code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("pandas/employees.csv")
df.head()
df.info()
df['Start Date'].head()
df['Start Date'] = pd.to_datetime(df['Start Date'])
df['Last Login Time'].head()
df['Last Login Time'] = pd.to_datetime(df['Last Login Time'])
df['Senior Management'].head()
df['Senior Management'] = df['Senior Management'].astype('bool')
df['Gender'].head()
df['Gender'] = df['Gender'].astype('category')
df.info()
49/62.6
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
df.info()
# ## Filter A `DataFrame` Based On A Condition
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
df[df['Gender'] == 'Male']
mask = df['Team'] == 'Finance'
df[mask]
df[(df['Team'] == 'Finance') & (df['Gender'] == 'Male')]
df[df['Senior Management']]
df[df['Team'] != 'Marketing']
df[df['Salary'] > 110000]
df[df['Bonus %'] < 1.5]
mask = df['Start Date'] <= '1985-01-01'
df[mask]
# ## Filter with More than One Condition (AND)
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
# +
mask1 = df["Gender"] == "Male"
mask2 = df["Team"] == "Marketing"
df[mask1 & mask2]
# -
# ## Filter with More than One Condition (OR)
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
# +
mask1 = df["Senior Management"]
mask2 = df["Start Date"] < "1990-01-01"
df[mask1 | mask2]
# +
mask1 = df["First Name"] == "Robert"
mask2 = df["Team"] == "Client Services"
mask3 = df["Start Date"] > "2016-06-01"
df[(mask1 & mask2) | mask3]
# -
# ## The `.isin()` Method
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
# +
mask1 = df["Team"] == "Legal"
mask2 = df["Team"] == "Sales"
mask3 = df["Team"] == "Product"
df[mask1 | mask2 | mask3]
# -
mask = df['Team'].isin(['Legal', 'Sales', 'Product'])
df[mask]
# ## The `.isnull()` and `.notnull()` Methods
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
# +
mask = df["Team"].isnull()
df[mask]
# +
condition = df["Gender"].notnull()
df[condition]
# -
# ## The `.between()` Method
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
df['Salary'].between(60000, 70000)
df[df["Salary"].between(60000, 70000)]
df[df["Bonus %"].between(2.0, 5.0)]
df[df["Start Date"].between("1991-01-01", "1992-01-01")]
df[df["Last Login Time"].between("08:30AM", "12:00PM")]
# ## The `.duplicated()` Method
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.sort_values("First Name", inplace = True)
df.head(3)
df['First Name']
df[df['First Name'].duplicated()]
df[df['First Name'].duplicated(keep='last')]
df[df['First Name'].duplicated(keep=False)]
ad = ~df['First Name'].duplicated(keep=False)
df[ad]
mask = ~df["First Name"].duplicated(keep = False)
af = df[mask]
# ## The `.drop_duplicates()` Method
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.sort_values("First Name", inplace = True)
df.head(3)
len(df)
len(df.drop_duplicates())
sf = df.drop_duplicates(subset=['First Name'], keep=False)
af == sf
df.drop_duplicates()
df.drop_duplicates(subset = ["First Name"], keep = False)
df.drop_duplicates(subset = ["First Name", "Team"], inplace = True)
df.head(2)
len(df)
# ## The `.unique()` and `.nunique()` Methods
df = pd.read_csv("pandas/employees.csv", parse_dates = ["Start Date", "Last Login Time"])
df["Senior Management"] = df["Senior Management"].astype("bool")
df["Gender"] = df["Gender"].astype("category")
df.head(3)
df['Gender']
df['Gender'].unique()
df['Team']
df['Team'].unique()
# +
df["Gender"].unique()
df["Team"].unique()
# -
len(df["Team"].unique())
df['Team'].nunique()
df["Team"].nunique(dropna = False)
| CompletedCourseFiles/DataFrames 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# # World Firearm Statistics
df = pd.read_csv("world_firearms.csv")
df.head(5)
df['Average total all civilian firearms'].sum()
df['Average total all civilian firearms'][df['Country/Territory']=='United States']/df['Average total all civilian firearms'].sum()
# Almost 42% of total civilian firearms in the world are in the United States.
# #### Obtain and merge human develompent index data from 2007 with matching firearm data
hdi = pd.read_csv("HDI.csv")
hdi['Country'] = hdi['Country'].apply(lambda x: x.lstrip(" "))
hdi = hdi[['Country', '2007']]
df1 = hdi.merge(df, right_on="Country/Territory", left_on="Country", how = "right")
df1 = df1.rename(columns = {"2007":"HDI", "Homicide by firearm rate per 100,000 pop":"Homicides_per_100",
"Average firearms per 100 people":"Firearms_per_100"})
df1.head()
df1['HDI_rank'] = df1.HDI.rank(ascending = False, method = 'min')
df1.to_csv('2007_world_guns.csv')
# ## Gun Control Laws
laws = pd.read_csv("gun_laws.csv")
laws2016 = laws[laws.year == 2016]
# Categories of Gun Control Laws to be analyzed:
#
# 3 - Limiting access to High Risk Individuals,
# 4 - Background Checks,
# 7 - Limiting Concealed Carry,
# 8 - Ban/Limit Assault weapons,
# 9 - Child Safety Laws,
def laws(x):
names = ['state','lawtotal']
for i in x:
if i.startswith('3'):
names.append(i)
if i.startswith('4'):
names.append(i)
if i.startswith('7'):
names.append(i)
if i.startswith('8'):
names.append(i)
if i.startswith('9'):
names.append(i)
return names
colnames = laws(laws2016.columns)
laws2016 = laws2016[colnames]
laws2016['high_risk'] = laws2016['3 - violentpartial']+laws2016['3 - violenth']+laws2016['3 - violent']+laws2016[
'3 - invoutpatient']+laws2016['3 - invcommitment']+laws2016['3 - felony']+laws2016['3 - drugmisdemeanor']+laws2016[
'3 - danger']+laws2016['3 - alctreatment']+laws2016['3 - alcoholism']
laws2016['background'] = laws2016['4 - universalpermith']+laws2016['4 - universalpermit']+laws2016['4 - universalh']+laws2016[
'4 - statechecks']+laws2016['4 - statechecksh']+laws2016['4 - universal']+laws2016['4 - threedaylimit']+laws2016[
'4 - mentalhealth']+laws2016['4 - gunshowh']+laws2016['4 - gunshow']+laws2016['4 - backgroundpurge']
laws2016['concealed'] = laws2016['7 - showing']+laws2016['7 - permitconcealed']+laws2016['7 - mayissue']+laws2016[
'7 - ccrevoke']+laws2016['7 - ccrenewbackground']+laws2016['7 - ccbackgroundnics']+laws2016['7 - ccbackground']
laws2016['assault'] = laws2016['8 - tenroundlimit']+laws2016['8 - magazinepreowned']+laws2016['8 - magazine']+laws2016[
'8 - assaulttransfer']+laws2016['8 - assaultregister']+laws2016['8 - assaultlist']+laws2016['8 - assault']
laws2016['child'] = laws2016['9 - cap14']+laws2016['9 - cap16']+laws2016['9 - cap18']+laws2016[
'9 - capaccess']+laws2016['9 - capliability']+laws2016['9 - capunloaded']+laws2016['9 - capuses']+laws2016[
'9 - lockd']+laws2016['9 - locked']+laws2016['9 - lockp']+laws2016['9 - lockstandards']
laws2016.to_csv('2016_gunlaws.csv')
# ### Violent Crime Stats by State
state_violence = pd.read_csv('gun-violence-13-18.csv')
state_violence['Year'] = state_violence.date.apply(lambda x: x[0:4])
state_violence2016 = state_violence[state_violence.Year == '2016']
# Load Population data from 2016 by state and merge to calcluate rates.
pop = pd.read_csv('2016_populations.csv')
pop['State'] = pop['State'].apply(lambda x:x.replace(".",""))
state_violence2016 = state_violence2016.merge(pop, left_on='state', right_on='State', how = 'left')
state_gb = state_violence2016.groupby('state').agg({'incident_id':'count',
'n_killed':'sum', 'n_injured':'sum', 'Population':'max', 'Firearms':'max'}).reset_index()
state_gb['firearms_per_100k'] = (state_gb['Firearms']/state_gb['Population'])*100000
state_gb['incidence_per_100k'] = (state_gb['incident_id']/state_gb['Population'])*100000
state_gb['deaths_per_100k'] = (state_gb['n_killed']/state_gb['Population'])*100000
state_gb['injured_per_100k'] = (state_gb['n_injured']/state_gb['Population'])*100000
state_gb['inj_killed_per_100k'] = ((state_gb['n_injured']+state_gb['n_killed'])/state_gb['Population'])*100000
state_gb.head(10)
state_gb.to_csv('2016_violence_by_state.csv')
| First_Version_Redesign/Data_Wrangling_Individual_Project_Tyler_Young.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:kaggle] *
# language: python
# name: conda-env-kaggle-py
# ---
# +
import albumentations as A
import numpy as np
import os
import random
import sys
import torch
import torch.nn.functional as F
import torchvision
import torchvision.transforms as T
from torch import nn, Tensor, IntTensor, LongTensor, FloatTensor
from functools import partial
from omegaconf import DictConfig, OmegaConf
from tqdm.notebook import tqdm
from typing import Any, Callable, Dict, Iterable, List, Tuple
# -
import cv2
import PIL
import kornia
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
BASE_DIR = '/home/dmitry/projects/dfdc'
SRC_DIR = os.path.join(BASE_DIR, 'src')
VID_DIR = '/media/dmitry/data/dfdc-videos/dfdc_train_part_38'
sys.path.insert(0, f'/home/{os.environ["USER"]}/projects/dfdc/vendors/Pytorch_Retinaface')
from data import cfg_mnet, cfg_re50
from layers.functions.prior_box import PriorBox
from models.retinaface import RetinaFace
from utils.nms.py_cpu_nms import py_cpu_nms
sys.path.insert(0, SRC_DIR)
sys.path.insert(0, './utils')
from detectors.retinaface import init_detector, prepare_imgs, decode_batch
from video import read_frames_cv2
from visualise import show_images
conf = OmegaConf.load('../config/predict.yaml')
device = torch.device('cuda')
def merge_detector_cfg(conf: DictConfig) -> Dict[str, any]:
cfg = cfg_mnet if conf.encoder == 'mnet' else cfg_re50
for key in "batch_size, score_thresh, nms_thresh, top_k, keep_top_k".split(", "):
if key not in conf or conf[key] is None:
raise AttributeError("Missing {} in detector config".format(key))
cfg = {**cfg, **conf}
return cfg
face_det_conf = merge_detector_cfg(conf['face-detection'])
detector = init_detector(
face_det_conf,
face_det_conf['weights'],
device
).to(device)
# detect_fn = partial(detect, model=detector, cfg=face_det_conf, device=device)
files = os.listdir(VID_DIR)
# +
path = os.path.join(VID_DIR, files[12])
sample_np = read_frames_cv2(path, 30)
sample_orig = torch.from_numpy(sample_np).to(device)
D, H, W, C = sample_orig.shape
# -
sample, scale = prepare_imgs(sample_orig)
sample.shape, scale
D, C, H, W = sample.shape
priorbox = PriorBox(face_det_conf, image_size=(H, W))
priors = priorbox.forward().to(device)
scale = scale.to(device)
with torch.no_grad():
locations, confidence, landmarks = detector(sample)
# +
def postproc_frame_torch(boxes: Tensor, scores: Tensor, conf: Dict[str, Any]) -> Tensor:
idxs = (scores > conf['score_thresh']).nonzero().squeeze_(1)
if idxs.size(0):
boxes = boxes[idxs]
scores = scores[idxs]
# keep top-K before NMS
top_k = conf['top_k']
scores, idxs = scores.sort(descending=True)
scores, idxs = scores[:top_k], idxs[:top_k]
boxes = boxes[idxs]
# do NMS
nms_thresh = conf['nms_thresh']
keep_top_k = conf['keep_top_k']
keep = torchvision.ops.nms(boxes, scores, nms_thresh)
boxes = boxes[keep][:keep_top_k]
scores = scores[keep][:keep_top_k]
scores = scores.unsqueeze_(1)
return torch.cat([boxes, scores], dim=1)
else:
return torch.empty(0, 5, device=boxes.device, dtype=torch.float32)
def postproc_detections(locations: Tensor, confidence: Tensor, priors: Tensor,
scale: Tensor, conf: Dict[str, any], resize=1) -> List[Tensor]:
boxes = decode_batch(locations, priors, conf['variance'])
boxes = boxes * scale / resize
scores = confidence[:, :, 1]
N = boxes.size(0)
out = []
for f in range(N):
boxes_f = postproc_frame_torch(boxes[f], scores[f], conf)
out.append(boxes_f)
return out
# -
dets = postproc_detections(locations, confidence, priors, scale, face_det_conf)
resize = 1
boxes = decode_batch(locations, priors, face_det_conf['variance'])
boxes = boxes * scale / resize
# +
def calc_axis_torch(c0: IntTensor, c1: IntTensor, pad: IntTensor,
cmax: int) -> Tuple[IntTensor, ...]:
c0 = max(0, c0 - pad)
c1 = min(cmax, c1 + pad)
return c0, c1, c1 - c0
def expand_bbox_torch(bbox: FloatTensor, pct: float) -> FloatTensor:
bbox = bbox.clone().detach()
bbox[:2] *= 1 - pct
bbox[2:] *= 1 + pct
return bbox
def fix_coords_torch(bbox: FloatTensor, img_width: int, img_height: int) -> Tuple[FloatTensor, ...]:
x0, y0, x1, y1 = bbox.int()
x0.clamp_min_(0)
y0.clamp_min_(0)
x1.clamp_max_(img_width)
y1.clamp_max_(img_height)
return x0, y0, x1, y1
def crop_square_torch(img: IntTensor, bbox: FloatTensor, pad_pct=0.05) -> IntTensor:
C, H, W = img.shape
if pad_pct > 0:
bbox = expand_bbox_torch(bbox, pad_pct)
x0, y0, x1, y1 = fix_coords_torch(bbox, W, H)
w, h = x1 - x0, y1 - y0
if w > h:
pad = (w - h) // 2
y0, y1, h = calc_axis_torch(y0, y1, pad, H)
elif h > w:
pad = (h - w) // 2
x0, x1, w = calc_axis_torch(x0, x1, pad, W)
size = min(w, h)
face = img[:, y0:y1, x0:x1][:, :size, :size]
return face
# -
sample.shape, len(dets)
f = -1
f += 1
crop = crop_square_torch(sample_orig[f].permute(2, 0, 1), dets[f][0, :4])
print(crop.shape)
crop1 = crop.cpu().numpy().transpose(1, 2, 0)
show_images(crop1[None, :])
from dataset.transforms import ResizeTensor, SpatialGradFilter
resize = ResizeTensor(256, normalize=True)
crop1 = resize(crop)
crop1.shape
pipe = T.Compose([
ResizeTensor(256),
SpatialGradFilter(3),
T.Normalize(mean=[0.5]*3, std=[0.5]*3)
])
files = list(map(lambda i: f'_{i}_', range(100)))
import pandas as pd
df = pd.DataFrame(files, columns=['filename'])
df['label'] = 0.5
df.loc[2, 'label'] = 2
df.head()
df.to_csv('kek.csv', index=False)
c = np.ones(30, dtype=np.uint8)
c[range(0,20,2)] = 0
a = [1,2,3,4,5,6,7,8,9,10]
| notebooks/detect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Compute the mean absolute error for the following line and points:
#
# line: y = 1.2x + 2
#
# points: (2, -2), (5, 6), (-4, -4), (-7, 1), (8, 14)
points = [(2, -2), (5, 6), (-4, -4), (-7, 1), (8, 14)]
m = len(points)
s = 0
for i, p in enumerate(points):
p_x = p[0]
p_y = p[1]
y_hat = (1.2 * p_x) + 2
s += abs(p_y - y_hat)
res = s/m
res
# Compute the mean squared error for the following line and points:
#
# line: y = 1.2x + 2
#
# points: (2, -2), (5, 6), (-4, -4), (-7, 1), (8, 14)
points = [(2, -2), (5, 6), (-4, -4), (-7, 1), (8, 14)]
m = len(points)
s = 0
for i, p in enumerate(points):
p_x = p[0]
p_y = p[1]
y_hat = (1.2 * p_x) + 2
s += pow((p_y - y_hat), 2)
res = s/(2*m)
res
| experiments/ErrorFunctions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # One Step Methods
# Given the first order equation
# \begin{equation}
# y^{'} = 2y-0.1y^2,
# \end{equation}
# with the initial condition
# \begin{equation}
# y(0) = 1.
# \end{equation}
#
# Aproximate the solution of the inital value problem for the interval $0 \leq t \leq 4$ using $N=8$.
# Using the:
# 1. Euler method:
# $$ w_{i+1}=w_i+hf(t_i,w_i);$$
#
# 2. Midpoint method:
# $$ w_{i+1}=w_i+hf(t_i+\frac{h}{2},w_i+\frac{h}{2}k1),$$
# where $k1=f(t_i,w_i)$;
#
# 3. 4th Order Runge Kutta method:
# $$ w_{i+1}=w_i+\frac{h}{6}(k1+2k2+2k3+k4),$$
# where $$k1=f(t_i,w_i),$$
# $$k2=f(t_i+\frac{h}{2},w_i+\frac{h}{2}k1),$$
# $$k3=f(t_i+\frac{h}{2},w_i+\frac{h}{2}k2),$$
# $$k4=f(t_i+h,w_i+hk3);$$
#
# 4. Imposter-Butler method:
# $$ w_{i+1}=w_i+\frac{h}{4}(k1+2k2),$$
# where $$k1=f(t_i,w_i),$$
# $$k2=f(t_i+h,w_i+hk1).$$
#
# What is wrong with the Imposter-Butler method?
# ### DECLARING LIBRARIES
# +
import numpy as np
import math
# %matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import matplotlib.gridspec as gridspec # subplots
import warnings
warnings.filterwarnings("ignore")
# -
# ### Outputting Table
class ListTable(list):
""" Overridden list class which takes a 2-dimensional list of
the form [[1,2,3],[4,5,6]], and renders an HTML Table in
IPython Notebook. """
def _repr_html_(self):
html = ["<table>"]
for row in self:
html.append("<tr>")
for col in row:
html.append("<td>{0}</td>".format(col))
html.append("</tr>")
html.append("</table>")
return ''.join(html)
# ## Setting up the discrete time axis
# $$ h=\frac{4-0}{N}=0.5 $$
# $$t_i=t_0+ih$$
# $$t_i=0+0.5i$$
# for $i=0,1,2,...,8.$
# +
N=8
x_end=4.0
h=((x_end-0)/N)
INITIALCONDITION=1
time=np.zeros(N+1)
table = ListTable()
table.append(['time', 'Euler', 'Mid', 'RK','Imposter'])
# -
# ### Defining the function
# From the initial value problem the function
# $$ f(t,y)= 2y-0.1y^2. $$
def myfun(w):
return 2*w-0.1*w*w
# ## NUMERICAL SOLUTION
time[0]=0
# ## Euler Method
# $$w_0=1$$
# $$ w_{i+1}=w_i+h(2w_i-0.1w_i^2)$$
# $$N=0,1,2,...7$$
# +
Euler=np.zeros(N+1)
Euler[0]=INITIALCONDITION
for i in range (0,N):
# Euler Method
Euler[i+1]=Euler[i]+h*myfun(Euler[i])
# -
# ## Midpoint Method
# $$w_0=1$$
# $$k=2w_i-0.1w_i^2$$
# $$ w_{i+1}=w_i+h(2(w_i+\frac{h}{2}k)-0.1(w_i+\frac{h}{2}k)^2)$$
# $$N=0,1,2,...7$$
# +
Midpoint=np.zeros(N+1)
Midpoint[0]=INITIALCONDITION
for i in range (0,N):
# Mid-point method
Midpoint[i+1]=Midpoint[i]+h*myfun(Midpoint[i]+h/2*myfun(Midpoint[i]))
# -
# ## 4th Order Runge Kutta Method
# $$w_0=1$$
# $$k1=2w_i-0.1w_i^2$$
# $$k2=2(w_i+\frac{h}{2}k1)-0.1(w_i+\frac{h}{2}k1)^2$$
# $$k3=2(w_i+\frac{h}{2}k2)-0.1(w_i+\frac{h}{2}k2)^2$$
# $$k4=2(w_i+hk)-0.1(w_i+hk)^2$$
# $$ w_{i+1}=w_i+\frac{h}{6}(k1+2k2+2k3+k4)$$
# $$N=0,1,2,...7$$
# +
RK4=np.zeros(N+1)
RK4[0]=INITIALCONDITION
for i in range (0,N):
# 4th Order Runge Kutta
k1=myfun(RK4[i])
k2=myfun(RK4[i]+h/2*k1)
k3=myfun(RK4[i]+h/2*k2)
k4=myfun(RK4[i]+h*k3)
RK4[i+1]=RK4[i]+h/6*(k1+2*k2+2*k3+k4)
# -
# ## Imposter Butler Method
# $$w_0=1$$
# $$k1=2w_i-0.1w_i^2$$
# $$k2=2(w_i+hk1)-0.1(w_i+hk1)^2$$
# $$ w_{i+1}=w_i+\frac{h}{4}(k1+2k2)$$
# $$N=0,1,2,...7$$
Imposter=np.zeros(N+1)
Imposter[0]=INITIALCONDITION
i=0
table.append([time[i], Euler[i], Midpoint[i], RK4[i],Imposter[i]])
for i in range (0,N):
#Imposter method
k1=myfun(Imposter[i])
k2=myfun(Imposter[i]+h*k3)
Imposter[i+1]=Imposter[i]+h/4*(k1+2*k2)
time[i+1]=time[i]+h
table.append([time[i+1], Euler[i+1], Midpoint[i+1], RK4[i+1],Imposter[i+1]])
# ## Plotting
fig = plt.figure(figsize=(8,4))
plt.plot(time,Euler,'o:',color='red',label='Euler')
plt.plot(time,Midpoint,'o:',color='blue',label='Midpoint')
plt.plot(time,RK4,'o:',color='green',label='RK4')
plt.plot(time,Imposter,'o:',color='yellow',label='Imposter')
plt.legend(loc=4)
plt.title('Numerical Solutions of initial value problem.')
v
| Chapter 01 - Euler Methods/.ipynb_checkpoints/Example Euler Method 2nd Order Equation-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%%HTML
<script>
function code_toggle() {
if (code_shown){
$('div.input').hide('500');
$('#toggleButton').val('Show Code')
} else {
$('div.input').show('500');
$('#toggleButton').val('Hide Code')
}
# code_shown = !code_shown
}
$( document ).ready(function(){
code_shown=false;
$('div.input').hide()
});
</script>
<form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>
<a href="https://github.com/a1rb4Ck/MECA654"><img style="position: absolute; top: 0; right: 0; border: 0;" src="https://s3.amazonaws.com/github/ribbons/forkme_right_darkblue_121621.png" alt="Fork me on GitHub"></a>
# +
# MECA654, March 2018, <NAME>
# <NAME>, SYMME lab, Savoie Mont Blanc University
from IPython.display import display, HTML, IFrame, Math
from IPython.core.interactiveshell import InteractiveShell
import numpy as np
import matplotlib as mpl
from matplotlib import pyplot as plt # Loading Pyplot
from scipy.interpolate import interp1d
# Setting Matplotlib render backend
# %matplotlib notebook
# # %matplotlib nbagg
# # %pylab inline
from sympy import * # Loading Sympy for symbolic calculus
from sympy.physics.vector import * # Everything needed for vector calculs, ReferenceFrame, dynamic
from sympy.physics.vector import init_vprinting # To print time varying vectors
init_printing()
init_vprinting(pretty_print=True)
# Display full output in Jupyter Notebook
InteractiveShell.ast_node_interactivity = "all"
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# __MECA654, March 2018, <NAME>__
# [This interactive notebook is on Github](https://github.com/a1rb4Ck/MECA654)
#
# ## Exercise 7 - Spherical robot
# ### Speeds composition and acceleration calculus
#
# A robot is composed of 3 bodies.
# The mechanical links between bodies are :
# $0 / 1$ : pivot with axis $O \vec{y_1}$
# $1 / 2$ : pivot with axis $O \vec{z_1}$
# $2 / 3$ : linear slide with axis $O \vec{x_2}$
#
# Those links are controlled by three motors. Thus angles $\theta_1$, $\theta_2$ and the lenght $r$ are variable.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# 
# -
IFrame('http://www.glowscript.org/#/user/a1rb4Ck/folder/MECA654/program/sphericalrobot', width=640, height=562)
# +
# dynamicsymbols is a shortcut function to create undefined functions of time within SymPy
r, theta_1, theta_2, t, OM = symbols('r theta_1 theta_2 t OM')
theta_1, theta_2 = dynamicsymbols('theta_1 theta_2')
theta_1dot = dynamicsymbols('theta_1', 1)
theta_2dot = dynamicsymbols('theta_2', 1)
# Our reference frames for each body:
B0 = ReferenceFrame('B0', indices=['i', 'j', 'k'])
B1 = ReferenceFrame('B1', indices=['i', 'j', 'k'])
B2 = ReferenceFrame('B2', indices=['i', 'j', 'k'])
# We define space transformation relations in our problem:
B1.orient(B0, 'Axis', [theta_1, B0.y]) # Rotation of axis (O, y1), same as (O, y0)
B2.orient(B1, 'Axis', [-theta_2, B1.z]) # Rotation of axis (O, z1)
# We can easily get the DCM of any reference frame!
# B1.dcm(B0)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **1. What is the movement of $(3)$ in relation to $(2)$?** _It's a translation of $r$._
#
# ---
# **Method for solving**
# 1. Differentiate the $\overrightarrow{OM}$ vector in the main reference frame, here it is $(2)$
# 2. Express the vector in the asked reference frame, here it is $(1)$
# ---
#
# - Differentiate $\overrightarrow{V_{23}(M)}$ in the $(2)$ reference frame:
# + button=false new_sheet=false run_control={"read_only": false}
# We define the OM vector translation
OM = r(t) * B2['i']
display(Math(r'\overrightarrow{OM}=%s \ %s' % (latex(OM), latex('in \ the \ B_2 \ base'))))
# -
# Differentiate a vector with respect to a variable in a reference frame:
# V_23 = OM.diff(t, B2)
V_23 = OM.dt(B2) # The reference frame for derivation is (2) because we want V_23 !
display(Math(r'\overrightarrow{V_{23}(M)}=%s \ %s' % (latex(V_23), latex('in \ the \ B_2 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# - Express $\overrightarrow{V_{23}(M)}$ in the $(1)$ reference frame:
# + button=false new_sheet=false run_control={"read_only": false}
# We project the x2 vector in the B1 reference frame
# x_2 = cos(theta_2) * x_1 + sin(theta_2) * y_1 + 0 * z_1
# y_2 = -sin(theta_2) * x_1 + cos(theta_2) * y_1 + 0 * z_1
# z_2 = z_1
# Express the vector that we originally wrote in the B2 frame, in the B1 frame:
# V_23.express(B1) # V_23 in the B1 base
display(Math(r'\overrightarrow{V_{23}(M)}=%s \ %s' % (latex(V_23.express(B1)), latex('in \ the \ B_1 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **2. What is the movement of (2) in relation to (1)?** _It's a rotation of angle $\theta_2$._
# - Differentiate $\overrightarrow{V_{12}(M)}$ in the $(1)$ reference frame:
# + button=false new_sheet=false run_control={"read_only": false}
# We define the OM rotation movement
OM = r * B2.x
display(Math(r'\overrightarrow{OM}=%s \ %s' % (latex(OM), latex('in \ the \ B_2 \ base'))))
# -
# Angular velocity of B2 with respect to B1
# B2.ang_vel_in(B1)
display(Math(r'\Omega_2=%s \ %s' % (latex(B2.ang_vel_in(B1)), latex('angular \ velocity \ of \ B_2 \ with \ respect \ to \ B_1'))))
V_12 = OM.dt(B1) # The reference frame for derivation is (1) because we want V_12!
# V_12 # V_12 in the B2 base
display(Math(r'\overrightarrow{V_{12}(M)}=%s \ %s' % (latex(V_12), latex('in \ the \ B_2 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# - Express $\overrightarrow{V_{12}(M)}$ in the $(1)$ reference frame:
# + button=false new_sheet=false run_control={"read_only": false}
# We project the y2 vector in the B1 reference frame
# V_12.express(B1) # V_12 in the B1 base
display(Math(r'\overrightarrow{V_{12}(M)}=%s \ %s' % (latex(V_12.express(B1)), latex('in \ the \ B_1 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **3. What is the movement of $(1)$ in relation to $(0)$?** _It's a rotation of angle $\theta_1$_
# - Differentiate $\overrightarrow{V_{01}(M)}$ in the $(0)$ reference frame:
# + button=false new_sheet=false run_control={"read_only": false}
# We define the OM rotation movement
OM = B1.x
display(Math(r'\overrightarrow{OM}=%s \ %s' % (latex(OM), latex('in \ the \ B_1 \ base'))))
# -
V_01 = OM.dt(B0) # The reference frame for derivation is (0) because we want V_01!
display(Math(r'\overrightarrow{V_{01}(M)}=%s \ %s' % (latex(V_01), latex('in \ the \ B_1 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **4. What is the speed of M in the reference frame $(0)$?**
# - $\overrightarrow{V_{03}(M)} = \overrightarrow{V_{01}(M)} + \overrightarrow{V_{12}(M)} + \overrightarrow{V_{23}(M)}$ in the same reference frame, here we choose $(1)$ :
# + button=false new_sheet=false run_control={"read_only": false}
V_03 = V_01.express(B1) + V_12.express(B1) + V_23.express(B1)
# V_03.to_matrix(B1)
display(Math(r'\overrightarrow{V_{03}(M)}=%s \ %s' % (latex(V_03.express(B1)), latex('in \ the \ B_1 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# - Express $\overrightarrow{V_{03}(M)}$ in the $(0)$ reference frame:
# + button=false new_sheet=false run_control={"read_only": false}
# V_03.to_matrix(B0)
display(Math(r'\overrightarrow{V_{03}(M)}=%s \ %s' % (latex(V_03.express(B0)), latex('in \ the \ B_0 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **5. What is the torseur of $(3)$ in relation to $(0)$ ?**
# - $\Gamma_{03}(M) = \Gamma_{01}(M) + \Gamma_{12}(M) + \Gamma_{23}(M)$ in the (0) reference frame:
# - $\Gamma_{03}(M) = V_{03}(O) + \Omega_{03} \wedge \overrightarrow{OM}$
# + button=false new_sheet=false run_control={"read_only": false}
# We use matrix calculus with the reference frame change formula:
# V_03(M) = V_03(O) + Rot_03 ^ OM
V_03_O = Matrix([diff(r(t)) * cos(theta_2), diff(r(t)) * sin(theta_2), 0]) # in the B1 base
display(Math(r'\overrightarrow{V_{03}(M)}=%s \ %s' % (latex(V_03_O), latex('in \ the \ B_1 \ base'))))
Rot_03 = Matrix([0, diff(theta_1), diff(theta_2)]) # in the B1 base
display(Math(r'\overrightarrow{\Omega_{03}(M)}=%s \ %s' % (latex(Rot_03), latex('in \ the \ B_1 \ base'))))
# print('V_03(M) = V_03(O) + Rot_03 ^ OM')
OM = Matrix([r * cos(theta_2), r * sin(theta_2), 0])
V_03_M_B1 = V_03_O + Rot_03.cross(OM) # .cross(OM) is the same as ^OM
display(Math(r'\overrightarrow{V_{03}(M)}=%s \ %s' % (latex(V_03_M_B1), latex('in \ the \ B_1 \ base'))))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **6. What is the acceleration of $M$ in relation to $(0)$ ?**
# + button=false new_sheet=false run_control={"read_only": false}
# print("V_03''")
# V_03.dt(B0).to_matrix(B0)
display(Math(r'\overrightarrow{A_{03}(M)}=%s \ %s' % (latex(V_03.dt(B0).to_matrix(B0)), latex('in \ the \ B_0 \ base'))))
# + button=false new_sheet=false run_control={"read_only": false}
# TODO: Why don't we find the same results ??!
print("\n= = = = = = = = = = = = = = = = = = = = = = = =")
print("Solve the exercise with another method:\n")
# Another method with O and M points:
# Define the Origin Point
O = Point('O')
O.set_vel(B0, 0) # Set velocity of O
O.set_acc(B0, 0) # Set acceleration of O
# Define the M point from the origin
M = O.locatenew('M', r(t) * B2.x)
# Check the OM vector:
print("OM vector definition:")
M.pos_from(O)
# Set velocity of M in the B2 reference frame
print("M velocity in the B2 frame: aka V_23 in B2")
M.set_vel(B2, M.pos_from(O).dt(B2))
M.vel(B2)
print("M velocity in the B1 frame: aka V_23 in B1")
M.vel(B2).express(B1)
print("M velocity in the B1 frame: aka V_12 in B1")
M.set_vel(B1, (r * B2.x).dt(B1)) # Set velocity in the B1 reference frame
M.vel(B1).express(B1)
print("M velocity in the B0 frame: aka V_01 in B0")
M.set_vel(B0, (B1.x).dt(B0)) # Set velocity in the B1 reference frame
M.vel(B0).express(B0)
print("M velocity in the B1 frame: aka V_01 in B1")
M.vel(B0).express(B1)
print("M acceleration in the B1 frame:")
M.acc(B1).to_matrix(B1)
print("M acceleration in the B0 frame:")
trigsimp(M.acc(B0).to_matrix(B0))
# +
# %%HTML
<script>
$(document).ready(function(){
$('div.prompt').hide();
$('div.back-to-top').hide();
$('nav#menubar').hide();
$('.breadcrumb').hide();
$('.hidden-print').hide();
});
</script>
<footer id="attribution" style="float:right; color:#999; background:#fff;">
Created with Jupyter, Sympy, GlowScript, Numpy   -   MECA654 - <NAME> - <NAME> - March 2018
</footer>
| Kinematic_Exercise 7_Spherical Robot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Collecting images of last two decades of 21st JUNE using NASA API.
# # NASA API => APOD( Astronomy Picture of the Day)
# +
# import libraries
import requests
from datetime import date, timedelta
import os
import urllib.request
# set the start and end date for collecting images.
start_date = date(2000, 6, 21)
end_date = date(2020, 6, 21)
# execution of loop:
while start_date <= end_date:
# converting starting year from "string" to integer.
year = int(start_date.strftime("%y"))
# checking condition for LEAP YEARS betweeen 2000 to 2020 and setting days:
if year == 99 or year == 3 or year == 7 or year == 11 or year == 15 or year == 19:
delta =timedelta(days=366)
else:
delta = timedelta(days=365)
# "key" holds the NASA API KEY:
key = '<KEY>'
# Passing the parameters required to acess the api.
params = {'date': start_date.strftime("%Y-%m-%d") , 'hd': True, 'api_key': key }
# Getting the requests and passing parameters for query.
r = requests.get('https://api.nasa.gov/planetary/apod', params= params)
#saving in .json format.
r = r.json()
#printing starting date
print (start_date.strftime("%Y-%m-%d"))
#print the title of image.
print(r['title'])
#print new blank line.
print()
#printing explanation for image.
print(r['explanation'])
# print 100 stars.
print('*' * 100)
#Use below line to retrieve(save image) with date format("YYYY-MM-DD").jpg
#urllib.request.urlretrieve(r['hdurl'], start_date.strftime("%Y-%m-%d")+'.jpg')
#used 'title' to save images with title names of images(title.jpg).
urllib.request.urlretrieve(r['hdurl'], r['title']+'.jpg')
#adding delta(interval between two images)
start_date += delta
#adding year by 1
year = year +1
# -
# The Metadata that you can extract.
r.keys()
| scrap(APOD).ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# Exp2021-10-03/2
# ## Sugar production under short- and long-day light regime
#
#
# ***Motivation***: Clock mutants can show different metabolic performance under different light conditions
#
# Author: <NAME>, <NAME>
#
# ---
#
#
# 
#
# Phenotypes of 3-week-old plants
# **Experiment details:**
# Growth protocol: https://bio-protocol.org/bio101/e126
# Measurement protocols: Chlorophyll and starch assay (https://doi.org/10.1038/nprot.2009.12), Quantification of starch (https://doi.org/10.1038/nprot.2006.232)
# * Metabolites reported per g of fresh weight of 6-week-old plant leaf rosettes
# * Study date range: 2019-12-05 to 2020-01-05
# * SD = short days 6h light
# * LD = long days 18h light
#
# **Tested lines:**
# * WT
# * PhyA-211
# * elf4
# ---
#
# > This file is part of the Jupyter Notebook
# > lesson within the [FAIR in (biological) practice DataCarpentry](https://carpentries-incubator.github.io/fair-bio-practice/)
# > course run by Ed-DaSH.
#
# ---
# +
#################################################
# #
# Hashtags are not run as code and are used for #
# annotation of code in R. #
#################################################
# load required libraries
library(ggplot2)
# load data frame with 'fake' data for each genotype
df <- read.delim(file = "light_results.txt", stringsAsFactors=TRUE, header = TRUE, sep = "\t", dec = ".")
# -
# print data frame
df
# **Table 1:** Genotypes (WT = wild type, PhyA-211, elf4-101) are listed with respective measurements for biomas (g), starch (mg/g FW), sucrose (mg/g FW) and chlorophyll (mg/g FW)under short- and long-day light regimes.
#
# **Note:**
# Not all measurements could be recorded for chlorophyl, as at one timepoint the building fire alarm went off and research labs had to be evacuated when chlorophyll short-day time measurements were due. Three data points for PhyA-211 chlorophyll on long-day light regimes are not available as the extraction assay failed.
# # Visualise data
# +
# Create ggplot subset by light_condition
# first change order of factors of the genotype table
# ggplots standard setting is set to alphabetical order so your plot would read elf, phyA, WT
# If you rerun the code with a hashtag in front of the line you can look
# at the difference in the plot!
df$genotype <- factor(df$genotype, levels = c("WT", "PhyA-211","elf4-101"))
# change size of plot so it fits the screen before plotting
options(repr.plot.width = 5, repr.plot.height = 4)
# plot your graph
ggplot(subset(df, light_condition %in% "SD"), # subset only SD from light condition column for plotting
mapping = aes(x = genotype, y = biomas, fill = genotype)) + # x-axis shows genotype, y-axis shows biomas
geom_boxplot(alpha=0.3) +
labs(title = "Biomas per Genotype on short days",
x = "Genotype", # Title of x-axis
y = "Biomas (g)") + # Title of y-axis
scale_fill_manual(values=c("#999999", "#E69F00", "#56B4E9")) + # change colour of groups
theme_bw() +
theme(legend.position="none")
# ggsave: save the last ggplot
ggsave("genotype_by_biomas.png")
# -
# **Figure 1** Biomas (g) per genotype on short days shows a much larger biomas production in PhyA-211 in comparison to wild-type. Elf4-101 has a marginally smaller biomas production than wild-type during short days.
#
# To quantify this we have explored our data and run a ONE-way ANOVA plus Tukey multiple pairwise test.
# # Statistical testing
# Run a one way anova - compute the variance of biomas between genotypes
# ONLY from data within short-day light condition
res.aov <- aov(biomas ~ genotype, data = subset(df, light_condition %in% "SD"))
# Summary of the analysis
summary(res.aov)
# **NOTE** The output includes the columns F value and Pr(>F) corresponding to the p-value of the test.
#
# Given the above p-value in the summary, we can conclude that there are significant differences between the groups highlighted with (*) in the model summary. To find out which groups differ from each other, we conduct a Tukey post-hoc test to find between which groups these differences lie.
# conduct Tukey multiple pairwise-comparison
TukeyHSD(res.aov)
# **NOTE**
# * diff: difference between means of the two groups
# * lwr, upr: the lower and the upper end point of the confidence interval at 95% (default)
# * p adj: p-value after adjustment for the multiple comparisons.
#
# **RESULT**
# We can see that PhyA-211 and WT are not statistically different.
# Whilst elf4-101 and WT also do not differ statistically, there is a significant difference in biomas between elf4-101 and PhyA-211 (*).
| student_notebook_light_conditions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.3
# language: julia
# name: julia-0.6
# ---
# + [markdown] colab_type="text" id="JndnmDMp66FL"
# This notebook is based on the file [Improving Neural Net Performance programming exercise](https://colab.research.google.com/notebooks/mlcc/improving_neural_net_performance.ipynb?utm_source=mlcc&utm_campaign=colab-external&utm_medium=referral&utm_content=improvingneuralnets-colab&hl=en), which is part of Google's [Machine Learning Crash Course](https://developers.google.com/machine-learning/crash-course/).
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="hMqWDc_m6rUC"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="eV16J6oUY-HN" slideshow={"slide_type": "slide"}
# # Improving Neural Net Performance
# + [markdown] colab_type="text" id="0Rwl1iXIKxkm"
# **Learning Objective:** Improve the performance of a neural network by normalizing features and applying various optimization algorithms
#
# **NOTE:** The optimization methods described in this exercise are not specific to neural networks; they are effective means to improve most types of models.
# + [markdown] colab_type="text" id="lBPTONWzKxkn"
# ## Setup
#
# First, we'll load the data.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="VtYVuONUKxko"
using Plots
using StatPlots
using Distributions
gr()
using DataFrames
using TensorFlow
import CSV
import StatsBase
using PyCall
sess=Session(Graph())
california_housing_dataframe = CSV.read("california_housing_train.csv", delim=",");
california_housing_dataframe = california_housing_dataframe[shuffle(1:size(california_housing_dataframe, 1)),:];
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="B8qC-jTIKxkr"
function preprocess_features(california_housing_dataframe)
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
[:latitude,
:longitude,
:housing_median_age,
:total_rooms,
:total_bedrooms,
:population,
:households,
:median_income]]
processed_features = selected_features
# Create a synthetic feature.
processed_features[:rooms_per_person] = (
california_housing_dataframe[:total_rooms] ./
california_housing_dataframe[:population])
return processed_features
end
function preprocess_targets(california_housing_dataframe)
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = DataFrame()
# Scale the target to be in units of thousands of dollars.
output_targets[:median_house_value] = (
california_housing_dataframe[:median_house_value] ./ 1000.0)
return output_targets
end
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Ah6LjMIJ2spZ"
# Choose the first 12000 (out of 17000) examples for training.
training_examples = preprocess_features(head(california_housing_dataframe,12000))
training_targets = preprocess_targets(head(california_housing_dataframe,12000))
# Choose the last 5000 (out of 17000) examples for validation.
validation_examples = preprocess_features(tail(california_housing_dataframe,5000))
validation_targets = preprocess_targets(tail(california_housing_dataframe,5000))
# Double-check that we've done the right thing.
println("Training examples summary:")
describe(training_examples)
println("Validation examples summary:")
describe(validation_examples)
println("Training targets summary:")
describe(training_targets)
println("Validation targets summary:")
describe(validation_targets)
# + [markdown] colab_type="text" id="NqIbXxx222ea"
# ## Train the Neural Network
#
# Next, we'll set up the neural network similar to the previous exercise.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="6k3xYlSg27VB"
function construct_columns(input_features):
"""Construct the TensorFlow Feature Columns.
Args:
input_features: DataFrame of the numerical input features to use.
Returns:
A set of feature columns
"""
out=convert(Array, input_features[:,:])
return convert.(Float64,out)
end
# +
function create_batches(features, targets, steps, batch_size=5, num_epochs=0)
"""Create batches.
Args:
features: Input features.
targets: Target column.
steps: Number of steps.
batch_size: Batch size.
num_epochs: Number of epochs, 0 will let TF automatically calculate the correct number
Returns:
An extended set of feature and target columns from which batches can be extracted.
"""
if(num_epochs==0)
num_epochs=ceil(batch_size*steps/size(features,1))
end
names_features=names(features);
names_targets=names(targets);
features_batches=copy(features)
target_batches=copy(targets)
for i=1:num_epochs
select=shuffle(1:size(features,1))
if i==1
features_batches=(features[select,:])
target_batches=(targets[select,:])
else
append!(features_batches, features[select,:])
append!(target_batches, targets[select,:])
end
end
return features_batches, target_batches
end
function next_batch(features_batches, targets_batches, batch_size, iter)
"""Next batch.
Args:
features_batches: Features batches from create_batches.
targets_batches: Target batches from create_batches.
batch_size: Batch size.
iter: Number of the current iteration
Returns:
An extended set of feature and target columns from which batches can be extracted.
"""
select=mod((iter-1)*batch_size+1, size(features_batches,1)):mod(iter*batch_size, size(features_batches,1));
ds=features_batches[select,:];
target=targets_batches[select,:];
return ds, target
end
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="De9jwyy4wTUT"
function my_input_fn(features_batches, targets_batches, iter, batch_size=5, shuffle_flag=1):
"""Prepares a batch of features and labels for model training.
Args:
features_batches: Features batches from create_batches.
targets_batches: Target batches from create_batches.
iter: Number of the current iteration
batch_size: Batch size.
shuffle_flag: Determines wether data is shuffled before being returned
Returns:
Tuple of (features, labels) for next data batch
"""
# Construct a dataset, and configure batching/repeating.
ds, target = next_batch(features_batches, targets_batches, batch_size, iter)
# Shuffle the data, if specified.
if shuffle_flag==1
select=shuffle(1:size(ds, 1));
ds = ds[select,:]
target = target[select, :]
end
# Return the next batch of data.
return ds, target
end
# -
# Now we can set up the neural network itself.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="W-51R3yIKxk4"
function train_nn_regression_model(my_optimizer,
steps,
batch_size,
hidden_units,
keep_probability,
training_examples,
training_targets,
validation_examples,
validation_targets)
"""Trains a neural network model of one feature.
Args:
my_optimizer: Optimizer function for the training step
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
hidden_units: A vector describing the layout of the neural network
keep_probability: A `float`, the probability of keeping a node active during one training step.
Returns:
p1: Plot of RMSE for the different periods
training_rmse: Training RMSE values for the different periods
validation_rmse: Validation RMSE values for the different periods
"""
periods = 10
steps_per_period = steps / periods
# Create feature columns.
feature_columns = placeholder(Float32, shape=[-1, size(construct_columns(training_examples),2)])
target_columns = placeholder(Float32, shape=[-1, size(construct_columns(training_targets),2)])
# Network parameters
push!(hidden_units,size(training_targets,2)) #create an output node that fits to the size of the targets
activation_functions = Vector{Function}(size(hidden_units,1))
activation_functions[1:end-1]=z->nn.dropout(nn.relu(z), keep_probability)
activation_functions[end] = identity #Last function should be idenity as we need the logits
# create network - professional template
Zs = [feature_columns]
for (ii,(hlsize, actfun)) in enumerate(zip(hidden_units, activation_functions))
Wii = get_variable("W_$ii"*randstring(4), [get_shape(Zs[end], 2), hlsize], Float32)
bii = get_variable("b_$ii"*randstring(4), [hlsize], Float32)
Zii = actfun(Zs[end]*Wii + bii)
push!(Zs, Zii)
end
y=Zs[end]
loss=reduce_sum((target_columns - y).^2)
features_batches, targets_batches = create_batches(training_examples, training_targets, steps, batch_size)
# Optimizer setup with gradient clipping
gvs = train.compute_gradients(my_optimizer, loss)
capped_gvs = [(clip_by_norm(grad, 5.), var) for (grad, var) in gvs]
my_optimizer = train.apply_gradients(my_optimizer,capped_gvs)
run(sess, global_variables_initializer())
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
println("Training model...")
println("RMSE (on training data):")
training_rmse = []
validation_rmse=[]
for period in 1:periods
# Train the model, starting from the prior state.
for i=1:steps_per_period
features, labels = my_input_fn(features_batches, targets_batches, convert(Int,(period-1)*steps_per_period+i), batch_size)
run(sess, my_optimizer, Dict(feature_columns=>construct_columns(features), target_columns=>construct_columns(labels)))
end
# Take a break and compute predictions.
training_predictions = run(sess, y, Dict(feature_columns=> construct_columns(training_examples)));
validation_predictions = run(sess, y, Dict(feature_columns=> construct_columns(validation_examples)));
# Compute loss.
training_mean_squared_error = mean((training_predictions- construct_columns(training_targets)).^2)
training_root_mean_squared_error = sqrt(training_mean_squared_error)
validation_mean_squared_error = mean((validation_predictions- construct_columns(validation_targets)).^2)
validation_root_mean_squared_error = sqrt(validation_mean_squared_error)
# Occasionally print the current loss.
println(" period ", period, ": ", training_root_mean_squared_error)
# Add the loss metrics from this period to our list.
push!(training_rmse, training_root_mean_squared_error)
push!(validation_rmse, validation_root_mean_squared_error)
end
println("Model training finished.")
# Output a graph of loss metrics over periods.
p1=plot(training_rmse, label="training", title="Root Mean Squared Error vs. Periods", ylabel="RMSE", xlabel="Periods")
p1=plot!(validation_rmse, label="validation")
#
println("Final RMSE (on training data): ", training_rmse[end])
println("Final RMSE (on validation data): ", validation_rmse[end])
return p1, training_rmse, validation_rmse
end
# -
# Train the model with a Gradient Descent Optimizer and a learning rate of 0.0007.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="KueReMZ9Kxk7"
p1, training_rmse, validation_rmse = train_nn_regression_model(
train.GradientDescentOptimizer(0.0007), #optimizer & learning rate
5000, #steps
70, #batch_size
[10, 10], #hidden_units
1.0, # keep probability
training_examples,
training_targets,
validation_examples,
validation_targets)
# -
plot(p1)
# + [markdown] colab_type="text" id="flxmFt0KKxk9"
# ## Linear Scaling
# It can be a good standard practice to normalize the inputs to fall within the range -1, 1. This helps SGD not get stuck taking steps that are too large in one dimension, or too small in another. Fans of numerical optimization may note that there's a connection to the idea of using a preconditioner here.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Dws5rIQjKxk-"
function linear_scale(series)
min_val = minimum(series)
max_val = maximum(series)
scale = (max_val - min_val) / 2.0
return (series .- min_val) ./ scale .- 1.0
end
# + [markdown] colab_type="text" id="MVmuHI76N2Sz" slideshow={"slide_type": "slide"}
# ## Task 1: Normalize the Features Using Linear Scaling
#
# **Normalize the inputs to the scale -1, 1.**
#
# As a rule of thumb, NN's train best when the input features are roughly on the same scale.
#
# Sanity check your normalized data. (What would happen if you forgot to normalize one feature?)
#
# + [markdown] colab_type="text" id="Ax_IIQVRx4gr"
# Since normalization uses min and max, we have to ensure it's done on the entire dataset at once.
#
# We can do that here because all our data is in a single DataFrame. If we had multiple data sets, a good practice would be to derive the normalization parameters from the training set and apply those identically to the test set.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="yD948ZgAM6Cx"
function normalize_linear_scale(examples_dataframe):
"""Returns a version of the input `DataFrame` that has all its features normalized linearly."""
processed_features = DataFrame()
processed_features[:latitude] = linear_scale(examples_dataframe[:latitude])
processed_features[:longitude] = linear_scale(examples_dataframe[:longitude])
processed_features[:housing_median_age] = linear_scale(examples_dataframe[:housing_median_age])
processed_features[:total_rooms] = linear_scale(examples_dataframe[:total_rooms])
processed_features[:total_bedrooms] = linear_scale(examples_dataframe[:total_bedrooms])
processed_features[:population] = linear_scale(examples_dataframe[:population])
processed_features[:households] = linear_scale(examples_dataframe[:households])
processed_features[:median_income] = linear_scale(examples_dataframe[:median_income])
processed_features[:rooms_per_person] = linear_scale(examples_dataframe[:rooms_per_person])
return processed_features
end
normalized_dataframe = normalize_linear_scale(preprocess_features(california_housing_dataframe))
normalized_training_examples = head(normalized_dataframe, 12000)
normalized_validation_examples = tail(normalized_dataframe, 5000)
p1, graddescent_training_rmse, graddescent_validation_rmse = train_nn_regression_model(
train.GradientDescentOptimizer(0.005),
2000,
50,
[10, 10],
1.0,
normalized_training_examples,
training_targets,
normalized_validation_examples,
validation_targets)
# -
describe(normalized_dataframe)
plot(p1)
# + [markdown] colab_type="text" id="MrwtdStNJ6ZQ" slideshow={"slide_type": "slide"}
# ## Task 2: Try a Different Optimizer
#
# **Use the Momentum and Adam optimizers and compare performance.**
#
# The Momentum optimizer is one alternative. The key insight of Momentum is that a gradient descent can oscillate heavily in case the sensitivity of the model to parameter changes is very different for different model parameters. So instead of just updating the weights and biases in the direction of reducing the loss for the current step, the optimizer combines it with the direction from the previous step. You can use Momentum by specifying `MomentumOptimizer` instead of `GradientDescentOptimizer`. Note that you need to give two parameters - a learning rate and a "momentum" - with Momentum.
#
# For non-convex optimization problems, Adam is sometimes an efficient optimizer. To use Adam, invoke the `train.AdamOptimizer` method. This method takes several optional hyperparameters as arguments, but our solution only specifies one of these (`learning_rate`). In a production setting, you should specify and tune the optional hyperparameters carefully.
# + [markdown] colab_type="text" id="X1QcIeiKyni4"
# First, let's try Momentum Optimizer.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Ntn4jJxnypGZ"
p1, momentum_training_rmse, momentum_validation_rmse = train_nn_regression_model(
train.MomentumOptimizer(0.005, 0.05),
2000,
50,
[10, 10],
1.0,
normalized_training_examples,
training_targets,
normalized_validation_examples,
validation_targets)
# -
plot(p1)
# + [markdown] colab_type="text" id="5JUsCdRRyso3"
# Now let's try Adam.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="lZB8k0upyuY8"
p1, adam_training_rmse, adam_validation_rmse = train_nn_regression_model(
train.AdamOptimizer(0.2),
2000,
50,
[10, 10],
1.0,
normalized_training_examples,
training_targets,
normalized_validation_examples,
validation_targets)
# -
plot(p1)
# + [markdown] colab_type="text" id="twYgC8FGyxm6"
# Let's print a graph of loss metrics side by side.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="8RHIUEfqyzW0"
p2=plot(graddescent_training_rmse, label="Gradient descent training", ylabel="RMSE", xlabel="Periods", title="Root Mean Squared Error vs. Periods")
p2=plot!(graddescent_validation_rmse, label="Gradient descent validation")
p2=plot!(adam_training_rmse, label="Adam training")
p2=plot!(adam_validation_rmse, label="Adam validation")
p2=plot!(momentum_training_rmse, label="Momentum training")
p2=plot!(momentum_validation_rmse, label="Momentum validation")
# + [markdown] colab_type="text" id="UySPl7CAQ28C" slideshow={"slide_type": "slide"}
# ## Task 3: Explore Alternate Normalization Methods
#
# **Try alternate normalizations for various features to further improve performance.**
#
# If you look closely at summary stats for your transformed data, you may notice that linear scaling some features leaves them clumped close to `-1`.
#
# For example, many features have a median of `-0.8` or so, rather than `0.0`.
# +
# I'd like a better solution to automate this, but all ideas for eval
# on quoted expressions failed :-()
hist1=histogram(normalized_training_examples[:latitude], bins=20, title="latitude" )
hist2=histogram(normalized_training_examples[:longitude], bins=20, title="longitude" )
hist3=histogram(normalized_training_examples[:housing_median_age], bins=20, title="housing_median_age" )
hist4=histogram(normalized_training_examples[:total_rooms], bins=20, title="total_rooms" )
hist5=histogram(normalized_training_examples[:total_bedrooms], bins=20, title="total_bedrooms" )
hist6=histogram(normalized_training_examples[:population], bins=20, title="population" )
hist7=histogram(normalized_training_examples[:households], bins=20, title="households" )
hist8=histogram(normalized_training_examples[:median_income], bins=20, title="median_income" )
hist9=histogram(normalized_training_examples[:rooms_per_person], bins=20, title="rooms_per_person" )
plot(hist1, hist2, hist3, hist4, hist5, hist6, hist7, hist8, hist9, layout=9, legend=false)
# + [markdown] colab_type="text" id="Xx9jgEMHKxlJ"
# We might be able to do better by choosing additional ways to transform these features.
#
# For example, a log scaling might help some features. Or clipping extreme values may make the remainder of the scale more informative.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="baKZa6MEKxlK"
function log_normalize(series)
return log.(series.+1.0)
end
function clip(series, clip_to_min, clip_to_max)
return min.(max.(series, clip_to_min), clip_to_max)
end
function z_score_normalize(series)
mean_val = mean(series)
std_dv = std(series, mean=mean_val)
return (series .- mean) ./ std_dv
end
function binary_threshold(series, threshold)
return map(x->(x > treshold ? 1 : 0), series)
end
# + [markdown] colab_type="text" id="-wCCq_ClKxlO"
# The block above contains a few additional possible normalization functions.
#
# Note that if you normalize the target, you'll need to un-normalize the predictions for loss metrics to be comparable.
# + [markdown] colab_type="text" id="OMoIsUMmzK9b"
# These are only a few ways in which we could think about the data. Other transformations may work even better!
#
# `households`, `median_income` and `total_bedrooms` all appear normally-distributed in a log space.
# -
hist10=histogram(log_normalize(california_housing_dataframe[:households]), title="households")
hist11=histogram(log_normalize(california_housing_dataframe[:total_rooms]), title="total_rooms")
hist12=histogram(log_normalize(training_examples[:rooms_per_person]), title="rooms_per_person")
plot(hist10, hist11, hist12, layout=3, legend=false)
# `latitude`, `longitude` and `housing_median_age` would probably be better off just scaled linearly, as before.
#
# `population`, `total_rooms` and `rooms_per_person` have a few extreme outliers. They seem too extreme for log normalization to help. So let's clip them instead.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="XDEYkPquzYCH"
function normalize_df(examples_dataframe)
"""Returns a version of the input `DataFrame` that has all its features normalized."""
processed_features = DataFrame()
processed_features[:households] = log_normalize(examples_dataframe[:households])
processed_features[:median_income] = log_normalize(examples_dataframe[:median_income])
processed_features[:total_bedrooms] = log_normalize(examples_dataframe[:total_bedrooms])
processed_features[:latitude] = linear_scale(examples_dataframe[:latitude])
processed_features[:longitude] = linear_scale(examples_dataframe[:longitude])
processed_features[:housing_median_age] = linear_scale(examples_dataframe[:housing_median_age])
processed_features[:population] = linear_scale(clip(examples_dataframe[:population], 0, 5000))
processed_features[:rooms_per_person] = linear_scale(clip(examples_dataframe[:rooms_per_person], 0, 5))
processed_features[:total_rooms] = linear_scale(clip(examples_dataframe[:total_rooms], 0, 10000))
return processed_features
end
normalized_dataframe = normalize_df(preprocess_features(california_housing_dataframe))
normalized_training_examples = head(normalized_dataframe,12000)
normalized_validation_examples = tail(normalized_dataframe,5000)
p1, adam_training_rmse, adam_validation_rmse = train_nn_regression_model(
train.AdamOptimizer(0.15),
2000,
50,
[10, 10],
1.0,
normalized_training_examples,
training_targets,
normalized_validation_examples,
validation_targets)
# -
plot(p1)
# + [markdown] colab_type="text" id="b7atJTbzU9Ca" slideshow={"slide_type": "slide"}
# ## Optional Challenge: Use only Latitude and Longitude Features
#
# **Train a NN model that uses only latitude and longitude as features.**
#
# Real estate people are fond of saying that location is the only important feature in housing price.
# Let's see if we can confirm this by training a model that uses only latitude and longitude as features.
#
# This will only work well if our NN can learn complex nonlinearities from latitude and longitude.
#
# **NOTE:** We may need a network structure that has more layers than were useful earlier in the exercise.
# + [markdown] colab_type="text" id="1hwaFCE71OPZ"
# It's a good idea to keep latitude and longitude normalized:
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="djKtt4mz1ZEc"
function location_location_location(examples_dataframe)
"""Returns a version of the input `DataFrame` that keeps only the latitude and longitude."""
processed_features = DataFrame()
processed_features[:latitude] = linear_scale(examples_dataframe[:latitude])
processed_features[:longitude] = linear_scale(examples_dataframe[:longitude])
return processed_features
end
lll_dataframe = location_location_location(preprocess_features(california_housing_dataframe))
lll_training_examples = head(lll_dataframe,12000)
lll_validation_examples = tail(lll_dataframe,5000)
p1, lll_training_rmse, lll_validation_rmse = train_nn_regression_model(
train.AdamOptimizer(0.15),
500,
100,
[10, 10, 5, 5],
1.0,
lll_training_examples,
training_targets,
lll_validation_examples,
validation_targets)
# -
plot(p1)
# + [markdown] colab_type="text" id="Dw2Mr9JZ1cRi"
# This isn't too bad for just two features. Of course, property values can still vary significantly within short distances.
# +
#EOF
| 9. Improving Neural Net Performance Julia.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ETL Project Name: Boston Airbnb market analysis
# Primary independent variable: Price of the listing
# Other dependent variables: availability, review, location, host since, response time, host acceptance rate, no. of host listing
# Plots
# Prices vs review (How to categorize a review as Good, Bad and Neutral?)
# Price vs host location
# Price vs host since
# Price vs host response time
# Price vs host acceptance rate
# Price vs No. of host listing
#
# ## Import Packages and Libraries
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import pymysql
pymysql.install_as_MySQLdb()
from sqlalchemy import create_engine
#from ET_mysql_key import ET_key
from IPython.core.interactiveshell import InteractiveShell
#InteractiveShell.ast_node_interactivity = "all"
# -
# ### Import csv files
# +
# File to Load
# 1.) Listing file had information of each listing in Boston area.
#There are 95 columns with listing info, space and features, pictures, availability, price, host reviews, policy etc.
listing_data_to_load = "data/listings.csv"
# 2.) Calender file has availibility of rentals. It has listing_id, date, available (t or f) and price
availability_data_to_load = "data/calendars.csv"
# 3.) Review file contanin information such as listing_id, id, date, reviewer_id, reviewer_name & comments
review_data_to_load = "data/reviews.csv"
# -
listing=pd.read_csv(listing_data_to_load)
listing.columns
# ## EXTRACTION
# +
# Reading files and then storing into Pandas data frame. # Create DataFrames
# used pd.read to read & give out DataFrame(or TextParser) for the variables assigned to the pandas DateFrame are
#availability_data_to_load & listing_data_to_load & review_data_to_load
# -
# Listing data
# Show number of row and columns.# Display the data table for preview. Used head() to get the three rows
listing_df = pd.read_csv(listing_data_to_load)
listing_df.shape
pd.set_option('display.max_rows', 3)
pd.set_option('display.max_columns', 10)
pd.set_option('display.width', 100)
pd.set_option('precision', 5)
listing_df.style.highlight_null(null_color='red')
#lising_df.style
listing_df.head(3)
listing_df
# Availability data.
# Show number of row and columns.# Display the data table for preview. Used head() to get the three rows
availability_df = pd.read_csv(availability_data_to_load)
availability_df.shape
#pd.set_option('display.max_rows', 3)
#pd.set_option('display.max_columns', 5)
#pd.set_option('display.width', 100)
pd.set_option('precision', 5)
listing_df.style.highlight_null(null_color='red')
availability_df.head(3)
availability_df
# Review data
# Show number of row and columns.Display the data table for preview. Used head() to get the three rows
review_df = pd.read_csv(review_data_to_load)
review_df.shape
pd.option_context('display.colheader_justify','left')
review_df.style.set_properties(**{'text-align': 'left'})
review_df.head(3)
listing_df.columns
# ## TRANSFORM: Cleaning and Merging
# +
# Listing data.
# Show number of row and columns.
# Show stats.
# Removing Replace NaN with zero. unnecssary columns, standardizing column names, and creating a new df
# -
listing_df = listing_df[["listing_id","minimum_nights","host_response_rate","host_acceptance_rate","price","host_response_time"]]
listing_df.head(5)
#merge_table.head(5)
# Combine the data into a single dataset
#used the pd.merge merge DataFrame city_pd and ride_pd based on the common list city
#ALso, used how left based on result table needed and also specified how as left
#Used head() to get the five rows
mergeal_table = pd.merge(availability_df,listing_df,how="inner",on=["listing_id","listing_id"])
mergealr_table = pd.merge(mergeal_table,review_df,how="inner",on=["listing_id","listing_id"])
# Display the data table for preview
mergealr_table.head(5)
#mergealr_table
mergealr_table['latitude']=listing['latitude']
mergealr_table['longitude']=listing['longitude']
mergealr_table.columns
#pd.set_option('display.expand_frame_repr', False)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
mergealr_table.describe()
#Rename column to show the units such as % or $
#df.rename(index=str, columns={"A": "a", "C": "c"})
mergealr_table = mergealr_table.rename(index=str, columns={
"price_x": "price_x($)",
"host_response_rate": "host_response_rate(%)",
"host_acceptance_rate": "host_acceptance_rate(%)"})
#Identify missin values. Note after the dataframe was created the blanks were assigned a NaN value
mergealr_table["price_x($)"].head(5)
#Fill the NaN with avergae value of the rest of the data and i.e. is $209. The command below to replace did not work for somw reason
#mergealr_table["price_x($)"] = mergealr_table["price_x($)"].replace(r'^\s+$', 208, regex=True)
mergealr_table['price_x($)'] = mergealr_table['price_x($)'].fillna(209)
#Verified that the values got replaced
#mergealr_table["price_x"].tolist()
mergealr_table["price_x($)"].head(5)
#Identify if host_acceptance_rate(%) has blanks or NaN
mergealr_table['host_acceptance_rate(%)'].head(5)
# Replace the blanks with average of the rest of the values which is 50%
mergealr_table['host_acceptance_rate(%)'] = mergealr_table['host_acceptance_rate(%)'].fillna(50)
#Verify if the blanks are filled in with 50%
mergealr_table['host_acceptance_rate(%)'].head(3)
#Identify if host_response_rate(%) has blanks or NaN
mergealr_table['host_response_rate(%)'].head(5)
# Replace the blanks with average of the rest of the values which is 95%
mergealr_table['host_response_rate(%)'] = mergealr_table['host_response_rate(%)'].fillna(95)
#Verify if the blanks are filled in with 50%
mergealr_table['host_response_rate(%)'].head(5)
#Change or strip the % as a float type for host_acceptance_rate(%) and host_response_rate(%)
mergealr_table['host_acceptance_rate(%)'] = mergealr_table['host_acceptance_rate(%)'].str.rstrip('%').astype('float')
#mergealr_table['price_x($)'] = mergealr_table['price_x($)'].str.rstrip('$').astype('float')
mergealr_table['price_x($)'] = mergealr_table['price_x($)'].str.replace('$', '')
mergealr_table['price_x($)'] = mergealr_table['price_x($)'].str.replace(',', '')
mergealr_table['price_x($)'] = mergealr_table['price_x($)'].astype('float')
price=mergealr_table['price_x($)']
host_acceptance_rate=mergealr_table['host_acceptance_rate(%)']
plt.scatter(host_acceptance_rate, price)
plt.show()
# +
#mergealr_table['host_response_time']
# -
y_pos=mergealr_table.groupby(['host_response_time']).count()
y_pos
No_of_reservation=y_pos["listing_id"]
x=y_pos.index
# Replace the blanks with average of the rest of the values which is 50%
mergealr_table['listing_id'] = mergealr_table['listing_id'].fillna(1234)
# +
#host_response_time=mergealr_table['host_response_time']
#host_response_time = ('within an hour', 'within a few hours', 'within a day', 'a few days or more')
#y_poss = np.arange(len(host_response_time))
#y_pos = groupby(mergealr_table['host_response_time']).count()
plt.bar(x,No_of_reservation,align='center', alpha=0.5)
#plt.xticks(y_pos, host_response_time)
plt.ylabel('No. of Reservations')
plt.title('host_response_time')
#price=mergealr_table['price_x($)']
#plt.bar(host_response_time,price)
plt.show
# -
mergealr_table['host_response_rate(%)'] = mergealr_table['host_response_rate(%)'].str.rstrip('%').astype('float')
price=mergealr_table['price_x($)']
host_response_rate=mergealr_table['host_response_rate(%)']
plt.scatter(host_response_rate, price)
plt.show()
mergealr_table.columns
mergealr_table.add
# ## LOADING
#
# + active=""
# df.to_csv(file_name, sep='\t', encoding='utf-8')
# -
mergealr_table.isnull().any()
# +
#mergealr_table = mergealr_table[np.isfinite(mergealr_table['price_x($)'])]
# +
#mergealr_table = mergealr_table[np.isfinite(mergealr_table['host_response_rate(%)'])]
# +
#mergealr_table=mergealr_table.drop('minimum_nights', axis=1).drop('reviewer_id', axis=1).drop('comments', axis=1).drop('comments', axis=1).drop('reviewer_name', axis=1).drop('host_response_time', axis=1)
# -
mergealr_table.head(5)
mergealr_table.isnull().any()
mergealr_table.columns
mergealr_table.dtypes
# +
#new_mergealr_table=mergealr_table.head(1000)
# -
mergealr_table.to_csv("mergealr6.csv", encoding='utf-8')
# #### Alternate method code is mentioned below
# +
# import mysql.connector
# db = mysql.connector.connect(
# host="localhost",
# user="root",
# passwd="<PASSWORD>"
# )
# print(db)
# +
# import pymysql
# conn = pymysql.connect(host='127.0.0.1',
# port=3306,
# user='root',
# passwd='<PASSWORD>',
# db='ETL_HW',
# charset='utf8')
# mergealr_table.to_sql(name='ETL_HW', con=conn, if_exists = 'replace', index=False)
# +
# # Creating database connection
# ET_key for ET's MySQL access only
# connection_string = ("root:{0}@localhost/mergealr_table").format(ET_key)
# engine = create_engine(f'mysql://{connection_string}')
# +
# Confirming tables
#engine.table_names()
# -
# Loading dataframes into database
#listing.to_sql(name='host_response_rate', con=engine, if_exists='replace', index=True)
#listing.to_sql(name='host_acceptance_rate', con=engine, if_exists='replace', index=True)
#availbility_Df.to_sql(name='price_x', con=engine, if_exists='replace', index=True)
# +
# Confirming that tables loaded successfully
#pd.read_sql("select * from price_x limit 3",con = engine)
#pd.read_sql("select * from host_response_rate",con = engine)
#pd.read_sql("select * from host_acceptance_rate limit 3",con = engine)
# -
| project2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''enterprise'': conda)'
# name: python3
# ---
# # Computing the optimal statistic with enterprise
#
# * In this notebook you will learn how to compute the optimal statistic.
# * The optimal statistic is a frequentist detection statistic for the stochastic background.
# * It assesses the significance of the cross-correlations and compares them to the Hellings-Downs curve.
#
# * For more information, see [Anholm et al. 2009](https://arxiv.org/abs/0809.0701), [Demorest et al. 2013](https://arxiv.org/abs/1201.6641), [Chamberlin et al. 2015](https://arxiv.org/abs/1410.8256), [Vigeland et al. 2018](https://arxiv.org/abs/1805.12188).
#
# * This notebook shows you how to compute the optimal statistic for the 12.5yr data set.
# +
from __future__ import (absolute_import, division,
print_function, unicode_literals)
import numpy as np
import pickle
import json
import glob
import os
import matplotlib.pyplot as plt
# %matplotlib inline
from enterprise.signals import signal_base
from enterprise.signals import gp_signals
from enterprise.pulsar import Pulsar
from enterprise_extensions import models
from enterprise_extensions.frequentist import optimal_statistic as opt_stat
import sys
sys.path.append("..")
from settings import matplotsettings
import matplotlib
matplotsettings()
# -
# ## Load pulsar data
psrlist = None # define a list of pulsar name strings that can be used to filter.
# set the data directory
datadir = '../data'
if not os.path.isdir(datadir):
datadir = '../../data'
print(datadir)
# +
# for the entire pta
parfiles = sorted(glob.glob(datadir + '/par/*par'))
timfiles = sorted(glob.glob(datadir + '/tim/*tim'))
# filter
if psrlist is not None:
parfiles = [x for x in parfiles if x.split('/')[-1].split('.')[0] in psrlist]
timfiles = [x for x in timfiles if x.split('/')[-1].split('.')[0] in psrlist]
# Make sure you use the tempo2 parfile for J1713+0747!!
# ...filtering out the tempo parfile...
parfiles = [x for x in parfiles if 'J1713+0747_NANOGrav_12yv3.gls.par' not in x]
# +
# check for file and load pickle if it exists:
pickle_loc = datadir + '/psrs.pkl'
if os.path.exists(pickle_loc):
with open(pickle_loc, 'rb') as f:
psrs = pickle.load(f)
# else: load them in slowly:
else:
psrs = []
ephemeris = 'DE438'
for p, t in zip(parfiles, timfiles):
psr = Pulsar(p, t, ephem=ephemeris)
psrs.append(psr)
# +
## Get parameter noise dictionary
noise_ng12 = datadir + '/channelized_12p5yr_v3_full_noisedict.json'
params = {}
with open(noise_ng12, 'r') as fp:
params.update(json.load(fp))
# -
# ### Use `enterprise_extensions` to make a model with a common red process
# (Note: It will take a few minutes to run this cell and may require at least ~4GB RAM)
pta = models.model_2a(psrs, noisedict=params, gamma_common=4.33, n_gwbfreqs=5)
# ### Initialize the optimal statistic object
ostat = opt_stat.OptimalStatistic(psrs, pta=pta, orf='hd')
ostat_dip = opt_stat.OptimalStatistic(psrs, pta=pta, orf='dipole')
ostat_mono = opt_stat.OptimalStatistic(psrs, pta=pta, orf='monopole')
# ## Load the maximum-likelihood values for the pulsars' red noise parameters and the common red process
# * These values come from the results of a Bayesian search (model 2A)
# * Once you have done your own Bayesian search, you can make your own parameter dictionary of maximum-likelihood values
with open(datadir + '/12p5yr_maxlike.json', 'r') as f:
ml_params = json.load(f)
# ## Optimal statistics with maximum likelihood noise parameters
# * The optimal statistic returns five quantities:
# - xi: an array of the angular separations between the pulsar pairs (in radians)
# - rho: an array of the cross-correlations between the pulsar pairs
# - sig: an array of the uncertainty in the cross-correlations
# - OS: the value of the optimal statistic
# - OS_sig: the uncertainty in the optimal statistic
#
# +
xi, rho, sig, OS, OS_sig = ostat.compute_os(params=ml_params)
print(OS, OS_sig, OS/OS_sig)
_, _, _, OS_dip, OS_sig_dip = ostat_dip.compute_os(params=ml_params)
print(OS_dip, OS_sig_dip, OS_dip/OS_sig_dip)
_, _, _, OS_mono, OS_sig_mono = ostat_mono.compute_os(params=ml_params)
print(OS_mono, OS_sig_mono, OS_mono/OS_sig_mono)
# -
# ### Plot the cross-correlations and compare to the Hellings-Downs curve
# +
def get_HD_curve(zeta):
coszeta = np.cos(zeta*np.pi/180.)
xip = (1.-coszeta) / 2.
HD = 3.*( 1./3. + xip * ( np.log(xip) -1./6.) )
return HD/2
## Before plotting, we need to bin the cross-correlations
def weightedavg(rho, sig):
weights, avg = 0., 0.
for r,s in zip(rho,sig):
weights += 1./(s*s)
avg += r/(s*s)
return avg/weights, np.sqrt(1./weights)
def bin_crosscorr(zeta, xi, rho, sig):
rho_avg, sig_avg = np.zeros(len(zeta)), np.zeros(len(zeta))
for i,z in enumerate(zeta[:-1]):
myrhos, mysigs = [], []
for x,r,s in zip(xi,rho,sig):
if x >= z and x < (z+10.):
myrhos.append(r)
mysigs.append(s)
rho_avg[i], sig_avg[i] = weightedavg(myrhos, mysigs)
return rho_avg, sig_avg
# sort the cross-correlations by xi
idx = np.argsort(xi)
xi_sorted = xi[idx]
rho_sorted = rho[idx]
sig_sorted = sig[idx]
# bin the cross-correlations so that there are the same number of pairs per bin
npairs = 66
xi_mean = []
xi_err = []
rho_avg = []
sig_avg = []
i = 0
while i < len(xi_sorted):
xi_mean.append(np.mean(xi_sorted[i:npairs+i]))
xi_err.append(np.std(xi_sorted[i:npairs+i]))
r, s = weightedavg(rho_sorted[i:npairs+i], sig_sorted[i:npairs+i])
rho_avg.append(r)
sig_avg.append(s)
i += npairs
xi_mean = np.array(xi_mean)
xi_err = np.array(xi_err)
# -
# ## Reproduce the bottom panel of Figure 5 in [https://arxiv.org/abs/2009.04496](https://arxiv.org/abs/2009.04496)
# +
# reproduce the bottom panel of Figure 5 in https://arxiv.org/abs/2009.04496
plt.figure(figsize=(8, 4))
(_, caps, _) = plt.errorbar(xi_mean*180/np.pi, rho_avg, xerr=xi_err*180/np.pi, yerr=sig_avg, marker='o', ls='',
color='0.1', capsize=4, elinewidth=1.2)
zeta = np.linspace(0.01,180,100)
HD = get_HD_curve(zeta+1)
plt.plot(zeta, OS*HD, ls='--', label='Hellings-Downs', color='C0', lw=1.5)
plt.plot(zeta, zeta*0.0+OS_mono, ls='--', label='Monopole', color='C1', lw=1.5)
plt.plot(zeta, OS_dip*np.cos(zeta*np.pi/180), ls='--', label='Dipole', color='C2', lw=1.5)
plt.xlim(0, 180);
plt.ylim(-3.5e-30, 3.5e-30);
plt.ylabel(r'$\hat{A}^2 \Gamma_{ab}(\zeta)$')
plt.xlabel(r'$\zeta$ (deg)');
plt.legend(loc=4);
plt.tight_layout();
plt.show();
# -
# ## Noise marginalized optimal statistics
# * To compute the noise-marginalized optimal statistic (Vigeland et al. 2018), you will need the chain from a Bayesian search for a common red process without spatial correlations (model 2A).
# * NOTE: This file requires the "model_2a, 5 frequency power law, fixed spectral index (gamma=4.33)" chain from `download_full_chains.ipynb`
# ### Load the samples from a Bayesian run
# +
# (Note: It will take a minute to run this cell)
chain = np.loadtxt('../../chains/downloaded_chains/12p5yr_DE438_model2a_cRN5freq_gammaFixed4.33_chain.gz')
params = np.loadtxt('../../chains/downloaded_chains/12p5yr_DE438_model2a_cRN5freq_gammaFixed4.33_pars.txt', dtype=str)
# -
# ### Take the median values from the chains and dump them into a `json` file
# +
param_dict = {}
params = list(params)
for p in params:
param_dict.update({p: np.median(chain[:, params.index(p)])})
with open('../../data/12p5yr_median.json', 'w') as f:
json.dump(param_dict, f)
# +
# (Note: It may take a few minutes to run this cell)
N = 1000 # number of times to compute the optimal statistic - official analysis used 10k - you can increase this if you don't mind the extra runtime
burn = int(0.25*chain.shape[0]) # estimate of when the chain has burned in
noisemarg_OS, noisemarg_OS_err = np.zeros(N), np.zeros(N)
noisemarg_OS_dip, noisemarg_OS_dip_err = np.zeros(N), np.zeros(N)
noisemarg_OS_mono, noisemarg_OS_mono_err = np.zeros(N), np.zeros(N)
for i in range(N):
if i%100==0:
print("Status: ", i, "/", N, " -- ", i/N*100, " %")
# choose a set of noise values from the chain
# make sure that you pull values from after the chain has burned in
idx = np.random.randint(burn, chain.shape[0])
# construct a dictionary with these parameter values
param_dict = {}
for p in params:
param_dict.update({p: chain[idx, params.index(p)]})
# compute the optimal statistic at this set of noise values and save in an array
_, _, _, noisemarg_OS[i], noisemarg_OS_err[i] = ostat.compute_os(params=param_dict)
_, _, _, noisemarg_OS_dip[i], noisemarg_OS_dip_err[i] = ostat_dip.compute_os(params=param_dict)
_, _, _, noisemarg_OS_mono[i], noisemarg_OS_mono_err[i] = ostat_mono.compute_os(params=param_dict)
# -
# ## Reproduce Figure 4 (bottom panel) from [https://arxiv.org/abs/2009.04496](https://arxiv.org/abs/2009.04496)
# +
plt.figure(figsize=(8, 4))
plt.hist(noisemarg_OS/noisemarg_OS_err, histtype='step', lw=2, label='Hellings-Downs', color='C0',
bins=20, density=True)
plt.hist(noisemarg_OS_mono/noisemarg_OS_mono_err, histtype='step', lw=2, label='Monopole', color='C1',
bins=20, density=True)
plt.hist(noisemarg_OS_dip/noisemarg_OS_dip_err, histtype='step', lw=2, label='Dipole', color='C2',
bins=20, density=True)
# plt.gca().axvline(x=np.mean(noisemarg_OS/noisemarg_OS_err), ls='--', color='C0')
# plt.gca().axvline(x=np.mean(noisemarg_OS_mono/noisemarg_OS_mono_err), ls='--', color='C1')
# plt.gca().axvline(x=np.mean(noisemarg_OS_dip/noisemarg_OS_dip_err), ls='--', color='C2')
# plt.gca().axvline(x=OS/OS_sig, ls=':', color='C0')
# plt.gca().axvline(x=OS_mono/OS_sig_mono, ls=':', color='C1')
# plt.gca().axvline(x=OS_dip/OS_sig_dip, ls=':', color='C2')
plt.legend(loc="upper right")
plt.xlabel("S/N")
plt.ylabel("PDF")
plt.show()
# -
# ## Reproduce Figure 4 (top panel)
# +
plt.figure(figsize=(8, 4))
plt.hist(noisemarg_OS, histtype='step', lw=2, label='Hellings-Downs', color='C0', bins=20, density=True)
plt.hist(noisemarg_OS_mono, histtype='step', lw=2, label='Monopole', color='C1', bins=20, density=True)
plt.hist(noisemarg_OS_dip, histtype='step', lw=2, label='Dipole', color='C2', bins=20, density=True)
plt.hist((10**(chain[:,params.index("gw_log10_A")]))**2, histtype='step', lw=2, label='Uncorrelated common process',
color='grey', ls='--', bins=50, density=True)
# plt.gca().axvline(x=np.mean(noisemarg_OS), ls='--', color='C0')
# plt.gca().axvline(x=np.mean(noisemarg_OS_mono), ls='--', color='C1')
# plt.gca().axvline(x=np.mean(noisemarg_OS_dip), ls='--', color='C2')
# plt.gca().axvline(x=np.mean(10**(2*chain[:,params.index("gw_log10_A")])), ls='--', color='grey')
# plt.gca().axvline(x=OS, ls=':', color='C0')
# plt.gca().axvline(x=OS_mono, ls=':', color='C1')
# plt.gca().axvline(x=OS_dip, ls=':', color='C2')
plt.xlim((-1e-30, 9e-30))
plt.legend(loc="upper right")
plt.xlabel(r'$\hat{A}^2$ and $A^2_{\rm CP}$')
plt.ylabel("PDF")
plt.show()
# -
# ## Remarks
# * The monopole correlation shows a large SNR compared to the other two correlations
# * However, only the amplitude of the HD correlation is consistent with the Bayesian samples
# * Both monopole and dipole correlations are disfavored from figure 3.
| tutorials/full_pta_gwb/optimal_stat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: p6
# language: python
# name: other-env
# ---
# +
"""setup"""
try:
from ..__init__ import activate
except:
import aye.activate
from aye import from_file
__file__ = globals().get('__file__', 'test_basics.ipynb')
from nbformat import v4
from pathlib import Path
try:
from . import test_basics
except:
import test_basics
"""local_value = 1000"""
local_value = 42
if __name__ == '__main__':
global_value = "Does not exist on imports."
# -
"""test_import"""
assert test_basics.__file__.endswith(__file__), "a notebook source file was not provided."
assert __import__('__main__') != test_basics, "the imported module is a different object entirely."
"""test_reload"""
from importlib import reload
assert reload(test_basics) is test_basics, "aye cannot reload test_basics"
"""test_attributes"""
if __name__ == '__main__':
assert test_basics.local_value is local_value, "the import probably barfed."
assert getattr(test_basics, 'global_value', None) is None, "__name__ is __main__"
"""test_source_format"""
from nbformat import validate, reads
from inspect import getsource
assert validate(reads(getsource(test_basics), 4)) or True
| aye/tests/test_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="historical-terminology"
import os
import pydot
import torch
from sklearn.model_selection import train_test_split
import time
# + id="covered-tulsa"
import glob
import os
import numpy as np
import pickle
import re
labels = []
sentences = []
for file in glob.glob('./txt/*.txt', recursive=True):
header =""
print(file)
with open(file, 'r') as f:
count = 0
sents = re.split(r'\.\s', f.read())
for sentence in sents:
if "Краткая информация" in sentence:
header = 0
elif "Диагностика" in sentence:
header = 1
elif "Лечение" in sentence:
header = 2
elif "Реабилитация" in sentence:
header = 3
elif "Профилактика" in sentetnce:
header = 4
if header != '':
labels.append(header)
sentences.append(sentence.replace('\n', ' '))
# -
# + id="selected-index"
X_train, X_test, y_train, y_test = train_test_split(sentences, labels, test_size=0.5)
y_train = torch.tensor(y_train, dtype=torch.long)
y_test = torch.tensor(y_test, dtype=torch.long)
# -
import pandas as pd
data = pd.DataFrame()
data['id'] = [i for i in range(len(y_train))]
data['label'] = y_train
data['text'] = X_train
# +
class MyDataset(torch.utils.data.Dataset):
def __init__(self, encodings, labels):
self.encodings = encodings
self.labels = labels
def __getitem__(self, idx):
item = {key: torch.tensor(val[idx]) for key, val in self.encodings.items()}
item['labels'] = self.labels[idx]
return item
def __len__(self):
return len(self.labels)
# -
from transformers import DistilBertTokenizerFast
tokenizer = DistilBertTokenizerFast.from_pretrained('DeepPavlov/rubert-base-cased-sentence', model_max_length=512)
train_encodings = tokenizer(X_train, truncation=True, padding=True)
val_encodings = tokenizer(X_test, truncation=True, padding=True)
print(train_encodings[0])
train_dataset = MyDataset(train_encodings, y_train)
val_dataset = MyDataset(val_encodings, y_test)
# +
from torch.utils.data import DataLoader
from transformers import BertForSequenceClassification, AdamW
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model = BertForSequenceClassification.from_pretrained('DeepPavlov/rubert-base-cased-sentence', num_labels=4)
model.to(device)
model.train()
train_loader = DataLoader(train_dataset, batch_size=16, shuffle=True)
optim = AdamW(model.parameters(), lr=5e-5)
running_loss = 0
total_train_loss = 0
start_time = time.time()
for i,batch in enumerate(train_loader):
optim.zero_grad()
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
labels = batch['labels'].to(device)
outputs = model(input_ids, attention_mask=attention_mask, labels=labels)
loss = outputs[0]
loss.backward()
optim.step()
running_loss += loss.data
total_train_loss += loss.data
print("Epoch {}, {:d}% \t train_loss: {:.2f} took: {:.2f}s".format(
epoch+1, int(100 * (i+1) / 16), running_loss, time.time() - start_time))
running_loss = 0.0
start_time = time.time()
model.eval()
# -
torch.save(model, "my.model")
# +
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
model = torch.load("../model-final2", map_location=device)
model.to(device)
model.eval()
# +
from torch.utils.data import DataLoader
import numpy
val_loader = DataLoader(val_dataset, batch_size=16, shuffle=True)
model.eval()
y = numpy.array([])
y_pred = numpy.array([])
with torch.no_grad():
for i, batch in enumerate(val_loader):
input_ids = batch['input_ids'].to(device)
attention_mask = batch['attention_mask'].to(device)
labels = batch['labels'].to(device)
outputs = model(input_ids, attention_mask=attention_mask)
y = numpy.append(y, labels.cpu().detach().numpy())
y_pred = numpy.append(y_pred, numpy.argmax(outputs[0].cpu().detach().numpy(), 1))
print(accuracy_score(y, y_pred))
# -
embeds = []
for s in sentences:
tokens = tokenizer([s], truncation=True, padding=True)
input_ids = torch.tensor(tokens.input_ids).to(device)
att = torch.tensor(tokens.attention_mask).to(device)
outputs = model(input_ids, attention_mask=att, output_hidden_states=True)
embeds.append(outputs.hidden_states[0])
import numpy as np
encoding = None
while True:
request = input()
encoding = tokenizer([request], truncation=True, padding=True)
input_ids = torch.tensor(encoding.input_ids).to(device)
att = torch.tensor(encoding.attention_mask).to(device)
outputs = model(input_ids, attention_mask=att)
label = np.argmax(outputs[0].detach().numpy(), 1)[0]
required_sentences = [y for x,y in list(zip(labels, sentences)) if x == label]
required_encodings = tokenizer(required_sentences, truncation=True, padding=True)
distances = []
encoding
from sklearn.metrics import classification_report
print(classification_report(y, y_pred))
| .ipynb_checkpoints/Finetuning-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulación Montecarlo
# > El método de Montecarlo es un método no determinista o estadístico numérico, usado para aproximar expresiones matemáticas complejas y costosas de evaluar con exactitud. El método se llamó así en referencia al Casino de Montecarlo (Mónaco) por ser “la capital del juego de azar”, al ser la ruleta un generador simple de números aleatorios. El nombre y el desarrollo sistemático de los métodos de Montecarlo datan aproximadamente de 1944 y se mejoraron enormemente con el desarrollo de la computadora.
#
# Referencia:
# - https://es.wikipedia.org/wiki/M%C3%A9todo_de_Montecarlo
# ___
# ## 1. Introducción
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/5/54/Monte_carlo_method.svg" width="300px" height="100px" />
#
# - Inventado por <NAME> y a <NAME>. Ulam ha explicado cómo se le ocurrió la idea mientras jugaba un solitario durante una enfermedad en 1946.
# - Advirtió que resulta mucho más simple tener una idea del resultado general del solitario haciendo pruebas múltiples con las cartas y contando las proporciones de los resultados que computar todas las posibilidades de combinación formalmente.
# - Se le ocurrió que esta misma observación debía aplicarse a su trabajo de Los Álamos sobre difusión de neutrones, para la cual resulta prácticamente imposible solucionar las ecuaciones íntegro-diferenciales que gobiernan la dispersión, la absorción y la fisión.
# - Dado que ya empezaban a estar disponibles máquinas de computación para efectuar las pruebas numéricas, el método cobró mucha fuerza.
# - El método de Montecarlo proporciona soluciones aproximadas a una gran variedad de problemas matemáticos posibilitando la realización de experimentos con muestreos de números pseudoaleatorios en una computadora. El método es aplicable a cualquier tipo de problema, ya sea estocástico o determinista.
# - El método de Montecarlo tiene un error absoluto de la estimación que decrece como $\frac{1}{\sqrt{N}}$ en virtud del teorema del límite central.
#
# ### Ejemplo
# Todos alguna vez hemos aplicado el método Montecarlo (inconscientemente). Como ejemplo, consideremos el juego de Astucia Naval.
#
# Normalmente, primero se realizan una serie de tiros a puntos aleatorios. Una vez se impacta en un barco, se puede utilizar un algoritmo determinista para identificar la posición del barco y así terminar de derrumbarlo.
#
#
# ___
# ## 2. Caminata aleatoria
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/d/da/Random_Walk_example.svg" width="300px" height="100px" />
#
# Una caminata aleatoria (*random walk* en inglés) es una formalización matemática de la trayectoria que resulta al hacer pasos sucesivos aleatorios. Un ejemplo elemental de caminata aleatoria es la caminata aleatoria en la línea de números enteros $\mathbb{Z}$, la cual comienza en $0$ y en cada paso se mueve $+1$ o $-1$ con igual probabilidad.
#
# Otros ejemplos:
# - Trayectoria de una molécula al viajar en un fluido (líquido o gas).
# - El camino que sigue un animal en su búsqueda de comida.
# - El precio fluctuante de una acción.
# - La situación de un apostador en un juego de azar.
#
# Todos pueden ser aproximados por caminatas aleatorias, aunque no sean en verdad procesos aleatorios.
#
# ### Caminata aleatoria en una dimensión
# Como dijimos, un ejemplo elemental de caminata aleatoria es la caminata aleatoria en la línea de números enteros $\mathbb{Z}$, la cual comienza en $0$ y a cada paso se mueve $+1$ o $-1$ con igual probabilidad.
#
# Esta caminata se puede ilustrar como sigue:
# - Se posiciona en $0$ en la línea de números enteros y una moneda justa se tira.
# - Si cae en **sol** nos moveremos una unidad a la derecha.
# - Si cae en **águila** nos moveremos una unidad a la izquierda.
#
# Notemos que después de $5$ pasos podremos estar en 1, −1, 3, −3, 5, or −5. Las posibilidades son las siguientes:
#
# <img style="float: center; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/0/05/Flips.svg" width="900px" height="300px" />
#
#
# Referencia:
# - https://en.wikipedia.org/wiki/Random_walk
#
# **Importante:** librería random.
#
# Referencia:
# - https://docs.python.org/3/library/random.html
# ## Caminata aleatoria
from IPython.display import YouTubeVideo
YouTubeVideo('Y77WnkLbT2Q')
# ## ¿Qué es una simulación montecarlo?
#
# Revisitamos el concepto de simulación montecarlo.
#
# > La idea de una simulación montecarlo es probar muchos resultados posibles. En la realidad, solo uno de esos resultados posibles se dará, pero, en términos de evaluación de riesgos, cualquiera de las posibilidades podría ocurrir.
#
# Los simuladores montecarlo se usan usualmente para evaluar el riesgo de una estrategia de negocios dada con opciones y acciones.
#
# Los simuladores montecarlo pueden ayudar a tomar decisiones exitosas, y que el resultado de una decisión no sea la única medida de si dicha decisión fue buena. Las decisiones no deben ser evaluadas después del resultado. Por el contrario, los riesgos y beneficios solo deben ser considerados en el momento en que se debe tomar la decisión, sin prejuicios retrospectivos. Un simulador montecarlo puede ayudar a visualizar muchos (o en algunos casos, todos) de los resultados potenciales para tener una mejor idea de los riesgos de una decisión.
#
# > ## <font color = "blue"> Tarea:
# > **Ver https://pythonprogramming.net/monte-carlo-simulator-python/**
# ### Usamos montecarlo para evaluar el resultado de la caminata aleatoria
#
# - Ver el valor esperado de la caminata después de N pasos.
#
# - Luego, evaluar el proceso utilizando montecarlo y comparar resultados.
# Hacer código acá
import random
import numpy as np
import matplotlib.pyplot as plt
from functools import reduce
# Caminata aleatorio método convencional
def randon_walk(N):
x = 0
xx = [x]
add_el = xx.append
for i in range(N):
z = random.choice([-1,1])
x += z
add_el(x)
return xx,x
# Caminata aleatoria vectorizada
def Vec_randon_walk(N:'Cantidad de pasos',x0:'Posición inicial'):
# Vector con todas las trayectorias
z = np.zeros(N)
z[0] = x0; z[1:] = np.random.choice([-1,1],N-1)
# Posición final en toda la trayectoria
x = reduce(lambda z1,z2: z1+z2,z)
# xx2 = [x0]
xx = [x0]
[xx.append(xx[-1]+z[i+1]) for i in range(N-1)]
# print(z,xx)
return xx
# +
# Usando el método ineficiente
N = 10000 # número de pasos
n = 100 # cantidad de trayectorias
final = []
# # %matplotlib inline
for j in range(n):
xx,x = randon_walk(N)
final.append(x)
plt.plot(xx)
plt.show()
print('En promedio el caminante esta en :',np.mean(final))
# +
# Usando el método vectorizado
N = 10000 # número de pasos
n = 100 # cantidad de trayectorias
x0 = 0 # Condición inicial
xx = np.asmatrix([Vec_randon_walk(N,x0) for i in range(n)])
plt.plot(xx.T)
plt.show()
# -
# ## Ejemplo
#
# Ahora analicemos el ejemplo básico del apostador.
#
# Referencia:
# - https://pythonprogramming.net/monte-carlo-simulator-python/
#
# Supongamos que estamos en un casino especial, donde el usuario puede tirar un *dado metafórico* que puede dar como resultado un número del uno (1) al número cien (100).
#
# Si el usuario tira cualquier número entre 1 y 50, el casino gana. Si el usuario tira cualquier número entre 51 y 99, el usuario gana. Si el usuario tira 100, pierde.
#
# Con esto, el casino mantiene un margen del 1%, el cual es mucho más pequeño que el margen típico en casinos, al igual que el margen de mercado cuando se incorporan costos por transacción.
#
# Por ejemplo, [Scottrade](https://www.scottrade.com/) cobra \$7 USD por transacción. Si se invierten \$1000 USD por acción, esto significa que tienes que pagar \$7 USD para entrar, y \$7 USD para salir, para un total de \$14 USD.
#
# Esto pone el margen en <font color ='red'> $1.4\%$ </font>. Esto significa, que a largo plazo, las ganancias tienen que ser mayores a $1.4\%$ en promedio, de otra manera se estará perdiendo dinero.
#
# De nuevo, con nuestro ejemplo en mente, 1-50, la casa gana. 51-99 el usuario gana. Un 100 significa que la casa gana.
#
# Ahora, comencemos. Primero tenemos que crear nuestro dado.
# Crear una función para que devuelva simplemente ganar(true) o perder(false)
def tirar_dado():
dado = random.randint(1,100)
if dado>0 and dado<= 50 or dado == 100:
return False
else:
return True
# +
# Probar la función creada para ver que funcione
N = 100
contador_ganar = 0
contador_perder = 0
for i in range(N):
if tirar_dado():
contador_ganar +=1
# print('FELICIDADES!!!!!')
else:
contador_perder+=1
# print('Sigue intentando')
print("Ganamos", contador_ganar, " veces y perdimos", contador_perder, " veces.")
# -
# Ahora, necesitamos crear un **apostador**. Empezaremos con uno extremadamente básico por ahora. Veremos, que aún con un apostador muy básico, veremos cosas muy reveladoras usando un simulador montecarlo.
# Crearemos un apostador simple. Las caracterísitcas son: se empieza con un capital inicial,
# siempre se apuesta lo mismo, y se va a apostar un número determinado de veces.
def apostador(cap_inicial, apuesta, n_apuestas):
capital = cap_inicial
ccapital = [capital]
for i in range(n_apuestas):
if tirar_dado():
capital += apuesta
ccapital.append(capital)
else:
capital -= apuesta
ccapital.append(capital)
return ccapital,capital
# Ver como evolucionan los fondos de nuestro apostador al jugar 100 veces
# # %matplotlib inline
ccapital,capital = apostador(10000, 100, 1000)
print('El capital luego del juego fue',capital)
plt.plot(ccapital)
plt.show()
# (Montecarlo) Simular varios (100) escenarios en que se apuestan
# 50, 100, 1000 y 10000 veces. ¿Qué pasa?
N = 100
n = [50,100,1000,10000]
final = []
i=1
capital = 10000
for j in n:
ccapital,capital2 = apostador(capital, 100, j)
final.append(capital2)
plt.figure(i)
plt.plot(ccapital,label= 'se apuestan %i veces' %j)
plt.legend()
i += 1
print('En promedio mi dinero fue:',np.mean(final))
plt.show()
# Graficar historial para ver el comportamiento del de los fondos para 1000 juegos distintos
plt.hist(final,10)
plt.show()
print(np.mean(final))
# Por esto los apostadores pierden. Normalmente las probabilidades no están evidentemente muy en contra de ellos, solo un poco. Los casinos únicamente entienden psicología básica: ganar es extremadamente adictivo. Por ello, los casino se construyen para mantenerte jugando.
#
# En el corto plazo, la mayoría de los jugadores no se dan cuenta que son más propensos a perder. Las veces que ganan y pierden son muy parejas. Estadísticamente, casi la mitad de las personas terminarán con ganancias después de jugar unas pocas veces. El problema es la adicción, y que ellos continuarán apostando, y por ende perdiendo sus ganancias. Es matemática extremadamente básica, pero la psicología humana es débil.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME> and modified by <NAME>.
# </footer>
| TEMA-2/Clase8_SimulacionMontecarlo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: mxnet
# language: python
# name: mxnet
# ---
import mxnet as mx
mnist = mx.test_utils.get_mnist()
batch_size = 100
train_iter = mx.io.NDArrayIter(mnist['train_data'], mnist['train_label'], batch_size, shuffle=True)
val_iter = mx.io.NDArrayIter(mnist['test_data'], mnist['test_label'], batch_size)
# The following source code defines a convolutional neural network architecture called LeNet. LeNet is a popular network known to work well on digit classification tasks. We will use a slightly different version from the original LeNet implementation, replacing the sigmoid activations with tanh activations for the neurons
#
# 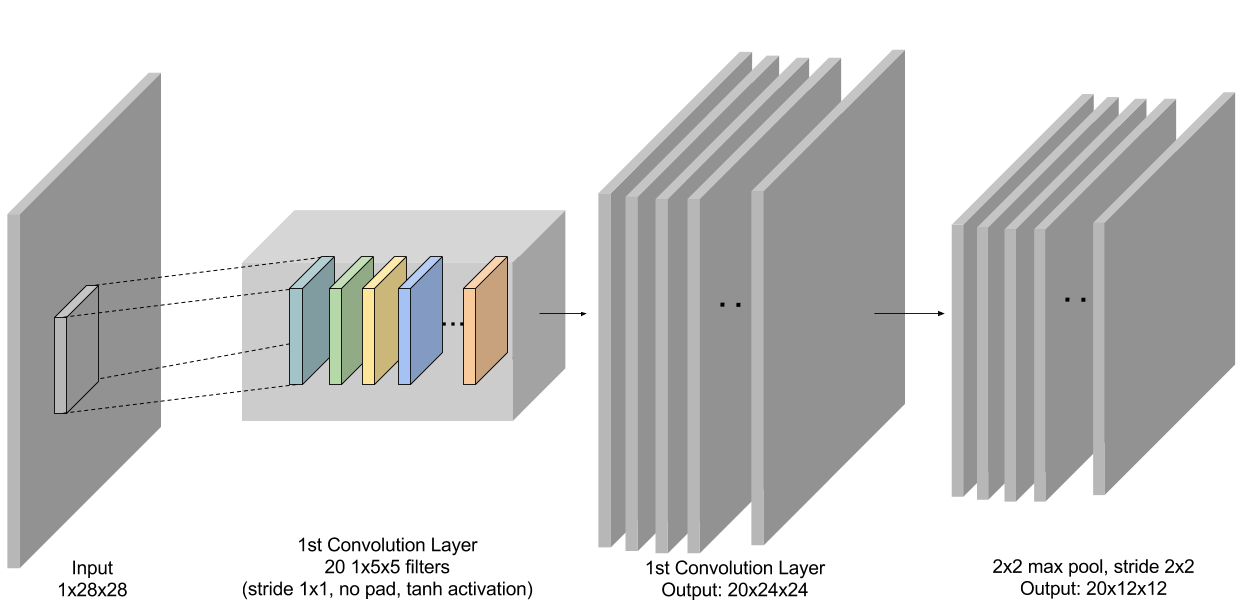
data = mx.sym.var('data')
# first conv layer
conv1 = mx.sym.Convolution(data=data, kernel=(5,5), num_filter=20)
tanh1 = mx.sym.Activation(data=conv1, act_type="tanh")
pool1 = mx.sym.Pooling(data=tanh1, pool_type="max", kernel=(2,2), stride=(2,2))
# second conv layer
conv2 = mx.sym.Convolution(data=pool1, kernel=(5,5), num_filter=50)
tanh2 = mx.sym.Activation(data=conv2, act_type="tanh")
pool2 = mx.sym.Pooling(data=tanh2, pool_type="max", kernel=(2,2), stride=(2,2))
# first fullc layer
flatten = mx.sym.flatten(data=pool2)
fc1 = mx.symbol.FullyConnected(data=flatten, num_hidden=500)
tanh3 = mx.sym.Activation(data=fc1, act_type="tanh")
# second fullc
fc2 = mx.sym.FullyConnected(data=tanh3, num_hidden=10)
# softmax loss
lenet = mx.sym.SoftmaxOutput(data=fc2, name='softmax')
# create a trainable module
lenet_model = mx.mod.Module(symbol=lenet, context=mx.gpu())
# train with the same
lenet_model.fit(train_iter,
eval_data=val_iter,
optimizer='sgd',
optimizer_params={'learning_rate':0.1},
eval_metric='acc',
batch_end_callback = mx.callback.Speedometer(batch_size, 100),
num_epoch=10)
test_iter = mx.io.NDArrayIter(mnist['test_data'], None, batch_size)
prob = lenet_model.predict(test_iter)
test_iter = mx.io.NDArrayIter(mnist['test_data'], mnist['test_label'], batch_size)
# predict accuracy for lenet
acc = mx.metric.Accuracy()
lenet_model.score(test_iter, acc)
print(acc)
assert acc.get()[1] > 0.98
# Let's try again with ReLu activations
data = mx.sym.var('data')
# first conv layer
conv1 = mx.sym.Convolution(data=data, kernel=(5,5), num_filter=20)
tanh1 = mx.sym.Activation(data=conv1, act_type="relu")
pool1 = mx.sym.Pooling(data=tanh1, pool_type="max", kernel=(2,2), stride=(2,2))
# second conv layer
conv2 = mx.sym.Convolution(data=pool1, kernel=(5,5), num_filter=50)
tanh2 = mx.sym.Activation(data=conv2, act_type="relu")
pool2 = mx.sym.Pooling(data=tanh2, pool_type="max", kernel=(2,2), stride=(2,2))
# first fullc layer
flatten = mx.sym.flatten(data=pool2)
fc1 = mx.symbol.FullyConnected(data=flatten, num_hidden=500)
tanh3 = mx.sym.Activation(data=fc1, act_type="relu")
# second fullc
fc2 = mx.sym.FullyConnected(data=tanh3, num_hidden=10)
# softmax loss
lenet = mx.sym.SoftmaxOutput(data=fc2, name='softmax')
# create a trainable module
lenet_model = mx.mod.Module(symbol=lenet, context=mx.gpu())
# train with the same
lenet_model.fit(train_iter,
eval_data=val_iter,
optimizer='sgd',
optimizer_params={'learning_rate':0.1},
eval_metric='acc',
batch_end_callback = mx.callback.Speedometer(batch_size, 100),
num_epoch=10)
test_iter = mx.io.NDArrayIter(mnist['test_data'], None, batch_size)
prob = lenet_model.predict(test_iter)
test_iter = mx.io.NDArrayIter(mnist['test_data'], mnist['test_label'], batch_size)
# predict accuracy for lenet
acc = mx.metric.Accuracy()
lenet_model.score(test_iter, acc)
print(acc)
assert acc.get()[1] > 0.98
# Add BatchNorm
data = mx.sym.var('data')
# first conv layer
conv1 = mx.sym.Convolution(data=data, kernel=(5,5), num_filter=20)
tanh1 = mx.sym.Activation(data=conv1, act_type="relu")
pool1 = mx.sym.Pooling(data=tanh1, pool_type="max", kernel=(2,2), stride=(2,2))
# second conv layer
conv2 = mx.sym.Convolution(data=pool1, kernel=(5,5), num_filter=50)
tanh2 = mx.sym.Activation(data=conv2, act_type="relu")
pool2 = mx.sym.Pooling(data=tanh2, pool_type="max", kernel=(2,2), stride=(2,2))
# first fullc layer
flatten = mx.sym.flatten(data=pool2)
fc1 = mx.sym.FullyConnected(data=flatten, num_hidden=4096)
tanh3 = mx.sym.Activation(data=fc1, act_type="relu")
bn1 = mx.sym.BatchNorm(data=tanh3)
dropout = mx.sym.Dropout(bn1, p = 0.2)
# second fullc
fc2 = mx.sym.FullyConnected(data=dropout, num_hidden=10)
# softmax loss
lenet = mx.sym.SoftmaxOutput(data=fc2, name='softmax')
# create a trainable module
lenet_model = mx.mod.Module(symbol=lenet, context=mx.gpu())
# train with the same
lenet_model.fit(train_iter,
eval_data=val_iter,
optimizer='sgd',
optimizer_params={'learning_rate':0.1},
eval_metric='acc',
batch_end_callback = mx.callback.Speedometer(batch_size, 100),
num_epoch=10)
test_iter = mx.io.NDArrayIter(mnist['test_data'], None, batch_size)
prob = lenet_model.predict(test_iter)
test_iter = mx.io.NDArrayIter(mnist['test_data'], mnist['test_label'], batch_size)
# predict accuracy for lenet
acc = mx.metric.Accuracy()
lenet_model.score(test_iter, acc)
print(acc)
assert acc.get()[1] > 0.98
| src/digits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# McKee_Scattering_2013
# <NAME>
# September 10, 2020
#
# McKee_Scattering_2013 corrects ac-s data for undetected scattering inside the absorption and attenuation tubes based
# on methods devised by McKee et al. (2013) (full citaiton below). These methods assumd different reflectance efficiencies
# (rw) of ac-s absorption tube. Therefore, the script outputs 10 different correction files, one for each of 10 reflectance
# efficiencies ranging from 1 (100%) reflectance to 0.95 (95% reflectance).
# IMPORTANT: this script cannot run without: MK13_coefFILE.h5
#
# Full citation:
# <NAME>., and <NAME> al. (2013) Evaluation and Improvement of an Iterative Scattering Correction Scheme for
# in situ absorption and attenuation measurements. Journal of atmospheric and ocean technology 30:1527-1541
# -
# Import libraries
import numpy as np
import h5py
from tkinter import filedialog as fd
from matplotlib import pyplot as plt
from datetime import datetime as dt
import os
from distutils.dir_util import copy_tree
# +
# 1. Define function for retrieving reflective tube specific coefficients.
def McKEE2013_COEFF(**kwargs):
"""This program is used to get the proper flow tube (r_w) specific McKee et al. (2013)
scattering coefficients
Keyword arguments (input):
r_w - reflection efficiency of ac-s absorption tube. This variable is qualitative and must be one
of the following values: 1, 0.999, 0.998, 0.995, 0.99, 0.985, 0.98, 0.97, 0.96 , & 0.95
Outputs:
coeff_array - tuple of absorption coefficients according to user-selected r_w. Based on McKee et al. (2013)"""
### 1. Create preliminary variables for acquiring correct scattering coefficients based on r_w
r_w = kwargs['r_w'] # Percent of scattering as input by user (based on keyword argument)
coeff_DICT = {} # Empty dictionary
pwd = os.getcwd()+'/' + 'MK13_coefFILE.h5'# Find present working directory and select MK13_coefFILE.h5
### 2. Open MK13_coefFILE.h5 and select correct set of coefficients
with h5py.File(pwd) as hf:
# Open the hdf5 file created in the previous cell
for m in hf.keys():
# for-loop fills in the dictionary
coeff_DICT[m] = hf[m][()] # Place coefficients into empty dictionary
### 3. Create an array of coefficients for the proper
# Creates a matrix of nans in the dimensions of the a coefficients and the number of
# reflection efficiencies (r_w's) (below)
COEFF_MAT = np.ones([len(coeff_DICT['a_0']),len(coeff_DICT.keys())-1])*np.nan
for i in np.arange(7):
# This for-loop fills in the matrix with the correct coefficient values (a_0-a_6)
COEFF_MAT[:,i] = coeff_DICT['a_' + str(i)] # Fill matrix by columns
booL_IND = coeff_DICT['r_w'] == r_w # bool index the correct r_w as input by the user
coeff_array = tuple(COEFF_MAT[booL_IND,:].reshape(7,)) # make a tuple of the correct matrix row
return(coeff_array) # Output a tuple
# +
# 2. Define functions needed for pre-processing ac-s data in preparation for McKee et al.'s
# (2013) scattering correction approach
def acs_matlab_LOAD(acs_fileNAME):
"""acs_matlab_LOAD loads .mat/hdf5 files of ac-s data that is partially processed by matlab.
It relies on the program 'acsPROCESS_INTERACTIVE.m.
Input variables:
acs_fileNAME - ac-s data. Must be hdf5 file (also formatted like .mat file)
Output variables:
A_CORR - absorption matrix
C_CORR - attenuation matrix
lambdA - array of wavelengths
deptH - array of depths (corresponding to spectra)"""
with h5py.File(acs_fileNAME) as data: # Read in data as a .mat/hdf5 file
A_CORR = data['A_CORR'][()].transpose() # Call in the absorption matrix
C_CORR = data['C_CORR'][()].transpose() # Call in the attenuation matrix
lamdA = data['a_wv'][()] # Call in the ac-s wavelengths (note: c is already interpolated to a wavelengths)
deptH = data['deptH'][0] # Call in depth array
return(A_CORR,C_CORR,lamdA,deptH)
def QAQC_acsDATA(A_CORR,C_CORR,lambdA,deptH):
"""QAQC_acsDATA performs QA/QC on ac-s data and removes spectra with at least one channel for which
a > c.
Input variables:
A_CORR - absorption matrix
C_CORR - attenuation matrix
lambdA - array of wavelengths
deptH - array of depths (corresponding to spectra)
Output variables:
A_CORR, C_CORR, lambdA - same variables as above, with contaminated spectra (a>c) removed"""
### 1. Create boolian indices and repository variables for QA/QC
wvl_BOOL = lambdA <= 715 # Use bool logic to find all wavelengths less than 715 nm
ac_IND = C_CORR > A_CORR # Use bool logic to find all instances where c > a (matrix)
ac_QUIVER = [] # Create a list for acs rows for which a > c at some point in the row.
### 2. Index rows (spectra) for which a > c
for b,aC in enumerate(ac_IND):
# For loop cycles through the bool logic matrix row by row in order to determine if
# a > c at any point (bool = 0)
neg_ARRAY = [i for i,j in enumerate(aC[wvl_BOOL[:,0]]) if j == 0] # Find channels where a > c
if neg_ARRAY != []:
# If at any point a > c in a given row
ac_QUIVER.append(b) # Add the index of the row to the "empty" list
ac_QUIVER.sort(reverse=True) # Sort all a>c rows into reverse order
### 3. Remove contaiminated spectra (a > c) from absorption, attenuation, and depth
for i in ac_QUIVER:
# This for-loop detetes rows from a and c matrices for which a > c at ANY channel.
# It uses indices in descending order so as not to shift them before their deletions.
A_CORR = np.delete(A_CORR, i, axis=0) # Delete row from a
C_CORR = np.delete(C_CORR, i, axis=0) # Delete row from c
deptH = np.delete(deptH, i, axis=0) # Delete depth from d
return(A_CORR,C_CORR,deptH) # Returns variables after contaminated rows removed
def QAQC_negative(A_CORR,C_CORR,lambdA,deptH):
"""QAQC_negative finds a/c values that are negative and removes their spectra from matrices
Input variables:
A_CORR - absorption matrix
C_CORR - attenuation matrix
lambdA - array of wavelengths
deptH - array of depths (corresponding to spectra)
Output variables:
A_CORR, C_CORR, lambdA - same variables as above, with contaminated spectra (a or c < 0) removed"""
wvl_BOOL = lambdA <= 715 # Use bool logic to find all wavelengths less than 715 nm
for i in range(2):
# For-loop goes through two cycles. Cycle 1 (i = 0) deals with attenuation. Cycle 2 (i = 1)
# deals with absorption
if i == 0:
ac_IND = C_CORR > 0 # Use bool logic to find all instances where c > a (matrix)
else:
ac_IND = A_CORR > 0 # Use bool logic to find all instances where c > a (matrix)
ac_QUIVER = [] # Create a list for acs rows for which a > c at some point.
for b,aC in enumerate(ac_IND):
# For loop cycles through the bool logic matrix row by row in order to determine if
# a > c at any point (bool = 0)
neg_ARRAY = [i for i,j in enumerate(aC[wvl_BOOL[:,0]]) if j == 0] # Find channels where a > c
if neg_ARRAY != []:
# If at any point a > c in a given row
ac_QUIVER.append(b) # Add the index of the row to the "empty" list
ac_QUIVER.sort(reverse=True) # Sort all a>c rows into reverse order
for i in ac_QUIVER:
# This for-loop detetes rows from a and c matrices for which a > c at ANY channel.
# It uses indices in descending order so as not to shift them before their deletions.
A_CORR = np.delete(A_CORR, i, axis=0) # Delete row from a
C_CORR = np.delete(C_CORR, i, axis=0) # Delete row from c
deptH = np.delete(deptH, i, axis=0) # Delete depth from d
return(A_CORR,C_CORR,deptH)
def acs_BINNING(A_CORR,C_CORR,lambdA,deptH,**parameters):
"""acs_BINNING bins ac-s data in to depth bins as specified by the user. User must specify
absorption
Input variables:
A_CORR - absorption matrix
C_CORR - attenuation matrix
lambdA - array of wavelengths
deptH - array of depths (corresponding to spectra)
Keyword arguments:
biN_size - specifiy the depth bin size to use for the binning process
Output variables:
deptH_BINS - array of depth bins (median values)
A_CORR_bin - depth-binned absorption spectra
C_CORR_bin - depth-binned attenuation spectra"""
biN_size = parameters['biN_size'] # Establish bin size from keyword arguments
maX_deptH = np.ceil(max(deptH)) # Find the limit for binned depths
deptH_BIN_EDGES = np.arange(0,maX_deptH+biN_size,biN_size) # Create an array of bin edges
deptH_BINS = (deptH_BIN_EDGES[:-1]+deptH_BIN_EDGES[1:])/2 # Find median bin values
A_CORR_bin = np.ones([len(deptH_BINS),len(lambdA)])*np.nan # Create nan matrix for binned a
C_CORR_bin = np.ones([len(deptH_BINS),len(lambdA)])*np.nan # Create nan matrix for binned c
for ii,d in enumerate(deptH_BINS):
# For-loop indexes individual depths according to the proper bin and averages them according
# to wavelength.
difF_deptH = deptH - d # Find differences between depths and bin medians
# Find indices of all depths that fall within the given bin
biN_IND = [i for i,j in enumerate(difF_deptH) if j > -(biN_size/2) and j <= (biN_size/2)]
naN_IND = [] # Create an empty array for potential non-existant bins (e.g. ac-s within a bin)
if biN_IND != []:
# If ac-s exist within a given depth bin
A_CORR_bin[ii,:] = np.mean(A_CORR[biN_IND,:],axis=0) # Average selected a spectra
C_CORR_bin[ii,:] = np.mean(C_CORR[biN_IND,:],axis=0) # Average selected c spectra
else:
# If no ac-s exist within a given depth bin
naN_IND.append(ii) # Put index into the naN_IND list. Do not calculate means
naN_IND.sort(reverse=True) # Reverse order of the empty bins
for n in naN_IND:
# Remove empty bins from the binned a & c matrices
A_CORR_bin = np.delete(A_CORR_bin, n, axis=0) # Delete nans for binned a matrix
C_CORR_bin = np.delete(C_CORR_bin, n, axis=0) # Delete nans for binned c matrix
deptH_BINS = np.delete(deptH_BINS, n, axis=0) # Delete median depths corresponding to empty bins
return(deptH_BINS,A_CORR_bin,C_CORR_bin)
# +
# 3. Define functions needed for pre-processing backscattering (HS6 data)
def bbp_ascii_LOAD(hs6_fileNAME):
"""bbp_ascii_LOAD reads a Hydrolight-compatible (binned) ascii hs6 file and places data
into numpy variables taylored for easy access.
Input variables:
hs6_fileNAME - Hydrolight-compatible ascii file. This file needs to be sigma-corrected PRIOR to use.
Output variables:
bbp_deptH - array of depth bins (median values)
bbp_data - depth-binned backscattering coefficients (sigma-corrected prior to use)
lambdA - wavelength of HS6 channels placed into an array"""
with open(hs6_fileNAME) as hs6_fid: # Open ascii file
bbp_dict = {} # Create empty dictionary for ascii columns
for i,ff in enumerate(hs6_fid):
# This for-loop reads in ascii data and organizes it into a dictionary
if i == 9:
# On the 10th line of the ascii header
fielDS = ff # Create variable for the line
fielDS = fielDS[:-1].split('\t') # Split the line by "tabs" and delete end of line character
elif i == 10:
# On the 11th line of the ascii header
lambdA = ff # Create variable for the 11th line of the header (wavelengths)
lambdA = lambdA.split('\t') # Split the wavelengths by "tabs"
lambdA.pop(0) # Eliminate the first value, which is not a wavelength
lambdA = np.asarray(lambdA,dtype=float) # Make wavelengths a numerical array
elif i > 10:
# On all lines below the wavelengths...
ff = ff.split('\t') # Split data by tabs
for g in fielDS:
# Index fielDS variable and apply the fields to the line of data. Index is used
# so assign bbp data to their correct fields (and dictionary key)
IND = fielDS.index(g) # Find the index of a field
if g in bbp_dict.keys():
# If the dictionary key has already been created
bbp_dict[g].append(ff[IND]) # Add data point to the dictionary key
else:
# If dictionary key is non-existant
bbp_dict[g] = [ff[IND]] # Create dictionary key and add data to it
bbp_deptH = np.asarray(bbp_dict['Depth'],dtype=float) # Convert bbp depths to numpy array
bbp_data = np.ones([len(bbp_deptH),len(lambdA)])*np.nan # Create a nan matrix for bbp data
for i,l in enumerate(lambdA):
# Column by column, for-loop fills in the nan matrix with dictionary data (see below)
bbp_data[:,i] = np.asarray(bbp_dict['bb' + str(int(l))],dtype=float)
# Convert strings to numbers and add to dictionary (see above)
return(bbp_deptH,bbp_data,lambdA)
def bbp_INTERPOLATE(bbp_deptH,bbp_data,bbp_lambdA,acs_lambdA,acs_deptH):
"""bbp_INTERPOLATE interpolates particulate for ac-s wavelenths. This function is a pre-requisite for
McKee et al.'s (2013) scattering correction approach.
Input Variables:
bbp_deptH - HS6 binned depths (medians)
bbp_data - depth-binned particulate backscattering spectra
bbp_lambdA - wavelengths of HS6 channels
acs_lambdA - ac-s channel wavelengths
acs_deptH - ac-s binned depths
Output Variables
bbp_acsINTERPOLATE - depth-binned particular backscattering spectra interpolated to ac-s wavelengths"""
### 1. Define functions within bbp_INTERPOLATE that will come in handy later on
def coeff_CALC(x_1,y_1,x_2,y_2):
"""coeff_CALC calculates slopes and y-intercepts for two ordered pairs (or arrays of
ordered pairs)
Input Variables:
x_1/x_2 - two wavelengths
y_1/y_2 - two backscattering coefficients that correspons to wavelengths
Output Variables:
sloPE - slope of the line (AKA ordered pairs)
y_int - y intercept of the line (AKA ordered pairs) or backscattering coefficient at theoretical wavelength of 0"""
sloPE = (y_2 - y_1)/(x_2 - x_1) # Calculates a slope
y_int = y_2 - (sloPE * x_2) # Calculates a y-intercept
return(sloPE,y_int)
def y_CALC_linear(sloPE,y_int,x_val):
"""y_CALC_linear calculates a y value along a line corresponding to an "x-value".
Input Variables:
sloPE - slope of the line
y_int - y-intercept of the line
x_val - any value of x along the line. y_val will correspond with this value
Output Variables:
y_val - y-value that corresponds to the x-value on the line"""
y_val = sloPE*x_val + y_int # Calculate y value using slope intercept form
return(y_val)
def acs_INDEXER(acs_d,acs_wvl,bbp_lambdA,bbp_deptH):
"""acs_INDEXER finds takes the depth and wavelength of a pair of ac-s absorption and attenuation values finds the index of the corresponding
backscattering coefficient.
Input Variables:
acs_d - depth of the ac-s spectrum in question
acs_wvl - wavelengths of the ac-s spectrum (channels)
bbp_lambdA - wavelengths of the backscattering spectrum
bbp_deptH - depth bins of the backscattering matrix
Output Variables:
wvl_IND - backscattering channel index for a pair of ac-s absorption/attenuation values
deptH_IND - depth index for a pair of ac-s absorption/attenuation values"""
deptH_diFF = np.abs(bbp_deptH - acs_d) # take absolute value of differences between ac-s and bbp depth bins
deptH_IND = np.where(deptH_diFF==min(deptH_diFF))[0][0] # Find depth index of the lowest absolute difference ac-s and backscattering depths
if acs_wvl < bbp_lambdA[0]:
# If ac-s wavelength is outside (<) the spectral range of backscattering coefficients
wvl_IND = 0 # Assign index of 0
elif acs_wvl >= bbp_lambdA[-1]:
# If ac-s wavelength is outside (>) or equal to the highest backscattering wavelength
wvl_IND = len(bbp_lambdA) - 2 # Assign the a specified index
else:
# If ac-s wavelenth is within the range of the backscattering spectrum (ideal scenario)
for i in np.arange(len(bbp_lambdA)-1):
# Search for the correct wavelength with the for-loop, which cycles through hs6
# wavelengths and determines whether the ac-s wavelength is inside.
if bbp_lambdA[i] < acs_wvl <= bbp_lambdA[i+1]:
wvl_IND = i # Assign the wavelength index
break # Break the for-loop
return(wvl_IND,deptH_IND)
### 2. Create matrices for slopes and y intercepts from which ac-s wavelengths will be interpolated
bbp_yINT_Table = np.ones([len(bbp_deptH), len(bbp_lambdA) - 1]) * np.nan # Create a nan matrix to put y intercepts
bbp_sloPE_Table = np.ones([len(bbp_deptH), len(bbp_lambdA) - 1]) * np.nan # Create a nan matrix to put slopes
for i,b in enumerate(bbp_data.transpose()):
# For-loop fills bbp_yINT_Table and bbp_sloPE_Table with slopes and y intercepts. These values differ
# depending on which hs6 wavelengths they fall between
if i == len(bbp_lambdA)-1:
# Indexing is over. Break the loop!
break
else:
# If indexing is NOT over (not the end of the line)
y_2 = b # Second y ordered pair (bbp)
y_1 = bbp_data[:,i+1] # First y ordered pair (bbp)
x_2 = bbp_lambdA[i] # Second x ordered pair (lambda)
x_1 = bbp_lambdA[i+1] # First x ordred pair (lambda)
bbp_sloPE_Table[:,i],bbp_yINT_Table[:,i] = coeff_CALC(x_1,y_1,x_2,y_2)
# Calculate slopes and y intercepts for each hs6 wavelength
### 3. Interpolate bbp for ac-s wavelengths
bbp_acsINTERPOLATE = np.ones([len(bbp_deptH),len(acs_lambdA)]) * np.nan # Create nan matrix for interpolated bbp
for i,d in enumerate(bbp_deptH):
for j,w in enumerate(acs_lambdA):
# This nested for-loop will interpolate bbp for each ac-s depth and wavelength input
# into the bbp_INTERPOLATE function.
wvl_IND, deptH_IND = acs_INDEXER(d,w,bbp_lambdA,bbp_deptH) # Finds index for slope and y-intercept tables
# Calculate interpolated bbp values (below)
bbp_acsINTERPOLATE[i,j] = w * bbp_sloPE_Table[int(deptH_IND),int(wvl_IND)] + bbp_yINT_Table[int(deptH_IND),int(wvl_IND)]
return(bbp_acsINTERPOLATE)
# +
# 4. Define the functions needed for calculating McKee-corrected absorption. This version of
# McKEE_SCATTER has the
def McKEE_SCATTER(a_m,c_m,b_bp,rw):
"""McKEE_SCATTER goes through the iterative process described in McKee et al. (2013). (see flowchart). The function
is an iterative process that accepts a single triplet consisting of an absorption value, attenuation value, and a
backscattering coefficient. THESE VALUES SHOULD ALL CORRESPOND TO THE SAVE WAVELENGTH!
Input variables:
a_m - single absorption value (uncorrected for scatter)
c_m - single attenuation value (uncorrected for scatter)
b_bp - single particulate backscattering value (sigma-corrected)
rw - reflectance efficiency of ac-s absorption tube (coefficients listed in McKee)
Output variables:
an1 - McKee-corrected absorption value
cn1 - McKee-corrected attenuation value
b_p2 - McKee-corrected scatter value"""
### 1. Define functions within a the function
def fa_calc(x):
"""fa_calc will compute f_a variable based on empirical relationship between f_a and
backscattering fraction (x)
Input Variables:
x - backscattering fraction (backscattering coefficient/scatter)
Output Variables:
f_a - fraction of scattered light undetected by absorption meter"""
### 1. Set up empirical coefficients (as constants) for fa equation
coeff0,coeff1,coeff2,coeff3,coeff4,coeff5,coeff6 = McKEE2013_COEFF(r_w=rw)
# Calculate fa based on McKee et al. (2008)'s empirical formula
f_a = coeff0 + coeff1*x + coeff2*(x**2) + coeff3*(x**3) + coeff4*(x**4) + coeff5*(x**5) + coeff6*(x**6)
return(f_a)
def fc_calc(x):
"""fc_calc will compute f_c variable based on empirical relationship between f_c and
backscattering fraction
Input Variables:
x - backscattering fraction (backscattering coefficient/scatter)
Output Variables:
f_c - fraction of scattered light undetected by attenuation meter"""
# Set up empirical coefficients (as constants) for f_c equation. These remain constant no matter what rw
coeff0 = 6.809e-3
coeff1 = 8.502e-3
coeff2 = -1.918e-2
# Calculate f_c based on McKee et al. (2013)'s empirical formula
f_c = coeff0/(coeff1 + x) + coeff2
return(f_c)
def eCOEFFac(x):
"""eCOEFFac calculates error weighing functions for absorption and attenuation from
backscattering fraction (x).
Input Variables:
x - backscattering fraction (backscattering coefficient/scatter)
Output Variables:
Ea - Calculate error weignt for absorption
Ec - alculate error weight for attenuation"""
fa = fa_calc(x) # Calculate fa1
fc = fc_calc(x) # Calculate fc1
Ea = fa/(1 - fc - fa) # Calculate error weignt for absorption
Ec = fc/(1 - fc - fa) # Calculate error weight for attenuation
return(Ea,Ec)
### 2. Before correcting for unmeasured ac-s scatter for iteration, determine starting variables
keY = 0 # provides a failsafe for the upcoming while loop
b_m = c_m - a_m # Calculate scatter from attenuation and absorption
b_p0 = b_m*1.5 # Correct for particulate scatter
b_frac0 = b_bp/b_p0 # Create initial backscattering fraction
fa0 = fa_calc(b_frac0) # Calculate fa0
fc0 = fc_calc(b_frac0) # Calculate fc0
#return(b_m,b_p0,b_frac0,fa0,fc0)
b_p1 = b_m/(1 - fa0 - fc0) # Calculate scatter #1 (b_p1)
b_frac1 = b_bp/b_p1 # Create first backscattering fraction
### 3. Run the iteration as presribed by McKee et al. (2013)
while 1:
Ea1, Ec1 = eCOEFFac(b_frac1) # Calculates error weighing functions
an1 = a_m - (Ea1 * b_m) # Correct absorption
cn1 = c_m + (Ec1 * b_m) # Correct attenuation
b_p2 = cn1 - an1 # Calculate scatter based on "Correct" absorption and attenuation
b_frac2 = b_bp/b_p2 # Calculate "Corrected" backscattering fraction
b_diFF = np.abs(b_frac2 - b_frac1) # Calculate difference between "Corrected" and first iteration fraction
if b_diFF < 0.001 or keY > 9:
if keY == 9:
print('Key')
# If the scattering coefficient was appropriately permutated, or if there were already
# 11 permutations
break # break the while loop!
else:
# If the scattering permutation is insufficient
b_frac1 = b_frac2 # Convert the second backscattering fraction to the first
keY+=1 # Record the iteration by increaing key
return(an1,cn1,b_p2) # Return calculated variables
# +
# 5. Define functions necessary for formatting scatter-corrected ac-s data.
def createFolder(directory):
""" createFolder searches for a dirctory specified by the user. If there is none, it creates one"""
try:
if not os.path.exists(directory): # If the folder doesn't exist
os.makedirs(directory) # Create a folder
except OSError: # If there is an error other than a non-existant folder
print ('Error: Creating directory. ' + directory) # report error and shut down createFolder
def Boostrap_FileWriter(BOOTSRAP_MATRIX,**parameters):
"""Creates a Hydrolight-compatible ascii file using depth-binned IOP data. File is output in a separate folder.
This folder is placed in the same directory as the original file containing non-binned IOP data.
Inputs:
BOOTSRAP_MATRIX - Depth-binned IOP matrix
keyword arguments (in order of appearance):
acs_file - the mat/hdf5 file from which binned data originated
TREAT - the sigma correction used on bbp data
hs6_file - the file containing binned bbp data
station - station number/name
BIN - depth bin size
lambdA - array of ac-s wavelengths
DIR - specify the directory to place the newly-created McKee-corrected ac-s file
Outputs:
Hyrolight-compatible file of binned ac-s data. This output is NOT a variable."""
### 1. Create a dictionary containing all important information for file header and column titles. This information
### is found in key-word arguments.
abs_str = '\ta' # Absorption marker for column headers
atn_str = '\tc' # Absorption marker for column headers
File_acs_new = parameters['acs_file'][:-7] + parameters['TREAT'] + '.txt' # Create name for new ac-s file (final product)
rW = str(parameters['r_w']) # Take r_w (reflectance efficiency) and convert into a string
headER = {}
headER['0'] = 'Total absorption (a) and attenuation (c) measurements\n' # Header line 1 (Data type)
headER['1'] = 'Corrected: ' + dt.now().strftime("%d/%m/%Y %H:%M:%S") + '\n' # Header line 2 (Processing date)
headER['2'] = 'Instrument: ac-s\n' # Header line 3 (Instrument type)
headER['3'] = 'ac-s file name: ' + parameters['acs_file'] + '\n' # Header Line 4 (Name of the original ac-s file)
headER['4'] = 'hs6 file name: ' + parameters['hs6_file'] + '\n' # Header Line 5 (Name of the backscattering coefficients file)
headER['5'] = 'Processed using McKee et al. (2013), r_w = ' + rW + '\n' # Header Line 6 (Name of the scatter correction method)
headER['6'] = 'Station: ' + str(parameters['station']) + '\n' # Header Line 7 (sampling station name)
headER['7'] = 'Bin Size = ' + str(parameters['BIN']) + ' m\n' # Header Line 8 (depth bin size)
headER['8'] = 'Data have been processed using code written and made available by <NAME> (email: <EMAIL>, GitHub: JesseBausell).\n' # Header Line 9 (shameless self promotion)
headER['9'] = 'Depth' # Header line 10. Start of column titles (more to be added later)
headER['10'] = str(len(parameters['lambdA'])) # Header Line 11. Actual number of ac-s channels (more to be added later)
### 2. Develop lines 10 and 11 of the file header.
fieldS_c = ''# Create an empty string for column headers/titles
for l in parameters['lambdA']:
# For-loop cycles through ac-s channels (wavelengths) and creates column headers for absorption and
# attenuation (Header Line 10). It also places wavelengths themselves on line 11 per Hydrolight formatting
# instructions
headER['9'] += abs_str + str(l[0]) # Add 'tab' + 'a' + wavelength to the Header Line 10
fieldS_c += atn_str + str(l[0]) # Add 'tab' + 'c' + wavelength to the segment to be added to Header Line 10
headER['10'] += '\t' + str(l[0]) # Add 'tab' + wavelength to segment to be added to Header Line 11
headER['9'] += fieldS_c + '\n' # Add attenuation column headers and end of line character to Header Line 10
headER['10'] += '\n' # Add end of line character to Header Line 10
### 3. Take absorption and attenuation data and create a Hydrolight-compatible ac-s file
with open(parameters['DIR']+File_acs_new, 'w', newline='') as txtfile: # Create a new ascii file
for i in np.arange(len(headER)):
# for-loop writes the header metadata into the new file
txtfile.write(headER[str(i)]) # Write header line into file
for nn in np.arange(len(BOOTSRAP_MATRIX[:,0])):
# for-loop writes the binned IOP data into the new file
if ~np.isnan(BOOTSRAP_MATRIX[nn,1]):
# if the last line is not reached
datUM = str(BOOTSRAP_MATRIX[nn,:].tolist())[1:-1] # Convert row of data from numpy array into comma-separated string. Remove brackets
datUM = datUM.replace(', ','\t') # replace commas with tabs
txtfile.write(datUM + '\n') # Write the data into the file
keY = nn # this variable preserves last row of actual values e.g. (not nan values)
BOOTSRAP_MATRIX[keY,0] = -1 # After data has been written into the file, replace the first column in the last line (depth) with -1
datUM = str(BOOTSRAP_MATRIX[nn,:].tolist())[1:-1] # Re-convert last row of data to comma-separated string
datUM = datUM.replace(', ','\t') # remove end of line characters
txtfile.write(datUM + '\n') # re-write the last line of code into the file
# -
acs_dir
# +
# 6. Now that all of the libraries are imported and the functions are defined,
### 1. Create starting variables to work with
# Create dictionary of file appendages (to be ID which rW value was used) along with different rW values described in
# McKee et al. (2013)
TREAT = {'_Mc13_1000':1,'_Mc13_0999':0.999,'_Mc13_0998':0.998,'_Mc13_0995':0.995,
'_Mc13_0990':0.99,'_Mc13_0985':0.985,'_Mc13_0980':0.98,'_Mc13_0970':0.97,
'_Mc13_0960':0.96,'_Mc13_0950':0.95}
depTHBIN = 0.5 # User specifies depth-bin.
station_name = 'CST18' # User specifies station
# Select ac-s file to process. Create variables that will be used down the road
acs_fileNAME = fd.askopenfilename() # Select ac-s file to apply McKee et al. (2013) scatter correction
acs_ind = acs_fileNAME.rfind('/') # Find the end of the directory and the beginning of the file name
acs_dir = acs_fileNAME[:acs_ind+1] # Separate directory from file pathway
acs_fileNAME = acs_fileNAME[acs_ind+1:] # Separate file name from file pathway
acs_ind2 = acs_file.rfind('.') # Find the beginning of the filename extension
acs_fileBASE = acs_fileNAME[:acs_ind2] # Create file base. This will be used to develop and create new files
McKee_DIR = acs_dir + '/McKee_2013/' # Create new directory as repository for newly created ac-s files (scatter corrected)
createFolder(McKee_DIR) # Create directory for aforementioned repository (see above line)
# Select binned (Hydrolight-compatible) backscattering coefficient file. If using HS6, this file should ALREADY be sigma-corrected
bbp_fileNAME = fd.askopenfilename() # Select bbp file to use in support of McKee et al.'s (2013) scatter correction approach
### 2. Load, process, and bin uncorrected ac-s data
A_CORR,C_CORR,acs_lambdA,acs_deptH = acs_matlab_LOAD(acs_dir+acs_fileNAME) # Load ac-s .mat/hdf5 file
A_CORR,C_CORR,deptH = QAQC_acsDATA(A_CORR,C_CORR,acs_lambdA,acs_deptH) # Flag and remove problematic ac-s spectra (a>c)
A_CORR,C_CORR,deptH = QAQC_negative(A_CORR,C_CORR,acs_lambdA,deptH) # Flag and remove problematic ac-s spectra (a or c<0)
deptH_BINS_acs,A_CORR_bin,C_CORR_bin = acs_BINNING(A_CORR,C_CORR,acs_lambdA,deptH,biN_size=depTHBIN) # Bin data into user-specified depth-bin
nanIND = [i for i,a in enumerate(A_CORR_bin[:,0]) if not np.isnan(a)] # locate and flag NaNs in binned ac-s data
A_CORR_bin = A_CORR_bin[nanIND,:] # Remove NaNs from binned absorption
C_CORR_bin = C_CORR_bin[nanIND,:] # Remove NaNs from binned attenuation
deptH_BINS_acs = deptH_BINS_acs[nanIND] # Remove NaNs from depth bins
### 3. Load and interpolate backscattering data to be consistent with use for
bbp_deptH,bbp_data,bbp_lambdA = bbp_ascii_LOAD(bbp_fileNAME) # Read in the HE53-formatted ascii file
bbp_acsINTERPOLATE = bbp_INTERPOLATE(bbp_deptH,bbp_data,bbp_lambdA,acs_lambdA,deptH_BINS_acs)
bbp_ind = bbp_fileNAME.rfind('/') # Find the end of the directory and the beginning of the file name
bbp_fileNAME = bbp_fileNAME[bbp_ind+1:] # Separate the file name from the directory
### 4. Apply McKee et al. (2013) Scatter Correction Approach to binned ac-s data
for t in TREAT.keys():
# for-loop goes through each of the 10 rw's, as laid out in McKee et al. (2013). For each rw, the loop applies
# McKee et al.'s (2013) approach, and then formats scatter-corrected ac-s data into a Hydrolight-compatible ascii file
# hs6_diR = acs_diR + t + '/' # create folder pathway for McKee et al. (2013) SCA
# createFolder(hs6_diR) # create folder for McKee et al. (2013 SCA) if not in existence
# copy_tree(acs_diR + '_Mc08/', hs6_diR) # Copy all contents from McKee_tools into new folder
# hs6_fileNAME = STN[s] + '_bbp_bin0.5_Mc08.txt' # Select directory containing m-files
# bbp_deptH,bbp_data,bbp_lambdA = bbp_ascii_LOAD(bbp_fileNAME) # Read in the HE53-formatted ascii file
# ### Interpolate bbp for ac-s wavelengths
# bbp_acsINTERPOLATE = bbp_INTERPOLATE(bbp_deptH,bbp_data,bbp_lambdA,acs_lambdA,deptH_BINS_acs)
## 4a. Create empty matrices to act as repositories for finished products and run nested for-loop!
A_CORR_McKee = np.ones(A_CORR_bin.shape)*np.nan # empty nan matrix for McKee absorption
C_CORR_McKee = np.ones(C_CORR_bin.shape)*np.nan # empty nan matrix for McKee attenuation
B_CORR_McKee = np.ones(C_CORR_bin.shape)*np.nan # empty nan matrix for McKee scatter
for acs_IND,d_acs in enumerate(deptH_BINS_acs):
# for-loop cycles through ac-s spectra, selecting them one at a time for McKee correction. Because the for-loop
# is nested, an additional for-loop takes absorption, attenuation, and backscattering coefficients from each
# spectrum one-by-one, and applies McKee et al. (2013) to them.
## 4b. Select the appropriate backscattering spectrum (same or similar depth) for selected depth-binned ac-s spectra
difF_acs = np.abs(bbp_deptH - d_acs) # Find absolute differences between backscattering depth bins and ac-s spectra
bbp_IND = np.where(difF_acs==min(difF_acs))[0][0] # Find index of the smallest absolute difference (AKA closest binned depth)
a_array = A_CORR_bin[acs_IND,:] # Select mean absorption spectrum at the appropriate depth
c_array = C_CORR_bin[acs_IND,:] # Select mean attenuation spectrum at the appropriate depth
bbp_array = bbp_acsINTERPOLATE[bbp_IND,:] # Select interpolated bbp spectrum based on the appropriate depth
for i,a_m in enumerate(a_array):
### 4c. This for-loop goes through the selected absorption, attenuation, and interpolated
# backscattering spectra channel by channel. It takes each uncorrected absorption, and its
# corresponding attenuation and backscattering coefficient and calculates McKee et al.
# (2013) scatter channel by channel.
c_m = c_array[i] # Find uncorrected attenuation based on absorption index
b_bp = bbp_array[i] # Find backscattering coefficient based on absorption index
an1,cn1,bn1 = McKEE_SCATTER(a_m,c_m,b_bp,TREAT[t]) # Correct for absorption, attenuation, and scatter
A_CORR_McKee[acs_IND,i] = an1 # Place absorption in the correct matrix
C_CORR_McKee[acs_IND,i] = cn1 # Place attenuation in the correct matrix
B_CORR_McKee[acs_IND,i] = bn1 # Place scatter in the correct matrix
IOP_MATRIX = np.round(np.append(A_CORR_McKee,C_CORR_McKee,axis=1),decimals=6) # Round answers to three decimal places
IOP_MATRIX = np.append(np.reshape(deptH_BINS_acs,[len(deptH_BINS_acs),1]),IOP_MATRIX,axis=1) # Attach binned depths to lefthand side
Boostrap_FileWriter(IOP_MATRIX,DIR=McKee_DIR,acs_file=acs_fileNAME,hs6_file=bbp_fileNAME,station=station_name,lambdA=acs_lambdA,BIN=depTHBIN,r_w=TREAT[t],TREAT =t)
# batch_FILE = 'I' + s + '_bin_05_PCT100' + t + '.txt'
# batch_FILE = 'I' + s + '_bin_05_PCT100' + '.txt' # Generic batch file name. This will be used to create proper batch file
# HE53_batch_WRITER(batch_diR+batch_FILE,STN=s,Station=STN[s],TRMT=t,TREATMENT='_bbp_bin0.5_Mc08.txt') #
| McKee_Scattering_2013.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # in some plots we are having 1 SOS with no EOS.
#
# Why nullify function did not work?
# +
# import warnings
# warnings.filterwarnings("ignore")
import csv
import numpy as np
import pandas as pd
# import geopandas as gpd
from IPython.display import Image
# from shapely.geometry import Point, Polygon
from math import factorial
import scipy
import scipy.signal
import os, os.path
from datetime import date
import datetime
import time
from statsmodels.sandbox.regression.predstd import wls_prediction_std
from sklearn.linear_model import LinearRegression
from patsy import cr
# from pprint import pprint
import matplotlib.pyplot as plt
import seaborn as sb
import sys
# +
sys.path.append('/Users/hn/Documents/00_GitHub/Ag/remote_sensing/python/')
import remote_sensing_core as rc
import remote_sensing_plot_core as rcp
data_dir = "/Users/hn/Documents/01_research_data/remote_sensing/test_Aeolus_data/"
# +
eleven_colors = ["gray", "lightcoral", "red", "peru",
"darkorange", "gold", "olive", "green",
"blue", "violet", "deepskyblue"]
indeks = "EVI"
given_county = "Grant"
SF_year = 2017
sos_thresh = 0.5
eos_thresh = 0.5
minFinderDetla = 0.4
f_name = "01_Regular_filledGap_Grant_SF_2017_EVI.csv"
a_df = pd.read_csv(data_dir + f_name, low_memory=False)
if 'Date' in a_df.columns:
if type(a_df.Date.iloc[0]) == str:
a_df['Date'] = pd.to_datetime(a_df.Date.values).values
a_df = a_df[a_df['county'] == given_county.replace("_", " ")] # Filter Grant
# a_df = rc.filter_out_NASS(a_df) # Toss NASS
# a_df = rc.filter_by_lastSurvey(a_df, year = SF_year) # filter by last survey date
a_df['SF_year'] = SF_year
if not('DataSrc' in a_df.columns):
print ("Data source is being set to NA")
a_df['DataSrc'] = "NA"
if not('CovrCrp' in a_df.columns):
print ("CovrCrp is being set to NA")
a_df['CovrCrp'] = "NA"
print (a_df.shape)
# +
print (a_df.shape)
a_df = rc.initial_clean(df = a_df, column_to_be_cleaned = indeks)
an_EE_TS = a_df.copy()
print (an_EE_TS.shape)
### List of unique polygons
polygon_list = np.sort(an_EE_TS['ID'].unique())
print ("_____________________________________")
print("len(polygon_list)")
print (len(polygon_list))
print ("_____________________________________")
counter = 0
# +
a_poly = "100106_WSDA_SF_2017"
a_poly in (polygon_list)
# +
if (counter%10 == 0):
print ("_____________________________________")
print ("counter: " + str(counter))
print (a_poly)
curr_field = an_EE_TS[an_EE_TS['ID']==a_poly].copy()
#
# filter just one year to have a clean SOS EOS stuff
#
curr_field = curr_field[curr_field.image_year == SF_year]
################################################################
# Sort by DoY (sanitary check)
curr_field.sort_values(by=['image_year', 'doy'], inplace=True)
# +
fig, axs = plt.subplots(1, 1, figsize=(10,6),
sharex='col', sharey='row',
gridspec_kw={'hspace': 0.1, 'wspace': .1});
(ax1) = axs;
ax1.grid(True); # ax2.grid(True);
dataAB = curr_field
idx=indeks
SG_params=[7, 3]
SFYr = SF_year
ax=ax1
deltA= minFinderDetla
onset_cut = sos_thresh
offset_cut = eos_thresh
rcp.SG_1yr_panels_clean_sciPy_My_Peaks_SOS_fineGranularity_1Year(dataAB = curr_field,
idx=indeks,
SG_params=SG_params,
SFYr = SF_year, ax=ax1, deltA= minFinderDetla,
onset_cut = sos_thresh,
offset_cut = eos_thresh);
fig_name = "/Users/hn/Documents/00_GitHub/Ag/remote_sensing/" + \
"python/Local_Jupyter_NoteBooks/scratches_to_experiment/1SOS.png"
# plt.savefig(fname = fig_name, dpi=400, bbox_inches='tight')
# -
dataAB = curr_field
idx = indeks
SG_params=[5, 1]
SFYr = SF_year
ax = ax1
deltA = minFinderDetla
onset_cut = sos_thresh
offset_cut = eos_thresh
# +
crr_fld = dataAB.copy()
if (not("human_system_start_time" in list(crr_fld.columns))):
crr_fld = rc.add_human_start_time(crr_fld)
eleven_colors = ["gray", "lightcoral", "red", "peru",
"darkorange", "gold", "olive", "green",
"blue", "violet", "deepskyblue"]
plant = crr_fld['CropTyp'].unique()[0]
# Take care of names, replace "/" and "," and " " by "_"
plant = plant.replace("/", "_")
plant = plant.replace(",", "_")
plant = plant.replace(" ", "_")
plant = plant.replace("__", "_")
county = crr_fld['county'].unique()[0]
ID = crr_fld['ID'].unique()[0]
y = crr_fld[idx].copy()
#############################################
###
### Smoothen
###
#############################################
# differences are minor, but lets keep using Pythons function
# my_savitzky_pred = rc.savitzky_golay(y, window_size=Sav_win_size, order=sav_order)
window_len = SG_params[0]
poly_order = SG_params[1]
SG_pred = scipy.signal.savgol_filter(y, window_length= window_len, polyorder=poly_order)
# SG might violate the boundaries. clip them:
SG_pred[SG_pred > 1 ] = 1
SG_pred[SG_pred < -1 ] = -1
crr_fld[idx] = SG_pred
#############################################
###
### fine granularity table
###
#############################################
# create the full calenadr to make better estimation of SOS and EOS.
fine_granular_table = rc.create_calendar_table(SF_year = SFYr)
fine_granular_table = pd.merge(fine_granular_table, crr_fld, on=['Date', 'SF_year', 'doy'], how='left')
###### We need to fill the NAs that are created because they were not created in fine_granular_table
fine_granular_table["image_year"] = crr_fld["image_year"].unique()[0]
fine_granular_table["ID"] = crr_fld["ID"].unique()[0]
fine_granular_table["Acres"] = crr_fld["Acres"].unique()[0]
fine_granular_table["county"] = crr_fld["county"].unique()[0]
fine_granular_table["CropGrp"] = crr_fld["CropGrp"].unique()[0]
fine_granular_table["CropTyp"] = crr_fld["CropTyp"].unique()[0]
fine_granular_table["DataSrc"] = crr_fld["DataSrc"].unique()[0]
fine_granular_table["ExctAcr"] = crr_fld["ExctAcr"].unique()[0]
fine_granular_table["IntlSrD"] = crr_fld["IntlSrD"].unique()[0]
fine_granular_table["Irrigtn"] = crr_fld["Irrigtn"].unique()[0]
fine_granular_table["LstSrvD"] = crr_fld["LstSrvD"].unique()[0]
fine_granular_table["Notes"] = crr_fld["Notes"].unique()[0]
fine_granular_table["RtCrpTy"] = crr_fld["RtCrpTy"].unique()[0]
fine_granular_table["Shap_Ar"] = crr_fld["Shap_Ar"].unique()[0]
fine_granular_table["Shp_Lng"] = crr_fld["Shp_Lng"].unique()[0]
fine_granular_table["TRS"] = crr_fld["TRS"].unique()[0]
fine_granular_table = rc.add_human_start_time_by_YearDoY(fine_granular_table)
# replace NAs with -1.5. Because, that is what the function fill_theGap_linearLine()
# uses as indicator for missing values
fine_granular_table.fillna(value={idx:-1.5}, inplace=True)
fine_granular_table = rc.fill_theGap_linearLine(regular_TS = fine_granular_table,
V_idx=idx,
SF_year=SFYr)
# update SG_pred so that we do not have to update too many other stuff.
SG_pred = fine_granular_table[idx].values.copy()
crr_fld = fine_granular_table
y = fine_granular_table[idx].copy()
# +
#############################################
###
### Form a data table of X and Y values
###
#############################################
if len(fine_granular_table['image_year'].unique()) == 2:
X = rc.extract_XValues_of_2Yrs_TS(fine_granular_table, SF_yr = SFYr)
elif len(fine_granular_table['image_year'].unique()) == 1:
X = fine_granular_table['doy']
d = {'DoY': X, 'Date': pd.to_datetime(fine_granular_table.human_system_start_time.values).values}
date_df = pd.DataFrame(data=d)
min_val_for_being_peak = 0.5
# -
crr_fld [idx] = SG_pred
crr_fld = rc.addToDF_SOS_EOS_White(pd_TS = crr_fld,
VegIdx = idx,
onset_thresh = onset_cut,
offset_thresh = offset_cut)
###
### Null_SOS_EOS_by_DoYDiff(pd_TS, min_season_length=40)
###
pd_TS = crr_fld.copy()
min_season_length=40
# +
pd_TS_DoYDiff = pd_TS.copy()
# find indexes of SOS and EOS
SOS_indexes = pd_TS_DoYDiff.index[pd_TS_DoYDiff['SOS'] != 0].tolist()
EOS_indexes = pd_TS_DoYDiff.index[pd_TS_DoYDiff['EOS'] != 0].tolist()
# -
SOS_indexes
EOS_indexes
| remote_sensing/python/Local_Jupyter_NoteBooks/scratches_to_experiment/null_SOS_failed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 17: Clustering (Unsupervised Classification)
# #### This notebook was developed by [<NAME>](http://faculty.washington.edu/ivezic/) for the 2021 data science class at the University of Sao Paulo and it is available from [github](https://github.com/ivezic/SaoPaulo2021/blob/main/notebooks/Lecture17.ipynb).
#
# Note: this notebook contains code developed by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME> and many others.
# ##### Resources for this notebook include:
# - [Textbook](http://press.princeton.edu/titles/10159.html) Chapters 6 and 9.
# <a id='toc'></a>
#
# ## This notebook includes:
#
#
# [Introduction to Clustering](#intro)
# - unsupervised vs. supervised classification
# - 1-D hypothesis testing
# - clustering with Gaussian Mixture models (GMM)
# - K-means clustering algorithm
# - hierarchical clustering algorithm
#
# [Discussion of Term Project](#project)
#
#
# ## Introduction to Clustering <a id='basics'></a>
# [Go to top](#toc)
# ### Clustering
#
# “Clustering” in astronomy refers to a number of different aspects of data analysis. Given a multivariate point data set, we can ask whether it displays any structure, that is, concentrations of points. Alternatively, when a density estimate is available we can search for “overdensities”. Another way to interpret clustering is to seek a partitioning or segmentation of data into smaller parts according to some criteria.
#
# Recall that in Activity 8 yesterday we had a simple 1-D example of
# using Gaussian Mixture Model and BIC to study the impact of sample size and measurement errors on ability to recognize structure in data. We were doing clustering even without knowing it!
#
# Here is an illustration of clustering in 2-D space:
#
# 
# ### Unsupervised vs. Supervised Classification
#
# In density estimation, we estimate joint probability distributions from multivariate data sets to identify the inherent clustering. This is essentially **unsupervised classification**. Here “unsupervised” means that there is no prior information about the number and properties of clusters.
# In other words, this method is search for unknown structure in your (multi-dimensional) dataset.
#
# If we have labels for some of these data points (e.g., an object is tall, short, red, or blue), we can develop a relationship between the label and the properties of a source. This is **supervised classification**. In other words, this method is finding objects in
# your (multi-dimensional) dataset that "look like" objects in your training set.
#
# Classification, regression, and density estimation are all related. For example, the regression function $\hat{y} = f(y|\vec{x})$ is the best estimated value of $y$ given a value of $\vec{x}$. In classification $y$ is categorical and $f(y|\vec{x})$ is called the _discriminant function_
#
# ## 1-D hypothesis testing <a id='1Dht'></a>
# [Go to top](#toc)
#
# How do we decide about the existance of a cluster? Let's start with
# the simplest but fundamental example: 1-D hypothesis testing.
#
#
# **Motivating question:** You just measured x = 3, with a negligible measurement error.
#
# You know that you could have drawn this value from one of two possible populations (e.g. stars and galaxies). One population can be described as N(0,2), and the other one as N(4,1).
#
# Which population is more likely, given your x?
#
# Naive (wrong) answer: 3 is closer to 4 (1 "sigma away") than to 0
# (1.5 "sigma away") so the second population is more likely.
#
# Let's see why this answer is wrong...
#
# 
# 
# 
# 
# ## Clustering with Gaussian Mixture models (GMM)
#
# We already addressed Gaussian Mixture models in Lecture 15 about Density Estimation and in Activity 8 (1-D example).
#
# We will see two more illustrative multi-dimensional examples later today in Activity 9.
# ## K-means clustering algorithm
#
# 
# 
# ## Hierarchical clustering algorithm
#
#
# 
# 
#
#
# ### We will apply this method to a real dataset later today in Activity 9.
#
#
# ## Discussion of Term Project <a id='project'></a>
# [Go to top](#toc)
#
#
# As you probably already know, your grade in this class will include a term project component. You will have about four weeks
# to apply your knowledge to a real dataset and thus demonstrate your mastery of the subject.
#
# ** The submission deadline is Aug 28, midnight Sao Paulo time. **
#
# Our last day of instruction will be Monday, Aug 23, and you will able to ask me last-minute questions. In addition, I will be available for discussion of your term project (and other class material) on Aug 20 and Aug 24 (9am Sao Paulo time).
#
# ### What do you need to do?
#
# There are only two requirements for your project:
# - it needs to be based on S-PLUS data
# - it needs to apply at least one of the methods we covered in these lectures (regression, density estimation, dimensionality reduction, clustering, classification)
#
# Each submitted notebook should start with a description of what you did (what question did you ask, how did you select your data, which
# algorithm you used and why, and what you concluded). Your code needs
# to be runnable (after receiving submission, I will run your notebook
# on my machine; please do not introduce new dependencies without a strong reason and if so, please emphasize it at the start of the notebook).
#
# Please email me (<EMAIL>) **link** to your notebook by Aug 28. Preferred method
# is to use your GitHub account, but other means (such as Dropbox) will
# be fine, too.
#
# You can find out **how to access S-PLUS data**, including examples of queries, at [splus.cloud](https://deepnote.com/project/S-PLUS-Meeting-1-3-June-2021-i5mav_NUQgO148fIhCC31A/%2FTutorial.ipynb)
#
| lectures/notes/Lecture7-density-estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `kickscore` basics
#
# This notebook introduces the library's API in the context of a simple example involving [Tom and Jerry](https://en.wikipedia.org/wiki/Tom_and_Jerry).
#
# 
#
# In this notebook, we will:
#
# 1. instantiate a `kickscore` model,
# 2. define items & express our prior beliefs on the temporal dynamics of the items' skill,
# 3. fit the model to a toy dataset, and
# 4. analyze the results.
#
# If you are not doing so already, you can [launch an interactive notebook on Google Colaboratory](https://colab.research.google.com/github/lucasmaystre/kickscore/blob/master/examples/kickscore-basics.ipynb)
# This is only needed for when running the notebook on Google Colaboratory.
# !pip install kickscore
import kickscore as ks
# %matplotlib inline
# ## Defining & fitting the model
#
# First, we define the model. In this example, we will use a _binary_ model, which is suitable for observations of the type "_A_ wins over _B_".
# Richer types of observations can be handled with `TernaryModel`, `CountModel` and `DifferenceModel`.
model = ks.BinaryModel()
# We will observe outcomes of "fights" between three contestants: Tom (the cat), Jerry (the mouse) and Spike (the dog).
# Each contestant is an _item_, in `kickscore`'s terminology.
#
# The key assumption behind the `kickscore` models is that the outcome probability of each fight is a function of the (latent) skill of each contestant.
# This skill can **change over time**.
# For each contestant, we need to describe how we expect their skill to change over time.
# We do this by specifying a **kernel function** (a.k.a. covariance function).
#
# - Many kernel functions have a variance parameter (usually called `var`). Roughly speaking, this defines the range of possible skill values.
# - Some kernel functions have a lengthscale parameter (usually called `lscale`). This roughly defines the time interval over which we expect the skill to stay similar.
# +
# Spike's skill does not change over time.
k_spike = ks.kernel.Constant(var=0.5)
# Tom's skill changes over time, with "jagged" (non-smooth) dynamics.
k_tom = ks.kernel.Exponential(var=1.0, lscale=1.0)
# Jerry's skill has a constant offset and smooth dynamics.
k_jerry = ks.kernel.Constant(var=1.0) + ks.kernel.Matern52(var=0.5, lscale=1.0)
# Now we are ready to add the items in the model.
model.add_item("Spike", kernel=k_spike)
model.add_item("Tom", kernel=k_tom)
model.add_item("Jerry", kernel=k_jerry)
# -
# Next, we add observations to the model. **Note that observations must be added in chronological order.** The `observe` method takes:
#
# - a list containing the items that _won_ the comparison (`winners`)
# - a list containing the items that _lost_ the comparison (`losers`)
# - the time of the observation (`t`)
#
# When `winners` or `losers` contains more than one item, `kickscore` considers the _sum of the corresponding items' skill_.
# +
# At first, Jerry beats Tom a couple of times.
model.observe(winners=["Jerry"], losers=["Tom"], t=0.0)
model.observe(winners=["Jerry"], losers=["Tom"], t=0.9)
# Then, Tom beats Spike, and then Jerry.
model.observe(winners=["Tom"], losers=["Spike"], t=1.7)
model.observe(winners=["Tom"], losers=["Jerry"], t=2.1)
# Finally, Jerry beats Tom, and then Tom + Spike.
model.observe(winners=["Jerry"], losers=["Tom"], t=3.0)
model.observe(winners=["Jerry"], losers=["Tom", "Spike"], t=3.5)
# -
# Now we are ready to fit the model (i.e., infer the skill parameters over time).
model.fit(verbose=True)
# ## Analyzing the results
#
# We can get a good understanding of what the model learned by plotting the skill as a function of time.
model.plot_scores(["Tom", "Jerry", "Spike"], figsize=(14, 5));
# As `kickscore` learns a posterior _distribution_ over the skills, we represent the mean skill (solid line) as well as the region corresponding to one standard deviation (the shaded area).
#
# Notice the following:
#
# - the skill of Tom is not smooth over time. Notice how the bends happen at moments corresponding to observations involving Tom.
# - in contrast, the skill of Jerry varies smoothly.
# - The skill of Spike is constant, as expected.
# We can also use the model to make predictions for any combination of items, at any time point.
# +
# We can predict a future outcome...
p_win, p_los = model.probabilities(["Jerry"], ["Tom"], t=4.0)
print("Chances that Jerry beats Tom at t = 4.0: {:.1f}%".format(100*p_win))
# ... or simulate what could have happened in the past.
p_win, p_los = model.probabilities(["Jerry"], ["Tom"], t=2.0)
print("Chances that Jerry beats Tom at t = 2.0: {:.1f}%".format(100*p_win))
p_win, p_los = model.probabilities(["Jerry"], ["Tom"], t=-1.0)
print("Chances that Jerry beats Tom at t = -1.0: {:.1f}%".format(100*p_win))
# -
# Finally, note that we had to make quite a few choices when we set up the model.
# In particular, we had to explicitly choose the kernel functions and their (hyper)paramaters.
#
# A principled way to compare various models and to eventually select the best one is to compare the log-marginal likelihood of the data (the higher, the better).
model.log_likelihood
# You can go back and change the kernels & their parameters. Can you find a better model than the one we picked?
| examples/kickscore-basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recommender system
#
# Two types of Recommender system
# 1) content based recommender sytem
# 2) collaborative recommender system
#
# # Import Libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # Get the data
col_names = ['user_id', 'item_id', 'rating', 'timestamp']
df = pd.read_csv('Desktop/u.data', sep='\t', names=col_names)
df.head()
df.columns
df.info()
# Now let's get the movie titles
#
movie_titles = pd.read_csv("Desktop/Movie_Id_Titles")
movie_titles.head()
# merge them together (df+movie _titles)
#
df=pd.merge(df,movie_titles)
df.head()
# Let's explore the data and look at most rated movies
import seaborn as sns
# Let's create a ratings dataframe with average rating and number of ratings:
df.groupby('title')['rating'].mean().sort_values(ascending=False).head()
df.groupby('title')['rating'].count().sort_values(ascending=False).head()
df.groupby('title')['rating'].count().sort_values(ascending=False).head()
ratings = pd.DataFrame(df.groupby('title')['rating'].mean())
ratings.head()
ratings['num of ratings'] = pd.DataFrame(df.groupby('title')['rating'].count())
ratings.head()
plt.figure(figsize=(10,4))
ratings['num of ratings'].hist(bins=70)
plt.xlabel('number of ratings')
sns.jointplot(x='rating',y='num of ratings',data=ratings,alpha=0.5)
# # Recommending similar movies
# create a matrix that has the user ids on one axis and the movie title on another axis. Each cell will then consist of the rating the user gave to that movie. Note there will be a lot of NaN values, because most people have not seen most of the movies.
#
#
movie_sim = df.pivot_table(index='user_id',columns='title',values='rating')
movie_sim
ratings.sort_values('num of ratings',ascending=False).head(10)
starwars_user_ratings = movie_sim['Star Wars (1977)']
contact_user_ratings = movie_sim['Contact (1997)']
starwars_user_ratings
contact_user_ratings
# We can then use corrwith() method to get correlations between two pandas series:
similar_to_starwars = movie_sim.corrwith(starwars_user_ratings)
similar_to_contact = movie_sim.corrwith(contact_user_ratings)
# Let's clean this by removing NaN values and using a DataFrame instead of a series:
corr_starwars = pd.DataFrame(similar_to_starwars,columns=['Correlation'])
corr_starwars.dropna(inplace=True)
corr_starwars
corr_contact = pd.DataFrame(similar_to_contact,columns=['Correlation'])
corr_contact.dropna(inplace=True)
corr_contact
corr_starwars.sort_values('Correlation',ascending=False).head(10)
corr_contact.sort_values('Correlation',ascending=False).head(10)
# Let's fix this by filtering out movies that have less than 100 reviews (this value was chosen based off the histogram from earlier).
corr_starwars = corr_starwars.join(ratings['num of ratings'])
corr_starwars.head()
# Now sort the values and notice how the titles make a lot more sense:
# +
corr_contact = corr_contact.join(ratings['num of ratings'])
corr_starwars[corr_starwars['num of ratings']>100].sort_values('Correlation',ascending=False).head()
# -
corr_contact = pd.DataFrame(similar_to_contact,columns=['Correlation'])
corr_contact.dropna(inplace=True)
corr_contact = corr_contact.join(ratings['num of ratings'])
corr_contact[corr_contact['num of ratings']>100].sort_values('Correlation',ascending=False).head()
| recommender system d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''census'': conda)'
# language: python
# name: python37664bitcensusconda6de06cb7d1124a39a673c8ac5ed43be4
# ---
# # Point Patterns
import numpy as np
from pysal.explore.pointpats import PointPattern
import libpysal as ps
# %matplotlib inline
import matplotlib.pyplot as plt
import geopandas as gpd
file = "https://opendata.arcgis.com/datasets/e295d0fe5def4102b42bdab776fe67d0_0.zip"
df = gpd.read_file(file)
df.head()
pp = PointPattern(points) #create a point pattern "pp" from list
pp.points
from pysal.explore.pointpats.centrography import hull, mbr, mean_center, weighted_mean_center, manhattan_median, std_distance,euclidean_median,ellipse
mc = mean_center(df.geometry)
mc
| notebooks/Point Pattern Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p style="font-family: Arial; font-size:3.50em;color:purple; font-style:bold">
# Course Prerequisites Structure Using Sankey
# </p>
# <br>
#
# Here, I am going to show flow from course to its prerequisites.
# <br>
# Notice that, I have made 1 change in File. I have filled "Passrate" with values which will be work as **value** for sankey diagram.
import plotly
import pandas as pd
import numpy as np
from plotly.graph_objs import Sankey
import random
data = pd.read_csv('FDU Electrical Engineering.csv', sep =",")
print(type(data))
print("Shape will show number of rows and columns in DataFrame (rows,columns):",data.shape)
data.head()
# ## Creating mapping for source and target.
# * In our case, column "Name" will work as source
# * column "Prerequisites" will work as target
# * and column "Passrate" will work as value
# <br>
#
# ** I have already explained process of mapping that is created for source and target with replacing those mapping in our DataFrame.**
# <br>
# #### What would be different here ?
#
# 1. Here I am going to replace "NaN" with value 0 (which will act as very first node.)
# 2. I will assign name to that node (which will be very first value in labels list.)
# 3. The reason behind that is to show all the subjects from each term. In sankey we can not show nodes without target and values. It is must to have target and value. (If value would be 0 then that connection won't be shown at all.)
source_name = data['Name'].tolist()
print(len(source_name))
source_name[:6]
source_zip = list(range(1,45))
source_mapping = dict(zip(source_name, source_zip))
source_mapping
# ### Creating copy of "data" DataFrame to perform mapping
df = data.copy()
df['Name'].replace(source_mapping, inplace = True)
df[-10:]
pre_requisites = data['Prerequisites'].dropna().unique().tolist()
print(len(pre_requisites))
pre_requisites[:5]
target_zip = list(range(45,66))
target_mapping = dict(zip(pre_requisites, target_zip))
target_mapping
# ### Replacing "NaN" with 0
df.replace(np.nan, 0, inplace=True)
df['Prerequisites'].replace(target_mapping, inplace = True)
df[:5]
# ### Creating labels and giving name to very first node which will represent all NaN values.
labels = ['No Pre_req']
labels = labels + source_name + pre_requisites
# # Creating sankey
# +
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
from plotly.graph_objs import *
init_notebook_mode(connected=True)
# +
data_trace = dict(
type='sankey',
domain = dict(
x = [0,1],
y = [0,1]
),
orientation = "h",
visible = True,
valueformat = ".0f",
valuesuffix = " Students",
node = dict(
pad = 15,
thickness = 30,
line = dict(
color = "black",
width = 0.5
),
label = labels,
#color = df['node_color']
),
link = dict(
source = df['Name'],
target = df['Prerequisites'],
value = df['Passrate'],
#color = df['link_color'],# Here we can add colors for each link which connected source to target.
)
)
layout = dict(
title = "Course Prerequisite Structure For BS Electrical Engineering",
height = 1000,
width = 1000,
font = dict(
size = 12
),
)
# -
fig = Figure(data=[data_trace], layout=layout)
plotly.offline.plot(fig, validate=False)
| Course Pre_req Flow for BS Electrical Engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Question Answering on a Knowledge Graph
#
# [](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial10_Knowledge_Graph.ipynb)
#
# Haystack allows storing and querying knowledge graphs with the help of pre-trained models that translate text queries to SPARQL queries.
# This tutorial demonstrates how to load an existing knowledge graph into haystack, load a pre-trained retriever, and execute text queries on the knowledge graph.
# The training of models that translate text queries into SPARQL queries is currently not supported.
# + pycharm={"name": "#%%\n"}
# Install the latest release of Haystack in your own environment
# #! pip install farm-haystack
# Install the latest master of Haystack
# !pip install --upgrade pip
# !pip install git+https://github.com/deepset-ai/haystack.git#egg=farm-haystack[colab,graphdb]
# + pycharm={"name": "#%%\n"}
# Here are some imports that we'll need
import subprocess
import time
from pathlib import Path
from haystack.nodes import Text2SparqlRetriever
from haystack.document_stores import GraphDBKnowledgeGraph
from haystack.utils import fetch_archive_from_http
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Downloading Knowledge Graph and Model
# + pycharm={"name": "#%%\n"}
# Let's first fetch some triples that we want to store in our knowledge graph
# Here: exemplary triples from the wizarding world
graph_dir = "../data/tutorial10_knowledge_graph/"
s3_url = "https://fandom-qa.s3-eu-west-1.amazonaws.com/triples_and_config.zip"
fetch_archive_from_http(url=s3_url, output_dir=graph_dir)
# Fetch a pre-trained BART model that translates text queries to SPARQL queries
model_dir = "../saved_models/tutorial10_knowledge_graph/"
s3_url = "https://fandom-qa.s3-eu-west-1.amazonaws.com/saved_models/hp_v3.4.zip"
fetch_archive_from_http(url=s3_url, output_dir=model_dir)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Launching a GraphDB instance
# + pycharm={"name": "#%%\n"}
# Unfortunately, there seems to be no good way to run GraphDB in colab environments
# In your local environment, you could start a GraphDB server with docker
# Feel free to check GraphDB's website for the free version https://www.ontotext.com/products/graphdb/graphdb-free/
print("Starting GraphDB ...")
status = subprocess.run(
[
"docker run -d -p 7200:7200 --name graphdb-instance-tutorial docker-registry.ontotext.com/graphdb-free:9.4.1-adoptopenjdk11"
],
shell=True,
)
if status.returncode:
raise Exception(
"Failed to launch GraphDB. Maybe it is already running or you already have a container with that name that you could start?"
)
time.sleep(5)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Creating a new GraphDB repository (also known as index in haystack's document stores)
# + pycharm={"name": "#%%\n"}
# Initialize a knowledge graph connected to GraphDB and use "tutorial_10_index" as the name of the index
kg = GraphDBKnowledgeGraph(index="tutorial_10_index")
# Delete the index as it might have been already created in previous runs
kg.delete_index()
# Create the index based on a configuration file
kg.create_index(config_path=Path(graph_dir + "repo-config.ttl"))
# Import triples of subject, predicate, and object statements from a ttl file
kg.import_from_ttl_file(index="tutorial_10_index", path=Path(graph_dir + "triples.ttl"))
print(f"The last triple stored in the knowledge graph is: {kg.get_all_triples()[-1]}")
print(f"There are {len(kg.get_all_triples())} triples stored in the knowledge graph.")
# + pycharm={"name": "#%%\n"}
# Define prefixes for names of resources so that we can use shorter resource names in queries
prefixes = """PREFIX rdf: <http://www.w3.org/1999/02/22-rdf-syntax-ns#>
PREFIX xsd: <http://www.w3.org/2001/XMLSchema#>
PREFIX hp: <https://deepset.ai/harry_potter/>
"""
kg.prefixes = prefixes
# Load a pre-trained model that translates text queries to SPARQL queries
kgqa_retriever = Text2SparqlRetriever(knowledge_graph=kg, model_name_or_path=model_dir + "hp_v3.4")
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Query Execution
#
# We can now ask questions that will be answered by our knowledge graph!
# One limitation though: our pre-trained model can only generate questions about resources it has seen during training.
# Otherwise, it cannot translate the name of the resource to the identifier used in the knowledge graph.
# E.g. "Harry" -> "hp:Harry_potter"
# + pycharm={"name": "#%%\n"}
query = "In which house is <NAME>?"
print(f'Translating the text query "{query}" to a SPARQL query and executing it on the knowledge graph...')
result = kgqa_retriever.retrieve(query=query)
print(result)
# Correct SPARQL query: select ?a { hp:Harry_potter hp:house ?a . }
# Correct answer: Gryffindor
print("Executing a SPARQL query with prefixed names of resources...")
result = kgqa_retriever._query_kg(
sparql_query="select distinct ?sbj where { ?sbj hp:job hp:Keeper_of_keys_and_grounds . }"
)
print(result)
# Paraphrased question: Who is the keeper of keys and grounds?
# Correct answer: <NAME>
print("Executing a SPARQL query with full names of resources...")
result = kgqa_retriever._query_kg(
sparql_query="select distinct ?obj where { <https://deepset.ai/harry_potter/Hermione_granger> <https://deepset.ai/harry_potter/patronus> ?obj . }"
)
print(result)
# Paraphrased question: What is the patronus of Hermione?
# Correct answer: Otter
# -
# ## About us
#
# This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
#
# We bring NLP to the industry via open source!
# Our focus: Industry specific language models & large scale QA systems.
#
# Some of our other work:
# - [German BERT](https://deepset.ai/german-bert)
# - [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
# - [FARM](https://github.com/deepset-ai/FARM)
#
# Get in touch:
# [Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
#
# By the way: [we're hiring!](https://www.deepset.ai/jobs)
| tutorials/Tutorial10_Knowledge_Graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### based on https://github.com/higgsfield/RL-Adventure and https://medium.com/swlh/introduction-to-reinforcement-learning-coding-sarsa-part-4-2d64d6e37617
# %matplotlib inline
import collections
import cv2
import gym
import matplotlib.pyplot as plot
import numpy as np
import random
import time
import torch as t
from IPython.display import clear_output
# +
class LazyFrames(object):
def __init__(self, frames):
"""This object ensures that common frames between the observations are only stored once.
It exists purely to optimize memory usage which can be huge for DQN's 1M frames replay
buffers.
This object should only be converted to numpy array before being passed to the model.
You'd not belive how complex the previous solution was."""
self._frames = frames
def __array__(self, dtype=None):
out = np.concatenate(self._frames, axis=0)
if dtype is not None:
out = out.astype(dtype)
return out
class ImageToPyTorch(gym.ObservationWrapper):
"""
Change image shape to CWH
"""
def __init__(self, env):
super(ImageToPyTorch, self).__init__(env)
old_shape = self.observation_space.shape
self.observation_space = gym.spaces.Box(low=0.0, high=1.0, shape=(old_shape[-1], old_shape[0], old_shape[1]))
def observation(self, observation):
return observation.transpose(2, 0, 1)
class FrameStack(gym.Wrapper):
def __init__(self, env, k):
"""Stack k last frames.
Returns lazy array, which is much more memory efficient.
See Also
--------
baselines.common.atari_wrappers.LazyFrames
"""
gym.Wrapper.__init__(self, env)
self.k = k
self.frames = collections.deque([], maxlen=k)
shp = env.observation_space.shape
self.observation_space = gym.spaces.Box(low=0, high=255, shape=(shp[0]*k, shp[1], shp[2]))
def reset(self):
ob = self.env.reset()
for _ in range(self.k):
self.frames.append(ob)
return self._get_ob()
def step(self, action):
ob, reward, done, info = self.env.step(action)
self.frames.append(ob)
return self._get_ob(), reward, done, info
def _get_ob(self):
assert len(self.frames) == self.k
return LazyFrames(list(self.frames))
class ResizeObservation(gym.ObservationWrapper):
def __init__(self, env):
super(ResizeObservation, self).__init__(env)
shp = env.observation_space.shape
self.observation_space = gym.spaces.Box(low=0, high=255, shape=(shp[0] // 2, shp[1] // 2, shp[2]))
self.resize_to = (shp[1] // 2, shp[0] // 2)
def observation(self, observation):
return cv2.resize(observation, self.resize_to, interpolation=cv2.INTER_AREA)
env = gym.make('PongDeterministic-v4')
env = ResizeObservation(env)
env = ImageToPyTorch(env)
env = FrameStack(env, 4)
# -
USE_CUDA = t.cuda.is_available()# and False
device = t.device('cuda') if USE_CUDA else t.device('cpu')
class Actor(object):
def __init__(self, env, model, eps, eps_final, eps_steps, initial_explore=0):
self.env = env
self.model = model
self.eps = eps
self.eps_final = eps_final
self.eps_decay = np.exp(np.log(eps_final / eps) / eps_steps)
self.initial_explore = initial_explore
def act(self, state):
if self.initial_explore > 0:
self.initial_explore -= 1
return self.env.action_space.sample()
self.eps = max(self.eps_final, self.eps * self.eps_decay)
if random.random() < self.eps:
return self.env.action_space.sample()
state = t.FloatTensor(np.array(state)).to(device)
q = self.model(state)
return q.argmax().item()
class NoisyLinear(t.nn.Module):
def __init__(self, in_features, out_features, std_init=0.4):
super(NoisyLinear, self).__init__()
self.in_features = in_features
self.out_features = out_features
self.std_init = std_init
self.weight_mu = t.nn.Parameter(t.FloatTensor(out_features, in_features))
self.weight_sigma = t.nn.Parameter(t.FloatTensor(out_features, in_features))
self.register_buffer('weight_epsilon', t.FloatTensor(out_features, in_features))
self.bias_mu = t.nn.Parameter(t.FloatTensor(out_features))
self.bias_sigma = t.nn.Parameter(t.FloatTensor(out_features))
self.register_buffer('bias_epsilon', t.FloatTensor(out_features))
self.reset_parameters()
self.reset_noise()
def forward(self, x):
if self.training:
weight = self.weight_mu + self.weight_sigma.mul(self.weight_epsilon)
bias = self.bias_mu + self.bias_sigma.mul(self.bias_epsilon)
else:
weight = self.weight_mu
bias = self.bias_mu
return t.nn.functional.linear(x, weight, bias)
def reset_parameters(self):
mu_range = 1 / np.sqrt(self.weight_mu.size(1))
self.weight_mu.data.uniform_(-mu_range, mu_range)
self.weight_sigma.data.fill_(self.std_init / np.sqrt(self.weight_sigma.size(1)))
self.bias_mu.data.uniform_(-mu_range, mu_range)
self.bias_sigma.data.fill_(self.std_init / np.sqrt(self.bias_sigma.size(0)))
def reset_noise(self):
epsilon_in = self._scale_noise(self.in_features)
epsilon_out = self._scale_noise(self.out_features)
self.weight_epsilon.copy_(epsilon_out.ger(epsilon_in))
self.bias_epsilon.copy_(self._scale_noise(self.out_features))
def _scale_noise(self, size):
x = t.randn(size)
x = x.sign().mul(x.abs().sqrt())
return x
class Model(t.nn.Module):
def __init__(self, input_shape, n_out):
super().__init__()
self.cnn = t.nn.Sequential(
t.nn.Conv2d(12, 32, kernel_size=8, stride=4), t.nn.PReLU(),
t.nn.Conv2d(32, 64, kernel_size=4, stride=2), t.nn.PReLU(),
t.nn.Conv2d(64, 64, kernel_size=3, stride=1), t.nn.PReLU(),
) # -> 64 9 6
self.fc = t.nn.Sequential(
NoisyLinear(self.feature_size(input_shape), 512), t.nn.PReLU(),
t.nn.Linear(512, n_out)
)
def feature_size(self, input_shape):
return self.cnn(t.zeros(1, *input_shape)).view(1, -1).size(1)
def forward(self, x):
if len(x.shape) < 4:
x = x.unsqueeze(0)
p = self.cnn(x)
p = p.view(p.size(0), -1)
return self.fc(p)
class Replay(object):
def __init__(self, maxlen):
self.memory = collections.deque(maxlen=maxlen)
def __len__(self):
return len(self.memory)
def add(self, state, action, next_state, reward, done):
self.memory.append((state, action, next_state, reward, done))
def sample(self, n):
with t.no_grad():
states, actions, next_states, rewards, masks = zip(*random.choices(self.memory, k=n))
actions = t.LongTensor(actions).to(device)
rewards = t.FloatTensor(rewards).to(device)
masks = 1 - t.FloatTensor(masks).to(device)
states = Replay.stack_states(states)
next_states = Replay.stack_states(next_states)
return states, actions, next_states, rewards, masks
@staticmethod
def stack_states(states):
s = np.concatenate([np.expand_dims(x, 0) for x in states])
return t.ByteTensor(s).to(device).float()
# +
def build_model():
return Model(env.observation_space.shape, env.action_space.n).to(device)
def reset_env(env):
state = env.reset()
if 'FIRE' in env.unwrapped.get_action_meanings():
state, _, _, _ = env.step(env.unwrapped.get_action_meanings().index('FIRE'))
return state
def plot_state():
clear_output(False)
plot.figure(figsize=(24,5))
plot.subplot(131)
plot.title('rewards (frame=%dk, %d episodes)' % (np.round(frame/1000), episode))
plot.plot(all_rewards[:-1])
plot.subplot(132)
plot.title('losses')
plot.plot(losses)
plot.subplot(133)
plot.title('random screen')
state, _, _, _, _ = replay.sample(1)
plot.imshow(state.squeeze(0)[-3:].permute(1, 2, 0).cpu().numpy() / 255)
plot.show();
def learn_on_replay():
states, actions, next_states, rewards, masks = replay.sample(batch_size)
q_values = model(states)
q_value = q_values.gather(1, actions.unsqueeze(1)).squeeze(1)
q_next_values = target_model(next_states)
q_next_value = q_next_values.max(1).values
target = rewards + gamma * masks * q_next_value
loss = loss_fn(q_value, target.detach())
opt.zero_grad()
loss.backward()
opt.step()
losses.append(loss.item())
DOUBLE_MODEL = True
model = build_model()
target_model = build_model() if DOUBLE_MODEL else model
opt = t.optim.Adam(model.parameters(), lr=1e-4)
loss_fn = t.nn.SmoothL1Loss()
replay = Replay(1000000)
actor = Actor(env, model, 1, 0.05, 50000, 0)
all_rewards = []
losses = collections.deque(maxlen=10000)
batch_size = 32
gamma = 0.99
frame = 0
for episode in range(10000):
all_rewards.append(0)
state, done = reset_env(env), False
while not done:
frame += 1
action = actor.act(state)
next_state, reward, done, info = env.step(action)
all_rewards[-1] += reward
replay.add(state, action, next_state, reward, done)
state = next_state
if len(replay) > 10000:
learn_on_replay()
if DOUBLE_MODEL and (frame % 1000 == 0):
target_model.load_state_dict(model.state_dict())
if frame % 1000 == 0:
plot_state()
if len(all_rewards) > 100 and np.mean(all_rewards[-100:]) > 18:
break
plot_state()
# -
import gzip
with gzip.open('model-pong-noisy-dqn.gz', 'wb') as f:
t.save(model.state_dict(), f, pickle_protocol=4)
# +
env = gym.make('PongDeterministic-v4')
env = gym.wrappers.Monitor(env, '.', force=True)
env = ResizeObservation(env)
env = ImageToPyTorch(env)
env = FrameStack(env, 4)
state, done = env.reset(), False
while not done:
state = t.FloatTensor(np.array(state)).to(device)
action = model(state).argmax().item()
state, _, done, _ = env.step(action)
env.close()
# !ls -lh
# + active=""
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
| jupyter/pong/pong-noisy-dqn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Feature Engineering
#
# In this notebook we test various statistical transformations of our data in an attempt to create new informative features.
# Global variables for testing changes to this notebook quickly
NUM_FOLDS = 5
MODEL_SEED = 0
NUM_TREES = 25000
EARLY_STOP = 250
SUBMIT = False
# +
# Essentials
import numpy as np
import pandas as pd
import scipy.stats as stats
import pyarrow
import pickle
import time
import matplotlib
# Models and Evaluation
from sklearn.metrics import roc_auc_score
from lightgbm import LGBMClassifier, plot_importance
# Hide warnings
import warnings
warnings.filterwarnings('ignore')
# List input files
import os
for dirname, _, filenames in os.walk('..\data'):
for filename in filenames:
if filename.endswith('.feather'):
print(os.path.join(dirname, filename))
# +
# %%time
train = pd.read_feather(f'../data/train.feather')
test = pd.read_feather(f'../data/test.feather')
features = [x for x in train.columns if x not in ['id','claim','kfold','3fold','4fold','5fold','6fold']]
print("Train Size (Mb):",
round(train.memory_usage().sum() / 1024 ** 2, 2))
print("Test Size (Mb):",
round(test.memory_usage().sum() / 1024 ** 2, 2))
train.head()
# -
# # Baseline
# +
# Scores, validation and test predictions
preds = np.zeros((test.shape[0],))
oof_preds = np.zeros((train.shape[0],))
scores = np.zeros(NUM_FOLDS)
for i in range(NUM_FOLDS):
start = time.time()
X_train = train[train[f'{NUM_FOLDS}fold'] != i][features].copy()
X_valid = train[train[f'{NUM_FOLDS}fold'] == i][features].copy()
y_train = train[train[f'{NUM_FOLDS}fold'] != i]['claim'].copy()
y_valid = train[train[f'{NUM_FOLDS}fold'] == i]['claim'].copy()
X_test = test[features].copy()
model = LGBMClassifier(random_state=MODEL_SEED,
n_estimators = NUM_TREES)
model = model.fit(X_train, y_train,
verbose = False,
eval_set = [(X_valid, y_valid)],
eval_metric = "auc",
early_stopping_rounds = EARLY_STOP
)
# Generate predictions on test set and validation set
valid_preds = model.predict_proba(X_valid)[:,1]
preds += model.predict_proba(X_test)[:, 1] / NUM_FOLDS
# Store scores and out-of-fold predictions
oof_preds[train[f'{NUM_FOLDS}fold'] == i] = valid_preds
scores[i] = roc_auc_score(y_valid, valid_preds)
end = time.time()
print(f"LightGBM Fold {i} (AUC):",
round(scores[i], 6), " ",
str(round(end-start, 3))+"s")
print("\nLightGBM (Avg):", round(scores.mean(), 6))
print("LightGBM (Min):", round(scores.min(), 6))
print(f'OOF AUC: ', roc_auc_score(train['claim'], oof_preds))
# -
# # Baseline Feature Importance
plot_importance(model, figsize=(15,30))
# # Static Transformations
#
# Here we test features which are calculated using only data found within a single row.
def create_row_stats(data):
new_data = data.copy()
new_data['nan_count'] = data.isnull().sum(axis=1)
new_data['nan_std'] = data.isnull().std(axis=1)
new_data['min'] = data.min(axis=1)
new_data['std'] = data.std(axis=1)
new_data['max'] = data.max(axis=1)
new_data['median'] = data.median(axis=1)
new_data['mean'] = data.mean(axis=1)
new_data['var'] = data.var(axis=1)
new_data['sum'] = data.sum(axis=1)
new_data['sem'] = data.sem(axis=1)
new_data['skew'] = data.skew(axis=1)
new_data['median_abs_dev'] = stats.median_abs_deviation(data, axis=1)
new_data['zscore'] = (np.abs(stats.zscore(data))).sum(axis=1)
return new_data
train = create_row_stats(train)
test = create_row_stats(test)
features = [x for x in train.columns if x not in ['id','claim','kfold','3fold','4fold','5fold','6fold']]
# # Training
# +
# Scores, validation and test predictions
preds = np.zeros((test.shape[0],))
oof_preds = np.zeros((train.shape[0],))
scores = np.zeros(NUM_FOLDS)
for i in range(NUM_FOLDS):
start = time.time()
X_train = train[train[f'{NUM_FOLDS}fold'] != i][features].copy()
X_valid = train[train[f'{NUM_FOLDS}fold'] == i][features].copy()
y_train = train[train[f'{NUM_FOLDS}fold'] != i]['claim'].copy()
y_valid = train[train[f'{NUM_FOLDS}fold'] == i]['claim'].copy()
X_test = test[features].copy()
model = LGBMClassifier(random_state=MODEL_SEED,
n_estimators = NUM_TREES)
model = model.fit(X_train, y_train,
verbose = False,
eval_set = [(X_valid, y_valid)],
eval_metric = "auc",
early_stopping_rounds = EARLY_STOP
)
# Generate predictions on test set and validation set
valid_preds = model.predict_proba(X_valid)[:,1]
preds += model.predict_proba(X_test)[:, 1] / NUM_FOLDS
# Store scores and out-of-fold predictions
oof_preds[train[f'{NUM_FOLDS}fold'] == i] = valid_preds
scores[i] = roc_auc_score(y_valid, valid_preds)
end = time.time()
print(f"LightGBM Fold {i} (AUC):",
round(scores[i], 6), " ",
str(round(end-start, 3))+"s")
print("\nLightGBM (Avg):", round(scores.mean(), 6))
print("LightGBM (Min):", round(scores.min(), 6))
print(f'OOF AUC: ', roc_auc_score(train['claim'], oof_preds))
# -
# # Feature Importance
plot_importance(model, figsize=(15,30))
# We notice that our new features not only result in better AUC scores but reduce the training time to roughly 2/3 of what it was previously. For the remaining notebooks we will use all of the transformations except those with less feature importance than any of the original features.
# New function
def create_row_stats(data):
new_data = data.copy()
new_data['nan_count'] = data.isnull().sum(axis=1)
#new_data['nan_std'] = data.isnull().std(axis=1)
new_data['min'] = data.min(axis=1)
#new_data['std'] = data.std(axis=1)
#new_data['max'] = data.max(axis=1)
new_data['median'] = data.median(axis=1)
#new_data['mean'] = data.mean(axis=1)
new_data['var'] = data.var(axis=1)
#new_data['sum'] = data.sum(axis=1)
#new_data['sem'] = data.sem(axis=1)
new_data['skew'] = data.skew(axis=1)
new_data['median_abs_dev'] = stats.median_abs_deviation(data, axis=1)
new_data['zscore'] = (np.abs(stats.zscore(data))).sum(axis=1)
return new_data
| tps-2021-09/notebooks/Notebook 3 - Feature Engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ToshikiShimizu/lecture-notebook/blob/master/Lesson9.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="werBe3xp97Vi" colab_type="text"
# # 指定した文字のカウント
# + id="3ANL7hon_jnB" colab_type="code" colab={}
s = 'hello world'
# + id="IKv17EsM_qNa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d715081f-2e55-4c28-f265-10b3e936440a"
print (s.count('o'))
# + id="pOP-PqPb_sX_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="59191d0c-992f-49b5-fb44-e68531030dca"
print (s.count('l'))
# + id="_9u_rPQv_uGu" colab_type="code" colab={}
| Lesson9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from basic_unet import UNet
from testnet import TestnetDataset
from plot_utils import plot_image_row
from torchvision import transforms
from pathlib import Path
import hdm
from sklearn.metrics import mean_squared_error
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model = UNet(in_channels=1, out_channels=1)
state_dict = torch.load('models/2_testnet_0490.pth')
model.load_state_dict(state_dict)
model = model.to(device)
transform = transforms.Compose([
transforms.Normalize([0.5, 0.5, 0.5, 0.5], [0.5, 0.5, 0.5, 0.5])
])
dataset = TestnetDataset(Path('testnet'), transform)
class DistanceMasks(hdm.HausdorffDistanceMasks):
def calculate_distance(self, image1, image2):
return mean_squared_error(image1, image2)
explainer = DistanceMasks(240, 240)
explainer.generate_masks(circle_size=15, offset=5, normalize=False)
def evaluate_plot(sample):
segment = sample['segment']
image = sample['input']
result = explainer.explain(model, image, segment, device)
raw = result.circle_map(hdm.RAW)
absolute = result.circle_map(hdm.ABSOLUTE)
better = result.circle_map(hdm.BETTER_ONLY)
worse = result.circle_map(hdm.WORSE_ONLY)
plot_image_row(
image,
title='Raw',
color_map='gray',
overlay=raw,
overlay_alpha=0.9,
)
plot_image_row(
image,
title='Absolute',
color_map='gray',
overlay=absolute,
overlay_alpha=0.9,
)
plot_image_row(
image,
title='Better (when this region is occluded, result is better)',
color_map='gray',
overlay=better,
overlay_alpha=0.9,
)
plot_image_row(
image,
title='Worse (when this region is occluded, result is worse)',
color_map='gray',
overlay=worse,
overlay_alpha=0.9,
)
evaluate_plot(dataset.get_sample('0'))
evaluate_plot(dataset.get_sample('1'))
evaluate_plot(dataset.get_sample('2'))
| brats/22b_testnet_hdm_circles_mse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Web Scraping Project
# This notebook scraps toronto's neighbourhood data from a wikipedia webpage
# ## Criteria for marking
# 1. Dataframe should consist of three columns: Postal code, borough and neighbourhood
# 2. All rows must have boroughs, ignore rows where borough is not assigned
# 3. Merge rows with same poastal code, and same borough but different neighbourhood by separating two negihbourhoods with comma
# 4. If the neighbourhood is missing, use borough as neighbourhood
# 5. Clean the dataframe
# 6. Print dimensions of table in the last cell
#
# Loading the required packages
# +
#installing urllib2 package
# #!conda install -c conda-forge urllib2 --yes
# -
from bs4 import BeautifulSoup as bs
import pandas as pd
import numpy as np
#import urllib2
import requests as request
print('Packages are loaded')
#Link for the wikipedia page
link = 'https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M'
wiki_page = request.get(link).text
html_page = bs(wiki_page, 'html.parser')
#print(html_page.prettify())
# +
#html_page.find_all('tbody')
# +
element_list = []
postal_code = []
borough = []
neighbourhood = []
for child in html_page.tbody.stripped_strings:
element_list.append(child)
#print(name_list)
for i in range(3, len(element_list)- 2, 3):
pcode = element_list[i]
bcode = element_list[i+1]
ncode = element_list[i+2]
postal_code.append(pcode)
borough.append(bcode)
neighbourhood.append(ncode)
df = pd.DataFrame(data = {'Postal Code': postal_code, 'Borough': borough, 'Neighbourhood': neighbourhood}, columns = ['Postal Code', 'Borough', 'Neighbourhood'])
df.head()
#print(postal_code)
# -
# Removing *Not assigned* values from Borough
df = df[df.Borough != 'Not assigned']
# Checking if there are any Not assigned values in Neighbourhood
df.iloc[:, 2][df.iloc[:,2] == 'Not assinged']
# The column Neighbourhood doesn't contain any *Not assigned* values.
# Merging the rows with same postal code, same borough but different Neigbourhood. For that, we will use groupby method available in pandas library. We will group the dataframe *df* by *Postal Code* and *Borough* and then join by comma.
grouped = df.groupby(['Postal Code', 'Borough'])['Neighbourhood'].apply(lambda x: ', '.join(x))
grouped = pd.DataFrame(grouped)
# Resetting the index to a series and include *Postal Code* and *Borough* in grouped dataframe
grouped = grouped.reset_index()
grouped.head()
# Printing the shape and rows of dataframe
print('Shape of dataframe: ', grouped.shape)
print('Number of rows in dataframe: ', grouped.shape[0])
# ## Part 2: Plotting the coordinates on the map
# Installing the geocoder pack
# +
# #!conda install -c conda-forge geocoder --yes
# -
#importing the geocoder package
import geocoder
# Getting coordinates using geocoder package
# +
#lat_long_coord = None
#latlng_list = []
#for pcode, ncode in zip(grouped['Postal Code'], grouped['Neighbourhood']):
# while (lat_long_coord is None):
# g = geocoder.google('{}, {}'.format(pcode, ncode))
# lat_long_coord = g.latlng
# latlng_list.append(lat_long_coord)
#coordinates = pd.DataFrame(data = latlng_list, columns = ['Latitude', 'Longitude'])
#coordinates
# -
# Loading coordinates data from csv file as not able to get coordinates from above code
coordinates = pd.read_csv(r'C:\Users\kswp234\Box Sync\3rd Rotation\Data Science\Coursera\IBM Data Science\IBM_DS_coursera\Geospatial_Coordinates.csv')
coordinates.head()
# +
#merging grouped (neighbourhood) data with coordinates
grouped = pd.merge(grouped, coordinates)
grouped.head()
# -
# ## Creating Visualizations
# +
# #!conda install -c conda-forge folium --yes
# -
import folium
# Creating a map of Canada
map_canada = folium.Map(location = [43.6532, -79.3832], zoom_start = 12, tiles = 'Stamen Terrain')
map_canada
# Superimposing neighbourhood labels on Canada Map
# +
for lat, lng, borough, neighborhood in zip(grouped['Latitude'], grouped['Longitude'], grouped['Borough'], grouped['Neighbourhood']):
label = '{}, {}'.format(borough, neighborhood)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_canada)
map_canada
# -
| Coursera/IBM Data Science Specialization/capstone project/Capstone Project Week 3 Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Example to Read / Write to Minio / S3 bucket with Spark
#
# NOTE: Read / Write from Minio like you would the local file system. The different is the path uses `s3a://`
import pyspark
from pyspark.sql import SparkSession
# MINIO CONFIGURATION
s3_host = "minio"
s3_url = f"http://{s3_host}:9000"
s3_key = "minio"
s3_secret = "SU2orange!"
s3_bucket = "minio-example"
# Spark init
spark = SparkSession.builder \
.master("local") \
.appName('jupyter-pyspark') \
.config("spark.jars.packages","org.apache.hadoop:hadoop-aws:3.1.2")\
.config("spark.hadoop.fs.s3a.endpoint", s3_url) \
.config("spark.hadoop.fs.s3a.access.key", s3_key) \
.config("spark.hadoop.fs.s3a.secret.key", s3_secret) \
.config("spark.hadoop.fs.s3a.fast.upload", True) \
.config("spark.hadoop.fs.s3a.path.style.access", True) \
.config("spark.hadoop.fs.s3a.impl", "org.apache.hadoop.fs.s3a.S3AFileSystem") \
.getOrCreate()
sc = spark.sparkContext
sc.setLogLevel("ERROR")
# read local data
df = spark.read.option("multiline","true").json("/home/jovyan/datasets/json-samples/stocks.json")
df.toPandas()
# Write to minio as CSV file - MAKE SURE BUCKET EXISTS !
df.write.mode("Overwrite").csv("s3a://minio-example/stocks.csv",header=True)
# read back from minio
df1 = spark.read.csv("s3a://minio-example/stocks.csv",header=True)
df1.toPandas()
| _old_/docker/all/work/connector-examples/Minio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Plot Model Components
# =====================
#
# Plotting power spectrum model parameters and components.
#
# +
# Import the FOOOFGroup object
from fooof import FOOOFGroup
# Import utilities to manage frequency band definitions
from fooof.bands import Bands
from fooof.analysis import get_band_peak_fg
# Import simulation utilities for making example data
from fooof.sim.gen import gen_group_power_spectra
from fooof.sim.params import param_jitter
# Import plotting function for model parameters and components
from fooof.plts.periodic import plot_peak_fits, plot_peak_params
from fooof.plts.aperiodic import plot_aperiodic_params, plot_aperiodic_fits
# -
# Experiment Set Up & Simulate Data
# ---------------------------------
#
# In this example, we will explore the plotting functions available to visualize
# model parameters and components from fitting power spectrum models.
#
# To do so, we will consider a hypothetical experiment in which we are compare power
# spectrum models between two groups of participants, and so we want to visualize differences
# between the groups. For simplicity, we will consider that we have one 'grand average'
# power spectrum per subject, which we can compare and visualize.
#
#
#
# Set up labels and colors for plotting
colors = ['#2400a8', '#00700b']
labels = ['Group-1', 'Group-2']
# +
# Set the number of 'subjects' per group
n_subjs = 20
# Define the frequency range for our simulations
freq_range = [1, 50]
# Define aperiodic parameters for each group, with some variation
g1_aps = param_jitter([1, 1.25], [0.5, 0.2])
g2_aps = param_jitter([1, 1.00], [0.5, 0.2])
# Define periodic parameters for each group, with some variation
g1_peaks = param_jitter([11, 1, 0.5], [0.5, 0.2, 0.2])
g2_peaks = param_jitter([9, 1, 0.5], [0.25, 0.1, 0.3])
# -
# Simulate some test data, as two groups of power spectra
freqs, powers1 = gen_group_power_spectra(n_subjs, freq_range, g1_aps, g1_peaks)
freqs, powers2 = gen_group_power_spectra(n_subjs, freq_range, g2_aps, g2_peaks)
# Fit Power Spectrum Models
# ~~~~~~~~~~~~~~~~~~~~~~~~~
#
# Now that we have our simulated data, we can fit our power spectrum models, using FOOOFGroup.
#
#
#
# Initialize a FOOOFGroup object for each group
fg1 = FOOOFGroup(verbose=False)
fg2 = FOOOFGroup(verbose=False)
# Parameterize neural power spectra
fg1.fit(freqs, powers1)
fg2.fit(freqs, powers2)
# Plotting Parameters & Components
# --------------------------------
#
# In the following, we will explore two visualization options:
#
# - plotting parameter values
# - plotting component reconstructions
#
# Each of these approaches can be done for either aperiodic or periodic parameters.
#
# All of the plots that we will use in this example can be used to visualize either
# one or multiple groups of data. As we will see, you can pass in a single group of
# parameters or components to visualize them, or pass in a list of group results to
# visualize and compare between groups.
#
# You can also pass in optional inputs `labels` and `colors` to all the following
# functions to add plot labels, and to set the colors used for each group.
#
#
#
# Periodic Components
# -------------------
#
# First, let's have a look at the periodic components.
#
# To do so, we will use the :obj:`~.Bands` object to store our frequency
# band definitions, which we can then use to sub-select peaks within bands of interest.
#
# We can then plot visualizations of the peak parameters, and the reconstructed fits.
#
#
#
# Define frequency bands of interest
bands = Bands({'theta' : [4, 8],
'alpha' : [8, 13],
'beta' : [13, 30]})
# Extract alpha peaks from each group
g1_alphas = get_band_peak_fg(fg1, bands.alpha)
g2_alphas = get_band_peak_fg(fg2, bands.alpha)
# Plotting Peak Parameters
# ~~~~~~~~~~~~~~~~~~~~~~~~
#
# The :func:`~.plot_peak_params` function takes in peak parameters,
# and visualizes them, as:
#
# - Center Frequency on the x-axis
# - Power on the y-axis
# - Bandwidth as the size of the circle
#
#
#
# Explore the peak parameters of Group 1's alphas
plot_peak_params(g1_alphas, freq_range=bands.alpha)
# Compare the peak parameters of alpha peaks between groups
plot_peak_params([g1_alphas, g2_alphas], freq_range=bands.alpha,
labels=labels, colors=colors)
# Plotting Peak Fits
# ~~~~~~~~~~~~~~~~~~
#
# The :func:`~.plot_peak_fits` function takes in peak parameters,
# and reconstructs peak fits.
#
#
#
# Plot the peak fits of the alpha fits for Group 1
plot_peak_fits(g1_alphas)
# Compare the peak fits of alpha peaks between groups
plot_peak_fits([g1_alphas, g2_alphas],
labels=labels, colors=colors)
# Aperiodic Components
# --------------------
#
# Next, let's have a look at the aperiodic components.
#
#
#
# Extract the aperiodic parameters for each group
aps1 = fg1.get_params('aperiodic_params')
aps2 = fg2.get_params('aperiodic_params')
# Plotting Aperiodic Parameters
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# The :func:`~.plot_aperiodic_params` function takes in
# aperiodic parameters, and visualizes them, as:
#
# - Offset on the x-axis
# - Exponent on the y-axis
#
#
#
# Plot the aperiodic parameters for Group 1
plot_aperiodic_params(aps1)
# Compare the aperiodic parameters between groups
plot_aperiodic_params([aps1, aps2], labels=labels, colors=colors)
# Plotting Aperiodic Fits
# ~~~~~~~~~~~~~~~~~~~~~~~
#
# The :func:`~.plot_aperiodic_fits` function takes in
# aperiodic parameters, and reconstructs aperiodic fits.
#
# Here again we can plot visualizations of the peak parameters, and the reconstructed fits.
#
#
#
# Plot the aperiodic fits for Group 1
plot_aperiodic_fits(aps1, freq_range, control_offset=True)
# The :func:`~.plot_aperiodic_fits` also has some additional options
# that can help to tune the visualization, including:
#
# - `control_offset` : whether the control for offset differences, by setting to zero
#
# - This can be useful to visualize if it's the exponent specifically that is changing
# - `log_freqs` : whether to log the frequency values, on the x-axis
#
#
#
# Plot the aperiodic fits for both groups
plot_aperiodic_fits([aps1, aps2], freq_range,
control_offset=True, log_freqs=True,
labels=labels, colors=colors)
# Conclusions
# -----------
#
# In this example, we explored plotting model parameters and components within and between
# groups of parameterized neural power spectra.
#
# If you check the simulation parameters used for the two groups, you can see that
# we set these groups to vary in their alpha center frequency and in the exponent value.
# Qualitatively, we can see those differences in the plots above, and this (in real data)
# would suggest there may be interesting differences between these groups. Follow up
# analyses in such a case could investigate whether there are statistically significant
# differences between these groups.
#
#
#
| doc/auto_examples/plots/plot_model_components.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from sklearn.neighbors import KNeighborsRegressor
import os
import pickle
np.set_printoptions(threshold=np.nan)
# from nn import nn_train, nn_eval
def midi_to_label(midi):
if (midi < strings[0]):
raise Error("Note " + note + " is not playable on a guitar in standard tuning.")
idealString = 0
for string, string_midi in enumerate(strings):
if (midi < string_midi):
break
idealString = string
label = [-1, -1, -1, -1, -1, -1]
label[idealString] = midi - strings[idealString];
return label
def process(filename):
label = int(filename.split('/')[-1])
#label = midi_to_label(int(filename.split('/')[-1]))
f = open(filename, 'r')
data = pickle.load(f)
f.close()
return data, label
strings = [40, 45, 50, 55, 60, 65];
def knn_train(process, files):
knn_models = []
for filename in files:
X_train, y_train = process(filename)
X_train = X_train[:len(X_train)*3/4]
y_train = [y_train]*len(X_train) #[np.asarray(y_train).reshape(1, 1)] * len(X_train)
print("X_train:")
print(X_train[:2])
print(len(X_train[0][1]))
print("y_train")
print(y_train)
print(len(X_train))
print(len(y_train))
neigh = KNeighborsRegressor(n_neighbors=1)
neigh.fit(X_train, y_train)
knn_models.append(neigh)
return knn_models
def knn_eval(neigh, process, files):
for filename in files:
X_test, y_test = process(filename)
X_test = X_test[len(X_test)*3/4:]
y_test = [y_test]*len(X_test) #np.asarray(y_test).reshape(1, 1)
print("X_test:")
print(X_test)
print("y_test")
print(y_test)
y_pred = neigh.predict(X_test)
print("y_pred")
print(y_pred)
# print("Testing Accuracy:", \
# sess.run(accuracy, feed_dict={X: X_test,
# Y: y_test}))
return
# def main():
# files = ["Data/" + f for f in os.listdir('Data') if os.path.isfile("Data/" + f) and "DS_Store" not in f]
# neigh = knn_train(process, files)
# knn_eval(neigh, process, files)
# main()
# + active=""
# df = pd.read_csv("data.csv")
#
# -
df.columns[-1]
# +
import pandas as pd
import numpy as np
from sklearn.neighbors import KNeighborsRegressor
from sklearn.metrics import mean_absolute_error, accuracy_score
from sklearn.neural_network import MLPClassifier
import os
import pickle
df = pd.read_csv("data2.csv", header=None)
df = df.sample(frac=1, random_state=50)
TARGET = [df.columns[-1]]
FEATS = [c for c in df.columns if c != TARGET[0]]
X_train = df[FEATS][:round(0.8*df.shape[0])]
y_train = df[TARGET][:round(0.8*df.shape[0])]
X_test = df[FEATS][round(0.8*df.shape[0]):]
y_test = df[TARGET][round(0.8*df.shape[0]):]
neigh = KNeighborsRegressor(n_neighbors=10)
neigh.fit(X_train, y_train.values.transpose()[0])
y_pred = neigh.predict(X_test.values)
mean_absolute_error(y_test, y_pred)
# for test, pred in zip(y_test.values, y_pred):
# print(test, pred)
# -
y_pred[0]
y_test["true"] = y_pred
y_test["err"] = abs(y_test[255]-y_test["true"])
y_test
# +
err = [abs(y_test.iloc[i].values[0]-y_pred[i]) for i in range(len(y_test))]
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(range(len(err)),err)
# print(y_train.iloc[i])
# plt.plot(x, np.cos(x))
plt.show()
# -
y_test["err"].hist(bins=50)
# +
for test, pred in zip(y_test.values, y_pred):
print(test, pred)
# -
X_train.iloc[0]
# +
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
for i in range(10):
plt.scatter(X_train.iloc[i].index,X_train.iloc[i])
print(y_train.iloc[i])
# plt.plot(x, np.cos(x))
plt.show()
# -
| sklearn_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kcisidro/OOP-58002/blob/main/Long_Quiz_1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="v2tLM9fGpiWV"
# #Long Quiz 1 (Temperature and Celsius)
# + colab={"base_uri": "https://localhost:8080/"} id="6R7Vxhlkp1wB" outputId="984de176-9161-4b33-b9ea-6a47033dcb4e"
class Temperature:
def __init__(self,Celsius):
self.Celsius = Celsius
def Fahrenheit_Conversion(self):
result = float((9 * self.Celsius) / 5 + 32)
return result
input_Temperature = float(input("Insert a Temperature in Celsius: "))
Temperature1 = Temperature(input_Temperature)
print(Temperature1.Fahrenheit_Conversion())
| Long_Quiz_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
t1 = torch.tensor(4.)
t1
t1.dtype
t2 = torch.tensor([1., 2, 3, 4])
t2
t3 = torch.tensor([[5., 6], [7, 8], [9, 10]])
t3
t4 = torch.tensor([[[11, 12, 13],
[13, 14, 15]],
[[15, 16, 17],
[17, 18, 19.]]])
t4
t1.shape
t2.shape
t3.shape
t4.shape
x = torch.tensor(3.)
w = torch.tensor(4., requires_grad=True)
b = torch.tensor(5., requires_grad=True)
y = w * x + b
y
y.backward()
print('dy/dx:', x.grad)
print('dy/dw:', w.grad)
print('dy/db:', b.grad)
import numpy as np
x = np.array([[1, 2], [3, 4.]])
x
y = torch.from_numpy(x)
y
x.dtype, y.dtype
z = y.numpy()
z
import jovian
jovian.commit()
| Untitled56.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # Chapter 09 code snippets
# This notebook contains all code snippets from chapter 9.
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Hyperparameter tuning using HyperDrive
# + gather={"logged": 1630833774903} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import (
Workspace, Environment
)
from azureml.core.conda_dependencies import \
CondaDependencies
import sklearn
ws = Workspace.from_config()
diabetes_env = Environment(name="diabetes-training-env")
diabetes_env.python.conda_dependencies = \
CondaDependencies.create(
conda_packages=[
f"scikit-learn=={sklearn.__version__}"],
pip_packages=["azureml-core",
"azureml-dataset-runtime[pandas]"])
target = ws.compute_targets['cpu-sm-cluster']
# + gather={"logged": 1630833776378} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import ScriptRunConfig
script = ScriptRunConfig(
source_directory='diabetes-training',
script='training.py',
environment=diabetes_env,
compute_target=target
)
# Note that you don't specify the --alpha argument.
# + gather={"logged": 1630833778917} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.train.hyperdrive import HyperDriveConfig
from azureml.train.hyperdrive import (
RandomParameterSampling, uniform, PrimaryMetricGoal
)
param_sampling = RandomParameterSampling({
'alpha': uniform(0.00001, 0.1),
}
)
hd_config = HyperDriveConfig(
run_config=script,
hyperparameter_sampling=param_sampling,
primary_metric_name="nrmse",
primary_metric_goal=
PrimaryMetricGoal.MINIMIZE,
max_total_runs=20,
max_concurrent_runs=4)
# + gather={"logged": 1630834973358} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import Experiment
experiment = Experiment(ws, "chapter09-hyperdrive")
hyperdrive_run = experiment.submit(hd_config)
hyperdrive_run.wait_for_completion(show_output=True)
# + gather={"logged": 1630834974037} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
best_run = hyperdrive_run.get_best_run_by_primary_metric()
best_run_metrics = best_run.get_metrics(name='nrmse')
parameter_values = best_run.get_details()[
'runDefinition']['arguments']
print('Best Run Id: ', best_run.id)
print('- NRMSE:', best_run_metrics['nrmse'])
print('- alpha:', parameter_values[1])
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Using the early termination policy
# + gather={"logged": 1630822462208} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import Workspace, ScriptRunConfig, Environment
ws = Workspace.from_config()
target = ws.compute_targets["cpu-sm-cluster"]
script = ScriptRunConfig(
source_directory="termination-policy-training",
script="training.py",
environment=Environment.get(ws, "AzureML-Minimal"),
compute_target=target,
)
# + gather={"logged": 1630822463391} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.train.hyperdrive import (
GridParameterSampling,
choice,
MedianStoppingPolicy,
HyperDriveConfig,
PrimaryMetricGoal
)
param_sampling = GridParameterSampling(
{
"a": choice(1, 2, 3, 4),
"b": choice(1, 2, 3, 4),
}
)
early_termination_policy = MedianStoppingPolicy(
evaluation_interval=1, delay_evaluation=5
)
# More aggressive alternative
# from azureml.train.hyperdrive import TruncationSelectionPolicy
# early_termination_policy = TruncationSelectionPolicy(
# truncation_percentage=50, evaluation_interval=1
#)
hd_config = HyperDriveConfig(
policy=early_termination_policy,
run_config=script,
hyperparameter_sampling=param_sampling,
primary_metric_name="fake_metric",
primary_metric_goal=PrimaryMetricGoal.MAXIMIZE,
max_total_runs=50,
max_concurrent_runs=4
)
# + gather={"logged": 1630823351465} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import Experiment
experiment = Experiment(ws, "chapter09-hyperdrive")
hyperdrive_run = experiment.submit(hd_config)
hyperdrive_run.wait_for_completion(show_output=True)
# + [markdown] nteract={"transient": {"deleting": false}}
# ## Running AutoML experiments via code
# + gather={"logged": 1628352313404} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core import Workspace, Dataset
from azureml.train.automl import AutoMLConfig
ws = Workspace.from_config()
compute_target = ws.compute_targets["cpu-sm-cluster"]
diabetes_dataset = Dataset.get_by_name(workspace=ws, name='diabetes')
train_ds,validate_ds = diabetes_dataset.random_split(percentage=0.8, seed=1337)
print(f"Train: {len(train_ds.to_pandas_dataframe().axes[0])} rows")
print(f"Validate: {len(validate_ds.to_pandas_dataframe().axes[0])} rows")
experiment_config = AutoMLConfig(
task = "regression",
primary_metric = 'normalized_root_mean_squared_error',
training_data = train_ds,
label_column_name = "target",
validation_data = validate_ds,
compute_target = compute_target,
experiment_timeout_hours = 0.25,
iterations = 4
)
# + gather={"logged": 1628353037974} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from azureml.core.experiment import Experiment
my_experiment = Experiment(ws, 'chapter09-automl-experiment')
run = my_experiment.submit(experiment_config, show_output=True)
# + gather={"logged": 1628353526686} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# To retrieve a previous run
# from azureml.train.automl.run import AutoMLRun
# run = AutoMLRun(my_experiment, run_id = 'AutoML_80833402-6e7e-4c25-b6aa-b6fd44d75d09')
# Get best run and model
best_run, best_model = run.get_output()
# Or with index
best_run = run.get_output()[0]
best_model = run.get_output()[1]
# + gather={"logged": 1628353536565} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Scaling/normalization and model algorythm
best_model.steps
# + gather={"logged": 1628353547828} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# Automated feature engineering
# Use 'timeseriestransformer' for task='forecasting',
# else use 'datatransformer' for 'regression' or 'classification' task.
print(best_model.named_steps['datatransformer'] \
.get_featurization_summary())
feature_names=best_model.named_steps['datatransformer'] \
.get_engineered_feature_names()
print("Engineered feature names:")
print(feature_names)
| chapter09/chapter09.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Only execute if you haven't already. Make sure to restart the kernel if these libraries have not been previously installed.
# !pip install xgboost==0.82 --user
# !pip install scikit-learn==0.20.4 --user
# **Note**: You may need to restart the kernel to use updated packages.
# # Import Python packages
#
# Execute the command below (__Shift + Enter__) to load all the python libraries we'll need for the lab.
# +
import datetime
import pickle
import os
import pandas as pd
import xgboost as xgb
import numpy as np
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import FeatureUnion, make_pipeline
from sklearn.utils import shuffle
from sklearn.base import clone
from sklearn.model_selection import train_test_split
from witwidget.notebook.visualization import WitWidget, WitConfigBuilder
import custom_transforms
import warnings
warnings.filterwarnings(action='ignore', category=DeprecationWarning)
# -
# Before we continue, note that we'll be using your Qwiklabs project id a lot in this notebook. For convenience, set it as an environment variable using the command below:
os.environ['QWIKLABS_PROJECT_ID'] = ''
# # Download and process data
#
# The models you'll build will predict the income level, whether it's less than or equal to $50,000 per year, of individuals given 14 data points about each individual. You'll train your models on this UCI [Census Income Dataset](https://archive.ics.uci.edu/ml/datasets/Adult).
#
# We'll read the data into a Pandas DataFrame to see what we'll be working with. It's important to shuffle our data in case the original dataset is ordered in a specific way. We use an sklearn utility called shuffle to do this, which we imported in the first cell:
# +
train_csv_path = 'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.data'
COLUMNS = (
'age',
'workclass',
'fnlwgt',
'education',
'education-num',
'marital-status',
'occupation',
'relationship',
'race',
'sex',
'capital-gain',
'capital-loss',
'hours-per-week',
'native-country',
'income-level'
)
raw_train_data = pd.read_csv(train_csv_path, names=COLUMNS, skipinitialspace=True)
raw_train_data = shuffle(raw_train_data, random_state=4)
# -
# `data.head()` lets us preview the first five rows of our dataset in Pandas.
raw_train_data.head()
# The `income-level` column is the thing our model will predict. This is the binary outcome of whether the individual makes more than $50,000 per year. To see the distribution of income levels in the dataset, run the following:
print(raw_train_data['income-level'].value_counts(normalize=True))
# As explained in [this paper](http://cseweb.ucsd.edu/classes/sp15/cse190-c/reports/sp15/048.pdf), each entry in the dataset contains the following information
# about an individual:
#
# * __age__: the age of an individual
# * __workclass__: a general term to represent the employment status of an individual
# * __fnlwgt__: final weight. In other words, this is the number of people the census believes
# the entry represents...
# * __education__: the highest level of education achieved by an individual.
# * __education-num__: the highest level of education achieved in numerical form.
# * __marital-status__: marital status of an individual.
# * __occupation__: the general type of occupation of an individual
# * __relationship__: represents what this individual is relative to others. For example an
# individual could be a Husband. Each entry only has one relationship attribute and is
# somewhat redundant with marital status.
# * __race__: Descriptions of an individual’s race
# * __sex__: the biological sex of the individual
# * __capital-gain__: capital gains for an individual
# * __capital-loss__: capital loss for an individual
# * __hours-per-week__: the hours an individual has reported to work per week
# * __native-country__: country of origin for an individual
# * __income-level__: whether or not an individual makes more than $50,000 annually
# An important concept in machine learning is train / test split. We'll take the majority of our data and use it to train our model, and we'll set aside the rest for testing our model on data it's never seen before. There are many ways to create training and test datasets. Fortunately, for our census data we can simply download a pre-defined test set.
test_csv_path = 'https://archive.ics.uci.edu/ml/machine-learning-databases/adult/adult.test'
raw_test_data = pd.read_csv(test_csv_path, names=COLUMNS, skipinitialspace=True, skiprows=1)
raw_test_data.head()
# Since we don't want to train a model on our labels, we're going to separate them from the features in both the training and test datasets. Also, notice that `income-level` is a string datatype. For machine learning, it's better to convert this to an binary integer datatype. We do this in the next cell.
# +
raw_train_features = raw_train_data.drop('income-level', axis=1).values
raw_test_features = raw_test_data.drop('income-level', axis=1).values
# Create training labels list
train_labels = (raw_train_data['income-level'] == '>50K').values.astype(int)
test_labels = (raw_test_data['income-level'] == '>50K.').values.astype(int)
# -
# Now you're ready to build and train your first model!
# # Build a First Model
#
# The model we build closely follows a template for the [census dataset found on AI Hub](https://aihub.cloud.google.com/p/products%2F526771c4-9b36-4022-b9c9-63629e9e3289). For our model we use an XGBoost classifier. However, before we train our model we have to pre-process the data a little bit. We build a processing pipeline using [Scikit-Learn's Pipeline constructor](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html). We appl some custom transformations that are defined in `custom_transforms.py`. Open the file `custom_transforms.py` and inspect the code. Out features are either numerical or categorical. The numerical features are `age-num`, and `hours-per-week`. These features will be processed by applying [Scikit-Learn's StandardScaler function](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html). The categorical features are `workclass`, `education`, `marital-status`, and `relationship`. These features are [one-hot encoded](https://machinelearningmastery.com/why-one-hot-encode-data-in-machine-learning/).
# +
numerical_indices = [0, 12]
categorical_indices = [1, 3, 5, 7]
p1 = make_pipeline(
custom_transforms.PositionalSelector(categorical_indices),
custom_transforms.StripString(),
custom_transforms.SimpleOneHotEncoder()
)
p2 = make_pipeline(
custom_transforms.PositionalSelector(numerical_indices),
StandardScaler()
)
p3 = FeatureUnion([
('numericals', p1),
('categoricals', p2),
])
# -
# To finalize the pipeline we attach an XGBoost classifier at the end. The complete pipeline object takes the raw data we loaded from csv files, processes the categorical features, processes the numerical features, concatenates the two, and then passes the result through the XGBoost classifier.
pipeline = make_pipeline(
p3,
xgb.sklearn.XGBClassifier(max_depth=4)
)
# We train our model with one function call using the fit() method. We pass the fit() method our training data.
pipeline.fit(raw_train_features, train_labels)
# Let's go ahead and save our model as a pickle file. Executing the command below will save the trained model in the file `model.pkl` in the same directory as this notebook.
with open('model.pkl', 'wb') as model_file:
pickle.dump(pipeline, model_file)
# # Save Trained Model to AI Platform
#
# We've got our model working locally, but it would be nice if we could make predictions on it from anywhere (not just this notebook!). In this step we'll deploy it to the cloud. For detailed instructions on how to do this visit [the official documenation](https://cloud.google.com/ai-platform/prediction/docs/exporting-for-prediction). Note that since we have custom components in our data pipeline we need to go through a few extra steps.
# ## Create a Cloud Storage bucket for the model
#
# We first need to create a storage bucket to store our pickled model file. We'll point Cloud AI Platform at this file when we deploy. Run this gsutil command to create a bucket. This will ensure the name of the cloud storage bucket you create will be globally unique.
# !gsutil mb gs://$QWIKLABS_PROJECT_ID
# ## Package custom transform code
#
# Since we're using custom transformation code we need to package it up and direct AI Platform to it when we ask it make predictions. To package our custom code we create a source distribution. The following code creates this distribution and then ports the distribution and the model file to the bucket we created. Ignore the warnings about missing meta data.
# + language="bash"
#
# python setup.py sdist --formats=gztar
#
# gsutil cp model.pkl gs://$QWIKLABS_PROJECT_ID/original/
# gsutil cp dist/custom_transforms-0.1.tar.gz gs://$QWIKLABS_PROJECT_ID/
# -
# ## Create and Deploy Model
#
# The following ai-platform gcloud command will create a new model in your project. We'll call this one `census_income_classifier`.
# !gcloud ai-platform models create census_income_classifier --regions us-central1
# Now it's time to deploy the model. We can do that with this gcloud command:
# + language="bash"
#
# MODEL_NAME="census_income_classifier"
# VERSION_NAME="original"
# MODEL_DIR="gs://$QWIKLABS_PROJECT_ID/original/"
# CUSTOM_CODE_PATH="gs://$QWIKLABS_PROJECT_ID/custom_transforms-0.1.tar.gz"
#
# gcloud beta ai-platform versions create $VERSION_NAME \
# --model $MODEL_NAME \
# --runtime-version 1.15 \
# --python-version 3.7 \
# --origin $MODEL_DIR \
# --package-uris $CUSTOM_CODE_PATH \
# --prediction-class predictor.MyPredictor \
# --region=global
# -
# While this is running, check the [models section](https://console.cloud.google.com/ai-platform/models) of your AI Platform console. You should see your new version deploying there. When the deploy completes successfully you'll see a green check mark where the loading spinner is. The deploy should take 2-3 minutes. You will need to click on the model name in order to see the spinner/checkmark. In the command above, notice we specify `prediction-class`. The reason we must specify a prediction class is that by default, AI Platform prediction will call a Scikit-Learn model's `predict` method, which in this case returns either 0 or 1. However, the What-If Tool requires output from a model in line with a Scikit-Learn model's `predict_proba` method. This is because WIT wants the probabilities of the negative and positive classes, not just the final determination on which class a person belongs to. Because that allows us to do more fine-grained exploration of the model. Consequently, we must write a [custom prediction routine](https://cloud.google.com/ai-platform/prediction/docs/custom-prediction-routines) that basically renames `predict_proba` as `predict`. The custom prediction method can be found in the file `predictor.py`. This file was packaged in the section __Package custom transform code__. By specifying `prediction-class` we're telling AI Platform to call our custom prediction method--basically, `predict_proba`-- instead of the default `predict` method.
# ## Test the deployed model
#
# To make sure your deployed model is working, test it out using gcloud to make a prediction. First, save a JSON file with one test instance for prediction:
# %%writefile predictions.json
[25, "Private", 226802, "11th", 7, "Never-married", "Machine-op-inspct", "Own-child", "Black", "Male", 0, 0, 40, "United-States"]
# Test your model by running this code:
# !gcloud ai-platform predict --model=census_income_classifier --json-instances=predictions.json --version=original --region=global
# You should see your model's prediction in the output. The first entry in the output is the model's probability that the individual makes under \\$50K while the second entry is the model's confidence that the individual makes over \\$50k. The two entries sum to 1.
# # What-If Tool
# To connect the What-if Tool to your AI Platform models, you need to pass it a subset of your test examples along with the ground truth values for those examples. Let's create a Numpy array of 2000 of our test examples.
# +
num_datapoints = 2000
test_examples = np.hstack(
(raw_test_features[:num_datapoints],
test_labels[:num_datapoints].reshape(-1,1)
)
)
# -
# Instantiating the What-if Tool is as simple as creating a WitConfigBuilder object and passing it the AI Platform model we built. Note that it'll take a minute to load the visualization.
# +
config_builder = (
WitConfigBuilder(test_examples.tolist(), COLUMNS)
.set_ai_platform_model(os.environ['QWIKLABS_PROJECT_ID'], 'census_income_classifier', 'original')
.set_target_feature('income-level')
.set_model_type('classification')
.set_label_vocab(['Under 50K', 'Over 50K'])
)
WitWidget(config_builder, height=800)
# -
# The default view on the What-if Tool is the __Datapoint editor__ tab. Here, you can click on any individual data point to see its features and even change feature values. Navigate to the __Performance & Fairness__ tab in the What-if Tool. By slicing on a feature you can view the model error for individual feature values. Finally, navigate to the __Features__ tab in the What-if Tool. This shows you the distribution of values for each feature in your dataset. You can use this tab to make sure your dataset is balanced. For example, if we only had Asians in a population, the model's predictions wouldn't necessarily reflect real world data. This tab gives us a good opportunity to see where our dataset might fall short, so that we can go back and collect more data to make it balanced.
#
# In the __Features__ tab, we can look to see the distribution of values for each feature in the dataset. We can see that of the 2000 test datapoints, 1346 are from men and 1702 are from caucasions. Women and minorities seem under-represented in this dataset. That may lead to the model not learning an accurate representation of the world in which it is trying to make predictions (of course, even if it does learn an accurate representation, is that what we want the model to perpetuate? This is a much deeper question still falling under the ML fairness umbrella and worthy of discussion outside of WIT). Predictions on those under-represented groups are more likely to be inaccurate than predictions on the over-represented groups.
#
# The features in this visualization can be sorted by a number of different metrics, including non-uniformity. With this sorting, the features that have the most non-uniform distributions are shown first. For numeric features, capital gain is very non-uniform, with most datapoints having it set to 0, but a small number having non-zero capital gains, all the way up to a maximum of 100k. For categorical features, country is the most non-uniform with most datapoints being from the USA, but there is a long tail of 40 other countries which are not well represented.
#
# Back in the __Performance & Fairness__ tab, we can set an input feature (or set of features) with which to slice the data. For example, setting this to `sex` allows us to see the breakdown of model performance on male datapoints versus female datapoints. We can see that the model is more accurate (has less false positives and false negatives) on females than males. We can also see that the model predicts high income for females much less than it does for males (8.0% of the time for females vs 27.1% of the time for males). __Note, your numbers will be slightly different due to the random elements of model training__.
#
# Imagine a scenario where this simple income classifier was used to approve or reject loan applications (not a realistic example but it illustrates the point). In this case, 28% of men from the test dataset have their loans approved but only 10% of women have theirs approved. If we wished to ensure than men and women get their loans approved the same percentage of the time, that is a fairness concept called "demographic parity". One way to achieve demographic parity would be to have different classification thresholds for males and females in our model.
#
# In this case, demographic parity can be found with both groups getting loans 16% of the time by having the male threshold at 0.67 and the female threshold at 0.31. Because of the vast difference in the properties of the male and female training data in this 1994 census dataset, we need quite different thresholds to achieve demographic parity. Notice how with the high male threshold there are many more false negatives than before, and with the low female threshold there are many more false positives than before. This is necessary to get the percentage of positive predictions to be equal between the two groups. WIT has buttons to optimize for other fairness constraints as well, such as "equal opportunity" and "equal accuracy". Note that the demographic parity numbers may be different from the ones in your text as the trained models are always a bit different.
#
# The use of these features can help shed light on subsets of your data on which your classifier is performing very differently. Understanding biases in your datasets and data slices on which your model has disparate performance are very important parts of analyzing a model for fairness. There are many approaches to improving fairness, including augmenting training data, building fairness-related loss functions into your model training procedure, and post-training inference adjustments like those seen in WIT. We think that WIT provides a great interface for furthering ML fairness learning, but of course there is no silver bullet to improving ML fairness.
# # Training on a more balanced dataset
#
# Using the What-If Tool we saw that the model we trained on the census dataset wouldn't be very considerate in a production environment. What if we retrained the model on a dataset that was more balanced? Fortunately, we have such a dataset. Let's train a new model on this balanced dataset and compare it to our original dataset using the What-If Tool.
# First, let's load the balanced dataset into a Pandas dataframe.
bal_data_path = 'https://storage.googleapis.com/cloud-training/dei/balanced_census_data.csv'
bal_data = pd.read_csv(bal_data_path, names=COLUMNS, skiprows=1)
bal_data.head()
# Execute the command below to see the distribution of gender in the data.
bal_data['sex'].value_counts(normalize=True)
# Unlike the original dataset, this dataset has an equal number of rows for both males and females. Execute the command below to see the distriubtion of rows in the dataset of both `sex` and `income-level`.
bal_data.groupby(['sex', 'income-level'])['sex'].count()
# We see that not only is the dataset balanced across gender, it's also balanced across income. Let's train a model on this data. We'll use exactly the same model pipeline as in the previous section. Scikit-Learn has a convenient utility function for copying model pipelines, `clone`. The `clone` function copies a pipeline architecture without saving learned parameter values.
# +
bal_data['income-level'] = bal_data['income-level'].isin(['>50K', '>50K.']).values.astype(int)
raw_bal_features = bal_data.drop('income-level', axis=1).values
bal_labels = bal_data['income-level'].values
# -
pipeline_bal = clone(pipeline)
pipeline_bal.fit(raw_bal_features, bal_labels)
# As before, we save our trained model to a pickle file. Note, when we version this model in AI Platform the model in this case must be named `model.pkl`. It's ok to overwrite the existing `model.pkl` file since we'll be uploading it to Cloud Storage anyway.
with open('model.pkl', 'wb') as model_file:
pickle.dump(pipeline_bal, model_file)
# Deploy the model to AI Platform using the following bash script:
# + language="bash"
#
# gsutil cp model.pkl gs://$QWIKLABS_PROJECT_ID/balanced/
#
# MODEL_NAME="census_income_classifier"
# VERSION_NAME="balanced"
# MODEL_DIR="gs://$QWIKLABS_PROJECT_ID/balanced/"
# CUSTOM_CODE_PATH="gs://$QWIKLABS_PROJECT_ID/custom_transforms-0.1.tar.gz"
#
# gcloud beta ai-platform versions create $VERSION_NAME \
# --model $MODEL_NAME \
# --runtime-version 1.15 \
# --python-version 3.7 \
# --origin $MODEL_DIR \
# --package-uris $CUSTOM_CODE_PATH \
# --prediction-class predictor.MyPredictor \
# --region=global
# -
# Now let's instantiate the What-if Tool by configuring a WitConfigBuilder. Here, we want to compare the original model we built with the one trained on the balanced census dataset. To achieve this we utilize the `set_compare_ai_platform_model` method. We want to compare the models on a balanced test set. The balanced test is loaded and then input to `WitConfigBuilder`.
bal_test_csv_path = 'https://storage.googleapis.com/cloud-training/dei/balanced_census_data_test.csv'
bal_test_data = pd.read_csv(bal_test_csv_path, names=COLUMNS, skipinitialspace=True)
bal_test_data['income-level'] = (bal_test_data['income-level'] == '>50K').values.astype(int)
# +
config_builder = (
WitConfigBuilder(bal_test_data.to_numpy()[1:].tolist(), COLUMNS)
.set_ai_platform_model(os.environ['QWIKLABS_PROJECT_ID'], 'census_income_classifier', 'original')
.set_compare_ai_platform_model(os.environ['QWIKLABS_PROJECT_ID'], 'census_income_classifier', 'balanced')
.set_target_feature('income-level')
.set_model_type('classification')
.set_label_vocab(['Under 50K', 'Over 50K'])
)
WitWidget(config_builder, height=800)
# -
# Once the WIT widget loads, click on the __Performance & Fairness__ tab. In the __Slice by__ field select `sex` and wait a minute for the graphics to load. For females, the model trained on the balanced dataset is over two times more likely to predict an income of over 50k than the model trained on the original dataset. While this results in a higher false positive rate, the false negative rate is decreased by a factor of three. This results in an improved overall accuracy of some 10 percentage points.
#
# How else does the model trained on balanced data perform differently when compared to the original model?
| quests/dei/census/income_xgboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from pathlib import Path
from glob import glob
import numpy as np
from datetime import datetime,timedelta
from tqdm.notebook import tqdm
# Import and sort all caravela apogee_SL510 eurec4a files, creating a dataframe from this:
# +
files_list = glob(r'E:\Eurec4a_master\Caravela\apogee_SP_110\*\*'+ '/*SP_110*')
files_list.sort()
li = []
for filename in tqdm(files_list):
df = pd.read_csv(filename, header=0, sep=',',index_col=False)
li.append(df)
sp = pd.concat(li, axis=0)
# -
sp
# Parse the timestamp to a datetime format
dt = []
for i in tqdm(sp['PC Timestamp[UTC]']):
dt.append(datetime.fromisoformat([i]))
sp['dt [UTC]'] = dt
sp = sp[(sp['dt [UTC]'] >= '2020-01-22 00:00:00.000')] #select data from Caravela's launch onwards
sp
# Tidy this up to drop the columns we dont need
sp = sp.reset_index()
sp = sp.drop(['index','PC Time Zone','PC Timestamp[UTC]' ],axis=1)
# check for any gaps in timeseries larger than 2 seconds
time_diff = sp['dt [UTC]'].values[1:] - sp['dt [UTC]'].values[:-1]
for i in np.arange(1, len(time_diff)):
if np.timedelta64(3599, 's') >= time_diff[i] > np.timedelta64(2, 's'):
print('gap starts at', sp['dt [UTC]'][i], 'and lasts for',
np.timedelta64(time_diff[i],'s'))
if time_diff[i] > np.timedelta64(3599, 's'):
print('gap starts at', sp['dt [UTC]'][i], 'and lasts for',
np.timedelta64(time_diff[i],'s'), ' - approximately', np.timedelta64(time_diff[i],'h'))
# Convert to iso time as this is a universally accepted format
a = []
for i in tqdm(range(0,len(sp['dt [UTC]']))):
a.append(sp['dt [UTC]'][i].isoformat())
sp['datetime [UTC]'] = a
sp = sp.drop(['dt [UTC]'], axis = 1)
sp.to_csv('CARAVELA_SP110.csv',index = None)
baa = pd.read_csv('CARAVELA_SP110.csv')# import file to test it
z = []
for i in tqdm(baa['datetime [UTC]']):
z.append(datetime.fromisoformat(i))
baa['dt'] = z
import matplotlib.pyplot as plt
fig,ax = plt.subplots(1,1, figsize=(18, 15))
ax.plot(baa['dt'], baa['SP-110-SS[W m-2]'])
ax.set_ylabel('SP-110-SS[W m-2]')
ax.set_xlabel('Date')
| apogee_SP110.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Circuit resource analysis for Gray code encoding
# +
import numpy as np
np.warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import sys
sys.path.append("../src/")
from qiskit_circuits import *
np.set_printoptions(precision=6, suppress=True)
# -
# Actually generate the ansatz circuits, and use Qiskit to collect the depth, and gate counts. Make sure that they match up with the formulas that we wrote down in the paper.
print("Jordan-Wigner ansatz resource counts (single circuit, computational basis measurement.)\n")
for N in range(2, 8):
t = np.random.randn(N-1)
c = sparse_variational_circuit(t, 0, 'statevector_simulator')
resources = c.decompose().count_ops()
print(f"N = {N}")
print(f"Depth = \t{c.depth()}, \t4N - 6 = \t{4*N - 6}")
print(f"CNOTs = \t{resources['cx']}, \t3N - 5 = \t{3*N - 5}")
print(f"SQ = \t{resources['u3'] + resources['r']}, \t2N - 2 = \t{2*N-2}")
print()
print("Gray-code ansatz resource counts (single circuit, computational basis measurement.)\n")
for N in range(2, 32):
η = int(np.ceil(np.log2(N)))
t = np.random.randn(2**η - 1)
c = dense_variational_circuit(t, 0, 'statevector_simulator')
#print(c.draw())
resources = c.decompose().count_ops()
depth = int(np.ceil((2**η - 1)/η))*(η+1) - 2*η + (2**η-1)%η
print(f"N = {N}")
print(f"Depth = \t{c.depth()}, \t expr = \t{depth}")
if 'cx' in resources:
print(f"CNOTs = \t{resources['cx']}, \t 2^η-η-1 = \t{2**η-η-1}")
print(f"SQ = \t{resources['r']}, \t 2^η-1 = \t{(2**η-1)}")
print()
# Let's check in terms of η
print("Gray-code ansatz resource counts (single circuit, computational basis measurement.)\n")
for η in range(2, 10):
t = np.random.randn(2**η - 1)
c = dense_variational_circuit(t, 0, 'statevector_simulator')
#print(c.draw())
resources = c.decompose().count_ops()
depth = int(np.ceil((2**η - 1)/η))*(η+1) - 2*η + (2**η-1)%η
print(f"η = {η}")
print(f"Depth = \t{c.depth()}, \t expr = \t{depth}")
if 'cx' in resources:
print(f"CNOTs = \t{resources['cx']}, \t 2^η-η-1 = \t{2**η-η-1}")
print(f"SQ = \t{resources['r']}, \t 2^η-1 = \t{(2**η-1)}")
print()
# ## Plots
# Now let's make some plots and do a comparison of total gate count and circuit depth.
# +
min_N = 2
max_N = 512
gates_jw = [18*N - 21 for N in range(min_N, max_N)]
gates_gc = []
for N in range(min_N, max_N):
η = np.ceil(np.log2(N))
gates_gc.append(2*(η+1)*(2**η-1) - η**2)
# -
plt.plot(range(min_N, max_N), gates_gc, label="GC")
plt.plot(range(min_N, max_N), gates_jw, label="JW")
plt.xlabel("N")
plt.ylabel("Total gate count")
plt.title("Total VQE gate count")
plt.legend();
# +
min_N = 2
max_N = 1024
cnots_jw = [3*(3*N - 5) for N in range(min_N, max_N)]
cnots_gc = []
for N in range(min_N, max_N):
η = np.ceil(np.log2(N))
cnots_gc.append((η+1)*(2**η-η-1))
# -
plt.plot(range(min_N, max_N), cnots_gc, label="GC")
plt.plot(range(min_N, max_N), cnots_jw, label="JW")
plt.xlabel("N")
plt.ylabel("CNOT count")
plt.title("VQE ansatz CNOT counts")
plt.legend();
# We notice that after $N=256$, the GC encoding uses consistently more gates Before that point, it still used less gates for $N$ that were powers of 2.
# Now the depth...
depth_jw = [4*N - 6 for N in range(min_N, max_N)]
depth_gc = []
for N in range(min_N, max_N):
η = np.ceil(np.log2(N))
depth_gc.append(int(np.ceil((2**η - 1)/η))*(η+1) - 2*η + (2**η-1)%η)
plt.plot(range(min_N, max_N), depth_gc, label="GC")
plt.plot(range(min_N, max_N), depth_jw, label="JW")
plt.xlabel("N")
plt.ylabel("Depth")
plt.title("VQE ansatz depth comparison")
plt.legend();
# The ansatz used for the GC encoding always has lower depth, even though the number of gates is higher.
| notebooks/Circuit-Complexity-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prep
#
# Setting up some prior functionality
import torch, torchvision
print(torch.__version__, torch.cuda.is_available())
# # Load a model
#
# First we have to decide if our model should be pretrained.
#
# This greatly depends on the size of a dataset. Smaller datasets rely more on finetuning.
# +
pretrained = True
if pretrained:
# Get pretrained weights
checkpoint = torch.hub.load_state_dict_from_url(
url='https://dl.fbaipublicfiles.com/detr/detr-r50-e632da11.pth',
map_location='cpu',
check_hash=True)
# Remove class weights
del checkpoint["model"]["class_embed.weight"]
del checkpoint["model"]["class_embed.bias"]
# SaveOGH
torch.save(checkpoint,
'detr-r50_no-class-head.pth')
# -
# # Dataset
#
# Our dataset should be loadable as a COCO format
#
# This allows us to use the pycocotools to load the data dict for the main python script
# +
dataset_file = "coco" # alternatively, implement your own coco-type dataset loader in datasets and add this "key" to datasets/__init__.py
dataDir='/COCO_dataset/' # should lead to a directory with a train2017 and val2017 folder as well as an annotations folder
num_classes = 91 # this int should be the actual number of classes + 1 (for no class)
outDir = 'outputs'
resume = "detr-r50_no-class-head.pth" if pretrained else ""
# -
# # Training
#
# We use the main.py script to run our training
# !python main.py \
# --dataset_file $dataset_file \
# --coco_path $dataDir \
# --output_dir $outDir \
# --resume $resume \
# --num_classes $num_classes \
# --lr 1e-5 \
# --lr_backbone 1e-6 \
# --epochs 1
# # Results
#
# Quick and easy overview of the training results
# +
from util.plot_utils import plot_logs
from pathlib import Path
log_directory = [Path(outDir)]
# +
fields_of_interest = (
'loss',
'mAP',
)
plot_logs(log_directory,
fields_of_interest)
# +
fields_of_interest = (
'loss_ce',
'loss_bbox',
'loss_giou',
)
plot_logs(log_directory,
fields_of_interest)
# +
fields_of_interest = (
'class_error',
'cardinality_error_unscaled',
)
plot_logs(log_directory,
fields_of_interest)
| finetune_detr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('survey_results_public.csv')
schema_df = pd.read_csv('survey_results_schema.csv')
pd.set_option('display.max_columns', 61)
pd.set_option('display.max_rows', 61)
df.head()
schema_df
df.shape
df.columns
df['Hobbyist']
df['Hobbyist'].value_counts()
df.loc[[0, 1, 2], 'Hobbyist']
df.loc[0:2, 'Hobbyist']
df.loc[0:5, 'Hobbyist':'Country']
| pandas/Pandas lab 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (base)
# language: python
# name: base
# ---
# <a href="https://colab.research.google.com/github/institutohumai/cursos-python/blob/master/AnalisisDeDatos/4_Data_Wrangling_Avanzado/data_wrangling_avanzado.ipynb"> <img src='https://colab.research.google.com/assets/colab-badge.svg' /> </a>
# <div align="center"> Recordá abrir en una nueva pestaña </div>
# # Data Wrangling Avanzado
#
# ## Tabla de Contenidos
#
# I. Data Wrangling Avanzado
#
# I. Tabla de Contenidos
#
# II. Table of Contents
#
# I. Introducción
# II. Pandas y Cadenas de caracteres
# I. count
# II. Ejercicio
# III. contains
# IV. lower, upper y title
# V. split
#
# III. ¿Cómo organizar la información? OLAP, OLTP y Tidy data
#
# I. OLTP y OLAP
# II. Tidy data
# I. Tuberculosis
# I. Tabla 1
# II. Tablas 2.a y 2.b
# III. Tabla 3
# IV. Tabla 4
# V. Tabla 5
# III. Melt
# I. Ejercicios
# II. Ejercicios
#
# IV. Series de Tiempo
#
# I. Datetime y metodos de manipulacion de series de tiempo
# II. Ventanas y Medias moviles
# III. Shift y Diff
# ## Introducción
#
# En esta clase vamos a ver cómo hacer manejo avanzado de datos a partir, tanto en cadenas de caracteres como en series de tiempo. El primer dataset que vamos a utilizar es del portal de datos abiertos de España.
# +
# # !pip install seaborn
# # !pip install numpy==1.18.5
# -
import pandas as pd
import numpy as np
df = pd.read_csv('https://datasets-humai.s3.amazonaws.com/datasets/parodesprov.csv')
df.head()
# Generalmente cuando en un dataset encontramos campos de texto es recomendable analizar si la carga se hizo con un buen criterio y no hay errores en la carga. Vamos a prestar atención a los campos Texto y PAD_1_COD_PROV.
df["Texto"].head()
# Noten que el primer valor incluye un "-" como separador
# Vamos a comenzar eliminando las filas que tienen null el campo Texto, para facilitar el análisis posterior.
df = df[~df["Texto"].isnull()]
# ## Pandas y Cadenas de caracteres
#
# Cuando una pd.Series es de tipo object Pandas permite acceder a métodos para operar sobre strings llamando al método **.str**.
# ### count
#
# El método **.str.count()** permite contar cuántas ocurrencias de un patrón hay en una Series. Para ello se puede pasar una regex. Las regex o expresiones regulares son una cadena de caracteres que define una búsqueda de un patrón. En esta clase no vamos a ahondar en regex pero sí podemos mencionar que:
# - Si pasamos un carácter en particular nos va a matchear cuántas veces aparece.
# - El carácter . funcion como comodín, con lo cual cualquier caracter va a matchear.
# Entonces, si queremos saber cuántas veces aparece el símbolo "-" en cada término podemos hacer:
df["Texto"].str.count("-").head()
# Ahora, nos llama la atención que el símbolo "-" aparece en la primera fila, ¿será algo común?
df["Texto"].str.count("-").value_counts()
# ### Ejercicio
#
# 1- Ver los casos que tiene 2 "-" en el nombre.
# 2- Graficar un histograma con la cantidad de caracteres en la columna Texto usando contains. Tip: pueden usar el método .hist.
import matplotlib.pyplot as plt
# %matplotlib inline
# 3- Mostrar que el resultado obtenido en el ejercicio anterior equivale a aplicar **.str.len()**. Para ello usen el método **pd.Series.equals()** y comparen el método usado en el ejercicio anterior con aplicar .str.len()
# ### contains
# Digamos que queremos encontrar nombres de ciudades vascas. Para ello sabemos que "herri" en Euskera significa lugar, con lo que esperamos encontrar "erri" en una gran cantidad de ciudades. Vamos a filtrar trayendo solamente las ciudades que incluyen "erri" dentro de su nombre con el método **.str.contains**.
df[df["Texto"].str.contains("erri",case=False)].head() # case es para considerar mayúsculas y minúsculas o no
df[df["Texto"].str.contains("erri",case=False)].shape
# ### lower, upper y title
# Supongamos que queremos estar seguros que las nombres están estandarizados... podemos elegir pasar todo a minúscula o a mayúscula de la siguiente manera:
df["Texto"].str.lower().head()
df["Texto"].str.upper().head()
# También podemos aplicar mayúscula a la primera letra y al resto minúscula, con **.str.title**. Este método toma en consideración espacios o símbolos no alfabéticos. Es decir, va a comenzar con mayúscula los términos después de un espacio, "-" o "/". Por ejemplo
df["Texto"][df["Texto"].str.count("-") == 2].str.title()
# ### split
# En el ejemplo anterior encontramos el uso de "/". En este caso se refiere a que el nombre de la ciudad (Oroz-Betelu/Orotz-Betelu) tiene dos nombres oficiales, uno en español y otro en vasco.
#
# Supongamos que queremos estandarizar el problema y quedarnos sólo con el nombre que aparece primero, ¿cómo podemos hacerlo?
# Primero veamos algunos ejemplos otros ejemplos de nombres con "/":
df["Texto"][df["Texto"].str.count("/") > 0].unique()[:10]
# Nota: en un primer momento uno podría pensar que en este dataset el primer elemento debería ser siempre el nombre en, por ejemplo, español y el segundo en vasco... Sin embargo, no es así :-S
#
# Vamos a usar **.str.split()**, vale la pena mencionar que el método devuelve una lista con un elemento por cada separación que pudo realizar. Para ello recibe como parámetro un string que es un patrón que va a usar, justamente, para dividir la cadena de caracteres. Comúnmente sólo vamos a pasarle un elemento por el cual splitear.
df["Texto"].str.split() # por default split separa por espacios en blanco
# Noten en el ejemplo anterior que si devuelve una lista con un solo elemento es que no matcheó y, por lo tanto, no dividió el string.
# Yendo a nuestro problema, separamos por el caracter '/'. Notar como se transforman los nombres compuestos
# MUCHO CUIDADO! Nuestros strings ahora son listas!! :o
df["Texto"].str.split("/").head()
# Entonces, nos quedamos con la primera versión del nombre al separar por "/"
# +
texto_para_split = df["Texto"][df["Texto"].str.count("/") > 0] # guardo en una variable casos con /
texto_para_split.str.split("/").apply(lambda x: x[0]) # para indexar las listas tengo que usar apply
# -
# Por otra parte, si quisiéramos guardar cada nombre en una columna separada podemos hacerlo usando la opción expand:
texto_para_split.str.split("/", expand=True)
# Podemos asignar el resultado de expand de la siguiente manera:
df[["nombre_1", "nombre_2"]] = df["Texto"].str.split("/", expand=True) # usamos el df original y expandimos
df[["nombre_1", "nombre_2"]].head() # vemos que hay nulls que nombre_2 porque tienen un solo nombre
df.loc[~df["nombre_2"].isnull(), ["nombre_1", "nombre_2"]].head() # noten el filtro booleano
# # ¿Cómo organizar la información? OLAP, OLTP y Tidy data
#
# Existen diversas formas de organizar la información. En general, qué vamos a hacer con la información es clave para entender cómo organizarla. Veamos algunos conceptos habituales a la hora de trabajar con datos.
# ## OLTP y OLAP
# La sigla OLTP viene de Online transaction processing y se refiere a transacciones que ocurren en tiempo real. Un ejemplo típico son los ATMs (automated teller machine), más conocidos como "cajeros automáticos".
#
# En este aspecto transacción tiene dos acepciones y ambas son válidas: por un lado, se procesan transacciones en término de bases de datos (que vamos a ver a continuación), por otro lado, se suele aplicar a transacciones económicas en donde se intercambian entidades económicas.
#
# Sistemas OLTP son la mayoría de los sistemas tradicionales que conocemos, especialmente los sistemas bancarios. Estos son sistemas transaccionales (en la primera acepción de arriba) porque intentan cumplir 4 objetivos (**ACID**):
#
# - Las transacciones son operaciones **atómicas**: se hacen por completo o no se hacen. Imagínense una transferencia bancaria, ésto requiere debitar en una cuenta y acreditar en otra. Si la operación no fuera atómica y fallara podríamos quedarnos en un estado inesperado en el cual se debite de la primera y no se acredite en la segunda. Para evitar ésto, la atomicidad garantiza que si la operación falla en alguna parte del proceso revertimos completamos la operación y no impactamos ningún cambio (hacemos *rollback*).
#
# - Toda transacción debe mantener la **consistencia** de la base de datos, es decir, debe respetar una serie de restricciones. Por ejemplo, podemos pensar que para que una tarjeta de crédito sea de extensión de una cuenta esa cuenta debe existir previamente. Otro ejemplo, puede ser que no pueden existir dos cuentas bancarias diferentes con el mismo número.
#
# - Además, los sistemas transaccionales (especialmente los sistemas OLTP) requieren garantizar el **aislamiento** de las operaciones. Es común que estos sistemas necesiten resolver miles de operaciones concurrentes (es decir, que suceden en simultáneo), el aislamiento consiste en que esas operaciones dejen en la base de datos el mismo estado que si las operaciones fueran secuenciales (es decir, una por vez). Incluso, si una de esas falla, no debería alterar el resultado.
#
#
# Para ésto, se implementan diferentes algoritmos a fin de asegurar que no se generen errores ni competencias entre usuarios que estan intentando acceder a los mismos registros a la vez.
# OLAP (Online Analytical Processing) por otra parte se refiere a todos los sistemas utilizados para analisis y reportes de negocios (Business Intelligence), en los cuales se realizan diferentes operaciones de agregacion sobre los datos, a fin de proveer a los usuarios con informacion relevante.
#
# Las herramientas OLAP permiten realizar análisis multidimensionales, tomando en consideración distintas dimensiones y métricas. En este sentido, está lo que se conoce como "cubo OLAP". El cubo OLAP es un array multidimensional que permite analizar la información vista desde distintos ángulos. Por ejemplo, podemos querer ver un reporte financiero por producto, por ciudad, por tiempo, etc. Cada uno de estos términos es una dimensión del análisis.
#
# Estos sistemas generalmente requieren procesos de carga y transformaciones masivas que pueden durar horas o días, y permiten presentan la información de un modo tal que el análisis es en tiempo real, no así la información que usa.
# ## Tidy data
# **Tidy data** es un trabajo escrito por <NAME> (de la empresa RStudio) que se ha difundido mucho, especialmente en la comunidad de R, sobre buenas prácticas a la hora de estructurar información tabular. Pueden consultar el trabajo acá: https://vita.had.co.nz/papers/tidy-data.pdf
#
# La información tabular consta de **filas** y **columnas**. Las columnas siempre tienen una etiqueta y las filas sólo a veces.
#
# Los *datasets* constan de **valores**, éstos pueden ser numéricos o no numéricos . En el caso en que sea un valor numérico representa una cantidad, si no es numérico es una cualidad. Además, los valores miden o caracterizan un determinado atributo (altura, peso, temperatura, etc.). Este atributo se conoce como **variable**. Por último, esa variable se corresponde con una determinada unidad observada. Generalmente llamamos a eso simplemente **observación** (por ejemplo, la persona a la que se le midió la altura).
#
# Dicho ésto, Wickham define un dataset *tidy* u ordenado como aquel que cumple la tercera forma normal de bases de datos, pero con un lenguaje más cercano al campo del análisis de datos y pensando en información contenida en una tabla, y no en una base de datos con muchas tablas. Las condiciones son:
#
# - Cada variable forma una columna.
# - Cada observación forma una fila.
# - Cada tipo de unidad observacional forma una tabla.
#
# Además, describe 5 de los errores más comunes a la hora de ordenar la información:
#
# - Los nombres de columna en vez de ser nombres de variables son valores
# - Muchas variables se guardan en una sola columna.
# - Las variables se guardan tanto en columnas como en filas.
# - Distintos tipos de unidades observacionales se guardan en una misma tabla.
# - Una única unidad observacional se almacena en distintas tablas.4
# ### Tuberculosis
#
# A continuación vamos a ver un dataset de tuberculosis representado de distintas maneras...
# #### Tabla 1
#
# En esta primera representación de la información vemos que una misma columna (type) contiene dos variables (cases y population)
# |country|year|type|count|
# | --- | --- | --- | --- |
# |Afghanistan|1999|cases|745|
# |Afghanistan|1999|population|19987071|
# |Afghanistan|2000|cases|2666|
# |Afghanistan|2000|population|20595360|
# |Brazil|1999|cases|37737|
# |Brazil|1999|population|172006362|
# #### Tablas 2.a y 2.b
# En este caso separamos la tabla en dos tablas. Una donde vemosla población y otra donde vemos la cantidad de casos... ¿Cuál es el problema acá?
#
# Si bien ésto puede parecer correcto noten que la unidad observacional en realidad es un país en un año determinado... Con lo cual, lo que estamos haciendo acá es tener la misma unidad observacional en dos tablas. Además, los valores de la variable year están como nombres de columnas...
#
# |country|1999|2000|
# | --- | --- | --- |
# |Afghanistan|745|2666|
# |Brazil|37737|80488|
# |China|212258|213766|
# |country|1999|2000|
# | --- | --- | --- |
# |Afghanistan|19987071|20595360|
# |Brazil|172006362|174504898|
# |China|1272915272|1280428583|
# #### Tabla 3
#
# En este caso, perdimos las variables cases y population y calculamos un ratio. Si bien ésto cumple con ser "ordenado" estamos perdiendo las variables originales.
# |country|year|rate|
# | --- | --- | --- |
# |Afghanistan|1999|745/19987071|
# |Afghanistan|2000|2666/20595360|
# |Brazil|1999|37737/172006362|
# |Brazil|2000|80488/174504898|
# |China|1999|212258/1272915272|
# |China|2000|213766/1280428583|
# #### Tabla 4
#
# En la tabla 4 vemos que la variable *year*, *cases* y *population* se juntaron, con lo cual los valores de *year* pasan a formar parte de los nombres de las columnas.
# |country|cases_1999|cases_2000|population_1999|population_2000|
# | --- | --- | --- | --- | --- |
# |Afghanistan|745|19987071|2666|20595360|
# |Brazil|37737|172006362|80488|174504898|
# |China|212258|1272915272|213766|1280428583|
# #### Tabla 5
#
# Finalmente, la tabla *tidy* es:
# |country|year|cases|population|
# | --- | --- | --- | --- |
# |Afghanistan|1999|745|19987071|
# |Afghanistan|2000|2666|20595360|
# |Brazil|1999|37737|172006362|
# |Brazil|2000|80488|174504898|
# |China|1999|212258|1272915272|
# |China|2000|213766|1280428583|
# ## Melt
#
# Una de los métodos más útiles de Pandas para pasar de un formato *wide* o *ancho* como el de la tabla 4 a uno *largo* como el de la tabla 5 es **.melt**.
#
# Para aplicar este método vamos a obtener un dataset de Billboard sobre las canciones mas escuchadas del 2000.
#
# Vamos a agregar la opcion de encoding para solucionar un problema con los caracteres usados.
billboard = pd.read_csv('https://raw.githubusercontent.com/hadley/tidy-data/master/data/billboard.csv', warn_bad_lines=False, error_bad_lines=False, encoding='iso-8859-1')
# ### Ejercicios
# 1- Analicen el dataset, vean cuáles son las variables, qué es la observación. ¿Es necesario separarlo en dos tablas? Nota: lo que se está midiendo en las columnas x1st.week a x76th.week es la posición en el ranking de las 100 canciones más escuchadas en esa semana. Es decir, x1st.week es la posición en el ranking durante la primera semana que esa canción fue top 100.
# 2- ¿Por qué creen que hay tantas columnas con valores nulos?
# 3- ¿Cómo podríamos hacer este dataset más ordenado? (no miren la continuación de la notebook :-S)
# Veamos primero como luce el dataset
billboard.head()
# Lo que querríamos es poder mantener todas las columnas excepto las columnas que incluyen "week" en el nombre. Para poder tener un formato "tidy" u "ordenado" deberíamos tener, por un lado, una columna week que indique el número de la semana (y preferentemente que sea un entero), y por otro lado una columna ranking con el valor del ranking en esa semana.
#
# Para lograr ésto, vamos a usar la función **.melt**. Esta función recibe el DataFrame y vamos a usar el parámetro id_vars para pasarle la lista de columnas que van a ser constantes, es decir, que no van a variar. Con las demás columnas .melt va a:
#
# 1- tomar los nombres de columnas no incluídas en id_vars y convertirlos en una columna
#
# 2- va a tomar los valores de esas columnas y convertirlos en una segunda columna.
pd.melt(billboard,
id_vars=["year", "artist.inverted", "track", "time", "genre", "date.entered", "date.peaked"]).head()
# Ahora, querríamos que variable se llame week y value se llame ranking, para eso hacemos...
billboard_2 = pd.melt(billboard,
id_vars=["year", "artist.inverted", "track", "time", "genre", "date.entered", "date.peaked"],
var_name="week",
value_name="ranking")
billboard_2.head()
# ### Ejercicios
# 1- Quédense sólo con la parte numérica de week sin usar regex, y conviertan a número los valores de la columna.
# 2- Conviertan los valores de la columna ranking a entero. Tip: prueben usar pd.isna() para saber si un valor es nulo o no.
# 3- Vean cuántas filas totales hay y eliminen las filas con nulos, ahora vuelvan a ver cuántas filas quedan.
# # Series de Tiempo
# Las series de tiempo representan uno de los problemas mas interesantes en la ciencia de datos ya que refiere a eventos continuos y ordenados los cuales pueden ser independientes o tener alguna correlacion entre si. En este modulo veremos los metodos mas utilizados para manipular series de tiempo en pandas
# ## Datetime y metodos de manipulacion de series de tiempo
#
# Para esta seccion vamos a utilizar el dataset de consumo energetico de Alemania.
df_energia = pd.read_csv('https://raw.githubusercontent.com/jenfly/opsd/master/opsd_germany_daily.csv')
# Primeramente vamos a evaluar el dataset
df_energia.columns = ['Fecha', 'Consumo', 'Eolica', 'Solar', 'Suma']
df_energia.tail(10)
# Las columnas son las siguientes:
# - Date — La fecha (yyyy-mm-dd)
# - Consumo — Electricidad Consumida en GWh
# - Eolica — Produccion de energia eolica en GWh
# - Solar — Produccion de energia solar en GWh
# - Suma — Suma de las dos anteriores GWh
# Antes de comenzar a explorar el dataset, veamos algunas funciones de pandas para crear fechas. El metodo to_datetime nos permite transformar un string en cierto formato a un objeto del tipo Timestamp, el cual consiste de una fecha y una hora. Podemos ver que acepta varios formatos
pd.to_datetime('2018-01-15 3:45pm')
pd.to_datetime('7/8/1952')
# Vamos a convertir el dtype de nuestra columna Fecha de object a datetime64
df_energia.info()
df_energia["Fecha"] = pd.to_datetime(df_energia["Fecha"])
df_energia.info()
# Convertir la columna *Fecha* en datetime nos permite filtrar usando la información temporal...
df_energia.head()
# Podemos traernos los casos posteriores a 2009, por default a partir del 1ero de enero...
df_energia[df_energia["Fecha"] > "2009"].head()
# Con el método **.between** podemos filtrar por un rango de fechas...
df_energia[df_energia["Fecha"].between("2008", "2010")].head()
# En vez de filtrar usando años podemos pasar la fecha completa (siempre tengan presente el formato de la fecha)...
df_energia[df_energia["Fecha"].between("2008-01-01", "2010-01-22")].head()
# Y como si fuera poco también podemos filtrar sólo usando el año y el mes, sin especificar el día...
df_energia[df_energia["Fecha"].between("2008-01", "2010-02")].head()
df_energia[df_energia["Fecha"].between("2008-01", "2010-02")].tail()
# Ahora, vamos a convertir nuestra columna Fecha en un índice temporal... Esto va a crear un nuevo tipo de objeto llamado DatetimeIndex
df_energia = df_energia.set_index("Fecha")
df_energia.tail(10)
df_energia.index
# Primeramente vamos a agregar columnas que nos proporcionen mas informacion sobre las fechas
df_energia['Anio'] = df_energia.index.year
df_energia['Mes'] = df_energia.index.month
df_energia['Dia'] = df_energia.index.day_name()
df_energia.sample(5, random_state=0)
# Como creamos el indice por las fechas, podemos localizar cualquier dia que querramos ahora
df_energia.loc['2017-08-10']
# Incluso podemos buscar rangos de fechas
df_energia.loc['2014-01-20':'2014-01-22']
# O buscar por algun mes en particular
df_energia.loc['2016-05']
# Ahora veamos como se ve nuestra data de consumo de energia
# #! pip install seaborn
import matplotlib.pyplot as plt
import seaborn as sns
df_energia['Consumo'].plot(linewidth=0.5);
# ## Ventanas y Medias moviles
#
# Las ventanas moviles se refiere a aplicar alguna operacion de agregacion, por ejemplo el promedio sobre un conjunto de datos ordenados a la vez, por ejemplo el promedio de los ultimos 6 dias sobre cada conjunto ordenado de 6 dias en el dataset. Veamos un ejemplo para entenderlo mejor
opsd_7d = df_energia["Consumo"].rolling(6).mean()
opsd_7d.head(10)
# Podemos observar como las primeras 5 mediciones son Nan, ya que utiliza los primeros 5 valores para calcular a partir del 6to dia la media. El valor en el 7mo dia va a ser calculado con los datos del 2do al 6to dia, y asi sucesivamente.
# Este tipo de metodos son particularmente utiles en analisis financiero.
# ## Shift y Diff
#
# La operacion de Shift, como su nombre lo dice traducido al castellano, desplazar los datos una cantidad N de periodos. Veamoslo con el ejemplo del consumo, suponiendo que quiero crear otra columna con el consumo total del periodo anterior para poder compararlo con el actual
df_energia['ConsumoAyer'] = df_energia['Suma'].shift(periods=1)
# Veamos como quedo ahora el dataset
df_energia.tail(10)
# Podemos observar como la columna 'ConsumoAyer' es el valor de la columna 'Suma' exactamente del periodo anterior. Esto se puede realizar con tantos periodos como uno desee, pero hay que tener cuidado porque al inicio de nuestro dataset van a quedar valores Nan por la cantidad de periodos que elijamos. Hay que evaluar con que valor completamos esos valores.
# Por ultimo supongamos que deseamos hacer la diferencia fila a fila del valor de ayer con el valor de hoy para el consumo total. Aqui nos conviene utilizar el metodo diff.
df_energia['Diferencia'] = df_energia['Suma'].diff(1)
df_energia.tail(10)
| AnalisisDeDatos/4_Data_Wrangling_Avanzado/data_wrangling_avanzado.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TEXT MINING for PRACTICE: Python Data Structure (자료구조)
# ---
# ## 1. List
# +
# List 선언 및 초기화
x = [] # 또는 list()로도 생성할 수 있습니다.
x = [1,'a',True]
print(x)
print(type(x))
# -
# List 원소 접근 (READ)
x = [1,2,3]
print(x[0])
print(type(x[0]))
# List 원소 추가 (WRITE)
x = [1,2,3]
print(x)
x.append(4)
print(x)
# List 원소 변경 (UPDATE)
x = [1,2,3,4]
print(x)
x[2] = 100
print(x)
# List 원소 삭제 (DELETE)
x = [1,2,3,4]
print(x)
del x[2]
print(x)
# 원소 존재여부 확인
1 in x
# ### 1-1 List간 연산
a = [1,2,3]
b = [4,5,6]
c = a+b
print(c)
c.append(b)
print(c)
x = [1,2,3,4,5]
# List Sub-list 접근-2
print(x[0:2])
print(type(x[0:2]))
x = [1,9,2,8,3,7,4,6,5]
print(x)
x=sorted(x)
print(x)
x=sorted(x,reverse=True)
print(x)
print(min(x))
print(max(x))
print(sum(x))
print(len(x))
# +
# Q. list의 평균은?
# +
# Q. 리스트에 숫자가 아닌 원소가 있을때 연산은 어떻게 될까?
# -
# ## 2. Dictionary
x = {} # x = dict()
print(x)
print(type(x))
x = {'a':1, 'b':2, 'c':3} # x = dict()
print(x)
print(x['a'])
x['d']=4
print(x)
x['d']=10
print(x)
del x['d']
print(x)
'd' in x
x = {'a':3, 'b':2, 'c':1}
print(x.keys())
print(x.values())
print(x.items())
sorted_x = sorted(x.items(), key=lambda kv: kv[0], reverse=True)
print(sorted_x)
sorted_x = sorted(x.items(), key=lambda kv: kv[1])
print(sorted_x)
# Q. Value가 오름차순으로 정렬하되, Value가 같을때는 Key를 기준으로 정렬 하려면?
x = {'a':3, 'b':2, 'c':1, 'd':3}
sorted_x = sorted(x.items(), key=lambda kv: (kv[1], kv[0]))
print(sorted_x)
# ## 3. Set
x = {1,2,3} # set()
type(x)
# READ
for a in x:
print(a)
# INSERT
x.add(1)
x.add(4)
x.add(5)
x.add(6)
print(x)
# UPDATE
x.update([1,2,7])
print(x)
# DELETE
x.remove(7)
print(x)
# +
a = {1,2,3,4,5}
b = {4,5,6,7,8}
#교집합
print(b.intersection(a))
print(a.intersection(b))
#합집합
print(a.union(b))
print(b.union(a))
#차집합
print(a.difference(b))
print(b.difference(a))
# -
x = ['Korea', 'Canada', 'Japan', 'USA', 'Korea', 'Italy']
print(x, type(x))
x = set(x)
print(x, type(x))
# ## 4. Tuple
x = () # x = tuple()
type(x)
x = (100, 10, 20)
print(x[0])
# INSERT
x = tuple()
x[0]=1
# UPDATE
x=(1,2)
x[0]=3
x = (1,2,3)
del x[0]
x = (1,2,3)
del x
print(x)
x = ("Apple", "Banana", "Orange", "Orange")
x.count('Orange')
x.index('Apple')
# ## 5. 자료구조간의 타입 캐스팅(casting)
# +
a = [1,2,3,4]
print(a, type(a))
set_a = set(a)
print(set_a, type(set_a))
tuple_a = tuple(a)
print(tuple_a, type(tuple_a))
dict_a = dict(a)
print(dict_a, type(dict_a))
# +
a = {'a':1, 'b':2, 'c':3}
print(a, type(a))
set_a = set(a)
print(set_a, type(set_a))
list_a = list(a)
print(list_a, type(list_a))
tuple_a = tuple(a)
print(tuple_a, type(tuple_a))
list_a = list(a.items())
set_a = set(a.items())
tuple_a = tuple(a.items())
print(list_a)
print(set_a)
print(tuple_a)
# -
| practice/week-01/W01_2_python_data_structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="P8wEOtbOLpvK"
# # Play with TensorBoard
# > TensorBoard provides the visualization and tooling needed for machine learning experimentation
#
# - toc: true
# - badges: true
# - author: <NAME>
# - categories: [tensorboard]
# + [markdown] id="zFlI6Q7bYo4q"
# TensorBoard is TensorFlow's visualization toolkit, enabling ones to track metrics like loss and accuracy, visualize the model graph, view histograms of weights, biases, or other tensors as they change over time, and much more.
#
# + id="w1VFbYXoIwVH"
#collapse-hide
# %load_ext tensorboard
import os
import numpy as np
import pandas as pd
import datetime
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow import keras
from tensorflow.keras import layers
pd.options.display.max_columns=25
# + colab={"base_uri": "https://localhost:8080/"} id="_M3cWtmeKvID" outputId="1c72110d-eb73-4b96-e0e8-f62a033ece7c"
df = pd.read_csv("kc_house_data.csv")
df.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 700} id="Us0b85OpLZCY" outputId="59b6b400-fc57-4fe4-e7a2-ffa2225cabda"
df.head().T
# + colab={"base_uri": "https://localhost:8080/"} id="-nrKrtubLdaA" outputId="3b76c216-7c98-4098-8b30-1d3fbd092f10"
df.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 731} id="EuMVLx_FMCFO" outputId="c1b9c5ba-7def-4d0a-cb84-b0502998eef3"
df['year'] = pd.to_numeric(df['date'].str.slice(0,4))
df['month'] = pd.to_numeric(df['date'].str.slice(4,6))
df['day'] = pd.to_numeric(df['date'].str.slice(6,8))
df.drop(['id', 'date'], axis="columns", inplace=True)
df.head().T
# + id="9LSp6RuQMnyf"
n = df.shape[0]
ids = np.random.permutation(n)
train_ids = ids[:int(n * .6)]
valid_ids = ids[int(n * .4) : int(n * .8)]
test_ids = ids[int(n * .8):]
train_data = df.loc[train_ids]
valid_data = df.loc[valid_ids]
test_data = df.loc[test_ids]
# + id="W3AwV47vNac1"
train_valid_data = pd.concat([train_data, valid_data])
mean = train_valid_data.mean()
std = train_valid_data.std()
train_data = (train_data - mean) / std
valid_data = (valid_data - mean) / std
# + id="EzDr_lgcN0jA"
train_x = np.array(train_data.drop('price', axis='columns')).astype('float32')
train_y = np.array(train_data['price']).astype('float32')
valid_x = np.array(valid_data.drop('price', axis='columns')).astype('float32')
valid_y = np.array(valid_data['price']).astype('float32')
# + colab={"base_uri": "https://localhost:8080/"} id="H9G0UNg1OKIj" outputId="2a79bb85-b9d2-434e-85c8-bda73f48e4a1"
train_x.shape, valid_x.shape
# + id="-DQhfNETOWN_"
model = tf.keras.Sequential(name='model-1')
model.add(tf.keras.layers.Dense(64, activation='relu', input_shape=(21,)))
model.add(tf.keras.layers.Dense(64, activation='relu'))
model.add(tf.keras.layers.Dense(1))
# + colab={"base_uri": "https://localhost:8080/"} id="-0hiWpigO0Y5" outputId="ab73512c-91b8-46ec-bfce-2889834c4682"
model.summary()
# + id="-XUOpbo6Oy7A"
model.compile(tf.keras.optimizers.Adam(0.001),
loss = tf.keras.losses.MeanSquaredError(),
metrics=[tf.keras.metrics.MeanAbsoluteError()])
# + id="lJXgPECIPIxm"
log_dir = "logs/model_1/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir)
checkpoint_callback = tf.keras.callbacks.ModelCheckpoint('models/best-model-1.h5',
monitor='val_mean_absolute_error',
save_best_only=True,
mode='min')
# + colab={"base_uri": "https://localhost:8080/"} id="g-lNKjs-UKcL" outputId="94b4ac99-0084-45dd-f00c-d13edb4ba021"
#collapse_output
history = model.fit(train_x, train_y,
batch_size=64,
epochs=300,
validation_data=(valid_x, valid_y),
callbacks=[tensorboard_callback,
checkpoint_callback])
# + colab={"base_uri": "https://localhost:8080/"} id="ft-cea13jLpZ" outputId="0c5a2a35-a794-4b02-f423-bff51cbbec14"
#hide
# !tensorboard dev upload \
# --logdir logs/model_1 \
# --name "Blog: Play with TensorBoard" \
# --description "Simple comparison of several hyperparameters" \
# --one_shot
# + colab={"base_uri": "https://localhost:8080/", "height": 820} id="-gewwKYjV1Yy" outputId="500cf5d9-4a3e-49a2-8f21-dc3894cb6b80"
#collapse
from IPython import display
display.IFrame(
src="https://tensorboard.dev/experiment/MVQyms8BSVym5wG2ETdDyA/",
width = "100%",
height="800px")
# + id="v0ME0PtGbZBR"
| _notebooks/2021-02-20-play-with-tensorboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # UTF-8 Embedding and Decoder Test with Autoencoder
#
# This file is dedicated to test Overfitting Autoencoder to compress the size of the input embedding from the multi-hot to a dense vector
#
#
import langmodels.utf8codec as utf8codec
from langmodels.utf8codec import *
import torch.nn.functional as F
import torch.nn as nn
import torch
# load the codebook and all the dictionaries mapping the data
code_matrix, txt2code, code2txt, txt2num, num2txt = utf8codec._load_codebook()
type(num2txt)
all_data = np.array(list(num2txt.keys()))
all_data = all_data.reshape((-1,1))
# +
# all_data
# +
# np.random.shuffle(all_data)
# +
# all_data
# -
def _prepare_overfit_batch(num2txt, batch_size):
"""
The idea is to prepare the list of all the numbers in batches, the batches are randomly mixed to avoid issues.
each batch contains:
(batch size, seq width, index) ??
(batch size, index) ??
:param num2txt: numeric index 2 string conversion dictionary containing the entire vocabulary
:return:
"""
# assert type(num2txt) == 'dict'
all_data = np.array(list(num2txt.keys()))
all_data = all_data.reshape((-1,1))
# print(all_data.shape)
# assume that we can hold all in memory
arr = []
for i in range(batch_size):
data = np.copy(all_data)
# print(data.shape)
np.random.shuffle(data)
# print(data.shape)
arr.append(data.transpose())
ret = np.stack(arr, axis=1)
ret = ret.reshape(batch_size,-1)
# print(ret.shape)
return ret
# +
# # %%time
# btch = _prepare_overfit_batch(num2txt, 100)
# btch = utf8codec._prepare_overfit_batch(num2txt, 100)
# +
def train_overfit(model, optimizer, loss_function, batches, epoch, device, log_interval=10):
# model.train()
train_loss = 0
batch_loss = []
batch_idx = 0
for b in batches:
tensor_data = torch.from_numpy(b).to(device).long() #.double() #.float()
optimizer.zero_grad()
# emb is obtained from the the pre-computed utf8codebook
emb, res = model(tensor_data)
# print(emb.shape,emb.dtype, res.shape, res.dtype)
loss = loss_function(emb, res)
loss.backward()
train_loss += loss.data.item() # [0]
optimizer.step()
if batch_idx % log_interval == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx , len(batches),
100. * batch_idx / len(tensor_data),
train_loss / len(tensor_data)))
batch_loss.append(train_loss)
batch_idx += 1
print('====> Epoch: {} Average loss: {:.8f}'.format(epoch, train_loss / len(batches)))
return batch_loss
def test(model, test_data, epoch, device):
model.eval()
test_loss = 0
for d in test_data:
tensor_data = torch.from_numpy(d).to(device)
res = model(data)
test_loss += loss_function(tensor_data, res).data.item() # [0]
test_loss /= len(test_data)
print('epoch: {}====> Test set loss: {:.4f}'.format(epoch, test_loss))
# -
# from https://stackoverflow.com/questions/434287/what-is-the-most-pythonic-way-to-iterate-over-a-list-in-chunks
def chunker(seq, size):
return (seq[pos:pos + size] for pos in range(0, len(seq), size))
# +
# def _get_activation_fn(activation):
# if activation == "sigmoid":
# return F.sigmoid
# elif activation == "tanh":
# return F.tanh
# elif activation == "relu":
# return F.relu
# elif activation == "gelu":
# return F.gelu
# else:
# return None
# # raise RuntimeError("activation should be sigmoid/tanh/relu/gelu, not %s." % activation)
# -
model = UTF8Autoencoder(code_matrix, dim=64)
# prepare many batches so I have everything ready to train
nbatches = 4000
batch_size = 64
batches = []
# %%time
for i in range(nbatches):
btch = _prepare_overfit_batch(num2txt, batch_size)
batches.append(btch)
len(batches)
# +
# encoder(batches[0])
# -
epochs = chunker(batches, batch_size)
device = device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model = model.to(device) #.float()
# optimizer = torch.optim.Adam(model.parameters(), lr=1e-6, weight_decay=0, amsgrad=False )
optimizer = torch.optim.AdamW(model.parameters())
# optimizer = torch.optim.AdamW(model.parameters(), lr=2e-4, weight_decay=1e-4)
loss_function = F.mse_loss
# loss_function = F.cross_entropy # nn.CrossEntropyLoss()
# loss_function = nn.NLLLoss()
# loss_function = F.kl_div # KL divergence
epoch_loss = []
# %%time
epoch_count = 1
for e in epochs:
eloss = train_overfit(model, optimizer, loss_function, e, epoch_count, device, log_interval=10)
epoch_count+=1
# if epoch_count == 20:
# print("epoch {} decreasing learning_rate to {}".format(epoch_count, 1e-5))
# optimizer = torch.optim.AdamW(model.parameters(), lr=1e-5, weight_decay=1e-4)
epoch_loss.append(eloss)
# ### For an encoding of dimension 32
# It seems to be processing about 1.5M chars/sec in my gtx1080
#
# In [61]: 4000*1984*100
# Out[61]: 793600000
#
# In [62]: _ / (60*8+49)
# Out[62]: 1500189.0359168241
#
# And wall time about 1.1M chars/sec:
#
# In [63]: 4000*1984*100
# Out[63]: 793600000
#
# In [64]: _ / (60*11+26)
# Out[64]: 1156851.3119533528
#
#
# ====> Epoch: 40 Average loss: 0.00016435
# CPU times: user 8min 49s, sys: 2min 38s, total: 11min 28s
# Wall time: 11min 26s
#
#
# ### For an encoding of dimension 48
#
# number of batches: 4000
#
# ====> Epoch: 63 Average loss: 0.00001490
# CPU times: user 8min 36s, sys: 2min 33s, total: 11min 10s
# Wall time: 11min 10s
#
# Loss has gotten down by a wide margin and processing time seems about the same for this network.
#
# The loss is much less with dimension 48 instead of 32 a lot less epochs, it seems that loss could be made less with more epochs
#
# ### For an encoding of dimension 64
#
# number of batches 8000
#
# ====> Epoch: 125 Average loss: 0.00000317
# CPU times: user 17min 38s, sys: 5min 17s, total: 22min 56s
# Wall time: 22min 56s
#
# Processing time rests the same while loss goes down by another order of magnitude.
#
#
# The issue with dimensionality is that good things start to happen with big dimensions due to the exponential growth in representational power, so for low dimensions many things won't work. The point is to find a balance with vectors of dimension big enough to make the representational power sufficient, and small enough to make it work in my PCs GPU once the network starts to grow with the next iterations on the complexity of the networks.
# +
# len(epoch_loss), len(epoch_loss[-1])
# -
model.save_model("2segments_d64", "trained_models")
# Now what needs to be worked on (with the current model already pre-trained to overfitting for the mapping) is to actually decode to index and character to see the kind of errors in decoding end-to-end.
#
# For this I have to make the decoder from the utf8codebook embedding to the code index and then to the utf-8 character to visually analyze the kind of errors.
#
# +
#TODO FIXME the actual loss reporting is broken and results are BAD, so I have to make something better and include tensorboard
# -
#
| predictors/sequence/text/utf8codec_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2 as cv
import numpy as np
import matplotlib.pyplot as plt
img = cv.imread('line.png')
gray_img = cv.cvtColor(img, cv.COLOR_RGB2GRAY)
thresh, thresh_img = cv.threshold(gray_img,127,255,cv.THRESH_BINARY_INV)
thresh_img = np.asarray(thresh_img)
thresh_img = thresh_img / 255
# -
# algorithm 4
def baseline_index(thresh_img):
projection = np.sum(thresh_img,axis=1).tolist()
print(projection)
plt.plot(projection)
plt.show()
peakValues = []
for i in range(1,len(projection)):
if i+1 < len(projection) and projection[i] > projection[i+1] and projection[i] > projection[i-1]:
peakValues.append(projection[i])
print(peakValues)
np.max(peakValues)
baseLineIndex = 0
# +
import skimage as io
listOfGaps = []
listOfGapsLength = []
Words = [] # list of segmented words
isWord = 0
projectionArray = np.asarray(projection)
for i in range(len(projectionArray)):
if projectionArray[i] != 0.0 and isWord == 0:
listOfGaps.append(i)
isWord = 1
elif projectionArray[i] == 0.0 and isWord == 1:
listOfGaps.append(i)
isWord = 0
print(len(listOfGaps))
word = []
listOfGapsIndex = 0
imgArray = np.asarray(img)
print(imgArray.shape)
for i in range(imgArray.shape[1]):
if i == listOfGaps[listOfGapsIndex]:
imgTemp = imgArray[:,listOfGaps[listOfGapsIndex]:listOfGaps[listOfGapsIndex + 1]]
word.append(imgTemp)
cv.imshow('label',imgTemp)
cv.waitKey(0)
cv.destroyAllWindows()
listOfGapsIndex += 2
Words.append(word)
if len(listOfGaps) <= listOfGapsIndex:
break
print(Words)
# -
| .ipynb_checkpoints/mina-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to OpenFermion
# Note that all the examples below must be run sequentially within a section.
# ## Initializing the FermionOperator data structure
#
# Fermionic systems are often treated in second quantization where arbitrary operators can be expressed using the fermionic creation and annihilation operators, $a^\dagger_k$ and $a_k$. The fermionic ladder operators play a similar role to their qubit ladder operator counterparts, $\sigma^+_k$ and $\sigma^-_k$ but are distinguished by the canonical fermionic anticommutation relations, $\{a^\dagger_i, a^\dagger_j\} = \{a_i, a_j\} = 0$ and $\{a_i, a_j^\dagger\} = \delta_{ij}$. Any weighted sums of products of these operators are represented with the FermionOperator data structure in OpenFermion. The following are examples of valid FermionOperators:
#
# $$
# \begin{align}
# & a_1 \nonumber \\
# & 1.7 a^\dagger_3 \nonumber \\
# &-1.7 \, a^\dagger_3 a_1 \nonumber \\
# &(1 + 2i) \, a^\dagger_4 a^\dagger_3 a_9 a_1 \nonumber \\
# &(1 + 2i) \, a^\dagger_4 a^\dagger_3 a_9 a_1 - 1.7 \, a^\dagger_3 a_1 \nonumber
# \end{align}
# $$
#
# The FermionOperator class is contained in $\textrm{ops/_fermion_operators.py}$. In order to support fast addition of FermionOperator instances, the class is implemented as hash table (python dictionary). The keys of the dictionary encode the strings of ladder operators and values of the dictionary store the coefficients. The strings of ladder operators are encoded as a tuple of 2-tuples which we refer to as the "terms tuple". Each ladder operator is represented by a 2-tuple. The first element of the 2-tuple is an int indicating the tensor factor on which the ladder operator acts. The second element of the 2-tuple is Boole: 1 represents raising and 0 represents lowering. For instance, $a^\dagger_8$ is represented in a 2-tuple as $(8, 1)$. Note that indices start at 0 and the identity operator is an empty list. Below we give some examples of operators and their terms tuple:
#
# $$
# \begin{align}
# I & \mapsto () \nonumber \\
# a_1 & \mapsto ((1, 0),) \nonumber \\
# a^\dagger_3 & \mapsto ((3, 1),) \nonumber \\
# a^\dagger_3 a_1 & \mapsto ((3, 1), (1, 0)) \nonumber \\
# a^\dagger_4 a^\dagger_3 a_9 a_1 & \mapsto ((4, 1), (3, 1), (9, 0), (1, 0)) \nonumber
# \end{align}
# $$
#
# Note that when initializing a single ladder operator one should be careful to add the comma after the inner pair. This is because in python ((1, 2)) = (1, 2) whereas ((1, 2),) = ((1, 2),). The "terms tuple" is usually convenient when one wishes to initialize a term as part of a coded routine. However, the terms tuple is not particularly intuitive. Accordingly, OpenFermion also supports another user-friendly, string notation below. This representation is rendered when calling "print" on a FermionOperator.
#
# $$
# \begin{align}
# I & \mapsto \textrm{""} \nonumber \\
# a_1 & \mapsto \textrm{"1"} \nonumber \\
# a^\dagger_3 & \mapsto \textrm{"3^"} \nonumber \\
# a^\dagger_3 a_1 & \mapsto \textrm{"3^}\;\textrm{1"} \nonumber \\
# a^\dagger_4 a^\dagger_3 a_9 a_1 & \mapsto \textrm{"4^}\;\textrm{3^}\;\textrm{9}\;\textrm{1"} \nonumber
# \end{align}
# $$
#
# Let's initialize our first term! We do it two different ways below.
# +
from openfermion.ops import FermionOperator
my_term = FermionOperator(((3, 1), (1, 0)))
print(my_term)
my_term = FermionOperator('3^ 1')
print(my_term)
# -
# The preferred way to specify the coefficient in openfermion is to provide an optional coefficient argument. If not provided, the coefficient defaults to 1. In the code below, the first method is preferred. The multiplication in the second method actually creates a copy of the term, which introduces some additional cost. All inplace operands (such as +=) modify classes whereas binary operands such as + create copies. Important caveats are that the empty tuple FermionOperator(()) and the empty string FermionOperator('') initializes identity. The empty initializer FermionOperator() initializes the zero operator.
# +
good_way_to_initialize = FermionOperator('3^ 1', -1.7)
print(good_way_to_initialize)
bad_way_to_initialize = -1.7 * FermionOperator('3^ 1')
print(bad_way_to_initialize)
identity = FermionOperator('')
print(identity)
zero_operator = FermionOperator()
print(zero_operator)
# -
# Note that FermionOperator has only one attribute: .terms. This attribute is the dictionary which stores the term tuples.
my_operator = FermionOperator('4^ 1^ 3 9', 1. + 2.j)
print(my_operator)
print(my_operator.terms)
# ## Manipulating the FermionOperator data structure
# So far we have explained how to initialize a single FermionOperator such as $-1.7 \, a^\dagger_3 a_1$. However, in general we will want to represent sums of these operators such as $(1 + 2i) \, a^\dagger_4 a^\dagger_3 a_9 a_1 - 1.7 \, a^\dagger_3 a_1$. To do this, just add together two FermionOperators! We demonstrate below.
# +
from openfermion.ops import FermionOperator
term_1 = FermionOperator('4^ 3^ 9 1', 1. + 2.j)
term_2 = FermionOperator('3^ 1', -1.7)
my_operator = term_1 + term_2
print(my_operator)
my_operator = FermionOperator('4^ 3^ 9 1', 1. + 2.j)
term_2 = FermionOperator('3^ 1', -1.7)
my_operator += term_2
print('')
print(my_operator)
# -
# The print function prints each term in the operator on a different line. Note that the line my_operator = term_1 + term_2 creates a new object, which involves a copy of term_1 and term_2. The second block of code uses the inplace method +=, which is more efficient. This is especially important when trying to construct a very large FermionOperator. FermionOperators also support a wide range of builtins including, str(), repr(), ==, !=, *=, *, /, /=, +, +=, -, -=, - and **. Note that since FermionOperators involve floats, == and != check for (in)equality up to numerical precision. We demonstrate some of these methods below.
# +
term_1 = FermionOperator('4^ 3^ 9 1', 1. + 2.j)
term_2 = FermionOperator('3^ 1', -1.7)
my_operator = term_1 - 33. * term_2
print(my_operator)
my_operator *= 3.17 * (term_2 + term_1) ** 2
print('')
print(my_operator)
print('')
print(term_2 ** 3)
print('')
print(term_1 == 2.*term_1 - term_1)
print(term_1 == my_operator)
# -
# Additionally, there are a variety of methods that act on the FermionOperator data structure. We demonstrate a small subset of those methods here.
# +
from openfermion.utils import commutator, count_qubits, hermitian_conjugated, normal_ordered
# Get the Hermitian conjugate of a FermionOperator, count its qubit, check if it is normal-ordered.
term_1 = FermionOperator('4^ 3 3^', 1. + 2.j)
print(hermitian_conjugated(term_1))
print(term_1.is_normal_ordered())
print(count_qubits(term_1))
# Normal order the term.
term_2 = normal_ordered(term_1)
print('')
print(term_2)
print(term_2.is_normal_ordered())
# Compute a commutator of the terms.
print('')
print(commutator(term_1, term_2))
# -
# ## The QubitOperator data structure
# The QubitOperator data structure is another essential part of openfermion. As the name suggests, QubitOperator is used to store qubit operators in almost exactly the same way that FermionOperator is used to store fermion operators. For instance $X_0 Z_3 Y_4$ is a QubitOperator. The internal representation of this as a terms tuple would be $((0, \textrm{"X"}), (3, \textrm{"Z"}), (4, \textrm{"Y"}))$. Note that one important difference between QubitOperator and FermionOperator is that the terms in QubitOperator are always sorted in order of tensor factor. In some cases, this enables faster manipulation. We initialize some QubitOperators below.
# +
from openfermion.ops import QubitOperator
my_first_qubit_operator = QubitOperator('X1 Y2 Z3')
print(my_first_qubit_operator)
print(my_first_qubit_operator.terms)
operator_2 = QubitOperator('X3 Z4', 3.17)
operator_2 -= 77. * my_first_qubit_operator
print('')
print(operator_2)
# -
# ## Jordan-Wigner and Bravyi-Kitaev
# openfermion provides functions for mapping FermionOperators to QubitOperators.
# +
from openfermion.ops import FermionOperator
from openfermion.transforms import jordan_wigner, bravyi_kitaev
from openfermion.utils import eigenspectrum, hermitian_conjugated
# Initialize an operator.
fermion_operator = FermionOperator('2^ 0', 3.17)
fermion_operator += hermitian_conjugated(fermion_operator)
print(fermion_operator)
# Transform to qubits under the Jordan-Wigner transformation and print its spectrum.
jw_operator = jordan_wigner(fermion_operator)
print('')
print(jw_operator)
jw_spectrum = eigenspectrum(jw_operator)
print(jw_spectrum)
# Transform to qubits under the Bravyi-Kitaev transformation and print its spectrum.
bk_operator = bravyi_kitaev(fermion_operator)
print('')
print(bk_operator)
bk_spectrum = eigenspectrum(bk_operator)
print(bk_spectrum)
# -
# We see that despite the different representation, these operators are iso-spectral. We can also apply the Jordan-Wigner transform in reverse to map arbitrary QubitOperators to FermionOperators. Note that we also demonstrate the .compress() method (a method on both FermionOperators and QubitOperators) which removes zero entries.
# +
from openfermion.transforms import reverse_jordan_wigner
# Initialize QubitOperator.
my_operator = QubitOperator('X0 Y1 Z2', 88.)
my_operator += QubitOperator('Z1 Z4', 3.17)
print(my_operator)
# Map QubitOperator to a FermionOperator.
mapped_operator = reverse_jordan_wigner(my_operator)
print('')
print(mapped_operator)
# Map the operator back to qubits and make sure it is the same.
back_to_normal = jordan_wigner(mapped_operator)
back_to_normal.compress()
print('')
print(back_to_normal)
# -
# ## Sparse matrices and the Hubbard model
# Often, one would like to obtain a sparse matrix representation of an operator which can be analyzed numerically. There is code in both openfermion.transforms and openfermion.utils which facilitates this. The function get_sparse_operator converts either a FermionOperator, a QubitOperator or other more advanced classes such as InteractionOperator to a scipy.sparse.csc matrix. There are numerous functions in openfermion.utils which one can call on the sparse operators such as "get_gap", "get_hartree_fock_state", "get_ground_state", etc. We show this off by computing the ground state energy of the Hubbard model. To do that, we use code from the openfermion.hamiltonians module which constructs lattice models of fermions such as Hubbard models.
# +
from openfermion.hamiltonians import fermi_hubbard
from openfermion.transforms import get_sparse_operator, jordan_wigner
from openfermion.utils import get_ground_state
# Set model.
x_dimension = 2
y_dimension = 2
tunneling = 2.
coulomb = 1.
magnetic_field = 0.5
chemical_potential = 0.25
periodic = 1
spinless = 1
# Get fermion operator.
hubbard_model = fermi_hubbard(
x_dimension, y_dimension, tunneling, coulomb, chemical_potential,
magnetic_field, periodic, spinless)
print(hubbard_model)
# Get qubit operator under Jordan-Wigner.
jw_hamiltonian = jordan_wigner(hubbard_model)
jw_hamiltonian.compress()
print('')
print(jw_hamiltonian)
# Get scipy.sparse.csc representation.
sparse_operator = get_sparse_operator(hubbard_model)
print('')
print(sparse_operator)
print('\nEnergy of the model is {} in units of T and J.'.format(
get_ground_state(sparse_operator)[0]))
# -
# ## Hamiltonians in the plane wave basis
# A user can write plugins to openfermion which allow for the use of, e.g., third-party electronic structure package to compute molecular orbitals, Hamiltonians, energies, reduced density matrices, coupled cluster amplitudes, etc using Gaussian basis sets. We may provide scripts which interface between such packages and openfermion in future but do not discuss them in this tutorial.
#
# When using simpler basis sets such as plane waves, these packages are not needed. openfermion comes with code which computes Hamiltonians in the plane wave basis. Note that when using plane waves, one is working with the periodized Coulomb operator, best suited for condensed phase calculations such as studying the electronic structure of a solid. To obtain these Hamiltonians one must choose to study the system without a spin degree of freedom (spinless), one must the specify dimension in which the calculation is performed (n_dimensions, usually 3), one must specify how many plane waves are in each dimension (grid_length) and one must specify the length scale of the plane wave harmonics in each dimension (length_scale) and also the locations and charges of the nuclei. One can generate these models with plane_wave_hamiltonian() found in openfermion.hamiltonians. For simplicity, below we compute the Hamiltonian in the case of zero external charge (corresponding to the uniform electron gas, aka jellium). We also demonstrate that one can transform the plane wave Hamiltonian using a Fourier transform without effecting the spectrum of the operator.
# +
from openfermion.hamiltonians import jellium_model
from openfermion.utils import eigenspectrum, fourier_transform, Grid
from openfermion.transforms import jordan_wigner
# Let's look at a very small model of jellium in 1D.
grid = Grid(dimensions=1, length=3, scale=1.0)
spinless = True
# Get the momentum Hamiltonian.
momentum_hamiltonian = jellium_model(grid, spinless)
momentum_qubit_operator = jordan_wigner(momentum_hamiltonian)
momentum_qubit_operator.compress()
print(momentum_qubit_operator)
# Fourier transform the Hamiltonian to the position basis.
position_hamiltonian = fourier_transform(momentum_hamiltonian, grid, spinless)
position_qubit_operator = jordan_wigner(position_hamiltonian)
position_qubit_operator.compress()
print('')
print (position_qubit_operator)
# Check the spectra to make sure these representations are iso-spectral.
spectral_difference = eigenspectrum(momentum_qubit_operator) - eigenspectrum(position_qubit_operator)
print('')
print(spectral_difference)
# -
# ## Basics of MolecularData class
#
# Data from electronic structure calculations can be saved in an OpenFermion data structure called MolecularData, which makes it easy to access within our library. Often, one would like to analyze a chemical series or look at many different Hamiltonians and sometimes the electronic structure calculations are either expensive to compute or difficult to converge (e.g. one needs to mess around with different types of SCF routines to make things converge). Accordingly, we anticipate that users will want some way to automatically database the results of their electronic structure calculations so that important data (such as the SCF integrals) can be looked up on-the-fly if the user has computed them in the past. OpenFermion supports a data provenance strategy which saves key results of the electronic structure calculation (including pointers to files containing large amounts of data, such as the molecular integrals) in an HDF5 container.
#
# The MolecularData class stores information about molecules. One initializes a MolecularData object by specifying parameters of a molecule such as its geometry, basis, multiplicity, charge and an optional string describing it. One can also initialize MolecularData simply by providing a string giving a filename where a previous MolecularData object was saved in an HDF5 container. One can save a MolecularData instance by calling the class's .save() method. This automatically saves the instance in a data folder specified during OpenFermion installation. The name of the file is generated automatically from the instance attributes and optionally provided description. Alternatively, a filename can also be provided as an optional input if one wishes to manually name the file.
#
# When electronic structure calculations are run, the data files for the molecule can be automatically updated. If one wishes to later use that data they either initialize MolecularData with the instance filename or initialize the instance and then later call the .load() method.
#
# Basis functions are provided to initialization using a string such as "6-31g". Geometries can be specified using a simple txt input file (see geometry_from_file function in molecular_data.py) or can be passed using a simple python list format demonstrated below. Atoms are specified using a string for their atomic symbol. Distances should be provided in angstrom. Below we initialize a simple instance of MolecularData without performing any electronic structure calculations.
# +
from openfermion.hamiltonians import MolecularData
# Set parameters to make a simple molecule.
diatomic_bond_length = .7414
geometry = [('H', (0., 0., 0.)), ('H', (0., 0., diatomic_bond_length))]
basis = 'sto-3g'
multiplicity = 1
charge = 0
description = str(diatomic_bond_length)
# Make molecule and print out a few interesting facts about it.
molecule = MolecularData(geometry, basis, multiplicity,
charge, description)
print('Molecule has automatically generated name {}'.format(
molecule.name))
print('Information about this molecule would be saved at:\n{}\n'.format(
molecule.filename))
print('This molecule has {} atoms and {} electrons.'.format(
molecule.n_atoms, molecule.n_electrons))
for atom, atomic_number in zip(molecule.atoms, molecule.protons):
print('Contains {} atom, which has {} protons.'.format(
atom, atomic_number))
# -
# If we had previously computed this molecule using an electronic structure package, we can call molecule.load() to populate all sorts of interesting fields in the data structure. Though we make no assumptions about what electronic structure packages users might install, we assume that the calculations are saved in OpenFermion's MolecularData objects. Currently plugins are available for [Psi4](http://psicode.org/) [(OpenFermion-Psi4)](http://github.com/quantumlib/OpenFermion-Psi4) and [PySCF](https://github.com/sunqm/pyscf) [(OpenFermion-PySCF)](http://github.com/quantumlib/OpenFermion-PySCF), and there may be more in the future. For the purposes of this example, we will load data that ships with OpenFermion to make a plot of the energy surface of hydrogen. Note that helper functions to initialize some interesting chemical benchmarks are found in openfermion.utils.
# +
# Set molecule parameters.
basis = 'sto-3g'
multiplicity = 1
bond_length_interval = 0.1
n_points = 25
# Generate molecule at different bond lengths.
hf_energies = []
fci_energies = []
bond_lengths = []
for point in range(3, n_points + 1):
bond_length = bond_length_interval * point
bond_lengths += [bond_length]
description = str(round(bond_length,2))
print(description)
geometry = [('H', (0., 0., 0.)), ('H', (0., 0., bond_length))]
molecule = MolecularData(
geometry, basis, multiplicity, description=description)
# Load data.
molecule.load()
# Print out some results of calculation.
print('\nAt bond length of {} angstrom, molecular hydrogen has:'.format(
bond_length))
print('Hartree-Fock energy of {} Hartree.'.format(molecule.hf_energy))
print('MP2 energy of {} Hartree.'.format(molecule.mp2_energy))
print('FCI energy of {} Hartree.'.format(molecule.fci_energy))
print('Nuclear repulsion energy between protons is {} Hartree.'.format(
molecule.nuclear_repulsion))
for orbital in range(molecule.n_orbitals):
print('Spatial orbital {} has energy of {} Hartree.'.format(
orbital, molecule.orbital_energies[orbital]))
hf_energies += [molecule.hf_energy]
fci_energies += [molecule.fci_energy]
# Plot.
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(0)
plt.plot(bond_lengths, fci_energies, 'x-')
plt.plot(bond_lengths, hf_energies, 'o-')
plt.ylabel('Energy in Hartree')
plt.xlabel('Bond length in angstrom')
plt.show()
# -
# The geometry data needed to generate MolecularData can also be retreived from the PubChem online database by inputting the molecule's name.
# +
from openfermion.utils import geometry_from_pubchem
methane_geometry = geometry_from_pubchem('methane')
print(methane_geometry)
# -
# ## InteractionOperator and InteractionRDM for efficient numerical representations
#
# Fermion Hamiltonians can be expressed as $H = h_0 + \sum_{pq} h_{pq}\, a^\dagger_p a_q + \frac{1}{2} \sum_{pqrs} h_{pqrs} \, a^\dagger_p a^\dagger_q a_r a_s$ where $h_0$ is a constant shift due to the nuclear repulsion and $h_{pq}$ and $h_{pqrs}$ are the famous molecular integrals. Since fermions interact pairwise, their energy is thus a unique function of the one-particle and two-particle reduced density matrices which are expressed in second quantization as $\rho_{pq} = \left \langle p \mid a^\dagger_p a_q \mid q \right \rangle$ and $\rho_{pqrs} = \left \langle pq \mid a^\dagger_p a^\dagger_q a_r a_s \mid rs \right \rangle$, respectively.
#
# Because the RDMs and molecular Hamiltonians are both compactly represented and manipulated as 2- and 4- index tensors, we can represent them in a particularly efficient form using similar data structures. The InteractionOperator data structure can be initialized for a Hamiltonian by passing the constant $h_0$ (or 0), as well as numpy arrays representing $h_{pq}$ (or $\rho_{pq}$) and $h_{pqrs}$ (or $\rho_{pqrs}$). Importantly, InteractionOperators can also be obtained by calling MolecularData.get_molecular_hamiltonian() or by calling the function get_interaction_operator() (found in openfermion.transforms) on a FermionOperator. The InteractionRDM data structure is similar but represents RDMs. For instance, one can get a molecular RDM by calling MolecularData.get_molecular_rdm(). When generating Hamiltonians from the MolecularData class, one can choose to restrict the system to an active space.
#
# These classes inherit from the same base class, PolynomialTensor. This data structure overloads the slice operator [] so that one can get or set the key attributes of the InteractionOperator: $\textrm{.constant}$, $\textrm{.one_body_coefficients}$ and $\textrm{.two_body_coefficients}$ . For instance, InteractionOperator[(p, 1), (q, 1), (r, 0), (s, 0)] would return $h_{pqrs}$ and InteractionRDM would return $\rho_{pqrs}$. Importantly, the class supports fast basis transformations using the method PolynomialTensor.rotate_basis(rotation_matrix).
# But perhaps most importantly, one can map the InteractionOperator to any of the other data structures we've described here.
#
# Below, we load MolecularData from a saved calculation of LiH. We then obtain an InteractionOperator representation of this system in an active space. We then map that operator to qubits. We then demonstrate that one can rotate the orbital basis of the InteractionOperator using random angles to obtain a totally different operator that is still iso-spectral.
# +
from openfermion.hamiltonians import MolecularData
from openfermion.transforms import get_fermion_operator, get_sparse_operator, jordan_wigner
from openfermion.utils import get_ground_state
import numpy
import scipy
import scipy.linalg
# Load saved file for LiH.
diatomic_bond_length = 1.45
geometry = [('Li', (0., 0., 0.)), ('H', (0., 0., diatomic_bond_length))]
basis = 'sto-3g'
multiplicity = 1
# Set Hamiltonian parameters.
active_space_start = 1
active_space_stop = 3
# Generate and populate instance of MolecularData.
molecule = MolecularData(geometry, basis, multiplicity, description="1.45")
molecule.load()
# Get the Hamiltonian in an active space.
molecular_hamiltonian = molecule.get_molecular_hamiltonian(
occupied_indices=range(active_space_start),
active_indices=range(active_space_start, active_space_stop))
# Map operator to fermions and qubits.
fermion_hamiltonian = get_fermion_operator(molecular_hamiltonian)
qubit_hamiltonian = jordan_wigner(fermion_hamiltonian)
qubit_hamiltonian.compress()
print('The Jordan-Wigner Hamiltonian in canonical basis follows:\n{}'.format(qubit_hamiltonian))
# Get sparse operator and ground state energy.
sparse_hamiltonian = get_sparse_operator(qubit_hamiltonian)
energy, state = get_ground_state(sparse_hamiltonian)
print('Ground state energy before rotation is {} Hartree.\n'.format(energy))
# Randomly rotate.
n_orbitals = molecular_hamiltonian.n_qubits // 2
n_variables = int(n_orbitals * (n_orbitals - 1) / 2)
numpy.random.seed(1)
random_angles = numpy.pi * (1. - 2. * numpy.random.rand(n_variables))
kappa = numpy.zeros((n_orbitals, n_orbitals))
index = 0
for p in range(n_orbitals):
for q in range(p + 1, n_orbitals):
kappa[p, q] = random_angles[index]
kappa[q, p] = -numpy.conjugate(random_angles[index])
index += 1
# Build the unitary rotation matrix.
difference_matrix = kappa + kappa.transpose()
rotation_matrix = scipy.linalg.expm(kappa)
# Apply the unitary.
molecular_hamiltonian.rotate_basis(rotation_matrix)
# Get qubit Hamiltonian in rotated basis.
qubit_hamiltonian = jordan_wigner(molecular_hamiltonian)
qubit_hamiltonian.compress()
print('The Jordan-Wigner Hamiltonian in rotated basis follows:\n{}'.format(qubit_hamiltonian))
# Get sparse Hamiltonian and energy in rotated basis.
sparse_hamiltonian = get_sparse_operator(qubit_hamiltonian)
energy, state = get_ground_state(sparse_hamiltonian)
print('Ground state energy after rotation is {} Hartree.'.format(energy))
# -
# ## Quadratic Hamiltonians and Slater determinants
#
# The general electronic structure Hamiltonian
# $H = h_0 + \sum_{pq} h_{pq}\, a^\dagger_p a_q + \frac{1}{2} \sum_{pqrs} h_{pqrs} \, a^\dagger_p a^\dagger_q a_r a_s$ contains terms that act on up to 4 sites, or
# is quartic in the fermionic creation and annihilation operators. However, in many situations
# we may fruitfully approximate these Hamiltonians by replacing these quartic terms with
# terms that act on at most 2 fermionic sites, or quadratic terms, as in mean-field approximation theory.
# These Hamiltonians have a number of
# special properties one can exploit for efficient simulation and manipulation of the Hamiltonian, thus
# warranting a special data structure. We refer to Hamiltonians which
# only contain terms that are quadratic in the fermionic creation and annihilation operators
# as quadratic Hamiltonians, and include the general case of non-particle conserving terms as in
# a general Bogoliubov transformation. Eigenstates of quadratic Hamiltonians can be prepared
# efficiently on both a quantum and classical computer, making them amenable to initial guesses for
# many more challenging problems.
#
# A general quadratic Hamiltonian takes the form
# $$H = \sum_{p, q} (M_{pq} - \mu \delta_{pq}) a^\dagger_p a_q + \frac{1}{2} \sum_{p, q} (\Delta_{pq} a^\dagger_p a^\dagger_q + \Delta_{pq}^* a_q a_p) + \text{constant},$$
# where $M$ is a Hermitian matrix, $\Delta$ is an antisymmetric matrix,
# $\delta_{pq}$ is the Kronecker delta symbol, and $\mu$ is a chemical
# potential term which we keep separate from $M$ so that we can use it
# to adjust the expectation of the total number of particles.
# In OpenFermion, quadratic Hamiltonians are conveniently represented and manipulated
# using the QuadraticHamiltonian class, which stores $M$, $\Delta$, $\mu$ and the constant. It is specialized to exploit the properties unique to quadratic Hamiltonians. Like InteractionOperator and InteractionRDM, it inherits from the PolynomialTensor class.
#
# The BCS mean-field model of superconductivity is a quadratic Hamiltonian. The following code constructs an instance of this model as a FermionOperator, converts it to a QuadraticHamiltonian, and then computes its ground energy:
# +
from openfermion.hamiltonians import mean_field_dwave
from openfermion.transforms import get_quadratic_hamiltonian
# Set model.
x_dimension = 2
y_dimension = 2
tunneling = 2.
sc_gap = 1.
periodic = True
# Get FermionOperator.
mean_field_model = mean_field_dwave(
x_dimension, y_dimension, tunneling, sc_gap, periodic=periodic)
# Convert to QuadraticHamiltonian
quadratic_hamiltonian = get_quadratic_hamiltonian(mean_field_model)
# Compute the ground energy
ground_energy = quadratic_hamiltonian.ground_energy()
print(ground_energy)
# -
# Any quadratic Hamiltonian may be rewritten in the form
# $$H = \sum_p \varepsilon_p b^\dagger_p b_p + \text{constant},$$
# where the $b_p$ are new annihilation operators that satisfy the fermionic anticommutation relations, and which are linear combinations of the old creation and annihilation operators. This form of $H$ makes it easy to deduce its eigenvalues; they are sums of subsets of the $\varepsilon_p$, which we call the orbital energies of $H$. The following code computes the orbital energies and the constant:
orbital_energies, constant = quadratic_hamiltonian.orbital_energies()
print(orbital_energies)
print()
print(constant)
# Eigenstates of quadratic hamiltonians are also known as fermionic Gaussian states, and they can be prepared efficiently on a quantum computer. One can use OpenFermion to obtain circuits for preparing these states. The following code obtains the description of a circuit which prepares the ground state (operations that can be performed in parallel are grouped together), along with a description of the starting state to which the circuit should be applied:
# +
from openfermion.utils import gaussian_state_preparation_circuit
circuit_description, start_orbitals = gaussian_state_preparation_circuit(quadratic_hamiltonian)
for parallel_ops in circuit_description:
print(parallel_ops)
print('')
print(start_orbitals)
# -
# In the circuit description, each elementary operation is either a tuple of the form $(i, j, \theta, \varphi)$, indicating the operation $\exp[i \varphi a_j^\dagger a_j]\exp[\theta (a_i^\dagger a_j - a_j^\dagger a_i)]$, which is a Givens rotation of modes $i$ and $j$, or the string 'pht', indicating the particle-hole transformation on the last fermionic mode, which is the operator $\mathcal{B}$ such that $\mathcal{B} a_N \mathcal{B}^\dagger = a_N^\dagger$ and leaves the rest of the ladder operators unchanged. Operations that can be performed in parallel are grouped together.
#
# In the special case that a quadratic Hamiltonian conserves particle number ($\Delta = 0$), its eigenstates take the form
# $$\lvert \Psi_S \rangle = b^\dagger_{1}\cdots b^\dagger_{N_f}\lvert \text{vac} \rangle,\qquad
# b^\dagger_{p} = \sum_{k=1}^N Q_{pq}a^\dagger_q,$$
# where $Q$ is an $N_f \times N$ matrix with orthonormal rows. These states are also known as Slater determinants. OpenFermion also provides functionality to obtain circuits for preparing Slater determinants starting with the matrix $Q$ as the input.
| examples/openfermion_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# 
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Azure ML Hardware Accelerated Models Quickstart
# This tutorial will show you how to deploy an image recognition service based on the ResNet 50 classifier using the Azure Machine Learning Accelerated Models service. Get more information about our service from our [documentation](https://docs.microsoft.com/en-us/azure/machine-learning/service/concept-accelerate-with-fpgas), [API reference](https://docs.microsoft.com/en-us/python/api/azureml-accel-models/azureml.accel?view=azure-ml-py), or [forum](https://aka.ms/aml-forum).
#
# We will use an accelerated ResNet50 featurizer running on an FPGA. Our Accelerated Models Service handles translating deep neural networks (DNN) into an FPGA program.
#
# For more information about using other models besides Resnet50, see the [README](./README.md).
#
# The steps covered in this notebook are:
# 1. [Set up environment](#set-up-environment)
# * [Construct model](#construct-model)
# * Image Preprocessing
# * Featurizer (Resnet50)
# * Classifier
# * Save Model
# * [Register Model](#register-model)
# * [Convert into Accelerated Model](#convert-model)
# * [Create Image](#create-image)
# * [Deploy](#deploy-image)
# * [Test service](#test-service)
# * [Clean-up](#clean-up)
# <a id="set-up-environment"></a>
# ## 1. Set up environment
import os
import tensorflow as tf
# ### Retrieve Workspace
# If you haven't created a Workspace, please follow [this notebook](https://github.com/Azure/MachineLearningNotebooks/blob/master/configuration.ipynb) to do so. If you have, run the codeblock below to retrieve it.
# +
from azureml.core import Workspace
ws = Workspace.from_config()
print(ws.name, ws.resource_group, ws.location, ws.subscription_id, sep = '\n')
# -
# <a id="construct-model"></a>
# ## 2. Construct model
#
# There are three parts to the model we are deploying: pre-processing, featurizer with ResNet50, and classifier with ImageNet dataset. Then we will save this complete Tensorflow model graph locally before registering it to your Azure ML Workspace.
#
# ### 2.a. Image preprocessing
# We'd like our service to accept JPEG images as input. However the input to ResNet50 is a tensor. So we need code that decodes JPEG images and does the preprocessing required by ResNet50. The Accelerated AI service can execute TensorFlow graphs as part of the service and we'll use that ability to do the image preprocessing. This code defines a TensorFlow graph that preprocesses an array of JPEG images (as strings) and produces a tensor that is ready to be featurized by ResNet50.
#
# **Note:** Expect to see TF deprecation warnings until we port our SDK over to use Tensorflow 2.0.
# +
# Input images as a two-dimensional tensor containing an arbitrary number of images represented a strings
import azureml.accel.models.utils as utils
tf.reset_default_graph()
in_images = tf.placeholder(tf.string)
image_tensors = utils.preprocess_array(in_images)
print(image_tensors.shape)
# -
# ### 2.b. Featurizer
# We use ResNet50 as a featurizer. In this step we initialize the model. This downloads a TensorFlow checkpoint of the quantized ResNet50.
from azureml.accel.models import QuantizedResnet50
save_path = os.path.expanduser('~/models')
model_graph = QuantizedResnet50(save_path, is_frozen = True)
feature_tensor = model_graph.import_graph_def(image_tensors)
print(model_graph.version)
print(feature_tensor.name)
print(feature_tensor.shape)
# ### 2.c. Classifier
# The model we downloaded includes a classifier which takes the output of the ResNet50 and identifies an image. This classifier is trained on the ImageNet dataset. We are going to use this classifier for our service. The next [notebook](./accelerated-models-training.ipynb) shows how to train a classifier for a different data set. The input to the classifier is a tensor matching the output of our ResNet50 featurizer.
classifier_output = model_graph.get_default_classifier(feature_tensor)
print(classifier_output)
# ### 2.d. Save Model
# Now that we loaded all three parts of the tensorflow graph (preprocessor, resnet50 featurizer, and the classifier), we can save the graph and associated variables to a directory which we can register as an Azure ML Model.
# +
# model_name must be lowercase
model_name = "resnet50"
model_save_path = os.path.join(save_path, model_name)
print("Saving model in {}".format(model_save_path))
with tf.Session() as sess:
model_graph.restore_weights(sess)
tf.saved_model.simple_save(sess, model_save_path,
inputs={'images': in_images},
outputs={'output_alias': classifier_output})
# -
# ### 2.e. Important! Save names of input and output tensors
#
# These input and output tensors that were created during the preprocessing and classifier steps are also going to be used when **converting the model** to an Accelerated Model that can run on FPGA's and for **making an inferencing request**. It is very important to save this information! You can see our defaults for all the models in the [README](./README.md).
#
# By default for Resnet50, these are the values you should see when running the cell below:
# * input_tensors = "Placeholder:0"
# * output_tensors = "classifier/resnet_v1_50/predictions/Softmax:0"
# + tags=["register model from file"]
input_tensors = in_images.name
output_tensors = classifier_output.name
print(input_tensors)
print(output_tensors)
# -
# <a id="register-model"></a>
# ## 3. Register Model
# You can add tags and descriptions to your models. Using tags, you can track useful information such as the name and version of the machine learning library used to train the model. Note that tags must be alphanumeric.
# + tags=["register model from file"]
from azureml.core.model import Model
registered_model = Model.register(workspace = ws,
model_path = model_save_path,
model_name = model_name)
print("Successfully registered: ", registered_model.name, registered_model.description, registered_model.version, sep = '\t')
# -
# <a id="convert-model"></a>
# ## 4. Convert Model
# For conversion you need to provide names of input and output tensors. This information can be found from the model_graph you saved in step 2.e. above.
#
# **Note**: Conversion may take a while and on average for FPGA model it is about 1-3 minutes and it depends on model type.
# + tags=["register model from file"]
from azureml.accel import AccelOnnxConverter
convert_request = AccelOnnxConverter.convert_tf_model(ws, registered_model, input_tensors, output_tensors)
if convert_request.wait_for_completion(show_output = False):
# If the above call succeeded, get the converted model
converted_model = convert_request.result
print("\nSuccessfully converted: ", converted_model.name, converted_model.url, converted_model.version,
converted_model.id, converted_model.created_time, '\n')
else:
print("Model conversion failed. Showing output.")
convert_request.wait_for_completion(show_output = True)
# -
# <a id="create-image"></a>
# ## 5. Package the model into an Image
# You can add tags and descriptions to image. Also, for FPGA model an image can only contain **single** model.
#
# **Note**: The following command can take few minutes.
# +
from azureml.core.image import Image
from azureml.accel import AccelContainerImage
image_config = AccelContainerImage.image_configuration()
# Image name must be lowercase
image_name = "{}-image".format(model_name)
image = Image.create(name = image_name,
models = [converted_model],
image_config = image_config,
workspace = ws)
image.wait_for_creation(show_output = False)
# -
# <a id="deploy-image"></a>
# ## 6. Deploy
# Once you have an Azure ML Accelerated Image in your Workspace, you can deploy it to two destinations, to a Databox Edge machine or to an AKS cluster.
#
# ### 6.a. Databox Edge Machine using IoT Hub
# See the sample [here](https://github.com/Azure-Samples/aml-real-time-ai/) for using the Azure IoT CLI extension for deploying your Docker image to your Databox Edge Machine.
#
# ### 6.b. Azure Kubernetes Service (AKS) using Azure ML Service
# We are going to create an AKS cluster with FPGA-enabled machines, then deploy our service to it. For more information, see [AKS official docs](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-deploy-and-where#aks).
#
# #### Create AKS ComputeTarget
# +
from azureml.core.compute import AksCompute, ComputeTarget
# Uses the specific FPGA enabled VM (sku: Standard_PB6s)
# Standard_PB6s are available in: eastus, westus2, westeurope, southeastasia
prov_config = AksCompute.provisioning_configuration(vm_size = "Standard_PB6s",
agent_count = 1,
location = "eastus")
aks_name = 'my-aks-pb6'
# Create the cluster
aks_target = ComputeTarget.create(workspace = ws,
name = aks_name,
provisioning_configuration = prov_config)
# -
# Provisioning an AKS cluster might take awhile (15 or so minutes), and we want to wait until it's successfully provisioned before we can deploy a service to it. If you interrupt this cell, provisioning of the cluster will continue. You can also check the status in your Workspace under Compute.
# %%time
aks_target.wait_for_completion(show_output = True)
print(aks_target.provisioning_state)
print(aks_target.provisioning_errors)
# #### Deploy AccelContainerImage to AKS ComputeTarget
# +
# %%time
from azureml.core.webservice import Webservice, AksWebservice
# Set the web service configuration (for creating a test service, we don't want autoscale enabled)
# Authentication is enabled by default, but for testing we specify False
aks_config = AksWebservice.deploy_configuration(autoscale_enabled=False,
num_replicas=1,
auth_enabled = False)
aks_service_name ='my-aks-service-1'
aks_service = Webservice.deploy_from_image(workspace = ws,
name = aks_service_name,
image = image,
deployment_config = aks_config,
deployment_target = aks_target)
aks_service.wait_for_deployment(show_output = True)
# -
# <a id="test-service"></a>
# ## 7. Test the service
# ### 7.a. Create Client
# The image supports gRPC and the TensorFlow Serving "predict" API. We will create a PredictionClient from the Webservice object that can call into the docker image to get predictions. If you do not have the Webservice object, you can also create [PredictionClient](https://docs.microsoft.com/en-us/python/api/azureml-accel-models/azureml.accel.predictionclient?view=azure-ml-py) directly.
#
# **Note:** If you chose to use auth_enabled=True when creating your AksWebservice, see documentation [here](https://docs.microsoft.com/en-us/python/api/azureml-core/azureml.core.webservice(class)?view=azure-ml-py#get-keys--) on how to retrieve your keys and use either key as an argument to PredictionClient(...,access_token=key).
# **WARNING:** If you are running on Azure Notebooks free compute, you will not be able to make outgoing calls to your service. Try locating your client on a different machine to consume it.
# +
# Using the grpc client in AzureML Accelerated Models SDK
from azureml.accel import client_from_service
# Initialize AzureML Accelerated Models client
client = client_from_service(aks_service)
# -
# You can adapt the client [code](https://github.com/Azure/aml-real-time-ai/blob/master/pythonlib/amlrealtimeai/client.py) to meet your needs. There is also an example C# [client](https://github.com/Azure/aml-real-time-ai/blob/master/sample-clients/csharp).
#
# The service provides an API that is compatible with TensorFlow Serving. There are instructions to download a sample client [here](https://www.tensorflow.org/serving/setup).
# ### 7.b. Serve the model
# To understand the results we need a mapping to the human readable imagenet classes
import requests
classes_entries = requests.get("https://raw.githubusercontent.com/Lasagne/Recipes/master/examples/resnet50/imagenet_classes.txt").text.splitlines()
# +
# Score image with input and output tensor names
results = client.score_file(path="./snowleopardgaze.jpg",
input_name=input_tensors,
outputs=output_tensors)
# map results [class_id] => [confidence]
results = enumerate(results)
# sort results by confidence
sorted_results = sorted(results, key=lambda x: x[1], reverse=True)
# print top 5 results
for top in sorted_results[:5]:
print(classes_entries[top[0]], 'confidence:', top[1])
# -
# <a id="clean-up"></a>
# ## 8. Clean-up
# Run the cell below to delete your webservice, image, and model (must be done in that order). In the [next notebook](./accelerated-models-training.ipynb) you will learn how to train a classfier on a new dataset using transfer learning and finetune the weights.
aks_service.delete()
aks_target.delete()
image.delete()
registered_model.delete()
converted_model.delete()
| how-to-use-azureml/deployment/accelerated-models/accelerated-models-quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
from context import uncertify
# +
import logging
from uncertify.log import setup_logging
setup_logging()
LOG = logging.getLogger(__name__)
# Matplotlib DEBUG logging spits out a whole bunch of crap
mpl_logger = logging.getLogger('matplotlib')
mpl_logger.setLevel(logging.WARNING)
# +
from pathlib import Path
import torch
import torchvision
from torchvision.transforms import Compose
from tqdm import tqdm
from uncertify.data.dataloaders import dataloader_factory, DatasetType
from uncertify.models.vae import VariationalAutoEncoder
from uncertify.models.encoder_decoder_baur2020 import BaurEncoder, BaurDecoder
from uncertify.evaluation.latent_space_analysis import sample_from_gauss_prior
#from uncertify.evaluation.inference import infer_latent_space_samples
from uncertify.data.default_dataloaders import default_dataloader_dict_factory
from uncertify.io.models import load_vae_baur_model
from uncertify.visualization.reconstruction import plot_vae_output
from uncertify.common import DATA_DIR_PATH
# -
# # Load model and Dataloaders
masked_model = load_vae_baur_model(Path('/mnt/2TB_internal_HD/lightning_logs/schedule_mask/version_1/checkpoints/last.ckpt'))
non_masked_model = load_vae_baur_model(Path('/mnt/2TB_internal_HD/lightning_logs/schedule_mask/version_6/checkpoints/last.ckpt'))
model = non_masked_model
model = load_vae_baur_model(Path('/mnt/2TB_internal_HD/lightning_logs/beta_test/version_2/checkpoints/last.ckpt'))
dataloader_dict = default_dataloader_dict_factory(batch_size=155,
num_workers=0,
shuffle_val=True)
# # Plot latent space behaviour
# ## Plot variance captured over one latent space dimension
from uncertify.visualization.latent_space_analysis import plot_latent_reconstructions_one_dim_changing
# +
change_dim_indices = [1, 80, 108]
for dim in change_dim_indices:
plot_latent_reconstructions_one_dim_changing(trained_model=model, change_dim_idx=dim, n_samples=32,
save_path=DATA_DIR_PATH / 'plots' / f'latent_sample_one_dim_{dim}.png');
# -
# ## Plot variance captured over all latent space dimensions having all others fixed
from uncertify.visualization.latent_space_analysis import plot_latent_reconstructions_multiple_dims
plot_latent_reconstructions_multiple_dims(model, latent_space_dims=128, n_samples_per_dim=16, save_path=DATA_DIR_PATH / 'plots' / 'latent_reconstruct_all_dims.png',
cmap='gray')
# ## Plot 2D grid varying values from -3 to 3 std's
from uncertify.visualization.latent_space_analysis import plot_latent_reconstructions_2d_grid
plot_latent_reconstructions_2d_grid(model, dim1=18, dim2=50, save_path=DATA_DIR_PATH / 'plots' / 'latent_space_2d_grid.png')
# # Plot latent space sample reconstructions from gaussian random samples
from uncertify.visualization.latent_space_analysis import plot_random_latent_space_samples
plot_random_latent_space_samples(model, n_samples=16, nrow=16, cmap='gray')
# # Plot Gaussian samples annulus distribution
from uncertify.visualization.latent_space_analysis import plot_gaussian_annulus_distribution
plot_gaussian_annulus_distribution(latent_space_dims=128, n_samples=1000);
# # Plot latent space sample reconstructions from different locations in latent space
# +
from uncertify.visualization.latent_space_analysis import plot_latent_samples_from_ring
radii = [(0, 1), (2, 3), (4, 5), (7, 9), (10, 12), (15, 17), (20, 30), (50, 60), (200, 210)]
for sample in radii:
inner_radius, outer_radius = sample
fig = plot_latent_samples_from_ring(model, n_samples=16, inner_radius=inner_radius, outer_radius=outer_radius, cmap='gray')
# -
# # Plot latent space embeddings UMAP
from uncertify.visualization.latent_space_analysis import plot_umap_latent_embedding
from uncertify.evaluation.inference import yield_inference_batches
print(dataloader_dict.keys())
# +
max_n_batches = 6
redisual_threshold = 0.67
select_dataloaders = ['CamCAN train', 'MNIST', 'Gaussian noise']
output_generators = []
for dataloader_name in select_dataloaders:
dataloader = dataloader_dict[dataloader_name]
output_generators.append(yield_inference_batches(dataloader, model, max_n_batches, redisual_threshold, progress_bar_suffix=f'{dataloader_name}'))
umap_fig = plot_umap_latent_embedding(output_generators, select_dataloaders, figsize=(14, 10))
# -
umap_fig.savefig(DATA_DIR_PATH / 'plots' / f'umap_latent_embedding_masked.png')
| notebooks/latent_space_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# A more interesting simulation allows us to study the behavior of the printing queue described earlier in this section. Recall that as students send printing tasks to the shared printer, the tasks are placed in a queue to be processed in a first-come first-served manner. Many questions arise with this configuration. The most important of these might be whether the printer is capable of handling a certain amount of work. If it cannot, students will be waiting too long for printing and may miss their next class.
# +
import random
class Queue:
def __init__(self):
self.items = []
def isEmpty(self):
return self.items == []
def enqueue(self, item):
self.items.insert(0,item)
def dequeue(self):
return self.items.pop()
def size(self):
return len(self.items)
class Printer:
def __init__(self, ppm):
self.pagerate = ppm
self.currentTask = None
self.timeRemaining = 0
def tick(self):
if self.currentTask != None:
self.timeRemaining = self.timeRemaining - 1
if self.timeRemaining <= 0:
self.currentTask = None
def busy(self):
if self.currentTask != None:
return True
else:
return False
def startNext(self,newtask):
self.currentTask = newtask
self.timeRemaining = newtask.getPages() * 60/self.pagerate
class Task:
def __init__(self,time):
self.timestamp = time
self.pages = random.randrange(1,21)
def getStamp(self):
return self.timestamp
def getPages(self):
return self.pages
def waitTime(self, currenttime):
return currenttime - self.timestamp
def simulation(numSeconds, pagesPerMinute):
labprinter = Printer(pagesPerMinute)
printQueue = Queue()
waitingtimes = []
for currentSecond in range(numSeconds):
if newPrintTask():
task = Task(currentSecond)
printQueue.enqueue(task)
if (not labprinter.busy()) and (not printQueue.isEmpty()):
nexttask = printQueue.dequeue()
waitingtimes.append( nexttask.waitTime(currentSecond))
labprinter.startNext(nexttask)
labprinter.tick()
averageWait=sum(waitingtimes)/len(waitingtimes)
print("Average Wait %6.2f secs %3d tasks remaining."%(averageWait,printQueue.size()))
def newPrintTask():
num = random.randrange(1,181)
if num == 180:
return True
else:
return False
for i in range(10):
simulation(3600,5)
| intermediate/Queue - Task Simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import myutil as mu
import torch
# -
# ---
# - 자동 미분(Autograd) 실습하기
# - 자동 미분에 대해서 실습을 통해 이해해봅시다.
# - 임의로 2w2+5라는 식을 세워보고, w에 대해 미분해보겠습니다.
#
# +
w = torch.tensor(2.0, requires_grad=True)
y = w ** 2
z = 2 * y + 5
z.backward()
print('수식을 w로 미분한 값 : {}'.format(w.grad))
| 0302_auto_grad.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Practical Computing
# ### Exercise 2: Write a simple, short program to calculate and print area of one of the biggest faces and volume of a rectangular cuboid with the dimensions given below. Use functions and loops if required. Lengths (l) = [30cm, 40cm, 50cm] Breadth (b) = 20cm Height (h) = 15cm
def bigestface(length,breath):
a= length* breath
#area of biggest face
return a
def volume(l,b,h):
volume= l*b*h
#Volume of cuboid
return volume
# +
length=30
breath=20
height=15
while(length < 60):
area= bigestface(length,breath)
vol= volume(length,breath,height)
print('Area of one of the biggest faces:',area)
#printing the Area of bggest Face for Lenght [30,40,50]
print("Volume of a rectangular cuboid: ",vol)
#Printind the Volume for Lenth [30, 40, 50]
length= length+10
# -
| Practical Computing Exercise 2 .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TensorFlow Regression Example
# ## Creating Data
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# 1 Million Points
x_data = np.linspace(0.0,10.0,1000000)
noise = np.random.randn(len(x_data))
# +
# y = mx + b + noise_levels
b = 5
y_true = (0.5 * x_data ) + 5 + noise
# -
my_data = pd.concat([pd.DataFrame(data=x_data,columns=['X Data']),pd.DataFrame(data=y_true,columns=['Y'])],axis=1)
my_data.head()
my_data.sample(n=250).plot(kind='scatter',x='X Data',y='Y')
# # TensorFlow
# ## Batch Size
#
# We will take the data in batches (1,000,000 points is a lot to pass in at once)
import tensorflow as tf
# Random 10 points to grab
batch_size = 8
# ** Variables **
m = tf.Variable(0.5)
b = tf.Variable(1.0)
# ** Placeholders **
xph = tf.placeholder(tf.float32,[batch_size])
yph = tf.placeholder(tf.float32,[batch_size])
# ** Graph **
y_model = m*xph + b
# ** Loss Function **
error = tf.reduce_sum(tf.square(yph-y_model))
# ** Optimizer **
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train = optimizer.minimize(error)
# ** Initialize Variables **
init = tf.global_variables_initializer()
# ### Session
with tf.Session() as sess:
sess.run(init)
batches = 1000
for i in range(batches):
rand_ind = np.random.randint(len(x_data),size=batch_size)
feed = {xph:x_data[rand_ind],yph:y_true[rand_ind]}
sess.run(train,feed_dict=feed)
model_m,model_b = sess.run([m,b])
model_m
model_b
# ### Results
y_hat = x_data * model_m + model_b
my_data.sample(n=250).plot(kind='scatter',x='X Data',y='Y')
plt.plot(x_data,y_hat,'r')
# ## tf.estimator API
#
# Much simpler API for basic tasks like regression! We'll talk about more abstractions like TF-Slim later on.
feat_cols = [tf.feature_column.numeric_column('x',shape=[1])]
estimator = tf.estimator.LinearRegressor(feature_columns=feat_cols)
# ### Train Test Split
#
# We haven't actually performed a train test split yet! So let's do that on our data now and perform a more realistic version of a Regression Task
from sklearn.model_selection import train_test_split
x_train, x_eval, y_train, y_eval = train_test_split(x_data,y_true,test_size=0.3, random_state = 101)
# +
print(x_train.shape)
print(y_train.shape)
print(x_eval.shape)
print(y_eval.shape)
# -
# ### Set up Estimator Inputs
# Can also do .pandas_input_fn
input_func = tf.estimator.inputs.numpy_input_fn({'x':x_train},y_train,batch_size=4,num_epochs=None,shuffle=True)
train_input_func = tf.estimator.inputs.numpy_input_fn({'x':x_train},y_train,batch_size=4,num_epochs=1000,shuffle=False)
eval_input_func = tf.estimator.inputs.numpy_input_fn({'x':x_eval},y_eval,batch_size=4,num_epochs=1000,shuffle=False)
# ### Train the Estimator
estimator.train(input_fn=input_func,steps=1000)
# ### Evaluation
train_metrics = estimator.evaluate(input_fn=train_input_func,steps=1000)
eval_metrics = estimator.evaluate(input_fn=eval_input_func,steps=1000)
print("train metrics: {}".format(train_metrics))
print("eval metrics: {}".format(eval_metrics))
# ### Predictions
input_fn_predict = tf.estimator.inputs.numpy_input_fn({'x':np.linspace(0,10,10)},shuffle=False)
list(estimator.predict(input_fn=input_fn_predict))
predictions = []# np.array([])
for x in estimator.predict(input_fn=input_fn_predict):
predictions.append(x['predictions'])
predictions
my_data.sample(n=250).plot(kind='scatter',x='X Data',y='Y')
plt.plot(np.linspace(0,10,10),predictions,'r')
# # Great Job!
| study_python/tensorflow/Tensorflow-Bootcamp-master/02-TensorFlow-Basics/04-TensorFlow-Regression-Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Tutorial 06: Creating Custom Scenarios
#
# This tutorial walks you through the process of generating custom scenarios. Scenarios define the network geometry of a task, as well as the constituents of the network, e.g. vehicles, traffic lights, etc... Various scenarios are available in Flow, depicting a diverse set of open and closed traffic networks such as ring roads, intersections/grids, straight highway merges, and more.
#
# In this exercise, we will recreate the ring road network, seen in the figure below.
#
# <img src="img/ring_scenario.png">
#
# In order to recreate this scenario, we will design a *scenario* class. This class creates the configuration files needed to produce a transportation network within the simulator. It also specifies the location of edge nodes in the network, as well as the positioning of vehicles at the start of a run.
#
# We begin by creating a class that inherits the methods of Flow's base scenario class. The separate methods are filled in in later sections.
# +
# import Flow's base scenario class
from flow.scenarios import Scenario
# define the scenario class, and inherit properties from the base scenario class
class myScenario(Scenario):
pass
# -
# The rest of the tutorial is organized as follows: sections 1 and 2 walk through the steps needed to specify custom traffic network geometry features and auxiliary features, respectively, while section 3 implements the new scenario in a simulation for visualization and testing purposes.
#
# ## 1. Specifying Traffic Network Features
#
# One of the core responsibilities of the scenario class is to to generate the necessary xml files needed to initialize a sumo instance. These xml files describe specific network features such as the position and directions of nodes and edges (see the figure above). Once the base scenario has been inherited, specifying these features becomes very systematic. All child classes are required to define at least the following three methods:
#
# * **specify_nodes**: specifies the attributes of nodes in the network
# * **specify_edges**: specifies the attributes of edges containing pairs on nodes in the network
# * **specify_routes**: specifies the routes vehicles can take starting from any edge
#
# Additionally, the following optional functions may also be defined:
#
# * **specify_types**: specifies the attributes of various edge types (if any exist)
# * **specify_connections**: specifies the attributes of connections. These attributes are used to describe how any specific node's incoming and outgoing edges/lane pairs are connected. If no connections are specified, sumo generates default connections.
#
# All of the functions mentioned above paragraph take in as input `net_params`, and output a list of dictionary elements, with each element providing the attributes of the component to be specified.
#
# This tutorial will cover the first three methods. For examples of `specify_types` and `specify_routes`, refer to source code located in `flow/scenarios/loop.py` and `flow/scenarios/bridge_toll.py`, respectively.
# ### 1.1 ADDITIONAL_NET_PARAMS
#
# The features used to parametrize the network are specified within the `NetParams` input, as discussed in tutorial 1. Specifically, for the sake of our network, the `additional_params` attribute within `NetParams` will be responsible for storing information on the radius, number of lanes, and speed limit within each lane, as seen in the figure above. Accordingly, for this problem, we define an `ADDITIONAL_NET_PARAMS` variable of the form:
ADDITIONAL_NET_PARAMS = {
"radius": 40,
"num_lanes": 1,
"speed_limit": 30,
}
# All scenarios presented in Flow provide a unique `ADDITIONAL_NET_PARAMS` component containing the information needed to properly define the network parameters of the scenario. We assume that these values are always provided by the user, and accordingly can be called from `net_params`. For example, if we would like to call the "radius" parameter, we simply type:
#
# radius = net_params.additional_params["radius"]
#
# ### 1.2 specify_nodes
#
# The nodes of a network are the positions of a select few points in the network. These points are connected together using edges (see section 1.4). In order to specify the location of the nodes that will be placed in the network, the function `specify_nodes` is used. This method returns a list of dictionary elements, where each dictionary depicts the attributes of a single node. These node attributes include:
# * **id**: name of the node
# * **x**: x coordinate of the node
# * **y**: y coordinate of the node
# * other sumo-related attributes, see: http://sumo.dlr.de/wiki/Networks/Building_Networks_from_own_XML-descriptions#Node_Descriptions
#
# Refering to the figure at the top of this tutorial, we specify four nodes at the bottom (0,-r), top (0,r), left (-r,0), and right (0,r) of the ring. This is done as follows:
class myScenario(myScenario): # update my scenario class
def specify_nodes(self, net_params):
# one of the elements net_params will need is a "radius" value
r = net_params.additional_params["radius"]
# specify the name and position (x,y) of each node
nodes = [{"id": "bottom", "x": 0, "y": -r},
{"id": "right", "x": r, "y": 0},
{"id": "top", "x": 0, "y": r},
{"id": "left", "x": -r, "y": 0}]
return nodes
# ### 1.3 specify_edges
#
# Once the nodes are specified, the nodes are linked together using directed edges. This done through the `specify_edges` method which, similar to `specify_nodes`, returns a list of dictionary elements, with each dictionary specifying the attributes of a single edge. The attributes include:
#
# * **id**: name of the edge
# * **from**: name of the node the edge starts from
# * **to**: the name of the node the edges ends at
# * **length**: length of the edge
# * **numLanes**: the number of lanes on the edge
# * **speed**: the speed limit for vehicles on the edge
# * other sumo-related attributes, see: http://sumo.dlr.de/wiki/Networks/Building_Networks_from_own_XML-descriptions#Edge_Descriptions.
#
# One useful additional attribute is **shape**, which specifies the shape of the edge connecting the two nodes. The shape consists of a series of subnodes (internal to sumo) that are connected together by straight lines to create a curved edge. If no shape is specified, the nodes are connected by a straight line. This attribute will be needed to create the circular arcs between the nodes in the system.
#
# We now create four arcs connected the nodes specified in section 1.2, with the direction of the edges directed counter-clockwise:
# +
# some mathematical operations that may be used
from numpy import pi, sin, cos, linspace
class myScenario(myScenario): # update my scenario class
def specify_edges(self, net_params):
r = net_params.additional_params["radius"]
edgelen = r * pi / 2
# this will let us control the number of lanes in the network
lanes = net_params.additional_params["num_lanes"]
# speed limit of vehicles in the network
speed_limit = net_params.additional_params["speed_limit"]
edges = [
{
"id": "edge0",
"numLanes": lanes,
"speed": speed_limit,
"from": "bottom",
"to": "right",
"length": edgelen,
"shape": [(r*cos(t), r*sin(t)) for t in linspace(-pi/2, 0, 40)]
},
{
"id": "edge1",
"numLanes": lanes,
"speed": speed_limit,
"from": "right",
"to": "top",
"length": edgelen,
"shape": [(r*cos(t), r*sin(t)) for t in linspace(0, pi/2, 40)]
},
{
"id": "edge2",
"numLanes": lanes,
"speed": speed_limit,
"from": "top",
"to": "left",
"length": edgelen,
"shape": [(r*cos(t), r*sin(t)) for t in linspace(pi/2, pi, 40)]},
{
"id": "edge3",
"numLanes": lanes,
"speed": speed_limit,
"from": "left",
"to": "bottom",
"length": edgelen,
"shape": [(r*cos(t), r*sin(t)) for t in linspace(pi, 3*pi/2, 40)]
}
]
return edges
# -
# ### 1.4 specify_routes
#
# The routes are the sequence of edges vehicles traverse given their current position. For example, a vehicle beginning in the edge titled "edge0" (see section 1.3) must traverse, in sequence, the edges "edge0", "edge1", "edge2", and "edge3", before restarting its path.
#
# In order to specify the routes a vehicle may take, the function `specify_routes` is used. The routes in this method can be specified in one of three ways:
#
# **1. Single route per edge:**
#
# In this case of deterministic routes (as is the case in the ring road scenario), the routes can be specified as dictionary where the key element represents the starting edge and the element is a single list of edges the vehicle must traverse, with the first edge corresponding to the edge the vehicle begins on. Note that the edges must be connected for the route to be valid.
#
# For this network, the available routes under this setting can be defined as follows:
class myScenario(myScenario): # update my scenario class
def specify_routes(self, net_params):
rts = {"edge0": ["edge0", "edge1", "edge2", "edge3"],
"edge1": ["edge1", "edge2", "edge3", "edge0"],
"edge2": ["edge2", "edge3", "edge0", "edge1"],
"edge3": ["edge3", "edge0", "edge1", "edge2"]}
return rts
# **2. Multiple routes per edge:**
#
# Alternatively, if the routes are meant to be stochastic, each element can consist of a list of (route, probability) tuples, where the first element in the tuple is one of the routes a vehicle can take from a specific starting edge, and the second element is the probability that vehicles will choose that route. Note that, in this case, the sum of probability values for each dictionary key must sum up to one.
#
# For example, modifying the code snippet we presented above, another valid way of representing the route in a more probabilistic setting is:
class myScenario(myScenario): # update my scenario class
def specify_routes(self, net_params):
rts = {"edge0": [(["edge0", "edge1", "edge2", "edge3"], 1)],
"edge1": [(["edge1", "edge2", "edge3", "edge0"], 1)],
"edge2": [(["edge2", "edge3", "edge0", "edge1"], 1)],
"edge3": [(["edge3", "edge0", "edge1", "edge2"], 1)]}
return rts
# **3. Per-vehicle routes:**
#
# Finally, if you would like to assign a specific starting route to a vehicle with a specific ID, you can do so by adding a element into the dictionary whose key is the name of the vehicle and whose content is the list of edges the vehicle is meant to traverse as soon as it is introduced to the network.
#
# As an example, assume we have a vehicle named "human_0" in the network (as we will in the later sections), and it is initialized in the edge names "edge_0". Then, the route for this edge specifically can be added through the `specify_routes` method as follows:
class myScenario(myScenario): # update my scenario class
def specify_routes(self, net_params):
rts = {"edge0": ["edge0", "edge1", "edge2", "edge3"],
"edge1": ["edge1", "edge2", "edge3", "edge0"],
"edge2": ["edge2", "edge3", "edge0", "edge1"],
"edge3": ["edge3", "edge0", "edge1", "edge2"],
"human_0": ["edge0", "edge1", "edge2", "edge3"]}
return rts
# In all three cases, the routes are ultimately represented in the class in the form described under the multiple routes setting, i.e.
#
# >>> print(scenario.rts)
#
# {
# "edge0": [
# (["edge0", "edge1", "edge2", "edge3"], 1)
# ],
# "edge1": [
# (["edge1", "edge2", "edge3", "edge0"], 1)
# ],
# "edge2": [
# (["edge2", "edge3", "edge0", "edge1"], 1)
# ],
# "edge3": [
# (["edge3", "edge0", "edge1", "edge2"], 1)
# ],
# "human_0": [
# (["edge0", "edge1", "edge2", "edge3"], 1)
# ]
# }
#
# where the vehicle-specific route is only included in the third case.
# ## 2. Specifying Auxiliary Scenario Features
#
# Other auxiliary methods exist within the base scenario class to help support vehicle state initialization and acquisition. Of these methods, the only required abstract method is:
#
# * **specify_edge_starts**: defines edge starts for road sections with respect to some global reference
#
# Other optional abstract methods within the base scenario class include:
#
# * **specify_internal_edge_starts**: defines the edge starts for internal edge nodes caused by finite length connections between road section
# * **specify_intersection_edge_starts**: defines edge starts for intersections with respect to some global reference frame. Only needed by environments with intersections.
# * **gen_custom_start_pos**: used to generate a user defined set of starting positions for vehicles in the network
#
# ### 2.2 Specifying the Starting Position of Edges
#
# All of the above functions starting with "specify" receive no inputs, and return a list of tuples in which the first element of the tuple is the name of the edge/intersection/internal_link, and the second value is the distance of the link from some global reference, i.e. [(link_0, pos_0), (link_1, pos_1), ...].
#
# The data specified in `specify_edge_starts` is used to provide a "global" sense of the location of vehicles, in one dimension. This is done either through the `get_x_by_id` method within an environment, or the `get_absolute_position` method in the `Vehicles` object within an environment. The `specify_internal_edge_starts` allows us to do the same to junctions/internal links when they are also located within the network (this is not the case for the ring road).
#
# In section 1, we created a network with 4 edges named: "edge0", "edge1", "edge2", and "edge3". We assume that the edge titled "edge0" is the origin, and accordingly the position of the edge start of "edge0" is 0. The next edge, "edge1", begins a quarter of the length of the network from the starting point of edge "edge0", and accordingly the position of its edge start is radius * pi/2. This process continues for each of the edges. We can then define the starting position of the edges as follows:
# +
# import some math functions we may use
from numpy import pi
class myScenario(myScenario): # update my scenario class
def specify_edge_starts(self):
r = self.net_params.additional_params["radius"]
edgestarts = [("edge0", 0),
("edge1", r * 1/2 * pi),
("edge2", r * pi),
("edge3", r * 3/2 * pi)]
return edgestarts
# -
# ## 3. Testing the New Scenario
# In this section, we run a new sumo simulation using our newly generated scenario class. For information on running sumo experiments, see `exercise01_sumo.ipynb`.
#
# We begin by defining some of the components needed to run a sumo experiment.
# +
from flow.core.params import VehicleParams
from flow.controllers import IDMController, ContinuousRouter
from flow.core.params import SumoParams, EnvParams, InitialConfig, NetParams
vehicles = VehicleParams()
vehicles.add(veh_id="human",
acceleration_controller=(IDMController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=22)
sumo_params = SumoParams(sim_step=0.1, render=True)
initial_config = InitialConfig(bunching=40)
# -
# For visualizing purposes, we use the environment `AccelEnv`, as it works on any given scenario.
# +
from flow.envs.loop.loop_accel import AccelEnv, ADDITIONAL_ENV_PARAMS
env_params = EnvParams(additional_params=ADDITIONAL_ENV_PARAMS)
# -
# Next, using the `ADDITIONAL_NET_PARAMS` component see created in section 1.1, we prepare the `NetParams` component.
additional_net_params = ADDITIONAL_NET_PARAMS.copy()
net_params = NetParams(additional_params=additional_net_params)
# We are ready now to create and run our scenario. Using the newly defined scenario classes, we create a scenario object and feed it into a `Experiment` simulation. Finally, we are able to visually confirm that are network has been properly generated.
# +
from flow.core.experiment import Experiment
scenario = myScenario( # we use the newly defined scenario class
name="test_scenario",
vehicles=vehicles,
net_params=net_params,
initial_config=initial_config
)
# AccelEnv allows us to test any newly generated scenario quickly
env = AccelEnv(env_params, sumo_params, scenario)
exp = Experiment(env)
# run the sumo simulation for a set number of time steps
_ = exp.run(1, 1500)
| tutorials/tutorial06_scenarios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/suhailnajeeb/ete-ice-399/blob/master/Experiment4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qfkmE669nHsj" colab_type="text"
# # Experiment No 4: Frequency & Phase Modulation (FM, PM)
# + id="AjItKPEyniAp" colab_type="code" colab={}
import numpy as np
from math import pi
from numpy import cos, absolute
import matplotlib.pyplot as plt
from numpy.fft import fft, fftshift
# + [markdown] id="yTw-VqxPnbX4" colab_type="text"
# ## FM Modulation
# + id="wVZZwgPAm7ep" colab_type="code" colab={}
fs = 8000
fc = 100
Ac = 1
fm = 5
Am = 1
kf = 100*pi
# + id="47WexCMQnkxx" colab_type="code" colab={}
t = np.arange(0,1,1/fs)
M = Am*cos(2*pi*fm*t)
C = Ac*cos(2*pi*fc*t)
# + id="2v5iB2a2nyBo" colab_type="code" colab={}
#M_int = np.zeros((1,len(t)))
M_int = []
for i in range(len(t)):
M_int.append(0.5*(1/fs)*(2*np.sum(M[:i]-M[0]-M[i])))
M_int = np.asarray(M_int)
# + id="U1HOBZGsrrUm" colab_type="code" colab={}
s_FM = Ac*cos(2*pi*fc*t + kf*M_int)
# + id="E_vMXYNmr-NZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 281} outputId="9c3e3116-e04e-4ce2-eb4a-54fcfeca761a"
fig, axs = plt.subplots(3,1)
axs[0].plot(t,M)
axs[0].set(ylabel = 'm(t)')
axs[0].set_title('Message Signal')
axs[1].plot(t,C)
axs[1].set(ylabel = 'c(t)')
axs[1].set_title('Carrier Signal')
axs[2].plot(t,s_FM,t,M)
axs[2].set(ylabel ='s_{FM}(t)')
axs[2].set_title('FM Signal')
plt.show()
# + id="Z8pQs0QFs3A-" colab_type="code" colab={}
M_f = fftshift(absolute(fft(M)))
f = np.linspace(-fs/2, fs/2, len(M_f))
# + id="P7cRoYC0wO_g" colab_type="code" colab={}
C_f = fftshift(absolute(fft(C)))
# + id="MDOGFEuKwRPA" colab_type="code" colab={}
S_FM_f = fftshift(absolute(fft(s_FM)))
beta = (kf*Am/(2*pi))/fm
BW = 2*fm*(beta+1)
BW_bound = np.zeros(len(f))
for i in range(len(f)):
if(abs(f[i])<(fc + BW/2) and abs(f[i]>(fc - BW/2))):
BW_bound[i] = 1
# + id="aosGn6PkwUqI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 423} outputId="171a4590-11b4-4347-afc6-02612e06a114"
fig, axs = plt.subplots(3,1)
axs[0].stem(f, M_f/M_f.max())
axs[0].set(xlabel = 'Frequency (Hz)', ylabel = 'Normalized |M(f)|', xlim = (-110,110))
axs[0].set_title('Spectrum of message Signal')
axs[1].stem(f, C_f/C_f.max())
axs[1].set(xlabel = 'Frequency (Hz)', ylabel = 'Normalized |C(f)|', xlim = (-110,110))
axs[1].set_title('Spectrum of carrier Signal')
axs[2].stem(f, S_FM_f/S_FM_f.max())
axs[2].plot(f, BW_bound, linewidth = 1)
axs[2].set(xlabel = 'Frequency (Hz)', ylabel = 'Normalized |S_{FM}(f)|', xlim = (-200,200))
axs[2].set_title('Spectrum of FM Signal')
plt.show()
| Experiment4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="gLB4anKXLM01" executionInfo={"status": "ok", "timestamp": 1620704471821, "user_tz": 420, "elapsed": 62111, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNaHwwca9YSThBL83juKHaVmkiQeJpIrJM_KI=s64", "userId": "10584158831815703022"}} outputId="4963de62-5131-4a59-c40d-75a856a99995"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="dFlTa5S7DRYZ"
# # Aux.ai Project
# **Milestone 3**
#
# ## Imports
# + id="nVNKpPqh_-ee"
from music21 import converter, instrument, note, chord, stream
from keras.utils.np_utils import to_categorical
from tensorflow import keras
from tensorflow.keras.callbacks import ModelCheckpoint
import numpy as np
import glob
import os
import json
PROJECT_PATH = "/content/drive/MyDrive/Term 8/DS2.4/final-project"
COMPOSER = "rachmaninov"
MIDI_PATH = os.path.join(PROJECT_PATH, "midi", COMPOSER)
NOTES_PATH = os.path.join(MIDI_PATH, "notes.json")
WEIGHTS_PATH = os.path.join(PROJECT_PATH, "weights")
# + [markdown] id="8VQvVKbXrW1C"
# ## Encode notes
# Because we are use MIDI files as input data, these have to be encoded into notes that we can feed the neural network. For this project I am only using the piano part and am encoding the notes as follows:
# - Single Note - pitch:duration
# - Chord - pitch,pitch,pitch:duration
# + colab={"base_uri": "https://localhost:8080/"} id="AOIYKQdnKSvG" executionInfo={"status": "ok", "timestamp": 1620721227183, "user_tz": 420, "elapsed": 1315, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNaHwwca9YSThBL83juKHaVmkiQeJpIrJM_KI=s64", "userId": "10584158831815703022"}} outputId="34055f70-58b3-4821-ec76-b86203cdea7c"
notes = []
song_limit = 100
if os.path.exists(NOTES_PATH):
# Notes already exist
print("Notes already exist")
with open(NOTES_PATH, 'r') as f:
data = json.load(f)
notes = data['notes']
else:
# Notes do not exist yet
print("Notes do not exist yet")
for i, file in enumerate(glob.glob(os.path.join(MIDI_PATH, "*.mid"))):
if i >= song_limit:
break
print(f"Looking at file: {file}")
try:
midi = converter.parse(file)
except:
print(f"Could not parse file: {file}")
notes_to_parse = None
try:
parts = instrument.partitionByInstrument(midi).parts
except:
print(f"Skipping {file} because of parts issue")
continue
# Find piano part index
piano_index = 0
for i, part in enumerate(parts):
if part.partName == "Piano":
piano_index = i
break
# Get notes from piano part
if parts:
notes_to_parse = parts[piano_index].recurse()
else:
notes_to_parse = midi.flat.notes
# Encode each note
for element in notes_to_parse:
if isinstance(element, note.Note):
notes.append(f"{element.pitch}:{element.duration.quarterLength}")
elif isinstance(element, chord.Chord):
notes.append(f"{','.join(str(n) for n in element.normalOrder)}:{element.duration.quarterLength}")
# Write notes.json file
with open(NOTES_PATH, 'w') as f:
data = {
"notes": notes
}
f.write(json.dumps(data))
print(f"Notes: {notes}")
# + [markdown] id="VPrfYxbArbhT"
# ## Create Network Input
#
# The model expects input in the form of sequences of encoded notes and the correct next note after that sequence. Therefore I define a sequence length and create a network input and output variables which essentially act like X_train, y_train. I do not split the data into test and validation sets because I do not have any evaluation established for the model besides loss.
#
# An important point here is that I am not using the string representation(encoded version) of the notes as input/output of the model, instead I convert them to numbers using a dictionary mapping. This is what note_to_int is.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="KT5Mrp3gK_xp" executionInfo={"status": "ok", "timestamp": 1620721229596, "user_tz": 420, "elapsed": 882, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNaHwwca9YSThBL83juKHaVmkiQeJpIrJM_KI=s64", "userId": "10584158831815703022"}} outputId="0204c3be-a4ea-43b4-e8a0-95560a31934f"
sequence_length = 40
# Create note to int dictionary
pitch_names = sorted(set(item for item in notes))
note_to_int = dict((note, number) for number, note in enumerate(pitch_names))
n_vocab = len(note_to_int.keys())
network_input = []
network_output = []
# Create network input sequences and corresponding outputs
for i in range(0, len(notes) - sequence_length, 1):
sequence_in = notes[i:i + sequence_length]
sequence_out = notes[i + sequence_length]
network_input.append([note_to_int[item] for item in sequence_in])
network_output.append(note_to_int[sequence_out])
n_patterns = len(network_input)
# Reshape and normalize input
network_input = np.reshape(network_input, (n_patterns, sequence_length, 1))
network_input = network_input / float(n_vocab)
print(f"Vocab Length: {n_vocab}")
network_output = to_categorical(network_output)
# + [markdown] id="Y1qNDApfE5-v"
# ## Create network_model_object which is used in the API.
# This is specific to each model/composer
# + id="6LGghbBdJOk1"
# Create network_object json for the API
input_json_model_object = {
"network_input": network_input.tolist()[:int(len(network_input)/4)],
"network_input_shape": network_input.shape,
"pitch_names": pitch_names,
"sequence_length": int(sequence_length),
"n_vocab": int(n_vocab)
}
with open(os.path.join(PROJECT_PATH, "models",f"{COMPOSER}_network_object_seqlen_{sequence_length}.json"), 'w') as f:
json.dump(input_json_model_object, f)
# + [markdown] id="SIUT5ubFFDrI"
# ## Define Model
# This is my LSTM model which has been proven to work pretty well for the input data that I have.
# + id="z8KeJjgmHOLK"
from tensorflow.keras import backend as K
class Attention(keras.layers.Layer):
def __init__(self, return_sequences=True):
self.return_sequences = return_sequences
super(Attention,self).__init__()
def build(self, input_shape):
self.W=self.add_weight(name="att_weight", shape=(input_shape[-1],1),
initializer="normal")
self.b=self.add_weight(name="att_bias", shape=(input_shape[1],1),
initializer="zeros")
super(Attention,self).build(input_shape)
def call(self, x):
e = K.tanh(K.dot(x,self.W)+self.b)
a = K.softmax(e, axis=1)
output = x*a
if self.return_sequences:
return output
return K.sum(output, axis=1)
def get_config(self):
config = super().get_config().copy()
config.update({
'return_sequences': self.return_sequences,
})
return config
# + id="Q5SODW3QW1kW"
model = keras.models.Sequential([
keras.layers.LSTM(512, input_shape=(network_input.shape[1], network_input.shape[2]), return_sequences=True),
keras.layers.Dropout(0.2),
Attention(return_sequences=True),
keras.layers.LSTM(512, return_sequences=True),
keras.layers.Dropout(0.2),
keras.layers.Flatten(input_shape=(sequence_length, 512)),
keras.layers.Dense(256),
keras.layers.Dropout(0.2),
keras.layers.Dense(n_vocab, activation='softmax'),
])
model.compile(loss='categorical_crossentropy', optimizer='rmsprop')
# + [markdown] id="A0cuzwfOFOO0"
# ## Fit the model
# + id="pPGjSFiZhAjU"
model_fn = f"{COMPOSER}_seqlen_{sequence_length}_2LSTM_1Attention_2Dense.hdf5"
model_fp = os.path.join(PROJECT_PATH, "models", COMPOSER, model_fn)
model.load_weights(model_fp)
# + colab={"base_uri": "https://localhost:8080/"} id="XbBo4P2ocr5F" executionInfo={"status": "ok", "timestamp": 1620722412202, "user_tz": 420, "elapsed": 1171531, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNaHwwca9YSThBL83juKHaVmkiQeJpIrJM_KI=s64", "userId": "10584158831815703022"}} outputId="335bdf6e-f885-4bc3-c01c-b47f38cc15a3"
# Add checkpoints
model.fit(network_input, network_output, epochs=200, batch_size=512)
model_fn = f"{COMPOSER}_seqlen_{sequence_length}_2LSTM_1Attention_2Dense.hdf5"
model_fp = os.path.join(PROJECT_PATH, "models", COMPOSER, model_fn)
model.save(model_fp)
# + [markdown] id="b0Jn_OFOFRsm"
# ## Generate Notes
#
# This part of the code generates notes using the model we just trained. All it needs as input is a random sequence of notes of the same sequence length as was used in training, and then I essentially use that as a starting point and generate n notes, giving the model sequences and getting a note one by one as the prediction.
# + colab={"base_uri": "https://localhost:8080/"} id="gYluFh2G-J3O" executionInfo={"status": "ok", "timestamp": 1620722948409, "user_tz": 420, "elapsed": 13903, "user": {"displayName": "<NAME>", "photoUrl": "<KEY>", "userId": "10584158831815703022"}} outputId="5174de48-373b-4a6e-81be-b7c9a348281e"
start = np.random.randint(0, len(network_input)-1)
print(f"start: {start}")
int_to_note = dict((number, note) for number, note in enumerate(pitch_names))
pattern = network_input[start]
prediction_output = []
# print(f"start pattern: {pattern}")
# Generate n notes
notes_to_generate = 400
for note_index in range(notes_to_generate):
prediction_input = np.reshape(pattern, (1, len(pattern), 1))
prediction = model.predict(prediction_input, verbose=0)[0]
prediction_index = np.argmax(prediction)
prediction_note = int_to_note[prediction_index]
prediction_output.append(prediction_note)
prediction_result = prediction_index / float(n_vocab)
# print(f"Pattern: {pattern[-5:]}")
pattern = np.append(pattern, prediction_result)
pattern = pattern[1:len(pattern)]
print(f"Output notes: {prediction_output}")
# + [markdown] id="Ak3I4XClFnfy"
# ## Convert back to MIDI file
#
# Here I decode the generates notes using the inverse of the note_to_int dictionary I had previously, and write a Midi file
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="2hjBzDtl-OI6" executionInfo={"status": "ok", "timestamp": 1620722950110, "user_tz": 420, "elapsed": 809, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjNaHwwca9YSThBL83juKHaVmkiQeJpIrJM_KI=s64", "userId": "10584158831815703022"}} outputId="d5478ea1-07a3-4322-ce85-2dce2f647913"
import random
offset = 0
output_notes = []
# create note and chord objects based on the values generated by the model
for pattern in prediction_output:
# pattern is a chord
if (',' in pattern) or pattern.isdigit():
pitch = pattern.split(":")[0]
duration = pattern.split(":")[1]
notes_in_chord = pitch.split(',')
chord_notes = []
for current_note in notes_in_chord:
new_note = note.Note(int(current_note))
new_note.storedInstrument = instrument.AltoSaxophone()
chord_notes.append(new_note)
new_chord = chord.Chord(chord_notes)
new_chord.offset = offset
output_notes.append(new_chord)
else:
pitch = pattern.split(":")[0]
duration = pattern.split(":")[1]
try:
new_note = note.Note(pitch)
except:
continue
new_note.offset = offset
new_note.storedInstrument = instrument.AltoSaxophone()
output_notes.append(new_note)
# increase offset each iteration so that notes do not stack
if '/' in duration:
duration = float(int(duration.split('/')[0])/int(duration.split('/')[1]))
offset += float(duration)
midi_stream = stream.Stream(output_notes)
midi_stream.write('midi', fp='test_output.mid')
# + [markdown] id="3x-6DHf_FwhK"
# ## A lot of work for this milestone was done on the Frontend of the application, and the link for the github repo is below:
#
# - Github repo: https://github.com/APNovichkov/aux-ai-frontend
# + id="4FNut4bjn2eq"
| notebooks/aux.io_V2_playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Road Following - Live demo
# 在这个笔记本中,我们将使用我们训练的模型,使Jetbot在轨道上平稳地移动。
#
# ### Load Trained Model
# 假设您已经按照“train_model.ipynb”笔记本中的说明将 ``best_steering_model_xy.pth`` 下载到工作站。现在,您应该将模型文件上传到JetBot的笔记本目录中。完成后,该笔记本的目录中应该有一个名为 ``best_steering_model_xy.pth`` 文件。
# > 在调用下一个单元格之前,请确保文件已完全上载
# 执行下面的代码来初始化PyTorch模型。从培训笔记本上看,这应该很熟悉。执行下面的代码来初始化PyTorch模型。从培训笔记本上看,这应该很熟悉。
# +
import torchvision
import torch
model = torchvision.models.resnet18(pretrained=False)
model.fc = torch.nn.Linear(512, 2)
# -
# 接下来,从上传的 ``best_steering_model_xy.pth`` 文件中加载训练过的权重。
model.load_state_dict(torch.load('best_steering_model_xy.pth'))
# 目前,模型权重位于CPU内存上,执行下面的代码传输到GPU设备。
device = torch.device('cuda')
model = model.to(device)
model = model.eval().half()
# ### Creating the Pre-Processing Function
# ### 创建预处理功能
# 现在已经加载模型,但有一个小问题。我们训练模型的格式与相机的格式不完全匹配。为此,我们需要做一些预处理。这包括以下步骤:
# 1. 从HWC布局转换为CHW布局
# 2. 使用与训练期间相同的参数进行规格化(相机提供[0,255]范围值和[0,1]范围的训练加载图像,因此我们需要缩放255.0
# 3. 将数据从CPU存储器传输到GPU存储器
# 4. 添加批次维度
# +
import torchvision.transforms as transforms
import torch.nn.functional as F
import cv2
import PIL.Image
import numpy as np
mean = torch.Tensor([0.485, 0.456, 0.406]).cuda().half()
std = torch.Tensor([0.229, 0.224, 0.225]).cuda().half()
def preprocess(image):
image = PIL.Image.fromarray(image)
image = transforms.functional.to_tensor(image).to(device).half()
image.sub_(mean[:, None, None]).div_(std[:, None, None])
return image[None, ...]
# -
# 令人惊叹的!我们现在已经定义了我们的预处理函数,它可以将图像从摄像机格式转换为神经网络输入格式。
# 现在,让我们开始展示我们的相机。你现在应该已经很熟悉了。
# +
from IPython.display import display
import ipywidgets
import traitlets
from jetbot import Camera, bgr8_to_jpeg
camera = Camera()
image_widget = ipywidgets.Image()
traitlets.dlink((camera, 'value'), (image_widget, 'value'), transform=bgr8_to_jpeg)
display(image_widget)
# -
# 我们还将创建我们的机器人实例,我们将需要驱动马达。
# +
from jetbot import Robot
robot = Robot()
# -
# 现在,我们将定义滑块来控制JetBot
# >注意:我们已经初始化已知配置的滑块值,但是这些值可能不适用于您的数据集,因此请根据您的设置和环境增加或减少滑块
#
# 1. 速度控制(speed_gain_slider):启动JetBot增加 ``speed_gain_slider`` `
# 2. 转向增益控制(steering_gain_sloder):如果你看到Jetbot抖动,你需要减小``steering_gain_slider`` 直到它平滑
# 3. 转向偏压控制(steering_bias_slider):如果您看到JetBot偏向轨迹的最右侧或最左侧,则应控制此滑块,直到JetBot开始沿着中心的直线或轨迹移动。这就解释了运动偏差和相机偏移
#
# >注意:您应该在上述滑块周围以较低的速度播放,以获得平滑的JetBot道路跟随行为。
# +
speed_gain_slider = ipywidgets.FloatSlider(min=0.0, max=1.0, step=0.01, description='speed gain')
steering_gain_slider = ipywidgets.FloatSlider(min=0.0, max=1.0, step=0.01, value=0.2, description='steering gain')
steering_dgain_slider = ipywidgets.FloatSlider(min=0.0, max=0.5, step=0.001, value=0.0, description='steering kd')
steering_bias_slider = ipywidgets.FloatSlider(min=-0.3, max=0.3, step=0.01, value=0.0, description='steering bias')
display(speed_gain_slider, steering_gain_slider, steering_dgain_slider, steering_bias_slider)
# -
# 接下来,让我们展示一些滑块,看看JetBot在想什么。x和y滑块将显示预测的x、y值。
#
# 转向滑块将显示我们估计的转向值。请记住,这个值不是目标的实际角度,而是几乎成比例。当实际角度为 ``0`` 时,该值为零,并随实际角度增加/减少。
# +
x_slider = ipywidgets.FloatSlider(min=-1.0, max=1.0, description='x')
y_slider = ipywidgets.FloatSlider(min=0, max=1.0, orientation='vertical', description='y')
steering_slider = ipywidgets.FloatSlider(min=-1.0, max=1.0, description='steering')
speed_slider = ipywidgets.FloatSlider(min=0, max=1.0, orientation='vertical', description='speed')
display(ipywidgets.HBox([y_slider, speed_slider]))
display(x_slider, steering_slider)
# -
# 接下来,我们将创建一个函数,每当相机的值发生变化时,该函数将被调用。此函数将执行以下步骤
# 1. 预处理摄像机图像
# 2. 执行神经网络
# 3. 计算近似转向值
# 4. 使用比例/微分控制(PD)控制电机
# +
angle = 0.0
angle_last = 0.0
def execute(change):
global angle, angle_last
image = change['new']
xy = model(preprocess(image)).detach().float().cpu().numpy().flatten()
x = xy[0]
y = (0.5 - xy[1]) / 2.0
x_slider.value = x
y_slider.value = y
speed_slider.value = speed_gain_slider.value
angle = np.arctan2(x, y)
pid = angle * steering_gain_slider.value + (angle - angle_last) * steering_dgain_slider.value
angle_last = angle
steering_slider.value = pid + steering_bias_slider.value
robot.left_motor.value = max(min(speed_slider.value + steering_slider.value, 1.0), 0.0)
robot.right_motor.value = max(min(speed_slider.value - steering_slider.value, 1.0), 0.0)
execute({'new': camera.value})
# -
# 太酷了!我们已经创建神经网络执行函数,现在需要将它附加到相机上进行处理。
#
# 我们通过 observe功能来实现这一点。
# >警告:此代码将移动机器人!!请确保您的机器人有净空,并且它在乐高或轨道上,您已经收集了数据。道路跟随者应该可以工作,但是神经网络只和它训练的数据一样好!
camera.observe(execute, names='value')
# 令人惊叹的!如果你的小车已经接入,现在应该会在每一个新的相机帧上产生新的命令。
#
# 你现在可以把JetBot放在乐高或者你收集到数据的轨道上,看看它是否可以跟踪轨道。
#
# 如果要停止此行为,可以通过执行下面的代码取消附加此回调。
camera.unobserve(execute, names='value')
robot.stop()
# ### 结论
# 这就是现场演示!希望你在看到你的Jetbot在赛道上沿着公路平稳地移动时玩得开心!!!
#
# 如果你的Jetbot跟不上路,试着找出它失败的地方。好处是我们可以为这些故障场景收集更多的数据,JetBot应该会变得更好:)
| notebooks/road_following/live_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
# -
# ### Download data
# + language="bash"
#
# wget -nc -O news-final.csv http://archive.ics.uci.edu/ml/machine-learning-databases/00432/Data/News_Final.csv
#
# wget -nc -O facebook-microsoft.csv http://archive.ics.uci.edu/ml/machine-learning-databases/00432/Data/Facebook_Microsoft.csv
# wget -nc -O googleplus-microsoft.csv http://archive.ics.uci.edu/ml/machine-learning-databases/00432/Data/GooglePlus_Microsoft.csv
# wget -nc -O linkedin-microsoft.csv http://archive.ics.uci.edu/ml/machine-learning-databases/00432/Data/LinkedIn_Microsoft.csv
#
# -
# ### Load data and set index
# +
news = pd.read_csv('news-final.csv')
news = news[news['Topic']=='microsoft']
news = news.drop('Topic', 1)
news['IDLink'] = news['IDLink'].astype(int)
news = news.set_index('IDLink')
facebook = pd.read_csv('facebook-microsoft.csv')
facebook = facebook.set_index('IDLink')
google = pd.read_csv('googleplus-microsoft.csv')
google = google.set_index('IDLink')
linkedin = pd.read_csv('linkedin-microsoft.csv')
linkedin = linkedin.set_index('IDLink')
# -
# ### Restrict to articles appearing on all 3 sites
# +
posted_everywhere = news[['Facebook', 'GooglePlus', 'LinkedIn']].min(axis=1) > -1
news = news[posted_everywhere]
# -
# ### Restrict to sources with at least 50 articles
# +
greater_than_50 = (news['Source'].value_counts() > 50)
sources_50 = greater_than_50[greater_than_50 > 0].index
print("Number of sources with at least 50 articles: {}".format(greater_than_50.sum()))
news = news[news['Source'].isin(sources_50)]
# -
# ### Process dates
# +
news.loc[:, 'PublishDate'] = pd.to_datetime(news['PublishDate'])
news.loc[:, 'Year'] = news['PublishDate'].dt.year
news.loc[:, 'Month'] = news['PublishDate'].dt.month
news.loc[:, 'DayOfWeek'] = news['PublishDate'].dt.weekday
news.loc[:, 'Hour'] = news['PublishDate'].dt.hour
# -
import datetime
# +
# datetime.datetime?
# +
news['PublishDate'].max()
may_1_2016 = datetime.datetime(2016, 5, 1)
news_historical = news[news['PublishDate'] <= may_1_2016]
news_new = news[news['PublishDate'] > may_1_2016]
# -
news_new.to_csv('news-new.csv')
# ### Summary viz
news.head(2)
# **Year published**
news['PublishDate'].dt.year.value_counts().plot.bar()
# **Day of week**
news['PublishDate'].dt.weekday.value_counts().plot.bar()
# **Sentiment**
news['ones'] = 1
# news.plot.scatter('SentimentTitle', 'ones', alpha=0.1, figsize=(14, 0.5))
#
# plt.axvline(news['SentimentTitle'].mean(), alpha = 0.4, ls='--')
# ## ML
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.preprocessing import OneHotEncoder
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn import base
class ColumnSelectTransformer(base.BaseEstimator, base.TransformerMixin):
def __init__(self, col_names):
self.col_names = col_names # We will need these in transform()
def fit(self, X, y=None):
# This transformer doesn't need to learn anything about the data,
# so it can just return self without any further processing
return self
def transform(self, X):
# Return an array with the same number of rows as X and one
# column for each in self.col_names
return X[self.col_names].values #REMOVE
df = news_historical.drop(['PublishDate', 'Title', 'Headline'], 1)
# +
source_pipe = Pipeline([
('cst', ColumnSelectTransformer(['Source'])),
('oh', OneHotEncoder())
])
source_pipe.fit_transform(df)
hour_pipe = Pipeline([
('cst', ColumnSelectTransformer(['Hour'])),
('oh', OneHotEncoder())
])
sentiment_pipe = Pipeline([
('cst', ColumnSelectTransformer(['SentimentTitle', 'SentimentHeadline']))
])
features = FeatureUnion([
('source', source_pipe),
('hour', hour_pipe),
('sentiment', sentiment_pipe)
])
features.fit_transform(df)
# +
facebook_model = Pipeline([
('features', features),
('model', LinearRegression())
])
facebook_model.fit(df, df['Facebook'])
google_model = Pipeline([
('features', features),
('model', LinearRegression())
])
google_model.fit(df, df['GooglePlus'])
linkedin_model = Pipeline([
('features', features),
('model', LinearRegression())
])
linkedin_model.fit(df, df['LinkedIn'])
# -
# ## Validate models
df_new = news_new
# +
print('training: {}'.format(facebook_model.score(df, df['Facebook'])))
print('testing: {}'.format(facebook_model.score(df_new, df_new['Facebook'])))
# +
print('training: {}'.format(google_model.score(df, df['GooglePlus'])))
print('testing: {}'.format(google_model.score(df_new, df_new['Facebook'])))
# -
linkedin_model.score(df_new, df_new['LinkedIn'])
# ## Use models
# +
facebook_predict = facebook_model.predict(df_new)
google_predict = google_model.predict(df_new)
linkedin_predict = linkedin_model.predict(df_new)
# +
df_new['FacebookPrediction'] = facebook_predict
df_new['GooglePrediction'] = google_predict
df_new['LinkedInPrediction'] = linkedin_predict
# -
# ### Compare predicted resutls
df_new[['Title', 'Source', 'PublishDate', 'FacebookPrediction', 'GooglePrediction', 'LinkedInPrediction']]\
.head()
| exploratory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib as mpl
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import matplotlib.pyplot as plt
import math
mpl.rcParams['legend.fontsize'] = 10
fig = plt.figure()
ax = fig.gca(projection="3d")
t = np.linspace(-3*np.pi,3*np.pi,100)
z = math.sqrt(2/15)*math.pi / 2 * t
x = np.sin(t)
y = np.cos(t)
ax.plot(x,y,z,label='spiral array')
ax.legend()
t = np.linspace(-3*np.pi,3*np.pi,100)
z1 = math.pi * t
x1 = np.sin(t)
y1 = np.cos(t)
ax.plot(x1,y1,z1,label='spiral array2')
ax.legend()
plt.show()
# -
mpl.rcParams['legend.fontsize'] = 10
# +
# fig = plt.figure()
# ax = fig.gca(projection="3d")
# theta = np.linspace(-4 * np.pi, 4* np.pi, 100)
# z = np.linspace(-2,2,100)
# r = z**2 + 1
# x = r * np.sin(theta)
# y = r * np.cos(theta)
# ax.plot(x,y,z,label='parametric curve')
# ax.legend()
# plt.show()
# -
fig = plt.figure()
ax = fig.gca(projection="3d")
t = np.linspace(-3*np.pi,3*np.pi,100)
z = t
x = np.sin(t)
y = np.cos(t)
ax.plot(x,y,z,label='spiral array')
ax.legend()
plt.show()
| tensorflow-GPU/tensionMusic/tensionPicture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vt-Nk3lQseTo"
# ## Welcome to my Notebook on Tide Times.
# With this notebook I hope to answer the following question:
#
#
# 1. If Japan is nearly at the other end of the world to us do they experience similar tide tables around the same time?
#
# For this I have data that I wrangled using an app I created in C# from multiple locations. I am going to use the data from Aberdeen and Hosojima, I took three years worth of measurements, to hopefully answer my question.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="8JsrK7o7q9TL" outputId="a966869b-88e7-4147-d02d-af2142391a9c"
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# !wget --no-check-certificate \
# https://github.com/xerscot/tidaltimesds/raw/63bc6c653623021b1e3dbfa4f331a626ddea7ab3/Files/CleanedData/Measurements.csv \
# -O /tmp/Measurements.csv
data_file = os.path.join('/tmp/Measurements.csv')
df = pd.read_csv(data_file)
print(df.info())
# + [markdown] id="KHJgkuC5zxtY"
# Let's have a look if we can plot the heights for a week's worth of data from the different locations in the file.
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="7Nh17Bsxz-3Q" outputId="0df5c126-e2ba-41a0-ff19-84d4487bb0be"
aberdeen = df.loc[df['Location'] == 'Aberdeen']
aberdeen_week = aberdeen.loc[(aberdeen['Date'] >= '1990/01/01') & (aberdeen['Date'] <= '1990/01/06')]
aberdeen_week.plot(kind="line", x="Date", y="Height", title='Aberdeen');
hosojima = df.loc[df['Location'] == 'Hosojima']
hosojima_week = hosojima.loc[(hosojima['Date'] >= '1990/01/01') & (hosojima['Date'] <= '1990/01/06')]
hosojima_week.plot(kind="line", x="Date", y="Height", title='Hosojima');
# + [markdown] id="ZD6rch1h6E_R"
# The time difference between Aberdeen and Hosojima is 8 hours. Let's look at a single day and see the heights per hour.
# + colab={"base_uri": "https://localhost:8080/", "height": 641} id="dRiTwTJP8M4S" outputId="a80fe103-884d-4d95-8a73-baeeb2a0c5f3"
aberdeen_day = aberdeen.loc[(aberdeen['Date'] >= '1990/06/09') & (aberdeen['Date'] < '1990/06/10')]
aberdeen_day.plot(kind="line", x="Time", y="Height", title='Aberdeen');
hosojima_day = hosojima.loc[(hosojima['Date'] >= '1990/06/09') & (hosojima['Date'] < '1990/06/10')]
hosojima_day.plot(kind="line", x="Time", y="Height", title='Hosojima');
print(aberdeen_day[aberdeen_day['Height']==aberdeen_day['Height'].max()])
print(hosojima_day[hosojima_day['Height']==hosojima_day['Height'].max()])
| Notebooks/TideTimesQ1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The TransformedDistribution class
#
# > In this post, we are going to take a look at transform distribution objects as a module. This is the summary of lecture "Probabilistic Deep Learning with Tensorflow 2" from Imperial College London.
#
# - toc: true
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Python, Coursera, Tensorflow_probability, ICL]
# - image: images/transformed_dist.png
# ## Packages
# +
import tensorflow as tf
import tensorflow_probability as tfp
import numpy as np
import matplotlib.pyplot as plt
tfd = tfp.distributions
tfpl = tfp.layers
tfb = tfp.bijectors
plt.rcParams['figure.figsize'] = (10, 6)
# -
print("Tensorflow Version: ", tf.__version__)
print("Tensorflow Probability Version: ", tfp.__version__)
# ## Overview
#
# The transformedDistribution is sort of distribution that can be defined by another base distribution and a bijector object. Tensorflow Probability offers [transformed distribution](https://www.tensorflow.org/probability/api_docs/python/tfp/distributions/TransformedDistribution) object with consistent API that can use same methods and properties of other distribution.
normal = tfd.Normal(loc=0., scale=1.)
z = normal.sample(3)
z
scale_and_shift = tfb.Chain([tfb.Shift(1.), tfb.Scale(2.)])
x = scale_and_shift.forward(z)
x
log_prob_z = normal.log_prob(z)
log_prob_z
log_prob_x = (log_prob_z - scale_and_shift.forward_log_det_jacobian(z, event_ndims=0))
log_prob_x
# Note that, the `event_ndims` argument means the number of rightmost dimensions of z make up the event shape. So in the above case, the log of the jacobian determinant is calculated for each element of the tensor z.
# Or we express it with the inverse of the bijective transformation.
log_prob_x = (log_prob_z + scale_and_shift.inverse_log_det_jacobian(x, event_ndims=0))
log_prob_x
# The result is the same as while using inverse of x.
log_prob_x = (normal.log_prob(scale_and_shift.inverse(x)) + scale_and_shift.inverse_log_det_jacobian(x, event_ndims=0))
log_prob_x
# You may notice that log probability of x can be calculated with only using z or x. In practice, most of cases uses second expression. The reason is that the z is from base distriubtion. So in terms of analysis, it is the latent variable. But x is from the data distribution, and it is the output from transformed distribution. While using mentioned approach, we can express transform object with bijector or invertible, it can be learned with best parameters for maximum likelihood.
# ```python
# # Base distribution Transformation Data distribution
# # z ~ P0 <=> x = f(z) <=> x ~ P1
#
# log_prob_x = (base_dist.log_prob(bijector.inverse(x)) + bijector.inverse_log_det_jacobian(x, event_ndims=0))
#
# ### Training
#
# x_sample = bijector.forward(base_dist.sample())
# ```
normal = tfd.Normal(loc=0., scale=1.)
z = normal.sample(3)
z
exp = tfb.Exp()
x = exp.forward(z)
x
log_normal = tfd.TransformedDistribution(normal, exp)
log_normal
# Above expression is same with like this,
log_normal = exp(normal)
log_normal
log_normal.sample()
log_normal.log_prob(x)
# We can also define specific `event_shape` and `batch_shape` for transformedDistribtion.
normal = tfd.Normal(loc=0., scale=1.)
scale_tril = [[1., 0.], [1., 1.]]
scale = tfb.ScaleMatvecTriL(scale_tril=scale_tril)
# Multivariate Normal distribution
mvn = tfd.TransformedDistribution(tfd.Sample(normal, sample_shape=[2]), scale)
mvn
# +
scale_tril = [[[1., 0.], [1., 1.]], [[0.5, 0.], [-1., 0.5]]]
scale = tfb.ScaleMatvecTriL(scale_tril=scale_tril)
mvn = tfd.TransformedDistribution(tfd.Sample(tfd.Normal(loc=[0., 0.], scale=1.), sample_shape=[2], ), scale)
mvn
# -
# ## Tutorial
# ### TransformedDistribution
# +
# Parameters
n = 10000
loc = 0
scale = 0.5
# +
# Normal distribution
normal = tfd.Normal(loc=loc, scale=scale)
# +
# Display event and batch shape
print('batch shape: ', normal.batch_shape)
print('event shape: ', normal.event_shape)
# -
# Exponential bijector
exp = tfb.Exp()
# +
# log normal transformed distribution using exp bijector and normal distribution
log_normal_td = exp(normal)
# +
# Display event and batch shape
print('batch shape: ', log_normal_td.batch_shape)
print('event shape: ', log_normal_td.event_shape)
# +
# Base distribution
z = normal.sample(n)
# -
# ### Plots
# +
# Plot z density
# -
plt.hist(z.numpy(), bins=100, density=True)
plt.show()
# +
# Transformed distribution
x = log_normal_td.sample(n)
# -
# Plot x density
plt.hist(x.numpy(), bins=100, density=True)
plt.show()
# +
# Define log normal distribution
log_normal = tfd.LogNormal(loc=loc, scale=scale)
l = log_normal.sample(n)
# -
plt.hist(l.numpy(), bins=100, density=True)
plt.show()
# ### Log probability
# +
# Log prob of LogNormal
log_prob = log_normal.log_prob(x)
# +
# Log prob of log normal transformed distribution
log_prob_td = log_normal_td.log_prob(x)
# +
# Check log probs
tf.norm(log_prob - log_prob_td)
# -
# ### Event shape and batch shape
# +
# Set a scaling lower triangular matrix
tril = tf.random.normal((2, 4, 4))
scale_low_tri = tf.linalg.LinearOperatorLowerTriangular(tril)
# +
# view of scale_low_tri
scale_low_tri.to_dense()
# +
# Define scale linear operator
scale_lin_op = tfb.ScaleMatvecLinearOperator(scale_low_tri)
# +
# Define scale linear operator transformed distribution with a batch and event shape
mvn = tfd.TransformedDistribution(tfd.Sample(tfd.Normal(loc=[0., 0.], scale=1.), sample_shape=[4]), scale_lin_op)
# +
# Display event and batch shape
print('batch shape: ', mvn.batch_shape)
print('event shape: ', mvn.event_shape)
# +
# Sample
y1 = mvn.sample(sample_shape=(n,))
print(y1.shape)
# +
# Define a MultivariateNormalLinearOperator distribution
mvn2 = tfd.MultivariateNormalLinearOperator(loc=0, scale=scale_low_tri)
mvn2
# -
# sample
y2 = mvn2.sample(sample_shape=(n, ))
y2.shape
# +
# Check
xn = normal.sample((n, 2, 4))
tf.norm(mvn.log_prob(xn) - mvn2.log_prob(xn)) / tf.norm(mvn.log_prob(xn))
| _notebooks/2021-09-07-01-the-transformedDistribution-class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (mmaction-lite)
# language: python
# name: pycharm-d922ce35
# ---
# + pycharm={"is_executing": false}
from mmaction.apis import init_recognizer, inference_recognizer
# + pycharm={"is_executing": false}
config_file = '../configs/recognition/tsn/tsn_r50_video_inference_1x1x3_100e_kinetics400_rgb.py'
# download the checkpoint from model zoo and put it in `checkpoints/`
checkpoint_file = '../checkpoints/tsn_r50_1x1x3_100e_kinetics400_rgb_20200614-e508be42.pth'
# + pycharm={"is_executing": false}
# build the model from a config file and a checkpoint file
model = init_recognizer(config_file, checkpoint_file, device='cpu')
# + pycharm={"is_executing": false}
# test a single video and show the result:
video = 'demo.mp4'
label = '../tools/data/kinetics/label_map_k400.txt'
results = inference_recognizer(model, video, label)
# + pycharm={"is_executing": false, "name": "#%%\n"}
# show the results
for result in results:
print(f'{result[0]}: ', result[1])
| demo/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
ROOT = r"D:\02PHBS_G2\PHBS_m2\bigDataAnalysis\big-data-hft\project"
os.chdir(ROOT)
# +
ref_morning_time = pd.date_range("9:31", "11:30", freq="1min").time
ref_afternoon_time = pd.date_range("13:01", "15:00", freq="1min").time
ref_time_arr = np.concatenate([ref_morning_time,ref_afternoon_time])
ref_time_str_list = [x.strftime("%H%M") for x in ref_time_arr]
# -
def my_fillna(df):
df[['Close']] = df[['Close']].fillna(method='ffill')
df[['Close','High','Low','Open']] = df[['Close','High','Low','Open']].fillna(method='ffill',axis=1)
df[['Amount','Count','Qty']] = df[['Amount','Count','Qty']].fillna(0)
return df
def process_mr_data(path):
raw_data = pd.read_csv(path, header=None)
raw_data.columns = ['raw']
raw_data = raw_data[raw_data["raw"].str.startswith("6") ]
tmp1 = raw_data["raw"].str.split("#")
raw_data["stock_code"] = tmp1.str[0]
tmp2 = tmp1.str[1].str.split("$")
raw_data["time_str"] = tmp2.str[0].str[8:]
tmp3 = tmp2.str[1].str.split("\t")
raw_data["feature_name"] = tmp3.str[0]
raw_data["feature_value"] = tmp3.str[1].astype("float32")
data = raw_data.iloc[:,1:]
min_data = data.pivot(index=['stock_code','time_str'],columns='feature_name',values='feature_value')
ref_code_list = min_data.index.unique(0)
ref_code_time_index =pd.MultiIndex.from_product([ref_code_list,ref_time_str_list],names=['stock_code','time_str'])
min_data_reindex = min_data.reindex(index=ref_code_time_index)
min_data_final = min_data_reindex.groupby("stock_code").apply(my_fillna)
min_data_final['Return'] = min_data_final['Close']/min_data_final['Open'] - 1
return min_data_final
raw_mr_data_dir = "./data/rawDataFromMR/"
cleaned_mins_data_dir = "./data/min_data_cleaned/"
for date_csv in os.listdir(raw_mr_data_dir):
data_path = os.path.join(raw_mr_data_dir, date_csv,date_csv,date_csv)
min_data_final = process_mr_data(data_path)
min_data_final.to_csv(os.path.join(cleaned_mins_data_dir,date_csv))
print(date_csv)
# raw data preview
pd.read_csv(data_path, header=None)
# cleaned data preview
cleaned_data = pd.read_csv(os.path.join(clearned_mins_data_dir,date_csv),index_col=[0,1])
cleaned_data
tmp = pd.read_pickle(r"D:\02PHBS_G2\PHBS_m2\bigDataAnalysis\big-data-hft\project\data\factor\tickcount_entropy_1m_l1.pkl",compression='gzip')
tmp.to_csv(r"D:\02PHBS_G2\PHBS_m2\bigDataAnalysis\big-data-hft\project\data\factor\tickcount_entropy_1m_l1.csv")
tmp=pd.read_csv(r"D:\02PHBS_G2\PHBS_m2\bigDataAnalysis\big-data-hft\project\data\factor\tickcount_entropy_1m_l1.csv",index_col=0)
tmp
tmp.index
tmp.columns
data = cleaned_data.copy()
data.index
def cal_Rvar(sample_data):
factor = np.sum(sample_data['Return']**2)
return factor
fac = data.groupby(level=0).apply(cal_Rvar)
fac.index = [str(x)+".SH" for x in fac.index]
fac.name = int(date_csv.split(".")[0])
fac
sample_data = cleaned_data.loc[600000]
sample_data.to_csv(r"D:\02PHBS_G2\PHBS_m2\bigDataAnalysis\big-data-hft\project\src\main\python\factorCalFunc\mins_data_1ticker.csv")
process_mr_data(os.path.join(ROOT,"data/20190102.csv"))
raw_data = pd.read_csv(os.path.join(ROOT,"data/20190102.csv"), header=None)
raw_data.columns = ['raw']
raw_data_bond = pd.read_csv(os.path.join(ROOT,"data/20190102.csv"), header=None)
raw_data_bond.columns = ['raw']
tmp1 = raw_data_bond["raw"].str.split("#")
raw_data_bond["stock_code"] = tmp1.str[0]
tmp2 = tmp1.str[1].str.split("$")
raw_data_bond["time_str"] = tmp2.str[0].str[8:]
tmp3 = tmp2.str[1].str.split("\t")
raw_data_bond["feature_name"] = tmp3.str[0]
raw_data_bond["feature_value"] = tmp3.str[1].astype("float32")
raw_data_bond
raw_data = raw_data[raw_data["raw"].str.startswith("6") ]
raw_data
tmp1 = raw_data["raw"].str.split("#")
raw_data["stock_code"] = tmp1.str[0]
tmp2 = tmp1.str[1].str.split("$")
raw_data["time_str"] = tmp2.str[0].str[8:]
tmp3 = tmp2.str[1].str.split("\t")
raw_data["feature_name"] = tmp3.str[0]
raw_data["feature_value"] = tmp3.str[1].astype("float32")
raw_data
data = raw_data.iloc[:,1:]
data
min_data = data.pivot(index=['stock_code','time_str'],columns='feature_name',values='feature_value')
min_data
# min_data['min_return'] = min_data['Close']/min_data['Open'] - 1
min_data
min_data.loc["600647"]
# +
ref_morning_time = pd.date_range("9:31", "11:30", freq="1min").time
ref_afternoon_time = pd.date_range("13:01", "15:00", freq="1min").time
ref_time_arr = np.concatenate([ref_morning_time,ref_afternoon_time])
ref_time_str_list = [x.strftime("%H%M") for x in ref_time_arr]
ref_code_list = min_data.index.unique(0)
ref_code_time_index =pd.MultiIndex.from_product([ref_code_list,ref_time_str_list],names=['stock_code','time_str'])
# -
ref_code_time_index
min_data_reindex = min_data.reindex(index=ref_code_time_index)
def my_fillna(df):
df[['Close']] = df[['Close']].fillna(method='ffill')
df[['Close','High','Low','Open']] = df[['Close','High','Low','Open']].fillna(method='ffill',axis=1)
df[['Amount','Count','Qty']] = df[['Amount','Count','Qty']].fillna(0)
return df
min_data_final = min_data_reindex.groupby("stock_code").apply(my_fillna)
min_data_final
min_data_final.to_csv("data/min_data_cleaned/sample_min_data_20190201.csv")
min_data_final['Return'] = min_data_final['Close']/min_data_final['Open'] - 1
gb_ret = min_data_final.groupby("stock_code")['Return']
feature_df = pd.DataFrame(index=min_data_final.index.unique(0))
min_data_final
data = pd.read_csv("data/min_data_cleaned/sample_min_data_20190201.csv")
data
data[data.stock_code==603999].to_csv("data/min_data_cleaned/min_data_ticker_20190201.csv")
sample_data = pd.read_csv("data/min_data_cleaned/min_data_ticker_20190201.csv",index_col=0)
sample_data
def cal_ret_stats(data):
factor = data["Return"].std()
return factor
cal_ret_stats(data)
feature_df['RVar'] = gb_ret.apply(lambda se: se.std()**2)
feature_df['RSkew'] = gb_ret.apply(lambda se: se.skew())
feature_df['RKurtosis'] = gb_ret.apply(lambda se: se.kurtosis())
feature_df
| project/src/main/python/tickOutput2minData/minDataPreparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import tools
reload(tools)
from tools import *
# +
source = "/home/walterms/mcmd/nn/data/unlbl/"
fnames = sorted([source+f for f in os.listdir(source)])
foutdir = "/home/walterms/mcmd/nn/data/pca/"
run = "nbrs_e3"
newnames = []
for f in fnames:
if f.endswith(".log") or f.endswith("swp"): continue
if f.startswith(source+"edge_3"):
newnames.append(f)
fnames = newnames
# Sort based on edge size
edges = []
for f in fnames:
edg = float(f[len(source)+len("edge_3_"):])
rh = 28**2 / (edg**2)
edges.append([edg,rh])
idxs = np.argsort(edges,axis=0)
edges = np.array([edges[i] for i in idxs[:,0]])
fnames = [fnames[i] for i in idxs[:,0]]
fnames, edges = fnames[::6], edges[::6]
nfiles = len(fnames)
# +
# FOR XTUD SETS
# Note, nProc = 10000, sweepEval = 100, nEquil = 0
source = "/home/walterms/mcmd/nn/data/train/"
fnames = [source+f for f in ["X","U","D","T"]]
foutdir = "/home/walterms/mcmd/nn/data/pca/"
run = "nbrs_xtud"
nfiles = 4
nrod = 28**2
edges = np.array([[6.324, nrod/(6.324**2)] for _ in range(nfiles)])
# -
edges
fidx = np.array([0,3,5,6,7,8])
fnames = np.asarray([fnames[i] for i in fidx])
edges = np.array([edges[i] for i in fidx])
nfiles = len(fnames)
print fnames
print edges
# +
# hyperparams
NBRS = np.array([10,15,20,30])
methods = ["random", "radial", "angular", "polar"]
nmethod = len(methods)
nrod = 28**2
nblskip = 10
sparse_bulk_factor = 1
NBL = 20
nsample = nrod*NBL*nfiles
sample_per_file = nsample // nfiles
print nfiles, nsample
# -
edges
# +
# Designed for handling
# the unlbl nn dataset
#
#
for n_nbr in NBRS:
for imeth, meth in enumerate(methods):
foutname = foutdir+run+"_"+str(n_nbr)+"_"+meth
print "Preparing samples for "+foutname
# Let's add an extra dimension being the density label to X
# Easy to extract later
# Adding probe rod info
X = np.empty((nsample,n_nbr+1))
isample = 0
for ifile,f in enumerate(fnames):
fin = open(f,'r')
params = {}
params.update({"ReducedRho": edges[ifile,1]})
params.update({"boxEdge": edges[ifile,0]})
print f,params["ReducedRho"]
edge = params["boxEdge"]
nbl = 0
rods = np.zeros((nrod,3))
irod = 0
isample_per_f = 0
for line in fin.readlines():
if nbl < nblskip:
if line == "\n": nbl+=1
continue
if line == "\n":
# Done a block
samples = get_nbrs(rods,n_nbr,edge,sparse_bulk_factor=sparse_bulk_factor,method=meth)
if isample_per_f + len(samples) > sample_per_file:
samples = samples[:sample_per_file]
if isample+len(samples) > nsample:
samples = samples[:nsample-isample]
len_samples = len(samples)
rh = np.ones((len_samples,1)) * params["ReducedRho"]
samples = np.append(rh,samples,axis=1)
X[isample:isample+len_samples] = samples
isample += len_samples
isample_per_f += len_samples
nbl+=1
if isample >= nsample:
break
if isample_per_f >= sample_per_file:
break
rods = np.zeros((nrod,3))
irod = 0
continue
if line.startswith("label"):
continue
rod = [float(s) for s in line.split()]
rod[0] *= edge
rod[1] *= edge
rod[2] *= twopi
rod[2] = myrotate(rod[2])
rods[irod] = rod
irod+=1
fin.close()
if isample >= nsample:
break
fout = open(foutname,'w')
print "Writing to "+foutname
for samp in X:
s = ""
for th in samp:
s += "%0.6f "%(th)
s += "\n"
fout.write(s)
fout.close()
print "Done"
# -
fnames
f_sample = fnames[0]
edge_sample = edges[0][0]
nfiles = 1
print f_sample, edge_sample
f_sample = "/home/walterms/mcmd/nn/data/unlbl/edge_3_11.71"
edge_sample = 11.71
nfiles = 1
# +
#
# This Chunk suited for looking at a single block of a single file
#
# This block is different
# It's for handling the FNN unlabeled dataset
NBL = 1
n_nbr = 30
nrod = 28**2
sparse_bulk_factor = 1
nblskip = 10
nsample = NBL*nrod*nfiles
methods = ["random", "radial", "angular", "polar"]
nmethod = len(methods)
# X = np.empty((nmethod,nsample,n_nbr))
# nbrs_full = np.empty(shape=(nmethod,nrod,n_nbr,4)) # first 4 is for each method
X = []
nbrs_full = []
alphas = []
for ifile,f in enumerate([f_sample]):
fin = open(f,'r')
print f, float(nrod)/(edge_sample*edge_sample)
nbl = 0
lambdacount = 0
rods = np.zeros((nrod,3))
irod = 0
for line in fin.readlines():
if nbl < nblskip:
if line == "\n": nbl+=1
continue
if line == "\n":
# Done a block
for im, meth in enumerate(methods):
x, nbs, alphs = \
get_nbrs(rods,n_nbr,edge_sample,sparse_bulk_factor=sparse_bulk_factor,method=meth,ret_nbrs=True)
X.append(x)
nbrs_full.append(nbs)
alphas.append(alphs)
# for im, meth in enumerate(methods):
# X[im,ifile:(ifile+1)*NBL*nrod], nbrs_full[im] = \
# get_nbrs(rods,n_nbr,edge,use_bulk=use_bulk,method=meth,ret_nbrs=True)
X = np.asarray(X)
nbrs_full = np.asarray(nbrs_full)
alphas = np.asarray(alphas)
nbl+=1
if nbl >= NBL+nblskip:
break
rods = np.zeros((nrod,3))
irod = 0
continue
if line.startswith("label"):
continue
rod = [float(s) for s in line.split()]
rod[0] *= edge_sample
rod[1] *= edge_sample
rod[2] *= twopi
rod[2] = myrotate(rod[2])
rods[irod] = rod
irod+=1
fin.close()
# print np.mean(lambdas[ifile])
# _=plt.plot(lambdas[ifile],'k.',markersize=2)
# _=plt.axis([0,nlambda,-1,1])
# plt.show()
# lvar = np.var(lambdas,axis=1)
# _ = plt.plot(edges[:,1],lvar,'o')
# _ = plt.xlabel(r'$\rho$')
# _ = plt.ylabel(r'$\sigma^2(\lambda)$',rotation="horizontal",labelpad=20)
# plt.show()
print "Done"
# +
f,ax = plt.subplots(4,2,figsize=(10,18))
irod = 200
for imeth, meth in enumerate(methods):
ax[imeth,0].set_aspect("equal")
# plotrods(rods,ax[imeth,0])
plotrods([rods[irod]],ax[imeth,0],col="r",lw=10.)
plotrods([rods[irod]],ax[imeth,0],col="k",lw=2.)
nbrs = nbrs_full[imeth,irod]
for ni, nbr in enumerate(nbrs[:,:3]):
col = (0,1-float(ni)/n_nbr,float(ni)/n_nbr)
plotrods([nbr],ax[imeth,0],col=col,lw=3.)
ax[imeth,0].set_title(meth)
ax[imeth,1].set_title(meth)
ax[imeth,1].set_ylabel(r'$feature$',rotation="vertical",labelpad=0,fontsize=14)
ax[imeth,1].plot(nbrs[:,3])
ax[imeth,1].set_ylim(-1,1)
# break
# -
| nn/pca/gen_nbrfiles.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
raw_data = pd.read_csv("seattleWeather_1948-2017.csv")
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
data = raw_data.drop("DATE",axis=1)
data["TAVG"]=(data["TMAX"]+data["TMIN"])/2
data = data.drop("PRCP", axis=1)
data = data.drop("RAIN", axis=1)
data
# Now let us import the processed data
pred_min = np.load("Min_Temp_pred.npy")
y_min = np.load("Min_Temp_y.npy")
pred_max = np.load("Max_Temp_pred.npy")
y_max = np.load("Max_Temp_y.npy")
pred_avg = np.load("Avg_Temp_pred.npy")
y_avg = np.load("Avg_Temp_y.npy")
# Now let us train the scalers
tmin = data.drop("TMAX", axis=1)
tmin = tmin.drop("TAVG", axis=1)
tmin
scaler.fit(tmin)
# And finally scale back the values
pred_min_scl = scaler.inverse_transform(pred_min)
y_min_scl = scaler.inverse_transform(y_min)
plt.scatter(range(len(pred_min_scl)), pred_min_scl)
plt.scatter(range(len(pred_min_scl)), y_min_scl)
plt.show()
# Let us see a plot in Celcius
def cel(data):
resul = (data-32)/1.8
return resul
pred_min_cel = cel(pred_min_scl)
y_min_cel = cel(y_min_scl)
plt.scatter(range(len(pred_min_cel)), pred_min_cel)
plt.scatter(range(len(pred_min_cel)), y_min_cel)
plt.show()
# And error of
import seaborn as sns
plt.hist(pred_min_cel-y_min_cel, bins = 50)
# Let us also do for the other features
tmax = data.drop("TMIN", axis=1)
tmax = tmax.drop("TAVG", axis=1)
scaler.fit(tmax)
pred_max_scl = scaler.inverse_transform(pred_max)
y_max_scl = scaler.inverse_transform(y_max)
pred_max_cel = cel(pred_max_scl)
y_max_cel = cel(y_max_scl)
plt.scatter(range(len(pred_min_cel)), pred_max_cel)
plt.scatter(range(len(pred_min_cel)), y_max_cel)
plt.show()
plt.hist(pred_max_cel-y_max_cel, bins = 50)
tavg = data.drop("TMAX", axis=1)
tavg = tavg.drop("TMIN", axis=1)
tavg
scaler.fit(tavg)
pred_avg_scl = scaler.inverse_transform(pred_avg)
y_avg_scl = scaler.inverse_transform(y_avg)
pred_avg_cel = cel(pred_avg_scl)
y_avg_cel = cel(y_avg_scl)
plt.scatter(range(len(pred_min_cel)), pred_avg_cel)
plt.scatter(range(len(pred_min_cel)), y_avg_cel)
plt.show()
plt.hist(pred_avg_cel-y_avg_cel, bins = 50)
# So we see that we are doinh super mega good! Most errors are in the max side, but more than 60% is within 1.5 C of difference, probably better than prediction
| SeattleResults.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ZEpieBvNXCME"
# # Shakespeare - text generation
# - doc: https://github.com/openaiknowledge/PR2
# - based on: https://www.kaggle.com/kutaykutlu/text-generation-guide-tensorflow-nlp-lstm
# + [markdown] id="H_R0LtNjV9sf"
# # Import Libraries
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" id="mrsqRw1HV9si" executionInfo={"status": "ok", "timestamp": 1622887079855, "user_tz": -120, "elapsed": 211, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import Embedding, LSTM, Dense, Dropout, Bidirectional
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import regularizers
from tensorflow import keras
import tensorflow.keras.utils
import tensorflow as tf
import numpy as np
import pandas as pd
import numpy as np
from google.colab import drive
# + [markdown] id="AGPtmIFhV9sj"
# # Load Dataset
# + id="ztBNwLfyYOBr" executionInfo={"status": "ok", "timestamp": 1622886757104, "user_tz": -120, "elapsed": 194, "user": {"displayName": "<NAME>\u00<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
BASE_FOLDER = '/content/drive/My Drive/openaiknowledge/pr2/'
DATA = BASE_FOLDER + 'data/1/' #version 1
MODEL = BASE_FOLDER + "model/1/"
# + colab={"base_uri": "https://localhost:8080/"} id="db_FgaRzYSZk" executionInfo={"status": "ok", "timestamp": 1622884254341, "user_tz": -120, "elapsed": 17948, "user": {"displayName": "<NAME>\u00<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="6ef24ee0-fe71-40df-8819-89a5a6940a00"
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 95} id="umBWMr87V9sk" executionInfo={"status": "ok", "timestamp": 1622884255883, "user_tz": -120, "elapsed": 866, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="c7237ccc-2efd-47ec-db6d-f4891714e59d"
#todo: use metada.json
datafile_name = DATA + "Shakespeare_data.csv"
print(datafile_name)
df = pd.read_csv(datafile_name)
df.head(1)
# + colab={"base_uri": "https://localhost:8080/"} id="reK_crwzV9sl" executionInfo={"status": "ok", "timestamp": 1622884256181, "user_tz": -120, "elapsed": 303, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="9a18bb28-077b-4068-aa49-3e8facea8f67"
import csv
corpus = []
#with open('/kaggle/input/shakespeare-plays/Shakespeare_data.csv') as f:
with open(datafile_name) as f:
reader = csv.reader(f, delimiter=',')
next(reader) # to pass first row,header
for row in reader:
corpus.append(row[5])
print(len(corpus))
print(corpus[:3])
# + [markdown] id="9afnAE31V9sl"
# # Data Cleaning
# + id="D0I30HFwV9sm" executionInfo={"status": "ok", "timestamp": 1622884256385, "user_tz": -120, "elapsed": 206, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
import string
def text_cleaner(text):
text = "".join(car for car in text if car not in string.punctuation).lower()
text = text.encode("utf8").decode("ascii",'ignore')
return text
corpus = [text_cleaner(line) for line in corpus]
# + colab={"base_uri": "https://localhost:8080/"} id="m4e8u3rnV9sm" executionInfo={"status": "ok", "timestamp": 1622884256578, "user_tz": -120, "elapsed": 195, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="bd1e53d4-808e-492f-f8d5-c3eec4a9f4d1"
# Tokenization is the process of splitting up a text into a list of individual words, or tokens.
# corpus is too big if you try with all data, you can see this message
# Your notebook tried to allocate more memory than is available. It has restarted.
corpus = corpus[:5000]
tokenizer = Tokenizer()
tokenizer.fit_on_texts(corpus)
word_index = tokenizer.word_index
total_words = len(word_index) + 1
total_words
# + [markdown] id="gfWjgXFvV9sn"
# 
# + id="cDUJ2pUwV9sn" executionInfo={"status": "ok", "timestamp": 1622884256863, "user_tz": -120, "elapsed": 287, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
# create input sequences using list of tokens
input_sequences =[]
for sentence in corpus:
token_list = tokenizer.texts_to_sequences([sentence])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
# + id="ir6zLKG_V9sn" executionInfo={"status": "ok", "timestamp": 1622884256863, "user_tz": -120, "elapsed": 6, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences,
maxlen=max_sequence_len,
padding='pre')) # as laste value is label.padding method is 'pre'
# + [markdown] id="XlYq3X1zV9so"
# - __Sequences__ and __n_Gram_squences__ for a sentence after __Padding__
#
# - Last value for every line is target, label.
#
# - line1 : 2
#
# - line2 : 66
#
# 
# + id="U2X8N6W7V9so" executionInfo={"status": "ok", "timestamp": 1622884257653, "user_tz": -120, "elapsed": 795, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
# create predictors and label
predictors, label = input_sequences[:,:-1],input_sequences[:,-1]
# create one-hot encoding of the labels
label = tensorflow.keras.utils.to_categorical(label, num_classes=total_words)
# + colab={"base_uri": "https://localhost:8080/"} id="R_BTXV2QV9so" executionInfo={"status": "ok", "timestamp": 1622884257654, "user_tz": -120, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="6fdcec25-0d69-4aa8-ae5f-1ed58c0cdca5"
print(label[0])
print(label[0].shape)
# + colab={"base_uri": "https://localhost:8080/"} id="mwIwaiBvV9sp" executionInfo={"status": "ok", "timestamp": 1622884266828, "user_tz": -120, "elapsed": 9177, "user": {"displayName": "<NAME>\u0<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="ae858fa7-c2cc-4ee0-fee5-f6efe9c1928f"
model = Sequential()
model.add(Embedding(total_words, 10, input_length=max_sequence_len-1))
model.add(Bidirectional(LSTM(512)))
model.add(Dropout(0.3))
model.add(Dense(total_words, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
# + _kg_hide-output=true colab={"base_uri": "https://localhost:8080/"} id="hB2CKWZgV9sp" executionInfo={"status": "ok", "timestamp": 1622885428171, "user_tz": -120, "elapsed": 1161358, "user": {"displayName": "<NAME>\u0<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="61ba2658-1b72-4ed0-9e4d-4ebc6ef61f17"
history = model.fit(predictors, label, epochs=50, verbose=1)
# + id="NTgVkuR3czMX" executionInfo={"status": "ok", "timestamp": 1622886778501, "user_tz": -120, "elapsed": 694, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
#save model
model_name = "shakespeare_text"
model_file_name = MODEL + model_name + '.h5'
model.save(model_file_name)
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="P-e5bSpkV9sp" executionInfo={"status": "ok", "timestamp": 1622886806163, "user_tz": -120, "elapsed": 937, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="cc1c3a31-243c-4a0b-fa3f-86c419655061"
import matplotlib.pyplot as plt
acc = history.history['accuracy']
loss = history.history['loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'b', label='Training accuracy')
plt.title('Training accuracy')
plt.figure()
plt.plot(epochs, loss, 'b', label='Training Loss')
plt.title('Training loss')
plt.legend()
plt.show()
# + id="Yp3Abx-zoein" executionInfo={"status": "ok", "timestamp": 1622887668654, "user_tz": -120, "elapsed": 175, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}}
next_words = 50
def generate_txt(model, seed_text):
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])[0]
token_list = pad_sequences([token_list], maxlen=max_sequence_len-1, padding='pre')
predicted = model_saved.predict_classes(token_list, verbose=0)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
if len(seed_text) % 10 == 0 :
seed_text+= '\n'
print(seed_text)
# + id="aaQz-D28o_-b"
model_saved = keras.models.load_model(model_file_name)
# + colab={"base_uri": "https://localhost:8080/"} id="6WwDUb2hpAzV" executionInfo={"status": "ok", "timestamp": 1622887674198, "user_tz": -120, "elapsed": 2032, "user": {"displayName": "<NAME>\u00e9<NAME>", "photoUrl": "", "userId": "00757604528325915378"}} outputId="776d243e-8e06-4cef-e285-a35f3f7fc4f4"
seed_text = "Somewhere in La Mancha,"
generate_txt(model_saved, seed_text)
| code/shakespeare-text-generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sklearn
print('The scikit-learn version is {}.'.format(sklearn.__version__))
# <div class="alert alert-block alert-info">
# Version should be greater than or equal to 0.19.1
# </div>
# +
#External Libs
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.metrics import classification_report
#External Algos
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
#Custom Libs
from process import getDataset
#For display
from IPython.display import HTML, display,Image
import tabulate
# -
# # Fetch Dataset
#Fetch Dataset
hr = getDataset('HR_comma_sep.csv')
# hr = pd.DataFrame(hr['data'],columns = hr['headers']).convert_objects(convert_numeric=True)
hr = pd.DataFrame(hr['data'],columns = hr['headers']).infer_objects()
print("DataSet:")
hr
# # PreProcessing
# +
#Encode Salary
salaryEncoder = LabelEncoder()
encodedSalary = salaryEncoder.fit_transform(hr['salary'])
#Representation in Encoder
table=[]
for i in list(salaryEncoder.classes_):
table.append([i,salaryEncoder.transform([i])])
display(HTML(tabulate.tabulate(table, headers=['Class', 'Label'],tablefmt='html')))
# -
#Create datasets
# X contains
# satisfaction_level
#,last_evaluation
#,number_project
#,average_montly_hours
#,time_spend_company
#,Work_accident
#,promotion_last_5years
#,salary
x = hr.drop(columns = ['left','sales'])#.infer_objects()
#Y contains Left
y = pd.to_numeric(hr['left'].values)
#Replace salary with it's encoded values
x['salary'] = encodedSalary
print("Encoded Inputs: ")
x
# Feature Scaling
sc = StandardScaler()
x = sc.fit_transform(x)
print("Feature Scaled Inputs: ")
x
# # CrossValidation
# Splitting the dataset into the Training set and Test set
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size = 0.25, random_state = 0)
# # Training
# ### DecisionTree Classifier with generic HyperParameters
# +
# Fitting Decision Tree Classifier to the Training set
classifier = DecisionTreeClassifier(criterion = 'entropy', random_state = 0, max_depth=3, min_samples_leaf=5)
classifier.fit(x_train, y_train)
# Predicting the Test set results
y_pred = classifier.predict(x_test)
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
acc = accuracy_score(y_test, y_pred)
acc
# +
## plot the importances ##
importances = classifier.feature_importances_
feat_names = hr.drop(['left','sales'],axis=1).columns
indices = np.argsort(importances)[::-1]
plt.figure(figsize=(12,6))
plt.title("Feature importances by DecisionTreeClassifier")
plt.bar(range(len(indices)), importances[indices], color='lightblue', align="center")
plt.step(range(len(indices)), np.cumsum(importances[indices]), where='mid', label='Cumulative')
plt.xticks(range(len(indices)), feat_names[indices], rotation='vertical',fontsize=14)
plt.xlim([-1, len(indices)])
plt.show()
# -
# ## GridSearch For most efficient D-Tree classifier
parameters = {
"criterion": ["gini","entropy"],
"random_state":[0],
"max_depth":list(range(3,11)),
"min_samples_leaf":list(range(1,11))
}
grid_search = GridSearchCV(DecisionTreeClassifier(), parameters)
grid_search.fit(x_train, y_train)
grid_search.best_params_
# ### Best Classifier Accuracy score
import pprint
pp = pprint.PrettyPrinter(indent=4,compact=True,width=10)
y_true, y_pred = y_test, grid_search.predict(x_test)
print("Classifier best hyper parameters")
pp.pprint(grid_search.best_params_)
print("Accuracy of the DecisionTree classifier = %s%%" % (accuracy_score(y_true, y_pred)*100))
# grid_search.grid_scores_ #Mean & STD dev for every combination
# +
## plot the importances ##
importances = grid_search.best_estimator_.feature_importances_
feat_names = hr.drop(['left','sales'],axis=1).columns
indices = np.argsort(importances)[::-1]
plt.figure(figsize=(12,6))
plt.title("Feature importances by DecisionTreeClassifier")
plt.bar(range(len(indices)), importances[indices], color='lightblue', align="center")
plt.step(range(len(indices)), np.cumsum(importances[indices]), where='mid', label='Cumulative')
plt.xticks(range(len(indices)), feat_names[indices], rotation='vertical',fontsize=14)
plt.xlim([-1, len(indices)])
plt.show()
# -
# # Visualize Best Decision Tree Classifier
# <div class="alert alert-block alert-info">
# Should have GraphViz Installed on the Jupyter Server
# </div>
# +
#Convert tree to image Dot format
from sklearn import tree
tree.export_graphviz(grid_search.best_estimator_, out_file='tree.dot', feature_names=hr.drop(columns = ['left','sales']).columns)
#Convert dot file to PNG
from subprocess import call
call(['dot', '-T', 'png', 'tree.dot', '-o', 'tree.png'])
Image(filename='tree.png')
# +
# benalexkeen.com/decision-tree-classifier-in-python-using-scikit-learn/
# http://scikit-learn.org/stable/modules/generated/sklearn.tree.DecisionTreeClassifier.html
| DecisionTree_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import requests
from summpy.lexrank import summarize
r = requests.get("https://medium.com/@arkbb3/%E7%8C%AB%E3%82%AB%E3%83%95%E3%82%A7%E3%81%AF%E3%81%A9%E3%81%93%E3%81%B8%E6%B6%88%E3%81%88%E3%81%9F-9486c89a432e")
soup = BeautifulSoup(r.content, 'html.parser')
for t in soup.findAll(class_='section-inner sectionLayout--insetColumn'):
text = t.get_text()
text_new = ''.join(text.splitlines())
#print(text_new)
sentences, debug_info = summarize(text_new, sent_limit=5, continuous=True, debug=True)
for sent in sentences:
print(sent.strip())
| medium_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
print("Hello World!")
# ## This is text shown in the notebook with markdown
#
#
# Python as a calculator:
2 + 3 * 5
# This is a comment, no markdown here!
x = 1
# x is a variable. It can change its value by assignment.
print(x)
x = x + 1
x
# You can have as many variables as you want...
y = 7
z = x * y
# Output of last statement is shown in console, if there is no assignment
z = z
# +
z
y
# -
# Print() can be used to show contents of variables. Works always, but messy. Better use logging (later in class?).
# +
print(z)
print(y)
# -
# # If statement
#
# An if statement assesses a boolean statement (TRUE or FALSE). If statement is TRUE, first command block is executed.
# IF statement is FALSE, second command block is executed.
if z > 3:
print("z is larger than 3")
else:
print("z is smaller than or equal to 3")
# # Correct indentation in Python is syntax, i.e. obligatory
#
if z > 3:
print("z is larger than 3")
# # Exercise 1
# Create a variable named `infected_03_17` and `infected_03_21`.
# Assign the values `1332` and `2785`, respectively.
#
# Assign `4` to the variable `diff_days`.
#
# Calculate the average daily discrete growth rate given by $\left(\frac{\textrm{infected_03_21}}{\textrm{infected_03_17}}\right)^{\frac{1}{\textrm{diff_days}}} - 1$ and assign it to the variable `growth_rate`. Note: the power function $x^2$ is written as `x**2` in Python code.
#
# Display the `growth_rate`.
# # Functions
# A function is used to put code into one container that can be reused. You will write some functions, but you will use even more functions.
#
# Most functions you are going to use are part of packages, i.e. containers of functions (more on that later!). For now you have to know that if you want to use a function, you first have to import the package. The package has to be available on your computer, so sometimes it has to be installed with conda first. Many important packages come in the conda standard installation already.
#
# Calculate the square root of a number:
# +
import math
math.sqrt(7)
# -
# Of course, you can also pass variables to functions:
# +
x = 7
math.sqrt(x)
# -
# For the moment, we do not write our own functions, but only use them. This will also be important for your home work, where you use functions to encrypt a text.
# # Exercise 2
#
# Calculate the logarithm of `infected_03_17` and `infected_03_21` with the Python library `math`. Use Google to find the correct function name in python. Or guess ;-)
#
# Display the result.
# # Exercise 3
#
#
#
# The code below assigns a random value in the range $[0, 50]$ to $x$. Add code to the cell that prints "Above 25" if the value of $x$ is above $25$ and "Below 25" if the value of $x$ is below $25$. Use an if statement for that purpose.
# +
import random
x = random.randrange(0, 50)
# -
# # Exercise 4
# The following code should calculate the doubling time of an exponential process from the growth_rate calculated before. However, something is wrong. Find the error!
# +
doubling_time = log(2) / log(1 + growth_rate)
doubling_time
# -
# # Writing your own functions
#
# Somewhere up here, we have calculated the growth rate of a process for two timesteps. It would be handy to reuse this code without having to retype the formula each time. This can be accompolished with your own user-defined function.
def growth_rate(value_t1, value_t2, time_diff):
result = (value_t2 / value_t1)**(1 / time_diff) - 1
return result
growth_rate
growth_rate(100, 200, 1)
# +
t1 = 100
t2 = 200
diff = 1
growth_rate(t1, t2, diff)
# +
value_t1 = 100
value_t2 = 200
time_diff = 1
growth_rate(value_t1, value_t2, time_diff)
# -
# # Exercise 5
#
# Write a function to determine the doubling time of an exponential process, given the growth rate.
# # Exercise 6
#
# This function should calculate the relative growth rate of a process given two discrete variables. The formula is given by $\textrm{relative_growth} = \frac{\textrm{value_t2} - \textrm{value_t1}}{\textrm{value_t2}}$. However, the function does not work. Find the problem!
def relative_growth(value_t1, value_t2):
result = (x_t2 - x_t1) / x_t1
return result
relative_growth(2, 1)
| lecture02-conda-python/my-first-notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Homework 2
# 😏
# **Welcome to homework 2!**
#
# We're continuing our journey of analyzing insurance costs data.
# This time, you have a checker cell for each task. It will tell you whether you did the task correctly or not.
# Good luck!
# **Task 1**
# We have two lists, names and insurance costs.
#
# The list `name`s stores the names of ten individuals, and `insurance_costs` stores their medical insurance costs.
#
# Let’s add additional data to these lists:
#
# Append a new individual, "Priscilla", to names.
# Append her insurance cost, 8320.0, to `insurance_costs`.
# +
names = ["Mohammed", "Sara", "Xia", "Paul", "Valentina", "Jide", "Aaron", "Emily", "Nikita", "Paul"]
insurance_costs = [13262.0, 48166.0, 6839.0, 5054.0, 14724.0, 5360.0, 7640.0, 6072.0, 2750.0, 12064.0]
# Add your code here
# -
def list_checker():
assert names[-1]=="Priscilla", "Did you add the name 'Priscilla' correctly?"
assert insurance_costs[-1]==8320.0, "Did you add the insurance cost correctly?"
print('Success! 🎉')
list_checker()
# **Task 2**
# In the previous homework, we manually calculated the estimated insurance cost for each individual. let's create a function which does the same.
# **The general formula is like this :**
# $$
# Insurance Cost=250\times age−128\times sex+370\times bmi+425\times Num Of Children+24000\times smoker−12500
# $$
#
# age: age of the individual in years
# ***
# sex: 0 for female, 1 for male
# ***
# bmi: individual’s body mass index
# ***
# num_of_children: number of children the individual has
# ***
# smoker: 0 for a non-smoker, 1 for a smoker
# - The function takes in 5 inputs : (age, sex, bmi, NumOfChildren, smoker).
# - The function returns a *float* which is the estimated insurance cost of that person.
def estimated_insurance_cost(age, sex, bmi, NumOfChildren, smoker):
pass
#enter your code here
def checking_function():
assert estimated_insurance_cost(20, 1,30, 1, 0)==3897, "hmm your function doesn't seem alright"
print('Success! 🎉')
checking_function()
# **Task 3**
# We're gonna create another function!
# With `estimated_insurance_cost()` function, we cannot input a list! we need to manually input each parameter.
# - Create a new function called `estimated_insurance()` which takes in **lst** as an input.
# - The function needs to use list indecies in **lst** and store each element in their respective name. (for example `age = lst[0]`)
# - The function should return the estimated insurance cost using the parameters and the formula given above
def estimated_insurance(lst):
pass
#enter your code here
test = [87, 0, 29, 0, 1]
def estimation_checker():
assert estimated_insurance(test)==43980, "hmm did you store the list indicies correctly?"
print('Success! 🎉')
estimation_checker()
# **Task 4**
# Using your `estimated_insurance()` function, loop through each person in the following list and calculate their `estimated insurance cost`, and store it in a new list called `estimatations`.
# 💡*hint* : use a **list comprehension**
people = [[39, 1, 28, 0, 0], [87, 0, 29, 0, 1], [67, 1, 35, 0, 0]]
#insert your code here
def checker():
assert estimations[0]==7482, "Your first calculated estimation doesn't seem right!"
assert estimations[1]==43980,"Your second calculated estimation doesn't seem right!"
assert estimations[2]==17072, "Your third calculated estimation doesn't seem right!"
print('Success! 🎉')
checker()
# **Task 5**
# Create a function which takes in a list of `insurance_costs` , and ouputs a new list in which only the insurance costs **below 5000** are included.
def insurance_range_checker(lst):
new_list = [cost for cost in insrance_costs if cost<5000]
#enter your code here
# +
test = [13262.0, 48166.0, 6839.0, 5054.0, 14724.0, 5360.0, 7640.0, 6072.0, 2750.0, 12064.0]
def insurance_range():
assert insurance_range_checker(test)==[2750.0], "did you place your conditionals correctly?"
print('Success! 🎉')
insurance_range()
# -
| homeworks/week-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 14. CNN with CIFAR10
# +
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import DataLoader
import torchvision.utils
import torchvision.datasets as dsets
import torchvision.transforms as transforms
import numpy as np
import os
# -
import matplotlib.pyplot as plt
# %matplotlib inline
# ## 14.1 Prepare Data
# +
train_data = dsets.CIFAR10(root='./data',
train=True,
download=True,
transform=transforms.ToTensor())
test_data = dsets.CIFAR10(root='./data',
train=False,
download=True,
transform=transforms.ToTensor())
# +
batch_size = 128
train_loader = DataLoader(train_data,
batch_size=batch_size,
shuffle=True)
test_loader = DataLoader(test_data,
batch_size=5,
shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck')
# -
def imshow(img, title):
img = torchvision.utils.make_grid(img, normalize=True)
npimg = img.numpy()
fig = plt.figure(figsize = (5, 15))
plt.imshow(np.transpose(npimg,(1,2,0)))
plt.title(title)
plt.axis('off')
plt.show()
# +
train_iter = iter(train_loader)
images, labels = train_iter.next()
imshow(images, "Train Image")
# -
images.shape
# ## 14.2 Define Model
# +
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.conv_layer = nn.Sequential(
nn.Conv2d(3, 32, 5),
nn.ReLU(),
nn.MaxPool2d(2,2),
nn.Conv2d(32, 64, 5),
nn.ReLU(),
nn.MaxPool2d(2,2)
)
self.fc_layer = nn.Sequential(
nn.Linear(64*5*5, 100),
nn.ReLU(),
nn.Linear(100, 10)
)
def forward(self, x):
out = self.conv_layer(x)
out = out.view(-1, 64*5*5)
out = self.fc_layer(out)
return out
model = CNN().cuda()
# -
loss = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=0.01)
# ## 14.3 Train Model
num_epochs = 10
for epoch in range(num_epochs):
total_batch = len(train_data) // batch_size
for i, (batch_images, batch_labels) in enumerate(train_loader):
X = batch_images.cuda()
Y = batch_labels.cuda()
pre = model(X)
cost = loss(pre, Y)
optimizer.zero_grad()
cost.backward()
optimizer.step()
if (i+1) % 200 == 0:
print('Epoch [%d/%d], lter [%d/%d], Loss: %.4f'
%(epoch+1, num_epochs, i+1, total_batch, cost.item()))
# ## 14.4 Test Model
# +
correct = 0
total = 0
for images, labels in test_loader:
images = images.cuda()
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels.cuda()).sum()
print('Accuracy of test images: %f %%' % (100 * float(correct) / total))
# +
images, labels = iter(test_loader).next()
outputs = model(images.cuda())
_, predicted = torch.max(outputs.data, 1)
print('Predicted: ', ' '.join('%5s' % classes[predicted[j]] for j in range(5)))
title = (' '.join('%5s' % classes[labels[j]] for j in range(5)))
imshow(images, title)
| Week4/14. CNN with CIFAR10.ipynb |
-- ---
-- jupyter:
-- jupytext:
-- text_representation:
-- extension: .hs
-- format_name: light
-- format_version: '1.5'
-- jupytext_version: 1.14.4
-- kernelspec:
-- display_name: Haskell
-- language: haskell
-- name: haskell
-- ---
-- # Programming R from Haskell
-- This notebook demonstrates integrating both R and Haskell in the same notebook, using `IHaskell-inline-r`.
-- ## Prelude
-- First, a bit of setup. We need to enable quasiquotation, to embed R code into Haskell expressions:
:ext QuasiQuotes
-- Next, we need to make sure that R is initialized properly:
import qualified H.Prelude as H
H.initialize H.defaultConfig
-- ## R computations
-- By default, computations in code cells are interpreted as Haskell code. For instance, here is a definition of the factorial function, in Haskell:
fact 0 = 1
fact n = n * fact (n - 1)
-- Here is a Haskell expression calling `fact`, together with its value:
fact 10
-- `inline-r` allows you to embed R expressions and R statements anywhere in Haskell code, using quasiquotation. The following is an IO action that asks R to print the value of the R code snipped embedded between the brackets:
[rprint| 1 + 1 |]
-- You can define the factorial function using R code, just as you can using Haskell code, so long as the R code is delineated within a quasiquote:
[r| fact <<- function(n) if(n == 0) 1 else n * fact(n - 1) |]
-- The `r` quasiquote is used for embedding R code that is only useful for its side effects. This is the case with the code above which has the side effect of binding `fact` in the toplevel environment. Applying the definition:
[rprint| fact(10) |]
-- ## R graphics
-- R has extremely powerful plotting facilities. They are available out-of-the-box:
[rgraph| plot(cars) |]
-- For effect, we can fit a straight line through our data set:
[rgraph| plot(cars); abline(lm(cars$dist ~ cars$speed), col="red") |]
-- R code snippets that have graphical output should be embedded using the `rgraph` quasiquote. The other quasiquotes ignore graphical output. For a more complex example, consider the following density plot (requires `ggplot2` to be installed):
[r| require("ggplot2") |]
[rgraph|
Xv <- c(rnorm (500, 10,3), rnorm (500, 50, 20), rnorm (500, 70, 20))
Yv <- c(rnorm (500, 10,3), rnorm (500, 70, 5), rnorm (500, 30, 5))
myd <- data.frame(Xv, Yv)
ggplot(myd, aes(x = Xv, y = Yv)) + geom_point() + geom_density2d() + theme_bw() |]
| IHaskell/examples/tutorial-ihaskell-inline-r.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### scrapy
#
# - xpath 기본문법
# - xpath 실습
# - 2가지 방법으로 가능 : scrapy shell(ipython), jupyter notebook
# - 네이버 키워드 데이터 수집, 다음 키워드 데이터 수집, gmarket 베스트 상품 데이터 수집
# - scrapy project
# - scrapy 파일 디렉토리 구조 및 각 파일에 대한 설명
# - crawler 라는 프로젝트
# - naver 현재 상영 영화 링크
# - 각각의 영화 링크에서 영화제목, 관객수를 크롤링
# - csv 파일로 저장
# - pipeline을 이용해서 csv 파일로 저장
#
import scrapy
# #### xpath 문법
# - 네이버 영화 제목 xpath: `//*[@id="content"]/div[1]/div[1]/div[3]/ul/li[1]/dl/dt/a`
#
# - `//*` : 가장 상위에서 하위를 모두 검색한다는 의미
#
# - `//` : 가장 상위 엘리먼트
# - `*` : css selector 하위 엘리먼트
# - `[@key=value]` : @가 attribute 속성값을 의미
# - `[@id="content]` : 속성 id가 content인 것
# - `/` : css selector 에서 `>`와 같다. 1단계 아래 하위 엘리먼트를 검색
# - `div` : 엘리먼트 이름
# - `[number]` : number 번째의 엘리먼트를 성택 (0부터 시작이 아니라 1부터 시작한다.)
# - `not` : `not(조건)`, 조건에는 @class="test" 이런 것들이 들어갈 수 있는데 class="test"가 아닌 엘리먼트를 선택한다.
# - `.` : 현재 엘리먼트를 의미
# #### 1. scrapy shell 이용
# - `$ scrapy shell "<url>"`
# - `In [1]: response.xpath(' ')`
#
# - 실습 내용
# - 네이버 실시간 검색어
# - 다음 실시간 검색어
# - gmarket 베스트 아이템
# +
# scrapy shell 실행
# cmd창에 : $ scrapy shell "https://naver.com"
# ipython으로 shell에 뜬다.
# 네이버 실시간 검색어 1위 xpath copy : //*[@id="PM_ID_ct"]/div[1]/div[2]/div[2]/div[1]/div/ul/li[1]/a/span[2]
# +
# 아래처럼 scrapy shell 이 실행된다.
# In [1] : response.xpath('//*[@id="PM_ID_ct"]/div[1]/div[2]/div[2]/div[1]/div/ul/li[1]/a/span')
# -
# #### 2. jupyter notebook에서 scrapy 사용
import requests
from scrapy.http import TextResponse
req = requests.get("http://naver.com")
response = TextResponse(req.url, body=req.text, encoding="utf-8") # response.
# 네이버 실시간 검색어 1위 객체 선택
response.xpath('//*[@id="PM_ID_ct"]/div[1]/div[2]/div[2]/div[1]/div/ul/li[1]/a/span[2]')
# 네이버 실시간 검색어 20개 객체 선택 li[1]에서 li로 변경하면 모두 다 나온다. > 문자열만 출력 /text()
response.xpath('//*[@id="PM_ID_ct"]/div[1]/div[2]/div[2]/div[1]/div/ul/li/a/span[2]/text()')
# 실시간 검색어 20개 객체에서 문자열만 출력
response.xpath('//*[@id="PM_ID_ct"]/div[1]/div[2]/div[2]/div[1]/div/ul/li/a/span[2]/text()').extract()
# - - -
# 다음 실시간 검색어 10개 출력하기
req = requests.get("http://daum.net")
response = TextResponse(req.url, body=req.text, encoding="utf-8") # response.
response.xpath('//*[@id="mArticle"]/div[2]/div[2]/div[2]/div[1]/ol/li/div/div[1]/span[2]/a/text()')
response.xpath('//*[@id="mArticle"]/div[2]/div[2]/div[2]/div[1]/ol/li/div/div[1]/span[2]/a/text()').extract()
# gmarket 베스트 아이템 셀렉트, not 사용하기, 링크 선택하기
req = requests.get("http://corners.gmarket.co.kr/Bestsellers")
response = TextResponse(req.url, body=req.text, encoding="utf-8") # response
# 아이디로 나올 경우 상위의 xpath 작성 한 후 아래에는 직접 마무리해서 작성하면 된다.
titles = response.xpath('//*//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li/a/text()').extract()
titles[:5]
# li 엘리먼트에서 class가 first인 데이터만 가지고 오기 [@ 사용해서 셀렉트 가능하다]
titles = response.xpath('//*//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li[@class="first"]/a/text()').extract()
len(titles), titles[:5]
# li 엘리먼트에서 class가 first인 데이터만 가지고 오기 [@ 사용해서 셀렉트 가능하다]
titles = response.xpath('//*//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li[@class="first"]/a/text()').extract()
len(titles), titles[:5]
# li 엘리먼트에서 class가 first인 데이터만 빼고 가지고 오기
titles = response.xpath('//*//*[@id="gBestWrap"]/div/div[3]/div[2]/ul/li[not(@class="first")]/a/text()').extract()
len(titles), titles[:5]
# ##### scrapy project
# - 프로젝트 생성
# - 프로젝트 만들 디렉토리로 이동 후 , `$ scrapy startproject <프로젝트명>`
# - 프로젝트 파일 설명
#
# ```
# ├── crawler
# │ ├── __init__.py
# │ ├── __pycache__
# │ ├── items.py
# │ ├── middlewares.py
# │ ├── pipelines.py
# │ ├── settings.py
# │ └── spiders
# │ ├── __init__.py
# │ └── __pycache__
# └── scrapy.cfg
# ```
#
# - crawler dir : 프로젝트 디렉토리
# - spiders dir (여러개 있을 수 있다.): 우리가 만들 크롤링실행할 클래스와 함수가 모여 있는 디렉토리, URL, links 들로 직접 들어가서 아이템 형태로 object를 만든다.
# - items.py : 크롤링을 할 때 가져오는 데이터를 정의하는 클래스(MVC - Model)(관객수, 영화 이름 등) 클래스 안에 새로운 데이터를 정의하는 것을 객체로 저장 해 둔다.
# - pipelines.py : 여러개의 링크에서 데이터를 가져올 때 실행하는 함수가 정의 되어 있는 클래스, item을 save csv 기능을 갖는다.
# - settings.py : scraping을 할때 정책과 같은 설정을 할 수 있는 파일 입니다. (예를 들어 robots.txt 정책을 따를 것 인지 안따를 것인지를 결정 할 수 있습니다.)
#
# - middlewares.py : 하드웨어 설정시 다루는 파일이고 여기서는 안다룬다.
#
#
#
# spider내의 링크 하나에서 데이터 수집할 것 만든다. item.py 클래스 내 오브젝트가 형성이 된다. 이 오브젝트를 pipeline으로 보내서 save csv를 통해 csv파일로 저장시킨다.
# multithread사용해서 뭐가 먼저 수집 될 지 모른다.
#
# - 네이버 영화에서 현재 상영중인 영화의 제목과 누적 관객수 데이터를 크롤링
req = requests.get("https://movie.naver.com/movie/running/current.nhn")
response = TextResponse(req.url, body=req.text, encoding="utf-8") # response
links = response.xpath('//*[@id="content"]/div[1]/div[1]/div[3]/ul/li/dl/dt/a/@href')[:10].extract()
for link in links:
link = response.urljoin(link) # urljoin : 도메인이 추가된 link 가 완성이 된다.
print(link)
# +
# full url로 만들기
req = requests.get("https://movie.naver.com/movie/bi/mi/basic.nhn?code=159892")
response = TextResponse(req.url, body=req.text, encoding="utf-8") # response
response.xpath('//*[@id="content"]/div[1]/div[2]/div[1]/h3/a[1]/text()').extract()[0]
# -
req = requests.get("https://movie.naver.com/movie/bi/mi/basic.nhn?code=159892")
response = TextResponse(req.url, body=req.text, encoding="utf-8") # response
response.xpath('//*[@id="content"]/div[1]/div[2]/div[1]/dl/dd[5]/div/p[2]/text()').extract()[0]
# +
# 프로젝트 디렉토리로 이동
# scrapy crawl NaverMovie 실행
# ROBOTSTXT_OBEY = False
# +
# yield
# 제너레이터 generator
# 함수 실행 순서에 따라서 return 을 바꿔준다
# -
def numbers():
yield 0
yield 1
yield 2 # yield 사용시 generator가 생성이 된다.
n = numbers()
n.__next__()
n.__next__()
n.__next__()
n.__next__()
# pipepline : item 던져서 csv로 차곡차곡 저장하는 것
import csv
csv.writer(open("NaverMovie.csv","w"))
csvwriter.writerow(["title","count"])
# cat 파일이름.csv
| web/08_scrapy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import pandas as pd
import sys
import matplotlib
print('Python version ' + sys.version)
print('Pandas version ' + pd.__version__)
print('Matplotlib version ' + matplotlib.__version__)
names = ['Bob','Jessica','Mary','John','Mel']
births = [968, 155, 77, 578, 973]
zip(names, births)
BabyDataSet = list (zip(names, births))
df = pd.DataFrame(data = BabyDataSet, columns=['Names', 'Births'])
df
water23 = pd.read_csv("../data/waterlevel/Water23.csv", index_col='date')
water23
ClimateWater = pd.read_csv("../data/waterlevel/ClimateWater.csv", index_col='date')
ClimateWater
# %pylab inline
#water23['upperlevel'].plot()
fig = plt.figure(figsize=(12,4))
ax1 = plt.subplot(111)
water23['downlevel'].plot(ax=ax1)
ax1_1 = ax1.twinx()
water23['upperlevel'].plot(ax=ax1_1)
#climate["Rainfall(mm)"].plot.bar(figsize=(12,5))
climate = pd.read_csv("../data/waterlevel/ClimateWater.csv", index_col='date')
#climate["WaterH1"].hist(bins=100)
water23['upperlevel'].hist(bins=500)
#water23['downlevel'].hist(bins=500)
pd=climate["WaterH1"]
pd
pd.iloc[5]
climate["WaterH1"].plot(figsize=(20,5))
climate["Rainfall(mm)"].plot(figsize=(20,5))
climate["WaterH1"].plot(figsize=(12,5))
water23['upperlevel'].plot(figsize=(12,5))
water23['downlevel'].plot()
#climate["Rainfall(mm)"].plot.bar(figsize=(12,5))
newindex = []
for ind in water23.index:
newindex.append(ind.split()[0])
vals, inds = np.unique(newindex, return_inverse=True)
upperh = np.ones(vals.size)*np.nan
downh = np.ones(vals.size)*np.nan
for i in range (vals.size):
active = inds==i
upperh[i] = water23["upperlevel"].values[active].sum() / active.sum()
downh[i] = water23["downlevel"].values[active].sum() / active.sum()
climate["WaterH1"].plot(figsize=(20,3))
grid(True)
water23['upperlevel'].plot(figsize=(20,3))
grid(True)
water23['downlevel'].plot(figsize=(20,3))
grid(True)
climate.keys()
climate["Moisture(%)"].plot(figsize=(20,3))
climate["SurfaceTemp(\xa1\xc6C)"].plot(figsize=(20,3))
grid(True)
climate["Rainfall(mm)"].plot(figsize=(20,3))
grid(True)
climate["Rainfall(mm)"].plot(figsize=(20,3)).bar
plt.plot(downh)
waterdataset = list (zip(vals, upperh, downh))
#df = pd.DataFrame(data = waterdataset, columns=['vals', 'upperh', 'downh'])
# +
# pd.DataFrame??
# -
df['upperH'].plot()
| notebook/DataStructure-lim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Regression Week 4: Ridge Regression (gradient descent)
# In this notebook, you will implement ridge regression via gradient descent. You will:
# * Convert an SFrame into a Numpy array
# * Write a Numpy function to compute the derivative of the regression weights with respect to a single feature
# * Write gradient descent function to compute the regression weights given an initial weight vector, step size, tolerance, and L2 penalty
# # Fire up graphlab create
# Make sure you have the latest version of GraphLab Create (>= 1.7)
import graphlab
# # Load in house sales data
#
# Dataset is from house sales in King County, the region where the city of Seattle, WA is located.
sales = graphlab.SFrame('kc_house_data.gl/')
# If we want to do any "feature engineering" like creating new features or adjusting existing ones we should do this directly using the SFrames as seen in the first notebook of Week 2. For this notebook, however, we will work with the existing features.
# # Import useful functions from previous notebook
# As in Week 2, we convert the SFrame into a 2D Numpy array. Copy and paste `get_numpy_data()` from the second notebook of Week 2.
import numpy as np # note this allows us to refer to numpy as np instead
def get_numpy_data(data_sframe,features,output):
data_sframe['constant']=1
features=['constant']+features
features_sframe=data_sframe.select_columns(features)
features_matrix = features_sframe.to_numpy()
output_sarray = data_sframe[output]
output_array = output_sarray.to_numpy()
return(features_matrix,output_array)
# Also, copy and paste the `predict_output()` function to compute the predictions for an entire matrix of features given the matrix and the weights:
def predict_output(feature_matrix,weights):
predictions = np.dot(feature_matrix,weights)
return predictions
# # Computing the Derivative
# We are now going to move to computing the derivative of the regression cost function. Recall that the cost function is the sum over the data points of the squared difference between an observed output and a predicted output, plus the L2 penalty term.
# ```
# Cost(w)
# = SUM[ (prediction - output)^2 ]
# + l2_penalty*(w[0]^2 + w[1]^2 + ... + w[k]^2).
# ```
#
# Since the derivative of a sum is the sum of the derivatives, we can take the derivative of the first part (the RSS) as we did in the notebook for the unregularized case in Week 2 and add the derivative of the regularization part. As we saw, the derivative of the RSS with respect to `w[i]` can be written as:
# ```
# 2*SUM[ error*[feature_i] ].
# ```
# The derivative of the regularization term with respect to `w[i]` is:
# ```
# 2*l2_penalty*w[i].
# ```
# Summing both, we get
# ```
# 2*SUM[ error*[feature_i] ] + 2*l2_penalty*w[i].
# ```
# That is, the derivative for the weight for feature i is the sum (over data points) of 2 times the product of the error and the feature itself, plus `2*l2_penalty*w[i]`.
#
# **We will not regularize the constant.** Thus, in the case of the constant, the derivative is just twice the sum of the errors (without the `2*l2_penalty*w[0]` term).
#
# Recall that twice the sum of the product of two vectors is just twice the dot product of the two vectors. Therefore the derivative for the weight for feature_i is just two times the dot product between the values of feature_i and the current errors, plus `2*l2_penalty*w[i]`.
#
# With this in mind complete the following derivative function which computes the derivative of the weight given the value of the feature (over all data points) and the errors (over all data points). To decide when to we are dealing with the constant (so we don't regularize it) we added the extra parameter to the call `feature_is_constant` which you should set to `True` when computing the derivative of the constant and `False` otherwise.
def feature_derivative_ridge(errors, feature, weight, l2_penalty, feature_is_constant):
# If feature_is_constant is True, derivative is twice the dot product of errors and feature
dotP = np.dot(feature,errors)
if feature_is_constant:
derivative = 2*dotP
# Otherwise, derivative is twice the dot product plus 2*l2_penalty*weight
else:
derivative = 2*dotP + 2*l2_penalty*weight
return derivative
# To test your feature derivartive run the following:
# +
(example_features, example_output) = get_numpy_data(sales, ['sqft_living'], 'price')
my_weights = np.array([1., 10.])
test_predictions = predict_output(example_features, my_weights)
errors = test_predictions - example_output # prediction errors
# next two lines should print the same values
print feature_derivative_ridge(errors, example_features[:,1], my_weights[1], 1, False)
print np.sum(errors*example_features[:,1])*2+20.
print ''
# next two lines should print the same values
print feature_derivative_ridge(errors, example_features[:,0], my_weights[0], 1, True)
print np.sum(errors)*2.
# -
# # Gradient Descent
# Now we will write a function that performs a gradient descent. The basic premise is simple. Given a starting point we update the current weights by moving in the negative gradient direction. Recall that the gradient is the direction of *increase* and therefore the negative gradient is the direction of *decrease* and we're trying to *minimize* a cost function.
#
# The amount by which we move in the negative gradient *direction* is called the 'step size'. We stop when we are 'sufficiently close' to the optimum. Unlike in Week 2, this time we will set a **maximum number of iterations** and take gradient steps until we reach this maximum number. If no maximum number is supplied, the maximum should be set 100 by default. (Use default parameter values in Python.)
#
# With this in mind, complete the following gradient descent function below using your derivative function above. For each step in the gradient descent, we update the weight for each feature before computing our stopping criteria.
def ridge_regression_gradient_descent(feature_matrix, output, initial_weights, step_size, l2_penalty, max_iterations=100):
weights = np.array(initial_weights) # make sure it's a numpy array
for j in range(100):
# compute the predictions based on feature_matrix and weights using your predict_output() function
predictions = predict_output(feature_matrix,weights)
# compute the errors as predictions - output
errors = predictions-output
for i in xrange(len(weights)): # loop over each weight
# Recall that feature_matrix[:,i] is the featurte column associaed with weights[i]
# compute the derivative for weight[i].
#(Remember: when i=0, you are computing the derivative of the constant!)
if i==0:
derivative=feature_derivative_ridge(errors,feature_matrix[:,i],weights[i],l2_penalty,True)
else:
derivative=feature_derivative_ridge(errors,feature_matrix[:,i],weights[i],l2_penalty,False)
# subtract the step size times the derivative from the current weight
weights[i]=weights[i]-step_size*derivative
return weights
# # Visualizing effect of L2 penalty
# The L2 penalty gets its name because it causes weights to have small L2 norms than otherwise. Let's see how large weights get penalized. Let us consider a simple model with 1 feature:
simple_features = ['sqft_living']
my_output = 'price'
# Let us split the dataset into training set and test set. Make sure to use `seed=0`:
train_data,test_data = sales.random_split(.8,seed=0)
# In this part, we will only use `'sqft_living'` to predict `'price'`. Use the `get_numpy_data` function to get a Numpy versions of your data with only this feature, for both the `train_data` and the `test_data`.
(simple_feature_matrix, output) = get_numpy_data(train_data, simple_features, my_output)
(simple_test_feature_matrix, test_output) = get_numpy_data(test_data, simple_features, my_output)
# Let's set the parameters for our optimization:
initial_weights = np.array([0., 0.])
step_size = 1e-12
max_iterations=1000
# First, let's consider no regularization. Set the `l2_penalty` to `0.0` and run your ridge regression algorithm to learn the weights of your model. Call your weights:
#
# `simple_weights_0_penalty`
#
# we'll use them later.
simple_weights_0_penalty = ridge_regression_gradient_descent(simple_feature_matrix,output,
initial_weights,step_size,0.0,max_iterations)
print simple_weights_0_penalty
# Next, let's consider high regularization. Set the `l2_penalty` to `1e11` and run your ridge regression algorithm to learn the weights of your model. Call your weights:
#
# `simple_weights_high_penalty`
#
# we'll use them later.
simple_weights_high_penalty = ridge_regression_gradient_descent(simple_feature_matrix,output,
initial_weights,step_size,1e11,max_iterations)
print simple_weights_high_penalty
# This code will plot the two learned models. (The blue line is for the model with no regularization and the red line is for the one with high regularization.)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(simple_feature_matrix,output,'k.',
simple_feature_matrix,predict_output(simple_feature_matrix, simple_weights_0_penalty),'b-',
simple_feature_matrix,predict_output(simple_feature_matrix, simple_weights_high_penalty),'r-')
# Compute the RSS on the TEST data for the following three sets of weights:
# 1. The initial weights (all zeros)
# 2. The weights learned with no regularization
# 3. The weights learned with high regularization
#
# Which weights perform best?
def get_residual_sum_of_squares(feature_matrix, weights,outcome):
predictions = predict_outcome(feature_matrix,weights)
residue = np.array(outcome) - predictions
RSS = residue*residue
return sum(RSS)
print get_residual_sum_of_squares(simple_test_feature_matrix,initial_weights,test_data['price'])
print get_residual_sum_of_squares(simple_test_feature_matrix,simple_weights_0_penalty,test_data['price'])
print get_residual_sum_of_squares(simple_test_feature_matrix,simple_weights_high_penalty,test_data['price'])
# ***QUIZ QUESTIONS***
# 1. What is the value of the coefficient for `sqft_living` that you learned with no regularization, rounded to 1 decimal place? What about the one with high regularization?
# 2. Comparing the lines you fit with the with no regularization versus high regularization, which one is steeper?
# 3. What are the RSS on the test data for each of the set of weights above (initial, no regularization, high regularization)?
#
# # Running a multiple regression with L2 penalty
# Let us now consider a model with 2 features: `['sqft_living', 'sqft_living15']`.
# First, create Numpy versions of your training and test data with these two features.
model_features = ['sqft_living', 'sqft_living15'] # sqft_living15 is the average squarefeet for the nearest 15 neighbors.
my_output = 'price'
(feature_matrix, output) = get_numpy_data(train_data, model_features, my_output)
(test_feature_matrix, test_output) = get_numpy_data(test_data, model_features, my_output)
# We need to re-inialize the weights, since we have one extra parameter. Let us also set the step size and maximum number of iterations.
initial_weights = np.array([0.0,0.0,0.0])
step_size = 1e-12
max_iterations = 1000
# First, let's consider no regularization. Set the `l2_penalty` to `0.0` and run your ridge regression algorithm to learn the weights of your model. Call your weights:
#
# `multiple_weights_0_penalty`
multiple_weights_0_penalty = ridge_regression_gradient_descent(feature_matrix,output,
initial_weights,step_size,0.0,max_iterations)
print multiple_weights_0_penalty
# Next, let's consider high regularization. Set the `l2_penalty` to `1e11` and run your ridge regression algorithm to learn the weights of your model. Call your weights:
#
# `multiple_weights_high_penalty`
multiple_weights_high_penalty = ridge_regression_gradient_descent(feature_matrix,output,
initial_weights,step_size,1e11,max_iterations)
print multiple_weights_high_penalty
# Compute the RSS on the TEST data for the following three sets of weights:
# 1. The initial weights (all zeros)
# 2. The weights learned with no regularization
# 3. The weights learned with high regularization
#
# Which weights perform best?
print get_residual_sum_of_squares(test_feature_matrix,initial_weights,test_data['price'])
print get_residual_sum_of_squares(test_feature_matrix,multiple_weights_0_penalty,test_data['price'])
print get_residual_sum_of_squares(test_feature_matrix,multiple_weights_high_penalty,test_data['price'])
# Predict the house price for the 1st house in the test set using the no regularization and high regularization models. (Remember that python starts indexing from 0.) How far is the prediction from the actual price? Which weights perform best for the 1st house?
prediction1 = predict_output(test_feature_matrix,multiple_weights_0_penalty)
print prediction1
prediction2 = predict_output(test_feature_matrix,multiple_weights_high_penalty)
print prediction2,test_data[0]['price']
# ***QUIZ QUESTIONS***
# 1. What is the value of the coefficient for `sqft_living` that you learned with no regularization, rounded to 1 decimal place? What about the one with high regularization?
# 2. What are the RSS on the test data for each of the set of weights above (initial, no regularization, high regularization)?
# 3. We make prediction for the first house in the test set using two sets of weights (no regularization vs high regularization). Which weights make better prediction <u>for that particular house</u>?
| Machine Learning/Regression/Week 4/week-4-ridge-regression-assignment-2-blank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Will Pueyo's Projection Come True?
#
# On **June 26, 2020**, <NAME> tweeted a warning about the worrysome trend in the growth of COVID-19 confirmed cases in the US over the previous two weeks. Pueyo's tweet includes a projection showing the US daily case growth rate hitting around 50,000 cases/day by 4th of July.
#
# 
#
# Source: https://twitter.com/tomaspueyo/status/1276464644429766672
#
# Pueyo's warning comes with the annotation:
#
# > What will happen over the next <br>
# > 3 weeks if the growth rate from <br>
# > the last week continues <br>
# > (it probably won't, it usually <br>
# > slows down a bit. But it gives us <br>
# > a sense of orders of magnitude) <br>
#
# This Jupyter notebook can be used to check the evolution of new cases in the US using the Johns Hopkins CSSE COVID-19 database availble at github and see if Pueyo's projection comes true. The notbook requires Python3, matplotlib, and pandas, but is otherwise free of package dependencies that can make using other-people's-Python-code painful.
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import matplotlib.ticker as mticker
import pandas as pd
# A utility function to plot 7-day rolling averages of the COVID-19 time-series data (after some minor tinkering)...
def plotRollingAvg(df, ax, names, title, logscale=False):
for name in names:
ax.plot(pd.to_datetime(df.loc[name].index),
df.loc[name].diff().rolling(window=7).mean(),
linewidth=2,
label=name)
_, ax2 = ax.get_xlim()
ax.set_xlim(ax2-7*17, ax2)
ax.xaxis.set_minor_locator(mdates.DayLocator(interval=1))
ax.xaxis.set_major_locator(mdates.DayLocator(interval=7))
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
plt.setp(ax.get_xticklabels(), rotation=30, horizontalalignment='right')
if logscale:
ax.set_yscale('log')
ax.yaxis.set_major_formatter(mticker.StrMethodFormatter('{x:,.0f}'))
ax.yaxis.set_minor_locator(plt.NullLocator())
ax.legend(loc='best', prop={'size': 12})
if title:
ax.title.set_text(title+', 7-Day Rolling Avg')
ax.grid(color='#d4d4d4')
# ### Let's massage the data...
# 1. Load time-series of confirmed cases directly from github repo into pandas dataframe.
# 2. Drop unused columns from the dataframe.
# 3. Group and aggregate (i.e. sum) results.
# +
# Load global time-series so we can compare US vs EU
df_global = pd.read_csv(
('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/'
'csse_covid_19_data/csse_covid_19_time_series/'
'time_series_covid19_confirmed_global.csv'))
df_global = df_global.drop(columns=['Province/State','Lat', 'Long'])
df_global = df_global.groupby('Country/Region').agg('sum')
# Add row for EU totals
eu = ['Austria', 'Belgium', 'Bulgaria', 'Croatia', 'Cyprus', 'Czechia', 'Denmark',
'Estonia', 'Finland', 'France', 'Germany', 'Greece', 'Hungary', 'Ireland',
'Italy', 'Latvia', 'Lithuania', 'Luxembourg', 'Malta', 'Netherlands', 'Poland',
'Portugal', 'Romania', 'Slovakia', 'Slovenia', 'Spain', 'Sweden']
df_global.loc['EU',:] = df_global.loc[eu].sum()
# Load US data so we can look at the four most populous states
df_us = pd.read_csv(
('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/'
'csse_covid_19_data/csse_covid_19_time_series/'
'time_series_covid19_confirmed_US.csv'))
df_us = df_us.drop(columns=[
'UID', 'iso2', 'iso3', 'code3', 'FIPS',
'Admin2', 'Country_Region','Lat', 'Long_'])
df_us = df_us.groupby('Province_State').agg('sum')
# Add row for US total (not needed or used)
df_us.loc['United States',:] = df_us.sum(axis=0)
# -
# ### Here are the raw numbers for the past 7 days...
countries = ['EU', 'US']
df_global.diff(axis=1).loc[countries].iloc[:,-7:]
states = ['California', 'Texas', 'Florida', 'New York']
df_us.diff(axis=1).loc[states].iloc[:,-7:]
# ### And the 7-day rolling average plots...
# 4. Plot overall results for the US and EU.
# 5. **Bonus**: A second plot shows individual state results for the four most populous states in the US.
pd.plotting.register_matplotlib_converters()
plt.style.use('fivethirtyeight')
fig, (ax1, ax2) = plt.subplots(2,1,figsize=(16, 12))
plotRollingAvg(df_global, ax1, countries, 'Daily New Cases')
plotRollingAvg(df_us, ax2, states, '')
fig.autofmt_xdate()
# ### Recommended Reading:
# - [Coronavirus: The Hammer and the Dance](https://medium.com/@tomaspueyo/coronavirus-the-hammer-and-the-dance-be9337092b56), <NAME>, Medium, 19 Mar 2020.
| pueyo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: opentree
# language: python
# name: opentree
# ---
# # How to get an updated tree of all bird families from Open Tree of Life
# In order to run these examples, you need to have installed the `python-opentree` package. Please see instructions at https://github.com/OpenTreeOfLife/python-opentree/blob/ms/INSTALL.md
#
# Currently (Nov 2020) this tutorial requires a newer version of `python-opentree` than is available on PyPi, please follow the instuctions for a local installation.
from opentree import OT, taxonomy_helpers, util
# To avoid typographical errors and confusing synonomies, OpenTree relies on unique identifiers to refer to taxa within OpenTree of Life's Taxonomy (OTT). These "OTT ids" can be found by searching on the website, or the python function:
aves = OT.get_ottid_from_name('Aves')
aves
# You can see more about this taxon at https://tree.opentreeoflife.org/taxonomy/browse?name=81461
# ## Fuzzy matching
# Using get_ottid_from_name requires an exact string match - if nothing is returned for your taxon of interest, you can try approximate matches.
typo = OT.get_ottid_from_name('Avez')
res = OT.tnrs_match(['avez'], do_approximate_matching=True)
res.response_dict
# # Getting a list of taxa in a group by rank
# While the OpenTree Taxonomy (OTT) is not rank focused, it does track rank information from component taxonomies, which we can use to capture families.
#
# The fastest way to access these data is to download the OTT directly.
#
# You can download the OTT by going to https://tree.opentreeoflife.org/about/taxonomy-version/ott3.2
# or by running the following commands.
#
# Set the `loc` argument to wherever you wany to store the taxonomy files.
taxonomy_helpers.download_taxonomy_file(version = 3.3, loc = '../..')
bird_families = taxonomy_helpers.get_ott_ids_group_and_rank(group_ott_id=aves,
rank='family',
taxonomy_file='../../ott3.2/taxonomy.tsv')
# By default this query prunes taxa that are not included in synth
# (usually bc they are extinct and we have no phylogentic input information)
# To get a list of taxa including those excluded from synth, run the same command with synth_only = False
#e.g.
# bird_families = taxonomy_helpers.get_ott_ids_group_and_rank(group_ott_id=aves,
# rank='family',
# synth_only = False,
# taxonomy_file='../../ott3.2/taxonomy.tsv')
len(bird_families)
# # Requesting a tree
# To get the relationships between these taxa we request a labelled induced synth tree
ret = taxonomy_helpers.labelled_induced_synth(ott_ids = bird_families, label_format="name_and_id")
# This return value packages togther a bunch of information in a dictionary. We can see what the keys are:
ret.keys()
# The labelled tree is the central output. It is a dendropy Tree object.
ret['labelled_tree'].print_plot()
ret['labelled_tree'].write(path="labelled_bird_families.tre", schema="newick")
# But lets dig in a bit deeper. Many of these tips are named 'MRCA of taxa in' a family. Those are tips that represent familes that are not-monophyletic, according to our phylogenetic inputs. These are seomtimes called 'broken' taxa. Information about them is returned under the key 'non-monophyletic_taxa'.
len(ret['non-monophyletic_taxa'])
# We can get more info on what is going on with these taxa by looking at that dictionary, e.g. for Sittidiae https://tree.opentreeoflife.org/taxonomy/browse?id=603925
ret['non-monophyletic_taxa']['603925']
# Any taxa that are are 'broken' have at least one study in the corpus that states that the mebers of that taxon are non-monophyletic. We can interactively view what published papers 'broke' Sittidae https://tree.opentreeoflife.org/opentree/argus/ottol@603925
#We can also accessthat information directly in python
resp = OT.synth_subtree('ott603925').response_dict['broken']['contesting_trees'].keys()
cites = OT.get_citations(resp)
print(cites)
# The call also returns a dictionary of any ids that weren't found the the current tree. However, in this case that dictionary is empty, as all search ID's were found.
ret['unknown_ids']
# But we should have the rest of our 194 bird families!
tips = [leaf.taxon.label for leaf in ret['labelled_tree'].leaf_node_iter()]
len(tips)
# # Why are we still missing families?
#
# Some of the non-monophyletic taxa map to internal nodes on our output tree. Input phylogenies are telling us that these 'families' are paraphyletic with respect to other families.
internal_node_fams = []
for tax in ret['non-monophyletic_taxa']:
if ret['non-monophyletic_taxa'][tax]['is_tip'] == False:
internal_node_fams.append(ret['non-monophyletic_taxa'][tax]['name'])
len(internal_node_fams)
print(internal_node_fams)
# We have a much more accurate tree than taxonomy would have geiven us, thanks to 64 published studies informating the relationships!!
len(ret['supporting_studies'])
print(OT.get_citations(ret['supporting_studies']))
| docs/notebooks/bird_families/TreeOfBirdFamilies.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# <h1 align=center><font size = 7>Peer-Graded Assignment: Segmenting and Clustering Neighborhoods in Toronto</font></h1>
# ## Import Necessary Libraries
# !pip install beautifulsoup4
# !pip install lxml
# !pip install html5lib
from bs4 import BeautifulSoup
import lxml
import html5lib
import numpy as np
import pandas as pd
import requests
print('imported')
# ## Download and Explore the Dataset
url = "https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M"
res = requests.get(url)
soup = BeautifulSoup(res.content,"html.parser")
table = soup.find_all('table')[0]
df = pd.read_html(str(table))[0]
# ## Data Cleaning
# Processing unassigned cells and setting the dataframe
df.columns = ['PostalCode','Borough','Neighborhood']
toronto_data = df[df['Borough']!= 'Not assigned']
toronto_data = toronto_data.reset_index(drop=True)
toronto_data = toronto_data.groupby("PostalCode").agg(lambda x:','.join(set(x)))
cond = toronto_data['Neighborhood'] == "Not assigned"
toronto_data.loc[cond, 'Neighborhood'] = toronto_data.loc[cond, 'Borough']
toronto_data.reset_index(inplace=True)
toronto_data.set_index(keys='PostalCode')
toronto_data
# +
url = 'http://cocl.us/Geospatial_data'
df_GeoData = pd.read_csv(url)
df_GeoData.rename(columns={'Postal Code':'PostalCode'},inplace=True)
df_GeoData.set_index(keys='PostalCode')
toronto_GeoData = pd.merge(toronto_data, df_GeoData, on='PostalCode' )
toronto_GeoData.head(15)
# -
# # Part 3 - Explore and cluster the neighborhoods in Toronto
import numpy as np
import pandas as pd
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import json
# !pip install geopy
from geopy.geocoders import Nominatim
import requests
from pandas.io.json import json_normalize
import matplotlib.cm as cm
import matplotlib.colors as colors
from sklearn.cluster import KMeans
# !pip install folium
import folium
print('imported!')
#work with only boroughs that contain the word Toronto
toronto_boroughs= toronto_GeoData[toronto_GeoData['Borough'].str.contains('Toronto', na = False)].reset_index(drop=True)
toronto_boroughs.head()
toronto_boroughs.shape
# The geograpical coordinate of Toronto are 43.6532° N, 79.3832° W
latitude = 43.6532
longitude = -79.3832
# +
# create map of Toronto using latitude and longitude values
map_toronto = folium.Map(location=[latitude, longitude], zoom_start=11)
# add markers to map
for lat, lng, label in zip(toronto_boroughs['Latitude'], toronto_boroughs['Longitude'],
toronto_boroughs['Neighborhood']):
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=5,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_toronto)
map_toronto
# -
# #### Define Foursquare Credentials and Version
CLIENT_ID = 'BEPFM1143I5AMGLPZB3VK0QRYPX1NYB1A3M424XL04RVKLRP' # your Foursquare ID
CLIENT_SECRET = '<KEY>' # your Foursquare Secret
VERSION = '20200523' # Foursquare API version
LIMIT = 100
# +
# A function to explore Toronto neighborhoods
def getNearbyVenues(names, latitudes, longitudes, radius=500):
venues_list=[]
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# make the GET request
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
# -
# #### The code to run the above function on each neighborhood and create a new dataframe called toronto_venues
toronto_venues = getNearbyVenues(names=toronto_boroughs['Neighborhood'],
latitudes=toronto_boroughs['Latitude'],
longitudes=toronto_boroughs['Longitude']
)
# ### Checking the dataframe
print(toronto_venues.shape)
toronto_venues.head()
# #### Let's check how many venues were returned for each neighborhood
toronto_venues.groupby('Neighborhood').count()
# #### Let's find out how many unique categories can be curated from all the returned venues
print('There are {} uniques categories.'.format(len(toronto_venues['Venue Category'].unique())))
# ## Analyze Each Neighborhood
# +
# one hot encoding
toronto_onehot = pd.get_dummies(toronto_venues[['Venue Category']], prefix="", prefix_sep="")
# add neighborhood column back to dataframe
toronto_onehot['Neighborhood'] = toronto_venues['Neighborhood']
# move neighborhood column to the first column
fixed_columns = [toronto_onehot.columns[-1]] + list(toronto_onehot.columns[:-1])
toronto_onehot = toronto_onehot[fixed_columns]
toronto_onehot.head()
# -
# #### let's examine the new dataframe size.
toronto_onehot.shape
toronto_grouped = toronto_onehot.groupby('Neighborhood').mean().reset_index()
toronto_grouped.head()
# #### let's confirm the dataframe size
toronto_grouped.shape
# Let's print each neighborhood along with the top 5 most common venues
# +
num_top_venues = 5
for hood in toronto_grouped['Neighborhood']:
print("----"+hood+"----")
temp = toronto_grouped[toronto_grouped['Neighborhood'] == hood].T.reset_index()
temp.columns = ['venue','freq']
temp = temp.iloc[1:]
temp['freq'] = temp['freq'].astype(float)
temp = temp.round({'freq': 2})
print(temp.sort_values('freq', ascending=False).reset_index(drop=True).head(num_top_venues))
print('\n')
# -
def return_most_common_venues(row, num_top_venues):
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
return row_categories_sorted.index.values[0:num_top_venues]
# +
num_top_venues = 10
indicators = ['st', 'nd', 'rd']
# create columns according to number of top venues
columns = ['Neighborhood']
for ind in np.arange(num_top_venues):
try:
columns.append('{}{} Most Common Venue'.format(ind+1, indicators[ind]))
except:
columns.append('{}th Most Common Venue'.format(ind+1))
# create a new dataframe
neighborhoods_venues_sorted = pd.DataFrame(columns=columns)
neighborhoods_venues_sorted['Neighborhood'] = toronto_grouped['Neighborhood']
for ind in np.arange(toronto_grouped.shape[0]):
neighborhoods_venues_sorted.iloc[ind, 1:] = return_most_common_venues(toronto_grouped.iloc[ind, :], num_top_venues)
neighborhoods_venues_sorted.head()
# +
# set number of clusters
kclusters = 5
toronto_grouped_clustering = toronto_grouped.drop('Neighborhood', 1)
kmeans = KMeans(n_clusters=kclusters, random_state=0).fit(toronto_grouped_clustering)
kmeans.labels_[0:10]
# +
toronto_boroughs_merged = toronto_boroughs
toronto_boroughs_merged['Cluster Labels'] = kmeans.labels_
toronto_boroughs_merged = toronto_boroughs_merged.join(neighborhoods_venues_sorted.set_index('Neighborhood'),on='Neighborhood')
toronto_boroughs_merged.head()
# -
# #### Finally, let's visualize the resulting clusters
# +
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
markers_colors = []
for lat, lon, poi, cluster in zip(toronto_boroughs_merged['Latitude'],
toronto_boroughs_merged['Longitude'],
toronto_boroughs_merged['Neighborhood'],
toronto_boroughs_merged['Cluster Labels']):
label = folium.Popup(str(poi) + ' Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
color=rainbow[cluster-1],
fill=True,
fill_color=rainbow[cluster-1],
fill_opacity=0.7).add_to(map_clusters)
map_clusters
# -
# #### Cluster 1
toronto_boroughs_merged.loc[toronto_boroughs_merged['Cluster Labels'] == 0,
toronto_boroughs_merged.columns[[1] +
list(range(5, toronto_boroughs_merged.shape[1]))]]
# #### Cluster 2
toronto_boroughs_merged.loc[toronto_boroughs_merged['Cluster Labels'] == 1,
toronto_boroughs_merged.columns[[1] +
list(range(5, toronto_boroughs_merged.shape[1]))]]
# #### Cluster 3
toronto_boroughs_merged.loc[toronto_boroughs_merged['Cluster Labels'] == 2,
toronto_boroughs_merged.columns[[1] +
list(range(5, toronto_boroughs_merged.shape[1]))]]
# #### Cluster 4
toronto_boroughs_merged.loc[toronto_boroughs_merged['Cluster Labels'] == 3,
toronto_boroughs_merged.columns[[1] +
list(range(5, toronto_boroughs_merged.shape[1]))]]
# #### Cluster 5
toronto_boroughs_merged.loc[toronto_boroughs_merged['Cluster Labels'] == 4,
toronto_boroughs_merged.columns[[1] +
list(range(5, toronto_boroughs_merged.shape[1]))]]
| Segmenting and Clustering Neighborhoods in Toronto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Making sure I can connect to MySQL and use basic commands
# +
import mysql.connector
file = open('config.txt', 'r')
lines = file.readlines()
host = lines[0].strip()
user = lines[1].strip()
password = lines[2].strip()
mydb = mysql.connector.connect(
host=host,
user=user,
password=password
)
mycursor = mydb.cursor()
mycursor.execute("SHOW DATABASES")
for x in mycursor:
print(x)
# -
# # --------------------------------
# This took me about 10 hours, and the issue was that python was reading in newline characters that I needed to strip. I uninstalled and reinstalled mysql twice, and I spend hours reading the documentation. I can't believe I just didn't check if the raw strings work. Oh well, know for next time.
| SQLTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import AiiDA archive file
#
{% if archive_url %}
# !verdi archive import "{{ archive_url }}"
{% else %}
# Optional
# !verdi archive import "https://path/to/archive.aiida"
{% endif %}
# # Query AiiDA provenance graph
# For more information see [the AiiDA documentation](https://aiida.readthedocs.io/projects/aiida-core/en/latest/howto/data.html)
from aiida import orm, load_profile
load_profile() # load default AiiDA profile
# start querying the database
qb = orm.QueryBuilder()
qb.append(orm.Node)
qb.count()
| aiida/notebooks/explore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from modules.vector_generator import *
from modules.vector_accuracy_checker import *
from modules.vector_distance import *
from modules.file_reader_french import *
from modules.file_reader_synonym import *
import time
from time import gmtime, strftime
accepted_rel = ["syn", "self_loop", "@", "~"] # TO be set: wordnet relation types to be considered
lang = "English"
all_pos = ["n","a","v","r"]
file_names = {"n":"data.noun","v":"data.verb","a":"data.adj","r":"data.adv"}
log_file = 'tmp.log'
log = open(log_file, "w+")
# +
all_data = {}
for pos in all_pos:
all_data.update({pos: data_file_reader(file_names[pos], lang)})
word_set, synset_wrd = word_extractor(all_pos, all_data, False, False, log)
# -
voc_file_path = 'wn_vocabulary.txt'
#file = open(voc_file_path, mode='w')
#file.write('\n'.join([word.split('_offset')[0] for word in word_set]))
#file.close()
file_names = {"wonef": "wonef-fscore-0.1.xml", "wolf": "wolf-1.0b4.xml"}
lang = "French"
all_pos = ["wolf"]
all_data = {}
all_data.update({all_pos[0]: data_file_reader_fr(file_names[all_pos[0]], lang)})
word_set, synset_wrd = word_extractor(all_pos, all_data, False, False, log)
word_set
synset_wrd
voc_file_path = 'wolf_vocabulary.txt'
#file = open(voc_file_path, mode='w')
#file.write('\n'.join([word.split('_offset')[0] for word in word_set]))
#file.close()
def query_synset(word, word_list, synset_wrd):
synset = set()
synset_dis = set()
for data in word_list:
w = data.split('_offset')[0]
key = data.split('\t')[1]+'_wolf'
if w == word:
print(key)
synset.update(synset_wrd[key])
for w in synset:
synset_dis.add(str(beautify(w)))
return synset_dis
def beautify(wn_str):
return wn_str.split('_offset')[0]
# +
query_list = ['livre', 'image', 'maison', 'proie', 'dormir', 'digestion', "chef-d'œuvre"]
for w in query_list:
print(w)
res = query_synset(w, word_set, synset_wrd)
print(query_synset(w, word_set, synset_wrd))
# -
rel_type_dict = {
'near_antonym': '!',
'hypernym': '@',
'instance_hypernym': '@i',
'hyponym': '~',
'instance_hyponym': '~i',
'be_in_state': '#s', # TO VERIFY
'eng_derivative': '+',
'subevent': '*', # TO VERIFY
'also_see': '^',
'verb_group': '$',
'category_domain': ';c',
'derived': '\\',
'similar_to': '&',
'usage_domain': ';u',
'region_domain': ';r',
'holo_part': '#p',
'holo_member': '#m',
'causes': '>',
'holo_portion': '#p', # TO VERIFY
'participle': '<'
}
# +
vocabulary_pos = set()
for sample in word_set:
sample_1s = sample.split('\t')
sample_token = sample_1s[0].split('_offset')[0] + '_' + sample_1s[1].split('-')[-1]
vocabulary_pos.add(sample_token)
# -
len(vocabulary_pos)
# +
#output_file = open('vocabulary_pos_wolf.txt', mode='w+')
#output_file.write('\n'.join(list(vocabulary_pos)));
#output_file.close()
# -
def query_wordlist(word, word_list):
word_set = set()
if word not in word_list:
for i in range(len(word)-1, 0, -1):
for w in word_list:
if w[:i] == word[:i] and w not in word_set:
print(w)
word_set.add(w)
else:
print(word)
# # Get Synset Words for Fine Tuned Vocabulary
test.append((2, 2,3))
test
vocabulary_input_file = open('./data/input/French_testset/tuned-vocabulary.dataset', mode='r')
vocabulary_input = vocabulary_input_file.read().split('\n')
vocabulary_input_file.close()
word_set_tuples = []
for candidate in word_set:
word_synset, synset, wolf = candidate.split('\t')
target_word = word_synset.split('_offset')[0]
target_pos = synset.rsplit('-', 1)[1]
word_set_tuples.append((target_word, target_pos, synset+'_'+wolf))
# +
vocabulary_synset_extend = set()
for word_pos in vocabulary_input:
try:
word, pos = word_pos.rsplit('_', 1)
except:
print(word_pos)
continue
for target_word, target_word, target_key in word_set_tuples:
if target_word == word and pos == target_pos:
for synset_word in synset_wrd[target_key]:
synset_word_word_synset, synset_word_synset, wolf = synset_word.split('\t')
word_to_add = synset_word_word_synset.split('_offset')[0]
pos_to_add = synset_word_synset.rsplit('-', 1)[1]
vocabulary_synset_extend.add(word_to_add+'_'+pos_to_add)
# -
vocabulary_synset_extend
len(vocabulary_synset_extend)
# +
overwrite_check = False
if overwrite_check:
vocabulary_output_file = open('./data/input/French_testset/tuned-vocabulary.dataset', mode='w')
vocabulary_output_file.write('\n'.join(list(vocabulary_synset_extend.union(vocabulary_input))))
vocabulary_output_file.close()
# -
| Voc Extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Video Game Sales Solution
# #### 1. Import pandas module as pd
import pandas as pd
# #### 2. Create variable vgs and read vgsales.csv file as dataframe in it
vgs = pd.read_csv('vgsales.csv')
# #### 3. Get first 10 rows from the dataframe
vgs.head(10)
# #### 4. Use info() method to know the information about number of entries in vgs dataframe
vgs.info()
# #### 5. Get average value of sales in Europe
vgs['EU_Sales'].mean()
# #### 6. Get the highest value of sales in Japan
vgs['JP_Sales'].max()
# #### 7. What is the genre of "Brain Age 2: More Training in Minutes a Day" video game?
vgs[vgs['Name']=='Brain Age 2: More Training in Minutes a Day']['Genre']
# #### 8. What is the number of sales "Grand Theft Auto: Vice City" video game around the world?
vgs[vgs['Name']=='Grand Theft Auto: Vice City']['Global_Sales']
# #### 9. Get the name of the video game which has the highest sales in North America
vgs[vgs['NA_Sales']== vgs['NA_Sales'].max()]['Name']
# или
# vgs.loc[vgs['NA_Sales'].idxmax()]
# #### 10. Get the name of video game which has the smallest sales around the world
vgs[vgs['Global_Sales']== vgs['Global_Sales'].min()]
# #### 11. What is the average value of sales of all video games in Japan?
vgs.groupby('Genre').mean()['JP_Sales']
# #### 12. How many unique names of video games in this dataframe?
vgs['Name'].nunique()
# #### 13. Get the 3 most common genres of video games worldwide
vgs['Genre'].value_counts().head(3)
# #### 14. How many video games have "Super" word in their names?
def super_in_name(name):
if 'super' in name.lower():
return True
else:
return False
sum(vgs['Name'].apply(lambda x: super_in_name(x)))
| notebooks/Python3-DataScience/03-Pandas/10-Video Game Sales Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Serbeld/Practicas-de-Python/blob/master/EvalFunc20.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Thr7AYL2Lc5C" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="fd1737f8-e374-420a-8918-3a5a1cec36ff"
# Metalmecanica QUENZA
# Promedio con inicializacion de numero de dias del mes en 30 dias
def Promedio(Unidades_Producidas_Por_Hora, mes = 30):
# Promedio entero
Promedio = int(Unidades_Producidas_Por_Hora/mes)
return Promedio
# Funcion Eficacia
def Eficacia(Produccion_real,Produccion_estandar):
# Se evalua la operacion matematica de Eficacia
# y se redondean los decimales a dos digitos
#Si la produccion estandar es mayor a 0 efectua la operacion
#Este condicional se usa para evitar divisiones por cero
if (Produccion_estandar > 0):
Eficacia = round((Produccion_real / Produccion_estandar) * 100, 2)
else:
Eficacia = 0
#Estandar de calidad del 85%
if (Eficacia >= 85):
print("La eficacia cumple con los estándares establecidos por parte de la empresa metalmecánica QUENZA")
else:
print("La eficacia no es lo suficientemente buena para cumplir los estándares pedidos por la empresa metalmecánica QUENZA")
return Eficacia
# Funcion Productividad con Piezas producidas referentes a
# una produccion de 121 unidades por hora
def Productividad(Maquinas, Trabajadores, PiezasProducidasACME121 = 121):
# Se evalua la operacion matematica de productividad en numeros enteros
#Este condicional se usa para evitar divisiones por cero
if (Trabajadores > 0):
productividad = int((Maquinas * PiezasProducidasACME121) / Trabajadores)
else:
productividad = 0
return productividad
# Funcion Maquinas_Para_Cumplir_Una_Meta
def Maquinas_Para_Cumplir_Una_Meta(Maquinas, Trabajadores, Meta):
# Encuentra el número de máquinas necesarias para cumplir con una
# meta de productividad establecida por la empresa
productividad = Productividad(Maquinas, Trabajadores)
# Mientras que la productividad sea menor a la meta
while(productividad < Meta):
# Aumenta una maquina
Maquinas = Maquinas + 1
productividad = Productividad(Maquinas, Trabajadores)
return Maquinas
def Menu():
respuesta = 0
# Mientras la opcion seleccionada no sea 5 Salir
while(respuesta != 5):
print("Bienvenido al Menu de inicializacion de la empresa metalmecanica QUENZA")
print() #Espacio
print("Escoge una de las siguientes opciones: ")
print("1 Hallar el promedio de produccion")
print("2 Hallar la eficacia de la maquina ACME121")
print("3 Hallar la productividad de la maquinaria")
print("4 Hallar el numero de maquinas necesarias para cumplir una productividad")
print("5 Salir")
print() #Espacio
# int(input()) pregunta por un valor entero
respuesta = int(input())
print() #Espacio
print("Opcion " + str(respuesta))
# Si se escoge la opcion 1
if respuesta == 1:
print("Ingrese las unidades totales producidas este mes: ")
parametro_Uno = int(input())
#Promedio(UnidadesTotalesProducidas)
promedio = Promedio(parametro_Uno)
print() #Espacio
print("El promedio de unidades producidas este mes fue de: " + str(promedio) + " Unidades por dia")
# Si se escoge la opcion 2
if respuesta == 2:
print("Ingrese las unidades producidas por hora: ")
parametro_Uno = int(input())
print("Ingrese la produccion estandar de las unidades producidas por hora indicadas por el proveedor: ")
parametro_Dos = int(input())
#Eficacia(Produccion_real,Produccion_estandar)
eficacia = Eficacia(parametro_Uno, parametro_Dos)
print() #Espacio
print("La eficacia de la maquina es de un: " + str(eficacia) + "%")
# Si se escoge la opcion 3
if respuesta == 3:
print("Ingrese el numero de maquinas disponibles: ")
parametro_Uno = int(input())
print("Ingrese el numero de trabajadores disponibles para operar las maquinas: ")
parametro_Dos = int(input())
#Productividad(Maquinas, Trabajadores, PiezasProducidasACME121 = 121)
productividad = Productividad(parametro_Uno, parametro_Dos)
print() #Espacio
print("La productividad de la maquina es de: " + str(productividad) + " Unidades por hora")
# Si se escoge la opcion 4
if respuesta == 4:
print("Ingrese el numero de maquinas disponibles: ")
parametro_Uno = int(input())
print("Ingrese el numero de trabajadores disponibles para operar las maquinas: ")
parametro_Dos = int(input())
print("Productividad deseada: ")
parametro_Tres = int(input())
#Maquinas_Para_Cumplir_Una_Meta(Maquinas, Trabajadores, Meta)
maquinas_Para_Cumplir_Una_Meta = Maquinas_Para_Cumplir_Una_Meta(parametro_Uno, parametro_Dos, parametro_Tres)
print() #Espacio
print("Para la productividad deseada se necesitan " + str(maquinas_Para_Cumplir_Una_Meta) + " maquinas para producir " + str(parametro_Tres) + " Unidades por hora")
print() #Espacio
print() #Espacio
print("El codigo ha finalizado!!!")
###############################################################################
## Promedio de unidades producidas del mes de Junio ###########################
UnidadesTotalesProducidasDadasPorElOperario = 1200
#Promedio(UnidadesTotalesProducidas)
promedio = Promedio(UnidadesTotalesProducidasDadasPorElOperario)
print("Promedio de unidades producidas")
print("Las unidades producidas en Junio fueron de " + str(UnidadesTotalesProducidasDadasPorElOperario))
print("El promedio de unidades producidas este mes fue de: " + str(promedio) + " Unidades por dia")
print()#Espacio
###############################################################################
## Eficacia ###################################################################
Produccion_real = 82 #82 unidades por hora
Produccion_estandar = 121 #Produccion estandar de la maquina ACME121
print("Eficacia de la maquina ACME 121")
print("La produccion real de la maquina es de " + str(Produccion_real) + " Unidades por hora")
print("La produccion estandar de la maquina es de " + str(Produccion_estandar) + " Unidades por hora")
#Eficacia(Produccion_real,Produccion_estandar)
eficacia = Eficacia(Produccion_real, Produccion_estandar)
print("La eficacia de la maquina es de un: " + str(eficacia) + "%")
print()#Espacio
###############################################################################
## Productividad ##############################################################
Maquinas = 2
Trabajadores = 2
#Productividad(Maquinas, Trabajadores, PiezasProducidasACME121 = 121)
productividad = Productividad(Maquinas, Trabajadores)
print("Productividad")
print("Se tienen " + str(Maquinas) + " maquinas ACME121")
print("Se tienen " + str(Trabajadores) + " trabajadores en las maquinas ACME121")
print("La productividad de la maquina es de: " + str(productividad) + " Unidades por hora")
print()
###############################################################################
## Maquinas_Para_Cumplir_Una_Meta ##############################################################
Maquinas = 2
Trabajadores = 2
Meta = 400
print("Maquinas necesarias para cumplir una productividad")
print("El numero de maquinas disponibles es de " + str(Maquinas))
print("El numero de trabajadores disponibles para operar las maquinas es de " + str(Trabajadores) + " trabajadores")
print("Productividad deseada es de " + str(Meta) + " Unidades por hora")
#Maquinas_Para_Cumplir_Una_Meta(Maquinas, Trabajadores, Meta)
maquinas_Para_Cumplir_Una_Meta = Maquinas_Para_Cumplir_Una_Meta(Maquinas, Trabajadores, Meta)
print("Para la productividad deseada se necesitan " + str(maquinas_Para_Cumplir_Una_Meta) + " maquinas para producir " + str(Meta) + " Unidades por hora")
print() #Espacio
# + id="11QVrAwSQItA" colab_type="code" colab={}
| Basico-Intermedio/EvalFunc20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# + [markdown] slideshow={"slide_type": "slide"}
# # Unsupervised Learning - Dimensionality Reduction
# + slideshow={"slide_type": "slide"}
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
iris = load_iris()
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, random_state=0)
print(X_train.shape)
# + [markdown] slideshow={"slide_type": "slide"}
# Principal Component Analysis
# ============================
# + [markdown] slideshow={"slide_type": "fragment"}
# PCA is a technique to reduce the dimensionality of the data, by creating a linear projection.
# That is, we find new features to represent the data that are a linear combination of the old data (i.e. we rotate it). Thus, we can think of PCA as a projection of our data onto a *new* feature space.
#
# The way PCA finds these new directions is by looking for the directions of maximum variance.
# Usually only few components that explain most of the variance in the data are kept. Here, the premise is to reduce the size (dimensionality) of a dataset while capturing most of its information. There are many reason why dimensionality reduction can be useful: It can reduce the computational cost when running learning algorithms, decrease the storage space, and may help with the so-called "curse of dimensionality," which we will discuss in greater detail later.
# + [markdown] slideshow={"slide_type": "slide"}
# To illustrate how a rotation might look like, we first show it on two-dimensional data and keep both principal components. Here is an illustration:
# + slideshow={"slide_type": "fragment"}
from figures import plot_pca_illustration
plot_pca_illustration()
# + [markdown] slideshow={"slide_type": "slide"}
# Now let's go through all the steps in more detail:
# We create a Gaussian blob that is rotated:
# + slideshow={"slide_type": "fragment"}
rnd = np.random.RandomState(5)
X_ = rnd.normal(size=(300, 2))
X_blob = np.dot(X_, rnd.normal(size=(2, 2))) + rnd.normal(size=2)
y = X_[:, 0] > 0
plt.scatter(X_blob[:, 0], X_blob[:, 1], c=y, linewidths=0, s=30)
plt.xlabel("feature 1")
plt.ylabel("feature 2");
# + [markdown] slideshow={"slide_type": "slide"}
# As always, we instantiate our PCA model. By default all directions are kept.
# + slideshow={"slide_type": "fragment"}
from sklearn.decomposition import PCA
pca = PCA()
# + [markdown] slideshow={"slide_type": "fragment"}
# Then we fit the PCA model with our data. As PCA is an unsupervised algorithm, there is no output ``y``.
# + slideshow={"slide_type": "fragment"}
pca.fit(X_blob)
# + [markdown] slideshow={"slide_type": "slide"}
# Then we can transform the data, projected on the principal components:
# + slideshow={"slide_type": "fragment"}
X_pca = pca.transform(X_blob)
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y, linewidths=0, s=30)
plt.xlabel("first principal component")
plt.ylabel("second principal component");
# + [markdown] slideshow={"slide_type": "slide"}
# On the left of the plot you can see the four points that were on the top right before. PCA found fit first component to be along the diagonal, and the second to be perpendicular to it. As PCA finds a rotation, the principal components are always at right angles ("orthogonal") to each other.
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-success">
# <b>EXERCISE</b>:
# <ul>
# <li>
# Visualize the iris dataset using the first two principal components, and compare this visualization to using two of the original features.
# </li>
# </ul>
# </div>
# + slideshow={"slide_type": "fragment"}
# # %load sols/01_iris_pca.py
from sklearn.datasets import load_iris
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
iris = load_iris()
X_train, X_test, y_train, y_test = train_test_split(iris.data,
iris.target,
random_state=0,
stratify=iris.target)
pca = PCA(n_components=2)
pca.fit(X_train)
X_train_pca = pca.transform(X_train)
X_test_pca = pca.transform(X_test)
for X, y in zip((X_train_pca, X_test_pca), (y_train, y_test)):
for i, annot in enumerate(zip(('Iris-setosa', 'Iris-versicolor', 'Iris-virginica'),
('blue', 'red', 'green'))):
plt.scatter(X[y==i, 0],
X[y==i, 1],
label=annot[0],
c=annot[1])
plt.xlabel('Principal Component 1')
plt.ylabel('Principal Component 2')
plt.legend(loc='best')
plt.tight_layout()
plt.show()
| notebooks/04 Unsupervised Learning - Dimensionality Reduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings('ignore') # 実行に影響のない warninig を非表示にします. 非推奨.
# # Chapter 5: 機械学習 回帰問題
# ## 5-1. 回帰問題を Pythonで解いてみよう
#
# 1. データセットの用意
# 2. モデル構築
# ### 5-1-1. データセットの用意
# 今回はwine-quality datasetを用いる.
# wine-quality dataset はワインのアルコール濃度や品質などの12要素の数値データ.
# 赤ワインと白ワイン両方あります。赤ワインの含まれるデータ数は1600ほど.
# まずはデータセットをダウンロードする.
# proxy下ではjupyter notebookに設定をしないと以下は動作しない.
# ! wget https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-red.csv ./data/winequality-red.csv
# jupyter notebook の設定が面倒な人へ.
# proxyの設定をしたshell、もしくはブラウザなどで以下のURIからダウンロードしてください.
# https://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/
import pandas as pd
wine = pd.read_csv("./data/winequality-red.csv", sep=";") # sepは区切り文字の指定
display(wine.head(5))
# まずは説明変数1つで回帰を行ってみよう. 今回はalcoholを目的変数 $t$ に, densityを説明変数 $x$ にする.
X = wine[["density"]].values
T = wine["alcohol"].values
# #### 前処理
# データを扱いやすいように中心化する.
X = X - X.mean()
T = T - T.mean()
# trainとtestに分割する.
X_train = X[:1000, :]
T_train = T[:1000]
X_test = X[1000:, :]
T_test = T[1000:]
# +
import matplotlib.pyplot as plt
# %matplotlib inline
fig, axes = plt.subplots(ncols=2, figsize=(12, 4))
axes[0].scatter(X_train, T_train, marker=".")
axes[0].set_title("train")
axes[1].scatter(X_test, T_test, marker=".")
axes[1].set_title("test")
fig.show()
# -
# train と test の分布がかなり違う.
# 予め shuffle して train と test に分割する必要があるようだ.
# XとTの対応関係を崩さず shuffle する方法は多々あるが、その1つが以下.
# +
import numpy as np
np.random.seed(0) # random の挙動を固定
p = np.random.permutation(len(X)) # random な index のリスト
X = X[p]
T = T[p]
# -
X_train = X[:1000, :]
T_train = T[:1000]
X_test = X[1000:, :]
T_test = T[1000:]
# +
fig, axes = plt.subplots(ncols=2, figsize=(12, 4))
axes[0].scatter(X_train, T_train, marker=".")
axes[0].set_title("train")
axes[1].scatter(X_test, T_test, marker=".")
axes[1].set_title("test")
fig.show()
# -
# ### 5-1-2. モデルの構築
# **今回は**, 目的変数 $t$ を以下の回帰関数で予測する.
# $$y=ax+b$$
# この時、損失が最小になるように, パラメータ$a,b$を定める必要がある. ここでは二乗損失関数を用いる.
# $$\mathrm{L}\left(a, b\right)
# =\sum^{N}_{n=1}\left(t_n - y_n\right)^2
# =\sum^{N}_{n=1}\left(t_n - ax_x-b\right)^2$$
#
# <span style="color: gray; ">※これは, 目的変数 $t$ が上記の回帰関数 $y$ を中心としたガウス分布に従うという仮定を置いて最尤推定することと等価.</span>
class MyLinearRegression(object):
def __init__(self):
"""
Initialize a coefficient and an intercept.
"""
self.a =
self.b =
def fit(self, X, y):
"""
X: data, array-like, shape (n_samples, n_features)
y: array, shape (n_samples,)
Estimate a coefficient and an intercept from data.
"""
return self
def predict(self, X):
"""
Calc y from X
"""
return y
# 上記の単回帰のクラスを完成させ, 以下の実行によって図の回帰直線が得られるはずだ.
# +
clf = MyLinearRegression()
clf.fit(X_train, T_train)
# 回帰係数
print("係数: ", clf.a)
# 切片
print("切片: ", clf.b)
fig, axes = plt.subplots(ncols=2, figsize=(12, 4))
axes[0].scatter(X_train, T_train, marker=".")
axes[0].plot(X_train, clf.predict(X_train), color="red")
axes[0].set_title("train")
axes[1].scatter(X_test, T_test, marker=".")
axes[1].plot(X_test, clf.predict(X_test), color="red")
axes[1].set_title("test")
fig.show()
# -
# もしdatasetをshuffleせずに上記の学習を行った時, 得られる回帰直線はどうなるだろう?
# 試してみてください.
# ## 5-2. scikit-learnについて
# ### 5-2-1. モジュールの概要
# [scikit-learn](http://scikit-learn.org/stable/)のホームページに詳しい情報がある.
#
# 実は scikit-learn に線形回帰のモジュールがすでにある.
#
# #### scikit-learn の特徴
# - scikit-learn(sklearn)には,多くの機械学習アルゴリズムが入っており,統一した形式で書かれているため利用しやすい.
# - 各手法をコードで理解するだけでなく,その元となる論文も紹介されている.
# - チュートリアルやどのように利用するのかをまとめたページもあり,似た手法が列挙されている.
import sklearn
print(sklearn.__version__)
# +
from sklearn.linear_model import LinearRegression
clf = LinearRegression()
# 予測モデルを作成
clf.fit(X_train, T_train)
# 回帰係数
print("係数: ", clf.coef_)
# 切片
print("切片: ", clf.intercept_)
# 決定係数
print("決定係数: ", clf.score(X_train, T_train))
fig, axes = plt.subplots(ncols=2, figsize=(12, 4))
axes[0].scatter(X_train, T_train, marker=".")
axes[0].plot(X_train, clf.predict(X_train), color="red")
axes[0].set_title("train")
axes[1].scatter(X_test, T_test, marker=".")
axes[1].plot(X_test, clf.predict(X_test), color="red")
axes[1].set_title("test")
fig.show()
# -
# 自分のコードと同じ結果が出ただろうか?
# また, データを shuffle せず得られた回帰直線のスコアと, shuffleした時の回帰直線のスコアの比較もしてみよう.
# scikit-learn の linear regression のコードは [github][1] で公開されている.
# コーディングの参考になると思うので眺めてみるといいだろう.
#
# ### 5-2-2. 回帰モデルの評価
# 性能を測るといっても,その目的によって指標を変える必要がある.
# どのような問題で,どのような指標を用いることが一般的か?という問いに対しては,先行研究を確認することを勧める.
# また,指標それぞれの特性(数学的な意味)を知っていることもその役に立つだろう.
# [参考][2]
#
# 回帰モデルの評価に用いられる指標は一般にMAE, MSE, 決定係数などが存在する.
#
# 1. MAE
# 2. MSE
# 3. 決定係数
#
# scikit-learn はこれらの計算をするモジュールも用意されている.
#
# [1]:https://github.com/scikit-learn/scikit-learn/blob/1495f69242646d239d89a5713982946b8ffcf9d9/sklearn/linear_model/base.py#L367
# [2]:https://scikit-learn.org/stable/modules/model_evaluation.html
# +
from sklearn import metrics
T_pred = clf.predict(X_test)
print("MAE: ", metrics.mean_absolute_error(T_test, T_pred))
print("MSE: ", metrics.mean_squared_error(T_test, T_pred))
print("決定係数: ", metrics.r2_score(T_test, T_pred))
# -
# ### 5-2-3. scikit-learn の他モデルを使ってみよう
# 1. データセットを用意する
from sklearn import datasets
iris = datasets.load_iris() # ここではIrisデータセットを読み込む
print(iris.data[0], iris.target[0]) # 1番目のサンプルのデータとラベル
# +
# 2.学習用データとテスト用データに分割する
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target)
# 3. 線形SVMという手法を用いて分類する
from sklearn.svm import SVC, LinearSVC
clf = LinearSVC()
clf.fit(X_train, y_train) # 学習
# -
# 4. 分類器の性能を測る
y_pred = clf.predict(X_test) # 予測
print(metrics.classification_report(y_true=y_test, y_pred=y_pred)) # 予測結果の評価
# ### 5-2-4. 分類モデルの評価
#
# 分類問題に対する指標について考えてみよう.一般的な指標だけでも以下の4つがある.
# 1. 正解率(accuracy)
# 2. 精度(precision)
# 3. 再現率(recall)
# 4. F値(F1-score)
#
# (精度,再現率,F値にはmacro, micro, weightedなどがある)
#
# 今回の実験でのそれぞれの値を見てみよう.
print('accuracy: ', metrics.accuracy_score(y_test, y_pred))
print('precision:', metrics.precision_score(y_test, y_pred, average='macro'))
print('recall: ', metrics.recall_score(y_test, y_pred, average='macro'))
print('F1 score: ', metrics.f1_score(y_test, y_pred, average='macro'))
# ## 5-3. 問題に合わせたコーディング
# ### 5-3-1. Irisデータの可視化
# Irisデータは4次元だったので,直接可視化することはできない.
# 4次元のデータをPCAによって圧縮して,2次元にし可視化する.
# +
from sklearn.decomposition import PCA
from sklearn import datasets
iris = datasets.load_iris()
pca = PCA(n_components=2)
X, y = iris.data, iris.target
X_pca = pca.fit_transform(X) # 次元圧縮
print(X_pca.shape)
# -
import matplotlib.pyplot as plt
# %matplotlib inline
plt.scatter(X_pca[:, 0], X_pca[:, 1], c=y);
# 次元圧縮したデータを用いて分類してみる
X_train, X_test, y_train, y_test = train_test_split(X_pca, iris.target)
clf = LinearSVC()
clf.fit(X_train, y_train)
y_pred2 = clf.predict(X_test)
from sklearn import metrics
print(metrics.classification_report(y_true=y_test, y_pred=y_pred2)) # 予測結果の評価
# ### 5-3-2. テキストに対する処理
#
# #### テキストから特徴量を設計
# テキストのカウントベクトルを作成し,TF-IDFを用いて特徴ベクトルを作る.
# いくつかの設計ができるが,例題としてこの手法を用いる.
#
# ここでは,20newsgroupsというデータセットを利用する.
# +
from sklearn.datasets import fetch_20newsgroups
categories = ['alt.atheism', 'soc.religion.christian','comp.graphics', 'sci.med']
news_train = fetch_20newsgroups(subset='train', categories=categories, shuffle=True, random_state=42)
# -
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
count_vec = CountVectorizer()
X_train_counts = count_vec.fit_transform(news_train.data)
tf_transformer = TfidfTransformer(use_idf=False).fit(X_train_counts)
X_train_tf = tf_transformer.transform(X_train_counts)
# #### Naive Bayseによる学習
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB().fit(X_train_tf, news_train.target)
docs = ["God is love.", "I study about Computer Science."]
X_test_counts = count_vec.transform(docs)
X_test_tf = tf_transformer.transform(X_test_counts)
preds = clf.predict(X_test_tf)
for d, label_id in zip(docs, preds):
print("{} -> {}".format(d, news_train.target_names[label_id]))
# このように文に対して,categoriesのうちのどれに対応するかを出力する学習器を作ることができた.
# この技術を応用することで,ある文がポジティブかネガティブか,スパムか否かなど自然言語の文に対する分類問題を解くことができる.
#
# ### 5-3-3. Pipelineによる結合
# +
from sklearn.pipeline import Pipeline
text_clf = Pipeline([('countvec', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB())])
text_clf.fit(news_train.data, news_train.target)
# -
for d, label_id in zip(docs, text_clf.predict(docs)):
print("{} -> {}".format(d, news_train.target_names[label_id]))
# ## 5.4 scikit-learn 準拠コーディング
# scikit-learn 準拠でコーディングするメリットは多数存在する.
# 1. scikit-learn の用意するgrid search や cross validation を使える.
# 2. 既存のscikit-learn の他手法と入れ替えが容易になる.
# 3. 他の人にみてもらいやすい。使ってもらいやすい.
# 4. <span style="color: gray; ">本家のコミッターになれるかも?</span>
#
# 詳しくは [Developer’s Guide][1] に書いてある.
#
# [1]:https://scikit-learn.org/stable/developers/#rolling-your-own-estimator
# scikit-learn ではモデルは以下の4つのタイプに分類されている.
#
# - Classifer
# - Naive Bayes Classifer などの分類モデル
# - Clusterring
# - K-mearns 等のクラスタリングモデル
# - Regressor
# - Lasso, Ridge などの回帰モデル
# - Transformer
# - PCA などの変数の変換モデル
#
# ***準拠コーディングでやるべきことは、***
#
# - sklearn.base.BaseEstimatorを継承する
# - 上記タイプに応じたMixinを多重継承する
#
# (予測モデルの場合)
# - fitメソッドを実装する
# - initでパラメータをいじる操作を入れるとgrid searchが動かなくなる(後述)
# - predictメソッドを実装する
#
# ### 5-4-1. リッジ回帰のscikit-learn 準拠コーディング
#
# 試しに今までにコーディングした MyLinearRegression を改造し, scikit-learn 準拠にコーディングし直してみよう.
# ついでにリッジ回帰の選択ができるようにもしてみよう.
from sklearn.base import BaseEstimator, RegressorMixin
from sklearn.utils.validation import check_X_y, check_is_fitted, check_array
# 回帰なので BaseEstimator と RegressorMixin の継承をする.
# さらにリッジ回帰のオプションも追加するため, initにハイパーパラメータも追加する.
# 入力のshapeやdtypeを整えるために```check_X_y```や```check_array```を用いる(推奨).
class MyLinearRegression(BaseEstimator, RegressorMixin):
def __init__(self, lam = 0):
"""
Initialize a coefficient and an intercept.
"""
self.a =
self.b =
self.lam = lam
def fit(self, X, y):
"""
X: array-like, shape (n_samples, n_features)
y: array, shape (n_samples,)
Estimate a coefficient and an intercept from data.
"""
X, y = check_X_y(X, y, y_numeric=True)
if self.lam != 0:
pass
else:
pass
self.a_ =
self.b_ =
return self
def predict(self, X):
"""
Calc y from X
"""
check_is_fitted(self, "a_", "b_") # 学習済みかチェックする(推奨)
X = check_array(X)
return y
# ***制約***
#
# - initで宣言する変数に全て初期値を定める
# - また引数の変数名とクラス内の変数名は一致させる
# - initにデータは与えない。データの加工なども(必要なら)fit内で行う
# - データから推定された値はアンダースコアをつけて区別する. 今回なら、a_と b_をfit関数内で新しく定義する.
# - アンダースコアで終わる変数をinit内では宣言しないこと.
# - init内で引数の確認, 加工をしてはいけない. 例えば```self.lam=2*lam```などをするとgrid searchができなくなる. [参考][1]
#
# > As model_selection.GridSearchCV uses set_params to apply parameter setting to estimators, it is essential that calling set_params has the same effect as setting parameters using the __init__ method. The easiest and recommended way to accomplish this is to not do any parameter validation in __init__. All logic behind estimator parameters, like translating string arguments into functions, should be done in fit.
#
# [github][2]のコードをお手本にしてみるのもいいだろう.
#
# [1]:https://scikit-learn.org/stable/developers/contributing.html#coding-guidelines
# [2]:https://github.com/scikit-learn/scikit-learn/blob/1495f69242646d239d89a5713982946b8ffcf9d9/sklearn/linear_model/base.py#L367
#
# ### 5-4-2. scikit-learn 準拠かどうか確認
#
# 自作のコードがちゃんとscikit-learn準拠かどうか確かめるには以下を実行する.
from sklearn.utils.estimator_checks import check_estimator
check_estimator(MyLinearRegression)
# 問題があれば指摘してくれるはずだ. なお上記を必ずパスする必要はない.
#
# #### Grid Search
# 準拠モデルを作ったなら, ハイパーパラメータの決定をscikit-learnでやってみよう.
# +
import numpy as np
from sklearn.model_selection import GridSearchCV
np.random.seed(0)
# Grid search
parameters = {'lam':np.exp([i for i in range(-30,1)])}
reg = GridSearchCV(MyLinearRegression(),parameters,cv=5)
reg.fit(X_train,T_train)
best = reg.best_estimator_
# 決定係数
print("決定係数: ", best.score(X_train, T_train)) # BaseEstimatorを継承しているため使える
# lambda
print("lam: ", best.lam)
fig, axes = plt.subplots(ncols=2, figsize=(12, 4))
axes[0].scatter(X_train, T_train, marker=".")
axes[0].plot(X_train, best.predict(X_train), color="red")
axes[0].set_title("train")
axes[1].scatter(X_test, T_test, marker=".")
axes[1].plot(X_test, best.predict(X_test), color="red")
axes[1].set_title("test")
fig.show()
# -
# ## [練習問題](./../exercise/questions.md#chapter-5)
| text/Chapter5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false}
# + active=""
# # to install
# !pip install /Users/johnmount/Documents/work/pyvtreat/pkg/dist/vtreat-0.2.1.tar.gz
# #!pip install https://github.com/WinVector/pyvtreat/raw/master/pkg/dist/vtreat-0.2.1.tar.gz
# + pycharm={"is_executing": false}
import numpy.random
import pandas
import seaborn
import vtreat # https://github.com/WinVector/pyvtreat
numpy.random.seed(235)
zip = ['z' + str(i+1).zfill(5) for i in range(15)]
d = pandas.DataFrame({'zip':numpy.random.choice(zip, size=1000)})
d["const"] = 1
d["const2"]= "b"
d.head()
# + pycharm={"is_executing": false}
transform = vtreat.UnsupervisedTreatment(
params=vtreat.unsupervised_parameters({
'indicator_min_fraction': 0.01,
}))
transform.params_
# + pycharm={"is_executing": false}
d_treated = transform.fit_transform(d)
d_treated.head()
# + pycharm={"is_executing": false}
transform.score_frame_
# + pycharm={"is_executing": false}
| Examples/OtherExamples/IndicatorVariables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **1**. Making wallpaper with `fromfunction`
#
# Adapted from [Circle Squared](http://igpphome.ucsd.edu/~shearer/COMP233/SciAm_Mandel.pdf)
#
# Create a $400 \times 400$ array using the function `lambda i, j: 0.27**2*(i**2 + j**2) % 1.5`. Use `imshow` from `matplotlib.pyplot` with `interpolation='nearest'` and the `YlOrBr` colormap to display the resulting array as an image.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
xs = np.fromfunction(lambda i, j: 0.27**2*(i**2 + j**2) % 1.5,
(400, 400),)
plt.figure(figsize=(8,8))
plt.imshow(xs, interpolation='nearest', cmap=plt.cm.YlOrBr)
plt.xticks([])
plt.yticks([])
pass
# **2**. Find t least squares solution for $\beta_0, \beta_1, \beta_2$ using the normal equations $\hat{\beta} = (X^TX)^{-1}x^Ty$.
#
# \begin{align}
# 10 &= \beta_0 + 3 \beta_1 + 7 \beta_2 \\
# 11 &= \beta_0 + 2 \beta_1 + 8 \beta_2 \\
# 9 &= \beta_0 + 3 \beta_1 + 7 \beta_2 \\
# 10 &= \beta_0 + 1 \beta_1 + 9 \beta_2 \\
# \end{align}
#
# You can find the inverse of a matrix by using `np.linalg.inv` and the transpose with `X.T`
y = np.array([10,11,9,10]).reshape(-1,1)
X = np.c_[np.ones(4), [3,2,3,1], [7,8,8,9]]
X
X.shape, y.shape
# Direct translation of normal equations
β = np.linalg.inv(X.T @ X) @ X.T @ y
β
# More numerically stable version
β = np.linalg.solve(X.T @ X, X.T @ y)
β
# Compare observed with fitted
np.c_[y, X @ β]
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111,projection='3d')
u = np.linspace(0, 4, 2)
v = np.linspace(6, 10, 2)
U, V = np.meshgrid(u, v)
Z = β[0] + U*β[1] +V*β[1]
ax.scatter(X[:,1] , X[:,2] , y, color='red', s=25)
ax.plot_surface(U, V, Z, alpha=0.2)
ax.view_init(elev=30, azim=-60)
| notebook/T03_Exercises_Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1 - Toffoli gate
# +
# Getting rid of unnecessary warnings
import warnings
from matplotlib.cbook import MatplotlibDeprecationWarning
warnings.filterwarnings('ignore', category=MatplotlibDeprecationWarning)
# Importing standard Qiskit libraries
from qiskit import QuantumCircuit, execute, Aer, IBMQ, QuantumRegister, ClassicalRegister
from qiskit.compiler import transpile, assemble
from qiskit.tools.jupyter import *
from qiskit.visualization import *
from ibm_quantum_widgets import *
# Useful to have pi
import math
pi=math.pi
# -
# ## The problem
#
# <div id='problem'></div>
# <div class="alert alert-block alert-success">
#
# We have seen above how to construct a Hadamard gate with our base set, and now we want to build a Toffoli gate as well. Why the Toffoli gate? As mentioned above, the Toffoli gate is also a universal gate for classical computation the same way the NAND gate is, but it is reversible. Further it builds a simple universal gate set for quantum computation if combined with the Hadamard gate.
#
# We have seen some examples on how to express more complex gates using basis gates, we now want to use the knowledge gained to construct a Toffoli gate only using our basis gates. In order to do solve this exercise the above examples on how to construct and use controlled rotations, will come in handy. The biggest challenge is to construct the needed controlled rotations.
#
# You can use the code below using the composer widget to construct your circuit.
#
# </div>
#
#
# <div class="alert alert-block alert-danger">
#
# As a reminder the basis gates for IBM Quantum systems are CX, RZ, SX and X gates, so no other gates are allowed.
#
# Of course we want also try to minimize the cost.
#
# $$
# Cost = 10 N_{CNOT} + N_{other}
# $$
#
# </div>
#
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit
from qiskit import IBMQ, Aer, execute
from ibm_quantum_widgets import CircuitComposer
editorEx = CircuitComposer()
editorEx
##### Build your quantum circuit here using the composer widget.
# +
# This code is being generated automatically by the IBM Quantum Circuit Composer widget.
# It changes in every update of the widget, so any modifications done in this cell will be lost.
# State: synchronized
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit
from numpy import pi
qreg_q = QuantumRegister(3, 'q')
creg_c = ClassicalRegister(3, 'c')
circuit = QuantumCircuit(qreg_q, creg_c)
circuit.rz(pi/2, qreg_q[2])
circuit.sx(qreg_q[2])
circuit.rz(pi/2, qreg_q[2])
circuit.cx(qreg_q[1], qreg_q[2])
circuit.rz(-pi/4, qreg_q[2])
circuit.cx(qreg_q[0], qreg_q[2])
circuit.rz(pi/4, qreg_q[2])
circuit.cx(qreg_q[1], qreg_q[2])
circuit.rz(-pi/4, qreg_q[2])
circuit.cx(qreg_q[0], qreg_q[2])
circuit.rz(pi/4, qreg_q[1])
circuit.rz(pi/4, qreg_q[2])
circuit.cx(qreg_q[0], qreg_q[1])
circuit.rz(pi/2, qreg_q[2])
circuit.rz(pi/4, qreg_q[0])
circuit.rz(-pi/4, qreg_q[1])
circuit.sx(qreg_q[2])
circuit.rz(pi/2, qreg_q[2])
circuit.cx(qreg_q[0], qreg_q[1])
# +
# Checking the resulting circuit
qc = circuit
#qc = circuit # Uncomment this line if you want to submit the circuit built using Qiskit code
qc.draw(output='mpl')
# -
# Check your answer using following code
from qc_grader import grade_ex1
grade_ex1(qc)
# Submit your answer. You can re-submit at any time.
from qc_grader import submit_ex1
submit_ex1(qc)
# ## Additional information
#
# **Created by:** <NAME>, <NAME>, <NAME>
#
# **Version:** 1.0.1
#
# ## Results
#
# **Score:** 73 CNOTs.
#
# **Solved by:** <NAME> (github MaldoAlberto)
| solutions by participants/ex1/ex1-AlbertoMaldonadoRomo-73cost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="0539Pwt1Ygwz" colab_type="text"
# # Project objective
# In this project, we try to identify best alpha in lasso machine learning method for predicting shares in social network popularity of Mashable using information regarding the articles published in a period of two years.
#
# This is a simple case of hyperparameter optimization. The selection is done by identifying best alpha comparing performance of the models in cross-validation setting.
#
# Information about the dataset, some technical details about the used machine learning method(s) and mathematical details of the quantifications approaches are provided in the code.
# + [markdown] id="VjtJFxdsNh05" colab_type="text"
# # Packages we work with in this notebook
# We are going to use the following libraries and packages:
#
# * **numpy**: NumPy is the fundamental package for scientific computing with Python. (http://www.numpy.org/)
# * **sklearn**: Scikit-learn is a machine learning library for Python programming language. (https://scikit-learn.org/stable/)
# * **pandas**: Pandas provides easy-to-use data structures and data analysis tools for Python. (https://pandas.pydata.org/)
#
# We also use **warnings** to stop the notebook from returning warning messages.
#
# + id="57oB2idEgr-g" colab_type="code" colab={}
import numpy as np
import pandas as pd
import sklearn as sk
import warnings
warnings.filterwarnings('ignore')
# + [markdown] id="Bb1Zm7ARN5D5" colab_type="text"
# # Introduction to the dataset
#
# **Name**: Online News Popularity Data Set
#
# **Summary**: "This dataset summarizes a heterogeneous set of features about articles published by Mashable in a period of two years. The goal is to predict the number of shares in social networks (popularity)." (UCI ML)
#
# **number of features**: 58 predictive features
#
# **Number of data points (instances)**: 39797
#
# **Link to the dataset**: https://archive.ics.uci.edu/ml/datasets/Online+News+Popularity
#
#
#
# + [markdown] id="QjBnejgpP0Gr" colab_type="text"
# ## Importing the dataset
# We can import the dataset in multiple ways
#
# **Colab Notebook**: You can download the dataset file (or files) from the link (if provided) and uploading it to your google drive and then you can import the file (or files) as follows:
#
# **Note.** When you run the following cell, it tries to connect the colab with google derive. Follow steps 1 to 5 in this link (https://www.marktechpost.com/2019/06/07/how-to-connect-google-colab-with-google-drive/) to complete the
# + id="RILQWrhjQUtF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="69c44f24-5278-4268-bb91-eb1fe8b850a1"
from google.colab import drive
drive.mount('/content/gdrive')
# This path is common for everybody
# This is the path to your google drive
input_path = '/content/gdrive/My Drive/'
# reading the data (target)
target_dataset = pd.read_csv(input_path + 'OnlineNewsPopularity.csv', index_col=0)
# + [markdown] id="MJgoRANkcTHs" colab_type="text"
# **Local directory**: In case you save the data in your local directory, you need to change "input_path" to the local directory you saved the file (or files) in.
#
# **GitHub**: If you use my GitHub (or your own GitHub) repo, you need to change the "input_path" to where the file (or files) exist in the repo. For example, when I clone ***ml_in_practice*** from my GitHub, I need to change "input_path" to 'data/' as the file (or files) is saved in the data dicretory in this repository.
#
# **Note.**: You can also clone my ***ml_in_practice*** repository (here: https://github.com/alimadani/ml_in_practice) and follow the same process.
# + [markdown] id="UIX-LbyLeEc6" colab_type="text"
# ## Data preparation
# We need to prepare the dataset for machine learnign modeling. Here we prepare the data in 2 steps:
#
# 1) Selecting target columns from the output dataframe (target_dataset_output)
# 2) Converting tissue names to integers (one for each tissue)
# + id="8GI52MUkePCR" colab_type="code" colab={}
# tissueid is the column that contains tissue type information
output_var = target_dataset[' shares']
# we would like to use all the features as input features of the model
input_features = target_dataset.drop([' timedelta', ' shares'], axis=1)
# + [markdown] id="qW4bVMDCdPVW" colab_type="text"
# ## Making sure about the dataset characteristics (number of data points and features)
# + id="TpSupKvgdS3e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="50ba1d76-0b96-4231-d7d6-565a3e0774d3"
print('number of features: {}'.format(input_features.shape[1]))
print('number of data points: {}'.format(input_features.shape[0]))
# + [markdown] id="qgejl_XWhWqN" colab_type="text"
# ## Splitting data to training and testing sets
#
# We need to split the data to train and test, if we do not have a separate dataset for validation and/or testing, to make sure about generalizability of the model we train.
#
# **test_size**: Traditionally, 30%-40% of the dataset cna be used for test set. If you split the data to train, validation and test, you can use 60%, 20% and 20% of teh dataset, respectively.
#
# **Note.**: We need the validation and test sets to be big enough for checking generalizability of our model. At the same time we would like to have as much data as possible in the training set to train a better model.
#
# **random_state** as the name suggests, is used for initializing the internal random number generator, which will decide the splitting of data into train and test indices in your case.
#
# + id="V3L9BbkSg2vp" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(input_features, output_var, test_size=0.30, random_state=5)
# + [markdown] id="d0Fi1jpVkbhC" colab_type="text"
# ## Building the supervised learning model
# We want to build a regression model as the output variable is continuous. Here we build multiple models using Lasso using different hyperparameter values.
#
#
# ### Lasso
# Lasso is a sparse learning algorithm to identify linear relationship between features an the output variable while trying to get rid of irrelevent(not a scientific term) features. The objective function of Lasso is to minimize:
#
# $$min_w {\frac {1}{2n_{sample}}}||Xw-y||_2^2+\alpha||w||_1$$
#
# where $\alpha$ is the model hyperparameter determining the level of sparsification. Larger $\alpha$ values results in fewer non-zero coefficients in the final model. $||w||_1$ is also the first norm ($l_1$) norm of the coefficient vector. The added term ($\alpha ||w||_1$) is a penalty term trying to constrained the coefficient values of the features in the final model.
#
#
# + [markdown] id="7a8ygNgNwCFB" colab_type="text"
# ## Cross-validation and checking generalizability of the model
# After training a machine learning model, we need to check its generalizability and making sure it is not only good in predicting the training set but is capable of predicting new data points. We splitted the data to 2 parts, training and test set. We can go one step further and repeat this splitting across the dataset so that every single data point is considered in one of the test (better to be said validation) sets. This process is called k-fold cross-validation. For example in case of 5-fold cross-validation, the dataset is splitted to 5 chunks and the model is trained in 4 out of 5 chunk and tested on the remianing chunk. The test chunk is then rotated so that every chunk is conisidered once for testing the model. Then we can get average performance of the model in the tested chunks.
#
# Here we use 5-fold cross-validation.
#
# Note. Lack (or low level) of generalizability of a trained model to new data points is called overfitting.
# + [markdown] id="PMM6m-IswFqS" colab_type="text"
# ## Hyperparameter selection
# We have parameters and hyperparameters that need to be determined to build a machine learning model. The parameters are determined in the optimization process in training set (this is hat happens when we train a model). The hyperparameters are those exist for the method (like $\alpha$ in lasso) irrespect of the data. But these hyperparameters can be optimized for the dataset at hand. The hyperparameter optimization is usually done in validation (or development) set. In cross-validation, we are technically assesing performanc of a model at hand on different validation sets we have in cross-validation setting. Hence, the performance in cross-validation setting can be compared to select the best hypeparameters.
#
# + id="fj3SSteMkxb2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 232} outputId="7c1aced4-8fad-4288-c9ee-75a9a2ee10b6"
from sklearn.model_selection import cross_val_score
from sklearn import linear_model
alpha_hyperparam = np.arange(0.1,1.1,0.1)
scores = []
for alpha_iter in alpha_hyperparam:
print('alpha: {}'.format(alpha_iter))
lasso = linear_model.Lasso(alpha=alpha_iter)
scores.append(-round(cross_val_score(lasso, X_train, y_train, cv=5, scoring='neg_mean_squared_error').mean()/(len(y_train)/5), 3))
# Create k nearest neighbour object
# average performance across all folds
print("Average cross-validation performance (mean squared error) in 5-fold cross validation for alpha values of 0.1 to 1 are {}, respectively.".format(scores))
# + id="12wuuVNKAzxH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d067d121-f8e6-429d-d317-f18a32dfe0e0"
print('best alpha value corresponding to the lowest MSE: {}'.format(alpha_hyperparam[np.argmin(scores)]))
# + [markdown] id="Xm8R0XRP8eo-" colab_type="text"
# We identified that $\alpha=1$ results in the best performance in 5-fold cross-validation setting. Now we use all the training data to refit a lasso model with $\alpha=1$ and then assess the performance of the model in the test set.
# + id="Od9xWVWhzp28" colab_type="code" colab={}
# Create k nearest neighbour object
lasso = linear_model.Lasso(alpha=1)
# Train the models using the training sets
lasso.fit(X_train, y_train)
# Make predictions using the testing set
y_pred_lass = lasso.predict(X_test)
# + [markdown] id="C-Uby3AV1OID" colab_type="text"
# ## Evaluating performance of the model
# Finally, we need to assess performance of the model using the predictions of the test set. We use mean squared error to assess the performance of our model. Here are their definitions:
#
# **Mean squared error (MSE)**:
#
# \begin{equation*} MSE = \frac{1}{n}\Sigma_{i=1}^n (Y_i-\hat{Y}_i)^2 \end{equation*}
#
# Note. By setting squared = False, we get the squared root of **MSE**.
#
# + id="kdDOtXow1CKT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5f481180-1306-408b-d445-e8358f5110ab"
from sklearn import metrics
print("normalized mean squared error of the predictions using lasso with alpha=1:", metrics.mean_squared_error(y_test, y_pred_lasso, squared = False))
| code/project15_lasso_hyperparameter_selection_onlinenews.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''procedure_det'': conda)'
# metadata:
# interpreter:
# hash: 4d83f2824dc1cb43aeaedf9feae7e4887829005d749a2c755ba5d31b1bd5b2cf
# name: python3
# ---
# # Binary Classification
#
# Now that we all the features for all the data/lists extracted from libre help and ubuntu help, let us classify them into procedures and non-procedures.
# +
import pandas as pd
import numpy as np
import pickle
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score, cross_val_predict, train_test_split, StratifiedShuffleSplit
from sklearn.preprocessing import normalize
from sklearn.metrics import accuracy_score, precision_score, recall_score, precision_recall_curve, f1_score
DATA_PATH = 'data'
# -
df = pd.read_csv(os.path.join(DATA_PATH, 'dense_features_train_procedures.csv'), encoding='utf-8')
feat = df[['Sents-No Subject', 'Sents-Starts with Verb', 'Avg Length', 'Gerunds', 'Infinitives']].values
labels = df['Labels']
clf = RandomForestClassifier(n_estimators=100)
N = 50
# We will evaluate our model with cross-validation. Since our dataset is class-imbalanced we will use StratifiedKFold. The folds are made by preserving the percentage of samples for each class.
skf = StratifiedShuffleSplit(n_splits=5, random_state=42)
scores = cross_val_score(clf, feat, labels, cv=skf)
scores
test_df = pd.read_csv(os.path.join(DATA_PATH, 'dense_features_test_procedures.csv'), encoding='utf-8')
X_test = df[['Sents-No Subject', 'Sents-Starts with Verb', 'Avg Length', 'Gerunds', 'Infinitives']].values
y_test = test_df['Labels']
clf.fit(feat, labels)
pickle.dump(clf, open('classify_proc.pkl','wb'), protocol=2)
clf = pickle.load(open('classify_proc.pkl', 'rb'))
y_predict = clf.predict(X_test)
print (y_predict)
test_acc = accuracy_score(y_test, y_predict)
print ('Test accuracy:', test_acc, 'F1 Score:', f1_score(y_test, y_predict))
| procedure_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''edna'': conda)'
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("../input/train_folds.csv")
df.head(45)
df[df["kfold"] == 0]
df[df["kfold"] == 1]
| notebooks/stratified.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import torch
import torch.nn.functional as F
from torch.autograd import Variable
# %matplotlib inline
# -
x = torch.Tensor(2, 8)
F.conv2d(x, torch.randn(2))
x = torch.Tensor(1, 8)
x
x = torch.rand(8, 1)
print('x: ', x)
print('x as 8x1: ', x.view(8,1))
print('x as 4x2: ', x.view(4,2))
print('x as 2x4: ', x.view(2,4))
[m, n] = x.size()
print(m)
print(n)
# +
x = torch.rand(8, 1)
x = x.resize_(9, 2)
print(x)
x = Variable(x)
y = Variable(torch.FloatTensor(2, 2))
print(y)
z = torch.cat((x, y), 0)
print(z)
z.data
# -
a = 2
a *= 3
a
1 // 2
# +
t = torch.linspace(0, 63, steps=64)
t = t.view(1, 64)
print('1 x 64: ', t)
t = t.transpose(0, 1).contiguous()
t = t.view(32, 2)
t = t.transpose(0, 1)
print('2 x 32: ', t)
t = t.transpose(0, 1).contiguous()
t = t.view(16, 4)
t = t.transpose(0, 1)
print('4 x 16: ', t)
t = t.transpose(0, 1).contiguous()
t = t.view(8, 8)
t = t.transpose(0, 1).contiguous()
print('8 x 8: ', t)
t = t.transpose(0, 1).contiguous()
t = t.view(1, 64)
print('1 x 64: ', t)
# -
t = t[:, 5:]
print(t)
t = torch.linspace(0, 15, steps=16)
s = t[-1:]
print(s)
s = t[-14:-1:1]
print(s)
s = t[-2:]
print(s)
s = s / 2
print(s)
print(torch.max(s))
| notebooks/tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Project Euler: Problem 6
# + [markdown] nbgrader={}
# https://projecteuler.net/problem=6
#
# The sum of the squares of the first ten natural numbers is,
#
# $$1^2 + 2^2 + ... + 10^2 = 385$$
#
# The square of the sum of the first ten natural numbers is,
#
# $$(1 + 2 + ... + 10)^2 = 552 = 3025$$
#
# Hence the difference between the sum of the squares of the first ten natural numbers and the square of the sum is 3025 − 385 = 2640.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
def sum_of_square(x): #PERSONAL NOTES FOR LATER CODING
n=0 #I may need another value holder(variable)
for i in range(1,101):
#need equations to define problem
n ** 2 + i == x #ignore this cell
print ()
# + deletable=false nbgrader={"checksum": "4a8ce9efca8c824de365eec816018842", "grade": true, "grade_id": "projecteuler6", "points": 10}
# This cell will be used for grading, leave it at the end of the notebook.
# -
def sum_stuff():
return sum(range(1,101))**2 - sum([n**2 for n in range(1,101)]) # just math, first term has sum of all numbers from
print(sum_stuff()) # from 1 to 100 and second term has the sum of the squares
# of numbers from 1 to 100
| assignments/assignment02/ProjectEuler6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ValerieLangat/DS-Unit-1-Sprint-4-Statistical-Tests-and-Experiments/blob/master/Valerie_Langat_(slightly_less_messy)_LS_DS_141_Statistics_Probability_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1WpJHMiiz7kq" colab_type="text"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 1 Sprint 3 Assignment 1*
#
# # Apply the t-test to real data
#
# Your assignment is to determine which issues have "statistically significant" differences between political parties in this [1980s congressional voting data](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records). The data consists of 435 instances (one for each congressperson), a class (democrat or republican), and 16 binary attributes (yes or no for voting for or against certain issues). Be aware - there are missing values!
#
# Your goals:
#
# 1. Load and clean the data (or determine the best method to drop observations when running tests)
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
#
# Note that this data will involve *2 sample* t-tests, because you're comparing averages across two groups (republicans and democrats) rather than a single group against a null hypothesis.
#
# Stretch goals:
#
# 1. Refactor your code into functions so it's easy to rerun with arbitrary variables
# 2. Apply hypothesis testing to your personal project data (for the purposes of this notebook you can type a summary of the hypothesis you formed and tested)
# + id="4cvhjnhcz7ks" colab_type="code" colab={}
import pandas as pd
import numpy as np
import seaborn as sns
# + id="l3N4B2tX0LU1" colab_type="code" colab={}
columns = ['Party', 'Handicapped_Infants', 'WaterProjectCostShare', 'BudgReso', 'PhysFeeFreeze', 'ElSalvAid', 'ReliGroupsinSchools',
'AntiSatTestBan', 'NicaraguanContrasAid',
'MxMissle', 'Immigration', 'SynfuelsCorpCutback', 'EdSpending', 'SuperfundRighttoSue', 'Crime', 'DutyFreeExports', 'ExportAdminActofSA']
data = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data', names=columns, na_values=['?'])
# + id="pUVYE3SG1gaf" colab_type="code" outputId="bf598280-3456-4795-beb7-6de1df9fc353" colab={"base_uri": "https://localhost:8080/", "height": 224}
data.head()
# + id="tE53ICjg1lVA" colab_type="code" colab={}
data.replace({'n':0, 'y':1, np.NaN:.5}, inplace=True)
# + id="CUNKhZF52rpE" colab_type="code" outputId="539d7302-fcc2-48ed-a4c5-c93c41c30044" colab={"base_uri": "https://localhost:8080/", "height": 34}
data.shape
# + id="wHB9rk3H3AfM" colab_type="code" colab={}
grouped = data.groupby('Party', axis='rows')
# + id="T8EejfjS4uWR" colab_type="code" colab={}
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
# + id="EaG2Yanl32lZ" colab_type="code" outputId="6723ed92-9ad1-428d-c680-6084e74eda7f" colab={"base_uri": "https://localhost:8080/", "height": 193}
grouped.describe()
# + id="wm9Qvymc4w5c" colab_type="code" colab={}
from scipy.stats import ttest_ind, ttest_ind_from_stats, ttest_rel
# + id="dSnw3mYp5NGA" colab_type="code" outputId="9d4da6cd-2d49-4e3d-c061-f6340799b434" colab={"base_uri": "https://localhost:8080/", "height": 269}
groupA = np.random.normal(0.112360, 0.400325, 267)
groupB = np.random.normal(1.005952, 0.172929, 168)
sns.kdeplot(groupA, color='b');
sns.kdeplot(groupB, color='r');
# + id="Qgr0cdbW5R52" colab_type="code" outputId="89bd102e-57b6-4b50-fb4e-d106d5a13e7f" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(groupA, groupB)
# + id="yu0U0I2b559B" colab_type="code" outputId="251673c8-35f7-4c29-861c-876657fec6f6" colab={"base_uri": "https://localhost:8080/", "height": 269}
groupA = np.random.normal(0.846442, 0.400677, 267)
groupB = np.random.normal(0.273810, 0.575990, 168)
sns.kdeplot(groupA, color='b');
sns.kdeplot(groupB, color='r');
# + id="YhzrP9qd6Zyb" colab_type="code" outputId="fe05bd02-f5a2-4dda-9af3-5a457edee2e6" colab={"base_uri": "https://localhost:8080/", "height": 34}
ttest_ind(groupA, groupB)
| Valerie_Langat_(slightly_less_messy)_LS_DS_141_Statistics_Probability_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Step 1: Data Preprocessing
#import library
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
#import dataset
dataset = pd.read_csv('dataset.csv')
#divide two column
X = dataset.iloc[ : , : 1 ].values
Y = dataset.iloc[ : , 1 ].values
# +
#train and test two separate model
X_train = X
Y_train = Y
X_test = np.array([12]).reshape(-1, 1)
Y_test = np.array([1]).reshape(-1, 1)
# -
# # Step 2: Fitting Simple Linear Regression Model to the training set
#import Linear Regression Libeary
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor = regressor.fit(X_train, Y_train)
# # Step 3: Predecting the Result
# +
#final result
Y_pred = regressor.predict(X_test)
print("coefficient :- ",regressor.coef_)
print("intercept :- ",regressor.intercept_)
print("Result :- ",Y_pred)
# -
# # Step 4: Visualization
# ## Visualising the Training results
plt.scatter(X_train , Y_train, color = 'blue')
plt.plot(X_train , regressor.predict(X_train), color ='red')
# ## Visualizing the test results
plt.scatter(X_test , Y_test, color = 'blue')
plt.plot(X_test , regressor.predict(X_test), color ='red')
| Semester V/Machine Learning (ML) (4659302)/7/ML_PR_7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Librerie
import math
import os
import random
import re
import sys
# +
#Birthday Cake Candles
def birthdayCakeCandles(candles):
# Write your code here
maxx = max(candles)
count = 0
for el in candles:
if maxx == el:
count += 1
return count
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
candles_count = int(input().strip())
candles = list(map(int, input().rstrip().split()))
result = birthdayCakeCandles(candles)
fptr.write(str(result) + '\n')
fptr.close()
# +
#Number Line Jumps
def kangaroo(x1, v1, x2, v2):
# Write your code here
if (x1 > x2 and v1 > v2) or (x2 > x1 and v2 > v1):
return "NO"
elif (v1-v2)!=0 and abs(x1-x2)%abs(v1-v2)==0:
return "YES"
else:
return "NO"
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
first_multiple_input = input().rstrip().split()
x1 = int(first_multiple_input[0])
v1 = int(first_multiple_input[1])
x2 = int(first_multiple_input[2])
v2 = int(first_multiple_input[3])
result = kangaroo(x1, v1, x2, v2)
fptr.write(result + '\n')
fptr.close()
# +
#Viral Advertising
def viralAdvertising(n):
# Write your code here
if n == 1:
return 2
like = [2]
for i in range(1,n):
like.append(math.floor((like[i-1]*3)/(2)))
return (sum(like))
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
n = int(input().strip())
result = viralAdvertising(n)
fptr.write(str(result) + '\n')
fptr.close()
# +
#Recursive Digit Sum
def superDigit(n, k):
# Write your code here
numbers = list(map(int, [char for char in n]))
somma = [char for char in str(sum(numbers)*k)] #piu efficiente sommare a tratti e moltiplicare
while(len(numbers) > 1):
numbers = list(map(int, [num for num in somma]))
if len(numbers) > 1:
somma = [char for char in str(sum(numbers))]
return numbers[0]
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
first_multiple_input = input().rstrip().split()
n = first_multiple_input[0]
k = int(first_multiple_input[1])
result = superDigit(n, k)
fptr.write(str(result) + '\n')
fptr.close()
# +
#Insertion Sort - Part 1
def insertionSort1(n, arr):
# Write your code here
store = arr[n-1]
i = n-2
while(store < arr[i]) and i>=0: #[0,n-2]
arr[i+1] = arr[i] #shift to right
print(" ".join(map(str,arr)))
i -= 1
arr[i+1] = store
print(" ".join(map(str,arr)))
if __name__ == '__main__':
n = int(input().strip())
arr = list(map(int, input().rstrip().split()))
insertionSort1(n, arr)
# +
#Insertion Sort - Part 2
def insertionSort1(n, arr):
store = arr[n-1]
i = n-2
while(store < arr[i]) and i>=0: #[0,n-2]
arr[i+1] = arr[i] #shift to right
i -= 1
arr[i+1] = store
print(" ".join(map(str,arr)))
def insertionSort2(n, arr):
# Write your code here
for i in range(2,n+1):
insertionSort1(i,arr)
if __name__ == '__main__':
n = int(input().strip())
arr = list(map(int, input().rstrip().split()))
insertionSort2(n, arr)
| Homework 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Previous: <a href = "keras_05.ipynb">1.5 Overfitting VS generalization</a>
# # <center> Keras </center>
# ## <center>1.6 Increasing/Decreasing number of layers</center>
# # Explanation
# There are three types of layers:
# - input layer
# - hidden layer
# - output layer
#
# One hidden layer is sufficient for the large majority of problems.
#
# There is no "magic" rule to calculate the number of hidden layers and nodes of Neural Network, but there are some tips and recommendations.
#
# The number of hidden nodes is based on a relationship between:
#
# - Number of input and output nodes
# - Amount of training data available
# - Complexity of the function that is trying to be learned
# - The training algorithm
# - To minimize the error and have a trained network that generalizes well, you need to pick an optimal number of hidden layers, as well as nodes in each hidden layer.
#
# - Too few nodes will lead to high error for your system as the predictive factors might be too complex for a small number of nodes to capture
#
# - Too many nodes will overfit to your training data and not generalize well
#
# <img src="img/hiddenlayers.png" width="60%" />
# +
#previously done
from keras.models import Sequential
from keras.layers.core import Dense, Dropout
from keras.optimizers import SGD, Adam, Adamax
from keras.utils import np_utils
from keras.utils.vis_utils import model_to_dot
from keras.datasets import mnist
from keras.utils import np_utils
# %matplotlib inline
import math
import random
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import SVG
#Load MNIST
(X_train, y_train), (X_test, y_test) = mnist.load_data()
#Reshape
X_train = X_train.reshape(60000, 784)
X_test = X_test.reshape(10000, 784)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
X_train /= 255
X_test /= 255
Y_train = np_utils.to_categorical(y_train, 10)
Y_test = np_utils.to_categorical(y_test, 10)
#Split
X_train = X_train[0:10000]
X_test = X_test[0:1000]
Y_train = Y_train[0:10000]
Y_test = Y_test[0:1000]
def plot_training_history(history):
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
#loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
# # Example
# +
model = Sequential()
model.add(Dense(input_dim=28*28, units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=500, activation='relu'))
model.add(Dense(units=10, activation='softmax'))
model.compile(loss='mse',
optimizer=SGD(lr=0.1),
metrics=['accuracy'])
BATCH_SIZE=100
NP_EPOCHS=10
history = model.fit(X_train, Y_train,
batch_size=BATCH_SIZE, epochs=NP_EPOCHS,
verbose=1, validation_data=(X_test, Y_test))
plot_training_history(history)
# +
model = Sequential()
model.add(Dense(input_dim=28*28, units=500, activation='relu'))
model.add(Dense(units=10, activation='softmax'))
model.compile(loss='mse',
optimizer=SGD(lr=0.1),
metrics=['accuracy'])
BATCH_SIZE=100
NP_EPOCHS=10
history = model.fit(X_train, Y_train,
batch_size=BATCH_SIZE, epochs=NP_EPOCHS,
verbose=1, validation_data=(X_test, Y_test))
plot_training_history(history)
# -
# # Task
# Name two disadvantages resulting from having to many layers.
# # Feedback
# ### Next: <a href = "keras_07.ipynb">1.7 Batch size and number of epochs</a>
| Keras/.ipynb_checkpoints/keras_06-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center><img src="http://i.imgur.com/sSaOozN.png" width="500"></center>
#
# ## Course: Computational Thinking for Governance Analytics
#
# ### Prof. <NAME>, PhD
# * Visiting Professor of Computational Policy at Evans School of Public Policy and Governance, and eScience Institute Senior Data Science Fellow, University of Washington.
# * Professor of Government and Political Methodology, Pontificia Universidad Católica del Perú.
#
# _____
#
# # Data Preprocessing in Python: Data Integration and Reshaping
# We all know collect data from different places. While the cleaning and formatting is done for each data source, we finally need to integrate all the sources into one to start the real analytical work.
# I will use several data sets in this material. Let me start with the one from CIA on internet users:
# +
import pandas as pd
interlink="https://www.cia.gov/the-world-factbook/field/internet-users/country-comparison"
ciainter=pd.read_html(interlink)[0]
# -
# The object **ciainter** is a data frame. Notice the **[0]** at the end of the **pd.read_html()** function, you should remember that functions returns a **list** of elements. The data frame was the first one (position zero). Let's see some info:
ciainter.info()
# The dataframe **ciainter** has four columns and 228 rows. The column **Dtype** informs the data type of each column in the data frame. Two columns are numbers (integers) and two are text (object). The third column has a weird name, let's see the data:
ciainter.head()
# The unnamed column is the one informing the amount of internet users, then let's rename it:
ciainter.rename(columns={'Unnamed: 2':'intusers'},inplace=True) #notice inplace which changes the data frame.
# I will not use the rank nor the date of information, so I will delete (drop) both:
bye = ['Rank','Date of Information']
ciainter.drop(columns=bye, inplace=True)
# Country is a **key** column. If I were to add more columns from other data frame, I will use this column for merging. I will make sure it has no trailing or leading spaces:
ciainter['Country']=ciainter['Country'].str.strip()
# The amount of users is influenced by the population size. So, it is better to make divide the users by the population. I need the population data, which the CIA also has:
poplink='https://www.cia.gov/the-world-factbook/field/population/country-comparison'
ciapop=pd.read_html(poplink)[0]
# Let's see what we got:
ciapop.info()
ciapop.head()
# These data needs the same as the previous one:
ciapop.rename(columns={'Unnamed: 2':'pob'},inplace=True)
ciapop['Country']=ciapop['Country'].str.strip()
ciapop.drop(columns=bye, inplace=True)
# ## Merging
# Integrating data sets needs the following considerations:
#
# * Merging is done on two data frames (you can prepare a function to merge more).
# * You need a common column to be used in both data frames. The column names can be different.
# * The merge can keep only the full coincidences, or also the values not matched, which will help you detect possible extra cleaning.
# * Pandas differentiates the **left** from the **right** data frames.
# Since I want to divide the number of internet users by the population, I need to **merge** both data frames. Let me show you several possibilities:
#
# * **Option one**: merge only the coincidences:
# +
ciainter.merge(ciapop)
# -
# The previous merge just got rid of any row that could not find the same country name in both data frames.
#
# * **Option two**: merge when the column keys are different:
#let me rename the key column in 'ciapop':
ciapop.rename(columns={'Country':'countries'},inplace=True)
# this will give you an error:
ciainter.merge(ciapop)
# this is the rigth code:
ciainter.merge(ciapop,left_on='Country',right_on='countries')
# You got the same result (with an extra column).
#
# * **Option three**: you want to keep all the rows in the **left** data frame:
ciainter.merge(ciapop,left_on='Country',right_on='countries',how='left')
# * **Option four**: you want to keep all the rows in the **right** data frame:
ciainter.merge(ciapop,left_on='Country',right_on='countries',how='right')
# * **Option five**: you want to keep all the rows from **both** data frames:
ciainter.merge(ciapop,left_on='Country',right_on='countries',how='outer',indicator='True')
# Notice that I included the argument **indicator=True**, which added a column telling if the row comes from both, or from the left or rigth data frame.
#
# ### Looking for improvements after merging
#
# Let me pay attention to this result again:
allRight=ciainter.merge(ciapop,left_on='Country',right_on='countries',how='left', indicator=True)
allRight
# The previous result is different from this one:
ciainter.merge(ciapop,left_on='Country',right_on='countries')
# There is **one** row difference, let me see:
allRight[allRight._merge!='both']
# I have found the only country that is not present in 'ciapop'. Imagine you had **The Antarctica** in *ciapop*, you could replace it like this:
# +
###dictionary of replacements:
#replacementscia={'The Antarctica':'Antarctica'}
### replacing
#ciapop.countries.replace(replacementscia,inplace=True)
# -
# ...and you will need to redo the merge.
#
# Let me keep the **allRight** dataframe, erasing the irrelevant columns and rows:
# dropping columns
byeCols=['countries','_merge']
allRight.drop(columns=byeCols,inplace=True)
# dropping rows
byeRows=[217]
allRight.drop(index=byeRows,inplace=True)
When you erase
# ____
# ____
#
#
# ### <font color="red">Saving File to Disk</font>
# #### For future use in Python:
allRight.to_pickle("allRight.pkl")
# you will need: DF=pd.read_pickle("interhdi.pkl")
# or:
# from urllib.request import urlopen
# DF=pd.read_pickle(urlopen("https://..../interhdi.pkl"),compression=None)
# #### For future use in R:
# +
from rpy2.robjects import pandas2ri
pandas2ri.activate()
from rpy2.robjects.packages import importr
base = importr('base')
base.saveRDS(allRight,file="allRight.RDS")
#In R, you call it with: DF = readRDS("interhdi.RDS")
#or, if iyou read from cloud: DF = readRDS(url("https://..../interhdi.RDS")
# -
# ## RESHAPING
#
# ### Wide and Long format
#
# The current format of **allRight** is known as the **WIDE** format. In this format, the variables are in every column, the most traditional one for spreadsheet users. Several functions are ready to use this format, for example:
# A scatter plot
allRight.plot.scatter(x='intusers', y='pob',grid=True)
# a boxplot
allRight.loc[:,['intusers','pob']].boxplot(vert=False,figsize=(15,5),grid=False)
# However, the wide format may be less efficient for some packages:
# +
# #!pip install plotnine
# +
import plotnine as p9
base=p9.ggplot(data=allRight)
base + p9.geom_boxplot(p9.aes(x=1,y='intusers')) + p9.geom_boxplot(p9.aes(x=2,y='pob'))
# -
# Let's see the **LONG** format:
allRight.melt(id_vars=['Country'])
# The amount of of rows multiplies, but **all** the variables in the wide format will use only **TWO** columns in the wide format (in its basic form). Notice the difference in this code:
allRightLONG=allRight.melt(id_vars=['Country'])
base=p9.ggplot(data=allRightLONG)
base + p9.geom_boxplot(p9.aes(x='variable',y='value'))
# ### Transposing
# We have two data sets on information about race, one for California and one for Washington State. These are the links:
# +
# California link
linkCa='https://github.com/EvansDataScience/data/raw/master/CaliforniaRace.xlsx'
# Washington link
linkWa='https://github.com/EvansDataScience/data/raw/master/WAraceinfo.xlsx'
# -
# You can realize from the links that both data are in Excel format ( _xlsx_ ). Let's fetch them:
raceca=pd.read_excel(linkCa,0) # first sheet
racewa=pd.read_excel(linkWa,1) # second sheet
# Let me see what **racewa** has:
racewa
# The rows give you information on geographical units (the **unit of analysis** is the county). It apparently starts with information of the whole state (Washington), and then county by county. Notice that units of analysis repeat by group age and by year.
# Now, let's see what **raceca** has:
raceca
# Notice that the data from California speaks of the same, but the **units of analysis** (counties) appear in the columns. Notice that while WA State only shows counts, CA State also shows percentages.
#
# The data from WA State is a standard format for data frames, while the one in CA State is not. However, a simple operation known as **transposing** will solve the situation:
raceca.transpose()
# Let's make the changes:
raceca=raceca.transpose()
# The transposed data frame requires several cleaning steps:
#
# * Move first row as column names:
# first row, where the columns names are.
raceca.columns=raceca.iloc[0,:].to_list()
# * Delete first row:
raceca.head()
# dropping first row effective immediatly
raceca.drop(index='Unnamed: 0',inplace=True)
# * Keep the columns about **race**:
# finding positions:
list(enumerate(raceca.columns))
# values needed:
[0]+ list(range(23,31))
# keeping the ones I want:
raceca=raceca.iloc[:,[0]+ list(range(23,31))]
raceca
# * Drop rows with missing values:
raceca.dropna(subset=['Statistics'],inplace=True)
# When we drop rows, we reset indexes:
raceca.reset_index(drop=True,inplace=True)
# currently
raceca
# This is a much simpler data frame.
# ### Aggregating
# The data from WA State has data from different years, while the one from CA is just from 2019. Let's keep that year for WA:
racewa.query('Year==2019',inplace=True)
# Now you have:
racewa
# Notice that the data is organized by age in WA:
racewa['Age Group'].to_list()
# There is a **Total** in **Age Group** that I will not use (that makes this work simpler).
racewa=racewa[racewa['Age Group']!='Total']
racewa
# The ages are organized in intervals, let's keep the consecutive ones:
# +
stay=['0-19', '20-64', '65+']
racewa=racewa[racewa['Age Group'].isin(stay)]
racewa
# -
# * We should keep the values that do not include '__Washington__'
racewa=racewa[racewa["Area Name"]!='Washington']
racewa
# The **aggregation** is used when you need to colapse rows. You can use different function for collapsing, in this case we will **sum** within each county, so I can get a total per county:
racewa=racewa.groupby(['Area Name','Area ID','Year']).sum()
racewa
# The **Age Group** is not used in the aggreting function **groupby()**, but it is the only **non-numeric** columns that is not used in this function. Notice that **Age Group** values have been concatenated, and the grouping variables are the **indexes** (row names).
#
# You can drop the age group now:
racewa.drop(columns=['Age Group'],inplace=True)
racewa
# ### Appending
# The units of analysis in both data frames are the same kind (counties) but they data from one data frame will not be a column for the other. In this situation, you do not **merge**, you **append**.
#
# The condition for appending is that both data frame have the same colum names. Let's see:
raceca.columns
racewa.columns
# The column 'Some Other Race Alone' in **raceca** has no similar value in **racewa**. Let's drop it:
raceca.drop(columns=['Some Other Race Alone'],inplace=True)
# The columns in WA State have values for male and female. Since CA State do not have that, we have to get rid of those:
# good names
[name for name in racewa.columns if 'Total' in name]
# then
racewa=racewa.loc[:,[name for name in racewa.columns if 'Total' in name]]
racewa
# We need to reciver the county names in racewa. They are part of the indexes. Let me use the **reset_index** funtion, but using the argument **drop=False**:
racewa.reset_index(drop=False,inplace=True)
# you have
racewa
# The columns "Area ID" and "Year" are not present in raceca, we should drop them:
racewa.drop(columns=["Area ID", "Year"],inplace=True)
racewa
# Let's see the names of both:
dict(zip(raceca.columns,racewa.columns))
# You can use that dictionary to alter the names in raceca:
#
raceca.rename(columns=dict(zip(raceca.columns,racewa.columns)),inplace=True)
raceca
# The **raceca** has the word "California"; since you are combining data from different states, it is better you keep that info:
# using 'expand'
raceca['Area Name'].str.split(pat=", ",expand=True)
# The fucntion **str.split** creates two columns, let me save them here:
twoCols=raceca['Area Name'].str.split(pat=", ",expand=True)
twoCols
# Use the columns to replace other columns:
raceca['Area Name']=twoCols[0]
raceca['State']=twoCols[1]
# we have
raceca
# Let's drop the las row (the 'TOTAL'):
raceca.drop(index=[41],axis=0,inplace=True)
# You can get rid of the 'County' string:
raceca['Area Name']=raceca['Area Name'].str.replace(" County","")
raceca
# The data frame **racewa** does not have a "State" column, let me create it:
racewa['State']='Washington'
# Let's check the coincidences in the column names:
racewa.columns==raceca.columns
# Now we can append:
racewaca=racewa.append(raceca, ignore_index=True)
racewaca
# Let's check the data types:
#checking
racewaca.info()
# There are several columns that are numeric, but they have the wrong Dtype. Let's solve this:
# this require formatting
racewaca[racewaca.columns[1:-1]]
# +
# let's do it:
racewaca[racewaca.columns[1:-1]]=racewaca[racewaca.columns[1:-1]].astype('float')
# -
# checking
racewaca.info()
| 2021/ComputationalThinking/Part1_PythonForPreProcessing/Integrating_and_Reshaping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PIC16B]
# language: python
# name: conda-env-PIC16B-py
# ---
# ## 1 Create a Database
#
# We are going to create a database with with three datasets to efficiently access information
# To do this, we will need the following imports
import pandas as pd
import numpy as np
import sqlite3
# ### Temperatures table
#
# The temperatures dataset includes the temperatures for each month of a year for thousands of stations. There is separate file for each decade worth of years, so we add that decade to the url. Let's look at the data table for the most recent decade.
interval = "2011-2020"
temps_url = f"https://raw.githubusercontent.com/PhilChodrow/PIC16B/master/datasets/noaa-ghcn/decades/{interval}.csv"
temperatures = pd.read_csv(temps_url)
temperatures.head(3)
# The way this data is set up is not tidy, since there are 12 temperatures values in each observation. To make it tidy, we want to have one row for each month. The temperature values are also in hundreths of a degree, so we want to divide them by degree. Let's make a function to do this.
def prep_temp_df(df):
"""
This function tidys and cleans the data.
It makes each observation be the temperature for a station, year, and month
and neatens the new columns of month and temp
Input: df - a dataframe of temperatures
Outputs a restructured dataframe of the temperatures
"""
#We want to keep ID and Year as columns in the transformed table
df = df.set_index(keys=["ID", "Year"])
#Convert table
df = df.stack()
df = df.reset_index()
#Make the new columns for Month and Temp look nice
df = df.rename(columns = {"level_2" : "Month" , 0 : "Temp"})
df["Month"] = df["Month"].str[5:].astype(int)
df["Temp"] = df["Temp"] / 100
return(df)
# Now we want to create a database.
conn = sqlite3.connect("climate.db")
# Now we want to add the data, one decade file at a time. So for each decade, we will read in data from the url, prepare it, and add it to the temperatures table.
# +
#List with the start of each decade
decades = np.arange(1901, 2021, 10)
for start in decades:
interval = str(start) + "-" + str(start+9)
temps_url = f"https://raw.githubusercontent.com/PhilChodrow/PIC16B/master/datasets/noaa-ghcn/decades/{interval}.csv"
#Create iterator to reach in chunk
df_iter = pd.read_csv(temps_url, chunksize = 100000)
#Iterate through the file, adding chunks
for df in df_iter:
cleaned = prep_temp_df(df)
cleaned.to_sql("temperatures", conn, if_exists = "append", index = False)
# -
# ### Stations table
#
# Now we will get our next table from its url.
stations_url = "https://raw.githubusercontent.com/PhilChodrow/PIC16B/master/datasets/noaa-ghcn/station-metadata.csv"
stations = pd.read_csv(stations_url)
stations.head(3)
# We will want to use this table to match countries. It turns out the first two characters of the ID match the FIPS 10-4 country code, so let's add that to our table
stations["FIPS"] = stations["ID"].str[0:2]
# Now we can add this table directly to the database
stations.to_sql("stations", conn, if_exists = "replace", index = False)
# ### Countries table
#
# Lastly we are going to add our countries table. We can get it from the url below.
countries_url = "https://raw.githubusercontent.com/mysociety/gaze/master/data/fips-10-4-to-iso-country-codes.csv"
countries = pd.read_csv(countries_url)
countries.head(3)
# We are going to run into problems with SQL if the column names have spaces, so let's shorten those. It might also make sense to change Name to Country.
countries = countries.rename(columns = {"FIPS 10-4":"FIPS", "ISO 3166":"ISO", "Name":"Country"})
countries.head(3)
# Then we can add this table to our database
countries.to_sql("countries", conn, if_exists = "replace", index = False)
# Since we are done adding to our database, we need to close the connection.
conn.close()
# ## 2 Write a Query Function
#
# Now we are going to write a function that will create table from the database using SQL
# We want to look at temperatures for a given countries for a given month in a range of years. We want the dataframe to have columns for the station name, latititude, longitude, and country, as well as year, month, and temperature of the reading.
#
# To get this we will make a command. The command will select all of the columns from the abreviated name of table they came from. It will join the temperatures table with station, on id, and countries, on FIPS in order to do this. In the WHERE statement, we will include the conditions.
def query_climate_database(country, year_begin, year_end, month):
"""
This function creates a dataframe of temperatures for stations in a country
for a given month and years from the climate database
Inputs:
country - the country to look in
year_begin - the first year to gather data for
year_end - the last year to gather data for
month - the month of tempterature to look at
Outputs a dataframe according to the specifications
"""
#The command pulls data from all three tables by joining them
cmd = """
SELECT S.name, S.latitude, S.longitude, C.country, T.year, T.month, T.temp
FROM temperatures T
LEFT JOIN stations S ON T.id = S.id
LEFT JOIN countries C ON C.FIPS = S.FIPS
WHERE C.country= \"""" + str(country) + """\"
AND T.year >=""" + str(year_begin) + """
AND T.year <=""" + str(year_end) + """
AND T.month =""" + str(month)
#Open the connection to execute the command
conn = sqlite3.connect("climate.db")
df = pd.read_sql_query(cmd, conn)
conn.close()
return df
# Lets do an example where we look at temperatures in August for stations in Germany from 2009 to 2012.
query_climate_database("Germany", 2009, 2012, 8).head(3)
# ## 3 Write a Geographic Scatter Function for Yearly Temperature Increases
#
# Our next task is to create a function our query that will address the following question:
#
# *How does the average yearly change in temperature vary within a given country?*
# To do this we are going to group the data by station and find the change using linear regression. For this we will need a function to use in the apply.
# +
from sklearn.linear_model import LinearRegression
def coef(data_group):
"""
This function finds the linear regression equation for a data group
Input: data_group - as data group from the groupby function
Output the rounded first coefficient of the equation
"""
x = data_group[["Year"]]
y = data_group["Temp"]
LR = LinearRegression()
LR.fit(x,y)
return round(LR.coef_[0],3)
# -
# We will do this with plotly.
from plotly import express as px
# Make function
def temperature_coefficient_plot(country, year_begin, year_end, month, min_obs, **kwargs):
df = query_climate_database(country, year_begin, year_end, month)
obs = df.groupby(["NAME"])["Month"].transform(np.sum) / month
df = df[obs >= min_obs]
df = df.reset_index()
coefs = df.groupby(["NAME", "LATITUDE", "LONGITUDE"]).apply(coef).reset_index()
coefs["Yearly\nIncrease"] = coefs[0]
title = "Yearly Temperature Increase in Month " + str(month)
title += "for stations in " + country +" "+ str(year_begin)
title += "-" + str(year_end)
fig = px.scatter_mapbox(coefs,
lat = "LATITUDE",
lon = "LONGITUDE",
color = "Yearly\nIncrease",
color_continuous_midpoint = 0,
hover_name = "NAME",
title = title,
mapbox_style = "carto-positron",
**kwargs)
return fig
# +
color_map = px.colors.diverging.RdGy_r # choose a colormap
fig = temperature_coefficient_plot("India", 1980, 2020, 1,
min_obs = 10,
zoom = 2,
#mapbox_style="carto-positron",
color_continuous_scale=color_map)
# -
from plotly.io import write_html
write_html(fig, "plot3.html")
| notebooks/blog_post_1-v3-test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Packed Padded Sequences, Masking and Inference
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torchtext.datasets import TranslationDataset, Multi30k
from torchtext.data import Field, BucketIterator
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import spacy
import random
import math
import time
# +
SEED = 1234
random.seed(SEED)
torch.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# -
spacy_de = spacy.load('de')
spacy_en = spacy.load('en')
# +
def tokenize_de(text):
return [tok.text for tok in spacy_de.tokenizer(text)]
def tokenize_en(text):
return [tok.text for tok in spacy_en.tokenizer(text)]
# +
# packed padded seqeuenceを使うときはpadする前の系列長の長さが必要
# torchtextには系列長を取得するオプションがある
SRC = Field(tokenize=tokenize_de,
init_token='<sos>',
eos_token='<eos>',
lower=True,
include_lengths=True)
TRG = Field(tokenize=tokenize_en,
init_token='<sos>',
eos_token='<eos>',
lower=True)
# -
train_data, valid_data, test_data = Multi30k.splits(exts=('.de', '.en'), fields=(SRC, TRG))
SRC.build_vocab(train_data, min_freq = 2)
TRG.build_vocab(train_data, min_freq = 2)
# +
# packed padded sequenceを使うにはミニバッチの各サンプルが
# 系列長の降順になっている必要がある
# BucketIteratorにはそのオプションがある
BATCH_SIZE = 128
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
train_iterator, valid_iterator, test_iterator = BucketIterator.splits(
(train_data, valid_data, test_data),
batch_size=BATCH_SIZE,
sort_within_batch=True,
sort_key=lambda x: len(x.src),
device=device)
# -
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, enc_hid_dim, dec_hid_dim, dropout):
super().__init__()
self.input_dim = input_dim
self.emb_dim = emb_dim
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.embedding = nn.Embedding(input_dim, emb_dim)
self.rnn = nn.GRU(emb_dim, enc_hid_dim, bidirectional=True)
self.fc = nn.Linear(enc_hid_dim * 2, dec_hid_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, src, src_len):
embedded = self.dropout(self.embedding(src))
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, src_len)
packed_outputs, hidden = self.rnn(packed_embedded)
outputs, _ = nn.utils.rnn.pad_packed_sequence(packed_outputs)
# bidirectionalの時はforwardとbackwardがあるのでfc層を通す
hidden = torch.tanh(self.fc(torch.cat((hidden[-2, :, :], hidden[-1, :, :]), dim=1)))
return outputs, hidden
class Attention(nn.Module):
def __init__(self, enc_hid_dim, dec_hid_dim):
super().__init__()
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.attn = nn.Linear((enc_hid_dim * 2) + dec_hid_dim, dec_hid_dim)
self.v = nn.Parameter(torch.rand(dec_hid_dim))
def forward(self, hidden, encoder_outputs, mask):
batch_size = encoder_outputs.shape[1]
src_len = encoder_outputs.shape[0]
hidden = hidden.unsqueeze(1).repeat(1, src_len, 1)
encoder_outputs = encoder_outputs.permute(1, 0, 2)
energy = torch.tanh(self.attn(torch.cat((hidden, encoder_outputs), dim=2)))
energy = energy.permute(0, 2, 1)
v = self.v.repeat(batch_size, 1).unsqueeze(1)
attention = torch.bmm(v, energy).squeeze(1)
# 入力のpadを無視するようにmaskをかける
attention = attention.masked_fill(mask == 0, -1e10)
return F.softmax(attention, dim=1)
class Decoder(nn.Module):
def __init__(self, output_dim, emb_dim, enc_hid_dim, dec_hid_dim, dropout, attention):
super().__init__()
self.emb_dim = emb_dim
self.enc_hid_dim = enc_hid_dim
self.dec_hid_dim = dec_hid_dim
self.output_dim = output_dim
self.attention = attention
self.embedding = nn.Embedding(output_dim, emb_dim)
self.rnn = nn.GRU((enc_hid_dim * 2) + emb_dim, dec_hid_dim)
self.out = nn.Linear((enc_hid_dim * 2) + dec_hid_dim + emb_dim, output_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, input, hidden, encoder_outputs, mask):
input = input.unsqueeze(0)
embedded = self.dropout(self.embedding(input))
# attention weights
# 1つ前のdecoderのhiddenとencoderの出力から計算
a = self.attention(hidden, encoder_outputs, mask)
a = a.unsqueeze(1)
# attention weightsでencoderの出力を重み付け
encoder_outputs = encoder_outputs.permute(1, 0, 2)
weighted = torch.bmm(a, encoder_outputs)
weighted = weighted.permute(1, 0, 2)
# Decoderの各ステップには重み付けされたencoder出力も入力される
rnn_input = torch.cat((embedded, weighted), dim=2)
output, hidden = self.rnn(rnn_input, hidden.unsqueeze(0))
assert (output == hidden).all()
embedded = embedded.squeeze(0)
output = output.squeeze(0)
weighted = weighted.squeeze(0)
output = self.out(torch.cat((output, weighted, embedded), dim=1))
return output, hidden.squeeze(0), a.squeeze(1)
class Seq2Seq(nn.Module):
def __init__(self, encoder, decoder, pad_idx, sos_idx, eos_idx, device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.pad_idx = pad_idx
self.sos_idx = sos_idx
self.eos_idx = eos_idx
self.device = device
def create_mask(self, src):
mask = (src != self.pad_idx).permute(1, 0)
return mask
def forward(self, src, src_len, trg, teacher_forcing_ratio=0.5):
# 推論時の設定
if trg is None:
assert teacher_forcing_ratio == 0, 'Must be zero during inference'
inference = True
trg = torch.zeros((100, src.shape[1])).long().fill_(self.sos_idx).to(src.device)
else:
inference = False
batch_size = src.shape[1]
max_len = trg.shape[0]
trg_vocab_size = self.decoder.output_dim
# decoder outputsを保存するテンソル
outputs = torch.zeros(max_len, batch_size, trg_vocab_size).to(self.device)
# attentionを保存するテンソル
attentions = torch.zeros(max_len, batch_size, src.shape[0]).to(self.device)
encoder_outputs, hidden = self.encoder(src, src_len)
# decoderへの最初の入力は<sos>
output = trg[0, :]
mask = self.create_mask(src)
for t in range(1, max_len):
output, hidden, attention = self.decoder(output, hidden, encoder_outputs, mask)
outputs[t] = output
attentions[t] = attention
teacher_force = random.random() < teacher_forcing_ratio
top1 = output.max(1)[1]
output = (trg[t] if teacher_force else top1)
if inference and output.item() == self.eos_idx:
return outputs[:t], attentions[:t]
return outputs, attentions
INPUT_DIM = len(SRC.vocab)
OUTPUT_DIM = len(TRG.vocab)
ENC_EMB_DIM = 256
DEC_EMB_DIM = 256
ENC_HID_DIM = 512
DEC_HID_DIM = 512
ENC_DROPOUT = 0.5
DEC_DROPOUT = 0.5
PAD_IDX = SRC.vocab.stoi['<pad>']
SOS_IDX = TRG.vocab.stoi['<sos>']
EOS_IDX = TRG.vocab.stoi['<eos>']
attn = Attention(ENC_HID_DIM, DEC_HID_DIM)
enc = Encoder(INPUT_DIM, ENC_EMB_DIM, ENC_HID_DIM, DEC_HID_DIM, ENC_DROPOUT)
dec = Decoder(OUTPUT_DIM, DEC_EMB_DIM, ENC_HID_DIM, DEC_HID_DIM, DEC_DROPOUT, attn)
model = Seq2Seq(enc, dec, PAD_IDX, SOS_IDX, EOS_IDX, device).to(device)
# +
def init_weights(m):
for name, param in m.named_parameters():
if 'weight' in name:
nn.init.normal_(param.data, mean=0, std=0.01)
else:
nn.init.constant_(param.data, 0)
model.apply(init_weights)
# +
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# -
optimizer = optim.Adam(model.parameters())
criterion = nn.CrossEntropyLoss(ignore_index=PAD_IDX)
def train(model, iterator, optimizer, criterion, clip):
model.train()
epoch_loss = 0
for i, batch in enumerate(iterator):
src, src_len = batch.src
trg = batch.trg
optimizer.zero_grad()
output, attention = model(src, src_len, trg)
output = output[1:].view(-1, output.shape[-1])
trg = trg[1:].view(-1)
loss = criterion(output, trg)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
def evaluate(model, iterator, criterion):
model.eval()
epoch_loss = 0
with torch.no_grad():
for i, batch in enumerate(iterator):
src, src_len = batch.src
trg = batch.trg
output, attention = model(src, src_len, trg, 0)
output = output[1:].view(-1, output.shape[-1])
trg = trg[1:].view(-1)
loss = criterion(output, trg)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# +
N_EPOCHS = 10
CLIP = 1
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion, CLIP)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'tut4-model.pt')
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. PPL: {math.exp(valid_loss):7.3f}')
# -
model.load_state_dict(torch.load('tut4-model.pt'))
test_loss = evaluate(model, test_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
| seq2seq/seq2seq4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
img = cv2.imread("../opencv-np-workshop/data/img/lane.jpg", 0)
plt.imshow(img, cmap="gray")
# **Goal:** detect edges in that image.
#
#
# - Edges $\approx$ derivatives
# ## Calculating first derivatives
dx = cv2.Sobel(img, -1, 1, 0, ksize=3)
dy = cv2.Sobel(img, -1, 0, 1, ksize=3)
# First argument: -1 = ddepth = type of array returned, if -1 keeps type of original array
# Second argument = derivative on x
# Third argument = derivative on y
plt.imshow(dx, cmap="gray")
plt.imshow(dy, cmap="gray")
laplacian = cv2.Laplacian(img, -1)
plt.imshow(laplacian, cmap="gray")
# ## Canny edge detection
edges = cv2.Canny(img, 200, 250, 3) #
plt.imshow(edges, cmap="gray")
| Day 2 - Edge detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] editable=true
# <!-- HTML file automatically generated from DocOnce source (https://github.com/doconce/doconce/)
# doconce format html hw1.do.txt --no_mako -->
# <!-- dom:TITLE: PHY321: Classical Mechanics 1 -->
# + [markdown] editable=true
# # PHY321: Classical Mechanics 1
# **Homework 1, due January 21 (midnight)**
#
# Date: **Jan 11, 2022**
# + [markdown] editable=true
# ### Practicalities about homeworks and projects
#
# 1. You can work in groups (optimal groups are often 2-3 people) or by yourself. If you work as a group you can hand in one answer only if you wish. **Remember to write your name(s)**!
#
# 2. Homeworks (final version) are available approximately ten days before the deadline.
#
# 3. How do I(we) hand in? You can hand in the paper and pencil exercises as a scanned document. For this homework this applies to exercises 1-5. You should upload the scan to D2L. Alternatively, you can hand in everyhting (if you are ok with typing mathematical formulae using say Latex) as a jupyter notebook at D2L. The numerical exercise (exercise 6 here) should always be handed in as a jupyter notebook by the deadline at D2L.
# + [markdown] editable=true
# ### Exercise 1 (12 pt), math reminder, properties of exponential function
#
# The first exercise is meant to remind ourselves about properties of
# the exponential function and imaginary numbers. This is highly
# relevant later in this course when we start analyzing oscillatory
# motion and some wave mechanics. As physicists we should thus feel comfortable with expressions that
# include $\exp{(\imath\omega t)}$. Here $t$ could be interpreted as time and $\omega$ as a frequency and $\imath$ is the imaginary unit number.
#
# * 1a (3pt): Perform Taylor expansions in powers of $\omega t$ of the functions $\cos{(\omega t)}$ and $\sin{(\omega t)}$.
#
# * 1b (3pt): Perform a Taylor expansion of $\exp{(i\omega t)}$.
#
# * 1c (3pt): Using parts (a) and (b) here, show that $\exp{(\imath\omega t)}=\cos{(\omega t)}+\imath\sin{(\omega t)}$.
#
# * 1d (3pt): Show that $\ln{(−1)} = \imath\pi$.
# + [markdown] editable=true
# ### Exercise 2 (12 pt), Vector algebra
#
# * 2a (6pt) One of the many uses of the scalar product is to find the angle between two given vectors. Find the angle between the vectors $\boldsymbol{a}=(1,2,4)$ and $\boldsymbol{b}=(4,2,1)$ by evaluating their scalar product.
#
# * 2b (6pt) For a cube with sides of length 1, one vertex at the origin, and sides along the $x$, $y$, and $z$ axes, the vector of the body diagonal from the origin can be written $\boldsymbol{a}=(1, 1, 1)$ and the vector of the face diagonal in the $xy$ plane from the origin is $\boldsymbol{b}=(1,1,0)$. Find first the lengths of the body diagonal and the face diagonal. Use then part (2a) to find the angle between the body diagonal and the face diagonal.
# + [markdown] editable=true
# ### Exercise 3 (10 pt), More vector mathematics
#
# * 3a (5pt) Show (using the fact that multiplication of reals is distributive) that $\boldsymbol{a}(\boldsymbol{b}+\boldsymbol{c})=\boldsymbol{a}\boldsymbol{b}+\boldsymbol{a}\boldsymbol{c}$.
#
# * 3b (5pt) Show that (using product rule for differentiating reals) $\frac{d}{dt}(\boldsymbol{a}\boldsymbol{b})=\boldsymbol{a}\frac{d\boldsymbol{b}}{dt}+\boldsymbol{b}\frac{d\boldsymbol{a}}{dt}$
# + [markdown] editable=true
# ### Exercise 4 (10 pt), Algebra of cross products
#
# * 4a (5pt) Show that the cross products are distribuitive $\boldsymbol{a}\times(\boldsymbol{b}+\boldsymbol{c})=\boldsymbol{a}\times\boldsymbol{b}+\boldsymbol{a}\times\boldsymbol{c}$.
#
# * 4b (5pt) Show that $\frac{d}{dt}(\boldsymbol{a}\times\boldsymbol{b})=\boldsymbol{a}\times\frac{d\boldsymbol{b}}{dt}+\frac{d\boldsymbol{a}}{dt}\times \boldsymbol{b}$. Be careful with the order of factors
# + [markdown] editable=true
# ### Exercise 5 (10 pt), Area of triangle and law of sines
#
# Exercise 1.18 in the textbook of Taylor, Classical Mechanics. Part (1.18a) gives 5pt and part (1.18b) gives also 5pt.
# + [markdown] editable=true
# ### Exercise 6 (40pt), Numerical elements, getting started with some simple data
#
# **This exercise should be handed in as a jupyter-notebook** at D2L. Remember to write your name(s).
#
# Our first numerical attempt will involve reading data from file or
# just setting up two vectors, one for position and one for time. Our data are from
# [Usain Bolt's world record 100m during the olympic games in Beijing in
# 2008](https://www.youtube.com/watch?v=93dC0o2aHto). The data show the time used in units of 10m (see below). Before we however
# venture into this, we need to repeat some basic Python syntax with an
# emphasis on
#
# * basic Python syntax for arrays
#
# * define and operate on vectors and matrices in Python
#
# * create plots for motion in 1D space
#
# For more information, see the [introductory slides](https://mhjensen.github.io/Physics321/doc/pub/week2/html/week2.html).
# Here are some of the basic packages we will be using this week
# + editable=true
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] editable=true
# The first exercise here deals with simply getting familiar with vectors and matrices.
#
# We will be working with vectors and matrices to get you familiar with them
#
# 1. Initalize two three-dimensional $xyz$ vectors in the below cell using np.array([x,y,z]). Vectors are represented through arrays in python
#
# 2. V1 should have x1=1, y1 =2, and z1=3.
#
# 3. Vector 2 should have x2=4, y2=5, and z2=6.
#
# 4. Print both vectors to make sure your code is working properly.
# + editable=true
V1 = np.array([1,2,3])
V2 = np.array([4,5,6])
print("V1: ", V1)
print("V2: ", V2)
# + [markdown] editable=true
# If this is not too familiar, here's a useful link for creating vectors in python
# <https://docs.scipy.org/doc/numpy-1.13.0/user/basics.creation.html>. Alternatively, look up the [introductory slides](https://mhjensen.github.io/Physics321/doc/pub/week2/html/week2.html).
#
# Now lets do some basic mathematics with vectors.
#
# Compute and print the following, and double check with hand calculations:
#
# * 6a (2pt) Calculate $\boldsymbol{V}_1-\boldsymbol{V}_2$.
#
# * 6b (2pt) Calculate $\boldsymbol{V}_2-\boldsymbol{V}_1$.
#
# * 6c (2pt) Calculate the dot product $\boldsymbol{V}_1\boldsymbol{V}_2$.
#
# * 6d (2pt) Calculate the cross product $\boldsymbol{V}_1\times\boldsymbol{V}_2$.
#
# Here is some useful explanation on numpy array operations if you feel a bit confused by what is happening,
# see <https://www.pluralsight.com/guides/overview-basic-numpy-operations>.
#
# The following code prints the first two exercises
# + editable=true
print(V1-V2)
print(V2-V1)
# + [markdown] editable=true
# For the dot product of V1 and V2 below we can use the **dot** function of **numpy** as follows
# + editable=true
print(V1.dot(V2))
# + [markdown] editable=true
# As a small challenge try to write your own function for the **dot** product of two vectors.
#
# Matrices can be created in a similar fashion in python. In this
# language we can work with them through the package numpy (which we
# have already imported)
# + editable=true
M1 = np.matrix([[1,2,3],
[4,5,6],
[7,8,9]])
M2 = np.matrix([[1,2],
[3,4],
[5,6]])
M3 = np.matrix([[9,8,7],
[4,5,6],
[7,6,9]])
# + [markdown] editable=true
# Matrices can be added in the same way vectors are added in python as shown here
# + editable=true
print("M1+M3: ", M1+M3)
# + [markdown] editable=true
# What happens if we try to do $M1+M2$?
#
# That's enough vectors and matrices for now. Let's move on to some physics problems! Yes, the actual subject we are studying for.
#
# We can opt for two different ways of handling the data. The data is listed in the table here and represents the total time Usain Bolt used in steps of 10 meters of distance. The label $i$ is just a counter and we start from zero since Python arrays are by default set from zero. The variable $t$ is time in seconds and $x$ is the position in meters.
#
# <table class="dotable" border="1">
# <thead>
# <tr><th align="center"> i </th> <th align="center"> 0 </th> <th align="center"> 1 </th> <th align="center"> 2 </th> <th align="center"> 3 </th> <th align="center"> 4 </th> <th align="center"> 5 </th> <th align="center"> 6 </th> <th align="center"> 7 </th> <th align="center"> 8 </th> <th align="center"> 9 </th> </tr>
# </thead>
# <tbody>
# <tr><td align="center"> x[m] </td> <td align="center"> 10 </td> <td align="center"> 20 </td> <td align="center"> 30 </td> <td align="center"> 40 </td> <td align="center"> 50 </td> <td align="center"> 60 </td> <td align="center"> 70 </td> <td align="center"> 80 </td> <td align="center"> 90 </td> <td align="center"> 100 </td> </tr>
# <tr><td align="center"> t[s] </td> <td align="center"> 1.85 </td> <td align="center"> 2.87 </td> <td align="center"> 3.78 </td> <td align="center"> 4.65 </td> <td align="center"> 5.50 </td> <td align="center"> 6.32 </td> <td align="center"> 7.14 </td> <td align="center"> 7.96 </td> <td align="center"> 8.79 </td> <td align="center"> 9.69 </td> </tr>
# </tbody>
# </table>
#
# * 6e (6pt) You can here make a file with the above data and read them in and set up two vectors, one for time and one for position. Alternatively, you can just set up these two vectors directly and define two vectors in your Python code.
#
# The following example code may help here
# + editable=true
# we just initialize time and position
x = np.array([10.0, 20.0, 30.0, 40.0, 50.0, 60.0, 70.0, 80.0, 90.0, 100.0])
t = np.array([1.85, 2.87, 3.78, 4.65, 5.50, 6.32, 7.14, 7.96, 8.79, 9.69])
plt.plot(t,x, color='black')
plt.xlabel("Time t[s]")
plt.ylabel("Position x[m]")
plt.title("Usain Bolt's world record run")
plt.show()
# + [markdown] editable=true
# * 6f (6pt) Plot the position as function of time
#
# * 6g (10pt) Compute thereafter the mean velocity for every interval $i$ and the total velocity (from $i=0$ to the given interval $i$) for each interval and plot these two quantities as function of time. Comment your results.
#
# * 6h (10pt) Finally, compute and plot the mean acceleration for each interval and the total acceleration. Again, comment your results. Can you see whether he slowed down during the last meters?
| doc/LectureNotes/_build/jupyter_execute/hw1.ipynb |