles' from geopackage and export as a CSV
layer = gpd.read_file("coastlines_cli_update (6).gpkg", layer="layer_styles")
layer.drop(['geometry'], axis=1).to_csv('coastlines/styles.csv', index=False)
# ## View output files on S3
# +
# # !aws s3 --no-sign-request --region=af-south-1 ls --recursive s3://deafrica-data-dev-af/coastlines/ | grep '.gpkg$'
# -
# # !aws s3 --no-sign-request --region=af-south-1 ls --recursive s3://deafrica-data-staging-af/coastlines/
# ## Run status per tile from Argo YAML
# +
import pandas as pd
import yaml
from yaml import SafeLoader
# Load Argo job status
with open('run_status.yaml') as f:
data = yaml.load(f, Loader=SafeLoader)
# Keep only jobs with valid inputs
data_cleaned = {a:b for a, b in data['status']['nodes'].items() if 'inputs' in b}
# Obtain error code or missing error code for each job
df = pd.DataFrame(
[
(b["inputs"]["parameters"][0]["value"], b["outputs"]["exitCode"])
if "outputs" in b
else (b["inputs"]["parameters"][0]["value"], None)
for a, b in data_cleaned.items()
],
columns=["id", "error"],
)
# Drop non-tiles
df = df.loc[~df.id.isin(['v0.2.3', 'https://deafrica-input-datasets.s3.af-south-1.amazonaws.com/deafrica-coastlines/32km_coastal_grid_deafrica.geojson'])]
# Export to CSV that can be merged with tile grid
df['id'] = df['id'].astype(int)
df.to_csv('tile_status.csv', index=False)
# -
# ***
#
# ## Additional information
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
# Digital Earth Africa data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
#
# **Contact:** For assistance with any of the Python code or Jupyter Notebooks in this repository, please post a [Github issue](https://github.com/GeoscienceAustralia/DEACoastLines/issues/new).
#
# **Last modified:** May 2022
| notebooks/DEAfricaCoastlines_utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Baisal89/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/LS_DS6_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="838Dmw1kM2LK" colab_type="text"
# # Lambda School Data Science Module 132
# ## Sampling, Confidence Intervals, and Hypothesis Testing
# + [markdown] id="dbcPKIo5M6Ny" colab_type="text"
# ## Prepare - examine other available hypothesis tests
#
# If you had to pick a single hypothesis test in your toolbox, t-test would probably be the best choice - but the good news is you don't have to pick just one! Here's some of the others to be aware of:
# + id="tlBel8j9M6tB" colab_type="code" outputId="fbe5c03c-228d-44e1-8769-00e6757c15e1" colab={"base_uri": "https://localhost:8080/", "height": 190}
import numpy as np
from scipy.stats import chisquare # One-way chi square test
# Chi square can take any crosstab/table and test the independence of rows/cols
# The null hypothesis is that the rows/cols are independent -> low chi square
# The alternative is that there is a dependence -> high chi square
# Be aware! Chi square does *not* tell you direction/causation
ind_obs = np.array([[1, 1], [2, 2]]).T
print(ind_obs)
print(chisquare(ind_obs, axis=None))
dep_obs = np.array([[16, 18, 16, 14, 12, 12], [32, 24, 16, 28, 20, 24]]).T
print(dep_obs)
print(chisquare(dep_obs, axis=None))
# + id="nN0BdNiDPxbk" colab_type="code" outputId="a7f44206-ea89-458e-aee2-c8623046c8b1" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Distribution tests:
# We often assume that something is normal, but it can be important to *check*
# For example, later on with predictive modeling, a typical assumption is that
# residuals (prediction errors) are normal - checking is a good diagnostic
from scipy.stats import normaltest
# Poisson models arrival times and is related to the binomial (coinflip)
sample = np.random.poisson(5, 1000)
print(normaltest(sample)) # Pretty clearly not normal
# + id="P5t0WhkDReFO" colab_type="code" outputId="705683ef-d267-44d4-f7f4-1286e9ed2156" colab={"base_uri": "https://localhost:8080/", "height": 52}
# Kruskal-Wallis H-test - compare the median rank between 2+ groups
# Can be applied to ranking decisions/outcomes/recommendations
# The underlying math comes from chi-square distribution, and is best for n>5
from scipy.stats import kruskal
x1 = [1, 3, 5, 7, 9]
y1 = [2, 4, 6, 8, 10]
print(kruskal(x1, y1)) # x1 is a little better, but not "significantly" so
x2 = [1, 1, 1]
y2 = [2, 2, 2]
z = [2, 2] # Hey, a third group, and of different size!
print(kruskal(x2, y2, z)) # x clearly dominates
# + [markdown] id="7pT3IP36Rh0b" colab_type="text"
# And there's many more! `scipy.stats` is fairly comprehensive, though there are even more available if you delve into the extended world of statistics packages. As tests get increasingly obscure and specialized, the importance of knowing them by heart becomes small - but being able to look them up and figure them out when they *are* relevant is still important.
# + [markdown] id="3JqroCQYQqhy" colab_type="text"
# ## T-test Assumptions
#
# <https://statistics.laerd.com/statistical-guides/independent-t-test-statistical-guide.php>
#
# - Independence of means
#
# Are the means of our voting data independent (do not affect the outcome of one another)?
#
# The best way to increase thel likelihood of our means being independent is to randomly sample (which we did not do).
#
# + id="sqy2hEFRZnvI" colab_type="code" colab={}
from scipy.stats import ttest_ind
# ?ttest_ind
# + [markdown] id="xI-PcK5sZ1A9" colab_type="text"
# - "Homogeneity" of Variance?
#
# Is the magnitude of the variance between the two roughly the same?
#
# I think we're OK on this one for the voting data, although it probably could be better, one party was larger than the other.
#
# If we suspect this to be a problem then we can use Welch's T-test
# + id="P02dL0waauN5" colab_type="code" colab={}
# ?ttest_ind
# + [markdown] id="tjgoHHwGayoC" colab_type="text"
# - "Dependent Variable" (sample means) are Distributed Normally
#
# <https://stats.stackexchange.com/questions/9573/t-test-for-non-normal-when-n50>
#
# Lots of statistical tests depend on normal distributions. We can test for normality using Scipy as was shown above.
#
# This assumption is often assumed even if the assumption is a weak one. If you strongly suspect that things are not normally distributed, you can transform your data to get it looking more normal and then run your test. This problem typically goes away for large sample sizes (yay Central Limit Theorem) and is often why you don't hear it brought up. People declare the assumption to be satisfied either way.
#
#
# + [markdown] id="bvvPV-RJN2vA" colab_type="text"
# ## Central Limit Theorem
#
#
# + id="FBLoOF8qOJeJ" colab_type="code" outputId="3e0e96e1-5782-41ff-d1d7-f8ff4182e4ab" colab={"base_uri": "https://localhost:8080/", "height": 72}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
sample_means = []
for x in range(0,3000):
coinflips = np.random.binomial(n=1, p=.5, size=30)
one_sample = coinflips
sample_means.append(coinflips.mean())
print(len(sample_means))
print(sample_means)
# + id="rfeA06evOT2K" colab_type="code" outputId="871ead8a-5331-4280-cbb4-3866dc72f002" colab={"base_uri": "https://localhost:8080/", "height": 198}
df = pd.DataFrame({'a': one_sample})
df.head()
# + id="GlMSNFX6OmBV" colab_type="code" outputId="453b56ed-b81a-4904-fa12-1f80504fdb7b" colab={"base_uri": "https://localhost:8080/", "height": 269}
df.a.hist();
# + id="Jie4ypgLOs5M" colab_type="code" outputId="293a19d0-cd80-4fbe-cd5f-5f7717aebb1b" colab={"base_uri": "https://localhost:8080/", "height": 296}
ax = plt.hist(sample_means, bins=30)
plt.title('Distribution of 3000 sample means \n (of 30 coinflips each)');
# + [markdown] id="LsEAjc4rOylm" colab_type="text"
# What does the Central Limit Theorem State? That no matter the initial distribution of the population, the distribution of sample means taken will approximate a normal distribution as $n \rightarrow \infty$.
#
# This has very important implications for hypothesis testing and is precisely the reason why the t-distribution begins to approximate the normal distribution as our sample size increases.
# + [markdown] id="EYqo5vZZSFUr" colab_type="text"
# ## Standard Error of the Mean
#
# What does it mean to "estimate"? the Population mean?
# + id="puGXH6vbSIE4" colab_type="code" outputId="6040e78b-f687-42c9-aff3-454f70c53ff5" colab={"base_uri": "https://localhost:8080/", "height": 69}
import numpy as np
import pandas as pd
lambda_heights = np.random.uniform(4,6.5, size=2000)
print(len(lambda_heights))
lambda_heights
# + id="fQlloeU4qwuI" colab_type="code" outputId="e94a6eb7-2f36-484a-c485-b86bfe30a90b" colab={"base_uri": "https://localhost:8080/", "height": 52}
print("Population Mean:", lambda_heights.mean())
print("Population Standard Deviation:", lambda_heights.std())
# + id="sOD7gQMxq3ib" colab_type="code" outputId="8df59c31-6f50-4c8e-a7e6-8f5fdd9667b9" colab={"base_uri": "https://localhost:8080/", "height": 215}
population = pd.DataFrame({'heights': lambda_heights})
print(population.shape)
population.head()
# + id="A1DEQgCAq75F" colab_type="code" outputId="73815115-060e-4be7-ec63-3417ba06c232" colab={"base_uri": "https://localhost:8080/", "height": 215}
sample = population.sample(100)
print(sample.shape)
sample.head()
# + id="IMjMwv2NrETa" colab_type="code" outputId="d6892eb8-6e18-4b9f-8a4f-5f6d21e928e4" colab={"base_uri": "https://localhost:8080/", "height": 35}
print("Sample Mean 1:", sample['heights'].mean())
# + id="SpMBMasFrJQK" colab_type="code" outputId="1b0d470c-e51d-43e2-b495-46d1b21fbb2a" colab={"base_uri": "https://localhost:8080/", "height": 215}
sample = population.sample(100)
print(sample.shape)
sample.head()
# + id="l7hKc-8hrK0a" colab_type="code" outputId="6c91016c-8084-4b24-ef47-c6642849637a" colab={"base_uri": "https://localhost:8080/", "height": 35}
print("Sample Mean 2:", sample['heights'].mean())
# + [markdown] id="nfdQf8QYUUmw" colab_type="text"
# ## Build and Interpret a Confidence Interval
#
# <img src="https://github.com/ryanallredblog/ryanallredblog.github.io/blob/master/img/Confidence_Interval.png?raw=true" width=400>
# + id="tBx71Kf0UjT3" colab_type="code" outputId="0b118ec2-baad-4d0b-9014-f007c5219d9c" colab={"base_uri": "https://localhost:8080/", "height": 52}
coinflips_100 = np.random.binomial(n=1, p=.5, size=100)
sample_std = np.std(coinflips_100)
print("sample standard deviation:", sample_std)
sample_size = len(coinflips_100)
print("sample size:", sample_size)
# + id="r9qmrQmvwALM" colab_type="code" outputId="29f37b3b-ab3c-48dd-bcfb-f0e43879b220" colab={"base_uri": "https://localhost:8080/", "height": 35}
standard_error = sample_std / (sample_size**(.5))
print("standard error:", standard_error)
# + id="oU4undqDwQvE" colab_type="code" outputId="c34025b3-a3dd-4cef-a296-fdba471e71c9" colab={"base_uri": "https://localhost:8080/", "height": 35}
from scipy import stats
stderr = stats.sem(coinflips_100, ddof=0)
stderr
# + [markdown] id="RkYC5rnUw914" colab_type="text"
# ### What confidence level do we want our confidence interval to represent?
#
# 95% confidence Interval? 99% confidence interval?
# + id="jze1zJsewQx_" colab_type="code" colab={}
t = stats.t.ppf(.975 , sample_size-1)
# + id="7YPoL8ID0RvM" colab_type="code" colab={}
sample_mean = coinflips_100.mean()
# + id="Xd7Cs1fUz9f0" colab_type="code" outputId="9805c65d-888f-41e2-e075-7759a1970ffe" colab={"base_uri": "https://localhost:8080/", "height": 69}
confidence_interval = (sample_mean - t*stderr, sample_mean + t*stderr)
margin_of_error = t*stderr
print("Sample Mean", sample_mean)
print("Margin of Error:", margin_of_error)
print("Confidence Interval:", confidence_interval)
# + id="bOUTSf4p090g" colab_type="code" outputId="9948a259-ed06-44fa-b74a-3ae4374fd712" colab={"base_uri": "https://localhost:8080/", "height": 35}
confidence_interval[0]
# + id="7_FZmjhZ1EIN" colab_type="code" outputId="50c0a118-a4f6-4300-ef86-8c2a9d51d978" colab={"base_uri": "https://localhost:8080/", "height": 35}
confidence_interval[1]
# + [markdown] id="C4rtc8luVUAK" colab_type="text"
# ## Graphically Represent a Confidence Interval
# + id="pz6F9_3_VmKr" colab_type="code" outputId="52afeaf6-180c-4d56-b5f3-9fce22303833" colab={"base_uri": "https://localhost:8080/", "height": 269}
import seaborn as sns
sns.kdeplot(coinflips_100)
plt.axvline(x=confidence_interval[0], color='red')
plt.axvline(x=confidence_interval[1], color='red')
plt.axvline(x=sample_mean, color='k');
# + [markdown] id="_oy0uoBGeoEb" colab_type="text"
# ## Relationship between Confidence Intervals and T-tests
#
# Confidence Interval == Bounds of statistical significance for our t-test
#
# A sample mean that falls inside of our confidence interval will "FAIL TO REJECT" our null hypothesis
#
# A sample mean that falls outside of our confidence interval will "REJECT" our null hypothesis
# + id="izIyVavzfCXS" colab_type="code" colab={}
from scipy.stats import t, ttest_1samp
# + id="Y7HwdMwDfL1N" colab_type="code" outputId="d398bcde-14ef-4f81-c203-b59c776cb235" colab={"base_uri": "https://localhost:8080/", "height": 55}
import numpy as np
coinflip_means = []
for x in range(0,100):
coinflips = np.random.binomial(n=1, p=.5, size=30)
coinflip_means.append(coinflips.mean())
print(coinflip_means)
# + id="nQDo-ZXlfOvR" colab_type="code" outputId="eaedb934-a2ef-4ab6-a856-3cb5db7f581d" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Sample Size
n = len(coinflip_means)
# Degrees of Freedom
dof = n-1
# The Mean of Means:
mean = np.mean(coinflip_means)
# Sample Standard Deviation
sample_std = np.std(coinflip_means, ddof=1)
# Standard Error
std_err = sample_std/n**.5
CI = t.interval(.95, dof, loc=mean, scale=std_err)
print("95% Confidence Interval: ", CI)
# + id="PiaALHSNfWou" colab_type="code" outputId="20556729-6765-4ad2-bc89-be77d8ea1a61" colab={"base_uri": "https://localhost:8080/", "height": 52}
'''You can roll your own CI calculation pretty easily.
The only thing that's a little bit challenging
is understanding the t stat lookup'''
# 95% confidence interval
t_stat = t.ppf(.975, dof)
print("t Statistic:", t_stat)
CI = (mean-(t_stat*std_err), mean+(t_stat*std_err))
print("Confidence Interval", CI)
# + [markdown] id="EamZNJhAf-fY" colab_type="text"
# A null hypothesis that's just inside of our confidence interval == fail to reject
#
#
# + id="cNpzYbjpfirR" colab_type="code" outputId="d635e9f7-8f37-46f7-d66d-feaace4ce822" colab={"base_uri": "https://localhost:8080/", "height": 35}
ttest_1samp(coinflip_means, .471714)
# + [markdown] id="hO34mbL9gHn1" colab_type="text"
# A null hypothesis that's just outside of our confidence interval == reject
#
#
# + id="N4SUjj82gKlv" colab_type="code" outputId="c459fc95-b59b-495b-a4c6-8392eef17925" colab={"base_uri": "https://localhost:8080/", "height": 35}
ttest_1samp(coinflip_means, .471713)
# + id="rQZvNu6B3b9b" colab_type="code" colab={}
def confidence_interval(data, confidence=0.95):
"""
Calculate a confidence interval around a sample mean for given data.
Using t-distribution and two-tailed test, default 95% confidence.
Arguments:
data - iterable (list or numpy array) of sample observations
confidence - level of confidence for the interval
Returns:
tuple of (mean, lower bound, upper bound)
"""
data = np.array(data)
mean = np.mean(data)
n = len(data)
stderr = stats.sem(data)
interval = stderr * stats.t.ppf((1 + confidence) / 2.0, n - 1)
return (mean, mean - interval, mean + interval)
# + [markdown] id="pTIzrkKdUaLl" colab_type="text"
# ## Run a $\chi^{2}$ Test "by hand" (Using Numpy)
# + id="DDsovHUyUj3v" colab_type="code" outputId="24c1e681-a8c3-4f2d-a043-d0f6cad1350e" colab={"base_uri": "https://localhost:8080/", "height": 356}
df = pd.read_csv('https://raw.githubusercontent.com/ryanleeallred/datasets/master/adult.csv', na_values=" ?")
print(df.shape)
df.head()
# + id="r2gf-s8L8cYd" colab_type="code" outputId="3c80544f-e434-40a6-d101-655abcbdf830" colab={"base_uri": "https://localhost:8080/", "height": 288}
df.describe()
# + id="FN8Sx1Ze8jNE" colab_type="code" outputId="be9d13fd-d4bd-430d-bf1d-a86007e504ab" colab={"base_uri": "https://localhost:8080/", "height": 168}
df.describe(exclude='number')
# + id="S8_eNSN48pY7" colab_type="code" outputId="c8873376-d615-448a-c44a-5ac87c5df84e" colab={"base_uri": "https://localhost:8080/", "height": 339}
cut_points = [0, 9, 19, 29, 39, 49, 1000]
label_names = ['0-9', '10-19', '20-29', '30-39', '40-49', '50+']
df['hours_per_week_categories'] = pd.cut(df['hours-per-week'], cut_points, labels=label_names)
df.head()
# + id="fuzbRKh687CW" colab_type="code" outputId="39b46413-00a0-45fa-c2b9-257c5bc43910" colab={"base_uri": "https://localhost:8080/", "height": 69}
df['sex'].value_counts()
# + id="_Z1lOGK888yu" colab_type="code" outputId="1de5017e-0605-4c13-8c02-162753992c8e" colab={"base_uri": "https://localhost:8080/", "height": 138}
df['hours_per_week_categories'].value_counts()
# + id="YVae1vOL9WzG" colab_type="code" outputId="b5fb69ab-770f-404c-bf9a-929fe74a79ba" colab={"base_uri": "https://localhost:8080/", "height": 339}
df = df.sort_values(by='hours_per_week_categories', ascending=True)
df.head()
# + id="lEDLMzaP9ERN" colab_type="code" outputId="c4b6e7d6-9377-45a3-fdde-507e654aa8e3" colab={"base_uri": "https://localhost:8080/", "height": 168}
contingency_table = pd.crosstab(df['sex'], df['hours_per_week_categories'], margins=True)
contingency_table
# + id="_eYw-Fq39dSg" colab_type="code" outputId="d948789b-8369-4863-cb09-813b97b1060f" colab={"base_uri": "https://localhost:8080/", "height": 35}
femalecount = contingency_table.iloc[0][0:6].values
femalecount
# + id="61jfs69T9t3x" colab_type="code" outputId="f1886731-ced3-4a76-9d5a-d8de2c91dd08" colab={"base_uri": "https://localhost:8080/", "height": 35}
malecount = contingency_table.iloc[1][0:6].values
malecount
# + id="xR0Nm24891Xd" colab_type="code" outputId="a19867f2-eb3f-4440-92fc-0d78db978b04" colab={"base_uri": "https://localhost:8080/", "height": 361}
import matplotlib.pyplot as plt
import seaborn as sns
#Plots the bar chart
fig = plt.figure(figsize=(10, 5))
sns.set(font_scale=1.8)
categories = ["0-9","10-19","20-29","30-39","40-49","50+"]
p1 = plt.bar(categories, malecount, 0.55, color='#d62728')
p2 = plt.bar(categories, femalecount, 0.55, bottom=malecount)
plt.legend((p2[0], p1[0]), ('Female', 'Male'))
plt.xlabel('Hours per Week Worked')
plt.ylabel('Count')
plt.show()
# + [markdown] id="uyw_hby7-OHF" colab_type="text"
# ## Expected Value Calculation
# \begin{align}
# expected_{i,j} =\frac{(row_{i} \text{total})(column_{j} \text{total}) }{(\text{total observations})}
# \end{align}
# + id="C11nWaal-acY" colab_type="code" outputId="392e1834-8f66-46f9-efc6-42985acb8aa9" colab={"base_uri": "https://localhost:8080/", "height": 52}
# Get Row Sums
row_sums = contingency_table.iloc[0:2, 6].values
col_sums = contingency_table.iloc[2, 0:6].values
print(row_sums)
print(col_sums)
# + id="XANdl4XR-LOw" colab_type="code" outputId="2df1f796-ccce-49c8-b31f-2bc7a754f478" colab={"base_uri": "https://localhost:8080/", "height": 35}
total = contingency_table.loc['All','All']
total
# + id="2bB64F9G-pzd" colab_type="code" outputId="f2b1a25c-8a2e-4991-bb57-caa51b1fe022" colab={"base_uri": "https://localhost:8080/", "height": 104}
expected = []
for i in range(len(row_sums)):
expected_row = []
for column in col_sums:
expected_val = column*row_sums[i]/total
expected_row.append(expected_val)
expected.append(expected_row)
expected = np.array(expected)
print(expected.shape)
print(expected)
# + id="LwLh6hSl-2aY" colab_type="code" outputId="e7cc7808-9e3e-454f-a46b-bfc167d13940" colab={"base_uri": "https://localhost:8080/", "height": 69}
observed = pd.crosstab(df['sex'], df['hours_per_week_categories']).values
print(observed.shape)
observed
# + [markdown] id="R6AWydhG_P4s" colab_type="text"
# ## Chi-Squared Statistic with Numpy
#
# \begin{align}
# \chi^2 = \sum \frac{(observed_{i}-expected_{i})^2}{(expected_{i})}
# \end{align}
#
# For the $observed$ values we will just use a version of our contingency table without the margins as a numpy array. In this way, if our observed values array and our expected values array are the same shape, then we can subtract them and divide them directly which makes the calculations a lot cleaner. No for loops!
# + id="o7YgaNij_cSo" colab_type="code" outputId="590fd6bb-e345-47f7-9162-af6b64009101" colab={"base_uri": "https://localhost:8080/", "height": 35}
chi_squared = ((observed - expected)**2/(expected)).sum()
print(f"Chi-Squared: {chi_squared}")
# + id="KkBhRm-aAHTS" colab_type="code" outputId="1b7c3492-08c8-46f5-d588-f968a145383f" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Calculate Degrees of Freedom
dof = (len(row_sums)-1)*(len(col_sums)-1)
print(f"Degrees of Freedom: {dof}")
# + [markdown] id="7Igz-XHcVbW3" colab_type="text"
# ## Run a $\chi^{2}$ Test using Scipy
# + id="kazgId8L9tYZ" colab_type="code" outputId="737b4e29-55bb-4e98-d520-063168756adb" colab={"base_uri": "https://localhost:8080/", "height": 155}
chi_squared, p_value, dof, expected = stats.chi2_contingency(observed)
print(f"Chi-Squared: {chi_squared}")
print(f"P-value: {p_value}")
print(f"Degrees of Freedom: {dof}")
print("Expected: \n", np.array(expected))
# + [markdown] id="TRtBEP3rA307" colab_type="text"
# Null Hypothesis: Hours worked per week bins is **independent** of sex.
#
# Due to a p-value of 0, we REJECT the null hypothesis that hours worked per week and sex are independent, and conclude that there is an association between hours worked per week and sex.
# + [markdown] id="11OzdxWTM7UR" colab_type="text"
# ## Assignment - Build a confidence interval
#
# A confidence interval refers to a neighborhood around some point estimate, the size of which is determined by the desired p-value. For instance, we might say that 52% of Americans prefer tacos to burritos, with a 95% confidence interval of +/- 5%.
#
# 52% (0.52) is the point estimate, and +/- 5% (the interval $[0.47, 0.57]$) is the confidence interval. "95% confidence" means a p-value $\leq 1 - 0.95 = 0.05$.
#
# In this case, the confidence interval includes $0.5$ - which is the natural null hypothesis (that half of Americans prefer tacos and half burritos, thus there is no clear favorite). So in this case, we could use the confidence interval to report that we've failed to reject the null hypothesis.
#
# But providing the full analysis with a confidence interval, including a graphical representation of it, can be a helpful and powerful way to tell your story. Done well, it is also more intuitive to a layperson than simply saying "fail to reject the null hypothesis" - it shows that in fact the data does *not* give a single clear result (the point estimate) but a whole range of possibilities.
#
# How is a confidence interval built, and how should it be interpreted? It does *not* mean that 95% of the data lies in that interval - instead, the frequentist interpretation is "if we were to repeat this experiment 100 times, we would expect the average result to lie in this interval ~95 times."
#
# For a 95% confidence interval and a normal(-ish) distribution, you can simply remember that +/-2 standard deviations contains 95% of the probability mass, and so the 95% confidence interval based on a given sample is centered at the mean (point estimate) and has a range of +/- 2 (or technically 1.96) standard deviations.
#
# Different distributions/assumptions (90% confidence, 99% confidence) will require different math, but the overall process and interpretation (with a frequentist approach) will be the same.
#
# Your assignment - using the data from the prior module ([congressional voting records](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records)):
#
#
# ### Confidence Intervals:
# 1. Generate and numerically represent a confidence interval
# 2. Graphically (with a plot) represent the confidence interval
# 3. Interpret the confidence interval - what does it tell you about the data and its distribution?
#
# ### Chi-squared tests:
# 4. Take a dataset that we have used in the past in class that has **categorical** variables. Pick two of those categorical variables and run a chi-squared tests on that data
# - By hand using Numpy
# - In a single line using Scipy
#
# Stretch goals:
#
# 1. Write a summary of your findings, mixing prose and math/code/results. *Note* - yes, this is by definition a political topic. It is challenging but important to keep your writing voice *neutral* and stick to the facts of the data. Data science often involves considering controversial issues, so it's important to be sensitive about them (especially if you want to publish).
# 2. Apply the techniques you learned today to your project data or other data of your choice, and write/discuss your findings here.
# 3. Refactor your code so it is elegant, readable, and can be easily run for all issues.
# + id="ywc0LC_y5rYL" colab_type="code" colab={}
from scipy.stats import norm
import numpy as np
# + id="8DEOWY9X5xx_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bb779d3d-1b33-4ac2-b411-6bdd132c61b0"
norm.ppf(0.975)## 95% of confidence level, inverse cumolative distribution function, this gives us Z critical critical value
# + id="fZOgivX355J_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5c4318cc-0d10-4d4d-eb92-07f22b98c339"
norm.ppf(0.995)#99%
# + id="g2QGIPbS5_gJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d8155020-7ea8-4a3f-ac1c-3c22f7eb7e7d"
norm.ppf(0.95)#90%
# + id="Ckcr4A4FM7cs" colab_type="code" colab={}
# TODO - your code!
# %matplotlib inline
import pandas as pd
import numpy as np
import scipy
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats
import math
from __future__ import division
# + [markdown] id="IxVzHoD-7zFi" colab_type="text"
# ###DATA
# + id="91EqPGqD7w51" colab_type="code" colab={}
df_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data'
df1 = pd.read_csv(df_url)
# + id="4L8USFJAcgj9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="7236cf51-597a-430f-b646-6c95066fcdf2"
print(df1.shape)
df1.head(5)
# + id="XakMrfkyckOZ" colab_type="code" colab={}
column_renamed = ['party', 'handicapped-infants',
'water-project-cost-sharing',
'adoption-of-the-budget-resolution',
'physician-fee-freeze',
'el-salvador-aid', 'religious-groups-in-schools',
'anti-satellite-test-ban', 'aid-to-nicaraguan-contras',
'mx-missele', 'immigration', 'synfuels-corporation-cutback',
'education-spending', 'superfund-right-to-sue', 'crime',
'duty-free-exports', 'export-administration-act-south-africa']
# + id="_w-4pN2sdI5G" colab_type="code" colab={}
df2 = pd.read_csv(df_url, names=column_renamed)
# + id="6BO3xOu2eO1W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="418063e9-d9db-41b2-af06-2bab8ff5c6d1"
df2.head()
# + id="zRjEEhLoeTKL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="1083b3d7-5fa3-469e-d77e-a8fb2ebc2b3c"
df3 = df2.replace(['y', 'n', '?'], [1, 0, 1])
df3.head()
# + id="4g6i6bEoew-M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="a2d9b530-bdac-41e7-f5c4-b2309200f667"
df3.isnull().sum()
# + id="Z2JgML5iSlUA" colab_type="code" colab={}
dem = df3.loc[df3['party']=='democrat']
rep = df3.loc[df3['party']=='republican']
# + id="5ZQmiCZZTLkY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="98232f09-0966-490b-ab84-4283015e0e35"
rep.describe()
# + id="qaXX682UezjO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 388} outputId="be2ce87b-3cba-4326-eeb2-304095d43a41"
dem.describe()
# + id="A7XoDn4vSa9q" colab_type="code" colab={}
st_d = pd.DataFrame({'Democrats': dem.mean(), 'Republicans': rep.mean()})
# + id="4AkO8BqfTd5J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="3d6b2613-962d-4eee-e2fc-eea8a554c15f"
st_d
# + id="aiLdjWma9TJ8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 354} outputId="a7a4216a-c8a5-4060-e844-a18a2f110f7e"
plt.hist(st_d.Democrats)
# + id="HEvFM19K9cXY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 354} outputId="93c98f84-e0ac-4147-9100-60bb414becc8"
plt.hist(st_d.Republicans)
# + id="US7pKB289Rm6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="54c2aeda-31be-46b1-ba0f-d2b373d40602"
sns.distplot(st_d.Republicans)
# + id="OxVZtshq9tFF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="3c4181d5-f7fc-456e-e3e7-0f12ae8819c6"
sns.distplot(st_d.Democrats)
# + id="Wk6tT6ad90Wa" colab_type="code" colab={}
n = len(st_d)
con_coef = .95
#alpha level
alpha = 1. - con_coef
# + id="4W-xgYuT90Ty" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b89dd76c-00d4-4b4e-a6ad-56d1dc4ea785"
d_bar_mean = st_d['Democrats'].mean()
d_bar_mean
# + id="YAhWT47T90Ra" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3eb9d4bf-490e-4e1b-b600-af67f53dc76a"
p_bar_mean = st_d['Republicans'].mean()
p_bar_mean
# + id="EiM5H0Ap90Oy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fd612932-aafa-4a3c-8d55-ff59678cbd39"
sigma = st_d['Democrats'].std()
sigma
# + id="-5sTmCcj90Md" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6d2e966e-55ac-418d-96a4-09e8d98b4fef"
sigma = st_d['Republicans'].std()
sigma
# + id="AUZUCMvg90JV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3fe6dd86-759e-4b1b-8589-36f1b41acce6"
#Here i will be looking at the Z critical Value
import scipy.stats as stats
z_critical = stats.norm.ppf(q = 0.975)
z_critical
# + id="yC-HqvGL_H54" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="15707918-497b-484f-8517-7eb5afcf41fe"
#now i have the z critical value,
#next step is to find the intevals
zinterval = stats.norm.interval(alpha=con_coef)
zinterval
# + id="sPzL-2sc_olF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="481901b3-ff95-4111-9ad7-c7c6c1d6f7d1"
#So here i will do standart error to calculate the bounds
stan_err = sigma / math.sqrt(n)
stan_err
# + id="SdZsho0N_897" colab_type="code" colab={}
conf_inte_lower = d_bar_mean - z_critical * stan_err
conf_inte_upper = p_bar_mean + z_critical * stan_err
# + id="8jDWOX5JAdKE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3bd9f7c7-ab39-4d73-a640-dec56b1e0aca"
conf_inte_lower, conf_inte_upper
# + [markdown] id="xNzd3l00ArQT" colab_type="text"
# ###Taking Sample
# + id="xRanWbhJApvV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 340} outputId="cb0ec2b0-b188-4cd1-87cb-7edc8b757775"
n_sample = 75
df_sample = st_d.ix[np.random.choice(st_d.index, n)]
df_sample.head()
# + id="A9b8B4I_BXHQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="29ea1e78-37ff-4eca-a7ea-7fb3796a5072"
d = sns.distplot(df_sample.Democrats)
# + id="hbfXtOJMBleg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="19e58fde-18ba-4349-fab9-522066ad303b"
p = sns.distplot(df_sample.Republicans)
# + id="BG5-5l0EByI_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="effec5c9-a8ce-4622-92fa-68a4894714f0"
#the mean
d_bar_sample = df_sample.Democrats.mean()
d_bar_sample
# + id="s2bRhQlqB_Gp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9b852c08-6d5c-4517-b310-d2dd79ff8867"
#the mean
p_bar_sample = df_sample.Republicans.mean()
p_bar_sample
# + id="b79UIIuzCpNp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ba1bb0d7-f393-4710-afb8-04bc8c49e418"
#standart deviation sigma of Democarats
sigma_sample_d = df_sample.Democrats.std()
sigma_sample_d
# + id="UgCFc3MOC5gO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="af1cc0cc-a2bb-42ef-d1d7-664633de232c"
#standart deviation sigma of Republicans
sigma_sample_p = df_sample.Republicans.std()
sigma_sample_p
# + id="K5LPEWbpDFB6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ef39ba0a-163c-4631-c7d5-5a210a9ec63d"
#calculating standart error
stan_err_sample = sigma_sample / math.sqrt(n_sample)
stan_err_sample
# + id="zSXclXanDyWl" colab_type="code" colab={}
#Upper and Lower for out sample
ci_lower_sample = d_bar_sample - z_critical * stan_err_sample
ci_upper_sample = p_bar_sample + z_critical * stan_err_sample
# + id="YxWABle4EQMX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f3bc401a-6283-4f12-d809-5e8b792d9d4b"
ci_lower_sample, ci_upper_sample
# + id="P_ifVjVrEQJv" colab_type="code" colab={}
# + [markdown] id="nyJ3ySr7R2k9" colab_type="text"
# ## Resources
#
# - [Interactive visualize the Chi-Squared test](https://homepage.divms.uiowa.edu/~mbognar/applets/chisq.html)
# - [Calculation of Chi-Squared test statistic](https://en.wikipedia.org/wiki/Pearson%27s_chi-squared_test)
# - [Visualization of a confidence interval generated by R code](https://commons.wikimedia.org/wiki/File:Confidence-interval.svg)
# - [Expected value of a squared standard normal](https://math.stackexchange.com/questions/264061/expected-value-calculation-for-squared-normal-distribution) (it's 1 - which is why the expected value of a Chi-Squared with $n$ degrees of freedom is $n$, as it's the sum of $n$ squared standard normals)
| LS_DS6_132_Sampling_Confidence_Intervals_and_Hypothesis_Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas
import os.path
import collections
Data = collections.namedtuple('Data', ['mnist', 'cifar'])
def load_dataset_files(filename, data_dirs={"cifar": "cifar-data", "mnist": "mnist-data"}, **kwargs):
dataframes = {
dataset_name: pandas.read_csv(os.path.join(data_dir, filename), **kwargs)
for dataset_name, data_dir in data_dirs.items()
}
return Data(**dataframes)
gcp_data = load_dataset_files("baseline-gcp.csv")
cache_data = load_dataset_files("baseline-cache.csv", converters={"Miss Rate": lambda percentage: float(percentage.rstrip('%'))/100})
disk_data = load_dataset_files("baseline-disk.csv")
fetch_size_data = load_dataset_files("fetch-size-results.csv", converters={"Miss Rate": lambda percentage: float(percentage.rstrip('%'))/100})
fetch_size_data = [df[df["Fetch Size"] == 1024] for df in fetch_size_data]
min_queue_size_data = load_dataset_files("min-queue-size.csv")
min_queue_size_data = [df[(df["Minimum Queue Size"] == 1024) & (df["Cache Size"] == 2048)] for df in min_queue_size_data]
# +
import collections
BATCH_SIZE = 512
def get_bucketed_data(data, bucket_field, *fields, average_over_bucket_field=True):
res = []
for epoch in data['Epoch'].drop_duplicates():
epoch_data = data.loc[(data['Epoch'] == epoch) & (data['Batch Size'] == BATCH_SIZE)][[bucket_field, *fields]]
if average_over_bucket_field:
epoch_data = epoch_data.groupby(bucket_field).mean()
res.append(epoch_data)
return res
# +
from matplotlib import pyplot as plt
import inflect
import numpy
import functools
import math
plt.rcParams.update({'font.size': 14})
BAR_WIDTH = 0.2
# HATCHES = ['', '...']
HATCHES = {'mnist': '', 'cifar': '...'}
COLORS = {"mnist": 'deeppink', "cifar": 'tab:cyan'}
COLORS2 = {
'mnist': ['lightskyblue', 'royalblue'],
'cifar': ['plum', 'magenta']
}
DATASET_DISPLAY_NAMES = {"mnist": "MNIST", "cifar": "CIFAR-10"}
inflect_engine = inflect.engine()
GraphEntry = collections.namedtuple('GraphEntry', ['average', 'error'])
def generate_epoch_name(epoch_index):
epoch_name = inflect_engine.number_to_words(inflect_engine.ordinal(epoch_index + 1)) + " Epoch"
epoch_name = epoch_name[0].upper() + epoch_name[1:]
return epoch_name
def make_bucketed_graph(data, bucket_field, dependent_field, labels_override=None, xlabel_override=None, custom_ordering=None):
bucketed_data = Data(*[
get_bucketed_data(item, bucket_field, dependent_field, average_over_bucket_field=False)
for item in data
])
"""
[
#epoch
{
'dataset_name': {size: entry}
}
]
"""
epoch_entries = []
plt.figure(figsize=(10,4))
for dataset_index, (data, dataset_name) in enumerate(zip(bucketed_data, bucketed_data._fields)):
for epoch, epoch_times in enumerate(data):
epoch_name = inflect_engine.number_to_words(inflect_engine.ordinal(epoch + 1)) + " Epoch"
epoch_name = epoch_name[0].upper() + epoch_name[1:]
bucket_items = collections.defaultdict(list)
for _, row in epoch_times.iterrows():
bucket = row[bucket_field]
dependent_res = row[dependent_field]
bucket_items[bucket].append(dependent_res)
bucket_iteration_list = sorted(bucket_items.items(), key=lambda item: item[0])
if custom_ordering:
bucket_iteration_list = sorted(zip(custom_ordering, bucket_items.items()), key=lambda item: item[0])
bucket_iteration_list = [item[1] for item in bucket_iteration_list]
entries = {
bucket: GraphEntry(
sum(items)/len(items),
numpy.std(items)
)
for bucket, items in bucket_iteration_list
}
if epoch >= len(epoch_entries):
epoch_entries.append({})
epoch_entries[epoch][dataset_name] = entries
all_num_entries = set()
for epoch, datasets in enumerate(epoch_entries):
# This is so gross but we're just ensuring that the same number of items are in all datasets
num_entries = len(datasets[list(datasets.keys())[0]].keys())
all_num_entries.add(num_entries)
assert all(num_entries == len(datasets[key].keys()) for key in datasets.keys())
assert len(all_num_entries) == 1
dataset_keys = {
dataset_name: list(dataset_entries.keys())
for dataset_name, dataset_entries in datasets.items()
}
epoch_name = generate_epoch_name(epoch)
for i in range(num_entries):
entries_at_i = [
(dataset_name, datasets[dataset_name][dataset_keys[dataset_name][i]])
for dataset_name, dataset_entries in datasets.items()]
entries_at_i.sort(key=lambda entry: entry[1].average, reverse=True)
for dataset_name, graph_entry in entries_at_i:
dataset_index = sorted(dataset_keys.keys(), reverse=True).index(dataset_name)
bar_shift = BAR_WIDTH * (epoch + dataset_index * 2) - 1.5 * BAR_WIDTH
plt.bar(
i + bar_shift,
graph_entry.average,
BAR_WIDTH,
capsize=4,
label=f"{epoch_name} {DATASET_DISPLAY_NAMES[dataset_name]}",
hatch=HATCHES[dataset_name],
color=COLORS2[dataset_name][epoch],
edgecolor="white",
linewidth=0
)
error_shift = 0
plt.errorbar(
i - error_shift + bar_shift,
graph_entry.average, yerr=graph_entry.error,
fmt='none',
barsabove=True,
color='black',
capsize=4
)
# https://stackoverflow.com/questions/13588920/stop-matplotlib-repeating-labels-in-legend
handles, labels = plt.gca().get_legend_handles_labels()
legend_labels = dict(zip(labels, handles))
legend_labels = {
"First Epoch (MNIST)": legend_labels["First Epoch MNIST"],
"Second Epoch (MNIST)": legend_labels["Second Epoch MNIST"],
"First Epoch (CIFAR-10)": legend_labels["First Epoch CIFAR-10"],
"Second Epoch (CIFAR-10)": legend_labels["Second Epoch CIFAR-10"],
}
plt.legend(legend_labels.values(), legend_labels.keys())
xlabels = labels_override
if xlabels is None:
xlabels = datasets[list(datasets.keys())[0]].keys()
assert all(xlabels == datasets[key].keys() for key in datasets.keys()), "You may want to use an xlabel override so that we don't have an ambiguity in labelling"
plt.xticks(numpy.arange(all_num_entries.pop()), xlabels)
plt.title(f"{dependent_field} vs {bucket_field}")
plt.xlabel(xlabel_override if xlabel_override else bucket_field)
plt.ylabel(dependent_field)
plt.grid(axis='y', alpha=0.5)
plt.savefig(f"Combined Baseline {dependent_field} vs {bucket_field}", bbox_inches="tight", dpi=300)
# + tags=[]
import math
corrected_cache_data = Data(*[data.copy() for data in cache_data])
for data in corrected_cache_data:
data["Cache Size"] = data["Cache Size"].replace(math.nan, math.inf)
make_bucketed_graph(corrected_cache_data, "Cache Size", "Miss Rate", ["25%", "50%", "75%", "inf"], "Cache Size (as percentage of node partition size)")
# + tags=[]
plt.rcParams.update({'font.size': 14})
unlimited_cache_items = Data(
*[
data
.drop(data[~data["Cache Size"].apply(math.isnan)].index)
for data in cache_data
]
)
for data in unlimited_cache_items:
data["Loading Method"] = "Unlimited \n Cache"
gcp_data_items = []
for data in gcp_data:
gcp_data_copy = data.copy()
gcp_data_copy["Loading Method"] = "GCP Bucket"
gcp_data_items.append(gcp_data_copy)
gcp_data_copy = Data(*gcp_data_items)
disk_data_items = []
for data in disk_data:
disk_data_copy = data.copy()
disk_data_copy["Loading Method"] = "Disk"
disk_data_items.append(disk_data_copy)
disk_data_copy = Data(*disk_data_items)
fetch_size_data_items = []
for data in fetch_size_data:
fetch_size_data_copy = data.copy()
fetch_size_data_copy["Loading Method"] = "Unlimited \n Cache \n w/ Prefetch \n(FS=1024)"
fetch_size_data_items.append(fetch_size_data_copy)
fetch_size_data_copy = Data(*fetch_size_data_items)
min_queue_size_data_items = []
for data in min_queue_size_data:
min_queue_size_data_copy = data.copy()
min_queue_size_data_copy["Loading Method"] = "50/50 Approach \n (FS=1024)"
min_queue_size_data_items.append(min_queue_size_data_copy)
min_queue_size_data_copy = Data(*min_queue_size_data_items)
concated_items = [
pandas.concat(
[gcp_data_copy[i], disk_data_copy[i], unlimited_cache_items[i], fetch_size_data_copy[i], min_queue_size_data_copy[i]]
)
for i in range(len(gcp_data_copy))
]
make_bucketed_graph(
concated_items,
"Loading Method",
"Data Loading Time (s)",
custom_ordering=[1, 0, 2, 3, 4]
)
plt.gcf().subplots_adjust(bottom=0.15)
# plt.savefig("./baselines.png", dpi=800)
# plt.savefig("./baselines.png", bbox_inches="tight", dpi=300)
| notebooks/base-analysis/Baseline and Unl. Cache Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
from fastai.nlp import *
from sklearn.linear_model import LogisticRegression
# -
# ## IMDB dataset and the sentiment classification task
# The [large movie review dataset](http://ai.stanford.edu/~amaas/data/sentiment/) contains a collection of 50,000 reviews from IMDB. The dataset contains an even number of positive and negative reviews. The authors considered only highly polarized reviews. A negative review has a score ≤ 4 out of 10, and a positive review has a score ≥ 7 out of 10. Neutral reviews are not included in the dataset. The dataset is divided into training and test sets. The training set is the same 25,000 labeled reviews.
#
# The **sentiment classification task** consists of predicting the polarity (positive or negative) of a given text.
#
# To get the dataset, in your terminal run the following commands:
#
# `wget http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz`
#
# `gunzip aclImdb_v1.tar.gz`
#
# `tar -xvf aclImdb_v1.tar`
# ### Tokenizing and term document matrix creation
PATH='data/aclImdb/'
names = ['neg','pos']
# %ls {PATH}
# %ls {PATH}train
# %ls {PATH}train/pos | head
trn,trn_y = texts_labels_from_folders(f'{PATH}train',names)
val,val_y = texts_labels_from_folders(f'{PATH}test',names)
# Here is the text of the first review
trn[0]
trn_y[0]
# [`CountVectorizer`](http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html) converts a collection of text documents to a matrix of token counts (part of `sklearn.feature_extraction.text`).
veczr = CountVectorizer(tokenizer=tokenize)
# `fit_transform(trn)` finds the vocabulary in the training set. It also transforms the training set into a term-document matrix. Since we have to apply the *same transformation* to your validation set, the second line uses just the method `transform(val)`. `trn_term_doc` and `val_term_doc` are sparse matrices. `trn_term_doc[i]` represents training document i and it contains a count of words for each document for each word in the vocabulary.
trn_term_doc = veczr.fit_transform(trn)
val_term_doc = veczr.transform(val)
trn_term_doc
trn_term_doc[0]
vocab = veczr.get_feature_names(); vocab[5000:5005]
w0 = set([o.lower() for o in trn[0].split(' ')]); w0
len(w0)
veczr.vocabulary_['absurd']
trn_term_doc[0,1297]
trn_term_doc[0,5000]
# ## Naive Bayes
# We define the **log-count ratio** $r$ for each word $f$:
#
# $r = \log \frac{\text{ratio of feature $f$ in positive documents}}{\text{ratio of feature $f$ in negative documents}}$
#
# where ratio of feature $f$ in positive documents is the number of times a positive document has a feature divided by the number of positive documents.
def pr(y_i):
p = x[y==y_i].sum(0)
return (p+1) / ((y==y_i).sum()+1)
# +
x=trn_term_doc
y=trn_y
r = np.log(pr(1)/pr(0))
b = np.log((y==1).mean() / (y==0).mean())
# -
# Here is the formula for Naive Bayes.
pre_preds = val_term_doc @ r.T + b
preds = pre_preds.T>0
(preds==val_y).mean()
# ...and binarized Naive Bayes.
# +
x=trn_term_doc.sign()
r = np.log(pr(1)/pr(0))
pre_preds = val_term_doc.sign() @ r.T + b
preds = pre_preds.T>0
(preds==val_y).mean()
# -
# ### Logistic regression
# Here is how we can fit logistic regression where the features are the unigrams.
m = LogisticRegression(C=1e8, dual=True)
m.fit(x, y)
preds = m.predict(val_term_doc)
(preds==val_y).mean()
m = LogisticRegression(C=1e8, dual=True)
m.fit(trn_term_doc.sign(), y)
preds = m.predict(val_term_doc.sign())
(preds==val_y).mean()
# ...and the regularized version
m = LogisticRegression(C=0.1, dual=True)
m.fit(x, y)
preds = m.predict(val_term_doc)
(preds==val_y).mean()
m = LogisticRegression(C=0.1, dual=True)
m.fit(trn_term_doc.sign(), y)
preds = m.predict(val_term_doc.sign())
(preds==val_y).mean()
# + [markdown] heading_collapsed=true
# ### Trigram with NB features
# + [markdown] hidden=true
# Our next model is a version of logistic regression with Naive Bayes features described [here](https://www.aclweb.org/anthology/P12-2018). For every document we compute binarized features as described above, but this time we use bigrams and trigrams too. Each feature is a log-count ratio. A logistic regression model is then trained to predict sentiment.
# + hidden=true
veczr = CountVectorizer(ngram_range=(1,3), tokenizer=tokenize, max_features=800000)
trn_term_doc = veczr.fit_transform(trn)
val_term_doc = veczr.transform(val)
# + hidden=true
trn_term_doc.shape
# + hidden=true
vocab = veczr.get_feature_names()
# + hidden=true
vocab[200000:200005]
# + hidden=true
y=trn_y
x=trn_term_doc.sign()
val_x = val_term_doc.sign()
# + hidden=true
r = np.log(pr(1) / pr(0))
b = np.log((y==1).mean() / (y==0).mean())
# + [markdown] hidden=true
# Here we fit regularized logistic regression where the features are the trigrams.
# + hidden=true
m = LogisticRegression(C=0.1, dual=True)
m.fit(x, y);
preds = m.predict(val_x)
(preds.T==val_y).mean()
# + [markdown] hidden=true
# Here is the $\text{log-count ratio}$ `r`.
# + hidden=true
r.shape, r
# + hidden=true
np.exp(r)
# + [markdown] hidden=true
# Here we fit regularized logistic regression where the features are the trigrams' log-count ratios.
# + hidden=true
x_nb = x.multiply(r)
m = LogisticRegression(dual=True, C=0.1)
m.fit(x_nb, y);
val_x_nb = val_x.multiply(r)
preds = m.predict(val_x_nb)
(preds.T==val_y).mean()
# -
# ## fastai NBSVM++
sl=2000
# Here is how we get a model from a bag of words
md = TextClassifierData.from_bow(trn_term_doc, trn_y, val_term_doc, val_y, sl)
learner = md.dotprod_nb_learner()
learner.fit(0.02, 1, wds=1e-6, cycle_len=1)
learner.fit(0.02, 2, wds=1e-6, cycle_len=1)
learner.fit(0.02, 2, wds=1e-6, cycle_len=1)
# + [markdown] heading_collapsed=true
# ## References
# + [markdown] hidden=true
# * Baselines and Bigrams: Simple, Good Sentiment and Topic Classification. <NAME> and <NAME> [pdf](https://www.aclweb.org/anthology/P12-2018)
# + hidden=true
| courses/ml1/lesson5-nlp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AxelAllen/Pre-trained-Multimodal-Text-Image-Classifier-in-a-Sparse-Data-Application/blob/master/run_bert_text_only.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CbEER_DL5uu1"
# # Run Text-Only Experiments
#
# This notebook shows the end-to-end pipeline to fine-tune pre-trained BERT model for text classification on our dataset.
#
# Parts of this pipeline are adapted from [McCormick's and Ryan's Tutorial on BERT Fine-Tuning](http://mccormickml.com/2019/07/22/BERT-fine-tuning/) and the
# Huggingface `run_mmimdb.py` script to execute the MMBT model. This code can
# be accessed [here.](https://github.com/huggingface/transformers/blob/8ea412a86faa8e9edeeb6b5c46b08def06aa03ea/examples/research_projects/mm-imdb/run_mmimdb.py#L305)
# + [markdown] id="zxeWeqpC5-CO"
# ## Skip unless on Google Colab
#
# + colab={"base_uri": "https://localhost:8080/"} id="gbcc0iwU2Z9n" outputId="3867d62b-1dbb-45d3-ec24-219fe3f5fcb4"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="wAwUUimTyz58" outputId="8f68542d-78b3-43cf-ad09-9d93c5038b4c"
# %pwd
# + [markdown] id="Kuai6mBTTCaK"
# ### Working Directory
# The notebook needs to be executed from the parent directory of the project, i.e. the `LAP` folder, which contains the notebooks, the data/, MMBT/, runs/, etc. directories.
#
# Change the cell below to reflace the correct path to the `LAP` folder in your drive.
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="kARAPI3_y9u7" outputId="ab6d0b91-a03e-424b-ffdb-e99ff5dbb618"
# %cd /content/drive/MyDrive/LAP
# %pwd
# + [markdown] id="A2QTP-Y3Ttn1"
# ### Checking Directory
# If you're in the correct directory, the command in the cell below should show the notebooks, MMBT/, data/, runs/, integrated_gradients/ directories. If you're not getting this outputk, you are not in the correct directory to run the subsequent cells in this notebook.
# + colab={"base_uri": "https://localhost:8080/"} id="XZbCJ9LdYG8R" outputId="9b4e5301-9c62-43ba-8dc0-5e582f0980c1"
# %ls
# + [markdown] id="HUHI4kEs6Dyi"
# ## Check GPU is Available
# + colab={"base_uri": "https://localhost:8080/"} id="3_3rHDZCzCTD" outputId="57db8ba5-e222-4758-b531-be1434289116"
import torch
# If there's a GPU available...
if torch.cuda.is_available():
# Tell PyTorch to use the GPU.
device = torch.device("cuda")
print('There are %d GPU(s) available.' % torch.cuda.device_count())
print('We will use the GPU:', torch.cuda.get_device_name(0))
# If not...
else:
print('No GPU available, using the CPU instead.')
device = torch.device("cpu")
# + [markdown] id="A4_WNIuW6KjT"
# ## Install Huggingface Trnasformers and WandB modules
#
# These should have been installed during your environment set-up; you only need to run these cells in Google Colab.
# + colab={"base_uri": "https://localhost:8080/"} id="5x-N8RUNzoDC" outputId="a021e1a7-13aa-4906-88f2-7d89d314b571"
# !pip install transformers
# + colab={"base_uri": "https://localhost:8080/"} id="wGCIhVcxaCis" outputId="4725e122-2133-4f8e-c8ab-e13a407a9ec4"
# %pip install wandb
# + [markdown] id="xV0Y4fNaOsew"
# ## Import Required Modules
# + id="2TvcJ0KYOsew"
from textBert_utils import (
get_train_val_test_data,
tokenize_and_encode_data,
make_tensor_dataset,
make_dataloader,
set_seed,
get_label_frequencies,
get_multiclass_criterion
)
# + id="qKv4TCojQ7uI"
from MMBT.mmbt_utils import get_multiclass_labels, get_labels
# + id="nljiGS-pTTKI"
import textBert_utils
# + id="B8t3xllQn1Qi"
import argparse
import pandas as pd
import os
import wandb
import glob
import numpy as np
# + id="8vvRk6_BOsex"
import logging
import json
# + id="3AHC-9ZjQjCT"
from transformers import (
WEIGHTS_NAME,
AutoConfig,
AutoModelForSequenceClassification,
AutoTokenizer,
)
# + [markdown] id="z_AbRMeB69ym"
# # Set-up Experiment Hyperparameters and Arguments
#
# Specify the training, validation, and test files to run the experiment on. The default here is running the model on both 'findings' and 'impression' texts.
#
# To re-make the training, validation, and test data, please refer to the information in the **data/** directory.
#
# Change the default values in the parser.add_argument function for the hyperparameters that you want to specify in the following cell or use the default option.
#
# For multiple experiment runs, please make sure to change the `output_dir` argument so that new results don't overwrit existing ones.
# + id="6p2ZlSYbh4BK"
#train_file = "image_labels_impression_frontal_train.csv"
#val_file = "image_labels_impression_frontal_val.csv"
#test_file = "image_labels_impression_frontal_test.csv"
#train_file = "image_multi_labels_major_findings_frontal_train.csv"
#val_file = "image_multi_labels_major_findings_frontal_val.csv"
#test_file = "image_multi_labels_major_findings_frontal_test.csv"
#train_file = "image_labels_major_findings_frontal_train.csv"
#val_file = "image_labels_major_findings_frontal_val.csv"
#test_file = "image_labels_major_findings_frontal_test.csv"
train_file = "image_labels_findings_frontal_train.csv"
val_file = "image_labels_findings_frontal_val.csv"
test_file = "image_labels_findings_frontal_test.csv"
# + id="QvzzL8Vuovw_"
parser = argparse.ArgumentParser(f'Project Hyperparameters and Other Configurations Argument Parser')
# + id="ibpA7E14oiZE"
parser = argparse.ArgumentParser()
# Required parameters
parser.add_argument(
"--data_dir",
default="data/csv",
type=str,
help="The input data dir. Should contain the .jsonl files.",
)
parser.add_argument(
"--model_name",
default="bert-base-uncased",
type=str,
help="model identifier from huggingface.co/models",
)
parser.add_argument(
"--output_dir",
default="10epochs_text_only_findings",
type=str,
help="The output directory where the model predictions and checkpoints will be written.",
)
parser.add_argument(
"--config_name", default="bert-base-uncased", type=str, help="Pretrained config name if not the same as model_name"
)
parser.add_argument(
"--tokenizer_name",
default="bert-base-uncased",
type=str,
help="Pretrained tokenizer name or path if not the same as model_name",
)
parser.add_argument("--train_batch_size", default=32, type=int, help="Batch size for training.")
parser.add_argument(
"--eval_batch_size", default=32, type=int, help="Batch size for evaluation."
)
parser.add_argument(
"--max_seq_length",
default=300,
type=int,
help="The maximum total input sequence length after tokenization. Sequences longer "
"than this will be truncated, sequences shorter will be padded.",
)
parser.add_argument(
"--num_image_embeds", default=3, type=int, help="Number of Image Embeddings from the Image Encoder"
)
parser.add_argument("--do_train", default=True, type=bool, help="Whether to run training.")
parser.add_argument("--do_eval", default=True, type=bool, help="Whether to run eval on the dev set.")
parser.add_argument(
"--evaluate_during_training", default=True, type=bool, help="Rul evaluation during training at each logging step."
)
parser.add_argument(
"--gradient_accumulation_steps",
type=int,
default=1,
help="Number of updates steps to accumulate before performing a backward/update pass.",
)
parser.add_argument("--learning_rate", default=5e-5, type=float, help="The initial learning rate for Adam.")
parser.add_argument("--weight_decay", default=0.1, type=float, help="Weight deay if we apply some.")
parser.add_argument("--adam_epsilon", default=1e-8, type=float, help="Epsilon for Adam optimizer.")
parser.add_argument("--max_grad_norm", default=1.0, type=float, help="Max gradient norm.")
parser.add_argument(
"--num_train_epochs", default=10.0, type=float, help="Total number of training epochs to perform."
)
parser.add_argument("--patience", default=5, type=int, help="Patience for Early Stopping.")
parser.add_argument(
"--max_steps",
default=-1,
type=int,
help="If > 0: set total number of training steps to perform. Override num_train_epochs.",
)
parser.add_argument("--warmup_steps", default=0, type=int, help="Linear warmup over warmup_steps.")
parser.add_argument("--logging_steps", type=int, default=25, help="Log every X updates steps.")
parser.add_argument("--save_steps", type=int, default=25, help="Save checkpoint every X updates steps.")
parser.add_argument(
"--eval_all_checkpoints",
default=True, type=bool,
help="Evaluate all checkpoints starting with the same prefix as model_name ending and ending with step number",
)
parser.add_argument("--num_workers", type=int, default=8, help="number of worker threads for dataloading")
parser.add_argument("--seed", type=int, default=42, help="random seed for initialization")
args = parser.parse_args("")
# Setup CUDA, GPU & distributed training
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
args.n_gpu = torch.cuda.device_count() if torch.cuda.is_available() else 0
args.device = device
# Setup Train/Val/Test filenames
args.train_file = train_file
args.val_file = val_file
args.test_file = test_file
# accomodatae multiclass labeling
args.multiclass = False
# + [markdown] id="BHXt2oRR76zo"
# ### Check that the Args dict contains correct configurations
# + colab={"base_uri": "https://localhost:8080/"} id="MeWSr2B02eY4" outputId="f0c903a5-1397-45ea-8f1b-41365fbc5bd1"
args.__dict__
# + [markdown] id="PUe1Aej96lbJ"
# ## Set-up WandB
#
# We are setting up our code to run more experiments later and would be tracking them in the WandB API. You need to sign up for an account first to continue.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} id="cwIuONkRcLsF" outputId="5cbc38ac-73ab-4251-a93e-0a17ef4a04e9"
wandb.login()
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="G82FnR5vtAbS" outputId="ff5edba6-973c-4cec-a984-e3ec997154a8"
wandb.init(name="Train_Findings_Texts_10", tags=['Findings', 'frontal'], project="Text_Only", notes="10 epochs 256 size and 32 batch", config=args.__dict__)
run_name = wandb.run.name
wandb_config = wandb.config
# + [markdown] id="n5abmm7V8FyP"
# ## Create Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="L18XRia4z_jc" outputId="b9cdf667-196c-4891-eaa8-041b474d5788"
train, val, test = get_train_val_test_data(wandb_config)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="HzKIwM7eCJee" outputId="0eaa09a5-6da6-432a-d9e0-03061384d44b"
train.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="_NfnWPR95Apc" outputId="3b30e9fe-5074-4219-c343-7006a7e846f6"
val.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="4dnH95_3VNMw" outputId="f270c6b5-e3e6-4218-99d7-d6d1a72954e0"
test.head()
# + [markdown] id="cyx0BCy9_z8u"
# # sentences and labels
# + id="xLSXvlOc_zT8"
train_sentences = train.text.values
train_labels = train.label.values
val_sentences = val.text.values
val_labels = val.label.values
test_sentences = test.text.values
test_labels = test.label.values
# + colab={"base_uri": "https://localhost:8080/"} id="bus3yFPvAWHX" outputId="a25f5168-da97-4c33-9add-357847d68721"
train_sentences[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="w9HUNbeWAb8e" outputId="4070467a-b4eb-4b0c-9fda-3930c6fc799b"
train_labels[:10]
# + [markdown] id="Pkoi56y3EX4e"
# # Tokenize and Encode with BERT encoder plus
# + [markdown] id="8Xe9z2opBU1W"
# The `tokenizer.encode_plus` function combines multiple steps for us:
#
# 1. Split the sentence into tokens.
# 2. Add the special `[CLS]` and `[SEP]` tokens.
# 3. Map the tokens to their IDs.
# 4. Pad or truncate all sentences to the same length.
# 5. Create the attention masks which explicitly differentiate real tokens from `[PAD]` tokens.
#
# These steps are performed inside the `make_tensor_dataset` function.
# + [markdown] id="49DUwTrmEl43"
# # Torch dataset and dataloader
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["21df3a41cd1949a28631d7ddcc942e2e", "802eb53a671f4d248a43728f9dd119d0", "f6d53fbe2885490e8e89d0235dbb44b5", "d7fdfc9c14774fc69be4ce1240eead6d", "8336558f99cc46888da931f428e9e21e", "30dd8b34d69b4328949a3674d3154a08", "<KEY>", "8f142d3230074d31beffe8a560895784", "44e5015ef3334ff68537619215d364d6", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "c9c74e48d95d4ece9dfec36ba41ecc2c", "4fb3cd6644be41df8386385f8df8866c", "<KEY>", "c0db84e3bf5f497780453755ee5e0739", "3cca2f2eea4b4977b4bab18f9622689b", "<KEY>", "07387a0aa5e14ce1a3a5652debd2f017", "aba5ef4c5fac44ec9de6f627421e0773", "ec9b23fbbe4940e58f01a1837717e221", "1030fbe306e64c1ebae2007cff2fb143"]} id="8-QTtgDFEkwD" outputId="2f2ad73c-0ac5-44da-aff0-38ef0fa83a05"
train_dataset = make_tensor_dataset(train_sentences, train_labels, wandb_config)
val_dataset = make_tensor_dataset(val_sentences, val_labels, wandb_config)
# + colab={"base_uri": "https://localhost:8080/"} id="m1j1bQcmEfdD" outputId="450a51cc-1435-440c-f8d3-e345f3665043"
print(f'{len(train_dataset):>5,} training samples')
print(f'{len(val_dataset):>5,} validation samples')
#print(f'{len(test_dataset):>5,} test samples')
# + colab={"base_uri": "https://localhost:8080/"} id="zTjcmuENFvBS" outputId="e5aeddaa-58e8-4c76-ddb2-6c8a93b88df8"
train_dataset[:3]
# + [markdown] id="Ray80zREF5nx"
# Create an iterator for the dataset using the torch DataLoader class.
# + id="zw8rZBerAJ-a"
data_loaders = {
'train' : make_dataloader(train_dataset, wandb_config, eval=False),
'train_size': len(train_dataset),
'eval' : make_dataloader(val_dataset, wandb_config, eval=True),
'eval_size' : len(val_dataset)
}
# + [markdown] id="zARn-95bGbYc"
# # Fine Tune BERT for Classification
# + [markdown] id="XtnoCExl97Ss"
# ## Setup Logging
# + id="5cd16bmFS1T5"
# Setup logging
logger = logging.getLogger(__name__)
if not os.path.exists(wandb_config.output_dir):
os.makedirs(wandb_config.output_dir)
logging.basicConfig(format="%(asctime)s - %(levelname)s - %(name)s - %(message)s",
datefmt="%m/%d/%Y %H:%M:%S",
filename=os.path.join(wandb_config.output_dir, f"{os.path.splitext(wandb_config.train_file)[0]}_logging.txt"),
level=logging.INFO)
logger.warning("device: %s, n_gpu: %s",
wandb_config.device,
wandb_config.n_gpu
)
# Set the verbosity to info of the Transformers logger (on main process only):
# Set seed
set_seed(wandb_config)
# + [markdown] id="jsLgoErr-CKX"
# ## Set up the Model and Train
#
# The Code will simply train and validate the specified train and validation sets.
#
# Outputs and saved checkpoints are saved in the specifed `--output_dir` argument.
# Tensorboard data are saved in the `runs/` directory with the date and time of the experiment as well as the filename of the train/test data file.
# + colab={"background_save": true, "base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["f1dd478ca17f4aacb3640ae6b2b0702a", "5c047fa462914813a79c506924ed618b", "fe302f48e5354d748b977a1e20b38c76", "<KEY>", "3592cf79c0ee4ef682954306f55524f8", "5f6f2b69804a4d75bebc883b770d7b54", "762150e077d340cf937d43bbe2b403ab", "89e0187e5afe44c590af5a1344420e08", "<KEY>", "<KEY>", "3f3d6cb26caf4e75a195dbec6abe0cb3", "<KEY>", "<KEY>", "4980d19561584f49b2575076ac214079", "82a184f6222c480a88d5c321a9e5c686", "7f187f02e68144b8ad714572bd02788c", "<KEY>"]} id="BQamR9mcRMFi" outputId="d2191cf4-8b14-4c2c-83ea-45dc00c68587"
# %pdb on
# set up model
if args.multiclass:
labels = get_multiclass_labels()
num_labels = len(labels)
else:
labels = get_labels()
num_labels = len(labels)
transformer_config = AutoConfig.from_pretrained(wandb_config.model_name, num_labels=num_labels)
tokenizer = AutoTokenizer.from_pretrained(
wandb_config.tokenizer_name,
do_lower_case=True,
cache_dir=None,
)
transformer_model = AutoModelForSequenceClassification.from_pretrained(wandb_config.model_name, config=transformer_config)
transformer_model.to(device)
logger.info(f"Training/evaluation parameters: {wandb_config}")
# Training
if wandb_config.do_train:
if wandb_config.multiclass:
criterion = get_multiclass_criterion(train_labels)
global_step, tr_loss = textBert_utils.train(data_loaders, wandb_config, transformer_model, criterion)
else:
global_step, tr_loss = textBert_utils.train(data_loaders, wandb_config, transformer_model)
logger.info(" global_step = %s, average loss = %s", global_step, tr_loss)
# Saving best-practices: if you use defaults names for the model, you can reload it using from_pretrained()
logger.info("Saving model checkpoint to %s", wandb_config.output_dir)
# Save a trained model, configuration and tokenizer using `save_pretrained()`.
# They can then be reloaded using `from_pretrained()`
model_to_save = (transformer_model.module if hasattr(transformer_model, "module") else transformer_model) # Take care of distributed/parallel training
torch.save(model_to_save.state_dict(), os.path.join(wandb_config.output_dir, WEIGHTS_NAME))
tokenizer.save_pretrained(wandb_config.output_dir)
transformer_config.save_pretrained(wandb_config.output_dir)
# Good practice: save your training arguments together with the trained model
torch.save(args, os.path.join(wandb_config.output_dir, "training_args.bin"))
# Load a trained model and vocabulary that you have fine-tuned
transformer_model = AutoModelForSequenceClassification.from_pretrained(wandb_config.model_name, config=transformer_config)
transformer_model.load_state_dict(torch.load(os.path.join(wandb_config.output_dir, WEIGHTS_NAME)))
tokenizer = AutoTokenizer.from_pretrained(wandb_config.output_dir)
transformer_model.to(device)
logger.info("***** Training Finished *****")
wandb.finish()
# + [markdown] id="Y3HT1vZmofkp"
# # Evaluation on Test set
# + [markdown] id="vOur0pn-oiay"
# ## tokenizer and prepare test dataset
#
# use the saved tokenizer from the training step
# + colab={"background_save": true} id="w-kuN61ooW1v" outputId="8d9bbd11-964b-4d04-ceca-e374b1265250"
wandb.init(name="Test_Findings_Texts_10", tags=['Findings', 'frontal'], project="Text_Only", notes="10 epochs 256 size and 32 batch", config=args.__dict__)
# wandb.tensorboard.patch(root_logdir="...")
run_name = wandb.run.name
wandb_config = wandb.config
# + colab={"background_save": true} id="L6K5T2jEpJQx" outputId="0720ee90-ce84-4242-e0f2-5fa2966ac68d"
test_dataset = make_tensor_dataset(test_sentences, test_labels, wandb_config, saved_model=True)
# + colab={"background_save": true} id="k2dCKWr-oheZ"
data_loaders['test'] = make_dataloader(test_dataset, wandb_config, eval=True)
data_loaders['test_size'] = len(test_dataset)
# + id="tFrWAi5AvjWs"
# Evaluation
results = {}
if wandb_config.do_eval:
checkpoints = [wandb_config.output_dir]
if wandb_config.eval_all_checkpoints:
checkpoints = list(os.path.dirname(c)
for c in sorted(glob.glob(wandb_config.output_dir + "/**/" +
WEIGHTS_NAME, recursive=False)))
# recursive=False because otherwise the parent diretory gets included
# which is not what we want; only subdirectories
logger.info("Evaluate the following checkpoints: %s", checkpoints)
for checkpoint in checkpoints:
global_step = checkpoint.split("-")[-1] if len(checkpoints) > 1 else ""
prefix = checkpoint.split("/")[-1] if checkpoint.find("checkpoint") != -1 else ""
transformer_model = AutoModelForSequenceClassification.from_pretrained(wandb_config.model_name, config=transformer_config)
checkpoint = os.path.join(checkpoint, 'pytorch_model.bin')
transformer_model.load_state_dict(torch.load(checkpoint))
transformer_model.to(wandb_config.device)
if wandb_config.multiclass:
result = textBert_utils.evaluate(data_loaders, wandb_config, transformer_model, prefix=prefix, test=True, criterion=criterion)
else:
result = textBert_utils.evaluate(data_loaders, wandb_config, transformer_model, prefix=prefix, test=True) # test=True uses the test_dataset not val_dataset
result = dict((k + "_{}".format(global_step), v) for k, v in result.items())
results.update(result)
logger.info("***** Evaluation on Test Data Finished *****")
wandb.finish()
# + [markdown] id="8PBlhF6N_GUj"
# ## Saving Test Eval Results
#
# The code automatically saved evaluation result from each checkpoint in its respective folder. This next cell simply saves all of them in one place.
# + id="oJq2ZUXBmsXW"
with open(os.path.join(args.output_dir, f"{os.path.splitext(args.test_file)[0]}_eval_results.txt"), mode='w', encoding='utf-8') as out_f:
print(results, file=out_f)
| run_bert_text_only.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pika
# create connection and channel
connection = pika.BlockingConnection(pika.ConnectionParameters('server'))
channel = connection.channel()
# create a new queue so that future messages can be received
queue_key='hello'
channel.queue_declare(queue=queue_key)
# -
# publish some message
channel.basic_publish(exchange='', routing_key=queue_key, body='Hello World!')
| 01-HelloWorld/Sender.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PROJ_irox_oer] *
# language: python
# name: conda-env-PROJ_irox_oer-py
# ---
# + jupyter={"source_hidden": true}
import os
import sys
import copy
import shutil
from pathlib import Path
from contextlib import contextmanager
# import pickle; import os
import pickle
import json
import pandas as pd
import numpy as np
from ase import io
import plotly.graph_objects as go
from pymatgen.io.ase import AseAtomsAdaptor
from pymatgen.analysis import local_env
# #########################################################
from misc_modules.pandas_methods import drop_columns
from methods import read_magmom_comp_data
import os
print(os.getcwd())
import sys
from IPython.display import display
import pandas as pd
pd.set_option("display.max_columns", None)
pd.options.display.max_colwidth = 20
# pd.set_option('display.max_rows', None)
# #########################################################
from methods import (
get_df_jobs_paths,
get_df_dft,
get_df_job_ids,
get_df_jobs,
get_df_jobs_data,
get_df_slab,
get_df_slab_ids,
get_df_jobs_data_clusters,
get_df_jobs_anal,
get_df_slabs_oh,
get_df_init_slabs,
get_df_magmoms,
get_df_ads,
get_df_atoms_sorted_ind,
get_df_rerun_from_oh,
get_df_slab_simil,
get_df_active_sites,
get_df_features_targets,
)
from methods import (
get_other_job_ids_in_set,
read_magmom_comp_data,
)
from methods import get_df_coord
from ase.visualize import view
# + jupyter={"source_hidden": true}
df_dft = get_df_dft()
df_job_ids = get_df_job_ids()
df_jobs = get_df_jobs(exclude_wsl_paths=True)
df_jobs_data = get_df_jobs_data(exclude_wsl_paths=True)
df_jobs_data_clusters = get_df_jobs_data_clusters()
df_slab = get_df_slab()
df_slab_ids = get_df_slab_ids()
df_jobs_anal = get_df_jobs_anal()
df_jobs_paths = get_df_jobs_paths()
df_slabs_oh = get_df_slabs_oh()
df_init_slabs = get_df_init_slabs()
df_magmoms = get_df_magmoms()
df_ads = get_df_ads()
df_atoms_sorted_ind = get_df_atoms_sorted_ind()
df_rerun_from_oh = get_df_rerun_from_oh()
magmom_data_dict = read_magmom_comp_data()
df_slab_simil = get_df_slab_simil()
df_active_sites = get_df_active_sites()
df_features_targets = get_df_features_targets()
# + jupyter={"source_hidden": true}
def display_df(df, df_name, display_head=True, num_spaces=3):
print(40 * "*")
print(df_name)
print("df_i.shape:", df_i.shape)
print(40 * "*")
if display_head:
display(df.head())
print(num_spaces * "\n")
df_list = [
("df_dft", df_dft),
("df_job_ids", df_job_ids),
("df_jobs", df_jobs),
("df_jobs_data", df_jobs_data),
("df_jobs_data_clusters", df_jobs_data_clusters),
("df_slab", df_slab),
("df_slab_ids", df_slab_ids),
("df_jobs_anal", df_jobs_anal),
("df_jobs_paths", df_jobs_paths),
("df_slabs_oh", df_slabs_oh),
("df_magmoms", df_magmoms),
("df_ads", df_ads),
("df_atoms_sorted_ind", df_atoms_sorted_ind),
("df_rerun_from_oh", df_rerun_from_oh),
("df_slab_simil", df_slab_simil),
("df_active_sites", df_active_sites),
]
# for name_i, df_i in df_list:
# display_df(df_i, name_i)
# print("")
# print("")
# for name_i, df_i in df_list:
# display_df(
# df_i,
# name_i,
# display_head=False,
# num_spaces=0)
# -
# ### Script Inputs
# +
# ('sherlock', 'telibose_95', 'oh', 35.0, 1)
# +
# dft_jobs/nd919pnr6q/111/01_attempt
# +
names_i = [
# ('slac', 'votafefa_68', 38.0),
# ('sherlock', 'kegidafu_92', 66.0),
# ('slac', 'bikoradi_95', 65., )
# ('sherlock', 'ramufalu_44', 54.0, )
('sherlock', 'telibose_95', 54.0, ),
]
# ('slac', 'bikoradi_95', 'o', 65, 1, False)
# ('sherlock', 'kegidafu_92', 'oh', 66.0, 1, True)
# job_id = "melifiwi_93"
job_id = "toberotu_75"
# ('slac', 'waloguhe_35', 65.0),
# ('sherlock', 'kesekodi_38', 50.0),
# ('slac', 'votafefa_68', 38.0),
# +
from methods import get_other_job_ids_in_set
df_oer_set = get_other_job_ids_in_set(job_id, df_jobs=df_jobs, oer_set=True)
# # get_other_job_ids_in_set?
# +
df = df_oer_set[["compenv", "slab_id", "ads", "active_site", "att_num", ]]
idx = pd.MultiIndex.from_tuples(
[tuple(x) for x in df.to_records(index=False)]
)
unique_idx = idx.unique()
long_indices = unique_idx.tolist()
# -
df_jobs_anal.loc[long_indices]
# +
# assert False
# +
df_ind = df_features_targets.index.to_frame()
df = df_ind
df = df[
(df["compenv"] == "slac") &
(df["slab_id"] == "bikoradi_95") &
# (df["active_site"] == 65.) &
[True for i in range(len(df))]
]
df
# +
from methods import create_name_str_from_tup
df_feat_i = df_features_targets.loc[names_i]
out_dir = os.path.join(
os.environ["PROJ_irox_oer"],
"__temp__/oer_sets",
)
try:
if os.path.exists(out_dir):
shutil.rmtree(out_dir)
except:
tmp = 42
job_ids = []
for index_i, row_i in df_feat_i.iterrows():
atoms_list = []
ads_list = ["o", "oh", "bare", ]
# ads_list = ["o", "bare", ]
# ads_list = ["oh", ]
for ads_i in ads_list:
job_id_i = row_i["data"]["job_id_" + ads_i][""]
# print("job_id_i:", job_id_i)
job_ids.append(job_id_i)
row_paths_i = df_jobs_paths.loc[job_id_i]
path_dir = os.path.join(
os.environ["PROJ_irox_oer_gdrive"],
row_paths_i.gdrive_path,
)
path_0 = os.path.join(path_dir, "final_with_calculator.json")
path_1 = os.path.join(path_dir, "out.cif")
atoms_i = io.read(path_0)
atoms_list.append(atoms_i)
out_dir = os.path.join(
os.environ["PROJ_irox_oer"],
"__temp__/oer_sets",
create_name_str_from_tup(index_i),
)
print(out_dir)
if not os.path.exists(out_dir):
os.makedirs(out_dir)
shutil.copy(
path_1,
os.path.join(
out_dir,
ads_i + ".cif"
)
)
view(atoms_list)
# -
print("job_ids:", job_ids)
# ### Print out local paths of jobs
# job_ids = ["pibuvule_81", ]
# job_ids = ["toberotu_75", ]
job_ids = ["wawehamu_56", ]
for job_id_i in job_ids:
row_paths_i = df_jobs_paths.loc[job_id_i]
gdrive_path_i = row_paths_i.gdrive_path
full_path_i = os.path.join(
os.environ["PROJ_irox_oer_gdrive"],
gdrive_path_i)
print(full_path_i)
# +
# frag_i = "slac/8p8evt9pcg/220/bare/active_site__24/01_attempt"
# frag_i = "slac/8p8evt9pcg/220/bare/active_site__24"
# frag_i = "8p8evt9pcg/220/bare"
# frag_i = "slac/8p8evt9pcg/220/bare/active_site__"
# frag_i = "slac/8p8evt9pcg/220/oh"
# frag_i = "dft_jobs/sherlock/v2blxebixh/2-10/bare/active_site__49/01_attempt"
# frag_i = "dft_workflow/run_slabs/run_oh_covered/out_data/dft_jobs/slac/b5cgvsb16w/111/oh/active_site__71/01_attempt/_02"
# frag_i = "dft_workflow/run_slabs/run_oh_covered/out_data/dft_jobs/zimixdvdxd/010/oh/active_site__26/00_attempt/_01"
# frag_i = "zimixdvdxd/010/oh/active_site__26"
# frag_i = "dft_workflow/run_slabs/run_oh_covered/out_data/dft_jobs/slac/zimixdvdxd/010/oh/active_site__26/00_attempt/_01"
# frag_i = "dft_jobs/nd919pnr6q/111/01_attempt"
frag_i = "nd919pnr6q/111"
for job_id_i, row_i in df_jobs_paths.iterrows():
gdrive_path_i = row_i.gdrive_path
if frag_i in gdrive_path_i:
print(job_id_i)
print(gdrive_path_i)
print("")
# +
df_jobs_paths
df_jobs.loc["seladuri_58"]
('sherlock', 'likeniri_51', 'o', 'NaN', 1)
# + active=""
#
#
#
#
# -
import os
import json
# +
root_path_i = os.path.join(
os.environ["PROJ_irox_oer_gdrive"],
"dft_workflow/run_slabs/run_o_covered/out_data/dft_jobs/slac",
"8p937183bh/10-12/active_site__38/01_attempt/_01")
# #########################################################
path_i = os.path.join(
root_path_i,
"out_data/init_magmoms.json")
with open(path_i, "r") as fle:
init_magmoms = json.load(fle)
# #########################################################
path_i = os.path.join(
root_path_i,
"init.traj")
atoms_init = io.read(path_i)
# #########################################################
path_i = os.path.join(
root_path_i,
"OUTCAR")
atoms_outcar = io.read(path_i, index=":")
# +
atoms_0 = atoms_outcar[0]
magmoms = atoms_0.get_magnetic_moments()
for atom, magmom_i in zip(atoms_0, magmoms):
print(
atom.index, "|",
atom.symbol, "|",
# atom.magmom, "|",
magmom_i, "|",
atom.position,
)
# -
np.sum(magmoms)
# magmoms
# +
# # io.read?
# +
# # MAGMOM = 1*1.0540 1*1.0010 1*-0.3030 1*-0.9940 1*-0.3480 1*0.3280 1*0.0070 1*-0.0020 1*-0.0370 1*-0.0240 1*-0.0270 1*-0.0440 1*- 0.0520 1*-0.0250 1*-0.0340 1*-0.0420 1*-0.0170 1*-0.0300 1*-0.0120 1*-0.0400 1*0.0050 1*-0.0070 1*0.0670 1*-0.0320 1*0.2310 1*-0. 0070 1*-0.0130 1*-0.0010 1*0.8900 1*-0.8390 1*-0.0530 1*-0.1740 1*-0.0140 1*-0.2610 1*-0.2180 1*-0.1320 1*0.0410 1*0.2170 1*-0. 0180
# 1.0540
# 1.0010
# -0.3030
# -0.9940
# -0.3480
# 0.3280
# 0.0070
# -0.0020
# -0.0370
# -0.0240
# -0.0270
# -0.0440
# -0.0520
# -0.0250
# -0.0340
# -0.0420
# -0.0170
# -0.0300
# -0.0120
# -0.0400
# 0.0050
# -0.0070
# 0.0670
# -0.0320
# 0.2310
# -0.0070
# -0.0130
# -0.0010
# 0.8900
# -0.8390
# -0.0530
# -0.1740
# -0.0140
# -0.2610
# -0.2180
# -0.1320
# 0.0410
# 0.2170
# -0.0180
# -
for atom in atoms_init:
print(
atom.index, "|",
atom.symbol, "|",
atom.magmom, "|",
atom.position,
# atom.position, "|",
)
# atoms_init.get_magnetic_moments()
atoms_init.get_initial_magnetic_moments()
np.array(init_magmoms)
| sandbox/manually_inspect_slabs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [TPV3] Off fault case: Desktop at Uni
# by <NAME> (Created on 02.12.2020)
#
# +
import os, sys, math, time
from glob import glob
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
sys.path.insert(0,"/import/freenas-m-03-geodynamics/jhayek/TEAR/processing/TEAR/PythonCodes/LibFolder")
from Lib_GeneralFunctions import *
from Lib_GeneralSignalProcNAnalysis import *
from Lib_ProfilePlotting import *
from Lib_ProfileProcessing import *
#=================== Plotting style ===================
plt.style.use('seaborn-whitegrid')
from matplotlib import cm
from matplotlib.colors import ListedColormap
from matplotlib.lines import Line2D
from matplotlib.gridspec import GridSpec
#definition of colormap
from palettable.scientific.sequential import Oleron_20
cmap = Oleron_20.mpl_colormap
plt.register_cmap(cmap=cmap)
plt.set_cmap('Oleron_20')
# -
# Timestamp variable
start_time = time.time()
# Save into a class the
class TPV3reference:
def __init__(self, filename, coordinates, RefSource="SEM2DPACK"):
line = pd.read_csv(filename.format("slip"), header=None)
self.Time = line[0]
self.Slip = line[1]
line = pd.read_csv(filename.format("sr"), header=None)
self.SlipRate = line[1]
self.Coord = coordinates #Only used for labels and
self.RefSource = RefSource
#end __init__
# Default object printing information
def __repr__(self):
return "The TPV3reference object was generated from: {} and the receiver is located at {}".format(self.RefSource, self.Coord)
#end __repr__
def __str__(self):
return "The TPV3reference object was generated from: {} and the receiver is located at {}".format(self.RefSource, self.Coord)
#end __str__
def PlotReference(self, ax, SlipSlipRate, filtering=True, **kwargs):
if SlipSlipRate=="Slip":
if(filtering):
ax.plot(self.Time, Butterworth(self.Slip, **kwargs), label = "", c = "k", ls = "--", zorder=1)
else:
ax.plot(self.Time, self.Slip, label = "", c = "k", ls = "--", zorder=1)
elif SlipSlipRate=="SlipRate":
if(filtering):
ax.plot(self.Time, Butterworth(self.SlipRate, **kwargs), label = "", c = "k", ls = "--", zorder=1)
else:
ax.plot(self.Time, self.SlipRate, label = "", c = "k", ls = "--", zorder=1)
return ax
# +
path = "/import/freenas-m-03-geodynamics/jhayek/TEAR/processing/TEAR/ProfilePicking/"
# Reference saved into a list of objects
RefList = [TPV3reference(path + "Output/Reference/sem2dpack/sem2d-{}-2.txt", "4km"),
TPV3reference(path + "Output/Reference/sem2dpack/sem2d-{}-3.txt", "6km"),
TPV3reference(path + "Output/Reference/sem2dpack/sem2d-{}-4.txt", "8km"),
]
# +
def format_axes(fig):
for i, ax in enumerate(fig.axes):
ax.set_xlim(0,7)
ax.set_ylim(-1,1)
Lines = fig.axes[-1].get_lines()[-4:]
legend2 = fig.axes[-1].legend(Lines, ['Reference', '4km', '6km', '8km'], loc=1)
fig.axes[-1].add_artist(legend2)
def GenericFigAxis():
fig = plt.figure(constrained_layout=True, figsize=[12,4])
gs = GridSpec(1, 2, figure=fig)
ax1 = fig.add_subplot(gs[0, 0])
ax2 = fig.add_subplot(gs[0, 1])
return fig, [ax1, ax2]
# -
def PlotReceiverFile(ax, ReceiverFile, ColIDX, OrderPeriodicity=8, NumReceivers=3, filtering=True, **kwargs):
ylabeldict={1:"Slip (m)", 2:"Slip Rate (m)", 3:"$\mu$"}
if(filtering):
SamplingFrequency = 1./(ReceiverFile[0][1]-ReceiverFile[0][0])
[ax.plot(ReceiverFile[0],
Butterworth(ReceiverFile[ColIDX+OrderPeriodicity*i],SamplingFrequency = SamplingFrequency, **kwargs),
zorder=2) for i in range(NumReceivers)]
else:
[ax.plot(ReceiverFile[0], ReceiverFile[ColIDX+OrderPeriodicity*i], zorder=2) for i in range(NumReceivers)]
ax.set_ylabel(ylabeldict[ColIDX])
ax.set_xlabel("time (s)")
return ax
# +
def PlotTimeProfileSetFlex(ax, Set,SlipSlipRate,title,Filtered = False, absolute = False, **kwargs):
UnitsDict = {"Slip" : "Slip [m]", "SlipRate" : "Slip Rate [m/s]"}
ax.set(xlabel = 'Time [s]', ylabel = UnitsDict[SlipSlipRate],
title = title)
OrdinateVariableList=[]
for idx,item in enumerate(Set):
if (SlipSlipRate == "Slip"):
OrdinateVariableList.append([a for a in item.DispX])
elif (SlipSlipRate == "SlipRate"):
OrdinateVariableList.append([a for a in item.VelX])
if (Filtered):
OrdinateVariableList[idx] = [a for a in Butterworth(OrdinateVariableList[idx])]
if (absolute):
OrdinateVariableList[idx] = [abs(a) for a in OrdinateVariableList[idx]]
for idx,item in enumerate(Set):
ax.plot(item.Time, OrdinateVariableList[idx], **kwargs)
def PlotFlexSpecificLegend(ax, ListOfFiles,SlipSlipRate,title,Filtered=True,**kwargs):
for iidx,SingleFile in enumerate(ListOfFiles):
head, tail = os.path.split(SingleFile)
File = LoadPickleFile(Filename = tail,FolderPath = head+"/")
PlotTimeProfileSetFlex(ax, File, SlipSlipRate, title,Filtered,
zorder = iidx + 2, c = cmap(iidx/(len(ListOfFiles)+1)),
**kwargs )
return ax
# -
CompletePath = [path+r"[TPV3]Results/20201202Duo/300dx-1p-300.3delta/TPList_t280_d300.1.pickle"]
# +
fig,axis=GenericFigAxis()
format_axes(fig)
PlotType = "Slip"
#[item.PlotReference(axis[0], PlotType, filtering=True) for item in RefList] #Reference
PlotFlexSpecificLegend(axis[0], CompletePath, "Slip", "Direct")
PlotType = "SlipRate"
#[item.PlotReference(axis[1], PlotType, filtering=True) for item in RefList] #Reference
PlotFlexSpecificLegend(axis[1], CompletePath,"SlipRate","Direct")
# +
head, tail = os.path.split(CompletePath[0])
File = LoadPickleFile(Filename = tail,FolderPath = head+"/")
aaa = File[0]
# -
plt.plot(aaa.Time,aaa.DispY)
plt.plot(aaa.Time,aaa.DispX)
plt.plot(aaa.Time,aaa.VelY)
plt.plot(aaa.Time,aaa.VelX)
| PythonCodes/[TPV3]-Plotting/SingleCase Off-fault.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mrartis1/MA-MachineLearning/blob/main/DNN1Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZSD57DiQhp1G"
# #DNN1 Testing
# + id="hBF4iyyshs_S"
from __future__ import print_function
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
np.random.seed(1337) # for reproducibility
from keras.preprocessing import sequence
from keras.utils import np_utils
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Embedding
from keras.layers import LSTM, SimpleRNN, GRU
from keras.datasets import imdb
from keras.utils.np_utils import to_categorical
from sklearn.metrics import (precision_score, recall_score,f1_score, accuracy_score,mean_squared_error,mean_absolute_error)
from sklearn import metrics
from sklearn.preprocessing import Normalizer
import h5py
from keras import callbacks
from keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau, CSVLogger
# + colab={"base_uri": "https://localhost:8080/"} id="TYZSHhhVi-7h" outputId="2fdbb2ba-89b9-423a-d778-fbfec4603561"
import tensorflow as tf
print("Num GPUs Available: ", len(tf.config.list_physical_devices('GPU')))
# + colab={"base_uri": "https://localhost:8080/"} id="81kMYJI3jAlI" outputId="0294bcdf-678c-44f4-e727-a53b49a1acfa"
gpus = tf.config.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
# + id="HgBDNfBNhvhk"
traindata = pd.read_csv('/content/drive/MyDrive/Colab Results/Data/Training.csv', header=None)
testdata = pd.read_csv('/content/drive/MyDrive/Colab Results/Data/Testing.csv', header=None)
# + id="Jspp-O40hzvt"
X = traindata.iloc[:,1:42]
Y = traindata.iloc[:,0]
C = testdata.iloc[:,0]
T = testdata.iloc[:,1:42]
trainX = np.array(X)
testT = np.array(T)
trainX.astype(float)
testT.astype(float)
scaler = Normalizer().fit(trainX)
trainX = scaler.transform(trainX)
scaler = Normalizer().fit(testT)
testT = scaler.transform(testT)
y_train = np.array(Y)
y_test = np.array(C)
X_train = np.array(trainX)
X_test = np.array(testT)
batch_size = 64
# + id="HGUmRRzdh2XV"
# 1. define the network
model = Sequential()
model.add(Dense(1024,input_dim=41,activation='relu'))
model.add(Dropout(0.01))
model.add(Dense(1))
model.add(Activation('sigmoid'))
# + colab={"base_uri": "https://localhost:8080/"} id="LkCpYxUkiNjZ" outputId="d1cdefe0-3a3f-4443-bb59-19ec26ba58e7"
# try using different optimizers and different optimizer configs
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
checkpointer = callbacks.ModelCheckpoint(filepath="kddresults/dnn1layer/checkpoint-{epoch:02d}.hdf5", verbose=1, save_best_only=True, monitor='loss')
csv_logger = CSVLogger('kddresults/dnn1layer/training_set_dnnanalysis.csv',separator=',', append=False)
model.fit(X_train, y_train, batch_size=batch_size, epochs=100, callbacks=[checkpointer,csv_logger])
model.save("kddresults/dnn1layer/dnn1layer_model.hdf5")
# + id="jy0PbA-0iOyP" colab={"base_uri": "https://localhost:8080/"} outputId="776b373f-c714-495a-d913-c1ff4a453248"
score = []
name = []
from sklearn.metrics import confusion_matrix
import os
for file in os.listdir("/content/drive/MyDrive/Colab Results/kddresults/dnn1layer/"):
if file.endswith(".hdf5"):
model.load_weights("/content/drive/MyDrive/Colab Results/kddresults/dnn1layer/"+file)
y_train1 = y_test
y_pred = (model.predict(X_test) > 0.5).astype("int32")
accuracy = accuracy_score(y_train1, y_pred)
recall = recall_score(y_train1, y_pred , average="binary")
precision = precision_score(y_train1, y_pred , average="binary")
f1 = f1_score(y_train1, y_pred, average="binary")
print("----------------------------------------------")
print("accuracy")
print("%.3f" %accuracy)
print("recall")
print("%.3f" %recall)
print("precision")
print("%.3f" %precision)
print("f1score")
print("%.3f" %f1)
score.append(accuracy)
name.append(file)
# + id="6TdCfBiyiVFz" colab={"base_uri": "https://localhost:8080/"} outputId="79593637-1c20-45c8-e505-9fbd5b78c1a3"
model.load_weights("/content/drive/MyDrive/Colab Results/kddresults/dnn1layer/"+name[score.index(max(score))])
pred = (model.predict(X_test) > 0.5).astype("int32")
#proba = model.predict_proba(X_test)
np.savetxt("/content/drive/MyDrive/Colab Results/dnnres/dnn1predicted.txt", pred)
#np.savetxt("/content/drive/MyDrive/Colab Results/dnnres/dnn1probability.txt", proba)
accuracy = accuracy_score(y_test, pred)
recall = recall_score(y_test, pred , average="binary")
precision = precision_score(y_test, pred , average="binary")
f1 = f1_score(y_test, pred, average="binary")
print("----------------------------------------------")
print("accuracy")
print("%.3f" %accuracy)
print("precision")
print("%.3f" %precision)
print("racall")
print("%.3f" %recall)
print("f1score")
print("%.3f" %f1)
# + id="3EkOSGh2iYu7" colab={"base_uri": "https://localhost:8080/"} outputId="bb083713-fee1-4a37-a9bb-d8edfa634f36"
model.load_weights("/content/drive/MyDrive/Colab Results/kddresults/dnn1layer/"+name[score.index(max(score))])
pred = (model.predict(X_test) > 0.5).astype("int32")
#proba = model.predict_proba(X_test)
np.savetxt("/content/drive/MyDrive/Colab Results/dnnres/dnn1predicted.txt", pred)
#np.savetxt("/content/drive/MyDrive/Colab Results/dnnres/dnn1probability.txt", proba)
accuracy = accuracy_score(y_test, pred)
recall = recall_score(y_test, pred , average="binary")
precision = precision_score(y_test, pred , average="binary")
f1 = f1_score(y_test, pred, average="binary")
print("----------------------------------------------")
print("accuracy")
print("%.3f" %accuracy)
print("precision")
print("%.3f" %precision)
print("racall")
print("%.3f" %recall)
print("f1score")
print("%.3f" %f1)
print(model.summary())
| DNN1Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import bz2
import os
import re
import mwparserfromhell
import numpy as np
import pandas as pd
import findspark
findspark.init('/usr/lib/spark2')
from pyspark.sql import SparkSession
# -
spark = (
SparkSession.builder
.appName('Pyspark notebook (isaacj -- pagerank)')
.master('yarn')
.config(
'spark.driver.extraJavaOptions',
' '.join('-D{}={}'.format(k, v) for k, v in {
'http.proxyHost': 'webproxy.eqiad.wmnet',
'http.proxyPort': '8080',
'https.proxyHost': 'webproxy.eqiad.wmnet',
'https.proxyPort': '8080',
}.items()))
.config('spark.jars.packages', 'graphframes:graphframes:0.6.0-spark2.3-s_2.11')
.config("spark.driver.memory", "4g")
.config('spark.dynamicAllocation.maxExecutors', 128)
.config("spark.executor.memory", "8g")
.config("spark.executor.cores", 4)
.config("spark.sql.shuffle.partitions", 512)
.getOrCreate()
)
spark
snapshot = '2020-07' # data will be current to this date -- e.g., 2020-07 means data is up to 30 June 2020 (at least)
wiki = 'enwiki' # wikidb you want to run pagerank for
# +
def getLinks(wikitext):
"""Extract list of links from wikitext for an article."""
try:
wt = mwparserfromhell.parse(wikitext)
return [str(l.title).partition('#')[0].replace(' ', '_') for l in wt.filter_wikilinks()]
except Exception:
return None
spark.udf.register('getLinks', getLinks, 'Array<String>')
# +
"""
Explanation of CTEs:
* title_to_id: mapping of page title to page ID, which is a more stable identifier
* redirects: resolve redirects so the network is much denser (~6M nodes instead of ~11M nodes)
* pagelinks: extract links from wikitext and explode so each row has one link
* pagelinks_reformatted: map link page titles to link page IDs
* final: resolve redirects and rename columns to match pagerank library expectations
"""
print_for_hive = False
do_execute = True
query = """
WITH title_to_id AS (
SELECT page_id,
page_title
FROM wmf_raw.mediawiki_page
WHERE snapshot = '{0}'
AND wiki_db = '{1}'
AND page_namespace = 0
),
redirects AS (
SELECT mr.rd_from AS rd_from,
tti.page_id AS rd_to
FROM wmf_raw.mediawiki_redirect mr
INNER JOIN title_to_id tti
ON (mr.rd_title = tti.page_title)
WHERE mr.snapshot = '{0}'
AND mr.wiki_db = '{1}'
AND mr.rd_namespace = 0
),
pagelinks AS (
SELECT wt.page_id AS pl_from,
explode(getLinks(wt.revision_text)) AS pl_title_to
FROM wmf.mediawiki_wikitext_current wt
LEFT ANTI JOIN redirects r
ON (wt.page_id = r.rd_from)
WHERE wt.snapshot = '{0}'
AND wt.wiki_db = '{1}'
AND wt.page_namespace = 0
),
pagelinks_reformatted AS (
SELECT pl.pl_from AS pl_from,
tti.page_id AS pl_to
FROM pagelinks pl
INNER JOIN title_to_id tti
ON (pl.pl_title_to = tti.page_title)
)
SELECT DISTINCT pl.pl_from AS src,
COALESCE(r.rd_to, pl.pl_to) AS dst
FROM pagelinks_reformatted pl
LEFT JOIN redirects r
ON (pl.pl_to = r.rd_from)
""".format(snapshot, wiki)
if print_for_hive:
print(re.sub(' +', ' ', re.sub('\n', ' ', query)).strip())
else:
print(query)
if do_execute:
src_dst = spark.sql(query)
src_dst.createOrReplaceTempView("src_dst")
# -
src_dst.show(n=10)
# en.wikipedia.org/wiki/?curid=21349232
spark.sql('SELECT * FROM src_dst WHERE src = 21349232').show(100, False)
# +
"""
Explanation of CTEs:
* all_pageids: get set of all page IDs that show up in src or dst columns
* pageid_to_title: gather titles for each page ID because its easier to interpret
* final: join the two together
"""
print_for_hive = False
do_execute = True
query = """
WITH all_pageids AS (
SELECT DISTINCT(page_id)
FROM (
SELECT src as page_id
FROM src_dst
UNION ALL
SELECT dst as page_id
FROM src_dst
) p
),
pageid_to_title AS (
SELECT page_id,
page_title
FROM wmf_raw.mediawiki_page mp
WHERE snapshot = '{0}'
AND wiki_db = '{1}'
AND page_namespace = 0
)
SELECT p.page_id as id,
t.page_title as page_title
FROM all_pageids p
LEFT JOIN pageid_to_title t
ON (p.page_id = t.page_id)
""".format(snapshot, wiki)
if print_for_hive:
print(re.sub(' +', ' ', re.sub('\n', ' ', query)).strip())
else:
print(query)
if do_execute:
nodes = spark.sql(query)
nodes.createOrReplaceTempView("nodes")
# -
nodes.show(n=10)
# ## Run PageRank
from graphframes import *
## create graph object
g = GraphFrame(nodes, src_dst)
g.inDegrees.show(n=10)
# See: https://graphframes.github.io/graphframes/docs/_site/api/python/graphframes.html#graphframes.GraphFrame.pageRank
# Hyperparameters:
# - resetProbability (inverse of damping factor: https://en.wikipedia.org/wiki/PageRank#Damping_factor)
# - most sources suggest it should be 0.15
# - maxIter is set to 40 here as that is the parameter used in: https://www.aifb.kit.edu/images/e/e5/Wikipedia_pagerank1.pdf
# - you could also set the tolerance to 0.01 but I don't know how long that takes to converge for enwiki
# This shouldn't take more than 20-30 minutes for English Wikipedia
# There will be k jobs you can track at https://yarn.wikimedia.org/cluster/scheduler where k is the number of iterations
pr = g.pageRank(resetProbability=0.15, maxIter=40)
result = pr.vertices.sort('pagerank', ascending=False)
result.createOrReplaceTempView('pagerank')
# write pagerank results to TSV
query = """
SELECT pr.id as page_id,
pr.pagerank as pagerank,
n.page_title as page_title
FROM pagerank pr
LEFT JOIN nodes n
ON (pr.id = n.id)
"""
results = spark.sql(query)
# this will write to 512 bzipped TSVs -- they can be easily compiled into 1 via Python or just use .coalesce(1) here
# to pull onto stat machines: stat100x$ hdfs dfs -copyToLocal /user/isaacj/pagerank-enwiki/part* .
results.write.csv(path="/user/isaacj/pagerank-{0}".format(wiki), compression="bzip2", header=True, sep="\t")
# !hdfs dfs -copyToLocal pagerank-enwiki/part* file_parts/
file_parts_dir = './file_parts/'
fns = [fn for fn in os.listdir(file_parts_dir) if fn.endswith('.csv.bz2')]
history_combined = 'enwiki_pagerank_notemplates.tsv'
print_every = 1
history_length = {}
skipped = 0
processed = 0
output_header = ['page_id', 'pagerank', 'page_title']
with open(history_combined, 'w') as fout:
fout.write('\t'.join(output_header) + '\n')
for i, fn in enumerate(fns, start=1):
with bz2.open(os.path.join(file_parts_dir, fn), 'rt') as fin:
# the quote symbol " is somehow a valid username character...
header = next(fin).strip().split('\t')
assert header == output_header
for line_no, line_str in enumerate(fin, start=1):
line = line_str.strip().split('\t')
assert len(line) == len(output_header)
pid = line[0]
pagerank = line[1]
page_title = line[2]
try:
int(pid)
except ValueError:
print("PID:", line_str)
skipped += 1
continue
try:
float(pagerank)
except ValueError:
print("PR:", line_str)
skipped += 1
continue
processed += 1
fout.write(line_str)
if i % print_every == 0:
print("{0} / {1} files processed.".format(i, len(fns)))
print_every = print_every * 2
print("{0} pages processed. {1} skipped.".format(processed, skipped))
df = pd.read_csv('./enwiki_pagerank_notemplates.tsv', sep='\t')
df.sort_values('pagerank', ascending=False).head(50).set_index('page_title')['pagerank']
| pagerank-spark/pagerank-enwiki_notemplates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="b4BBN50oyiwG"
# 
#
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/jupyter/training/english/dictionary-sentiment/sentiment.ipynb)
#
# ## 0. Colab Setup
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" executionInfo={"elapsed": 64719, "status": "ok", "timestamp": 1589641853953, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14469489166467359317"}, "user_tz": -120} id="nTH23Yu1yqfD" outputId="7a6fa535-d869-48be-e49f-e4e36d7e9059"
import os
# Install java
# ! apt-get update -qq
# ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
# ! java -version
# Install pyspark
# ! pip install --ignore-installed pyspark==2.4.4
# Install Spark NLP
# ! pip install --ignore-installed spark-nlp
# + [markdown] colab_type="text" id="4Ow6rjyOyiwN"
# ## Rule-based Sentiment Analysis
#
# In the following example, we walk-through a simple use case for our straight forward SentimentDetector annotator.
#
# This annotator will work on top of a list of labeled sentences which can have any of the following features
#
# positive
# negative
# revert
# increment
# decrement
#
# Each of these sentences will be used for giving a score to text
# + [markdown] colab_type="text" id="K_1aCdWNyiwQ"
# #### 1. Call necessary imports and set the resource path to read local data files
# + colab={} colab_type="code" id="jIH8pFdPyiwS"
#Imports
import sys
sys.path.append('../../')
import sparknlp
from pyspark.sql import SparkSession
from pyspark.ml import Pipeline
from pyspark.sql.functions import array_contains
from sparknlp.annotator import *
from sparknlp.common import RegexRule
from sparknlp.base import DocumentAssembler, Finisher
# + [markdown] colab_type="text" id="58CQiS99yiwh"
# #### 2. Load SparkSession if not already there
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 132364, "status": "ok", "timestamp": 1589641921624, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14469489166467359317"}, "user_tz": -120} id="Ub7u0Z2yyiwj" outputId="bc752875-2714-41d1-8eef-483c7481ae79"
import sparknlp
spark = sparknlp.start()
print("Spark NLP version: ", sparknlp.version())
print("Apache Spark version: ", spark.version)
# + colab={"base_uri": "https://localhost:8080/", "height": 595} colab_type="code" executionInfo={"elapsed": 148313, "status": "ok", "timestamp": 1589641937579, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14469489166467359317"}, "user_tz": -120} id="XYXJ8Lrhyiwz" outputId="e3ee6935-80e6-43a6-8787-db65564c832e"
# ! rm /tmp/sentiment.parquet.zip
# ! rm -rf /tmp/sentiment.parquet
# ! wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/en/sentiment.parquet.zip -P /tmp
# ! wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/en/lemma-corpus-small/lemmas_small.txt -P /tmp
# ! wget -N https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/resources/en/sentiment-corpus/default-sentiment-dict.txt -P /tmp
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 152121, "status": "ok", "timestamp": 1589641941394, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14469489166467359317"}, "user_tz": -120} id="zu_lzjvXyiw6" outputId="aa12f375-7d0e-48e6-95f8-7c3299987d78"
# ! unzip /tmp/sentiment.parquet.zip -d /tmp/
# + colab={"base_uri": "https://localhost:8080/", "height": 459} colab_type="code" executionInfo={"elapsed": 160363, "status": "ok", "timestamp": 1589641949644, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14469489166467359317"}, "user_tz": -120} id="8ycCJ0Vyyiw_" outputId="8a2515c3-66ab-4ed5-9f46-7e0ad1944c7a"
data = spark. \
read. \
parquet("/tmp/sentiment.parquet"). \
limit(10000).cache()
data.show()
# + [markdown] colab_type="text" id="HPH7HLK8yixE"
# #### 3. Create appropriate annotators. We are using Sentence Detection, Tokenizing the sentences, and find the lemmas of those tokens. The Finisher will only output the Sentiment.
# + colab={} colab_type="code" id="rPDSRAXtyixG"
document_assembler = DocumentAssembler() \
.setInputCol("text")
sentence_detector = SentenceDetector() \
.setInputCols(["document"]) \
.setOutputCol("sentence")
tokenizer = Tokenizer() \
.setInputCols(["sentence"]) \
.setOutputCol("token")
lemmatizer = Lemmatizer() \
.setInputCols(["token"]) \
.setOutputCol("lemma") \
.setDictionary("/tmp/lemmas_small.txt", key_delimiter="->", value_delimiter="\t")
sentiment_detector = SentimentDetector() \
.setInputCols(["lemma", "sentence"]) \
.setOutputCol("sentiment_score") \
.setDictionary("/tmp/default-sentiment-dict.txt", ",")
finisher = Finisher() \
.setInputCols(["sentiment_score"]) \
.setOutputCols(["sentiment"])
# + [markdown] colab_type="text" id="3tYe_QijyixO"
# #### 4. Train the pipeline, which is only being trained from external resources, not from the dataset we pass on. The prediction runs on the target dataset
# + colab={} colab_type="code" id="o53EAomsyixQ"
pipeline = Pipeline(stages=[document_assembler, sentence_detector, tokenizer, lemmatizer, sentiment_detector, finisher])
model = pipeline.fit(data)
result = model.transform(data)
# + [markdown] colab_type="text" id="MgvlQ7TiyixV"
# #### 5. filter the finisher output, to find the positive sentiment lines
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" executionInfo={"elapsed": 163295, "status": "ok", "timestamp": 1589641952604, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14469489166467359317"}, "user_tz": -120} id="FD8jYLEsyixW" outputId="47d7fcc6-08e0-40ed-9de2-38947ba01c13"
result.where(array_contains(result.sentiment, "positive")).show(10,False)
# + colab={} colab_type="code" id="j8pjkB7Zyixd"
| jupyter/training/english/dictionary-sentiment/sentiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# Assessment from https://classroom.udacity.com/courses/ud730/lessons/6370362152/concepts/63703142310923
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
from __future__ import print_function
import matplotlib.pyplot as plt
import numpy as np
import os
import sys
import tarfile
from IPython.display import display, Image
from scipy import ndimage
from sklearn.linear_model import LogisticRegression
from six.moves.urllib.request import urlretrieve
from six.moves import cPickle as pickle
# Config the matplotlib backend as plotting inline in IPython
# %matplotlib inline
# +
url = 'http://commondatastorage.googleapis.com/books1000/'
last_percent_reported = None
def download_progress_hook(count, blockSize, totalSize):
"""A hook to report the progress of a download. This is mostly intended for users with
slow internet connections. Reports every 5% change in download progress.
"""
global last_percent_reported
percent = int(count * blockSize * 100 / totalSize)
if last_percent_reported != percent:
if percent % 5 == 0:
sys.stdout.write("%s%%" % percent)
sys.stdout.flush()
else:
sys.stdout.write(".")
sys.stdout.flush()
last_percent_reported = percent
def maybe_download(filename, expected_bytes, force=False):
"""Download a file if not present, and make sure it's the right size."""
if force or not os.path.exists(filename):
print('Attempting to download:', filename)
filename, _ = urlretrieve(url + filename, filename, reporthook=download_progress_hook)
print('\nDownload Complete!')
statinfo = os.stat(filename)
if statinfo.st_size == expected_bytes:
print('Found and verified', filename)
else:
raise Exception(
'Failed to verify ' + filename + '. Can you get to it with a browser?')
return filename
train_filename = maybe_download('notMNIST_large.tar.gz', 247336696)
test_filename = maybe_download('notMNIST_small.tar.gz', 8458043)
# +
num_classes = 10
np.random.seed(133)
def maybe_extract(filename, force=False):
root = os.path.splitext(os.path.splitext(filename)[0])[0] # remove .tar.gz
if os.path.isdir(root) and not force:
# You may override by setting force=True.
print('%s already present - Skipping extraction of %s.' % (root, filename))
else:
print('Extracting data for %s. This may take a while. Please wait.' % root)
tar = tarfile.open(filename)
sys.stdout.flush()
tar.extractall()
tar.close()
data_folders = [
os.path.join(root, d) for d in sorted(os.listdir(root))
if os.path.isdir(os.path.join(root, d))]
if len(data_folders) != num_classes:
raise Exception(
'Expected %d folders, one per class. Found %d instead.' % (
num_classes, len(data_folders)))
print(data_folders)
return data_folders
train_folders = maybe_extract(train_filename)
test_folders = maybe_extract(test_filename)
# +
image_size = 28 # Pixel width and height.
pixel_depth = 255.0 # Number of levels per pixel.
def load_letter(folder, min_num_images):
"""Load the data for a single letter label."""
image_files = os.listdir(folder)
dataset = np.ndarray(shape=(len(image_files), image_size, image_size),
dtype=np.float32)
print(folder)
num_images = 0
for image in image_files:
image_file = os.path.join(folder, image)
try:
image_data = (ndimage.imread(image_file).astype(float) -
pixel_depth / 2) / pixel_depth
if image_data.shape != (image_size, image_size):
raise Exception('Unexpected image shape: %s' % str(image_data.shape))
dataset[num_images, :, :] = image_data
num_images = num_images + 1
except IOError as e:
print('Could not read:', image_file, ':', e, '- it\'s ok, skipping.')
dataset = dataset[0:num_images, :, :]
if num_images < min_num_images:
raise Exception('Many fewer images than expected: %d < %d' %
(num_images, min_num_images))
print('Full dataset tensor:', dataset.shape)
print('Mean:', np.mean(dataset))
print('Standard deviation:', np.std(dataset))
return dataset
def maybe_pickle(data_folders, min_num_images_per_class, force=False):
dataset_names = []
for folder in data_folders:
set_filename = folder + '.pickle'
dataset_names.append(set_filename)
if os.path.exists(set_filename) and not force:
# You may override by setting force=True.
print('%s already present - Skipping pickling.' % set_filename)
else:
print('Pickling %s.' % set_filename)
dataset = load_letter(folder, min_num_images_per_class)
try:
with open(set_filename, 'wb') as f:
pickle.dump(dataset, f, pickle.HIGHEST_PROTOCOL)
except Exception as e:
print('Unable to save data to', set_filename, ':', e)
return dataset_names
train_datasets = maybe_pickle(train_folders, 45000)
test_datasets = maybe_pickle(test_folders, 1800)
# +
def make_arrays(nb_rows, img_size):
if nb_rows:
dataset = np.ndarray((nb_rows, img_size, img_size), dtype=np.float32)
labels = np.ndarray(nb_rows, dtype=np.int32)
else:
dataset, labels = None, None
return dataset, labels
def merge_datasets(pickle_files, train_size, valid_size=0):
num_classes = len(pickle_files)
valid_dataset, valid_labels = make_arrays(valid_size, image_size)
train_dataset, train_labels = make_arrays(train_size, image_size)
vsize_per_class = valid_size // num_classes
tsize_per_class = train_size // num_classes
start_v, start_t = 0, 0
end_v, end_t = vsize_per_class, tsize_per_class
end_l = vsize_per_class+tsize_per_class
for label, pickle_file in enumerate(pickle_files):
try:
with open(pickle_file, 'rb') as f:
letter_set = pickle.load(f)
# let's shuffle the letters to have random validation and training set
np.random.shuffle(letter_set)
if valid_dataset is not None:
valid_letter = letter_set[:vsize_per_class, :, :]
valid_dataset[start_v:end_v, :, :] = valid_letter
valid_labels[start_v:end_v] = label
start_v += vsize_per_class
end_v += vsize_per_class
train_letter = letter_set[vsize_per_class:end_l, :, :]
train_dataset[start_t:end_t, :, :] = train_letter
train_labels[start_t:end_t] = label
start_t += tsize_per_class
end_t += tsize_per_class
except Exception as e:
print('Unable to process data from', pickle_file, ':', e)
raise
return valid_dataset, valid_labels, train_dataset, train_labels
train_size = 200000
valid_size = 10000
test_size = 10000
valid_dataset, valid_labels, train_dataset, train_labels = merge_datasets(
train_datasets, train_size, valid_size)
_, _, test_dataset, test_labels = merge_datasets(test_datasets, test_size)
print('Training:', train_dataset.shape, train_labels.shape)
print('Validation:', valid_dataset.shape, valid_labels.shape)
print('Testing:', test_dataset.shape, test_labels.shape)
# -
def randomize(dataset, labels):
permutation = np.random.permutation(labels.shape[0])
shuffled_dataset = dataset[permutation,:,:]
shuffled_labels = labels[permutation]
return shuffled_dataset, shuffled_labels
train_dataset, train_labels = randomize(train_dataset, train_labels)
test_dataset, test_labels = randomize(test_dataset, test_labels)
valid_dataset, valid_labels = randomize(valid_dataset, valid_labels)
# +
pickle_file = 'notMNIST.pickle'
try:
f = open(pickle_file, 'wb')
save = {
'train_dataset': train_dataset,
'train_labels': train_labels,
'valid_dataset': valid_dataset,
'valid_labels': valid_labels,
'test_dataset': test_dataset,
'test_labels': test_labels,
}
pickle.dump(save, f, pickle.HIGHEST_PROTOCOL)
f.close()
except Exception as e:
print('Unable to save data to', pickle_file, ':', e)
raise
# -
statinfo = os.stat(pickle_file)
print('Compressed pickle size:', statinfo.st_size)
#
# +
train_r = train_dataset.reshape(train_dataset.shape[0],-1)
print(np.shape(train_r))
train_idx = np.lexsort(train_r.T)
print(np.shape(train_idx))
train_dataset_sanitized = train_dataset[train_idx][np.append(True,(np.diff(train_r[train_idx],axis=0)!=0).any(1))]
print(np.shape(train_dataset_sanitized))
train_labels_sanitized = train_labels[train_idx][np.append(True,(np.diff(train_r[train_idx],axis=0)!=0).any(1))]
print(np.shape(train_labels_sanitized))
valid_r = valid_dataset.reshape(valid_dataset.shape[0],-1)
valid_idx = np.lexsort(valid_r.T)
valid_dataset_sanitized = valid_dataset[valid_idx][np.append(True,(np.diff(valid_r[valid_idx],axis=0)!=0).any(1))]
valid_labels_sanitized = valid_labels[valid_idx][np.append(True,(np.diff(valid_r[valid_idx],axis=0)!=0).any(1))]
test_r = test_dataset.reshape(test_dataset.shape[0],-1)
test_idx = np.lexsort(test_r.T)
test_dataset_sanitized = test_dataset[test_idx][np.append(True,(np.diff(test_r[test_idx],axis=0)!=0).any(1))]
test_labels_sanitized = test_labels[test_idx][np.append(True,(np.diff(test_r[test_idx],axis=0)!=0).any(1))]
del train_r, valid_r, test_r
print('Training dataset has', train_dataset_sanitized.shape[0],'unique images.')
print('Validation dataset has', valid_dataset_sanitized.shape[0],'unique images.')
print('Test dataset has', test_dataset_sanitized.shape[0],'unique images.\n')
train_r = train_dataset_sanitized.reshape(train_dataset_sanitized.shape[0],-1)
valid_r = valid_dataset_sanitized.reshape(valid_dataset_sanitized.shape[0],-1)
test_r = test_dataset_sanitized.reshape(test_dataset_sanitized.shape[0],-1)
valid_dup = []
test_dup = []
train_r = {tuple(row):i for i,row in enumerate(train_r)}
for i,row in enumerate(valid_r):
if tuple(row) in train_r:
valid_dup.append(i)
for i,row in enumerate(test_r):
if tuple(row) in train_r:
test_dup.append(i)
print('Validation dataset has', len(valid_dup), 'duplicate images to training dataset.')
print('Test dataset has', len(test_dup), 'duplicate images to training dataset.\n')
valid_dataset_sanitized = np.delete(valid_dataset_sanitized, np.asarray(valid_dup), 0)
valid_labels_sanitized = np.delete(valid_labels_sanitized, np.asarray(valid_dup), 0)
test_dataset_sanitized = np.delete(test_dataset_sanitized, np.asarray(test_dup), 0)
test_labels_sanitized = np.delete(test_labels_sanitized, np.asarray(test_dup), 0)
print('Sanitized train dataset has', train_dataset_sanitized.shape[0],'images.')
print('Sanitized validation dataset has', valid_dataset_sanitized.shape[0],'images.')
print('Sanitized test dataset has', test_dataset_sanitized.shape[0],'images.')
# +
pickle_file = 'notMNIST_sanitized.pickle'
try:
f = open(pickle_file, 'wb')
save = {
'train_dataset': train_dataset_sanitized,
'train_labels': train_labels_sanitized,
'valid_dataset': valid_dataset_sanitized,
'valid_labels': valid_labels_sanitized,
'test_dataset': test_dataset_sanitized,
'test_labels': test_labels_sanitized,
}
pickle.dump(save, f, pickle.HIGHEST_PROTOCOL)
f.close()
print('Sanitized data saved to', pickle_file);
except Exception as e:
print('Unable to save data to', pickle_file, ':', e)
raise
# +
from sklearn.metrics import classification_report, confusion_matrix
def train_predict(clf, n_data, train_data, train_label, test_data, test_label):
clf.fit(train_data[:n_data,:,:].reshape(n_data,-1), train_label[:n_data])
# Predict
expected = test_label
predicted = clf.predict(test_data.reshape(test_data.shape[0],-1))
# Print Results
print('Classification Report of',n_data,'training samples:\n', classification_report(expected, predicted))
#print('Confusion Matrix of',n_data,'training samples:\n', confusion_matrix(expected, predicted))
# Create a Logistic Regression Classifier
clf = LogisticRegression(penalty='l2', tol=0.0001, C=1.0, random_state=133, solver='sag', max_iter=100, multi_class='ovr', verbose=0, n_jobs=4)
print('-------')
print(np.shape(train_dataset))
print(np.shape(train_labels))
print(np.shape(test_dataset))
print(np.shape(test_labels))
print(np.shape(valid_dataset))
print(np.shape(valid_labels))
print('-------_sanitized')
print(np.shape(train_dataset_sanitized))
print(np.shape(train_labels_sanitized))
print(np.shape(test_dataset_sanitized))
print(np.shape(test_labels_sanitized))
print(np.shape(valid_dataset_sanitized))
print(np.shape(valid_labels_sanitized))
print('-------')
# -
train_predict(clf, 50, train_dataset, train_labels, test_dataset, test_labels)
train_predict(clf, 100, train_dataset, train_labels, test_dataset, test_labels)
train_predict(clf, 100, train_dataset_sanitized, train_labels_sanitized, test_dataset_sanitized, test_labels_sanitized)
train_predict(clf, 1000, train_dataset, train_labels, test_dataset, test_labels)
print('RAW')
train_predict(clf, 5000, train_dataset, train_labels, test_dataset, test_labels)
print('SANITIZED')
train_predict(clf, 5000, train_dataset_sanitized, train_labels_sanitized, test_dataset_sanitized, test_labels_sanitized)
# Train and predict sanitized datasets
print('Starting to train on entire sanitized dataset. samples=%d' % train_dataset_sanitized.shape[0])
train_predict(clf, train_dataset_sanitized.shape[0], train_dataset_sanitized, train_labels_sanitized, test_dataset_sanitized, test_labels_sanitized)
| study/udacity-deep-learning/assignment1-datapreparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Woodwork Typing in Featuretools
#
# Featuretools relies on having consistent typing across the creation of EntitySets, Primitives, Features, and feature matrices. Previously, Featuretools used its own type system that contained objects called Variables. Now and moving forward, Featuretools will use an external data typing library for its typing: [Woodwork](https://woodwork.alteryx.com/en/stable/index.html).
#
# Understanding the Woodwork types that exist and how Featuretools uses Woodwork's type system will allow users to:
# - build EntitySets that best represent their data
# - understand the possible input and return types for Featuretools' Primitives
# - understand what features will get generated from a given set of data and primitives.
#
# Read the [Understanding Woodwork Logical Types and Semantic Tags](https://woodwork.alteryx.com/en/stable/guides/logical_types_and_semantic_tags.html) guide for an in-depth walkthrough of the available Woodwork types that are outlined below.
#
# For users that are familiar with the old `Variable` objects, the [Transitioning to Featuretools Version 1.0](../resources/transition_to_ft_v1.0.ipynb) guide will be useful for converting Variable types to Woodwork types.
#
# ## Physical Types
# Physical types define how the data in a Woodwork DataFrame is stored on disk or in memory. You might also see the physical type for a column referred to as the column’s `dtype`.
#
# Knowing a Woodwork DataFrame's physical types is important because Pandas, Dask, and Koalas rely on these types when performing DataFrame operations. Each Woodwork `LogicalType` class has a single physical type associated with it.
#
# ## Logical Types
# Logical types add additional information about how data should be interpreted or parsed beyond what can be contained in a physical type. In fact, multiple logical types have the same physical type, each imparting a different meaning that's not contained in the physical type alone.
#
# In Featuretools, a column's logical type informs how data is read into an EntitySet and how it gets used down the line in Deep Feature Synthesis.
#
# Woodwork provides many different logical types, which can be seen with the `list_logical_types` function.
# +
import featuretools as ft
ft.list_logical_types()
# -
# Featuretools will perform type inference to assign logical types to the data in EntitySets if none are provided, but it is also possible to specify which logical types should be set for any column (provided that the data in that column is compatible with the logical type).
#
# To learn more about how logical types are used in EntitySets, see the [Creating EntitySets](using_entitysets.ipynb) guide.
#
# To learn more about setting logical types directly on a DataFrame, see the Woodwork guide on [working with Logical Types](https://woodwork.alteryx.com/en/stable/guides/working_with_types_and_tags.html#Working-with-Logical-Types).
#
# ## Semantic Tags
# Semantic tags provide additional information to columns about the meaning or potential uses of data. Columns can have many or no semantic tags. Some tags are added by Woodwork, some are added by Featuretools, and users can add additional tags as they see fit.
#
# To learn more about setting semantic tags directly on a DataFrame, see the Woodwork guide on [working with Semantic Tags](https://woodwork.alteryx.com/en/stable/guides/working_with_types_and_tags.html#Working-with-Semantic-Tags).
#
# ### Woodwork-defined Semantic Tags
#
# Woodwork will add certain semantic tags to columns at initialization. These can be standard tags that may be associated with different sets of logical types or index tags. There are also tags that users can add to confer a suggested meaning to columns in Woodwork.
#
# To get a list of these tags, you can use the `list_semantic_tags` function.
ft.list_semantic_tags()
# Above we see the semantic tags that are defined within Woodwork. These tags inform how Featuretools is able to interpret data, an example of which can be seen in the `Age` primitive, which requires that the `date_of_birth` semantic tag be present on a column.
#
# The `date_of_birth` tag will not get automatically added by Woodwork, so in order for Featuretools to be able to use the `Age` primitive, the `date_of_birth` tag must be manually added to any columns to which it applies.
#
# ### Featuretools-defined Semantic Tags
#
# Just like Woodwork specifies semantic tags internally, Featuretools also defines a few tags of its own that allow the full set of Features to be generated. These tags have specific meanings when they are present on a column.
#
# - `'last_time_index'` - added by Featuretools to the last time index column of a DataFrame. Indicates that this column has been created by Featuretools.
# - `'foreign_key'` - used to indicate that this column is the child column of a relationship, meaning that this column is related to a corresponding index column of another dataframe in the EntitySet.
#
#
# ## Woodwork Throughout Featuretools
#
# Now that we've described the elements that make up Woodwork's type system, lets see them in action in Featuretools.
#
# ### Woodwork in EntitySets
# For more information on building EntitySets using Woodwork, see the [EntitySet guide](using_entitysets.ipynb).
#
# Let's look at the Woodwork typing information as it's stored in a demo EntitySet of retail data:
es = ft.demo.load_retail()
es
# Woodwork typing information is not stored in the EntitySet object, but rather is stored in the individual DataFrames that make up the EntitySet. To look at the Woodwork typing information, we first select a single DataFrame from the EntitySet, and then access the Woodwork information via the `ww` namespace:
df = es['products']
df.head()
df.ww
# Notice how the three columns showing this DataFrame's typing information are the three elements of typing information outlined at the beginning of this guide. To reiterate: By defining physical types, logical types, and semantic tags for each column in a DataFrame, we've defined a DataFrame's Woodwork schema, and with it, we can gain an understanding of the contents of each column.
#
# This column-specific typing information that exists for every column in every DataFrame in an EntitySet is an integral part of Deep Feature Synthesis' ability to generate features for an EntitySet.
#
# ### Woodwork in DFS
# As the units of computation in Featuretools, Primitives need to be able to specify the input types that they allow as well as have a predictable return type. For an in-depth explanation of Primitives in Featuretools, see the [Feature Primitives](primitives.ipynb) guide. Here, we'll look at how the Woodwork types come together into a `ColumnSchema` object to describe Primitive input and return types.
#
# Below is a Woodwork `ColumnSchema` that we've obtained from the `'product_id'` column in the `products` DataFrame in the retail EntitySet.
products_df = es['products']
product_ids_series = products_df.ww['product_id']
column_schema = product_ids_series.ww.schema
column_schema
# This combination of logical type and semantic tag typing information is a `ColumnSchema`. In the case above, the `ColumnSchema` describes the **type definition** for a single column of data.
#
# Notice that there is no physical type in a `ColumnSchema`. This is because a `ColumnSchema` is a collection of Woodwork types that doesn't have any data tied to it and therefore has no physical representation. Because a `ColumnSchema` object is not tied to any data, it can also be used to describe a **type space** into which other columns may or may not fall.
#
# This flexibility of the `ColumnSchema` class allows `ColumnSchema` objects to be used both as type definitions for every column in an EntitySet as well as input and return type spaces for every Primitive in Featuretools.
#
# Let's look at a different column in a different DataFrame to see how this works:
order_products_df = es['order_products']
order_products_df.head()
quantity_series = order_products_df.ww['quantity']
column_schema = quantity_series.ww.schema
column_schema
# The `ColumnSchema` above has been pulled from the `'quantity'` column in the `order_products` DataFrame in the retail EntitySet. This is a **type definition**.
#
# If we look at the Woodwork typing information for the `order_products` DataFrame, we can see that there are several columns that will have similar `ColumnSchema` type definitions. If we wanted to describe subsets of those columns, we could define several `ColumnSchema` **type spaces**
es['order_products'].ww
# Below are several `ColumnSchema`s that all would include our `quantity` column, but each of them describes a different type space. These `ColumnSchema`s get more restrictive as we go down:
#
# ##### Entire DataFrame
# No restrictions have been placed; any column falls into this definition. This would include the whole DataFrame.
# +
from woodwork.column_schema import ColumnSchema
ColumnSchema()
# -
# An example of a Primitive with this `ColumnSchema` as its input type is the `IsNull` transform primitive.
#
# ##### By Semantic Tag
# Only columns with the `numeric` tag apply. This can include Double, Integer, and Age logical type columns as well. It will not include the `index` column which, despite containing integers, has had its standard tags replaced by the `'index'` tag.
ColumnSchema(semantic_tags={'numeric'})
df = es['order_products'].ww.select(include='numeric')
df.ww
# And example of a Primitive with this `ColumnSchema` as its input type is the `Mean` aggregation primitive.
#
# ##### By Logical Type
# Only columns with logical type of `Integer` are included in this definition. Does not require the `numeric` tag, so an index column (which has its standard tags removed) would still apply.
# +
from woodwork.logical_types import Integer
ColumnSchema(logical_type=Integer)
# -
df = es['order_products'].ww.select(include='Integer')
df.ww
# ##### By Logical Type and Semantic Tag
# The column must have logical type `Integer` and have the `numeric` semantic tag, excluding index columns.
ColumnSchema(logical_type=Integer, semantic_tags={'numeric'})
df = es['order_products'].ww.select(include='numeric')
df = df.ww.select(include='Integer')
df.ww
# In this way, a `ColumnSchema` can define a type space under which columns in a Woodwork DataFrame can fall. This is how Featuretools determines which columns in a DataFrame are valid for a Primitive in building Features during DFS.
#
# Each Primitive has `input_types` and a `return_type` that are described by a Woodwork `ColumnSchema`. Every DataFrame in an EntitySet has Woodwork initialized on it. This means that when an EntitySet is passed into DFS, Featuretools can select the relevant columns in the DataFrame that are valid for the Primitive's `input_types`. We then get a Feature that has a `column_schema` property that indicates what that Feature's typing definition is in a way that lets DFS stack features on top of one another.
#
# In this way, Featuretools is able to leverage the base unit of Woodwork typing information, the `ColumnSchema`, and use it in concert with an EntitySet of Woodwork DataFrames in order to build Features with Deep Feature Synthesis.
| docs/source/getting_started/woodwork_types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Example quantitative plots
#
# How to plot the results of the quantitative evaluation.
# +
import numpy as np
import os
import fnmatch
import pandas as pd
import sklearn.metrics as sm
import scipy.stats as ss
import matplotlib.pyplot as plt
import dense_correspondence_manipulation.utils.utils as utils
utils.add_dense_correspondence_to_python_path()
from dense_correspondence.evaluation.evaluation import DenseCorrespondenceEvaluationPlotter as DCEP
# -
# If you have multiple networks trained, you can add them to the `nets_list` below, and they will be plotted together.
# +
folder_name = "tutorials"
path_to_nets = os.path.join("code/data_volume/pdc/trained_models", folder_name)
path_to_nets = utils.convert_to_absolute_path(path_to_nets)
all_nets = sorted(os.listdir(path_to_nets))
nets_to_plot = []
nets_list = ["caterpillar_3"]
for net in nets_list:
nets_to_plot.append(os.path.join(folder_name,net))
# -
# # Training
# Evaluate the network on the training scenes. Correspondences are all within scene
# +
p = DCEP()
dc_source_dir = utils.getDenseCorrespondenceSourceDir()
network_name = nets_to_plot[0]
path_to_csv = os.path.join(dc_source_dir, "data_volume", "pdc", "trained_models", network_name, "analysis/train/data.csv")
fig_axes = DCEP.run_on_single_dataframe(path_to_csv, label=network_name, save=False)
for network_name in nets_to_plot[1:]:
path_to_csv = os.path.join(dc_source_dir, "data_volume", "pdc", "trained_models", network_name, "analysis/train/data.csv")
fig_axes = DCEP.run_on_single_dataframe(path_to_csv, label=network_name, previous_fig_axes=fig_axes, save=False)
_, axes = fig_axes
# axes[0].set_title("Training Set")
plt.show()
# -
# # Test
# Evaluate the network on the test scenes. Correspondences are all within scene
# +
p = DCEP()
dc_source_dir = utils.getDenseCorrespondenceSourceDir()
network_name = nets_to_plot[0]
path_to_csv = os.path.join(dc_source_dir, "data_volume", "pdc", "trained_models", network_name, "analysis/test/data.csv")
fig_axes = DCEP.run_on_single_dataframe(path_to_csv, label=network_name, save=False)
for network_name in nets_to_plot[1:]:
path_to_csv = os.path.join(dc_source_dir, "data_volume", "pdc", "trained_models", network_name, "analysis/test/data.csv")
fig_axes = DCEP.run_on_single_dataframe(path_to_csv, label=network_name, previous_fig_axes=fig_axes, save=False)
_, axes = fig_axes
# axes[0].set_title("Test Set")
plt.show()
# -
# ## Cross Scene Single Object
# Evaluate the network on correspondences that come from different scenes. These correspondences were manually annotated only for evaluation purposes.
# +
p = DCEP()
dc_source_dir = utils.getDenseCorrespondenceSourceDir()
network_name = nets_to_plot[0]
path_to_csv = os.path.join(dc_source_dir, "data_volume", "pdc", "trained_models", network_name, "analysis/cross_scene/data.csv")
fig_axes = DCEP.run_on_single_dataframe(path_to_csv, label=network_name, save=False)
for network_name in nets_to_plot[1:]:
path_to_csv = os.path.join(dc_source_dir, "data_volume", "pdc", "trained_models", network_name, "analysis/cross_scene/data.csv")
fig_axes = DCEP.run_on_single_dataframe(path_to_csv, label=network_name, previous_fig_axes=fig_axes, save=False)
_, axes = fig_axes
# axes[0].set_title("Cross Scene Set")
plt.show()
| dense_correspondence/evaluation/evaluation_quantitative_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Author: <NAME>
# email: <EMAIL>
# Date: 19.01.2019
# State: Stable
# # Demonstration of Finnish Business Portal
# This file demonstrates the use of Finnish Business Portal Python API. In order to replicate the demo, please make sure you have appropriate version of Python 3 and the libraries Pandas and Requests and their dependencies. After this, download this application from the source.
#
# ## Navigation:
# * [Setup](#H0)
# * [Import the module](#H0_imports)
# * [Parameters](#H1)
# * [Example Searches](#H20)
# * [Example Application](#H3)
# ## Setup<a class="anchor" id="H0"></a>
# +
import datetime
import platform
import os
import sys
print(f'Python version: {platform.python_version()}')
print(f'''
Time of Run:
Year: {datetime.datetime.now().year}
Month: {datetime.datetime.now().month}
Day: {datetime.datetime.now().day}
''')
# -
# ### Import the module<a class="anchor" id="H0_imports"></a>
import finnish_business_portal as busportal
# ## Parameters<a class="anchor" id="H1"></a>
# The API options can be found using "api_infos" or "api_options" attributes of the class. You can copy-paste the string as an initial argument for the class initiation. This is the only required argument.
busportal.SearchModel.api_infos
busportal.SearchModel.api_options
portal = busportal.SearchModel("BisCompany")
# You can get allowed parameters for your selected API using method ".parameter_options". For more detailed information, use ".parameter_infos".
portal.parameter_options
# Returning only two parameters for clarity
{key: val for i, (key, val) in enumerate(portal.parameter_infos.items()) if i < 2}
# There is also a method ".help()" to give thorough information of the APIs.
# There are also other keyword arguments you can use in the class initiation.
#
# These are:
# - wait_time: seconds to wait before each call to slow down querying. Be nice to the APIs. (default: 2)
# - loop_results: True to loop all found entries (causing multiple additional queries), False to return the first query (default: False)
# - deep: True to go through all detailed URLs giving more data but more queries (default: False)
# ## Example Searches<a class="anchor" id="H20"></a>
portal = busportal.SearchModel("BisCompany")
portal.search(name=["Fortum", "Nokia"], companyForm="oy")
# The search returned itself and no any data. The data is in the attribute "results" but by default this is just a list of dictionaries. To turn the data to table format, use to_frame() (requires Pandas) and then access the results attribute.
portal.to_frame().results
# The class does not loop the results by default and is showing only the number of companies specified in max_results. To get all of the companies,
# a) increase "max_results" (may crash the query if too large) or
# b) set "loop_results_" to True
portal = busportal.SearchModel("BisCompany", loop_results=True)
portal.search(name=["Fortum", "Nokia"], companyForm="oy")
portal.to_frame().results
portal = busportal.SearchModel("BisCompany", loop_results=False, deep=True)
portal.search(name=["Fortum", "Nokia"], companyForm="oy", total_results="True")
portal.to_frame().results
# And finally testing with other API.
portal.api = "TradeRegisterCompany"
portal.parameter_options
portal.search(name="Renk", company_registration_from="2000-01-01")
portal.to_frame().results
# Some of the columns may seem to be confusing as they are list of dictionaries. You can "flat" them passing True to the to_frame method.
portal.to_frame(True).results
# But on the other hands, they were list of dicts for a reason.
# ## Example Application<a class="anchor" id="H3"></a>
import pandas as pd
# Using a bit of data manipulation with Pandas, one can put the data any desirable format.
portal = busportal.SearchModel("BisCompany", loop_results=False, deep=True)
portal.search(name="Fortum", companyForm="oy", companyRegistrationFrom="2000-01-01")
df_fortum = portal.to_frame(True).results
# If case you need to get street addresses from a company/companies, you can do it like the following:
first_level = "address"
second_level = ("street", "postCode", "city", "country", "endDate")
(
df_fortum[
[col for col in df_fortum.columns
if col[0].startswith(first_level)
and col[1] in second_level]
]
.stack(0)
).join(df_fortum["name"])
# What happens here is that we take only the columns that have word "address" in the first level (ie. addresses 0) and the second level must in the list "street", "postCode", "city", "country" or "endDate". Then we put the first level as index level ("stack(0)") and join the name column to the results.
#
# Note that the data is straight from the API and may have confusing fields (ie. there might not be a city named as "Fortum"). This has nothing to do with this software.
#
# Easy and convenient way to transform the data. We can also use the same method with other columns:
first_level = "businessIdChanges"
(
df_fortum[
[col for col in df_fortum.columns if col[0].startswith(first_level)]
]
.stack(0)
.dropna(axis=1, how="all")
)
# The codes can also easily be turned in the full name. Please see the API documentation for the codings.
(
df_fortum[
[col for col in df_fortum.columns if col[0].startswith("contactDetails")]
]
.stack(0)
.query('language == "EN" & endDate != endDate')
.dropna(how="all", axis=1)
)
# Thanks to the versatileness of Pandas, you can also query the data in single line. Note that NaN values are never equal to themselves thus you can query them with using the logic.
# ## Additional Links
# The meaning of "entryCodes" and "typeOfRegistration" can be found from the API's website.
# - Entry codes: http://avoindata.prh.fi/tr-codes_v1.fi.txt
# - Type of Registration: http://avoindata.prh.fi/tr-type_v1.fi.txt
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:wildfires] *
# language: python
# name: conda-env-wildfires-py
# ---
# ## Setup
from specific import *
# ### Get shifted data
(
endog_data,
exog_data,
master_mask,
filled_datasets,
masked_datasets,
land_mask,
) = get_offset_data()
client = get_client()
client
# ### Define the training and test data
# +
@data_split_cache
def get_split_data():
X_train, X_test, y_train, y_test = train_test_split(
exog_data, endog_data, random_state=1, shuffle=True, test_size=0.3
)
return X_train, X_test, y_train, y_test
X_train, X_test, y_train, y_test = get_split_data()
# -
# ### Specific model training without grid seach
rf = get_model(X_train, y_train)
# +
rf.n_jobs = get_ncpus()
with parallel_backend("threading", n_jobs=get_ncpus()):
y_pred = rf.predict(X_test)
y_train_pred = rf.predict(X_train)
print("Test R2:", r2_score(y_test, y_pred))
print("Test MSE:", mean_squared_error(y_test, y_pred))
print("Train R2:", r2_score(y_train, y_train_pred))
print("Train MSE:", mean_squared_error(y_train, y_train_pred))
| analyses/seasonality_paper/no_temporal_shifts/model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 2 - Making materials from elements
#
# As we saw in Part 1, materials can be defined in OpenMC using isotopes. However, materials can also be made from elements - this is more concise and still supports isotopic enrichment.
#
# This python notebook allows users to create different materials from elements using OpenMC.
# The following code block is a simple example of creating a material (water H2O) from elements. (Note how Hydrogen and Oxygen elements have been specified rather than each specific isotope).
# +
import openmc
# Making water from elements
water_mat = openmc.Material(name='water')
water_mat.add_element('H', 2.0, percent_type='ao')
water_mat.add_element('O', 1.0, percent_type='ao')
water_mat.set_density('g/cm3', 0.99821)
water_mat
# -
# The next code block is an example of making a ceramic breeder material.
# +
# Making Li4SiO4 from elements
Li4SiO4_mat = openmc.Material(name='lithium_orthosilicate')
Li4SiO4_mat.add_element('Li', 4.0, percent_type='ao')
Li4SiO4_mat.add_element('Si', 1.0, percent_type='ao')
Li4SiO4_mat.add_element('O', 4.0, percent_type='ao')
Li4SiO4_mat.set_density('g/cm3', 2.32)
Li4SiO4_mat
# -
# It is also possible to enrich specific isotopes while still benefitting from the concise code of making materials from elements.
#
# Here is an example of making the same ceramic breeder material but this time with Li6 enrichment.
# +
# Making enriched Li4SiO4 from elements with enrichment of Li6 enrichment
Li4SiO4_mat = openmc.Material(name='lithium_orthosilicate')
Li4SiO4_mat.add_element('Li', 4.0, percent_type='ao',
enrichment=60,
enrichment_target='Li6',
enrichment_type='ao'
)
Li4SiO4_mat.add_element('Si', 1.0, percent_type='ao')
Li4SiO4_mat.add_element('O', 4.0, percent_type='ao')
Li4SiO4_mat.set_density('g/cm3', 2.32) # this would actually be lower than 2.32 g/cm3 but this would need calculating
Li4SiO4_mat
# -
# In the case of materials that can be represented as a chemical formula (e.g. 'H2O', 'Li4SiO4') there is an even more concise way of making these materials by using their chemical formula.
# +
# making Li4SiO4 from a formula
Li4SiO4_mat = openmc.Material(name='lithium_orthosilicate')
Li4SiO4_mat.add_elements_from_formula('Li4SiO4')
Li4SiO4_mat
# -
# This add_elements_from_formula (which was added to OpenMC source code by myself) can also support enrichment.
# +
# making Li4SiO4 from a formula with enrichment
Li4SiO4_mat = openmc.Material(name='lithium_orthosilicate')
Li4SiO4_mat.add_elements_from_formula('Li4SiO4',
enrichment=60,
enrichment_target='Li6',
enrichment_type='ao'
)
Li4SiO4_mat
# -
# Making more detailed materials such as a low activation steel Eurofer would require about 20 elements. While this is fewer user inputs than making the material from isotopes it is still quite a lot of coding for the user. Unfortunately, they cannot be input as a chemical formula either.
# **Learning Outcomes for Part 2:**
# - Materials can be made in OpenMC using element fractions and densities.
# - Making materials from elements is more concise than making materials from isotopes.
# - If materials can be represented as a chemical formula, OpenMC also offers a way to construct those materials from that.
# - Making materials from elements also supports isotopic enrichment.
| tasks/task_02_making_materials/2_example_materials_from_elements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
# + [markdown] origin_pos=0
# # 多GPU的简洁实现
# :label:`sec_multi_gpu_concise`
#
# 每个新模型的并行计算都从零开始实现是无趣的。此外,优化同步工具以获得高性能也是有好处的。下面我们将展示如何使用深度学习框架的高级API来实现这一点。数学和算法与 :numref:`sec_multi_gpu`中的相同。不出所料,你至少需要两个GPU来运行本节的代码。
#
#
# + origin_pos=2 tab=["pytorch"]
# 在notebook下面单GPU可以执行,多GPU会提示使用python -m paddle.distributed.launch xx.py格式方式执行。
import paddle
from paddle import nn
import d2l.paddle as d2l
# 导入必要并行训练的依赖包
import paddle.distributed as dist
paddle.set_device("gpu") # 多卡并行需要设置这句
# + [markdown] origin_pos=3
# ## [**简单网络**]
#
# 让我们使用一个比 :numref:`sec_multi_gpu`的LeNet更有意义的网络,它依然能够容易地和快速地训练。我们选择的是 :cite:`He.Zhang.Ren.ea.2016`中的ResNet-18。因为输入的图像很小,所以稍微修改了一下。与 :numref:`sec_resnet`的区别在于,我们在开始时使用了更小的卷积核、步长和填充,而且删除了最大汇聚层。
#
# + origin_pos=5 tab=["pytorch"]
#@save
def resnet18(num_classes, in_channels=1):
"""稍加修改的ResNet-18模型"""
def resnet_block(in_channels, out_channels, num_residuals,
first_block=False):
blk = []
for i in range(num_residuals):
if i == 0 and not first_block:
blk.append(d2l.Residual(in_channels, out_channels,
use_1x1conv=True, strides=2))
else:
blk.append(d2l.Residual(out_channels, out_channels))
return nn.Sequential(*blk)
# 该模型使用了更小的卷积核、步长和填充,而且删除了最大汇聚层
net = nn.Sequential(
nn.Conv2D(in_channels, 64, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2D(64),
nn.ReLU())
net.add_sublayer("resnet_block1", resnet_block(
64, 64, 2, first_block=True))
net.add_sublayer("resnet_block2", resnet_block(64, 128, 2))
net.add_sublayer("resnet_block3", resnet_block(128, 256, 2))
net.add_sublayer("resnet_block4", resnet_block(256, 512, 2))
net.add_sublayer("global_avg_pool", nn.AdaptiveAvgPool2D((1, 1)))
net.add_sublayer("fc", nn.Sequential(nn.Flatten(),
nn.Linear(512, num_classes)))
return net
# + [markdown] origin_pos=6
# ## 网络初始化
#
# + [markdown] origin_pos=8 tab=["pytorch"]
# 我们将在训练代码部分初始化网络。
# ```
# # 初始化并行计算环境
# dist.init_parallel_env()
#
# net = resnet18(10) # resnet10分类
# # 模型设置为数据并行模式
# net = paddle.DataParallel(net)
# # 模型weight参数初始化,飞桨里默认初始化效果也不错,可以省略
# init_normal = nn.initializer.Normal(mean=0.0, std=0.01)
# for i in net.sublayers():
# if type(i) in [nn.Linear, nn.Conv2D]:
# init_normal(i.weight)
#
# # 配置优化器
# trainer = paddle.optimizer.SGD(parameters=net.parameters(), learning_rate=lr)
# # 配置loss
# loss = nn.CrossEntropyLoss()
#
#
# ```
# + [markdown] origin_pos=17
# ## [**训练**]
#
# 如前所述,用于训练的代码需要执行几个基本功能才能实现高效并行:
#
# * 需要在所有设备上初始化网络参数。
# * 在数据集上迭代时,要将小批量数据分配到所有设备上。
# * 跨设备并行计算损失及其梯度。
# * 聚合梯度,并相应地更新参数。
#
# 最后,并行地计算精确度和发布网络的最终性能。除了需要拆分和聚合数据外,训练代码与前几章的实现非常相似。
#
# 当然,飞桨已经通过并行训练套件帮我们实现了,我们只需要导入和初始化数据并行训练,配置网络模型数据并行,然后写入一个py文件,最后在命令行下使用`python -m paddle.distributed.launch xx.py`直接执行即可。
#
# + origin_pos=19 tab=["pytorch"]
def train(batch_size, lr):
devices = d2l.try_all_gpus() # 获得GPU字符串,只是为了最后显示GPU列表,不影响实际训练。
# 初始化并行计算环境
dist.init_parallel_env()
net = resnet18(10) # resnet10分类
# 模型设置为数据并行模式
net = paddle.DataParallel(net)
# 模型weight参数初始化,飞桨里默认初始化效果也不错,可以省略
init_normal = nn.initializer.Normal(mean=0.0, std=0.01)
for i in net.sublayers():
if type(i) in [nn.Linear, nn.Conv2D]:
init_normal(i.weight)
# 配置优化器
trainer = paddle.optimizer.SGD(parameters=net.parameters(), learning_rate=lr)
# 配置loss
loss = nn.CrossEntropyLoss()
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
# 设置训练循环次数为10
timer, num_epochs = d2l.Timer(), 10
animator = d2l.Animator('epoch', 'test acc', xlim=[1, num_epochs])
for epoch in range(num_epochs):
net.train()
timer.start()
for batch_id, data in enumerate(train_iter):
img, label = data
label.stop_gradient = True
out = net(img)
l = loss(input=out, label=label)
avg_loss = paddle.mean(x=l)
# 以下两句是为了终端输出精度数据,去掉相关代码不影响实际训练
acc_top1 = paddle.metric.accuracy(input=out, label=label, k=1)
acc_top5 = paddle.metric.accuracy(input=out, label=label, k=5)
if batch_id % 50 == 0: # 每50个batch显示训练信息
print("[Epoch %d, batch %d] loss: %.5f, acc1: %.5f, acc5: %.5f" % (
epoch, batch_id, avg_loss, acc_top1, acc_top5))
l.backward()
trainer.step()
trainer.clear_gradients()
timer.stop()
animator.add(epoch + 1, (d2l.evaluate_accuracy_gpu(net, test_iter),))
print(f'测试精度:{animator.Y[0][-1]:.2f},{timer.avg():.1f}秒/轮,'
f'在{str(devices)}')
# + [markdown] origin_pos=20
# 让我们看看这在实践中是如何运作的。我们先[**在单个GPU上训练网络**]进行预热。
#
# 在notebook环境下使用`train(batch_size=256, lr=0.1)`。
#
#
# + origin_pos=22 tab=["pytorch"]
# 单卡测试
train(batch_size=256, lr=0.1)
# + [markdown] origin_pos=23
# 接下来我们[**使用2个GPU进行训练**]。与 :numref:`sec_multi_gpu`中评估的LeNet相比,ResNet-18的模型要复杂得多。这就是显示并行化优势的地方,计算所需时间明显大于同步参数需要的时间。因为并行化开销的相关性较小,因此这种操作提高了模型的可伸缩性。
#
# 飞桨的多卡训练,需要在命令行下使用`python -m paddle.distributed.launch xx.py`方式执行。同时飞桨的多卡并行非常智能,会自动识别GPU卡的数量,默认使用全部GPU卡进行训练!
#
# 下面我们使用`%%writefile multigpu.py`魔法命令将代码写入文件,然后在notebook中使用`!python -m paddle.distributed.launch multigpu.py`执行即可。
#
# 刚才我们知道了,飞桨会自动识别GPU卡的数量,那么神奇的地方就是,在不修改代码和参数,也不修改执行命令的情况下,我们前面写的代码在单卡环境、双卡环境和多卡环境下都能正常运行!
#
#
# + origin_pos=25 tab=["pytorch"]
# %%writefile multigpu.py
# 飞桨多GPU程序,不管单GPU环境还是多GPU环境,都可以在不修改代码,不修改参数,不修改执行命令的情况下正常执行。
import paddle
from paddle import nn
import d2l.paddle as d2l
# 导入并行训练的依赖包
import paddle.distributed as dist
paddle.set_device("gpu") # 多卡并行需要设置这句
def resnet18(num_classes, in_channels=1):
"""稍加修改的ResNet-18模型"""
def resnet_block(in_channels, out_channels, num_residuals,
first_block=False):
blk = []
for i in range(num_residuals):
if i == 0 and not first_block:
blk.append(d2l.Residual(in_channels, out_channels,
use_1x1conv=True, strides=2))
else:
blk.append(d2l.Residual(out_channels, out_channels))
return nn.Sequential(*blk)
# 该模型使用了更小的卷积核、步长和填充,而且删除了最大汇聚层
net = nn.Sequential(
nn.Conv2D(in_channels, 64, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2D(64),
nn.ReLU())
net.add_sublayer("resnet_block1", resnet_block(
64, 64, 2, first_block=True))
net.add_sublayer("resnet_block2", resnet_block(64, 128, 2))
net.add_sublayer("resnet_block3", resnet_block(128, 256, 2))
net.add_sublayer("resnet_block4", resnet_block(256, 512, 2))
net.add_sublayer("global_avg_pool", nn.AdaptiveAvgPool2D((1,1)))
net.add_sublayer("fc", nn.Sequential(nn.Flatten(),
nn.Linear(512, num_classes)))
return net
# 训练
def train(batch_size, lr):
devices = d2l.try_all_gpus() # 获得GPU字符串,只是为了最后显示GPU列表,不影响实际训练。
# 初始化并行计算环境
dist.init_parallel_env()
net = resnet18(10) # resnet10分类
# 模型设置为数据并行模式
net = paddle.DataParallel(net)
# 模型weight参数初始化,飞桨里默认初始化效果也不错,可以省略
init_normal = nn.initializer.Normal(mean=0.0, std=0.01)
for i in net.sublayers():
if type(i) in [nn.Linear, nn.Conv2D]:
init_normal(i.weight)
# 配置优化器
trainer = paddle.optimizer.SGD(parameters=net.parameters(), learning_rate=lr)
# 配置loss
loss = nn.CrossEntropyLoss()
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size)
# 设置训练循环次数为10
timer, num_epochs = d2l.Timer(), 10
animator = d2l.Animator('epoch', 'test acc', xlim=[1, num_epochs])
for epoch in range(num_epochs):
net.train()
timer.start()
for batch_id, data in enumerate(train_iter):
img, label = data
label.stop_gradient = True
out = net(img)
l = loss(input=out, label=label)
avg_loss = paddle.mean(x=l)
# 以下两句是为了终端输出精度数据,去掉相关代码不影响实际训练
acc_top1 = paddle.metric.accuracy(input=out, label=label, k=1)
acc_top5 = paddle.metric.accuracy(input=out, label=label, k=5)
if batch_id % 50 == 0: # 每50个batch显示训练信息
print("[Epoch %d, batch %d] loss: %.5f, acc1: %.5f, acc5: %.5f" % (
epoch, batch_id, avg_loss, acc_top1, acc_top5))
l.backward()
trainer.step()
trainer.clear_gradients()
timer.stop()
animator.add(epoch + 1, (d2l.evaluate_accuracy_gpu(net, test_iter),))
print(f'测试精度:{animator.Y[0][-1]:.2f},{timer.avg():.1f}秒/轮,'
f'在{str(devices)}')
if __name__ == "__main__" :
# 若为多GPU,学习率需要乘以相应倍数以达到较好效果,比如2GPU lr=0.2 4GPU lr=0.4
train(batch_size=256, lr=0.2)
# -
# 不管单卡还是多卡环境,下面命令都可以正常执行!
# !python -m paddle.distributed.launch multigpu.py
# + [markdown] origin_pos=26
# ## 小结
#
# + [markdown] origin_pos=28 tab=["pytorch"]
# * 飞桨的多卡并行训练非常简单方便,不用调整原来的网络结构,只要加上三处代码即可。
#
# ```
# 导入并行训练需要的依赖包。
#
# 初始化并行环境。
#
# 设置数据并行的模型。
#
# ```
#
# * 所有的数据并行处理,以及GPU卡的管理,统一由并行模块处理,省心省力。
# * 飞桨的多卡并行训练代码不用任何修改,就可以适应单卡、双卡以及多卡的各种环境。
#
# + [markdown] origin_pos=29
# ## 练习
#
# + [markdown] origin_pos=31 tab=["pytorch"]
# 1. 本节使用ResNet-18,请尝试不同的迭代周期数、批量大小和学习率,以及使用更多的GPU进行计算。如果使用$16$个GPU(例如,在AWS p2.16xlarge实例上)尝试此操作,会发生什么?
# 1. 有时候不同的设备提供了不同的计算能力,我们可以同时使用GPU和CPU,那应该如何分配工作?为什么?
#
# + [markdown] origin_pos=33 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/2803)
#
| Dive-into-DL-paddlepaddle/docs/12_computational-performance/12.6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Nn885eRyu7ND"
# ### Easy string manipulation
# + colab={} colab_type="code" id="7CPBUMfm7Ljn"
x = 'a string'
y = "a string"
if x == y:
print("they are the same")
# + colab={} colab_type="code" id="Ani3HhQk7LkF"
fox = "tHe qUICk bROWn fOx."
# + [markdown] colab_type="text" id="Lr1xNTks7LkN"
# To convert the entire string into upper-case or lower-case, you can use the ``upper()`` or ``lower()`` methods respectively:
# + colab={} colab_type="code" id="hLEjP_-r7LkP"
fox.upper()
# + colab={} colab_type="code" id="J_sWSEtc7LkV"
fox.lower()
# + [markdown] colab_type="text" id="gDJiQgTK7LkZ"
# A common formatting need is to capitalize just the first letter of each word, or perhaps the first letter of each sentence.
# This can be done with the ``title()`` and ``capitalize()`` methods:
# + colab={} colab_type="code" id="-VFGL3Hr7Lka"
fox.title()
# + colab={} colab_type="code" id="cV3ghJce7Lkr"
fox.capitalize()
# + [markdown] colab_type="text" id="2CjcAH_47Lkw"
# The cases can be swapped using the ``swapcase()`` method:
# + colab={} colab_type="code" id="9VqNWnnG7Lkx"
fox.swapcase()
# + colab={} colab_type="code" id="VXnRg_RV7Lk8"
line = ' this is the content '
line.strip()
# + [markdown] colab_type="text" id="JGI2dfcr7LlA"
# To remove just space to the right or left, use ``rstrip()`` or ``lstrip()`` respectively:
# + colab={} colab_type="code" id="wNtxgkpd7LlC"
line.rstrip()
# + colab={} colab_type="code" id="mqAp9fhT7LlH"
line.lstrip()
# + [markdown] colab_type="text" id="0Y1XeOHR7LlK"
# To remove characters other than spaces, you can pass the desired character to the ``strip()`` method:
# + colab={} colab_type="code" id="4ZoSwbiO7LlL"
num = "000000000000435"
num.strip('0')
# + colab={} colab_type="code" id="fDyZR0R97LlX"
line = 'the quick brown fox jumped over a lazy dog'
line.find('fox')
# + colab={} colab_type="code" id="BIArRU7l7Llh"
line.index('fox')
# + colab={} colab_type="code" id="D58dXOqH7Llb"
line[16:21]
# + [markdown] colab_type="text" id="h85MmPIU7Llm"
# The only difference between ``find()`` and ``index()`` is their behavior when the search string is not found; ``find()`` returns ``-1``, while ``index()`` raises a ``ValueError``:
# + colab={} colab_type="code" id="_1KC6QOU7Lln"
line.find('bear')
# + colab={} colab_type="code" id="L-3u--pe7Lls"
line.index('bear')
# + colab={} colab_type="code" id="broQs8Q27Ll-"
line.partition('fox')
# + [markdown] colab_type="text" id="52RL7jri7Ll_"
# The ``rpartition()`` method is similar, but searches from the right of the string.
#
# The ``split()`` method is perhaps more useful; it finds *all* instances of the split-point and returns the substrings in between.
# The default is to split on any whitespace, returning a list of the individual words in a string:
# + colab={} colab_type="code" id="XgNMTET57LmA"
line_list = line.split()
print(line_list)
# + colab={} colab_type="code" id="UL5ebfOXzAN-"
print(line_list[1])
# + [markdown] colab_type="text" id="oG-08k1P7LmC"
# A related method is ``splitlines()``, which splits on newline characters.
# Let's do this with a Haiku, popularly attributed to the 17th-century poet Matsuo Bashō:
# + colab={} colab_type="code" id="Xhoq3dov7LmD"
haiku = """matsushima-ya
aah matsushima-ya
matsushima-ya"""
haiku.splitlines()
# + [markdown] colab_type="text" id="nvAwkkOY7LmF"
# Note that if you would like to undo a ``split()``, you can use the ``join()`` method, which returns a string built from a splitpoint and an iterable:
# + colab={} colab_type="code" id="x6dnl1We7LmG"
'--'.join(['1', '2', '3'])
# + [markdown] colab_type="text" id="1AV-VUtK7LmI"
# A common pattern is to use the special character ``"\n"`` (newline) to join together lines that have been previously split, and recover the input:
# + colab={} colab_type="code" id="iPGNvfi17LmI"
print("\n".join(['matsushima-ya', 'aah matsushima-ya', 'matsushima-ya']))
# + colab={} colab_type="code" id="ATNMLZ1c7LmK"
pi = 3.14159
str(pi)
# + colab={} colab_type="code" id="IAXuKoSj7LmL"
print ("The value of pi is " + pi)
# + [markdown] colab_type="text" id="lxow04Rc7LmN"
# Pi is a float number so it must be transform to sting.
# + colab={} colab_type="code" id="n9XHEO0V7LmN"
print( "The value of pi is " + str(pi))
# + [markdown] colab_type="text" id="gLxjpyku7LmP"
# A more flexible way to do this is to use *format strings*, which are strings with special markers (noted by curly braces) into which string-formatted values will be inserted.
# Here is a basic example:
# + colab={} colab_type="code" id="TnpyntFp7LmQ"
"The value of pi is {}".format(pi)
# + [markdown] colab_type="text" id="fwhJjvfvvC_P"
# ### Easy regex manipulation!
# + colab={} colab_type="code" id="FgEQCSC5vQki"
import re
# + colab={} colab_type="code" id="7xYEVDt77Lmb"
line = 'the quick brown fox jumped over a lazy dog'
# + [markdown] colab_type="text" id="2ARgmkQP7Lmd"
# With this, we can see that the ``regex.search()`` method operates a lot like ``str.index()`` or ``str.find()``:
# + colab={} colab_type="code" id="mAPbNKrk7Lmd"
line.index('fox')
# + colab={} colab_type="code" id="PrQPx1bw7Lme"
regex = re.compile('fox')
match = regex.search(line)
match.start()
# + [markdown] colab_type="text" id="4NlxHgi57Lmg"
# Similarly, the ``regex.sub()`` method operates much like ``str.replace()``:
# + colab={} colab_type="code" id="Gx3ieKh47Lmg"
line.replace('fox', 'BEAR')
# + colab={} colab_type="code" id="Df1ZhHR97Lmi"
regex.sub('BEAR', line)
# + [markdown] colab_type="text" id="vAj0YV25vro6"
# The following is a table of the repetition markers available for use in regular expressions:
#
# | Character | Description | Example |
# |-----------|-------------|---------|
# | ``?`` | Match zero or one repetitions of preceding | ``"ab?"`` matches ``"a"`` or ``"ab"`` |
# | ``*`` | Match zero or more repetitions of preceding | ``"ab*"`` matches ``"a"``, ``"ab"``, ``"abb"``, ``"abbb"``... |
# | ``+`` | Match one or more repetitions of preceding | ``"ab+"`` matches ``"ab"``, ``"abb"``, ``"abbb"``... but not ``"a"`` |
# | ``.`` | Any character | ``.*`` matches everything |
# | ``{n}`` | Match ``n`` repetitions of preeeding | ``"ab{2}"`` matches ``"abb"`` |
# | ``{m,n}`` | Match between ``m`` and ``n`` repetitions of preceding | ``"ab{2,3}"`` matches ``"abb"`` or ``"abbb"`` |
# + colab={} colab_type="code" id="aQrRIY4nu_lv"
bool(re.search(r'ab', "Boabab"))
# + colab={} colab_type="code" id="Jl-dqlZ_vBjX"
bool(re.search(r'.*ma.*', "Ala ma kota"))
# + colab={} colab_type="code" id="MzfXrZA1wGvK"
bool(re.search(r'.*(psa|kota).*', "Ala ma kota"))
# + colab={} colab_type="code" id="QNwLgTchwXfU"
bool(re.search(r'.*(psa|kota).*', "Ala ma psa"))
# + colab={} colab_type="code" id="rEfi1FLzwcHa"
bool(re.search(r'.*(psa|kota).*', "Ala ma chomika"))
# + colab={} colab_type="code" id="FBH8XwJcwf_k"
zdanie = "Ala ma kota."
wzor = r'.*' #pasuje do każdego zdania
zamiennik = r"Ala ma psa."
# + colab={} colab_type="code" id="4_dDCJ1iwy3s"
re.sub(wzor, zamiennik, zdanie)
# + colab={} colab_type="code" id="Ouy86Z0txPvr"
wzor = r'(.*)kota.'
zamiennik = r"\1 psa."
# + colab={} colab_type="code" id="nKtdRi9bxhww"
re.sub(wzor, zamiennik, zdanie)
# + colab={} colab_type="code" id="deshsKUaxipk"
wzor = r'(.*)ma(.*)'
zamiennik = r"\1 posiada \2"
# + colab={} colab_type="code" id="bSc0LkQDx_z6"
re.sub(wzor, zamiennik, zdanie)
| Some_strings_and_regex_operation_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Surface Temperature Analysis
# You are asked to visualize the surface temperature change for the northern hemisphere for the past years. Data from the GISS Surface Temperature Analysis is used which contains estimates of global surface temperature change (in degree Celsius) for every month. The dataset contains temperature anomalies for every month from 1880 till present. Temperature anomalies indicate how much warmer or colder it is than normal. For the GISS analysis, normal means the average over the 30-year period 1951-1980.
# The goal of this exercise is to choose an appropriate color palette for the given data.
# Import the necessary modules and enable plotting within a Jupyter Notebook.
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# Use the pandas read_csv() function to read the dataset 'northern_surface_temperature.csv' located in the Dataset folder. After successful loading, transpose the dataset so that the dataset is in a suitable structure.
data = pd.read_csv("../../Datasets/northern_surface_temperature.csv", index_col=['Year'])
data = data.transpose()
# Create a custom diverging palette that diverges to blue (240 degrees on the hue wheel) for low values and to red (15 degrees on the hue wheel) for high values. Set the saturation s=99. Make sure that the diverging_palette() function returns a colormap by setting as_cmap=True.
heat_colormap = sns.diverging_palette(240, 15, s=99, as_cmap=True)
# Plot the heatmap for every 5 years. To ensure that the neutral color corresponds to no temperature change (value is zero), set center=0.
plt.figure(dpi=200)
sns.heatmap(data.iloc[:, ::5], cmap=heat_colormap, center=0)
plt.title("Temperature Changes from 1880 to 2015 (base period 1951-1980)")
plt.show()
| Chapter04/Exercise4.02/Exercise4.02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/brayannmb/python_basics_letscode_santander/blob/main/modulo_01.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="iS7KBom6O0jI"
# # **Módulo 01 - Santander Coders - Let´s Code - Data Science**
# + [markdown] id="MMBlRyMrQGWl"
# ##**Estrutura de Repetição While**
#
# A estrutura de repetição while executa uma ou mais tarefas até que uma enquanto uma condição é atendida. Quando a condição não for mais atendida, o programa sai do looping e continua executando o restante do código.
# + colab={"base_uri": "https://localhost:8080/"} id="va0petdEQO1G" outputId="0a84bbe8-624a-481b-b0d6-b322908229b2"
contador = 0 #deve estar no escopo global
while contador < 10:
contador+= 1 # o incremento do contador deve ser sempre uma das primeiras declarações dentro do bloco
if contador == 1:
print(contador, "item limpo")
else:
print(contador, "itens limpos")
print("Fim do looping while")
# + [markdown] id="Iuc-byeGQRGW"
# Looping infinito (while True), a estrutura de repetição while também pode ser utilizada em como looping infinito, necessariamente esse loop deve ser pausado em algum determinado momento do código. Caso não seja passado, o programa irá continuar executando o programa para sempre.
# + colab={"base_uri": "https://localhost:8080/"} id="OJrGfoCRQeH4" outputId="2a68bdfe-e342-41fb-93b0-f7d4c7dbc1b4"
contador = 0
while True: #looping infinito
if contador < 10:
contador+= 1
if contador == 1:
print(contador, "item limpo")
else:
print(contador, "itens limpos")
else:
break #comando para pausar o looping
# + [markdown] id="1CyLxbGmQhRM"
# Uma aplicação muito comum utilizando o while são os sistemas de verificação. Esses sistemas funcionam com a verificação da entrada de um usúario, caso a entrada que o usuário inseriu seja diferente da condição especificada na estrutura de repetição, o usuário será solicitado uma nova entrada até que aquela entrada seja igual a condição do while.
# + colab={"base_uri": "https://localhost:8080/"} id="j7RJKUJlQlcM" outputId="64ee93dc-0f29-4ca1-9d0c-92571fc550ed"
texto = input("Digite a sua senha: ")
while texto != "LetsCode":
texto = input("Acesso negado. Digite novamente: ")
print("Acesso permitido")
# + colab={"base_uri": "https://localhost:8080/"} id="qnVjiyZrQnEo" outputId="43c912dd-cad5-4594-b99f-dedf718157f1"
contador = 0
while contador < 10:
contador+= 1
if contador == 1:
continue #este comando ignora condição do if acima e pula para próxima linha.
print(contador, "itens limpos")
print("Fim do looping while")
# + [markdown] id="eLg-TxZ-QtC0"
# ###**Exemplos feitos em aula:**
# + colab={"base_uri": "https://localhost:8080/"} id="Sg5S1YZMQ05F" outputId="ff27e4b3-7ffd-4471-ce97-323b2a264a18"
#EXEMPLO 01:
horario = int(input('Qual horario é agora? '))
# Testando a condição uma única vez com o if:
if 0 < horario < 6:
print('Você está no horario da madrugada')
else:
print('Você nao está no horario da madrugada')
# Testando a condição em loop com o while:
while 0 < horario < 6:
print('Você está no horario da madrugada')
horario = horario + 1
else:
print('Você nao está no horario da madrugada')
# + colab={"base_uri": "https://localhost:8080/"} id="K-CI7kPPQ4Ch" outputId="a4093b91-db64-4477-baa5-b671669b5a13"
#EXEMPLO 02
# O while permite continuar decrementando o número de pipocas até chegar em 0:
num_pipocas = int(input('Digite o numero de pipocas: '))
while num_pipocas > 0:
print('O numero de pipocas é: ', num_pipocas)
num_pipocas = num_pipocas - 1
# + colab={"base_uri": "https://localhost:8080/"} id="HV6mzRE3Q8dI" outputId="8de16f54-b05d-4aa6-c394-6d9f317278bc"
#EXEMPLO 03
horario = int(input('Qual horario é agora? '))
while horario > 0 and horario < 6:
print('Você está no horario da madrugada')
horario = horario + 1
else:
print('Você nao está no horario da madrugada')
# + [markdown] id="Y4Al6BhEQ_Wa"
# ###**Exemplos com sistemas de verificação**
# + colab={"base_uri": "https://localhost:8080/"} id="PCzjY7AUREwc" outputId="7c7a1617-233c-41c3-b5f7-99ca05aa133e"
#EXEMPLO 01
# o exemplo abaixo não aceita um salário menor do que o mínimo atual:
salario = float(input('Digite seu salario: '))
while salario < 998.0:
salario = float(input('Entre com um salario MAIOR DO QUE 998.0: '))
else:
print('O salario que você entrou foi: ', salario)
# + colab={"base_uri": "https://localhost:8080/"} id="y6hGL9xvRGDW" outputId="ac9618d0-0359-4463-b2ec-310184a7cf0f"
#EXEMPLO 02
# o exemplo abaixo só sai do loop quando o usuário digitar "OK":
resposta = input('Digite OK: ')
while resposta != 'OK':
resposta = input('Não foi isso que eu pedi, digite OK: ')
print('Agora sim!')
# + [markdown] id="rVuqV6PTROIq"
# ###**Exemplo de código utilizando o comando break**
# + colab={"base_uri": "https://localhost:8080/"} id="fYrW2jYkRjG-" outputId="5ad81099-5933-4a18-9923-f8fd2b908872"
while True:
resposta = input('Digite OK: ')
if resposta == 'OK':
break
# + id="odMwEKKIRmfz"
| modulo_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# pip install numpy
# -
import numpy as np
np.array([1,2,3])
np.zeros(9)
np.ones(3)
np.zeros((3,4))
np.ones((3,7))
np.ones((2,2), dtype=np.int16)
np.empty((3,4))
np.eye(5)
np.arange(6)
np.arange(3,5)
# +
#. 0, 1 , 2 , 3 , 4
#. ]3,5]
# -
np.arange(3,15,2)
np.array([(1,2,3),(4,5,6)]).shape
np.array([1,2,3,4]).shape
np.arange(6).reshape(2,3)
np.arange(20).reshape(10,2)
np.ones_like(np.arange(6).reshape(3,2))
np.arange(6).reshape(3,2)
np.arange(1000000).reshape(10000,100)
# Suma
a = np.array([10,10,10])
b = np.array([5,5,5])
a + b
# Resta
a - b
# Multiplicacion
a * b
# Division
a / b
a
# Mod
a % 2
# Expresiones
a = np.array([10,5,12])
a < 11
A = np.array([[0,1],[2,3]])
B = np.array([[4,5],[6,7]])
print(A)
print(B)
# print(A * B)
# A.dot(B)
np.dot(A,B)
a
a *= 3
a
arreglo_numeros = np.arange(1,11)
arreglo_numeros
arreglo_numeros.sum()
arreglo_numeros.min()
arreglo_numeros.max()
numeros = np.arange(12).reshape(3,4)
numeros
numeros.sum(axis=0) # Columnas
numeros.sum(axis=1) # Filas
angulos = np.array([0,30,45,60,90])
angulos
angulos_en_radianes = angulos * (np.pi/180)
angulos_en_radianes
np.radians(angulos)
np.degrees(angulos_en_radianes)
# Seno
np.sin(angulos)
# Seno
np.sin(angulos_en_radianes)
# Coseno
np.cos(angulos_en_radianes)
# Tangente
np.tan(angulos_en_radianes)
notas = np.array([1,2,4,3,2,1,2,4,5,6,7,8,9,8,7,6,5,4,3,2,1,2,4,6,9])
notas.mean()
np.median(notas)
np.std(notas)
np.var(notas)
salarios = np.genfromtxt("./salarios.csv", delimiter=",")
salarios
salarios.mean()
arreglo_once = np.arange(11)**2
arreglo_once
print(arreglo_once[0:5])
print(arreglo_once[-10:-2])
print(arreglo_once[:8])
print(arreglo_once[5:])
print(arreglo_once[2:8:4])
print(arreglo_once[::-2])
print(arreglo_once[::2])
estudiantes = np.array([
['Adrian','Vicente','Wendy','Carolina'],
[1,2,3,4],
[5,6,7,8]
])
for i in estudiantes:
print(f"Valor: {i}")
for i in estudiantes.flatten(order='F'):
print(f"Valor individual: {i}")
estudiantes.flatten().reshape(3,4)
# nditer
arreglo_tres_cuatro = np.arange(12).reshape(3,4)
arreglo_tres_cuatro
for i in np.nditer(arreglo_tres_cuatro,
order="F",
flags=["external_loop"]):
print(f"Valor: {i}")
arreglo_split = np.array([1,2,3,4,5,6,7,8,9,10,11,12])
print(np.split(arreglo_split,6))
print(np.split(arreglo_split,[9,12]))
a = np.arange(11)
a.sort(axis=-1)
a
randomico = np.random.randint(0,3, size=10)
randomico
| 07_Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
import numpy as np
from pprint import pprint
from nltk.tokenize import sent_tokenize, word_tokenize
import os
#The OS module in Python provides a way of using operating system dependent functionality.
#The functions that the OS module provides allows you to interface with the underlying operating system
#that Python is running on – be that Windows, Mac or Linux.
from os import listdir
from os.path import isfile, join
# -
import nltk as nltk
nltk.download('stopwords')
# +
stopwords = nltk.corpus.stopwords.words('french')
stopwords.append('’')
stopwords.append('«')
stopwords.append('»')
# -
def tokenize(text):
tokens = word_tokenize(text)
tokens = _pre_clean(tokens)
tokens = [token for token in tokens if len(token) > 2]
tokens = [token for token in tokens if token not in stopwords]
#tokens = [get_lemma(token) for token in tokens]
return tokens
def _pre_clean(list_of_text):
'''
preliminary cleaning of the text
- remove new line character i.e. \n or \r
- remove tabs i.e. \t
- remove extra spaces
'''
cleaned_list = []
for text in list_of_text:
# print("original:", text)
text = text.replace('\\n', ' ')
text = text.replace('\\r', ' ')
text = text.replace('\\t', ' ')
pattern = re.compile(r'\s+')
text = re.sub(pattern, ' ', text)
text = text.strip()
text = text.lower()
# check for empty strings
if text != '' and text is not None:
cleaned_list.append(text)
return cleaned_list
pwd
# +
HOME = os.getcwd()
# +
TEXTS_DIR = HOME + "/Chaire_Altissia_0/"
filelabels = {}
texts_data = []
files = [f for f in os.listdir(TEXTS_DIR) if os.path.isfile(os.path.join(TEXTS_DIR, f))]
import string
from string import punctuation
remove_punct_map = dict.fromkeys(map(ord, string.punctuation))
tokens_total = []
count = -1
os.chdir(TEXTS_DIR)
for f in files:
#os.chdir(TEXTS_DIR)
with open(f, "r", encoding='utf-8', errors = 'ignore') as openf:
tokens = []
count = count + 1
filelabels[count] = os.path.basename(openf.name)
for line in openf:
sent_text = nltk.sent_tokenize(line)
for sentence in sent_text:
tokens1 = tokenize(sentence)
tokens1 = [item.translate(remove_punct_map)
for item in tokens1]
#filter_object = filter(lambda x: x != "", tokens1)
tokens1 = [x for x in tokens1 if x!= ""]
for token in tokens1:
tokens.append(token)
tokens_total.append(token)
#if random.random() > .99:
#print(tokens)
#print(tokens_total)
texts_data.append(tokens)
print(filelabels)
# +
len(tokens_total)
# -
tokens_total = [x for x in tokens_total if x not in stopwords]
# +
len(tokens_total)
# +
from collections import Counter
Count_total = Counter(tokens_total)
# +
print(Count_total)
# +
import pyperclip as clip
# -
clip.copy(f"{Count_total}")
# +
# Command+V into a page/word/txt file [or clip.paste() to print it here, but in this case it is too large a list to print]
# +
for i in range(6):
print(len(texts_data[i]))
# +
for i in range(6):
texts_data[i] = [x for x in texts_data[i] if x not in stopwords]
# -
for i in range(6):
print(len(texts_data[i]))
# +
Count_total_0 = Counter(texts_data[0])
# -
print(Count_total_0)
# +
Count_total_6 = Counter(texts_data[6])
# +
Count_total_5 = Counter(texts_data[5])
# +
print(Count_total_5)
# -
| JupyterNotebook/Chaire_Altissia_French_Tokenization_&_Term_Count_Chris_Margento.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ATLAS Assignment
# ## Autoencoders for Data Compression
# ### Implementing Autoencoder network for compressing 4 variables data to 3 variables.
#Fixing the number of columns for consideriung all the objects per event and not skipping any rows.
columns=[]
for i in range(21):
columns.append(str(i))
# Reading data from the CSV file, with delimiter as ';' as data for events are stored in the format of event ID; process ID; event weight; MET; METphi; obj1, E1, pt1, eta1, phi1; obj2,
# E2, pt2, eta2, phi2; . . . and filling the missing values with 0
# +
import pandas as pd
import io
df = pd.read_csv('monojet_Zp2000.0_DM_50.0_chan3.csv', header=None, delimiter=';',error_bad_lines=False, names=columns).fillna(0)
print(df)
# -
#Raw dataset
df
# #### Processing dataset step
# Iterating over the rows of raw dataset, checking every object from 5th column, as before that the cells have values of event ID; process ID; event weight; MET; METphi;
# Checking if per event we have more than one object of jet-type and then forming the processes dataset.
# +
row_count,col_count=df.shape
data=pd.DataFrame(columns = [ 'E', 'pt', 'eta', 'phi'])
for i in range(row_count):
for j in range(5,21):
val=str(df.iloc[i,j]).split(",") #spliting the dataframe cell value to form the list
if(val[0]=='j'): #checking whether object is of j-type
data = data.append({'E' : float(val[1]), #appending processed dataset with the values
'pt' : float(val[2]),
'eta': float(val[3]),
'phi': float(val[4])},
ignore_index = True)
# -
# Random sampling of training and testing dataset, divided into 80% as training data and remaining as testing data
#
train = data.sample(frac = 0.80)
test = data.drop(train.index)
print("Entries in the train dataset", len(train))
print("Entries in the test dataset", len(test))
train
# Plotting the data using Matplot library for the objects present.
# +
import os
save_dir = "plotOutput"
if not os.path.exists(save_dir):
os.makedirs(save_dir)
import numpy as np
import matplotlib.pyplot as plt
unit_list = ['[log(GeV)]', '[log(GeV)]', '[rad/3]', '[rad/3]']
variable_list = [r'$E$', r'$p_T$', r'$\eta$', r'$phi$']
branches=["E","pt","eta","phi"]
n_bins = 100
for kk in range(0,4):
n_hist_data, bin_edges, _ = plt.hist(train[branches[kk]], color='gray', label='Input', alpha=1, bins=n_bins)
plt.xlabel(xlabel=variable_list[kk] + ' ' + unit_list[kk])
plt.ylabel('# of events')
plt.show()
# -
# #### Normalization
# Normalization is to scaling the values by modifying to use a common scale, without distorting differences in the ranges of values or losing information. We use standarization(z-score normalization) or max-min normalization for the scaling purposes.
#
# In max-min normalization we scale by,
# 
#
# In z-score normalization, we scale by,
# 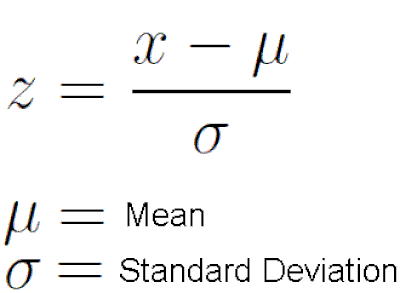
# +
mean = train.mean()
std = train.std()
train_data = (train - mean) / std #z-score normalizing training input data
test_data = (test - mean) / std #z-score normalizing testing input data
# +
unit_list = ['[log(GeV)]', '[log(GeV)]', '[rad/3]', '[rad/3]']
variable_list = [r'$E$', r'$p_T$', r'$\eta$', r'$phi$']
branches=["E","pt","eta","phi"]
n_bins = 100
for kk in range(0,4):
n_hist_data, bin_edges, _ = plt.hist(train_data[branches[kk]], color='gray', label='Input', alpha=1, bins=n_bins)
plt.xlabel(xlabel=variable_list[kk] + ' ' + unit_list[kk])
plt.ylabel('# of events')
plt.show()
# -
# #### Setting up the network
# ##### Preparing the data
# Forming the TensorDatasets from adding the two datasets using PyTorch. also importing other libraries and packages.
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.utils.data
from torch.autograd import Variable
from torch.utils.data import TensorDataset
from torch.utils.data import DataLoader
from fastai import learner
from fastai.data import core
train_x = train_data
test_x = test_data
train_y = train_x # y = x since we are building an autoencoder
test_y = test_x
# Constructs a tensor object of the data and wraps them in a TensorDataset object.
train_ds = TensorDataset(torch.tensor(train_x.values, dtype=torch.float), torch.tensor(train_y.values, dtype=torch.float))
valid_ds = TensorDataset(torch.tensor(test_x.values, dtype=torch.float), torch.tensor(test_y.values, dtype=torch.float))
# -
# Fixing the batch size and further dividing into batches.Batch Size is a hyperparameter specifying the number of samples to work or pass into the network before updating the model parameters.
# +
bs = 256
# Converts the TensorDataset into a DataLoader object and combines into one DataLoaders object (a basic wrapper
# around several DataLoader objects).
train_dl = DataLoader(train_ds, batch_size=bs, shuffle=True)
valid_dl = DataLoader(valid_ds, batch_size=bs * 2)
dls = core.DataLoaders(train_dl, valid_dl)
# -
# ### Preparing the Network
# Here we design the required neural network for the autoencoder. Here we have a LeakyReLU, tanh activation function, which is used to add non-linearity. Layers goes from 4 to 200 to 20 to 3 for the encoder part and same in reverse order for the decoder part, where the 4 variables are compressed to 3 after the encoder and can be again decompressed using the decoder network.
#
# Below figure shows a diagrammatic view of the created neural network where the encoder part is till Hidden layer 4 and after that the decoder network starts.
#
# Layers for 200 nodes are scaled to 50 for easier visual representation and understanding.
# 
#
#
# +
class AE_3D_200_LeakyReLU(nn.Module):
def __init__(self, n_features=4):
super(AE_3D_200_LeakyReLU, self).__init__()
self.en1 = nn.Linear(n_features, 200)
self.en2 = nn.Linear(200, 200)
self.en3 = nn.Linear(200, 20)
self.en4 = nn.Linear(20, 3)
self.de1 = nn.Linear(3, 20)
self.de2 = nn.Linear(20, 200)
self.de3 = nn.Linear(200, 200)
self.de4 = nn.Linear(200, n_features)
self.tanh = nn.Tanh()
def encode(self, x):
return self.en4(self.tanh(self.en3(self.tanh(self.en2(self.tanh(self.en1(x)))))))
def decode(self, x):
return self.de4(self.tanh(self.de3(self.tanh(self.de2(self.tanh(self.de1(self.tanh(x))))))))
def forward(self, x):
z = self.encode(x)
return self.decode(z)
def describe(self):
return 'in-200-200-20-3-20-200-200-out'
model = AE_3D_200_LeakyReLU()
model.to('cpu')
# -
# Now we define a loss function. Here we choose MSE loss which stands for Mean Squared Error loss, defined as
# 
#
# MSE Loss is appropriate for a compression autoencoder as it reflects the [(input-output)/input] physical quantity that we want to minimize.
#
# +
from fastai.metrics import mse
loss_func = nn.MSELoss()
#bn_wd = False # Don't use weight decay for batchnorm layers
#true_wd = True # weight decay will be used for all optimizers
wd = 1e-6
recorder = learner.Recorder()
learn = learner.Learner(dls, model=model, wd=wd, loss_func=loss_func, cbs=recorder)
# -
# ### Training the network
# Before training the neural network, we will find the best learning rate. The learning rate is a hyper-paramater that sets how much the weights of the network will change each step with respect to the loss gradient.
#
# Then we plot the loss versus the learning rates. We're interested in finding a good order of magnitude of learning rate, so we plot with a log scale.
#
# A good value for the learning rates is then either:
#
# one tenth of the minimum before the divergence
# when the slope is the steepest
# +
from fastai.callback import schedule
lr_min, lr_steep = learn.lr_find()
print('Learning rate with the minimum loss:', lr_min)
print('Learning rate with the steepest gradient:', lr_steep)
# -
# Now we want to run the training!
#
# User-chosen variables:
#
# n_epoch: The number of epochs, i.e how many times the to run through all of the training data once (i.e the 6000 entries, see cell 2)
# lr: The learning rate. Either choose lr_min, lr_steep from above or set your own.
# +
import time
start = time.perf_counter() # Starts timer
learn.fit_one_cycle(100)
end = time.perf_counter() # Ends timer
delta_t = end - start
print('Training took', delta_t, 'seconds')
# -
# Then we plot the loss as a function of batches and epochs to check if we reach a plateau.
recorder.plot_loss()
# Finding the MSE Error after network training
learn.validate()
# +
plt.close('all')
unit_list = ['[log(GeV)]', '[log(GeV)]', '[rad/3]', '[rad/3]']
variable_list = [r'$E$', r'$p_T$', r'$\eta$', r'$\phi$']
line_style = ['--', '-']
colors = ['orange', 'c']
markers = ['*', 's']
model.to('cpu')
save = False # Option to save figure
# Histograms
idxs = (0, 100000) # Choose events to compare
data = torch.tensor(test_data[idxs[0]:idxs[1]].values, dtype=torch.float)
pred = model(data)
pred = pred.detach().numpy()
data = data.detach().numpy()
data_df = pd.DataFrame(data, columns=test.columns)
pred_df = pd.DataFrame(pred, columns=test.columns)
alph = 0.8
n_bins = 200
for kk in np.arange(4):
plt.figure()
n_hist_data, bin_edges, _ = plt.hist(data[:, kk], color=colors[1], label='Input', alpha=1, bins=n_bins)
n_hist_pred, _, _ = plt.hist(pred[:, kk], color=colors[0], label='Output', alpha=alph, bins=bin_edges)
plt.suptitle(test.columns[kk])
plt.xlabel(test.columns[kk])
plt.ylabel('Number of events')
plt.yscale('log')
if save:
plt.savefig(os.path.join(save_dir,test.columns[kk]+'.png'))
plt.legend()
# -
# #### Experimenting with Batch Normalization Layers
# We will modify the neural network and add Batch Normalization layers, which normalizes batches after layers. Basically, rather than normalizing the values once in the beginning, we normalize all over the network.
# Batch Normalization helps in improving training time and decreases the time of convergence and the problem of vanishing gradients and acts as regularization too.
#
# +
#Also modify the network by adding another layer each in encoder and decoder part
class AE_3D_200_LeakyReLU_BatchNorm(nn.Module):
def __init__(self, in_features=4):
super().__init__()
self.encoder = nn.Sequential(
nn.Linear(in_features, 200),
nn.BatchNorm1d(200),
nn.Tanh(),
nn.Linear(200, 200),
nn.BatchNorm1d(200),
nn.Tanh(),
nn.Linear(200, 20),
nn.BatchNorm1d(20),
nn.Tanh(),
nn.Linear(20, 3)
)
self.decoder = nn.Sequential(
nn.Tanh(),
nn.Linear(3, 20),
nn.BatchNorm1d(20),
nn.Tanh(),
nn.Linear(20, 200),
nn.BatchNorm1d(200),
nn.Tanh(),
nn.Linear(200, 200),
nn.BatchNorm1d(200),
nn.Tanh(),
nn.Linear(200, 4)
)
def forward(self, x):
encoded = self.encoder(x)
return self.decoder(encoded)
def describe(self):
return 'in-200-200-20-3-20-200-200-out'
model = AE_3D_200_LeakyReLU_BatchNorm()
model.to('cpu')
# +
from fastai.metrics import mse
loss_func = nn.MSELoss()
#bn_wd = False # Don't use weight decay for batchnorm layers
#true_wd = True # weight decay will be used for all optimizers
wd = 1e-6
recorder = learner.Recorder()
learn = learner.Learner(dls, model=model, wd=wd, loss_func=loss_func, cbs=recorder)
# +
from fastai.callback import schedule
lr_min, lr_steep = learn.lr_find()
print('Learning rate with the minimum loss:', lr_min)
print('Learning rate with the steepest gradient:', lr_steep)
# +
import time
start = time.perf_counter() # Starts timer
learn.fit_one_cycle(100,lr_min)
end = time.perf_counter() # Ends timer
delta_t = end - start
print('Training took', delta_t, 'seconds')
# -
recorder.plot_loss()
learn.validate()
# +
plt.close('all')
unit_list = ['[log(GeV)]', '[log(GeV)]', '[rad/3]', '[rad/3]']
variable_list = [r'$E$', r'$p_T$', r'$\eta$', r'$\phi$']
line_style = ['--', '-']
colors = ['orange', 'c']
markers = ['*', 's']
model.to('cpu')
save = False # Option to save figure
# Histograms
idxs = (0, 100000) # Choose events to compare
data = torch.tensor(test_data[idxs[0]:idxs[1]].values, dtype=torch.float)
pred = model(data)
pred = pred.detach().numpy()
data = data.detach().numpy()
data_df = pd.DataFrame(data, columns=test.columns)
pred_df = pd.DataFrame(pred, columns=test.columns)
alph = 0.8
n_bins = 200
for kk in np.arange(4):
plt.figure()
n_hist_data, bin_edges, _ = plt.hist(data[:, kk], color=colors[1], label='Input', alpha=1, bins=n_bins)
n_hist_pred, _, _ = plt.hist(pred[:, kk], color=colors[0], label='Output', alpha=alph, bins=bin_edges)
plt.suptitle(test.columns[kk])
plt.xlabel(test.columns[kk])
plt.ylabel('Number of events')
plt.yscale('log')
if save:
plt.savefig(os.path.join(save_dir,test.columns[kk]+'.png'))
plt.legend()
# -
| ATLAS Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Come scrivere comodamente comandi SQL in un notebook
#
# Viene sfruttato il componente <https://github.com/catherinedevlin/ipython-sql> che rende disponibile un comando `magic` dedicato.
import pandas as pd
# %load_ext sql
# ## Connessione
# + language="sql"
# sqlite:///db.sqlite
# -
# ## Query
# + language="sql"
# select * from tabella
# + language="sql"
# SELECT load_extension('mod_spatialite');
# -
# ## Convesione in dataframe
# converto in dataframe l'output
# result = %sql select * from tabella
dataframe = result.DataFrame()
dataframe
# # Connessione a db sqlite e conversione in dataframe
import sqlite3
with sqlite3.connect('db.sqlite') as conn:
dataframe = pd.io.sql.read_sql("""
SELECT *
FROM tabella;""", conn)
# ## Abilitare modulo spatialite in sqlite
conna=sqlite3.connect('db.sqlite')
conna.enable_load_extension(True)
conna.load_extension('mod_spatialite')
conna.execute('select InitSpatialMetadata(1)')
conna.execute("SELECT AddGeometryColumn('tabella', 'geom', 4326, 'POINT', 2);")
conna.execute('''
UPDATE tabella SET
geom = GeomFromText(('POINT(13 38)'),4326);
''')
conna.commit()
conna.close()
# + language="sql"
# select * from tabella
# -
| notebook/03 query SQL su db sqlite/sqlite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.5 64-bit
# language: python
# name: python36564bit5b328c96e5964481b5634e4dffc91a39
# ---
import pandas as pd
import numpy as np
import pandas as pd
import keras
from keras import backend as K
from keras.layers import Input, Dense, Dropout
from keras.layers.advanced_activations import PReLU
from keras.models import Model
from keras.optimizers import Adamax
import math
import matplotlib
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
import matplotlib
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
# +
# Model fixed parameters
NInputs = 4 # Number of features (model inputs)
NOutputs = 3 # Number of targets (model outputs)
# +
# Import training data (model inputs & outputs)
Feature_Train = pd.read_csv('processed_data/SL_Train_Features.csv', header = 0)
Feature_Train = Feature_Train[['GR', 'RHOB','NPHI','RPCELM_l10']]
Feature_Train = np.asarray(Feature_Train)
print(Feature_Train.shape)
Target_Train = pd.read_csv('processed_data/SL_Train_Target.csv', header = 0)
Target_Train = Target_Train[['SW', 'PHIF', 'VSH']]
Target_Train = np.asarray(Target_Train)
print(Target_Train.shape)
print(Feature_Train[0])
# Import validation data (model inputs & outputs)
Feature_Val = pd.read_csv('processed_data/SL_Val_Features.csv', header = 0)
Feature_Val = Feature_Val[['GR', 'RHOB','NPHI','RPCELM_l10']]
Feature_Val = np.asarray(Feature_Val)
print(Feature_Val.shape)
Target_Val = pd.read_csv('processed_data/SL_Val_Target.csv', header = 0)
Target_Val = Target_Val[['SW', 'PHIF', 'VSH']]
Target_Val = np.asarray(Target_Val)
print(Target_Val.shape)
# Import test data (model inputs & outputs)
Feature_Test = pd.read_csv('processed_data/SL_Test_Features.csv', header = 0)
Feature_Test = Feature_Test[['GR', 'RHOB','NPHI','RPCELM_l10']]
Feature_Test = np.asarray(Feature_Test)
print(Feature_Test.shape)
Target_Test = pd.read_csv('processed_data/SL_Test_Target.csv', header = 0)
Target_Test = Target_Test[['SW', 'PHIF', 'VSH']]
Target_Test = np.asarray(Target_Test)
print(Target_Test.shape)
# -
Feature_Test
# +
# Defining neural network
rate = 0.1
def nn():
NN_ip = Input(shape=(NInputs,))
x = Dense(1000)(NN_ip)
x = PReLU()(x)
x = Dropout(rate)(x)
x = Dense(1000)(x)
x = PReLU()(x)
x = Dropout(rate)(x)
NN_op = Dense(NOutputs)(x)
NN_model = Model(NN_ip, NN_op)
def loss_mse(true, pred):
mean = pred[:, :NOutputs]
return K.mean(K.square(true - mean), -1)
def metric_mse(y_true, y_pred):
mean = y_pred[:, :NOutputs]
return K.mean(K.square(y_true - mean), -1)
opt = Adamax(1e-3)
NN_model.compile(loss = loss_mse, optimizer = opt, metrics = [metric_mse])
return NN_model
NN = nn()
NN.summary()
# +
NN = nn()
NN.load_weights('NNWeights.h5')
history = NN.fit(Feature_Train, Target_Train,
epochs = 1000,
batch_size = int(Feature_Train.shape[0]/8),
shuffle = True,
validation_data = (Feature_Val, Target_Val))
plt.plot(history.history['val_loss'][2:],'r')
plt.plot(history.history['loss'][2:],'k')
# -
#3.1 Plot the predicted saturation versus the Interpreted saturation
PredictedTarget = NN.predict(Feature_Test)
[PredictedSw, PredictedPHIF, PredictedVSH] = [PredictedTarget[:,0], PredictedTarget[:,1], PredictedTarget[:,2]]
PredictedTarget
PredictedTarget.shape
Target_Test.shape
plt.figure(figsize = (6,6))
plt.style.use('seaborn-white')
plt.scatter(Target_Test[:,0], PredictedSw)
plt.plot(np.linspace(0, 1, 2), np.linspace(0, 1, 2), color = 'r', alpha = 1)
plt.xlabel('Interpreted Sw', FontSize = 16)
plt.ylabel('Predicted Sw', FontSize = 16)
plt.show()
plt.figure(figsize = (6,6))
plt.style.use('seaborn-white')
plt.scatter(Target_Test[:,1], PredictedPHIF)
plt.plot(np.linspace(0, .4, 2), np.linspace(0, .4, 2), color = 'r', alpha = 1)
plt.xlabel('Interpreted PHIF', FontSize = 16)
plt.ylabel('Predicted PHIF', FontSize = 16)
plt.show()
plt.figure(figsize = (6,6))
plt.style.use('seaborn-white')
plt.scatter(Target_Test[:,2], PredictedVSH)
plt.plot(np.linspace(0, .8, 2), np.linspace(0, .8, 2), color = 'r', alpha = 1)
plt.xlabel('Interpreted VSH', FontSize = 16)
plt.ylabel('Predicted VSH', FontSize = 16)
plt.show()
output_df = pd.read_csv('processed_data/SL_Test_Target.csv', header = 0)
type(output_df)
output_df.insert( 4,"PredictedSw", PredictedSw)
output_df.insert( 6,"PredictedVSH", PredictedVSH)
output_df.insert( 8,"PredictedPHIF", PredictedPHIF)
output_df
output_df.to_csv('processed_data/SL_PredictedProperties.csv', index=True)
output_df['VSHDifference'] = abs(output_df['PredictedVSH'] - output_df['VSH'])
output_df['PHIFDifference'] = abs(output_df['PredictedPHIF'] - output_df['PHIF'])
output_df['SwDifference'] = abs(output_df['PredictedSw'] - output_df['SW'])
# ## Plotting the Prediction Results
# After the prediction has completed, we can view the predicted result against the actual measurement on the test well: 15/9-F-11 B
# +
fig, ax = plt.subplots(3, 1, figsize=(20,10))
ax1 = plt.subplot2grid((2,1), (0,0))
ax2 = plt.subplot2grid((2,1), (1,0))
ax1.plot(output_df['MD'], output_df['VSH'], label='Interpreted', color='black')
ax1.plot(output_df['MD'], output_df['PredictedVSH'], label='Predicted', color='red')
ax1.set_ylabel('VSH', fontsize=16, fontweight='bold', labelpad=30)
ax1.set_xlabel('Depth (m)', fontsize=16, fontweight='bold', labelpad=30)
ax1.tick_params(which='both', labelsize=14)
ax2.plot(output_df['MD'], output_df['VSHDifference'], label='Interpreted', color='black')
ax2.set_ylim(0,1)
ax2.set_ylabel('Abs. Prediction Difference', fontsize=16, fontweight='bold', labelpad=30)
for ax in [ax1, ax2]:
ax.grid()
ax.set_axisbelow(True)
ax.set_xlabel('Depth (m)', fontsize=16, fontweight='bold', labelpad=30)
ax.tick_params(which='both', labelsize=14)
ax1.legend(fontsize=14, facecolor='white', frameon=True)
plt.show()
# +
fig, ax = plt.subplots(3, 1, figsize=(20,10))
ax1 = plt.subplot2grid((2,1), (0,0))
ax2 = plt.subplot2grid((2,1), (1,0))
ax1.plot(output_df['MD'], output_df['PHIF'], label='Interpreted', color='black')
ax1.plot(output_df['MD'], output_df['PredictedPHIF'], label='Predicted', color='red')
ax1.set_ylabel('PHIF', fontsize=16, fontweight='bold', labelpad=30)
ax1.set_xlabel('Depth (m)', fontsize=16, fontweight='bold', labelpad=30)
ax1.tick_params(which='both', labelsize=14)
ax2.plot(output_df['MD'], output_df['PHIFDifference'], label='Interpreted', color='black')
ax2.set_ylim(0,1)
ax2.set_ylabel('Abs. Prediction Difference', fontsize=16, fontweight='bold', labelpad=30)
for ax in [ax1, ax2]:
ax.grid()
ax.set_axisbelow(True)
ax.set_xlabel('Depth (m)', fontsize=16, fontweight='bold', labelpad=30)
ax.tick_params(which='both', labelsize=14)
ax1.legend(fontsize=14, facecolor='white', frameon=True)
plt.show()
# +
fig, ax = plt.subplots(3, 1, figsize=(20,10))
ax1 = plt.subplot2grid((2,1), (0,0))
ax2 = plt.subplot2grid((2,1), (1,0))
ax1.plot(output_df['MD'], output_df['SW'], label='Interpreted', color='black')
ax1.plot(output_df['MD'], output_df['PredictedSw'], label='Predicted', color='red')
ax1.set_ylabel('Water Saturation (dec)', fontsize=16, fontweight='bold', labelpad=30)
ax1.set_xlabel('Depth (m)', fontsize=16, fontweight='bold', labelpad=30)
ax1.tick_params(which='both', labelsize=14)
ax2.plot(output_df['MD'], output_df['SwDifference'], label='Interpreted', color='black')
ax2.set_ylim(0,1)
ax2.set_ylabel('Abs. Prediction Difference', fontsize=16, fontweight='bold', labelpad=30)
for ax in [ax1, ax2]:
ax.grid()
ax.set_axisbelow(True)
ax.set_xlabel('Depth (m)', fontsize=16, fontweight='bold', labelpad=30)
ax.tick_params(which='both', labelsize=14)
ax1.legend(fontsize=14, facecolor='white', frameon=True)
# -
| 2 - Predict_Properties.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:kmer-hashing]
# language: python
# name: conda-env-kmer-hashing-py
# ---
# ! aws s3 ls s3://olgabot-maca/sourmash/2018-08-10_test_kmer_sizes_gencode/
# +
import seaborn as sns
import matplotlib.pyplot as plt
import altair as alt
import pandas as pd
import numpy as np
import altair as alt
alt.renderers.enable('default')
# %matplotlib inline
# -
alt.__version__
# +
prefix = "s3://olgabot-maca/sourmash/2018-08-10_test_kmer_sizes_gencode"
TRIMMED_TEMPLATE = f"{prefix}/A1-D041914-3_8_M-1-1.trimmed" + ".k{ksize}.dist"
def logify(dist):
dist['abundance_log10'] = np.log10(dist['abundance'])
dist['count_log10'] = np.log10(dist['count'])
return dist
def get_trimmed_dist(ksize, template=TRIMMED_TEMPLATE):
dist = pd.read_csv(template.format(ksize=ksize))
dist['ksize'] = ksize
dist['is_trimmed'] = 'trimmed'
dist = logify(dist)
return dist
k21_trimmed = get_trimmed_dist(21)
k27_trimmed = get_trimmed_dist(27)
trimmed_dist = pd.concat([k21_trimmed, k27_trimmed])
print(trimmed_dist.shape)
trimmed_dist.head()
# +
TEMPLATE = f"{prefix}/A1-D041914-3_8_M-1-1_S269_"+"R{read_number}_001.k{ksize}.dist"
def get_untrimmed_dist(ksize, template=TEMPLATE):
r1 = pd.read_csv(template.format(read_number=1, ksize=ksize), index_col=0)
r2 = pd.read_csv(template.format(read_number=2, ksize=ksize), index_col=0)
dist = r1.add(r2[['count', 'cumulative']])
dist = dist.reset_index()
dist['cumulative_fraction'] = dist['cumulative'] / dist['count'].sum()
dist['ksize'] = ksize
dist['is_trimmed'] = 'untrimmed'
dist = logify(dist)
return dist
k21_untrimmed = get_untrimmed_dist(21)
print(k21_untrimmed.shape)
k21_untrimmed.head()
# -
k27_untrimmed = get_untrimmed_dist(27)
print(k27_untrimmed.shape)
k27_untrimmed.head()
untrimmed = pd.concat([k27_untrimmed, k21_untrimmed])
untrimmed['is_trimmed'] = 'untrimmed'
untrimmed.head()
g = sns.FacetGrid(untrimmed, hue='ksize')
g.map(sns.lineplot, 'abundance_log10', 'count_log10', scaley='log')
g.add_legend()
combined_dists = pd.concat([trimmed_dist, untrimmed], ignore_index=True)
print(combined_dists.shape)
combined_dists.head()
g = sns.FacetGrid(combined_dists, hue='is_trimmed', col='ksize')
g.map(sns.lineplot, 'abundance_log10', 'count_log10')
g.add_legend()
untrimmed
# +
# # The basic line
# line = alt.Chart(untrimmed).mark_line().encode(
# x='abundance:Q',
# y='count:Q',
# color='ksize:N'
# )
# line
# -
r2_k21 = pd.read_csv("s3://olgabot-maca/sourmash/2018-08-10_test_kmer_sizes_gencode/A1-D041914-3_8_M-1-1_S269_R2_001.k21.dist", index_col=0)
r2_k21.head()
k21 = r1_k21.add(r2_k21[['count', 'cumulative']])
k21['cumulative_fraction'] = k21['cumulative'] / k21['count'].sum()
k21.head()
annotations = pd.read_csv('https://github.com/czbiohub/tabula-muris/raw/master/00_data_ingest/18_global_annotation_csv/annotations_facs.csv',
index_col='cell')
annotations.index = annotations.index.str.replace('.', '-')
annotations['sample_id'] = annotations.index
annotations = annotations.fillna("NA")
print(annotations.shape)
annotations.head()
cell_id = 'A1-D041914-3_8_M-1-1'
annotations.loc[cell_id]
# +
ngenes_ncells = pd.read_csv("https://raw.githubusercontent.com/czbiohub/tabula-muris/master/00_data_ingest/13_ngenes_ncells_facs/Bladder_nreads_ngenes.csv", index_col=0)
ngenes_ncells.index = ngenes_ncells.index.str.replace('.', '-')
ngenes_ncells['sample_id'] = ngenes_ncells.index
ngenes_ncells = ngenes_ncells.fillna("NA")
ngenes_ncells.head()
# -
ngenes_ncells.loc[cell_id]
| notebooks/n_kmers_per_single_cell.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from disan_keras_layers import DISAN
from disan_keras import get_attn, plot_attn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from keras.models import Model
from keras.layers import Input, Dense, Reshape
from keras.optimizers import Adadelta
from keras.callbacks import ReduceLROnPlateau, ModelCheckpoint, TensorBoard, EarlyStopping
import pandas_datareader.data as web
from sklearn.preprocessing import MinMaxScaler
import os
# %matplotlib inline
df = web.get_data_yahoo('SPY')
df.head()
ts = df[['Adj Close', 'Open', 'High', 'Low']].dropna()
ts.plot(subplots=True);
# +
t = 10
train_perc = 0.9
#ts_scaled = MinMaxScaler().fit_transform(ts.values.reshape(-1, 1))[:, 0]
ts_scaled = np.diff(np.log(ts.values + 1), axis=0)
print(ts_scaled.min(), ts_scaled.max(), ts_scaled.shape)
X, y = [], []
for i in range(len(ts_scaled)):
if i >= t:
X.append(ts_scaled[i-t:i])
y.append(ts_scaled[i, 0])
X, y = np.asarray(X), np.asarray(y)
train_len = int(len(X) * train_perc)
X_train, y_train = X[:train_len, ...], y[:train_len, ...]
X_test, y_test = X[train_len:, ...], y[train_len:, ...]
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
assert not np.isnan(ts_scaled).any() and not np.isnan(ts_scaled).any() and not np.isinf(ts_scaled).any() and not np.isinf(ts_scaled).any()
# +
n = 64
ts_input = Input(shape=(t, 4))
#ts_reshape = Reshape((t, 1))(ts_input)
disan = DISAN(n, dropout=0.2)
rep_2d = disan(ts_input)
output = Dense(1, activation='linear')(rep_2d)
model = Model(ts_input, output)
opt = Adadelta(lr=0.5)
model.compile(loss='mse', optimizer=opt, metrics=['mae'])
model.summary()
# +
filepath = 'models'
callbacks = [
ReduceLROnPlateau(patience=0, verbose=1),
EarlyStopping(patience=3, verbose=1),
#ModelCheckpoint(filepath, verbose=1, save_best_only=True, save_weights_only=True),
#TensorBoard(os.path.join(filepath, 'logs'), histogram_freq=1)
]
model.fit(X_train, y_train, validation_split=0.1, epochs=100, verbose=1, callbacks=callbacks)
print(model.evaluate(X_test, y_test, batch_size=len(X_test)))
# +
attn = get_attn(X_test[np.newaxis, np.random.randint(len(X_test)), ...], model, disan.attn_dict)
for k, v in attn.items():
print(k, v.shape)
plot_attn(attn)
# -
y_pred = model.predict(X_test)[:, 0]
y_pred.shape
plt.scatter(y_test, y_pred);
| disan_ts_pred.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # “But it wouldn’t be an encyclopedia; it would be a wiki”: the changing imagined affordances of wikis, 1995-2002
#
# ## <NAME> (@staeiou, <EMAIL>)
# ### UC-Berkeley, Berkeley Institute for Data Science
# ### Association of Internet Researchers (IR17)
# ### Tartu, Estonia | 19 Oct 2017
# + [markdown] slideshow={"slide_type": "slide"}
# ## About me
#
# - Bio
# - Grew up on the internets (while living in Waco, Texas)
# - Ph.D from UC-Berkeley's [School of Information](http://ischool.berkeley.edu)
# - Now an "ethnographer/postdoc" at the [Berkeley Institute for Data Science](http://bids.berkeley.edu)
# + [markdown] slideshow={"slide_type": "fragment"}
# - Disciplines/fields
# - Science and Technology Studies
# - Communication and Media Studies
# - Critical Theory / Cultural Studies
# - Social Informatics / Computer-Supported Cooperative Work
#
# + [markdown] slideshow={"slide_type": "fragment"}
# - Methods/methodologies
# - Ethnography / trace ethnography
# - Grounded theory / qualitative contextual inquiry
# - Computationally-supported analysis of trace data
# - Archival and historical analysis / historiographical hermenutics
# + [markdown] slideshow={"slide_type": "slide"}
# # The socio-technical governance of Wikipedia
# + [markdown] slideshow={"slide_type": "subslide"}
# # Why does Wikipedia look like this?
#
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# # Instead of this?
#
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # Wikipedia is produced out of the tension to be both a “wiki-” and a “-pedia”
#
# (also see [Rosenzweig 2006](https://www.sfu.ca/cmns/courses/2012/801/1-Readings/Rosenzweig-%20Can%20history%20be%20open%20source%20.pdf), [Lih 2009](https://en.wikipedia.org/wiki/The_Wikipedia_Revolution), [Reagle 2010](https://mitpress.mit.edu/books/good-faith-collaboration), [Niederer & <NAME> 2010](http://journals.sagepub.com/doi/pdf/10.1177/1461444810365297), [Ford 2014](https://papers.ssrn.com/sol3/papers.cfm?abstract_id=2490454), [Wadewitz 2014](https://www.hastac.org/blogs/paigecm/2014/01/05/01-reading-and-writing-wikipedia-prospective-syllabus), Tkacz 2015)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Wikipedia was neither the first encyclopedia nor the first wiki
#
# Diderot and D'Alembert's [Encyclopedie](https://en.wikipedia.org/wiki/Encyclop%C3%A9die) | <NAME>ham's [WikiWikiWeb](http://wiki.c2.com/?WikiWikiWeb)
# :-------------------------:|:-------------------------:
#  | 
# + [markdown] slideshow={"slide_type": "skip"}
# 
#
# Image by [Wittylama](https://en.wikipedia.org/wiki/User:Wittylama), CC BY-SA 3.0
# From [Wikimedia Commons](https://commons.wikimedia.org/wiki/File:Denis_Diderot_conference_room.jpg)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Wikipedians started out with the same platform as late 1990s wikis
#
# 
#
# (archived from [archive.org](https://web.archive.org/web/20010405142805/http://www.wikipedia.com:80/wiki/Critical_Theory))
# + [markdown] slideshow={"slide_type": "subslide"}
# # But Wikipedia became a quite different kind of wiki
#
# 
# + [markdown] slideshow={"slide_type": "fragment"}
# # As Wikipedians worked out how to write an encyclopedia in a wiki, they revised the meanings and materialities of both.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# # Imagined affordances ([Nagy & Neff 2015](http://journals.sagepub.com/doi/abs/10.1177/2056305115603385)):
#
# # “Imagined affordances emerge between users’ perceptions, attitudes, and expectations; between the materiality and functionality of technologies; and between the intentions and perceptions of designers.”
# + [markdown] slideshow={"slide_type": "slide"}
# # The pre-history of Wikipedia
# + [markdown] slideshow={"slide_type": "fragment"}
# ## 1999-2000: [Nupedia](https://web.archive.org/web/20001203141800/http://www.nupedia.com:80/): Nupedia was a volunteer-based project to write a freely licensed encyclopedia.
#
# ## It was highly-structured, expert-based, and...
#
#
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# ## ...a failure. After 18 months & $250,000 USD, only 12 articles.
#
# 
#
# (From [archive.org](https://web.archive.org/web/20010331191028/http://www.nupedia.com:80/newest.phtml), layout edited to fit)
# + [markdown] slideshow={"slide_type": "subslide"}
# # January 2001: <NAME> & <NAME> learn about WikiWikiWeb, Jimmy asks them a simple question:
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "fragment"}
# 
#
# (archived from [WikiWikiWeb/WikiPedia](wiki.c2.com/?WikiPedia), as told in [Sanger's memoir](https://features.slashdot.org/story/05/04/18/164213/the-early-history-of-nupedia-and-wikipedia-a-memoir))
# + [markdown] slideshow={"slide_type": "fragment"}
# ### (They go ahead and make Wikipedia anyway)
# + [markdown] slideshow={"slide_type": "slide"}
# # So what was a wiki pre-Wikipedia?
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## The fast, quick, read-write version of the WorldWideWeb
#
# 
#
# (the php wiki, archived by [<NAME>](https://www.wired.com/2010/03/0325wikiwikiweb-first-wiki/))
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Originally, wikis did not:
# + [markdown] slideshow={"slide_type": "fragment"}
# - keep full revision histories (with UIs for browsing history)
# - separate article content from discussions about content
# - be able to temporarily protect pages from public editing
# - use or have built-in support academic-style references
# - templates for specialized formatting and task signalling __[citation needed]__
#
# + [markdown] slideshow={"slide_type": "fragment"}
# ## Pre-Wikipedia "wikizens" linked software features (or lack thereof) to ideas about what "the wiki way" demanded from them
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # Imagined affordance #1:
#
# ## The "WikiNow" versus a full version control system
# + [markdown] slideshow={"slide_type": "fragment"}
# ### "WhyWikiWorks" from WikiWikiWeb:
#
# 
#
# (archived on [archive.org](https://web.archive.org/web/20030401184429/http://c2.com:80/cgi/wiki?WhyWikiWorks))
# + [markdown] slideshow={"slide_type": "subslide"}
# 
#
# (archived on [archive.org](https://web.archive.org/web/20020717030429/http://c2.com:80/cgi/wiki?PerpetualNow))
# + [markdown] slideshow={"slide_type": "subslide"}
# # To Wikipedians, the lack of a full revision history was a bug
#
# 
#
# (from a [wikipedia-l discussion](https://lists.wikimedia.org/pipermail/wikipedia-l/2001-September/000494.html))
# + [markdown] slideshow={"slide_type": "subslide"}
# # Controversies over admins deleting pages from the database with no record
#
# 
#
# (from a [wikipedia-l discussion](https://lists.wikimedia.org/pipermail/wikipedia-l/2001-November/))
# + [markdown] slideshow={"slide_type": "subslide"}
# # Soon, the codebase was patched to support full histories and logging
#
# 
#
# (archived on [archive.org](https://web.archive.org/web/20011216233557/http://www.wikipedia.com:80/wiki.cgi?action=history&id=Creationism))
# + [markdown] slideshow={"slide_type": "slide"}
# # Imagined affordance #2:
# ## Wiki pages as stream of consciousness vs. separate pages for content and discussion
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# # Wikipedia's talk pages, separating content from discussion
#
# Wikipedia article | Wikipedia talk page
# :-------------------------:|:-------------------------:
#  | 
# + [markdown] slideshow={"slide_type": "slide"}
# # I'm going to have to skip over some slides now
# -
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # Wikipedia would become both an encyclopedia and a wiki, as Wikipedians revised both what encyclopedias and wikis were.
# + [markdown] slideshow={"slide_type": "fragment"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
# # Theoretical takeaways
#
# - Software platforms are not reducible to their code/features
# - Platforms are also constituted by their users' shared cultural imaginaries
# - Affordances are not equivilent to code/features
# - People attach different meanings/implications to the same features
# - "What the technology demands" is often fluid and negotiable
# - Tensions between tailoring society to code and code to society
# -
| aoir2017/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Here is an example of how to get access to all data
# ### Import your security token from file
import os
# env = %env
token_name = "<PASSWORD>"
token_file = token_name
has_error = False
if os.path.isfile(token_file):
__token__ = open(token_file, "r").read()
if len(__token__) == 0:
has_error = True
else:
has_error = True
if has_error:
print("\x1b[31m")
print("!!! [_Security_Token_Error_] !!!")
print("Please copy and save a valid PIC-SURE authentication token value into the file \""+token_name+"\".")
print("This file is located in the current Notebook directory.")
open(token_file, "w").write("")
else:
print("\x1b[32m")
print("[_Security_Token_Imported_Correctly_]")
# ### Connect to the NHANES data resource using the HPDS Adapter
# +
import PicSureClient
import PicSureHpdsLib
client = PicSureClient.Client()
connection = client.connect("https://copdgene-dev.hms.harvard.edu/picsure/", __token__, allowSelfSignedSSL=True)
adapter = PicSureHpdsLib.Adapter(connection)
adapter.list()
# -
resource = adapter.useResource("b6ef7b1a-56f6-11e9-8958-0242c0a83007")
# ### Download all data dictionary items
all_terms = resource.dictionary().find()
term_list = all_terms.keys()
term_list.sort()
term_list
all_terms.DataFrame()
# ### Get all subject records
query = resource.query()
query.help()
# +
query.select().add(all_terms.keys())
# add the correct consent group
query.filter().add("\\00 Consent groups\\", ["COPD_HMB","COPD_DS-CS-RD"])
# -
query.getResultsDataFrame()
| jupyter-notebooks/For_COPD_HMB_COPD_DS-CS-RD_Users/MISC+-+Download+All+Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="g_nWetWWd_ns"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="2pHVBk_seED1"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="M7vSdG6sAIQn"
# # TensorFlow Lite Model Analyzer
# + [markdown] id="fwc5GKHBASdc"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/lite/guide/model_analyzer"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/guide/model_analyzer.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/guide/model_analyzer.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/tensorflow/tensorflow/lite/g3doc/guide/model_analyzer.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="9ee074e4"
# TensorFlow Lite Model Analyzer API provides a way to analyze the given tflite_model with dumping model structure.
#
# + [markdown] id="JKwW0VfDKMWS"
# ## Model Analyzer API
#
# The following API is available for the TensorFlow Lite Model Analyzer.
#
# ```
# tf.lite.experimental.Analyzer.analyze(model_path=None,
# model_content=None,
# gpu_compatibility=False)
# ```
#
# You can find the API details from https://www.tensorflow.org/api_docs/python/tf/lite/experimental/Analyzer or run `help(tf.lite.experimental.Analyzer.analyze)` from a Python terminal.
#
# + [markdown] id="qi8Vk4_065jN"
# ## Basic usage with simple Keras model
#
# The following code shows basic usage of Model Analyzer. It shows contents of the converted Keras model in TFLite model content, formatted as a flatbuffer object.
# + id="_jkg6UNtdz8c"
import tensorflow as tf
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(128, 128)),
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10)
])
fb_model = tf.lite.TFLiteConverter.from_keras_model(model).convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model)
# + [markdown] id="pe_ZU5Zy7PeH"
# ## Basic usage with MobileNetV3Large Keras model
#
# This API works with large models such as MobileNetV3Large. Since the output is large, you might want to browse it with your favorite text editor.
# + id="QFywJ_g56VW5"
model = tf.keras.applications.MobileNetV3Large()
fb_model = tf.lite.TFLiteConverter.from_keras_model(model).convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model)
# + [markdown] id="4BGqG2j9yqRf"
# ## Check GPU delegate compatibility
#
# The ModelAnalyzer API provides a way to check the [GPU delegate](https://www.tensorflow.org/lite/performance/gpu) compatibility of the given model by providing `gpu_compatibility=True` option.
#
# + [markdown] id="sVGC1oX33RkV"
# ### Case 1: When model is incompatibile
#
# The following code shows a way to use `gpu_compatibility=True` option for simple tf.function which uses `tf.slice` with a 2D tensor and `tf.cosh` which are not compatible with GPU delegate.
#
# You will see `GPU COMPATIBILITY WARNING` per every node which has compatibility issue(s).
# + id="9GEg5plIzD-3"
import tensorflow as tf
@tf.function(input_signature=[
tf.TensorSpec(shape=[4, 4], dtype=tf.float32)
])
def func(x):
return tf.cosh(x) + tf.slice(x, [1, 1], [1, 1])
converter = tf.lite.TFLiteConverter.from_concrete_functions(
[func.get_concrete_function()], func)
converter.target_spec.supported_ops = [
tf.lite.OpsSet.TFLITE_BUILTINS,
tf.lite.OpsSet.SELECT_TF_OPS,
]
fb_model = converter.convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model, gpu_compatibility=True)
# + [markdown] id="BFU7HYb_2a8M"
# ### Case 2: When model is compatibile
#
# In this example, the given model is compatbile with GPU delegate.
#
# **Note:** Even though the tool doesn't find any compatibility issue, it doesn't guarantee that your model works well with GPU delegate on every device. There could be some runtime incompatibililty happen such as missing `CL_DEVICE_IMAGE_SUPPORT` feature by target OpenGL backend.
#
# + id="85RgG6tQ3ABT"
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(128, 128)),
tf.keras.layers.Dense(256, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10)
])
fb_model = tf.lite.TFLiteConverter.from_keras_model(model).convert()
tf.lite.experimental.Analyzer.analyze(model_content=fb_model, gpu_compatibility=True)
| site/en-snapshot/lite/guide/model_analyzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
from sklearn import tree
# +
## Set up initial data
###In this case, we make the data categorical instead of numeric
###Data often needs to be transformed before it is usable
# -
features_data = [[140, 'bumpy'], [130, 'bumpy'], [150, 'smooth'], [170, 'smooth']]
labels_data = ['orange', 'orange', 'apple', 'apple']
# +
## Transform the feature and label data
### Many algorithms do not deal with categorical data
### So we encode the categories as a series of 0/1 fields
# +
for feature in features_data:
if feature[1] == 'bumpy':
feature[1] = 1
if feature[1] == 'smooth':
feature[1] = 0
features = features_data
labels = []
for label in labels_data:
if label == 'orange':
labels.append(0)
if label == 'apple':
labels.append(1)
print (features)
print (labels)
# -
clf = tree.DecisionTreeClassifier()
clf = clf.fit(features, labels)
print (clf.predict([[160,0]]))
| notebooks/ML Workshop 01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Data Structures
#
# To keep this documentation generic we typically use dimensions `x` or `y`, but this should *not* be seen as a recommendation to use these labels for anything but actual positions or offsets in space.
#
# ## Variable
#
# ### Basics
#
# [scipp.Variable](../generated/classes/scipp.Variable.rst#scipp.Variable) is a labeled multi-dimensional array.
# A variable has the following key properties:
#
# - `values`: a multi-dimensional array of values, e.g., a [numpy.ndarray](https://numpy.org/doc/stable/reference/generated/numpy.ndarray.html#numpy.ndarray)
# - `variances`: a (optional) multi-dimensional array of variances for the array values
# - `dims`: a list of dimension labels (strings) for each axis of the array
# - `unit`: a (optional) physical unit of the values in the array
#
# Note that variables, unlike [DataArray](data-structures.ipynb#DataArray) and its eponym [xarray.DataArray](http://xarray.pydata.org/en/stable/user-guide/data-structures.html#dataarray), variables do *not* have coordinate dicts.
import numpy as np
import scipp as sc
# Variables should generally be created using one of the available [creation functions](../reference/creation-functions.rst#creation-functions).
# For example, we can create a variable from a numpy array:
var = sc.array(dims=['x', 'y'], values=np.random.rand(2, 4))
# <div class="alert alert-info">
#
# **Note:**
#
# Internally scipp is not using numpy, so the above makes a *copy* of the numpy array of values into an internal buffer.
#
# </div>
# We can inspect the created variable as follows:
sc.show(var)
var
var.unit
var.values
print(var.variances)
# Variances must have the same shape as values, and units are specified using the [scipp.units](../reference/units.rst) module or with a string:
var = sc.array(dims=['x', 'y'],
unit='m/s',
values=np.random.rand(2, 4),
variances=np.random.rand(2, 4))
sc.show(var)
var
var.variances
# ### 0-D variables (scalars)
#
# A 0-dimensional variable contains a single value (and an optional variance).
# The most convenient way to create a scalar variable is by multiplying a value by a unit:
scalar = 1.2 * sc.units.m
sc.show(scalar)
scalar
# Singular versions of the `values` and `variances` properties are provided:
print(scalar.value)
print(scalar.variance)
# An exception is raised from the `value` and `variance` properties if the variable is not 0-dimensional.
# Note that a variable with one or more dimension extent(s) of 1 contains just a single value as well, but the `value` property will nevertheless raise an exception.
# Creating scalar variables with variances or with custom `dtype` or variances is possible using [scipp.scalar](../generated/functions/scipp.scalar.rst#scipp.scalar):
var_0d = sc.scalar(value=1.0, variance=0.5, dtype=sc.dtype.float32, unit='kg')
var_0d
var_0d.value = 2.3
var_0d.variance
# ## DataArray
#
# ### Basics
#
# [scipp.DataArray](../generated/classes/scipp.DataArray.rst#scipp.DataArray) is a labeled array with associated coordinates.
# A data array is essentially a [Variable](../generated/classes/scipp.Variable.rst#scipp.Variable) object with attached dicts of coordinates, masks, and attributes.
#
# A data array has the following key properties:
#
# - `data`: the variable holding the array data.
# - `coords`: a dict-like container of coordinates for the array, accessed using a string as dict key.
# - `masks`: a dict-like container of masks for the array, accessed using a string as dict key.
# - `attrs`: a dict-like container of "attributes" for the array, accessed using a string as dict key.
#
# The key distinction between coordinates (added via the `coords` property) and attributes (added via the `attrs` property) is that the former are required to match ("align") in operations between data arrays whereas the latter are not.
#
# `masks` allows for storing boolean-valued masks alongside data.
# All four have items that are internally a [Variable](../generated/classes/scipp.Variable.rst#scipp.Variable), i.e., they have a physical unit and optionally variances.
a = sc.DataArray(
data = sc.array(dims=['y', 'x'], values=np.random.rand(2, 3)),
coords={
'y': sc.array(dims=['y'], values=np.arange(2.0), unit='m'),
'x': sc.array(dims=['x'], values=np.arange(3.0), unit='m')},
attrs={
'aux': sc.array(dims=['x'], values=np.random.rand(3))})
sc.show(a)
# Note how the `'aux'` attribute is essentially a secondary coordinate for the x dimension.
# The dict-like `coords` and `masks` properties give access to the respective underlying variables:
a.coords['x']
a.attrs['aux']
# Access to coords and attrs in a unified manner is possible with the `meta` property.
# Essentially this allows us to ignore whether a coordinate is aligned or not:
a.meta['x']
a.meta['aux']
# Unlike `values` when creating a variable, `data` as well as entries in the meta data dicts (`coords`, `masks`, and `attrs`) are *not* deep-copied on insertion into a data array.
# To avoid unwanted sharing, call the `copy()` method.
# Compare:
x2 = sc.zeros(dims=['x'], shape=[3])
a.coords['x2_shared'] = x2
a.coords['x2_copied'] = x2.copy()
x2 += 123
a
# Meta data can be removed in the same way as in Python dicts:
del a.attrs['aux']
# ### Distinction between dimension coords and non-dimension coords, and coords and attrs
#
# When the name of a coord matches its dimension, e.g., if `d.coord['x']` depends on dimension `'x'` as in the above example, we call this coord *dimension coordinate*.
# Otherwise it is called *non-dimension coord*.
# It is important to highlight that for practical purposes (such as matching in operations) **dimension coords and non-dimension coords are handled equivalently**.
# Essentially:
#
# - **Non-dimension coordinates are coordinates**.
# - There is at most one dimension coord for each dimension, but there can be multiple non-dimension coords.
# - Operations such as value-based slicing that accept an input dimension and require lookup of coordinate values will only consider dimension coordinates.
#
# As mentioned above, the difference between coords and attrs is "alignment", i.e., only the former are compared in operations.
# The concept of dimension coords is unrelated to the distinction between `coords` or `attrs`.
# In particular, dimension coords could be made attrs if desired, and non-dimension coords can (and often are) "aligned" coords.
# ## Dataset
#
# [scipp.Dataset](../generated/classes/scipp.Dataset.rst#scipp.Dataset) is a dict-like container of data arrays.
# Individual items of a dataset ("data arrays") are accessed using a string as a dict key.
#
# In a dataset the coordinates of the sub-arrays are enforced to be *aligned*.
# That is, a dataset is not actually just a dict of data arrays.
# Instead, the individual arrays share their coordinates.
# It is therefore not possible to combine *arbitrary* data arrays into a dataset.
# If, e.g., the extents in a certain dimension mismatch, or if coordinate values mismatch, insertion of the mismatching data array will fail.
#
# Often a dataset is not created from individual data arrays.
# Instead we may provide a dict of variables (the data of the items), and dicts for coords:
d = sc.Dataset(
data={
'a': sc.array(dims=['y', 'x'], values=np.random.rand(2, 3)),
'b': sc.array(dims=['y'], values=np.random.rand(2)),
'c': sc.scalar(value=1.0)},
coords={
'x': sc.array(dims=['x'], values=np.arange(3.0), unit='m'),
'y': sc.array(dims=['y'], values=np.arange(2.0), unit='m'),
'aux': sc.array(dims=['x'], values=np.random.rand(3))})
sc.show(d)
d
d.coords['x'].values
# The name of a data item serves as a dict key.
# Item access returns a new data array which is a view onto the data in the dataset and its corresponding coordinates, i.e., no deep copy is made:
sc.show(d['a'])
d['a']
# Use the `copy()` method to turn the view into an independent object:
copy_of_a = d['a'].copy()
copy_of_a += 17 # does not change d['a']
d
# Each data item is linked to its corresponding coordinates, masks, and attributes.
# These are accessed using the `coords` , `masks`, and `attrs` properties.
# The variable holding the data of the dataset item is accessible via the `data` property:
d['a'].data
# For convenience, properties of the data variable are also properties of the data item:
d['a'].values
d['a'].variances
d['a'].unit
# Coordinates of a data item include only those that are relevant to the item's dimensions, all others are hidden.
# For example, when accessing `'b'`, which does not depend on the `'y'` dimension, the coord for `'y'` as well as the `'aux'` coord are not part of the item's `coords`:
sc.show(d['b'])
# Similarly, when accessing a 0-dimensional data item, it will have no coordinates:
sc.show(d['c'])
# All variables in a dataset must have consistent dimensions.
# Thanks to labeled dimensions, transposed data is supported:
d['d'] = sc.array(dims=['x', 'y'], values=np.random.rand(3, 2))
sc.show(d)
d
# When inserting a data array or variable into a dataset ownership is shared by default.
# Use the `copy()` method to avoid this if undesirable:
d['a_shared'] = a
d['a_copied'] = a.copy()
a += 1000
d
# The usual `dict`-like methods are available for `Dataset`:
for name in d:
print(name)
'a' in d
# + tags=[]
'e' in d
| docs/user-guide/data-structures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# name: python392jvsc74a57bd0aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# ---
# +
#Source : https://geohub.lacity.org/datasets/bus-stop-benches/
# -
import json
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import base64
CSV_FILE = 'bus_stops'
with open('dataRaw/'+ CSV_FILE +'.csv') as f:
df=pd.read_csv(f, delimiter=',')
# REMOVE COLUMNS
# BETTER DOCUMENTATION ON TYPES OF BUS STOPS is needed
df = df.drop(columns=['X', 'Y', 'TOOLTIP', 'MEDIA_TYPE', 'UNIT_TYPE'])
df.head(10)
df.to_csv('dataClean/'+ CSV_FILE +'.csv', index=False)
| bus_stops.ipynb |
#!/usr/bin/env python
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2-parameters discriminant analysis
#
# Python notebook for constructing a Fisher disciminant from two 2D Gaussianly distributed correlated variables. The notebook creates artificial random data for two different types of processes, and the goal is then to separate these by constructing a Fisher discriminant.
#
# ### Authors:
# - <NAME> (Niels Bohr Institute)
# - <NAME> (Niels Bohr Institute)
#
# ### Date:
# - 15-12-2021 (latest update)
#
# ### References:
# - <NAME>, Statistical Data Analysis, pages 51-57
# - http://en.wikipedia.org/wiki/Linear_discriminant_analysis
#
# ***
import numpy as np # Matlab like syntax for linear algebra and functions
import matplotlib.pyplot as plt # Plots and figures like you know them from Matlab
from numpy.linalg import inv
r = np.random # Random generator
r.seed(42) # Set a random seed (but a fixed one)
save_plots = False # For now, don't save plots (once you trust your code, switch on)
# ### Functions:
#
# Function to calculate the separation betweem two lists of numbers (see equation at the bottom of the script).
#
# __Note__: You need to fill in this function!
def calc_separation(x, y):
print("calc_separation needs to be filled out")
d = 0
return d
# ## Define parameters:
#
#
# Number of species, their means and widths, correlations and the number of observations of each species:
# +
# Number of 'species': signal / background
n_spec = 2
# Species A, mean and width for the two dimensions/parameters
mean_A = [15.0, 50.0]
width_A = [ 2.0, 6.0]
# Species B, mean and width for the two dimensions/parameters
mean_B = [12.0, 55.0]
width_B = [ 3.0, 6.0]
# Coefficient of correlation
corr_A = 0.8
corr_B = 0.9
# Amount of data you want to create
n_data = 2000
# -
# ## Generate data:
#
# For each "species", produce a number of $(x_0,x_1)$ points which are (linearly) correlated:
# +
# The desired covariance matrix.
V_A = np.array([[width_A[0]**2, width_A[0]*width_A[1]*corr_A],
[width_A[0]*width_A[1]*corr_A, width_A[1]**2]])
V_B = np.array([[width_B[0]**2, width_B[0]*width_B[1]*corr_B],
[width_B[0]*width_B[1]*corr_B, width_B[1]**2]])
# Generate the random samples.
spec_A = np.random.multivariate_normal(mean_A, V_A, size=n_data)
spec_B = np.random.multivariate_normal(mean_B, V_B, size=n_data)
# -
# ***
#
# ## Plot your generated data:
#
# We plot the 2D-data as 1D-histograms (basically projections) in $x_0$ and $x_1$:
# +
fig_1D, ax_1D = plt.subplots(ncols=2, figsize=(14, 6))
ax_1D[0].hist(spec_A[:, 0], 50, (0, 25), histtype='step', label='Species A', color='Red', lw=1.5)
ax_1D[0].hist(spec_B[:, 0], 50, (0, 25), histtype='step', label='Species B', color='Blue', lw=1.5)
ax_1D[0].set(title='Parameter x0', xlabel='x0', ylabel='Counts', xlim=(0,25))
ax_1D[0].legend(loc='upper left')
# uncomment later
#ax_1D[0].text(1, 176, fr'$\Delta_{{x0}} = {calc_separation(spec_A[:, 0], spec_B[:, 0]):.3f}$', fontsize=16)
ax_1D[1].hist(spec_A[:, 1], 50, (20, 80), histtype='step', label='Species A', color='Red', lw=1.5)
ax_1D[1].hist(spec_B[:, 1], 50, (20, 80), histtype='step', label='Species B', color='Blue', lw=1.5)
ax_1D[1].set(title='Parameter x1', xlabel='x1', ylabel='Counts', xlim=(20, 80))
ax_1D[1].legend(loc='upper left')
# uncomment later
#ax_1D[1].text(22, 140, fr'$\Delta_{{x1}} = {calc_separation(spec_A[:, 1], spec_B[:, 1]):.3f}$', fontsize=16)
fig_1D.tight_layout()
if save_plots :
fig_1D.savefig('InputVars_1D.pdf', dpi=600)
# -
# NOTE: Wait with drawing the 2D distribution, so that you think about the 1D distributions first!
#
# ***
# From the two 1D figures, it seems that species A and B can be separated to some degree, but not very well. If you were to somehow select cases of species A, then I can imagine a selection as follows:
# - If (x0 > 16) or (x1 < 46) or (x0 > 13 and x1 < 52), then guess / select as A.
#
# Think about this yourself, and discuss with your peers, how you would go about separating A from B based on x0 and x1.
#
# ----------------------- 5-10 minutes later -----------------------
#
# As it is, this type of selection is hard to optimise, especially with more dimensions (i.e. more variables than just x0 and x1). That is why Fisher's linear discriminant, $F$, is very useful. It makes the most separating linear combination of the input variables, and the coefficients can be calculated analytically. Thus, it is fast, efficient, and transparent. And it takes linear correlations into account.
# +
# fig_corr, ax_corr = plt.subplots(figsize=(14, 8))
# ax_corr.scatter(spec_A[:, 0], spec_A[:, 1], color='Red', s=10, label='Species A')
# ax_corr.scatter(spec_B[:, 0], spec_B[:, 1], color='Blue', s=10, label='Species B')
# ax_corr.set(xlabel='Parameter x0', ylabel='Parameter x1', title='Correlation');
# ax_corr.legend();
# fig_corr.tight_layout()
#if save_plots :
# fig_corr.savefig('InputVars_2D.pdf', dpi=600)
# -
# ## Fisher Discriminant calculation:
#
# We want to find $\vec{w}$ defined by:
#
# $$\vec{w} = \left(\Sigma_A + \Sigma_B\right)^{-1} \left(\vec{\mu}_A - \vec{\mu}_B\right)$$
#
# which we use to project our data into the best separating plane (line in this case) given by:
#
# $$ \mathcal{F} = w_0 + \vec{w} \cdot \vec{x} $$
#
# We start by finding the means and covariance of the individuel species: (__fill in yourself!__)
mu_A = 0 # fill in yourself
mu_B = 0 # fill in yourself
mu_A
cov_A = 0 # fill in yourself
cov_B = 0 # fill in yourself
cov_sum = cov_A + cov_B
cov_sum
# where `cov_sum` is the sum of the all of the species' covariance matrices. We invert this using scipy's `inv` function. __Note__: fill in yourself!
# +
# Delete the definition below of cov_sum when you have filled in the cells above:
cov_sum = np.diag([1, 2])
# Inverts cov_sum
cov_sum_inv = inv(cov_sum)
cov_sum_inv
# -
# We calculate the fisher weights, $\vec{w}$. __Note__: fill in yourself:
wf = np.ones(2) # fill in yourself
wf
# We calculate the fisher discriminant, $\mathcal{F}$. __Note__: fill in yourself:
fisher_data_A = spec_A[:, 0] * (-1.4) + 10 # fill in yourself
fisher_data_B = spec_B[:, 0] * (-1.4) + 10 # fill in yourself
# and plot it:
# +
fig_fisher, ax_fisher = plt.subplots(figsize=(12, 8))
ax_fisher.hist(fisher_data_A, 200, (-22, 3), histtype='step', color='Red', label='Species A')
ax_fisher.hist(fisher_data_B, 200, (-22, 3), histtype='step', color='Blue', label='Species B')
ax_fisher.set(xlim=(-22, 3), xlabel='Fisher-discriminant')
ax_fisher.legend()
# ax_fisher.text(-21, 60, fr'$\Delta_{{fisher}} = {calc_separation(fisher_data_A, fisher_data_B):.3f}$', fontsize=16)
fig_fisher.tight_layout()
if save_plots:
fig_fisher.savefig('FisherOutput.pdf', dpi=600)
# -
# It is easy to visually see the increased seperation (when done correctly). We can also compare $\Delta_{fisher}$ to $\Delta_{x0}$ or $\Delta_{x1}$ and see it clearly.
# # Questions
#
# As always, make sure that you know what the code is doing so far, and what the aim of the exercise is (i.e. which problem to solve, and how). Then start to expand on it.
#
# 1. Look at the 1D distributions of the two discriminating variables for the two species, and see how well you can separate them by eye. It seems somewhat possible, but certainly far from perfect... Once you consider the 2D distribution (scatter plot - to be uncommented by you!), then it is clear, that some cut along a line at an angle will work much better. This exercise is about finding that optimal line, and thus the perpendicular axis to project the data onto!
#
# 2. Calculate from the data the mean, widths (std) and covariance of each discriminating variable (pair of variables for covariance) for each species, and put these into the matrices defined.
#
# 3. From the inverted summed covariance matrices and vectors of means, calculate the two Fisher coefficients, and given these, calculate the Fisher discriminant for the two species in question, i.e. $ \mathcal{F} = \vec{w} \cdot \vec{x} = w_x \cdot x + w_y \cdot y $ for each point (x,y).
#
# 4. What separation did you get, and is it notably better than what you obtain by eye? Also, do your weights make sense? I.e. are they comparable to the widths of the
# corresponding variable? As a simple measure of how good the separation obtained is, we consider the "distance" $\Delta$ between the two distributions as a measure of goodness:
#
# $$\Delta = \frac{|\mu_A-\mu_B|}{\sqrt{\sigma_A^2+\sigma_B^2}}$$
#
# Compare the separation you get from each of the two 1D histograms of $x_0$ and $x_1$ with what you get from the Fisher discriminant, using the above formula. Of course the ultimate comparison should be done using ROC curves!
| AppStat2022/Week5/original/MVA_part1/2par_discriminant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>AutomationDirect P3-530 PLC</h1>
#
# This device is a programmable logic controller. The firmware can be found here:
#
# https://ftp.automationdirect.com/firmware/FirmwareOnly/P3-530_1.2.7.39.adfw
#
# When unpacked with Binwalk, there are several binary files that can be analyzed. The file analyzed here is `P3_530.bin`. When a signature scan of this file is run, the output is as follows:
#
# ```
# $ binwalk P3_530.bin
#
# DECIMAL HEXADECIMAL DESCRIPTION
# --------------------------------------------------------------------------------
# 1140488 0x116708 Copyright string: "Copyright MGC 2003 - Nucleus PLUS - MPC824x Metrowerks v. 1.14"
# 1141840 0x116C50 Base64 standard index table
# 1143036 0x1170FC HTML document header
# 1157304 0x11A8B8 DES SP1, big endian
# 1157560 0x11A9B8 DES SP2, big endian
# 1160064 0x11B380 SHA256 hash constants, big endian
# 1225624 0x12B398 CRC32 polynomial table, big endian
# 1300760 0x13D918 HTML document header
# 1301521 0x13DC11 HTML document footer
# 1301644 0x13DC8C HTML document header
# 1301793 0x13DD21 HTML document footer
# 1301804 0x13DD2C HTML document header
# 1301892 0x13DD84 HTML document footer
# 1313328 0x140A30 HTML document header
# 1314231 0x140DB7 HTML document footer
# 1314532 0x140EE4 HTML document header
# 1315095 0x141117 HTML document footer
# 1315104 0x141120 HTML document header
# 1315182 0x14116E HTML document footer
# 1315192 0x141178 HTML document header
# 1315279 0x1411CF HTML document footer
# 1319268 0x142164 PEM certificate
# 1319324 0x14219C PEM certificate request
# 1319512 0x142258 PEM RSA private key
# 1319708 0x14231C PEM EC private key
# 1319776 0x142360 PEM DSA private key
#
# ```
#
# Time to use Centrifuge.
# +
import sys
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [16, 9]
sys.path[0:0] = ['.', '..']
from centrifuge.binfile import BinFile
# -
file_handle = open("_P3-530_172.16.58.3.adfw.extracted/P3_530.bin", "rb")
plc_bin = BinFile(file_handle)
plc_bin.slice_file()
# We can look at a couple of plots to get our bearings:
plc_bin.plot_file_entropy()
plc_bin.plot_file_feature("zeroes", "black")
# There seem to be 2 large discrete areas with a few smaller ones. Let's see what can be found with clustering:
plc_bin.cluster_DBSCAN(epsilon=0.7,
minimum_samples=10,
find_optimal_epsilon=True)
plc_bin.plot_DBSCAN_results()
results = plc_bin.identify_cluster_data_types()
# Turns out that there is PowerPC machine code present. For fun, we can take a look at the UTF-8 data in cluster 2:
_, cluster_byte_values = plc_bin.extract_clusters()
bytes(cluster_byte_values[2])[:1000]
# Full results:
results
| notebooks/Analyzing Firmware with Centrifuge Example 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Young Star Magnetic Models
#
# Convergence of magnetic models is improved with a [new treatment of the peak magnetic field strength](http://nbviewer.ipython.org/github/gfeiden/Notebook/blob/master/Daily/20150728_peak_magnetic_field.ipynb) definition. Previously, it was defined as either $R_{\rm peak} = 0.5R_{\star}$ or $R_{\rm peak} = R_{\rm tach}$, where $R_{\rm tach}$ is the radial location of the interface region between the stellar radiation and convection zones (i.e., the tachocline). This caused problems for young star models as models start off fully convective but develop radiative cores as the central temperature increases throughout its gravitational contraction. Magnetic fields therefore jumped rapidly from a fully convective treatment to a partially convective treatment, leading to excessively large interior magnetic field strengths. To avoid this problem, the peak magnetic field strength is treated as either $R_{\rm peak} = 0.5R_{\star}$ or $R_{\rm peak} = R_{\rm tach}$, _whichever is larger_, in all cases.
#
# Two small grids of magnetic models are computed with GS98 and GAS07 solar abundances. These may be incorporated into the Young Star manuscript, where we present models of young stars that have now been used in several publications (e.g., [Malo et al. 2014](http://adsabs.harvard.edu/abs/2014arXiv1406.6750M); [Herczeg & Hillenbrand 2015](http://adsabs.harvard.edu/abs/2015arXiv150506518H)). However, these models are computed, specifically, at the request of <NAME>, who wishes to incorporate magnetic models into an analysis. The tracks themselves will not be incorporated into the GitHub repo, as publishing the full grid would require too much disk space, but they are available [upon request by creating an "issue"](https://github.com/gfeiden/Notebook/issues).
#
# __Update__: raw magnetic mass tracks are contained in a tarball in the [`files/` directory](https://github.com/gfeiden/Notebook/tree/master/Daily/files) with the extension `_mtrks.tgz`.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# ## Magnetic Mass Tracks
#
# We'll start with loading mass tracks from the GAS07 solar abundance subset. These adopt surface boundary conditions from the MARCS model atmosphere structures. While we typically recommend surface boundary conditions be attached at an optical depth where $\tau_{\rm ross} \ge 50$, the magnetic models are computed by fitting surface boundary conditions where $\tau_{\rm ross} \ge 10$. Magnetic fields largely affect the super-adiabiatic layers near the stellar surface, with deeper field strenghts playing a less critical role ([Feiden & Chaboyer 2013](http://adsabs.harvard.edu/abs/2013ApJ...779..183F), [2014](http://adsabs.harvard.edu/abs/2014A%26A...571A..70F)). However, the motivation for attaching the boundary conditions at larger optical depths is to provide a better treatment of super-adiabatic layers where radiation and convection are both significnat contributors to the total energy flux ([Chabrier & Baraffe 1997](http://adsabs.harvard.edu/abs/1997A%26A...327.1039C)), which is in opposition to our efforts of including the effects of magnetic fields.
#
# We provide a compromise by fixing the surface boundary conditions at higher layer in the star. This provides a sufficiently large super-adiabatic layer to give the magnetic field a reasonable influence, while still providing a reliable estimate of the surface conditions that help set the overall thermal structure of the star.
masses = np.arange(0.1, 0.96, 0.05) # list of masses
# ## Magnetic Isochrones
#
# Process the magnetic mass tracks into isochrones. Since mass tracks are computed with a relatively course mass resolution ($0.05 M_{\odot}$), spline interpolation is used to smooth the resulting isochrones with a finer mass resolution.
#
# Below, a grid of isochrones is computed from 5 to 30 Myr in steps of 1 Myr.
# +
from scipy.interpolate import interp1d
ages = np.arange(5.0e6, 3.1e7, 1.0e6) # ages requested
# -
# ### Dartmouth & MARCS; Solar abundance: Grevesse, Asplund, & Sauval 2007
# +
# open output file objects
output_files = [open('files/dmestar_{:07.1f}myr_gas07_z+0.00_a+0.00_mag25kG.iso'.format(age/1.0e6), 'w')
for age in ages]
trk_directory = '../../evolve/dmestar/trk/gas07/p000/a0/amlt2040/mag25kG'
for mass in masses:
trk_filename = 'm{:04.0f}_GAS07_p000_p0_y26_mlt2.040_mag25kG.trk'.format(mass*1000.)
try:
gas07_trk = np.genfromtxt('{0}/{1}'.format(trk_directory, trk_filename), usecols=(0, 1, 2, 3, 4, 8))
except IOError:
continue
# extract only relevant age chunk for easier interpolation
gas07_trk = np.array([time_step for time_step in gas07_trk if 1.0e6 <= time_step[0] <= 5.0e7])
# generate linear interpolation curve as a function of age
try:
icurve = interp1d(gas07_trk[:, 0], gas07_trk[:, 1:], kind='linear', axis=0)
except IndexError:
continue
# extract properties at the requested age
trk_props = icurve(ages)
i = 0
for props in trk_props:
s = '{:6.3f}'.format(mass)
for prop in props:
if np.isnan(prop) or prop < -12.0:
prop = -12.0
s += '{:14.6f}'.format(prop)
s += '\n'
output_files[i].write(s)
i += 1
#print "{:4.2f} Mo Track Processed.".format(mass)
# close output files
for f in output_files:
f.close()
# -
# Interpolate isochrones onto a finer mass grid.
fine_mass_grid = np.arange(0.1, 0.95, 0.02)
for age in ages:
iso_filename = 'files/dmestar_{:07.1f}myr_gas07_z+0.00_a+0.00_mag25kG.iso'.format(age/1.0e6)
isochrone = np.genfromtxt(iso_filename)
# generate interpolation curve
icurve = interp1d(isochrone[:,0], isochrone[:,1:], axis=0, kind='slinear')
# interpolate onto a finer mass grid
fine_isochrone = icurve(fine_mass_grid)
fine_isochrone = np.column_stack((fine_mass_grid, fine_isochrone))
# write header
header = 'Dartmouth Stellar Evolution Model: Quick Isochrone \n\n'
header += 'Age = {:7.1f} Myr [Fe/H] = {:+5.2f} [a/Fe] = {:+5.2f} \n\n'.format(age/1.e6, 0.0, 0.0)
header += '{:^14} {:^14} {:^14} {:^14} {:^14} {:^14}'.format('Mass', 'log(Teff)', 'log(g)', 'log(L/Lo)',
'log(R/Ro)', 'A(Li)')
# overwrite original file
np.savetxt(iso_filename, fine_isochrone, fmt='%14.6f', header=header)
# Magnetic isochrones are stored in the directory [`files/`](https://github.com/gfeiden/Notebook/tree/master/Daily/files/) and follow the format outline in the two code snippets above. We can take a quick look at some of the proeprties of these isochrones and how they compare to standard stellar evolution isochrones (i.e., without a magnetic perturbation).
#
# A tarball with all of the above computed isochrones can be found in [`files/dmestar_gas07_z+0.00_a+0.00_mag25kG.tgz`](https://github.com/gfeiden/Notebook/tree/master/Daily/files/dmestar_gas07_z+0.00_a+0.00_mag25kG.tgz).
# ### Dartmouth & PHOENIX; Solar abundance: Grevesse & Sauval 1998
masses = np.arange(0.10, 0.86, 0.05) # higher masses did not converge (investigating)
# +
# open output file objects
output_files = [open('files/dmestar_{:07.1f}myr_gs98_z+0.00_a+0.00_mag25kG.iso'.format(age/1.0e6), 'w')
for age in ages]
trk_directory = '../../evolve/dmestar/trk/gs98/p000/a0/amlt1884/mag25kG'
for mass in masses:
trk_filename = 'm{:04.0f}_GS98_p000_p0_y28_mlt1.884_mag25kG.trk'.format(mass*1000.)
try:
gs98_trk = np.genfromtxt('{0}/{1}'.format(trk_directory, trk_filename), usecols=(0, 1, 2, 3, 4, 8))
except IOError:
continue
# extract only relevant age chunk for easier interpolation
gs98_trk = np.array([time_step for time_step in gs98_trk if 1.0e6 <= time_step[0] <= 5.0e7])
# generate linear interpolation curve as a function of age
try:
icurve = interp1d(gs98_trk[:, 0], gs98_trk[:, 1:], kind='linear', axis=0)
except IndexError:
continue
# extract properties at the requested age
trk_props = icurve(ages)
i = 0
for props in trk_props:
s = '{:6.3f}'.format(mass)
for prop in props:
if np.isnan(prop) or prop < -12.0:
prop = -12.0
s += '{:14.6f}'.format(prop)
s += '\n'
output_files[i].write(s)
i += 1
#print "{:4.2f} Mo Track Processed.".format(mass)
# close output files
for f in output_files:
f.close()
# -
# Interpolate onto a finer mass grid,
fine_mass_grid = np.arange(0.1, 0.85, 0.02)
for age in ages:
iso_filename = 'files/dmestar_{:07.1f}myr_gs98_z+0.00_a+0.00_mag25kG.iso'.format(age/1.0e6)
isochrone = np.genfromtxt(iso_filename)
# generate interpolation curves
icurve = interp1d(isochrone[:,0], isochrone[:,1:], axis=0, kind='slinear')
# interpolate onto a finer mass grid
fine_isochrone = icurve(fine_mass_grid)
fine_isochrone = np.column_stack((fine_mass_grid, fine_isochrone))
# write header
header = 'Dartmouth Stellar Evolution Model: Quick Isochrone \n\n'
header += 'Age = {:7.1f} Myr [Fe/H] = {:+5.2f} [a/Fe] = {:+5.2f} \n\n'.format(age/1.e6, 0.0, 0.0)
header += '{:^14} {:^14} {:^14} {:^14} {:^14} {:^14}'.format('Mass', 'log(Teff)', 'log(g)', 'log(L/Lo)',
'log(R/Ro)', 'A(Li)')
# overwrite original file
np.savetxt(iso_filename, fine_isochrone, fmt='%14.6f', header=header)
# Magnetic isochrones are stored in the directory [`files/`](https://github.com/gfeiden/Notebook/tree/master/Daily/files/) and follow the format outline in the two code snippets above. We can take a quick look at some of the proeprties of these isochrones and how they compare to standard stellar evolution isochrones (i.e., without a magnetic perturbation).
#
# A tarball with all of the above computed isochrones can be found in [`files/dmestar_gs98_z+0.00_a+0.00_mag25kG.tgz`](https://github.com/gfeiden/Notebook/tree/master/Daily/files/dmestar_gs98_z+0.00_a+0.00_mag25kG.tgz).
#
# ### Simple Diagnostic Plots
#
# Here are some simple diagnostic figures to assess that isochrones look smooth and do not deviate too significantly from expectation (i.e., they're smooth and properties change monotonically). Plot a few isochrones: 5 Myr, 12 Myr, and 30 Myr.
# +
# GS98 isochrones
gs98_05 = np.genfromtxt('files/dmestar_00005.0myr_gs98_z+0.00_a+0.00_mag25kG.iso')
gs98_12 = np.genfromtxt('files/dmestar_00012.0myr_gs98_z+0.00_a+0.00_mag25kG.iso')
gs98_30 = np.genfromtxt('files/dmestar_00030.0myr_gs98_z+0.00_a+0.00_mag25kG.iso')
# GAS07 isochrones
gas07_05 = np.genfromtxt('files/dmestar_00005.0myr_gas07_z+0.00_a+0.00_mag25kG.iso')
gas07_12 = np.genfromtxt('files/dmestar_00012.0myr_gas07_z+0.00_a+0.00_mag25kG.iso')
gas07_30 = np.genfromtxt('files/dmestar_00030.0myr_gas07_z+0.00_a+0.00_mag25kG.iso')
# +
fig, ax = plt.subplots(1, 2, figsize=(12, 6))
ax[0].set_title('GAS07 Series', fontsize=22.)
ax[1].set_title('GS98 Series', fontsize=22.)
for axis in ax:
axis.set_xlabel('Effective Temperature (K)', fontsize=20.)
axis.set_ylabel('$\\log (L / L_{\\odot})$', fontsize=20.)
axis.set_xlim(4500., 2500.)
axis.set_ylim(-2.5, 0.0)
axis.tick_params(which='major', axis='both', length=10., labelsize=16.)
# GAS07 series
ax[0].plot(10.0**gas07_05[:, 1], gas07_05[:, 3], '-', lw=2, color='#1e90ff')
ax[0].plot(10.0**gas07_12[:, 1], gas07_12[:, 3], '--', lw=2, color='#1e90ff')
ax[0].plot(10.0**gas07_30[:, 1], gas07_30[:, 3], '-.', lw=2, color='#1e90ff')
# GS98 series
ax[1].plot(10.0**gs98_05[:, 1], gs98_05[:, 3], '-', lw=2, color='#1e90ff')
ax[1].plot(10.0**gs98_12[:, 1], gs98_12[:, 3], '--', lw=2, color='#1e90ff')
ax[1].plot(10.0**gs98_30[:, 1], gs98_30[:, 3], '-.', lw=2, color='#1e90ff')
fig.tight_layout()
# -
# There looks to be some noise in the GS98 isochrones at the highest temperatures, which is likely related to the convergence issues with those above $0.90 M_{\odot}$. Nevertheless, the isochrones appear quite smooth.
#
# Quick look at Li depletion curves. ~~(note: due to issues with NaNs in the 28+ Myr isochrones, switching from 30 Myr to 27 Myr.)~~
# +
fig, ax = plt.subplots(1, 2, figsize=(12, 6))
ax[0].set_title('GAS07 Series', fontsize=22.)
ax[1].set_title('GS98 Series', fontsize=22.)
for axis in ax:
axis.set_xlabel('Effective Temperature (K)', fontsize=20.)
axis.set_ylabel('A(Li)', fontsize=20.)
axis.set_xlim(4500., 2500.)
axis.set_ylim(2.5, 3.5)
axis.tick_params(which='major', axis='both', length=10., labelsize=16.)
axis.plot([4500., 2500.], [3.30, 3.30], '--', lw=1, color="#555555")
# GAS07 series
ax[0].plot(10.0**gas07_05[:, 1], gas07_05[:, 5], '-', lw=2, color='#1e90ff')
ax[0].plot(10.0**gas07_12[:, 1], gas07_12[:, 5], '--', lw=2, color='#1e90ff')
ax[0].plot(10.0**gas07_30[:, 1], gas07_30[:, 5], '-.', lw=2, color='#1e90ff')
# GS98 series
ax[1].plot(10.0**gs98_05[:, 1], gs98_05[:, 5], '-', lw=2, color='#1e90ff')
ax[1].plot(10.0**gs98_12[:, 1], gs98_12[:, 5], '--', lw=2, color='#1e90ff')
ax[1].plot(10.0**gs98_30[:, 1], gs98_30[:, 5], '-.', lw=2, color='#1e90ff')
fig.tight_layout()
# -
# There are interpolation issues for ages greater than 28 Myr, as at least one of the models in the course grid had A(Li) = NaN. This leads to all interpolate values coming back as NaNs. And A(Li) does not appear to be so smooth, with a random bump at low temperatures that is clearly indicative of an artifact of the spline interpolation.
#
# __Update 01__: NaNs have now been removed. The gap in the GS98 A(Li) figure at 30 Myr is due to values listed as `-inf`.
#
# __Update 02__: `-inf` values have now been replaced by actual values.
# ## References
#
# If you use these models, please consider citing the following papers depending on the context of your work. Each potential reference is preceeded by a brief description of the paper.
#
# Original inspiration and basis for the magnetic Dartmouth stellar evolution models:
#
# > Lydon & Sofia (1995), ApJS, 101, 357 ([ADS](http://adsabs.harvard.edu/abs/1995ApJS..101..357L)).
#
# Framework for the magnetic models is the Dartmouth stellar evolution program (DSEP):
#
# > <NAME>, Jevremovic, _et al._ (2008), ApJS, 178, 89 ([ADS](http://adsabs.harvard.edu/abs/2008ApJS..178...89D)).
#
# Description and first demonstration of the magnetic Dartmouth stellar evolution code:
#
# > Feiden & Chaboyer (2012), ApJ, 761, 30 ([ADS](http://adsabs.harvard.edu/abs/2012ApJ...761...30F)).
#
# Demonstration of the magnetic code on three main sequence eclipsing binary systems whose stars are believed to possess a radiative core. Showed that magnetic field perturbation in the super-adiabatic region governs how the model is affected by the presence of global magnetic perturbation:
#
# > Feiden & Chaboyer (2013), ApJ, 779, 183 ([ADS](http://adsabs.harvard.edu/abs/2013ApJ...779..183F)).
#
# Demonstration of the magnetic code on two main sequence eclipsing binary systems whose stars are believed to be fully convective. Instituted the fixed peak magnetic field strength at $0.15 R_{\odot}$ for fully convective stars:
#
# > Feiden & Chaboyer (2014), ApJ, 786, 53 ([ADS](http://adsabs.harvard.edu/abs/2014ApJ...786...53F)).
#
# First application of magnetic Dartmouth stellar evolution models to young stars. Implemented the condition that the peak magnetic field strength occurs at $0.50 R_{\odot}$ for fully convective stars:
#
# > Malo, Doyon, Feiden, _et al._ (2014), ApJ, 792, 37 ([ADS](http://adsabs.harvard.edu/abs/2014ApJ...792...37M))
#
| Daily/20150729_small_magnetic_grid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Lab 07<h1>
# +
class Cab:
def __init__(self,kms,type_of_cab,year):
self.__kms = kms
self.__type_of_cab = type_of_cab
self.__year = year
def get_kms(self):
return self.__kms
def get_type_of_car(self):
return self.__type_of_cab
def get_year(self):
return self.__year
def __gt__(self,other):
return self.__year > other.get_year()
def __eq__(self,other):
return self.__year == other.get_year() and self.__type_of_cab == other.get_type_of_car()
def __repr__(self):
print(f"kms: {self.__kms} type of car {self.__type_of_cab} year : {self.__year}")
class Sedan(Cab):
def __init__(self,kms,type_of_cab,year):
Cab.__init__(self,kms,type_of_cab,year)
self.__price_per_km = 2.5
def calculate_fare(self):
return Cab.get_kms(self) * self.__price_per_km
class Hatchback(Cab):
def __init__(self,kms,type_of_cab,year):
Cab.__init__(self,kms,type_of_cab,year)
self.__price_per_km = 2.2
def calculate_fare(self):
return Cab.get_kms(self) * self.__price_per_km
s1 = Sedan(10000,'Sedan',2004)
s1.calculate_fare()
s1.get_kms()
# -
# ### Script
# +
def read_file(f1):
file_one = open(f1,'r')
l = []
for line in file_one:
l_split = line.split(';')
if l_split[0] == "Sedan":
l.append(Sedan(int(l_split[1]),l_split[0],int(l_split[2])))
elif l_split[0] == "Hatchback":
l.append(Hatchback(int(l_split[1]),l_split[0],int(l_split[2])))
return l
def find_greater(l,cab,year):
counter_year = 0
temp_km = 0
for i in l:
if cab == 'Sedan':
if i.get_type_of_car() == cab and i > s1:
counter_year += 1
elif cab == "Hatchback":
if i == h1 :
temp_km += i.get_kms()
if cab == 'Sedan':
return f'There are {counter_year} {cab} cars newer than 2015.'
else:
return f'All {cab} cars of {year} have travelled {temp_km} kms.'
list_cars = read_file("cabs.txt")
counter = 1
cab = Hatchback(10000,'Hatchback',2020)
s1 = Sedan(10000,'Sedan',2015)
for i in range(len(list_cars)):
if list_cars[i].get_type_of_car() == "Sedan":
print("Sedan ",counter, "will pay",list_cars[i].calculate_fare())
counter += 1
print("")
print(find_greater(list_cars,"Sedan",2015))
print("")
print(find_greater(list_cars,"Hatchback",2020))
# -
| LAB07_BALCI_MustafaCankan/Lab07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats as sts
# %matplotlib inline
# # Нормальное распределение
# Вот так можно сгенерировать выборку из нормально распределённой случайной величины с параметрами $\mu=2.0$ и $\sigma=0.5$:
# +
mu = 2.0
sigma = 0.5
# зададим нормально распределенную случайную величину
norm_rv = sts.norm(loc=mu, scale=sigma)
# сгенерируем 10 значений
norm_rv.rvs(size=10)
# -
# Параметр ```loc``` задаёт $\mu$, ```scale``` — среднеквадратичное отклонение $\sigma$, ```size``` — размер выборки. РРјСЏ параметра ```size``` РїСЂРё вызове функции ```rvs``` можно РЅРµ писать.
#
# Следующая функция возвращает значение функции распределения нормальной случайной величины в точке, соответствующей её аргументу:
norm_rv.cdf(2)
# Построим график функции распределения:
x = np.linspace(0,4,100)
cdf = norm_rv.cdf(x) # функция может принимать и вектор (x)
plt.plot(x, cdf)
plt.ylabel('$F(x)$')
plt.xlabel('$x$')
# А так можно вычислить значение функции плотности вероятности нормального распределения в заданной точке:
norm_rv.pdf(3)
# Построим график функции плотности вероятности:
# +
x = np.linspace(0,4,100)
pdf = norm_rv.pdf(x)
plt.plot(x, pdf)
plt.ylabel('$f(x)$')
plt.xlabel('$x$')
# -
# # Равномерное распределение на отрезке
# Вот так можно сгенерировать выборку из случайной величины, имеющей равномерное распределение на отрезке $[a,b]$:
# +
a = 1
b = 4
# обратите внимание, что в этой функции задается левая граница и масштаб, а не левая и правая границы:
uniform_rv = sts.uniform(a, b-a)
uniform_rv.rvs(10)
# -
# А так — вычислять значения функций распределения и плотностей:
# +
x = np.linspace(0,5,100)
cdf = uniform_rv.cdf(x)
plt.plot(x, cdf)
plt.ylabel('$F(x)$')
plt.xlabel('$x$')
# +
x = np.linspace(0,5,1000)
pdf = uniform_rv.pdf(x)
plt.plot(x, pdf)
plt.ylabel('$f(x)$')
plt.xlabel('$x$')
# -
# # Распределение Бернулли
# Генерация выборок из распределения Бернулли с заданным параметром $p$:
# +
bernoulli_rv = sts.bernoulli(0.7)
bernoulli_rv.rvs(10)
# -
# # Биномиальное распределение
# Генерация выборок из биномиального распределения:
binomial_rv = sts.binom(20, 0.7)
binomial_rv.rvs(10)
# Первый аргумент функции binom — значение параметра $n$, второй — параметра $p$.
#
# Функция распределения:
# +
x = np.linspace(0,20,21)
cdf = binomial_rv.cdf(x)
plt.step(x, cdf)
plt.ylabel('$F(x)$')
plt.xlabel('$x$')
# -
# Функция вероятности ```pmf``` для дискретных случайных величин заменяет функцию плотности ```pdf```:
# +
x = np.linspace(0,20,21)
pmf = binomial_rv.pmf(x)
plt.plot(x, pmf, 'o')
plt.ylabel('$P(X=x)$')
plt.xlabel('$x$')
# -
# Посмотрим, как ведут себя биномиально распределенные величины при разных значениях параметров:
# +
x = np.linspace(0,45,46)
for N in [20, 30]:
for p in [0.2, 0.7]:
rv = sts.binom(N, p)
cdf = rv.cdf(x)
plt.step(x, cdf, label="$N=%s, p=%s$" % (N,p))
plt.legend()
plt.title("CDF (binomial)")
plt.ylabel('$F(X)$')
plt.xlabel('$x$')
# +
x = np.linspace(0,45,46)
symbols = iter(['o', 's', '^', '+'])
for N in [20, 30]:
for p in [0.2, 0.8]:
rv = sts.binom(N, p)
pmf = rv.pmf(x)
plt.plot(x, pmf, next(symbols), label="$N=%s, p=%s$" % (N,p))
plt.legend()
plt.title("PMF (binomial)")
plt.ylabel('$P(X=x)$')
plt.xlabel('$x$')
# -
# # Распределение Пуассона
# Генерация выборок из распределения Пуассона с параметром $\lambda$:
poisson_rv = sts.poisson(5)
poisson_rv.rvs(10)
# +
x = np.linspace(0,30,31)
for l in [1, 5, 10, 15]:
rv = sts.poisson(l)
cdf = rv.cdf(x)
plt.step(x, cdf, label="$\lambda=%s$" % l)
plt.legend()
plt.title("CDF (poisson)")
plt.ylabel('$F(x)$')
plt.xlabel('$x$')
# +
x = np.linspace(0,30,31)
symbols = iter(['o', 's', '^', '+'])
for l in [1, 5, 10, 15]:
rv = sts.poisson(l)
pmf = rv.pmf(x)
plt.plot(x, pmf, next(symbols), label="$\lambda=%s$" % l)
plt.legend()
plt.title("PMF (poisson)")
plt.ylabel('$P(X=x)$')
plt.xlabel('$x$')
# -
# # Дискретное распределение общего вида
# Чтобы сгенерировать дискретную случайную величину общего вида, нужно задать множество её значений и соответствующих вероятностей и использовать функцию ```numpy.random.choice```:
elements = np.array([1, 5, 12])
probabilities = [0.05, 0.7, 0.25]
np.random.choice(elements, 10, p=probabilities)
# # Другие распределения
# Существует большое количество других стандартных семейств распределений, многие из которых также можно генерировать в Питоне.
# Например, распределение хи-квадрат $\chi^2_k$, имеющее натуральный параметр $k$, который называется числом степеней свободы:
x = np.linspace(0,30,100)
for k in [1, 2, 3, 4, 6, 9]:
rv = sts.chi2(k)
cdf = rv.cdf(x)
plt.plot(x, cdf, label="$k=%s$" % k)
plt.legend()
plt.title("CDF ($\chi^2_k$)")
x = np.linspace(0,30,100)
for k in [1, 2, 3, 4, 6, 9]:
rv = sts.chi2(k)
pdf = rv.pdf(x)
plt.plot(x, pdf, label="$k=%s$" % k)
plt.legend()
plt.title("PDF ($\chi^2_k$)")
# Полный список функций SciPy для работы со всеми распределениями можно найти тут: http://docs.scipy.org/doc/scipy-0.14.0/reference/stats.html
| Course_1/St1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Loading Cobra package
import cobra
# +
# Loading Functions
def input_output_reacs(model):
"Returns all input and output reactions for the given model"
reacs = []
nums = []
counter = 0
for r in model.reactions:
coeffs = []
for i in r.metabolites:
c = r.get_coefficient(i.id)
coeffs.append(c)
if all(i > 0.0 for i in coeffs) or all(i < 0.0 for i in coeffs) :
reacs.append(r)
nums.append(counter)
counter = counter + 1
return(reacs)
def deg_free(model):
"Prints degrees of freedom of the model"
no_r = 0
for r in model.reactions:
if r.lower_bound == 0.0:
no_r = no_r +1
else:
no_r = no_r +2
no_m = len(model.metabolites)
no_IO = len(input_output_reacs(mod))
print("Inner degrees of freedom: {}".format(no_r-no_m))
print("Outer degrees of freedom: {}".format(no_r-no_m-no_IO))
# -
# Loading the three different models
Arnold2014 = cobra.io.read_sbml_model("Arnold2014.xml")
Poolman2009 = cobra.io.read_sbml_model("Poolman2009.xml")
Gomes2010 = cobra.io.read_sbml_model("Gomes2010.xml")
# Model Setup
robjs = ['Bio_opt','ATPase','BIO_L']
i = 0
for mod in [Arnold2014,Poolman2009,Gomes2010]:
print(mod.objective)
print(mod.optimize().objective_value)
robj = mod.reactions.get_by_id(robjs[i])
print(robj.bounds)
deg_free(mod)
i = i+1
| ComparingAlgorithms/ExploringModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/probml/pyprobml/blob/master/book1/mlp/lrschedule_tf.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DK0_Km7Zj-mN"
# # Illustrate various learning rate schedules
# Based on
# https://github.com/ageron/handson-ml2/blob/master/11_training_deep_neural_networks.ipynb
#
# + id="i_w02PIIkNSd"
import numpy as np
import matplotlib.pyplot as plt
import os
try:
import tensorflow
except ModuleNotFoundError:
# %pip install tensorflow
import tensorflow
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
K = tf.keras.backend
# + id="Qs0gPskvk3gF"
def save_fig(fname, *args, **kwargs):
"""Save current plot window to the figures directory."""
if "PYPROBML" in os.environ:
root = os.environ["PYPROBML"]
figdir = os.path.join(root, "figures")
else:
figdir = "../figures"
print("cannot find environment variable PYPROBML, writing to {}".format(figdir))
if not os.path.exists(figdir):
os.mkdir(figdir)
fname_full = os.path.join(figdir, fname)
print("saving image to {}".format(fname_full))
plt.tight_layout()
plt.savefig(fname_full, *args, **kwargs)
# + id="kvrxOigmk_QT"
(X_train_full, y_train_full), (X_test, y_test) = tf.keras.datasets.fashion_mnist.load_data()
X_train_full = X_train_full / 255.0
X_test = X_test / 255.0
# There are 60k training examples. For speed, we use 10k for training
# and 10k for validation.
n_train = 1000
n_valid = 1000
X_valid, X_train = X_train_full[:n_valid], X_train_full[n_valid : n_valid + n_train]
y_valid, y_train = y_train_full[:n_valid], y_train_full[n_valid : n_valid + n_train]
pixel_means = X_train.mean(axis=0, keepdims=True)
pixel_stds = X_train.std(axis=0, keepdims=True)
X_train_scaled = (X_train - pixel_means) / pixel_stds
X_valid_scaled = (X_valid - pixel_means) / pixel_stds
X_test_scaled = (X_test - pixel_means) / pixel_stds
# + id="8TCzm0qjkBhC"
n_epochs = 20
lr0 = 0.01
batch_size = 32
n_steps_per_epoch = len(X_train) // batch_size
epochs = np.arange(n_epochs)
def make_model(lr0=0.01, momentum=0.9):
optimizer = tf.keras.optimizers.SGD(lr=lr0, momentum=momentum)
model = tf.keras.models.Sequential(
[
tf.keras.layers.Flatten(input_shape=[28, 28]),
tf.keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
tf.keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
tf.keras.layers.Dense(10, activation="softmax"),
]
)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
return model
# Power scheduling
# lr = lr0 / (1+ steps/s)**c
# Keras.optimizer.SGD uses power scheduling with c=1 and s=1/decay
def power_decay(lr0, s, c=1):
def power_decay_fn(epoch):
return lr0 / (1 + epoch / s) ** c
return power_decay_fn
power_schedule = tf.keras.callbacks.LearningRateScheduler(power_decay(lr0=lr0, s=20))
# Exponential scheduling
# lr = lr0 * 0.1**(epoch / s)
def exponential_decay(lr0, s):
def exponential_decay_fn(epoch):
return lr0 * 0.1 ** (epoch / s)
return exponential_decay_fn
exponential_schedule = tf.keras.callbacks.LearningRateScheduler(exponential_decay(lr0=lr0, s=20))
# Piecewise constant
def piecewise_constant(boundaries, values):
boundaries = np.array([0] + boundaries)
values = np.array(values)
def piecewise_constant_fn(epoch):
return values[np.argmax(boundaries > epoch) - 1]
return piecewise_constant_fn
piecewise_schedule = tf.keras.callbacks.LearningRateScheduler(piecewise_constant([5, 15], [0.01, 0.005, 0.001]))
# Performance scheduling
perf_schedule = tf.keras.callbacks.ReduceLROnPlateau(factor=0.5, patience=5)
# Make plots
schedules = {
"power": power_schedule,
"exp": exponential_schedule,
"piecewise": piecewise_schedule,
"perf": perf_schedule,
}
def ema(y, beta):
"""Exponentially weighted average."""
n = len(y)
zs = np.zeros(n)
z = 0
for i in range(n):
z = beta * z + (1 - beta) * y[i]
zs[i] = z
return zs
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="y81v3tfvkX8e" outputId="a26ad25f-531b-419d-cfa9-6501c31200eb"
for name, lr_scheduler in schedules.items():
tf.random.set_seed(42)
np.random.seed(42)
model = make_model()
history = model.fit(
X_train_scaled, y_train, epochs=n_epochs, validation_data=(X_valid_scaled, y_valid), callbacks=[lr_scheduler]
)
plt.figure()
plt.plot(history.epoch, history.history["lr"], "o-")
# plt.axis([0, n_epochs - 1, 0, 0.011])
plt.xlabel("Epoch")
plt.ylabel("Learning rate")
plt.title(name, fontsize=14)
plt.grid(True)
if name == "perf":
ax2 = plt.gca().twinx()
hist = history.history["val_loss"]
ax2.plot(history.epoch, hist, "r^-")
ax2.set_ylabel("Validation Loss", color="r")
ax2.tick_params("y", colors="r")
# plt.figure()
# plt.plot(history.epoch, ema(hist, 0.95))
fname = "lrschedule-{}.pdf".format(name)
save_fig(fname)
plt.show()
# + id="lBvcUdDwlHS4"
# + [markdown] id="DbnxOURzmR1o"
# # One-cycle heuristic
#
# Illustrate the learning rate finder and 1cycle heuristic from <NAME>
# It is described in this WACV'17 paper (https://arxiv.org/abs/1506.01186)
# and this blog post:
# https://sgugger.github.io/how-do-you-find-a-good-learning-rate.html
#
# The code below is modified from
# [here](https://github.com/ageron/handson-ml2/blob/master/11_training_deep_neural_networks.ipynb)
#
#
# It trains an MLP on FashionMNIST
#
# + colab={"base_uri": "https://localhost:8080/", "height": 492} id="Z1QAJN7SmgvH" outputId="a1e6fea7-6a82-43e9-fe8b-d8be7457d25a"
class OneCycleScheduler(keras.callbacks.Callback):
def __init__(self, iterations, max_rate, start_rate=None, last_iterations=None, last_rate=None):
self.iterations = iterations
self.max_rate = max_rate
self.start_rate = start_rate or max_rate / 10
self.last_iterations = last_iterations or iterations // 10 + 1
# self.half_iteration = (iterations - self.last_iterations) // 2
self.half_iteration = self.last_iterations // 2
self.last_rate = last_rate or self.start_rate / 1000
self.iteration = 0
self.rate_hist = []
def _interpolate_broken(self, iter1, iter2, rate1, rate2):
return (rate2 - rate1) * (iter2 - self.iteration) / (iter2 - iter1) + rate1
def _interpolate(self, iter1, iter2, rate1, rate2):
return (rate2 - rate1) * (self.iteration - iter1) / (iter2 - iter1) + rate1
def on_batch_begin(self, batch, logs):
if self.iteration < self.half_iteration:
rate = self._interpolate(0, self.half_iteration, self.start_rate, self.max_rate)
elif self.iteration < 2 * self.half_iteration:
rate = self._interpolate(self.half_iteration, 2 * self.half_iteration, self.max_rate, self.start_rate)
else:
rate = self._interpolate(2 * self.half_iteration, self.iterations, self.start_rate, self.last_rate)
rate = max(rate, self.last_rate)
self.iteration += 1
K.set_value(self.model.optimizer.lr, rate)
self.rate_hist.append(K.get_value(self.model.optimizer.lr))
# https://stackoverflow.com/questions/48198031/keras-add-variables-to-progress-bar/48206009#48206009
n_epochs = 5
n_steps_per_epoch = len(X_train) // batch_size
onecycle = OneCycleScheduler(n_steps_per_epoch * n_epochs, max_rate=0.05)
history = model.fit(
X_train_scaled,
y_train,
epochs=n_epochs,
batch_size=batch_size,
validation_data=(X_valid_scaled, y_valid),
callbacks=[onecycle],
)
# lr_hist = history.history["lr"] # only stored by LRScheduler
lr_hist = onecycle.rate_hist
plt.figure()
plt.plot(lr_hist, "o-")
plt.xlabel("Step")
plt.ylabel("Learning rate")
plt.title("onecycle", fontsize=14)
plt.grid(True)
save_fig("lrschedule-onecycle.pdf")
plt.show()
# + [markdown] id="ym5_voi7m4qr"
# # Loss vs learning rate
# + colab={"base_uri": "https://localhost:8080/"} id="lv7D9QhAm6C4" outputId="3d8afee9-863e-46d5-aa1f-025794d00d88"
class ExponentialLearningRate(keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.rates = []
self.losses = []
def on_batch_end(self, batch, logs):
self.rates.append(K.get_value(self.model.optimizer.lr))
self.losses.append(logs["loss"])
K.set_value(self.model.optimizer.lr, self.model.optimizer.lr * self.factor)
def find_learning_rate(model, X, y, epochs=1, batch_size=32, min_rate=10**-5, max_rate=10):
init_weights = model.get_weights()
iterations = len(X) // batch_size * epochs
factor = np.exp(np.log(max_rate / min_rate) / iterations)
init_lr = K.get_value(model.optimizer.lr)
K.set_value(model.optimizer.lr, min_rate)
exp_lr = ExponentialLearningRate(factor)
history = model.fit(X, y, epochs=epochs, batch_size=batch_size, callbacks=[exp_lr])
K.set_value(model.optimizer.lr, init_lr)
model.set_weights(init_weights)
return exp_lr.rates, exp_lr.losses
def plot_lr_vs_loss(rates, losses):
plt.plot(rates, losses)
plt.gca().set_xscale("log")
plt.hlines(min(losses), min(rates), max(rates))
plt.axis([min(rates), max(rates), min(losses), (losses[0] + min(losses)) / 2])
plt.xlabel("Learning rate")
plt.ylabel("Loss")
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential(
[
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="elu"),
keras.layers.Dense(100, activation="elu"),
keras.layers.Dense(10, activation="softmax"),
]
)
"""
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
"""
model.compile(loss="sparse_categorical_crossentropy", optimizer=keras.optimizers.SGD(lr=1e-3), metrics=["accuracy"])
batch_size = 128
rates, losses = find_learning_rate(model, X_train_scaled, y_train, epochs=1, batch_size=batch_size)
# + colab={"base_uri": "https://localhost:8080/"} id="_YwWO-RFnv6e" outputId="deb6c516-a932-4962-835b-dc67b2edf91b"
print(rates)
print(losses)
# + id="GmfVEI83nRiQ"
plt.figure()
plot_lr_vs_loss(rates, losses)
save_fig("lrfinder-raw.pdf")
plt.show()
# https://sites.google.com/site/hardwaremonkey/blog/ewmafilterexmpleusingpandasandpython
# x = np.linspace(0, 2 * np.pi, 100)
# y = 2 * np.sin(x) + 0.1 * np.random.normal(x)
x = rates
y = np.array(losses, dtype=np.float64)
df = pd.Series(y)
filtered = pd.Series.ewm(df, span=10).mean()
plt.figure() # figsize=(10,6))
plt.plot(x, y)
# plt.plot(x, filtered)
plt.gca().set_xscale("log")
plt.xlabel("Learning rate")
plt.ylabel("Loss")
save_fig("lrfinder-unfiltered.pdf")
plt.show()
plt.figure() # figsize=(10,6))
# plt.plot(x, y)
plt.plot(x, filtered)
plt.gca().set_xscale("log")
plt.xlabel("Learning rate")
plt.ylabel("Loss")
save_fig("lrfinder-filtered.pdf")
plt.show()
plt.figure() # figsize=(10,6))
plt.plot(x, y)
plt.plot(x, filtered)
plt.gca().set_xscale("log")
plt.xlabel("Learning rate")
plt.ylabel("Loss")
save_fig("lrfinder-filtered-both.pdf")
plt.show()
| notebooks/book1/08/lrschedule_tf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import math
from PIL import Image
# -
train_folder = '/home/nicholas/Workspace/Resources/Camera/train'
# +
sample_image = '/home/nicholas/Workspace/Resources/Camera/train/Motorola-X/(MotoX)2.jpg'
im = Image.open(sample_image)
# -
plt.imshow(np.array(im))
# +
grid_size = 512
print(im.size)
crops = []
for col in range(im.size[0] // grid_size):
for row in range(im.size[1] // grid_size):
x1 = col * grid_size
y1 = row * grid_size
x2 = x1 + grid_size
y2 = y1 + grid_size
crops.append(im.crop((x1, y1, x2, y2)))
fig = plt.figure()
rows = math.ceil(len(crops) / 6)
for idx, i in enumerate(crops):
ax = fig.add_subplot(rows, 6, idx + 1)
ax.imshow(i)
| Visualize Images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: brainio_contrib
# language: python
# name: brainio_contrib
# ---
from pathlib import Path
import brainio_contrib
from mkgu_packaging.movshon import aperture_correct
from brainio_collection import list_stimulus_sets, list_assemblies, get_stimulus_set, get_assembly
from brainio_collection.knownfile import KnownFile as kf
from tqdm import tqdm
import xarray as xr
list_stimulus_sets()
mov_pub = get_stimulus_set("movshon.FreemanZiemba2013-public")
mov_pub
mov_pri = get_stimulus_set("movshon.FreemanZiemba2013-private")
mov_pri
# ls /braintree/home/jjpr/.brainio/image_movshon_FreemanZiemba2013-public/
mov_pub_ap = aperture_correct.main()
# ls /braintree/home/jjpr/.brainio/
mov_pub_ap
converted_image_ids = {}
for image_id in tqdm(mov_pub_ap['image_id'], total=len(mov_pub_ap), desc='apply cosine aperture'):
converted_image_id = kf(mov_pub_ap.get_image(image_id)).sha1
converted_image_ids[image_id] = converted_image_id
converted_image_ids
mov_pub_ap["image_id_old"] = mov_pub_ap["image_id"]
mov_pub_ap["image_id"] = mov_pub_ap["image_id"].map(converted_image_ids)
mov_pub_ap
source_path = Path('/Users/jjpr/.brainio/image_movshon_stimuli/movshon_stimuli/noise-320x320-im38-smp5.png')
target_path = Path('/Users/jjpr/.brainio/image_movshon_stimuli/movshon_stimuli_aperture/noise-320x320-im38-smp5.png')
# ls {source_path.parent}
# ls {target_path.parent}
list_assemblies()
a_mov_pub = get_assembly("movshon.FreemanZiemba2013.public")
a_mov_pri = get_assembly("movshon.FreemanZiemba2013.private")
a_mov_pub
import pandas as pd
a_mov_pub["image_id_converted"] = ("presentation", pd.Series(a_mov_pub["image_id"]).map(converted_image_ids))
a_mov_pub
list(a_mov_pub.indexes.keys())
a_mov_pub.reset_index("presentation", inplace=True)
a_mov_pub
[k for k, v in a_mov_pub.coords.variables.items() if v.dims == ("presentation",)]
mov_pub_ap.columns
[k for k, v in a_mov_pub.coords.variables.items() if v.dims == ("presentation",) and k not in mov_pub_ap.columns]
image_level = [k for k, v in a_mov_pub.coords.variables.items() if v.dims == ("presentation",) and k in mov_pub_ap.columns and k != "image_id"]
image_level
[k for k in a_mov_pub.attrs]
d_mov_pub = xr.DataArray(a_mov_pub)
| packaging/notebooks/2020-02-21_movshon_aperture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="scU9JVUodThH"
# # Cube Imaging
# This notebook will demonstrate how to create a cube dirty image with natural weighting using ngCASA. The resulting image will be compared with an image created by CASA. The Dask visualize tool will also be introduced.
#
# For this demonstration data from the ALMA First Look at Imaging CASAguide (https://casaguides.nrao.edu/index.php/First_Look_at_Imaging) will be used. The measurement set has been converted to vis.zarr (using convert_ms in cngi.conversion).
#
# + [markdown] colab_type="text" id="znAYKj-Ym-xK"
# ## Installation and Dataset Download
#
# + colab_type="code" id="rFkS-UIfm5xA" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a68f206c-447e-485e-b8eb-43b6d1b2ba1e"
import os
os.system("pip install --extra-index-url https://test.pypi.org/simple/ cngi-prototype==0.0.58rc2")
#https://drive.google.com/file/d/1QAEHs2OwP5h37WZId23nmzXSeQ4qt_LG/view?usp=sharing
#https://drive.google.com/file/d/1UeNywNIU-AEwIloWNCpUHZbxuV6RE3lz/view?usp=sharing
for id in ['<KEY>_LG', '1UeNywNIU-AEwIloWNCpUHZbxuV6RE3lz']:
os.system('curl -c ./cookie -s -L "https://drive.google.com/uc?export=download&id=%s"' % id)
os.system('curl -Lb ./cookie "https://drive.google.com/uc?export=download&confirm=`awk \'/download/ {print $NF}\' ./cookie`&id=%s" -o ms.tar.gz' % id)
os.system('tar -xzf ms.tar.gz')
print('complete')
# + [markdown] colab_type="text" id="m5S4JYLxAi87"
# ## Load Dataset
#
# Two datasets are are needed for this notebook:
# - sis14_twhya_chan_avg_field_5_lsrk_pol_xx.vis.zarr
# - casa_twhya_standard_gridder_lsrk_cube_natural.img.zarr
#
# (for more information about the img.zarr format go [here](https://cngi-prototype.readthedocs.io/en/latest/visibilities.html) and for the vis.zarr format go [here](https://cngi-prototype.readthedocs.io/en/latest/images.html)).
#
# The sis14_twhya_chan_avg_field_5_lsrk_pol_xx.vis.zarr dataset is used to create a cube image. The dataset was created by using the ```mstransform``` command in CASA
#
# ```python
# mstransform('sis14_twhya_calibrated_flagged.ms'
# outputvis='sis14_twhya_chan_avg_field_5_lsrk_pol_xx.ms',
# regridms=True, outframe='LSRK', datacolumn='data',
# correlation='XX', field='5', nchan=7)
# ```
#
# and then convert_ms in cngi.conversion
#
# ```python
# infile = 'sis14_twhya_chan_avg_field_5_lsrk_pol_xx.ms'
# outfile = 'sis14_twhya_chan_avg_field_5_lsrk_pol_xx.vis.zarr'
# chunk_shape=(270, 210, 1, 1)
# convert_ms(infile, outfile=outfile, chunk_shape=chunk_shape)
# ```
#
# The conversion to 'LSRK' is necessary because cngi does not currently have an implementation and tclean does a conversion to 'LSRK' before imaging.
#
# To check the ngcasa imaging results the casa_twhya_standard_gridder_lsrk_cube_natural.img.zarr dataset is used. This dataset was generated by running ```tclean``` in CASA
#
# ```python
# tclean(vis='sis14_twhya_chan_avg_field_5_lsrk_pol_xx.ms',
# imagename='twhya_standard_gridder_lsrk_cube_natural',
# specmode='cube',
# deconvolver='hogbom',
# imsize=[200,400],
# cell=['0.08arcsec'],
# weighting='natural',
# threshold='0mJy',
# niter=0,stokes='XX')
# ```
#
# and then ```image_ms``` in cngi.conversion
#
# ```python
# infile = 'cube_image/twhya_standard_gridder_lsrk_cube_natural.image'
# outfile = 'casa_twhya_standard_gridder_lsrk_cube_natural.img.zarr'
# convert_image(infile=infile,outfile=outfile)
# ```
# + colab_type="code" id="F1hf_AtTAi88" colab={"base_uri": "https://localhost:8080/", "height": 747} outputId="b985db8c-8600-4bf8-fa70-8ce515b76757"
import xarray as xr
from cngi.dio import read_vis, read_image
xr.set_options(display_style="html")
vis_dataset = read_vis("sis14_twhya_chan_avg_field_5_lsrk_pol_xx.vis.zarr", 0)
vis_dataset
# + id="777UmJd4fpLD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 411} outputId="b8395cad-3126-4e28-ff87-84914257cabc"
casa_image_dataset = read_image("casa_twhya_standard_gridder_lsrk_cube_natural.img.zarr")
casa_image_dataset
# + [markdown] id="J1KUtSxXPigJ" colab_type="text"
# Note that the chunks parameter in cngi and ngcasa functions specifies the size of a chunk and not the number of chunks (in CASA ```tclean``` chanchunks refers to the number of channel chunks).
#
# The dimensionality of the sis14_twhya_chan_avg_field_5_lsrk_pol_xx.vis.zarr dataset is (time:270,baseline:210,chan:7,pol:1) and a zarr chunk size of (time:270,baseline:210,chan:1,pol:1) was chosen. The dask chunk size was chosen to be the same as the zarr chunk size. For more information concerning chunking go to [here](https://cngi-prototype.readthedocs.io/en/latest/design.html).
# + [markdown] colab_type="text" id="8-2NdsqRe_ry"
# ## Flag Data and Create Imaging Weights
#
# The ```applyflags``` cngi.vis function sets all values that should be flagged to nan. The ngcasa.imaging code does not internally apply flags but does ignore nan values. [applyflags documentation](https://cngi-prototype.readthedocs.io/en/latest/_api/api/cngi.vis.applyflags.html#cngi.vis.applyflags)
#
# The ```make_imaging_weight``` cngi.imaging function takes the WEIGHT or WEIGHT_SPECTRUM data variables and creates IMAGING_WEIGHT data variable that has dimensions time x baseline x chan x pol (matches the visibility DATA variable). Weighting schemes that are supported include natural, uniform, briggs, briggs_abs. Using imaging_weights_parms['chan_mode'] = 'cube' is equavalent to perchanweightdensity=True in CASA. [make_imaging_weight documentation](https://cngi-prototype.readthedocs.io/en/latest/_api/api/imaging.make_imaging_weight.html#ngcasa.imaging.make_imaging_weight)
#
# When ```storage_parms['to_disk']``` is False no execution will occur only a graph will be generated.
# + colab_type="code" id="NEzRyI0De_rz" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="bc5ef732-4baa-452c-b0e5-3c8e1c15785f"
from cngi.vis import applyflags
from ngcasa.imaging import make_imaging_weight
vis_dataset_flagged = applyflags(vis_dataset, flags=['FLAG', 'FLAG_ROW'])
imaging_weights_parms = {}
imaging_weights_parms['weighting'] = 'natural'
imaging_weights_parms['chan_mode'] = 'cube'
imaging_weights_parms['imsize'] = [200,400]
imaging_weights_parms['cell'] = [0.08, 0.08]
storage_parms = {}
storage_parms['to_disk'] = False
vis_dataset_flagged = make_imaging_weight(vis_dataset_flagged, imaging_weights_parms, storage_parms)
# + [markdown] colab_type="text" id="gSnob3jPe_r2"
# ## Create Dirty Cube Image
# The ```make_image``` cngi.imaging function grids the data (using the prolate spheroidal function as an anti-aliasing filter), fast Fourier transform the gridded data to an image and normalizes the image. The ```make_pb``` function currently supports rotationally symmetric airy disk primary beams.
#
# [make_pb documentation](https://cngi-prototype.readthedocs.io/en/latest/_api/api/imaging.make_pb.html#ngcasa.imaging.make_pb)
#
# [make_image documentation](https://cngi-prototype.readthedocs.io/en/latest/_api/api/imaging.make_image.html)
#
# To create an image of the execution graph the [dask.visualize](https://docs.dask.org/en/latest/api.html#dask.visualize) method can be used. By keeping ```storage_parms['to_disk'] ``` False the image_dataset returned by ```make_image``` will contain a graph for flagging, ```applyflags```, ```make_imaging_weight```, ```make_image``` and ```make_pb```.
#
# Changing the ```storage_parms['to_disk']``` to True will trigger a compute.
# + colab_type="code" id="EQ6Ziz1he_r3" colab={"base_uri": "https://localhost:8080/", "height": 620} outputId="9679ab5a-9bb8-4685-db20-234f4b436c74"
from ngcasa.imaging import make_image
from ngcasa.imaging import make_pb
from cngi.dio import write_zarr
import dask
grid_parms = {}
grid_parms['chan_mode'] = 'cube'
grid_parms['imsize'] = [200,400]
grid_parms['cell'] = [0.08, 0.08]
storage_parms = {}
#Create Dask execution graph
storage_parms['to_disk'] = False
image_dataset = make_image(vis_dataset_flagged,grid_parms,storage_parms)
make_pb_parms = {}
make_pb_parms['function'] = 'airy'
make_pb_parms['cell'] = [0.08,0.08]
make_pb_parms['imsize'] = [200,400]
make_pb_parms['list_dish_diameters'] = [10.7]
make_pb_parms['list_blockage_diameters'] = [0.75]
make_pb_parms['pb_name'] = 'PB'
image_dataset = make_pb(image_dataset,make_pb_parms, storage_parms)
dask.visualize(image_dataset,filename='cube_image_graph.png')
#Trigger compute on graph and store result on disk
image_dataset = write_zarr(image_dataset, outfile='twhya_standard_gridder_lsrk_cube_natural.img.zarr', graph_name='make_imaging_weights, make_image and make_pb')
image_dataset
# + [markdown] id="_1t5cOBIPigT" colab_type="text"
# ## Dask Visualization
#
# The Dask execution graph below shows how the images for each channel are computed in parallel. Each image is written to disk independently and Dask along with Zarr handles the virtual concatenation (the resulting img.zarr is chunked by channel). This allows for processing cubes that are larger than memory.
#
#
# 
# + [markdown] colab_type="text" id="i8AgTPSoe_r5"
# ## Plot and Compare With CASA
# + colab_type="code" id="X05Y4vZLe_r9" colab={"base_uri": "https://localhost:8080/", "height": 464} outputId="772dbf24-8c67-4dce-d146-ce75f00b1ec2"
import matplotlib.pylab as plt
import numpy as np
from ipywidgets import interactive
def comparison_plots(chan):
print('Frequency',image_dataset.chan[chan].values, 'Hz')
dirty_image = image_dataset.DIRTY_IMAGE[:,:,chan,0]
casa_dirty_image = casa_image_dataset['RESIDUAL'].values[:, :, chan, 0]
fig0, ax0 = plt.subplots(1, 2, sharey=True)
im0 = ax0[0].imshow(casa_dirty_image)
im1 = ax0[1].imshow(dirty_image)
ax0[0].title.set_text('CASA Dirty Image')
ax0[1].title.set_text('CNGI Dirty Image')
fig0.colorbar(im0, ax=ax0[0], fraction=0.046, pad=0.04)
fig0.colorbar(im1, ax=ax0[1], fraction=0.046, pad=0.04)
plt.show()
plt.figure()
plt.imshow(casa_dirty_image - dirty_image)
plt.title('Difference Dirty Image')
plt.colorbar()
plt.show()
dirty_image = dirty_image / np.max(np.abs(dirty_image))
casa_dirty_image = casa_dirty_image / np.max(np.abs(casa_dirty_image))
# Calculate max error
max_error_dirty_image = np.max(np.abs(dirty_image - casa_dirty_image)).values
print('Max Error',max_error_dirty_image)
# Calculate root mean square error
rms_error_dirty_image = np.linalg.norm(dirty_image - casa_dirty_image, 'fro')
print('RMS Error',rms_error_dirty_image)
#interactive_plot = interactive(comparison_plots, chan=(0, 6))
#output = interactive_plot.children[-1]
#output.layout.height = '550px'
#interactive_plot
comparison_plots(3)
# + [markdown] id="bHN-Gb_YPigY" colab_type="text"
# The first channel (channel 0) is flagged by both ngCASA and CASA. Why CASA is flagging the last channel and ngCASA is not needs further investigation. Checking sis14_twhya_chan_avg_field_5_lsrk_pol_xx.ms with browsetable in CASA shows that only the first channel is flagged.
#
# The reason for the small difference between ngCASA and CASA, in channels 1 to 5, is due to ngCASA using a different implementation of the Fast Fourier Transform.
# + id="OvcNUvtQPigY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="14684b9c-dbb0-4c04-86ac-6776d248b499"
import matplotlib.pylab as plt
import numpy as np
from ipywidgets import interactive
#### Primary Beam Corrected Images ####
def comparison_plots(chan):
print('Frequency',image_dataset.chan[chan].values, 'Hz')
pb_limit = 0.2
primary_beam = image_dataset.PB[100,:,chan,0,0].where(image_dataset.PB[100,:,chan,0,0] > pb_limit,other=0.0)
dirty_image_pb_cor = image_dataset.DIRTY_IMAGE[:,:,chan,0]/image_dataset.PB[:,:,chan,0,0]
dirty_image_pb_cor = dirty_image_pb_cor.where(image_dataset.PB[:,:,chan,0,0] > pb_limit,other=np.nan)
casa_primary_beam = casa_image_dataset['PB'][100, :, chan, 0] #Primary beam created by CASA
casa_dirty_image_pb_cor = (casa_image_dataset['IMAGE.PBCOR'][:, :, chan, 0]).where(casa_image_dataset['PB'][:, :, chan, 0] > pb_limit,other=np.nan) #Image created by CASA
#Plot Primary Beams
fig0, ax0, = plt.subplots(1, 2, sharey=True,figsize=(10, 5))
im0 = ax0[0].plot(casa_primary_beam)
im1 = ax0[1].plot(primary_beam)
ax0[0].title.set_text('CASA Primary Beam Cross Section')
ax0[1].title.set_text('ngCASA Primary Beam Cross Section')
plt.show()
plt.figure()
plt.plot(casa_primary_beam-primary_beam)
plt.title('Difference Primary Beam')
plt.show()
#Plotting Images
fig0, ax0 = plt.subplots(1, 2, sharey=True,figsize=(10, 5))
im0 = ax0[0].imshow(casa_dirty_image_pb_cor)
im1 = ax0[1].imshow(dirty_image_pb_cor)
ax0[0].title.set_text('CASA PB Corrected Dirty Image')
ax0[1].title.set_text('ngCASA PB Corrected Dirty Image')
fig0.colorbar(im0, ax=ax0[0], fraction=0.046, pad=0.04)
fig0.colorbar(im1, ax=ax0[1], fraction=0.046, pad=0.04)
plt.show()
plt.figure()
plt.imshow(casa_dirty_image_pb_cor - dirty_image_pb_cor)
plt.title('Difference Dirty Image')
plt.colorbar()
plt.show()
dirty_image_pb_cor = dirty_image_pb_cor / np.nanmax(np.abs(dirty_image_pb_cor))
casa_dirty_image_pb_cor = casa_dirty_image_pb_cor / np.nanmax(np.abs(casa_dirty_image_pb_cor))
norm_diff_image_pb_cor = dirty_image_pb_cor - casa_dirty_image_pb_cor
# Calculate max error
max_error_dirty_image = np.nanmax(np.abs(norm_diff_image_pb_cor))
print('Max Normalized Error',max_error_dirty_image)
# Calculate root mean square error
rms_error_dirty_image = np.sqrt(np.nansum(np.square(norm_diff_image_pb_cor)))
print('RMS Normalized Error',rms_error_dirty_image)
#interactive_plot = interactive(comparison_plots, chan=(0, 6))
#output = interactive_plot.children[-1]
#output.layout.height = '1100px'
#interactive_plot
comparison_plots(3)
# + [markdown] id="V3lbww_SPigc" colab_type="text"
# The frequency does not change enough for the primary beam to vary significantly.
#
# The difference in primary beam is due to CASA using a sampled 1D function while ngCASA calculates the PB for each pixel. If it is found that PB creation becomes a bottleneck for ngCASA the implementation will be changed to match CASA.
| docs/prototypes/cube_imaging_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Applying algorithm to full dataset
# +
from gensim.models.doc2vec import Doc2Vec, TaggedDocument
from gensim.models.deprecated.doc2vec import LabeledSentence
from gensim.models.word2vec import Word2Vec
from gensim.models.phrases import Phraser, Phrases
from gensim.parsing.porter import PorterStemmer
from gensim.parsing.preprocessing import STOPWORDS
from gensim.parsing.preprocessing import remove_stopwords
from string import digits
import pandas as pd
import numpy as np
import string
import re
import random
import os
import csv
import pickle
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
import nltk
nltk.download('stopwords')
from sklearn import metrics
from sklearn import svm
from sklearn.naive_bayes import GaussianNB
from sklearn.ensemble import GradientBoostingClassifier, RandomForestClassifier
from sklearn.calibration import CalibratedClassifierCV
from sklearn.metrics import confusion_matrix, precision_recall_curve, plot_precision_recall_curve, auc, average_precision_score,classification_report, confusion_matrix, accuracy_score, average_precision_score, precision_score, f1_score, recall_score
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.utils.multiclass import unique_labels
from sklearn.model_selection import cross_val_score, cross_validate, RepeatedStratifiedKFold, train_test_split,KFold, cross_val_score, GridSearchCV
from sklearn.feature_extraction.text import TfidfTransformer, CountVectorizer
from sklearn.linear_model import LogisticRegression, SGDClassifier
from modAL.models import ActiveLearner
from modAL.uncertainty import uncertainty_sampling
from modAL.uncertainty import entropy_sampling
from modAL.density import information_density
from scipy.stats import entropy
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# # Training
# +
#### preprocessing -------------------------------
punctuation_dictionary = {s:None for s in list(string.punctuation)}
punctuation_translator = str.maketrans(punctuation_dictionary)
stop_words = set(stopwords.words('english'))
# (remove punctuation, numbers, lowercase, stop words)
def text_cleaner_all(text, punctuation_translator):
text = text.replace('c("', '')
text = str(text).translate(punctuation_translator)
text = text.lower()
remove_digits = str.maketrans('', '', digits)
text = text.translate(remove_digits)
word_tokens = word_tokenize(text)
filtered_text = [w for w in word_tokens if not w.lower() in stop_words]
text = ' '.join(filtered_text)
return(text)
# (remove punctuation, lowercase, stop words)
def text_cleaner_mod(text, punctuation_translator):
text = text.replace('c("', '')
text = str(text).translate(punctuation_translator)
text = text.lower()
word_tokens = word_tokenize(text)
filtered_text = [w for w in word_tokens if not w.lower() in stop_words]
text = ' '.join(filtered_text)
return(text)
# (remove punctuation, lowercase)
def text_cleaner_min(text, punctuation_translator):
text = text.replace('c("', '')
text = str(text).translate(punctuation_translator)
text = text.lower()
return(text)
# +
#data
clas_dat = pd.read_csv("/Users/carlyknight/Dropbox/PROJECTS/Forecasting Downturns/data/coded_sample_final.csv")
clas_dat = clas_dat.drop_duplicates()
clas_dat.shape
# +
#clean
clas_dat["clean_text"] = clas_dat["text"].apply(lambda x: text_cleaner_all(x, punctuation_translator))
# find phrases
phrases1 = Phrases(map(lambda x: x.split(), clas_dat["clean_text"].tolist())) #bigram
phrases2 = Phrases(phrases1[map(lambda x: x.split(), clas_dat["clean_text"].tolist())]) #trigram
clas_dat["phrased_text"] = clas_dat["clean_text"].apply(lambda x: " ".join(phrases2[phrases1[x.split()]]))
# +
# vectorize
vectorizer = CountVectorizer(min_df=5)
tfidfconverter = TfidfTransformer()
X = vectorizer.fit_transform(clas_dat["phrased_text"]).toarray()
X_tf = tfidfconverter.fit_transform(X).toarray()
y = np.array(clas_dat['final_code'])
# +
#training set
X_train, X_test, y_train, y_test = train_test_split(X_tf, y, test_size=0.2)
model = LogisticRegression()
solvers = ['newton-cg', 'lbfgs', 'liblinear']
penalty = ['l2']
c_values = [100, 10, 1.0, 0.1, 0.01]
# define grid search
scoring = ['accuracy', 'precision']
grid = dict(solver=solvers,penalty=penalty,C=c_values)
cv = RepeatedStratifiedKFold(n_splits=10, n_repeats = 5, random_state=1)
grid_search = GridSearchCV(estimator=model, param_grid=grid, n_jobs=-1, cv=cv, scoring= ['accuracy', 'precision'],refit = "accuracy")
grid_result = grid_search.fit(X_train, y_train)
# summarize results
print("Best Accuracy: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
precisions = grid_result.cv_results_['mean_test_precision']
accuracys = grid_result.cv_results_['mean_test_accuracy']
std_prec = grid_result.cv_results_['std_test_precision']
std_acc = grid_result.cv_results_['std_test_accuracy']
params = grid_result.cv_results_['params']
for prec, acc, param in zip(precisions, accuracys, params):
print("Precision: %f (Accuracy: %f) with: %r" % (prec, acc, param))
# +
y_pred = grid_search.best_estimator_.predict(X_test)
print('Accuracy: ', accuracy_score(y_test, y_pred))
print('Precision: ', precision_score(y_test, y_pred))
print('Recall: ', recall_score(y_test, y_pred))
print('F1: ', f1_score(y_test, y_pred))
# -
#save
import joblib
joblib.dump(grid_search.best_estimator_, '/Users/carlyknight/Dropbox/PROJECTS/Forecasting Downturns/data/best_estimator_1200-8-16-21.pkl')
# # Entire dataset
#open all files
fulldat = pd.read_csv("/Users/carlyknight/Dropbox/PROJECTS/Forecasting Downturns/data/all_text.csv")
# +
#create a list of ys
y_pred = []
#iterate over the dataframe 1000 times (chunks of 200)
i=0
for chunk in np.array_split(fulldat, 1000):
print("Working on chunk: ", str(i))
#clean
chunk["clean_text"] = chunk["text"].apply(lambda x: text_cleaner_all(x, punctuation_translator))
#find phrases (this will take a long time)
phrases1 = Phrases(map(lambda x: x.split(), chunk["clean_text"].tolist())) #bigram
phrases2 = Phrases(phrases1[map(lambda x: x.split(), chunk["clean_text"].tolist())]) #trigram
chunk["phrased_text"] = chunk["clean_text"].apply(lambda x: " ".join(phrases2[phrases1[x.split()]]))
#vectorize
X = vectorizer.transform(chunk["phrased_text"]).toarray()
X_tf = tfidfconverter.transform(X).toarray()
#predict
ystar = grid_search.best_estimator_.predict(X_tf)
y_pred.append(ystar)
i+=1
# +
#add column
y_pred_list = [item for items in y_pred for item in items]
fulldat['prediction'] = y_pred_list
#keep id and prediction and output
output = fulldat[["id", "prediction"]]
output.to_csv("/Users/carlyknight/Dropbox/PROJECTS/Forecasting Downturns/data/text_predictions_8-19-21.csv")
| codes/Full data predictions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# Define a function for which we'd like to find the roots
def function_for_roots(x):
a = 1.01
b = -3.04
c= 2.07
return a*x**2 + b*x + c #get the roots of ax^2 +bx + c
def check_initial_values(f,x_min,x_max,tol):
y_min = f(x_min)
y_max = f(x_max)
if(y_min*y_max>=0.0):
print("No zero crossing found in the range =",x_min,x_max)
s = "f(%f) = %f, f(%f) = %f" % (x_min,y_min,x_max,y_max)
print(s)
return 0
# if x_min is a root, then return flag == 1
if(np.fabs(y_min)<tol):
return 1
#if x_max is a root, then return flag == 2
if(np.fabs(y_max)<tol):
return 2
#if we reach this point, the bracket is valid
#and we will return 3
return 3
| Bisection Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1 - Quais casas o CEO da House Rocket deveria comprar e por qual preço de compra?
#
# ## 2 - Uma vez a casa em posse da empresa, qual seria o preço da venda?
#
# ## 3 - A House Rocket deveria fazer uma reforma para aumentar o preço da venda? Quais seriam as sugestões de mudanças? Qual o incremento no preço dado por cada opção de reforma?
import pandas as pd
import numpy as np
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score
# # Passo 1: Importar os dados e criar o modelo
tabela = pd.read_csv('kc_house_data.csv')
modelo = RandomForestRegressor()
tabela2 = tabela
# # Passo 2: Verificar o estado dos dados
tabela.info()
# # Passo 3: Limpeza e Organização
tabela = tabela.drop(['date', 'id'], axis=1)
tabela.floors = tabela.floors.astype(int)
tabela.price = tabela.price.astype(int)
tabela.bathrooms = tabela.bathrooms.astype(int)
tabela.price = tabela.price.round(-3)
display(tabela)
# # Passo 4: Modelagem
X = tabela.drop('price', axis=1)
y = tabela['price']
x_train, x_test, y_train, y_test = train_test_split(X, y, train_size=0.3, random_state=52)
# # Passo 5: Treinamento do algoritmo
modelo.fit(x_train, y_train)
pred = modelo.predict(x_test)
r2_score(y_test, pred)
# # Passo 6: Exportando modelo
import joblib
joblib.dump(modelo, 'model2.pkl')
teste = np.array([[3,1,1180,5650,1,0,0,3,7,1180,0,1955,0,98178,47.5112,-122.257,1340,5650]])
modelo.predict(teste)
| Projeto House Rocket/ProjetoHouseRocket_MachineLearning.ipynb |