code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="XhdK9tifRPYR"
# This is a toy example of using empirical calibration for survey calibration.
# + cellView="form" id="NBeKkRlhq1-Q"
#@title Copyright 2019 The Empirical Calibration Authors.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ============================================================================
# + [markdown] id="rLVCPSCxCJez"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/google/empirical_calibration/blob/master/notebooks/survey_calibration_simulated.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/google/empirical_calibration/blob/master/notebooks/survey_calibration_simulated.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="VGDMBD7TRIf4"
# ## Imports
# + id="hzy_rdy7WXyS" outputId="7d80089c-1972-4dc4-c121-76462cb9216d" colab={"base_uri": "https://localhost:8080/", "height": 90}
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
sns.set_style('whitegrid')
# %config InlineBackend.figure_format='retina'
# install and import ec
# !pip install -q git+https://github.com/google/empirical_calibration
import empirical_calibration as ec
# + [markdown] id="D8orhXBdZq9l"
# ## Selection Bias
# + [markdown] id="N2BGbpoHcLOc"
# Generate a population of size 1000.
# + id="aJ9oaJ4VeD2b" outputId="8ebf10de-c658-4724-ad46-c5f3cd051d05" colab={"base_uri": "https://localhost:8080/", "height": 34}
np.random.seed(123)
N = 1000 # population size.
df_population = pd.DataFrame({
'x1': np.repeat(['F', 'F', 'M', 'M'], repeats=N),
'x2': np.repeat(['H', 'L', 'H', 'L'], repeats=N),
'y': (np.repeat([1e1, 2e1, 3e1, 4e1], repeats=N) +
np.random.normal(size=4*N)),
'response_rate': np.repeat([1.0e-2, 1.1e-2, 1.2e-2, 1.3e-2],
repeats=N)
})
true_mean = df_population['y'].mean()
print('true population mean is {}.'.format(true_mean))
# + [markdown] id="utqFsTLJEUjP"
# Define a function that draws a sample and calculates the estimates of mean.
# + id="CkmBwMobdm2A"
def simulate(seed):
"""Draws sample and calculates (un)weighted estimates."""
np.random.seed(seed)
responded = (np.random.uniform(size=4*N) <=
df_population['response_rate'])
df_sample = df_population.loc[responded]
weights, _ = ec.from_formula(
formula='~x1+x2',
df=df_sample,
baseline_weights = np.repeat(1.0 / df_sample.shape[0], df_sample.shape[0]),
target_df=df_population,
objective=ec.Objective.QUADRATIC)
return pd.Series({
'n': df_sample.shape[0],
'unweighted': df_sample['y'].mean(),
'weighted': df_sample['y'].mul(weights).sum()})
# + id="CRB-gGkpWcNu" outputId="c1d9d36b-5c27-4336-ae07-3feaf8282e83" colab={"base_uri": "https://localhost:8080/", "height": 87}
# Try it out.
simulate(123)
# + id="BlwdzFnfetYi"
# Do it 1000 times and collect results.
result = pd.DataFrame(simulate(seed) for seed in range(1000))
# + id="aaZYjzvn_w7s" outputId="2c37d85b-d983-4b81-9a04-90cb2c2c6914" colab={"base_uri": "https://localhost:8080/", "height": 52}
# Calculate rmse.
print('unweighted rmse is {}'.format(
np.sqrt(np.mean((result['unweighted']-true_mean)**2))))
print('weighted rmse is {}'.format(
np.sqrt(np.mean((result['weighted']-true_mean)**2))))
# + id="qyTeGKSF6dMT" outputId="00f93779-529b-489f-aba2-e138813a322c" colab={"base_uri": "https://localhost:8080/", "height": 494}
# visually compare unweighted vs weighted estimates.
def compare_estimates(unweighted, weighted, truth,
figsize=(12, 8),
fontsize=22,
output_path=None):
f = plt.figure(figsize=figsize)
plt.rcParams.update({'font.size': fontsize})
plt.hist(unweighted,
alpha=0.4,
color='orange',
edgecolor='none',
label='unweighted')
plt.hist(weighted,
alpha=0.4,
color='blue',
edgecolor='none',
label='weighted')
plt.axvline(truth, linestyle='dashed', color='red')
plt.legend(loc='upper left')
plt.show()
if output_path is not None:
f.savefig(output_path, bbox_inches='tight')
compare_estimates(unweighted=result['unweighted'],
weighted=result['weighted'],
truth=true_mean)
| notebooks/survey_calibration_simulated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] state="normal"
# # Lab 01: Basic Python
# + [markdown] state="normal"
# ## Exercise 1: Perfect Numbers
#
# Most of the lab notebooks you'll be working on for this class will come with a fair bit of skeleton code --- i.e., stubbed out classes and functions that you need to complete or modify to get working correctly.
#
# In the cell below, for instance, you'll find a stubbed out function named `is_perfect`, which should return `True` if the number passed to it is a "perfect" number, and `False` otherwise.
#
# A perfect number is a postive integer whose value is equal to the sum of its proper divisors (i.e., its factors excluding the number itself). 6 is the first perfect number, as its divisors 1, 2, and 3 add up to 6.
#
# Fill in your own implementation of the function below:
# + deletable=false id="is_perfect" nbgrader={"grade": false, "grade_id": "is_perfect", "locked": false, "schema_version": 1, "solution": true} starter_code="def is_perfect(n):\n pass" state="graded"
def is_perfect(n):
s = 0
for i in range(1, n):
if n % i == 0:
s += i
result = True if s == n else False
return result
# + [markdown] state="normal"
# Each exercise will also be accompanied by one or more *unit test* cells, each of which is meant to test some aspect of your implementation. When you run the unit test cell(s) after evaluating your implementation, you'll either find errors reported, which should help you identify what you need to fix, or they will complete silently, which means you've passed the test(s).
#
# **It's important that you ensure your implementation and test cell(s) actually run to completion before moving on** --- there's a big difference between a cell not producing an error and not completing! (A "`In [*]`" marker next to the cell means that it's still being evaluated by the interpreter.)
# + deletable=false editable=false id="is_perfect_test" nbgrader={"grade": true, "grade_id": "is_perfect_test", "locked": true, "points": 3, "schema_version": 1, "solution": false} state="read_only"
# (3 points)
import unittest
tc = unittest.TestCase()
for n in (6, 28, 496):
tc.assertTrue(is_perfect(n), '{} should be perfect'.format(n))
for n in (1, 2, 3, 4, 5, 10, 20):
tc.assertFalse(is_perfect(n), '{} should not be perfect'.format(n))
for n in range(30, 450):
tc.assertFalse(is_perfect(n), '{} should not be perfect'.format(n))
# + [markdown] state="normal"
# ## Exercise 2: Multiples of 3 and 5
#
# If we list all the natural numbers below 10 that are multiples of 3 or 5, we get 3, 5, 6 and 9. The sum of these multiples is 23.
#
# Complete the following function, which finds the sum of all the multiples of 3 or 5 below the argument `n`.
# + deletable=false id="multiples_of_3_and_5" nbgrader={"grade": false, "grade_id": "multiples_of_3_and_5", "locked": false, "schema_version": 1, "solution": true} starter_code="def multiples_of_3_and_5(n):\n pass" state="graded"
def multiples_of_3_and_5(n):
s = 0
for i in range(1, n):
if (i % 3 == 0 or i % 5 == 0):
s += i
return s
# + deletable=false editable=false id="multiples_of_3_and_5_test" nbgrader={"grade": true, "grade_id": "multiples_of_3_and_5_test", "locked": true, "points": 3, "schema_version": 1, "solution": false} state="read_only"
# (3 points)
import unittest
tc = unittest.TestCase()
tc.assertEqual(multiples_of_3_and_5(10), 23)
tc.assertEqual(multiples_of_3_and_5(500), 57918)
tc.assertEqual(multiples_of_3_and_5(1000), 233168)
# + [markdown] state="normal"
# ## Exercise 3: Integer Right Triangles
#
# Given a perimeter of 60, we can find two right triangles with integral length sides: [(10, 24, 26), (15, 20, 25)]. Complete the following function, which takes an integer `p` and returns the number of unique integer right triangles with perimeter `p`.
#
# Note that your solution should take care to limit the number of triangles it tests --- **your function must complete in under 3 seconds for all values of `p` used in the test cells below to earn credit.**
# + deletable=false id="integer_right_triangles" nbgrader={"grade": false, "grade_id": "integer_right_triangles", "locked": false, "schema_version": 1, "solution": true} starter_code="def integer_right_triangles(p):\n " state="graded"
def integer_right_triangles(p):
s = 0
for i in range(1, int((p/3)+1)):
j = p-p*p/(2*p-2*i)
if i < j and j - int(j) < 1e-5:
s += 1
return s
# -
# time test
import time
start = time.time()
integer_right_triangles(100000)
end = time.time()
print("take times %.3fs" % (end-start))
# + deletable=false editable=false id="integer_right_triangles_tests_1" nbgrader={"grade": true, "grade_id": "integer_right_triangles_tests_1", "locked": true, "points": 1, "schema_version": 1, "solution": false} state="read_only"
# (2 points)
import unittest
tc = unittest.TestCase()
tc.assertEqual(integer_right_triangles(60), 2)
tc.assertEqual(integer_right_triangles(100), 0)
tc.assertEqual(integer_right_triangles(180), 3)
# + [markdown] state="normal"
# ## Exercise 4: Simple ASCII Art
#
# For this next exercise, you'll need to complete the function `gen_pattern`, which, when called with a string of length $\ge$ 1, will print an ASCII art pattern of concentric diamonds using those characters. The following are examples of patterns printed by the function (note the newline at the end of the last line!):
#
# > gen_pattern('X')
#
# X
#
# > gen_pattern('XY')
#
# ..Y..
# Y.X.Y
# ..Y..
#
# > gen_pattern('WXYZ')
#
# ......Z......
# ....Z.Y.Z....
# ..Z.Y.X.Y.Z..
# Z.Y.X.W.X.Y.Z
# ..Z.Y.X.Y.Z..
# ....Z.Y.Z....
# ......Z......
#
# You ought to find the string [`join`](https://docs.python.org/3.6/library/stdtypes.html#str.join) and [`center`](https://docs.python.org/3.6/library/stdtypes.html#str.center) methods helpful in your implementation. They are demonstrated here:
#
# > '*'.join('abcde')
#
# 'a*b*c*d*e'
#
# > 'hello'.center(11, '*')
#
# '***hello***'
#
# Complete the `gen_pattern` function, below:
# + deletable=false id="gen_pattern" nbgrader={"grade": false, "grade_id": "gen_pattern", "locked": false, "schema_version": 1, "solution": true} starter_code="def gen_pattern(chars):\n pass" state="graded"
def gen_pattern(chars):
gen_str = lambda x: x[::-1]+x[1:]
gen_join = lambda x: '.'.join(gen_str(x))
gen_line = lambda x: gen_join(x).center(len(gen_join(chars)), '.')
gen_list_str = list(map(lambda x: chars[x:], list(range(len(chars)))))
gen_list_str = gen_list_str[1:][::-1] + gen_list_str
list(map(lambda x: print(gen_line(x)), gen_list_str))
# + deletable=false editable=false id="gen_pattern1" state="read_only"
# (1 point) output:
# @
gen_pattern('@')
# + deletable=false editable=false id="gen_pattern2" state="read_only"
# (1 point) output:
# ..%..
# %.@.%
# ..%..
gen_pattern('@%')
# + deletable=false editable=false id="gen_pattern3" state="read_only"
# (1 point) output:
# ....C....
# ..C.B.C..
# C.B.A.B.C
# ..C.B.C..
# ....C....
gen_pattern('ABC')
# + deletable=false editable=false id="gen_pattern5" state="read_only"
# (1 point) output:
# ........#........
# ......#.#.#......
# ....#.#.#.#.#....
# ..#.#.#.#.#.#.#..
# #.#.#.#.#.#.#.#.#
# ..#.#.#.#.#.#.#..
# ....#.#.#.#.#....
# ......#.#.#......
# ........#........
gen_pattern('#####')
# + deletable=false editable=false id="gen_pattern_tests16" nbgrader={"grade": true, "grade_id": "gen_pattern_tests_2", "locked": true, "points": 2, "schema_version": 1, "solution": false} state="read_only"
# (2 points) output:
# ..............................p..............................
# ............................p.o.p............................
# ..........................p.o.n.o.p..........................
# ........................p.o.n.m.n.o.p........................
# ......................p.o.n.m.l.m.n.o.p......................
# ....................p.o.n.m.l.k.l.m.n.o.p....................
# ..................p.o.n.m.l.k.j.k.l.m.n.o.p..................
# ................p.o.n.m.l.k.j.i.j.k.l.m.n.o.p................
# ..............p.o.n.m.l.k.j.i.h.i.j.k.l.m.n.o.p..............
# ............p.o.n.m.l.k.j.i.h.g.h.i.j.k.l.m.n.o.p............
# ..........p.o.n.m.l.k.j.i.h.g.f.g.h.i.j.k.l.m.n.o.p..........
# ........p.o.n.m.l.k.j.i.h.g.f.e.f.g.h.i.j.k.l.m.n.o.p........
# ......p.o.n.m.l.k.j.i.h.g.f.e.d.e.f.g.h.i.j.k.l.m.n.o.p......
# ....p.o.n.m.l.k.j.i.h.g.f.e.d.c.d.e.f.g.h.i.j.k.l.m.n.o.p....
# ..p.o.n.m.l.k.j.i.h.g.f.e.d.c.b.c.d.e.f.g.h.i.j.k.l.m.n.o.p..
# p.o.n.m.l.k.j.i.h.g.f.e.d.c.b.a.b.c.d.e.f.g.h.i.j.k.l.m.n.o.p
# ..p.o.n.m.l.k.j.i.h.g.f.e.d.c.b.c.d.e.f.g.h.i.j.k.l.m.n.o.p..
# ....p.o.n.m.l.k.j.i.h.g.f.e.d.c.d.e.f.g.h.i.j.k.l.m.n.o.p....
# ......p.o.n.m.l.k.j.i.h.g.f.e.d.e.f.g.h.i.j.k.l.m.n.o.p......
# ........p.o.n.m.l.k.j.i.h.g.f.e.f.g.h.i.j.k.l.m.n.o.p........
# ..........p.o.n.m.l.k.j.i.h.g.f.g.h.i.j.k.l.m.n.o.p..........
# ............p.o.n.m.l.k.j.i.h.g.h.i.j.k.l.m.n.o.p............
# ..............p.o.n.m.l.k.j.i.h.i.j.k.l.m.n.o.p..............
# ................p.o.n.m.l.k.j.i.j.k.l.m.n.o.p................
# ..................p.o.n.m.l.k.j.k.l.m.n.o.p..................
# ....................p.o.n.m.l.k.l.m.n.o.p....................
# ......................p.o.n.m.l.m.n.o.p......................
# ........................p.o.n.m.n.o.p........................
# ..........................p.o.n.o.p..........................
# ............................p.o.p............................
# ..............................p..............................
gen_pattern('abcdefghijklmnop')
# -
| cs-master/cs401/Lab01-Basic Python_done.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# -*- coding: utf-8 -*-
import os
import re
import glob
import argparse
import datetime
import zipfile
import numpy as np
import pandas as pd
# -
def get_radar_archive_file(date, rid):
"""
Return the archive containing the radar file for a given radar ID and a
given date.
Parameters:
===========
date: datetime
Date.
Returns:
========
file: str
Radar archive if it exists at the given date.
"""
if type(rid) is not str:
rid = f"{rid:02}"
datestr = date.strftime("%Y%m%d")
file = f"/g/data/rq0/admin/level_1b/grid_150km/{rid}/{date.year}/{rid}_{datestr}_level1b_grid_150km.zip"
if not os.path.exists(file):
print(f"{file} does not exist.")
return None
return file
def extract_zip(inzip, date, path="/scratch/kl02/vhl548/unzipdir"):
"""
Extract file in a daily archive zipfile for a specific datetime.
Parameters:
===========
inzip: str
Input zipfile
date: pd.Timestamp
Which datetime we want to extract.
path: str
Path where we want to temporarly store the output file.
Returns:
========
grfile: str
Output ground radar file.
"""
def get_zipfile_name(namelist, date):
datestr = [re.findall("[0-9]{8}_[0-9]{6}", n)[0] for n in namelist]
timestamps = np.array([datetime.datetime.strptime(dt, "%Y%m%d_%H%M%S") for dt in datestr], dtype="datetime64")
pos = np.argmin(np.abs(timestamps - date.to_numpy()))
delta = np.abs(pd.Timestamp(timestamps[pos]) - date).seconds
grfile = namelist[pos]
return grfile
with zipfile.ZipFile(inzip) as zid:
namelist = zid.namelist()
file = get_zipfile_name(namelist, date)
zid.extract(file, path=path)
grfile = os.path.join(path, file)
return grfile
def get_cpol_file(date):
datestr = date.strftime("%Y%m%d")
path = f"/scratch/kl02/vhl548/cpol_level_1b/v2020/gridded/grid_150km_1000m/{date.year}/{datestr}/*.nc"
namelist = sorted(glob.glob(path))
if len(namelist) == 0:
raise FileNotFoundError(f"No CPOL file found for this date {datestr}.")
datelist = [re.findall("[0-9]{8}.[0-9]{6}", n)[0] for n in namelist]
timestamps = np.array([datetime.datetime.strptime(dt, "%Y%m%d.%H%M%S") for dt in datelist], dtype="datetime64")
pos = np.argmin(np.abs(timestamps - date.to_numpy()))
delta = np.abs(pd.Timestamp(timestamps[pos]) - date).seconds
grfile = namelist[pos]
return grfile
input_date = datetime.datetime(2014, 12, 1, 6, 30)
date = [pd.Timestamp(input_date)]
date.append(date[0] + pd.Timedelta("10Min"))
# +
files = [get_cpol_file(d) for d in date]
zips = [get_radar_archive_file(d, 64) for d in date]
for z, d in zip(zips, date):
files.append(extract_zip(z, d))
# -
with open("r3dbrc", "w+") as fid:
fid.write("\n".join(files))
pd.Timestamp("201701011210")
| notebook/generate_3dwinds_config.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# # Visualize channel over epochs as an image
#
#
# This will produce what is sometimes called an event related
# potential / field (ERP/ERF) image.
#
# 2 images are produced. One with a good channel and one with a channel
# that does not see any evoked field.
#
# It is also demonstrated how to reorder the epochs using a 1d spectral
# embedding as described in:
#
# Graph-based variability estimation in single-trial event-related neural
# responses <NAME>, <NAME>, <NAME>, 2010,
# Biomedical Engineering, IEEE Trans. on, vol. 57 (5), 1051-1061
# https://hal.inria.fr/inria-00497023
#
#
# +
# Authors: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne import io
from mne.datasets import sample
print(__doc__)
data_path = sample.data_path()
# -
# Set parameters
#
#
# +
raw_fname = data_path + '/MEG/sample/sample_audvis_filt-0-40_raw.fif'
event_fname = data_path + '/MEG/sample/sample_audvis_filt-0-40_raw-eve.fif'
event_id, tmin, tmax = 1, -0.2, 0.5
# Setup for reading the raw data
raw = io.read_raw_fif(raw_fname)
events = mne.read_events(event_fname)
# Set up pick list: EEG + MEG - bad channels (modify to your needs)
raw.info['bads'] = ['MEG 2443', 'EEG 053']
picks = mne.pick_types(raw.info, meg='grad', eeg=False, stim=True, eog=True,
exclude='bads')
# Read epochs
epochs = mne.Epochs(raw, events, event_id, tmin, tmax, proj=True,
picks=picks, baseline=(None, 0), preload=True,
reject=dict(grad=4000e-13, eog=150e-6))
# -
# Show event related fields images
#
#
# +
# and order with spectral reordering
# If you don't have scikit-learn installed set order_func to None
from sklearn.cluster.spectral import spectral_embedding # noqa
from sklearn.metrics.pairwise import rbf_kernel # noqa
def order_func(times, data):
this_data = data[:, (times > 0.0) & (times < 0.350)]
this_data /= np.sqrt(np.sum(this_data ** 2, axis=1))[:, np.newaxis]
return np.argsort(spectral_embedding(rbf_kernel(this_data, gamma=1.),
n_components=1, random_state=0).ravel())
good_pick = 97 # channel with a clear evoked response
bad_pick = 98 # channel with no evoked response
plt.close('all')
mne.viz.plot_epochs_image(epochs, [good_pick, bad_pick], sigma=0.5, vmin=-100,
vmax=250, colorbar=True, order=order_func, show=True)
| 0.12/_downloads/plot_channel_epochs_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (cv)
# language: python
# name: cv
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Introduction to image similarity
#
# Image similarity methods can be used to build Image Retrieval systems where, given a query image, the goal is to find all similar images in a reference set. These systems can be used e.g. on a shopping website to suggest related products.
#
# In this tutorial we build an image retrieval system based on leveraging DNNs trained for image classification. Representing images as the output of a DNN is a powerful approach and shown to give good results on a wide variety of tasks. Given a query image, we find the most similar images in the reference set by computing the pairwise distances as illustrated below, and by returning the images with the lowest distance to the query image.
#
# The distance between two images is computed by:
# <img src="media/image_similarity_diagram2.jpg" width="600px" />
#
# - Representing each image using the output of a DNN which was pre-trained on millions of images and fine-tuned on images from the same domain. The input to the DNN is an image, and the output is the penultimate layer which, for the ResNet-18 model used in this tutorial, consists of 512-floating point values. Note that we use only a single DNN which takes each of the two images independently as input.
# - Normalizing each of the 512-floats DNN image representations (aka. embeddings or features) to be unit vectors, i.e. have an L2 norm of one.
# - Finally, obtaining the distance between the two input images by computing the L2 distance between their respective DNN representations.
#
# This notebook starts by loading a dataset and splitting it into a training and a validation set. The training set is used to refine an ImageNet pre-trained ResNet-18 DNN, which is then used to compute the DNN features for each image. The validation set is used in an image retrieval example where, given a query image, the top similar images are displayed. This is followed by a quantitative evaluation of the proposed image similarity system.
# ## Initialization
# Ensure edits to libraries are loaded and plotting is shown in the notebook.
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
# +
# Regular python libraries
import sys
import numpy as np
from pathlib import Path
import random
import scrapbook as sb
# fast.ai
import fastai
from fastai.vision import (
accuracy,
cnn_learner,
DatasetType,
ImageList,
imagenet_stats,
models,
partial,
)
# Computer Vision repository
sys.path.extend([".", "../.."]) # to access the utils_cv library
from utils_cv.classification.data import Urls
from utils_cv.classification.model import TrainMetricsRecorder
from utils_cv.common.data import unzip_url
from utils_cv.common.gpu import which_processor, db_num_workers
from utils_cv.similarity.data import comparative_set_builder
from utils_cv.similarity.metrics import (
compute_distances,
positive_image_ranks,
recall_at_k,
)
from utils_cv.similarity.model import compute_features, compute_features_learner
from utils_cv.similarity.plot import (
plot_comparative_set,
plot_distances,
plot_ranks_distribution,
plot_recalls,
)
from utils_cv.similarity.widget import RetrievalWidget
# -
print(f"Fast.ai version = {fastai.__version__}")
which_processor()
# ## Data preparation
# We start with parameter specifications and data preparation. We use the *Fridge objects* dataset, which is composed of 134 images, divided into 4 classes: can, carton, milk bottle and water bottle. To train your own image retrieval systems, simply change the `DATA_PATH` variable below to point to a different (single-label) dataset.
# + tags=["parameters"]
# Set dataset, model and evaluation parameters
DATA_PATH = unzip_url(Urls.fridge_objects_path, exist_ok=True)
# DNN configuration and learning parameters
EPOCHS_HEAD = 4
EPOCHS_BODY = 12
LEARNING_RATE = 10* 1e-4
BATCH_SIZE = 16
ARCHITECTURE = models.resnet18
IM_SIZE = 300
# -
# We can now build our training data object, and split it to get a certain percentage (here 20%) assigned to a validation set.
# +
# Load images into fast.ai's ImageDataBunch object
random.seed(642)
data = (
ImageList.from_folder(DATA_PATH)
.split_by_rand_pct(valid_pct=0.2, seed=20)
.label_from_folder()
.transform(size=IM_SIZE)
.databunch(bs=BATCH_SIZE, num_workers = db_num_workers())
.normalize(imagenet_stats)
)
print(f"""\
Training set: {len(data.train_ds.x)} images
Validation set: {len(data.valid_ds.x)} images\
"""
)
# Display example images
data.show_batch(rows=3, figsize=(6, 6))
# -
# ## Classification model fine-tuning
#
# We begin by retrieving a [ResNet18](https://arxiv.org/pdf/1512.03385.pdf) CNN from fast.ai's library which is pre-trained on ImageNet, and fine-tune the model on our training set. We use the same training parameters and take the same approach as what we did in our [classification notebooks](https://github.com/microsoft/ComputerVision/tree/master/classification/notebooks), training first the (new) last layer only, and then the full DNN.
#
# Note how we train the DNN here on an image classification task but will use it as featurizer later for image similarity.
learn = cnn_learner(
data,
ARCHITECTURE,
metrics=[accuracy],
callback_fns=[partial(TrainMetricsRecorder, show_graph=True)],
ps=0 #Leave dropout at zero. Higher values tend to perform significantly worse
)
# Train the last layer using a larger rate since most of the DNN is fixed.
learn.fit_one_cycle(EPOCHS_HEAD, 10* LEARNING_RATE)
# Let's now unfreeze all the layers and fine-tuning the model more
learn.unfreeze()
learn.fit_one_cycle(EPOCHS_BODY, LEARNING_RATE)
# ## Feature extraction
#
# Before computing the feature representation for each image, let's look at its architecture and in particular the last layers. Fast.ai's ResNet-18 model is composed of a different set of final layers (here: `(1): Sequential`). As discussed at the start of this notebook, we use the output of the penultimate layer (here: `(6): BatchNorm1d`) as our image representation.
learn.model
# The following line will allow us to extract the penultimate layer (ie 512 floating points vector) after running an image through the model.
# Use penultimate layer as image representation
embedding_layer = learn.model[1][-2]
print(embedding_layer)
#Compute DNN features for all validation images
valid_features = compute_features_learner(data, DatasetType.Valid, learn, embedding_layer)
# ## Image Retrieval Example
# The cell below shows how to find and display the most similar images in the validation set for a given query image (which we also select from the validation set). This example is similar to the one shown in the [00_webcam.ipynb](https://github.com/microsoft/ComputerVision/tree/master/similarity/notebooks/00_webcam.ipynb) notebook.
#
# We use the L2 distance which is defined as <sup> $ \sqrt{\sum_{i=1}^{n}{(F_{q}[i] - F_{r}[i])^{2}}} $ </sup> where $F_{q}$ and $F_{r}$ are the features of a query image and a reference image respectively, and $n=512$ is their dimensionality. By default, we normalize the feature vectors $F_{q}$ and $F_{r}$ to be unit-length, i.e. have a magnitude ||$F$|| of 1, before computing the L2 distance. One could also use other distances measures, such as L1 or cosine similarity, however L2 with unit-length normalized feature vectors seems to work well in practice.
# +
# Get the DNN feature for the query image
query_im_path = str(data.valid_ds.items[1])
query_feature = valid_features[query_im_path]
print(f"Query image path: {query_im_path}")
print(f"Query feature dimension: {len(query_feature)}")
assert len(query_feature) == 512
# Compute the distances between the query and all reference images
distances = compute_distances(query_feature, valid_features)
plot_distances(distances, num_rows=1, num_cols=7, figsize=(15,5))
# -
# The `RetrievalWidget` below can be used to interactively inspect the ranking output of our image retrieval system. Given a query image, the most similar images are shown with their respective distance to the query image in brackets. A new image can be selected as query by clicking its yellow box above the image. Note that Jupyter widgets are quite unstable - if the widget below does not show then see the "Troubleshooting" section in this [FAQ](../classification/FAQ.md) for possible fixes.
# <img src="./media/retrieval_ui1.jpg" width="600"/>
# <center><i>Image retrieval widget</i></center>
w_results = RetrievalWidget(
ds=data.valid_ds,
features=valid_features,
rows=2,
cols=5,
)
display(w_results.show())
# ## Quantitative evaluation
#
# To measure accuracy of our image retrieval system, we create so called `comparative sets` from the validation images. Each comparative set consists of a `query` image, a `positive` image (with the same label as the query image), and 99 `negative` images (different label). When sorting the 101 reference images according to their distance to the query image, a perfect image similarity system would place the positive image at the top before all negative images i.e. at rank 1.
#
#
# ### Comparative sets creation
#
# In the cell below, we construct 1000 comparative sets from the validation set, each with 99 negative images (and one positive image).
# Build multiple sets of comparative images from the validation images
comparative_sets = comparative_set_builder(data.valid_ds, num_sets = 1000, num_negatives = 99)
print(f"Generated {len(comparative_sets)} comparative image sets.")
# Plot the query image, the positive image, and some of the negative images of the first comparative set
plot_comparative_set(comparative_sets[0], 7, figsize=(15,5))
# For each comparative set compute the distances between the query image and all reference images
for cs in comparative_sets:
cs.compute_distances(valid_features)
# ### Evaluation
#
# To measure accuracy of our image retrieval system, we compute these two statistics:
# - Median rank: median rank of the positive example across all comparative sets. A value of 1 being best, 100 worst, and 50 random chance guessing.
# - Recall at k: percentage of positive examples within the top k, ie. with rank <= k. A value of 100% being best, 0% worst, and at k=1 or k=5 random chance guessing of 1% or 5% respectively.
# Compute the median rank of the positive example over all comparative sets
ranks = positive_image_ranks(comparative_sets)
median_rank = np.median(ranks)
random_rank = np.median([(len(cs.neg_im_paths)+1)/2.0 for cs in comparative_sets])
print(f"The positive example ranks {median_rank}, as a median, \
across our {len(ranks)} comparative sets. Random chance rank is {random_rank}")
# +
# Compute recall at k=1, 5, and 10
print(f"""The positive image is:
--> {recall_at_k(ranks, 1)}% of the time the most similar to the query
--> {recall_at_k(ranks, 5)}% of the time in the top 5 images
--> {recall_at_k(ranks, 10)}% of the time in the top 10 images""")
# Plot recall versus k
plot_recalls(ranks)
# -
# Display the distribution of positive ranks among the comparative sets
plot_ranks_distribution(ranks)
# Write trained model to disk
learn.export("image_similarity_01_model")
print(f"Exported model to directory {learn.path}")
# ## Fine-tuning parameters <a name="finetuning"></a>
#
# Using the provided default parameters, one can get good results across a wide variety of datasets. However, as in most machine learning projects, getting the best possible results for a new dataset often requires tuning the parameters further.
#
# See the image classification [03_training_accuracy_vs_speed.ipynb](../../classification/notebooks/03_training_accuracy_vs_speed.ipynb) notebook for guidelines on optimizing for accuracy, inference speed, or model size for a given dataset. In addition, the notebook also goes through the parameters that will make the largest impact on your model as well as the parameters that may not be worth modifying.
#
# The notebook [11_exploring_hyperparameters.ipynb](11_exploring_hyperparameters.ipynb) in this directory is provided to run run sweeps to find the parameters with best possible image retrieval (ie rank) performance. Below is an example where, to identify good default parameters for this repository, different learning rates where tried on diverse datasets. Note that lower ranks is better, and learning rates between $1e-4$ and $1e-3$ performed best.
# <img src="media/sweep_learning_rates.jpg" width="600px" />
#
# Log some outputs using scrapbook which are used during testing to verify correct notebook execution
sb.glue("median_rank", median_rank)
sb.glue("random_rank", random_rank)
| scenarios/similarity/01_training_and_evaluation_introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install numpy
import numpy as np
import copy
# 
# +
R1 = np.matrix([[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,0,-1,-1,-2,-2,0,-2,-1,-2,-2,-2,-1,-1,-1,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,0,-1,0,-1,-2,-2,-1,-2,-1,-2,-2,-2,-1,-1,-1,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,0,-1,0,-2,-2,-1,-2,0,-2,-2,-2,-1,-1,-1,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,-1,0,-1,-2,-2,-1,-2,-1,-2,-2,-2,-1,-1,-1,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,0,-1,-1,-1,-2,-2,-1,-2,-1,-2,-2,-2,0,-1,-1,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,-1,0,-1,-2,-2,-1,-2 ,-1,-2,-2,-2,-1,-1,0,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,-1,-1,-1,-2,-2,0,-2,-1,-2,-2,-2,-1,0,-1,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,-1,-1,-1,-2,-2,-1,-2,-1,-2,-2,-2,0,-1,0,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,-1,-1,-1,-2,-2,-1,-2,0,-2,-2,-2,-1,0,-1,0,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-1,-1,-1,-1,-2,-2,-1,-2 ,-1,-2,-2,-2,-1,-1,0,-1,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2],
[-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2,-2]])
#[-2,-2,-2,-2,-2,-2,-2, , , , ,-2,-2, ,-2 , ,-2,-2,-2, , , , ,-2,-2,-2,-2,-2,-2,-2]
# -
R=copy.deepcopy(R1)
Q = np.matrix (np.zeros([30,30]))
gamma = 0.8
initial_state = 8
# +
valid=[7,8,9,10,13,15,19,20,21,22]
print('The valid Locations in the map are 7,8,9,10,13,15,19,20,21 and 22')
print('The Obstacles are at position 14 and Danger is at 16')
while True:
cur=int(input('Enter the current location :'))
des=int(input('Enter the destination location :'))
print()
if cur in valid and des in valid:
break
else:
print('You have enter one of wrong values. Please Enter Again.')
print()
continue
# -
dic={7:[8,13],8:[7,9],9:[8,10,15],10:[9],13:[7,19],15:[9,21],19:[13,20],20:[19,21],21:[15,20,22],22:[21]}
y=dic[des]
R[des,des]=100
for x in y:
R[x,des]=100
# +
initial_state=cur
def available_actions(state):
current_state_row = R[state,]
av_act = np.where(current_state_row >= 0)[1]
return av_act
available_act = available_actions(initial_state)
def sample_next_action(available_actions_range):
next_action = int(np.random.choice(available_act,1))
return next_action
action = sample_next_action(available_act)
# +
def update(current_state, action, gamma):
max_index = np.where(Q[action,] == np.max(Q[action,]))[1]
if max_index.shape[0] > 1:
max_index = int(np.random.choice(max_index, size = 1))
else:
max_index = int(max_index)
max_value = Q[action, max_index]
Q[current_state, action] = R[current_state, action] + gamma * max_value
update(initial_state,action,gamma)
print(Q)
# -
import random
for i in range(10000):
current_state = random.choice(valid)
available_act = available_actions(current_state)
action = sample_next_action(available_act)
update(current_state,action,gamma)
print(Q)
print("Trained Q matrix:")
print(Q/np.max(Q)*100)
current_state = cur
steps = [current_state]
while current_state != des:
next_step_index = np.where(Q[current_state,] == np.max(Q[current_state,]))[1]
if next_step_index.shape[0] > 1:
next_step_index = int(np.random.choice(next_step_index, size=1))
else:
next_step_index = int(next_step_index)
steps.append(next_step_index)
current_state = next_step_index
print("Selected path:")
print(steps)
| Basic Reinforcement Learning/.ipynb_checkpoints/BasicReinforcement-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Methods Using Turtle
#
# Here we introduce you to using Python methods by making a method for drawing a star, then repeatedly calling that method from the main program. You can copy and paste this code into your IDE to see the result.
# +
# This is a program that will draw stars with random values
# all over the window. The stars are drawn using a method
# with parameters.
# Program Structure Note:
# imports come first
# methods come next
# main program comes last
import turtle as t
import random
# method to draw a star
def draw_star(size, num_rays, color, x, y):
# go to the right location
t.goto(x,y)
# set the proper color
if( color % 4 == 0 ):
t.color("red")
if( color % 4 == 1 ):
t.color("green")
if( color % 4 == 2 ):
t.color("yellow")
if( color % 4 == 3 ):
t.color("blue")
# draw the star
t.pendown()
for i in range(num_rays): # loop num_rays number of times
t.left( 360 / num_rays ) # num_rays lines in our star
t.forward(size)
t.backward(size)
t.penup()
# end of the draw_star method
# main program
t.reset() # clear the window for drawing
t.shape("turtle")
t.penup()
t.pensize(3)
num_stars = 20
for i in range(num_stars):
# setup star values
star_size = random.randrange(10,100) # ray size from 10 to 100 px
star_rays = random.randrange(5,10) # num rays from 5 through 9
star_color = random.randrange(4) # there will be 4 possible color values
star_x = random.randrange(-300, 300) # set the star's x location
star_y = random.randrange(-300, 300) # set the star's y location
draw_star(star_size, star_rays, star_color, star_x, star_y)
# end of for loop
t.hideturtle()
| turtle-draw-using-methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Accession: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE133344
# +
import gzip
import os
import re
import pandas as pd
import numpy as np
from anndata import AnnData
from scipy.io import mmread
from scipy.sparse import coo_matrix
from utils import download_binary_file
# Gene program lists obtained by cross-referencing the heatmap here
# https://github.com/thomasmaxwellnorman/Perturbseq_GI/blob/master/GI_optimal_umap.ipynb
# with Figure 2b in Norman 2019
G1_CYCLE = [
"CDKN1C+CDKN1B",
"CDKN1B+ctrl",
"CDKN1B+CDKN1A",
"CDKN1C+ctrl",
"ctrl+CDKN1A",
"CDKN1C+CDKN1A",
"CDKN1A+ctrl",
]
ERYTHROID = [
"BPGM+SAMD1",
"ATL1+ctrl",
"UBASH3B+ZBTB25",
"PTPN12+PTPN9",
"PTPN12+UBASH3A",
"CBL+CNN1",
"UBASH3B+CNN1",
"CBL+UBASH3B",
"UBASH3B+PTPN9",
"PTPN1+ctrl",
"CBL+PTPN9",
"CNN1+UBASH3A",
"CBL+PTPN12",
"PTPN12+ZBTB25",
"UBASH3B+PTPN12",
"SAMD1+PTPN12",
"SAMD1+UBASH3B",
"UBASH3B+UBASH3A",
]
PIONEER_FACTORS = [
"ZBTB10+SNAI1",
"FOXL2+MEIS1",
"POU3F2+CBFA2T3",
"DUSP9+SNAI1",
"FOXA3+FOXA1",
"FOXA3+ctrl",
"LYL1+IER5L",
"FOXA1+FOXF1",
"FOXF1+HOXB9",
"FOXA1+HOXB9",
"FOXA3+HOXB9",
"FOXA3+FOXA1",
"FOXA3+FOXL2",
"POU3F2+FOXL2",
"FOXF1+FOXL2",
"FOXA1+FOXL2",
"HOXA13+ctrl",
"ctrl+HOXC13",
"HOXC13+ctrl",
"MIDN+ctrl",
"TP73+ctrl",
]
GRANULOCYTE_APOPTOSIS = [
"SPI1+ctrl",
"ctrl+SPI1",
"ctrl+CEBPB",
"CEBPB+ctrl",
"JUN+CEBPA",
"CEBPB+CEBPA",
"FOSB+CEBPE",
"ZC3HAV1+CEBPA",
"KLF1+CEBPA",
"ctrl+CEBPA",
"CEBPA+ctrl",
"CEBPE+CEBPA",
"CEBPE+SPI1",
"CEBPE+ctrl",
"ctrl+CEBPE",
"CEBPE+RUNX1T1",
"CEBPE+CEBPB",
"FOSB+CEBPB",
"ETS2+CEBPE",
]
MEGAKARYOCYTE = [
"ctrl+ETS2",
"MAPK1+ctrl",
"ctrl+MAPK1",
"ETS2+MAPK1",
"CEBPB+MAPK1",
"MAPK1+TGFBR2",
]
PRO_GROWTH = [
"CEBPE+KLF1",
"KLF1+MAP2K6",
"AHR+KLF1",
"ctrl+KLF1",
"KLF1+ctrl",
"KLF1+BAK1",
"KLF1+TGFBR2",
]
def download_norman_2019(output_path: str) -> None:
"""
Download Norman et al. 2019 data and metadata files from the hosting URLs.
Args:
----
output_path: Output path to store the downloaded and unzipped
directories.
Returns
-------
None. File directories are downloaded to output_path.
"""
file_urls = (
"https://ftp.ncbi.nlm.nih.gov/geo/series/GSE133nnn/GSE133344/suppl"
"/GSE133344_filtered_matrix.mtx.gz",
"https://ftp.ncbi.nlm.nih.gov/geo/series/GSE133nnn/GSE133344/suppl"
"/GSE133344_filtered_genes.tsv.gz",
"https://ftp.ncbi.nlm.nih.gov/geo/series/GSE133nnn/GSE133344/suppl"
"/GSE133344_filtered_barcodes.tsv.gz",
"https://ftp.ncbi.nlm.nih.gov/geo/series/GSE133nnn/GSE133344/suppl"
"/GSE133344_filtered_cell_identities.csv.gz",
)
for url in file_urls:
output_filename = os.path.join(output_path, url.split("/")[-1])
download_binary_file(url, output_filename)
def read_norman_2019(file_directory: str) -> coo_matrix:
"""
Read the expression data for Norman et al. 2019 in the given directory.
Args:
----
file_directory: Directory containing Norman et al. 2019 data.
Returns
-------
A sparse matrix containing single-cell gene expression count, with rows
representing genes and columns representing cells.
"""
with gzip.open(
os.path.join(file_directory, "GSE133344_filtered_matrix.mtx.gz"), "rb"
) as f:
matrix = mmread(f)
return matrix
# +
download_path = "./norman2019/"
download_norman_2019(download_path)
matrix = read_norman_2019(download_path)
# List of cell barcodes. The barcodes in this list are stored in the same order
# as cells are in the count matrix.
cell_barcodes = pd.read_csv(
os.path.join(download_path, "GSE133344_filtered_barcodes.tsv.gz"),
sep="\t",
header=None,
names=["cell_barcode"],
)
# IDs/names of the gene features.
gene_list = pd.read_csv(
os.path.join(download_path, "GSE133344_filtered_genes.tsv.gz"),
sep="\t",
header=None,
names=["gene_id", "gene_name"],
)
# Dataframe where each row corresponds to a cell, and each column corresponds
# to a gene feature.
matrix = pd.DataFrame(
matrix.transpose().todense(),
columns=gene_list["gene_id"],
index=cell_barcodes["cell_barcode"],
dtype="int32",
)
# Dataframe mapping cell barcodes to metadata about that cell (e.g. which CRISPR
# guides were applied to that cell). Unfortunately, this list has a different
# ordering from the count matrix, so we have to be careful combining the metadata
# and count data.
cell_identities = pd.read_csv(
os.path.join(download_path, "GSE133344_filtered_cell_identities.csv.gz")
).set_index("cell_barcode")
# This merge call reorders our metadata dataframe to match the ordering in the
# count matrix. Some cells in `cell_barcodes` do not have metadata associated with
# them, and their metadata values will be filled in as NaN.
aligned_metadata = pd.merge(
cell_barcodes,
cell_identities,
left_on="cell_barcode",
right_index=True,
how="left",
).set_index("cell_barcode")
adata = AnnData(matrix)
adata.obs = aligned_metadata
# Filter out any cells that don't have metadata values.
rows_without_nans = [
index for index, row in adata.obs.iterrows() if not row.isnull().any()
]
adata = adata[rows_without_nans, :]
# Remove these as suggested by the authors. See lines referring to
# NegCtrl1_NegCtrl0 in GI_generate_populations.ipynb in the Norman 2019 paper's
# Github repo https://github.com/thomasmaxwellnorman/Perturbseq_GI/
adata = adata[adata.obs["guide_identity"] != "NegCtrl1_NegCtrl0__NegCtrl1_NegCtrl0"]
# We create a new metadata column with cleaner representations of CRISPR guide
# identities. The original format is <Guide1>_<Guide2>__<Guide1>_<Guide2>_<number>
adata.obs["guide_merged"] = adata.obs["guide_identity"]
control_regex = re.compile(r"NegCtrl(.*)_NegCtrl(.*)+NegCtrl(.*)_NegCtrl(.*)")
for i in adata.obs["guide_merged"].unique():
if control_regex.match(i):
# For any cells that only had control guides, we don't care about the
# specific IDs of the guides. Here we relabel them just as "ctrl".
adata.obs["guide_merged"].replace(i, "ctrl", inplace=True)
else:
# Otherwise, we reformat the guide label to be <Guide1>+<Guide2>. If Guide1
# or Guide2 was a control, we replace it with "ctrl".
split = i.split("__")[0]
split = split.split("_")
for j, string in enumerate(split):
if "NegCtrl" in split[j]:
split[j] = "ctrl"
adata.obs["guide_merged"].replace(i, f"{split[0]}+{split[1]}", inplace=True)
guides_to_programs = {}
guides_to_programs.update(dict.fromkeys(G1_CYCLE, "G1 cell cycle arrest"))
guides_to_programs.update(dict.fromkeys(ERYTHROID, "Erythroid"))
guides_to_programs.update(dict.fromkeys(PIONEER_FACTORS, "Pioneer factors"))
guides_to_programs.update(
dict.fromkeys(GRANULOCYTE_APOPTOSIS, "Granulocyte/apoptosis")
)
guides_to_programs.update(dict.fromkeys(PRO_GROWTH, "Pro-growth"))
guides_to_programs.update(dict.fromkeys(MEGAKARYOCYTE, "Megakaryocyte"))
guides_to_programs.update(dict.fromkeys(["ctrl"], "Ctrl"))
adata.obs["gene_program"] = [guides_to_programs[x] if x in guides_to_programs else "N/A" for x in adata.obs["guide_merged"]]
adata.obs["good_coverage"] = adata.obs["good_coverage"].astype(bool)
# -
adata.write('Norman_2019_raw.h5ad')
| datasets/Norman_2019_curation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Sampling of Dataset : 1250
import pandas as pd
data = data.drop("Unnamed: 110", axis=1)
data = data.drop("Unnamed: 111", axis=1)
list_vocabulary=list(set_vocabulary)
list_vocabulary
# +
#moving samples to a new location
# importing os module
import os
# importing shutil module
import shutil
root_dir_aa="/home/jimitcard/Desktop/OMR/dataset/primusCalvoRizoAppliedSciences2018/package_aa/"
destination="/home/jimitcard/Desktop/OMR/sample_1250"
for sample in sample_list:
#source
source=root_dir_aa+sample
dest = shutil.move(source, destination)
print(os.listdir(destination))
# -
# now remove the irrelevant vocabulary
frequency_of_notes=sample_1250.sum(numeric_only=True, axis = 0, skipna = True)
sorted_frequency=frequency_of_notes.sort_values(axis=0, ascending=False, inplace=False, kind='quicksort', na_position='last')
sorted_frequency
len(sample_list)
dropnote='note-E4_eighth' #note-E4_eighth 142
drop_list=[]
for sample in sample_list:
if sample_1250.loc[sample,dropnote]>=1:
drop_list.append(sample)
len(drop_list)
sample_1250=sample_1250.drop(drop_list)
sample_list=sample_1250.index
sample_1250=sample_1250.drop(dropnote, axis=1)
sample_1250.shape
sample_1250.index
sample_1250=sample_1250.drop('000100497-1_1_1')
sample_list=sample_1250.index
sample_1250.shape
sorted_frequency.describe()
# +
#moving samples to a new location
# importing os module
import os
# importing shutil module
import shutil
root_dir_aa="/home/jimitcard/Desktop/OMR/sample_1250/"
destination="/home/jimitcard/Desktop/OMR/sampled_1250"
for sample in sample_list:
#source
source=root_dir_aa+sample
dest = shutil.move(source, destination)
print(os.listdir(destination))
# -
# # Visualization of sampled dataset
# +
# we are using the inline backend
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
print ('Matplotlib version: ', mpl.__version__) # >= 2.0.0
# -
# Line Plot
sorted_frequency.head()
# +
sorted_frequency.plot(kind='line')
plt.title('Frequency Distribution of Notes in Sampled Dataset')
plt.ylabel('Frequency')
plt.xlabel('Notes')
plt.xticks([])
plt.savefig('frequencyVsNotes.png')
#plt.plot([5,0], [5,4500], 'k-')
#plt.yscale('log')
#median
#10-90
plt.show()
# -
# ## Bins
# +
sorted_frequency.plot(kind='line', label='')
plt.title('Bins formation with first and last 10 Percentile')
plt.ylabel('Frequency')
plt.xlabel('Notes arranged in ascending order of frequency of occurrence')
plt.xticks([])
plt.savefig('bins.png')
#plt.plot([5,0], [5,4500], 'k-')
plt.yscale('log')
#plt.plot([30], [1000], marker='o', markersize=3, color="red")
plt.plot((30,30),(50,4000),label="Median",linestyle='dashed')
plt.plot((6,6),(50,4000),label="HF 10 %tile",linestyle='dashed')
plt.plot((54,54),(50,4000),label="LF 10 %tile",linestyle='dashed')
plt.legend(bbox_to_anchor=(1.05, 1), loc='upper left', borderaxespad=0.)
plt.savefig('bins.png')
#median
#10-90
plt.show()
# +
SMALL_SIZE = 14
MEDIUM_SIZE = 9
BIGGER_SIZE = 12
plt.rc('font', size=SMALL_SIZE) # controls default text sizes
plt.rc('axes', titlesize=SMALL_SIZE) # fontsize of the axes title
plt.rc('axes', labelsize=BIGGER_SIZE) # fontsize of the x and y labels
plt.rc('xtick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
plt.rc('ytick', labelsize=MEDIUM_SIZE) # fontsize of the tick labels
plt.rc('legend', fontsize=SMALL_SIZE) # legend fontsize
plt.rc('figure', titlesize=BIGGER_SIZE) # fontsize of the figure title
# -
# ## Box Plot
# +
red_square = dict(markerfacecolor='r', marker='^')
fig6, ax6 = plt.subplots()
ax6.set_title('Outliers representing LF and HF samples')
ax6.boxplot(sorted_frequency, flierprops=red_square, vert=False, whis=0.1)
plt.yticks([])
plt.xscale('log')
#plt.xlabel('Frequency of Symbol')
#plt.savefig('boxPlot.png')
#exponential scale?
#editing
# -
# ## Waffle Chart
# +
i=0
propotion_hf=0
propotion_normal=0
propotion_lf=0
for frequency in sorted_frequency:
if i<=5:
propotion_hf=propotion_hf+frequency
elif i<=53:
propotion_normal=propotion_normal+frequency
else:
propotion_lf=propotion_lf+frequency
i=i+1
propotion_hf=float(propotion_hf/6.0)
propotion_normal=float(propotion_normal/48.0)
propotion_lf=float(propotion_lf/6.0)
print(str(propotion_hf)+' '+str(propotion_normal)+" "+str(propotion_lf))
# -
total=propotion_hf+propotion_normal+propotion_lf
propotion_hf=float(propotion_hf/total)
propotion_normal=float(propotion_normal/total)
propotion_lf=float(propotion_lf/total)
print(str(propotion_hf)+' '+str(propotion_normal)+" "+str(propotion_lf))
# +
width = 40 # width of chart
height = 10 # height of chart
total_num_tiles = width * height # total number of tiles
print ('Total number of tiles is ', total_num_tiles)
# -
propotion_hf=round(propotion_hf*400)
propotion_normal=round(propotion_normal*400)
propotion_lf=round(propotion_lf*400)
print(str(propotion_hf)+' '+str(propotion_normal)+" "+str(propotion_lf))
# +
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
# Let's make a default data frame with catagories and values.
df = pd.DataFrame({ 'catagories': ['HF', 'Normal', 'LF'],
'values': [323, 64, 13] })
# Now, we define a desired height and width.
waffle_plot_width = 40
waffle_plot_height =10
classes = df['catagories']
values = df['values']
def waffle_plot(classes, values, height, width, colormap):
# Compute the portion of the total assigned to each class.
class_portion = [float(v)/sum(values) for v in values]
# Compute the number of tiles for each catagories.
total_tiles = width * height
tiles_per_class = [round(p*total_tiles) for p in class_portion]
# Make a dummy matrix for use in plotting.
plot_matrix = np.zeros((height, width))
# Popoulate the dummy matrix with integer values.
class_index = 0
tile_index = 0
# Iterate over each tile.
for col in range(waffle_plot_width):
for row in range(height):
tile_index += 1
# If the number of tiles populated is sufficient for this class...
if tile_index > sum(tiles_per_class[0:class_index]):
# ...increment to the next class.
class_index += 1
# Set the class value to an integer, which increases with class.
plot_matrix[row, col] = class_index
# Create a new figure.
fig = plt.figure()
# Using matshow solves your "non-square" problem.
plt.matshow(plot_matrix, cmap=colormap)
plt.colorbar()
# Get the axis.
ax = plt.gca()
# Minor ticks
ax.set_xticks(np.arange(-.5, (width), 1), minor=True);
ax.set_yticks(np.arange(-.5, (height), 1), minor=True);
# Gridlines based on minor ticks
ax.grid(which='minor', color='w', linestyle='-', linewidth=2)
# Manually constructing a legend solves your "catagorical" problem.
legend_handles = []
for i, c in enumerate(classes):
lable_str = c + " (" + str(values[i]) + ")"
color_val = colormap(float(i+1)/len(classes))
legend_handles.append(mpatches.Patch(color=color_val, label=lable_str))
# Add the legend. Still a bit of work to do here, to perfect centering.
plt.legend(handles=legend_handles, loc=1, ncol=len(classes),
bbox_to_anchor=(0., -0.1, 0.95, .10))
plt.xticks([])
plt.yticks([])
plt.title('Domination of different type of samples during Training')
# Call the plotting function.
waffle_plot(classes, values, waffle_plot_height, waffle_plot_width,
plt.cm.coolwarm)
plt.savefig('waffleChart.png')
# -
# ## Word Cloud
wordCloud = pd.DataFrame({'word':sorted_frequency.index ,
'count': counts})
counts=[]
for frequency in sorted_frequency:
counts.append(frequency)
wordCloud
# +
data = dict(zip(wordCloud['word'].tolist(), wordCloud['count'].tolist()))
print(data)
# +
from wordcloud import WordCloud
wc = WordCloud(width=800, height=400, max_words=200,background_color='white').generate_from_frequencies(data)
# +
import matplotlib.pyplot as plt
plt.figure(figsize=(15, 15))
plt.imshow(wc, interpolation='bilinear')
plt.axis('off')
plt.savefig('wordCloud.png')
plt.show()
| scripts/sampling_250_for_imbalance_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Bi-directional LSTM with average pooling
#
# Author: <NAME>
#
# November 2019
#
# We make use of a bi-directional LSTM networks that extends the modelling capabilities of the vanilla LSTM. This approach is similar to that of InferSent (Conneau et al. 2017) where the authors combine bi-directional LSTM models with pooling layers to produce high-quality sentence embeddings. In addition to InferSent, we attach a dense classification layer after the pooling layers.
import sys
import os
#sys.path.append(os.path.join(\"..\")) # path to source relative to current directory"
import numpy as np
import gensim
import preprocess_data
import pandas as pd
import tensorflow as tf
physical_devices = tf.config.experimental.list_physical_devices("GPU")
tf.config.experimental.set_memory_growth(physical_devices[0], True)
from tensorflow.keras.models import Model
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import GlobalAveragePooling1D, GlobalMaxPooling1D
from tensorflow.keras.layers import Dense, Dropout, Embedding, LSTM, Bidirectional, TimeDistributed, Input, Flatten, AdditiveAttention
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
data = pd.read_csv('dataset_7B', delimiter = ';', engine = 'python')
data_text = data.loc[data['set'] == 'Train'][['helpdesk_question']]
number_of_classes = data.loc[data['set'] == 'Train']['helpdesk_reply'].value_counts().shape[0]
data = data[['helpdesk_question', 'helpdesk_reply', 'set', 'low_resource']]
responses = pd.DataFrame(data.loc[data['set'] == 'Train']['helpdesk_reply'].value_counts()).reset_index()
responses['reply'] = responses['index']
responses['index'] = responses.index
responses = dict(responses.set_index('reply')['index'])
len(responses)
data_text['index'] = data_text.index
documents = data_text
dictionary = preprocess_data.create_dictionary(data_text, 1, 0.25, 95000) #our entire vocabulary
# +
df_train = data.loc[data['set'] == 'Train']
df_train = df_train.reset_index()[['helpdesk_question', 'helpdesk_reply']]
df_valid = data.loc[data['set'] == 'Valid']
df_valid = df_valid.reset_index()[['helpdesk_question', 'helpdesk_reply']]
df_test = data.loc[data['set'] == 'Test']
df_test = df_test.reset_index()[['helpdesk_question', 'helpdesk_reply']]
df_LR = data.loc[(data['set'] == 'Test') & (data['low_resource'] == 'True') ]
df_LR = df_LR.reset_index()[['helpdesk_question', 'helpdesk_reply']]
# -
df_train.shape
unique_words = dictionary
len(unique_words) + 1
max_length = 30
min_token_length = 0
word_to_id, id_to_word = preprocess_data.create_lookup_tables(unique_words)
# #### Transforming the input sentence into a sequence of word IDs
# +
train_x_word_ids = []
for question in df_train['helpdesk_question'].apply(preprocess_data.preprocess_question,
args = [unique_words, min_token_length]):
word_ids = preprocess_data.transform_sequence_to_word_ids(question, word_to_id)
train_x_word_ids.append(np.array(word_ids, dtype = float))
train_x_word_ids = np.stack(train_x_word_ids)
print(train_x_word_ids.shape)
val_x_word_ids = []
for question in data['helpdesk_question'].loc[data['set'] == 'Valid'].apply(preprocess_data.preprocess_question,
args = [unique_words, min_token_length]):
word_ids = preprocess_data.transform_sequence_to_word_ids(question, word_to_id)
val_x_word_ids.append(np.array(word_ids, dtype = float))
val_x_word_ids = np.stack(val_x_word_ids)
test_x_word_ids = []
for question in data['helpdesk_question'].loc[data['set'] == 'Test'].apply(preprocess_data.preprocess_question,
args = [unique_words, min_token_length]):
word_ids = preprocess_data.transform_sequence_to_word_ids(question, word_to_id)
test_x_word_ids.append(np.array(word_ids, dtype = float))
test_x_word_ids = np.stack(test_x_word_ids)
LR_x_word_ids = []
for question in data['helpdesk_question'].loc[(data['set'] == 'Test') &
(data['low_resource'] == 'True')].apply(preprocess_data.preprocess_question,
args = [unique_words, min_token_length]):
word_ids = preprocess_data.transform_sequence_to_word_ids(question, word_to_id)
LR_x_word_ids.append(np.array(word_ids, dtype = float))
LR_x_word_ids = np.stack(LR_x_word_ids)
# -
def get_dummies(reply, all_responses):
""" Constructs a one-hot vector for replies
Args:
reply: query item
all_responses: dict containing all the template responses with their corresponding IDs
Return:
a one-hot vector where the corresponding ID of the reply is the one-hot index
"""
Y = np.zeros(len(all_responses), dtype = int)
Y[all_responses[reply]] += 1
return Y
train_y = np.array(list(df_train['helpdesk_reply'].apply(get_dummies, args = [responses])))
valid_y = np.array(list(df_valid['helpdesk_reply'].apply(get_dummies, args = [responses])))
test_y = np.array(list(df_test['helpdesk_reply'].apply(get_dummies, args = [responses])))
LR_y = np.array(list(df_LR['helpdesk_reply'].apply(get_dummies, args = [responses])))
train_x_word_ids = train_x_word_ids.reshape(train_x_word_ids.shape[:-1])
val_x_word_ids = val_x_word_ids.reshape(val_x_word_ids.shape[:-1])
test_x_word_ids = test_x_word_ids.reshape(test_x_word_ids.shape[:-1])
LR_x_word_ids = LR_x_word_ids.reshape(LR_x_word_ids.shape[:-1])
# #### Transform vectors where the input sentence yields a sequence of length 0
train_zero_vectors = np.where(train_x_word_ids.sum(axis = 1) == 0.0)[0]
for t in range(train_zero_vectors.shape[0]):
train_x_word_ids[train_zero_vectors[t]][0] += 1
val_zero_vectors = np.where(val_x_word_ids.sum(axis = 1) == 0.0)[0]
for t in range(val_zero_vectors.shape[0]):
val_x_word_ids[val_zero_vectors[t]][0] += 1
# #### Bi-directional LSTM with average pooling
#
# The network consists of an embedding layer, followed by a dropout layer. This is followed by an bi-directional LSTM layer that outputs a variable-length sequence of embedding vectors. To construct a single sentence embedding from the sequence we use average pooling. The sentence embedding is then fed to a classification layer. We train with a dropout rate of 0.5 and batch size of 32. During training we use early stopping and Adadelta as our optimization algorithm. This network has an embedding of size 300 and 512 hidden units in the biLSTM network.
def bilstm_avg_pooling_network(max_features, input_length=30, embed_dim=100, lstm_units=512):
""" Constructs a bi-directional LSTM network with average pooling
Args:
max_features: size of vocabulary
input_length: length of input sequence
embed_dim: dimension of the embedding vector
lstm_units: number of hidden units in biLSTM
Returns:
An biLSTM model
"""
inputs = Input(shape=(input_length, ))
x = Embedding(max_features, output_dim=embed_dim, input_length=input_length, mask_zero=True)(inputs)
x = (Dropout(rate = 0.5))(x)
x = Bidirectional(LSTM(lstm_units, activation = 'tanh', return_sequences=True,
dropout=0.25, recurrent_dropout=0.5))(x)
x = GlobalAveragePooling1D()(x)
outputs = Dense(89, activation='softmax')(x)
return Model(inputs=inputs, outputs=outputs)
max_features = len(unique_words) + 1
model = bilstm_avg_pooling_network(max_features, embed_dim=300, input_length=30, lstm_units = 512)
model.summary()
# ### Training
es = EarlyStopping(monitor='val_accuracy', verbose=1, restore_best_weights=True, patience=10)
model.compile(loss='categorical_crossentropy', #optimizer = 'sgd',
optimizer=tf.keras.optimizers.Adadelta(learning_rate=0.5, rho=0.95),
metrics=['accuracy'])
model.fit(train_x_word_ids, train_y,
batch_size=32,
epochs=500,
callbacks=[es],
validation_data=[val_x_word_ids, valid_y])
# ### Test score
def classifier_score_top_1(word_ids, y_true, model):
""" Computes top-1 classification accuracy for model.
Args:
word_ids: matrix where each row is
y_true: true labels
model: trained model
Returns:
None
"""
score = 0
probs = model.predict(word_ids)
for i in range(word_ids.shape[0]):
if y_true[i].argmax() == np.argsort(probs[i])[-1]:
score += 1
print("Overall Accuracy:", score/word_ids.shape[0])
classifier_score_top_1(test_x_word_ids, test_y, model)
# ### LR test score
classifier_score_top_1(LR_x_word_ids, LR_y, model)
# ### Top-5 accuracy
def classifier_score_top_5(word_ids, y_true, model):
""" Computes top-5 classification accuracy for model.
Args:
word_ids: matrix where each row is
y_true: true labels
model: trained model
Returns:
None
"""
score = 0
probs = model.predict(word_ids)
for i in range(word_ids.shape[0]):
if y_true[i].argmax() in np.argsort(probs[i])[-5:]:
score += 1
print("Overall Accuracy:", score/word_ids.shape[0])
classifier_score_top_5(test_x_word_ids, test_y, model)
classifier_score_top_5(LR_x_word_ids, LR_y, model)
| experiments/bilstm_512_avg_pooling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from kanren import run, fact, eq, Relation, var
coastal = Relation()
adjacent = Relation()
file_coastal = 'coastal_states.txt'
with open(file_coastal, 'r') as f:
line = f.read()
coastal_states = line.split(',')
for state in coastal_states:
fact(coastal, state)
file_adjacent = 'adjacent_states.txt'
with open(file_adjacent, 'r') as f:
adjlist = [line.strip().split(',') for line in f
if line and line[0].isalpha()]
for L in adjlist:
head, tail = L[0], L[1:]
for state in tail:
fact(adjacent, head, state)
# -
x = var()
output = run(0, x, adjacent('Nevada', 'Louisiana'))
print('Yes' if len(output) else 'No')
output = run(0, x, adjacent('Oregon', x))
for item in output:
print(item)
output = run(0, x, adjacent('Mississippi', x), coastal(x))
for item in output:
print(item)
y= var()
output = run(7, x, coastal(y), adjacent(x, y))
for item in output:
print(item)
output = run(0, x, adjacent('Arkansas', x), adjacent('Kentucky', x))
for item in output:
print(item)
| artificial-intelligence-with-python-ja-master/Chapter 6/states.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 64-bit
# language: python
# name: python37664bitffa1473ee06547deb7ef2d0693223843
# ---
# +
# project configurations
import sqlalchemy as sa
import numpy as np
import pandas as pd
from pandas import ExcelWriter
from pandas import ExcelFile
import datetime
import os
import zipfile
import uuid
import matplotlib
import seaborn as sns
from os.path import basename
from datetime import datetime, date, time, timedelta
client_pg = sa.create_engine('postgresql://localhost:5432/postgres')
# +
# question number one part two
number_one_part_two = """
WITH processed_transactions AS (
(select u.id, ((t.amount / power(10, cd.exponent)) * fx.rate) AS amount, t.created_date, t.type, t.state
from users u
join transactions t on t.user_id = u.id
JOIN fx_rates fx ON (fx.ccy = t.currency AND fx.base_ccy = 'USD')
JOIN currency_details cd ON cd.currency = t.currency
and t.currency != 'USD')
union
(SELECT u.id, (t.amount / power(10, cd.exponent)) as amount, t.created_date, t.type, t.state
from users u
join transactions t on t.user_id = u.id
join currency_details cd ON cd.currency = t.currency
and t.currency = 'USD')
order by created_date asc
)
select * from
processed_transactions pt
where pt.amount > 10
and pt.type = 'CARD_PAYMENT'
and pt.state = 'COMPLETED'
and pt.id IN (select distinct on (pt.id) pt.id from processed_transactions pt);
"""
df_first = pd.read_sql(number_one_part_two, client_pg)
df_first.head()
| Financial-Crime-Analysis/Challenge One.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic example
from quickdef import quickdef, quickargs
calib = dict(
a=0.1,
b=0.5,
c=0.1
)
# +
@quickdef(calib)
def fun(x,y):
return x**2*a + x*b + c
# gets translated into
# def fun(x,y,calib):
# a = calib['a']
# b = calib['b']
# c = calib['c']
# return x**2*a + x*b + c
fun(0,1, calib)
# -
# # Same with vector arguments
# +
from collections import OrderedDict as odict
calib = odict(
a=0.1,
b=0.5,
c=0.1
)
# +
@quickdef(calib, sub_type='array')
def fun(x,y):
return x**2*a + x*b + c
# gets translated into
# def fun(x,y,calib):
# a = calib[0]
# b = calib[1]
# c = calib[2]
# return x**2*a + x*b + c
calib_vec = [e for e in calib.values()]
fun(0,1, calib_vec)
# -
# # Dolo functions
from collections import OrderedDict
import quickdef
# +
d = OrderedDict(
states = ['α','β'],
controls = ['x', 'y']
)
@quickargs(d)
def fun():
h = α*x + β*y
return h
# gets translated into
# def fun(states, controls):
# α = states[0]
# β = states[1]
# x = controls[0]
# y = controls[1]
# h = α*x + β*y
# return h
import numpy
v_states = numpy.array([1.0, 2.0])
v_controls = numpy.array([1.0, 1.0])
fun(v_states,v_controls)
| experiments/quickdef/Example (python).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Final NLP project (University of Guilan)
# In this repository, we implemented a Statistical NLP model to predict news agency, news tags, etc as final project of NLP course in university of Guilan.
# # Contents
# * Libraries and Constants
# * Importing Data
# * Preprocessing
# * Creating Model for the First Task
# * Creating Model for the Second Task
# * Creating Model for the Third Task
# +
from __future__ import unicode_literals
import json
import os
import re
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy.optimize as opt
from functools import reduce
from operator import add
from hazm import *
from utils.preprocessing import *
from utils.models import *
from copy import deepcopy
# +
# Data root path
data_root = 'data'
fars_file = 'farsnews.jsonl'
asriran_file = 'asriran.jsonl'
# Dataset dataframe column names
keys = None
# News headline tags
not_valid_labels = None
# News agencies
news_agencies = None
# -
# # Importing Data
with open(os.path.join(data_root, asriran_file), encoding='utf-8') as jd:
asriran = [json.loads(line) for line in jd]
asriran = pd.DataFrame(asriran)
print('Number of Datapoints: {}'.format(len(asriran)))
with open(os.path.join(data_root, fars_file), encoding='utf-8') as jd:
fars = [json.loads(line) for line in jd]
fars = pd.DataFrame(fars)
print('Number of Datapoints: {}'.format(len(fars)))
# # Preprocessing
# Finding Valid Labels:
asr_labels = list(set(reduce(np.append, asriran.newsPathLinks.apply(lambda x: tuple(x.keys())))))
fars_labels = list(set(reduce(np.append, fars.newsPathLinks.apply(lambda x: list(x.keys())))))
set((list(asr_labels) + list(fars_labels)))
# Some labels are not valid so:
not_valid_labels = [
'دانلود',
'ساير حوزه ها',
'سایر حوزه ها',
'دیگر رسانه ها',
'نامشخص',
'پیامک',
'صفحه نخست',
'عصرايران دو',
]
valid_labels = list(filter(lambda x: x not in not_valid_labels, list(set((list(asr_labels) + list(fars_labels))))))
# Creating Documents & Labels:
asriran_tags = asriran['tags'].apply(lambda x: ' '.join(list(x.keys())))
fars_tags = fars['tags'].apply(lambda x: ' '.join(list(x.keys())))
title_count = 2
tag_count = 10
documents = np.append(asriran['body'] + ' ' + asriran['title'] * title_count + asriran_tags*tag_count,
fars['body'] + ' ' + fars['title'] * title_count + fars_tags*tag_count)
raw_labels = np.append(asriran.newsPathLinks.apply(lambda x: tuple(x.keys())),
fars.newsPathLinks.apply(lambda x: tuple(x.keys())))
org_labels = np.append( ['AsrIran'] * len(asriran), ['Fars'] * len(fars)) # For the third task
# Removing Documents which are emtpy:
none_zero_docs = list(map(lambda x: len(x) > 1, documents))
documents = documents[none_zero_docs]
raw_labels = cleans_labels(raw_labels[none_zero_docs], valid_labels)
org_labels = org_labels[none_zero_docs]
# Duplicating documents for each of their labels:
proc_documents, proc_labels = extend_labels(documents, raw_labels, valid_labels)
# Normalizing & Tokenizing & Removing Stopwords Documents:
normalizer = Normalizer()
word_filter = WordFilter()
documents = list(pd.Series(documents).apply(normalizer.normalize).apply(tokenize).apply(word_filter.filter_words))
proc_documents = list(proc_documents.apply(normalizer.normalize).apply(tokenize).apply(word_filter.filter_words))
# Replacing words with less than 2 occurances with unknown word
documents = make_unknown(documents)
proc_documents = make_unknown(proc_documents)
# Making documents one hot encoded
label_set, proc_labels = one_hot_encoder(proc_labels)
label_set_th, org_labels = one_hot_encoder(org_labels)
# Deviding document to train and test datasets:
x_train, y_train, x_test, y_test = train_test_split(proc_documents , proc_labels, train_size = 0.80, random_state=85)
x_train_th, y_train_th, x_test_th, y_test_th = train_test_split(documents , org_labels, train_size = 0.80, random_state=85)
# # Creating Model for the First Task
# Training:
nb = NaiveBayes()
nb.fit(x_train, y_train)
# Train Evaluation:
nb.evaluate(x_train, y_train, label_set=label_set)
# Test Evaluation:
nb.evaluate(x_test, y_test, label_set=label_set)
# # Creating Model for the Second Task
t = []
for i, raw_label in enumerate(raw_labels):
l = []
for j, label in enumerate(raw_label):
l.append(np.argmax(label == label_set))
t.append(l)
nb.evaluate(documents, t, label_set, eval_type='multiple')
# # Creating Model for the Third Task
# Training:
nb_th = NaiveBayes()
nb_th.fit(x_train_th, y_train_th)
# Train Evaluation:
nb_th.evaluate(x_train_th, y_train_th, label_set_th)
# Test Evaluation:
nb_th.evaluate(x_test_th, y_test_th, label_set_th)
| Project Final Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="14W3B3KZJ7J_" outputId="29405fdf-dc74-44da-954e-a3673194835d"
# Instalação do Plotly
# !pip install Plotly==4.12
# Instalação do Dash
# !pip install dash
# !pip install dash-html-components
# !pip install dash-core-components
# !pip install dash-table
# + id="czJEHl_sUiuF"
# Pacotes Python Pandas e Plotly
import numpy as np
import pandas as pd
import plotly.express as px
# + id="Stjha8niUkUI"
# Pacotes para criação de processos para suportar o serviço HTTP externo do Ngrok
import os.path
import sys, json
import requests
import subprocess
# Pacotes para o utilizar as propriedades de conexões HTTP
from requests.exceptions import RequestException
from requests.adapters import HTTPAdapter
from requests.packages.urllib3.util.retry import Retry
from collections import namedtuple
# + id="DFurrObpUno1"
# Definição da função de Download do Ngrok
def download_ngrok():
if not os.path.isfile('ngrok'):
# !wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# !unzip -o ngrok-stable-linux-amd64.zip
pass
# + id="lsbLmVqQUzkK"
# Criação de um Túnel entre uma aplicação no Colab e uma URL externa no Ngrok
# Função para pegar a referência nesse túnel
Response = namedtuple('Response', ['url', 'error'])
def get_tunnel():
try:
Tunnel = subprocess.Popen(['./ngrok','http','8050'])
session = requests.Session()
retry = Retry(connect=3, backoff_factor=0.5)
adapter = HTTPAdapter(max_retries=retry)
session.mount('http://', adapter)
res = session.get('http://localhost:4040/api/tunnels')
res.raise_for_status()
tunnel_str = res.text
tunnel_cfg = json.loads(tunnel_str)
tunnel_url = tunnel_cfg['tunnels'][0]['public_url']
return Response(url=tunnel_url, error=None)
except RequestException as e:
return Response(url=None, error=str(e))
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="pfLsy2swU7vM" outputId="8b9e9b35-e1f0-49ee-fc36-6ee5a9a918c0"
df = px.data.gapminder()
fig1 = px.scatter(df, x="gdpPercap", y="lifeExp", animation_frame="year", animation_group="country",
size="pop", color="continent", hover_name="country",
log_x=True, size_max=55, range_x=[100,100000], range_y=[25,90])
fig1.show()
# + colab={"base_uri": "https://localhost:8080/"} id="iuixcjPbVD21" outputId="0591db70-3fe3-40a1-cf0c-0485a490aff2"
# Geração do programa exemplo_dash_1.py
# %%writefile covid-vacine-progression-analysis.py
import dash
import dash_core_components as dcc
import dash_html_components as html
import pandas as pd
external_stylesheets = ['https://codepen.io/chriddyp/pen/bWLwgP.css']
app = dash.Dash(__name__, external_stylesheets=external_stylesheets)
df = pd.read_csv('https://plotly.github.io/datasets/country_indicators.csv')
available_indicators = df['Indicator Name'].unique()
app.layout = html.Div([
html.Div([
html.Div([
dcc.Dropdown(
id='crossfilter-xaxis-column',
options=[{'label': i, 'value': i} for i in available_indicators],
value='Fertility rate, total (births per woman)'
),
dcc.RadioItems(
id='crossfilter-xaxis-type',
options=[{'label': i, 'value': i} for i in ['Linear', 'Log']],
value='Linear',
labelStyle={'display': 'inline-block'}
)
],
style={'width': '49%', 'display': 'inline-block'}),
html.Div([
dcc.Dropdown(
id='crossfilter-yaxis-column',
options=[{'label': i, 'value': i} for i in available_indicators],
value='Life expectancy at birth, total (years)'
),
dcc.RadioItems(
id='crossfilter-yaxis-type',
options=[{'label': i, 'value': i} for i in ['Linear', 'Log']],
value='Linear',
labelStyle={'display': 'inline-block'}
)
], style={'width': '49%', 'float': 'right', 'display': 'inline-block'})
], style={
'borderBottom': 'thin lightgrey solid',
'backgroundColor': 'rgb(250, 250, 250)',
'padding': '10px 5px'
}),
html.Div([
dcc.Graph(
id='crossfilter-indicator-scatter',
hoverData={'points': [{'customdata': 'Japan'}]}
)
], style={'width': '49%', 'display': 'inline-block', 'padding': '0 20'}),
html.Div([
dcc.Graph(id='x-time-series'),
dcc.Graph(id='y-time-series'),
], style={'display': 'inline-block', 'width': '49%'}),
html.Div(dcc.Slider(
id='crossfilter-year--slider',
min=df['Year'].min(),
max=df['Year'].max(),
value=df['Year'].max(),
marks={str(year): str(year) for year in df['Year'].unique()},
step=None
), style={'width': '49%', 'padding': '0px 20px 20px 20px'})
])
@app.callback(
dash.dependencies.Output('crossfilter-indicator-scatter', 'figure'),
[dash.dependencies.Input('crossfilter-xaxis-column', 'value'),
dash.dependencies.Input('crossfilter-yaxis-column', 'value'),
dash.dependencies.Input('crossfilter-xaxis-type', 'value'),
dash.dependencies.Input('crossfilter-yaxis-type', 'value'),
dash.dependencies.Input('crossfilter-year--slider', 'value')])
def update_graph(xaxis_column_name, yaxis_column_name,
xaxis_type, yaxis_type,
year_value):
dff = df[df['Year'] == year_value]
return {
'data': [dict(
x=dff[dff['Indicator Name'] == xaxis_column_name]['Value'],
y=dff[dff['Indicator Name'] == yaxis_column_name]['Value'],
text=dff[dff['Indicator Name'] == yaxis_column_name]['Country Name'],
customdata=dff[dff['Indicator Name'] == yaxis_column_name]['Country Name'],
mode='markers',
marker={
'size': 15,
'opacity': 0.5,
'line': {'width': 0.5, 'color': 'white'}
}
)],
'layout': dict(
xaxis={
'title': xaxis_column_name,
'type': 'linear' if xaxis_type == 'Linear' else 'log'
},
yaxis={
'title': yaxis_column_name,
'type': 'linear' if yaxis_type == 'Linear' else 'log'
},
margin={'l': 40, 'b': 30, 't': 10, 'r': 0},
height=450,
hovermode='closest'
)
}
def create_time_series(dff, axis_type, title):
return {
'data': [dict(
x=dff['Year'],
y=dff['Value'],
mode='lines+markers'
)],
'layout': {
'height': 225,
'margin': {'l': 20, 'b': 30, 'r': 10, 't': 10},
'annotations': [{
'x': 0, 'y': 0.85, 'xanchor': 'left', 'yanchor': 'bottom',
'xref': 'paper', 'yref': 'paper', 'showarrow': False,
'align': 'left', 'bgcolor': 'rgba(255, 255, 255, 0.5)',
'text': title
}],
'yaxis': {'type': 'linear' if axis_type == 'Linear' else 'log'},
'xaxis': {'showgrid': False}
}
}
@app.callback(
dash.dependencies.Output('x-time-series', 'figure'),
[dash.dependencies.Input('crossfilter-indicator-scatter', 'hoverData'),
dash.dependencies.Input('crossfilter-xaxis-column', 'value'),
dash.dependencies.Input('crossfilter-xaxis-type', 'value')])
def update_y_timeseries(hoverData, xaxis_column_name, axis_type):
country_name = hoverData['points'][0]['customdata']
dff = df[df['Country Name'] == country_name]
dff = dff[dff['Indicator Name'] == xaxis_column_name]
title = '<b>{}</b><br>{}'.format(country_name, xaxis_column_name)
return create_time_series(dff, axis_type, title)
@app.callback(
dash.dependencies.Output('y-time-series', 'figure'),
[dash.dependencies.Input('crossfilter-indicator-scatter', 'hoverData'),
dash.dependencies.Input('crossfilter-yaxis-column', 'value'),
dash.dependencies.Input('crossfilter-yaxis-type', 'value')])
def update_x_timeseries(hoverData, yaxis_column_name, axis_type):
dff = df[df['Country Name'] == hoverData['points'][0]['customdata']]
dff = dff[dff['Indicator Name'] == yaxis_column_name]
return create_time_series(dff, axis_type, yaxis_column_name)
if __name__ == '__main__':
app.run_server(debug=True)
# + id="4OjoLRwKVRDu"
download_ngrok()
# + colab={"base_uri": "https://localhost:8080/"} id="Jgv1tV84Vbsk" outputId="c78538f1-edab-4cda-a8a4-a46a2e53dd8b"
retorno = get_tunnel()
print(retorno) # Para podermos saber o endereço da URL da Aplicação
# !python covid-vacine-progression-analysis.py
| src/install_dash_plotly_ngrok.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Find the lowest common ancestor in a binary tree.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Is this a binary search tree?
# * No
# * Can we assume the two nodes are in the tree?
# * No
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# <pre>
# _10_
# / \
# 5 9
# / \ / \
# 12 3 18 20
# / \ /
# 1 8 40
# </pre>
#
# * 0, 5 -> None
# * 5, 0 -> None
# * 1, 8 -> 3
# * 12, 8 -> 5
# * 12, 40 -> 10
# * 9, 20 -> 9
# * 3, 5 -> 5
# ## Algorithm
#
# Refer to the [Solution Notebook](http://nbviewer.jupyter.org/github/donnemartin/interactive-coding-challenges/blob/master/graphs_trees/tree_lca/tree_lca_solution.ipynb). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
class Node(object):
def __init__(self, key, left=None, right=None):
self.key = key
self.left = left
self.right = right
def __repr__(self):
return str(self.key)
class BinaryTree(object):
def lca(self, root, node1, node2):
# TODO: Implement me
pass
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_lca.py
import unittest
class TestLowestCommonAncestor(unittest.TestCase):
def test_lca(self):
node10 = Node(10)
node5 = Node(5)
node12 = Node(12)
node3 = Node(3)
node1 = Node(1)
node8 = Node(8)
node9 = Node(9)
node18 = Node(18)
node20 = Node(20)
node40 = Node(40)
node3.left = node1
node3.right = node8
node5.left = node12
node5.right = node3
node20.left = node40
node9.left = node18
node9.right = node20
node10.left = node5
node10.right = node9
root = node10
node0 = Node(0)
binary_tree = BinaryTree()
self.assertEqual(binary_tree.lca(root, node0, node5), None)
self.assertEqual(binary_tree.lca(root, node5, node0), None)
self.assertEqual(binary_tree.lca(root, node1, node8), node3)
self.assertEqual(binary_tree.lca(root, node12, node8), node5)
self.assertEqual(binary_tree.lca(root, node12, node40), node10)
self.assertEqual(binary_tree.lca(root, node9, node20), node9)
self.assertEqual(binary_tree.lca(root, node3, node5), node5)
print('Success: test_lca')
def main():
test = TestLowestCommonAncestor()
test.test_lca()
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook]() for a discussion on algorithms and code solutions.
| graphs_trees/tree_lca/tree_lca_challenge.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
# -
# ## 1. Array and Slicing
# ### Axis: selected by row/column(axis=0:column, axis=1:row)
arr=np.array([[0,1,2],[3,4,5],[6,7,8]])
print(arr)
# +
#rank:number of dimensions of array
print(arr.ndim)
#shape
print(arr.shape)# 3 x 3
#slicing array
print(arr[2:4])# 6,7,8
print(arr[:2])#
# -
# ## 2. Matmul, Multiply
# +
matrix1=tf.constant([[1,2],[3,4]])
matrix2=tf.constant([[1],[2]])
#Matmul: multiply of matrix1 and matrix2
print("Matmul:")
print(tf.matmul(matrix1, matrix2).numpy())
#Multiply:
print("Multiply:")
print((matrix1 * matrix2).numpy())#this output came out because of broadcasting
# -
# ## 3. Broadcasting
# :make calculation of different-shaped matrix available
# +
mat1=tf.constant([[1,2]])
mat2=tf.constant([[3,4]])
print("Multiply:")#when mat1,mat2 is same shape
print((mat1+mat2).numpy())
mat1=tf.constant([[1,2]])
mat2=tf.constant([[3]])
print("Broadcasting:")#when mat1,mat2 has different shape
print((mat1+mat2).numpy())#broadcasting use [3,3] shape to calculate
# -
# ## 4-1.Reduce Mean
# :return mean of matrix selected by axis
# +
x=tf.constant([[1.,2.],[3.,4.]])
#reduce_mean(): return minimum values of each axis
print(tf.reduce_mean(x,axis=0).numpy())#output:[mean(1,3),mean(2,4)]
print(tf.reduce_mean(x,axis=1).numpy())#output:[mean(1,2),mean(3,4)]
print(tf.reduce_mean(x).numpy())#if axis is not entered, return just mean(x)
# -
# ## 4-2.Reduce Sum
# :return sum of matrix selected by axis
print(tf.reduce_sum(x , axis=0).numpy())
print(tf.reduce_sum(x , axis=1).numpy())
print(tf.reduce_sum(x , axis=-1).numpy()) # same axis = 1
# ## 5.Argmax
# :return position of maximum value by axis
# +
x=[[0,1,2],[2,1,0]]
print(tf.argmax(x,axis=0).numpy()) #position of (2,1,2)
print(tf.argmax(x,axis=1).numpy()) #position of (2,2)
print(tf.argmax(x,axis=-1).numpy()) #same as axis=1
# -
# ## 6.Reshape(reshape,squeeze,expand)
# :return reshaped array of input matrix
x = tf.constant([[[0, 1, 2],
[3, 4, 5]],
[[6,7,8],
[9,10,11]]
])
print(x.shape) #2*(2*3)
# +
print("Reshape:")
print(tf.reshape(x,shape=[-1,3]).numpy())
#If you want more, try these
#print(x.numpy()) #result expired after reshape function call
#print(tf.reshape(x,shape=[-1,1,3]).numpy()) #add 1 more rank
# +
print("Squeeze:") #simplify the unused rank
print(tf.squeeze([[[0],[1],[2]]]).numpy())
print("\nExpand_dims:") #expand dimension of matrix
print(tf.expand_dims([0,1,2], 1).numpy())
# -
# ## 7.One-Hot
# : express numbers by one-hot encoding.
# #### ex) [0,1,2,3] will be same as below.
# 0 -> [1,0,0,0]<br>
# 1 -> [0,1,0,0]<br>
# 2-> [0,0,1,0]<br>
# 3 -> [0,0,0,1]<br>
# 0 -> [1,0,0,0]<br>
print("example:")
arr = [0, 1, 2, 3]
print(tf.one_hot(arr, depth=4).numpy())
# ## 8.Type-casting
# :type casting array
arr = [1.8, 2.2, 4.1, 0.5]
print(tf.cast(arr,tf.int32).numpy()) #type-casting float array to int32
print(tf.cast([True,False,1==1,0==1],tf.int32).numpy()) #casting boolean to int32
# ## 9.Stack
# : return stack of input arrays by directed by axis
# +
x = [1, 4]
y = [2, 5]
z = [3, 6]
print("Stack_axis0:") #axis 0: stacked in a row
print(tf.stack([x,y,z],axis=0).numpy()) #axis=0
print("Stack_axis1:") #axis 1: stacked in a column
print(tf.stack([x,y,z],axis=1).numpy()) #axis=1
# -
# ## 10.Ones,Zeros
# : return new arrays initialized with 1(ones) or 0(zeros)
print("Ones:")
print(tf.ones([2,3]).numpy()) #create new array of ones in size(2*3)
print("Zeros:")
print(tf.zeros([3,4]).numpy()) #create new array of zeros in size(3*4)
# ## 11.Zip
# :return 2 arrays binded by index
# +
a = [1, 2, 3]
b = [4, 5, 6]
c = [7, 8, 9]
print("zip(a,b):")
for x,y in zip(a,b):
print(x,y)
print("zip(a,b,c):")
for x,y,z in zip(a,b,c):
print(x,y,z)
# -
| day8/day8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import matplotlib.pyplot as plt
import matplotlib.animation as animation
import cv2
from tqdm import tqdm
import random
# # %matplotlib inline
from keras.models import Model
from keras.layers import *
from keras.optimizers import *
from keras.callbacks import *
from keras import backend as K
from keras.layers.advanced_activations import *
from keras import metrics
from keras.applications import *
from keras.preprocessing import image
from keras.activations import *
import tensorflow as tf
from IPython.display import Image, display
from keras.utils.vis_utils import plot_model
# -
class HighwayDriving_Process:
def imread(self , name):
return cv2.cvtColor(cv2.imread(name) , cv2.COLOR_BGR2RGB)
def image2class(self , image):
img = np.zeros((image.shape[0],image.shape[1]))
for i,t in enumerate(self.color_list):
img[np.all(image == t[0] , axis = -1)] = i
return img
def class2onehot(self , classm):
return (np.arange(self.nb_classes) == classm[...,None]).astype(np.int8)
def class2image(self , classm):
img = np.zeros((classm.shape[0],classm.shape[1],3))
for i,t in enumerate(self.color_list):
img[classm == i] = np.array(t[0])
return img.astype(np.uint8)
def image_file(self , name):
return self.images_dir+'/'+name
def label_file(self , name):
n , _ , _ , s3 = name.split(self.split_char)
return self.labels_dir+'/'+self.split_char.join([n,self.label_suffix ,s3])
def __init__(self ,images_dir , labels_dir , classes = None):
'''
Original Video sampling by cameras was at 30 Hz
i.e 30 frames per second
This then turned into a 1Hz sampling for 1 frame
per second. The sampling is done for getting rid of
reptitive frames
for our experiment we create these kind of data set
organisation
1. frame => label (training segmentation network)
2. frame => future label (training sgemententation with optical flow)
i.e capture future intuition
Try with 1second , Try with 2second , Try with 3 second
3. frame_seq => frame_seq_label (training segmentation sequence model)
4. frame_seq => future_frame_seq_label (shifted by two or three frames)
using all these the frame vision field can be limited
'''
self.color_list = [ ([255, 255, 255] , 'Undefined'), ## The position is their class so void stays at 0
([0, 255, 255] , 'Sky'),
([128,128,128] , 'Road'),
([255,255,0] , 'Lane'),
([255,0,0] , 'Fence'),
([128, 0, 128] , 'Construction'),
([255, 0, 128] , 'Traffic sign'),
([0, 0, 128] , 'Car'),
([0,128,128] , 'Truck'),
([0,128,0] , 'vegetation'),
]
if classes is not None:
assert(isinstance(classes,list) ) ,"log:give a list of classes, check dataset or code"
color_list = [self.color_list[0]]
for t in self.color_list[1:]:
if t[1] in classes:
color_list.append(t)
self.color_list = color_list
self.nb_classes = len(self.color_list)
print("log: after filtering there are ",self.nb_classes," classes for training")
self.label_suffix = 'ColorLabel' ## dataset specific
self.split_char = '_' ## dataset specific
image_name_list = os.listdir(images_dir)
image_name_list.sort()
labels_name_list = os.listdir(labels_dir)
image_sequence_dict = {}
for name in image_name_list:
# print(name)
n , s1 , s2 , s3 = name.split(self.split_char)
image_sequence_dict.setdefault(n , [])
image_sequence_dict[n].append(self.split_char.join([s1,s2,s3]))
self.images_dir = images_dir
self.labels_dir = labels_dir
self.image_name_list = image_name_list
self.labels_name_list = labels_name_list
self.image_sequence_dict = image_sequence_dict
def frame2label_val_datagen(self , size = None , normalise = True):
'''
Using this makes sense when validation is True
'''
val_images = self.image_name_list[int(len(self.image_name_list)*0.9):]
random.shuffle(val_images)
batch_size = len(val_images)
itr = len(val_images) // batch_size
for i in range(itr+1):
IMAGES = []
LABELS = []
beg = i*batch_size
end = (i*batch_size + batch_size) if (i*batch_size + batch_size) < len(val_images) else -1
for img_file in val_images[beg:end]:
image = self.imread(self.image_file(img_file))
label = self.image2class(self.imread(self.label_file(img_file)))
if size != None:
assert( len(size) == 2) , "log: give (y , x) format size"
image = cv2.resize(image , size , interpolation = cv2.INTER_NEAREST)
label = cv2.resize(label , size , interpolation = cv2.INTER_NEAREST)
label = self.class2onehot(label)
IMAGES.append(image)
LABELS.append(label)
if len(IMAGES) == 0 or len(LABELS) == 0:
continue
IMAGES = np.array(IMAGES)
LABELS = np.array(LABELS)
if normalise:
IMAGES = IMAGES / 255
yield np.array(IMAGES),np.array(LABELS)
def frame2label_train_datagen(self ,size = None , batch_size = 16 , validation = True , normalise = True):
train_images = self.image_name_list
if validation == True:
train_images = self.image_name_list[:int(len(self.image_name_list)*0.9)]
random.shuffle(train_images)
itr = len(train_images) // batch_size
for i in range(itr+1):
IMAGES = []
LABELS = []
beg = i*batch_size
end = (i*batch_size + batch_size) if (i*batch_size + batch_size) < len(train_images) else -1
for img_file in train_images[beg:end]:
image = self.imread(self.image_file(img_file))
label = self.image2class(self.imread(self.label_file(img_file)))
if size != None:
assert( len(size) == 2) , "log: give (y , x) format size"
image = cv2.resize(image , size , interpolation = cv2.INTER_NEAREST)
label = cv2.resize(label , size , interpolation = cv2.INTER_NEAREST)
label = self.class2onehot(label)
IMAGES.append(image)
LABELS.append(label)
IMAGES = np.array(IMAGES)
LABELS = np.array(LABELS)
if normalise:
IMAGES = IMAGES / 255
yield np.array(IMAGES),np.array(LABELS)
def jaccard_index(y_pred , y):
intersection = np.logical_and(y, y_pred)
union = np.logical_or(y, y_pred)
iou_score = np.sum(intersection) / np.sum(union)
return iou_score
def get_class_weights(self , c=1.02):
_ , labels = next(self.frame2label_train_datagen(size = (256 , 256) , batch_size = len(self.image_name_list) ,
validation = False , normalise = False))
labels = np.argmax(labels , axis = -1)
print(labels.shape)
all_labels = labels.flatten()
each_class = np.bincount(all_labels, minlength=self.nb_classes)
prospensity_score = each_class / len(all_labels)
class_weights = 1 / (np.log(c + prospensity_score))
return class_weights
def frameSequence_train_datagen(self , size = None , batch_size = 16, time_steps = 4 , skip = 4, normalise = True , log=False):
assert(time_steps >= 2) , 'log: give more than or equal to two time steps'
assert(skip >= 1), 'log: give time skip >= 1'
for sequence in self.image_sequence_dict.keys():
if log:
print("log: sequence started:",sequence)
train_images = self.image_sequence_dict[sequence]
assert(len(train_images) <= batch_size+time_steps*skip), "log: batch_size + time_steps*skip exceeds max sequence length of video"
IMAGES = []
LABELS_FUTURE = []
train_batch = train_images
images_numpy = []
labels_numpy = []
for img_file in train_batch:
if size != None:
images_numpy.append(
cv2.resize(self.imread(
self.image_file(
self.split_char.join([sequence,img_file])))
,size , interpolation = cv2.INTER_NEAREST))
labels_numpy.append(self.class2onehot(
cv2.resize(self.image2class(
self.imread(self.label_file(
self.split_char.join([sequence,img_file]))))
, size , interpolation = cv2.INTER_NEAREST)))
else:
images_numpy.append(self.imread(
self.image_file(
self.split_char.join([sequence,img_file]))))
labels_numpy.append(self.class2onehot(self.image2class(
self.imread(self.label_file(
self.split_char.join([sequence,img_file]))))))
itr = (len(train_batch) - time_steps*skip)
for j in range(itr):
frames =[]
labels = []
for k in range(time_steps):
frames.append( images_numpy[j+k*skip] )
labels.append( labels_numpy[j+(k+1)*skip])
IMAGES.append(frames)
LABELS_FUTURE.append(labels)
if (j+1)%batch_size == 0 or (j+1) == itr:
IMAGES = np.array(IMAGES)
LABELS_FUTURE = np.array(LABELS_FUTURE)
if normalise:
IMAGES = IMAGES / 255
x = IMAGES
y = LABELS_FUTURE
IMAGES = []
LABELS_FUTURE = []
yield x , y
def frame2futurelabel_train_datagen(self , size=None , batch_size = 16, time_step = 3 , normalise = True):
assert(batch_size > time_step) ,'log: batch size must be greater than time step'
batch_size = batch_size+time_step
for sequence in self.image_sequence_dict.keys():
train_images = self.image_sequence_dict[sequence]
itr = len(train_images) // batch_size
for i in range(itr+1):
IMAGES = []
IMAGES_FUTURE = []
LABELS_FUTURE = []
beg = i*batch_size
end = (i*batch_size + batch_size) if (i*batch_size + batch_size) < len(train_images) else -1
train_batch = train_images[beg:end ]
for i in range(len(train_batch) - time_step):
frame1 = self.image_file(self.split_char.join([sequence,train_batch[i]]))
frame2 = self.image_file(self.split_char.join([sequence,train_batch[i+time_step]]))
label2 = self.label_file(self.split_char.join([sequence,train_batch[i+time_step]]))
frame1_image = self.imread(frame1)
frame2_image = self.imread(frame2)
label2_classim = self.image2class(self.imread(label2))
if size != None:
assert( len(size) == 2) , "log: give (y , x) format size"
frame1_image = cv2.resize(frame1_image , size , interpolation = cv2.INTER_NEAREST)
frame2_image = cv2.resize(frame2_image , size , interpolation = cv2.INTER_NEAREST)
label2_classim = cv2.resize(label2_classim , size , interpolation = cv2.INTER_NEAREST)
label2_onehote = self.class2onehot(label2_classim)
IMAGES.append(frame1_image)
IMAGES_FUTURE.append(frame2_image)
LABELS_FUTURE.append(label2_onehote)
if(len(IMAGES) == 0):
continue
IMAGES = np.array(IMAGES)
IMAGES_FUTURE = np.array(IMAGES_FUTURE)
LABELS_FUTURE = np.array(LABELS_FUTURE)
if normalise:
IMAGES_FUTURE = IMAGES_FUTURE / 255
IMAGES = IMAGES / 255
yield IMAGES , IMAGES_FUTURE , LABELS_FUTURE
# +
data_path = "highway/HighwayDataset/images"
labels_path = "highway/HighwayDataset/label"
proc = HighwayDriving_Process(data_path , labels_path , classes = None)
nb_classes = proc.nb_classes
# +
class Sequenced_Unet:
def jaccard_distance(self , y_true, y_pred, smooth=100):
intersection = K.sum(K.abs(y_true * y_pred), axis=-1)
sum_ = K.sum(K.abs(y_true) + K.abs(y_pred), axis=-1)
jac = (intersection + smooth) / (sum_ - intersection + smooth)
return (1 - jac) * smooth
def conv2d(self , x, n_filters=64 , kernel_size = 3, batchnorm = True , name = None):
x = TimeDistributed(Conv2D(filters = n_filters, kernel_size = (kernel_size, kernel_size),
kernel_initializer = 'glorot_uniform', padding = 'same', activation=None ) , name = name)(x)
if batchnorm:
x = TimeDistributed(BatchNormalization() , name = name+'_bn')(x)
x = TimeDistributed(Activation('relu') , name = name+'_relu')(x)
return x
def conv2d_block(self , x , n_filters=64 , kernel_size = 3, batchnorm = True , name = None):
x = self.conv2d(x , n_filters , kernel_size , batchnorm , name= name+"_conv1")
x = self.conv2d(x , n_filters , kernel_size , batchnorm , name= name+"_conv2")
return x
def convlstm2d_block(self , x , filters = 64 , kernel_size = 3 , name = None , return_sequence = True):
x = ConvLSTM2D(filters = n_filters , kernel_size = kernel_size , strides = 1 , padding = 'same' , return_sequences = True , name = name+"_1")(x)
x = ConvLSTM2D(filters = n_filters , kernel_size = kernel_size , strides = 1 , padding = 'same' , return_sequences = return_sequence , name = name+"_2")(x)
return x
def __init__(self ,nb_classes , size = (512 , 512 , 3) , n_filters = 16 , time_step = 4, batchnorm = True):
self.size = size
self.nb_class = nb_classes
inp = Input(shape = (time_step,)+size)
c1 = self.conv2d_block(inp, n_filters * 1, kernel_size = 3, batchnorm = batchnorm , name = 'c1')
seqc1 = self.convlstm2d_block(c1 , filters = n_filters * 1 , kernel_size = 3 , name = 'seqc1')
p1 = TimeDistributed(MaxPooling2D((2, 2)) , name='mp1')(seqc1)
c2 = self.conv2d_block(p1, n_filters * 2, kernel_size = 3, batchnorm = batchnorm , name="c2")
seqc2 = self.convlstm2d_block(c2 , filters = n_filters * 2 , kernel_size = 3 , name = 'seqc2')
p2 = TimeDistributed(MaxPooling2D((2, 2)) , name='mp2')(seqc2)
c3 = self.conv2d_block(p2, n_filters * 4, kernel_size = 3, batchnorm = batchnorm , name = "c3")
seqc3 = self.convlstm2d_block(c3 , filters = n_filters * 4 , kernel_size = 3 , name = 'seqc3')
p3 = TimeDistributed(MaxPooling2D((2, 2)), name='mp3')(seqc3)
c4 = self.conv2d_block(p3, n_filters * 8, kernel_size = 3, batchnorm = batchnorm , name = "c4")
seqc4 = self.convlstm2d_block(c4 , filters = n_filters * 8 , kernel_size = 3 , name = 'seqc4')
p4 = TimeDistributed(MaxPooling2D((2, 2)), name='mp4')(seqc4)
c5 = self.conv2d_block(p4, n_filters * 16, kernel_size = 3, batchnorm = batchnorm , name = "c5")
seq = self.convlstm2d_block(c5 , filters = n_filters * 16 , kernel_size = 3 , name = 'seq') ## Remember since we are intending to recreate the frames we are intending it to be an identity function i.e use (1,1) filters
u6 = TimeDistributed(Conv2DTranspose(n_filters * 8, (3, 3), strides = (2, 2), padding = 'same') , name = "up1")(seq)
u6 = Concatenate(axis = -1)([u6, seqc4])
c6 = self.conv2d_block(u6, n_filters * 8, kernel_size = 3, batchnorm = batchnorm , name = "c6")
u7 = TimeDistributed(Conv2DTranspose(n_filters * 4, (3, 3), strides = (2, 2), padding = 'same') , name = "up2")(c6)
u7 = Concatenate(axis = -1)([u7, seqc3])
c7 = self.conv2d_block(u7, n_filters * 4, kernel_size = 3, batchnorm = batchnorm , name = "c7")
u8 = TimeDistributed(Conv2DTranspose(n_filters * 2, (3, 3), strides = (2, 2), padding = 'same') , name = "up3")(c7)
u8 = Concatenate(axis = -1)([u8, seqc2])
c8 = self.conv2d_block(u8, n_filters * 2, kernel_size = 3, batchnorm = batchnorm , name = "c8")
u9 = TimeDistributed(Conv2DTranspose(n_filters * 1, (3, 3), strides = (2, 2), padding = 'same' ) , name = "up4")(c8)
u9 = Concatenate(axis = -1)([u9, seqc1])
c9 = self.conv2d_block(u9, n_filters * 1, kernel_size = 3, batchnorm = batchnorm , name = "c9")
out = TimeDistributed(Conv2D(filters = nb_classes, kernel_size = 1 ,kernel_initializer = 'glorot_uniform', padding = 'same' , activation='softmax') , name='output')(c9)
u_model = Model(inputs = inp , outputs = out)
model = Model(inputs = inp , outputs = out , name='u_net_segmentation')
self.model = model
model.summary()
def train(self , epochs = 30 ,lr = 1e-4 , batch_size = 16 , mem_rate = 10 , time_steps = 4 , skip = 4, dataset = None ,
pretrained = False , class_weights = None , model_name="time_model.h5"):
assert (dataset is not None),'log: give a dataset class with generator function for validation and training'
if pretrained:
self.model.load_weights(model_name)
self.model.compile(optimizer = Adam(lr = lr), loss = 'categorical_crossentropy', metrics = [self.jaccard_distance])
train_losses = []
val_losses = []
for i in range(epochs):
print("Epoch :", i)
print("Training .....")
for x , y in dataset.frameSequence_train_datagen(size=self.size[:-1] ,
batch_size= batch_size*mem_rate , time_steps = time_steps , skip = skip , normalise = True):
x[:, :, :, 0:28, :] = 0
x[:, :, :, 228:256, :] = 0
hist = self.model.fit(x = x , y = y , batch_size=batch_size , epochs = 1 , verbose = 1 , class_weight = class_weights)
train_losses.append(hist.history['loss'][0])
self.model.save_weights("saved_models/future_segmentation_generate/" + model_name + "_e" + str(i))
x , y = next(dataset.frameSequence_train_datagen(size = self.size[:-1] ,batch_size = 4 , time_steps = 4 , skip = 4 , normalise = True))
index = random.randint(0,x.shape[0])
p = self.model.predict(np.array([x[index]]))
plt.figure(figsize = (time_steps*time_steps,time_steps*time_steps))
for i in range(time_steps):
plt.subplot(3,time_steps,i+1)
plt.imshow(x[index][i])
plt.title("frame "+str(i+1))
plt.subplot(3,time_steps,i+time_steps+1)
plt.imshow(dataset.class2image(np.argmax(y[index][i] , axis = -1)))
plt.title("label "+str(i+1))
plt.subplot(3,time_steps,i+2*time_steps+1)
plt.imshow(dataset.class2image(np.argmax(p[0][i] , axis = -1)))
plt.title("label predicted "+str(i+1))
plt.show()
plt.figure(figsize = (10,10))
plt.plot(train_losses)
plt.show()
pickle_out = open("saved_models/future_segmentation_generate/history_future_segmentation_generate_e30.pickle","wb")
pickle.dump(train_losses, pickle_out)
pickle_out.close()
size = (256 , 256 , 3)
n_filters = 32
time_step = 4
skip = 4
nb_classes
net = Sequenced_Unet(nb_classes, size , n_filters = n_filters , time_step = time_step)
# -
epochs = 30
batch_size = 4
lr = 1e-4
net.train(epochs=epochs , lr = 1e-4 , dataset = proc , batch_size = batch_size , time_steps = time_step , skip = skip
, pretrained = False , mem_rate = 11 , class_weights = None , model_name = "time_model.h5")
# +
checkpointer = ModelCheckpoint(
filepath=os.path.join('saved_models', 'future_segmentation_generate', 'future_segmentation_generate' + \
'.{epoch:03d}-{loss:.3f}.hdf5'),
verbose=1,
save_weights_only=True,
save_best_only=True)
proc = HighwayDriving_Process(data_path, labels_path, classes = None)
datagen = proc.frameSequence_train_datagen(size = (256, 256))
history = u_model.fit_generator(generator = datagen, steps_per_epoch = 60, epochs = 5, verbose = 1, workers = 4, shuffle = False, callbacks = [checkpointer], initial_epoch = 0)
# +
proc = HighwayDriving_Process(data_path, labels_path, classes = None)
for data in proc.frameSequence_train_datagen(size = (256, 256)):
x, y = data
print(str(x.shape) + str(y.shape))
# -
| sequence models/notebooks/future_segmentation_generate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Note: This notebook about 40 MB of embedded output data, which should be put under version control.
# Please make sure to clear all cell output using the Cell>All Output>Clear command from the menu befor committing changes!
# Use Cell>Run all to reproduce the output.
#
# 注意:这个笔记本大约有40mb的嵌入式输出数据,应该放在版本控制下。
#
# 在提交更改之前,请确保使用菜单中的单元格> all output > clear命令清除所有单元格输出使用单元>全部运行以重现输出。
# # 前期准备
# +
import sys
sys.path.append(r'your_path')
sys.path.append(r'e:\360movedata\users\zx305\documents\github')
sys.path.append(r'c:\users\zx305\appdata\roaming\python\python36\site-packages')
print(sys.path)
# 这个很有用
# -
# !pip install msgpack --user
# #!python -m pip install --upgrade pip
# #!python3 -m pip install --upgrade pip
# !python3 -m numpy install --upgrade numpy
# !pip install librosa --user
# !pip install mir_eval --user
# !pip install pylearn2 --user
# # !pip install deepthought --user #这段不是这么安装的!但还没有解决
# !pip uninstall deepthought
# !pip list
# !cd/d E:\360MoveData\Users\zx305\Documents\GitHub\pylearn2
# !python setup.py install
# # 程序正式开始段
# +
import sys
sys.path.append(r'your_path')
sys.path.append(r'd:\python\anaconda3\lib\site-packages')
print(sys.path)
# 这个很有用
# -
import numpy as np
import librosa #音频处理用的
import mir_eval
import matplotlib.pyplot as plt
# %matplotlib inline
# +
import IPython.display
from IPython.display import display
import os
import deepthought
from deepthought.deepthought.datasets.openmiir.metadata import load_stimuli_metadata, save_beat_times
STIMULI_VERSION = 2 # change to 1 for older stimuli version
data_root = os.path.join(deepthought.DATA_PATH, 'OpenMIIR')
default_save_beat_times = False # change to True to save beat time to txt file
# +
def play_beats(y, sr, beats):
if y is None:
# Sonify the beats only
y_beat = mir_eval.sonify.clicks(beats, sr, length=len(y))
else:
# Sonify the beats and add them to y
y_beat = y + mir_eval.sonify.clicks(beats, sr, length=len(y))
return IPython.display.Audio(data=y_beat, rate=sr)
def visualize(y, sr, title=None, playback=True, beats=None):
# show playback widget above figure
if playback:
if title is not None:
print title
if beats is None:
display(IPython.display.Audio(data=y, rate=sr))
else:
beat_times = librosa.frames_to_time(beats, sr=sr, hop_length=64)
display(play_beats(y, sr, beat_times))
# Let's make and display a mel-scaled power (energy-squared) spectrogram
# We use a small hop length of 64 here so that the frames line up with the beat tracker example below.
S = librosa.feature.melspectrogram(y, sr=sr, n_fft=2048, hop_length=64, n_mels=128)
# Convert to log scale (dB). We'll use the peak power as reference.
log_S = librosa.logamplitude(S, ref_power=np.max)
# Make a new figure
plt.figure(figsize=(12,4))
# Display the spectrogram on a mel scale
# sample rate and hop length parameters are used to render the time axis
librosa.display.specshow(log_S, sr=sr, hop_length=64, x_axis='time', y_axis='mel')
# Put a descriptive title on the plot
if title is not None:
plt.title('mel power spectrogram ({})'.format(title))
if beats is not None:
# Let's draw lines with a drop shadow on the beat events
plt.vlines(beats, 0, log_S.shape[0], colors='k', linestyles='-', linewidth=2.5)
plt.vlines(beats, 0, log_S.shape[0], colors='w', linestyles='-', linewidth=1.5)
# draw a color bar
plt.colorbar(format='%+02.0f dB')
# Make the figure layout compact
plt.tight_layout()
# This make sure the figures are plotted in place and not after text and audio
plt.show()
plt.close()
def _analyze_beats(audio_filepath, bpm, label=None, tightness=400, offset=0, duration=None, vy=True, vh=True, vp=True, vb=True):
print audio_filepath
# load audio file
# sr = 22050 # default
sr = 44100 # slower but gives better results for Harry Potter Theme
y, sr = librosa.load(audio_filepath, sr=sr, offset=offset, duration=duration)
if label is not None:
print label
if vy:
visualize(y, sr, 'original')
# split into harmonic and percussive component
y_harmonic, y_percussive = librosa.effects.hpss(y)
if vh:
visualize(y_harmonic, sr, 'harmonic component')
if vp:
visualize(y_percussive, sr, 'percussive component')
# Now, let's run the beat tracker
# We'll use the percussive component for this part
# By default, the beat tracker will trim away any leading or trailing beats that don't appear strong enough.
# To disable this behavior, call beat_track() with trim=False.
tempo, beats = librosa.beat.beat_track(y=y_percussive, sr=sr, hop_length=64, trim=False, start_bpm=bpm, tightness=tightness)
# Let's re-draw the spectrogram, but this time, overlay the detected beats
if vb:
visualize(y, sr, 'with beats', beats=beats)
print 'Offset: %.4f s' % offset
print 'Expected tempo: %.2f BPM' % bpm
print 'Estimated tempo: %.2f BPM' % tempo
print 'First 5 beat frames: ', beats[:5]
# Frame numbers are great and all, but when do those beats occur?
print 'First 5 beat times: ', librosa.frames_to_time(beats[:5], sr=sr, hop_length=64)
return tempo, beats, librosa.frames_to_time(beats, sr=sr, hop_length=64)
def get_audio_filepath(meta):
return os.path.join(data_root, 'audio', 'full.v{}'.format(STIMULI_VERSION), meta['audio_file'])
def analyze_beats(meta, tightness=400, save=default_save_beat_times, **kwargs):
tempo, beat_frames, beat_times = _analyze_beats(
audio_filepath=get_audio_filepath(meta),
label=meta['label'],
bpm=meta['bpm'],
tightness=tightness,
offset=meta['length_of_cue'],
**kwargs
)
if save:
offset = meta['length_of_cue']
save_beat_times(beat_times, stimulus_id=meta['id'], offset=offset, version=STIMULI_VERSION)
return tempo, beat_frames, beat_times
# -
# NOTE: this is experimental
def analyze_onsets(meta):
audio_filepath=os.path.join(data_root, 'audio', 'full.v{}'.format(STIMULI_VERSION), meta['audio_file'])
sr = 44100 # slower but gives better results for Harry Potter Theme
offset=meta['length_of_cue']
duration=None
print sr
y, sr = librosa.load(audio_filepath, sr=sr, offset=offset, duration=duration)
"""
# Get onset times from a signal
onset_frames = librosa.onset.onset_detect(y=y, sr=sr, hop_length=64)
onset_times = librosa.frames_to_time(onset_frames, sr, hop_length=64)
# Or use a pre-computed onset envelope
o_env = librosa.onset.onset_strength(y, sr=sr)
onset_frames = librosa.onset.onset_detect(onset_envelope=o_env, sr=sr)
onset_times = librosa.frames_to_time(onset_frames, sr, hop_length=64)
"""
onset_frames = librosa.onset.onset_detect(y=y, sr=sr, hop_length=64)
print onset_frames
visualize(y, sr, 'with beats', beats=onset_frames)
o_env = librosa.onset.onset_strength(y, sr=sr)
plt.plot(o_env)
onset_frames = librosa.onset.onset_detect(onset_envelope=o_env, sr=sr)
print onset_frames
visualize(y, sr, 'with beats', beats=onset_frames*7)
meta = load_stimuli_metadata(data_root, version=STIMULI_VERSION)
# print meta
# # some examples
# run this to analyze onsets for stimulus 22
analyze_onsets(meta[22])
# run this to analyze onsets for stimulus 1, specify tightness
tempo, beat_frames, beat_times = analyze_beats(meta[1], tightness=800)
print beat_times
print beat_frames
# analyze beginning of stimulus 22,
# suppress visualization of original signal (vy) and harmonic (vh) and percussive (vp) component
_analyze_beats(get_audio_filepath(meta[22]),
bpm=166, tightness=250, offset=2.182, duration=4.0, vy=False, vh=False, vp=False);
# test different tightness settings on stimulus 22
_analyze_beats(get_audio_filepath(meta[22]),
bpm=166, tightness=250, offset=0, duration=None, vy=False, vh=False, vp=False);
_analyze_beats(get_audio_filepath(meta[22]),
bpm=166, tightness=400, offset=0, duration=None, vy=False, vh=False, vp=False);
_analyze_beats(get_audio_filepath(meta[22]),
bpm=166, tightness=800, offset=0, duration=None, vy=False, vh=False, vp=False);
# # analyze stimuli
_ = analyze_beats(meta[1], tightness=1000, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[2], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[3], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[4], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[11], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[12], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[13], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[14], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[21], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[22], tightness=300, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[23], tightness=800, vy=False, vh=False, vp=False)
_ = analyze_beats(meta[24], tightness=800, vy=False, vh=False, vp=False)
# # analyze cue click tracks
# +
from deepthought.datasets.openmiir.constants import STIMULUS_IDS
for stimulus_id in STIMULUS_IDS:
tempo, beat_frames, beat_times = _analyze_beats(
audio_filepath=os.path.join(data_root, 'audio', 'cues.v{}'.format(STIMULI_VERSION), meta[stimulus_id]['cue_file']),
label=meta[stimulus_id]['label'],
bpm=meta[stimulus_id]['cue_bpm'],
tightness=10000, vy=False, vh=False, vp=False
)
if default_save_beat_times:
save_beat_times(beat_times, stimulus_id=stimulus_id, cue=True, version=STIMULI_VERSION)
| .ipynb_checkpoints/Beat and Onset Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: MindSpore-1.0.1
# language: python
# name: mindspore-1.0.1
# ---
# # <center/>同步训练和验证模型体验
# ## 概述
# 在面对复杂网络时,往往需要进行几十甚至几百次的epoch训练。而在训练之前,往往很难掌握在训练到第几个epoch时,模型的精度能达到满足要求的程度。所以经常会采用一边训练的同时,在相隔固定epoch的位置对模型进行精度验证,并保存相应的模型,等训练完毕后,通过查看对应模型精度的变化就能迅速地挑选出相对最优的模型,本文将采用这种方法,以LeNet网络为样本,进行示例。
# 整体流程如下:
# 1. 数据集准备。
# 2. 构建神经网络。
# 3. 定义回调函数EvalCallBack。
# 4. 定义训练网络并执行。
# 5. 定义绘图函数并对不同epoch下的模型精度绘制出折线图。
#
# > 本文档适用于CPU、GPU和Ascend环境。
# ## 数据准备
# ### 数据集的下载
# 下载并解压数据集数据。
# !wget -N https://obs.dualstack.cn-north-4.myhuaweicloud.com/mindspore-website/notebook/datasets/MNIST_Data.zip
# !unzip -o MNIST_Data.zip -d ./datasets
# !tree ./datasets/MNIST_Data/
# ### 数据集的增强操作
# 下载下来后的数据集,需要通过`mindspore.dataset`处理成适用于MindSpore框架的数据,再使用一系列框架中提供的工具进行数据增强操作来适应LeNet网络的数据处理需求。
# +
import os
import mindspore.dataset as ds
import mindspore.dataset.vision.c_transforms as CV
import mindspore.dataset.transforms.c_transforms as C
from mindspore.dataset.vision import Inter
from mindspore import dtype as mstype
def create_dataset(data_path, batch_size=32, repeat_size=1,
num_parallel_workers=1):
# define dataset
mnist_ds = ds.MnistDataset(data_path)
# define map operations
resize_op = CV.Resize((32, 32), interpolation=Inter.LINEAR)
rescale_nml_op = CV.Rescale(1 / 0.3081, -1 * 0.1307 / 0.3081)
rescale_op = CV.Rescale(1/255.0, 0.0)
hwc2chw_op = CV.HWC2CHW()
type_cast_op = C.TypeCast(mstype.int32)
# apply map operations on images
mnist_ds = mnist_ds.map(operations=type_cast_op, input_columns="label", num_parallel_workers=num_parallel_workers)
mnist_ds = mnist_ds.map(operations=[resize_op,rescale_op,rescale_nml_op,hwc2chw_op],
input_columns="image", num_parallel_workers=num_parallel_workers)
# apply DatasetOps
buffer_size = 10000
mnist_ds = mnist_ds.shuffle(buffer_size=buffer_size)
mnist_ds = mnist_ds.batch(batch_size, drop_remainder=True)
mnist_ds = mnist_ds.repeat(repeat_size)
return mnist_ds
# -
# ## 构建神经网络
# LeNet网络属于7层神经网络,其中涉及卷积层,全连接层,函数激活等算法,在MindSpore中都已经建成相关算子只需导入使用,如下先将卷积函数,全连接函数,权重等进行初始化,然后在LeNet5中定义神经网络并使用`construct`构建网络。
# +
import mindspore.nn as nn
from mindspore.common.initializer import Normal
class LeNet5(nn.Cell):
"""Lenet network structure."""
# define the operator required
def __init__(self, num_class=10, num_channel=1):
super(LeNet5, self).__init__()
self.conv1 = nn.Conv2d(num_channel, 6, 5, pad_mode='valid')
self.conv2 = nn.Conv2d(6, 16, 5, pad_mode='valid')
self.fc1 = nn.Dense(16 * 5 * 5, 120, weight_init=Normal(0.02))
self.fc2 = nn.Dense(120, 84, weight_init=Normal(0.02))
self.fc3 = nn.Dense(84, num_class, weight_init=Normal(0.02))
self.relu = nn.ReLU()
self.max_pool2d = nn.MaxPool2d(kernel_size=2, stride=2)
self.flatten = nn.Flatten()
# use the preceding operators to construct networks
def construct(self, x):
x = self.max_pool2d(self.relu(self.conv1(x)))
x = self.max_pool2d(self.relu(self.conv2(x)))
x = self.flatten(x)
x = self.relu(self.fc1(x))
x = self.relu(self.fc2(x))
x = self.fc3(x)
return x
# -
# ## 定义回调函数EvalCallBack
# 实现思想:每隔n个epoch验证一次模型精度,需要在自定义回调函数中实现,如需了解自定义回调函数的详细用法,请参考[API说明](https://www.mindspore.cn/doc/api_python/zh-CN/master/mindspore/mindspore.train.html#mindspore.train.callback.Callback)。
#
# 核心实现:回调函数的`epoch_end`内设置验证点,如下:
#
# `cur_epoch % eval_per_epoch == 0`:即每`eval_per_epoch`个epoch结束时,验证一次模型精度。
#
# - `cur_epoch`:当前训练过程的`epoch`数值。
# - `eval_per_epoch`:用户自定义数值,即验证频次。
#
# 其他参数解释:
#
# - `model`:MindSpore中的`Model`类。
# - `eval_dataset`:验证数据集。
# - `epoch_per_eval`:记录验证模型的精度和相应的epoch数,其数据形式为`{"epoch":[],"acc":[]}`。
# +
from mindspore.train.callback import Callback
class EvalCallBack(Callback):
def __init__(self, model, eval_dataset, eval_per_epoch, epoch_per_eval):
self.model = model
self.eval_dataset = eval_dataset
self.eval_per_epoch = eval_per_epoch
self.epoch_per_eval = epoch_per_eval
def epoch_end(self, run_context):
cb_param = run_context.original_args()
cur_epoch = cb_param.cur_epoch_num
if cur_epoch % self.eval_per_epoch == 0:
acc = self.model.eval(self.eval_dataset, dataset_sink_mode=False)
self.epoch_per_eval["epoch"].append(cur_epoch)
self.epoch_per_eval["acc"].append(acc["Accuracy"])
print(acc)
# -
# ## 定义训练网络并执行
# 在保存模型的参数`CheckpointConfig`中,需计算好单个`epoch`中的`step`数,根据保存模型参数`ckpt`文件时,需要间隔的`step`数来设置,本次示例每个`epoch`有1875个`step`,按照每两个`epoch`验证一次的思想,这里设置`save_checkpoint_steps=eval_per_epoch*1875`,
# 其中变量`eval_per_epoch`等于2。
#
# 参数解释:
#
# - `train_data_path`:训练数据集地址。
# - `eval_data_path`:验证数据集地址。
# - `train_data`:训练数据集。
# - `eval_data`:验证数据集。
# - `net_loss`:定义损失函数。
# - `net-opt`:定义优化器函数。
# - `config_ck`:配置保存模型信息。
# - `save_checkpoint_steps`:每多少个step保存一次模型权重参数`ckpt`文件。
# - `keep_checkpoint_max`:设置保存模型的权重参数`cpkt`文件的数量上限。
# - `ckpoint_cb`:配置模型权重参数`ckpt`文件保存名称的前缀信息及保存路径信息。
# - `model`:MindSpore中的`Model`类。
# - `model.train`:`Model`类的执行训练函数。
# - `epoch_per_eval`:定义收集`epoch`数和对应模型精度信息的字典。
# +
from mindspore.train.callback import ModelCheckpoint, CheckpointConfig, LossMonitor
from mindspore import context, Model
from mindspore.nn.metrics import Accuracy
from mindspore.nn.loss import SoftmaxCrossEntropyWithLogits
import os
if __name__ == "__main__":
context.set_context(mode=context.GRAPH_MODE, device_target="CPU")
train_data_path = "./datasets/MNIST_Data/train"
eval_data_path = "./datasets/MNIST_Data/test"
model_path = "./models/ckpt/mindspore_evaluate_the_model_during_training/"
# clean up old run files before in Linux
os.system('rm -f {}*.ckpt {}*.meta {}*.pb'.format(model_path, model_path, model_path))
epoch_size = 10
eval_per_epoch = 2
repeat_size = 1
network = LeNet5()
train_data = create_dataset(train_data_path, repeat_size=repeat_size)
eval_data = create_dataset(eval_data_path, repeat_size=repeat_size)
# define the loss function
net_loss = SoftmaxCrossEntropyWithLogits(sparse=True, reduction='mean')
# define the optimizer
net_opt = nn.Momentum(network.trainable_params(), learning_rate=0.01, momentum=0.9)
config_ck = CheckpointConfig(save_checkpoint_steps=eval_per_epoch*1875, keep_checkpoint_max=15)
ckpoint_cb = ModelCheckpoint(prefix="checkpoint_lenet", directory=model_path, config=config_ck)
model = Model(network, net_loss, net_opt, metrics={"Accuracy": Accuracy()})
epoch_per_eval = {"epoch": [], "acc": []}
eval_cb = EvalCallBack(model, eval_data, eval_per_epoch, epoch_per_eval)
model.train(epoch_size, train_data, callbacks=[ckpoint_cb, LossMonitor(375), eval_cb],
dataset_sink_mode=False)
# -
# 在同一目录的文件夹中可以看到`lenet_ckpt`文件夹中,保存了5个模型,和一个计算图相关数据,其结构如下:
# !tree ./models/ckpt/mindspore_evaluate_the_model_during_training
# ## 绘制不同epoch下模型的精度
# 定义绘图函数`eval_show`,将`epoch_per_eval`载入到`eval_show`中,绘制出不同`epoch`下模型的验证精度折线图。
# +
import matplotlib.pyplot as plt
def eval_show(epoch_per_eval):
plt.xlabel("epoch number")
plt.ylabel("Model accuracy")
plt.title("Model accuracy variation chart")
plt.plot(epoch_per_eval["epoch"], epoch_per_eval["acc"], "red")
plt.show()
eval_show(epoch_per_eval)
# -
# 从上图可以一目了然地挑选出需要的最优模型权重参数`ckpt`文件。
# ## 总结
# 本例使用MNIST数据集通过卷积神经网络LeNet5进行训练,着重介绍了利用回调函数在进行模型训练的同时进行模型的验证,保存对应`epoch`的模型权重参数`ckpt`文件,并从中挑选出最优模型的方法。
| tutorials/notebook/mindspore_evaluate_the_model_during_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"<NAME>"
greeting = "Hello World" # this is a string variable (<- this is a comment)
greeting
greeting = "Hello World"
greeting
print(greeting)
a = 2
b = 15
a + b
my_sum = a + b # get sum of a & b
# print sum
my_sum
my_product = a * b
my_product
import numpy as np
ages_list = [25, 30, 35, 40, 45] # this is a Python list
ages = np.array(ages_list) # this is a numpy array
ages
# you can also do this in a single line
ages = np.array([25, 30, 35, 40, 45, 50, 55])
ages
ages[0]
ages[6]
ages[-1] # this is the last element
ages[0:4] # goes from index = 0 to 3 (first 4 elements)
ages[:4] # goes from beginning to index 3 (same result as above)
ages[1:4] # goes from index = 1 to 3 (2nd to 4th elements)
ages[1:] #goes from index = 1 to last element (same result as above)
ages[:] # goes from first to last element (same as ages)
ages[::2] # keeps every 2nd element only
ages[::-1] # reverses order of elements
ages[ages<40]
ages[ages<=40]
len(ages)
ages_mean = np.mean(ages)
ages_mean
np.min(ages)
np.max(ages)
| session-4/scripts/1_Intro_to_Python.ipynb |
# +
#@title Copyright 2021 The Earth Engine Community Authors { display-mode: "form" }
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# -
# # Mathematical Operations
#
#
# Image math can be performed using operators like `add()` and `subtract()`, but for complex computations with more than a couple of terms, the `expression()` function provides a good alternative. See the following sections for more information on [operators](https://developers.google.com#operators) and [expressions](https://developers.google.com#expressions).
#
# ## Setup
#
# ### Earth Engine setup
import ee
ee.Authenticate()
ee.Initialize()
# ### Folium setup (for interactive map display)
# +
import folium
def add_ee_layer(self, ee_image_object, vis_params, name):
map_id_dict = ee.Image(ee_image_object).getMapId(vis_params)
folium.raster_layers.TileLayer(
tiles=map_id_dict['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
name=name,
overlay=True,
control=True
).add_to(self)
folium.Map.add_ee_layer = add_ee_layer
# -
# ## Operators
#
# Math operators perform basic arithmetic operations on image bands. They take two inputs: either two images or one image and a constant term, which is interpreted as a single-band constant image with no masked pixels. Operations are performed per pixel for each band.
#
# As a simple example, consider the task of calculating the Normalized Difference Vegetation Index (NDVI) using Landsat imagery, where `add()`, `subtract()`, and `divide()` operators are used:
# +
# Load a 5-year Landsat 7 composite 1999-2003.
landsat_1999 = ee.Image('LANDSAT/LE7_TOA_5YEAR/1999_2003')
# Compute NDVI.
ndvi_1999 = (landsat_1999.select('B4').subtract(landsat_1999.select('B3'))
.divide(landsat_1999.select('B4').add(landsat_1999.select('B3'))))
# -
# **Note:** the normalized difference operation is available as a shortcut method: [`normalizedDifference()`](https://developers.google.com/earth-engine/apidocs/ee-image-normalizeddifference).
#
# Only the intersection of unmasked pixels between the two inputs are considered and returned as unmasked, all else are masked. In general, if either input has only one band, then it is used against all the bands in the other input. If the inputs have the same number of bands, but not the same names, they're used pairwise in the natural order. The output bands are named for the longer of the two inputs, or if they're equal in length, in the first input's order. The type of the output pixels is the union of the input types.
#
# The following example of multi-band image subtraction demonstrates how bands are matched automatically, resulting in a “change vector” for each pixel for each co-occurring band.
# +
# Load a 5-year Landsat 7 composite 2008-2012.
landsat_2008 = ee.Image('LANDSAT/LE7_TOA_5YEAR/2008_2012')
# Compute multi-band difference between the 2008-2012 composite and the
# previously loaded 1999-2003 composite.
diff = landsat_2008.subtract(landsat_1999)
# Compute the squared difference in each band.
squared_difference = diff.pow(2)
# Define a map centered on Australia.
map_diff = folium.Map(location=[-24.003, 133.565], zoom_start=5)
# Add the image layers to the map and display it.
map_diff.add_ee_layer(diff,
{'bands': ['B4', 'B3', 'B2'], 'min': -32, 'max': 32},
'diff.')
map_diff.add_ee_layer(squared_difference,
{'bands': ['B4', 'B3', 'B2'], 'max': 1000},
'squared diff.')
display(map_diff.add_child(folium.LayerControl()))
# -
# In the second part of this example, the squared difference is computed using `image.pow(2)`. For the complete list of mathematical operators handling basic arithmetic, trigonometry, exponentiation, rounding, casting, bitwise operations and more, see the [API documentation](https://developers.google.com/earth-engine/apidocs).
#
# ## Expressions
#
# To implement more complex mathematical expressions, consider using `image.expression()`, which parses a text representation of a math operation. The following example uses `expression()` to compute the Enhanced Vegetation Index (EVI):
# +
# Load a Landsat 8 image.
image = ee.Image('LANDSAT/LC08/C01/T1_TOA/LC08_044034_20140318')
# Compute the EVI using an expression.
evi = image.expression(
'2.5 * ((NIR - RED) / (NIR + 6 * RED - 7.5 * BLUE + 1))', {
'NIR': image.select('B5'),
'RED': image.select('B4'),
'BLUE': image.select('B2')
})
# Define a map centered on San Francisco Bay.
map_evi = folium.Map(location=[37.4675, -122.1363], zoom_start=9)
# Add the image layer to the map and display it.
map_evi.add_ee_layer(
evi, {'min': -1, 'max': 1, 'palette': ['FF0000', '00FF00']}, 'evi')
display(map_evi)
# -
# Observe that the first argument to `expression()` is the textual representation of the math operation, the second argument is a dictionary where the keys are variable names used in the expression and the values are the image bands to which the variables should be mapped. Bands in the image may be referred to as `b("band name")` or `b(index)`, for example `b(0)`, instead of providing the dictionary. Bands can be defined from images other than the input when using the band map dictionary. Note that `expression()` uses "floor division", which discards the remainder and returns an integer when two integers are divided. For example `10 / 20 = 0`. To change this behavior, multiply one of the operands by `1.0`: `10 * 1.0 / 20 = 0.5`. Only the intersection of unmasked pixels are considered and returned as unmasked when bands from more than one source image are evaluated. Supported expression operators are listed in the following table.
#
# Operators for `expression()`:
#
# Type | Symbol | Name
# ---|---|---
# **Arithmetic** | \+ - * / % ** | Add, Subtract, Multiply, Divide, Modulus, Exponent
# **Relational** | == != < > <= >= | Equal, Not Equal, Less Than, Greater than, etc.
# **Logical** | && || ! ^ | And, Or, Not, Xor
# **Ternary** | ? : | If then else
| guides/ipynb/image_math.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: blog
# language: python
# name: blog
# ---
# # Statistics essentials
#
# - hide: false
# - toc: true
# - comments: true
# - categories: [stats]
# ## Distributions
# My notes from working through section 2, data and sampling distributions, of [Practical statistics for data science](https://learning.oreilly.com/library/view/practical-statistics-for/9781492072935/), to revise consepts and get comfortable implementing them in Python.
# Terminology
# - Stochastic is a synonym for random. A stochastic process is a random process. The distinction between *stochastics* and *statistics* is that a stochastic processes generate the data we analyse in statistics.
# Sampling
#
# - We rely on a sample to learn about a larger population.
# - We thus need to make sure that the sampling procedure is free of bias, so that units in the sample are representative of those in the population.
# - While representativeness cannot be achieved perfectly, it's important to ensure that non-representativeness is due to random error and not due to systematic bias.
# - Random errors produce deviations that vary over repeated samples, while systematic bias persists. Such selection bias can lead to misleading and ephemeral conclusions.
# - Two basic sampling procedures are simple random sampling (randomly select $n$ units from a population of $N$) and stratified random sampling (randomly select $n_s$ from each stratum $S$ of a population of $N$).
# - The mean outcome of the sample is denoted $\bar{x}$, that of the population $\mu$.
#
# Selection bias
#
# - Using the data to answer many questions will eventually reveal something interesting by mere chance (if 20,000 people flip a coin 10 times, some will have 10 straight heads). This is sometimes called the Vast Search Effect.
# - Common types of selection bias in data science:
# - The vast search effect
# - Nonrandom sampling
# - Cherry-picking data
# - Selecting specific time-intervals
# - Stopping experiments prematurely
# - Ways to guard against selection bias: have one or many holdout datasets to confirm your results.
# - Regression to the mean results form a particular kind of selection bias in a setting where we measure outcomes repeatedly over time: when luck and skill combine to determine outcomes, winners of one period will be less lucky next period and perform closer to the mean performer.
# Sampling distributions
#
# - A sampling distribution is the distribution of a statistic (e.g. the mean) over many repeated samples. Classical statistics is much concerned with making inferences from samples about the population based on such statistics.
# - When we measure an attribute of the population based on a sample using a statistic, the result will vary over repeated samples. To capture by how much it varies, we are concerned with the sampling variability.
#
# - Key distinctions:
# - The data distribution is the distribution of the data in the sample, the sampling distribution is the distribution of the sample statistic.
# - The standard deviation is a measure of spread of the data distribution, the standard error a measure of spread of the sampling distribution.
# +
import pandas as pd
import numpy as np
import seaborn as sns
from scipy.stats import norm
import matplotlib.pyplot as plt
mean, sd, N = 0, 1, 1_000_000
full_data = norm.rvs(mean, sd, N)
sample_data = pd.DataFrame({
'income': np.random.choice(full_data, 1000),
'type': 'Data'
})
mof1 = pd.DataFrame({
'income': [np.random.choice(full_data, 1).mean() for _ in range(1000)],
'type':'Mean of 1'
})
mof5 = pd.DataFrame({
'income': [np.random.choice(full_data, 5).mean() for _ in range(1000)],
'type':'Mean of 5'
})
mof20 = pd.DataFrame({
'income': [np.random.choice(full_data, 20).mean() for _ in range(1000)],
'type':'Mean of 20'
})
mof100 = pd.DataFrame({
'income': [np.random.choice(full_data, 100).mean() for _ in range(1000)],
'type':'Mean of 100'
})
results = pd.concat([sample_data, mof1, mof5, mof20, mof100])
g = sns.FacetGrid(results, col='type')
g.map(plt.hist, 'income', bins=40)
g.set_axis_labels('Income', 'Count')
g.set_titles('{col_name}');
# -
# Plots show that:
# - Data distribution has larger spread than sampling distributions (each data point is a special case of a sample with n = 1)
# - The spread of sampling distributions decreases with increasing sample size
# Degrees of freedom
#
# - The number of parameters you had to estimate en route to calculate the desired statistic ([source](http://onlinestatbook.com/2/estimation/df.html)). If you calculate sample variance with an estimated mean rather than a known mean, you have to estimate the sample mean first and thus loose 1 degree of freedom. Hence, you'd divide the sum of squared deviations from the (estimated) mean by n-1 rather than n.
# Central limit theorem
#
# - The second point above is an instance of the central limit theorem, which states that means from multiple samples are normally distributed even if the underlying distribution is not normal, provied that the sample size is large enough.
#
# - More precisely: Suppose that we have a sequence of independent and identically distributed (iid) random variables $\{x_1, ..., x_n\}$ drawn from a distribution with expected value $\mu$ and finite variance given by $\sigma^2$, and we are interested in the mean value $\bar{x} = \frac{x_1 + ... + x_n}{n}$. By the law of large numbers, $\bar{x}$ converges to $\mu$. The central limite theorem describes the shape of the random variation of $\bar{x}$ around $\mu$ during this convergence. In particular, for large enough $n$, the distribution of $\bar{x}$ will be close to a normal distribution with mean $\mu$ and standard deviation $\sigma/n$.
#
# - This is useful because it means that irrespective of the underlying distribution (i.e. the distribution of the values in our sequence above), we can use the normal distribution and approximations to it (such as the t-distribution) to calculate sample distributions when we do inference. Because of this, the CLT is at the heart of the theory of hypothesis testing and confidence intervals, and thus of much of classical statistics.
#
# - For experiments, this means that our estiamted treatment effect is normally distributed, which is what allows us to draw inferences from our experimental setting ot the population as a whole. The CLT is thus at the heart of the experimental approach.
# +
# CLT demo
from scipy.stats import norm, gamma
import matplotlib.pyplot as plt
def means(n):
return [np.mean(norm.rvs(0, 2, 10)) for _ in range(n)]
plt.subplots(figsize=(10,10))
plt.subplot(441)
plt.hist(means(100), bins=30)
plt.subplot(442)
plt.hist(means(1000), bins=30)
plt.subplot(443)
plt.hist(means(10000), bins=30);
# -
# Standard error
#
# - The standard error is a measure for the variability of the sampling distribution.
# - It is related to the standard deviation of the observations, $\sigma$ and the sample size $n$ in the following way:
#
# $$
# se = \frac{\sigma}{\sqrt{n}}
# $$
#
# - The relationship between sample size and se is sometimes called the "Square-root of n rule", since reducing the $se$ by a factor of 2 requires an increase in the sample size by a factor of 4.
# Bootstrap
#
# - In practice, we often use the bootstrap to calculate standard errors of model parameters or statistics.
# - Conceptually, the bootstrap works as follows: 1) we draw an original sample and calculate our statistic, 2) we then create a blown-up version of that sample by duplicating it many times, 3) we then draw repeated samples from the large sample, recalculate our statistic, and calculate the standard deviation of these statistics to get the standard error.
# - To achieve this easily, we can skip step 2) by simply sampling with replacement from the original distribution in step 3).
# - The full procedure makes clear what the bootstrap results tell us, however: they tell us how lots of additional samples would behave if they were drawn from a population like our original sample.
# - Hence, if the original sample is not representative of the population of interest, then bootstrap results are not informative about that population either.
# - The bootstrap can also be used to improve the performance of classification or regression trees by fitting multiple trees on bootstrapped sample and then averaging their predictions. This is called "bagging", short for "bootstrap aggregating".
#
# +
# A simple bootstrap implementation
from sklearn.utils import resample
mean, sd, N = 0, 5, 1000
original_sample = norm.rvs(mean, sd, N)
results = []
for nrepeat in range(1000):
sample = resample(original_sample)
results.append(np.median(sample))
print('Bootstrap Statistics:')
print(f'Original: {np.median(original_sample)}')
print(f'Bias: {np.median(results) - np.median(original_sample)}')
print(f'Std. error: {np.std(results)}')
# -
# Confidence intervals
#
# - A CI is another way to learn about the variability of a test statistic.
# - It can be calculated using the (standard) normal distribution or the t-distribution (if sample sizes are small).
# - But for data science purposes we can compute a x percent CI from the bootstrap, following this algorithm: 1) Draw a large number of bootstrap samples and calculate the statistic of interest, 2) Trim [(100-x)/2] percent of the bootstrap results on either end of the distribution, 3) the trim points are the end point of the CI.
# The normal distribution
#
# - Useful not mainly because data is often normally distributed, but because sample distributions of statistics (as well as errors) often are.
# - But rely on normality assumption only as a last resort if using empirical distributions or bootstrap is not available.
# Q-Q plots
#
# - Q-Q plots (for quantile-quantile plot) help us compare the quantiles in our dataset to the quantiles of a theoretical distribution to see whether our data follows this distribution (I'll refer to the normal distribution below to fix ideas).
# - In general, the x percent quantile is a point in the data such that x percent of the data fall below it (this point is also the xth percentile).
# - To create a Q-Q plot, we proceed as follows: First, we split the data into quantiles such that each data point represents its own quantiles. Second, we split the normal distribution into an equal number of quantiles (for the normal distribution, quantiles are intervals of equal probability mass). Third, we mark the quantiles for the data on the y-axis and for the normal distribution on the x-axis. Finally, we use these points as coordinates for each quantile in the plot. (See [this](https://www.youtube.com/watch?v=okjYjClSjOg) helpful video for more details on how to construct Q-Q plots, and [this](https://towardsdatascience.com/explaining-probability-plots-9e5c5d304703) useful article for details on probability plots more generally.)
# +
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from statsmodels.api import ProbPlot
from scipy import stats
# %config InlineBackend.figure_format ='retina'
sns.set_style('darkgrid')
sns.mpl.rcParams['figure.figsize'] = (10.0, 6.0)
# Comparing skew normal and standard normal
n = 10000
rv_std_normal = np.random.normal(size=n)
rv_skew_normal = stats.skewnorm.rvs(a=5, size=n)
fig, ax = plt.subplots(1, 3, figsize=(15,5))
ProbPlot(rv_std_normal).qqplot(line='s', ax=ax[0])
ax[0].set_title('Q-Q plot for std. normal - std. normal')
ProbPlot(rv_skew_normal).qqplot(line='s', ax=ax[1])
ax[1].set_title('Q-Q plot for skew normal - std. normal')
sns.histplot(rv_skew_normal, kde=False, label='Skew normal', ax=ax[2])
sns.histplot(rv_std_normal, kde=False, label='Std. normal', ax=ax[2])
ax[2].set_title('Histograms')
ax[2].legend();
# -
# As expected, data from a standard normal distribution fits almost perfectly onto standard normal quantiles, while data from our positively skewed distribution does not -- it has more probability mass for lower values, as well as more extreme higher values.
# +
# Comparing Google stock returns to standard normal
# import os
# import pandas_datareader as pdr
# from dotenv import load_dotenv
# from datetime import datetime
# load_dotenv()
# start = datetime(2019, 1, 1)
# end = datetime(2019, 12, 31)
# key = os.getenv('tiingo_api_key')
# goog = np.log(pdr.get_data_tiingo('GOOG', start, end, api_key=key)['close']).diff().dropna()
# fix, ax = plt.subplots(1, 2)
# ProbPlot(nflx).qqplot(line='s', ax=ax[0])
# ax[0].set_title('Q-Q plot for Google returns - std. normal')
# sns.distplot(nflx, norm_hist=True, ax=ax[1]);
# -
# The above graph shows clearly that Google's daily stock returns are not normally distributed. While the inner part of the distribution fits a normal distribution relatively well, the returns distribution has (very) fat tails.
# Chi-Squared distribution
#
# - To assess goodness of fit.
# F distribution
#
# - Can be used to measure whether means of different treatment groups differ from control condition.
# - F-statistic is calculated as the ratio of the variance between groups and the variance within groups (ANOVA).
# - F distribution gives all values that would be produced if between variance were zero (i.e. under the null model).
# - Df is given by the number of groups we compare.
# Poisson distribution
#
# - Useful to model processes that randomly generate outcomes at a constant rate (e.g. processes like arrivals that vary over time, or number of defects or typos that vary over space).
# - The parameter of the distribution is lambda, which is both the rate per unit of time and the variance.
# - The poisson and exponential distribution can be very useful when modelling, say, arrivals and waiting times. It's important, though, to remember the three key assumptions: 1) lambda remains constant across intervals, 2) events are independent, and 3) two events cannot occur at the same time.
# - To account for 1), defining the intervals such that they are sufficiently homogenous often helps.
# +
# Comparing Poisson distributions
x = np.random.poisson(2, 1000000)
y = np.random.poisson(6, 1000000)
plt.hist(x, alpha=0.5, label='$\\lambda = 2$', bins=np.arange(min(x), max(x))-0.5)
plt.hist(y, alpha=0.5, label='$\\lambda = 6$', bins=np.arange(min(y), max(y))-0.5)
plt.legend();
# -
# Exponential distribution
#
# - Takes the same parameter lambda as the Poisson distribution, but can be used to model the time between random events occuring at a frequent rate lambda (i.e. the time/space difference between Poisson events).
# +
# Comparing exponential distributions
n = 100000
x = np.random.exponential(2, n)
y = np.random.exponential(6, n)
plt.hist(x, alpha=0.5, label='$\\lambda = 2$', bins=np.arange(min(x), max(x))-0.5)
plt.hist(y, alpha=0.5, label='$\\lambda = 6$', bins=np.arange(min(y), max(y))-0.5)
plt.legend();
# -
# Weibull distribution
#
# - Used to model events for which the event rate changes during the time of the interval, and thus violates the poisson and exponential assumption.
# - An example is mechanical failure, where the probability of failure increases as time goas by.
# - Parameters of the distribution are $\eta$, the scale parameter, and $\beta$, the shape parameter ($\beta > 1$ indicates increasing probability of an event over time, $\beta < 1$ decreasing probability).
# Plotting distributions in Seaborn
# +
# Generating random samples
n = 10000
rv_std_normal = np.random.normal(size=n)
rv_normal = np.random.normal(1, 2.5, n)
rv_skew_normal = stats.skewnorm.rvs(a=5, size=n)
# +
# Drawing histogram, pdf, and cdf of std normal sample
x = np.linspace(min(rv_std_normal), max(rv_std_normal), 1000);
pdf = stats.norm.pdf(x)
cdf = stats.norm.cdf(x)
ax = sns.distplot(rv_std_normal, kde=False, norm_hist=True, label='Data')
ax.plot(x, pdf, lw=2, label='PDF')
ax.plot(x, cdf, lw=2, label='CDF')
ax.set_title('Standard normal distribution')
ax.legend();
# +
# Compare three distributions
ax = sns.distplot(rv_std_normal, kde=False, norm_hist=True, label='Standard normal')
ax = sns.distplot(rv_normal, kde=False, norm_hist=True, label='N(1, 2.5)')
ax = sns.distplot(rv_skew_normal, kde=False, norm_hist=True, label='Skew normal, $\\alpha$=5')
ax.set_title('Comparison of different distributions')
ax.legend();
| content/post/basic-stats/.ipynb_checkpoints/basic-stats-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Surrogate model
#
#
# The objective of this work is to develop a surrogate model for the Rayleigh-Bénard convection problem, which is a type of natural convection occurring in a horizontal plane layer of fluid heated from the bottom. The surrogate model to be developed will be based on Neural Networks and dimensionality reduction. Thus, it is necessary to have data that describes the flow in the problem to train the neural networks.
#
# The data will be generated using the [OpenLB](https://www.openlb.net/) library, which is a library that implements the lattice Boltzmann method. Therefore, different conditions for the Rayleigh-Bénard convection will be simulated using this library, and then, the simulated output will be used to train the neural networks. The Rayleigh-Bénard convection problem simulation using the OpenLB is shown.
#
# 
#
# The implementation of the Rayleigh-Bénard convection problem used was an adaptation from the examples of the OpenLB. It was adapted to accept the Rayleigh number, Prantdl number and wall and fluid temperature as input and it can be accessed
# at [`rayleighBenard2d.cpp`](rayleighBenard2d.cpp)
#
#
# ## Test matrix
#
#
# It is known from the design of experiments that the pseudorandom sampling in higher dimensional spaces does not generate a uniform projections, and are use Latin Hypercube Sampling (LHS), Halton sequence or Sobol sequence are frequently used. The difference between the Sobol sequence and pseudorandom sampling is show in the figure below:
#
# Sobol sequence | Pseudorandom sampling
# :-------------------------:|:-------------------------:
#  | 
#
# Fig.1 - Sobol sequence [[Wikipedia]](https://en.wikipedia.org/wiki/Sobol_sequence)
#
# To simulate the Rayleigh-Bénard convection problem using the OpenLB it is necessary the Rayleigh number, Prantdl number and wall and fluid temperature as input. Then, it was determined the range of each variable, which is presented below.
#
#
# | | Min. | Max. |
# |-------------------|----------------|-----------------|
# | Rayleigh number | $10^3$ | $10^6$ |
# | Prantdl number | $0.1$ | $70$ |
# | Fluid temperature | $10 C$ | $100 C$ |
# | $\Delta T$ | $1 C$ | $10 C$ |
#
#
# Finally, using the Sobol sequence with the ranges of the variables presented above, it was initially generated a test matrix of $30$ experiments. It is generated using the Python script [`test_matrix.py`](test_matrix.py).
#
#
# ### Run test matrix
#
#
# The [`test_matrix.py`](test_matrix.py) script generates a JSON file type called `args.json` that contains the experiments to be performed. This file is then used as an argument for the Python script [`multi_thread.py`](multi_thread.py). It runs runs the simulations using the compiled version of the[`rayleighBenard2d.cpp`](rayleighBenard2d.cpp), and automatically convert the OpenLB output format, `.vtk`, to an HDF5 file.
#
# The [`multi_thread.py`](multi_thread.py) script arguments are:
#
# * `save_file`: file to save the simulations;
# * `case_name`: where the OpenLB simulations are being stored;
# * `command`: command to run the simulation;
# * `arg_file`: the path to a JSON file containing the parameters for the simulations;
# * `n_threads`: number of threads to use.
#
# An example of the command to run the simulation:
#
# ```bash
# python multi_thread.py data.h5 rayleighBenard2d ./rayleighBenard2d args.json 2
# ```
#
#
# ### Preprocessing the test matrix
#
#
# As commonly known, the preprocessing data step is important for neural networks and machine learning algorithms. Thus, after generated the HDF5 file by [`multi_thread.py`](multi_thread.py) script, the data was preprocessed to remove cases where there were positive or negative infinity values or not a number values in any of the time steps of a given simulation. Moreover, after pruning the cases with invalid values, the remained cases were standardized, according to:
#
# $$x_{std} = \frac{x - \mu}{\sigma},$$
#
# where $\mu$ and $\sigma$ are the mean and standard deviation of $x$. These steps are executed by the Python script [`pre_proc_open_lb.py`](pre_proc_open_lb.py). It outputs a new HDF5 file where the scaled and unscaled data are stored. The script mix the data from all study cases as a single dataset. In this case, the dataset has four dimensions, the first dimension is the sample, the second and third are the spatial distribution and the third the problem output variables, which are in this order pressure, temperature and velocity. After preprocessed the simulated data from OpenLB, $16$ simulations were found to have some problems related to positive or negative infinity values or not a number values. Therefore, they were removed from the analysis, and only $14$ were kept.
#
#
# ## Dimensionality reduction
#
#
# The dimensionality reduction development can be found at [`Autoencoder.ipynb`](Autoencoder.ipynb). In there, a few neural networks architectures are evaluated and compared for the dimensionality reduction problem for the Rayleigh-Bénard problem. The architectures evaluated were:
#
# 1. [Convolutional autoencoder](./tests/jupyter-notebooks/train_ae_conv.ipynb);
# 2. [Convolutional autoencoder with dropout](./tests/jupyter-notebooks/train_ae_conv_drop.ipynb);
# 3. [VQ-VAE-2 based autoencoder](./tests/jupyter-notebooks/train_ae_add.ipynb);
# 4. [Depthwise convolutional autoencoder](./tests/jupyter-notebooks/train_ae_depth.ipynb);
# 5. [Dual convolutional autoencoder](./tests/jupyter-notebooks/train_dual_model.ipynb).
#
# The development of the hyperparmeter optimisation for the autoencoder neural networks can be found at [`Autoencoder_Hyperparameter_Optimisation.ipynb`](Autoencoder_Hyperparameter_Optimisation.ipynb).
#
# Also, there are the Python script files that uses the [Independent Component Analysis](dimensionality-reduction/ica.py), [Kernel PCA](dimensionality-reduction/kernel_pca.py) and [PCA](dimensionality-reduction/pca.py) for dimensionality reduction. They are designed to run using [Visual Studio Code](https://code.visualstudio.com/) or the [Spyder](https://www.spyder-ide.org/).
#
#
# ## Surrogate model development
#
#
# The development of the surrogate model starts by evaluating the prediction performance of a FCNN, LSTM and Decoder Predictor neural networks, this part is developed at [`Prediction.ipynb`](Prediction.ipynb). After evaluated and their hyperparameter optmised, the surrogate models are assembled and evaluated at [`Surrogate_model.ipynb`](Surrogate_model.ipynb).
#
#
# ## Workflow
#
#
# The proposed surrogate models follows the workflow presented below:
#
# 1. Determine the test matrix;
# * Run the Python script [`test_matrix.py`](test_matrix.py);
# 2. Simulate the test matrix:
# * Run the Python script [`multi_thread.py`](multi_thread.py);
# 3. Preprocess the simulated test matrix:
# * Run the Python script [`pre_proc_open_lb.py`](pre_proc_open_lb.py);
# 4. Train, analyse and optmise the Autoencoder neural networks for dimensionality reduction:
# * Overall analysis in a Python notebook [`Autoencoder.ipynb`](Autoencoder.ipynb);
# * Hyperparameter optmisation in a Python notebook: [`Autoencoder_Hyperparameter_Optimisation.ipynb`](Autoencoder_Hyperparameter_Optimisation.ipynb);
# 5. Analyse and build the surrogate models:
# * Overall analysis of the predictors in a Python notebook [`Prediction.ipynb`](Prediction.ipynb);
# * Final surrogate models in a Python notebook [`Surrogate_model.ipynb`](Surrogate_model.ipynb).
#
#
# # Extras
#
#
# The notebooks used to train the autoencoder neural networks are also available in a Python script file. They can be used with [Visual Studio Code](https://code.visualstudio.com/) as a notebook. They are found at `tests` folder.
#
# Some examples of how to read the `.vtk` or how to run a process in parallel can be found at the folder `examples`.
#
# where 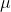 and 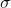 are the mean and standard deviation of $x$. These steps are executed by the Python script [`pre_proc_open_lb.py`](pre_proc_open_lb.py). It outputs a new HDF5 file where the scaled and unscaled data are stored. The script mix the data from all study cases as a single dataset. In this case, the dataset has four dimensions, the first dimension is the sample, the second and third are the spatial distribution and the third the problem output variables, which are in this order pressure, temperature and velocity. After preprocessed the simulated data from OpenLB, $16$ simulations were found to have some problems related to positive or negative infinity values or not a number values. Therefore, they were removed from the analysis, and only $14$ were kept.
#
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## DataFrame
# A DataFrame represents a rectangular table of data and contains an ordered collec‐
# tion of columns, each of which can be a different value type (numeric, string,
# boolean, etc.). The DataFrame has both a row and column index; it can be thought of
# as a dict of Series all sharing the same index. Under the hood, the data is stored as one
# or more two-dimensional blocks rather than a list, dict, or some other collection of
# one-dimensional arrays
# There are many ways to construct a DataFrame, though one of the most common is
# from a dict of equal-length lists or NumPy arrays:
# +
import pandas as pd
data = {'state': ['Ohio', 'Ohio', 'Ohio', 'Nevada', 'Nevada', 'Nevada'],
'year': [2000, 2001, 2002, 2001, 2002, 2003],
'pop': [1.5, 1.7, 3.6, 2.4, 2.9, 3.2]}
frame = pd.DataFrame(data)
frame
# -
# Selecting the head and the tail of the objects
#
frame.head()# default, no parameter pass gives first five heads in the data. You specify the no of heads to get
frame.tail(3)
# If you specify a sequence of columns, the DataFrame’s columns will be arranged in
# that order:
pd.DataFrame(data, columns=[ 'state', 'pop','year'])
# Let's change the default index to something better
# +
pd.DataFrame(data, columns=[ 'state', 'pop','year'], index = ['245','244','765', '897','765','897'])
# -
# Access columns
Frame1 = pd.DataFrame(data, columns=[ 'state', 'pop','year'], index = ['245','244','765', '897','765','897'])
# Frame1=['state']
Frame1.state
#Rows can also be retrieved by position or name with the special loc attribute
Frame1.loc['245']
# Columns can be modified by assignment. For example, the empty 'debt' column
# could be assigned a scalar value or an array of values:
# +
import numpy as np
Frame1 = pd.DataFrame(data, columns=[ 'state', 'pop','year'], index = ['245','244','765', '897','765','897'])
Frame1.index.name = "country code"#name the index
Frame1['debt'] = np.arange(6.)# adding another colum
Frame1
| Python/Pandas/data_frames.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cmip6
# language: python
# name: cmip6
# ---
# # Day 2: Pearson Correlation Coefficient vs. Lead Time
# This notebook creates a plot for each observation source (GPCC, CHIRPS, MSWEP) showing the Pearson correlation coefficient for each lead time for each DCPP model. The significant coefficient values at 95th percentile level is shown as grey horizontal dashed lines. The same six panel format is used and explained in 00_DCPP_GPCC_visualisation.ipynb.
# +
import numpy as np
import xarray as xr
import glob as glob
import matplotlib.pyplot as plt
from scipy.stats import t
def calc_r_crit(n):
"""
:param int n: Number of data points used to calculate Pearson correlation coefficient
:return tuple: Lower critical correlation coefficient, Upper critical correlation coefficient
"""
t_upper = t.ppf(0.975, df=n-1)
t_lower = t.ppf(0.025, df=n-1)
# https://en.wikipedia.org/wiki/Pearson_correlation_coefficient#Testing_using_Student's_t-distribution
r_crit_upper = t_upper / np.sqrt(n - 2 + t_upper**2)
r_crit_lower = t_lower / np.sqrt(n - 2 + t_lower**2)
return r_crit_lower, r_crit_upper
model_namelist = ['CanESM5', 'EC-Earth3', 'HadGEM3-GC31-MM', 'IPSL-CM6A-LR', 'MIROC6', 'MPI-ESM1-2-HR', 'NorCPM1']
obs_namelist = ['GPCC', 'CHIRPS', 'MSWEP']
dcpp_trbindices_directory = '/gws/pw/j05/cop26_hackathons/bristol/project02/data/DCPP/TRBindices'
obs_trbindices_directory = '/gws/pw/j05/cop26_hackathons/bristol/project02/data/obs/TRBindices'
for obs_name in obs_namelist:
fig, axs = plt.subplots(3, 2, figsize=(10,12), sharey=True)
lead_time_years = np.arange(1,11,1)
obs_filelist = glob.glob(obs_trbindices_directory+'/'+obs_name+'/*.nc')
obs_filelist.sort()
# Selection conditions
obs_data = xr.open_mfdataset(obs_filelist)
obs_data_indices = obs_data.gaussian_params
is_djf = (obs_data_indices['time'].dt.season == 'DJF')
is_jja = (obs_data_indices['time'].dt.season == 'JJA')
obs_data_djf = obs_data_indices.isel(time=is_djf)
obs_data_jja = obs_data_indices.isel(time=is_jja)
obs_data_mean_djf = obs_data_djf.groupby('time.year').mean('time')
obs_data_mean_djf_1981_2016 = obs_data_mean_djf.sel(year=slice(1981,2016))
obs_data_mean_djf_1981_2016_overallmean = obs_data_mean_djf_1981_2016.mean(dim='year')
obs_data_mean_djf_1981_2016_overallstdev = obs_data_mean_djf_1981_2016.std(dim='year', ddof=1)
obs_data_mean_djf_1981_2016_zscores = (obs_data_mean_djf_1981_2016 - obs_data_mean_djf_1981_2016_overallmean)/obs_data_mean_djf_1981_2016_overallstdev
obs_data_mean_jja = obs_data_jja.groupby('time.year').mean('time')
obs_data_mean_jja_1981_2016 = obs_data_mean_jja.sel(year=slice(1981,2016))
obs_data_mean_jja_1981_2016_overallmean = obs_data_mean_jja_1981_2016.mean(dim='year')
obs_data_mean_jja_1981_2016_overallstdev = obs_data_mean_jja_1981_2016.std(dim='year', ddof=1)
obs_data_mean_jja_1981_2016_zscores = (obs_data_mean_jja_1981_2016 - obs_data_mean_jja_1981_2016_overallmean)/obs_data_mean_jja_1981_2016_overallstdev
# Calculate critical r values
r_lower, r_upper = calc_r_crit(obs_data_mean_jja_1981_2016_zscores.shape[0])
for model_name in model_namelist:
print(model_name)
model_filelist = glob.glob(dcpp_trbindices_directory+'/'+model_name+'/*.nc')
model_filelist.sort()
zscores_list_djf = list()
zscores_list_jja = list()
for lead_time_year in lead_time_years:
model_ensembles_all_djf = list()
model_ensembles_all_jja = list()
# Combining all forecasts
for model_file in model_filelist:
model_dataset = xr.open_dataset(model_file)
# Selection conditions
is_djf = (model_dataset['time'].dt.season == 'DJF')
is_jja = (model_dataset['time'].dt.season == 'JJA')
model_ensemble_mean = model_dataset.gaussian_params.mean(dim='ensemble')
model_ensemble_mean_djf = model_ensemble_mean.isel(time=is_djf)
model_ensemble_mean_djf = model_ensemble_mean_djf.groupby('time.year').mean('time').isel(year=lead_time_year-1)
model_ensembles_all_djf.append(model_ensemble_mean_djf)
model_ensemble_mean_jja = model_ensemble_mean.isel(time=is_jja)
model_ensemble_mean_jja = model_ensemble_mean_jja.groupby('time.year').mean('time').isel(year=lead_time_year-1)
model_ensembles_all_jja.append(model_ensemble_mean_jja)
lead_time_time_series_djf = xr.concat(model_ensembles_all_djf, dim='year').sel(year=slice(1981,2016))
lead_time_time_series_djf_overallmean = lead_time_time_series_djf.mean(dim='year')
lead_time_time_series_djf_stdev = lead_time_time_series_djf.std(dim='year', ddof=1)
lead_time_time_series_djf_zscore = (lead_time_time_series_djf - lead_time_time_series_djf_overallmean) / lead_time_time_series_djf_stdev
lead_time_time_series_jja = xr.concat(model_ensembles_all_jja, dim='year').sel(year=slice(1981,2016))
lead_time_time_series_jja_overallmean = lead_time_time_series_jja.mean(dim='year')
lead_time_time_series_jja_stdev = lead_time_time_series_jja.std(dim='year', ddof=1)
lead_time_time_series_jja_zscore = (lead_time_time_series_jja - lead_time_time_series_jja_overallmean) / lead_time_time_series_jja_stdev
zscores_this_leadtime_djf = xr.corr(obs_data_mean_djf_1981_2016_zscores, lead_time_time_series_djf_zscore, dim='year')
zscores_list_djf.append(zscores_this_leadtime_djf)
zscores_this_leadtime_jja = xr.corr(obs_data_mean_jja_1981_2016_zscores, lead_time_time_series_jja_zscore, dim='year')
zscores_list_jja.append(zscores_this_leadtime_jja)
this_model_correlation_djf = xr.concat(zscores_list_djf, dim='lead_time')
this_model_correlation_jja = xr.concat(zscores_list_jja, dim='lead_time')
axs[0,0].scatter(lead_time_years, this_model_correlation_djf[:,0], label=model_name, s=20.0)
axs[1,0].scatter(lead_time_years, this_model_correlation_djf[:,1], label=model_name, s=20.0)
axs[2,0].scatter(lead_time_years, this_model_correlation_djf[:,2], label=model_name, s=20.0)
axs[0,1].scatter(lead_time_years, this_model_correlation_jja[:,0], label=model_name, s=20.0)
axs[1,1].scatter(lead_time_years, this_model_correlation_jja[:,1], label=model_name, s=20.0)
axs[2,1].scatter(lead_time_years, this_model_correlation_jja[:,2], label=model_name, s=20.0)
axs[0,1].legend(bbox_to_anchor=(1.05, 1.03), loc='upper left')
axs[0,0].set_title('Rainfall Intensity DJF (mm/day)')
axs[0,0].set_xticks(lead_time_years)
axs[0,0].axhline(r_lower, linestyle='dashed', color='grey')
axs[0,0].axhline(r_upper, linestyle='dashed', color='grey')
axs[1,0].set_title(r'TRB Mean Location DJF ($^\circ$)')
axs[1,0].set_xticks(lead_time_years)
axs[1,0].axhline(r_lower, linestyle='dashed', color='grey')
axs[1,0].axhline(r_upper, linestyle='dashed', color='grey')
axs[2,0].set_title(r'TRB Width DJF ($^\circ$)')
axs[2,0].set_xticks(lead_time_years)
axs[2,0].axhline(r_lower, linestyle='dashed', color='grey')
axs[2,0].axhline(r_upper, linestyle='dashed', color='grey')
axs[2,0].set_xlabel('Lead Time (Years)')
axs[0,1].set_title('Rainfall Intensity JJA (mm/day)')
axs[0,1].set_xticks(lead_time_years)
axs[0,1].axhline(r_lower, linestyle='dashed', color='grey')
axs[0,1].axhline(r_upper, linestyle='dashed', color='grey')
axs[1,1].set_title(r'TRB Mean Location JJA ($^\circ$)')
axs[1,1].set_xticks(lead_time_years)
axs[1,1].axhline(r_lower, linestyle='dashed', color='grey')
axs[1,1].axhline(r_upper, linestyle='dashed', color='grey')
axs[2,1].set_title(r'TRB Width JJA ($^\circ$)')
axs[2,1].set_xlabel('Lead Time (Years)')
axs[2,1].set_xticks(lead_time_years)
axs[2,1].axhline(r_lower, linestyle='dashed', color='grey')
axs[2,1].axhline(r_upper, linestyle='dashed', color='grey')
fig.suptitle(f"DCPP to {obs_name} Pearson Correlation (1981 - 2016)", fontsize=16, y=0.95)
plt.show()
# plt.savefig(f'plots/pearson_correlation_analysis/correlation_DCPP_to_{obs_name}.pdf', bbox_inches='tight')
# -
| notebooks/subgroup3/04_CorrelationLeadTime.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import yt
import numpy as np
import matplotlib.pyplot as plt
from galaxy_analysis import Galaxy
colors = {'massive_star_winds' : 'black', 'AGB_winds' : 'C1', 'SN' : 'C4', 'other_stars' : 'black'}
markers = {'massive_star_winds' : '*', 'AGB_winds' : 'D', 'SN' : '*', 'other_stars' : '.'}
ps = {'massive_star_winds' : 20, 'AGB_winds' : 100, 'SN' : 40, 'other_stars' : 10}
# +
#ds = yt.load('./../example_data/DD0401/DD0401')
#data = ds.all_data()
gal = Galaxy('DD0401', wdir = './../example_data/')
# -
data = gal.df
gal.ds.derived_field_list
# +
M = data['birth_mass']
t_o = data['creation_time'].convert_to_units('Myr')
MS_lifetime = data[('io','particle_model_lifetime')].to('Myr')
MS_death = t_o + MS_lifetime
px = (data['particle_position_x'] - gal.ds.domain_center[0]).to('pc')
py = (data['particle_position_y'] - gal.ds.domain_center[1]).to('pc')
pz = (data['particle_position_z'] - gal.ds.domain_center[2]).to('pc')
recent_death = (MS_death > ds.current_time - dt) * (MS_death <= ds.current_time + 0.001*yt.units.Myr)
alive = MS_death > ds.current_time + 0.001*yt.units.Myr
AGB = M < 8.0
massive_star = (M > 8.0) * (M < 25.0)
# +
width = 500.0
boxdim = np.array([width*1.25,width*1.25,30.0])*yt.units.pc
region = gal.ds.box(ds.domain_center - boxdim*0.5, ds.domain_center + boxdim*0.5)
proj = yt.ProjectionPlot(gal.ds, 'z', ['number_density','N_over_O','O_over_H'],
weight_field = 'number_density', data_source = region, width = (width,'pc'))
proj.set_unit('number_density','cm**(-3)')
proj.set_cmap('number_density','viridis')
proj.set_cmap('N_over_O','magma')
proj.set_zlim('N_over_O',-2,2)
proj.set_log('N_over_O',False)
proj.set_cmap('O_over_H','cubehelix')
proj.set_zlim('O_over_H',-5,0)
proj.set_log('O_over_H',False)
dt = 5.0 * yt.units.Myr
in_image = (np.abs(pz) <= boxdim[2]*0.5) * (np.abs(px) <= width*0.5) * (np.abs(py) <= width*0.5)
proj.show()
# +
pp = {}
pp['massive_star_winds'] = in_image * alive * massive_star
pp['AGB_winds'] = in_image * recent_death * AGB
pp['SN'] = in_image * recent_death * massive_star
#pp['other_stars'] = in_image * alive * (np.logical_not(pp['massive_star_winds']))
for k in proj.plots.keys():
image = proj.plots[k]
#
# Now select and annotate the points we want
#
for s in pp.keys():
if np.size(px[pp[s]].value) > 0:
print np.size(px[pp[s]]), 'Particles in ', s, px[pp[s]], py[pp[s]]
image.axes.scatter(px[pp[s]].value,py[pp[s]].value, s = ps[s], marker = markers[s], color = colors[s])
else:
print 'No particles in ', s
proj.show()
# -
np.min(data['particle_position_x'])
| notebooks/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: widgets-tutorial
# language: python
# name: widgets-tutorial
# ---
# <!--NAVIGATION-->
# < [Jupyter Widgets Tutorial Introduction](00.00-introduction.ipynb) | [Contents](00.00-index.ipynb) | [Widgets without writing widgets: interact](02.00-Using-Interact.ipynb) >
# # Overview
#
# What you can accomplish with just Python has increased quite a bit in the last years as more sophisticated tools that plug in to the Jupyter widget ecosystem have been written.
#
# One of those tools is [bqplot](https://github.com/bloomberg/bqplot/blob/master/examples/Index.ipynb), which provides a plotting tool in which the plot, and the lines, markers, labels and legend, all act as widgets. That required both Python *and* JavaScript. On the JavaScript side bqplot uses [d3](https://d3js.org/) to do the drawing in the browser.
#
# The widely-used plotting library [matplotlib](https://matplotlib.org/3.2.2/contents.html) also has a widget interface. Use `%matplotlib widget` in the notebook to have interactive plots that are widgets. For more control, look at the documentation for [ipympl](https://github.com/matplotlib/ipympl) for more details on using it as a widget.
#
# Another example is [ipyvolume](https://ipyvolume.readthedocs.io/en/latest/), which does three-dimensional renderings of point or volumetric data in the browser. It has both Python and JavaScript pieces but using requires only Python.
#
# One last addition is in `ipywidgets` itself: the new `Output` widget can display any content which can be rendered in a Jupyter notebook. That means that anything you can show in a notebook you can include in a widget using only Python.
# ## Example 1: COVID dashboard (pure Python)
#
# + Dashboard: http://jupyter.mnstate.edu/COVID
# + Code: https://github.com/JuanCab/COVID_DataViz (see `Dashboard.ipynb`)
#
# Orange boxes are [ipympl](); magenta box is [ipyleaflet](https://ipyleaflet.readthedocs.io/en/latest/); remaining widgets are from [ipywidgets](https://ipywidgets.readthedocs.io/en/stable/).
#
# | | |
# |----|----|
#  | 
# ## Example 2: Binary star simulation (pure Python)
#
# + Green: [pythreejs](https://github.com/jupyter-widgets/pythreejs)
# + Blue: [bqplot](https://github.com/bloomberg/bqplot/blob/master/examples/Index.ipynb)
# + Everything else: [ipywidgets](https://github.com/jupyter-widgets/ipywidgets)
# + Serving it up to users during development on [mybinder.org](https://mybinder.org/)
#
# 
#
# ### Source for this example (including links to binder): https://github.com/JuanCab/AstroInteractives
#
# [Video](https://youtu.be/kbgST0uifvM)
# ## Remainder of this tutorial
#
# + Widget basics and core ipywidgets
# + Widget styling and layout
# + Widget events
# + Other widget libraries
# <!--NAVIGATION-->
# < [Jupyter Widgets Tutorial Introduction](00.00-introduction.ipynb) | [Contents](00.00-index.ipynb) | [Widgets without writing widgets: interact](02.00-Using-Interact.ipynb) >
| notebooks/01.00-overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm
import pandas as pd
import seaborn as sns
from sklearn import svm
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn import neighbors, datasets
from sklearn.model_selection import cross_val_score
from sklearn.datasets import make_blobs
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import ExtraTreesClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.ensemble import GradientBoostingClassifier
from scipy.spatial import ConvexHull
from tqdm import tqdm
import random
plt.style.use('ggplot')
import pickle
from sklearn import tree
from sklearn.tree import export_graphviz
from joblib import dump, load
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
# %matplotlib inline
from sklearn.impute import SimpleImputer
# +
def getAuc(X,y,test_size=0.25,max_depth=None,n_estimators=100,
minsplit=4,FPR=[],TPR=[],VERBOSE=False, USE_ONLY=None):
'''
get AUC given training data X, with target labels y
'''
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size)
CLASSIFIERS=[DecisionTreeClassifier(max_depth=max_depth, min_samples_split=minsplit,class_weight='balanced'),
RandomForestClassifier(n_estimators=n_estimators,
max_depth=max_depth,min_samples_split=minsplit,class_weight='balanced'),
ExtraTreesClassifier(n_estimators=n_estimators,
max_depth=max_depth,min_samples_split=minsplit,class_weight='balanced'),
AdaBoostClassifier(n_estimators=n_estimators),
GradientBoostingClassifier(n_estimators=n_estimators,max_depth=max_depth),
svm.SVC(kernel='rbf',gamma='scale',class_weight='balanced',probability=True)]
if USE_ONLY is not None:
if isinstance(USE_ONLY, (list,)):
CLASSIFIERS=[CLASSIFIERS[i] for i in USE_ONLY]
if isinstance(USE_ONLY, (int,)):
CLASSIFIERS=CLASSIFIERS[USE_ONLY]
for clf in CLASSIFIERS:
clf.fit(X_train,y_train)
y_pred=clf.predict_proba(X_test)
#print(X_test,y_pred)
fpr, tpr, thresholds = metrics.roc_curve(y_test,y_pred[:,1], pos_label=1)
auc=metrics.auc(fpr, tpr)
if VERBOSE:
print(auc)
FPR=np.append(FPR,fpr)
TPR=np.append(TPR,tpr)
points=np.array([[a[0],a[1]] for a in zip(FPR,TPR)])
hull = ConvexHull(points)
x=np.argsort(points[hull.vertices,:][:,0])
auc=metrics.auc(points[hull.vertices,:][x,0],points[hull.vertices,:][x,1])
return auc,CLASSIFIERS
def saveFIG(filename='tmp.pdf',AXIS=False):
'''
save fig for publication
'''
import pylab as plt
plt.subplots_adjust(top = 1, bottom = 0, right = 1, left = 0,
hspace = 0, wspace = 0)
plt.margins(0,0)
if not AXIS:
plt.gca().xaxis.set_major_locator(plt.NullLocator())
plt.gca().yaxis.set_major_locator(plt.NullLocator())
plt.savefig(filename,dpi=300, bbox_inches = 'tight',
pad_inches = 0,transparent=False)
return
# +
def getCoverage(model,verbose=True):
'''
return how many distinct items (questions)
are used in the model set.
This includes the set of questions being
covered by all forms that may be
generated by the model set
'''
FS=[]
for m in model:
for count in range(len(m.estimators_)):
clf=m.estimators_[count]
fs=clf.tree_.feature[clf.tree_.feature>0]
FS=np.array(list(set(np.append(FS,fs))))
if verbose:
print("Number of items used: ", FS.size)
return FS
def getConfusion(X,y,test_size=0.25,max_depth=None,n_estimators=100,
minsplit=4,CONFUSION={},VERBOSE=False, USE_ONLY=None,target_names = None):
'''
get AUC given training data X, with target labels y
'''
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size)
CLASSIFIERS=[DecisionTreeClassifier(max_depth=max_depth, min_samples_split=minsplit),
RandomForestClassifier(n_estimators=n_estimators,class_weight='balanced',
max_depth=max_depth,min_samples_split=minsplit),
ExtraTreesClassifier(n_estimators=n_estimators,class_weight='balanced',
max_depth=max_depth,min_samples_split=minsplit),
AdaBoostClassifier(n_estimators=n_estimators),
GradientBoostingClassifier(n_estimators=n_estimators,max_depth=max_depth),
svm.SVC(kernel='rbf',gamma='scale',class_weight='balanced',probability=True)]
if USE_ONLY is not None:
if isinstance(USE_ONLY, (list,)):
CLASSIFIERS=[CLASSIFIERS[i] for i in USE_ONLY]
if isinstance(USE_ONLY, (int,)):
CLASSIFIERS=CLASSIFIERS[USE_ONLY]
for clf in CLASSIFIERS:
clf.fit(X_train,y_train)
y_pred=clf.predict(X_test)
print(y_test,y_pred)
cmat=confusion_matrix(y_test, y_pred)
acc=accuracy_score(y_test, y_pred)
CONFUSION[clf]=cmat
if VERBOSE:
print(classification_report(y_test, y_pred, target_names=target_names))
print('Confusion MAtrix:\n', cmat)
print(' ')
print('Accuracy:', acc)
return CONFUSION,acc
# -
df=pd.read_csv('bsnip.csv',index_col=0)
df.head()
df.Biotype.value_counts()
# 3 is HC
#df=df[df['Biotype']==3]
df=df.dropna()
df0=df
#df=df0[df0.Biotype.isin([1,5])]
df=df0
X=df.iloc[:,2:].values
y=df.Biotype.values#.astype(str)
y=[(int(x)==5)+0 for x in y ]
CF,acc=getConfusion(X,y,test_size=0.2,max_depth=None,n_estimators=500,
minsplit=2,CONFUSION={},VERBOSE=False, USE_ONLY=[2],target_names = None)
CF
ACC=[]
CLFh={}
for run in tqdm(np.arange(500)):
auc,CLFS=getAuc(X,y,test_size=0.2,max_depth=10,n_estimators=2,
minsplit=2,VERBOSE=False, USE_ONLY=[2])
ACC=np.append(ACC,auc)
if auc > 0.75:
CLFh[auc]=CLFS
sns.distplot(ACC)
np.median(ACC)
CLFstar=CLFh[np.array([k for k in CLFh.keys()]).max()][0]
from scipy import interpolate
from scipy.interpolate import interp1d
auc_=[]
ROC={}
fpr_ = np.linspace(0, 1, num=20, endpoint=True)
for run in np.arange(1000):
clf=CLFstar
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5)
y_pred=clf.predict_proba(X_test)
fpr, tpr, thresholds = metrics.roc_curve(y_test,y_pred[:,1], pos_label=1)
f = interp1d(fpr, tpr)
auc_=np.append(auc_,metrics.auc(fpr_, f(fpr_)))
ROC[metrics.auc(fpr, tpr)]={'fpr':fpr_,'tpr':f(fpr_)}
sns.distplot(auc_)
auc_.mean()
TPR=[]
for a in ROC.keys():
#print(a)
#break
plt.plot(ROC[a]['fpr'],ROC[a]['tpr'],'-k',alpha=.05)
TPR=np.append(TPR,ROC[a]['tpr'])
TPR=TPR.reshape(int(len(TPR)/len(fpr_)),len(fpr_))
plt.plot(fpr_,np.median(TPR,axis=0),'-r')
metrics.auc(fpr_,np.median(TPR,axis=0))
plt.gca().set_title('B3 vs others')
plt.text(.6,.65,'AUC: '+str(metrics.auc(fpr_,np.median(TPR,axis=0)))[:5],color='r')
#plt.text(.6,.31,'AUC: '+str(metrics.auc(fpr_,np.median(tprA,axis=0)))[:5],color='b')
#plt.text(.6,.19,'AUC: '+str(metrics.auc(fpr_,np.median(tprB,axis=0)))[:5],color='g')
plt.gca().set_xlabel('1-specificity')
plt.gca().set_ylabel('sensitivity')
saveFIG('bsnip001_L5.pdf',AXIS=True)
| code/notebooks/bsnip001-Copy2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import xarray as xr
from os.path import join, abspath
import os
from scipy import quantile
RUN = abspath(os.getenv('RUN', '../models/12/test'))
data_2d_path = join(RUN, 'OUT_2D', '*.nc')
data_3d_path = join(RUN, 'OUT_3D', '*.nc')
data_stat_path = join(RUN, 'OUT_STAT', '*.nc')
print(data_2d_path)
data_2d = xr.open_mfdataset(data_2d_path)
data_3d = xr.open_mfdataset(data_3d_path)
stat = xr.open_mfdataset(data_stat_path)
data_3d['FQTNN'] *= 86400
data_3d['FQTNN'].attrs['units'] = 'g/kg/d'
data_3d['FSLINN'] *= 86400
data_3d['FSLINN'].attrs['units'] = 'K/d'
# -
data_2d.PW[::12].plot(col='time', col_wrap=3)
data_2d.W500[::12].plot(col='time', col_wrap=3)
# # Forcing in a Single Location
for loc in [dict(x=0, y=32), dict(x=0, y=10)]:
for key in ['FQTNN', 'FSLINN', 'SLI', 'QT']:
plt.figure()
data_3d[key].isel(**loc).plot.contourf(x='time', levels=11)
# # Zonal-Means
data_3d['FQTNN']
# +
def get_plot_kwargs(x):
kwargs = {}
a,b = quantile(x, [0.02, .98])
if a * b < 0:
cmap = 'RdBu_r'
m = max(abs(a), abs(b))
a = -m
b = m
else:
cmap = 'viridis'
return dict(levels=np.linspace(a, b, 11), cmap=cmap)
for key in ['FQTNN', 'FSLINN', 'SLI', 'QT']:
plt.figure()
x = data_3d[key][::12].mean('x')
x.plot.contourf(col='time', **get_plot_kwargs(x))
| notebooks/templates/sam-run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AI-Frameworks
# <center>
# <a href="http://www.insa-toulouse.fr/" ><img src="http://www.math.univ-toulouse.fr/~besse/Wikistat/Images/logo-insa.jpg" style="float:left; max-width: 120px; display: inline" alt="INSA"/></a>
# <a href="http://wikistat.fr/" ><img src="http://www.math.univ-toulouse.fr/~besse/Wikistat/Images/wikistat.jpg" width=400, style="max-width: 150px; display: inline" alt="Wikistat"/></a>
# <a href="http://www.math.univ-toulouse.fr/" ><img src="http://www.math.univ-toulouse.fr/~besse/Wikistat/Images/logo_imt.jpg" width=400, style="float:right; display: inline" alt="IMT"/> </a>
# </center>
# # LAB 5 Introduction to Recommendation System with Collaborative Filtering - Part 3 : Latent Vector-Based Methods with `Keras` Python Library.
#
# The objectives of this notebook are the following :
#
# * Build Keras models to learn embedding space for user and item data.
# * Visualize these space.
# * Use results of algorithm to apply recommendation.
# # Library
# +
import numpy as np
import pandas as pd
import tensorflow.keras.layers as kl
import tensorflow.keras.models as km
import sklearn.metrics as sm
import sklearn.decomposition as sdec
import matplotlib.pyplot as plt
import seaborn as sb
# -
# # Data
#
# We download the updated ratings data generated in the first notebook. `1-Python-Neighborhood-MovieLens.ipynb`
DATA_DIR = "movielens_small/"
rating = pd.read_csv(DATA_DIR + "ratings_updated.csv")
nb_entries = rating.shape[0]
print("Number of entries : %d " %nb_entries)
rating.head(5)
# We first create two new columns. The column **user_id** (resp. **item_id**) rearange the userId (resp. MovieId) columns in order that these columns lies in the range(0,609) (resp. range(0,0723)).
userIdToNormUserId = {k:v for v,k in enumerate(rating.userId.unique())}
rating["user_id"] = [userIdToNormUserId[x] for x in rating.userId.values]
itemIdToNormItemId = {k:v for v,k in enumerate(rating.movieId.unique())}
rating["item_id"] = [itemIdToNormItemId[x] for x in rating.movieId.values]
rating.head()
movies = pd.read_csv(DATA_DIR + "movies.csv")
id_movie_to_title = dict(movies[["movieId","title"]].values)
id_item_to_title = {itemIdToNormItemId[k]:v for k,v in id_movie_to_title.items() if k in itemIdToNormItemId}
print("Number of movies in the dictionary : %d" %(len(id_item_to_title)))
movies.head()
# We now create the same train/test dataset that the one in the first notebook.
# +
train = rating[rating.test_train=="train"]
user_id_train = train['user_id']
item_id_train = train['item_id']
rating_train = train['rating']
print(train.shape)
test = rating[rating.test_train=="test"]
user_id_test = test['user_id']
item_id_test = test['item_id']
rating_test = test['rating']
print(test.shape)
# -
# # Neural Recommender System
#
# We first build a very simple recommender according to this architecture:
#
# 
#
# Let's decompose the construction of this network.
#
#
# We first create the inputs layer, which will take as entry the id of the user and the id of the item.
# For each sample we input the integer identifiers of a single user and a single item
user_id_input = kl.Input(shape=[1], name='user')
item_id_input = kl.Input(shape=[1], name='item')
# This id we will then be converted in their embedding space. This can be easily done with the `Embedding` layer object of Keras.
max_user_id= rating.user_id.max()
max_item_id= rating.item_id.max()
embedding_size = 30
user_embedding = kl.Embedding(output_dim=embedding_size, input_dim=max_user_id + 1,
input_length=1, name='user_embedding')(user_id_input)
item_embedding = kl.Embedding(output_dim=embedding_size, input_dim=max_item_id + 1,
input_length=1, name='item_embedding')(item_id_input)
# We compute the dot product of the two vectors which are the vectors representation in the embedding space of the user and the item given in input.
# +
# reshape from shape: (batch_size, input_length, embedding_size)
# to shape: (batch_size, input_length * embedding_size) which is
# equal to shape: (batch_size, embedding_size)
user_vecs = kl.Flatten()(user_embedding)
item_vecs = kl.Flatten()(item_embedding)
y = kl.Dot(axes=1)([user_vecs, item_vecs])
# -
# We now have the complete model.
model = km.Model(inputs=[user_id_input, item_id_input], outputs=y)
model.compile(optimizer='adam', loss='mse')
model.summary()
# The prediction can now be applied by giving the list of user and item ids that we want to compute.
initial_train_preds = model.predict([user_id_train, item_id_train])
initial_train_preds.shape
# Of course, as the model has not been traine, the Model error is quite bad.
print("Random init MSE: %0.3f" % sm.mean_squared_error(initial_train_preds, rating_train))
print("Random init MAE: %0.3f" % sm.mean_absolute_error(initial_train_preds, rating_train))
# Let's fit the model
history = model.fit([user_id_train, item_id_train], rating_train,
batch_size=64, epochs=10, validation_split=0.1,
shuffle=True)
# **Questions**:
#
# - Why is the train loss higher than the first loss in the first few epochs?
# - Why is Keras not computing the train loss on the full training set at the end of each epoch as it does on the validation set?
#
#
# Now that the model is trained, the model MSE and MAE look nicer:
test_preds = model.predict([user_id_test, item_id_test])
print("Final test MSE: %0.3f" % sm.mean_squared_error(test_preds, rating_test))
print("Final test MAE: %0.3f" % sm.mean_absolute_error(test_preds, rating_test))
train_preds = model.predict([user_id_train, item_id_train])
print("Final train MSE: %0.3f" % sm.mean_squared_error(train_preds, rating_train))
print("Final train MAE: %0.3f" % sm.mean_absolute_error(train_preds, rating_train))
# **Q** What do you think about those results?
# # A Deep recommender model
#
# Let's know compute a deeper architecture in order to improve those results.
#
# 
#
#
# **Exercise** : Implement a model similar to the previous one with:
#
# * A concatenate layer (look at the kl.Concatenate function)
# * A dropout layer (rate=0.5) after the concatenate layer.
# * only one Hidden layer with 64 neurons and relu activation function.
#
# +
# # %load solutions/exercise_3_1.py
# -
history = model.fit([user_id_train, item_id_train], rating_train,
batch_size=64, epochs=5, validation_split=0.1,
shuffle=True)
train_preds = model.predict([user_id_train, item_id_train])
print("Final train MSE: %0.3f" % sm.mean_squared_error(train_preds, rating_train))
print("Final train MAE: %0.3f" % sm.mean_absolute_error(train_preds, rating_train))
test_preds = model.predict([user_id_test, item_id_test])
print("Final test MSE: %0.3f" % sm.mean_squared_error(test_preds, rating_test))
print("Final test MAE: %0.3f" % sm.mean_absolute_error(test_preds, rating_test))
# **Question** What can you say about those results?
# # Exploiting the model
#
# In this section we will see how to explore both the model and the embedding space.
# ## Finding similar items and user.
#
# We want to find the K closest element of an item or a user. The model e build can't be used directly as it take into account a user and a item and not two user nor two items.
#
# But we can't easily build a method based on the constructed embedding space. Let's first get the embedding matices of the user and the movies.
weights = model.get_weights()
user_embeddings = weights[0]
print("User embedding matrix dimension : %s" %str(user_embeddings.shape))
item_embeddings = weights[1]
print("item embedding matrix dimension : %s" %str(item_embeddings.shape))
# For the id of an item we compute the distance (*cosine*, *euclidean*, etc.) of its embedding vector to all embedding vectors of the items.
#
# (The procedure would be the same for the user, but the results are easier to interpreted with the movies)
idx = 1027
X = np.expand_dims(item_embeddings[idx],axis=0)
distX = sm.pairwise_distances(X, item_embeddings, metric="cosine")[0]
# The top 10 items of the item "idx" are then the ten items that are the closest to this items.
print("Top 10 items similar to movies %s" %str(id_item_to_title[idx]))
mostSimilarItem = pd.DataFrame([[id_item_to_title[x], distX[x],x] for x in distX.argsort()[:10]])
mostSimilarItem
# **Question** What do you think of these results? Unfortunalty the dataset is to small to really get good meanings.
# ## Visualizing Items
pcaItems = sdec.PCA(n_components=2)
items_pca_embeddings = pcaItems.fit_transform(item_embeddings)
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(1,1,1)
ax.plot(items_pca_embeddings[:,0], item_embeddings[:,1], linestyle="None", marker=".")
ax.plot(items_pca_embeddings[mostSimilarItem[2].values,0], item_embeddings[mostSimilarItem[2].values,1], linestyle="None", marker=".", markersize=10)
# ## A recommendation function for a given user
#
# Once the model is trained, the system can be used to recommend a few items for a user, that he/she hasn't already seen:
#
# First let's select a user and display the movies he likes or dislikes.
user_id = 0
rating_user = rating[rating["user_id"]==user_id]
rating_user_sorted = rating_user.sort_values("rating")
print("10 best rated movies by user %d" %user_id)
display(rating_user_sorted[-10:][["movie","rating"]])
print("10 worst rated movies by user %d" %user_id)
display(rating_user_sorted[:10][["movie","rating"]])
# **Exercise** Use the model to compute the estimated rates that the user would give to the movies he hasn't seen. Display the 10 movies you would recommend to him.
# +
# # %load solutions/exercise_3_2.py
# -
# # Complete dataset
#
# The following code perform the same model on the complete dataset.
# It would take too much time if you don't have a GPU.
#
DATA_DIR = "ml-25/"
rating = pd.read_csv(DATA_DIR + "ratings_updated.csv")
nb_entries = rating.shape[0]
print("Number of entries : %d " %nb_entries)
rating.head(5)
movies = pd.read_csv(DATA_DIR + "movies.csv")
id_movie_to_title = dict(movies[["movieId","title"]].values)
id_item_to_title = {itemIdToNormItemId[k]:v for k,v in id_movie_to_title.items() if k in itemIdToNormItemId}
print("Number of movies in the dictionary : %d" %(len(id_item_to_title)))
movies.head()
userIdToNormUserId = {k:v for v,k in enumerate(rating.userId.unique())}
rating["user_id"] = [userIdToNormUserId[x] for x in rating.userId.values]
itemIdToNormItemId = {k:v for v,k in enumerate(rating.movieId.unique())}
rating["item_id"] = [itemIdToNormItemId[x] for x in rating.movieId.values]
# +
train = rating[rating.test_train=="train"]
user_id_train = train['user_id']
item_id_train = train['item_id']
rating_train = train['rating']
print(train.shape)
test = rating[rating.test_train=="test"]
user_id_test = test['user_id']
item_id_test = test['item_id']
rating_test = test['rating']
print(test.shape)
# +
user_id_input = kl.Input(shape=[1], name='user')
item_id_input = kl.Input(shape=[1], name='item')
embedding_size = 30
max_user_id= rating.user_id.max()
max_item_id= rating.item_id.max()
user_embedding = kl.Embedding(output_dim=embedding_size, input_dim=max_user_id + 1,
input_length=1, name='user_embedding')(user_id_input)
item_embedding = kl.Embedding(output_dim=embedding_size, input_dim=max_item_id + 1,
input_length=1, name='item_embedding')(item_id_input)
# reshape from shape: (batch_size, input_length, embedding_size)
# to shape: (batch_size, input_length * embedding_size) which is
# equal to shape: (batch_size, embedding_size)
user_vecs = kl.Flatten()(user_embedding)
item_vecs = kl.Flatten()(item_embedding)
input_vecs = kl.Concatenate()([user_vecs, item_vecs])
input_vecs = kl.Dropout(0.5)(input_vecs)
x = kl.Dense(64, activation='relu')(input_vecs)
y = kl.Dense(1)(x)
model = km.Model(inputs=[user_id_input, item_id_input], outputs=y)
model.compile(optimizer='adam', loss='mae')
model.summary()
# -
history = model.fit([user_id_train, item_id_train], rating_train,
batch_size=2048, epochs=10, validation_split=0.1,
shuffle=True)
weights = model.get_weights()
user_embeddings = weights[0]
print("User embedding matrix dimension : %s" %str(user_embeddings.shape))
item_embeddings = weights[1]
print("item embedding matrix dimension : %s" %str(item_embeddings.shape))
idx = 283
X = np.expand_dims(item_embeddings[idx],axis=0)
distX = sm.pairwise_distances(X, item_embeddings, metric="cosine")[0]
print("Top 10 items similar to movies %s" %str(id_item_to_title[idx]))
mostSimilarItem = pd.DataFrame([[id_item_to_title[x], distX[x],x] for x in distX.argsort()[:10]])
mostSimilarItem
pcaItems = sdec.PCA(n_components=2)
items_pca_embeddings = pcaItems.fit_transform(item_embeddings)
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(1,1,1)
ax.plot(items_pca_embeddings[:,0], item_embeddings[:,1], linestyle="None", marker=".")
ax.plot(items_pca_embeddings[mostSimilarItem[2].values,0], item_embeddings[mostSimilarItem[2].values,1], linestyle="None", marker=".", markersize=10)
user_id = 1
rating_user = rating[rating["user_id"]==user_id]
rating_user_sorted = rating_user.sort_values("rating")
print("10 best rated movies by user %d" %user_id)
display(rating_user_sorted[-10:][["movie","rating"]])
print("10 worst rated movies by user %d" %user_id)
display(rating_user_sorted[:10][["movie","rating"]])
#Run prediction for all movies
prediction = model.predict([[user_id for _ in range(max_item_id)], [x for x in range(max_item_id)]])
#Concatenate results with id of the movie
prediction_with_id = zip(prediction, [x for x in range(max_item_id)])
# Filter on unseen movie, get the title and sort the results according to predicted rate
prediction_of_unseen_movie = sorted([[p[0],id_item_to_title[x]] for p,x in prediction_with_id if not(x in seen_movie)], key=lambda x :x[0], reverse = True)
#Display it.
pd.DataFrame(prediction_of_unseen_movie)
| RecomendationSystem/3-Python-Neural-MovieLens.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mincloud1501/Bokeh/blob/master/googlesheet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qjjuLLUzUwfW" colab_type="text"
# # Getting setup
#
# Install boketh `pip install bokeh`.
# Setup-test, run the next cell. Hopefully you should see output that looks something like this.
# + id="asztQ5dMVDBB" colab_type="code" outputId="3e625a3d-4deb-4076-ad81-a1bdce04bf08" colab={"base_uri": "https://localhost:8080/", "height": 71}
from IPython import __version__ as ipython_version
from pandas import __version__ as pandas_version
from bokeh import __version__ as bokeh_version
print("IPython - %s" % ipython_version)
print("Pandas - %s" % pandas_version)
print("Bokeh - %s" % bokeh_version)
# + [markdown] id="w8eaAUE5suk8" colab_type="text"
# # Basic plotting with Bokeh
#
# + id="TZuXfMYqhaAh" colab_type="code" colab={}
# Import figure from bokeh.plotting
import numpy as np
# Import pandas
import pandas as pd
# Import output_file and show from bokeh.io
from bokeh.plotting import figure, show
# Import figure from bokeh.plotting
from bokeh.io import output_file, output_notebook
output_notebook()
# + [markdown] id="Oxs7lktMf-OC" colab_type="text"
# # Google Sheets
#
# Our examples below use the open-source [`gspread`](https://github.com/burnash/gspread) library for interacting with Google Sheets.
#
# First, install the package using `pip`.
# + id="Nvh5GRcYgMXn" colab_type="code" outputId="eac1522c-e997-488b-c4dd-2b2592716361" colab={"base_uri": "https://localhost:8080/", "height": 215}
# !pip install --upgrade --quiet gspread
# !pip install --upgrade oauth2client
# !pip install PyOpenSSL
# !pip install -U -q PyDrive
# + [markdown] id="s6dBcoABgX3Y" colab_type="text"
# Import the library, authenticate, and create the interface to Sheets.
#
# + id="9zAw7Xb2ghm2" colab_type="code" colab={} cellView="code"
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
import gspread
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
gc = gspread.authorize(GoogleCredentials.get_application_default())
# + id="EMTvwIEqQ5Pq" colab_type="code" colab={}
sh = gc.create('My Test')
# + id="DmjnBHF5ekG6" colab_type="code" outputId="042b7e67-4164-430f-f462-1c17f1118cad" colab={"base_uri": "https://localhost:8080/", "height": 35}
from os import path
from google.colab import drive
notebook_dir_name = 'Colab Notebooks'
drive.mount('/content/drive')
notebook_base_dir = path.join('./drive/My Drive/', notebook_dir_name)
# + id="q5TSl_JYPBe-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1dc5b39e-6b4a-4b8d-b032-a79376f2fef1"
with open('/content/drive/My Drive/foo.txt', 'w') as f:
f.write('Hello Google Drive!')
# !cat /content/drive/My\ Drive/foo.txt
# + [markdown] id="uo7rhToLhfyS" colab_type="text"
# ## Downloading data from a sheet into Python as a Pandas DataFrame
#
# + id="JrPSILQ0hP5G" colab_type="code" colab={}
worksheet = gc.open('My Test').sheet1
# get_all_values gives a list of rows.
data = worksheet.get_all_values()
headers = data.pop(0)
df = pd.DataFrame(data, columns=headers)
# + [markdown] id="m33Vf_g_TxhN" colab_type="text"
#
# + [markdown] id="5PDNcs1GEv-Y" colab_type="text"
# ## Plotting with glyphs
# https://docs.bokeh.org/en/latest/docs/user_guide/plotting.html
#
# + id="EuF-7sPFha5U" colab_type="code" outputId="f34a306d-a5e1-48fa-f3b2-5d9c390493df" colab={"base_uri": "https://localhost:8080/", "height": 517}
plot = figure(plot_width=800, plot_height=500, tools='pan,box_zoom,')
plot.circle([10,20,33,4,5, 100], [8,6,5,2,3, 300])
output_file('circles.html')
show(plot)
# + [markdown] id="oj59pOriG9s3" colab_type="text"
# ## What are glyphs?
# * Visual shapes
# * circles, squares, triangles
# * rectangles, lines, wedges
# * With properties a!ached to data
# * coordinates (x,y)
# * size, color, transparency
#
# ### Glyph properties
# * Lists, arrays, sequences of values
# * Single fixed values
#
# ```
# plot = figure()
# plot.circle(x=10, y=[2,5,8,12], size=[10,20,30,40])
# ```
#
# ### Markers
# https://docs.bokeh.org/en/latest/docs/gallery/markers.html
# + [markdown] id="O0dTJQ4ce-hk" colab_type="text"
#
# + [markdown] id="7qSA9yOpHBEz" colab_type="text"
# ## A simple scatter plot
#
# In this example, you're going to make a scatter plot of female literacy vs fertility using data from the European Environmental Agency. This dataset highlights that countries with low female literacy have high birthrates. The x-axis data has been loaded for you as fertility and the y-axis data has been loaded as female_literacy.
#
# Your job is to create a figure, assign x-axis and y-axis labels, and plot female_literacy vs fertility using the circle glyph.
#
# After you have created the figure, in this exercise and the ones to follow, play around with it! Explore the different options available to you on the tab to the right, such as "Pan", "Box Zoom", and "Wheel Zoom". You can click on the question mark sign for more details on any of these tools.
#
# Note: You may have to scroll down to view the lower portion of the figure.
#
# Import the figure function from bokeh.plotting, and the output_file and show functions from bokeh.io.
# Create the figure p with figure(). It has two parameters: x_axis_label and y_axis_label.
# Add a circle glyph to the figure p using the function p.circle() where the inputs are, in order, the x-axis data and y-axis data.
# Use the output_file() function to specify the name 'fert_lit.html' for the output file.
# Create and display the output file using show() and passing in the figure p.
# + id="u9xy8uc2RxkP" colab_type="code" colab={}
sh1 = gc.create('My Test1')
# + id="kMvn8vt0TyaC" colab_type="code" colab={}
worksheet = gc.open('My Test1').sheet1
# get_all_values gives a list of rows.
data = worksheet.get_all_values()
# get head
headers = data.pop(0)
# make dataframe
df = pd.DataFrame(data, columns=headers)
# get column data
population = df['population']
fertility = df['fertility']
# print(fertility)
female_literacy = df['female literacy']
# + id="kGemJ-DaHKRS" colab_type="code" outputId="9d59ad4f-c6cf-4107-b706-a725e3c914b8" colab={"base_uri": "https://localhost:8080/", "height": 617}
# Create the figure: p
p = figure(x_axis_label='여성 1인명 당 자녀 수', y_axis_label ='population (% 인구율)')
# Add a circle glyph to the figure p
p.circle(fertility, female_literacy)
# Call the output_file() function and specify the name of the file
output_file('fert_lit.html')
# Display the plot
show(p)
# + [markdown] id="nqQiwF1JHk1U" colab_type="text"
# ## Customizing your scatter plots
#
# The three most important arguments to customize scatter glyphs are color, size, and alpha. Bokeh accepts colors as hexadecimal strings, tuples of RGB values between 0 and 255, and any of the 147 CSS color names. Size values are supplied in screen space units with 100 meaning the size of the entire figure.
#
# The alpha parameter controls transparency. It takes in floating point numbers between 0.0, meaning completely transparent, and 1.0, meaning completely opaque.
#
# In this exercise, you'll plot female literacy vs fertility for Africa and Latin America as red and blue circle glyphs, respectively.
#
# Using the Latin America data (fertility_latinamerica and female_literacy_latinamerica), add a blue circle glyph of size=10 and alpha=0.8 to the figure p. To do this, you will need to specify the color, size and alpha keyword arguments inside p.circle().
# Using the Africa data (fertility_africa and female_literacy_africa), add a red circle glyph of size=10 and alpha=0.8 to the figure p.
#
#
#
#
# + id="PxMKVFbFHmQM" colab_type="code" outputId="f7081d57-29d6-4113-e143-c823f061dc36" colab={"base_uri": "https://localhost:8080/", "height": 617}
fertility = df['fertility']
female_literacy = df['female literacy']
fertility_latinamerica = df.loc[df['continent'] == "LAT"]['fertility']
female_literacy_latinamerica = df.loc[df['continent'] == "LAT"]['female literacy']
fertility_africa = df.loc[df['continent'] == "AF"]['fertility']
female_literacy_africa = df.loc[df['continent'] == "AF"]['female literacy']
# Create the figure: p
p = figure(x_axis_label='fertility (children per woman)', y_axis_label='female_literacy (% population)')
# Add a blue circle glyph to the figure p
p.circle(fertility_latinamerica, female_literacy_latinamerica, color='blue', size=20, alpha=0.8)
# Add a red circle glyph to the figure p
p.circle(fertility_africa, female_literacy_africa, color='red', size=20, alpha=0.8)
# Specify the name of the file
output_file('fert_lit_separate_colors.html')
# Display the plot
show(p)
# + [markdown] id="dmFD6F91I8TS" colab_type="text"
# ## Lines
#
#
# + id="CxUi90ZoRP4_" colab_type="code" outputId="ea8107dc-f915-471f-bbe0-ff3ed3bd6727" colab={"base_uri": "https://localhost:8080/", "height": 617}
x = [1, 2, 3, 4, 5]
y = [8, 6, 5, 2, 3]
plot = figure()
plot.line(x, y, line_width=3)
output_file('line.html')
show(plot)
# + [markdown] id="XYNgfemiIVTW" colab_type="text"
# ## Lines and markers
# + id="_0sCGnwES-CI" colab_type="code" outputId="247b5bb0-7d9f-4a21-da5e-f2eb95ac2dc3" colab={"base_uri": "https://localhost:8080/", "height": 617}
plot = figure()
plot.line(x, y, line_width=2)
plot.circle(x, y, fill_color='white', size=10)
output_file('line.html')
show(plot)
# + [markdown] id="NlVzCc8swVlL" colab_type="text"
#
# + [markdown] id="WrxI80SJH0Gm" colab_type="text"
# ## Patches
# * Userful for showing geographic regions
# * Data given as "list of lists"
#
# + id="GEyFWDPlUH97" colab_type="code" outputId="e72a237a-c8d2-416d-d27c-bec4c7c53c89" colab={"base_uri": "https://localhost:8080/", "height": 635}
xs = [ [1,1,2,2], [2,2,4], [2,2,3,3] ]
ys = [ [2,5,5,2], [3,5,5], [2,3,4,2] ]
plot = figure()
plot.patches(xs, ys, fill_color= ['red', 'blue','green', 'yellow'], line_color='white')
output_file('patches.html')
show(plot)
# + id="xxxiQtdGXXho" colab_type="code" colab={}
| googlesheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Create a NLP Pipeline to 'Clean' Reviews Data
# - Load Input File and Read Reviews
# - Tokenize
# - Remove Stopwords
# - Perform Stemming
# - Write cleaned data to output file
sample_text = """
First things first, <NAME> did a fantastic, believable job as a Cambodian hit-man,
born and bred in the dumps and a gladiatorial ring, where he honed his craft of savage battery in order to survive,
living on the mantra of kill or be killed. In a role that had little dialogue, or at least a few lines in Cambodian/Thai,
his performance is compelling, probably what should have been in the Jet Li vehicle Danny the Dog,
where a man is bred for the sole purpose of fighting, and on someone else's leash.<br /><br />Like Danny the Dog,
the much talked about bare knuckle fight sequences are not choreographed stylistically, but rather designed as normal,
brutal fisticuffs, where everything goes. This probably brought a sense of realism and grit when you see the characters
slug it out at each other's throats, in defending their own lives while taking it away from others. It's a grim, gritty
and dark movie both literally and figuratively, and this sets it apart from the usual run off the mill cop thriller
production.<br /><br />Edison plays a hired gun from Cambodia, who becomes a fugitive in Hong Kong, on the run from the
cops as his pickup had gone awry. Leading the chase is the team led by <NAME>, who has to contend with maverick
member Inspector Ti (<NAME>), who's inclusion and acceptance in the team had to do with the sins of his father.
So begins a cat and mouse game in the dark shades and shadows of the seedier looking side of Hong Kong.<br /><br />
The story itself works on multiple levels, especially in the character studies of the hit-man, and the cop.
On opposite sides of the law, we see within each character not the black and white, but the shades of grey.
With the hit-man, we see his caring side when he got hooked up and developed feelings of love for a girl (Pei Pei),
bringing about a sense of maturity, tenderness, and revealing a heart of gold. The cop, with questionable tactics and
attitudes, makes you wonder how one would buckle when willing to do anything it takes to get the job done.
There are many interesting moments of moral questioning, on how anti-hero, despicable strategies are adopted.
You'll ask, what makes a man, and what makes a beast, and if we have the tendency to switch sides depending on
circumstances - do we have that dark inner streak in all of us, transforming from man to dog, and dog to man?
Dog Bite Dog grips you from the start and never lets go until the end, though there are points mid way through
that seemed to drag, especially on its tender moments, and it suffered too from not knowing when to end.
If I should pick a favourite scene, then it must be the one in the market food centre - extremely well controlled and
delivered, a suspenseful edge of your seat moment. Listen out for the musical score too, and you're not dreaming if you
hear growls of dogs.<br /><br />Highly recommended, especially if you think that you've seen about almost everything from
the cop thriller genre."""
# #### NLTK
from nltk.tokenize import RegexpTokenizer
from nltk.stem.porter import PorterStemmer
from nltk.corpus import stopwords
import sys
# Initialize Objects
tokenizer = RegexpTokenizer(r'\w+')
en_stopwords = set(stopwords.words('english'))
ps = PorterStemmer()
def getStemmedReview(review):
review = review.lower()
review = review.replace('<br /><br />',' ')
# Tokenize
tokens = tokenizer.tokenize(review)
new_tokens = [token for token in tokens if token not in en_stopwords]
stemmed_tokens = [ps.stem(tokens) for tokens in new_tokens]
cleaned_review = ' '.join(stemmed_tokens)
return cleaned_review
getStemmedReview(sample_text)
| .ipynb_checkpoints/NLP - Pipeline-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
from matplotlib import pyplot as plt
from xdgmm import XDGMM
from sklearn.model_selection import validation_curve
from sklearn.model_selection import ShuffleSplit
from test_plots import *
'''
Due to AstroML still using the deprecated GMM class from
scikit-learn (instead of GaussianMixture), this demo will
throw numerous errors whenever the XDGMM object calls
an AstroML method, such as fit. The lines below will
suppress these warnings; comment them out to see everything.
This XDGMM class has been updated to use GaussianMixture
instead of GMM when necessary, but since it uses an AstroML
XDGMM object to store and manipulate the model, it is
dependent on AstroML. These warnings will continue to occur
until the XDGMM class from AstroML has been updated.
'''
import warnings
warnings.filterwarnings('ignore')
# -
# Next, generate some data to use for our fitting and plotting.
# +
## Generate synthetic data
N,D = 100, 3 # number of points and dimenstinality
if D == 2:
#set gaussian ceters and covariances in 2D
means = np.array([[1.5, 0.0],
[0, 0],
[-1.0, -1.0],
[-1.0, 0.5]])
covs = np.array([np.diag([0.01, 0.01]),
np.diag([0.08, 0.01]),
np.diag([0.01, 0.05]),
np.diag([0.03, 0.07])])
elif D == 3:
# set gaussian ceters and covariances in 3D
means = np.array([[1.5, 0.0, 0.0],
[0.0, 0.0, 0.0],
[-1.0, -1.0, -1.0],
[-1.0, 1.0, 1.0]])
covs = np.array([np.diag([0.05, 0.05, 0.10]), #np.diag()输出矩阵的对角线元素,或将一维数组变成以一维数组为对角线元素的矩阵
np.diag([0.08, 0.03, 0.03]),
np.diag([0.20, 0.05, 0.15]),
np.diag([0.03, 0.07, 0.15]),])
n_gaussians = means.shape[0] #shape结果为显示矩阵为(n,m)如(1,4), shape[0]显示列(第二维的长度)
points = []
for i in range(len(means)):
x = np.random.multivariate_normal(means[i], covs[i], N )
points.append(x)
X = np.concatenate(points)
fig = plt.figure(figsize=(8, 8))
axes = fig.add_subplot(111, projection='3d')
axes.scatter(X[:, 0], X[:, 1], X[:, 2], s = 2.0, alpha = 0.5)
axes.set_xlim3d(-2,2)
axes.set_ylim3d(-2,2)
axes.set_zlim3d(-2,2)
# add noise to get the "observed" distribution
dx = 0.5 + 0.6 * np.random.random(400)
dy = 0.5 + 0.7 * np.random.random(400)
dz = 0.5 + 0.8 * np.random.random(400)
x_noise = X[:, 0].T + np.random.normal(0, dx)
y_noise = X[:, 1].T + np.random.normal(0, dy)
z_noise = X[:, 2].T + np.random.normal(0, dz)
X_noise = np.vstack([x_noise, y_noise, z_noise]).T
X_all = np.concatenate((X,X_noise), axis=0)
fig_noise = plt.figure(figsize=(8, 8))
axes_noise = fig_noise.add_subplot(111, projection='3d')
axes_noise.scatter(X_all[:, 0], X_all[:, 1], X_all[:, 2], s = 2.0, alpha = 0.5)
axes_noise.set_xlim3d(-2,2)
axes_noise.set_ylim3d(-2,2)
axes_noise.set_zlim3d(-2,2)
Xerr = 0.05*np.zeros(X.shape + X.shape[-1:])
# diag = np.arange(X.shape[-1])
# Xerr[:, diag, diag] = np.vstack([dx ** 2, dy ** 2]).T
# -
# ## Component Number Selection
#
# We could fit this data directly, but one problem is that we don't know the optimal number of components (Gaussians) to use in the fit. Knowing the optimal number of components to fit allows us to obtain a good fit in the smallest amount of time without overfitting the data.
#
# ### BIC
# One way this can be determined is by fitting the model with different numbers of components and calculating the Bayesian information criterion (BIC) for each model. The BIC incorporates the number of components in the model, the sample size, and the likelihood of the data under the model, and the model with the lowest score is the optimal model to use.
# +
# Instantiate an XDGMM model:
xdgmm = XDGMM(method='Bovy')
# Define the range of component numbers, and get ready to compute the BIC for each one:
param_range = np.array([1,2,3,4,5,6,7])
# Loop over component numbers, fitting XDGMM model and computing the BIC:
bic, optimal_n_comp, lowest_bic = xdgmm.bic_test(X, Xerr, param_range)
# -
plot_bic(param_range, bic, optimal_n_comp)
# ## Model Fitting
#
# Now that we know the best number of components to use, we can fit the data. First set the number of components, then perform the fit.
# +
xdgmm.n_components = optimal_n_comp
xdgmm = xdgmm.fit(X, Xerr)
xdgmm.save_model('test_model.fit')
# -
def plot_sphere(w=0, center=[0,0,0], r=[1, 1, 1], rotation=[1,1,1], ax=None):
'''
plot a sphere surface
Input:
c: 3 elements list, sphere center
r: 3 element list, sphere original scale in each axis ( allowing to draw elipsoids)
subdiv: scalar, number of subdivisions (subdivision^2 points sampled on the surface)
是椭球的分辨率
ax: optional pyplot axis object to plot the sphere in.
sigma_multiplier: sphere additional scale (choosing an std value when plotting gaussians)
Output:
ax: pyplot axis object
'''
if ax is None:
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
u = np.linspace(0, 2 * np.pi, 30) #np.linspace 取等差数列
v = np.linspace(0, np.pi, 30)
x = r[0] * np.outer(np.cos(u), np.sin(v))
y = r[1] * np.outer(np.sin(u), np.sin(v))
z = r[2] * np.outer(np.ones(np.size(u)), np.cos(v))
for i in range(len(x)):
for j in range(len(x)):
#[x[i, j], y[i, j], z[i, j]] = [x[i, j], y[i, j], z[i, j]] + center #spherical专用
[x[i, j], y[i, j], z[i, j]] = np.dot([x[i, j], y[i, j], z[i, j]], rotation.T) + center
ax.plot_surface(x, y, z, alpha=0.3, linewidth=0)
return ax
# +
fig = plt.figure(figsize=(8, 8))
axes = fig.add_subplot(111, projection='3d')
axes.set_zlim3d(-2, 2)
axes.set_xlim3d(-2, 2)
axes.set_ylim3d(-2, 2)
# axes.view_init(-90, 90)
axes.scatter(X_all[:, 0], X_all[:, 1], X_all[:, 2], s = 1.0, alpha = 0.5)
for i in range(optimal_n_comp):
covariances = xdgmm.V[i][:3, :3]
v, u = np.linalg.eigh(covariances)
r = np.sqrt(v)
plot_sphere(xdgmm.weights[i], xdgmm.mu.T[:3,i], r, u, ax=axes)
# +
from sklearn.mixture import GaussianMixture
gmm = GaussianMixture(n_components= 4, covariance_type='full')
gmm.fit(X_all)
figsk = plt.figure(figsize=(8, 8))
axessk = figsk.add_subplot(111, projection='3d')
axessk.set_zlim3d(-2, 2)
axessk.set_xlim3d(-2, 2)
axessk.set_ylim3d(-2, 2)
axessk.scatter(X_all[:, 0], X_all[:, 1], X_all[:, 2], s = 1.0, alpha = 0.5)
for i in range(0,4):
covariances = gmm.covariances_[i][:3, :3]
v, u = np.linalg.eigh(covariances)
r = np.sqrt(v)
plot_sphere(gmm.weights_[i], gmm.means_.T[:3,i], r, u, ax=axessk)
# +
sample = xdgmm.sample(N*3)
plot_sample(X[:,0], X[:,1], X_all[:,0], X_all[:,1], gmm, xdgmm)
# -
def plot_sample3D(x_true, y_true, z_true, x, y, z, gmm, xdgmm):
setup_text_plots(fontsize=16, usetex=True)
plt.clf()
fig = plt.figure(figsize=(12, 12))
fig.subplots_adjust(left=0.1, right=0.95,
bottom=0.1, top=0.95,
wspace=0.02, hspace=0.02)
ax1 = fig.add_subplot(221, projection='3d')
ax1.scatter(x_true, y_true, z_true, s=4, lw=0, c='k')
ax2 = fig.add_subplot(222, projection='3d')
ax2.scatter(x, y, z, s=4, lw=0, c='k')
ax3 = fig.add_subplot(223, projection='3d')
for i in range(gmm.n_components):
covariances = gmm.covariances_[i][:3, :3]
v, u = np.linalg.eigh(covariances)
r = np.sqrt(v)
plot_sphere(gmm.weights_[i], gmm.means_.T[:3,i], r, u, ax=ax3)
ax4 = fig.add_subplot(224, projection='3d')
for i in range(xdgmm.n_components):
covariances = xdgmm.V[i][:3, :3]
v, u = np.linalg.eigh(covariances)
r = np.sqrt(v)
plot_sphere(xdgmm.weights[i], xdgmm.mu.T[:3,i], r, u, ax=ax4)
titles = ["True Distribution", "Noisy Distribution",
"Gaussian Mixture Models",
"Extreme Deconvolution GMM"]
ax = [ax1, ax2, ax3, ax4]
for i in range(4):
ax[i].set_xlim3d(-2, 2)
ax[i].set_ylim3d(-2, 2)
ax[i].set_zlim3d(-2, 2)
ax[i].xaxis.set_major_locator(plt.MultipleLocator(2))
ax[i].yaxis.set_major_locator(plt.MultipleLocator(2))
ax[i].zaxis.set_major_locator(plt.MultipleLocator(2))
ax[i].set_title(titles[i])
# ax[i].text(0.05, 0.95, titles[i],
# ha='left', va='top', transform=ax[i].transAxes)
if i in (0, 1):
ax[i].xaxis.set_major_formatter(plt.NullFormatter())
else:
ax[i].set_xlabel('$x$', fontsize = 18)
if i in (0, 2):
ax[i].zaxis.set_major_formatter(plt.NullFormatter())
else:
ax[i].set_zlabel('$z$', fontsize = 18)
if i in (0, 1):
ax[i].yaxis.set_major_formatter(plt.NullFormatter())
else:
ax[i].set_ylabel('$y$', fontsize = 18)
plot_sample3D(X[:,0], X[:,1], X[:,2], X_all[:,0], X_all[:,1], X_all[:,2], gmm, xdgmm)
# ## Conditioning the Model
#
# One of the key advantages of our implementation of the XDGMM class is that it contains a method for producing a model for the conditional PDF $P(x | y)$ from the original GMM model for the joint PDF $P(x, y)$. The conditional PDF is also a GMM with the same number of components, but with adjusted weights, means and variance. This is useful for predicting new $x$ values given test $y$ values: we simply sample from the conditioned model.
#
# For our example dataset, let's assume that we have measured the $y$ value to be $1.5\pm0.05$. We can produce a new model for $x$ that is conditioned on $y=1.5\pm0.05$, and use that to sample values of $x$. When `condition()` is called, we set all parameters that we do not want included in the conditioning to `NaN`. The model can be conditioned on values either with uncertainties or without; to condition without uncertainty either do not pass an error array, or set all the uncertainties equal to $0$. If provided, the uncertainties are interpreted as being Gaussian and uncorrelated, and thus providing a simple $P(y)$ to be combined with the joint PDF.
# +
cond_X = np.array([np.nan, 1.5])
cond_Xerr = np.array([0.0,0.05])
cond_xdgmm = xdgmm.condition(X_input = cond_X,Xerr_input = cond_Xerr)
# Compare the conditioned model to the original:
print(xdgmm.weights)
print(cond_xdgmm.weights)
print("\n")
print(xdgmm.mu)
print(cond_xdgmm.mu)
# -
# Note how the number of components in the conditioned model is the same as in the original joint model, but that the weights of the components have changed, and the mu array is now 1-dimensional (since $y$ has been conditioned out).
#
# A list of labels for each parameter in the dataset can also be stored in the XDGMM object, and then a dictionary object can be used to condition the model using the labels. This removes the need for the user to remember which index corresponds to which parameter. When using a dictionary, it is only necessary to pass values for the labels that you want to condition on. If floats are given with each label, then `condition()` assumes that these are data values and that the uncertainty is 0. If tuples are given, the function assumes that the tuples contain (data, uncertainty) pairs. The new, conditioned XDGMM object will retain the labels of any parameters that were not conditioned out. This functionality is demonstrated below.
#
# If the data used to fit the XDGMM model are in the form of a Pandas DataFrame and the columns have been labelled, the XDGMM object will automatically save the column labels in the `labels` array for future use.
# +
# First, set the labels in the XDGMM object
xdgmm.labels = np.array(['x','y'])
# The dictionary can pass either floats or tuples
cond_dict = {'y':(1.5,0.05)}
cond_xdgmm2 = xdgmm.condition(X_dict = cond_dict)
# Print the weights and means of the new model.
print(cond_xdgmm2.weights)
print(cond_xdgmm2.mu)
print(cond_xdgmm2.labels)
# -
# As expected, the conditioning results are the same. Labels will also be saved to and read from files when the `save_model()` and `read_model()` functions are used.
#
# Below we plot the original Gaussian components in the $x-y$ plane, a blue line showing the condition $y=1.5$, and a red line showing the new conditional distribution for $x$ given $y=1.5\pm0.05$.
plot_cond_model(xdgmm, cond_xdgmm, 1.5)
# If we sample 1000 points from this conditional distribution, we would get something like this:
cond_sample = cond_xdgmm.sample(1000)
y = np.ones(1000)*1.5
plot_cond_sample(cond_sample,y)
# ## Conditional Prediction
#
# As stated before, one potential use of a conditioned model is to create a "prediction engine" that can predict some parameters using an `XDGMM` model conditioned on known values of other parameters.
#
# To demonstrate this, let's sample 1000 data points from our joint model to create a dataset to be compared with our predictions. Now, at the same $y$ values as this "observed" dataset, we'll condition the model on each $y$ value in turn, and draw a single predicted $x$ value. These predicted $x$'s should follow the same distribution as the observed $x$'s.
# +
# Simulate a dataset:
true_sample = xdgmm.sample(1000)
true_x = true_sample[:,0]
y = true_sample[:,1]
# Predict x values given y values:
predicted_x = np.array([])
for this_y in y:
# Specify y-conditioning to apply to P(x,y):
on_this = np.array([np.nan,this_y])
# Compute conditional PDF P(x|y):
cond_gmm = xdgmm.condition(on_this)
# Draw a sample x value from this PDF, and add it to the growing list
predicted_x = np.append(predicted_x, cond_gmm.sample())
# Plot the two datasets, to compare the true x and the predicted x:
plot_conditional_predictions(y, true_x, predicted_x)
| Test/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2 style="color:blue" align="center"> Missing Values : K NEAREST NEIGHBORS (KNN Imputer) </h2>
# -------------------------------------------------------------------------------------------------------------------------------
import pandas as pd
# +
url = "C:/Users/deepusuresh/Documents/Data Science/10. Missing Values/3. Techniques to Handle Missing Data/3. KNN Imputer"
df = pd.read_csv("KNN imputer.csv")
df
# -
x = df.iloc[:,:-1].values
y = df.iloc[:,-1].values
from sklearn.impute import KNNImputer
imputer = KNNImputer(n_neighbors=2, weights='uniform')
z = imputer.fit_transform(x)
imputer = KNNImputer(missing_values=7, n_neighbors=2, weights='uniform')
| 10_Missing_Values/2_Imputation_Sklearn/3_KNN_Imputer/KNN_Imputer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.3.8
# language: julia
# name: julia-0.3
# ---
include("asinclude.jl")
# +
## Demo
asinclude.@asinclude("mymodule", quote
import Base.start
import Base.next
import Base.done
export MyType, IterType, bar
type MyType
x
baz::Int
end
bar(x) = 2x
show(io, a::MyType) = print(io, "MyType $(a.baz)")
type IterType
baz
end
function start(c::IterType)
N = c.baz
state = 0
return state
end
function next(c::IterType, state)
return state + 1, state + 1
end
function done(c::IterType, state)
if isempty(state)
return true
end
state += 1
i = 1
if state > c.baz
return true
end
return false
end
end)
println(mymodule.MyType(2, 5))
for i = mymodule.IterType(5)
println(i)
end
# +
asinclude.@asinclude("mymodule2", quote
import Base: start, done, next
export MyType, IterType, bar
type MyType
baz::Int
end
bar(x) = 2x
show(io, a::MyType) = print(io, "MyType $(a.baz)")
type IterType
baz
end
function start(c::IterType)
N = c.baz
state = 0
return state
end
function next(c::IterType, state)
return state + 1, state + 1
end
function done(c::IterType, state)
if isempty(state)
return true
end
state += 1
i = 1
if state > c.baz
return true
end
return false
end
end)
println(mymodule.MyType(10))
for i = mymodule.IterType(5)
println(i)
end
# -
macro dynamic_import(modules)
(modules = eval(modules))::Vector{Symbol}
ex = Expr(:toplevel)
for m in modules
push!(ex.args, Expr(:import, m))
end
return ex
end
eval(Expr(:import, symbol("mymodule")))
names(eval(symbol("mymodule")))
toks = split(string(quote
import Base: start, next
end), ':')[3:end]
specialform_handler["toplevel"](toks...)
specialform_handler = {
"import" => (args...) -> begin
args = map(x -> replace(x, r",|\s|\)|(#.*$)", ""), args)
args = filter(x->length(x)>0, [args...])
return (args[1] * " " * join(args[2:end], '.'))
end,
"export" => (args...) -> begin
args = map(x -> replace(x, r",|\s|\)|(#.*$)", ""), args)
args = filter(x->length(x)>0, [args...])
return (args[1] * " " * join(args[2:end], ','))
end,
"toplevel" => (args...) -> begin
args = map(x -> replace(x, r",|\s|\)|(#.*$)", ""), args)
entries = String[]
currententry = String[]
for token in args
println(token)
if contains(token, "\$")
push!(entries, specialform_handler[currententry[1]](currententry...))
currententry = []
end
push!(currententry, token)
end
return join(map(x -> specialform_handler[x[1]](x), entries), "\n")
end
}
| asincludetest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys, collections, functools, itertools, heapq, pprint, bisect, operator, math, random
# # 1. List
# List is perhaps the most frequently used python built-in data structure. Basically you can treat it as a resizable array which is similar to vector in STL and ArrayList in Java.
# initialization
lst1 = [1, 45, 10]
lst2 = list(range(5, 10)) # initialize to a range
lst3 = [0] * 10 # initialize to same thing, make sure you understand what happens here, check the below example
print(lst1)
print(lst2)
print(lst3)
## high level
lst4 = [[]] * 3 # when you use [item] * num to init a list and item is mutable, basically the list contains num references pointing to one object
print(lst4)
lst4[0].append(10)
print(lst4)
# sometimes this is used for some reason
lst = list(range(20))
iters = [iter(lst)] * 4
print(list(zip(*iters)))
# length of list, list is zero-indexed
lst = [0] * 10
print('length of lst is {:d}.'.format(len(lst)))
lst[0] = 1
print(lst)
# traverse list
lst = list("apple")
for item in lst:
print(item)
for i in range(len(lst)):
print(lst[i])
# if you want the index, item at the same time
for idx, item in enumerate(lst):
print('lst[{}] = {}'.format(idx, item))
# you can specify the start index in enumerate
for idx, item in enumerate(lst, 1):
print('index = {}, item = {}'.format(idx, item))
# +
# how to sort a list
lst = [random.randint(0, 10) for _ in range(20)]
print("Before sort:")
print(lst)
print("After sort:")
lst.sort() # this will change lst
print(lst)
# If you do not want to change the original list, you can use sorted() function
lst = [random.randint(0, 10) for _ in range(20)]
print("Sorted list:")
print(sorted(lst))
print("Original list:")
print(lst)
# -
# sort the list in descending order
lst = [3, 4, 1, 5, 2]
lst.sort(reverse=True)
print(lst)
# list could be used to simulate the behavior of stack
lst = [1, 2, 3, 4]
print(lst.pop())
lst.append(5)
print(lst)
# +
# some useful methods of Python list
# 1. insert an element at index
lst = [1, 2, 3, 4]
lst.insert(0, 9) # note that this is a
print(lst)
# 2. delete an element at index
del lst[1]
print(lst)
lst.pop(0)
print(lst)
# 3. count the occurance of an element
lst = [1, 2, 3, 1, 2, 3, 1]
print(lst)
print("number of 1 is {}".format(lst.count(1)))
# 4. first index of an element
print("index of 1 is {}".format(lst.index(1)))
# you can specify the start index to search
print(lst.index(1, 2))
# 5. append a list to a list
lst = [1, 2, 3]
print(lst)
lst.extend([4, 5, 6])
print(lst)
# or you can use
lst += [7, 8]
print(lst)
# 6. remove the first occurance of an element
lst = [1, 2, 1, 1]
print(lst)
lst.remove(1)
print(lst)
# 7. reverse the list
lst = [1, 2, 3]
lst.reverse()
print(lst)
# +
# slice [start:end:step]
lst = list(range(10))
print(lst)
# elements at even indexes
evens = lst[::2]
# elements at odd indexes
odds = lst[1::2]
print(evens)
print(odds)
print(lst[::-1]) # create a reverse copy of lst
# -
# # copy a list
a = [1, 2, 3]
b = a # this is not copy
b[0] = 0
print(a)
b = a.copy()
b[1] = 10
print(a)
b = a[:]
print(b)
# +
# list comprehension
# -
| IPython-notebooks/python_linear_data_structure.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Data Science - Lecture 2 - Version Control
# *COMP 5360 / MATH 4100, University of Utah, http://datasciencecourse.net/*
#
# In this lecture, we will learn about version control. We'll look at a couple of general principles and then go into the specifics of git and also GitHub. We'll also look at features of GitHub such as issue tracking. We strongly recommend that you use proper version control for your final project.
#
#
# ## Installation
#
# See the [official documentation](https://git-scm.com/book/en/v2/Getting-Started-Installing-Git) on how to install git on your operating system.
#
# On Mac, install the XCode package from the app store.
#
# On Windows, see the above link, or install [GitHub Desktop](https://desktop.github.com/) which includes a git shell.
# ## Why version Control?
#
# * **Keep copies of multiple states of files**
# By committing you record a state of the file to which you can go back any time.
# * **Create alternative states**
# Imagine you just want to try out something, but you realize you have to modify multiple files. You're not sure whether it works or is worth it. With version control you can just create a **branch** where you can experiment or develop new features without changing the main or other branches.
# * **Collaborate in teams**
# Nobody wants to send code via e-mail or share via Dropbox. If two people work on a file at the same time it's unclear how to merge the code. Version control lets you keep your code in a shared central location and has dedicated ways to merge and deal with conflicts.
# * **Keep your work safe**
# Your hard drive breaks. Your computer is stolen. But your code is safe because you store it not only on your computer but also on a remote server.
# * **Share**
# You developed something awesome and want to share it. But not only do you want to make it available, you're also happy about contributions from others!
#
#
#
# ## Types of Version Control: Central Repository
#
# 
#
# * Everybody needs to write to one server
# * All operations (history, commit, branches) require server connection
# * The traditional model: CVS, SVN, etc.
# * Pros:
# * Simple
# * Cons:
# * Complex for larger and community projects
# * Who is allowed to write?
# * How do you apply changes that someone outside your team made?
#
#
#
# ## Types of Version Control: Distributed Version Control
#
#
# 
#
# * Everybody has a full history of the repository locally
# * No dedicated server - every node is equal.
# * In practice: often server is used for one "official" copy of code.
# But: server by convention only, no technical difference.
# * Pros:
# * No access issues
# * Make a copy and hack away
# * Ask if partner wants to accept your changes
# * Everything is local
# * Fast!
# * No internet connection required
# * Commit often model (once per feature) - don't sync all the time.
# * Cons:
# * Extra effort to distinguish between committing and pushing/pulling (synchronizing).
#
# ## Implementations
#
# * Centralized
# * CVS
# * SVN
# * Team Foundation Server
# * ...
# * Distributed
# * git
# * Mercurial
# * ...
# * We will be using git in this lecture.
#
# ## git
#
# * Created by <NAME>ds, 2005
# * Meaning: British English slang roughly equivalent to "unpleasant person".
# * git – the stupid content tracker.
#
# *I'm an egotistical bastard, and I name all my projects after myself. First 'Linux', now 'git'. -- <NAME>*
#
# ## Why git?
#
# * Popular ([~60-90% of open source projects](https://rhodecode.com/insights/version-control-systems-2016))
# * Truly distributed
# * Very fast
# * Everything is local
# * Free
# * Safe against corruptions
# * GitHub!
#
# ## git model
#
# A git repository is essentially a large graph.
#
# 
#
# # Git tutorial
#
# This is a quick intro to git, used in combination with GitHub. This is not a complete tutorial, but will use the most important git features.
#
# We start by configuring git
#
# ```bash
# $ git config --global user.name "<NAME>"
# $ git config --global user.email "YOUR EMAIL ADDRESS"
# ```
# We recommend that you set this to your official school address and your correct name!
#
# **Create a folder for your project**
#
# ```bash
# $ mkdir myProject
# $ cd myProject/
# ```
#
# **Initalize the git repository**
#
# ```bash
# $ git init
# Initialized empty Git repository in ../myProject/.git/
# ```
#
# **What does git do to your file system?**
#
# ```bash
# # Let's look at what git creates
# $ ls .git/
# branches config description HEAD hooks info objects refs
#
# # The interesting stuff is in the config file
# $ cat .git/config
# [core]
# repositoryformatversion = 0
# filemode = true
# bare = false
# logallrefupdates = true
#
#
# # More interesting for a project with branches and remotes
# $ cat .git/config
# [core]
# repositoryformatversion = 0
# filemode = true
# bare = false
# logallrefupdates = true
# ignorecase = true
# precomposeunicode = true
# [remote "origin"]
# url = https://github.com/dataviscourse/2016-dataviscourse-website
# fetch = +refs/heads/*:refs/remotes/origin/*
# [branch "master"]
# remote = origin
# merge = refs/heads/master
#
# ```
#
# **Now let's create a file**
# ```bash
# $ echo 'Hello World' > demo.txt
# $ cat demo.txt
# Hello World
# ```
#
# **Let's add it to version control**
# ```bash
# $ git add demo.txt
# ```
#
#
# **Let's look at what is going on with the repository**
# ```bash
# $ git status
# # On branch master
# #
# # Initial commit
# #
# # Changes to be committed:
# # (use "git rm --cached <file>..." to unstage)
# #
# # new file: demo.txt
# #
# ```
#
# That means: git knows that it's supposed to track this file, but it's not yet versioned.
#
# **Let's commit the file.** Once a file is committed, it's state is recorded and you can go back to previous versions at any time.
#
# ```bash
# # The -m option specifies the commit message. If you don't use it you'll go into an editor to enter your commit message.
# $ git commit -m "Committing the test file"
# [master (root-commit) 3be5e8c] Wrote to demo
# 1 file changed, 1 insertion(+)
# create mode 100644 demo.txt
#
# # Did it work?
# $ git status
# # On branch master
# nothing to commit, working directory clean
# ```
#
# That means that now the file is tracked and committed to git. But it's still only stored on this one computer!
#
# **Next, we change a file and commit it again.**
#
# ```bash
# # Now let's change something
# $ echo 'Are you still spinning?' >> demo.txt
# $ cat demo.txt
# Hello World!
# Are you still spinning?
#
# # Let's check the status of git!
# $ git status
# # On branch master
# # Changes not staged for commit:
# # (use "git add <file>..." to update what will be committed)
# # (use "git checkout -- <file>..." to discard changes in working directory)
# #
# # modified: demo.txt
# #
# no changes added to commit (use "git add" and/or "git commit -a")
#
# # So git knows that something has changed, but hasn't recorded it. Let's commit.
# $ git commit -m "Added a line to the demo file"
# On branch master
# Changes not staged for commit:
# modified: demo.txt
# ```
#
# That didn't work! You have to add all the files you want to commit every time. There is a shorthand that you can use to add all the tracked files: append '-a'.
#
# ```bash
# $ git commit -a -m "added a line to the demo file"
# [master b03178f] added a line to the demo file
# 1 file changed, 1 insertion(+)
#
# # Better. Now, let's look at what happened up to now
# $ git log
# commit bf92da7ad772480f7fe5f28ef105227383e07a45
# Author: <NAME> <<EMAIL>>
# Date: Wed Aug 24 14:39:45 2016 -0600
#
# Added another line to the demo file
#
# commit <PASSWORD>
# Author: <NAME> <<EMAIL>>
# Date: Wed Aug 24 14:37:06 2016 -0600
#
# added demo file
# ```
#
# Through this cycle of editing, adding and committing, you can develop software in a linear fashion. Now let's see how we can create alternate versions.
# ## Branching
#
# **Now let's create a branch**
#
# ```bash
# $ git branch draft
#
# # This created a branch with the name draft. Let's look at all the other branches
# $ git branch
# draft
# * master
#
# ```
#
# We have two branches, draft and master. The * tells us the active branch (the HEAD).
#
# The files in your folders are in the state as they are stored in the active branch. When you change the branch the files are changed, removed or added to the state of the target branch.
#
# **Let's switch the branch.**
#
# ```bash
# $ git checkout draft
# Switched to branch 'draft'
# ```
#
# Let's see if there is something different
# ```bash
# $ cat demo.txt
# Hello World!
# Are you still spinning?
# ```
#
# No - it's the same! Now let's edit.
#
# ```bash
# $ echo "Spinning round and round" >> demo.txt
# $ cat demo.txt
# Hello World!
# Are you still spinning?
# Spinning round and round
# ```
#
#
# And commit
# ```bash
# $ git commit -a
# [draft 059daaa] Confirmed, spinning
# 1 file changed, 1 insertion(+)
# ```
#
# We have now written changes to the new branch, `draft`. The master branch should remain unchanged. Let's see if that's true.
#
# ```bash
# # Now let's switch back to master again
# $ git checkout master
# Switched to branch 'master'
#
# $ cat demo.txt
# Hello World!
# Are you still spinning?
# ```
#
# The text we added isn't here, as expected! Next we're going to change something in the main branch and thus cause a conflict.
#
# ```bash
# # Writing something to the front and to the end in an editor
# $ cat demo.txt
# I am here!
# Hello World!
# Are you still spinning?
# Indeed!
#
# # committing again
# $ git commit -a
# [master 8437327] Front and back
# 1 file changed, 2 insertions(+)
# ```
#
# At this point we have changed the file in two different branches of the repository. This is great for working on new features without breaking a stable codebase, but it can result in conflicts.
#
# **Let's try to merge those two branches.**
#
# ```bash
# # the git merge command merges the specified branch into the currently active one. "master" is active, and we want to merge in "draft".
# $ git merge draft
# # Auto-merging demo.txt
# # CONFLICT (content): Merge conflict in demo.txt
# # Automatic merge failed; fix conflicts and then commit the result.
#
# # The result
# $ cat demo.txt
# I am here!
# Hello World!
# Are you still spinning?
# <<<<<<< HEAD
# Indeed!
# =======
# Spinning round and round
# >>>>>>> draft
# ```
#
# The first line was merged without problems, The final line, where we have two alternative versions **is a conflict**. We have to manually resolve the conflict.
#
# Once this conflict is resolved, we can commit again.
#
# ```bash
# $ git commit -a
# [master 4dad82f] Merge branch 'draft'
#
# # Everything back in order.
# $ git status
# On branch master
# nothing to commit, working directory clean
# ```
#
# These are the basics of git on a local server. Now we'll learn how to sync with other people. This can be done with just git, but we'll be using GitHub as we're also using GitHub in the homeworks.
# ## Ignore Files
#
# When developing software, it's quite common that there are a lot of temporary files, e.g., created by Jupyter notebook to save temporary states. We shouldn't track temporary files, there is no reason to store them, and the can create conflicts.
#
# When you work with git on the command line, you have to manually add files you want to commit. But some GUI tools just add everything, so it's easy to add files you don't want.
#
# A good approach to avoid that is to use a `.gitignore` file. A gitignore file is a hidden file that contains file extensions that shouldn't be added to a git repository. For Jupyter notebooks, this is a minimal .gitignore file:
#
# ```bash
# # IPython Notebook
# .ipynb_checkpoints
# ```
# ## Other Files
#
# You should always add a `README.md` file that describes what the code in the repository does and how to run it.
#
# You should always add a license to your code. We recommend the BSD or MIT license, which are non-viral open source licenses.
# ## Jupyter Notebooks and Git
#
# Unfortunately, Jupyter Notebooks aren't handled well by git, as they mix code and output in the jupyter notebook file.
#
# Let's take a quick look at a notebook file [this is edited and cut]:
#
# ```json
# {
# "cells": [
# {
# "cell_type": "markdown",
# "metadata": {},
# "source": [
# "# Introduction to Data Science, CS 5963 / Math 3900\n",
# "*CS 5963 / MATH 3900, University of Utah, http://datasciencecourse.net/* \n",
# "\n",
# "## Lab 10: Classification\n",
# "\n",
# "In this lab, we will use the [scikit-learn](http://scikit-learn.org/) library to revisit the three classification methods we introduced: K-nearest neighbor, decision trees, and support vector machines. We will use a [dataset on contraceptive methods in Indonesia](https://archive.ics.uci.edu/ml/datasets/Contraceptive+Method+Choice).\n"
# ]
# },
# {
# "cell_type": "markdown",
# "metadata": {},
# "source": [
# "## The Data\n",
# "\n",
# "We will explore a dataset about the use of contraception in Indonesia. The dataset has 1473 records and the following attributes:\n",
# "\n",
# "1. Woman's age (numerical) \n",
# "2. Woman's education (categorical) 1=low, 2, 3, 4=high \n",
# "3. Husband's education (categorical) 1=low, 2, 3, 4=high \n",
# "4. Number of children ever born (numerical) \n",
# "5. Woman's religion (binary) 0=Non-Islam, 1=Islam \n",
# "6. Employed? (binary) 0=Yes, 1=No \n",
# "7. Husband's occupation (categorical) 1, 2, 3, 4 \n",
# "8. Standard-of-living index (categorical) 1=low, 2, 3, 4=high \n",
# "9. Media exposure (binary) 0=Good, 1=Not good \n",
# "10. Contraceptive method used (class attribute) 1=No-use, 2=Long-term, 3=Short-term"
# ]
# },
# {
# "cell_type": "markdown",
# "metadata": {},
# "source": [
# "### Hypothesis\n",
# "\n",
# "Write down which features do you think have the most impact on the use of contraception."
# ]
# },
# {
# "cell_type": "code",
# "execution_count": 2,
# "metadata": {
# "collapsed": false
# },
# "outputs": [
# {
# "data": {
# "text/html": [
# "<div>\n",
# "<table border=\"1\" class=\"dataframe\">\n",
# " <thead>\n",
# " <tr style=\"text-align: right;\">\n",
# " <th></th>\n",
# " <th>Age</th>\n",
# " <th>Education</th>\n",
# " <th>Husband-Education</th>\n",
# " <th>Children</th>\n",
# " <th>Religion</th>\n",
# ```
#
# Things like "outputs" and "execution_count" can change without any change to the notebooks functionality.
#
# So, what can you do?
#
# * Only commit clean notebooks, i.e., run "Restart and Clear Output" before committing pusing. This gets tedious, of course, if your script takes a long time to run.
# * Deal with conflicts (it's not too hard).
# * Work in pure python (not encouraged for project).
# * Synchronize with your collaborators over chat (...).
# * More sophisticated solutions [such as this one](https://gist.github.com/pbugnion/ea2797393033b54674af) (untested).
# * Hope and wait that Jupyter notebook will at some point separate input from output. (It's [looking good](https://github.com/jupyter/roadmap/blob/master/companion-files.md)).
#
# # Working with GitHub
#
# First, we'll create a new repository on github by going to [https://github.com/new](https://github.com/new).
#
# 
#
# Now let's clone the repository from GitHub.
#
# ```bash
# $ git clone https://github.com/alexsb/Demo.git
# ```
#
# Let's see how the config looks for this one.
# ```bash
# $ cat .git/config
# [core]
# repositoryformatversion = 0
# filemode = true
# bare = false
# logallrefupdates = true
# ignorecase = true
# precomposeunicode = true
# [remote "origin"]
# url = https://github.com/alexsb/demo.git
# fetch = +refs/heads/*:refs/remotes/origin/*
# [branch "master"]
# remote = origin
# merge = refs/heads/master
# ```
#
#
# This creates a local copy of the (empty) GitHub repository. We will just start working with that and commit and push the code to the server. If you'd like to add an existing repository to GitHub, follow [these instructions](https://help.github.com/articles/adding-an-existing-project-to-github-using-the-command-line/).
#
#
# ```bash
# # What's currently in the repository?
# $ ls
# LICENSE README.md
# ```
# Write something to demo.txt.
#
# ```bash
# $ echo "Hello world!" > demo.txt
# # echo "Hello world" > demo.txt
# ```
# Add demo.txt to the repository.
# ```bash
# $ git add demo.txt
# ```
# Commit the file to the repository.
#
# ```bash
# $ git commit -a -m "added demo file"
# [master 2e1918d] added demo file
# 1 file changed, 1 insertion(+)
# create mode 100644 demo.txt
# ```
#
# **Pushing it to the server!**
#
# ```bash
# $ git push
# Counting objects: 3, done.
# Delta compression using up to 8 threads.
# Compressing objects: 100% (2/2), done.
# Writing objects: 100% (3/3), 324 bytes | 0 bytes/s, done.
# Total 3 (delta 0), reused 0 (delta 0)
# To https://github.com/alexsb/demo.git
# 8e1ecd1..2e1918d master -> master
# ```
#
# We have now committed a file locally and pushed it to the server, i.e., our local copy is in sync with the server copy.
#
# Note that the `git push` command uses the origin defined in the config file. You can also push to other repositories!
#
# Next, we will make changes at another place. We'll use the **GitHub web interface** to do that.
#
# Once these changes are done, our local repository is out of sync with the remote repository. To get these changes locally, we have to pull from the repository:
#
# ```bash
# $ git pull
# remote: Counting objects: 3, done.
# remote: Compressing objects: 100% (2/2), done.
# remote: Total 3 (delta 1), reused 0 (delta 0), pack-reused 0
# Unpacking objects: 100% (3/3), done.
# From https://github.com/alexsb/demo
# 2e1918d..5dd3090 master -> origin/master
# Updating 2e1918d..5dd3090
# Fast-forward
# demo.txt | 1 +
# 1 file changed, 1 insertion(+)
# ```
#
# Let's see whether the changes are here
# ```bash
# $ cat demo.txt
# Hello world
# Are you still spinning?
# ```
# ### Other GitHub Features
#
# #### GitHub Issues
#
# Github Issues are a great way to keep track of open tasks and problems. You can tag people, add labels, add screenshots, etc.
#
# Issues can be referenced and closed from commits.
#
# ```
# git commit -a -m "added line, fixes #1"
# ```
#
# This is a very transparent way of coding, as it exactly tracks which which commit addresses wich task or issue.
#
#
# #### Forking
#
# Forking is essentially making use of the distributed nature of git, while having the benefits of a server. When you fork a repository you make a clone of someone else's code that you are not allowed to read. The repository appears in your github account and you can start editing the code. If you think you improved the code, you can send a "pull request" to the original owner. The owner can then review your code and merge your modifications into his main repository. Forking is hence virtually the same as branching, with the exception that it resolves issues with write permissions.
#
# ### GUI Clients
#
# * **GitHub Desktop**
# Good option if you want a GUI client. [Download here](https://desktop.github.com/)
# * **Integrated in IDEs**
# Many operations can be done out of a IDE such as WebStorm
#
# ### Getting updates to the lectures and homeworks
#
# The labs and homeworks are hosted in a git repository. Every time we release a lecture or homework we will just update the repository. You can then pull from that repository to get the latest lecture/homework on your computer.
#
# To get the homework repository, run the following:
#
# ```bash
# $ git clone https://github.com/datascience-course/2018-datascience-homeworks -o homework
# $ cd 2018-datascience-homeworks
# ```
#
# Note that by using the `-o homework` option we're not using the default remote `origin` but a user-defined remote called `homework`.
#
# Now, you can manage your own implementations in the homeworks, while keeping in sync with the releases by create a new repository on the Github.
#
# Ensure your new repository is private and don't click the option to "Initialize the repository with a README".
#
# Run the two commands described on GitHub under the heading "Push an existing repository from the command line". For my repository these are:
#
# ```bash
# $ git remote add origin https://github.com/alexsb/datascience-hw-lex-alexander.git
# $ git push -u origin master
# ```
#
# Now your homework repository is all set!
#
# #### Committing
#
# While working on homework assignments, periodically run the following:
#
# ```bash
# $ git add -A
# $ git commit -m "Describe your changes"
# $ git push
# ```
#
# Remember, the `git commit` operation takes a snapshot of your code at that point in time but doesn't write to the server.
#
# The `git push` operation pushes your local commits to the remote repository.
#
# You should do this frequently: as often as you have an incremental, standalone improvement.
#
# #### Getting new homework assignments
#
# When we release a new assignment we will simply add it to the homework github repository.
#
# To get the latest homework assignments and potential updates or corrections to the assignment, run the following.
#
# ```bash
# $ git pull homework master
# ```
#
# Make sure to have all your changes committed before you do that.
| 02-basic-python/lecture-02-version-control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sweetpand/Algorithms/blob/master/Amazon_130_Leetcode.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="1E9t7j7k1VKo" colab_type="text"
#
#
# ```
# Given a 2D board containing 'X' and 'O' (the letter O), capture all regions surrounded by 'X'.
#
# A region is captured by flipping all 'O's into 'X's in that surrounded region.
#
# Example:
#
# X X X X
# X O O X
# X X O X
# X O X X
# After running your function, the board should be:
#
# X X X X
# X X X X
# X X X X
# X O X X
# Explanation:
#
# Surrounded regions shouldn’t be on the border, which means that any 'O' on the border of the board are not flipped to 'X'. Any 'O' that is not on the border and it is not connected to an 'O' on the border will be flipped to 'X'. Two cells are connected if they are adjacent cells connected horizontally or vertically.
# ```
#
#
# + [markdown] id="rbwCun3B8lJK" colab_type="text"
# Solution 1. Using BFS
# + id="lnMYCgRM0yXM" colab_type="code" colab={}
from typing import List
from collections import deque
class Solution:
paths = [[-1, 0], [1, 0], [0, -1], [0, 1]]
def solve(self, board: List[List[str]]) -> None:
if not board or not board[0]:
return []
m = len(board)
n = len(board[0])
q = deque()
visited = [[0]*n for _ in range(m)]
for i in range(m):
if board[i][0] == 'O':
q.append([i, 0])
visited[i][0] = 1
if board[i][n-1] == 'O':
q.append([i, n-1])
visited[i][n-1] = 1
for i in range(n):
if board[0][i] == 'O':
q.append([0, i])
visited[0][i] = 1
if board[m-1][i] == 'O':
q.append([m - 1, i])
visited[m - 1][i] = 1
while q:
x, y = q.popleft()
board[x][y] = 'F'
for a, b in self.paths:
nx, ny = x + a, y + b
if 0 <= nx < m and 0 <= ny < n and board[nx][ny] == 'O' and not visited[nx][ny]:
q.append([nx, ny])
visited[nx][ny] = 1
for i in range(m):
for j in range(n):
if board[i][j] == 'O':
board[i][j] = 'X'
if board[i][j] == 'F':
board[i][j] = 'O'
# + [markdown] id="UHMRyuE_8q67" colab_type="text"
# Solution 2. Using DFS (depth first search)
# + id="75ZrAevG8wvl" colab_type="code" colab={}
from typing import List
class Solution:
paths = [[-1, 0], [1, 0], [0, -1], [0, 1]]
def dfs(self, board, i, j, m, n):
if 0 <= i < m and 0 <= j < n and board[i][j] == 'O':
board[i][j] = 'F'
for x, y in self.paths:
ni, nj = i + x, j + y
board = self.dfs(board, ni, nj, m, n)
return board
def solve(self, board: List[List[str]]) -> None:
"""
Do not return anything, modify board in-place instead.
"""
if not board or not board[0]:
return []
m = len(board)
n = len(board[0])
# print(board)
if m == 0 or n == 0:
return []
for i in range(m):
if board[i][0] == 'O':
board = self.dfs(board, i, 0, m, n)
if board[i][n -1] == 'O':
board = self.dfs(board, i, n - 1, m, n)
for i in range(n):
if board[0][i] == 'O':
board = self.dfs(board, 0, i, m, n)
if board[m-1][i] == 'O':
board = self.dfs(board, m - 1, i, m, n)
for i in range(m):
for j in range(n):
if board[i][j] == 'O':
board[i][j] = 'X'
if board[i][j] == 'F':
board[i][j] = 'O'
| Amazon_130_Leetcode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##https://towardsdatascience.com/simple-and-multiple-linear-regression-in-python-c928425168f9
import statsmodels.api as sm
from sklearn import datasets ## imports datasets from scikit-learn
data = datasets.load_boston()
import numpy as np
import pandas as pd
# +
# define the data/predictors as the pre-set feature names
df = pd.DataFrame(data.data, columns=data.feature_names)
# Put the target (housing value -- MEDV) in another DataFrame
target = pd.DataFrame(data.target, columns=["MEDV"])
# +
X = df["RM"] ## X usually means our input variables (or independent variables)
y = target["MEDV"] ## Y usually means our output/dependent variable
X = sm.add_constant(X) ## let's add an intercept (beta_0) to our model
# Note the difference in argument order
model = sm.OLS(y, X).fit() ## sm.OLS(output, input)
predictions = model.predict(X)
# Print out the statistics
model.summary()
# -
| code/linear_model.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # balance_group_sat
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/examples/balance_group_sat.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/python/balance_group_sat.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010-2021 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""We are trying to group items in equal sized groups.
Each item has a color and a value. We want the sum of values of each group to
be as close to the average as possible.
Furthermore, if one color is an a group, at least k items with this color must
be in that group.
"""
from ortools.sat.python import cp_model
# Create a solution printer.
class SolutionPrinter(cp_model.CpSolverSolutionCallback):
"""Print intermediate solutions."""
def __init__(self, values, colors, all_groups, all_items, item_in_group):
cp_model.CpSolverSolutionCallback.__init__(self)
self.__solution_count = 0
self.__values = values
self.__colors = colors
self.__all_groups = all_groups
self.__all_items = all_items
self.__item_in_group = item_in_group
def on_solution_callback(self):
print('Solution %i' % self.__solution_count)
self.__solution_count += 1
print(' objective value = %i' % self.ObjectiveValue())
groups = {}
sums = {}
for g in self.__all_groups:
groups[g] = []
sums[g] = 0
for item in self.__all_items:
if self.BooleanValue(self.__item_in_group[(item, g)]):
groups[g].append(item)
sums[g] += self.__values[item]
for g in self.__all_groups:
group = groups[g]
print('group %i: sum = %0.2f [' % (g, sums[g]), end='')
for item in group:
value = self.__values[item]
color = self.__colors[item]
print(' (%i, %i, %i)' % (item, value, color), end='')
print(']')
# Data.
num_groups = 10
num_items = 100
num_colors = 3
min_items_of_same_color_per_group = 4
all_groups = range(num_groups)
all_items = range(num_items)
all_colors = range(num_colors)
# Values for each items.
values = [1 + i + (i * i // 200) for i in all_items]
# Color for each item (simple modulo).
colors = [i % num_colors for i in all_items]
sum_of_values = sum(values)
average_sum_per_group = sum_of_values // num_groups
num_items_per_group = num_items // num_groups
# Collect all items in a given color.
items_per_color = {}
for c in all_colors:
items_per_color[c] = []
for i in all_items:
if colors[i] == c:
items_per_color[c].append(i)
print('Model has %i items, %i groups, and %i colors' %
(num_items, num_groups, num_colors))
print(' average sum per group = %i' % average_sum_per_group)
# Model.
model = cp_model.CpModel()
item_in_group = {}
for i in all_items:
for g in all_groups:
item_in_group[(i, g)] = model.NewBoolVar('item %d in group %d' %
(i, g))
# Each group must have the same size.
for g in all_groups:
model.Add(
sum(item_in_group[(i, g)]
for i in all_items) == num_items_per_group)
# One item must belong to exactly one group.
for i in all_items:
model.Add(sum(item_in_group[(i, g)] for g in all_groups) == 1)
# The deviation of the sum of each items in a group against the average.
e = model.NewIntVar(0, 550, 'epsilon')
# Constrain the sum of values in one group around the average sum per group.
for g in all_groups:
model.Add(
sum(item_in_group[(i, g)] * values[i]
for i in all_items) <= average_sum_per_group + e)
model.Add(
sum(item_in_group[(i, g)] * values[i]
for i in all_items) >= average_sum_per_group - e)
# color_in_group variables.
color_in_group = {}
for g in all_groups:
for c in all_colors:
color_in_group[(c, g)] = model.NewBoolVar(
'color %d is in group %d' % (c, g))
# Item is in a group implies its color is in that group.
for i in all_items:
for g in all_groups:
model.AddImplication(item_in_group[(i, g)],
color_in_group[(colors[i], g)])
# If a color is in a group, it must contains at least
# min_items_of_same_color_per_group items from that color.
for c in all_colors:
for g in all_groups:
literal = color_in_group[(c, g)]
model.Add(
sum(item_in_group[(i, g)] for i in items_per_color[c]) >=
min_items_of_same_color_per_group).OnlyEnforceIf(literal)
# Compute the maximum number of colors in a group.
max_color = num_items_per_group // min_items_of_same_color_per_group
# Redundant contraint: The problem does not solve in reasonable time without it.
if max_color < num_colors:
for g in all_groups:
model.Add(
sum(color_in_group[(c, g)] for c in all_colors) <= max_color)
# Minimize epsilon
model.Minimize(e)
model.ExportToFile('balance_group_sat.pbtxt')
solver = cp_model.CpSolver()
solution_printer = SolutionPrinter(values, colors, all_groups, all_items,
item_in_group)
status = solver.Solve(model, solution_printer)
if status == cp_model.OPTIMAL:
print('Optimal epsilon: %i' % solver.ObjectiveValue())
print('Statistics')
print(' - conflicts : %i' % solver.NumConflicts())
print(' - branches : %i' % solver.NumBranches())
print(' - wall time : %f s' % solver.WallTime())
else:
print('No solution found')
| examples/notebook/examples/balance_group_sat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 给定树的前序和中序遍历,构造二叉树。
#
# 注意:
# 您可以假设树中不存在重复项。
# -
# <img src='105.jpg' width=500>
# <img src='105_1.jpg' width=500>
# + jupyter={"source_hidden": true}
class TreeNode:
def __init__(self, val=0, left=None, right=None):
self.val = val
self.left = left
self.right = right
class Solution:
def buildTree(self, preorder, inorder) -> TreeNode:
root_val = preorder[0]
root = TreeNode(root_val)
index = inorder.index(root_val)
self.count = 0
self.helper(root, preorder, inorder[:index], inorder[index+1:])
return root
def helper(self, root, preorder, left, right):
if self.count == len(preorder) - 1:
return
val = preorder[self.count + 1]
if val in left: # val的值在左边
print(val, self.count)
root.left = TreeNode(val)
self.count += 1
index = left.index(val)
self.helper(root.left, preorder, left[:index], left[index+1:])
val = preorder[self.count + 1]
if val in right: # val的值在右边
root.right = TreeNode(val)
self.count += 1
index = right.index(val)
self.helper(root, preorder, right[:index], right[index+1:])
# +
class TreeNode:
def __init__(self, val=0, left=None, right=None):
self.val = val
self.left = left
self.right = right
class Solution:
def buildTree(self, preorder, inorder) -> TreeNode:
if inorder:
val = preorder.pop(0)
root = TreeNode(val)
index = inorder.index(val) # 当前节点在 inorder中的下标
root.left = self.buildTree(preorder, inorder[:index]) # 先构建左子树
root.right = self.buildTree(preorder, inorder[index+1:])
return root
else:
return None
# -
# +
t1 = TreeNode(3)
t2 = TreeNode(9)
t3 = TreeNode(20)
t4 = TreeNode(15)
t5 = TreeNode(7)
t1.left = t2
t1.right = t3
t3.left = t4
t3.right = t5
root_ = t1
solution = Solution()
solution.buildTree([3, 9, 20, 15, 7], [9, 3, 15, 20, 7])
# -
a = []
if not a:
print(a)
| Tree/0829/105. Construct Binary Tree from Preorder and Inorder Traversal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial - Plotting F' and F'' Schematas
# This tutorial shows how to plot Prime Implicants (F') and Two-Symbol (F'') schematas
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.text import Text
from matplotlib.patches import Circle, Rectangle, RegularPolygon
from matplotlib.collections import PatchCollection
from copy import copy
from cana.datasets.bio import THALIANA #, DROSOPHILA, BUDDING_YEAST
N = THALIANA()
#N = DROSOPHILA()
#N = BUDDING_YEAST()
print(N)
def plot_schemata(n):
# Init values from BooleanNode
k = n.k if n.k>=1 else 1
inputs = n.inputs if not n.constant else [n.name]
inputlabels = [n.network.get_node_name(i)[0] if n.network is not None else i for i in inputs]
pi0s = n._prime_implicants.get('0', [])
pi1s = n._prime_implicants.get('1', [])
ts0s = n._two_symbols[0]
ts1s = n._two_symbols[1]
# Count number of PI and TS
n_pi = sum(len(pis) for pis in [pi0s,pi1s])
n_ts = sum(len(tss) for tss in [ts0s,ts1s])
# Schemata Cell Width and spacing
cwidth = 60.
cxspace = 0
cyspace = 6
border = 1
sepcxspace = 21
sepcyspace = 15
dpi = 150.
# Margins
top, right, bottom, left, hs = 160, 25, 25, 60, 60
# Axes Width & Height
ax1width = ((k*(cwidth+cxspace))+sepcxspace+(cwidth))
ax1height = (n_pi*(cwidth+cyspace)+sepcyspace-cyspace)
ax2width = ((k*(cwidth+cxspace))+sepcxspace+(cwidth))
ax2height = (n_ts*(cwidth+cyspace)+sepcyspace-cyspace)
# Figure Width & Height
fwidth = (left + ax1width + hs + ax2width + right)
fheight = (bottom + max(ax1height,ax2height) + top)
# Percentages for Axes location
_ax1w = ((ax1width*100) / fwidth) / 100
_ax2w = ((ax2width*100) / fwidth) / 100
_ax1h = ((ax1height*100) / fheight) / 100
_ax2h = ((ax2height*100) / fheight) / 100
_bottom = ((bottom*100) / fheight) / 100
_left = ((left*100) / fwidth) / 100
_hs = ((hs*100) / fwidth) / 100
# Init Figure
fig = plt.figure(figsize=(fwidth/dpi,fheight/dpi), facecolor='w', dpi=dpi)
ax1 = fig.add_axes((_left,_bottom,_ax1w,_ax1h), aspect=1, label='PI')
ax2 = fig.add_axes((_left+_ax1w+_hs,_bottom,_ax2w,_ax1h), aspect=1, label='TS')
### PI Plot ###
yticks = []
patches = []
x,y = 0.,0.
#
for out,pis in zip([1,0],[pi1s,pi0s]):
for pi in pis:
x = 0.
xticks = []
for input in pi:
if input == '0':
facecolor = 'white'
textcolor = 'black'
elif input == '1':
facecolor = 'black'
textcolor = 'white'
elif input == '#':
facecolor = '#cccccc'
textcolor = 'black'
text = '%s'%(input) if (input!='2') else '#'
ax1.add_artist(Text(x+cwidth/2,y+cwidth/10*4, text=text, color=textcolor, va='center', ha='center',fontsize=14,family='serif'))
r = Rectangle((x,y), width=cwidth, height=cwidth, facecolor=facecolor, edgecolor='black')
patches.append(r)
xticks.append(x+cwidth/2)
x += cwidth + cxspace
x += sepcxspace
r = Rectangle((x,y), width=cwidth, height=cwidth, facecolor='black' if (out==1) else 'white', edgecolor='black')
ax1.add_artist(Text(x-(sepcxspace/2)-(cxspace/2),y+cwidth/10*4, text=':', color='black', va='center', ha='center',fontsize=14,weight='bold',family='serif'))
ax1.add_artist(Text(x+(cwidth/2),y+cwidth/10*4, text=out, color='white' if (out==1) else 'black', va='center', ha='center',fontsize=14,family='serif'))
patches.append(r)
xticks.append(x+cwidth/2)
yticks.append(y+cwidth/2)
y += cwidth + cyspace
y += sepcyspace
ax1.add_collection(PatchCollection(patches,match_original=True))
#
ax1.set_yticks(yticks)
ax1.set_yticklabels([r"$f^{'}_{%d}$"%(i+1) for i in range(n_pi)[::-1]], fontsize=14)
ax1.set_xticks(xticks)
ax1.set_xticklabels(inputlabels + ['%s'%(n.name)], rotation=90, fontsize=14)
#
ax1.xaxis.tick_top()
# Remove Tick
ax1.tick_params(which='major',pad=7)
for tic in ax1.xaxis.get_major_ticks():
tic.tick1On = tic.tick2On = False
for tic in ax1.yaxis.get_major_ticks():
tic.tick1On = tic.tick2On = False
# Remove Border
ax1.spines['top'].set_visible(False)
ax1.spines['right'].set_visible(False)
ax1.spines['bottom'].set_visible(False)
ax1.spines['left'].set_visible(False)
# Limits
ax1.set_xlim(-border,ax1width+border)
ax1.set_ylim(-border,ax1height+border)
#ax1.invert_yaxis()
## TS ##
t = 0
x,y = 0.,0.
yticks = []
boxes,symbols = [], []
#
tssymbols = [
Circle((0, 0), radius=5, facecolor='white', edgecolor='black'),
RegularPolygon((0, 0),numVertices=3, radius=5, orientation=0, facecolor='white', edgecolor='black'),
]
#
for out,tss in zip([1,0],[ts1s,ts0s]):
for ts,pss,sss in tss:
x = 0.
xticks = []
for i,input in enumerate(ts):
if input == '0':
facecolor = 'white'
textcolor = 'black'
elif input == '1':
facecolor = 'black'
textcolor = 'white'
elif input == '2':
facecolor = '#cccccc'
textcolor = 'black'
if len(pss):
# TODO: If there are several symbols in the same input position, place them side-by-side
iinpss = [j for j,ps in enumerate(pss) if i in ps]
xpos = np.linspace(x,x+cwidth, len(iinpss)+2)
for z,j in enumerate(iinpss,start=1):
s = copy(tssymbols[j])
s.xy = (xpos[z],y+cwidth*0.8)
s.center = xpos[z],y+cwidth*0.8 # A hack for circles only
s.set_edgecolor('#a6a6a6' if (input=='1') else 'black')
symbols.append(s)
ax2.add_patch(s)
text = '%s'%(input) if (input!='2') else '#'
ax2.add_artist(Text(x+cwidth/2,y+cwidth/10*4, text=text, color=textcolor, va='center', ha='center',fontsize=14,family='serif'))
r = Rectangle((x,y), width=cwidth, height=cwidth, facecolor=facecolor, edgecolor='#4c4c4c',zorder=2)
boxes.append(r)
xticks.append(x+cwidth/2)
x += cwidth + cxspace
x += sepcxspace
r = Rectangle((x,y), width=cwidth, height=cwidth, facecolor='black' if (out==1) else 'white', edgecolor='#4c4c4c')
ax2.add_artist(Text(x-(sepcxspace/2)-(cxspace/2),y+cwidth/2, text=':', color='black', va='center', ha='center',fontsize=14,weight='bold',family='serif'))
ax2.add_artist(Text(x+(cwidth/2),y+cwidth/10*4, text=out, color='white' if (out==1) else 'black', va='center', ha='center',fontsize=14,family='serif'))
boxes.append(r)
xticks.append(x+cwidth/2)
yticks.append(y+cwidth/2)
y += cwidth + cyspace
t += 1
y += sepcyspace
if len(boxes):
ax2.add_collection(PatchCollection(boxes,match_original=True))
if len(symbols):
ax2.add_collection(PatchCollection(symbols,match_original=True))
#
ax2.set_yticks(yticks)
ax2.set_yticklabels([r"$f^{''}_{%d}$"%(i+1) for i in range(n_ts)[::-1]], fontsize=14)
ax2.set_xticks(xticks)
ax2.set_xticklabels(inputlabels + ['%s'%(n.name)], rotation=90, fontsize=14)
#
ax2.xaxis.tick_top()
# Remove Tick
ax2.tick_params(which='major',pad=7)
for tic in ax2.xaxis.get_major_ticks():
tic.tick1On = tic.tick2On = False
for tic in ax2.yaxis.get_major_ticks():
tic.tick1On = tic.tick2On = False
# Remove Border
ax2.spines['top'].set_visible(False)
ax2.spines['right'].set_visible(False)
ax2.spines['bottom'].set_visible(False)
ax2.spines['left'].set_visible(False)
# Limits
ax2.set_xlim(-border,ax2width+border)
ax2.set_ylim(-border,ax2height+border)
# FileName
filename = n.name.replace('/','_')
filename = filename.replace(',','_')
## Display ##
display(fig)
plt.close()
node = N.nodes[2]
# Compute Prime Implicants & Two-Symbol schematas
node._check_compute_canalization_variables(prime_implicants=True)
node._check_compute_canalization_variables(two_symbols=True)
# Plot
plot_schemata(node)
| tutorials/Canalization - BioModels - Schematas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Collin-Campbell/DS-Unit-2-Linear-Models/blob/master/module2-regression-2/Lecture_2_notes.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="E5WTvJFCWQjx"
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + [markdown] id="TGelVAREWyrN"
# # 1. Wrangle Data
# + [markdown] id="wzyKDCxhdABb"
# ## 1.2. Import
# + id="htqZ_oVnWxUl"
# Import
import pandas as pd
df = pd.read_csv(DATA_PATH+'elections/bread_peace_voting.csv', index_col='Year')
# + [markdown] id="GeUEq0ARdB-l"
# ## EDA
# + id="gplXOI-Jnlgb" outputId="41b94147-fc98-4e4a-bd49-a74627113c55" colab={"base_uri": "https://localhost:8080/", "height": 618}
df
# + id="Xn8y2BfupUNM"
incomes = 'Average Recent Growth in Personal Incomes'
fatalities = 'US Military Fatalities per Million'
votes = 'Incumbent Party Vote Share'
# + [markdown] id="xvI8_xb5pslR"
# Is there a linear relationship between incomes and votes?
# + id="duDOtHvsprla" outputId="55f2e2ed-f048-417f-8674-01fb538080f4" colab={"base_uri": "https://localhost:8080/", "height": 282}
import matplotlib.pyplot as plt
plt.scatter(df[incomes], df[votes])
plt.xlabel('Recent Growth in Personal Incomes')
plt.ylabel('Incumbent Party Vote Share [%]');
# + id="Vt1-R6oFqfn2" outputId="26af7ba3-9fbd-4f51-e70e-58f6327d37da" colab={"base_uri": "https://localhost:8080/", "height": 282}
plt.scatter(df[fatalities], df[votes])
plt.xlabel('US Military Fatalities per Million')
plt.ylabel('Incumbent Party Vote Share [%]');
# + [markdown] id="l688_ftdeT6O"
# # 2. Split Data
# + [markdown] id="n4sFhCGRrBvy"
# ## 2.1 Split our **target vector** from our **feature matrix**
# + id="d-4U9smurAvs"
y = df[votes]
X = df[[incomes, fatalities]]
# + [markdown] id="D3GFerxQrpIA"
# ## 2.2. Split our data into **training** and **test** sets
#
# Two strategies:
#
# - Random train-test split using [`train_test_split`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html). Generally we use 80% of the data for training, and 20% of the data for testing.
# - If you have **timeseries**, then you need to do a "cutoff" split.
# + id="MAdcxm22rkfo"
# Train on data before 2008 and test on data 2008-present
cutoff = 2008
mask = X.index < 2008
X_train, y_train = X.loc[mask], y.loc[mask]
X_test, y_test = X.loc[~mask], y.loc[~mask]
# + id="Wr_WWRqxtfP_"
assert len(X) == len(X_train) + len(X_test)
# + [markdown] id="FivqV88YlglV"
# # Establish Baseline
#
# - **Q**: Is this a regression or a classification problem?
# - **A**: Regression
# + id="4gkloq_ZwgNt" outputId="34291d7b-24c6-4720-f9a2-c800070b29bd" colab={"base_uri": "https://localhost:8080/", "height": 35}
from sklearn.metrics import mean_absolute_error
y_pred = [y_train.mean()] * len(y_train)
print('Baseline MAE:', mean_absolute_error(y_train, y_pred))
# + id="vQCBB1G6yL2j" outputId="838a2351-6d27-44dd-d9f9-a2efffd5debf" colab={"base_uri": "https://localhost:8080/", "height": 282}
col = 'Average Recent Growth in Personal Incomes'
plt.scatter(X_train[col], y_train)
plt.plot(X_train[col],
y_pred,
color='grey',
label='Baseline Model')
plt.xlabel('Recent Growth in Personal Incomes')
plt.ylabel('INCUMBENT Vote Share')
plt.legend();
# + [markdown] id="wRx_D8Vjoyq9"
# # Build Model
#
# To start, let's use one feature only.
# + id="86X8WTZay3H3" outputId="04d5b26e-cf54-4d13-ef0e-1022655a6013" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Step 1: Import our predictor
from sklearn.linear_model import LinearRegression
# Step 2: Instantiate our predictor
model = LinearRegression()
# Step 3: Fit our model to the TRAINING DATA
model.fit(X_train[[col]], y_train)
# Step 4: Make predictions... see below
# + [markdown] id="A-VQpOHgqlae"
# # Check Metrics
#
# ## Mean Absolute Error
# + id="wL3nCzy8zja8" outputId="5bad3bc1-4452-4f0a-c37f-7aed1d17afe7" colab={"base_uri": "https://localhost:8080/", "height": 52}
print('Training MAE:', mean_absolute_error(y_train, model.predict(X_train[[col]])))
print('Testing MAE:', mean_absolute_error(y_test, model.predict(X_test[[col]])))
# + [markdown] id="MuPtGBMGvSA4"
# ## Root Mean Squared Error
# + id="tMc04Jv-1aXN" outputId="744ce4f3-a620-4e9f-9531-4784d7859a88" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.metrics import mean_squared_error
print('Training RMSE:', mean_squared_error(y_train, model.predict(X_train[[col]]), squared=False))
print('Testing RMSE:', mean_squared_error(y_test, model.predict(X_test[[col]]), squared=False))
# + [markdown] id="BnSZ4IOJvWxD"
# ## $R^2$
# + id="qZhhhU9Wu4kD" outputId="a9782ee1-eb9b-4c6d-8792-1d8189994fe0" colab={"base_uri": "https://localhost:8080/", "height": 383, "referenced_widgets": ["6d6923e272a645218b7a6cfad3db7f3b", "<KEY>", "<KEY>", "b49c8ce5c5054a95974e41c3149ff0be", "de231edf52834d7886adcc0876889ca0", "47597f86046941d2ad01e8038c6f9cb6", "20e8bac452aa483e8ddd1180f0e113f6", "128f39a70fa5426fbcef87f38ede1624", "4aeadb1c3d7c4bf6af55a44be86acaad", "123ffd48a6c544c8b8ba12baeeeed994"]}
from ipywidgets import interactive, IntSlider, FloatSlider
from math import floor, ceil
from sklearn.base import BaseEstimator, RegressorMixin
import numpy as np
class BruteForceRegressor(BaseEstimator, RegressorMixin):
def __init__(self, m=0, b=0):
self.m = m
self.b = b
self.mean = 0
def fit(self, X, y):
self.mean = np.mean(y)
return self
def predict(self, X, return_mean=True):
if return_mean:
return [self.mean] * len(X)
else:
return X * self.m + self.b
def plot(slope, intercept):
# Assign data to variables
x = df['Average Recent Growth in Personal Incomes']
y = df['Incumbent Party Vote Share']
# Create figure
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,6))
# Set ax limits
mar = 0.2
x_lim = floor(x.min() - x.min()*mar), ceil(x.max() + x.min()*mar)
y_lim = floor(y.min() - y.min()*mar), ceil(y.max() + y.min()*mar)
# Instantiate and train model
bfr = BruteForceRegressor(slope, intercept)
bfr.fit(x, y)
# ax1
## Plot data
ax1.set_xlim(x_lim)
ax1.set_ylim(y_lim)
ax1.scatter(x, y)
## Plot base model
ax1.axhline(bfr.mean, color='orange', label='baseline model')
## Plot residual lines
y_base_pred = bfr.predict(x)
ss_base = mean_squared_error(y, y_base_pred) * len(y)
for x_i, y_i, yp_i in zip(x, y, y_base_pred):
ax1.plot([x_i, x_i], [y_i, yp_i],
color='gray', linestyle='--', alpha=0.75)
## Formatting
ax1.legend()
ax1.set_title(f'Sum of Squares: {np.round(ss_base, 2)}')
ax1.set_xlabel('Growth in Personal Incomes')
ax1.set_ylabel('Incumbent Party Vote Share [%]')
# ax2
ax2.set_xlim(x_lim)
ax2.set_ylim(y_lim)
## Plot data
ax2.scatter(x, y)
## Plot model
x_model = np.linspace(*ax2.get_xlim(), 10)
y_model = bfr.predict(x_model, return_mean=False)
ax2.plot(x_model, y_model, color='green', label='our model')
for x_coord, y_coord in zip(x, y):
ax2.plot([x_coord, x_coord], [y_coord, x_coord * slope + intercept],
color='gray', linestyle='--', alpha=0.75)
ss_ours = mean_squared_error(y, bfr.predict(x, return_mean=False)) * len(y)
## Formatting
ax2.legend()
ax2.set_title(f'Sum of Squares: {np.round(ss_ours, 2)}')
ax2.set_xlabel('Growth in Personal Incomes')
ax2.set_ylabel('Incumbent Party Vote Share [%]')
y = df['Incumbent Party Vote Share']
slope_slider = FloatSlider(min=-5, max=5, step=0.5, value=0)
intercept_slider = FloatSlider(min=int(y.min()), max=y.max(), step=2, value=y.mean())
interactive(plot, slope=slope_slider, intercept=intercept_slider)
# + id="5Z4LUDqF5Sgf" outputId="ca1912fd-68fc-475b-8d55-83841a9612d1" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.metrics import r2_score
print('Training R^2:', r2_score(y_train, model.predict(X_train[[col]])))
print('Testing R^2:', r2_score(y_test, model.predict(X_test[[col]])))
# + [markdown] id="LER3Ft-kz_Jf"
# # And finally... Multiple Linear Regression
# + id="4RbzxHhE0W67" outputId="25c6ba08-4653-4d55-a295-8492099ec8f9" colab={"base_uri": "https://localhost:8080/", "height": 35}
# Step 2: Instantiate our predictor
model2 = LinearRegression()
# Step 3: Fit our model to the TRAINING DATA
model2.fit(X_train, y_train)
# + id="7FNqfoxI8hxG" outputId="7b0688d1-4128-4ff9-8316-d80434c8920e" colab={"base_uri": "https://localhost:8080/", "height": 52}
print('Training R^2:', model2.score(X_train, y_train))
print('Testing R^2:', model2.score(X_test, y_test))
# + [markdown] id="ndH-ZVZE84Tt"
# # Explain Model
# + id="hvs90SVu9Mq9" outputId="92d646ee-6c87-4091-a2f3-01d06c1a9e80" colab={"base_uri": "https://localhost:8080/", "height": 35}
X_train.columns
# + id="hJaJAg2E8xg6" outputId="446eaa9f-b6e6-4324-c995-88313681f39a" colab={"base_uri": "https://localhost:8080/", "height": 35}
model2.coef_
# + id="AsJf46A-9CT1" outputId="5239f035-1784-4983-ab58-58593a113d27" colab={"base_uri": "https://localhost:8080/", "height": 35}
model2.intercept_
# + id="HDP5XRcg9Fw-"
| module2-regression-2/Lecture_2_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: crypten-pycon
# language: python
# name: crypten-pycon
# ---
# # Training a Model on Encrypted Data
# +
import sys
import torch
import torchvision
import matplotlib.pyplot as plt
# python 3.7 is required
assert sys.version_info[0] == 3 and sys.version_info[1] == 7, "python 3.7 is required"
import crypten
crypten.init()
# %matplotlib inline
# -
# ## MNIST
# +
digits = torchvision.datasets.MNIST(root='/tmp/data',
train=True,
transform=torchvision.transforms.ToTensor(),
download=True)
digits_test = torchvision.datasets.MNIST(root='/tmp/data',
train=True,
transform=torchvision.transforms.ToTensor(),
download=True)
# -
plt.imshow(digits[0][0][0], cmap='gray', interpolation='none')
print("label for image is ", digits[0][1])
# ## Preprocess Into Tensors
def take_samples(digits, n_samples=1000):
"""Returns images and labels based on sample size"""
images, labels = [], []
for i, digit in enumerate(digits):
if i == n_samples:
break
image, label = digit
images.append(image)
label_one_hot = torch.nn.functional.one_hot(torch.tensor(label), 10)
labels.append(label_one_hot)
images = torch.cat(images)
labels = torch.stack(labels)
return images, labels
images, labels = take_samples(digits, n_samples=100)
print(images.shape)
print(labels.shape)
images_enc = crypten.cryptensor(images)
labels_enc = crypten.cryptensor(labels)
images_enc[0]
# test set
images_test, labels_test = take_samples(digits_test, n_samples=20)
images_test_enc = crypten.cryptensor(images_test)
labels_test_enc = crypten.cryptensor(labels_test)
# ## Logistic Regression Model
#
# (multiclass logistic regression)
# +
class LogisticRegression(crypten.nn.Module):
def __init__(self):
super().__init__()
# images are 28x28 pixels
self.linear = crypten.nn.Linear(28 * 28, 10)
def forward(self, x):
return self.linear(x)
# -
model = LogisticRegression().encrypt()
model(images_enc)
# ### Train Model on Encrypted Images
def train_model(model, X, y, epochs=10, learning_rate=0.05):
criterion = crypten.nn.CrossEntropyLoss()
for epoch in range(epochs):
model.zero_grad()
output = model(X)
loss = criterion(output, y)
print(f"epoch {epoch} loss: {loss.get_plain_text()}")
loss.backward()
model.update_parameters(learning_rate)
return model
model = train_model(model, images_enc, labels_enc)
# ### Decrypt Prediction
prediction = model(images_enc[3].unsqueeze(0))
prediction.get_plain_text().argmax()
plt.imshow(images[3], cmap='gray', interpolation='none')
# ### Test Model Accuracy
def avg_test_accuracy(model, X, y):
output = model(X).get_plain_text().softmax(0)
predicted = output.argmax(1)
labels = y.get_plain_text().argmax(1)
correct = (predicted == labels).sum().float()
return float(correct / y.shape[0])
avg_test_accuracy(model, images_enc, labels_enc)
# ## Training a CNN
# based on https://github.com/pytorch/examples/blob/master/mnist/main.py
class CNN(crypten.nn.Module):
def __init__(self):
super().__init__()
self.conv1 = crypten.nn.Conv2d(1, 32, 3, 1)
self.conv2 = crypten.nn.Conv2d(32, 64, 3, 1)
self.dropout1 = crypten.nn.Dropout2d(0.25)
self.dropout2 = crypten.nn.Dropout2d(0.5)
self.fc1 = crypten.nn.Linear(9216, 128)
self.fc2 = crypten.nn.Linear(128, 10)
def forward(self, x):
x = x.unsqueeze(1)
x = self.conv1(x)
x = x.relu()
x = self.conv2(x)
x = x.relu()
x = x.max_pool2d(2)
x = self.dropout1(x)
x = self.fc1(x)
x = x.relu()
x = self.dropout2(x)
x = self.fc2(x)
return x
model = CNN().encrypt()
x = images_enc[0].unsqueeze(0)
print(x.shape)
model(x)
model = train_model(model, images_enc[:10, ], labels_enc[:10,], epochs=3)
prediction = model(images_enc[3].unsqueeze(0)).argmax()
# # Importing PyTorch Models
# +
import torch.nn as nn
import torch.nn.functional as F
class PyTorchModel(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(1, 32, 3, 1)
self.conv2 = nn.Conv2d(32, 64, 3, 1)
self.dropout1 = nn.Dropout2d(0.25)
self.dropout2 = nn.Dropout2d(0.5)
self.fc1 = nn.Linear(9216, 128)
self.fc2 = nn.Linear(128, 10)
def forward(self, x):
x = x.unsqueeze(1)
x = self.conv1(x)
x = F.relu(x)
x = self.conv2(x)
x = F.relu(x)
x = F.max_pool2d(x, 2)
x = self.dropout1(x)
x = torch.flatten(x, 1)
x = self.fc1(x)
x = F.relu(x)
x = self.dropout2(x)
x = self.fc2(x)
output = F.log_softmax(x, dim=1)
return output
# -
pytorch_model = PyTorchModel()
dummy_input = torch.empty(images.shape)
crypten_model = crypten.nn.from_pytorch(pytorch_model, dummy_input)
crypten_model.encrypt()
prediction = crypten_model(images_enc[3].unsqueeze(0))
print(prediction)
print(prediction.get_plain_text())
prediction.get_plain_text().argmax()
crypten_model = train_model(crypten_model, images_enc[:10, ], labels_enc[:10,], epochs=3)
| pycon-workshop-2020/2 - Training a Model on Encrypted Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import animation, rc
from IPython.display import HTML
# +
# First set up the figure, the axis, and the plot element we want to animate
fig, ax = plt.subplots()
ax.set_xlim(( 0, 2))
ax.set_ylim((-2, 2))
line, = ax.plot([], [], lw=2)
# -
# initialization function: plot the background of each frame
def init():
line.set_data([], [])
return (line,)
# animation function. This is called sequentially
def animate(i):
x = np.linspace(0, 2, 1000)
y = np.sin(2 * np.pi * (x - 0.01 * i))
line.set_data(x, y)
return (line,)
# call the animator. blit=True means only re-draw the parts that have changed.
anim = animation.FuncAnimation(fig, animate, init_func=init,
frames=100, interval=20, blit=True)
from IPython.display import HTML
# +
# import imageio
# imageio.plugins.ffmpeg.download()
# -
HTML(anim.to_html5_video())
| FAI_old/lesson2/animation_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ajdillhoff/Linear-Algebra-Examples/blob/main/compute_affine_transform.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="h-gpXkZKGxP_"
# # Computing Affine Transformations
#
# One important result of solving linear systems is to compute a transformation between two different sets of points.
#
# Given a set of points from the original coordinate system and another set of the same points transformed by $A$, how do we solve for that transformation matrix?
#
# We are looking for some $A$ such that
#
# $$
# A \mathbf{x} = \hat{\mathbf{x}}.
# $$
#
# In a perfect world, $\hat{\mathbf{x}}$ will match the target points $\mathbf{y}$. In other words,
#
# $$
# \|\hat{\mathbf{x}} - \mathbf{y}\|_2 = 0.
# $$
#
# **For this problem, we will assume this is the case.**
#
# For an affine transformation, we would have some matrix of the form
#
# $$
# A = \begin{bmatrix}
# a_{11} & a_{12} & a_{13}\\
# a_{21} & a_{22} & a_{23}
# \end{bmatrix}.
# $$
#
# Given a homogeneous 2D coordinate $\mathbf{x}$, we can compute each component $A\mathbf{x}$ as
#
# \begin{align*}
# \hat{x}_1 &= a_{11} * x_1 + a_{12} * x_2 + a_{13} * 1\\
# \hat{x}_2 &= a_{21} * x_1 + a_{22} * x_2 + a_{23} * 1\\
# \end{align*}
#
# We can fit this using a least squares approach by the following construction.
#
# \begin{equation*}
# \begin{bmatrix}
# x_1^{(1)} & x_2^{(1)} & 1 & 0 & 0 & 0\\
# 0 & 0 & 0 & x_1^{(1)} & x_2^{(1)} & 1\\
# && \vdots\\
# x_1^{(n)} & x_2^{(n)} & 1 & 0 & 0 & 0\\
# 0 & 0 & 0 & x_1^{(n)} & x_2^{(n)} & 1\\
# \end{bmatrix}
# \begin{bmatrix}
# a_{11}\\
# a_{12}\\
# a_{13}\\
# a_{21}\\
# a_{22}\\
# a_{23}\\
# \end{bmatrix}=
# \begin{bmatrix}
# \hat{x}_1^{(1)}\\
# \hat{x}_2^{(1)}\\
# \vdots\\
# \hat{x}_1^{(n)}\\
# \hat{x}_2^{(n)}\\
# \end{bmatrix}
# \end{equation*}
#
# We can solve this using the *normal equations*:
#
# $$
# A^T A \mathbf{x} = A^T \mathbf{b}.
# $$
#
# + id="e4maPg_sJ4Yy"
import numpy as np
import matplotlib.pyplot as plt
# + id="_DLoMbxyIylI"
def compute_affine_transform(src, dst):
"""Computes the affine transformation matrix `A` such that A @ src = dst."""
num_samples = src.shape[0]
# Convert to homogeneous coordinates
src_affine = np.concatenate((src, np.ones((num_samples, 1))), axis=1)
zero_buffer = np.zeros_like(src_affine)
r1 = np.concatenate((src_affine, zero_buffer), axis=1)
r2 = np.concatenate((zero_buffer, src_affine), axis=1)
# Construct our design matrix X by interleaving the two equations required
# for each point correspondence.
X = np.empty((r1.shape[0] + r2.shape[0], r1.shape[1]), dtype=r1.dtype)
X[0::2] = r1
X[1::2] = r2
# Flatten the target points into a vector (this is the b vector from above)
y = dst.ravel()
# TODO: Compute M by using the normal equations.
# Here, M corresponds to the solution vector x from above.
# Resahpe our solution vector (x in the equations above) into a 3x3
# affine transformation matrix.
M = np.reshape(M, (2, 3))
M = np.vstack((M, np.zeros((1, M.shape[1]))))
M[2, 2] = 1
return M
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="57x1uU78Jw56" outputId="2c57ee22-73b7-4339-dc0a-4bfe9d62620c"
# Randomly sample a pattern and then transform it by some random rotation.
num_points = 100
x = np.random.rand(num_points, 2) * 2 - 1
# Create a random rotation matrix
theta = np.random.uniform(0, np.pi)
R = np.array([
[np.cos(theta), -np.sin(theta), 0],
[np.sin(theta), np.cos(theta), 0],
[0, 0, 1]
])
y = R @ np.concatenate((x, np.ones((100, 1))), axis=1).T
y = y.T[:, :2]
# Call `compute_affine_transform` to estimate the rotation matrix
M = compute_affine_transform(x, y)
# Compute the norm between the true matrix and estimated matrix.
# This should be a very small number close to 0.
estimation_error = np.linalg.norm(R - M)
print(f"Error = {estimation_error}")
plt.scatter(x[:, 0], x[:, 1], c='b')
plt.scatter(y[:, 0], y[:, 1], c='r')
| compute_affine_transform.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SLSTR spatial plotting, quality control and data interrogation
# Version: 2.0
# Date: 10/04/2019
# Author: <NAME> and <NAME> (Plymouth Marine Laboratory)
# Credit: This code was developed for EUMETSAT under contracts for the Copernicus
# programme.
# License: This code is offered as free-to-use in the public domain, with no warranty.
#
# This routine has been designed to work with SLSTR L2 NRT data, which is available as
# tiles. It can be easily adapted for application to L1 data, but care should be taken
# if using it to analyse L2 NTC data, as this is delivered as a half orbit PDU.
# This routine shows examples of how to use python netcdf libraries to ingest Level 2 SLSTR data, mask it according to quality control variables, correct for bias, select only for dual view data and compare it against other, coincident geo-physical variables.
# In Python, we usually have two sections of code at the top that occur before we enter the main programme. These
# sections typically include:
# 1. importing modules
# 2. defining functions that are called in the main programme
# Firstly, we begin by importing all of the external modules that we will be using in this script; they are annotated with brief explanations of what they do.
# +
# import tools that allow us access tot system functions, e.g. get working directory, check path.
import os
import shutil
# import high level python system functions
import sys
import warnings
# import tools that let us manipulate dates and times
import datetime
# import tools that let us manipulate arrays (makes Python more like Matlab for matrix operations)
import numpy as np
# import tools that facilitate string pattern matching
import fnmatch
# import tools that let us create log files to write to
import logging
# import tools for netCFD4 manipulation
import xarray as xr
# import tools for plotting, making subplots, and utilising map projections
import matplotlib.pyplot as plt
from matplotlib import gridspec
import cartopy.crs as ccrs
import cartopy.feature as cfeature
# turn off warnings. Bad practice, but we don't want to see them here....
warnings.filterwarnings("ignore")
# -
# We will quickly get some map backgrounds for our later plots...
land_resolution = '50m'
land_poly = cfeature.NaturalEarthFeature('physical', 'land', land_resolution,
edgecolor='k',
facecolor=cfeature.COLORS['land'])
# The next section is defines two paths for where we read our input data from ("MYPATH"), and where we write our log files to (again, current working directory). To help to find your data, please complete the MYPATH variable below with the output generated by the /Configuration_Testing/Data_Path_Checker.ipynb Jupyter notebook in the Configuration_Testing folder.
# +
# e.g. MYPATH = os.path.join("C:/","Users","me","Desktop")
MYPATH = "<please insert your path from Data_Path_Checker.ipynb here, removing the quotes and chevrons>"
DEFAULT_ROOT_DIR = os.path.join(MYPATH,'SLSTR_test_data')
DEFAULT_LOG_PATH = os.getcwd()
# we will be looking for all files that match this pattern
DEFAULT_FILE_FILTER = '*SLSTR*.nc'
# -
# #----------------------------------------------------PRE-AMBLE COMPLETE-----------------------------------------------
#
# After all of that preparation, we are at the main entrance point for our code. We begin by defining our logfile so we can record any errors here if things go wrong, or any info/debug messages we may want along the way if we don't want them printed to screen (or to the console in interactive mode).
# +
#-main-------------------------------------------------------------------------
# preliminary stuff
logfile = os.path.join(DEFAULT_LOG_PATH,"SLSTR_test_plot_"+datetime.datetime.now().strftime('%Y%m%d_%H%M')+".log")
# we define a verbose flag to control how much info we want to see. It can also be useful to define a debug flag
# for even more information.
verbose=False
# this option will stop plotting to screen, and instead plot to file
no_show=True
if no_show:
plt.ioff()
# -
# We have defined a log file above, and here we set up how python will use it. Note that this is the first time we use the 'print' command. Print will output its contents to the screen, and here, this output will appear below the box when we run it.
# set file logger
try:
if os.path.exists(logfile):
os.remove(logfile)
print("logging to: "+logfile)
logging.basicConfig(filename=logfile,level=logging.DEBUG)
except:
print("Failed to set logger")
# So, lets proceed with loading some SLSTR data. The first thing we need to do is find the relevant netCDF files that contain the data that we want to plot. This next block of code collects the names of all netCDF files in our DEFAULT_ROOT_DIR path. We can make this more specific by adapting the DEFAULT_FILE_FILTER variable from "*.nc".
#
# We begin by setting up an empty "list" variables called nc_files, and append to this list as we proceed through a series of loops, defined by the "for" statements.
# -get the files-------------------------------------------------------------
nc_files=[]
for root, _, filenames in os.walk(DEFAULT_ROOT_DIR):
for filename in fnmatch.filter(filenames, DEFAULT_FILE_FILTER):
nc_files.append(os.path.join(root, filename))
if verbose:
print('Found: '+filename)
logging.info('Found: '+os.path.join(root, filename))
# Lets check what files we have by looping through the list...
#
# In python you can loop through the values in a list using "for item in list:". We only have 1 file in this cacse, though
for nc_file in nc_files:
print(nc_file)
# For now, we will just work with the first file we find, which is held in nc_files[0].
nc_file = nc_files[0]
# This next line opens our netCDF file. It does not read any data in yet, just makes the contents accessible. We should remember to close this link, especially if we are opening lots of files.
nc_fid = xr.open_dataset(nc_file)
# We start by loading our coordinate variables, using the function that we defined at the top of the script.
#
# note: python is very accepting of white space, but the next line would flag as a problem in a code-checker like pylint. It is spaced like this to make it easy to read.
LON = nc_fid.lon
LAT = nc_fid.lat
TIME = nc_fid.adi_dtime_from_sst
# And finally we load our data variables and close the netCDF file link. SST is stored as a three dimensional variable, with dimensions of time, lat and lon. The time dimension has length of one, and in order to plot SST as an image, we need to remove this "singleton" dimension. The numpy method "squeeze" will take care of this for us.
SST_raw = np.squeeze(nc_fid.sea_surface_temperature.data)
SST = np.squeeze(nc_fid.sea_surface_temperature.data)
SST_STD_DEV = np.squeeze(nc_fid.sses_standard_deviation.data)
SST_BIAS = np.squeeze(nc_fid.sses_bias.data)
SST_ALG_TYPE = np.squeeze(nc_fid.sst_algorithm_type.data)
WIND_SPEED = np.squeeze(nc_fid.wind_speed.data)
# Now we load the quality and masking variables
L2P_FLAGS = np.squeeze(nc_fid.l2p_flags.data)
QUALITY_LEVEL = np.squeeze(nc_fid.quality_level.data)
nc_fid.close()
# Now, lets set up our figure and make an initial plot of our SST data
fig = plt.figure(figsize=(6,6), dpi=150)
plt.imshow(SST)
if no_show:
fig.savefig('Plot1_Initial.png',bbox_inches='tight')
else:
plt.show()
# Plot successful, but this is ugly, not very helpful as it is not on any geographical map, incorrect as we have not taken abias into account, and has not been quality controlled! Lets take steps to improve this by:
# 1. reprojecting the data onto a map
# 2. make a contour plot against our LON and LAT data (we use contourf here as it is faster, but pcolor is more appropriate)
# 3. apply a more sensible colour bar for SST data
# 4. adding a colour bar
#
# Then:
# 1. Masking our data for specific features and qulity values
# 2. Correcting our the SST for bias and considering the standard deviation
# 3. Considering dual SST only.
#
# Lastly:
# 1. Checking associated variables; e.g. wind speed
#
# So, lets perform the steps in the first list. The following code block does exactly that; reprojects (using the basemap toolkit), makes a filled contour plot (using contourf), applies a colour scale (cmap) and adds a colour bar (plt.colorbar).
# +
fig = plt.figure(figsize=(10,10), dpi=300)
# set projection
m = plt.axes(projection=ccrs.PlateCarree(central_longitude=0.0))
# set my vertical plotting order and fontsize
zordcoast=0
fsz=12
SST_plot = SST.copy()
vmin=np.nanmean(SST_plot)-3*np.nanstd(SST_plot)
vmax=np.nanmean(SST_plot)+3*np.nanstd(SST_plot)
SST_plot[SST_plot<vmin] = np.nan
SST_plot[SST_plot>vmax] = np.nan
# plot the data
p1 = plt.pcolormesh(LON,LAT,SST_plot,cmap=plt.cm.jet,vmin=vmin,vmax=vmax)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', linewidth=1)
m.add_feature(land_poly)
g1 = m.gridlines(draw_labels = True)
g1.xlabels_top = False
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
cbar = plt.colorbar(p1, orientation='horizontal')
cbar.set_label('SST [K]',fontsize=fsz);
plt.title('SLSTR SST [K]', fontsize=fsz);
# -
if no_show:
fig.savefig('Plot2_SST.png',bbox_inches='tight')
# A definite improvement, our data is now accompanied by a scale, with units, and is reprojected so we can relate it to a map.
#
# However, we still have not interrogated our data.
#
# One of the most important things we need to do with SST data is consider the quality level - so lets start by doing that.
#
# The next code bloack will display the values of the quality level, stored in the quality level variables in the L2 WST product. Usually, we only consider the product viable where the quality flag is five, but can use quality level 4 in some circumstances.
fig = plt.figure(figsize=(10*int(np.nanmax(QUALITY_LEVEL))+1,10), dpi=150)
gs = gridspec.GridSpec(1, int(np.nanmax(QUALITY_LEVEL))+1)
contour_vals = np.arange(np.nanmin(QUALITY_LEVEL)-1,np.nanmax(QUALITY_LEVEL)+1,1)
gs.update(wspace=0.1, hspace=0.1)
# loop through each algorithm
for ii in np.arange(0,int(np.nanmax(QUALITY_LEVEL))+1):
m = plt.subplot(gs[0,ii], projection=ccrs.PlateCarree(central_longitude=0.0))
MASKED_QUALITY_LEVEL = QUALITY_LEVEL.astype('float')
MASKED_QUALITY_LEVEL[MASKED_QUALITY_LEVEL != float(ii)] = np.nan
# plot the data
plt.pcolormesh(LON,LAT,np.ma.masked_invalid(MASKED_QUALITY_LEVEL),vmin=0,vmax=5)
plt.text(33,-17.5,ii,fontweight='bold',fontsize=fsz*2)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', linewidth=1)
m.add_feature(land_poly)
g1 = m.gridlines(draw_labels = False)
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
if no_show:
fig.savefig('Plot3_Quality.png',bbox_inches='tight')
# So, lets mask out any data that have a quality value of 2 or lower...
SST[QUALITY_LEVEL<=4]=np.nan
# Next, the SST field has two associated measurements that we need to consider, the bias, and the standard deviation. So lets plot these...
# +
fig = plt.figure(figsize=(20,20), dpi=300)
gs = gridspec.GridSpec(1, 2)
fsz = 20
m = plt.subplot(gs[0,0], projection=ccrs.PlateCarree(central_longitude=0.0))
# plot the data
SST_plot = SST_BIAS.copy()
vmin=np.nanmean(SST_plot)-3*np.nanstd(SST_plot)
vmax=np.nanmean(SST_plot)+3*np.nanstd(SST_plot)
SST_plot[SST_plot<vmin] = np.nan
SST_plot[SST_plot>vmax] = np.nan
p1 = plt.pcolormesh(LON,LAT,SST_plot,cmap=plt.cm.jet,vmin=vmin,vmax=vmax)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', linewidth=1)
m.add_feature(land_poly)
g1 = m.gridlines(draw_labels = True)
g1.xlabels_top = False
g1.ylabels_right = False
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
cbar = plt.colorbar(p1, orientation='horizontal', pad=0.05)
cbar.set_label('SST bias [K]',fontsize=fsz)
m = plt.subplot(gs[0,1], projection=ccrs.PlateCarree(central_longitude=0.0))
# plot the data
SST_plot = SST_STD_DEV.copy()
vmin=np.nanmean(SST_plot)-3*np.nanstd(SST_plot)
vmax=np.nanmean(SST_plot)+3*np.nanstd(SST_plot)
SST_plot[SST_plot<vmin] = np.nan
SST_plot[SST_plot>vmax] = np.nan
p1 = plt.pcolormesh(LON,LAT,SST_plot,cmap=plt.cm.jet,vmin=vmin,vmax=vmax)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', linewidth=1)
m.add_feature(land_poly)
g1 = m.gridlines(draw_labels = True)
g1.xlabels_top = False
g1.ylabels_right = False
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
cbar = plt.colorbar(p1, orientation='horizontal', pad=0.05)
cbar.set_label('SST standard deviation [K]',fontsize=fsz);
# -
if no_show:
fig.savefig('Plot4_SSES.png',bbox_inches='tight')
# The SST value we are interested in needs to be corrected for the bias, so lets do that:
SST = SST + SST_BIAS
# We can see from the bias and standard deviation plots that there are some sharp lines across the image. SLSTR uses 5 algorithms to estimate SST. Some of these algorithms use the nadir view only, while some take advantage of the 'Dual View' capability of the sensor. Dual view takes two images of the surface, one at nadir and one at an oblique angle. This allows it to better characterise the effects of the atmosphere. In practice, the nadir view is wider than the dual view, which results in a stripe along the middle of the swath. We can check which algorithms were used to derive the SST estimate by checking the SST_ALG_TYPE variable, as below:
# +
fig = plt.figure(figsize=(10*int(np.nanmax(SST_ALG_TYPE))+1,10), dpi=150)
gs = gridspec.GridSpec(1, 6)
gs.update(wspace=0.1, hspace=0.1)
fsz = 12
contour_vals = np.arange(np.nanmin(SST_ALG_TYPE)-1,np.nanmin(SST_ALG_TYPE)+1,1)
# loop through each algorithm
for ii in np.arange(0,int(np.nanmax(SST_ALG_TYPE))+1):
m = plt.subplot(gs[0,ii], projection=ccrs.PlateCarree(central_longitude=0.0))
MASKED_ALG_TYPE = SST_ALG_TYPE.astype('float')
MASKED_ALG_TYPE[MASKED_ALG_TYPE != float(ii)] = np.nan
# plot the data
plt.pcolormesh(LON,LAT,np.ma.masked_invalid(MASKED_ALG_TYPE),cmap=plt.cm.jet,vmin=0,vmax=5)
if ii == 0:
plt.text(33,-17.5,'No retrieval',fontweight='bold',fontsize=fsz*2)
if ii == 1:
plt.text(33,-17.5,'N2',fontweight='bold',fontsize=fsz*2)
if ii == 2:
plt.text(33,-17.5,'N3R',fontweight='bold',fontsize=fsz*2)
if ii == 3:
plt.text(33,-17.5,'N3',fontweight='bold',fontsize=fsz*2)
elif ii == 4:
plt.text(33,-17.5,'D2',fontweight='bold',fontsize=fsz*2)
elif ii == 5:
plt.text(33,-17.5,'D3',fontweight='bold',fontsize=fsz*2)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', linewidth=1)
m.add_feature(land_poly)
g1 = m.gridlines(draw_labels = False)
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
# -
if no_show:
fig.savefig('Plot5_Algorithms.png',bbox_inches='tight')
# We should remember that, just because a measurement is Nadir view only, it does not mean that it is bad! Sometimes the nadir view is the best to use. Here, though, lets finally plot our nadir+dual and our dual view data, corrected for bias, and masked for a quality level of 3 or greater. We will overlay the plot with contours from contemporaneous ECMWF wind data, that is included in with SLSTR L2 WAT products, as part of the GHRSST specification.
# +
# %%capture
SST_C = SST-273.15
fig = plt.figure(figsize=(20,20), dpi=150)
gs = gridspec.GridSpec(3, 1, height_ratios=[20,0.5,1])
gs.update(wspace=0.01, hspace=0.01)
# set my vertical plotting order and fontsize
zordcoast=0
fsz=20
# plot the data
SST_plot = SST_C.copy()
vmin=int(np.nanmean(SST_plot)-3*np.nanstd(SST_plot))-1
vmax=int(np.nanmean(SST_plot)+3*np.nanstd(SST_plot))+1
SST_plot[SST_plot<vmin] = np.nan
SST_plot[SST_plot>vmax] = np.nan
m = plt.subplot(gs[0,0], projection=ccrs.PlateCarree(central_longitude=0.0))
p1 = plt.contourf(LON,LAT,SST_plot,100,cmap=plt.cm.jet,vmin=vmin,vmax=vmax,zorder=-1)
CS = plt.contour(LON,LAT,WIND_SPEED,10,linewidths=1.0,cmap=plt.get_cmap('Greys'),zorder=0)
plt.clabel(CS, fontsize=10, inline=1)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', linewidth=1)
m.add_feature(land_poly)
g1 = m.gridlines(draw_labels = True)
g1.xlabels_top = False
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
# add colorbar
axes0 = plt.subplot(gs[2,0])
cbar = plt.colorbar(p1, cax=axes0, orientation='horizontal')
cbar.ax.tick_params(labelsize=fsz)
cbar.set_label('Bias corrected, quality controlled, whole view SST [$^{o}$C]',fontsize=fsz)
# -
bbox_inches='tight'
fig.savefig('Plot6_SLSTR_All_SST.png',bbox_inches='tight')
# +
# %%capture
SST_C = SST-273.15
SST_C[SST_ALG_TYPE<4] = np.nan
fig = plt.figure(figsize=(20,20), dpi=150)
gs = gridspec.GridSpec(3, 1, height_ratios=[20,1,1])
gs.update(wspace=0.01, hspace=0.01)
# set my vertical plotting order and fontsize
zordcoast=0
fsz=20
# plot the data
SST_plot = SST_C.copy()
vmin=int(np.nanmean(SST_plot)-3*np.nanstd(SST_plot))-1
vmax=int(np.nanmean(SST_plot)+3*np.nanstd(SST_plot))+1
SST_plot[SST_plot<vmin] = np.nan
SST_plot[SST_plot>vmax] = np.nan
m = plt.subplot(gs[0,0], projection=ccrs.PlateCarree(central_longitude=0.0))
p1 = m.contourf(LON,LAT,SST_plot,100,cmap=plt.cm.jet,vmin=vmin,vmax=vmax,zorder=-1)
CS = m.contour(LON,LAT,WIND_SPEED,10,linewidths=1.0,cmap=plt.get_cmap('Greys'),zorder=0)
plt.clabel(CS, fontsize=14, inline=1)
# add embelishments
m.coastlines(resolution=land_resolution, color='black', zorder=3)
m.add_feature(land_poly, zorder = 2)
g1 = m.gridlines(draw_labels = True)
g1.xlabels_top = False
g1.xlabel_style = {'size': 16, 'color': 'gray'}
g1.ylabel_style = {'size': 16, 'color': 'gray'}
# add colorbar
axes0 = plt.subplot(gs[2,0])
cbar = plt.colorbar(p1, cax=axes0, orientation='horizontal')
cbar.ax.tick_params(labelsize=fsz)
cbar.set_label('Bias corrected, quality controlled, dual view SST [$^{o}$C]',fontsize=fsz)
# -
bbox_inches='tight'
fig.savefig('Plot7_SLSTR_Dual_SST_demo.png',bbox_inches='tight')
# <br> <a href="./16_OLCI_spectral_AC_L1_L2_comparison.ipynbb"><< 16 - OLCI - atmospheric correction effects: Level-1B to Level-2 spectral comparison</a><span style="float:right;"><a href="../STM/31_SRAL_SLAs.ipynb">31 - Compute corrected sea surface heights & anomalies from Sentinel-3 >></a> <hr> <p style="text-align:left;">This project is licensed under the <a href="/edit/LICENSE">MIT License</a> <span style="float:right;"><a href="https://gitlab.eumetsat.int/eo-lab-usc-open/ocean">View on GitLab</a> | <a href="https://training.eumetsat.int/">EUMETSAT Training</a> | <a href=mailto:<EMAIL>>Contact</a></span></p>
| SLSTR/21_SLSTR_spatial_interrogation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## The great 1 dimensional bandstructure function
# **Autors:** *<NAME>* and *<NAME>* with a vital teoretical support of *<NAME>*, *<NAME>* and *Hofmanns* book *Solid State Physics*
#
# **Year:** *2019*
#
# * If you are looking at this file from the universities computer which is used for the lab, then please leave everything exactly the way it was before.
#
# * The file is structured to be able to run cell by cell. If any of the variable is unclear, it is possible to create a new cell by pressing *Esc* and then *A* and play around. The shortcut to delete cell: *Esc* and then *D+D*.
#
# * In the following document, the one dimensional bandstructure of a crystal is solved using the Plane wave basis approximation (further described in the lab manual, here only a summary of presented, stating main variables).
#
# * In this case the atomic potential is approximated by the Gaussian potential $V(x)=-U\sum_n e^{-(x-na)^2/2\sigma^2}$. This potential can be written in the Fourier components as $V(x)=\sum_{G_j}-U\sqrt {2\pi}\frac{\sigma}{a}e^{-G^2_j \sigma^2/2} e^{i G_j x} \equiv \sum_{G_j} V_{G_j} \cdot e^{i G_j x}$,
# where $G_j$ is a reciprocal lattice vector.
#
# * The ampliture of $V_G$, we denote as $A$, that is $A \equiv -U\sqrt {2\pi}\frac{\sigma}{a}$ and $V_G = A e^{-G^2 \sigma^2/2} $.
#
# * In order to obtain the energy spectrum or the band structure we need to solve the following eigenvalue equation:
# $\Big( \frac{\hbar^2 (k+K)^2}{2m} - \varepsilon \Big) c_{k+K} +\sum_G c_{k+K-G} V_G$ and search for $\varepsilon$ - the energy eigenvalues.
#
# ### Necessary imports
# +
# Usual imports
import matplotlib.pyplot as plt
# %matplotlib notebook
import numpy as np
# pretty print all cell's output and not just the last one
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# Widgets imports
import ipywidgets as widgets
from ipywidgets import interact, interactive
from IPython.display import display
from ipywidgets import HBox, Label
from ipywidgets import IntSlider
# -
# There are many variables involving *k* but it is important that you note the differences;
# * *K* the reciprocal lattice vector and *k* the wave vector $k\in [-\pi/a; \pi/a]$
# * The number of *k* vectors in our lattice is *Len_k*
# * *MaxK* is the maximal reciprocal lattice vector we consider to include in the sum ($K$ or $K-G$) thus the summation goes from *-MaxK* to *MaxK*.
# ### Definition of initial values
a = 3 # Lattice constant in Ångstrongs
sigma = 0.5 # Very arbitrary value
U = 11 # eV
maxK = 6
Len_k = 15 # How fine to calculate the band structure
NrOfK = maxK*2+1
A = -U*np.sqrt(2 * np.pi)*sigma/a
# *BandStructureFunction.ipynb* file contains a function *BandStructF* which creates the matrix for the eigenvalue equation described above and solves for the eigenvalues - energies.
# *%run* is one of the so-called magic functions and does the import of the function found in the file *BandStructureFunction.ipynb*.
# %run BandStructureFunction.ipynb
# Solve eigenvalue equation for each wave number *k*
# * Put k vector into right magnitude. Before *k* is some integer $k =.. -2,-1,0,1..$. Save the obtained value in *kVect*
# * Save calculated energies in *Energies*
# * Uncomment "%%time" to measure the calculation time of the cell
# # %%time
Energies = np.zeros(shape=(Len_k*2+1, NrOfK))
# Vector for plotting containing all of the k values in 1/Angstrom
kVect = [0]*(Len_k*2+1)
for ki in range(-Len_k, Len_k+1): # k vector for which we are calculating
k = np.pi/(a)*ki/Len_k
# %run BandStructureFunction.ipynb
E = BandStructF()
Energies[ki+Len_k] = np.real(E)
kVect[ki+Len_k] = k # in 1/Angstrom
# ## Plots
# ### Plot potential
#
# Plots are hidden so that they would not appear when running the interface. To make the plots visible either comment out the lines containg *%%capture* or comment out the cell containing *fig*.
# +
# %%capture
from matplotlib import pyplot as plt
# If you want to display the figure here, then uncomment previous line
# %matplotlib notebook
# %matplotlib notebook
fig, [ax, ax2] = plt.subplots(nrows=1, ncols=2) # Two plots in the same figure
fig.set_size_inches(9.5, 3.5)
fig.subplots_adjust(wspace=0.4, bottom=0.2) # Margins around the subplots
x = np.linspace(-5*a, 5*a, 401)
# length of the x vector should be an odd number so that 0 is included,
# otherwise it does not plot the peak correctly
Vx = -U*np.exp(-x**2/(2*sigma**2))
VxSum = [0]*len(x)
for n in range(-5, 6):
Vx1 = -U*np.exp(-(x-n*a)**2/(2*sigma**2))
ax.plot(x, Vx1, 'y--')
VxSum = VxSum+Vx1
ax.plot(x, VxSum)
ax.plot(x, Vx)
ax.set(xlabel='x, ($\AA$)', ylabel='V (eV)', title='Atomic potential')
# -
# ### Plot energies
# %%capture
ax2.plot(kVect, Energies[:, 0:5], color='purple')
ax2.set(xlabel='k ($1/ \AA$)', ylabel='E (eV)', title='Band structure')
# +
# fig
# -
# ### Creating an interactive user interface
#
# Function *f* repeats the steps above whenever some of the widgets are going to be used. Function *interactive* creates these widgets and connects them to the function *f*.
# +
def f(Lattice_Constant=a, Amplitude=U, NumKVec=maxK, PlotK=5):
# So that BandStructureFunction.ipynb knows the variables used in the function f
global a, A, k, Len_k, maxK, U, NrOfK
a = Lattice_Constant
U = Amplitude
maxK = NumKVec
A = -U*np.sqrt(2 * np.pi)*sigma/a # np.sqrt(2*sigma)
NrOfK = maxK*2+1
Energies = np.zeros(shape=(Len_k*2+1, NrOfK))
kVect = [0]*(Len_k*2+1)
for ki in range(-Len_k, Len_k+1):
k = np.pi/(a)*ki/Len_k
E = BandStructF()
# %run BandStructureFunction.ipynb
Energies[ki+Len_k] = np.real(E)
kVect[ki+Len_k] = k
ax.cla()
ax.set(xlabel='x, ($\AA$)', ylabel='V (eV)', title='Atomic potential')
x = np.linspace(-5*a, 5*a, 401)
Vx = -U*np.exp(-x**2/(2*sigma**2))
VxSum = [0]*len(x)
for n in range(-5, 6):
Vx1 = -U*np.exp(-(x-n*a)**2/(2*sigma**2))
ax.plot(x, Vx1, 'y--')
VxSum = VxSum+Vx1
ax.plot(x, VxSum)
ax.plot(x, Vx)
ax2.cla()
ax2.set(xlabel='k ($1/ \AA$)', ylabel='E (eV)', title='Band structure')
ax2.plot(kVect, Energies[:, 0:PlotK], color='purple')
# return Energies
The_Interaction = interactive(f, Lattice_Constant=(1, 10, 0.5), Amplitude=(
0, 100, 1), NumKVec=(1, 20, 1), PlotK=(0, 10, 1))
# -
# ### Creating boxes for widgets that are to be displayed
# +
for widg in The_Interaction.children[:-1]:
widg.description = ""
widg.continuous_update = False
Lattice_Const, Potential_Amp, Num_Of_K, PlotK = [
The_Interaction.children[i] for i in range(4)]
# display(Lattice_Const,Potential_Amp ,Num_Of_K ) #by uncommenting the widgets will be displayed here
FirstBox = widgets.HBox([Label(r'Lattice constant (Å)'), Lattice_Const, Label(
r'Potential depth, (eV)'), Potential_Amp, ])
SecondBox = widgets.HBox(
[Label(r'N of K vectors'), Num_Of_K, Label(r'How many bands to plot'), PlotK])
# -
# The number of calculated bands depends on *maxK* as $2\cdot maxK + 1$. So we set the maximal possible band to be plotted accordingly.
# +
def PlotKMax(*args):
PlotK.max = 2*Num_Of_K.value+1
Num_Of_K.observe(PlotKMax, 'value')
| 1D/User_interface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
matrix1 = [[5, 1, 6], [7, 3, 6], [9, 0, 4]]
matrix2 = [[6, 3, 7], [4, 1, 7], [9, 5, 3]]
matrix3 = [[5, 3, 7], [8, 4, 3], [7, 4, 7]]
matrix1 = np.matrix(matrix1)
matrix2 = np.matrix(matrix2)
matrix3 = np.matrix(matrix3)
print(matrix1)
print(matrix2)
matrix_t = matrix1.transpose()
print(matrix_t)
matrix_double_t = matrix_t.transpose()
print(matrix_double_t == matrix1)
matrix_sum = matrix1 + matrix2
sum_t = matrix_sum.transpose()
matrix_sum_t = matrix1.transpose() + matrix2.transpose()
print( sum_t == matrix_sum_t)
matrix_proiz = matrix2 * matrix1
proiz_t = matrix_sum.transpose()
matrix_proiz_t = matrix1.transpose() * matrix2.transpose()
print( sum_t == matrix_sum_t)
det = np.linalg.det(matrix1)
det_t = np.linalg.det(matrix1.transpose())
print(det)
print(det_t)
print(matrix1)
print(matrix1 * 5)
print(matrix1)
print(matrix1 * 0)
s = 5 + 3 + 4
print( matrix1 * s)
print( matrix1 * 5 + matrix1 * 3 + matrix1 * 4)
p = 5 * 6
print(matrix1 * p)
print(matrix1 * 6 * 5)
print((matrix1 + matrix2) * 3)
print(matrix1 * 3 + matrix2 * 3)
print(matrix1 + matrix2)
print(matrix2 + matrix1)
print(matrix1 + (matrix2 + matrix3))
print((matrix1 + matrix2) + matrix3)
print(matrix1 + (-1) * matrix1)
print(matrix1 * (matrix2 * matrix3))
print((matrix1 * matrix2) * matrix3)
print(matrix1 * (matrix2 + matrix3))
print(matrix1 * matrix2 + matrix1 * matrix3)
print(matrix1 * matrix2)
print(matrix2 * matrix1)
e = np.matrix([[1, 0, 0], [0, 1, 0], [0, 0, 1]])
print(e * matrix1 == matrix1 * e)
print(e * matrix1 == matrix1)
print(e * matrix1)
z = np.matrix([[0, 0, 0], [0, 0, 0], [0, 0, 0]])
print(z * matrix1 == matrix1 * z)
print(z * matrix1 == z)
print(z * matrix1)
print(np.linalg.det(matrix1))
print(np.linalg.det(matrix1.transpose()))
A = np.matrix([[6, 3, 5], [0, 0, 0], [7, 3, 6]])
print(np.linalg.det(A))
a = [[5, 3, 7], [8, 4, 3], [7, 4, 7]]
a = np.matrix(a)
b = [[5, 3, 7], [7, 4, 7], [8, 4, 3]]
b = np.matrix(b)
print(np.linalg.det(a))
print(np.linalg.det(b))
b = [[5, 3, 7], [7, 4, 7], [7, 4, 7]]
b = np.matrix(b)
print(np.linalg.det(b))
b = [[5, 3, 7], [7, 4, 7], [8, 4, 3]]
b = np.matrix(b)
print(np.linalg.det(b))
b[0] *= 5
print(np.linalg.det(b))
| MyExamples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
from fastai.basics import *
# # Rossmann
# ## Data preparation / Feature engineering
# In addition to the provided data, we will be using external datasets put together by participants in the Kaggle competition. You can download all of them [here](http://files.fast.ai/part2/lesson14/rossmann.tgz). Then you shold untar them in the dirctory to which `PATH` is pointing below.
#
# For completeness, the implementation used to put them together is included below.
PATH=Path('data/rossmann/')
table_names = ['train', 'store', 'store_states', 'state_names', 'googletrend', 'weather', 'test']
tables = [pd.read_csv(PATH/f'{fname}.csv', low_memory=False) for fname in table_names]
train, store, store_states, state_names, googletrend, weather, test = tables
len(train),len(test)
# We turn state Holidays to booleans, to make them more convenient for modeling. We can do calculations on pandas fields using notation very similar (often identical) to numpy.
train.StateHoliday = train.StateHoliday!='0'
test.StateHoliday = test.StateHoliday!='0'
# `join_df` is a function for joining tables on specific fields. By default, we'll be doing a left outer join of `right` on the `left` argument using the given fields for each table.
#
# Pandas does joins using the `merge` method. The `suffixes` argument describes the naming convention for duplicate fields. We've elected to leave the duplicate field names on the left untouched, and append a "\_y" to those on the right.
def join_df(left, right, left_on, right_on=None, suffix='_y'):
if right_on is None: right_on = left_on
return left.merge(right, how='left', left_on=left_on, right_on=right_on,
suffixes=("", suffix))
# Join weather/state names.
weather = join_df(weather, state_names, "file", "StateName")
# In pandas you can add new columns to a dataframe by simply defining it. We'll do this for googletrends by extracting dates and state names from the given data and adding those columns.
#
# We're also going to replace all instances of state name 'NI' to match the usage in the rest of the data: 'HB,NI'. This is a good opportunity to highlight pandas indexing. We can use `.loc[rows, cols]` to select a list of rows and a list of columns from the dataframe. In this case, we're selecting rows w/ statename 'NI' by using a boolean list `googletrend.State=='NI'` and selecting "State".
googletrend['Date'] = googletrend.week.str.split(' - ', expand=True)[0]
googletrend['State'] = googletrend.file.str.split('_', expand=True)[2]
googletrend.loc[googletrend.State=='NI', "State"] = 'HB,NI'
# The following extracts particular date fields from a complete datetime for the purpose of constructing categoricals.
#
# You should *always* consider this feature extraction step when working with date-time. Without expanding your date-time into these additional fields, you can't capture any trend/cyclical behavior as a function of time at any of these granularities. We'll add to every table with a date field.
def add_datepart(df, fldname, drop=True, time=False):
"Helper function that adds columns relevant to a date."
fld = df[fldname]
fld_dtype = fld.dtype
if isinstance(fld_dtype, pd.core.dtypes.dtypes.DatetimeTZDtype):
fld_dtype = np.datetime64
if not np.issubdtype(fld_dtype, np.datetime64):
df[fldname] = fld = pd.to_datetime(fld, infer_datetime_format=True)
targ_pre = re.sub('[Dd]ate$', '', fldname)
attr = ['Year', 'Month', 'Week', 'Day', 'Dayofweek', 'Dayofyear',
'Is_month_end', 'Is_month_start', 'Is_quarter_end', 'Is_quarter_start', 'Is_year_end', 'Is_year_start']
if time: attr = attr + ['Hour', 'Minute', 'Second']
for n in attr: df[targ_pre + n] = getattr(fld.dt, n.lower())
df[targ_pre + 'Elapsed'] = fld.astype(np.int64) // 10 ** 9
if drop: df.drop(fldname, axis=1, inplace=True)
add_datepart(weather, "Date", drop=False)
add_datepart(googletrend, "Date", drop=False)
add_datepart(train, "Date", drop=False)
add_datepart(test, "Date", drop=False)
# The Google trends data has a special category for the whole of the Germany - we'll pull that out so we can use it explicitly.
trend_de = googletrend[googletrend.file == 'Rossmann_DE']
# Now we can outer join all of our data into a single dataframe. Recall that in outer joins everytime a value in the joining field on the left table does not have a corresponding value on the right table, the corresponding row in the new table has Null values for all right table fields. One way to check that all records are consistent and complete is to check for Null values post-join, as we do here.
#
# *Aside*: Why note just do an inner join?
# If you are assuming that all records are complete and match on the field you desire, an inner join will do the same thing as an outer join. However, in the event you are wrong or a mistake is made, an outer join followed by a null-check will catch it. (Comparing before/after # of rows for inner join is equivalent, but requires keeping track of before/after row #'s. Outer join is easier.)
store = join_df(store, store_states, "Store")
len(store[store.State.isnull()])
joined = join_df(train, store, "Store")
joined_test = join_df(test, store, "Store")
len(joined[joined.StoreType.isnull()]),len(joined_test[joined_test.StoreType.isnull()])
joined = join_df(joined, googletrend, ["State","Year", "Week"])
joined_test = join_df(joined_test, googletrend, ["State","Year", "Week"])
len(joined[joined.trend.isnull()]),len(joined_test[joined_test.trend.isnull()])
joined = joined.merge(trend_de, 'left', ["Year", "Week"], suffixes=('', '_DE'))
joined_test = joined_test.merge(trend_de, 'left', ["Year", "Week"], suffixes=('', '_DE'))
len(joined[joined.trend_DE.isnull()]),len(joined_test[joined_test.trend_DE.isnull()])
joined = join_df(joined, weather, ["State","Date"])
joined_test = join_df(joined_test, weather, ["State","Date"])
len(joined[joined.Mean_TemperatureC.isnull()]),len(joined_test[joined_test.Mean_TemperatureC.isnull()])
for df in (joined, joined_test):
for c in df.columns:
if c.endswith('_y'):
if c in df.columns: df.drop(c, inplace=True, axis=1)
# Next we'll fill in missing values to avoid complications with `NA`'s. `NA` (not available) is how Pandas indicates missing values; many models have problems when missing values are present, so it's always important to think about how to deal with them. In these cases, we are picking an arbitrary *signal value* that doesn't otherwise appear in the data.
for df in (joined,joined_test):
df['CompetitionOpenSinceYear'] = df.CompetitionOpenSinceYear.fillna(1900).astype(np.int32)
df['CompetitionOpenSinceMonth'] = df.CompetitionOpenSinceMonth.fillna(1).astype(np.int32)
df['Promo2SinceYear'] = df.Promo2SinceYear.fillna(1900).astype(np.int32)
df['Promo2SinceWeek'] = df.Promo2SinceWeek.fillna(1).astype(np.int32)
# Next we'll extract features "CompetitionOpenSince" and "CompetitionDaysOpen". Note the use of `apply()` in mapping a function across dataframe values.
for df in (joined,joined_test):
df["CompetitionOpenSince"] = pd.to_datetime(dict(year=df.CompetitionOpenSinceYear,
month=df.CompetitionOpenSinceMonth, day=15))
df["CompetitionDaysOpen"] = df.Date.subtract(df.CompetitionOpenSince).dt.days
# We'll replace some erroneous / outlying data.
for df in (joined,joined_test):
df.loc[df.CompetitionDaysOpen<0, "CompetitionDaysOpen"] = 0
df.loc[df.CompetitionOpenSinceYear<1990, "CompetitionDaysOpen"] = 0
# We add "CompetitionMonthsOpen" field, limiting the maximum to 2 years to limit number of unique categories.
for df in (joined,joined_test):
df["CompetitionMonthsOpen"] = df["CompetitionDaysOpen"]//30
df.loc[df.CompetitionMonthsOpen>24, "CompetitionMonthsOpen"] = 24
joined.CompetitionMonthsOpen.unique()
# Same process for Promo dates. You may need to install the `isoweek` package first.
# +
# If needed, uncomment:
# # ! pip install isoweek
# -
from isoweek import Week
for df in (joined,joined_test):
df["Promo2Since"] = pd.to_datetime(df.apply(lambda x: Week(
x.Promo2SinceYear, x.Promo2SinceWeek).monday(), axis=1).astype(pd.datetime))
df["Promo2Days"] = df.Date.subtract(df["Promo2Since"]).dt.days
for df in (joined,joined_test):
df.loc[df.Promo2Days<0, "Promo2Days"] = 0
df.loc[df.Promo2SinceYear<1990, "Promo2Days"] = 0
df["Promo2Weeks"] = df["Promo2Days"]//7
df.loc[df.Promo2Weeks<0, "Promo2Weeks"] = 0
df.loc[df.Promo2Weeks>25, "Promo2Weeks"] = 25
df.Promo2Weeks.unique()
joined.to_pickle(PATH/'joined')
joined_test.to_pickle(PATH/'joined_test')
# ## Durations
# It is common when working with time series data to extract data that explains relationships across rows as opposed to columns, e.g.:
# * Running averages
# * Time until next event
# * Time since last event
#
# This is often difficult to do with most table manipulation frameworks, since they are designed to work with relationships across columns. As such, we've created a class to handle this type of data.
#
# We'll define a function `get_elapsed` for cumulative counting across a sorted dataframe. Given a particular field `fld` to monitor, this function will start tracking time since the last occurrence of that field. When the field is seen again, the counter is set to zero.
#
# Upon initialization, this will result in datetime na's until the field is encountered. This is reset every time a new store is seen. We'll see how to use this shortly.
def get_elapsed(fld, pre):
day1 = np.timedelta64(1, 'D')
last_date = np.datetime64()
last_store = 0
res = []
for s,v,d in zip(df.Store.values,df[fld].values, df.Date.values):
if s != last_store:
last_date = np.datetime64()
last_store = s
if v: last_date = d
res.append(((d-last_date).astype('timedelta64[D]') / day1))
df[pre+fld] = res
# We'll be applying this to a subset of columns:
columns = ["Date", "Store", "Promo", "StateHoliday", "SchoolHoliday"]
#df = train[columns]
df = train[columns].append(test[columns])
# Let's walk through an example.
#
# Say we're looking at School Holiday. We'll first sort by Store, then Date, and then call `add_elapsed('SchoolHoliday', 'After')`:
# This will apply to each row with School Holiday:
# * A applied to every row of the dataframe in order of store and date
# * Will add to the dataframe the days since seeing a School Holiday
# * If we sort in the other direction, this will count the days until another holiday.
fld = 'SchoolHoliday'
df = df.sort_values(['Store', 'Date'])
get_elapsed(fld, 'After')
df = df.sort_values(['Store', 'Date'], ascending=[True, False])
get_elapsed(fld, 'Before')
# We'll do this for two more fields.
fld = 'StateHoliday'
df = df.sort_values(['Store', 'Date'])
get_elapsed(fld, 'After')
df = df.sort_values(['Store', 'Date'], ascending=[True, False])
get_elapsed(fld, 'Before')
fld = 'Promo'
df = df.sort_values(['Store', 'Date'])
get_elapsed(fld, 'After')
df = df.sort_values(['Store', 'Date'], ascending=[True, False])
get_elapsed(fld, 'Before')
# We're going to set the active index to Date.
df = df.set_index("Date")
# Then set null values from elapsed field calculations to 0.
columns = ['SchoolHoliday', 'StateHoliday', 'Promo']
for o in ['Before', 'After']:
for p in columns:
a = o+p
df[a] = df[a].fillna(0).astype(int)
# Next we'll demonstrate window functions in pandas to calculate rolling quantities.
#
# Here we're sorting by date (`sort_index()`) and counting the number of events of interest (`sum()`) defined in `columns` in the following week (`rolling()`), grouped by Store (`groupby()`). We do the same in the opposite direction.
bwd = df[['Store']+columns].sort_index().groupby("Store").rolling(7, min_periods=1).sum()
fwd = df[['Store']+columns].sort_index(ascending=False
).groupby("Store").rolling(7, min_periods=1).sum()
# Next we want to drop the Store indices grouped together in the window function.
#
# Often in pandas, there is an option to do this in place. This is time and memory efficient when working with large datasets.
bwd.drop('Store',1,inplace=True)
bwd.reset_index(inplace=True)
fwd.drop('Store',1,inplace=True)
fwd.reset_index(inplace=True)
df.reset_index(inplace=True)
# Now we'll merge these values onto the df.
df = df.merge(bwd, 'left', ['Date', 'Store'], suffixes=['', '_bw'])
df = df.merge(fwd, 'left', ['Date', 'Store'], suffixes=['', '_fw'])
df.drop(columns,1,inplace=True)
df.head()
# It's usually a good idea to back up large tables of extracted / wrangled features before you join them onto another one, that way you can go back to it easily if you need to make changes to it.
df.to_pickle(PATH/'df')
df["Date"] = pd.to_datetime(df.Date)
df.columns
joined = pd.read_pickle(PATH/'joined')
joined_test = pd.read_pickle(PATH/f'joined_test')
joined = join_df(joined, df, ['Store', 'Date'])
joined_test = join_df(joined_test, df, ['Store', 'Date'])
# The authors also removed all instances where the store had zero sale / was closed. We speculate that this may have cost them a higher standing in the competition. One reason this may be the case is that a little exploratory data analysis reveals that there are often periods where stores are closed, typically for refurbishment. Before and after these periods, there are naturally spikes in sales that one might expect. By ommitting this data from their training, the authors gave up the ability to leverage information about these periods to predict this otherwise volatile behavior.
joined = joined[joined.Sales!=0]
# We'll back this up as well.
joined.reset_index(inplace=True)
joined_test.reset_index(inplace=True)
joined.to_pickle(PATH/'train_clean')
joined_test.to_pickle(PATH/'test_clean')
| nbs/dl1/rossman_data_clean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import re
#laad data
with open('input.txt') as t:
text = t.read()
# **Deel 1**
# maak een nieuw bestand met per regel opeenvogende waarden en een enter bij een witregel
new = ''
for line in text.split('\n'):
# lege regel: voeg een enter toe
if line == '':
new += '\n'
# voeg een spatie aan het einde toe voor het plakken van waarden die van verschillende regels komen
else:
new += line + ' '
# maak een lijst, een item is een paspoort (string)
textlist = list(new.split('\n'))
lwb = []
for line in textlist:
items = line.split()
# maak een dictonary per paspoort
wb = {}
for item in items:
key, val = item.split(':')
wb[key] = val
#maak een lijst van deze dictonaries
lwb.append(wb.copy())
# maak een dataframe met de categorieen als kolommen en de paspoorten als rijen
df = pd.DataFrame(lwb)
# verwijder de optionele 'cid' kolom en verwijder de rijen met missende waarden in de overige kolommen
df_valid = df.drop("cid", 1).dropna()
# tel het aantal geldige paspoorten
print(f'Aantal geldige paspoorten: {len(df_valid.index)}')
# **Deel 2**
# defineer functie voor de jaartal condities
df_check = df_valid.copy()
def test_rules(column, length, minv, maxv):
li = []
for item in column:
rules = [len(item) == length, pd.to_numeric(item) >= minv, pd.to_numeric(item) <= maxv]
if all(rules):
li.append(True)
else:
li.append(False)
return(li)
# +
# byr (Birth Year) - four digits; at least 1920 and at most 2002.
# iyr (Issue Year) - four digits; at least 2010 and at most 2020.
# eyr (Expiration Year) - four digits; at least 2020 and at most 2030.
df_check['byr'] = test_rules(df_valid['byr'], 4, 1920, 2002)
df_check['iyr'] = test_rules(df_valid['iyr'], 4, 2010, 2020)
df_check['eyr'] = test_rules(df_valid['eyr'], 4, 2020, 2030)
# +
# hgt (Height) - a number followed by either cm or in:
# If cm, the number must be at least 150 and at most 193.
# If in, the number must be at least 59 and at most 76.
r = []
for item in df_valid['hgt']:
s = re.split('(cm|in)', item)
if len(s) >= 2:
if ((s[1] == 'cm') & ((int(s[0]) >= 150) & (int(s[0]) <= 193))):
r.append(True)
elif ((s[1] == 'in') & ((int(s[0]) >= 59) & (int(s[0]) <= 76))):
r.append(True)
else:
r.append(False)
else:
r.append(False)
df_check['hgt'] = r
# -
# hcl (Hair Color) - a # followed by exactly six characters 0-9 or a-f.
hcl_re = r'(^#[0-9a-z]{6}$)'
df_check['hcl'] = pd.isna(df_valid['hcl'].str.extract(hcl_re)) == False
# ecl (Eye Color) - exactly one of: amb blu brn gry grn hzl oth.
ecl_re = r'(amb|blu|brn|gry|grn|hzl|oth)'
df_check['ecl'] = pd.isna(df_valid['ecl'].str.extract(ecl_re)) == False
# pid (Passport ID) - a nine-digit number, including leading zeroes.
pid_re = r'(^\d{9}$)'
df_check['pid'] = pd.isna(df_valid['pid'].str.extract(pid_re)) == False
# tel rijen waar alle waarden 'True' zijn (true = 1, dus gemiddelde is dan 1)
print(f'Aantal geldige paspoorten: {len(df_check.loc[df_check.mean(axis = 1) == 1,:])}')
| Dag4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/skywalker0803r/c620/blob/main/notebook/Modeling_ICG_svr.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="BM6iz_tLAk2g"
import torch
from torch import nn
import numpy as np
import pandas as pd
import joblib
import matplotlib.pyplot as plt
# !pip install catboost > log.txt
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import StandardScaler
from sklearn.svm import SVR
from sklearn.pipeline import Pipeline
import random
torch.manual_seed(11)
np.random.seed(11)
random.seed(11)
# + id="pqXAl8LnGPD_"
def get_col(df,name):
return df.columns[df.columns.str.contains(name)]
# + [markdown] id="bxlX2azhhYbt"
# 
#
#
# + [markdown] id="m6HLNV1kFw-i"
# # load data
# + id="qVAUSM1Rstlj" colab={"base_uri": "https://localhost:8080/"} outputId="11b1749c-f9c1-420e-e758-36779b7e67e9"
c = joblib.load('/content/drive/MyDrive/台塑輕油案子/data/c620/col_names/icg_col_names.pkl')
c.keys(),c['x'],c['y'],len(c['x']),len(c['y'])
# + id="PKGO3l1qFWeQ" colab={"base_uri": "https://localhost:8080/", "height": 374} outputId="a8873cca-54da-4273-9f1c-277e72d468f2"
df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/c620_c670.csv',index_col=0)
df.head()
# + id="hIUVgPKydpqt" outputId="74cf9989-5504-4b5f-f28f-ab47ab110bb7" colab={"base_uri": "https://localhost:8080/"}
idx = df.index[:5]
idx
# + id="_vpjn562eBh4"
df.loc[idx,c['x']+c['y']].to_excel('/content/drive/MyDrive/台塑輕油案子/data/c620/Data_format_example/icg_data.xlsx')
# + [markdown] id="pRbtjflMDPSQ"
# # Distillate Rate_m3/hr == 0.01的部份 改成 0
# + id="bClY80IpC3iW"
df.loc[df['Simulation Case Conditions_C620 Distillate Rate_m3/hr']==0.01,'Simulation Case Conditions_C620 Distillate Rate_m3/hr'] = 0
# + id="GI8lnQIyPr39" colab={"base_uri": "https://localhost:8080/"} outputId="e78f7de7-9b29-4f34-cd17-3ef2a0dc79e1"
df[c['y']].max(),df[c['y']].min()
# + [markdown] id="1mMOhGQyU81L"
# # modeling 不要限制輸出 可以有負值
# + id="KisJWGajI8HN"
import catboost
from sklearn.model_selection import train_test_split
X = df[c['x']]
y = df[c['y']]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# + id="9q6vVxh1PiDA" colab={"base_uri": "https://localhost:8080/"} outputId="dab6c353-bc55-4096-b7c3-fbabf5a2b45f"
c620_icg = Pipeline([
('scaler',StandardScaler()),
('lr',SVR())])
c620_icg.fit(X_train,y_train)
# + [markdown] id="pyjRDUyLGwXM"
# # evaluate
# + id="DhD4BMaIKBe8"
from sklearn.metrics import r2_score,mean_squared_error
def mape(y_true, y_pred, e = 1e-8):
y_true, y_pred = np.array(y_true), np.array(y_pred)
mask = y_true > e
y_true, y_pred = y_true[mask], y_pred[mask]
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def show_metrics(y_real,y_pred,e=1e-8):
res = pd.DataFrame(index=y_pred.columns,columns=['R2','MSE','MAPE'])
for i in y_pred.columns:
res.loc[i,'R2'] = np.clip(r2_score(y_real[i],y_pred[i]),0,1)
res.loc[i,'MSE'] = mean_squared_error(y_real[i],y_pred[i])
res.loc[i,'MAPE'] = mape(y_real[i],y_pred[i],e)
res.loc['AVG'] = res.mean(axis=0)
return res
# + id="snJ7eOfyZ_vL" colab={"base_uri": "https://localhost:8080/", "height": 108} outputId="e9c7718b-e0a1-4428-b5ad-9720dfde1247"
y_real = y_test
y_pred = pd.DataFrame(c620_icg.predict(X_test),index=y_real.index,columns=y_real.columns)
show_metrics(y_real,y_pred)
# + [markdown] id="4cQXjWtBcbxY"
# # ICG while 迴圈
# + id="cD2SO3IvBvWe"
def ICG(Input):
while True:
output = pd.DataFrame(c620_icg.predict(Input.values),index=Input.index,columns=['Simulation Case Conditions_C620 Distillate Rate_m3/hr'])
dist_rate = output['Simulation Case Conditions_C620 Distillate Rate_m3/hr'].values[0]
na_in_benzene = Input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'].values[0]
print('current Distillate Rate_m3/hr:{} NA in Benzene_ppmw:{}'.format(dist_rate,na_in_benzene))
if output['Simulation Case Conditions_C620 Distillate Rate_m3/hr'].values[0] > 0:
return output,Input
else:
Input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'] -= 30
print('NA in Benzene_ppmw -= 30')
# + id="4NRN-RakiARY" colab={"base_uri": "https://localhost:8080/"} outputId="fc3ac174-a9f7-470e-9e8c-396698dd75ec"
print(df.shape)
df = df[df['Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%']==70.0]
df = df[df['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw']==980.0]
print(df.shape)
# + id="vSsgf_sNcGuc" colab={"base_uri": "https://localhost:8080/"} outputId="2bf430d7-3857-4a40-8f30-6a160d0a56da"
for i in range(10):
Input = df[c['x']].sample(1)
output,Input = ICG(Input)
print(i)
# + [markdown] id="3fHjLubVFDgn"
# # 特別資料測試
# + id="KvdUPKwkFDo8" colab={"base_uri": "https://localhost:8080/"} outputId="db56f1a2-3e9e-4a17-9164-8161e8a6d3b2"
Input = {
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm_m3/hr': {0: 183.71834522298502},
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm NA_wt%': {0: 2.06},
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm Benzene_wt%': {0: 11.91},
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm Toluene_wt%': {0: 37.48},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist_m3/hr': {0: 18.599028347620738},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist NA_wt%': {0: 2.51},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist Benzene_wt%': {0: 69.02},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist Toluene_wt%': {0: 20.62},
'Simulation Case Conditions_Feed Rate_Feed from T651_m3/hr': {0: 103.76780384010259},
'Simulation Case Conditions_Feed Rate_Feed from T651 NA_wt%': {0: 0.685},
'Simulation Case Conditions_Feed Rate_Feed from T651 Benzene_wt%': {0: 46.628},
'Simulation Case Conditions_Feed Rate_Feed from T651 Toluene_wt%': {0: 40.868},
'Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%': {0: 89.39},
'Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw': {0: 849.0}}
Input = pd.DataFrame(Input)
output,Input = ICG(Input)
# + id="UXaPFveeFahS" colab={"base_uri": "https://localhost:8080/"} outputId="42cecb3f-ab67-47ec-ffe9-a31d35c4a365"
Input = {
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm_m3/hr': {0: 171.58942010720452},
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm NA_wt%': {0: 1.971},
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm Benzene_wt%': {0: 12.146},
'Simulation Case Conditions_Feed Rate_Feed from V615 Btm Toluene_wt%': {0: 37.791},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist_m3/hr': {0: 19.669237218735567},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist NA_wt%': {0: 2.56},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist Benzene_wt%': {0: 68.421},
'Simulation Case Conditions_Feed Rate_Feed from C820 Dist Toluene_wt%': {0: 21.33},
'Simulation Case Conditions_Feed Rate_Feed from T651_m3/hr': {0: 101.95451869681709},
'Simulation Case Conditions_Feed Rate_Feed from T651 NA_wt%': {0: 0.777},
'Simulation Case Conditions_Feed Rate_Feed from T651 Benzene_wt%': {0: 50.531},
'Simulation Case Conditions_Feed Rate_Feed from T651 Toluene_wt%': {0: 35.435},
'Simulation Case Conditions_Spec 1 : Benzene in C620 Sidedraw_wt%': {0: 89.851},
'Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw': {0: 1193.0}}
Input = pd.DataFrame(Input)
output,Input = ICG(Input)
# + id="cm0RGUQ3R7fn" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="c7fc9422-ea77-46fd-ca47-feab2c0ee5fd"
df = pd.read_csv('/content/drive/MyDrive/台塑輕油案子/data/c620/cleaned/c620_c670.csv',index_col=0)
Input = df.loc['024-008',c['x']].to_frame().T
Input
# + id="LalwMjRBS3CL" colab={"base_uri": "https://localhost:8080/", "height": 185} outputId="58824d62-1a0a-4fd0-fe25-e39b46286cd9"
Input['Simulation Case Conditions_Spec 2 : NA in Benzene_ppmw'] = 920
Input
# + id="IruICO2PS-iS" colab={"base_uri": "https://localhost:8080/"} outputId="df7ca93d-80b8-4697-a71b-7e7517897e75"
c620_icg.predict(Input.values)
# + id="SwwsnjGlTI7K" colab={"base_uri": "https://localhost:8080/"} outputId="bf1331cf-007b-4fa7-82d8-6dea3e93d596"
_,_ = ICG(Input)
# + id="Bg1-IZygimjg" colab={"base_uri": "https://localhost:8080/"} outputId="b4c4a004-4661-42ab-871d-b64a66704363"
joblib.dump(c620_icg,'/content/drive/MyDrive/台塑輕油案子/data/c620/model/c620_icg_svr.pkl')
# + id="237TVoFnCfm0"
| notebook/Modeling_ICG_svr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Adversarial Robustness Toolbox (ART) and scikit-learn Pipeline
# This notebook contains an example of generating adversarial samples using a black-box attack against a scikit-learn pipeline consisting of principal component analysis (PCA) and a support vector machine classifier (SVC), but any other valid pipeline would work too. The pipeline is first optimised using grid search with cross validation. The adversarial samples are created with black-box `HopSkipJump` attack. The training data is MNIST, becasue of its intuitive visualisation, but any other dataset including tabular data would be suitable too.
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
from sklearn.decomposition import PCA
from sklearn.svm import SVC
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from art.utils import load_dataset
from art.estimators.classification import SklearnClassifier
from art.attacks.evasion import HopSkipJump
import warnings
warnings.filterwarnings('ignore')
# -
# ## Load the training and testing dataset
# +
n_features = 28*28
(x_train, y_train), (x_test, y_test), _, _ = load_dataset('mnist')
x_train = x_train.reshape((x_train.shape[0], n_features))
x_test = x_test.reshape((x_test.shape[0], n_features))
y_train = np.argmax(y_train, axis=1)
y_test = np.argmax(y_test, axis=1)
# Select a smaller set of samples to accelerate notebook example, remove for higher accuracy
x_train = x_train[0:1000]
x_test = x_test[0:100]
y_train = y_train[0:1000]
y_test = y_test[0:100]
# -
# ## Create a pipeline containing PCA and SVC classifier
svc = SVC(C=1.0, kernel='rbf')
pca = PCA()
pipeline = Pipeline(steps=[('pca', pca), ('svc', svc)])
# ## Grid search and cross validation to optimise number of PCA components and error term penalty
param_grid = {'pca__n_components': [5, 20, 30, 40, 50, 64],
'svc__C': np.logspace(-4, 4, 5)}
search = GridSearchCV(estimator=pipeline, param_grid=param_grid, iid=False, cv=5)
search.fit(x_train, y_train)
print("Best parameter (CV score=%0.3f):" % search.best_score_)
print(search.best_params_)
# ## Create a black-box attack using ART
classifier = SklearnClassifier(model=search.best_estimator_)
attack = HopSkipJump(classifier=classifier, targeted=False, norm=np.inf, max_iter=100, max_eval=100,
init_eval=100, init_size=100)
# ## Generate adversarial samples
x_test_adv = attack.generate(x_test)
# ## Evaluate benign accuracy
accuracy_test_benign = search.score(x_test, y_test)
print('Accuracy on benign test samples {}%:'.format(accuracy_test_benign * 100))
# ## Evaluate adversarial accuracy
accuracy_test_adversarial = search.score(x_test_adv, y_test)
print('Accuracy on adversarial test samples {}%:'.format(accuracy_test_adversarial * 100))
# ## Inspect a benign test sample
plt.matshow(x_test[0].reshape((28, 28)));
print('Predicted label:', search.predict(x_test[0:1])[0])
# ## Inspect an adversarial test sample
plt.matshow(x_test_adv[0].reshape((28, 28)));
print('Predicted label:', search.predict(x_test_adv[0:1])[0])
print('L_Infinity-norm:', np.linalg.norm(x_test_adv[0] - x_test[0], ord=np.inf))
| notebooks/classifier_scikitlearn_pipeline_pca_cv_svc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
import pickle
import pandas as pd
import numpy as np
target_protein = 'AAL (100 ug/ml)'
target_result_file = 'Results/eval-EXP-20201129-195018-model.pkl'
a = pickle.load(open(target_result_file, 'rb'))
y_iupac = a['y_iupac']
y_true = a['y_label']
y_pred = a['y_pred']
plt.rcParams['figure.dpi'] = 300
plt.rcParams['figure.figsize'] = [12.0, 4.0]
iupac_data = pd.read_csv('./Data/IUPAC.csv')
iupacs = iupac_data['IUPAC'].tolist()
mscore_data = pd.read_csv('./Data/MScore_useful.csv')
mscore = mscore_data[target_protein].tolist()
cv_order = sorted(list(zip(y_iupac, y_true, y_pred)))
excel_order = sorted(list(zip(iupacs, mscore)))
iupacs, y_true, y_pred = list(zip(*cv_order))
_, mscore = list(zip(*excel_order))
pack = sorted(list(zip(mscore, y_true, y_pred, iupacs)))
mscore, y_true, y_pred, iupacs = list(zip(*pack))
subject_range = range(len(y_true))
fig, ax1 = plt.subplots()
ax1.plot(subject_range, y_true, "ro", markersize=4, zorder=3, label="True Class")
ax1.plot(subject_range, y_pred, "go", markersize=8, zorder=2, label="Predict Class")
ax1.set_xlim([-0.5, len(y_true)])
ax1.set_xlabel("Proteins")
ax1.set_ylabel("Labels and Prediction Results")
ax2 = ax1.twinx()
ax2.plot(subject_range, mscore, label="M Score")
ax2.set_ylabel("M Score")
ax2.hlines(y=2.0, xmin=300, xmax=500, colors='purple', linestyles='-', lw=2, label='Threshold for Binary Classification')
# ax2.legend(loc="upper left")
# To combine two legend together
ax1.plot(np.nan, color='purple', label='Threshold for Binary Classification')
ax1.legend(loc="upper left")
plt.show()
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
import numpy as np
# Draw Bar Chart for monos
plt.rcParams['figure.dpi'] = 300
plt.rcParams["figure.figsize"] = (5, 3)
column_label = ['GlcNAc', 'Gal', 'End', 'Man', 'Fuc', 'Neu5Ac', 'GalNAc', 'Glc', 'Others']
occurrence = [1230, 1026, 945, 485, 283, 220, 208, 103, 32]
x = np.arange(len(column_label))
fig, ax = plt.subplots()
ax.bar(x, occurrence)
ax.set_xticks(x)
ax.set_xticklabels(column_label)
fig.autofmt_xdate()
plt.ylim([0, 1400])
plt.show()
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
import numpy as np
# Draw Bar Chart for links
plt.rcParams['figure.dpi'] = 300
plt.rcParams["figure.figsize"] = (5, 3)
column_label = [r'$(\beta 1-4)$', 'End-Linkage', 'Start-Linkage', r'$(\beta 1-3)$', r'$(\alpha 1-3)$', r'$(\beta 1-2)$', r'$(\beta 1-6)$', r'$(\beta 1-2)$', r'$(\beta 2-3)$', r'$(\beta 1-6)$', r'$(\beta 2-6)$', r'$(\beta 1-4)$', 'Others']
occurrence = [1086, 945, 600, 564, 332, 261, 196, 159, 129, 99, 76, 63, 32]
x = np.arange(len(column_label))
fig, ax = plt.subplots()
ax.bar(x, occurrence)
ax.set_xticks(x)
ax.set_xticklabels(column_label)
fig.autofmt_xdate()
plt.ylim([0, 1200])
plt.show()
# + pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
import numpy as np
import pickle
h2 = pickle.load(open('Saved/2-Class/eval-states_2_folds_5-20201202-171820-model.pkl', 'rb'))['Metrics']
h4 = pickle.load(open('Saved/2-Class/eval-states_4_folds_5-20201202-171820-model.pkl', 'rb'))['Metrics']
h6 = pickle.load(open('Saved/2-Class/eval-states_6_folds_5-20201202-171820-model.pkl', 'rb'))['Metrics']
h8 = pickle.load(open('Saved/2-Class/eval-states_8_folds_5-20201202-171820-model.pkl', 'rb'))['Metrics']
h2_f1 = h2['Normal']['F1'][4][0]
h2_prior_f1 = h2['Posterior']['F1'][4][0]
h4_f1 = h4['Normal']['F1'][4][0]
h4_prior_f1 = h4['Posterior']['F1'][4][0]
h6_f1 = h6['Normal']['F1'][4][0]
h6_prior_f1 = h6['Posterior']['F1'][4][0]
h8_f1 = h8['Normal']['F1'][4][0]
h8_prior_f1 = h8['Posterior']['F1'][4][0]
h2_f1_error = h2['Normal']['F1'][4][1] * 0.87653
h2_prior_f1_error = h2['Posterior']['F1'][4][1] * 0.87653
h4_f1_error = h4['Normal']['F1'][4][1] * 0.87653
h4_prior_f1_error = h4['Posterior']['F1'][4][1] * 0.87653
h6_f1_error = h6['Normal']['F1'][4][1] * 0.87653
h6_prior_f1_error = h6['Posterior']['F1'][4][1] * 0.87653
h8_f1_error = h8['Normal']['F1'][4][1] * 0.87653
h8_prior_f1_error = h8['Posterior']['F1'][4][1] * 0.87653
states = [2, 4, 6, 8]
f1s = [h2_f1, h4_f1, h6_f1, h8_f1]
f1s_error = [h2_f1_error, h4_f1_error, h6_f1_error, h8_f1_error]
f1s_prior = [h2_prior_f1, h4_prior_f1, h6_prior_f1, h8_prior_f1]
f1s_prior_error = [h2_prior_f1_error, h4_prior_f1_error, h6_prior_f1_error, h8_prior_f1_error]
plt.rcParams['figure.dpi'] = 300
plt.rcParams["figure.figsize"] = (5, 4)
plt.rcParams["errorbar.capsize"] = 5
plt.figure()
plt.errorbar(states, f1s, yerr=f1s_error, color='red', ecolor='red', capthick=2, label='Without Prior')
plt.errorbar(states, f1s_prior, yerr=f1s_prior_error, color='blue', ecolor='blue', capthick=2, label='With Prior')
plt.xticks(range(2, 9))
plt.xlabel('Number of Hidden States')
plt.ylabel('F1 Score (Mean)')
plt.title('Without Linkage')
plt.legend()
plt.show()
# + pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
import numpy as np
import pickle
edge_h2 = pickle.load(open('Saved/2-Class/eval-states_2_folds_5_use_edge-20201202-171820-model.pkl', 'rb'))['Metrics']
edge_h4 = pickle.load(open('Saved/2-Class/eval-states_4_folds_5_use_edge-20201202-171820-model.pkl', 'rb'))['Metrics']
edge_h6 = pickle.load(open('Saved/2-Class/eval-states_6_folds_5_use_edge-20201202-171820-model.pkl', 'rb'))['Metrics']
edge_h8 = pickle.load(open('Saved/2-Class/eval-states_8_folds_5_use_edge-20201202-171820-model.pkl', 'rb'))['Metrics']
edge_h2_f1 = edge_h2['Normal']['F1'][4][0]
edge_h2_prior_f1 = edge_h2['Posterior']['F1'][4][0]
edge_h4_f1 = edge_h4['Normal']['F1'][4][0]
edge_h4_prior_f1 = edge_h4['Posterior']['F1'][4][0]
edge_h6_f1 = edge_h6['Normal']['F1'][4][0]
edge_h6_prior_f1 = edge_h6['Posterior']['F1'][4][0]
edge_h8_f1 = edge_h8['Normal']['F1'][4][0]
edge_h8_prior_f1 = edge_h8['Posterior']['F1'][4][0]
edge_h2_f1_error = edge_h2['Normal']['F1'][4][1] * 0.87653
edge_h2_prior_f1_error = edge_h2['Posterior']['F1'][4][1] * 0.87653
edge_h4_f1_error = edge_h4['Normal']['F1'][4][1] * 0.87653
edge_h4_prior_f1_error = edge_h4['Posterior']['F1'][4][1] * 0.87653
edge_h6_f1_error = edge_h6['Normal']['F1'][4][1] * 0.87653
edge_h6_prior_f1_error = edge_h6['Posterior']['F1'][4][1] * 0.87653
edge_h8_f1_error = edge_h8['Normal']['F1'][4][1] * 0.87653
edge_h8_prior_f1_error = edge_h8['Posterior']['F1'][4][1] * 0.87653
states = [2, 4, 6, 8]
edge_f1s = [edge_h2_f1, edge_h4_f1, edge_h6_f1, edge_h8_f1]
edge_f1s_error = [edge_h2_f1_error, edge_h4_f1_error, edge_h6_f1_error, edge_h8_f1_error]
edge_f1s_prior = [edge_h2_prior_f1, edge_h4_prior_f1, edge_h6_prior_f1, edge_h8_prior_f1]
edge_f1s_prior_error = [edge_h2_prior_f1_error, edge_h4_prior_f1_error, edge_h6_prior_f1_error, edge_h8_prior_f1_error]
plt.rcParams['figure.dpi'] = 300
plt.rcParams["figure.figsize"] = (5, 4)
plt.rcParams["errorbar.capsize"] = 5
plt.figure()
plt.errorbar(states, edge_f1s, yerr=edge_f1s_error, color='green', ecolor='green', capthick=2, label='Without Prior')
plt.errorbar(states, edge_f1s_prior, yerr=edge_f1s_prior_error, color='orange', ecolor='orange', capthick=2, label='With Prior')
plt.xticks(range(2, 9))
plt.xlabel('Number of Hidden States')
plt.ylabel('F1 Score (Mean)')
plt.title('With Linkage')
plt.legend()
plt.show()
| Draw_figures_SQ.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# # Table tennis simulation
#
# This example shows the usage of `DiscreteEvents.jl` with event driven state machines.
#
# We implement players as timed state machines and thus need definitions of states and events and some data describing the players:
# +
using DiscreteEvents, Random, Printf
abstract type PState end
struct Idle <: PState end
struct Wait <: PState end
struct Unalert <: PState end
abstract type PEvent end
struct Start <: PEvent end
struct Serve <: PEvent end
struct Return <: PEvent end
struct Miss <: PEvent end
mutable struct Player
name::AbstractString
opp::Union{Number,Player}
state::PState
accuracy::Float64
attentiveness::Float64
score::Int64
Player(name, acc, att) = new(name, 0, Idle(), acc, att, 0)
end
# -
# Then we define some physical facts and a function to randomize them:
# +
const dist = 3 # distance for ball to fly [m]
const vs = 10 # serve velocity [m/s]
const vr = 20 # return velocity [m/s]
rd(s::Float64) = randn()*s + 1
# -
# Next we must describe the behaviour of our players. They are modeled as finite state machines, which have known states and react to known events. This is done with the `step!` function. Julia's multiple dispatch allows to give multiple definitions of `step!` for different combinations of states and events.
#
# The `serve` and `ret`-functions, used for describing serving and return of players are used to randomize the time and the behaviour of players. The players thus act probabilistically as Markov automata.
# +
function init!(p::Player, opp::Player)
p.opp = opp
if rand() ≤ p.attentiveness
p.state = Wait()
else
p.state = Unalert()
end
end
function serve(p::Player)
ts = 3 + dist*rd(0.15)/(vs*rd(0.25))
if (rand() ≤ p.accuracy) && (p.state == Wait())
event!(𝐶, SF(step!, p.opp, Serve()), after, ts)
@printf("%5.2f: %s serves %s\n", tau()+ts, p.name, p.opp.name)
else
event!(𝐶, SF(step!, p.opp, Miss()), after, ts)
@printf("%5.2f: %s serves and misses %s\n", tau()+ts, p.name, p.opp.name)
end
if rand() ≥ p.attentiveness
p.state = Unalert()
end
end
function ret(p::Player)
tr = dist*rd(0.15)/(vr*rd(0.25))
if rand() ≤ p.accuracy
event!(𝐶, SF(step!, p.opp, Return()), after, tr)
@printf("%5.2f: %s returns %s\n", tau()+tr, p.name, p.opp.name)
else
event!(𝐶, SF(step!, p.opp, Miss()), after, tr)
@printf("%5.2f: %s returns and misses %s\n", tau()+tr, p.name, p.opp.name)
end
if rand() ≥ p.attentiveness
p.state = Unalert()
end
end
# -
# The actual behaviour of a player is implemented as a state machine via the `step!`--function.
# +
"default transition for players"
step!(p::Player, q::PState, σ::PEvent) =
println("undefined transition for $(p.name), $q, $σ")
"player p gets a start command"
step!(p::Player, ::Union{Wait, Unalert}, ::Start) = serve(p)
"player p is waiting and gets served or returned"
step!(p::Player, ::Wait, ::Union{Serve, Return}) = ret(p)
"player p is unalert and gets served or returned"
function step!(p::Player, ::Unalert, ::Union{Serve, Return})
@printf("%5.2f: %s looses ball\n", τ(), p.name)
p.opp.score += 1
p.state = Wait()
serve(p)
end
"player p is waiting or unalert and gets missed"
function step!(p::Player, ::Union{Wait, Unalert}, ::Miss)
p.score += 1
p.state = Wait()
serve(p)
end
"simplified `step!` call"
step!(p::Player, σ::PEvent) = step!(p, p.state, σ)
# -
# In order to setup a simulation, we have to create and initialize the players, to start and run the game:
# +
ping = Player("Ping", 0.90, 0.90)
pong = Player("Pong", 0.90, 0.90)
init!(ping, pong)
init!(pong, ping)
step!(ping, Start())
Random.seed!(123)
println(run!(𝐶, 30))
println("Ping scored $(ping.score)")
println("Pong scored $(pong.score)")
# -
# Finally we reset `𝐶` for further simulations.
reset!(𝐶)
| notebooks/tabletennis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/YeonKang/Tensorflow-with-Colab/blob/master/Lab10_3_mnist_NN_dropout.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="aIDt3eMVHWr1" colab={"base_uri": "https://localhost:8080/"} outputId="8b45491e-b9b7-441e-90ff-09e6fa5af37f"
import tensorflow as tf
import numpy as np
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.datasets import mnist
from time import time
import os
print(tf.__version__)
# + [markdown] id="X3fZtzr8qmOA"
# **Checkpoint function**
# + id="UnQs_gD5qo4J"
def load(model, checkpoint_dir):
print(" [*] Reading checkpoints...")
ckpt = tf.train.get_checkpoint_state(checkpoint_dir)
if ckpt :
ckpt_name = os.path.basename(ckpt.model_checkpoint_path)
checkpoint = tf.train.Checkpoint(dnn=model)
checkpoint.restore(save_path=os.path.join(checkpoint_dir, ckpt_name))
counter = int(ckpt_name.split('-')[1])
print(" [*] Success to read {}".format(ckpt_name))
return True, counter
else:
print(" [*] Failed to find a checkpoint")
return False, 0
def check_folder(dir):
if not os.path.exists(dir):
os.makedirs(dir)
return dir
# + [markdown] id="2BGyGMYfqrF6"
# **Data load & pre-processing function**
# + id="yF6D1Cf6qtu2"
def load_mnist() :
(train_data, train_labels), (test_data, test_labels) = mnist.load_data()
train_data = np.expand_dims(train_data, axis=-1) #[N, 28, 28] -> [N, 28, 28, 1]
test_data = np.expand_dims(test_data, axis=-1) #[N, 28, 28] -> [N, 28, 28, 1]
train_data, test_data = normalize(train_data, test_data)
train_labels = to_categorical(train_labels, 10) #[N,] -> [N, 10]
test_labels = to_categorical(test_labels, 10) #[N,] -> [N, 10]
return train_data, train_labels, test_data, test_labels
def normalize(train_data, test_data):
train_data = train_data.astype(np.float32) / 255.0
test_data = test_data.astype(np.float32) / 255.0
return train_data, test_data
# + [markdown] id="Yf3W0L-_rD82"
# **Performance function**
# + id="8qdJTGk4rGbr"
def loss_fn(model, images, labels):
logits = model(images, training=True)
loss = tf.reduce_mean(tf.keras.losses.categorical_crossentropy(y_pred=logits, y_true=labels,
from_logits=True))
return loss
def accuracy_fn(model, images, labels):
logits = model(images, training=False)
prediction = tf.equal(tf.argmax(logits, -1), tf.argmax(labels, -1))
accuracy = tf.reduce_mean(tf.cast(prediction, tf.float32))
return accuracy
def grad(model, images, labels):
with tf.GradientTape() as tape:
loss = loss_fn(model, images, labels)
return tape.gradient(loss, model.variables)
# + [markdown] id="tu2fffyKrInT"
# **Model function**
# + id="msE5bL5vrK2T"
def flatten() :
return tf.keras.layers.Flatten()
def dense(label_dim, weight_init) :
return tf.keras.layers.Dense(units=label_dim, use_bias=True, kernel_initializer=weight_init)
def relu() :
return tf.keras.layers.Activation(tf.keras.activations.relu)
def dropout(rate) :
return tf.keras.layers.Dropout(rate)
# + [markdown] id="s9LfwMinPQ5h"
# **Create model (class version)**
# + id="mUGQRMbcPVfb"
class create_model_class(tf.keras.Model):
def __init__(self, label_dim):
super(create_model_class, self).__init__()
weight_init = tf.keras.initializers.glorot_uniform()
self.model = tf.keras.Sequential()
self.model.add(flatten())
for i in range(4):
self.model.add(dense(512, weight_init))
self.model.add(relu())
self.model.add(dropout(rate=0.5))
self.model.add(dense(label_dim, weight_init))
def call(self, x, training=None, mask=None):
x = self.model(x)
return x
# + [markdown] id="urFtJvg6PX2r"
# **Create model (function version)**
# + id="fizHkWAoPacm"
def create_model_function(label_dim) :
weight_init = tf.keras.initializers.glorot_uniform()
model = tf.keras.Sequential()
model.add(flatten())
for i in range(4) :
model.add(dense(512, weight_init))
model.add(relu())
model.add(dropout(rate=0.5))
model.add(dense(label_dim, weight_init))
return model
# + [markdown] id="YBhV76EkPciW"
# **Define data & hyper-parameter**
# + colab={"base_uri": "https://localhost:8080/"} id="sXVM45mlPez4" outputId="f2efbde8-d76b-453b-d813-712b1e0729fb"
""" dataset """
train_x, train_y, test_x, test_y = load_mnist()
""" parameters """
learning_rate = 0.001
batch_size = 128
training_epochs = 1
training_iterations = len(train_x) // batch_size
label_dim = 10
train_flag = True
""" Graph Input using Dataset API """
train_dataset = tf.data.Dataset.from_tensor_slices((train_x, train_y)).\
shuffle(buffer_size=100000).\
prefetch(buffer_size=batch_size).\
batch(batch_size, drop_remainder=True)
test_dataset = tf.data.Dataset.from_tensor_slices((test_x, test_y)).\
shuffle(buffer_size=100000).\
prefetch(buffer_size=len(test_x)).\
batch(len(test_x))
# + [markdown] id="1l2ixVduPh-J"
# **Define model & optimizer & writer**
# + id="rNcWjpgJPkZj"
""" Model """
network = create_model_function(label_dim)
""" Training """
optimizer = tf.keras.optimizers.Adam(learning_rate=learning_rate)
""" Writer """
checkpoint_dir = 'checkpoints'
logs_dir = 'logs'
model_dir = 'nn_dropout'
checkpoint_dir = os.path.join(checkpoint_dir, model_dir)
check_folder(checkpoint_dir)
checkpoint_prefix = os.path.join(checkpoint_dir, model_dir)
logs_dir = os.path.join(logs_dir, model_dir)
# + [markdown] id="t9gNJQBlPnmL"
# **Restore checkpoint & start train or test phase**
# + colab={"base_uri": "https://localhost:8080/"} id="Z9a7OKrSPsDS" outputId="7d0cd8a4-30a5-4c86-b0e4-ce8495634d5b"
if train_flag :
checkpoint = tf.train.Checkpoint(dnn=network)
#create writer for tensorboard
summary_writer = tf.summary.create_file_writer(logdir=logs_dir)
start_time = time()
#restore check-point if it exits
could_load, checkpoint_counter = load(network, checkpoint_dir)
if could_load:
start_epoch = (int)(checkpoint_counter / training_iterations)
counter = checkpoint_counter
print(" [*] Load SUCCESS")
else:
start_epoch = 0
start_iteration = 0
counter = 0
print(" [!] Load failed...")
#train phase
with summary_writer.as_default(): #for tensorboard
for epoch in range(start_epoch, training_epochs):
for idx, (train_input, train_label) in enumerate(train_dataset):
grads = grad(network, train_input, train_label)
optimizer.apply_gradients(grads_and_vars=zip(grads, network.variables))
train_loss = loss_fn(network, train_input, train_label)
train_accuracy = accuracy_fn(network, train_input, train_label)
for test_input, test_label in test_dataset:
test_accuracy = accuracy_fn(network, test_input, test_label)
tf.summary.scalar(name='train_loss', data=train_loss, step=counter)
tf.summary.scalar(name='train_accuracy', data=train_accuracy, step=counter)
tf.summary.scalar(name='test_accuracy', data=test_accuracy, step=counter)
print(
"Epoch: [%2d] [%5d/%5d] time: %4.4f, train_loss: %.8f, train_accuracy: %.4f, test_Accuracy: %.4f" \
% (epoch, idx, training_iterations, time() - start_time, train_loss, train_accuracy,
test_accuracy))
counter += 1
checkpoint.save(file_prefix=checkpoint_prefix + '-{}'.format(counter))
#test phase
else :
_, _ = load(network, checkpoint_dir)
for test_input, test_label in test_dataset:
test_accuracy = accuracy_fn(network, test_input, test_label)
print("test_Accuracy: %.4f" % (test_accuracy))
| Lab10_3_mnist_NN_dropout.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interval based time series classification in sktime
#
# Interval based approaches look at phase dependant intervals of the full series, calculating summary statistics from selected subseries to be used in classification.
#
# Currently 5 univariate interval based approaches are implemented in sktime. Time Series Forest (TSF) \[1\], the Random Interval Spectral Ensemble (RISE) \[2\], Supervised Time Series Forest (STSF) \[3\], the Canonical Interval Forest (CIF) \[4\] and the Diverse Representation Canonical Interval Forest (DrCIF). Both CIF and DrCIF have multivariate capabilities.
#
# In this notebook, we will demonstrate how to use these classifiers on the ItalyPowerDemand and JapaneseVowels datasets.
#
# #### References:
#
# \[1\] <NAME>., <NAME>., <NAME>., & <NAME>. (2013). A time series forest for classification and feature extraction. Information Sciences, 239, 142-153.
#
# \[2\] <NAME>., <NAME>., & <NAME>. (2019). The contract random interval spectral ensemble (c-RISE): the effect of contracting a classifier on accuracy. In International Conference on Hybrid Artificial Intelligence Systems (pp. 381-392). Springer, Cham.
#
# \[3\] <NAME>., <NAME>., <NAME>., & <NAME>. (2020). Fast and Accurate Time Series Classification Through Supervised Interval Search. In IEEE International Conference on Data Mining.
#
# \[4\] <NAME>., <NAME>., & <NAME>. (2020). The Canonical Interval Forest (CIF) Classifier for Time Series Classification. arXiv preprint arXiv:2008.09172.
#
# \[5\] <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2019). catch22: CAnonical Time-series CHaracteristics. Data Mining and Knowledge Discovery, 33(6), 1821-1852.
# ## 1. Imports
# +
from sklearn import metrics
from sktime.classification.interval_based import (
RandomIntervalSpectralForest,
SupervisedTimeSeriesForest,
TimeSeriesForestClassifier,
)
from sktime.classification.interval_based._cif import CanonicalIntervalForest
from sktime.classification.interval_based._drcif import DrCIF
from sktime.datasets import load_italy_power_demand
from sktime.datasets.base import load_japanese_vowels
# -
# ## 2. Load data
# +
X_train, y_train = load_italy_power_demand(split="train", return_X_y=True)
X_test, y_test = load_italy_power_demand(split="test", return_X_y=True)
X_test = X_test[:50]
y_test = y_test[:50]
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
X_train_mv, y_train_mv = load_japanese_vowels(split="train", return_X_y=True)
X_test_mv, y_test_mv = load_japanese_vowels(split="test", return_X_y=True)
X_train_mv = X_train_mv[:50]
y_train_mv = y_train_mv[:50]
X_test_mv = X_test_mv[:50]
y_test_mv = y_test_mv[:50]
print(X_train_mv.shape, y_train_mv.shape, X_test_mv.shape, y_test_mv.shape)
# -
# ## 3. Time Series Forest (TSF)
#
# TSF is an ensemble of tree classifiers built on the summary statistics of randomly selected intervals.
# For each tree sqrt(series_length) intervals are randomly selected.
# From each of these intervals the mean, standard deviation and slope is extracted from each time series and concatenated into a feature vector.
# These new features are then used to build a tree, which is added to the ensemble.
# +
tsf = TimeSeriesForestClassifier(n_estimators=200, random_state=47)
tsf.fit(X_train, y_train)
tsf_preds = tsf.predict(X_test)
print("TSF Accuracy: " + str(metrics.accuracy_score(y_test, tsf_preds)))
# -
# ## 4. Random Interval Spectral Ensemble (RISE)
#
# RISE is a tree based interval ensemble aimed at classifying audio data. Unlike TSF, it uses a single interval for each tree, and it uses spectral features rather than summary statistics.
# + pycharm={"name": "#%%\n"}
rise = RandomIntervalSpectralForest(n_estimators=200, random_state=47)
rise.fit(X_train, y_train)
rise_preds = rise.predict(X_test)
print("RISE Accuracy: " + str(metrics.accuracy_score(y_test, rise_preds)))
# -
# ## 5. Supervised Time Series Forest (STSF)
#
# STSF makes a number of adjustments from the original TSF algorithm. A supervised method of selecting intervals replaces random selection. Features are extracted from intervals generated from additional representations in periodogram and 1st order differences. Median, min, max and interquartile range are included in the summary statistics extracted.
# + pycharm={"name": "#%%\n"}
stsf = SupervisedTimeSeriesForest(n_estimators=200, random_state=47)
stsf.fit(X_train, y_train)
stsf_preds = stsf.predict(X_test)
print("STSF Accuracy: " + str(metrics.accuracy_score(y_test, stsf_preds)))
# -
# ## 6. Canonical Interval Forest (CIF)
#
# ### Univariate
#
# CIF extends from the TSF algorithm. In addition to the 3 summary statistics used by TSF, CIF makes use of the features from the `Catch22` \[5\] transform.
# To increase the diversity of the ensemble, the number of TSF and catch22 attributes is randomly subsampled per tree.
# + pycharm={"name": "#%%\n"}
cif = CanonicalIntervalForest(n_estimators=500, att_subsample_size=8, random_state=47)
cif.fit(X_train, y_train)
cif_preds = cif.predict(X_test)
print("CIF Accuracy: " + str(metrics.accuracy_score(y_test, cif_preds)))
# -
# ### Multivariate
# + pycharm={"name": "#%%\n"}
cif_m = CanonicalIntervalForest(n_estimators=500, att_subsample_size=8, random_state=47)
cif_m.fit(X_train, y_train)
cif_m_preds = cif_m.predict(X_test)
print("CIF Accuracy: " + str(metrics.accuracy_score(y_test, cif_m_preds)))
# -
# ## 6. Diverse Representation Canonical Interval Forest (DrCIF)
#
# ### Univariate
#
# DrCIF makes use of the periodogram and differences representations used by STSF as well as the addition summary statistics in CIF.
# + pycharm={"name": "#%%\n"}
drcif = DrCIF(n_estimators=500, att_subsample_size=10, random_state=47)
drcif.fit(X_train, y_train)
drcif_preds = drcif.predict(X_test)
print("DrCIF Accuracy: " + str(metrics.accuracy_score(y_test, drcif_preds)))
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Multivariate
# + pycharm={"name": "#%%\n"}
drcif_m = DrCIF(n_estimators=500, att_subsample_size=10, random_state=47)
drcif_m.fit(X_train, y_train)
drcif_m_preds = drcif_m.predict(X_test)
print("DrCIF Accuracy: " + str(metrics.accuracy_score(y_test, drcif_m_preds)))
| examples/interval_based_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MEGI001-2101033 Introduction to Earth System Data
#
# ## Task 6.2 - Data Handling Analysis (2-D)
#
# Created on: Jan 24, 2019 by <NAME> <<EMAIL>>
# ## T6.2 change histogram bin
#
# * please produce histograms of the above topography over Europe using different bins.
# * Considering that the resolution of the topography is about 1.8 km, how much is the area of the imaged part of europe and africa between 0 and 1000 meters of altitude in square kilometers?
# +
# -*- coding: utf-8 -*-
"""
Created on Jan 24 2019
@author: <NAME>, <NAME>
"""
# Import relevant libraries
import numpy as np
import xarray as xr
import netCDF4
from matplotlib import pyplot as plt
from PIL import Image
from scipy.interpolate import griddata
# fname = 'http://thredds.ucar.edu/thredds/dodsC/grib/NCEP/GFS/Global_onedeg/Best' # Remote OPeNDAP Dataset
filename = '../assets/data/geospatial-raster/etopo1.nc' # Local NetCDF file
# f = netCDF4.Dataset(filename)
f = xr.open_dataset(filename)
# # dsloc = ds.sel(lon=230.5,lat=55.0,method='nearest')
for v in f.variables: print(v)
print(f.Band1.data)
# -
#plotting
plt.imshow(f.Band1.data)
plt.show()
flipped = np.flipud(f.Band1.data)# it's flipped!
#plotting
plt.imshow(flipped)
plt.show()
# +
print("minumum latitude is: ", min(f.Band1.lat.values))
print("maximum latitude is: ", max(f.Band1.lat.values))
print("minumum longitude is: ", min(f.Band1.lon.values))
print("maximum longitude is: ", max(f.Band1.lon.values))
minlon = min(f.Band1.lon.values)
maxlon = max(f.Band1.lon.values)
minlat = min(f.Band1.lat.values)
maxlat = max(f.Band1.lat.values)
# +
# plot with legend, x an y axis labels
fig, ax = plt.subplots()
#Ticks for y-axis (latitude)
ax.axes.yaxis.set_ticks([0,flipped.shape[0]])
ax.axes.yaxis.set_ticklabels([maxlat,minlat])
plt.ylabel('Latitude')
#Ticks for x-axis (latitude)
ax.axes.xaxis.set_ticks([0,flipped.shape[1]])
ax.axes.xaxis.set_ticklabels([minlon,maxlon])
plt.xlabel('Longitude')
plt.imshow(flipped)
plt.colorbar()
plt.show()
# -
# plot with legend, x an y axis labels
fig, ax = plt.subplots()
# plt.imshow(flipped, cmap='rainbow')
plt.imshow(flipped, cmap='magma')
# plt.imshow(flipped, cmap='viridis') # the default one from above
plt.colorbar()
#Ticks for y-axis (latitude)
ax.axes.yaxis.set_ticks([0,flipped.shape[0]])
ax.axes.yaxis.set_ticklabels([maxlat,minlat])
plt.ylabel('Latitude')
#Ticks for x-axis (latitude)
ax.axes.xaxis.set_ticks([0,flipped.shape[1]])
ax.axes.xaxis.set_ticklabels([minlon,maxlon])
plt.xlabel('Longitude')
plt.show()
# histogramm of plot
plt.hist(flipped.ravel(), bins=10, range=(-6000, 6000), fc='k', ec='k')
plt.xlabel('Height above mean sea level')
plt.ylabel('Number of pixels')
plt.show()
# histogramm of plot
plt.hist(flipped.ravel(), bins=20, range=(0, 1000), fc='k', ec='k')
plt.xlabel('Height above mean sea level')
plt.ylabel('Number of pixels')
plt.show()
| notebooks/task_6/florent-ralph_task6.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Geonotebook (Python 2)
# language: python
# name: geonotebook2
# ---
# # Welcome to the Geonotebook
# GeoNotebook is an application that provides client/server enviroment with inteactive visualization and analysis capabilities using Jupyter notebook, GeoJS and other open source tools.
#
# The example notesbooks in this directory will walk you through several of the features that the ```geonotebook``` plugin to Jupyter makes available. The first thing to know about is...
#
# ### The geonotebook object
#
# The ```M``` object is inserted into the kernel automatically once the notebook is started. This object lives inside the Python kernel's namespace and communicates information to (and receives information from) the GeoJS map. Note that nothing has been imported, but the ```M``` variable is still available.
#
# **Note:** If you are viewing a static version of this notebook you will NOT see the GeoJS map that is dynamically added to a running notebook. Please see this [Screen shot](https://raw.githubusercontent.com/OpenGeoscience/geonotebook/master/screenshots/geonotebook.png) to get a sense of the running interface.
M
# ### Set the map's center
#
# The M object exposes a number of different functions for interacting with the map (which should be located to the right of standard jupyter python cells).
#
# Executing the following cell should set the center of the map to New York State.
#
# set_center's arguments are longitude, latitude, and zoom level
M.set_center(-74, 43, 6)
# ### What just happened?
#
# It is important to understand that ```M.set_center(...)``` is a Python statement being made inside the Python kernel. It is using a remote procedure call to change the javascript map's location.
#
# The standard Jupyter notebook has three components, (1) the client that makes up the notebook cells, (2) a web server that lists notebook files, directories and serves notebook assets like HTML and CSS (3) a kernel that executes commands in the chosen language (in our case Python).
#
# 
#
# When you executed the previous cell the string "M.set_center(-74, 43, 6)" was transmitted over a web socket to the webserver, then proxied through ZeroMQ to the IPykernel where it was evaluated as a Python expression. This is the standard way in which Jupyter notebook takes code from a web browser, and executes it in an interactive shell (kernel). M is an object in the kernel, and it has a function *set_center*. That function executed and returned a [promise](https://pypi.python.org/pypi/promise), which is why you see something in the cell output like ```<promise.promise.Promise at 0x7f567dd8f290>```
#
# While the ```set_center``` function returns a promise, it also has a side effect. This side effect uses a custom jupyter communication channel (or 'Comm') to tell the map to change its view port so the center is at (in this case) -74.0 latitude, 43.0 longitude, with a zoom level of 6.
#
#
# ## Widget example
#
# One question you may immediately ask yourself is, why not have the notebook cell talk to the map directly? Why get python involved at all? Well, because ```M.set_center``` is just a Python function, it can do things like leverage the existing widget extension to the notebook.
# +
from ipywidgets import interact
import ipywidgets as widgets
def map_widgets(lat=0.0, lon=0.0, zoom=4):
M.set_center(lon, lat, zoom)
interact(map_widgets, lat=(-90.0, 90.0), lon=(-180.0, 180.0), zoom=(1, 9))
# -
# # Annotations
#
# In addition to supporting Python to Map communications, Geonotebook allows objects and events on the map to communicate back to the Python kernel. One of the primary ways in which this is used is through geonotebook annotations.
#
# On the toolbar, next to the "CellToolbar" button, there should be three additional buttons with a circle, square and a polygon. Hovering over each of these reveals that they can be used to start a point, rectangle or polygon annotation.
#
# ### Point annotations
# Try clicking on the circle icon. Notice that the cursor, when it hovers over the map, is now a cross rather than an arrow. Click on the map and a circle annotation should appear.
#
# ### Rectangle Annotations
# Try clicking on the square icon. If you click on the map and hold down the left mouse button, then drag the mouse and release the left mouse button you should be able to create a rectangular annotation.
#
# ### Polygon annotations
# Try clicking on the polygon icon. Single click on a series of points to begin creating a polygon annotation. Double click on a point and the final segment will be added completing the annotation.
#
# Annotations inherit from [shapely](http://toblerity.org/shapely/manual.html) geometries, this means they support a wide range of spatial functions.
p = M.layers.annotation.polygons[0]
p
# You can get a list of coordinates for the polygon expressed in latitude and longitude
# List the exterior coordinates of the annotation
# Expressed in latitude and longitude point pairs
list(p.exterior.coords)
# Other properties like 'centroid' and 'area' are also available, keeping in mind that all coordinates are measured in latitude/longitude. This means properties like 'area' will not have much meaning. You can look at Shapely's [transform](http://toblerity.org/shapely/manual.html#shapely.ops.transform) method for information on how to translate these into to something more useful
list(p.centroid.coords)
# Here is an example of using shapely's transform method to convert coordinates from latitude/longitude (EPSG:4326) to Albers equal area (AEA). The resulting object gives area in units of meters squared
# +
import pyproj
import shapely.ops as ops
from functools import partial
project = partial(pyproj.transform,
pyproj.Proj(init='EPSG:4326'),
pyproj.Proj(proj='aea',
lat1=p.bounds[1],
lat2=p.bounds[3]))
ops.transform(project, p).area
# -
M.layers.annotation.clear_annotations()
# ## National Land Cover Dataset Example
# %matplotlib inline
from matplotlib.pylab import plt
import numpy as np
import pandas as pd
# +
legend = pd.DataFrame([
(11, "Open Water", "#476BA0"),
(12, "Perennial Ice/Snow", "#D1DDF9"),
(21, "Developed, Open Space","#DDC9C9"),
(22, "Developed, Low Intensity", "#D89382"),
(23, "Developed, Medium Intensity", "#ED0000"),
(24, "Developed High Intensity", "#AA0000"),
(31, "Barren Land (Rock/Sand/Clay)", "#B2ADA3"),
(41, "Deciduous Forest", "#68AA63"),
(42, "Evergreen Forest", "#1C6330"),
(43, "Mixed Forest", "#B5C98E"),
(51, "Dwarf Scrub", "#A58C30"),
(52, "Shrub/Scrub", "#CCBA7C"),
(71, "Grassland/Herbaceous", "#E2E2C1"),
(72, "Sedge/Herbaceous", "#C9C977"),
(73, "Lichens", "#99C147"),
(74, "Moss", "#77AD93"),
(81, "Pasture/Hay", "#DBD83D"),
(82, "Cultivated Crops", "#AA7028"),
(90, "Woody Wetlands", "#BAD8EA"),
(95, "Emergent Herbaceous Wetlands","#70A3BA")],
columns=["Code", "Desc", "Color"])
def highlight(e):
return 'background-color: {}'.format(e)
# +
from geonotebook.wrappers import RasterData
rd = RasterData("/data/kotfic/nlcd_2011_landcover_2011_edition_2014_10_10.tif")
colormap = legend[["Code", "Color"]].rename(columns={
"Code": "quantity", "Color": "color"}).to_dict("records")
# -
M.add_layer(rd[1], colormap=colormap, opacity=0.7)
# ### What just happened here?
#
# 
# ## National Land Cover Dataset
styles = [
dict(selector="th,td", props=[("font-size", "150%")])
]
legend.set_index("Code", inplace=True)
legend.style.applymap(highlight).set_table_styles(styles)
len(legend)
# !du -sh /data/kotfic/nlcd_2011_landcover_2011_edition_2014_10_10.tif
# +
import fiona
fh = fiona.open("/data/kotfic/nynta-wgs84/nynta-wgs84.shp")
# -
for feature in fh:
if feature['geometry']['type'] == "Polygon" and feature['properties']['BoroName'] == 'Manhattan':
M.add_annotation('polygon', feature['geometry']['coordinates'][0], feature['properties'])
p = M.layers.annotation.polygons[7]
p
p.NTAName
l, d = next(p.data)
d
# +
from collections import Counter
counts = zip(*np.unique(next(p.data)[1].data, return_counts=True))
print(p.NTAName)
data, index = zip(*[(num, legend.loc[c, 'Desc']) for c, num in counts if c != 0])
pd.Series(data, index=index, name="Count").to_frame()\
.sort_values("Count", ascending=False)\
.style.set_table_styles(styles)
# -
df = pd.DataFrame([(p.NTAName, n) for p in M.layers.annotation.polygons
for n in next(p.data)[1].compressed()],
columns=["Neighborhood", "Code"])
# +
n_idx = df['Code'].isin([24])
d_idx = df['Code'].isin([21, 22, 23, 24])
high_dev_codes = df[n_idx].groupby('Neighborhood').sum()
all_codes = df.groupby('Neighborhood').sum()
ddf = (high_dev_codes / all_codes).fillna(0.0).rename(columns={"Code": "High/All"})
ddf.sort_values("High/All", ascending=False).style.set_table_styles(styles)
# -
# ### Don't forget to take a screen shot!
M.layers.annotation.clear_annotations()
M.remove_layer(M.layers[0])
# # Raster operations on the map
#
# In this section we'll take a look at using the built in tile server to render raster data to the map. The tile server used is based on [KTile](https://github.com/OpenGeoscience/KTile) a fork of TileStache and is directly integrated into the Jupyter Notebook. The GeoJS map uses this tile server to render data efficiently to the map for visualization.
# Set the center of the map to the location the data
M.set_center(-120.32, 47.84, 7)
# +
from geonotebook.wrappers import RasterData
rd = RasterData('file:///data/kotfic/L57.Globe.month09.2010.hh09vv04.h6v1.doy247to273.NBAR.v3.0.tiff')
rd
# -
# ### Adding a single band with JET colormap
M.add_layer(rd[4], opacity=0.8)
# ### Something a little less agressive
# +
M.remove_layer(M.layers[0])
cmap = plt.get_cmap('winter', 10)
M.add_layer(rd[4], colormap=cmap, opacity=0.8)
# -
# ### Something more appropriate for NDVI
# +
from matplotlib.colors import LinearSegmentedColormap
M.remove_layer(M.layers[0])
# Divergent Blue to Beige to Green colormap
cmap =LinearSegmentedColormap.from_list(
'ndvi', ['blue', 'beige', 'green'], 20)
# Add layer with custom colormap
M.add_layer(rd[4], colormap=cmap, opacity=0.8, min=-1.0, max=1.0)
# -
# # What can I do with this data?
#
# We will address the use of annotations for analysis and data comparison in a separate notebook. For now Let's focus on a small agricultural area north of I-90:
M.layers.annotation.clear_annotations()
M.set_center(-119.25618502500376, 47.349300631765104, 11)
layer, data = next(M.layers.annotation.rectangles[0].data)
data
# As a sanity check we can prove the data is the region we've selected by plotting the data with matplotlib's [imshow](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.imshow) function:
#
# *Note* The scale of the matplotlib image may seem slightly different than the rectangle you've selected on the map. This is because the map is displaying in [Web Mercator](https://en.wikipedia.org/wiki/Web_Mercator) projection (EPSG:3857) while imshow is simply displaying the raw data, selected out of the geotiff (you can think of it as being in a 'row', 'column' projection).
# +
import numpy as np
fig, ax = plt.subplots(figsize=(16, 16))
ax.imshow(data, interpolation='none', cmap=cmap, clim=(-1.0, 1.0))
# -
# ### NDVI Segmentation analysis
#
# Once we have this data we can run arbitrary analyses on it. In the next cell we use a sobel filter and a watershed transformation to generate a binary mask of the data. We then use an implementation of marching cubes to vectorize the data, effectively segmenting green areas (e.g. fields) from surrounding areas.
#
# This next cell requires both [scipy](https://www.scipy.org/) and [scikit-image](http://scikit-image.org/). Check your operating system documentation for how best to install these packages.
# +
# Adapted from the scikit-image segmentation tutorial
# See: http://scikit-image.org/docs/dev/user_guide/tutorial_segmentation.html
import numpy as np
from skimage import measure
from skimage.filters import sobel
from skimage.morphology import watershed
from scipy import ndimage as ndi
WATER_MIN = 0.2
WATER_MAX = 0.6
def print_segments(data, THRESHOLD = 20):
fig, ax = plt.subplots(figsize=(10., 10.))
edges = sobel(data)
markers = np.zeros_like(data)
markers[data > WATER_MIN] = 2
markers[data > WATER_MAX] = 1
mask = (watershed(edges, markers) - 1).astype(bool)
seg = np.zeros_like(mask, dtype=int)
seg[~mask] = 1
# Fill holes
seg = ndi.binary_fill_holes(seg)
# Ignore entities smaller than THRESHOLD
label_objects, _ = ndi.label(seg)
sizes = np.bincount(label_objects.ravel())
mask_sizes = sizes > THRESHOLD
mask_sizes[0] = 0
clean_segs = mask_sizes[label_objects]
# Find contours of the segmented data
contours = measure.find_contours(clean_segs, 0)
ax.imshow(data, interpolation='none', cmap=cmap, clim=(-1.0, 1.0))
ax.axis('tight')
for n, contour in enumerate(contours):
ax.plot(contour[:, 1], contour[:, 0], linewidth=4)
print_segments(data)
# -
# ### Select a different region
print_segments(next(M.layers.annotation.rectangles[1].data)[1].data)
M.layers.annotation.clear_annotations()
M.remove_layer(M.layers[0])
| notebooks/demos/JupyterCon 2017 Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to *Merge $k$ sorted lists* (in Python 3)
#
# The question comes from [this nice blog on programming](https://tianrunhe.wordpress.com/2012/11/04/merge-k-sorted-lists/).
# I will solve it without giving much details, and test it quickly.
# ## Solution
# +
def merge_two(list1, list2):
if len(list1) == 0:
return list2[:]
elif len(list2) == 0:
return list1[:]
else:
if list1[0] < list2[0]:
return [list1[0]] + merge_two(list1[1:], list2)
else:
return [list2[0]] + merge_two(list1, list2[1:])
def merge(*lists):
head = []
for list_i in lists:
head = merge_two(head, list_i)
return head
# -
# ## Tests
# +
import random
def random_sorted_list(size):
return sorted([random.randint(0, 100) for _ in range(size)])
def issorted(alist):
return alist == sorted(alist)
for size in [10, 20, 30]:
for k in range(2, 20):
lists = [random_sorted_list(size) for _ in range(k)]
merged_list = merge(*lists)
assert issorted(merged_list)
# -
# ## Complexity
# One can prove that the algorithm we proposed is:
#
# - correctly merging $k$ sorted list into a sorted list containing the values from all the list,
# - and does so with an extra memory of at most $\mathcal{O}(k n)$ if all the lists have size at most $n$,
# - and does so with a time complexity of at most $\mathcal{O}(k n)$ if all the lists have size at most $n$.
# ## Conclusion
# *Et voilà.*
| Merge-k-sorted-lists.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Principal Component Analysis(PCA)
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# %matplotlib inline
from sklearn.datasets import load_breast_cancer
cancer=load_breast_cancer()
cancer.keys()
print(cancer['DESCR'])
df=pd.DataFrame(cancer['data'],columns=cancer['feature_names'])
df.head(5)
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
scaler=StandardScaler()
scaler.fit(df)
scaled_data=scaler.transform(df)
scaled_data
from sklearn.decomposition import PCA
pca=PCA(n_components=2)
pca.fit(scaled_data)
x_pca=pca.transform(scaled_data)
scaled_data.shape
x_pca.shape
scaled_data
x_pca
# +
plt.figure(figsize=(8,6))
plt.scatter(x_pca[:,0],x_pca[:,1],c=cancer['target'])
plt.xlabel('First principle component')
plt.ylabel('Second principle component')
# -
| PCA/PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="axF7OBsg-AJs"
# #Transformer
# + id="MiFdiUPY8CAr" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888569244, "user_tz": -540, "elapsed": 20787, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="22769aa4-770a-48aa-b0c1-9e93aaa2598f"
from google.colab import drive
drive.mount('/content/drive')
# + id="PtFKWnKP7mKd" colab={"base_uri": "https://localhost:8080/", "height": 269} executionInfo={"status": "ok", "timestamp": 1632888573219, "user_tz": -540, "elapsed": 3981, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="5ecf3d2d-94eb-4f6a-e305-0ff8b6ae0c2a"
# informer, ARIMA, Prophet, LSTMa와는 다른 형식의 CSV를 사용한다.(Version2)
# !pip install pandas
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
df = pd.read_csv('/content/drive/MyDrive/Colab Notebooks/Data/삼성전자_6M_NonST_Version2.csv', encoding='cp949')
df.head()
# + id="Z3_TiLg07mKk" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888573220, "user_tz": -540, "elapsed": 34, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="0ec7fd42-bcec-47b2-ae36-830208ac5163"
df.info()
# + id="m6hUoeDr7mKm" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888573220, "user_tz": -540, "elapsed": 29, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="d58219ed-ba58-48bb-c662-914c400869b1"
data_start_date = df.columns[1]
data_end_date = df.columns[-1]
print('Data ranges from %s to %s' % (data_start_date, data_end_date))
# + [markdown] id="-SsCTE-O7mKo"
# ### Train and Validation Series Partioning
#
# + id="OYUANKZx7mKs" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888573222, "user_tz": -540, "elapsed": 27, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="1a075be6-3110-411c-a4c7-ac4b1c8f763d"
######################## CHECK #########################
# 기준시간이 hour이므로, 7일 예측한다면 7*24로 설정한다.
from datetime import timedelta
pred_steps = 24*30+23
pred_length=timedelta(hours = pred_steps)
first_day = pd.to_datetime(data_start_date)
last_day = pd.to_datetime(data_end_date)
val_pred_start = last_day - pred_length + timedelta(1)
val_pred_end = last_day
print(val_pred_start, val_pred_end)
train_pred_start = val_pred_start - pred_length
train_pred_end = val_pred_start - timedelta(days=1)
print(train_pred_start, train_pred_end)
# + id="Yc-Nwfy07mKt" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888573223, "user_tz": -540, "elapsed": 24, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="2f01e1aa-0ed3-4deb-b7ba-4db31c81f063"
enc_length = train_pred_start - first_day
print(enc_length)
train_enc_start = first_day
train_enc_end = train_enc_start + enc_length - timedelta(1)
val_enc_start = train_enc_start + pred_length
val_enc_end = val_enc_start + enc_length - timedelta(1)
print(train_enc_start, train_enc_end)
print(val_enc_start, val_enc_end)
# + id="_w1ZjY4o7mKv" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888573224, "user_tz": -540, "elapsed": 20, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="f3788dcb-51d0-4109-ec21-bf8588471d63"
# 최종적으로 Val prediction 구간을 예측하게 된다.
print('Train encoding:', train_enc_start, '-', train_enc_end)
print('Train prediction:', train_pred_start, '-', train_pred_end, '\n')
print('Val encoding:', val_enc_start, '-', val_enc_end)
print('Val prediction:', val_pred_start, '-', val_pred_end)
print('\nEncoding interval:', enc_length.days)
print('Prediction interval:', pred_length.days)
# + [markdown] id="5k8nzx9H7mKw"
# ## Data Formatting
# + id="YYDpaFD47mKx" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888573608, "user_tz": -540, "elapsed": 393, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="62f84875-9032-428d-f8e4-d3e92194c63e"
#np.log 1p 해준다.
date_to_index = pd.Series(index=pd.Index([pd.to_datetime(c) for c in df.columns[1:]]),
data=[i for i in range(len(df.columns[1:]))])
series_array = df[df.columns[1:]].values.astype(np.float32)
print(series_array)
def get_time_block_series(series_array, date_to_index, start_date, end_date):
inds = date_to_index[start_date:end_date]
return series_array[:,inds]
def transform_series_encode(series_array):
series_array = np.log1p(np.nan_to_num(series_array)) # filling NaN with 0
series_mean = series_array.mean(axis=1).reshape(-1,1)
series_array = series_array - series_mean
series_array = series_array.reshape((series_array.shape[0],series_array.shape[1], 1))
return series_array, series_mean
def transform_series_decode(series_array, encode_series_mean):
series_array = np.log1p(np.nan_to_num(series_array)) # filling NaN with 0
series_array = series_array - encode_series_mean
series_array = series_array.reshape((series_array.shape[0],series_array.shape[1], 1))
return series_array
# + id="feu8hw4N7mKy"
# sample of series from train_enc_start to train_enc_end
encoder_input_data = get_time_block_series(series_array, date_to_index,
train_enc_start, train_enc_end)
encoder_input_data, encode_series_mean = transform_series_encode(encoder_input_data)
# sample of series from train_pred_start to train_pred_end
decoder_target_data = get_time_block_series(series_array, date_to_index,
train_pred_start, train_pred_end)
decoder_target_data = transform_series_decode(decoder_target_data, encode_series_mean)
encoder_input_val_data = get_time_block_series(series_array, date_to_index, val_enc_start, val_enc_end)
encoder_input_val_data, encode_series_mean = transform_series_encode(encoder_input_val_data)
decoder_target_val_data = get_time_block_series(series_array, date_to_index, val_pred_start, val_pred_end)
decoder_target_val_data = transform_series_decode(decoder_target_val_data, encode_series_mean)
#for d in encoder_input_data:
# print(d.shape)
#train_dataset = tf.data.Dataset.from_tensor_slices((encoder_input_data, decoder_target_data))
#train_dataset = train_dataset.batch(54)
#for d in train_dataset:
# #print(f'features:{features_tensor} target:{target_tensor}')
# print("-----")
# print(d)
# + [markdown] id="xZe8F9KP7mKz"
# ### Transformer model
# + id="2HymE_Lx7mK1" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888577917, "user_tz": -540, "elapsed": 4313, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="8c095bdb-2efb-44c8-aa8e-30876dbd7aa2"
# !pip install tensorflow_datasets
import tensorflow_datasets as tfds
import tensorflow as tf
import time
import numpy as np
import matplotlib.pyplot as plt
# + id="p9HiWaBB7mK3"
train_dataset = tf.data.Dataset.from_tensor_slices((encoder_input_data, decoder_target_data))
val_dataset = tf.data.Dataset.from_tensor_slices((encoder_input_val_data, decoder_target_val_data))
# + id="Mp70_nYj7mK3"
### position
def get_angles(pos, i, d_model):
angle_rates = 1 / np.power(10000, (2 * (i//2)) / np.float32(d_model))
return pos * angle_rates
def positional_encoding(position, d_model):
angle_rads = get_angles(np.arange(position)[:, np.newaxis],
np.arange(d_model)[np.newaxis, :],
d_model)
# apply sin to even indices in the array; 2i
sines = np.sin(angle_rads[:, 0::2])
# apply cos to odd indices in the array; 2i+1
cosines = np.cos(angle_rads[:, 1::2])
pos_encoding = np.concatenate([sines, cosines], axis=-1)
pos_encoding = pos_encoding[np.newaxis, ...]
return tf.cast(pos_encoding, dtype=tf.float32)
# + id="yqDIhFln7mK3" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888581144, "user_tz": -540, "elapsed": 20, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="68f67afa-044d-41a1-be42-86b679d94230"
# Masking
def create_padding_mask(seq):
seq = tf.cast(tf.math.equal(seq, 0), tf.float32)
# add extra dimensions so that we can add the padding
# to the attention logits.
return seq[:, tf.newaxis, tf.newaxis, :] # (batch_size, 1, 1, seq_len)
x = tf.constant([[7, 6, 0, 0, 1], [1, 2, 3, 0, 0], [0, 0, 0, 4, 5]])
print(create_padding_mask(x))
def create_look_ahead_mask(size):
mask = 1 - tf.linalg.band_part(tf.ones((size, size)), -1, 0)
return mask # (seq_len, seq_len)
x = tf.random.uniform((1, 4))
temp = create_look_ahead_mask(x.shape[1])
print(temp)
# + id="bZhflMS47mK4"
# Scaled dot product attention
def scaled_dot_product_attention(q, k, v, mask):
"""Calculate the attention weights.
q, k, v must have matching leading dimensions.
The mask has different shapes depending on its type(padding or look ahead)
but it must be broadcastable for addition.
Args:
q: query shape == (..., seq_len_q, depth)
k: key shape == (..., seq_len_k, depth)
v: value shape == (..., seq_len_v, depth)
mask: Float tensor with shape broadcastable
to (..., seq_len_q, seq_len_k). Defaults to None.
Returns:
output, attention_weights
"""
matmul_qk = tf.matmul(q, k, transpose_b=True) # (..., seq_len_q, seq_len_k)
# scale matmul_qk
dk = tf.cast(tf.shape(k)[-1], tf.float32)
scaled_attention_logits = matmul_qk / tf.math.sqrt(dk)
# add the mask to the scaled tensor.
if mask is not None:
scaled_attention_logits += (mask * -1e9)
# softmax is normalized on the last axis (seq_len_k) so that the scores
# add up to 1.
attention_weights = tf.nn.softmax(scaled_attention_logits, axis=-1) # (..., seq_len_q, seq_len_k)
output = tf.matmul(attention_weights, v) # (..., seq_len_v, depth)
return output, attention_weights
# + id="wTun4caK7mK7" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888582873, "user_tz": -540, "elapsed": 1744, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="89aef004-0602-4e96-daca-2477c073fae2"
# scaled dot product attetion test
def print_out(q, k, v):
temp_out, temp_attn = scaled_dot_product_attention(
q, k, v, None)
print ('Attention weights are:')
print (temp_attn)
print ('Output is:')
print (temp_out)
np.set_printoptions(suppress=True)
temp_k = tf.constant([[10,0,0],
[0,10,0],
[0,0,10],
[0,0,10]], dtype=tf.float32) # (4, 3)
temp_v = tf.constant([[ 1,0],
[ 10,0],
[ 100,5],
[1000,6]], dtype=tf.float32) # (4, 3)
# This `query` aligns with the second `key`,
# so the second `value` is returned.
temp_q = tf.constant([[0, 10, 0]], dtype=tf.float32) # (1, 3)
print_out(temp_q, temp_k, temp_v)
# + id="Tf9CrG-j7mK8"
# Multi Head Attention
class MultiHeadAttention(tf.keras.layers.Layer):
def __init__(self, d_model, num_heads):
super(MultiHeadAttention, self).__init__()
self.num_heads = num_heads
self.d_model = d_model
assert d_model % self.num_heads == 0
self.depth = d_model // self.num_heads
self.wq = tf.keras.layers.Dense(d_model)
self.wk = tf.keras.layers.Dense(d_model)
self.wv = tf.keras.layers.Dense(d_model)
self.dense = tf.keras.layers.Dense(d_model)
def split_heads(self, x, batch_size):
x = tf.reshape(x, (batch_size, -1, self.num_heads, self.depth))
return tf.transpose(x, perm=[0, 2, 1, 3])
def call(self, v, k, q, mask):
batch_size = tf.shape(q)[0]
q = self.wq(q)
k = self.wk(k)
v = self.wv(v) # (batch_size, seq_len, d_model)
q = self.split_heads(q, batch_size)
k = self.split_heads(k, batch_size)
v = self.split_heads(v, batch_size) #(batch_size, num_head, seq_len_v, depth)
# scaled_attention.shape == (batch_size, num_heads, seq_len_v, depth)
# attention_weights.shape == (batch_size, num_heads, seq_len_q, seq_len_k)
scaled_attention, attention_weights = scaled_dot_product_attention(
q, k, v, mask)
scaled_attention = tf.transpose(scaled_attention, perm=[0, 2, 1, 3]) # (batch_size, seq_len_v, num_heads, depth)
concat_attention = tf.reshape(scaled_attention,
(batch_size, -1, self.d_model)) # (batch_size, seq_len_v, d_model)
output = self.dense(concat_attention) # (batch_size, seq_len_v, d_model)
return output, attention_weights
# + id="vrKTDbzp7mK9" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888583305, "user_tz": -540, "elapsed": 21, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="6a5e4a69-8b32-4900-d41f-3edd1a10da2f"
# multhead attention test
temp_mha = MultiHeadAttention(d_model=512, num_heads=8)
y = tf.random.uniform((1, 60, 512)) # (batch_size, encoder_sequence, d_model)
out, attn = temp_mha(y, k=y, q=y, mask=None)
out.shape, attn.shape
# + id="h4jCVAuU7mK-"
# activation – the activation function of encoder/decoder intermediate layer, relu or gelu (default=relu).
# Point wise feed forward network
def point_wise_feed_forward_network(d_model, dff):
return tf.keras.Sequential([
tf.keras.layers.Dense(dff, activation='relu'), # (batch_size, seq_len, dff)
tf.keras.layers.Dense(d_model) # (batch_size, seq_len, d_model)
])
# + id="kR78pzaU7mK_" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888583305, "user_tz": -540, "elapsed": 14, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="28fd321c-6c2b-4360-d6ea-e78ac6bc0512"
# Point wise feed forward network test
sample_ffn = point_wise_feed_forward_network(512, 2048)
sample_ffn(tf.random.uniform((64, 50, 512))).shape
# + [markdown] id="0JHQxZvp7mK_"
# ### Encoder and Decoder
# + id="y_ja4MiQ7mLC"
# Encoder Layer
class EncoderLayer(tf.keras.layers.Layer):
def __init__(self, d_model, num_heads, dff, rate=0.1):
super(EncoderLayer, self).__init__()
self.mha = MultiHeadAttention(d_model, num_heads)
self.ffn = point_wise_feed_forward_network(d_model, dff)
self.layernorm1 = tf.keras.layers.BatchNormalization(epsilon=1e-6)
self.layernorm2 = tf.keras.layers.BatchNormalization(epsilon=1e-6)
self.dropout1 = tf.keras.layers.Dropout(rate)
self.dropout2 = tf.keras.layers.Dropout(rate)
def call(self, x, training, mask):
attn_output, _ = self.mha(x, x, x, mask) # (batch_size, input_seq_len, d_model)
attn_output = self.dropout1(attn_output, training=training)
out1 = self.layernorm1(x + attn_output)
ffn_output = self.ffn(out1) # (batch_size, input_seq_len, d_model)
ffn_output = self.dropout2(ffn_output, training=training)
out2 = self.layernorm2(out1 + ffn_output) # (batch_size, input_seq_len, d_model)
return out2
# + id="msUwz7Bx7mLD" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888583306, "user_tz": -540, "elapsed": 11, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="c85f5e7f-cf23-4249-ac6c-dfaf9c1a683d"
# Encoder Layer Test
sample_encoder_layer = EncoderLayer(512, 8, 2048)
sample_encoder_layer_output = sample_encoder_layer(
tf.random.uniform((64, 43, 512)), False, None)
sample_encoder_layer_output.shape # (batch_size, input_seq_len, d_model)
# + id="V-c2G1Cp7mLE"
# Decoder Layer
class DecoderLayer(tf.keras.layers.Layer):
def __init__(self, d_model, num_heads, dff, rate=0.1):
super(DecoderLayer, self).__init__()
self.mha1 = MultiHeadAttention(d_model, num_heads)
self.mha2 = MultiHeadAttention(d_model, num_heads)
self.ffn = point_wise_feed_forward_network(d_model, dff)
self.layernorm1 = tf.keras.layers.BatchNormalization(epsilon=1e-6)
self.layernorm2 = tf.keras.layers.BatchNormalization(epsilon=1e-6)
self.layernorm3 = tf.keras.layers.BatchNormalization(epsilon=1e-6)
self.dropout1 = tf.keras.layers.Dropout(rate)
self.dropout2 = tf.keras.layers.Dropout(rate)
self.dropout3 = tf.keras.layers.Dropout(rate)
def call(self, x, enc_output, training,
look_ahead_mask, padding_mask):
# enc_output.shape == (batch_size, input_seq_len, d_model)
attn1, attn_weights_block1 = self.mha1(x, x, x, look_ahead_mask)
attn1 = self.dropout1(attn1, training=training)
out1 = self.layernorm1(attn1 + x)
attn2, attn_weights_block2 = self.mha2(
enc_output, enc_output, out1, padding_mask)
attn2 = self.dropout2(attn2, training=training)
out2 = self.layernorm2(attn2 + out1)
ffn_output = self.ffn(out2)
ffn_output = self.dropout3(ffn_output, training=training)
out3 = self.layernorm3(ffn_output + out2)
return out3, attn_weights_block1, attn_weights_block2
# + id="s2HBk8KS7mLE" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888583611, "user_tz": -540, "elapsed": 18, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="59ced80b-f113-46b0-86bf-32c4bfa08ade"
# Decoder layer test
sample_decoder_layer = DecoderLayer(512, 8, 2048)
sample_decoder_layer_output, _, _ = sample_decoder_layer(
tf.random.uniform((64, 50, 512)), sample_encoder_layer_output,
False, None, None)
sample_decoder_layer_output.shape # (batch_size, target_seq_len, d_model)
# + id="-X496L3Y7mLF"
# Encoder
class Encoder(tf.keras.layers.Layer):
def __init__(self, num_layers, d_model, num_heads, dff, max_len=5000,
rate=0.1):
super(Encoder, self).__init__()
self.d_model = d_model
self.num_layers = num_layers
self.embedding = tf.keras.layers.Dense(d_model, use_bias=False)
self.pos_encoding = positional_encoding(max_len, self.d_model)
self.enc_layers = [EncoderLayer(d_model, num_heads, dff, rate)
for _ in range(num_layers)]
self.dropout = tf.keras.layers.Dropout(rate)
def call(self, x, training, mask):
seq_len = tf.shape(x)[1]
# adding embedding and position encoding
x = self.embedding(x)
# (batch_size, input_seq_len, d_model)
x *= tf.math.sqrt(tf.cast(self.d_model, tf.float32))
x += self.pos_encoding[:, :seq_len, :]
x = self.dropout(x, training=training)
for i in range(self.num_layers):
x = self.enc_layers[i](x, training, mask)
return x
# + id="YSO73qPR7mLF" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888583612, "user_tz": -540, "elapsed": 10, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="39ad32b0-bd79-4946-f744-d5256d2b5daa"
sample_encoder = Encoder(num_layers=2, d_model=512, num_heads=8,
dff=2048)
sample_encoder_output = sample_encoder(tf.random.uniform((64, 62,1)),
training=False, mask=None)
print (sample_encoder_output.shape) # (batch_size, input_seq_len, d_model)
# + id="iApaMNvC7mLG"
# Decoder
class Decoder(tf.keras.layers.Layer):
def __init__(self, num_layers, d_model, num_heads, dff, max_len=5000, rate=0.1):
super(Decoder, self).__init__()
self.d_model = d_model
self.num_layers = num_layers
self.embedding = tf.keras.layers.Dense(d_model, use_bias=False)
self.pos_encoding = positional_encoding(max_len, self.d_model)
self.dec_layers = [DecoderLayer(d_model, num_heads, dff, rate)
for _ in range(num_layers)]
self.dropout = tf.keras.layers.Dropout(rate)
def call(self, x, enc_output, training,
look_ahead_mask, padding_mask):
seq_len = tf.shape(x)[1]
attention_weights = {}
x = self.embedding(x)
x *= tf.math.sqrt(tf.cast(self.d_model, tf.float32))
x += self.pos_encoding[:, :seq_len, :]
x = self.dropout(x, training=training)
for i in range(self.num_layers):
x, block1, block2 = self.dec_layers[i](x, enc_output, training,
look_ahead_mask, padding_mask)
attention_weights['decoder_layer{}_block1'.format(i+1)] = block1
attention_weights['decoder_layer{}_block2'.format(i+1)] = block2
return x, attention_weights
# + id="lyz4sJBn7mLG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888583893, "user_tz": -540, "elapsed": 11, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="b131884c-78cb-4c9e-9331-185581e86ad8"
sample_decoder = Decoder(num_layers=2, d_model=512, num_heads=8,
dff=2048)
output, attn = sample_decoder(tf.random.uniform((64, 26,3)),
enc_output=sample_encoder_output,
training=False, look_ahead_mask=None,
padding_mask=None)
output.shape, attn['decoder_layer2_block2'].shape
# + [markdown] id="cDMZbb2D7mLH"
# ### Transfomer for TS
#
# + id="3Wff4gY-7mLH"
class Transformer(tf.keras.Model):
def __init__(self, num_layers, d_model, num_heads, dff, out_dim, max_len=5000,
rate=0.1):
super(Transformer, self).__init__()
self.encoder = Encoder(num_layers, d_model, num_heads, dff,
max_len, rate)
self.decoder = Decoder(num_layers, d_model, num_heads, dff,
max_len, rate)
self.final_layer = tf.keras.layers.Dense(out_dim)
def call(self, inp, tar, training, enc_padding_mask,
look_ahead_mask, dec_padding_mask):
enc_output = self.encoder(inp, training, enc_padding_mask)
dec_output, attention_weights = self.decoder(
tar, enc_output, training, look_ahead_mask, dec_padding_mask)
final_output = self.final_layer(dec_output)
return final_output, attention_weights
# + id="zy2jyCC27mLH" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888584688, "user_tz": -540, "elapsed": 799, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="e1c7069f-e937-4a69-ed4b-081257045fac"
sample_transformer = Transformer(
num_layers=2, d_model=512, num_heads=8, dff=2048,
out_dim=1)
temp_input = tf.random.uniform((64, 62,1))
temp_target = tf.random.uniform((64, 23,1))
fn_out, _ = sample_transformer(temp_input, temp_target,training=False,
enc_padding_mask=None,
look_ahead_mask=None,
dec_padding_mask=None)
fn_out.shape
# + id="vkBfkgGy7mLI"
# Set hyperparameters
# 트랜스포머 기준으로 바꿔볼까?
# d_model – the number of expected features in the encoder/decoder inputs (default=512).
# nhead – the number of heads in the multiheadattention models (default=8).
# num_encoder_layers – the number of sub-encoder-layers in the encoder & decoder (default=6).
# num_decoder_layers – the number of sub-decoder-layers in the decoder (default=6).
# dff(dim_feedforward) – the dimension of the feedforward network model (default=2048).
# dropout – the dropout value (default=0.1).
num_layers = 1
d_model = 64
dff = 256
num_heads = 8
dropout_rate = 0.1
input_sequence_length = 4320-(24*30+23) # Length of the sequence used by the encoder
target_sequence_length = 24*30+23 # Length of the sequence predicted by the decoder
batch_size = 2**11
train_dataset = train_dataset.batch(batch_size)
val_dataset = val_dataset.batch(batch_size)
# + id="dwNIQKry7mLI"
# Optimizizer
class CustomSchedule(tf.keras.optimizers.schedules.LearningRateSchedule):
def __init__(self, d_model, warmup_steps=4000):
super(CustomSchedule, self).__init__()
self.d_model = d_model
self.d_model = tf.cast(self.d_model, tf.float32)
self.warmup_steps = warmup_steps
def __call__(self, step):
arg1 = tf.math.rsqrt(step)
arg2 = step * (self.warmup_steps ** -1.5)
return tf.math.rsqrt(self.d_model) * tf.math.minimum(arg1, arg2)
# + id="EHuh2wWR7mLI"
learning_rate = CustomSchedule(64)
optimizer = tf.keras.optimizers.Adam(learning_rate, beta_1=0.9, beta_2=0.98,
epsilon=1e-9)
# + id="ftwg7b_Z7mLI" colab={"base_uri": "https://localhost:8080/", "height": 296} executionInfo={"status": "ok", "timestamp": 1632888584696, "user_tz": -540, "elapsed": 29, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="eb7e2b29-69aa-4f68-dd85-9b91<PASSWORD>cd"
temp_learning_rate_schedule = CustomSchedule(512)
plt.plot(temp_learning_rate_schedule(tf.range(40000, dtype=tf.float32)))
plt.ylabel("Learning Rate")
plt.xlabel("Train Step")
# + id="Tn59NBXK7mLI"
# Loss and metrics
loss_object = tf.keras.losses.MeanAbsoluteError()
# + id="0BwT16Z77mLJ"
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
# + id="vlFkoDfS7mLJ"
train_loss = tf.keras.metrics.Mean(name='train_loss')
#train_accuracy = tf.keras.metrics.mean_absolute_error()
test_loss = tf.keras.metrics.Mean(name='test_loss')
# + id="wpZb8zHP7mLJ"
# Training and checkpoint
transformer = Transformer(num_layers, d_model, num_heads, dff,
out_dim=1, rate=dropout_rate)
# + id="shPhCuBM7mLJ"
def create_masks(inp, tar):
inp = inp.reshape()
# Encoder padding mask
enc_padding_mask = create_padding_mask(inp)
# Used in the 2nd attention block in the decoder.
# This padding mask is used to mask the encoder outputs.
dec_padding_mask = create_padding_mask(inp)
# Used in the 1st attention block in the decoder.
# It is used to pad and mask future tokens in the input received by
# the decoder.
look_ahead_mask = create_look_ahead_mask(tf.shape(tar)[1])
dec_target_padding_mask = create_padding_mask(tar)
combined_mask = tf.maximum(dec_target_padding_mask, look_ahead_mask)
return enc_padding_mask, combined_mask, dec_padding_mask
# + id="r9ENhH-s7mLK"
# check point
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(transformer=transformer,
optimizer=optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
# if a checkpoint exists, restore the latest checkpoint.
if ckpt_manager.latest_checkpoint:
ckpt.restore(ckpt_manager.latest_checkpoint)
print ('Latest checkpoint restored!!')
# + id="zf4S65Ud7mLK"
# EPOCHS
EPOCHS=3000
# + id="dN2lAMeP7mLK"
@tf.function
def train_step(inp, tar):
last_inp = tf.expand_dims(inp[:,0,:],-1)
tar_inp = tf.concat([last_inp, tar[:,:-1,:]], axis=1)
tar_real = tar
#enc_padding_mask, combined_mask, dec_padding_mask = create_masks(inp, tar_inp)
#print(enc_padding_mask)
look_ahead_mask = create_look_ahead_mask(tf.shape(tar)[1])
with tf.GradientTape() as tape:
predictions, _ = transformer(inp, tar_inp,
True,
None,
look_ahead_mask,
None)
loss = loss_function(tar_real, predictions)
gradients = tape.gradient(loss, transformer.trainable_variables)
optimizer.apply_gradients(zip(gradients, transformer.trainable_variables))
train_loss(loss)
#train_accuracy(tar_real, predictions)
# + id="aFzrcnAU7mLK"
@tf.function
def test_step(inp, tar):
#print(inp)
#print(tar)
last_inp = tf.expand_dims(inp[:,0,:],-1)
#print(last_inp)
tar_inp = tf.concat([last_inp, tar[:,:-1,:]], axis=1)
tar_real = tar
look_ahead_mask = create_look_ahead_mask(tf.shape(tar)[1])
with tf.GradientTape() as tape:
predictions, _ = transformer(inp, tar_inp,
False,
None,
look_ahead_mask,
None)
loss = loss_function(tar_real, predictions)
gradients = tape.gradient(loss, transformer.trainable_variables)
optimizer.apply_gradients(zip(gradients, transformer.trainable_variables))
test_loss(loss)
# + id="VU9sFkFu7mLL" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888762685, "user_tz": -540, "elapsed": 177591, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="252c643b-dcb2-44e4-e40f-8bd7c3ac4c1e"
# Val_dataset을 돌려서 Val_prediction 구간을 예측한다
for epoch in range(EPOCHS):
start = time.time()
train_loss.reset_states()
test_loss.reset_states()
# validation:
for (batch, (inp, tar)) in enumerate(val_dataset):
#print(inp, tar)
test_step(inp, tar)
if (epoch + 1) % 5 == 0:
ckpt_save_path = ckpt_manager.save()
print ('Saving checkpoint for epoch {} at {}'.format(epoch+1,
ckpt_save_path))
#print ('Epoch {} Train Loss {:.4f}'.format(epoch + 1,
#train_loss.result()))
#train_accuracy.result()))
print ('Epoch {} Test Loss {:.4f}'.format(epoch + 1,
test_loss.result()))
print ('Time taken for 1 epoch: {} secs\n'.format(time.time() - start))
# + id="cU4kkZys7mLL"
MAX_LENGTH = target_sequence_length
def evaluate(inp):
encoder_input = inp
#print(encoder_input)
output = tf.expand_dims(encoder_input[:,-1,:],-1)
#print(output)
for i in range(MAX_LENGTH):
look_ahead_mask = create_look_ahead_mask(tf.shape(output)[1])
predictions, attention_weights = transformer(encoder_input,
output,
False,
None,
look_ahead_mask,
None)
# select the last word from the seq_len dimension
predictions = predictions[: ,-1:, :] # (batch_size, 1)
#print("pred:", predictions) #
output = tf.concat([output, predictions], axis=1)
#print(output)
return tf.squeeze(output, axis=0), attention_weights
# + id="8gnQAWpsA0Vs"
def mape(y_pred, y_true):
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
# + id="7khHQRegMByM"
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
# + id="y8O2fiewMEaG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888788744, "user_tz": -540, "elapsed": 25684, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="2cd998ad-21a3-47f0-955e-a3580899cb6a"
encode_series = encoder_input_val_data[0:1,:,:]
#print(encode_series)
pred_series, _ = evaluate(encode_series)
pred_series = np.array([pred_series])
encode_series = encode_series.reshape(-1,1)
pred_series = pred_series.reshape(-1,1)[1:,:]
target_series = decoder_target_val_data[0,:,:1].reshape(-1,1)
encode_series_tail = np.concatenate([encode_series[-999:],target_series[:1]])
x_encode = encode_series_tail.shape[0]
print(mape(pred_series[:24*30+23-23]*80846+81652.04075, target_series*80846+81652.04075))
print(mean_squared_error(target_series*80846+81652.04075, pred_series[:24*30+23-23]*80846+81652.04075))
print(mean_absolute_error(target_series*80846+81652.04075, pred_series[:24*30+23-23]*80846+81652.04075))
# + id="gRxmWFiyPIOG" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888788745, "user_tz": -540, "elapsed": 18, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="ec92d016-6189-4a7f-e7b4-cb5950556e7e"
x_encode
# + id="pzq3ApNZMHx6" colab={"base_uri": "https://localhost:8080/", "height": 408} executionInfo={"status": "ok", "timestamp": 1632888789488, "user_tz": -540, "elapsed": 747, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="439714f0-0e39-4fd1-fc22-ae12903c5b9f"
# 실제와 가격차이가 어떻게 나는지 비교해서 보정한다.
plt.figure(figsize=(20,6))
plt.plot(range(1,x_encode+1),encode_series_tail*80846+81652.04075)
plt.plot(range(x_encode,x_encode+pred_steps-23),target_series*80846+81652.04075,color='orange')
plt.plot(range(x_encode,x_encode+pred_steps-23),pred_series[:24*30+23-23]*80846+81652.04075,color='teal',linestyle='--')
plt.title('Encoder Series Tail of Length %d, Target Series, and Predictions' % 1000)
plt.legend(['Encoding Series','Target Series','Predictions'])
# + [markdown] id="FHfEpRQj_7jQ"
# #Prophet
# + id="GxabcXNaR_HX"
import pandas as pd
from fbprophet import Prophet
import matplotlib.pyplot as plt
import numpy as np
# + id="KMFg8RedAAXL" colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"status": "ok", "timestamp": 1632888869283, "user_tz": -540, "elapsed": 554, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="32b7bae7-67e4-4b07-efeb-180659e520dc"
df = pd.read_csv("/content/drive/MyDrive/Colab Notebooks/Data/삼성전자_6M_NonST_Version1.csv", encoding='CP949')
df = df.drop(df.columns[0], axis=1)
df.columns = ["ds","y"]
df["ds"] = pd.to_datetime(df["ds"], dayfirst = True)
df.head()
# + id="xv9FXHpaAELb" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632888875329, "user_tz": -540, "elapsed": 6049, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="1d87dfe2-d645-4828-91b1-bb37c4b68deb"
m = Prophet()
m.fit(df[:-24*30])
# + id="imkZ1wN0AGgF" colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"status": "ok", "timestamp": 1632888875331, "user_tz": -540, "elapsed": 23, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="560f006d-983f-4d04-89de-5f6d2ceb3d71"
future = m.make_future_dataframe(freq='H',periods=24*30)
future.tail()
# + id="xiGIX82FAIvT" colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"status": "ok", "timestamp": 1632888879766, "user_tz": -540, "elapsed": 4456, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="3adaa287-6480-4f33-d3f3-15a1e57ee2b7"
forecast = m.predict(future)
forecast[['ds', 'yhat']].tail()
# + id="L-HWlN-ZAN-f" colab={"base_uri": "https://localhost:8080/", "height": 320} executionInfo={"status": "ok", "timestamp": 1632888879772, "user_tz": -540, "elapsed": 28, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="1e62f21f-4e9c-4ec8-ab01-78265de7a382"
plt.figure(figsize=(20,5))
plt.plot(df["y"][1784:], label="real")
plt.plot(range(4320-24*30,4320),forecast['yhat'][-24*30:], label="Prophet")
plt.plot(range(4320-24*30,4320),pred_series[:24*30+23-23]*80846+81652.04075, label="Transformer")
plt.legend()
plt.show()
# + [markdown] id="UUMRI2j6Bocb"
# #LSTMa
# + id="7WLcYZZYBqhX"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import trange
import random
# + id="hGUOrhKuBscl" colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"status": "ok", "timestamp": 1632888879778, "user_tz": -540, "elapsed": 29, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="d05fe285-f289-4241-d410-f80cb31a8bd3"
data = pd.read_csv("/content/drive/MyDrive/Colab Notebooks/Data/삼성전자_6M_NonST_Version1.csv", encoding='CP949')
data.head()
# + id="MR7XsjsxBtY_"
from sklearn.preprocessing import MinMaxScaler
min_max_scaler = MinMaxScaler()
data["종가"] = min_max_scaler.fit_transform(data["종가"].to_numpy().reshape(-1,1))
# + id="LzqEQL_2Byuf"
train = data[:-24*30]
train = train["종가"].to_numpy()
test = data[-24*30:]
test = test["종가"].to_numpy()
# + id="3Q6VbWpwB09L"
import torch
import torch.nn as nn
from torch import optim
import torch.nn.functional as F
device = torch.device("cuda", index=0)
# + id="Vsgdq7a1B2Zf"
class lstm_encoder(nn.Module):
def __init__(self, input_size, hidden_size, num_layers = 1):
super(lstm_encoder, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.lstm = nn.LSTM(input_size = input_size, hidden_size = hidden_size, num_layers = num_layers, batch_first=True)
def forward(self, x_input):
lstm_out, self.hidden = self.lstm(x_input)
return lstm_out, self.hidden
# + id="g3V7hzfIB4ua"
class lstm_decoder(nn.Module):
def __init__(self, input_size, hidden_size, num_layers = 1):
super(lstm_decoder, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.num_layers = num_layers
self.lstm = nn.LSTM(input_size = input_size, hidden_size = hidden_size,num_layers = num_layers, batch_first=True)
self.linear = nn.Linear(hidden_size, input_size)
def forward(self, x_input, encoder_hidden_states):
lstm_out, self.hidden = self.lstm(x_input.unsqueeze(-1), encoder_hidden_states)
output = self.linear(lstm_out)
return output, self.hidden
# + id="khG35pB8B8Zb"
class lstm_encoder_decoder(nn.Module):
def __init__(self, input_size, hidden_size):
super(lstm_encoder_decoder, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.encoder = lstm_encoder(input_size = input_size, hidden_size = hidden_size)
self.decoder = lstm_decoder(input_size = input_size, hidden_size = hidden_size)
def forward(self, inputs, targets, target_len, teacher_forcing_ratio):
batch_size = inputs.shape[0]
input_size = inputs.shape[2]
outputs = torch.zeros(batch_size, target_len, input_size)
_, hidden = self.encoder(inputs)
decoder_input = inputs[:,-1, :]
for t in range(target_len):
out, hidden = self.decoder(decoder_input, hidden)
out = out.squeeze(1)
if random.random() < teacher_forcing_ratio:
decoder_input = targets[:, t, :]
else:
decoder_input = out
outputs[:,t,:] = out
return outputs
def predict(self, inputs, target_len):
inputs = inputs.unsqueeze(0)
self.eval()
batch_size = inputs.shape[0]
input_size = inputs.shape[2]
outputs = torch.zeros(batch_size, target_len, input_size)
_, hidden = self.encoder(inputs)
decoder_input = inputs[:,-1, :]
for t in range(target_len):
out, hidden = self.decoder(decoder_input, hidden)
out = out.squeeze(1)
decoder_input = out
outputs[:,t,:] = out
return outputs.detach().numpy()[0,:,0]
# + id="euepwoDsB_AS"
from torch.utils.data import DataLoader, Dataset
class windowDataset(Dataset):
def __init__(self, y, input_window=80, output_window=20, stride=5):
#총 데이터의 개수
L = y.shape[0]
#stride씩 움직일 때 생기는 총 sample의 개수
num_samples = (L - input_window - output_window) // stride + 1
#input과 output
X = np.zeros([input_window, num_samples])
Y = np.zeros([output_window, num_samples])
for i in np.arange(num_samples):
start_x = stride*i
end_x = start_x + input_window
X[:,i] = y[start_x:end_x]
start_y = stride*i + input_window
end_y = start_y + output_window
Y[:,i] = y[start_y:end_y]
X = X.reshape(X.shape[0], X.shape[1], 1).transpose((1,0,2))
Y = Y.reshape(Y.shape[0], Y.shape[1], 1).transpose((1,0,2))
self.x = X
self.y = Y
self.len = len(X)
def __getitem__(self, i):
return self.x[i], self.y[i]
def __len__(self):
return self.len
# + id="fyRG-o1DCB_6"
iw = 24*60
ow = 24*30
train_dataset = windowDataset(train, input_window=iw, output_window=ow, stride=1)
train_loader = DataLoader(train_dataset, batch_size=64)
# y_train_loader = DataLoader(y_train, batch_size=5)
# + id="o6RRY-yGCEFL"
model = lstm_encoder_decoder(input_size=1, hidden_size=16).to(device)
# model.train_model(X_train.to(device), y_train.to(device), n_epochs=100, target_len=ow, batch_size=5, training_bprediction="mixed_teacher_forcing", teacher_forcing_ratio=0.6, learning_rate=0.01, dynamic_tf=False)
# + id="Zh7o6HY4CGC8"
#5000으로 할 경우 시간도 오래걸리고 에러도 커서 100으로 줄인다.
learning_rate=0.01
epoch = 100
optimizer = optim.Adam(model.parameters(), lr = learning_rate)
criterion = nn.MSELoss()
# + id="zoaTXiKbCHce" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632890117027, "user_tz": -540, "elapsed": 1229103, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="ba995f34-63c7-4787-88e3-bcdefca10225"
from tqdm import tqdm
model.train()
with tqdm(range(epoch)) as tr:
for i in tr:
total_loss = 0.0
for x,y in train_loader:
optimizer.zero_grad()
x = x.to(device).float()
y = y.to(device).float()
output = model(x, y, ow, 0.6).to(device)
loss = criterion(output, y)
loss.backward()
optimizer.step()
total_loss += loss.cpu().item()
tr.set_postfix(loss="{0:.5f}".format(total_loss/len(train_loader)))
# + id="Shr1ZdBvCJfq"
predict = model.predict(torch.tensor(train_dataset[0][0]).to(device).float(), target_len=ow)
real = train_dataset[0][1]
# + id="6Y_bM2yLCMSN"
predict = model.predict(torch.tensor(train[-24*30*2:]).reshape(-1,1).to(device).float(), target_len=ow)
real = data["종가"].to_numpy()
predict = min_max_scaler.inverse_transform(predict.reshape(-1,1))
real = min_max_scaler.inverse_transform(real.reshape(-1,1))
# + id="AcWSBCdLCOeU" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632890117705, "user_tz": -540, "elapsed": 14, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="b013f3a0-d5bd-46df-a2f5-23c40f90a739"
real.shape
# + id="Qe3V9gtwCPsl" colab={"base_uri": "https://localhost:8080/", "height": 322} executionInfo={"status": "ok", "timestamp": 1632890118420, "user_tz": -540, "elapsed": 720, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="cbfe1258-3355-4e00-d9b8-32d59ff490c5"
plt.figure(figsize=(20,5))
plt.plot(range(3319,4320), real[3320:], label="real")
plt.plot(range(4320-24*30,4320), predict[-24*30:], label="LSTMa")
plt.plot(range(4320-24*30,4320),forecast['yhat'][-24*30:], label="Prophet")
plt.plot(range(4320-24*30,4320),pred_series[:24*30+23-23]*80846+81652.04075, label="Transformer")
plt.legend()
plt.show()
# + [markdown] id="ZHCM0K5uDzwc"
# #Informer
# + id="Ighg_VhkDrwT" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632890119734, "user_tz": -540, "elapsed": 1327, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="ae58a8b9-e01e-4343-a13f-ebece69492e9"
# !git clone https://github.com/zhouhaoyi/Informer2020.git
# + id="2yxRebPpD1nz" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632890119735, "user_tz": -540, "elapsed": 17, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="d1a999c9-44ec-4427-8b4f-fbedcc38ce46"
from google.colab import drive
drive.mount('/content/drive')
# + id="Q_nMD1gGD5ND"
import sys
if not 'Informer2020' in sys.path:
sys.path += ['Informer2020']
# + id="iTVBZNqqD6sX"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import MinMaxScaler
from datetime import timedelta
import torch
from torch import nn
from torch import optim
from torch.utils.data import DataLoader, Dataset
from tqdm import tqdm
from models.model import Informer
# + id="jjqkV_XfD8jj"
class StandardScaler():
def __init__(self):
self.mean = 0.
self.std = 1.
def fit(self, data):
self.mean = data.mean(0)
self.std = data.std(0)
def transform(self, data):
mean = torch.from_numpy(self.mean).type_as(data).to(data.device) if torch.is_tensor(data) else self.mean
std = torch.from_numpy(self.std).type_as(data).to(data.device) if torch.is_tensor(data) else self.std
return (data - mean) / std
def inverse_transform(self, data):
mean = torch.from_numpy(self.mean).type_as(data).to(data.device) if torch.is_tensor(data) else self.mean
std = torch.from_numpy(self.std).type_as(data).to(data.device) if torch.is_tensor(data) else self.std
return (data * std) + mean
def time_features(dates, freq='h'):
dates['month'] = dates.date.apply(lambda row:row.month,1)
dates['day'] = dates.date.apply(lambda row:row.day,1)
dates['weekday'] = dates.date.apply(lambda row:row.weekday(),1)
dates['hour'] = dates.date.apply(lambda row:row.hour,1)
dates['minute'] = dates.date.apply(lambda row:row.minute,1)
dates['minute'] = dates.minute.map(lambda x:x//15)
freq_map = {
'y':[],'m':['month'],'w':['month'],'d':['month','day','weekday'],
'b':['month','day','weekday'],'h':['month','day','weekday','hour'],
't':['month','day','weekday','hour','minute'],
}
return dates[freq_map[freq.lower()]].values
def _process_one_batch(batch_x, batch_y, batch_x_mark, batch_y_mark):
batch_x = batch_x.float().to(device)
batch_y = batch_y.float()
batch_x_mark = batch_x_mark.float().to(device)
batch_y_mark = batch_y_mark.float().to(device)
dec_inp = torch.zeros([batch_y.shape[0], pred_len, batch_y.shape[-1]]).float()
dec_inp = torch.cat([batch_y[:,:label_len,:], dec_inp], dim=1).float().to(device)
outputs = model(batch_x, batch_x_mark, dec_inp, batch_y_mark)
batch_y = batch_y[:,-pred_len:,0:].to(device)
return outputs, batch_y
# + id="ftToXwQzD_XZ"
class Dataset_Pred(Dataset):
def __init__(self, dataframe, size=None, scale=True):
self.seq_len = size[0]
self.label_len = size[1]
self.pred_len = size[2]
self.dataframe = dataframe
self.scale = scale
self.__read_data__()
def __read_data__(self):
self.scaler = StandardScaler()
df_raw = self.dataframe
df_raw["date"] = pd.to_datetime(df_raw["date"])
delta = df_raw["date"].iloc[1] - df_raw["date"].iloc[0]
if delta>=timedelta(hours=1):
self.freq='h'
else:
self.freq='t'
border1 = 0
border2 = len(df_raw)
cols_data = df_raw.columns[1:]
df_data = df_raw[cols_data]
if self.scale:
self.scaler.fit(df_data.values)
data = self.scaler.transform(df_data.values)
else:
data = df_data.values
tmp_stamp = df_raw[['date']][border1:border2]
tmp_stamp['date'] = pd.to_datetime(tmp_stamp.date)
pred_dates = pd.date_range(tmp_stamp.date.values[-1], periods=self.pred_len+1, freq=self.freq)
df_stamp = pd.DataFrame(columns = ['date'])
df_stamp.date = list(tmp_stamp.date.values) + list(pred_dates[1:])
data_stamp = time_features(df_stamp, freq=self.freq)
self.data_x = data[border1:border2]
self.data_y = data[border1:border2]
self.data_stamp = data_stamp
def __getitem__(self, index):
s_begin = index
s_end = s_begin + self.seq_len
r_begin = s_end - self.label_len
r_end = r_begin + self.label_len + self.pred_len
seq_x = self.data_x[s_begin:s_end]
seq_y = self.data_y[r_begin:r_end]
seq_x_mark = self.data_stamp[s_begin:s_end]
seq_y_mark = self.data_stamp[r_begin:r_end]
return seq_x, seq_y, seq_x_mark, seq_y_mark
def __len__(self):
return len(self.data_x) - self.seq_len- self.pred_len + 1
# + id="HOXR317aEHvb" colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"status": "ok", "timestamp": 1632890119741, "user_tz": -540, "elapsed": 16, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="58c6e976-11d1-4d81-df29-78482003cffc"
data = pd.read_csv("/content/drive/MyDrive/Colab Notebooks/Data/삼성전자_6M_NonST_Version1.csv", encoding='CP949')
data.head()
# + id="cWQ9usQTEKUl"
data["date"] = data["날짜"]
data["date"] = pd.to_datetime(data["date"], dayfirst = True)
data["value"] = data["종가"]
min_max_scaler = MinMaxScaler()
data["value"] = min_max_scaler.fit_transform(data["value"].to_numpy().reshape(-1,1)).reshape(-1)
data = data[["date", "value"]]
data_train = data.iloc[:-24*30].copy()
# + id="nrnJWkCXENIX"
pred_len = 24*30
seq_len = pred_len#인풋 크기
label_len = pred_len#디코더에서 참고할 크기
pred_len = pred_len#예측할 크기
batch_size = 10
shuffle_flag = True
num_workers = 0
drop_last = True
dataset = Dataset_Pred(dataframe=data_train ,scale=True, size = (seq_len, label_len,pred_len))
data_loader = DataLoader(dataset,batch_size=batch_size,shuffle=shuffle_flag,num_workers=num_workers,drop_last=drop_last)
# + id="4rWuHee2EPMC"
enc_in = 1
dec_in = 1
c_out = 1
device = torch.device("cuda:0")
model = Informer(enc_in, dec_in, c_out, seq_len, label_len, pred_len, device = device).to(device)
learning_rate = 1e-4
criterion = nn.MSELoss()
model_optim = optim.Adam(model.parameters(), lr=learning_rate)
# + id="3OuiFBlHESYz" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632896970587, "user_tz": -540, "elapsed": 6850320, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="ba74fc05-5ef0-422d-e982-dc61d7647aef"
# Informer는 error를 100하는게 시간도 덜 걸리고 에러도 적다.
train_epochs = 100
model.train()
progress = tqdm(range(train_epochs))
for epoch in progress:
train_loss = []
for i, (batch_x,batch_y,batch_x_mark,batch_y_mark) in enumerate(data_loader):
model_optim.zero_grad()
pred, true = _process_one_batch(batch_x, batch_y, batch_x_mark, batch_y_mark)
loss = criterion(pred, true)
train_loss.append(loss.item())
loss.backward()
model_optim.step()
train_loss = np.average(train_loss)
progress.set_description("loss: {:0.6f}".format(train_loss))
# + id="s0pez33IETCg" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632896972538, "user_tz": -540, "elapsed": 1958, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="fdf49bb3-db40-4aa1-b7df-c6407a983b21"
import time
now = time.time()
scaler = dataset.scaler
df_test = data_train.copy()
df_test["value"] = scaler.transform(df_test["value"])
df_test["date"] = pd.to_datetime(df_test["date"].values)
delta = df_test["date"][1] - df_test["date"][0]
for i in range(pred_len):
df_test = df_test.append({"date":df_test["date"].iloc[-1]+delta}, ignore_index=True)
df_test = df_test.fillna(0)
df_test_x = df_test.iloc[-seq_len-pred_len:-pred_len].copy()
df_test_y = df_test.iloc[-label_len-pred_len:].copy()
df_test_numpy = df_test.to_numpy()[:,1:].astype("float")
test_time_x = time_features(df_test_x, freq=dataset.freq) #인풋 타임 스템프
test_data_x = df_test_numpy[-seq_len-pred_len:-pred_len] #인풋 데이터
test_time_y = time_features(df_test_y, freq=dataset.freq) #아웃풋 타임스템프
test_data_y =df_test_numpy[-label_len-pred_len:]
test_data_y[-pred_len:] = np.zeros_like(test_data_y[-pred_len:]) #예측하는 부분을 0으로 채워준다.
test_time_x = test_time_x
test_time_y = test_time_y
test_data_y = test_data_y.astype(np.float64)
test_data_x = test_data_x.astype(np.float64)
_test = [(test_data_x,test_data_y,test_time_x,test_time_y)]
_test_loader = DataLoader(_test,batch_size=1,shuffle=False)
preds = []
with torch.no_grad():
for i, (batch_x,batch_y,batch_x_mark,batch_y_mark) in enumerate(_test_loader):
batch_x = batch_x.float().to(device)
batch_y = batch_y.float().to(device)
batch_x_mark = batch_x_mark.float().to(device)
batch_y_mark = batch_y_mark.float().to(device)
outputs = model(batch_x, batch_x_mark, batch_y, batch_y_mark)
preds = outputs.detach().cpu().numpy()
preds = scaler.inverse_transform(preds[0])
df_test.iloc[-pred_len:, 1:] = preds
print(time.time() - now)
# + id="B3mPZPrRKuVp"
# + id="MeRtX-qAD2ww"
# + id="XkN2Q60bRlyp"
# + id="Nw9690NE8_b1"
# + id="UKVKmhZLEWXz" colab={"base_uri": "https://localhost:8080/", "height": 320} executionInfo={"status": "ok", "timestamp": 1632896972540, "user_tz": -540, "elapsed": 17, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="4a8d8869-ef73-49f6-ce91-7ccadada159c"
import matplotlib.pyplot as plt
real = data["value"].to_numpy()
result = df_test["value"].iloc[-24*30:].to_numpy()
real = min_max_scaler.inverse_transform(real.reshape(-1,1)).reshape(-1)
result = min_max_scaler.inverse_transform(result.reshape(-1,1)).reshape(-1)
plt.figure(figsize=(20,5))
plt.plot(range(3319,4320),real[3320:], label="real")
plt.plot(range(4320-24*30,4320),result, label="Informer")
plt.plot(range(4320-24*30,4320), predict[-24*30:], label="LSTMa")
plt.plot(range(4320-24*30,4320),forecast['yhat'][-24*30:], label="Prophet")
plt.plot(range(4320-24*30,4320),pred_series[:24*30+23-23]*80846+81652.04075, label="Transformer")
plt.legend()
plt.show()
# + [markdown] id="oyQIUFttGo_w"
# #ARIMA
# + id="hiLPHVi8FtJA"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + id="U3r5vY8HFudT" colab={"base_uri": "https://localhost:8080/", "height": 206} executionInfo={"status": "ok", "timestamp": 1632896972945, "user_tz": -540, "elapsed": 412, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="d9db0517-5255-499b-9823-abb9764259a6"
df = pd.read_csv("/content/drive/MyDrive/Colab Notebooks/Data/삼성전자_6M_NonST_Version1.csv", encoding='CP949')
df = df.drop(df.columns[0], axis=1)
df.columns = ["ds","y"]
df.head()
# + id="3H-pHMqLF1ND"
df_train = df.iloc[:-24*30]
# + id="vVJjfCniF1xb" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632896972949, "user_tz": -540, "elapsed": 27, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="34a38d88-1b99-4022-920b-e0765a73480a"
from statsmodels.tsa.seasonal import seasonal_decompose
# + id="1AzDqmYFF3qS" colab={"base_uri": "https://localhost:8080/", "height": 506} executionInfo={"status": "ok", "timestamp": 1632896973629, "user_tz": -540, "elapsed": 690, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="28122f7d-cb38-4eb6-aab0-515f7b623db3"
import statsmodels.api as sm
fig = plt.figure(figsize=(20,8))
ax1 = fig.add_subplot(211)
fig = sm.graphics.tsa.plot_acf(df_train["y"], lags=20, ax=ax1)
fig = plt.figure(figsize=(20,8))
ax1 = fig.add_subplot(212)
fig = sm.graphics.tsa.plot_pacf(df_train["y"], lags=20, ax=ax1)
# + id="ljPV71OzF64s"
from statsmodels.tsa.arima_model import ARIMA
from statsmodels.tsa.statespace.sarimax import SARIMAX
import itertools
from tqdm import tqdm
# + id="rtlXujmwF60T" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632897101507, "user_tz": -540, "elapsed": 127899, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="4d7077e7-75b0-4c6b-b7d3-08de9e491969"
p = range(0,3)
d = range(1,2)
q = range(0,6)
m = 24
pdq = list(itertools.product(p,d,q))
seasonal_pdq = [(x[0],x[1], x[2], m) for x in list(itertools.product(p,d,q))]
aic = []
params = []
with tqdm(total = len(pdq) * len(seasonal_pdq)) as pg:
for i in pdq:
for j in seasonal_pdq:
pg.update(1)
try:
model = SARIMAX(df_train["y"], order=(i), season_order = (j))
model_fit = model.fit()
# print("SARIMA:{}{}, AIC:{}".format(i,j, round(model_fit.aic,2)))
aic.append(round(model_fit.aic,2))
params.append((i,j))
except:
continue
# + id="t77yJtlmGCGl" colab={"base_uri": "https://localhost:8080/", "height": 557} executionInfo={"status": "ok", "timestamp": 1632897109374, "user_tz": -540, "elapsed": 7884, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="4929dd83-128e-4dd4-c635-5f16ad8e6b07"
optimal = [(params[i],j) for i,j in enumerate(aic) if j == min(aic)]
model_opt = SARIMAX(df_train["y"], order = optimal[0][0][0], seasonal_order = optimal[0][0][1])
model_opt_fit = model_opt.fit()
model_opt_fit.summary()
# + id="PojlDVwHGDm9" colab={"base_uri": "https://localhost:8080/", "height": 458} executionInfo={"status": "ok", "timestamp": 1632897118016, "user_tz": -540, "elapsed": 8649, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="a6ac3c47-c501-4f03-a140-325edaa7755a"
model = SARIMAX(df_train["y"], order=optimal[0][0][0], seasonal_order=optimal[0][0][1])
model_fit = model.fit(disp=0)
ARIMA_forecast = model_fit.forecast(steps=24*30)
plt.figure(figsize=(20,5))
plt.plot(range(0,4320), df["y"].iloc[1:], label="Real")
plt.plot(ARIMA_forecast, label="ARIMA")
plt.plot(range(4320-24*30,4320),result, label="Informer")
plt.plot(range(4320-24*30,4320), predict[-24*30:], label="LSTMa")
plt.plot(range(4320-24*30,4320),forecast['yhat'][-24*30:], label="Prophet")
plt.plot(range(4320-24*30,4320),pred_series[:24*30+23-23]*80846+81652.04075, label="Transformer")
plt.legend()
plt.show()
# + id="b0hfHNU0ec52" colab={"base_uri": "https://localhost:8080/", "height": 320} executionInfo={"status": "ok", "timestamp": 1632897118607, "user_tz": -540, "elapsed": 616, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="724e4bcd-d1e7-4949-ed4e-d446ba28b996"
plt.figure(figsize=(20,5))
plt.plot(range(3319,4320), df["y"].iloc[3320:], label="Real")
plt.plot(ARIMA_forecast, label="ARIMA")
plt.plot(range(4320-24*30,4320),result, label="Informer")
plt.plot(range(4320-24*30,4320), predict[-24*30:], label="LSTMa")
plt.plot(range(4320-24*30,4320),forecast['yhat'][-24*30:], label="Prophet")
plt.plot(range(4320-24*30,4320),pred_series[:24*30+23-23]*80846+81652.04075, label="Transformer")
plt.legend()
plt.show()
# + id="a3VzMw4tIcni" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632897118608, "user_tz": -540, "elapsed": 58, "user": {"displayName": "\uae40\ud658\uc601", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "01699788118209796333"}} outputId="9ad848fe-d66a-4d96-c2aa-262eb9bba26a"
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_absolute_error
def MAPEval(y_pred, y_true):
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
def MSE(y_true, y_pred):
return np.mean(np.square((y_true - y_pred)))
def MAE(y_true, y_pred):
return np.mean(np.abs((y_true - y_pred)))
print('Transformer')
print('-' * 40)
print('MAPE: {} |\nMSE: {} |\nMAE : {}\n'.format(mape(pred_series[:24*30+23-23]*80846+81652.04075, target_series*80846+81652.04075), mean_squared_error(target_series*80846+81652.04075, pred_series[:24*30+23-23]*80846+81652.04075), mean_absolute_error(target_series*80846+81652.04075, pred_series[:24*30+23-23]*80846+81652.04075)))
print('Informer')
print('-' * 40)
print('MAPE: {} |\nMSE: {} |\nMAE : {}\n'.format(mape(result, real[-24*30:]), mean_squared_error(real[-24*30:], result), mean_absolute_error(real[-24*30:], result)))
print('ARIMA')
print('-' * 40)
print('MAPE: {} |\nMSE: {} |\nMAE : {}\n'.format(mape(ARIMA_forecast, df["y"].iloc[-24*30:]), mean_squared_error(df["y"].iloc[-24*30:], ARIMA_forecast), mean_absolute_error(df["y"].iloc[-24*30:], ARIMA_forecast)))
print('Prophet')
print('-' * 40)
print('MAPE: {} |\nMSE: {} |\nMAE : {}\n'.format(mape(forecast['yhat'][4320-24*30:],df["y"][4320-24*30:]), mean_squared_error(df["y"][4320-24*30:], forecast['yhat'][4320-24*30:]), mean_absolute_error(df["y"][4320-24*30:], forecast['yhat'][4320-24*30:])))
print('LSTMa')
print('-' * 40)
print('MAPE: {} |\nMSE: {} |\nMAE : {}\n'.format(mape(predict[-24*30:],real[-24*30:]), mean_squared_error(real[-24*30:], predict[-24*30:]), mean_absolute_error(real[-24*30:], predict[-24*30:])))
| Model/6M_720H.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Adding colors
# +
# Notice we are importing the color class!
import numpy as np
import color
# %matplotlib inline
# Auto-reload function so that this notebook keeps up with
# changes in the class file
# %load_ext autoreload
# %autoreload 2
# -
# ### Define colors and print them out
color1 = color.Color(250, 0, 0)
print(color1)
color2 = color.Color(0, 50, 200)
print(color2)
# ### Add the two colors and visualize the result!
#
# Once you've implemented the `__add__` function in the color class, you should be able to add colors with a `+` operator and display the result!
#
# Remember, to go back to all your files, click on the orange Jupyter icon at the top left of this notebook!
#
# Add the two colors to create a *new* color object
new_color = color1 + color2
print(new_color)
| 4_5_State_and_Motion_old/.ipynb_checkpoints/6. Add color playground-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import json
import argparse
import pprint
import datetime
import numpy as np
import matplotlib.pyplot as plt
import torch
from torch.utils import data
from bnaf import *
from tqdm import trange
from data.generate2d import sample2d, energy2d
# standard imports
import torch
import torch.nn as nn
from sklearn.datasets import make_moons
# from generate2d import sample2d, energy2d
# FrEIA imports
import FrEIA.framework as Ff
import FrEIA.modules as Fm
| Tutorials/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import os
# ## En datos/properati deben estar todos los archivos de ventas provistos por properati
filesProperati = filter( lambda f: not f.startswith('.'), os.listdir("datos/properati"))
# +
dataframes = []
for file_name in filesProperati:
newDataFrame = pd.read_csv("datos/properati/"+file_name, error_bad_lines = False)
dataframes.append(newDataFrame)
# -
# ## ¿Como filtramos por las zonas pedidas?
nulosStateName = 0
nulosPlaceWParentNames = 0
noCuentaConStateName = 0
noCuentaConPlaceWParentNames = 0
for df in dataframes:
try:
nulosStateName += df.state_name.isnull().sum()
except:
noCuentaConStateName += 1
try:
nulosPlaceWParentNames += df.place_with_parent_names.isnull().sum()
except:
noCuentaConPlaceWParentNames += 1
print "Cantidad de registros con valor nulo en state_name: "+str(nulosStateName)
print "Cantidad de registros con valor nulo en place_w_parent_names: "+str(nulosPlaceWParentNames)
print "Cantidad de archivos que no cuentan con la columna state_name: "+str(noCuentaConStateName)
print "Cantidad de archivos que no cuentan con la columna place_w_parent_names: "+str(noCuentaConPlaceWParentNames)
# ## Dado los resultados, para quedarnos solo con las propiedades de Capital Federal y GBA tendremos que utilizar la columna 'place_with_parent_names'
frecuencias = {}
for df in dataframes:
parentNames = df['place_with_parent_names']
for padres in parentNames:
zona = padres.split("|")[2]
if zona not in frecuencias:
frecuencias[zona] = 1
else:
frecuencias[zona]+=1
for zona in frecuencias:
print zona,frecuencias[zona]
# ## Interesan: Bs.As. G.B.A. Zona Norte, Bs.As. G.B.A. Zona Sur, Bs.As. G.B.A. Zona Oeste, Capital Federal
#
def filtrarCapitalYGBA(serie):
deseados = ["Bs.As. G.B.A. Zona Norte", "Bs.As. G.B.A. Zona Sur", "Bs.As. G.B.A. Zona Oeste", "Capital Federal"]
booleans = []
for item in serie:
if item.split("|")[2] in deseados:
booleans.append(True)
else:
booleans.append(False)
return booleans
filtrados =[]
for df in dataframes:
df = df[filtrarCapitalYGBA(df['place_with_parent_names'])]
filtrados.append(df)
# ### Por las dudas chequeo que ahora las frecuencias sean las mismas y solo queden las zonas que interesan
frecuencias = {}
for df in filtrados:
parentNames = df['place_with_parent_names']
for padres in parentNames:
zona = padres.split("|")[2]
if zona not in frecuencias:
frecuencias[zona] = 1
else:
frecuencias[zona]+=1
for key in frecuencias:
print key, frecuencias[key]
# # Ahora recupero datos, elimino algunas columnas, y demas.
# # Primero miro como estan los datos, para ver si conviene trabajar con los precios en pesos argentinos o dolares.
def a(df, column_name):
try:
return(df[column_name].isnull().sum(),0)
except:
return (0,1)
# +
nulsPriceAproxLocalCurr = 0
nulsPriceAproxUsd = 0
nulsPricePerM2 = 0
nulsPricePerM2usd = 0
priceAproxLocalCurrInex = 0
priceAproxUsdInex = 0
pricePerM2Inex = 0
pricePerM2usdInex = 0
for df in filtrados:
nuls, inex = a(df, "price_aprox_local_currency")
nulsPriceAproxLocalCurr += nuls
priceAproxLocalCurrInex += inex
nuls,inex = a(df, "price_aprox_usd")
nulsPriceAproxUsd += nuls
priceAproxUsdInex += inex
nuls, inex = a(df, "price_per_m2")
nulsPricePerM2 += nuls
pricePerM2Inex += inex
nuls,inex = a(df, "price_usd_per_m2")
nulsPricePerM2usd += nuls
pricePerM2usdInex += inex
# -
print str(nulsPriceAproxLocalCurr)+" Nulls priceAproxLocalCurr"
print str(nulsPriceAproxUsd)+" Nulls priceAproxUsd"
print str(nulsPricePerM2)+" Nulls Price Per M2"
print str(nulsPricePerM2usd)+" Nulls pricePerM2USD"
print str(priceAproxLocalCurrInex)+" Inexistentes priceAproxLocalCurr"
print str(priceAproxUsdInex)+" Inexistentes priceAproxUsd"
print str(pricePerM2Inex)+" Inexistentes pricePerM2"
print str(pricePerM2usdInex)+" Inexistentes pricePerM2usd"
# # Tambien miro los datos acerca de las superficies
# ## Observo que algunos archivos solo tienen una columna 'surface_in_m2', no distingue entre si es cubierta o si es total, suponemos que es la superficie total.
nulsSuperficieTotal = 0
for df in filtrados:
try:
nulsSuperficieTotal += df.surface_in_m2.isnull().sum()
except:
nulsSuperficieTotal += df.surface_total_in_m2.isnull().sum()
nulsSuperficieTotal
#Cuenta cuantas propiedades tienen el precio por m2 en dolares nulo,
#pero el precio local no nulo
count = 0
for df in filtrados:
try:
count += len(df[~df.price_per_m2.isnull() & df.price_usd_per_m2.isnull()])
except:
#Algunos dataframes no tienen el precio por m2 en pesos
continue
print "Cantidad de propiedades de las que se conoce el precio por m2 (En pesos)\
pero no en dolares: "+str(count)
# # ¿La columna price, tiene el mismo precio que price_aprox_usd o el mismo precio que price_aprox_local_curr? solo analizamos para el caso en el que no este el precio en dolares
# +
mismo_usd = 0
mismo_aprox_local_curr = 0
for df in filtrados:
mismo_aprox_local_curr += len(df[(df["price_aprox_local_currency"] == df["price"]) &\
(df["price"] != df["price_aprox_usd"]) & \
~df["price_aprox_usd"].isnull()])
mismo_usd += len(df[df["price_aprox_usd"] == df["price"]])
print mismo_aprox_local_curr
print mismo_usd
# -
# # Salvo pocos casos, la columna price contiene los mismos valores que la columna price_aprox_usd. Descarto la columna price. ¿Puedo recuperar precio de propiedades en dolares a partir del precio aproximado local?
# +
coincidencias = 0
for df in filtrados:
coincidencias += len(df[(~df["price_aprox_local_currency"].isnull() & df["price_aprox_usd"].isnull()) |\
(~df["price"].isnull() & df["price_aprox_usd"].isnull())])
print coincidencias
# -
# # Son tan solo dos casos en los que el precio aproximado local no es nulo y el precio en dolares lo es. No tenerlo en cuenta no deberia afectarnos
# # Se pueden recuperar datos de los precios en dolares por m2 realizando la correspondiente conversion. De igual manera, antes estudio para ver que no se pueda recuperar de otra manera.
# ## Antes de realizar conversiones o algunos calculos para poder recuperar el precio en USD, investigo si con la descripcion puedo recuperar algo de esto
sum(len(x) for x in filtrados) #cuantos registros tenemos en total?
for df in filtrados:
if "description" in df.columns:
df["description"] = df["description"].str.lower()
if "title" in df.columns:
df["title"] = df["title"].str.lower()
if "extra" in df.columns:
df["extra"] = df["extra"].str.lower()
total_no_price_usd = 0
total_recuperable_info = 0
for df in filtrados:
df_no_price_usd = df[df["price_aprox_usd"].isnull()]
cantidad = len(df_no_price_usd)
cols_considerar = []
if "extra" in df.columns:
cols_considerar.append("extra")
if "title" in df.columns:
cols_considerar.append("title")
if "description" in df.columns:
cols_considerar.append("description")
posibles_recuperables = 0
for col in cols_considerar:
menciona_dolares = df_no_price_usd[df_no_price_usd[col].str.contains("dolares")]
menciona_dolares_2 = df_no_price_usd[df_no_price_usd[col].str.contains("dólares")]
menciona_usd = df_no_price_usd[df_no_price_usd[col].str.contains("usd")]
posibles_recuperables += (len(menciona_dolares) + len(menciona_dolares_2) + len(menciona_usd))
total_no_price_usd += cantidad
total_recuperable_info += posibles_recuperables
print "Se encontro que "+str(total_no_price_usd)+" registros no tienen informacion del precio en USD"
print "Se encontro tambien que en "+str(total_recuperable_info)+" registros se podria recuperar el precio en USD de otras columnas"
print ""
# ## Hay algunos precios en dolar que podemos recuperar de la descripcion! hacemos una vista previa a el sector de la descripcion donde se menciona USD o dolares
for df in filtrados:
df_no_price_usd = df[df["price_aprox_usd"].isnull()]
cantidad = len(df_no_price_usd)
cols_considerar = []
if "extra" in df.columns:
cols_considerar.append("extra")
if "title" in df.columns:
cols_considerar.append("title")
if "description" in df.columns:
cols_considerar.append("description")
posibles_recuperables = 0
for col in cols_considerar:
menciona_dolares = df_no_price_usd[df_no_price_usd[col].str.contains("dolares")].head(1)
menciona_dolares_2 = df_no_price_usd[df_no_price_usd[col].str.contains("dólares")].head(1)
menciona_usd = df_no_price_usd[df_no_price_usd[col].str.contains("usd")].head(1)
dfs = [menciona_dolares, menciona_dolares_2, menciona_usd]
for d in dfs:
for index, row in d.iterrows():
text= row['description']
print ""
print text
break
# # Lamentablemente se observan cosas del estilo '1 usd null' o precios concatenados con otros strings, precios en distintos formatos y ademas a veces se menciona dolares/usd y se podria estar hablando de otra cosa, si bien se podria hacer el trabajo seria muy arduo y no vale la pena en este caso para 25000 registros de un total de 1797813.. de todos modos algunos de estos precios podrian ser recuperados posteriormente de otras formas que iremos viendo a continuacion:
# # ¿A cuantos se les desconoce el precio por m2 en USD y no se conoce el precio total (en USD) o la superficie total?
# +
coincidencias = 0
for df in filtrados:
if "price_per_m2" in df.columns:
coincidencias += len(df[(~df["price_per_m2"].isnull() & df["price_usd_per_m2"].isnull())\
& (df["price_aprox_usd"].isnull() | df["surface_total_in_m2"].isnull())])
print coincidencias
# -
# # 94025 de 639946 registros pueden ser recuperados si realizamos la conversion
# # Los datos para las conversiones fueron encontrados en:
# ### www.bcra.gob.ar/Pdfs/PublicacionesEstadisticas/com3500.xls
# # Los mismos fueron formateados de manera conveniente por como se nos presentan los datos de properati, usando LibreOffice Calc
# +
datosDolar = pd.read_csv("info_dolar.csv",index_col = "Fecha")
# -
from datetime import datetime
from datetime import timedelta
def recuperar_precio_m2_en_dolares(rowProperati):
if ~pd.isnull(rowProperati["price_per_m2"]) & pd.isnull(rowProperati["price_usd_per_m2"]):
fechaCreacionPropiedad = rowProperati["created_on"]
anho = int(fechaCreacionPropiedad.split("-")[0])
mes = int(fechaCreacionPropiedad.split("-")[1])
dia = int(fechaCreacionPropiedad.split("-")[2])
fechaConCotizacion = datetime(anho, mes, dia)
fechaEncontrada = False
while not(fechaEncontrada):
try:
cotizFechaCreacProp = float(datosDolar.ix[fechaConCotizacion.strftime('%Y-%m-%d')]["Cotizacion Dolar"])
fechaEncontrada = True
except:
#En esa fecha no se habia publicado informacion del dolar (no vario)
#Me fijo en el dia anterior
fechaConCotizacion = fechaConCotizacion - timedelta(days = 1)
return cotizFechaCreacProp*rowProperati["price_per_m2"]
else:
return rowProperati["price_per_m2"]
for df in filtrados:
if "price_per_m2" in df.columns:
df["price_usd_per_m2"] = df.apply(lambda row: recuperar_precio_m2_en_dolares(row),axis=1)
# # Corroboro
# +
coincidencias = 0
for df in filtrados:
if "price_per_m2" in df.columns:
coincidencias += len(df[(~df["price_per_m2"].isnull() & df["price_usd_per_m2"].isnull())\
& (df["price_aprox_usd"].isnull() | df["surface_total_in_m2"].isnull())])
print coincidencias
# -
# # Se recuperaron 94025 registros mediante la conversion a dolares del precio del metro cuadrado
# # ¿Se puede recuperar mas informacion?
#Primero le pongo el mismo nombre a la superficie total
for df in filtrados:
try:
df['surface_total_in_m2']
except:
df.rename(columns={'surface_in_m2':'surface_total_in_m2'},inplace=True)
#S 'se tiene informacion de' N 'no se tiene informacion de'
S_ppm2_S_sup_N_precio = 0
S_ppm2_S_precio_N_sup = 0
S_sup_S_precio_N_ppm2 = 0
for df in filtrados:
S_ppm2_S_sup_N_precio += len(df[~df["price_usd_per_m2"].isnull() &\
~df["surface_total_in_m2"].isnull() &\
df["price_aprox_usd"].isnull()])
S_ppm2_S_precio_N_sup += len(df[~df["price_usd_per_m2"].isnull() &\
~df["price_aprox_usd"].isnull() &\
df["surface_total_in_m2"].isnull()])
S_sup_S_precio_N_ppm2 += len(df[~df["surface_total_in_m2"].isnull() &\
~df["price_aprox_usd"].isnull() &\
df["price_usd_per_m2"].isnull()])
print S_ppm2_S_sup_N_precio
print S_ppm2_S_precio_N_sup
print S_sup_S_precio_N_ppm2
# # Recupero esta informacion
from utils_recuperacion_datos import recuperar_superficie,recuperar_precio_usd,recuperar_ppm2
for df in filtrados:
df['surface_total_in_m2'] = df.apply(lambda row: recuperar_superficie(row),axis=1)
df['price_aprox_usd'] = df.apply(lambda row: recuperar_precio_usd(row),axis=1)
df['price_usd_per_m2'] = df.apply(lambda row: recuperar_ppm2(row),axis=1)
# ## Chequeo que se hayan recuperado datos
#S 'se tiene informacion de' N 'no se tiene informacion de'
S_ppm2_S_sup_N_precio = 0
S_ppm2_S_precio_N_sup = 0
S_sup_S_precio_N_ppm2 = 0
for df in filtrados:
S_ppm2_S_sup_N_precio += len(df[~df["price_usd_per_m2"].isnull() &\
~df["surface_total_in_m2"].isnull() &\
df["price_aprox_usd"].isnull()])
S_ppm2_S_precio_N_sup += len(df[~df["price_usd_per_m2"].isnull() &\
~df["price_aprox_usd"].isnull() &\
df["surface_total_in_m2"].isnull()])
S_sup_S_precio_N_ppm2 += len(df[~df["surface_total_in_m2"].isnull() &\
~df["price_aprox_usd"].isnull() &\
df["price_usd_per_m2"].isnull()])
print S_ppm2_S_sup_N_precio
print S_ppm2_S_precio_N_sup
print S_sup_S_precio_N_ppm2
# ## A continuacion unifico y voy eliminando duplicados, teniendo en cuenta distintos subgrupos de columnas
unificacion = pd.concat(filtrados, axis=0, ignore_index=True)
len(unificacion)
#Elimino los registros que tengan todas las columnas identicas
a = unificacion.drop_duplicates(keep = "first")
len(a)
#Los que no tienen ni precio aproximado en dolares ya no me interesan
b = a[~a["price_aprox_usd"].isnull()]
len(b)
links_repetidos = b["properati_url"].value_counts()
# # Observo un poco que informacion encuentro en estas publicaciones con links repetidos
links_repetidos.head(2)
b[b["properati_url"] == "http://www.properati.com.ar/95fn_venta_departamento_capital-federal"].head(5)
b[b["properati_url"] == "http://www.properati.com.ar/7v2b_venta_departamento_olivos_avenida-del-libertador-gral-san-martin_2300"].head(5)
# # Observo que hay publicaciones con misma URL y que parecen ser las mismas propiedades, pero el inconveniente es que los campos vacios de cada uno de estos registros varia, debo filtrar de una forma mas compleja
# # Antes de hacer esto voy a eliminar columnas que no interesan
len(b[~b["lat"].isnull() & ~b["lon"].isnull() & b["lat-lon"].isnull()])
#Puedo eliminar latitud y longitud, ya que lat-lon tiene su informacion
#tambien, country_name, image_thumbnail, operation, price, price aprox local curr, currency
#ya que solo trabajaremos con precios en dolares
b.columns
data = b.drop(["country_name","lat","lon","image_thumbnail",\
"operation","price",'price_per_m2',"price_aprox_local_currency","currency"],axis=1)
len(data[data.duplicated(subset=["property_type","properati_url","price_aprox_usd","price_usd_per_m2","rooms","floor","place_name","surface_covered_in_m2","surface_total_in_m2","lat-lon","expenses","description","title","extra"])])
#Elimino registros con misma url de properati, mismo precio y misma cantidad de habitaciones y piso
data = data.drop_duplicates(subset=["property_type","properati_url","price_aprox_usd","price_usd_per_m2","rooms","floor","place_name","surface_covered_in_m2","surface_total_in_m2","lat-lon","expenses","description","title","extra"])
len(data)
# # Vamos a intentar recuperar el piso y la cantidad de habitaciones de los registros que no guarden esta informacion pero que la guarden en su descripcion
sin_rooms = data[data["rooms"].isnull()]
print len(sin_rooms)
print len(sin_rooms["description"].str.contains("habitaciones"))
print len(sin_rooms["description"].str.contains("ambiente"))
# ### capaz podemos llegar a recuperar todas las cantidades de habitaciones!
# ### Como se presenta esta informacion?
descripciones = sin_rooms["description"].values
print len(sin_rooms["title"].str.contains("ambientes"))
# +
posibles_descripciones_ambientes = {"1 ambiente": 1, "monoambiente": 1, "mono ambiente":1, "2 ambientes": 2,"3 ambientes": 3,
"4 ambientes": 4,"5 ambientes": 5, "6 ambientes": 6, "7 ambientes": 7,"8 ambientes": 8,
"9 ambientes": 1, "10 ambientes": 10,
"ambientes: 1": 1, "ambientes: 2": 2, "ambientes: 3": 3, "ambientes: 4": 4, "ambientes: 5": 5,
"ambientes: 6": 6, "ambientes: 7": 7, "un ambiente": 1, "dos ambientes": 2, "tres ambientes": 3,
"cuatro ambientes": 4,"cinco ambientes": 5," seis ambientes": 6, "siete ambientes": 7,
"2 impecables ambientes": 2, "3 impecables ambientes": 3, "4 amplios ambientes": 4, "2 amplios ambientes": 2,
"3 amplios ambientes": 3, "5 amplios ambientes": 5, "2 ambientes": 2, "3 ambientes": 3,
"2 grandes ambientes": 2, "6 amplios ambientes": 6, "1 ambientes": 1
}
encuentros = 0
for d in descripciones:
found = False
for k in posibles_descripciones_ambientes:
if not(pd.isnull(d)):
if k in d:
encuentros += 1
found = True
break
# if not(found) and not(pd.isnull(d)):
# split = d.split(" ")
# for i in range(0,len(split)):
# if split[i] == "ambientes":
# try:
# string = [split[i-2], split[i-1], split[i], split[i+1], split[i + 2]]
# for strr in string:
# if strr.isdigit():
# print " ".join(string)
# except:
# print d
print encuentros
# -
# # vemos que podemos recuperar al menos 65727 registros (porque esto no solo lo aplicamos a descripcion, a titulo y extra tambien) de los que no se tiene su habitacion asignada. Se podrian buscar mas casos extraños para agregar al diccionario pero al parecer no cambian demasiado el total de registros que se pueden recuperar.
# ## Hago lo mismo para el piso
sin_piso = data[data["floor"].isnull()]
print len(sin_piso)
print len(sin_piso["description"].str.contains("piso"))
descripciones = sin_piso["description"]
# +
posibles_descripciones_piso = {'1° piso': 1,
'2° piso': 2,
'3° piso': 3,
'4° piso': 4,
'5° piso': 5,
'6° piso': 6,
'7° piso': 7,
'8° piso': 8,
'9° piso': 9,
'10° piso': 10,
'11° piso': 11,
'12° piso': 12,
'13° piso': 13,
'14° piso': 14,
'15° piso': 15,
'16° piso': 16,
'17° piso': 17,
'18° piso': 18,
'19° piso': 19,
'20° piso': 20,
'21° piso': 21,
'22° piso': 22,
'23° piso': 23,
'24° piso': 24,
'25° piso': 25,
'26° piso': 26,
'27° piso': 27,
'28° piso': 28,
'29° piso': 29,
'30° piso': 30,
'31° piso': 31,
'32° piso': 32,
'33° piso': 33,
'34° piso': 34,
'35° piso': 35,
'36° piso': 36,
'37° piso': 37,
'38° piso': 38,
'39° piso': 39,
'40° piso': 40,
'41° piso': 41,
'42° piso': 42,
'43° piso': 43,
'44° piso': 44,
'45° piso': 45,
'46° piso': 46,
'47° piso': 47,
'48° piso': 48,
'49° piso': 49,
'50° piso': 50}
encuentros = 0
for d in descripciones:
found = False
for k in posibles_descripciones_piso:
if not(pd.isnull(d)):
if k in d:
encuentros += 1
found = True
break
# if not(found) and not(pd.isnull(d)):
# split = d.split(" ")
# for i in range(0,len(split)):
# if split[i] == "piso":
# try:
# string = [split[i- 4], split[i - 3],split[i-2], split[i-1], split[i], split[i+1], split[i + 2], split[i + 3], split[i + 4]]
# for strr in string:
# if strr.isdigit():
# print " ".join(string)
# except:
# print d
print encuentros
# +
#Se recuperian bastante menos, es mas complicado recuperar los pisos
# +
# Antes de agregar lo que vimos recien arreglamos un par de cosas que son que se tiene a barrios inexistentes en las distintas zonas como
# barrio, ej: capital federal en CABA
# -
barrios_caba = data[data["place_with_parent_names"].str.contains("Capital Federal")]["place_name"].value_counts().index
barrios_caba.delete(0) #Elimino capital federal
barrios_zn = []
barrios_zo = []
barrios_zs = []
data_barrios = data["place_with_parent_names"]
for v in data_barrios:
if v.split("|")[2] == "Bs.As. G.B.A. Zona Oeste":
try:
if v.split("|")[4] not in barrios_zo:
barrios_zo.append(v.split("|")[4])
except:
if v.split("|")[3] not in barrios_zo:
barrios_zo.append(v.split("|")[3])
elif v.split("|")[2] == "Bs.As. G.B.A. Zona Sur":
try:
if v.split("|")[4] not in barrios_zs:
barrios_zs.append(v.split("|")[4])
except:
if v.split("|")[3] not in barrios_zs:
barrios_zs.append(v.split("|")[3])
elif v.split("|")[2] == "Bs.As. G.B.A. Zona Norte":
try:
if v.split("|")[4] not in barrios_zn:
barrios_zn.append(v.split("|")[4])
except:
if v.split("|")[3] not in barrios_zn:
barrios_zn.append(v.split("|")[3])
barrios_zn.remove("")
barrios_zo.remove("")
barrios_zs.remove("")
#¿cuantos hay incorrectos?
bad_places = ["Capital Federal", "Bs.As. G.B.A. Zona Norte", "Bs.As. G.B.A. Zona Sur", "Bs.As. G.B.A. Zona Oeste"]
for place in bad_places:
print len(data[data["place_name"] == place])
barrios = {"Capital Federal": barrios_caba, "Bs.As. G.B.A. Zona Oeste": barrios_zo, "Bs.As. G.B.A. Zona Sur":barrios_zs,
"Bs.As. G.B.A. Zona Norte": barrios_zn}
# +
def recuperar_barrio(row):
place_actual = row["place_name"]
#Si es nulo hacemos una busqueda mas exhaustiva
descripcion = row["description"]
titulo = row["title"]
extra = row["extra"]
if (pd.isnull(place_actual)):
zona = "Capital Federal"
if row["place_with_parent_names"] == "Bs.As. G.B.A. Zona Norte":
zona = "Bs.As. G.B.A. Zona Norte"
elif row["place_with_parent_names"] == "Bs.As. G.B.A. Zona Sur":
zona = "Bs.As. G.B.A. Zona Sur"
elif row["place_with_parent_names"] == "Bs.As. G.B.A. Zona Oeste":
zona = "Bs.As. G.B.A. Zona Oeste"
barrios_posibles = barrios[zona]
if not(pd.isnull(descripcion)):
for b in barrios_posibles:
if b in descripcion:
return b
elif row["place_name"] in barrios:
barrios_posibles = barrios[row["place_name"]]
if not(pd.isnull(descripcion)):
for b in barrios_posibles:
if b in descripcion:
return b
else:
return row["place_name"]
# -
data["place_name"] = data.apply(lambda row: recuperar_barrio(row), axis = 1)
len(data[data["place_name"] == "Bs.As. G.B.A. Zona Norte"])
len(data[data["place_name"] == "Bs.As. G.B.A. Zona Sur"])
len(data[data["place_name"] == "Bs.As. G.B.A. Zona Oeste"])
len(data[data["place_name"] == "Capital Federal"])
# +
# Recupero los pisos y cantidad de ambientes que podamos
# -
def recuperar_atributo(row, atributo, info_atributo):
atributo_act = row[atributo]
if not(pd.isnull(atributo_act)):
return atributo_act
descripcion = row["description"]
extra = row["extra"]
title = row["title"]
atributo_corregido = atributo_act
cols = [descripcion, extra, title]
for col in cols:
if not(pd.isnull(col)):
for k in info_atributo:
if k in col:
atributo_corregido = info_atributo[k]
return atributo_corregido
return atributo_corregido #Si no se logro encontro nunca, devuelvo el NaN
data["floor"] = data.apply(lambda row: recuperar_atributo(row, "floor", posibles_descripciones_piso), axis = 1)
data["rooms"] = data.apply(lambda row: recuperar_atributo(row, "rooms", posibles_descripciones_ambientes), axis = 1)
data.isnull().sum()
# +
data.to_csv("training_data.csv", compression = "gzip")
# -
| recuperaciones_data_properati.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Imports
#
# Importing all required modules.
# %% pycharm={"name": "#%%\n"}
# %load_ext autoreload
# %autoreload 2
import datetime as dt
import os
from typing import Dict
import pandas as pd
import helpers.s3 as hs3
import im.kibot.data.load as kdl
import im.kibot.data.load.kibot_file_path_generator as fpgen
import im.common.data.types as types
# %% [markdown] pycharm={"name": "#%% md\n"}
# Define helper functions to calculate the report.
# %% pycharm={"name": "#%%\n"}
def slice_price_data(
price_df_dict: Dict[str, pd.DataFrame], last_years: int
) -> Dict[str, pd.DataFrame]:
"""Slice DataFrames for each symbol to contain records only for the
last_years years.
:param price_df_dict: {symbol: prices_for_symbol_df}
:param last_years: Number of years data is averaged to.
:return: {symbol: prices_for_symbol_df} sliced.
"""
now = dt.datetime.now()
# TODO(vr): check if dateutils.relativedate is better?
before = now - dt.timedelta(days=last_years * 365)
sliced_price_df_dict = {
symbol: prices.loc[before:now] for symbol, prices in price_df_dict.items()
}
return sliced_price_df_dict
def get_start_date(price_df_dict: Dict[str, pd.DataFrame]) -> pd.DataFrame:
"""Extract start dates for each time series.
:param price_df_dict: {symbol: prices_for_symbol_df}
:return: pd.DataFrame indexed by symbol
"""
start_date_dict = {
symbol: prices.index[0].strftime("%Y-%m-%d")
for symbol, prices in price_df_dict.items()
}
start_date_df = pd.DataFrame.from_dict(
start_date_dict, orient="index", columns=["start_date"]
)
return start_date_df
def get_price_data(
price_df_dict: Dict[str, pd.DataFrame],
price_col: str,
agg_func: str,
last_years: int,
) -> pd.DataFrame:
"""Get grouped prices for each symbol.
:param price_df_dict: {symbol: prices_for_symbol_df}
:param price_col: The name of the price column
:param agg_func: The name of the aggregation function that needs to
be applied to the prices for each symbol
:param last_years: Number of years data is averaged to.
:return: pd.DataFrame indexed by symbol
"""
price_dict = {
symbol: getattr(prices[price_col], agg_func)()
for symbol, prices in price_df_dict.items()
}
price_df = pd.DataFrame.from_dict(
price_dict,
orient="index",
columns=[f"{agg_func}_{last_years}y_{price_col}"],
)
price_df.index.name = "symbol"
return price_df
# %% [markdown] pycharm={"name": "#%% md\n"}
# Define main method to generate the report for a dataset.
# %% pycharm={"name": "#%%\n"}
def generate_report(
contract_type: types.ContractType, frequency: types.Frequency
) -> pd.DataFrame:
"""Generate a report for a dataset.
:param frequency: `D` or `T` for daily or minutely data respectively
:param contract_type: `continuous` or `expiry`
:return: a dataframe with the report
"""
dataset_aws_path = fpgen.FilePathGenerator().generate_file_path(
frequency, contract_type, "ROOT", ext=types.Extension.CSV
)
dataset_aws_directory = os.path.dirname(dataset_aws_path)
# Get a list of payloads (symbols) in format XYZ.csv.gz.
payloads = hs3.listdir(dataset_aws_directory, mode="non-recursive")
# Get only first n-rows.
n_rows = 100
# Get only symbols list.
symbols = tuple(
payload.replace(".csv.gz", "") for payload in payloads[:n_rows]
)
# Read dataframes.
kibot_data_loader = kdl.KibotDataLoader()
price_df_dict = kibot_data_loader.read_data(frequency, contract_type, symbols)
# Get avg. vol for the last 1 year
price_1y_df_dict = slice_price_data(price_df_dict, last_years=1)
mean_1y_vol = get_price_data(price_1y_df_dict, "vol", "mean", last_years=1)
# Get avg. vol for the last 3 years
price_3y_df_dict = slice_price_data(price_df_dict, last_years=3)
mean_3y_vol = get_price_data(price_3y_df_dict, "vol", "mean", last_years=3)
# Get avg. vol for the last 5 years
price_5y_df_dict = slice_price_data(price_df_dict, last_years=5)
mean_5y_vol = get_price_data(price_5y_df_dict, "vol", "mean", last_years=5)
# Get start date for each symbol.
start_date_df = get_start_date(price_df_dict)
# Get report for dataset.
report = pd.concat(
[start_date_df, mean_1y_vol, mean_3y_vol, mean_5y_vol],
axis=1,
join="inner",
)
report.index.name = "symbol"
report.fillna(0, inplace=True)
return report
# %% [markdown] pycharm={"name": "#%% md\n"}
# Report for all_futures_contracts_1min
# %% pycharm={"name": "#%%\n"}
dataset_report = generate_report(
types.ContractType.Expiry, types.Frequency.Minutely
)
dataset_report
# %% [markdown]
# Report for all_futures_contracts_daily
# %% pycharm={"name": "#%%\n"}
dataset_report = generate_report(types.ContractType.Expiry, types.Frequency.Daily)
dataset_report
# %% [markdown]
# Report for futures_continuous_contracts_1min
# %% pycharm={"is_executing": true, "name": "#%%\n"}
dataset_report = generate_report(
types.ContractType.Continuous, types.Frequency.Minutely
)
dataset_report
# %% [markdown]
# Report for futures_continuous_contracts_daily
# %% pycharm={"is_executing": true, "name": "#%%\n"}
dataset_report = generate_report(
types.ContractType.Continuous, types.Frequency.Daily
)
dataset_report
| im/kibot/notebooks/kibot_volume_statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow_cpu]
# language: python
# name: conda-env-tensorflow_cpu-py
# ---
from core.dataset import Dataset
from tqdm import tqdm
from matplotlib import pyplot as plt
trainset = Dataset('train')
type(trainset)
pbar = tqdm(trainset)
for train_data in pbar:
print(type(train_data))
print(train_data[0].shape)
print(train_data[1].shape)
print(train_data[4].shape)
train_data = trainset.__next__()
# 6 is the batch size defined in config.py
train_data[0].shape
plt.imshow(train_data[0][0])
train_data[3].shape
train_data[3][0, :, :, :, 4:5].sum(axis=-1).shape
plt.imshow(train_data[3][0, :, :, :, 4:5].sum(axis=-1))
plt.imshow(train_data[1][0, :, :, :, 4:5].sum(axis=-1))
plt.imshow(train_data[2][0, :, :, :, 4:5].sum(axis=-1))
train_data[4].shape
train_data[5].shape
plt.imshow(train_data[0][3])
plt.imshow(train_data[3][3, :, :, :, 4:5].sum(axis=-1))
plt.imshow(train_data[2][3, :, :, :, 4:5].sum(axis=-1))
# it looks like for this picture, two scales have the ground-truth bounding boxes for the same objects. Why?
plt.imshow(train_data[1][3, :, :, :, 4:5].sum(axis=-1))
plt.imshow(train_data[0][4])
plt.imshow(train_data[3][4, :, :, :, 4:5].sum(axis=-1))
plt.imshow(train_data[2][4, :, :, :, 4:5].sum(axis=-1))
plt.imshow(train_data[1][4, :, :, :, 4:5].sum(axis=-1))
| dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import COLLAB_Prediction as cp
# The following function allows you to train the provided models with a feature-augmented version of the datasets.
#
# zr.perform_collab_prediction(Dataset, Model, random_seed)
#
# The available options for the dataset are: "OGBL-COLLAB", "OGBL-COLLAB_3Cl", "OGBL-COLLAB_4Cl",
# "OGBL-COLLAB_5Cl", "OGBL-COLLAB_34Cl", "OGBL-COLLAB_345Cl"
#
# The available models are "GAT", "GCN", "MoNet", "GraphSage" and "GatedGCN_E_PE".
#
# A random seed of choise needs to be chosen as well (the results in the paper are obtained by averaging over 41, 42, 43 and 44).
#
# An example (to reproduce the best results for the model) is provided below:
cp.perform_collab_prediction("OGBL-COLLAB_5Cl", "GCN", 43)
| COLLAB_Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Export an ImageCollection to Google Drive
import ee
ee.Initialize()
import geetools
import ipygee as ui
# ## Define an ImageCollection
site = ee.Geometry.Point([-72, -42]).buffer(1000)
collection = ee.ImageCollection("LANDSAT/LC08/C01/T1_SR").filterBounds(site).limit(5)
# ## Set parameters
help(geetools.batch.Export.imagecollection.toDrive)
bands = ['B2', 'B3', 'B4']
scale = 30
name_pattern = '{sat}_{system_date}_{WRS_PATH:%d}-{WRS_ROW:%d}'
date_pattern = 'ddMMMy' # dd: day, MMM: month (JAN), y: year
folder = 'MYFOLDER'
data_type = 'uint32'
extra = dict(sat='L8SR')
region = site
# ## Export
tasks = geetools.batch.Export.imagecollection.toDrive(
collection=collection,
folder=folder,
region=site,
namePattern=name_pattern,
scale=scale,
dataType=data_type,
datePattern=date_pattern,
extra=extra,
verbose=True,
maxPixels=int(1e13)
)
| notebooks/batch/ImageCollectionToDrive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Question answering with TensorFlow
# ## Using advanced neural networks to tackle challenging natural language tasks.
#
# ##### By <NAME>
#
# A question answering (QA) system is a system designed to answer questions posed in natural language. Some QA systems draw information from a source such as text or an image in order to answer a specific question. These "sourced" systems can be partitioned into two major subcategories: open domain, in which the questions can be virtually anything, but aren't focused on specific material, and closed domain, in which the questions have concrete limitations, in that they relate to some predefined source (e.g., a provided context or a specific field, like medicine).
#
# This article will guide you through the task of creating and coding a question answering system using [TensorFlow](https://www.tensorflow.org/). We'll create a QA system that is based on a neural network, and sourced using a closed domain. In order to do this, we'll use a simplified version of a model known as a dynamic memory network (DMN), introduced by Kumar, et al, in their paper ["Ask Me Anything: Dynamic Memory Networks for Natural Language Processing."](https://arxiv.org/abs/1506.07285)
#
#
# ### Before we get started
#
# In addition to installing [TensorFlow](https://www.tensorflow.org/) version 1.2+ in Python 3, make sure you’ve installed each of the following:
# - [Jupyter](http://jupyter.org)
# - [Numpy](http://www.numpy.org)
# - [Matplotlib](http://matplotlib.org)
#
# Optionally, you can install TQDM to view training progess and get training speed metrics, but it's not required. The code and Jupyter Notebook for this article is [on GitHub](https://github.com/Steven-Hewitt/QA-with-Tensorflow), and I encourage you to grab it and follow along. If this is your first time working with TensorFlow, I recommend that you first check out <NAME>'s ["Hello, TensorFlow"](https://www.oreilly.com/learning/hello-tensorflow) for a quick overview of what TensorFlow is and how it works. If this is your first time using TensorFlow for natural language tasks, I would also encourage you to check out ["Textual Entailment with TensorFlow"](#link not live yet), as it introduces several concepts that will be used to help construct this network.
#
# Let's start by importing all of the relevant libraries:
# +
# %matplotlib inline
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import urllib
import sys
import os
import zipfile
import tarfile
import json
import hashlib
import re
import itertools
# -
# ### Exploring bAbI
#
# For this project, we will be using the [bAbI data set](https://research.fb.com/downloads/babi) created by Facebook. This data set, like all QA data sets, contains questions. Questions in bAbI are very straightforward, although some are trickier than others. All of the questions in this data set have an associated context, which is a sequence of sentences guaranteed to have the details necessary to answer the question. In addition, the data set provides the correct answer to each question.
#
# Questions in the bAbI data set are partitioned into 20 different tasks based on what skills are required to answer the question. Each task has its own set of questions for training, and a separate set for testing. These tasks test a variety of standard natural language processing abilities, including time reasoning (task #14) and inductive logic (task #16). To get a better idea of this, let's consider a concrete example of a question that our QA system will be expected to answer, as shown in Figure 1.
#
# <img src="questionex.png">
# <center><em>Figure 1.</em> An example of bAbI's data, with the context in blue, question in gold, and answer in green. Credit: <NAME>.</center>
#
# This task (#5) tests the network's understanding of actions where there are relationships between three objects. Grammatically speaking, it tests to see if the system can distinguish between the subject, direct object, and indirect object. In this case, the question asks for the indirect object in the last sentence -- the person who received the milk from Jeff. The network must make the distinction between the fifth sentence, in which Bill was the subject and Jeff was the indirect object, and the sixth sentence, in which Jeff was the subject. Of course, our network doesn't receive any explicit training on what a subject or object is, and has to extrapolate this understanding from the examples in the training data.
#
# Another minor problem the system must solve is understanding the various synonyms used throughout the data set. Jeff "handed" the milk to Bill, but he could have just as easily "gave" it or "passed" it to him. In this regard, though, the network doesn't have to start from scratch. It gets some assistance in the form of word vectorization, which can store infomation about the definition of words and their relations to other words. Similar words have similar vectorizations, which means that the network can treat them as nearly the same word. For word vectorization, we'll use Stanford’s Global Vectors for Word Representation (GloVe), which I’ve discussed previously in more detail [here](https://www.oreilly.com/learning/textual-entailment-with-tensorflow).
#
# Many of the tasks have a restriction that forces the context to contain the exact word used for the answer. In our example, the answer "Bill" can be found in the context. We will use this restriction to our advantage, as we can search the context for the word closest in meaning to our final result.
#
# Note: It might take a few minutes to download and unpack all of this data, so run the next three code snippets to get that started as quickly as possible. As you run the code, it will download bAbI and GloVe, and unpack the necessary files from those data sets so they can be used in our network.
# +
glove_zip_file = "glove.6B.zip"
glove_vectors_file = "glove.6B.50d.txt"
# 15 MB
data_set_zip = "tasks_1-20_v1-2.tar.gz"
#Select "task 5"
train_set_file = "qa5_three-arg-relations_train.txt"
test_set_file = "qa5_three-arg-relations_test.txt"
train_set_post_file = "tasks_1-20_v1-2/en/"+train_set_file
test_set_post_file = "tasks_1-20_v1-2/en/"+test_set_file
# -
try: from urllib.request import urlretrieve, urlopen
except ImportError:
from urllib import urlretrieve
from urllib2 import urlopen
#large file - 862 MB
if (not os.path.isfile(glove_zip_file) and
not os.path.isfile(glove_vectors_file)):
urlretrieve ("http://nlp.stanford.edu/data/glove.6B.zip",
glove_zip_file)
if (not os.path.isfile(data_set_zip) and
not (os.path.isfile(train_set_file) and os.path.isfile(test_set_file))):
urlretrieve ("https://s3.amazonaws.com/text-datasets/babi_tasks_1-20_v1-2.tar.gz",
data_set_zip)
def unzip_single_file(zip_file_name, output_file_name):
"""
If the output file is already created, don't recreate
If the output file does not exist, create it from the zipFile
"""
if not os.path.isfile(output_file_name):
with open(output_file_name, 'wb') as out_file:
with zipfile.ZipFile(zip_file_name) as zipped:
for info in zipped.infolist():
if output_file_name in info.filename:
with zipped.open(info) as requested_file:
out_file.write(requested_file.read())
return
def targz_unzip_single_file(zip_file_name, output_file_name, interior_relative_path):
if not os.path.isfile(output_file_name):
with tarfile.open(zip_file_name) as un_zipped:
un_zipped.extract(interior_relative_path+output_file_name)
unzip_single_file(glove_zip_file, glove_vectors_file)
targz_unzip_single_file(data_set_zip, train_set_file, "tasks_1-20_v1-2/en/")
targz_unzip_single_file(data_set_zip, test_set_file, "tasks_1-20_v1-2/en/")
#
#
# ### Parsing GloVe and handling unknown tokens
#
# In "[Textual Entailment with TensorFlow](https://www.oreilly.com/learning/textual-entailment-with-tensorflow)," I discuss `sentence2sequence`, which is a function that would turn a string into a matrix, based on the mapping defined by GloVe. This function split up the string into tokens, which are smaller strings that are roughly equivalent to punctuation, words, or parts of words. For example, in "Bill traveled to the kitchen," there are six tokens: five that correspond to each of the words, and the last for the period at the end. Each token gets individually vectorized, resulting in a list of vectors corresponding to each sentence, as shown in Figure 2.
#
# <img src="vectorize.png">
# <center><em>Figure 2.</em> The process of turning a sentence into vectors. Credit: <NAME>.</center>
#
# In some tasks in bAbI, the system will encounter words that are not in the GloVe word vectorization. In order for the network to be capable of processing these unknown words, we need to maintain a consistent vectorization of those words. Common practice is to replace all unknown tokens with a single `<UNK>` vector, but this isn't always effective. Instead, we can use randomization to draw a new vectorization for each unique unknown token.
#
# The first time we run across a new unknown token, we simply draw a new vectorization from the (Gaussian-approximated) distribution of the original GloVe vectorizations, and add that vectorization back to the GloVe word map . To gather the distribution hyperparameters, Numpy has functions that automatically calculate variance and mean.
#
# `fill_unk` will take care of giving us a new word vectorization whenever we need one.
# Deserialize GloVe vectors
glove_wordmap = {}
with open(glove_vectors_file, "r", encoding="utf8") as glove:
for line in glove:
name, vector = tuple(line.split(" ", 1))
glove_wordmap[name] = np.fromstring(vector, sep=" ")
# +
wvecs = []
for item in glove_wordmap.items():
wvecs.append(item[1])
s = np.vstack(wvecs)
# Gather the distribution hyperparameters
v = np.var(s,0)
m = np.mean(s,0)
RS = np.random.RandomState()
def fill_unk(unk):
global glove_wordmap
glove_wordmap[unk] = RS.multivariate_normal(m,np.diag(v))
return glove_wordmap[unk]
# -
# ### Known or unknown
#
# The limited vocabulary of bAbI tasks means the network can learn the relationships between words even without knowing what the words mean. However, for speed of learning, we should choose vectorizations that have inherent meaning when we can. To do this, we use a greedy search for words that exist in Stanford's GLoVe word vectorization data set, and if the word does not exist, then we fill in the entire word with an unknown, randomly created, new representation.
#
# Under that model of word vectorization, we can define a new `sentence2sequence`:
def sentence2sequence(sentence):
"""
- Turns an input paragraph into an (m,d) matrix,
where n is the number of tokens in the sentence
and d is the number of dimensions each word vector has.
TensorFlow doesn't need to be used here, as simply
turning the sentence into a sequence based off our
mapping does not need the computational power that
TensorFlow provides. Normal Python suffices for this task.
"""
tokens = sentence.strip('"(),-').lower().split(" ")
rows = []
words = []
#Greedy search for tokens
for token in tokens:
i = len(token)
while len(token) > 0:
word = token[:i]
if word in glove_wordmap:
rows.append(glove_wordmap[word])
words.append(word)
token = token[i:]
i = len(token)
continue
else:
i = i-1
if i == 0:
# word OOV
# https://arxiv.org/pdf/1611.01436.pdf
rows.append(fill_unk(token))
words.append(token)
break
return np.array(rows), words
# Now we can package all the data together needed for each question, including the vectorization of the contexts, questions, and answers. In bAbI, contexts are defined by a numbered sequence of sentences, which `contextualize` deserializes into a list of sentences associated with one context. Questions and answers are on the same line, separated by tabs, so we can use tabs as a marker of whether a specific line refers to a question or not. When the numbering resets, future questions will refer to the new context (note that often there is more than one question corresponding to a single context). Answers also contain one other piece of information that we keep but don't need to use: the number(s) corresponding to the sentences needed to answer the question, in the reference order. In our system, the network will teach itself which sentences are needed to answer the question.
def contextualize(set_file):
"""
Read in the dataset of questions and build question+answer -> context sets.
Output is a list of data points, each of which is a 7-element tuple containing:
The sentences in the context in vectorized form.
The sentences in the context as a list of string tokens.
The question in vectorized form.
The question as a list of string tokens.
The answer in vectorized form.
The answer as a list of string tokens.
A list of numbers for supporting statements, which is currently unused.
"""
data = []
context = []
with open(set_file, "r", encoding="utf8") as train:
for line in train:
l, ine = tuple(line.split(" ", 1))
# Split the line numbers from the sentences they refer to.
if l is "1":
# New contexts always start with 1,
# so this is a signal to reset the context.
context = []
if "\t" in ine:
# Tabs are the separator between questions and answers,
# and are not present in context statements.
question, answer, support = tuple(ine.split("\t"))
data.append((tuple(zip(*context))+
sentence2sequence(question)+
sentence2sequence(answer)+
([int(s) for s in support.split()],)))
# Multiple questions may refer to the same context, so we don't reset it.
else:
# Context sentence.
context.append(sentence2sequence(ine[:-1]))
return data
train_data = contextualize(train_set_post_file)
test_data = contextualize(test_set_post_file)
final_train_data = []
def finalize(data):
"""
Prepares data generated by contextualize() for use in the network.
"""
final_data = []
for cqas in data:
contextvs, contextws, qvs, qws, avs, aws, spt = cqas
lengths = itertools.accumulate(len(cvec) for cvec in contextvs)
context_vec = np.concatenate(contextvs)
context_words = sum(contextws,[])
# Location markers for the beginnings of new sentences.
sentence_ends = np.array(list(lengths))
final_data.append((context_vec, sentence_ends, qvs, spt, context_words, cqas, avs, aws))
return np.array(final_data)
final_train_data = finalize(train_data)
final_test_data = finalize(test_data)
# ### Defining hyperparameters
#
# At this point, we have fully prepared our training data and our testing data. The next task is to construct the network we'll use to understand the data. Let's start by clearing out the TensorFlow default graph so we always have the option to run the network again if we want to change something.
tf.reset_default_graph()
# Since this is the beginning of the actual network, let's also define all the constants we'll need for the network. We call these "hyperparameters," as they define how the network looks and trains:
# +
# Hyperparameters
# The number of dimensions used to store data passed between recurrent layers in the network.
recurrent_cell_size = 128
# The number of dimensions in our word vectorizations.
D = 50
# How quickly the network learns. Too high, and we may run into numeric instability
# or other issues.
learning_rate = 0.005
# Dropout probabilities. For a description of dropout and what these probabilities are,
# see Entailment with TensorFlow.
input_p, output_p = 0.5, 0.5
# How many questions we train on at a time.
batch_size = 128
# Number of passes in episodic memory. We'll get to this later.
passes = 4
# Feed Forward layer sizes: the number of dimensions used to store data passed from feed-forward layers.
ff_hidden_size = 256
weight_decay = 0.00000001
# The strength of our regularization. Increase to encourage sparsity in episodic memory,
# but makes training slower. Don't make this larger than leraning_rate.
training_iterations_count = 400000
# How many questions the network trains on each time it is trained.
# Some questions are counted multiple times.
display_step = 100
# How many iterations of training occur before each validation check.
# -
# ### Network structure
#
# With the hyperparameters out of the way, let's describe the network structure. The structure of this network is split loosely into four modules and is described in [Ask Me Anything: Dynamic Memory Networks for Natural Language Processing](https://arxiv.org/abs/1506.07285).
#
# The network is designed around having a recurrent layer's memory be set dynamically, based on other information in the text, hence the name dynamic memory network (DMN). DMNs are loosely based on an understanding of how a human tries to answer a reading-comprehension-type question. The person gets a chance, first of all, to read the context and create memories of the facts inside. With those facts in mind, they then read the question, and re-examine the context specifically searching for the answer to that question, comparing the question to each of the facts.
#
# Sometimes, one fact guides us to another. In the bAbI data set, the network might want to find the location of a football. It might search for sentences about the football to find that John was the last person to touch the football, then search for sentences about John to find that John had been in both the bedroom and the hallway. Once it realizes that John had been last in the hallway, it can then answer the question and confidently say that the football is in the hallway.
#
# <img src="model.png">
# <center><em>Figure 2.</em> The modules inside the network as they work together to answer a bAbI question. In each episode, new facts are attended to so they can help come up with the answer. Kumar notes that the network incorrectly puts some weight in sentence 2, which makes sense since John has been there, even though at the time he did not have the football. Credit: <NAME>, et al., used with permission.
# </center>
#
# #### Input
#
# The input module is the first of the four modules that a dynamic memory network uses to come up with its answer, and consists of a simple pass over the input with a gated recurrent unit, or GRU, (TensorFlow's [tf.contrib.nn.GRUCell](https://www.tensorflow.org/versions/r1.3/api_docs/python/tf/contrib/rnn/GRUCell)) to gather pieces of evidence. Each piece of evidence, or *fact*, corresponds to a single sentence in the context, and is represented by the output at that timestep. This requires a bit of non-TensorFlow preprocessing so we can gather the locations of the ends of sentences and pass that in to TensorFlow for use in later modules.
#
# We'll take care of that external processing later on, when we get to training. We can use that processed data with TensorFlow's `gather_nd` to select the corresponding outputs. The function `gather_nd` is an extraordinarily useful tool, and I'd suggest you review the [API documentation](https://www.tensorflow.org/versions/r1.2/api_docs/python/tf/gather_nd) to learn how it works.
# +
# Input Module
# Context: A [batch_size, maximum_context_length, word_vectorization_dimensions] tensor
# that contains all the context information.
context = tf.placeholder(tf.float32, [None, None, D], "context")
context_placeholder = context # I use context as a variable name later on
# input_sentence_endings: A [batch_size, maximum_sentence_count, 2] tensor that
# contains the locations of the ends of sentences.
input_sentence_endings = tf.placeholder(tf.int32, [None, None, 2], "sentence")
# recurrent_cell_size: the number of hidden units in recurrent layers.
input_gru = tf.contrib.rnn.GRUCell(recurrent_cell_size)
# input_p: The probability of maintaining a specific hidden input unit.
# Likewise, output_p is the probability of maintaining a specific hidden output unit.
gru_drop = tf.contrib.rnn.DropoutWrapper(input_gru, input_p, output_p)
# dynamic_rnn also returns the final internal state. We don't need that, and can
# ignore the corresponding output (_).
input_module_outputs, _ = tf.nn.dynamic_rnn(gru_drop, context, dtype=tf.float32, scope = "input_module")
# cs: the facts gathered from the context.
cs = tf.gather_nd(input_module_outputs, input_sentence_endings)
# to use every word as a fact, useful for tasks with one-sentence contexts
s = input_module_outputs
# -
# #### Question
#
# The question module is the second module, and arguably the simplest. It consists of another GRU pass, this time over the text of the question. Instead of pieces of evidence, we can simply pass forward the end state, as the question is guaranteed by the data set to be one sentence long.
# +
# Question Module
# query: A [batch_size, maximum_question_length, word_vectorization_dimensions] tensor
# that contains all of the questions.
query = tf.placeholder(tf.float32, [None, None, D], "query")
# input_query_lengths: A [batch_size, 2] tensor that contains question length information.
# input_query_lengths[:,1] has the actual lengths; input_query_lengths[:,0] is a simple range()
# so that it plays nice with gather_nd.
input_query_lengths = tf.placeholder(tf.int32, [None, 2], "query_lengths")
question_module_outputs, _ = tf.nn.dynamic_rnn(gru_drop, query, dtype=tf.float32,
scope = tf.VariableScope(True, "input_module"))
# q: the question states. A [batch_size, recurrent_cell_size] tensor.
q = tf.gather_nd(question_module_outputs, input_query_lengths)
# -
# #### Episodic memory
# Our third module, the episodic memory module, is where things begin to get interesting. It uses attention to do multiple passes, each pass consisting of GRUs iterating over the input. Each iteration inside each pass has a weighted update on current memory, based on how much attention is being paid to the corresponding fact at that time.
#
# ##### Attention
# Attention in neural networks was originally designed for image analysis, especially for cases where parts of the image are far more relevant than others. Networks use attention to determine the best locations in which to do further analysis when performing tasks, such as finding locations of objects in images, tracking objects that move between images, facial recognition, or other tasks that benefit from finding the most pertinent information for the task within the image.
#
# The main problem is that attention, or at least hard attention (which attends to exactly one input location) is not easily optimizable. As with most other neural networks, our optimization scheme is to compute the derivative of a loss function with respect to our inputs and weights, and hard attention is simply not differentiable, thanks to its binary nature. Instead, we are forced to use the real-valued version known as “soft attention,” which combines all the input locations that could be attended to using some form of weighting. Thankfully, the weighting is fully differentiable and can be trained normally. While it is possible to learn hard attention, it’s much more difficult and sometimes performs worse than soft attention. Thus, we’ll stick with soft attention for this model. Don't worry about coding the derivative; TensorFlow's optimization schemes do it for us.
#
# We calculate attention in this model by constructing similarity measures between each fact, our current memory, and the original question. (Note that this is different from normal attention, which only constructs similarity measures between facts and current memory.) We pass the results through a two-layer feed-forward network to get an attention constant for each fact. We then modify the memory by doing a weighted pass with a GRU over the input facts (weighted by the corresponding attention constant). In order to avoid adding incorrect information into memory when the context is shorter than the full length of the matrix, we create a mask for which facts exist and don't attend at all (i.e., retain the same memory) when the fact does not exist.
#
# Another notable aspect is that the attention mask is nearly always wrapped around a representation used by a layer. For images, that wrapping is most likely to happen around a convolutional layer (most likely one with a direct mapping to locations in the image), and for natural language, that wrapping is most likely to happen around a recurrent layer. Wrapping attention around a feed-forward layer, while technically possible, is usually not useful -- at least not in a way that can’t be more easily simulated by further feed-forward layers.
#
# +
# Episodic Memory
# make sure the current memory (i.e. the question vector) is broadcasted along the facts dimension
size = tf.stack([tf.constant(1),tf.shape(cs)[1], tf.constant(1)])
re_q = tf.tile(tf.reshape(q,[-1,1,recurrent_cell_size]),size)
# Final output for attention, needs to be 1 in order to create a mask
output_size = 1
# Weights and biases
attend_init = tf.random_normal_initializer(stddev=0.1)
w_1 = tf.get_variable("attend_w1", [1,recurrent_cell_size*7, recurrent_cell_size],
tf.float32, initializer = attend_init)
w_2 = tf.get_variable("attend_w2", [1,recurrent_cell_size, output_size],
tf.float32, initializer = attend_init)
b_1 = tf.get_variable("attend_b1", [1, recurrent_cell_size],
tf.float32, initializer = attend_init)
b_2 = tf.get_variable("attend_b2", [1, output_size],
tf.float32, initializer = attend_init)
# Regulate all the weights and biases
tf.add_to_collection(tf.GraphKeys.REGULARIZATION_LOSSES, tf.nn.l2_loss(w_1))
tf.add_to_collection(tf.GraphKeys.REGULARIZATION_LOSSES, tf.nn.l2_loss(b_1))
tf.add_to_collection(tf.GraphKeys.REGULARIZATION_LOSSES, tf.nn.l2_loss(w_2))
tf.add_to_collection(tf.GraphKeys.REGULARIZATION_LOSSES, tf.nn.l2_loss(b_2))
def attention(c, mem, existing_facts):
"""
Custom attention mechanism.
c: A [batch_size, maximum_sentence_count, recurrent_cell_size] tensor
that contains all the facts from the contexts.
mem: A [batch_size, maximum_sentence_count, recurrent_cell_size] tensor that
contains the current memory. It should be the same memory for all facts for accurate results.
existing_facts: A [batch_size, maximum_sentence_count, 1] tensor that
acts as a binary mask for which facts exist and which do not.
"""
with tf.variable_scope("attending") as scope:
# attending: The metrics by which we decide what to attend to.
attending = tf.concat([c, mem, re_q, c * re_q, c * mem, (c-re_q)**2, (c-mem)**2], 2)
# m1: First layer of multiplied weights for the feed-forward network.
# We tile the weights in order to manually broadcast, since tf.matmul does not
# automatically broadcast batch matrix multiplication as of TensorFlow 1.2.
m1 = tf.matmul(attending * existing_facts,
tf.tile(w_1, tf.stack([tf.shape(attending)[0],1,1]))) * existing_facts
# bias_1: A masked version of the first feed-forward layer's bias
# over only existing facts.
bias_1 = b_1 * existing_facts
# tnhan: First nonlinearity. In the original paper, this is a tanh nonlinearity;
# choosing relu was a design choice intended to avoid issues with
# low gradient magnitude when the tanh returned values close to 1 or -1.
tnhan = tf.nn.relu(m1 + bias_1)
# m2: Second layer of multiplied weights for the feed-forward network.
# Still tiling weights for the same reason described in m1's comments.
m2 = tf.matmul(tnhan, tf.tile(w_2, tf.stack([tf.shape(attending)[0],1,1])))
# bias_2: A masked version of the second feed-forward layer's bias.
bias_2 = b_2 * existing_facts
# norm_m2: A normalized version of the second layer of weights, which is used
# to help make sure the softmax nonlinearity doesn't saturate.
norm_m2 = tf.nn.l2_normalize(m2 + bias_2, -1)
# softmaxable: A hack in order to use sparse_softmax on an otherwise dense tensor.
# We make norm_m2 a sparse tensor, then make it dense again after the operation.
softmax_idx = tf.where(tf.not_equal(norm_m2, 0))[:,:-1]
softmax_gather = tf.gather_nd(norm_m2[...,0], softmax_idx)
softmax_shape = tf.shape(norm_m2, out_type=tf.int64)[:-1]
softmaxable = tf.SparseTensor(softmax_idx, softmax_gather, softmax_shape)
return tf.expand_dims(tf.sparse_tensor_to_dense(tf.sparse_softmax(softmaxable)),-1)
# facts_0s: a [batch_size, max_facts_length, 1] tensor
# whose values are 1 if the corresponding fact exists and 0 if not.
facts_0s = tf.cast(tf.count_nonzero(input_sentence_endings[:,:,-1:],-1,keep_dims=True),tf.float32)
with tf.variable_scope("Episodes") as scope:
attention_gru = tf.contrib.rnn.GRUCell(recurrent_cell_size)
# memory: A list of all tensors that are the (current or past) memory state
# of the attention mechanism.
memory = [q]
# attends: A list of all tensors that represent what the network attends to.
attends = []
for a in range(passes):
# attention mask
attend_to = attention(cs, tf.tile(tf.reshape(memory[-1],[-1,1,recurrent_cell_size]),size),
facts_0s)
# Inverse attention mask, for what's retained in the state.
retain = 1-attend_to
# GRU pass over the facts, according to the attention mask.
while_valid_index = (lambda state, index: index < tf.shape(cs)[1])
update_state = (lambda state, index: (attend_to[:,index,:] *
attention_gru(cs[:,index,:], state)[0] +
retain[:,index,:] * state))
# start loop with most recent memory and at the first index
memory.append(tuple(tf.while_loop(while_valid_index,
(lambda state, index: (update_state(state,index),index+1)),
loop_vars = [memory[-1], 0]))[0])
attends.append(attend_to)
# Reuse variables so the GRU pass uses the same variables every pass.
scope.reuse_variables()
# -
# #### Answer
# The final module is the answer module, which regresses from the question and episodic memory modules' outputs using a fully connected layer to a "final result" word vector, and the word in the context that is closest in distance to that result is our final output (to guarantee the result is an actual word). We calculate the closest word by creating a "score" for each word, which indicates the final result's distance from the word. While you can design an answer module that can return multiple words, it is not needed for the bAbI tasks we attempt in this article.
# +
# Answer Module
# a0: Final memory state. (Input to answer module)
a0 = tf.concat([memory[-1], q], -1)
# fc_init: Initializer for the final fully connected layer's weights.
fc_init = tf.random_normal_initializer(stddev=0.1)
with tf.variable_scope("answer"):
# w_answer: The final fully connected layer's weights.
w_answer = tf.get_variable("weight", [recurrent_cell_size*2, D],
tf.float32, initializer = fc_init)
# Regulate the fully connected layer's weights
tf.add_to_collection(tf.GraphKeys.REGULARIZATION_LOSSES,
tf.nn.l2_loss(w_answer))
# The regressed word. This isn't an actual word yet;
# we still have to find the closest match.
logit = tf.expand_dims(tf.matmul(a0, w_answer),1)
# Make a mask over which words exist.
with tf.variable_scope("ending"):
all_ends = tf.reshape(input_sentence_endings, [-1,2])
range_ends = tf.range(tf.shape(all_ends)[0])
ends_indices = tf.stack([all_ends[:,0],range_ends], axis=1)
ind = tf.reduce_max(tf.scatter_nd(ends_indices, all_ends[:,1],
[tf.shape(q)[0], tf.shape(all_ends)[0]]),
axis=-1)
range_ind = tf.range(tf.shape(ind)[0])
mask_ends = tf.cast(tf.scatter_nd(tf.stack([ind, range_ind], axis=1),
tf.ones_like(range_ind), [tf.reduce_max(ind)+1,
tf.shape(ind)[0]]), bool)
# A bit of a trick. With the locations of the ends of the mask (the last periods in
# each of the contexts) as 1 and the rest as 0, we can scan with exclusive or
# (starting from all 1). For each context in the batch, this will result in 1s
# up until the marker (the location of that last period) and 0s afterwards.
mask = tf.scan(tf.logical_xor,mask_ends, tf.ones_like(range_ind, dtype=bool))
# We score each possible word inversely with their Euclidean distance to the regressed word.
# The highest score (lowest distance) will correspond to the selected word.
logits = -tf.reduce_sum(tf.square(context*tf.transpose(tf.expand_dims(
tf.cast(mask, tf.float32),-1),[1,0,2]) - logit), axis=-1)
# -
# ### Optimizing optimization
#
# Gradient descent is the default optimizer for a neural network. Its goal is to decrease the network's "loss," which is a measure of how poorly the network performs. It does this by finding the derivative of loss with respect to each of the weights under the current input, and then "descends" the weights so they’ll reduce the loss. Most of the time, this works well enough, but often it’s not ideal. There are various schemes that use “momentum” or other approximations of the more direct path to the optimal weights. One of the most useful of these optimization schemes is known as adaptive moment estimation, or *Adam*.
#
# Adam estimates the first two moments of the gradient by calculating an exponentially decaying average of past iterations' gradients and squared gradients, which correspond to the estimated mean and the estimated variance of these gradients. The calculations use two additional hyperparameters to indicate how quickly the averages decay with the addition of new information. The averages are initialized as zero, which leads to bias toward zero, especially when those hyperparameters near one.
#
# In order to counteract that bias, Adam computes bias-corrected moment estimates that are greater in magnitude than the originals. The corrected estimates are then used to update the weights throughout the network. The combination of these estimates make Adam one of the best choices overall for optimization, especially for complex networks. This applies doubly to data that is very sparse, such as is common in natural language processing tasks.
#
# In TensorFlow, we can use Adam by creating a `tf.train.AdamOptimizer`.
# +
# Training
# gold_standard: The real answers.
gold_standard = tf.placeholder(tf.float32, [None, 1, D], "answer")
with tf.variable_scope('accuracy'):
eq = tf.equal(context, gold_standard)
corrbool = tf.reduce_all(eq,-1)
logloc = tf.reduce_max(logits, -1, keep_dims = True)
# locs: A boolean tensor that indicates where the score
# matches the minimum score. This happens on multiple dimensions,
# so in the off chance there's one or two indexes that match
# we make sure it matches in all indexes.
locs = tf.equal(logits, logloc)
# correctsbool: A boolean tensor that indicates for which
# words in the context the score always matches the minimum score.
correctsbool = tf.reduce_any(tf.logical_and(locs, corrbool), -1)
# corrects: A tensor that is simply correctsbool cast to floats.
corrects = tf.where(correctsbool, tf.ones_like(correctsbool, dtype=tf.float32),
tf.zeros_like(correctsbool,dtype=tf.float32))
# corr: corrects, but for the right answer instead of our selected answer.
corr = tf.where(corrbool, tf.ones_like(corrbool, dtype=tf.float32),
tf.zeros_like(corrbool,dtype=tf.float32))
with tf.variable_scope("loss"):
# Use sigmoid cross entropy as the base loss,
# with our distances as the relative probabilities. There are
# multiple correct labels, for each location of the answer word within the context.
loss = tf.nn.sigmoid_cross_entropy_with_logits(logits = tf.nn.l2_normalize(logits,-1),
labels = corr)
# Add regularization losses, weighted by weight_decay.
total_loss = tf.reduce_mean(loss) + weight_decay * tf.add_n(
tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES))
# TensorFlow's default implementation of the Adam optimizer works. We can adjust more than
# just the learning rate, but it's not necessary to find a very good optimum.
optimizer = tf.train.AdamOptimizer(learning_rate)
# Once we have an optimizer, we ask it to minimize the loss
# in order to work towards the proper training.
opt_op = optimizer.minimize(total_loss)
# +
# Initialize variables
init = tf.global_variables_initializer()
# Launch the TensorFlow session
sess = tf.Session()
sess.run(init)
# -
# ### Train the network
#
# With everything set and ready, we can begin batching our training data to train our network! While the system is training, we should check on how well the network is doing in terms of accuracy. We do this with a validation set, which is taken from testing data so it has no overlap with the training data.
#
# Using a validation set based on testing data allows us to get a better sense of how well the network can generalize what it learns and apply it to other contexts. If we validate on the training data, the network may overfit -- in other words, learn specific examples and memorize the answers to them, which doesn't help the network answer new questions.
#
# If you installed TQDM, you can use it to keep track of how long the network has been training and receive an estimate of when training will finish. You can stop the training at any time if you feel the results are good enough by interrupting the Jupyter Notebook kernel.
def prep_batch(batch_data, more_data = False):
"""
Prepare all the preproccessing that needs to be done on a batch-by-batch basis.
"""
context_vec, sentence_ends, questionvs, spt, context_words, cqas, answervs, _ = zip(*batch_data)
ends = list(sentence_ends)
maxend = max(map(len, ends))
aends = np.zeros((len(ends), maxend))
for index, i in enumerate(ends):
for indexj, x in enumerate(i):
aends[index, indexj] = x-1
new_ends = np.zeros(aends.shape+(2,))
for index, x in np.ndenumerate(aends):
new_ends[index+(0,)] = index[0]
new_ends[index+(1,)] = x
contexts = list(context_vec)
max_context_length = max([len(x) for x in contexts])
contextsize = list(np.array(contexts[0]).shape)
contextsize[0] = max_context_length
final_contexts = np.zeros([len(contexts)]+contextsize)
contexts = [np.array(x) for x in contexts]
for i, context in enumerate(contexts):
final_contexts[i,0:len(context),:] = context
max_query_length = max(len(x) for x in questionvs)
querysize = list(np.array(questionvs[0]).shape)
querysize[:1] = [len(questionvs),max_query_length]
queries = np.zeros(querysize)
querylengths = np.array(list(zip(range(len(questionvs)),[len(q)-1 for q in questionvs])))
questions = [np.array(q) for q in questionvs]
for i, question in enumerate(questions):
queries[i,0:len(question),:] = question
data = {context_placeholder: final_contexts, input_sentence_endings: new_ends,
query:queries, input_query_lengths:querylengths, gold_standard: answervs}
return (data, context_words, cqas) if more_data else data
# +
# Use TQDM if installed
tqdm_installed = False
try:
from tqdm import tqdm
tqdm_installed = True
except:
pass
# Prepare validation set
batch = np.random.randint(final_test_data.shape[0], size=batch_size*10)
batch_data = final_test_data[batch]
validation_set, val_context_words, val_cqas = prep_batch(batch_data, True)
# training_iterations_count: The number of data pieces to train on in total
# batch_size: The number of data pieces per batch
def train(iterations, batch_size):
training_iterations = range(0,iterations,batch_size)
if tqdm_installed:
# Add a progress bar if TQDM is installed
training_iterations = tqdm(training_iterations)
wordz = []
for j in training_iterations:
batch = np.random.randint(final_train_data.shape[0], size=batch_size)
batch_data = final_train_data[batch]
sess.run([opt_op], feed_dict=prep_batch(batch_data))
if (j/batch_size) % display_step == 0:
# Calculate batch accuracy
acc, ccs, tmp_loss, log, con, cor, loc = sess.run([corrects, cs, total_loss, logit,
context_placeholder,corr, locs],
feed_dict=validation_set)
# Display results
print("Iter " + str(j/batch_size) + ", Minibatch Loss= ",tmp_loss,
"Accuracy= ", np.mean(acc))
train(30000,batch_size) # Small amount of training for preliminary results
# -
# After a little bit of training, let's peek inside and see what kinds of answers we're getting from the network. In the diagrams below, we visualize attention over each of the episodes (rows) for all the sentences (columns) in our context; darker colors represent more attention paid to that sentence on that episode.
#
# You should see attention change between at least two episodes for each question, but sometimes attention will be able to find the answer within one, and sometimes it will take all four episodes. If the attention appears to be blank, it may be saturating and paying attention to everything at once. In that case, you can try training with a higher `weight_decay` in order to discourage that from happening. Later on in training, saturation becomes extremely common.
# +
ancr = sess.run([corrbool,locs, total_loss, logits, facts_0s, w_1]+attends+
[query, cs, question_module_outputs],feed_dict=validation_set)
a = ancr[0]
n = ancr[1]
cr = ancr[2]
attenders = np.array(ancr[6:-3])
faq = np.sum(ancr[4], axis=(-1,-2)) # Number of facts in each context
limit = 5
for question in range(min(limit, batch_size)):
plt.yticks(range(passes,0,-1))
plt.ylabel("Episode")
plt.xlabel("Question "+str(question+1))
pltdata = attenders[:,question,:int(faq[question]),0]
# Display only information about facts that actually exist, all others are 0
pltdata = (pltdata - pltdata.mean()) / ((pltdata.max() - pltdata.min() + 0.001)) * 256
plt.pcolor(pltdata, cmap=plt.cm.BuGn, alpha=0.7)
plt.show()
#print(list(map((lambda x: x.shape),ancr[3:])), new_ends.shape)
# -
# In order to see what the answers for the above questions were, we can use the location of our distance score in the context as an index and see what word is at that index.
# +
# Locations of responses within contexts
indices = np.argmax(n,axis=1)
# Locations of actual answers within contexts
indicesc = np.argmax(a,axis=1)
for i,e,cw, cqa in list(zip(indices, indicesc, val_context_words, val_cqas))[:limit]:
ccc = " ".join(cw)
print("TEXT: ",ccc)
print ("QUESTION: ", " ".join(cqa[3]))
print ("RESPONSE: ", cw[i], ["Correct", "Incorrect"][i!=e])
print("EXPECTED: ", cw[e])
print()
# -
# Let's keep training! In order to get good results, you may have to train for a long period of time (on my home desktop, it took about 12 hours), but you should eventually be able to reach very high accuracies (over 90%). Experienced users of Jupyter Notebook should note that at any time, you can interrupt training and still save the progress the network has made so far, as long as you keep the same `tf.Session`; this is useful if you want to visualize the attention and answers the network is currently giving.
train(training_iterations_count, batch_size)
# Final testing accuracy
print(np.mean(sess.run([corrects], feed_dict= prep_batch(final_test_data))[0]))
# Once we're done viewing what our model is returning, we can close the session to free up system resources.
sess.close()
# ### Looking for more?
#
# There's a lot still left to be done and experimented with:
#
# - *Other tasks in bAbI.* We've only sampled the many tasks that bAbI has to offer. Try changing the preprocessing to fit another task and see how our dynamic memory network performs on it. You may want to retrain the network before you try running it on the new task, of course. If the task doesn't guarantee the answer is inside the context, you may want to compare output to a dictionary of words and their corresponding vectors instead. (Those tasks are 6-10 and 17-20.) I recommend trying task 1 or 3, which you can do by changing the values of `test_set_file` and `train_set_file`.
#
# - *Supervised training.* Our attention mechanism is *unsupervised*, in that we don't specify explicitly which sentences should be attended to and instead let the network figure that out on its own. Try adding loss to the network that encourages the attention mechanism toward attending to the right sentences.
#
# - *Coattention.* Instead of attending simply over the input sentences, some researchers have found success in what they call ["dynamic coattention networks"](https://arxiv.org/pdf/1611.01604.pdf), which attends over a matrix representing two locations in two sequences simultaneously.
#
# - *Alternate vectorization schemes and sources.* Try making more intelligent mappings between sentences and vectors, or maybe use a different data set. GloVe offers larger corpi of up to eight-hundred-forty billion distinct tokens, of three hundred dimensions each.
#
#
# <p><em>This post is a collaboration between O'Reilly and </em><a href="https://www.tensorflow.org/"><em>TensorFlow</em></a><em>. </em><a href="http://www.oreilly.com/about/editorial_independence.html"><em>See our statement of editorial independence</em></a><em>.</em></p>
| TF-Keras_CHATBOT/QA with Tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda3]
# language: python
# name: conda-env-anaconda3-py
# ---
# +
# load dependencies
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler, Normalizer
from sklearn.decomposition import PCA as sklearnPCA
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.svm import SVC
import time
# +
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
cancer = pd.read_csv('breast_cancer.csv')
X_train, X_test, y_train, y_test = train_test_split(cancer.loc[:,"mean radius":"worst fractal dimension"],cancer['target'], stratify=cancer['target'], random_state=0)
svm_model = SVC()
svm_model.fit(X_train, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm_model.score(X_train, y_train)))
print('The accuracy on the testing subset: {:.3f}'.format(svm_model.score(X_test, y_test)))
# +
min_train = X_train.min(axis=0)
range_train = (X_train - min_train).max(axis=0)
X_train_scaled = (X_train - min_train)/range_train
print('Minimum per feature\n{}'.format(X_train_scaled.min(axis=0)))
print('Maximum per feature\n{}'.format(X_train_scaled.max(axis=0)))
# +
X_test_scaled = (X_test - min_train)/range_train
svm = SVC()
svm.fit(X_train_scaled, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm.score(X_train_scaled, y_train)))
print('The accuracy on the test subset: {:.3f}'.format(svm.score(X_test_scaled, y_test)))
# +
svm = SVC(C=800)
svm.fit(X_train_scaled, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm.score(X_train_scaled, y_train)))
print('The accuracy on the test subset: {:.3f}'.format(svm.score(X_test_scaled, y_test)))
# +
svm = SVC(C=1000)
svm.fit(X_train_scaled, y_train)
print('The accuracy on the training subset: {:.3f}'.format(svm.score(X_train_scaled, y_train)))
print('The accuracy on the test subset: {:.3f}'.format(svm.score(X_test_scaled, y_test)))
# -
svm
prediction = svm.predict(X_test_scaled)
prediction
# +
import matplotlib.pyplot as plt
plt.style.use('Solarize_Light2')
plt.figure(figsize = (6,4))
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
plt.figure(figsize = (8,4))
benign_count = len(prediction[prediction == 1])
malignant_count = len(prediction[prediction == 0])
objects = ('Benign','Malignent')
y_pos = np.arange(len(objects))
performance = [benign_count, malignant_count ]
plt.bar(y_pos, performance, align='center', alpha=0.5, color = ['r', 'g'])
plt.xticks(y_pos, objects)
plt.ylabel('Count')
plt.title('Predicted Results')
plt.show()
# -
predict = svm.decision_function(X_test_scaled)
predict
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
plt.figure(figsize = (6,4))
benign = list(predict[predict > 0])
malignant = list(predict[predict < 0])
malignantIndex = []
for i in malignant:
malignantIndex.append(malignant.index(i))
benignIndex = []
for i in benign:
benignIndex.append(benign.index(i))
plt.scatter(benignIndex, benign, color = "green", alpha=0.5)
plt.ylabel('Confident Value')
plt.title(f'Predicted Results (benign): Count of value: {len(benignIndex)}')
plt.show()
plt.figure(figsize = (6,4))
plt.scatter(malignantIndex,malignant , color = "Red", alpha=0.5)
plt.ylabel('Confident Value')
plt.title(f'Predicted Results (malignant): Count of value: {len(malignantIndex)}')
plt.show()
| Final_Project/Prediction/Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # PRMT-1984 [HYPOTHESIS] The spread of EMIS-EMIS pending (without error) transfers is even across GP Practices
# NB: Could this be extended to Integrated Late??
#
# ### Hypothesis
# We believe that EMIS-EMIS pending transfers are spread evenly across practices
# We will know this to be true when we see in the data that for these transfers, they are consistently spread cross the majority of practices rather than clustered in a smaller group of practices
#
#
#
# ### Subsequent questions (later hypotheses)
#
# If transfers are in a cluster of smaller practices, what do the integrations look like for those practices? I.e. are they bad at integration and therefore makes sense that they have poor pending?
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
transfer_file_location = "s3://prm-gp2gp-data-sandbox-dev/transfers-duplicates-hypothesis/"
transfer_files = [
"9-2020-transfers.parquet",
"10-2020-transfers.parquet",
"11-2020-transfers.parquet",
"12-2020-transfers.parquet",
"1-2021-transfers.parquet",
"2-2021-transfers.parquet"
]
transfer_input_files = [transfer_file_location + f for f in transfer_files]
transfers_raw = pd.concat((
pd.read_parquet(f)
for f in transfer_input_files
))
# This is only needed when using transfers-duplicates-hypothesis datasets
transfers_raw = transfers_raw.drop(["sending_supplier", "requesting_supplier"], axis=1)
asid_lookup_file = "s3://prm-gp2gp-data-sandbox-dev/asid-lookup/asidLookup-Mar-2021.csv.gz"
asid_lookup = pd.read_csv(asid_lookup_file)
# Given the findings in PRMT-1742 - many duplicate EHR errors are misclassified, the below reclassifies the relevant data
successful_transfers_bool = transfers_raw['request_completed_ack_codes'].apply(lambda x: True in [(np.isnan(i) or i==15) for i in x])
transfers = transfers_raw.copy()
transfers.loc[successful_transfers_bool, "status"] = "INTEGRATED"
transfers.loc[transfers['status']=='FAILED','sla_duration'].dropna()
pending_sender_error_codes=[6,7,10,24,30,23,14,99]
transfers_with_pending_sender_code_bool=transfers['sender_error_code'].isin(pending_sender_error_codes)
transfers_with_pending_with_error_bool=transfers['status']=='PENDING_WITH_ERROR'
transfers_which_need_pending_to_failure_change_bool=transfers_with_pending_sender_code_bool & transfers_with_pending_with_error_bool
transfers.loc[transfers_which_need_pending_to_failure_change_bool,'status']='FAILED'
eight_days_in_seconds=8*24*60*60
transfers_after_sla_bool=transfers['sla_duration']>eight_days_in_seconds
transfers_with_integrated_bool=transfers['status']=='INTEGRATED'
transfers_integrated_late_bool=transfers_after_sla_bool & transfers_with_integrated_bool
transfers.loc[transfers_integrated_late_bool,'status']='INTEGRATED LATE'
twenty_eight_days_in_seconds=28*24*60*60
transfers_after_month_bool=transfers['sla_duration']>twenty_eight_days_in_seconds
transfers_pending_at_month_bool=transfers_after_month_bool & transfers_integrated_late_bool
transfers.loc[transfers_pending_at_month_bool,'status']='PENDING'
transfers_with_early_error_bool=(~transfers.loc[:,'sender_error_code'].isna()) |(~transfers.loc[:,'intermediate_error_codes'].apply(len)>0)
transfers.loc[transfers_with_early_error_bool & transfers_pending_at_month_bool,'status']='PENDING_WITH_ERROR'
# +
# Supplier name mapping
supplier_renaming = {
"EGTON MEDICAL INFORMATION SYSTEMS LTD (EMIS)":"EMIS",
"IN PRACTICE SYSTEMS LTD":"Vision",
"MICROTEST LTD":"Microtest",
"THE PHOENIX PARTNERSHIP":"TPP",
None: "Unknown"
}
lookup = asid_lookup[["ASID", "MName", "NACS"]]
transfers = transfers.merge(lookup, left_on='requesting_practice_asid',right_on='ASID',how='left').drop("NACS", axis=1)
transfers = transfers.rename({'MName': 'requesting_supplier', 'ASID': 'requesting_supplier_asid'}, axis=1)
transfers = transfers.merge(lookup, left_on='sending_practice_asid',right_on='ASID',how='left')
transfers = transfers.rename({'MName': 'sending_supplier', 'ASID': 'sending_supplier_asid', 'NACS': 'sending_ods_code'}, axis=1)
transfers["sending_supplier"] = transfers["sending_supplier"].replace(supplier_renaming.keys(), supplier_renaming.values())
transfers["requesting_supplier"] = transfers["requesting_supplier"].replace(supplier_renaming.keys(), supplier_renaming.values())
# -
# ### Select just relevant data: EMIS-EMIS
# +
both_suppliers_EMIS_bool=(transfers['requesting_supplier']=='EMIS') & (transfers['sending_supplier']=='EMIS')
relevant_transfers=transfers.loc[both_suppliers_EMIS_bool]
# +
emis_emis_status_table=relevant_transfers['status'].value_counts()
emis_emis_status_table_pc=(100*emis_emis_status_table/emis_emis_status_table.sum()).round(2)
pd.concat([emis_emis_status_table.rename('Volume'),emis_emis_status_table_pc.rename('%')],axis=1)
# -
# ### Pivot table of practice vs status volumes
practice_status_table=pd.pivot_table(relevant_transfers,index='requesting_practice_asid',columns='status',values='conversation_id',aggfunc='count').fillna(0)
practice_status_table['TOTAL']=practice_status_table.sum(axis=1)
practice_status_pc=100*practice_status_table.div(practice_status_table['TOTAL'],axis=0).drop('TOTAL',axis=1)
practice_status_pc.columns=practice_status_pc.columns + " %"
practice_status_pc=practice_status_pc.round(2)
practice_status_table=pd.concat([practice_status_table,practice_status_pc],axis=1)
print(practice_status_table.shape[0])
print(practice_status_table['PENDING %'].median())
print(practice_status_table['PENDING %'].mean())
ax =practice_status_table['PENDING %'].plot.hist(bins=range(101),ylim=[0,50])
ax.set_xlabel('% Pending')
ax.set_ylabel('Number of GP Practices')
ax.set_title('Distribution of % transfers pending across GP Practices')
practice_status_table['PENDING %'].quantile(np.arange(0.1, 1.1, 0.1))
plt.gcf().savefig('EMIS_EMIS_pending_distribution.png')
practice_status_table.plot.scatter(x='TOTAL',y='PENDING %')
practice_status_table.plot.scatter(x='TOTAL',y='PENDING %',xlim=[0,1000])
practice_status_table.plot.scatter(x='PENDING %',y='INTEGRATED LATE %',xlim=[0,0.1])
practice_status_table[["FAILED %","INTEGRATED %","INTEGRATED LATE %","PENDING %","PENDING_WITH_ERROR %"]].corr().round(2)
practice_status_table.plot.scatter(x='PENDING %',y='FAILED %',ylim=[0,0.2])
practice_status_table.plot.scatter(x='TOTAL',y='PENDING')
practice_status_table.corr()
practice_status_table['PENDING'].max()/practice_status_table['PENDING'].sum()
practice_status_table=practice_status_table.sort_values(by='PENDING',ascending=False)
full_practice_status_table=asid_lookup.merge(practice_status_table, left_on='ASID', right_index=True, how='right')
full_practice_status_table=full_practice_status_table.drop(['MName','PName'],axis=1)
full_practice_status_table.head()
# +
def generate_gp_table(gp_asid):
gp_practice_transfers=transfers.loc[transfers['requesting_practice_asid']==gp_asid].copy()
gp_practice_transfers.loc[:,'Month']=gp_practice_transfers['date_requested'].copy().dt.to_period('M')
gp_totals=pd.pivot_table(gp_practice_transfers,index='Month',columns='status',values='conversation_id',aggfunc='count').fillna(0)
gp_percentages=pd.pivot_table(gp_practice_transfers,index='Month',columns='status',values='conversation_id',aggfunc='count').fillna(0).div(gp_practice_transfers['Month'].value_counts(),axis=0)
gp_percentages=(100*gp_percentages).round(2)
gp_percentages.columns='% '+gp_percentages.columns
return pd.concat([gp_totals, gp_percentages], axis=1)
# -
# # Break down by user profile:
#
# ## Ie is the issue related to a handful of practices or distributed?
#
# Consider for each practice, it's total number of transfers, it's outcomes
#
# - Rank practices by the total number of issues (eg Pending)
# - Break into quantiles based on number of issues
# - Identify what % of users and what % of total transfers each quantile represents
# +
practice_profile_data=practice_status_table.copy()
cumulative_percentage=practice_status_table['PENDING'].cumsum()/practice_status_table['PENDING'].sum()
quantiles=5
practice_profile_data['Quantile Group']=np.ceil(cumulative_percentage*quantiles)
practice_profile_data=practice_profile_data.groupby('Quantile Group').agg({'PENDING':'sum','TOTAL':'sum','INTEGRATED':'count'}).astype(int)
practice_profile_data=practice_profile_data.rename({'PENDING':'Total Pending','TOTAL':'Total Transfers','INTEGRATED':'Total Practices'},axis=1)
practice_profile_data_percentages=(100*practice_profile_data/practice_profile_data.sum()).round(2)
practice_profile_data_percentages.columns= "% " + practice_profile_data_percentages.columns
practice_profile_table=pd.concat([practice_profile_data,practice_profile_data_percentages],axis=1)
practice_profile_table
# -
ax=practice_status_table['PENDING'].cumsum().reset_index(drop=True).plot(figsize=(8,5))
ax.set_ylabel('Number of Pending without error Transfers')
ax.set_xlabel('Number of GP Practices')
ax.set_title('Cumulative graph of total pending transfers for GP Practices')
plt.gcf().savefig('EMIS_EMIS_cumulative_pending_transfers.jpg')
# +
def make_autopct(values):
def my_autopct(pct):
total = sum(values)
val = int(round(pct*total/100.0))
return '{p:.2f}% ({v:d})'.format(p=pct,v=val)
return my_autopct
practices_by_total_pending=practice_status_table['PENDING'].value_counts()
practices_by_total_pending.index=practices_by_total_pending.index.astype(int)
reduced_df=practices_by_total_pending.head(3)
reduced_df['3+']=practices_by_total_pending.tail(-3).sum()
reduced_df.index=reduced_df.index.astype(str) + ' pending transfers'
practices_by_total_pending=reduced_df
#practices_by_total_pending.plot.pie(autopct=make_autopct(values))
plt.pie(practices_by_total_pending.values, labels=practices_by_total_pending.index, autopct=make_autopct(practices_by_total_pending.values))
plt.gcf().set_size_inches(7,7)
plt.gca().set_title('Number of pending Transfers by % of GP Practices')
#plt.show()
plt.gcf().savefig('EMIS_EMIS_pie_number_pending_pc_practices.jpg')
# -
# # Output to Excel
# +
writer = pd.ExcelWriter('prmt-1984-emis-emis-practice-level-statuses.xlsx', engine='xlsxwriter')
practice_profile_table.to_excel(writer, sheet_name='Breakdown')
full_practice_status_table.to_excel(writer, sheet_name='Overview')
asid_of_interest=full_practice_status_table.head(10)['ASID'].values
[generate_gp_table(gp_asid).to_excel(writer,sheet_name=gp_asid) for gp_asid in asid_of_interest]
writer.save()
# -
| notebooks/26-PRMT-1984--emis-emis-practice-level-behaviour.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
# %load_ext autoreload
# %autoreload 2
import os
import runner
# +
qfib_dir = '/home/d/Dropbox/TRAKODATA/ISMRM2015/ISMRM_2015_Tracto_challenge_ground_truth_bundles_TCK_v2/'
qfib_ext = '.tck'
tko_dir = '/home/d/Dropbox/TRAKODATA/ISMRM2015/ISMRM_2015_Tracto_challenge_ground_truth_bundles_VTK_v2/'
tko_ext = '.vtk'
files = []#['CA', 'CC', 'CP']
for f in os.listdir(tko_dir):
if f.endswith('vtk'):
files.append(f.split('.')[0])
qfib_files = [(qfib_dir, f+qfib_ext) for f in files]
qfib_bits = [8, 16]
tko_files = [(tko_dir, f+tko_ext) for f in files]
tko_bits = [6,7,8,9,10,11,12,13,14]
# -
qfib_sizes = [0]*len(qfib_bits)
qfib_errors = [0]*len(qfib_bits)
qfib_stds = [0]*len(qfib_bits)
for f in qfib_files:
for i,b in enumerate(qfib_bits):
rundata = runner.Runner.qfib(f[0], f[1], bits=b, force=False)
c_time = rundata[0]
d_time = rundata[1]
sizestats = rundata[2]
compressedsize = sizestats[1]
meanerror = rundata[3][2]
stderror = rundata[3][3]
if meanerror > 2:
continue
qfib_sizes[i] += compressedsize
qfib_errors[i] += meanerror
qfib_stds[i] += stderror
# qfib_sizes.append(compressedsize)
# qfib_errors.append(meanerror)
# qfib_stds.append(stderror)
for i,b in enumerate(qfib_bits):
qfib_sizes[i] /= len(qfib_files)
qfib_errors[i] /= len(qfib_files)
qfib_stds[i] /= len(qfib_files)
tko_sizes = [0]*len(tko_bits)
tko_errors = [0]*len(tko_bits)
tko_stds = [0]*len(tko_bits)
fails = [len(tko_files)]*len(tko_bits)
for j,f in enumerate(tko_files):
for i,b in enumerate(tko_bits):
config = {
'POSITION': {
'position':True,
'sequential':True,
'quantization_bits':b,
'compression_level':10,
'quantization_range':-1,
'quantization_origin':None
},
'INDICES': {
'position':False,
'sequential':True,
'quantization_bits':b,
'compression_level':10,
'quantization_range':-1,
'quantization_origin':None
},
'name': 'qbi'+str(b)
}
try:
rundata = runner.Runner.tko(f[0], f[1], config=config, coords_only=True, force=False)
except:
fails[i] -= 1
continue
c_time = rundata[0]
d_time = rundata[1]
sizestats = rundata[2]
compressedsize = sizestats[1]
meanerror = rundata[3][2]
stderror = rundata[3][3]
tko_sizes[i] += compressedsize
tko_errors[i] += meanerror
tko_stds[i] += stderror
# tko_sizes.append(compressedsize)
# tko_errors.append(meanerror)
# tko_stds.append(stderror)
for i,b in enumerate(tko_bits):
tko_sizes[i] /= fails[i]#len(tko_files)
tko_errors[i] /= fails[i]#len(tko_files)
tko_stds[i] /= fails[i]#len(tko_files)
# +
figure(num=None, figsize=(8, 6), dpi=80, facecolor='w', edgecolor='k')
tko_sizes_ = np.array(tko_sizes) / 1000
qfib_sizes_ = np.array(qfib_sizes) / 1000
plt.scatter(tko_sizes_, tko_errors, s = np.array(tko_stds)*1000, color='blue', alpha=1, label='TRAKO')
plt.errorbar(tko_sizes_, tko_errors, tko_stds, fmt='|', color='blue', alpha=.5)
plt.scatter(qfib_sizes_, qfib_errors, s = np.array(qfib_stds)*1000, color='red', alpha=1, label='QFib')
plt.errorbar(qfib_sizes_, qfib_errors, qfib_stds, fmt='|', color='red', alpha=.5)
plt.ticklabel_format(style='plain', axis='both', scilimits=(0, 0))
import matplotlib.ticker as mticker
plt.gca().xaxis.set_major_formatter(mticker.FormatStrFormatter('%d MB'))
plt.xlabel('Data Size')
plt.ylabel('Error')
plt.legend()
plt.show()
# -
| IPY/MICCAI/SimpleStats_ISMRM2015.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="kQRtLPG5YR3O"
# # Load data from backend
# + id="0Tz2J0CfXeR7"
import requests
import json
headers = {'Cookie': '_orca_session=C1iXOgy3Ot1RBjOoS8Jlk%2Fnco4XfPEl00dvzT898KjlrQCVGyVkTg1QX5lh1Bs1Up4mTHsyGkzJ43s18envJxLmoadBxo0Cn58Y6m4YP0ru3dR9jv5mhsKHfeYxCwYwAiaFKJ%2FgkU5MW%2Fa0n8gMXJHHnn1aNvLZFPDMEMagTbAaFnXpManKfUTr%2F8sfkFkxD1S2Ev%2FeBd4ZRhcL5CumwbAqxkiRk%2F53EkSczz8d8%2FWOMAQJ%2Bz01zEsBQ1PVp1Q2h97NMbmAW1cwH2hH7xO7DIGzeS6S57HFLgH1FHC4P2W%2FNEuYoboHzqQJEkHh1qA8wtd%2BFO3%2BGpENxyDxPnl7dmtnpeHa9--Dm3Fwrgb8nKYxNBq--jJAWAAqO5%2FmcCmyDae7HzQ%3D%3D'}
UNIT_URL = "https://orca-develop.herokuapp.com/units.json"
ACTIVTY_URL = "https://orca-develop.herokuapp.com/activities.json"
# get activities from orca
r = requests.get(ACTIVTY_URL, headers=headers)
activities = r.json()
r = requests.get(UNIT_URL, headers=headers)
units = r.json()
# + [markdown] id="UxwEYaO2YP9p"
# # Class Definitions
# + id="fbzH2FRRacR6"
class Activity:
def __init__(self, id, label, category, activity_executions):
self.id = id
self.label = label
self.category = category
self.activity_executions = activity_executions
def __repr__(self):
return "id: " + str(self.id) + "\n label: " + self.label
# executions which are yet not full
def getAvailableExecutions(self, unit):
def singleMovaActivityPerDay(unit, activity_execution):
for unit_activity_execution in unit.assigned_activities_executions:
if unit_activity_execution.starts_at <= activity_execution.starts_at and \
unit_activity_execution.ends_at >= activity_execution.starts_at or \
unit_activity_execution.starts_at <= activity_execution.ends_at and \
unit_activity_execution.ends_at >= activity_execution.ends_at:
return False
return True
def noOutOfCampAcitivities(unit, activity_execution):
if unit.starts_at <= activity_execution.starts_at and unit.ends_at >= activity_execution.ends_at:
return True
else:
return False
possible_activity_executions = []
for activity_execution in self.activity_executions:
print(activity_execution.hasLanguage(unit.language))
print(activity_execution.amount_participants > unit.participant_count)
print(singleMovaActivityPerDay(unit, activity_execution))
print(noOutOfCampAcitivities(unit, activity_execution))
print(possible_activity_executions.append(activity_execution))
if activity_execution.hasLanguage(unit.language) and \
activity_execution.amount_participants > unit.participant_count and \
singleMovaActivityPerDay(unit, activity_execution) and \
noOutOfCampAcitivities(unit, activity_execution) and \
transportCapcitynotOverreached() and \ # respect transport only with ausflug
maxMovaBookingsCheck() # is activity still allowed to be assigned with respect to mova rules
possible_activity_executions.append(activity_execution)
return possible_activity_executions
# + id="TzI2N6V8am6Z"
class ActivityExecutions:
def __init__(self, id, amount_participants, starts_at, ends_at, languages, spot, transport):
self.id = id
self.amount_participants = amount_participants
self.starts_at = starts_at
self.ends_at = ends_at
self.languages = languages
self.spot = spot
self.transport = transport
self.assigned_units = []
def addUnit(self, unit):
self.assigned_units.append(unit)
def hasLanguage(self, language):
if language in self.languages:
return True
else:
return False
# + id="qRgYRy0fanwr"
class Unit:
def __init__(self, id, district, starts_at, ends_at, language, participant_count, leader_count, stufe, selected_activities):
self.id = id
self.district = district
self.starts_at = starts_at
self.ends_at = ends_at
self.language = language
self.participant_count = participant_count
self.leader_count = leader_count
self.stufe = stufe
self.selected_activities = selected_activities
self.assigned_activities_executions = []
def __repr__(self):
return "id" + str(self.id)
def getHappyScore(self):
happyScore = 0
return happyScore
def satisfied():
# mova rules
#
def getNextBooking(self):
# satisfied and next highest priority booking
for selected_activity in self.selected_activities:
if not any(x.id == selected_activity.id for x in self.assigned_activities_executions) and satisfied():
return selected_activity
return []
def addActivityExecutions(self, activity_execution):
self.assigned_activities_executions.append(activity_execution)
# + id="Wc95GD8Nc7af"
class SelectedActivity:
def __init__(self, id, priority):
self.id = id
self.priority = priority
# + id="uKjFr0lfeBb9"
class Scheduler:
def __init__(self, ):
self.acitivites = []
self.units = []
def addActivity(self, activity):
self.acitivites.append(activity)
def addUnit(self, unit):
self.units.append(unit)
def getHappyScore(self):
return sum([unit.getHappyScore() for unit in self.units]) / len(self.units)
def getActivity(self, activity_id):
for activity in self.acitivites:
if activity.id == activity_id:
return activity
def schedule(self, iterations):
# iterate, always start with "worst unit so far"
for n in range(iterations):
units_sorted = sorted(self.units, key=Unit.getHappyScore, reverse=True)
for unit_sorted in units_sorted:
# get activity which unit has selected and has not yet been assigned, get one with highest prio
selected_activity = unit_sorted.getNextBooking()
if selected_activity:
# get executions of that activity and check which execution the unit can attend based:
# - unit size
# - unit language
# - unit availability (camp date)
# - unit no overlapping other acitivities
activity = self.getActivity(selected_activity.id)
if activity:
available_executions = activity.getAvailableExecutions(unit_sorted)
if available_executions:
# assign the unit to the activity
unit.assignAcitivtyExecutions(available_executions)
available_executions.addUnit(unit_sorted)
print("Assigned")
else:
print("Assignment for activity " + str(selected_activity.id) + " and unit " + str(unit_sorted.id) + " not possible")
else:
print("Activity " + str(selected_activity.id) + " not found")
else:
print("Unit has all selected activities assigned or no activity executions")
# units which are unsatisfied, because booking possibilities didn't or didn't make any bookings
# + [markdown] id="8lhdCIZtYN05"
# # Load scheduler with data
# + id="fQeM2KANUn0u"
scheduler = Scheduler()
# create activities
for activity in activities:
act_executions = activity.get("activity_executions")
executions = []
for act_exe in act_executions:
# respect already assigend units to executions
executions.append(ActivityExecutions(
id = act_exe.get("id"),
amount_participants = act_exe.get("amount_participants"),
starts_at = act_exe.get("starts_at"),
ends_at = act_exe.get("ends_at"),
languages = act_exe.get("languages"),
spot = act_exe.get("spot"),
transport = act_exe.get("transport")))
act = Activity(activity.get("id"), activity.get("activity_category").get("code"), executions)
scheduler.addActivity(act)
# create units
for unit in units:
sel_activities = unit.get("unit_activities")
selected_activities = []
for sel_act in sel_activities:
selected_activities.append(SelectedActivity(sel_act.get("activity_id"), sel_act.get("priority")))
selected_activities = sorted(selected_activities, key=lambda selected_activity: selected_activity.priority)
un = Unit(
id = unit.get("id"),
district = unit.get("district"),
starts_at = unit.get("starts_at"),
ends_at = unit.get("ends_at"),
language = unit.get("language"),
participant_count = unit.get("participant_count"),
stufe = unit.get("stufe"),
selected_activities = selected_activities
)
scheduler.addUnit(un)
# + [markdown] id="udY_04CtYWBj"
# # Scheduling
# + colab={"base_uri": "https://localhost:8080/"} id="zyPiViLkYXtA" executionInfo={"status": "ok", "timestamp": 1633895445079, "user_tz": -120, "elapsed": 371, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhFWWmIVAhK1sY9EtdYjxL4CJsfGJ4fbr59V7AO=s64", "userId": "14424118282442031312"}} outputId="1850b56d-fc80-4545-eaa5-255729174411"
scheduler.schedule(1)
# + [markdown] id="FLTm4t9XYzAk"
# # Get Happy Score
# + colab={"base_uri": "https://localhost:8080/"} id="JZniVdRIY0kk" executionInfo={"status": "ok", "timestamp": 1633895053619, "user_tz": -120, "elapsed": 24, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhFWWmIVAhK1sY9EtdYjxL4CJsfGJ4fbr59V7AO=s64", "userId": "14424118282442031312"}} outputId="f9fee908-4894-4f05-b532-e8d9a7b11f16"
# get current assignment happy score
scheduler.getHappyScore()
| report/ActivityAssignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0.0. IMPORTS
#Importar bibliotecas
import bs4
import selenium
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
from selenium import webdriver
from bs4 import BeautifulSoup
from collections import Counter
# %matplotlib inline
from nltk.corpus import stopwords
# ## 1.0. Web Scraping
#cria um driver de conexão, abre uma nova janela do navegador
#não feche a aba
driver = webdriver.Chrome( './chromedriver' )
#criar um dataframe para receber os dados
dados = pd.DataFrame( columns = [ 'titulo','localidade', 'empresa', 'salario', 'tipo_pesquisa','desc' ] )
#abre a conexão com o site e faz a pesquisa
driver.get( 'https://br.indeed.com/jobs?q=data+science&l=Brasil' )
driver.implicitly_wait( 1 )
#gravando o resultado da pesquisa
resultado = driver.find_elements_by_class_name( 'result' )
# + code_folding=[]
#loop pelos resultados do scraping e extração dos dados nas tags
for vaga in resultado:
#Primeiro coletamos o elemento HTML interno.
result_html = vaga.get_attribute( 'innerHTML' )
#Fazemos então o parser do código HTML
soup = BeautifulSoup( result_html, 'html.parser' )
#Buscamos as tags para analise.
#usaremos blocos try/except para evitar erros na execução
#titulo da vaga
try:
title = soup.find( 'a', class_ = 'jobtitle' ).text.replace( '\n', '' )
except:
title = 'None'
#Localidade
try:
location = soup.find( class_ = 'location' ).text
except:
location = 'None'
#Empresa
try:
company = soup.find( class_ = 'company' ).text.replace( '\n', '' )
except:
location = 'None'
#Salario
try:
salary = soup.find( 'a', class_ = 'salary' ).text.replace( '\n', '' )
except:
salary = 'None'
#Tipo de pesquisa (organica ou patrocinada)
try:
sponsored = soup.find( 'a', class_ = 'sponsoredGray').text
sponsored = 'Sponsored'
except:
sponsored = 'Organic'
try:
data = soup.find( class_ = 'date' ).text
except:
location = 'None'
#Descição de vaga
job_desc = soup.find_all( name = 'div', attr={'id':'vjs-desc'} )
#job_desc = driver.find_element_by_css_selector('vjs-desc').text
#gravamos o resultado em nosso dataframe
dados = dados.append( {'titulo': title,'localidade':location,'empresa':company,'salario':salary,'tipo_pesquisa':sponsored,'date':data,'desc':job_desc}, ignore_index = True )
dados
# -
#aqui você consegue salvar o arquivo se já tiver um arqui no csv no mesmo diretorio
guardar = dados.to_csv( 'vagas1.csv' )
#aqui você consegue salvar o arquivo em qualquer diretorio e ele cria um novo arquivo
dados.to_csv ( r'C:\Users\mateu\Desktop\Web_Scraping\ teste1.csv ', index = True )
| web_scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create Simulated Data Set from Kinetic Model
# This notebook currently creates a simulated limonene data set for use in the KineticLearning.ipynb file. This file can be time consuming to run.
# ## 1. Setup & Import
# Set Parameters for Creation of Simulated DataSet
# +
# Set Parameters
kinetic_name = 'Full' # Choose Kinetic Model To Generate Data For
numStrains = 10000 # Number of Strains to Generate
use_kinetic_model_fit = False # Use the Fit Values for the Kinetic Data as starting parameters
# Set Output Time Points
timePoints = [0,2,4,6,8,10,12,14,16,18,20,24,28,32,36,40,44,48]
timePoints = list(range(70))
# measurement_noise_percent = 0.00005
# sigma = measurement_noise_percent**(0.5)
# -
# Import Required Modules
# +
import csv
import math
import numpy as np
from scipy.integrate import odeint,ode
#Ploting Utilities
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from IPython.display import display
# -
# Define Helper Functions
def odeintz(fun,y0,times):
maxDelta = 10
f = lambda t,x: fun(x,t)
r = ode(f).set_integrator('dop853',nsteps=1000,atol=1e-6)
r.set_initial_value(y0,times[0])
#progress bar
#f = FloatProgress(min=0, max=max(times))
#display(f)
#Perform Integration
x = [y0,]
curTime = times[0]
for nextTime in times[1:]:
#while r.successful() and r.t < nextTime:
while r.t < nextTime:
if nextTime-curTime < maxDelta:
dt = nextTime-curTime
else:
dt = maxDelta
value = r.integrate(r.t + dt)
curTime = r.t
#print(curTime, end='\r')
#sleep(0.001)
f.value = curTime
x.append(value)
return x
# +
#Create Differential Equation
def kineticOde(x,t,k):
def f(e1,e2,e3,e4,e5,e6,e7,e8,e9,
s1,s2,s3,s4,s5,s6,s7,s8,s9,s10,
k11,k12,k13,k21,k22,k23,k24,k31,k32,k33,k34,k35,
k41,k42,k43,k44,k45,k51,k52,k61,k62,k63,k64,k65,
k71,k72,k81,k82,k83,k84,k91,k92,Vin,ks3):
r1 = Vin - (k11*e1*s1)/(k12 + k13*s1) - k21*e2*s1*s2*ks3 / (k22*s2 + k23*s1 + k24*s1*s2)
r2 = (k11*e1*s1)/(k12 + k13*s1) - k21*e2*s1*s2*ks3 / (k22*s2 + k23*s1 + k24*s1*s2)
r3 = k21*e2*s1*s2*ks3 / (k22*s2 + k23*s1 + k24*s1*s2) - k31*e3*s3 / (k32*s1 + k33*s2 + k34*s3 + k35)
r4 = k31*e3*s3 / (k32*s1 + k33*s2 + k34*s3 + k35) - k41*e4*s4 / (k42*s9 + k43*s5 + k44*s4 + k45)
r5 = k41*e4*s4 / (k42*s9 + k43*s5 + k44*s4 + k45) - k51*e5*s5 / (k52 + s5)
r6 = k51*e5*s5 / (k52 + s5) - k61*e6*s6 / (k62*s5 + k63*s4 + k64*s6 + k65)
r7 = k61*e6*s6 / (k62*s5 + k63*s4 + k64*s6 + k65) - k71*e7*s7 / (k72 + s7) - k81*e8*s7*s8 / (k82 + k83*s7 + k84*s8 + s7*s8)
r8 = k71*e7*s7 / (k72 + s7) - k81*e8*s7*s8 / (k82 + k83*s7 + k84*s8 + s7*s8)
r9 = k81*e8*s7*s8 / (k82 + k83*s7 + k84*s8 + s7*s8) - k91*e9*s9 / (k92 + s9)
r10 = k91*e9*s9 / (k92 + s9)
return [r1,r2,r3,r4,r5,r6,r7,r8,r9,r10]
#print('len of param vec',len(k))
#print('KineticCoefficients',k[3*9:],len(k[3*9:]))
#print('Proteomics',k[:3*9],len(k[:3*9]))
ke = k[:3*9]
kr = k[3*9:]
e = proteomicsData(t,ke)
#print(len(e),len(x),len(kr))
return f(*e,*x,*kr)
def kinetic_model(e1,e2,e3,e4,e5,e6,e7,e8,e9,
s1,s2,s3,s4,s5,s6,s7,s8,s9,s10,
k11,k12,k13,k21,k22,k23,k24,k31,k32,k33,k34,k35,
k41,k42,k43,k44,k45,k51,k52,k61,k62,k63,k64,k65,
k71,k72,k81,k82,k83,k84,k91,k92,Vin,ks3):
r1 = Vin - (k11*e1*s1)/(k12 + k13*s1) - k21*e2*s1*s2*ks3 / (k22*s2 + k23*s1 + k24*s1*s2)
r2 = (k11*e1*s1)/(k12 + k13*s1) - k21*e2*s1*s2*ks3 / (k22*s2 + k23*s1 + k24*s1*s2)
r3 = k21*e2*s1*s2*ks3 / (k22*s2 + k23*s1 + k24*s1*s2) - k31*e3*s3 / (k32*s1 + k33*s2 + k34*s3 + k35)
r4 = k31*e3*s3 / (k32*s1 + k33*s2 + k34*s3 + k35) - k41*e4*s4 / (k42*s9 + k43*s5 + k44*s4 + k45)
r5 = k41*e4*s4 / (k42*s9 + k43*s5 + k44*s4 + k45) - k51*e5*s5 / (k52 + s5)
r6 = k51*e5*s5 / (k52 + s5) - k61*e6*s6 / (k62*s5 + k63*s4 + k64*s6 + k65)
r7 = k61*e6*s6 / (k62*s5 + k63*s4 + k64*s6 + k65) - k71*e7*s7 / (k72 + s7) - k81*e8*s7*s8 / (k82 + k83*s7 + k84*s8 + s7*s8)
r8 = k71*e7*s7 / (k72 + s7) - k81*e8*s7*s8 / (k82 + k83*s7 + k84*s8 + s7*s8)
r9 = k81*e8*s7*s8 / (k82 + k83*s7 + k84*s8 + s7*s8) - k91*e9*s9 / (k92 + s9)
r10 = k91*e9*s9 / (k92 + s9)
return [r1,r2,r3,r4,r5,r6,r7,r8,r9,r10]
def simple_kinetics(x,t,k):
#Parameters
def f(e,s,k):
e0,e1 = e
s0,s1,s2 = s
v_in,v_out,K_cat,K_s,K_i,K_cat1,K_m = k
v_0 = K_cat*e0*(s0/K_s)/(1 + (s0/K_s) + (s2/K_i))
v_1 = K_cat1*e1*s1/(K_m)
r1 = v_in - v_0
r2 = v_0 - v_1
r3 = v_1 - v_in*s2
return [r1,r2,r3]
e = proteomicsData(t,k[0:6])
return f(e,x,k[6:])
def proteomicsData(t,k):
e = []
for i in range(int(len(k)/3)):
#Sorting the gains to ensure proteins only increase
#gains = sorted(k[3*i:3*(i+1)],reverse=True)
gains = k[3*i:3*(i+1)]
e.append(leaky_hill_fcn(t,*gains))
return e
def leaky_hill_fcn(t,kf,km,kl):
return kf*t/(km + t) + kl
def applyMeasurementNoise(data,noise):
return [[value + np.random.normal(scale=(noise*value)**(0.5)) for value in line] for line in data]
# -
# ## 2. Extract Protein Time Series Curves From Real Data and Fit to Hill Function
# +
if kinetic_name == 'Small':
kinetic_fcn = simple_kinetics
proteins = 2
parameters = 7
substrates = 3
#CSV Header
CSV = [['Strain','Time','e0','e1','s0','s1','s2']]
elif kinetic_name == 'Full':
kinetic_fcn = kineticOde
proteins = 9
substrates = 10
parameters = 34
#CSV Header
CSV = [['Strain','Time','AtoB','HMGS','HMGR','MK','PMK','PMD','Idi','GPPS','LS',
'Acetyl-CoA','Acetoacetyl-CoA','HMG-CoA','Mev','MevP','MevPP','IPP','DMAPP','GPP','Limonene']]
# -
# A Few bits of code to help understand the parameters...
#
# **Kinetic Fit:**
# [ 5.27715270e+08 4.27024131e+02 5.23121317e+08 3.49139373e+08
# 1.53108505e+03 5.01679469e+01 6.65986265e+08 9.96515901e+08
# 1.65236177e+01 2.87429118e+08 4.59767573e+08 1.00320436e+08
# 2.55724875e+08 6.32539506e+06 7.12783803e+08 9.60680172e+07
# 4.13852542e+08 3.18329880e+04 5.52989871e+08 2.47237593e+08
# 7.17889114e+08 7.50755996e+08 1.00000000e-12 6.48850195e+08
# 1.42191830e+01 3.33891377e+08 5.85448828e+08 4.55254785e+08
# 1.01295189e+08 2.76330022e+08 6.90161521e+08 4.98233858e+02
# 8.63101820e+07 9.65536186e+08 8.07997571e+08 1.06576294e-04
# 6.27655014e+08 2.28120610e-02 2.91790555e-01]
# +
def cost_fcn(free_params):
cost = 0
for x_val,y_vals in zip(X,y):
params = []
params.extend(x_val[0:6]) # AtoB to PMD Values
params.extend(free_params[0:2]) # Keep Constant GPPS and IDI levels as free parameters
params.extend(x_val[6:8]) # LS and Acetyl-CoA
params.append(free_params[2]) # AcetoAcetyl-CoA as a free Param
params.extend(x_val[8:11]) # HMG-CoA & Mev & MevP measured
params.append(free_params[3]) #MevPP
params.extend([x_val[11],x_val[11]]) #DMAPP & IDI Measured
params.extend([free_params[4],x_val[12]]) #GPP as a Free Parameter #Measured Limonene Synthase
params.extend(free_params[5:]) # Remaining Kinetic Free Parameters
mp = kinetic_model(*params)
prediction = [mp[0],mp[2],mp[3],mp[4],mp[6]+mp[7],mp[9]]
cost += sum([(fx_val - y_val)**2 for fx_val,y_val in zip(prediction,y_vals)])
return cost
#Figure out what Fit Value is associated with what kinetic model parameter Parameter
#kinetic_fit = [5.27715270e+08, # GPPS Enzyme
# 4.27024131e+02, # IDI Enzyme
# 5.23121317e+08, # AcetoAcetyl-CoA
# 3.49139373e+08, # MevPP
# 1.53108505e+03, # GPP
# 5.01679469e+01, # Remaining Kinetic Parameters
# 6.65986265e+08,
# 9.96515901e+08,
# 1.65236177e+01 2.87429118e+08 4.59767573e+08 1.00320436e+08 2.55724875e+08 6.32539506e+06 7.12783803e+08 9.60680172e+07 4.13852542e+08 3.18329880e+04 5.52989871e+08 2.47237593e+08 7.17889114e+08 7.50755996e+08 1.00000000e-12 6.48850195e+08 1.42191830e+01 3.33891377e+08 5.85448828e+08 4.55254785e+08 1.01295189e+08 2.76330022e+08 6.90161521e+08 4.98233858e+02 8.63101820e+07 9.65536186e+08 8.07997571e+08 1.06576294e-04 6.27655014e+08 2.28120610e-02 2.91790555e-01]
fit_kinetic_parameters = [7.51679469e+01, 6.65986265e+08, 9.96515901e+08, #ATOB Params
1.65236177e+02, 2.87429118e+08, 4.59767573e+08, 1.00320436e+08, #HMGS PArams
4.55724875e+06, 6.32539506e+06, 7.12783803e+08, 9.60680172e+07, 4.13852542e+08, #HMGR
3.18329880e+05, 5.52989871e+08, 2.47237593e+08,
7.17889114e+08, 7.50755996e+08, 1.00000000e1, 6.48850195e+08, 1.42191830e+01,
3.33891377e+08, 5.85448828e+08, 4.55254785e+08, 1.01295189e+08, 2.76330022e+08,
6.90161521e+08, 4.98233858e+02, 8.63101820e+07, 9.65536186e+08, 8.07997571e+08,
1.06576294e-04, 6.27655014e+08, 2.28120610e-1, 2.91790555e2]
fit_kinetic_parameters = [2096.3009981450155, 7874322774.19455, 49306379427.56338, 132500.5312217570943, 20586148840.691242, 23802328389.43773, 2087040054.0574365, 4439677505.252835, 266857338.26716882, 11636187181.118937, 2830617721.9279895, 38700521939.961815, 1994251.5089258063, 13495068871.5197, 5635309295.377245, 8823936481.806189, 33150292760.489136, 51.02190165102573, 19498380183.677044, 601.2843909320609, 15921655319.058493, 43094446860.82067, 42028337180.13576, 8642163412.320396, 4525784432.95358, 39055142600.75094, 37460.65655172646, 1681760830.1612275, 25648330078.428032, 48445445838.558846, 0.006281218505564503, 10867785463.5929, 1.321769162835513, 18.343026702997715]
#print(len(fit_kinetic_parameters))
# -
# Set Kinetic Parameters
#Create 34 reaction kinetic constants
if use_kinetic_model_fit and kinetic_name == 'Full':
kr = fit_kinetic_parameters
else:
parameter_bounds = [(1e-1,1e2)] + [(1e7,1e9)]*31 + [(1e-5,1e2)]*2
parameter_bounds = [(1e-15,1e11)]*34
low_fit = [param[0] for param in parameter_bounds]
high_fit = [param[1] for param in parameter_bounds]
kr = np.random.uniform(low=low_fit,high=high_fit,size=(parameters)).tolist()
# Set Enzyme Parameters
from scipy.optimize import differential_evolution
#Maximize Limonene Production in Fit Model by searching the Protein expression space
bounds = [(1e6,1e7),(0.1,1),(1e-10,1e1)]*9 #+ [(1e-12,1e9)]*34
#bounds = [(1e-10,1e1),(0.001,1),(1e-10,1e0)]*9
lower_bound = [bound[0] for bound in bounds]
upper_bound = [bound[1] for bound in bounds]
ke = np.random.uniform(low=lower_bound,high=upper_bound)
# Set Strain Initial Conditions
y0 = [0.2]*(substrates-1) + [0]
# Generate Fit Based on Parameter Ranges.
# +
# Generate a Set of Proteomics
from scipy.interpolate import interp1d
# Create Metabolite Standard Curves
bounds = (0,2)
num_points_range = (2,4)
standard_curves = []
for metabolite in range(10):
# Generate a Standard Curve at Random
num_points = np.random.randint(num_points_range[0],high=num_points_range[-1]+1) + 2
times = np.linspace(timePoints[0],timePoints[-1],num_points)
concs = np.random.uniform(low=bounds[0],high=bounds[1],size=num_points)
standard_curves.append(interp1d(times,concs,kind='cubic'))
num_strains = 5
X = []
Y = []
dx = timePoints[1] - timePoints[0]
for i in range(num_strains):
# Enzyme Parameters for Strain
ke = np.random.uniform(low=lower_bound,high=upper_bound)
#ke = [ke[i:i + 3] for i in range(0, len(ke), 3)]
# Generate Enzyme Profiles for Each Strain
for t in timePoints:
# Generate Protein Curves
x = proteomicsData(t,ke)
# Generate Metabolite Values at Time T
z = [(x_l + x_h)/1e7 for x_l,x_h in zip(x+[0,],[0,]+x)]
for i,curve in enumerate(standard_curves):
x.append(z[i]*curve(t))
#Each Timepoint is a new line
X.append(x)
# Compute the Derivative and append to Y
Y = [np.gradient(np.array(X)[:,i],1) for i in range(9,19)]
#plt.plot(np.array(X)[:,9])
#plt.show()
#plt.plot(Y[0])
#plt.show()
Y = np.transpose(np.array(Y))
print(Y.shape)
print(np.array(X).shape)
# Solve for Kinetic Parameters which Minimize Difference Between Parameters using data set
def cost_fcn(free_params):
cost = 0
for x_val,y_vals in zip(X,Y):
params = np.append(x_val, free_params)
mp = kinetic_model(*params)
cost += sum([(fx_val - y_val)**2 for fx_val,y_val in zip(mp,y_vals)])
return cost
# Use Differtial Evolution to find Optimal Parameters
parameter_bounds = [(1e-15,1e11)]*34
sol = differential_evolution(cost_fcn,parameter_bounds,disp=True,maxiter=10000)
print(sol.x,sol.fun)
kr = sol.x
# -
# Optimize Kinetic Parameters to Maximize "Interesting" Dynamics.
if use_kinetic_model_fit:
def cost(kr):
#Create Strain ODE
kOde = lambda x,t: kinetic_fcn(x,t,kp.tolist() + kr.tolist())
#Solve Strain OD for Final Time Point
sol = odeintz(kOde,y0,[0,0.1])
sol = list(map(list, zip(*sol)))
total_cost = 0
for ts in sol:
if max(ts) > 5:
total_cost += math.exp((5-max(ts))**2)
if min(ts) < 0.001:
total_cost += math.exp((0.001 - min(ts))**2)
print(total_cost)
return total_cost
bounds = [(param*0.001,param*100) for param in fit_kinetic_parameters]
#bounds = [(1e-12,1e9) for param in fit_kinetic_parameters]
sol = differential_evolution(cost,bounds,disp=True)
print(sol.x,sol.fun)
kr = sol.x
# ## 3. Generate Strains
# +
#high_vals = [val*10 for val in sol.x]
#What data structure for output??? - CSV, what format?
#Strain,Time,Enzyme1,Enzyme2,...,EnzymeN,Metabolite1,...,MetaboliteN
#CSV = [['Strain','Time','AtoB','HMGS','HMGR','MK','PMK','PMD','Idi','GPPS','LS',
# 'Acetyl-CoA','Acetoacetyl-CoA','HMG-CoA','Mev','MevP','MevPP','IPP','DMAPP','GPP','Limonene']]
for strain in range(numStrains):
#Create 27 Enzyme Profile Strain Parameters
if use_kinetic_model_fit and kinetic_name == 'Full':
ke = np.random.uniform(low=lower_bound,high=upper_bound).tolist()
else:
ke = np.random.uniform(low=lower_bound,high=upper_bound,size=(proteins*3)).tolist()
#Create Kinetic ODE with Strain Specific Coeficients
#print('Kinetic Coefficients',kr)
#print('Proteomics',ke)
kOde = lambda x,t: kinetic_fcn(x,t,np.append(ke, kr))
#Solve ODE
sol = odeint(kOde,y0,timePoints).tolist()
#Calculate Time Series Proteomics Data
for i,timePoint in enumerate(timePoints):
e = proteomicsData(timePoint,ke)
sol[i] = e + sol[i]
#Add Noise to Data 0 noise currently
#noisyData = applyMeasurementNoise(sol,measurement_noise_percent)
noisyData = sol
#Add Lines to CSV
lines = [[strain+1,timePoints[i]] + line for i,line in enumerate(noisyData)]
CSV = CSV + lines
# +
#Plot Distributions for each Metabolite...
df = pd.DataFrame(CSV[1:],columns=CSV[0])
#Generate Column Wise Dist Plots (With Legend...)
column = lambda table,i: [entry[i] for entry in table[1:]]
for i in range(2,len(CSV[1])):
print(df.columns[i])
try:
sns.distplot(column(CSV,i))
plt.title(CSV[0][i])
plt.show()
except:
pass
#Generate Trace Plots
unique_strains = list(set(df['Strain'].values))
species = df.columns[2:]
print(species)
for element in species:
#display(df[['Time','Strain',element]])
sns.tsplot(df,time='Time',unit='Strain',value=element,err_style='unit_traces')
plt.show()
# -
# ## 4. Write out Generated Strains to CSV File
#Print CSV to File
fileName = kinetic_name + 'd' + str(numStrains) + 'n0.csv'
with open(fileName, 'w', newline='') as csvfile:
writer = csv.writer(csvfile)
writer.writerows(CSV)
| notebooks/Create Data Set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finetuning of the pretrained Japanese BERT model
#
# Finetune the pretrained model to solve multi-class classification problems.
# This notebook requires the following objects:
# - trained sentencepiece model (model and vocab files)
# - pretraiend Japanese BERT model
#
# Dataset is livedoor ニュースコーパス in https://www.rondhuit.com/download.html.
# We make test:dev:train = 2:2:6 datasets.
# Results:
#
# - Full training data
# - BERT with SentencePiece
# ```
# precision recall f1-score support
#
# dokujo-tsushin 0.98 0.94 0.96 178
# it-life-hack 0.96 0.97 0.96 172
# kaden-channel 0.99 0.98 0.99 176
# livedoor-homme 0.98 0.88 0.93 95
# movie-enter 0.96 0.99 0.98 158
# peachy 0.94 0.98 0.96 174
# smax 0.98 0.99 0.99 167
# sports-watch 0.98 1.00 0.99 190
# topic-news 0.99 0.98 0.98 163
#
# micro avg 0.97 0.97 0.97 1473
# macro avg 0.97 0.97 0.97 1473
# weighted avg 0.97 0.97 0.97 1473
# ```
# - sklearn GradientBoostingClassifier with MeCab
# ```
# precision recall f1-score support
#
# dokujo-tsushin 0.89 0.86 0.88 178
# it-life-hack 0.91 0.90 0.91 172
# kaden-channel 0.90 0.94 0.92 176
# livedoor-homme 0.79 0.74 0.76 95
# movie-enter 0.93 0.96 0.95 158
# peachy 0.87 0.92 0.89 174
# smax 0.99 1.00 1.00 167
# sports-watch 0.93 0.98 0.96 190
# topic-news 0.96 0.86 0.91 163
#
# micro avg 0.92 0.92 0.92 1473
# macro avg 0.91 0.91 0.91 1473
# weighted avg 0.92 0.92 0.91 1473
# ```
#
# - Small training data (1/5 of full training data)
# - BERT with SentencePiece
# ```
# precision recall f1-score support
#
# dokujo-tsushin 0.97 0.87 0.92 178
# it-life-hack 0.86 0.86 0.86 172
# kaden-channel 0.95 0.94 0.95 176
# livedoor-homme 0.82 0.82 0.82 95
# movie-enter 0.97 0.99 0.98 158
# peachy 0.89 0.95 0.92 174
# smax 0.94 0.96 0.95 167
# sports-watch 0.97 0.97 0.97 190
# topic-news 0.94 0.94 0.94 163
#
# micro avg 0.93 0.93 0.93 1473
# macro avg 0.92 0.92 0.92 1473
# weighted avg 0.93 0.93 0.93 1473
# ```
# - sklearn GradientBoostingClassifier with MeCab
# ```
# precision recall f1-score support
#
# dokujo-tsushin 0.82 0.71 0.76 178
# it-life-hack 0.86 0.88 0.87 172
# kaden-channel 0.91 0.87 0.89 176
# livedoor-homme 0.67 0.63 0.65 95
# movie-enter 0.87 0.95 0.91 158
# peachy 0.70 0.78 0.73 174
# smax 1.00 1.00 1.00 167
# sports-watch 0.87 0.95 0.91 190
# topic-news 0.92 0.82 0.87 163
#
# micro avg 0.85 0.85 0.85 1473
# macro avg 0.85 0.84 0.84 1473
# weighted avg 0.86 0.85 0.85 1473
# ```
# +
import configparser
import glob
import os
import pandas as pd
import subprocess
import sys
import tarfile
from urllib.request import urlretrieve
CURDIR = os.getcwd()
CONFIGPATH = os.path.join(CURDIR, os.pardir, 'config.ini')
config = configparser.ConfigParser()
config.read(CONFIGPATH)
# -
# ## Data preparing
#
# You need execute the following cells just once.
FILEURL = config['FINETUNING-DATA']['FILEURL']
FILEPATH = config['FINETUNING-DATA']['FILEPATH']
EXTRACTDIR = config['FINETUNING-DATA']['TEXTDIR']
# Download and unzip data.
# +
# %%time
urlretrieve(FILEURL, FILEPATH)
mode = "r:gz"
tar = tarfile.open(FILEPATH, mode)
tar.extractall(EXTRACTDIR)
tar.close()
# -
# Data preprocessing.
def extract_txt(filename):
with open(filename) as text_file:
# 0: URL, 1: timestamp
text = text_file.readlines()[2:]
text = [sentence.strip() for sentence in text]
text = list(filter(lambda line: line != '', text))
return ''.join(text)
# +
categories = [
name for name
in os.listdir( os.path.join(EXTRACTDIR, "text") )
if os.path.isdir( os.path.join(EXTRACTDIR, "text", name) ) ]
categories = sorted(categories)
# -
categories
table = str.maketrans({
'\n': '',
'\t': ' ',
'\r': '',
})
# +
# %%time
all_text = []
all_label = []
for cat in categories:
files = glob.glob(os.path.join(EXTRACTDIR, "text", cat, "{}*.txt".format(cat)))
files = sorted(files)
body = [ extract_txt(elem).translate(table) for elem in files ]
label = [cat] * len(body)
all_text.extend(body)
all_label.extend(label)
# -
df = pd.DataFrame({'text' : all_text, 'label' : all_label})
df.head()
df = df.sample(frac=1, random_state=23).reset_index(drop=True)
df.head()
# Save data as tsv files.
# test:dev:train = 2:2:6. To check the usability of finetuning, we also prepare sampled training data (1/5 of full training data).
# +
df[:len(df) // 5].to_csv( os.path.join(EXTRACTDIR, "test.tsv"), sep='\t', index=False)
df[len(df) // 5:len(df)*2 // 5].to_csv( os.path.join(EXTRACTDIR, "dev.tsv"), sep='\t', index=False)
df[len(df)*2 // 5:].to_csv( os.path.join(EXTRACTDIR, "train.tsv"), sep='\t', index=False)
### 1/5 of full training data.
# df[:len(df) // 5].to_csv( os.path.join(EXTRACTDIR, "test.tsv"), sep='\t', index=False)
# df[len(df) // 5:len(df)*2 // 5].to_csv( os.path.join(EXTRACTDIR, "dev.tsv"), sep='\t', index=False)
# df[len(df)*2 // 5:].sample(frac=0.2, random_state=23).to_csv( os.path.join(EXTRACTDIR, "train.tsv"), sep='\t', index=False)
# -
# ## Finetune pre-trained model
#
# It will take a lot of hours to execute the following cells on CPU environment.
# You can also use colab to recieve the power of TPU. You need to uplode the created data onto your GCS bucket.
#
# [](https://colab.research.google.com/drive/1zZH2GWe0U-7GjJ2w2duodFfEUptvHjcx)
PRETRAINED_MODEL_PATH = '../model/model.ckpt-1000000'
FINETUNE_OUTPUT_DIR = '../model/livedoor_output'
# +
# %%time
# It will take many hours on CPU environment.
# !python3 ../src/run_classifier.py \
# --task_name=livedoor \
# --do_train=true \
# --do_eval=true \
# --data_dir=../data/livedoor \
# --model_file=../model/wiki-ja.model \
# --vocab_file=../model/wiki-ja.vocab \
# --init_checkpoint={PRETRAINED_MODEL_PATH} \
# --max_seq_length=512 \
# --train_batch_size=4 \
# --learning_rate=2e-5 \
# --num_train_epochs=10 \
# --output_dir={FINETUNE_OUTPUT_DIR}
# -
# ## Predict using the finetuned model
#
# Let's predict test data using the finetuned model.
# +
import sys
sys.path.append("../src")
import tokenization_sentencepiece as tokenization
from run_classifier import LivedoorProcessor
from run_classifier import model_fn_builder
from run_classifier import file_based_input_fn_builder
from run_classifier import file_based_convert_examples_to_features
from utils import str_to_value
# +
sys.path.append("../bert")
import modeling
import optimization
import tensorflow as tf
# +
import configparser
import json
import glob
import os
import pandas as pd
import tempfile
bert_config_file = tempfile.NamedTemporaryFile(mode='w+t', encoding='utf-8', suffix='.json')
bert_config_file.write(json.dumps({k:str_to_value(v) for k,v in config['BERT-CONFIG'].items()}))
bert_config_file.seek(0)
bert_config = modeling.BertConfig.from_json_file(bert_config_file.name)
# -
output_ckpts = glob.glob("{}/model.ckpt*data*".format(FINETUNE_OUTPUT_DIR))
latest_ckpt = sorted(output_ckpts)[-1]
FINETUNED_MODEL_PATH = latest_ckpt.split('.data-00000-of-00001')[0]
class FLAGS(object):
'''Parameters.'''
def __init__(self):
self.model_file = "../model/wiki-ja.model"
self.vocab_file = "../model/wiki-ja.vocab"
self.do_lower_case = True
self.use_tpu = False
self.output_dir = "/dummy"
self.data_dir = "../data/livedoor"
self.max_seq_length = 512
self.init_checkpoint = FINETUNED_MODEL_PATH
self.predict_batch_size = 4
# The following parameters are not used in predictions.
# Just use to create RunConfig.
self.master = None
self.save_checkpoints_steps = 1
self.iterations_per_loop = 1
self.num_tpu_cores = 1
self.learning_rate = 0
self.num_warmup_steps = 0
self.num_train_steps = 0
self.train_batch_size = 0
self.eval_batch_size = 0
FLAGS = FLAGS()
processor = LivedoorProcessor()
label_list = processor.get_labels()
# +
tokenizer = tokenization.FullTokenizer(
model_file=FLAGS.model_file, vocab_file=FLAGS.vocab_file,
do_lower_case=FLAGS.do_lower_case)
tpu_cluster_resolver = None
is_per_host = tf.contrib.tpu.InputPipelineConfig.PER_HOST_V2
run_config = tf.contrib.tpu.RunConfig(
cluster=tpu_cluster_resolver,
master=FLAGS.master,
model_dir=FLAGS.output_dir,
save_checkpoints_steps=FLAGS.save_checkpoints_steps,
tpu_config=tf.contrib.tpu.TPUConfig(
iterations_per_loop=FLAGS.iterations_per_loop,
num_shards=FLAGS.num_tpu_cores,
per_host_input_for_training=is_per_host))
# +
model_fn = model_fn_builder(
bert_config=bert_config,
num_labels=len(label_list),
init_checkpoint=FLAGS.init_checkpoint,
learning_rate=FLAGS.learning_rate,
num_train_steps=FLAGS.num_train_steps,
num_warmup_steps=FLAGS.num_warmup_steps,
use_tpu=FLAGS.use_tpu,
use_one_hot_embeddings=FLAGS.use_tpu)
estimator = tf.contrib.tpu.TPUEstimator(
use_tpu=FLAGS.use_tpu,
model_fn=model_fn,
config=run_config,
train_batch_size=FLAGS.train_batch_size,
eval_batch_size=FLAGS.eval_batch_size,
predict_batch_size=FLAGS.predict_batch_size)
# +
predict_examples = processor.get_test_examples(FLAGS.data_dir)
predict_file = tempfile.NamedTemporaryFile(mode='w+t', encoding='utf-8', suffix='.tf_record')
file_based_convert_examples_to_features(predict_examples, label_list,
FLAGS.max_seq_length, tokenizer,
predict_file.name)
predict_drop_remainder = True if FLAGS.use_tpu else False
predict_input_fn = file_based_input_fn_builder(
input_file=predict_file.name,
seq_length=FLAGS.max_seq_length,
is_training=False,
drop_remainder=predict_drop_remainder)
# -
result = estimator.predict(input_fn=predict_input_fn)
# +
# %%time
# It will take a few hours on CPU environment.
result = list(result)
# -
result[:2]
# Read test data set and add prediction results.
import pandas as pd
test_df = pd.read_csv("../data/livedoor/test.tsv", sep='\t')
test_df['predict'] = [ label_list[elem['probabilities'].argmax()] for elem in result ]
test_df.head()
sum( test_df['label'] == test_df['predict'] ) / len(test_df)
# A littel more detailed check using `sklearn.metrics`.
# !pip install scikit-learn
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
print(classification_report(test_df['label'], test_df['predict']))
print(confusion_matrix(test_df['label'], test_df['predict']))
# ### Simple baseline model.
import pandas as pd
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
train_df = pd.read_csv("../data/livedoor/train.tsv", sep='\t')
dev_df = pd.read_csv("../data/livedoor/dev.tsv", sep='\t')
test_df = pd.read_csv("../data/livedoor/test.tsv", sep='\t')
# !apt-get install -q -y mecab libmecab-dev mecab-ipadic mecab-ipadic-utf8
# !pip install mecab-python3==0.7
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.ensemble import GradientBoostingClassifier
import MeCab
m = MeCab.Tagger("-Owakati")
train_dev_df = pd.concat([train_df, dev_df])
# +
train_dev_xs = train_dev_df['text'].apply(lambda x: m.parse(x))
train_dev_ys = train_dev_df['label']
test_xs = test_df['text'].apply(lambda x: m.parse(x))
test_ys = test_df['label']
# -
vectorizer = TfidfVectorizer(max_features=750)
train_dev_xs_ = vectorizer.fit_transform(train_dev_xs)
test_xs_ = vectorizer.transform(test_xs)
# The following set up is not exactly identical to that of BERT because inside Classifier it uses `train_test_split` with shuffle.
# In addition, parameters are not well tuned, however, we think it's enough to check the power of BERT.
# +
# %%time
model = GradientBoostingClassifier(n_estimators=200,
validation_fraction=len(train_df)/len(dev_df),
n_iter_no_change=5,
tol=0.01,
random_state=23)
### 1/5 of full training data.
# model = GradientBoostingClassifier(n_estimators=200,
# validation_fraction=len(dev_df)/len(train_df),
# n_iter_no_change=5,
# tol=0.01,
# random_state=23)
model.fit(train_dev_xs_, train_dev_ys)
# -
print(classification_report(test_ys, model.predict(test_xs_)))
print(confusion_matrix(test_ys, model.predict(test_xs_)))
| notebook/finetune-to-livedoor-corpus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# * [data.karta.orebro.se/](http://data.karta.orebro.se)
urlbase = "http://data.karta.orebro.se/opendataapi/v1/layers"
layer = "10" # layer number
import json, urllib3
import pandas as pd
| Öppen data orebro.se.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: fastai-example
# kernelspec:
# display_name: fastai-example
# language: python
# name: fastai-example
# ---
# ### This example illustrates how to use mlflow to track parameters, metrics, artifacts and models while training a simple MNIST model.
# + gather={"logged": 1645819857086}
import mlflow.fastai
from fastai.vision.all import (
CategoryBlock,
DataBlock,
GrandparentSplitter,
ImageBlock,
PILImage,
URLs,
)
from fastai.vision.all import cnn_learner, get_image_files, parent_label, resnet18, untar_data
import numpy as np
# -
# #### Splitting data between training and validation
# + gather={"logged": 1645819861074}
splitter = GrandparentSplitter(train_name="training", valid_name="testing")
# Prepare DataBlock which is a generic container to quickly build Datasets and DataLoaders
mnist = DataBlock(
blocks=(ImageBlock(PILImage), CategoryBlock),
get_items=get_image_files,
splitter=splitter,
get_y=parent_label,
)
# -
# #### Downloading MNIST data set and creating DataLoader from Datablock
# + gather={"logged": 1645820059423}
data_path = untar_data(URLs.MNIST)
image_files = get_image_files(data_path)
print('Data located at:', data_path)
data = mnist.dataloaders(data_path, bs=256, num_workers=0)
data.train.show_batch()
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1645820312256}
data.valid_ds[0]
# -
# ### Creating signature
# #### This helps to define what input does our model take. With this info we can easily share model without users having to ponder on what input does a model take
#
# We will register our model with a given signature.This means that our inputs will be tensors of shape (IMAGE_HEIGHT, IMAGE_WIDTH, CHANNELS). Since the model expects batches of images, the right input shape is (BATCH_SIZE, IMAGE_HEIGHT, IMAGE_WIDTH, CHANNELS).
#
# **input_schema**: Our model can handle variable image sizes.Then, we can indicate a variable input size using -1 in height and width. The same applies to batch size, since we can handle variable sizes.
#
# **output_schema**: Our model will return class probabilities. Since we have 10 classes, then the out of the model has shape (BATCH_SIZE, NUM_CLASSES) or (-1, 10).
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}}
# + gather={"logged": 1645820560513}
import numpy as np
from mlflow.models.signature import ModelSignature
from mlflow.types.schema import Schema, TensorSpec
input_schema = Schema([
TensorSpec(np.dtype(np.uint8), (-1, -1, -1, 3)),
])
output_schema = Schema([
TensorSpec(np.dtype(np.float32), (-1, 10)),
])
signature = ModelSignature(inputs=input_schema, outputs=output_schema)
# -
# ##### Enabling autologging from mlflow
# + gather={"logged": 1645577187840}
# mlflow.fastai.autolog()
# -
# ##### Create Learner model
# Once our dataset is ready, it's time to create our NN. CNN represents a very convenient way to solve Computer Vision problems, specially when combined with transfer learning. We use transfer learning with a pretrained image classification models
# + gather={"logged": 1645820565229}
learn = cnn_learner(data, resnet18)
# -
# ##### Using different loading function
# +
# %%writefile fastai_model_loader.py
import os
import numpy as np
import pandas as pd
class _FastaiModelWrapperPatch:
def __init__(self, learner):
self.learner = learner
def predict(self, dataframe):
dl = self.learner.dls.test_dl(dataframe)
preds, _ = self.learner.get_preds(dl=dl)
return pd.Series(map(np.array, preds.numpy())).to_frame('predictions')
def _load_model(path):
from fastai.learner import load_learner
return load_learner(os.path.abspath(path))
def _load_pyfunc(path):
"""
Load PyFunc implementation. Called by ``pyfunc.load_pyfunc``.
:param path: Local filesystem path to the MLflow Model with the ``fastai`` flavor.
"""
print("Model loaded from:",path)
return _FastaiModelWrapperPatch(_load_model(path))
# -
# ##### Training model
# + gather={"logged": 1645820685016}
saved_model = "/home/azureuser/.fastai/model.fastai"
with mlflow.start_run() as run:
# Train and fit with default or supplied command line arguments
learn.fit_one_cycle(1, 0.1)
learn.export(saved_model)
model = mlflow.pyfunc.log_model("model",
registered_model_name="digits_cnn_model",
data_path=saved_model,
code_path=["./fastai_model_loader.py"],
loader_module="fastai_model_loader",
signature=signature)
# -
# #### Loading trained model from the run
# + gather={"logged": 1645820720044}
model_uri = "runs:/{}/model".format(run.info.run_id)
loaded_model = mlflow.pyfunc.load_model(model_uri)
model_uri
# + [markdown] nteract={"transient": {"deleting": false}}
# #### Creating a sample batch for predcition to a rest endpoint
# + gather={"logged": 1645820722395} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
from fastai.vision.core import load_image
# sample_img = load_image(image_files[1])
sample_img_arr = np.array(data.train_ds[0][0])
sample_batch = sample_img_arr.reshape(1, sample_img.height, sample_img.width,3)
# + gather={"logged": 1645820725436} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
data.train_ds[0][0]
# -
# ##### Local prediction
# + gather={"logged": 1645820728340} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
loaded_model.predict(sample_batch)
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Running the model as a local endpoint
# + gather={"logged": 1645820748049} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
import json
import requests
inference_request = {
"instances": sample_batch.tolist()
}
# + gather={"logged": 1645820768098} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
endpoint = "http://localhost:5000/invocations"
response = requests.post(endpoint, json=inference_request)
response.json()
| examples/fastai/mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process DeepSea datset
#
# In this notebook, the DeepSea dataset is acquired and parsed to generate a smaller transcription factor dataset, consisting of CTCF, GABP, SP1, SRF, and YY1, for K562 and HepG2 celltypes. The dataset is first downloaded directly from DeepSea webserver and then custom scripts convert these into a h5py file.
import os, sys, h5py, scipy.io
import numpy as np
import subprocess as sp
# # download DeepSea dataset
# download deepsea dataset into data folder, if it does not exist
if not os.path.isdir('../../data/deepsea_train'):
print('downloading DeepSea dataset')
os.system('wget http://deepsea.princeton.edu/media/code/deepsea_train_bundle.v0.9.tar.gz -O ../data/deepsea_train_bundle.v0.9.tar.gz')
print('decompressing DeepSea dataset')
os.system('tar xzvf ../../data/deepsea_train_bundle.v0.9.tar.gz -C ../data ')
os.system('rm ../../data/deepsea_train_bundle.v0.9.tar.gz')
# # define functions
# +
def load_DeepSea_subset(filepath, class_range=range(918)):
""" function to load DeepSea's dataset of specific transcription factors specified
by class_range. The output is a h5py file with the sequences represented
as a 4D tensor for input into Lasagne/Theano convolution layers. The labels
is a 2D matrix where each row corresponds to a new sequence. """
def data_subset(y, class_range):
" gets a subset of data in the class_range"
data_index = []
for i in class_range:
index = np.where(y[:, i] == 1)[0]
data_index = np.concatenate((data_index, index), axis=0)
unique_index = np.unique(data_index)
return unique_index.astype(int)
print("loading training data")
trainmat = h5py.File(os.path.join(filepath,'train.mat'), 'r')
y_train = np.transpose(trainmat['traindata'], axes=(1,0))
index = data_subset(y_train, class_range)
y_train = y_train[:,class_range]
y_train = y_train[index,:]
X_train = np.transpose(trainmat['trainxdata'], axes=(2,1,0))
X_train = X_train[index,:,:]
X_train = X_train[:,[0,2,1,3],:]
X_train = np.expand_dims(X_train, axis=3)
train = (X_train.astype(np.int8), y_train.astype(np.int8))
print("loading validation data")
validmat = scipy.io.loadmat(os.path.join(filepath,'valid.mat'))
y_valid = np.array(validmat['validdata'])
index = data_subset(y_valid,class_range)
y_valid = y_valid[:, class_range]
y_valid = y_valid[index,:]
X_valid = np.transpose(validmat['validxdata'], axes=(0,1,2))
X_valid = X_valid[index,:,:]
X_valid = X_valid[:,[0,2,1,3],:]
X_valid = np.expand_dims(X_valid, axis=3)
valid = (X_valid.astype(np.int8), y_valid.astype(np.int8))
print("loading test data")
testmat = scipy.io.loadmat(os.path.join(filepath,'test.mat'))
y_test = np.array(testmat['testdata'])
index = data_subset(y_test,class_range)
y_test = y_test[:, class_range]
y_test = y_test[index,:]
X_test = np.transpose(testmat['testxdata'], axes=(0,1,2))
X_test = X_test[index,:,:]
X_test = X_test[:,[0,2,1,3],:]
X_test = np.expand_dims(X_test, axis=3)
test = (X_test.astype(np.int8), y_test.astype(np.int8))
return train, valid, test
def process_DeepSea_subset(train, valid, valid_percentage=0.1):
"""merge training and validation data, shuffle, and reallocate
based on 90% training and 10% cross-validation """
X_train = np.vstack([train[0], valid[0]])
Y_train = np.vstack([train[1], valid[1]])
index = np.random.permutation(X_train.shape[0])
cutoff = np.round(X_train.shape[0]*valid_percentage).astype(int)
valid = (X_train[:cutoff], Y_train[:cutoff])
train = (X_train[cutoff:], Y_train[cutoff:])
return train, valid
def save_DeepSea_subset(grp, train, valid, test):
""" save to h5py dataset """
print("saving datset")
X_train = grp.create_dataset('X_train', data=train[0], dtype='int8', compression="gzip")
Y_train = grp.create_dataset('Y_train', data=train[1], dtype='int8', compression="gzip")
X_valid = grp.create_dataset('X_valid', data=valid[0], dtype='int8', compression="gzip")
Y_valid = grp.create_dataset('Y_valid', data=valid[1], dtype='int8', compression="gzip")
X_test = grp.create_dataset('X_test', data=test[0], dtype='int8', compression="gzip")
Y_test = grp.create_dataset('Y_test', data=test[1], dtype='int8', compression="gzip")
# -
# # parse subset of DeepSea dataset
# +
core_names = ['Arid3a', 'CEBPB', 'FOSL1', 'Gabpa', 'MAFK', 'MAX',
'MEF2A', 'NFYB', 'SP1', 'SRF', 'STAT1', 'YY1']
core_index = [592, 602, 344, 345, 635, 636, 349, 642, 359, 361, 661, 369]
#core_index = [547, 602, 344, 345, 635, 636, 218, 642, 237, 238, 535, 369]
# save datasets in a hdf5 file under groups HepG2 and K562
data_path = '../../data/deepsea_train/'
# load deep sea dataset
train, valid, test = load_DeepSea_subset(data_path, class_range=core_index)
# -
print("number of training samples for each class")
np.sum(train[1], axis=0)
# # save dataset
#train, valid = process_DeepSea_subset(train, valid, valid_percentage=0.1)
with h5py.File('../../data/invivo_dataset.h5', 'w') as fout:
X_train = fout.create_dataset('X_train', data=train[0], dtype='int8', compression="gzip")
Y_train = fout.create_dataset('Y_train', data=train[1], dtype='int8', compression="gzip")
X_valid = fout.create_dataset('X_valid', data=valid[0], dtype='int8', compression="gzip")
Y_valid = fout.create_dataset('Y_valid', data=valid[1], dtype='int8', compression="gzip")
X_test = fout.create_dataset('X_test', data=test[0], dtype='int8', compression="gzip")
Y_test = fout.create_dataset('Y_test', data=test[1], dtype='int8', compression="gzip")
| code/generate_data/task2_generate_invivo_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 03 - Prequential Expanding
# Quebra o conjunto original em folders. Na primeira tentativa, pega o primeiro bloco para treino e o segundo bloco para teste. Na segunda tentativa, pega o primeiro e o segundo bloco para treino e o terceiro bloco para teste e assim vai fazendo para as demais tentativas, até chegar no último bloco. Os dados de treino vão sempre expandindo.
import pandas as pd
import numpy as np
from lightgbm import LGBMRegressor
data = pd.read_csv('data-processed/train.csv')
# ## Prequential Expanding
# +
data['block'] = np.trunc(data['era']*.1).astype(int)
data.loc[data['block'] == 12, 'block'] = 11
data['block'].value_counts().sort_index()
# +
results_val = []
for block in range(1,12):
print("Train blocks 0-{} - Validation Block {}".format(block - 1, block))
train = data[data['block'] < block]
val = data[data['block'] == block]
X_train = train.filter(regex=r'feature')
X_val = val.filter(regex=r'feature')
y_train = train['target']
y_val = val['target']
mdl = LGBMRegressor(max_depth=5, num_leaves=2**5, learning_rate=0.01, n_estimators=2000, colsample_bytree=0.1, random_state=0)
mdl.fit(X_train, y_train)
predictions = pd.Series(mdl.predict(X_val))
ranked_predictions = predictions.rank(pct=True, method="first")
correlation = np.corrcoef(y_val, ranked_predictions)[0, 1]
#print(correlation)
results_val.append(correlation)
print("Correlation {}".format(correlation))
print()
# -
np.median(results_val)
np.min(results_val)
np.max(results_val)
np.mean(results_val)
len(results_val)
# ## Prequential Expanding With Gap
# Inserimos gaps entre os blocos de treino e validação. Isso é interessante quando não temos os dados imediatamente antes dos dados que precisamos prever.
# +
results_val = []
for block in range(2, 12):
print("Train blocks 0-{} - Gap Block {} - Validation Block {}".format(block - 2, block - 1, block))
train = data[data['block'] < block-1]
val = data[data['block'] == block]
X_train = train.filter(regex=r'feature')
X_val = val.filter(regex=r'feature')
y_train = train['target']
y_val = val['target']
mdl = LGBMRegressor(max_depth=5, num_leaves=2**5, learning_rate=0.01, n_estimators=2000, colsample_bytree=0.1, random_state=0)
mdl.fit(X_train, y_train)
predictions = pd.Series(mdl.predict(X_val))
ranked_predictions = predictions.rank(pct=True, method="first")
correlation = np.corrcoef(y_val, ranked_predictions)[0, 1]
#print(correlation)
results_val.append(correlation)
print("Correlation {}".format(correlation))
print()
# -
np.median(results_val)
np.min(results_val)
np.max(results_val)
np.mean(results_val)
len(results_val)
# # Fim
| 03 - Prequential Expanding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow2_latest_p37)
# language: python
# name: conda_tensorflow2_latest_p37
# ---
# %load_ext autoreload
# %autoreload 2
print('\n')
print('=============loading external modules now=========')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
# tf.enable_eager_execution()
# print(f'tensorflow version: {tf.__version__}')
from tensorflow.keras import layers
import os, glob, math, random, time, datetime, PIL, imageio
from collections import defaultdict
from tqdm import tqdm, tqdm_notebook
from PIL import Image
from IPython import display
import xml.etree.ElementTree as ET
# print('\n')
# print('=============loading user-defined modules now=========')
# print('\n')
start_=time.time()
start=time.time()
from optimizers import *
# print(f'\t\toptimizers took {time.time()-start} s in loading')
# print('======================')
# print('\n')
start=time.time()
from losses import *
# print(f'\t\tlosses took {time.time()-start} s in loading')
# print('======================')
# print('\n')
start=time.time()
from g_model import *
# print(f'\t\tgenerator model took {time.time()-start} s in loading')
# print('======================')
# print('\n')
start=time.time()
from d_model import *
# print(f'\t\tdiscriminator model took {time.time()-start} s in loading')
# print('======================')
# print('\n')
start=time.time()
from static import *
# print(f'\t\tstatic took {time.time()-start} s in loading')
# print('======================')
# print('\n')
start=time.time()
from onboarding import *
# print(f'\t\tonboarding took {time.time()-start} s in loading')
# print('======================')
# print('\n')
start=time.time()
from generate_and_save_images import *
# print(f'\t\tgenerate_and_save_images took {time.time()-start} s in loading')
print('======================')
from train_step import *
# print(f'\tall user-defined modules took {time.time()-start_} s in loading')
tf.__version__
import keras
from keras.callbacks import TensorBoard
images_inputs, normalized_image_vectors, breeds_names = onboarding(all_images, all_breeds)
BUFFER_SIZE = 20000
BATCH_SIZE = 16
# imagesIn = tf.cast(normalized_image_vectors, 'float32')
ds = tf.data.Dataset.from_tensor_slices((normalized_image_vectors,)).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
print(ds)
from tensorflow.python.framework import ops
ops.convert_to_tensor()
# # Generative Network
#
# The purpose of a generative network is to use a deconvolution network and generate a random image from a random vector.
# Reference: https://arxiv.org/pdf/1505.04366.pdf
# +
# %%time
generator = g_model()
noise = tf.random.normal([1, 100])
generated_image = generator(noise, training=False)
print(type(generated_image))
plt.imshow((generated_image[0, :, :, 0]))
generator.predict(noise, steps=10).shape
generator_loss(generator.predict(noise, steps=10))
# -
discriminator = d_model()
decision = discriminator(generated_image)
print(decision)
discriminator
discriminator, generator
def write_log(callback, names, logs, batch_no):
for name, value in zip(names, logs):
summary = tf.Summary()
summary_value = summary.value.add()
summary_value.simple_value = value
summary_value.tag = name
callback.writer.add_summary(summary, batch_no)
callback.writer.flush()
log_path = './logs'
callback = TensorBoard(log_path)
# This method returns a helper function to compute cross entropy loss
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
checkpoint_prefix = os.path.join('./training-checkpoints-6', "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
type(imagesIn), type(ds)
# # Training Loop
# +
# Notice the use of `tf.function`
# This annotation causes the function to be "compiled".
# @tf.function
def train_step(images):
noise = tf.random.normal([BATCH_SIZE, noise_dim])
G_loss_list, D_loss_list = [], []
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise, training=True)
real_output = discriminator(images, training=True)
fake_output = discriminator(generated_images, training=True)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
print(gen_loss, disc_loss)
# G_loss_list.append(gen_loss.numpy())
# D_loss_list.append(disc_loss.numpy())
# gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
# gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
# generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
# discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
return gen_loss, disc_loss
# -
g,d = train_step(imagesIn[:3,:, :, :])
discriminator(ds)
def train(dataset, epochs):
for epoch in range(epochs):
start = time.time()
for image_batch in dataset:
train_step(image_batch)
if (epoch % 10 == 0):
display.clear_output(wait=True)
generate_and_save_images(generator,epoch + 1,seed)
# Save the model every 15 epochs
if (epoch + 1) % 15 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time for epoch {} is {} sec'.format(epoch + 1, time.time()-start))
# Generate after the final epoch
display.clear_output(wait=True)
generate_and_save_images(generator,epochs,seed)
def generate_and_save_images(model, epoch, test_input):
# Notice `training` is set to False.
# This is so all layers run in inference mode (batchnorm).
predictions = model(test_input, training=False)
fig = plt.figure(figsize=(8,8))
for i in range(predictions.shape[0]):
plt.subplot(4, 4, i+1)
plt.imshow(predictions[i, :, :, 0] * 127.5 + 127.5, cmap='gray')
plt.axis('off')
plt.savefig('../training_results_6/image_at_epoch_{:04d}.png'.format(epoch))
plt.show()
# %%time
tf.enable_eager_execution()
print('Starting training')
train_dataset=ds
EPOCHS = 12
train(train_dataset, EPOCHS)
generator.summary()
checkpoint_dir='training_checkpoints'
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
status = checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
for _ in range(10):
discriminator_optimizer.minimize() # Variables will be restored on creation.
status.assert_consumed() # Optional sanity checks.
checkpoint.save(file_prefix=checkpoint_prefix)
# +
import glob, imageio
anim_file = 'dcgan-2.gif'
with imageio.get_writer(anim_file, mode='I') as writer:
filenames = glob.glob('../iter2/image*.png')
filenames = sorted(filenames)
for filename in filenames:
image = imageio.imread(filename)
writer.append_data(image)
image = imageio.imread(filename)
writer.append_data(image)
# -
pip show protobuf
pip install --upgrade protobuf
anim_file
pwd
int(4.5)
| src/eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import pickle
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from catboost import CatBoostRegressor
# ## Загрузка данных и их разделение на обучающую и тестовые выборки
load = pd.read_csv('main_train.csv')
x = load.iloc[:, :-1]
y = load.iloc[:, -1]
x = x.iloc[:, 1:]
x = x.fillna(0)
test_size = round(y.size * 0.3)
test_data = x.sample(test_size)
train_data = x.drop(x.sample(test_size).index, axis=0)
test_y = y[test_data.index].to_numpy()
train_y = y[train_data.index].to_numpy()
# # Linear Regression
lin_reg = LinearRegression().fit(train_data, train_y)
sum(abs(np.round(lin_reg.predict(test_data)) - test_y))/ test_y.size
pkl_filename = './models/linear_regression.pkl'
with open(pkl_filename, 'wb') as file:
pickle.dump(lin_reg, file)
# # Catboost
# ### Сначала построим простую модель и оценим важность признаков
catboost_reg = CatBoostRegressor(iterations=200, loss_function='MAE', depth=10, learning_rate=0.03, l2_leaf_reg=3)
catboost_reg = catboost_reg.fit(train_data, train_y, verbose=False)
sum(abs(catboost_reg.predict(test_data) - test_y)) / test_y.size
# +
feature_importance = catboost_reg.get_feature_importance()
features = catboost_reg.feature_names_
deleted = []
for feature_id in feature_importance.argsort()[::-1]:
name = features[feature_id]
importance = feature_importance[feature_id]
if importance < 1.1:
deleted.append(name)
print(name, importance)
deleted
# -
test_data_del = test_data.drop(deleted, axis=1)
train_data_del = train_data.drop(deleted, axis=1)
# ### Воспользуемся несколькими моделями и подберем к ним гиперпараметры
# ## Catboost
grid_search = pd.DataFrame(columns=['test_mae', 'train_mae', 'learning_rate', 'depth', 'regularization'])
learning_rate = [0.01, 0.03, 0.1, 0.3]
depth = [4, 6, 8, 10, 12]
l2_leaf_reg = [1, 3, 5, 7, 9]
n = 0
for lr in learning_rate:
for d in depth:
for l in l2_leaf_reg:
print(n)
m = CatBoostRegressor(iterations=1000, loss_function='MAE', depth=d, learning_rate=lr, l2_leaf_reg=l)
m = m.fit(train_data, train_y, verbose=False)
grid_search.loc[n] = [sum(abs(m.predict(test_data) - test_y)) / test_y.size, sum(abs(m.predict(train_data) - train_y)) / train_y.size, lr, d, l]
n += 1
grid_search.head()
grid_search.sort_values(by='test_mae').head()
# ## RandomForest
grid_search_randfor = pd.DataFrame(columns=['test_mae', 'train_mae', 'estimators', 'depth'])
estimators = [100, 200, 50]
depth = [5, 8, 10, None]
n = 0
for est in estimators:
for d in depth:
print(n)
m = RandomForestRegressor(n_estimators=est, max_depth=d, criterion='mae').fit(train_data, train_y)
grid_search_randfor.loc[n] = [sum(abs(m.predict(test_data) - test_y)) / test_y.size, sum(abs(m.predict(train_data) - train_y)) / train_y.size, est, d]
n += 1
grid_search_randfor.head()
grid_search_randfor.sort_values(by='test_mae').head()
# ## Neural Net
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="iCtMBiq0WFWI" outputId="11a207e4-d4b6-45d6-a07c-e308d1fbcb3f"
grid_search_nn = pd.DataFrame(columns=['test_mae', 'train_mae', 'neurons', 'epochs', 'lr_start'])
neurons = [20, 40, 60]
epochs = [50, 100, 150, 200]
learning_rate = [0.3, 0.1, 0.03, 0.01]
n = 0
for neur in neurons:
for ep in epochs:
for ler in learning_rate:
print(n)
model = Sequential()
model.add(Dense(neur, input_shape=(train_data.shape[1],)))
model.add(Activation('relu'))
model.add(Dense(neur))
model.add(Activation('relu'))
model.add(Dense(neur))
model.add(Activation('relu'))
model.add(Dense(1, activation='relu'))
model.compile(optimizer=Adam(lr=ler), loss="mae", metrics=['accuracy'])
annealer = LearningRateScheduler(lambda x: ler * 0.95 ** x)
model.fit(train_data, train_y, batch_size=32, epochs = ep, callbacks=[annealer], validation_data=(test_data, test_y), verbose=0)
model.save('nn-' + '-'.join([str(neur), str(ep), str(ler)]))
grid_search_nn.loc[n] = [sum(abs(model.predict(test_data)[:, 0] - test_y)) / test_y.size, sum(abs(model.predict(train_data)[:, 0] - train_y)) / train_y.size, neur, ep, ler]
n += 1
grid_search_nn.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 206} colab_type="code" id="Pms_RR0fiDb0" outputId="cf7dc270-7e84-4e61-a308-0629b2719b45"
grid_search_nn.sort_values(by='test_mae').head()
# -
# Аналогично посмотрим для даты без нескольких столбцов (нейронную сеть и catboost можно не смотреть, так как у них результаты значительно хуже случайного леса)
# ## Random Forest
# + colab={"base_uri": "https://localhost:8080/", "height": 415} colab_type="code" id="rfdCBEuwWFWu" outputId="75807a4f-d345-42c3-cb38-29af65bffc79"
grid_search_randfor = pd.DataFrame(columns=['test_mae', 'train_mae', 'estimators', 'depth'])
estimators = [100, 200, 50]
depth = [5, 8, 10, None]
n = 0
for est in estimators:
for d in depth:
print(n)
m = RandomForestRegressor(n_estimators=est, max_depth=d, criterion='mae').fit(train_data_del, train_y)
grid_search_randfor.loc[n] = [sum(abs(m.predict(test_data_del) - test_y)) / test_y.size, sum(abs(m.predict(train_data_del) - train_y)) / train_y.size, est, d]
n += 1
grid_search_randfor.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 426} colab_type="code" id="kLAJuYbZnKMr" outputId="59e99a2d-f7f6-46e5-d4db-bcb1aa743665"
grid_search_randfor.sort_values(by='test_mae')
# -
# Так как качество модели ухудшилось совсем на незначительное значение, то мы можем ее спокойно использовать
# # Сохранение предсказаний
test = pd.read_csv('main_test.csv')
test = test.iloc[:, 1:]
test = test.fillna(0).drop(deleted, axis=1)
final_model = RandomForestRegressor(n_estimators=200, criterion='mae').fit(train_data_del, train_y)
pkl_filename = './models/random_forest.pkl'
with open(pkl_filename, 'wb') as file:
pickle.dump(final_model, file)
final_res = np.round(final_model.predict(test)).astype(np.int64)
pd.DataFrame(final_res, columns=['Pred_kcal']).to_csv('Pred_main_2.csv', index=False)
| 3/.ipynb_checkpoints/models-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="L6nlob9OmuDv"
# # Aprendizaje basado en Ensembles: Voting, Bagging y Boosting
# + [markdown] id="_bmHMrt1muDw"
# En este notebook vamos a ver diferentes técnicas para hacer ensemble de modelos, es decir, combinaciones de diferentes modelos para mejorar nuestras predicciones. Aquí podremos ver cómo implementar el Voting, qué es el Bagging, el Boosting; así como implementaciones de modelos basadas en estas configuraciones como Random Forest o XGBoost.
# + [markdown] id="lC4HhuNBmuDw"
# ## Configuración del entorno
# + id="MZpFS5xZmuDy"
import sys
import sklearn
import numpy as np
import os
# Semilla para poder reproducir los mismos resultados:
np.random.seed(42)
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# La función que hemos definido en otros ssitios para pintar:
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "ensembles"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# + [markdown] id="yXNzBMdZmuD1"
# ## Voting: selección de la salida por votación
#
# La técnica más utilizada para realizar la combinación de varios modelos consiste en realizar una votación. Cada modelo, entrenado con sus correspondientes datos, devolverá una predicción (o una probabilidad de la misma) y se hará la votación:
# - **Hard voting**: los clasificadores votan una respuesta, y la decisión final lo determinará la respuesta más votada.
# - **Soft voting**: está basado en las probabilidades de las respuestas de los clasificadores. Suele funcionar mejor.
# + id="DVuI23_lmuD8"
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_moons
# Creamos un dataset de clasificación con la función make_moons:
X_moons, y_moons = make_moons(n_samples=500, noise=0.30, random_state=42)
# Dividimos en train/test:
X_train, X_test, y_train, y_test = train_test_split(X_moons, y_moons, random_state=42)
# + id="DhPtKT1FmuD_"
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import VotingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
# Nos vamos a crear diferentes algoritmos:
# Regresión Logística:
logreg = LogisticRegression(solver="lbfgs", random_state=42)
# Árbol de decisión:
dt = DecisionTreeClassifier(max_depth=10, random_state=42)
# SVC:
svc = SVC(gamma="scale", random_state=42)
# Y nos creamos el objeto de Voting (Hard Voting), que se encargará de gestionar por nosotros el entrenamiento y predicción con todos los algoritmos:
voting_clf = VotingClassifier(
estimators=[('lr', logreg), ('dt', dt), ('svc', svc)], voting='hard')
# + id="MBT4z1u6muEC" outputId="5a0b9d89-8451-40c0-9877-229efb166e57"
# Etrenamos nuestro sistema de votación:
voting_clf.fit(X_train, y_train)
# + id="EZD7VxEDmuEF" outputId="0998b8e0-3825-4082-9caf-230384248505"
from sklearn.metrics import accuracy_score
# Realizamos la predicción con cada algoritmo de forma individual y con el sistema de votación de todos ellos:
for clf in (logreg, dt, svc, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, accuracy_score(y_test, y_pred))
# + id="9_OyiVkamuEI" outputId="8b166f3d-74d0-4d3b-c91a-311aefeef0c7"
# Repetimos lo mismo pero creando un sistema de votación "soft" (el resto es exactamente igual, podíamos haber usado los modelos definidos antes):
log_clf = LogisticRegression(solver="lbfgs", random_state=42)
dt_clf = DecisionTreeClassifier(random_state=42)
svm_clf = SVC(gamma="scale", probability=True, random_state=42)
voting_clf = VotingClassifier(
estimators=[('lr', log_clf), ('dt', dt_clf), ('svc', svm_clf)],
voting='soft')
voting_clf.fit(X_train, y_train)
# + id="HReL1oI3muEK" outputId="68b429e2-260f-4e9e-ef95-a4465c7c3a1f"
from sklearn.metrics import accuracy_score
for clf in (log_clf, dt_clf, svm_clf, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, f"Train: {accuracy_score(y_train, clf.predict(X_train))}", f"Test: {accuracy_score(y_test, y_pred)}")
# -
# ### EJERCICIO
#
# Por mantener cierta coherencia con lo que hemos visto en clases pasadas, vamos a repetir estos ejercicios sobre el mismo dataset, que ya lo tenemos más que conocido.
#
# 1. Prueba a combinar los algoritmos Regresión Logística, Árbol de decisión y KNN, y qué salida consigues:
import pandas as pd
df = pd.read_csv("../../../data/titanic.csv", sep='\t')
df
# +
# El que queda de Embarked, lo vamos a rellenar con "S" porque es el mayoritario.
df['Embarked'] = df['Embarked'].fillna('S')
# En Age, vamos a completar los valores nulos con la media de los valores de Age.
df['Age'] = df['Age'].fillna(df['Age'].mean())
# En Pclass, vamos a convertirlo a string para luego hacer el One Hot Encoding.
df['Pclass'] = df['Pclass'].astype(str)
# Separamos las variables que podemos utilizar para predecir de la que queremos predecir:
y_col = 'Survived'
X_cols = [col for col in df.columns if col not in ['PassengerId', 'Name', 'Ticket', 'Cabin', 'Survived']]
X = df[X_cols]
y = df[y_col]
# Tratamiento categóricas:
X = pd.get_dummies(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3)
# +
# Repetimos lo mismo pero creando un sistema de votación "soft" (el resto es exactamente igual, podíamos haber usado los modelos definidos antes):
log_clf = LogisticRegression(max_iter=2000, solver="lbfgs", random_state=42)
dt_clf = DecisionTreeClassifier(random_state=42)
# svm_clf = SVC(gamma="scale", probability=True, random_state=42)
voting_clf = VotingClassifier(
estimators=[('lr', log_clf), ('dt', dt_clf), ('svc', svm_clf)],
voting='soft')
voting_clf.fit(X_train, y_train)
# +
from sklearn.metrics import accuracy_score
for clf in (log_clf, dt_clf, svm_clf, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, f"\n\tTrain: {accuracy_score(y_train, clf.predict(X_train))}", f"\n\tTest: {accuracy_score(y_test, y_pred)}")
# + [markdown] id="rSmuM9kHmuEN"
# ## Bagging ensembles
#
# El "bagging" sería el siguiente paso, que consiste en utilizar unas cuantas instancias del mismo modelo pero entrenados con muestreos diferentes de los datos originales. Tras ello, cada predicción se hará mediante la votación de cada uno de los modelos que acabos de entrenar, de modo que cada uno pueda aportar "su punto de vista":
# -
# Antes de nada, volvemos a crearnos X_train y X_test, por si las hemos sobreescrito en el ejercicio anterior:
X = X_moons
y = y_moons
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# + id="igA2O8rxmuEN"
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
# Nos creamos el bagging con árboles de decisión:
bag_clf = BaggingClassifier(
DecisionTreeClassifier(random_state=42),
n_estimators=500,
max_samples=100,
bootstrap=True,
random_state=42)
# Lo ajustamos:
bag_clf.fit(X_train, y_train)
# Y predecimos:
y_pred = bag_clf.predict(X_test)
# + id="YjKdfEFOmuEQ" outputId="54801ad4-a87c-401f-903e-60ba02bb748c"
from sklearn.metrics import accuracy_score
# Y calculamos el accuracy:
print(accuracy_score(y_test, y_pred))
# + id="Fm72Hw8-muES" outputId="9baf9034-3b0b-4cd3-c93d-a8e1b5c5463f"
# Si lo comparamos con un solo decisor:
tree_clf = DecisionTreeClassifier(random_state=42)
tree_clf.fit(X_train, y_train)
y_pred_tree = tree_clf.predict(X_test)
print(accuracy_score(y_test, y_pred_tree))
# -
# A continuación, vamos a representar un par de figuras para entender qué está pasando, pero no hay que saberse cómo se hace esta representación. Lo importante es lo anterior: saber crearse los objetos y entender qué están haciendo.
# + id="keVwZYAAmuEV"
from matplotlib.colors import ListedColormap
def plot_decision_boundary(clf, X, y, axes=[-1.5, 2.5, -1, 1.5], alpha=0.5, contour=True):
x1s = np.linspace(axes[0], axes[1], 100)
x2s = np.linspace(axes[2], axes[3], 100)
x1, x2 = np.meshgrid(x1s, x2s)
X_new = np.c_[x1.ravel(), x2.ravel()]
y_pred = clf.predict(X_new).reshape(x1.shape)
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=custom_cmap)
if contour:
custom_cmap2 = ListedColormap(['#7d7d58','#4c4c7f','#507d50'])
plt.contour(x1, x2, y_pred, cmap=custom_cmap2, alpha=0.8)
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo", alpha=alpha)
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs", alpha=alpha)
plt.axis(axes)
plt.xlabel(r"$x_1$", fontsize=18)
plt.ylabel(r"$x_2$", fontsize=18, rotation=0)
# + id="qo1U1r4KmuEX" outputId="dbb1fda3-2ce0-45e2-b0ab-c3ab34b56991"
plt.figure(figsize=(11,4))
plt.subplot(121)
plot_decision_boundary(tree_clf, X, y)
plt.title("Decision Tree", fontsize=14)
plt.subplot(122)
plot_decision_boundary(bag_clf, X, y)
plt.title("Decision Trees con Bagging", fontsize=14)
save_fig("decision_tree_without_and_with_bagging_plot")
plt.show()
# -
# ### EJERCICIO
#
# 1. Prueba a utilizar la técnica de bagging con árboles de decisión y con regresores logísticos, variando el número de instancias utilizadas para el bagging. Prueba pocas combinaciones, como 20, 50 y 100, por ejemplo, para cada uno de los algoritmos. ¿Con cuál obtienes mejor score sobre test?
#
# Utiliza el dataset del Titanic.
import pandas as pd
df = pd.read_csv("../../../data/titanic.csv", sep='\t')
df
# +
# El que queda de Embarked, lo vamos a rellenar con "S" porque es el mayoritario.
df['Embarked'] = df['Embarked'].fillna('S')
# En Age, vamos a completar los valores nulos con la media de los valores de Age.
df['Age'] = df['Age'].fillna(df['Age'].mean())
# En Pclass, vamos a convertirlo a string para luego hacer el One Hot Encoding.
df['Pclass'] = df['Pclass'].astype(str)
# Separamos las variables que podemos utilizar para predecir de la que queremos predecir:
y_col = 'Survived'
X_cols = [col for col in df.columns if col not in ['PassengerId', 'Name', 'Ticket', 'Cabin', 'Survived']]
X = df[X_cols]
y = df[y_col]
# Tratamiento categóricas:
X = pd.get_dummies(X)
# +
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import MinMaxScaler
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
bag_clf = BaggingClassifier(
base_estimator=DecisionTreeClassifier(random_state=42),
n_estimators=500,
max_samples=100,
bootstrap=True,
random_state=42)
pipe = Pipeline([('scaler', MinMaxScaler()), ('bag', bag_clf)])
params = {
'bag__base_estimator': [LogisticRegression(max_iter=2000, C=2, solver='liblinear'), DecisionTreeClassifier(max_depth=5)],
'bag__n_estimators': [20, 50, 100]
}
gs_pipe = GridSearchCV(pipe,
params,
cv = 10,
scoring='accuracy',
n_jobs=-1,
verbose=1)
# -
gs_pipe.fit(X_train, y_train)
gs_pipe.best_estimator_.score(X_test, y_test)
gs_pipe.best_params_
X_train.shape
# Otra opción con Pipelines:
# +
from sklearn.pipeline import Pipeline
from sklearn.model_selection import GridSearchCV
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import SelectKBest
bag_clf = BaggingClassifier(
base_estimator=DecisionTreeClassifier(random_state=42),
n_estimators=500,
max_samples=100,
bootstrap=True,
random_state=42)
pipe = Pipeline([('scaler', StandardScaler()), ('selector', SelectKBest()), ('bag', bag_clf)])
params = {
'selector__k': [1]
}
gs_pipe = GridSearchCV(pipe,
params,
cv = 10,
scoring='accuracy',
n_jobs=-1,
verbose=1)
# -
gs_pipe.fit(X_train, y_train)
# gs_pipe.best_score_
gs_pipe.best_estimator_.score(X_test, y_test)
# + [markdown] id="8st00lihmuEZ"
# # Random Forests
#
# Esta fórmula, aplicada a los árboles de decisión (como acabamos de ver), y haciendo las divisiones de los árboles de decisión de manera aleatoria (y no eligiendo la mejor en cada corte), es lo que da origen al modelo Random Forest:
# + id="voEaXAS0muEa"
# Implementamos RandomForest, creando un bagging de n árboles de decisión seleccionando variables de forma aleatoria:
bag_clf = BaggingClassifier(
DecisionTreeClassifier(splitter="random", max_leaf_nodes=16, random_state=42),
n_estimators=500, max_samples=1.0, bootstrap=True, random_state=42)
# + id="dd7j8TTvmuEc"
# Entrenamos y predecimos:
bag_clf.fit(X_train, y_train)
y_pred = bag_clf.predict(X_test)
# -
bag_clf.score(X_test, y_test)
# + id="vu85ut28muEg"
from sklearn.ensemble import RandomForestClassifier
# Y ahora probaremos con el Radnom Forest de sklearn (que, pese a que se basa en la misma idea, tiene ligeras mejoras):
rnd_clf = RandomForestClassifier(n_estimators=500, max_leaf_nodes=16, random_state=42)
rnd_clf.fit(X_train, y_train)
y_pred_rf = rnd_clf.predict(X_test)
# -
rnd_clf.score(X_test, y_test)
# + [markdown] id="Lei_Kjr7muE5"
# ## Feature importance (Importancia de variables)
#
# Anteriormente, hemos visto cómo realizar una selección de variables. Sin embargo, el modelo RandomForest es capaz de obtener la importancia de las diferentes variables, en base a cómo le afectan a cada uno de sus árboles:
# + id="ZDSbVWwymuEl" outputId="12da4216-271b-4b98-d0dc-1ca16864663a"
from sklearn.datasets import load_iris
# Leemos el dataset de sklearn:
iris = load_iris()
# Creamos el Random Forest:
rnd_clf = RandomForestClassifier(n_estimators=500, random_state=42)
# Entrenamos:
rnd_clf.fit(iris["data"], iris["target"])
# Obtenemos la importancia de las variables, iterando junto las columnas para mantener la relación, y las vamos imprimiendo:
for name, score in zip(iris["feature_names"], rnd_clf.feature_importances_):
print(name, score)
# + id="Q0UZLYXImuEo" outputId="5b94f074-4ba5-425e-90d6-de55eed84a42"
pd.DataFrame({'importances': rnd_clf.feature_importances_, 'variables': iris.feature_names}).sort_values(by='importances', ascending=False)
# -
# Veamos un ejemplo para un set de imágenes con trazos de números:
# + id="57HVLzbamuE6"
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1)
mnist.target = mnist.target.astype(np.uint8)
# + id="kANyaw-WmuE8" outputId="a0f43737-7ab3-4988-db44-9cdaa238f04a"
rnd_clf = RandomForestClassifier(n_estimators=100, random_state=42)
rnd_clf.fit(mnist["data"], mnist["target"])
# + id="nNmHdytHmuE_"
def plot_digit(data):
image = data.reshape(28, 28)
plt.imshow(image, cmap = mpl.cm.hot,
interpolation="nearest")
plt.axis("off")
# + id="4qyK5NALmuFB" outputId="40e0ad8b-0ee3-4097-99b5-51396340c18e"
plot_digit(rnd_clf.feature_importances_)
cbar = plt.colorbar(ticks=[rnd_clf.feature_importances_.min(), rnd_clf.feature_importances_.max()])
cbar.ax.set_yticklabels(['Not important', 'Very important'])
save_fig("mnist_feature_importance_plot")
plt.show()
# -
# ### EJERCICIO
#
# 1. Prueba este modelo sobre el dataset del Titanic para ver si una persona ha sobrevivido o no.
# 2. Obtén la importancia de las variables.
import pandas as pd
df = pd.read_csv("../../../data/titanic.csv", sep='\t')
df
# +
# El que queda de Embarked, lo vamos a rellenar con "S" porque es el mayoritario.
df['Embarked'] = df['Embarked'].fillna('S')
# En Age, vamos a completar los valores nulos con la media de los valores de Age.
df['Age'] = df['Age'].fillna(df['Age'].mean())
# En Pclass, vamos a convertirlo a string para luego hacer el One Hot Encoding.
df['Pclass'] = df['Pclass'].astype(str)
df['Sex'] = df['Sex'].apply(lambda x: x=='male')
# Separamos las variables que podemos utilizar para predecir de la que queremos predecir:
y_col = 'Survived'
X_cols = [col for col in df.columns if col not in ['PassengerId', 'Name', 'Ticket', 'Cabin', 'Survived']]
X = df[X_cols]
y = df[y_col]
# Tratamiento categóricas:
X = pd.get_dummies(X)
# Dividimos en train/test:
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.3, random_state=42)
# +
# Creamos el modelo:
rnd_clf = RandomForestClassifier(n_estimators=100, max_leaf_nodes=32, random_state=42)
# Entrenamos:
rnd_clf.fit(X_train, y_train)
# -
# Calculamos score:
rnd_clf.score(X_test, y_test)
# Calculamos score:
rnd_clf.score(X_train, y_train)
pd.DataFrame({'importances': rnd_clf.feature_importances_, 'variables': X.columns}).sort_values(by='importances', ascending=False)
# + [markdown] id="i0yaLopnmuFD"
# ## AdaBoost
#
# Otros de los modelos de ensemble más utilziados es AdaBoost, que se trata de un algoritmo de boosting, donde se colocan en serie los modelos para generar su salida en base a la salida del anterior:
# -
# Antes de nada, volvemos a crearnos X_train y X_test, por si las hemos sobreescrito en el ejercicio anterior:
X = X_moons
y = y_moons
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# + id="Fmx5d6NMmuFD" outputId="3c9e9027-3758-47f3-fd22-d4495c4d4837"
from sklearn.ensemble import AdaBoostClassifier
# Creamos clasificador:
ada_clf = AdaBoostClassifier(
DecisionTreeClassifier(max_depth=3),
n_estimators=5000,
algorithm="SAMME.R",
learning_rate=1,
random_state=42)
# Entrenamos:
ada_clf.fit(X_train, y_train)
# + id="KwXQVe-BmuFF" outputId="e1914478-03fa-47fa-d375-a2c00f66e572"
# Utilizamos la función diseñada anteriormente para pintar (pero no te aprendas esa función, esto es algo puntual)
plot_decision_boundary(ada_clf, X, y)
# + id="yO7092QNmuFH" outputId="34b0a733-a95f-4f61-f59e-b63f474ef4b8"
# Y nos creamos otra visualización variando diferentes parámetros:
m = len(X_train)
plt.figure(figsize=(11, 4))
for subplot, learning_rate in ((121, 1), (122, 0.5)):
sample_weights = np.ones(m)
plt.subplot(subplot)
for i in range(5):
svm_clf = SVC(kernel="rbf", C=0.05, gamma="scale", random_state=42)
svm_clf.fit(X_train, y_train, sample_weight=sample_weights)
y_pred = svm_clf.predict(X_train)
sample_weights[y_pred != y_train] *= (1 + learning_rate)
plot_decision_boundary(svm_clf, X, y, alpha=0.2)
plt.title("learning_rate = {}".format(learning_rate), fontsize=16)
if subplot == 121:
plt.text(-0.7, -0.65, "1", fontsize=14)
plt.text(-0.6, -0.10, "2", fontsize=14)
plt.text(-0.5, 0.10, "3", fontsize=14)
plt.text(-0.4, 0.55, "4", fontsize=14)
plt.text(-0.3, 0.90, "5", fontsize=14)
save_fig("boosting_plot")
plt.show()
# -
# ### EJERCICIO
#
# ¿Qué resultados conseguimos si utilizamos un modelo AdaBoost sobre nuestros datos del Titanic?
import pandas as pd
df = pd.read_csv("../../../data/titanic.csv", sep='\t')
df
# +
# El que queda de Embarked, lo vamos a rellenar con "S" porque es el mayoritario.
df['Embarked'] = df['Embarked'].fillna('S')
# En Age, vamos a completar los valores nulos con la media de los valores de Age.
df['Age'] = df['Age'].fillna(df['Age'].mean())
# En Pclass, vamos a convertirlo a string para luego hacer el One Hot Encoding.
df['Pclass'] = df['Pclass'].astype(str)
# Separamos las variables que podemos utilizar para predecir de la que queremos predecir:
y_col = 'Survived'
X_cols = [col for col in df.columns if col not in ['PassengerId', 'Name', 'Ticket', 'Cabin', 'Survived']]
X = df[X_cols]
y = df[y_col]
# Tratamiento categóricas:
X = pd.get_dummies(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# +
from sklearn.ensemble import AdaBoostClassifier
# Creamos clasificador:
ada_clf = AdaBoostClassifier(
DecisionTreeClassifier(max_depth=3),
n_estimators=500,
algorithm="SAMME.R",
learning_rate=0.85,
random_state=42)
pipe = Pipeline([('booster', ada_clf)])
params = {
'booster__n_estimators': [10, 50, 75, 100, 200, 500, 1000],
'booster__learning_rate': np.arange(0.5, 1, 0.05)
}
# Entrenamos:
gs_ada = GridSearchCV(pipe,
params,
cv=10,
scoring='accuracy',
n_jobs=-1,
verbose=1)
gs_ada.fit(X_train, y_train)
# -
gs_ada.score(X_test, y_test)
# +
from sklearn.ensemble import RandomForestClassifier
# Y ahora probaremos con el Radnom Forest de sklearn (que, pese a que se basa en la misma idea, tiene ligeras mejoras):
rnd_clf = RandomForestClassifier(n_estimators=500, max_leaf_nodes=16, random_state=42)
rnd_clf.fit(X_train, y_train)
# +
from sklearn.ensemble import AdaBoostClassifier
# Creamos clasificador:
ada_clf = AdaBoostClassifier(
DecisionTreeClassifier(), n_estimators=100,
algorithm="SAMME.R", learning_rate=1, random_state=42)
# Entrenamos:
ada_clf.fit(X_train, y_train)
print('score_train :', ada_clf.score(X_train, y_train))
print('score_test :',ada_clf.score(X_test, y_test))
# -
ada_clf.fit(X_train, y_train)
ada_clf.feature_importances_
# + [markdown] id="CRow6YWsmuFL"
# ## Gradient Boosting
#
# Otro modelo es el Gradient Boosting, que a diferencia del AdaBoost, no se encarga de actualizar los pesos de cada registro en cada iteración, sino que busca minimizar errores:
# + id="Zh1dlOnAmuFL"
# Nos creamos datos pseudo-aleatorios:
np.random.seed(42)
X = np.random.rand(100, 1) - 0.5
y = 3*X[:, 0]**2 + 0.05 * np.random.randn(100)
# -
# Vamos a hacerlo de forma "manual", entrenando uno a uno de forma secuencial. Además, crearemos un modelo de regresión, que se obtendrá con la media de los valores:
# + id="NUaOw9_AmuFN" outputId="fa442317-ac92-4c4f-9afb-e470c974a653"
from sklearn.tree import DecisionTreeRegressor
# Nos creamos el primer árbol de decisión:
tree_reg1 = DecisionTreeRegressor(max_depth=2, random_state=42)
# Y entrenamos:
tree_reg1.fit(X, y)
# -
# Ahora nos entrará el error de la salida (predicción) del primero:
y2 = y - tree_reg1.predict(X)
y2
# + id="oV8o2rM0muFP" outputId="edb7c45b-a6df-4809-fd2d-f8391da6ac53"
# Nos creamos otro árbol:
tree_reg2 = DecisionTreeRegressor(max_depth=2, random_state=42)
# Y lo entrenamos:
tree_reg2.fit(X, y2)
# -
# Ahora hacemos lo propio para el siguiente modelo:
y3 = y2 - tree_reg2.predict(X)
y3
# + id="Cislkt7HmuFS" outputId="4c65f876-16c2-4b8b-85f3-8918b777af58"
tree_reg3 = DecisionTreeRegressor(max_depth=2, random_state=42)
tree_reg3.fit(X, y3)
# -
# Y así podríamos seguir hasta que queramos. Vamos a probar con un ejemplo:
# + id="bGp-DSIumuFU"
X_new = np.array([[0.8]])
# + id="ZrgoEcG7muFW"
y_pred = sum(tree.predict(X_new) for tree in (tree_reg1, tree_reg2, tree_reg3))
# + id="yaO527VxmuFY" outputId="6d1388c2-792c-4c9a-cc75-47fa5ecfcd5f"
y_pred
# -
# Y representacmos gráficamente (no te tienes que aprender esto, repito):
# + id="Wgls3RIpmuFb"
def plot_predictions(regressors, X, y, axes, label=None, style="r-", data_style="b.", data_label=None):
x1 = np.linspace(axes[0], axes[1], 500)
y_pred = sum(regressor.predict(x1.reshape(-1, 1)) for regressor in regressors)
plt.plot(X[:, 0], y, data_style, label=data_label)
plt.plot(x1, y_pred, style, linewidth=2, label=label)
if label or data_label:
plt.legend(loc="upper center", fontsize=16)
plt.axis(axes)
# + id="OlBxhrBlmuFe" outputId="ab322637-71b1-4961-e431-2dc269ed4061"
plt.figure(figsize=(11,11))
plt.subplot(321)
plot_predictions([tree_reg1], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h_1(x_1)$", style="g-", data_label="Training set")
plt.ylabel("$y$", fontsize=16, rotation=0)
plt.title("Residuals and tree predictions", fontsize=16)
plt.subplot(322)
plot_predictions([tree_reg1], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h(x_1) = h_1(x_1)$", data_label="Training set")
plt.ylabel("$y$", fontsize=16, rotation=0)
plt.title("Ensemble predictions", fontsize=16)
plt.subplot(323)
plot_predictions([tree_reg2], X, y2, axes=[-0.5, 0.5, -0.5, 0.5], label="$h_2(x_1)$", style="g-", data_style="k+", data_label="Residuals")
plt.ylabel("$y - h_1(x_1)$", fontsize=16)
plt.subplot(324)
plot_predictions([tree_reg1, tree_reg2], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h(x_1) = h_1(x_1) + h_2(x_1)$")
plt.ylabel("$y$", fontsize=16, rotation=0)
plt.subplot(325)
plot_predictions([tree_reg3], X, y3, axes=[-0.5, 0.5, -0.5, 0.5], label="$h_3(x_1)$", style="g-", data_style="k+")
plt.ylabel("$y - h_1(x_1) - h_2(x_1)$", fontsize=16)
plt.xlabel("$x_1$", fontsize=16)
plt.subplot(326)
plot_predictions([tree_reg1, tree_reg2, tree_reg3], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h(x_1) = h_1(x_1) + h_2(x_1) + h_3(x_1)$")
plt.xlabel("$x_1$", fontsize=16)
plt.ylabel("$y$", fontsize=16, rotation=0)
save_fig("gradient_boosting_plot")
plt.show()
# -
# También podemos utilizar el objeto ya definido en ``sklearn``:
# + id="Alzq19vZmuFi" outputId="6054f854-e70a-4712-aff7-5905e5f38497"
from sklearn.ensemble import GradientBoostingRegressor
gbrt = GradientBoostingRegressor(max_depth=2, n_estimators=3, learning_rate=1.0, random_state=42)
gbrt.fit(X, y)
# +
from sklearn.metrics import mean_squared_error
mean_squared_error(y, gbrt.predict(X))
# + id="_-PdUJyXmuFj" outputId="c4fef0e0-adcd-4106-c32f-67353b04d338"
gbrt_slow = GradientBoostingRegressor(max_depth=2, n_estimators=200, learning_rate=0.1, random_state=42)
gbrt_slow.fit(X, y)
# -
mean_squared_error(y, gbrt_slow.predict(X))
# + id="GbmgmAqcmuFn" outputId="1fe8b54f-9158-4333-976f-c327d63fa58e"
plt.figure(figsize=(11,4))
plt.subplot(121)
plot_predictions([gbrt], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="Ensemble predictions")
plt.title("learning_rate={}, n_estimators={}".format(gbrt.learning_rate, gbrt.n_estimators), fontsize=14)
plt.subplot(122)
plot_predictions([gbrt_slow], X, y, axes=[-0.5, 0.5, -0.1, 0.8])
plt.title("learning_rate={}, n_estimators={}".format(gbrt_slow.learning_rate, gbrt_slow.n_estimators), fontsize=14)
save_fig("gbrt_learning_rate_plot")
plt.show()
# + [markdown] id="vfM62GztmuF6"
# ## XGBoost
#
# Finalmente, tenemos XGBoost, otro de los algoritmos de boosting más utilizados, basado en el descenso de gradiente par encontrar su mejor versión:
# + id="CqqQKGdomuF6"
import xgboost
# + id="9WZDMC11muF8" outputId="e1a05a5b-2f88-4a5c-b453-051f6a41d39d"
from sklearn.metrics import mean_squared_error
xgb_reg = xgboost.XGBRegressor(random_state=42)
xgb_reg.fit(X_train, y_train)
y_pred = xgb_reg.predict(X_test)
val_error = mean_squared_error(y_test, y_pred)
print("Validación MSE:", val_error)
# + id="NvdVVtxOmuGA" outputId="61579536-fd02-4357-9599-c198d0be9afd"
xgb_reg.fit(X_train, y_train, eval_set=[(X_test, y_test)], early_stopping_rounds=2)
y_pred = xgb_reg.predict(X_test)
val_error = mean_squared_error(y_test, y_pred)
print("Validación MSE:", val_error)
# -
# ### EJERCICIO
#
# El modelo xgboost también tiene versión de clasificación, xgboost.XGBClassifier. Investiga y utiliza este algoritmo sobre el dataset del Titanic que ya tenemos muy visto para predecir la gente que ha muerto:
import pandas as pd
df = pd.read_csv("../../../data/titanic.csv", sep='\t')
df
# +
# El que queda de Embarked, lo vamos a rellenar con "S" porque es el mayoritario.
df['Embarked'] = df['Embarked'].fillna('S')
# En Age, vamos a completar los valores nulos con la media de los valores de Age.
df['Age'] = df['Age'].fillna(df['Age'].mean())
# En Pclass, vamos a convertirlo a string para luego hacer el One Hot Encoding.
df['Pclass'] = df['Pclass'].astype(str)
# Separamos las variables que podemos utilizar para predecir de la que queremos predecir:
y_col = 'Survived'
X_cols = [col for col in df.columns if col not in ['PassengerId', 'Name', 'Ticket', 'Cabin', 'Survived']]
X = df[X_cols]
y = df[y_col]
# Tratamiento categóricas:
X = pd.get_dummies(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# -
y_pred
# + active=""
#
# +
from sklearn.metrics import mean_squared_error, accuracy_score
xgb_reg = xgboost.XGBClassifier(use_label_encoder=False, eval_metric='logloss', random_state=42)
pipe = Pipeline([("model", xgb_reg)])
parameters = {
'model__n_estimators': [1, 10, 25, 50, 100, 200, 500],
'model__max_depth': np.arange(1, 3),
'model__learning_rate': np.arange(0.01, 0.1, 0.01)
}
gs = GridSearchCV(pipe,
parameters,
cv=10,
scoring='neg_log_loss')
gs.fit(X_train, y_train)
# -
gs.best_params_
# +
from sklearn.metrics import accuracy_score
xgb_reg = xgboost.XGBClassifier(use_label_encoder=False, random_state=42)
xgb_reg.fit(X_train, y_train, eval_set=[(X_test, y_test)], early_stopping_rounds=2)
y_pred = xgb_reg.predict(X_test)
val_error = accuracy_score(y_test, y_pred)
print("Validación MSE:", val_error)
# -
| Bloque 3 - Machine Learning/01_Supervisado/9-Ensemble/01_Ensemble_Modelos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SPThole/Pytorch-Tutorials/blob/master/Eng_Hindi_transliteration_attention_up.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="KozZ0Acj5o2b"
# # Importing Required Libraries
# + id="6zDt0PeAbzsm"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import dataset, DataLoader
from torch.utils.data import random_split
from tqdm import tqdm_notebook as tqdm
from sklearn.metrics import r2_score, accuracy_score
from sklearn.preprocessing import StandardScaler,OneHotEncoder
from sklearn.model_selection import train_test_split
# + [markdown] id="-H7mx1Zdtb7f"
# # Geting data
# + id="AVThAKdXb9kS"
df = pd.read_csv('/content/crowd_transliterations.hi-en.txt',sep='\t',header=None)
# + colab={"base_uri": "https://localhost:8080/", "height": 195} id="zkSUaAEhcIWh" outputId="c49331e2-f190-4b13-c991-c092ec51205f"
df.head()
# + [markdown] id="F7rcReowthT4"
# # Preprocessing
# + id="I_nl33XccXkb"
d = {'<SOS>':1,
'<EOS>':2}
# + id="ds3gd6OlcwJ2"
df['eng_len'] = df[0].apply(lambda x: len(list(x)))
df['hin_len'] = df[1].apply(lambda x: len(list(x)))
# + id="m58S5wj_db3Z"
max_input = df['hin_len'].max()
max_output = df['eng_len'].max()
# + colab={"base_uri": "https://localhost:8080/"} id="67cev0bedoPY" outputId="72e3531c-bcb9-4641-cadb-563116d3bb6a"
max_input, max_output
# + id="n9c8rycbcmQu"
def name_to_idx(name,d):
name1 = name
for c in name1:
if c not in list(d.keys()):
d[c] = max(list(d.values()))+1
return d
def form_dict(names,d,max_len):
for n in tqdm(names):
n1 = input_create(n,max_len)
d = name_to_idx(n1,d)
return d
def input_create(name,max_len):
name1 = list(name)
name1 = name1 + ['<EOS>']
# for i in range(max_len -len(name1)):
# name1.append('<PAD>')
return name1
def input_create1(name,max_len):
name1 = list(name)
name1 = ['<SOS>'] + name1
name1 = name1 + ['<EOS>']
# for i in range(max_len -len(name1)):
# name1.append('<PAD>')
return name1
def one_hot(c,d):
one_h = [0 for i in range(len(d))]
one_h[d[c]] = 1
return d[c]
# + colab={"base_uri": "https://localhost:8080/", "height": 165, "referenced_widgets": ["9ae6abb4c31249e79e0f6b235a74fdef", "<KEY>", "8339ea9a2d074abaaa88f8ff21746f4a", "906508e146834107982002a76fe1c485", "<KEY>", "c76f26fd52324527a0e1d1ca3d4fd8da", "a3339392de344c8e8dea1c80a2c224a3", "<KEY>", "a9f525fe8d1e40909f36378995caaa65", "<KEY>", "<KEY>", "42ced9fd79c64289ba63d706a485e895", "<KEY>", "a516fc5617c043558e514148a8aad52a", "f6b0fc6a9f2a40cbb52ce2a43a2131ce", "3276e86db0e14b62904a7664d4c5dfbf"]} id="3FG3V0LQcpSD" outputId="8b7b00f7-cb69-4e97-a413-7e2bed8030dd"
d_input = form_dict(df[0],
{'<SOS>':1,
'<EOS>':2,
'<PAD>':0},
21)
d_output = form_dict(df[1],
{'<SOS>':1,
'<EOS>':2,
'<PAD>':0},
21)
# + id="K1THQy7EeIBG"
class mydata(DataLoader):
def __init__(self,eng,hin):
self.eng = eng
self.hin = hin
def __getitem__(self,idx):
e = self.eng[idx]
h = self.hin[idx]
X = torch.tensor([one_hot(c,d_input) for c in input_create(list(e),21)]).float()
X1= torch.tensor([one_hot(c,d_output) for c in input_create1(list(h),21)]).float()[:-1]
Y = torch.tensor([d_output[s] for s in input_create1(list(h),21)])[1:]
# Y = torch.tensor(X[1:]).float()
return X,X1,Y
def __len__(self):
return len(self.eng)
# + id="7AgqNapQ0OrA"
EOSI = torch.tensor(one_hot('<EOS>',d_input)).float()
SOSI = torch.tensor(one_hot('<SOS>',d_input)).float()
EOSO = torch.tensor(one_hot('<EOS>',d_output)).float()
SOSO = torch.tensor(one_hot('<SOS>',d_output)).float()
# + [markdown] id="TJcRdXnGtou7"
# # Dataset
# + id="gA5kjcoGQ7Gi"
from torch.nn.utils.rnn import pad_sequence,pad_packed_sequence
from torch.nn.utils.rnn import pack_padded_sequence
def padder(batch):
(xx, xx1, yy) = zip(*batch)
xx_lens = [len(xs) for xs in xx]
xx1_lens = [len(xs) for xs in xx1]
yy_lens = [len(yy) for ys in yy]
xx_padded = pad_sequence(xx, batch_first= True, padding_value = 0)
yy_padded = pad_sequence(yy, batch_first= True, padding_value = 0)
xx1_padded = pad_sequence(xx1, batch_first= True, padding_value = 0)
return xx_padded, xx1_padded, yy_padded, \
xx_lens, xx1_lens, yy_lens
# + id="e2CidKozfOLl"
data = mydata(df[0].values,df[1].values)
# + id="ezdJFGtUDKmS"
train_dataset, test_dataset = random_split(data,[12000,2919])
train_loader = DataLoader(train_dataset,128,collate_fn=padder,drop_last=True)
test_loader = DataLoader(test_dataset,128,collate_fn=padder,drop_last=True)
# + colab={"base_uri": "https://localhost:8080/"} id="djCw_3tzVPAV" outputId="8e88ac51-373a-4c94-bddd-4c41e93ff5cf"
for Z in train_loader:
print(Z[2].shape)
break
# + [markdown] id="2m3jZY3YtvTf"
# # Models
# + id="kzjqajZGr56y"
class encoder(nn.Module):
def __init__(self,
vocab_size,
input_size,
layers,
hidden_size,
output_size,
batch_size):
super(encoder, self).__init__()
self.vocab_size = vocab_size
self.input_size = input_size
self.layers = layers
self.hidden_size = hidden_size
self.output_size = output_size
self.batch_size = batch_size
self.emb = nn.Embedding(self.vocab_size,
self.input_size)
self.rnn = nn.LSTM(input_size=input_size,
hidden_size = hidden_size,
dropout = 0.2,
num_layers = self.layers,
batch_first = True)
self.fc = nn.Linear(self.hidden_size,self.output_size)
def forward(self,x,x_lens):
# print('ENCODER')
h0 = torch.randn((self.layers,self.batch_size,self.hidden_size))
c0 = torch.randn((self.layers,self.batch_size,self.hidden_size))
# print('ENCODER H0,C0 INIT',h0.shape,c0.shape)
x = self.emb(x.long())
# print('EMBEDDED INPUT',x.shape)
x_packed = pack_padded_sequence(x,
x_lens,
batch_first=True,
enforce_sorted=False)
out, (h1,c1) = self.rnn(x_packed,(h0,c0))
out, _ = pad_packed_sequence(out, batch_first=True)
# print('RNN OUTPUT',out.shape,h1.shape,c1.shape)
out = self.fc(out)
# print('FINAL ENCODER OUTPUT',out.shape)
return out,h1,c1
class attn(nn.Module):
def __init__(self,
encoder_out_size,
decoder_hidden_size,
decoder_input_size):
super(attn, self).__init__()
self.enc_out = encoder_out_size
self.dec_hid = decoder_hidden_size
self.dec_in = decoder_input_size
self.U = nn.Linear(self.enc_out,self.dec_in)
self.W = nn.Linear(self.dec_hid,self.dec_in)
self.attn_f = nn.Linear(self.dec_in,1)
def forward(self,enc_out,dec_hidn):
# print('ATTENTION')
# print('Enc out shape',enc_out.shape,'>>')
encU = self.U(enc_out)
# print('EncU',encU.shape)
dec_hidn = dec_hidn.permute(1,0,2)
# print('dec_hidden shape',dec_hidn.shape,'>>')
decW = self.W(dec_hidn)
# print('decW',decW.shape)
# print(encU.shape,decW.shape)
UW = nn.functional.tanh(encU+decW)
# print('UW',UW.shape,'>>')
scores = nn.functional.softmax(self.attn_f(UW),dim=1)
# print('scores',scores.shape,'>>')
return scores
class decoderattn(nn.Module):
def __init__(self,
input_size,
hidden_size,
output_size,
attn):
super(decoderattn, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.output_size = output_size
self.emb = nn.Embedding(self.input_size,self.input_size)
self.rnn = nn.LSTM(input_size = 2*self.input_size,
hidden_size = self.hidden_size,
num_layers = 1,
batch_first = True)
self.attn = attn
self.fc = nn.Linear(self.hidden_size,self.output_size)
def forward(self,enc_out,x,h0,c0):
# print("DECODER")
# print("ENC OUT,X",enc_out.shape,x.shape)
a = self.attn(enc_out,h0)
# print('scores',a.shape)
context = a*enc_out
context = context.sum(axis=1)
# print('context',context.shape)
# print('DECODER EMB INPUT',x.long().shape)
x = self.emb(x.long())
# print('DECODER EMBEDDED INPUT',x.shape,'>>')
# print('DECODER CONCAT STEP',context.unsqueeze(1).shape,x.shape)
context_cat_lho = torch.cat((context.unsqueeze(1),
x.unsqueeze(1)),axis=-1)
# print('context_cat_lho',context_cat_lho.shape)
o,(h1,c1) = self.rnn(context_cat_lho,(h0,c0))
o = self.fc(o)
# print('OUTPUT',o.shape)
return o,a,h1,c1
class myencdec(nn.Module):
def __init__(self,enc,dec):
super(myencdec, self).__init__()
self.enc = enc
self.dec = dec
self.attn = attn
def forward(self,X):
h0 = torch.randn((self.enc.layers,128,self.enc.hidden_size))
c0 = torch.randn((self.enc.layers,128,self.enc.hidden_size))
# OUT = []
# print('ITERATION ENCODER INPUT',X[0].shape,'>>')
enc_out,h0,c0 = self.enc(X[0],X[3])
# print('ITERATION ENCODER OUTPUT',enc_out.shape,h0.shape,c0.shape,'>>')
# print('ITERATION DECODER INPUT',enc_out.shape,X[1][:,0].shape,h0.shape,c0.shape,'>>')
o,a,h1,c1 = self.dec(enc_out,X[1][:,0],h0,c0)
# print('ITERATION DECODER INIT OUTPUT',o.shape)
OUT = [o]
for i in range(1,X[1].size(1)):
o = torch.argmax(o,axis=-1).squeeze(1)
# print('ITERATION DEC',i,'th',o.shape)
o,a,h1,c1 = dec(enc_out,
X[1][:,i],
h1,
c1)
OUT.append(o)
return torch.stack(OUT).permute(2,1,0,3).squeeze(0)
# + colab={"base_uri": "https://localhost:8080/"} id="R7o9jyLGW7_a" outputId="92d554bd-3ffa-432e-d0fb-f98545ccb329"
#ENCODER
VOCAB_SIZE = len(d_input)
INPUT_SIZE = np.int(1.5*VOCAB_SIZE)
HIDDEN_SIZE = 100
OUTPUT_SIZE = 82
BATCH_SIZE = 128
enc = encoder(VOCAB_SIZE,INPUT_SIZE,1,HIDDEN_SIZE,OUTPUT_SIZE,BATCH_SIZE)
# ATTENTION
DECODER_INPUT_SIZE = len(d_output)
DECODER_HIDDEN_SIZE = 100
att = attn(OUTPUT_SIZE,DECODER_HIDDEN_SIZE,DECODER_INPUT_SIZE)
# # DECODER
dec = decoderattn(DECODER_INPUT_SIZE,DECODER_HIDDEN_SIZE,DECODER_INPUT_SIZE,att)
# + id="bs_OJWTWYGQN"
# for X in train_loader:
# out,h1, c1 = enc(X)
# break
# print(X[0].shape,out.shape)
# print(att(out,torch.rand(1,128,DECODER_HIDDEN_SIZE)).shape)
# o,a,h,c = dec(out,torch.rand((128,1,DECODER_INPUT_SIZE)),h1,c1)
# + id="KX-sifdoT-yI"
# o.shape,a.shape,h.shape
# + id="P_ec3x2EtqQN"
# enc = encoder(100,4,200,100)
# dec = decoder(100,4,100,82)
# + id="q5GR2f0Rwkuv"
h0 = torch.randn((enc.layers,128,enc.hidden_size))
c0 = torch.randn((enc.layers,128,enc.hidden_size))
# + id="i6ApCPI1LpPJ"
model = myencdec(enc,dec)
# + id="5aeGQosUu71j" colab={"base_uri": "https://localhost:8080/"} outputId="dc59dc75-98c2-4cdb-9b4c-76053b460c8f"
for X in train_loader:
out =model(X)
break
# + colab={"base_uri": "https://localhost:8080/"} id="jyUTRXETnGWx" outputId="fbf8e3fb-a479-45e3-ac9e-119b800c8465"
out.shape,len(X[2][0])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="BV-kz2XRADie" outputId="4cc5559d-5080-4ea1-a49c-e967c11b6700"
inv_dict = dict(zip(d_output.values(), d_output.keys()))
''.join([inv_dict[j.item()] for j in torch.argmax(out[5],axis=1)])
# + id="--J9cewrGOEo"
optimizer = optim.AdamW(model.parameters(), lr = 0.001)
loss_func = nn.CrossEntropyLoss()
# + [markdown] id="DjDwS569t2JQ"
# # Training
# + id="bD5uWdRMiKEe"
class train_model:
def __init__(self,
train_loader,
test_loader,
task,
model,
optimizer,
criterion,
epochs):
self.train_loader = train_loader
self.test_loader = test_loader
self.task = task
self.optimizer = optimizer
self.criterion = criterion
self.epochs = epochs
self.model = model
def train_it(self):
"""
"""
history = {'train_loss':[],
'test_loss':[],
'train_score':[],
'test_score':[]}
for epoch in range(self.epochs):
model.train()
for loader in self.train_loader:
self.optimizer.zero_grad()
xs = loader
ys = torch.tensor(xs[2]).reshape(-1,)
preds= self.predict(xs)
# print(preds.shape,ys.shape)
loss = self.criterion(self.predict(xs).reshape(-1,82),ys.long())
loss.backward()
self.optimizer.step()
model.eval()
history['train_loss'].append(self.calc_loss(self.train_loader))
history['test_loss'].append(self.calc_loss(self.test_loader))
history['train_score'].append(self.calc_score(self.train_loader))
history['test_score'].append(self.calc_score(self.test_loader))
if epoch%5==0:
print(f"Iteration : {epoch}")
fig = plt.figure(figsize = (12,7))
plt.subplot(1, 2, 1)
plt.plot(history['train_loss'], color = 'red', label = 'Training Loss')
plt.plot(history['test_loss'], color = 'green', label = 'Validation Loss')
plt.legend()
plt.subplot(1, 2, 2)
plt.plot(history['train_score'], color = 'red', label = 'Training Score')
plt.plot(history['test_score'], color = 'green', label = 'Validation Score' )
plt.legend()
plt.show()
# fig = plt.figure()
# ax = fig.add_subplot(111, projection = '3d')
# ax.scatter(X[:,0], X[:,1], model(torch.tensor(X).float())[:,0].detach().numpy())
# plt.show()
return self.model, history
def r2(self, xs, ys):
return r2_score(ys.reshape(-1,), self.predict(xs).numpy())
def acc(self, ys, preds):
maxs, cls = torch.max(preds, axis = 1)
return torch.sum(cls==ys.reshape(-1,)).item()/ys.shape[0]
def predict(self,xs):
return self.model(xs)
def calc_loss(self,loader):
loss = []
for idx, data in enumerate(loader):
xs = data
ys = torch.tensor(xs[2]).reshape(-1,)
loss.append(self.criterion(self.predict(xs).reshape(-1,82),ys.long()).item())
return np.mean(loss)
def calc_score(self,loader):
scores = []
for data in loader:
xs = data
ys = torch.tensor(xs[2]).reshape(-1,)
if self.task == 'reg':
scores.append(self.r2(ys,self.predict(xs).reshape(-1,82)))
else:
scores.append(self.acc(ys,self.predict(xs).reshape(-1,82)))
return np.mean(scores)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="qbJtbHiqjNP4" outputId="a889eb7f-6e62-4da0-e368-fb77ab4f50b6"
trainer = train_model(train_loader,
test_loader,
'clf',
model,
optimizer,
loss_func,
16)
model, h = trainer.train_it()
# + [markdown] id="Oc4IUrPSt8xd"
# # Evaluating
# + id="02CBnugyooY7" colab={"base_uri": "https://localhost:8080/"} outputId="b1b8f35d-5923-4fc0-dcda-137cadeb504f"
model.eval()
# + id="18Gj1T3oH0R8" colab={"base_uri": "https://localhost:8080/"} outputId="594e6d04-f9f7-4bfb-b81b-4a4216b170fe"
for X in test_loader:
out =model(X)
for idx in list(np.arange(127)):
inv_dict = dict(zip(d_output.values(), d_output.keys()))
print(''.join([inv_dict[j.item()] for j in torch.argmax(out[idx],axis=1)]))
print(''.join([inv_dict[j.item()] for j in X[2][idx]]))
print('==============================================')
# + id="Nvf7R59ooscj" colab={"base_uri": "https://localhost:8080/"} outputId="5a7c5e64-53b8-4b72-dcad-42f06b95df99"
for idx in list(np.arange(127)):
inv_dict = dict(zip(d_output.values(), d_output.keys()))
print(''.join([inv_dict[j.item()] for j in torch.argmax(out[idx],axis=1)]))
print(''.join([inv_dict[j.item()] for j in X[2][idx]]))
print('==============================================')
# + [markdown] id="dVgeEYKcuBYH"
# # Testing
# + id="tLvxj11DbEMX" colab={"base_uri": "https://localhost:8080/"} outputId="0090c89b-c55e-4b2c-8862-346b0853841b"
for letter in 'index'.split(' '):
fortest = mydata([letter],['1111111111111'])
fortestload = DataLoader(fortest,1,collate_fn=padder)
s = ''
for X in fortestload:
out,h0,c0 = model.enc(X[0],X[3])
print(X[1].shape)
o,a,h1,c1 = model.dec(out,X[1][:,0],h0,c0)
# o = torch.argmax(o,axis=-1).squeeze(1)
s = inv_dict[torch.argmax(o).item()]
A = a
for i in range(10):
# print(o)
o = torch.argmax(o,axis=-1).squeeze(1)
o,a,h1,c1 = model.dec(out,o,h1,c1)
id = torch.argmax(o)
s = s + inv_dict[torch.argmax(o).item()]
A =torch.cat((A,a))
print(a.shape)
print(s)
# + id="t7ntZMeDGt2o"
import skimage.io as io
# + colab={"base_uri": "https://localhost:8080/"} id="llpAQVGl_LH6" outputId="712a4947-d75a-4ec0-f0fe-2bbad7186eb1"
A.squeeze(2).detach().numpy().shape
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="dkbEven_-Zhe" outputId="9f3e0df7-cc50-4d40-a6f0-cdf90fbe1014"
io.imshow(A.squeeze(2).permute(1,0).detach().numpy())
# + id="_umR3Nmt_EL9"
A.squeeze(2).detach().numpy()
# + id="mqfyNwfyPP1y"
| Sequential/Eng_Hindi_transliteration_attention_up.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# import the pymongo library - this is the Python driver!
import pymongo
# replace "uri" with your Atlas URI string - should look like mongodb+srv://...
uri = "mongodb+srv://m220student:<EMAIL>@mf<EMAIL>.mongodb.net/sample_mflix?retryWrites=true&w=majority"
client = pymongo.MongoClient(uri)
mflix = client.sample_mflix
# movies is our collection handle - it refers to the sample_mflix.movies collection
movies = mflix.movies
# -
# find one document in the "movies" collection - does not matter which one
movies.find_one()
# find one document in the "movies" collection - must have "Salma Hayek" in the "cast"
movies.find_one( { "cast": "Salma Hayek" } )
# find all the documents in the "movies" collection with "Salma Hayek" in the "cast"
# this returns a cursor, which IS a Python iterable, but is NOT a document!
movies.find( { "cast": "Salma Hayek" } )
# return the count of movies with "Salma Hayek" in the "cast"
movies.find( { "cast": "Salma Hayek" } ).count()
# find all movies with Salma Hayek
# then pretty print
cursor = movies.find( { "cast": "Salma Hayek" } )
from bson.json_util import dumps
print(dumps(cursor, indent=2))
# find all movies with Salma Hayek, but only project the "_id" and "title" fields
cursor = movies.find( { "cast": "Salma Hayek" }, { "title": 1 } )
print(dumps(cursor, indent=2))
# find all movies with Salma Hayek, but only project the "title" field
cursor = movies.find( { "cast": "Salma Hayek" }, { "title": 1, "_id": 0 } )
print(dumps(cursor, indent=2))
| mflix-python/notebooks/your_first_read.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="N7ITxKLUkX0v"
# ##### Copyright 2020 The TensorFlow Authors.
# + cellView="form" id="yOYx6tzSnWQ3"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="6xgB0Oz5eGSQ"
# # グラフと関数の基礎
# + [markdown] id="w4zzZVZtQb1w"
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://www.tensorflow.org/guide/intro_to_graphs"><img src="https://www.tensorflow.org/images/tf_logo_32px.png"> TensorFlow.orgで表示</a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ja/guide/intro_to_graphs.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab で実行</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ja/guide/intro_to_graphs.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub でソースを表示{</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ja/guide/intro_to_graphs.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">ノートブックをダウンロード/a0}</a></td>
# </table>
# + [markdown] id="RBKqnXI9GOax"
# # グラフと `tf.function` の基礎
#
# このガイドは、TensorFlow の仕組みを説明するために、TensorFlow と Keras 基礎を説明します。今すぐ Keras に取り組みたい方は、[Keras のガイド一覧](keras/)を参照してください。
#
# このガイドでは、グラフ取得のための単純なコード変更、格納と表現、およびモデルの高速化とエクスポートを行うための使用方法について、TensorFlow の中核的な仕組みを説明します。
#
# 注意: TensorFlow 1.x のみの知識をお持ちの場合は、このガイドでは、非常に異なるグラフビューが紹介されています。
#
# これは、基礎を概説したガイドです。これらの概念の徹底ガイドについては、[`tf.function` ガイド](function)を参照してください。
#
# + [markdown] id="v0DdlfacAdTZ"
# ## グラフとは?
#
# 前回の 3 つのガイドでは、TensorFlow の **Eager** execution について説明しました。これは、TensorFlow 演算が演算ごとにPythonによって実行され、結果を Python に返すことを意味します。Eager TensorFlow は GPU を活用し、変数、テンソル、さらには演算を GPU と TPU に配置することができます。また、デバックも簡単に行えます。
#
# 一部のユーザーは、Python から移動する必要はありません。
#
# ただし、TensorFlow を Python で演算ごとに実行すると、ほかの方法では得られない多数の高速化機能が利用できなくなります。Python からテンソルの計算を抽出できる場合は、*グラフ* にすることができます。
#
# **グラフとは、計算のユニットを表す一連の `tf.Operation` オブジェクトと、演算間を流れるデータのユニットを表す `tf.Tensor` オブジェクトを含むデータ構造です。** `tf.Graph` コンテキストで定義されます。これらのグラフはデータ構造であるため、元の Python コードがなくても、保存、実行、および復元することができます。
#
# 次は、TensorBoard で視覚化された単純な二層グラフです。
#
# + [markdown] id="FvQ5aBuRGT1o"
# 
# + [markdown] id="DHpY3avXGITP"
# ## グラフのメリット
#
# グラフを使用すると、柔軟性が大幅に向上し、モバイルアプリケーション。組み込みデバイス、バックエンドサーバーといった Python インタプリタのない環境でも TensorFlow グラフを使用できます。TensorFlow は、Python からエクスポートされた場合に、保存されるモデルの形式としてグラフを使用します。
#
# また、グラフは最適化を簡単に行えるため、コンパイラは次のような変換を行えます。
#
# - 計算に定数ノードを畳み込むで、テンソルの値を統計的に推論します*(「定数畳み込み」)*。
# - 独立した計算のサブパートを分離し、スレッドまたはデバイスに分割します。
# - 共通部分式を取り除き、算術演算を単純化します。
#
# + [markdown] id="o1x1EOD9GjnB"
# これやほかの高速化を実行する [Grappler](./graph_optimization.ipynb) という総合的な最適化システムがあります。
#
# まとめると、グラフは非常に便利なもので、**複数のデバイス**で、TensorFlow の**高速化**、**並列化**、および効率化を期待することができます。
#
# ただし、便宜上、Python で機械学習モデル(またはその他の計算)を定義した後、必要となったときに自動的にグラフを作成することをお勧めします。
# + [markdown] id="pSZebVuWxDXu"
# # グラフのトレース
#
# TensorFlow でグラフを作成する方法は、直接呼出しまたはデコレータのいずれかとして `tf.function` を使用することです。
# + id="goZwOXp_xyQj"
import tensorflow as tf
import timeit
from datetime import datetime
# + id="HKbLeJ1y0Umi"
# Define a Python function
def function_to_get_faster(x, y, b):
x = tf.matmul(x, y)
x = x + b
return x
# Create a `Function` object that contains a graph
a_function_that_uses_a_graph = tf.function(function_to_get_faster)
# Make some tensors
x1 = tf.constant([[1.0, 2.0]])
y1 = tf.constant([[2.0], [3.0]])
b1 = tf.constant(4.0)
# It just works!
a_function_that_uses_a_graph(x1, y1, b1).numpy()
# + [markdown] id="MT7U8ozok0gV"
# `tf.function` 化された関数は、[Python コーラブル]()で、Python 相当と同じように機能します。特定のクラス(`python.eager.def_function.Function`)を使用しますが、ユーザーにとっては、トレースできないものと同じように動作します。
#
# `tf.function` は、それが呼び出す Python 関数を再帰的にトレースします。
# + id="rpz08iLplm9F"
def inner_function(x, y, b):
x = tf.matmul(x, y)
x = x + b
return x
# Use the decorator
@tf.function
def outer_function(x):
y = tf.constant([[2.0], [3.0]])
b = tf.constant(4.0)
return inner_function(x, y, b)
# Note that the callable will create a graph that
# includes inner_function() as well as outer_function()
outer_function(tf.constant([[1.0, 2.0]])).numpy()
# + [markdown] id="P88fOr88qgCj"
# TensorFlow 1.x を使用したことがある場合は、`Placeholder` または `tf.Sesssion` をまったく定義する必要がないことに気づくでしょう。
# + [markdown] id="wfeKf0Nr1OEK"
# ## フローの制御と副次的影響
#
# フロー制御とループは、デフォルトで `tf.autograph` によって TensorFlow に変換されます。Autograph は、ループコンストラクトの標準化、アンロール、および [AST](https://docs.python.org/3/library/ast.html) マニピュレーションなどのメソッドを組み合わせて使用します。
#
# + id="PFObpff1BMEb"
def my_function(x):
if tf.reduce_sum(x) <= 1:
return x * x
else:
return x-1
a_function = tf.function(my_function)
print("First branch, with graph:", a_function(tf.constant(1.0)).numpy())
print("Second branch, with graph:", a_function(tf.constant([5.0, 5.0])).numpy())
# + [markdown] id="hO4DBUNZBMwQ"
# Autograph 変換を直接呼び出して、Python が TensorFlow 演算に変換される様子を確認することができます。これはほとんど解読不能ですが、変換を確認することができます。
# + id="8x6RAqza1UWf"
# Don't read the output too carefully.
print(tf.autograph.to_code(my_function))
# + [markdown] id="GZ4Ieg6tBE6l"
# Autograph は、`if-then` 句、ループ、 `break`、`return`、`continue` などを自動的に変換します。
#
# ほとんどの場合、Autograph の動作に特別な考慮はいりませんが、いくつかの注意事項があり、これについては [tf.function ガイド](./function.ipynb)のほか、[Autograph 完全リファレンス](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/autograph/g3doc/reference/index.md)が役立ちます。
# + [markdown] id="A6NHDp7vAKcJ"
# ## 高速化の確認
#
# tensor-using 関数を `tf.function` でラッピングするだけでは、コードは高速化しません。単一のマシンで数回呼び出された小さな関数では、グラフまたはグラフの一部の呼び出しにかかるオーバーヘッドによってランタイムが占有されてしまうことがあります。また、GPU 大きな負荷をかける畳み込みのスタックなど、計算のほとんどがすでにアクセラレータで発生している場合は、グラフの高速化をあまり確認できません。
#
# 複雑な計算については、グラフによって大幅な高速化を得ることができます。これは、グラフが Python からデバイスへの通信や一部の高速化の実装を減らすためです。
#
# 次のコードは、小さな密のレイヤーでの数回の実行にかかる時間を計測します。
# + id="zbNndv-0BeO4"
# Create an oveerride model to classify pictures
class SequentialModel(tf.keras.Model):
def __init__(self, **kwargs):
super(SequentialModel, self).__init__(**kwargs)
self.flatten = tf.keras.layers.Flatten(input_shape=(28, 28))
self.dense_1 = tf.keras.layers.Dense(128, activation="relu")
self.dropout = tf.keras.layers.Dropout(0.2)
self.dense_2 = tf.keras.layers.Dense(10)
def call(self, x):
x = self.flatten(x)
x = self.dense_1(x)
x = self.dropout(x)
x = self.dense_2(x)
return x
input_data = tf.random.uniform([60, 28, 28])
eager_model = SequentialModel()
graph_model = tf.function(eager_model)
print("Eager time:", timeit.timeit(lambda: eager_model(input_data), number=10000))
print("Graph time:", timeit.timeit(lambda: graph_model(input_data), number=10000))
# + [markdown] id="kNGuLnjK1c5U"
# ### 多層型関数
#
# 関数をトレースする場合、**多層型**の `Function` オブジェクトを作成します。多層型関数は Pythonコーラブルで、1つの API の背後にあるいくつかの具象関数グラフをカプセル化します。
#
# この `Function` は、あらゆる `dtypes` と形状に使用できます。新しい引数シグネチャでそれを呼び出すたびに、元の関数が新しい引数で再トレースされます。`Function` は、そのトレースに対応する `tf.Graph` を `concrete_function` に格納します。関数がすでにそのような引数でトレースされている場合は、トレース済みのグラフが取得されます。
#
# 概念的に、次のようになります。
#
# - **`tf.Graph`** は計算を説明する未加工のポータブルなデータ構造である
# - **`Function`** は、ConcreteFunctions のキャッシュ、トレース、およびディスパッチャーである
# - **`ConcreteFunction`** は、Python からグラフを実行できるグラフの Eager 対応ラッパーである
#
# ### 多層型関数の検査
#
# `a_function` を検査できます。これはPython 関数 `my_function` に対して `tf.function` を呼び出した結果です。この例では、3 つの引数で `a_function` を呼び出すことで、3 つの具象関数を得られています。
#
# + id="7heuYuwn2edE"
print(a_function)
print("Calling a `Function`:")
print("Int:", a_function(tf.constant(2)))
print("Float:", a_function(tf.constant(2.0)))
print("Rank-1 tensor of floats", a_function(tf.constant([2.0, 2.0, 2.0])))
# + id="s1c8db0cCW2k"
# Get the concrete function that works on floats
print("Inspecting concrete functions")
print("Concrete function for float:")
print(a_function.get_concrete_function(tf.TensorSpec(shape=[], dtype=tf.float32)))
print("Concrete function for tensor of floats:")
print(a_function.get_concrete_function(tf.constant([2.0, 2.0, 2.0])))
# + id="JLTNuv_CCZXK"
# Concrete functions are callable
# Note: You won't normally do this, but instead just call the containing `Function`
cf = a_function.get_concrete_function(tf.constant(2))
print("Directly calling a concrete function:", cf(tf.constant(2)))
# + [markdown] id="PTHNiHLXH9es"
# この例では、スタックの非常に奥を調べています。具体的にトレースを管理していない限り、通常は、ここに示されるように具象関数を呼び出す必要はありません。
# + [markdown] id="V11zkxU22XeD"
# # Eager execution でのデバッグ
#
# スタックトレースが長い場合、特に `tf.Graph` または `with tf.Graph().as_default()` の参照が含まれる場合、グラフコンテキストで実行している可能性があります。TensorFlow のコア関数は Keras の `model.fit()` などのグラフコンテキストを使用します。
#
# Eager execution をデバッグする方がはるかに簡単であることがよくあります。スタックトレースは比較的に短く、理解しやすいからです。
#
# グラフのデバックが困難な場合は、Eager execution に戻ってデバックすることができます。
#
# Eager で実行していることを確認するには、次を行います。
#
# - メソッドとレイヤーを直接コーラブルとして呼び出す
#
# - Keras compile/fit を使用している場合、コンパイル時に **`model.compile(run_eagerly=True)`** を使用する
#
# - **`tf.config.experimental_run_functions_eagerly(True)`** でグローバル実行モードを設定する
#
# + [markdown] id="iTHvdQfRICJb"
# ### `run_eagerly=True` を使用する
# + id="kqzBV2rSzvpC"
# Define an identity layer with an eager side effect
class EagerLayer(tf.keras.layers.Layer):
def __init__(self, **kwargs):
super(EagerLayer, self).__init__(**kwargs)
# Do some kind of initialization here
def call(self, inputs):
print("\nCurrently running eagerly", str(datetime.now()))
return inputs
# + id="5DFvc9ySr7t3"
# Create an override model to classify pictures, adding the custom layer
class SequentialModel(tf.keras.Model):
def __init__(self):
super(SequentialModel, self).__init__()
self.flatten = tf.keras.layers.Flatten(input_shape=(28, 28))
self.dense_1 = tf.keras.layers.Dense(128, activation="relu")
self.dropout = tf.keras.layers.Dropout(0.2)
self.dense_2 = tf.keras.layers.Dense(10)
self.eager = EagerLayer()
def call(self, x):
x = self.flatten(x)
x = self.dense_1(x)
x = self.dropout(x)
x = self.dense_2(x)
return self.eager(x)
# Create an instance of this model
model = SequentialModel()
# Generate some nonsense pictures and labels
input_data = tf.random.uniform([60, 28, 28])
labels = tf.random.uniform([60])
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
# + [markdown] id="U3-hcwmpI3Sv"
# まず、Eager を使用せずにモデルをコンパイルします。モデルはトレースされません。名前にも関わらず、`compile` は、損失関数、最適化、およびトレーニングパラメータのセットアップしか行いません。
# + id="w2GdwhB_KQlw"
model.compile(run_eagerly=False, loss=loss_fn)
# + [markdown] id="WLMXk1uxKQ44"
# ここで、`fit` を呼び出し、関数がトレース(2 回)されると Eager 効果が実行しなくなるのを確認します。
# + id="VCoLlZDythZ8"
model.fit(input_data, labels, epochs=3)
# + [markdown] id="jOk6feLOK1pR"
# ただし、エポックを 1 つでも Eager で実行すると、Eager の副次的作用が 2 回現れます。
# + id="MGIYwrKpK06e"
print("Running eagerly")
# When compiling the model, set it to run eagerly
model.compile(run_eagerly=True, loss=loss_fn)
model.fit(input_data, labels, epochs=1)
# + [markdown] id="qwq_cnc8Lwf8"
# ### `experimental_run_functions_eagerly` を使用する
#
# また、すべてを Eager で実行するよにグローバルに設定することができます。これは、トレースし直した場合にのみ機能することに注意してください。トレースされた関数は、トレースされたままとなり、グラフとして実行します。
# + id="oFSxRtcptYpe"
# Now, globally set everything to run eagerly
tf.config.experimental_run_functions_eagerly(True)
print("Run all functions eagerly.")
# First, trace the model, triggering the side effect
polymorphic_function = tf.function(model)
# It was traced...
print(polymorphic_function.get_concrete_function(input_data))
# But when you run the function again, the side effect happens (both times).
result = polymorphic_function(input_data)
result = polymorphic_function(input_data)
# + id="pD-AQxEhua4E"
# Don't forget to set it back when you are done
tf.config.experimental_run_functions_eagerly(False)
# + [markdown] id="sm0bNFp8PX53"
# # トレースとパフォーマンス
#
# トレースにはある程度のオーバーヘッドがかかります。小さな関数のトレースは素早く行えますが、大規模なモデルであればかなりの時間がかかる場合があります。パフォーマンスが上昇するとこの部分の時間は迅速に取り戻されますが、大規模なモデルのトレーニングの最初の数エポックでは、トレースによって遅延が発生する可能性があることに注意しておくことが重要です。
#
# モデルの規模に関係なく、頻繁にトレースするのは避けたほうがよいでしょう。[tf.function ガイドのこのセクション](function.ipynb#when_to_retrace)では、入力仕様を設定し、テンソル引数を使用して再トレースを回避する方法について説明しています。フォーマンスが異常に低下している場合は、誤って再トレースしていないかどうかを確認することをお勧めします。
#
# eager-only の副次的効果(Python 引数の出力など)を追加して、関数がいつトレースされているかを確認できます。ここでは、新しい Python 引数が常に再トレースをトリガするため、余分な再トレースが発生していることを確認できます。
# + id="jsGQ4GQAP2Ve"
# Use @tf.function decorator
@tf.function
def a_function_with_python_side_effect(x):
print("Tracing!") # This eager
return x * x + tf.constant(2)
# This is traced the first time
print(a_function_with_python_side_effect(tf.constant(2)))
# The second time through, you won't see the side effect
print(a_function_with_python_side_effect(tf.constant(3)))
# This retraces each time the Python argument chances
# as a Python argument could be an epoch count or other
# hyperparameter
print(a_function_with_python_side_effect(2))
print(a_function_with_python_side_effect(3))
# + [markdown] id="D1kbr5ocpS6R"
# # 次のステップ
#
# より詳しい説明については、`tf.function` API リファレンスページと[ガイド](./function.ipynb)を参照してください。
| site/ja/guide/intro_to_graphs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Flopy MODFLOW Boundary Conditions
#
# Flopy has a new way to enter boundary conditions for some MODFLOW packages. These changes are substantial. Boundary conditions can now be entered as a list of boundaries, as a numpy recarray, or as a dictionary. These different styles are described in this notebook.
#
# Flopy also now requires zero-based input. This means that **all boundaries are entered in zero-based layer, row, and column indices**. This means that older Flopy scripts will need to be modified to account for this change. If you are familiar with Python, this should be natural, but if not, then it may take some time to get used to zero-based numbering. Flopy users submit all information in zero-based form, and Flopy converts this to the one-based form required by MODFLOW.
#
# The following MODFLOW packages are affected by this change:
#
# * Well
# * Drain
# * River
# * General-Head Boundary
# * Time-Variant Constant Head
#
# This notebook explains the different ways to enter these types of boundary conditions.
#
# +
#begin by importing flopy
import os
import sys
import numpy as np
# run installed version of flopy or add local path
try:
import flopy
except:
fpth = os.path.abspath(os.path.join('..', '..'))
sys.path.append(fpth)
import flopy
workspace = os.path.join('data')
#make sure workspace directory exists
if not os.path.exists(workspace):
os.makedirs(workspace)
print(sys.version)
print('numpy version: {}'.format(np.__version__))
print('flopy version: {}'.format(flopy.__version__))
# -
# ## List of Boundaries
# Boundary condition information is passed to a package constructor as stress_period_data. In its simplest form, stress_period_data can be a list of individual boundaries, which themselves are lists. The following shows a simple example for a MODFLOW River Package boundary:
stress_period_data = [
[2, 3, 4, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 5, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 6, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
]
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data)
m.write_input()
# If we look at the River Package created here, you see that the layer, row, and column numbers have been increased by one.
# !head -n 10 'data/test.riv'
# If this model had more than one stress period, then Flopy will assume that this boundary condition information applies until the end of the simulation
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
dis = flopy.modflow.ModflowDis(m, nper=3)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data)
m.write_input()
# !head -n 10 'data/test.riv'
# ## Recarray of Boundaries
#
# Numpy allows the use of recarrays, which are numpy arrays in which each column of the array may be given a different type. Boundary conditions can be entered as recarrays. Information on the structure of the recarray for a boundary condition package can be obtained from that particular package. The structure of the recarray is contained in the dtype.
riv_dtype = flopy.modflow.ModflowRiv.get_default_dtype()
print(riv_dtype)
# Now that we know the structure of the recarray that we want to create, we can create a new one as follows.
stress_period_data = np.zeros((3), dtype=riv_dtype)
stress_period_data = stress_period_data.view(np.recarray)
print('stress_period_data: ', stress_period_data)
print('type is: ', type(stress_period_data))
# We can then fill the recarray with our boundary conditions.
stress_period_data[0] = (2, 3, 4, 10.7, 5000., -5.7)
stress_period_data[1] = (2, 3, 5, 10.7, 5000., -5.7)
stress_period_data[2] = (2, 3, 6, 10.7, 5000., -5.7)
print(stress_period_data)
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data)
m.write_input()
# !head -n 10 'data/test.riv'
# As before, if we have multiple stress periods, then this recarray will apply to all of them.
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
dis = flopy.modflow.ModflowDis(m, nper=3)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data)
m.write_input()
# !head -n 10 'data/test.riv'
# ## Dictionary of Boundaries
#
# The power of the new functionality in Flopy3 is the ability to specify a dictionary for stress_period_data. If specified as a dictionary, the key is the stress period number (**as a zero-based number**), and the value is either a nested list, an integer value of 0 or -1, or a recarray for that stress period.
#
# Let's say that we want to use the following schedule for our rivers:
# 0. No rivers in stress period zero
# 1. Rivers specified by a list in stress period 1
# 2. No rivers
# 3. No rivers
# 4. No rivers
# 5. Rivers specified by a recarray
# 6. Same recarray rivers
# 7. Same recarray rivers
# 8. Same recarray rivers
#
sp1 = [
[2, 3, 4, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 5, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
[2, 3, 6, 10.7, 5000., -5.7], #layer, row, column, stage, conductance, river bottom
]
print(sp1)
riv_dtype = flopy.modflow.ModflowRiv.get_default_dtype()
sp5 = np.zeros((3), dtype=riv_dtype)
sp5 = sp5.view(np.recarray)
sp5[0] = (2, 3, 4, 20.7, 5000., -5.7)
sp5[1] = (2, 3, 5, 20.7, 5000., -5.7)
sp5[2] = (2, 3, 6, 20.7, 5000., -5.7)
print(sp5)
sp_dict = {0:0, 1:sp1, 2:0, 5:sp5}
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
dis = flopy.modflow.ModflowDis(m, nper=8)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=sp_dict)
m.write_input()
# !head -n 10 'data/test.riv'
# ## MODFLOW Auxiliary Variables
#
# Flopy works with MODFLOW auxiliary variables by allowing the recarray to contain additional columns of information. The auxiliary variables must be specified as package options as shown in the example below.
#
# In this example, we also add a string in the last column of the list in order to name each boundary condition. In this case, however, we do not include boundname as an auxiliary variable as MODFLOW would try to read it as a floating point number.
#create an empty array with an iface auxiliary variable at the end
riva_dtype = [('k', '<i8'), ('i', '<i8'), ('j', '<i8'),
('stage', '<f4'), ('cond', '<f4'), ('rbot', '<f4'),
('iface', '<i4'), ('boundname', object)]
riva_dtype = np.dtype(riva_dtype)
stress_period_data = np.zeros((3), dtype=riva_dtype)
stress_period_data = stress_period_data.view(np.recarray)
print('stress_period_data: ', stress_period_data)
print('type is: ', type(stress_period_data))
stress_period_data[0] = (2, 3, 4, 10.7, 5000., -5.7, 1, 'riv1')
stress_period_data[1] = (2, 3, 5, 10.7, 5000., -5.7, 2, 'riv2')
stress_period_data[2] = (2, 3, 6, 10.7, 5000., -5.7, 3, 'riv3')
print(stress_period_data)
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data, dtype=riva_dtype, options=['aux iface'])
m.write_input()
# !head -n 10 'data/test.riv'
# ## Working with Unstructured Grids
#
# Flopy can create an unstructured grid boundary condition package for MODFLOW-USG. This can be done by specifying a custom dtype for the recarray. The following shows an example of how that can be done.
#create an empty array based on nodenumber instead of layer, row, and column
rivu_dtype = [('nodenumber', '<i8'), ('stage', '<f4'), ('cond', '<f4'), ('rbot', '<f4')]
rivu_dtype = np.dtype(rivu_dtype)
stress_period_data = np.zeros((3), dtype=rivu_dtype)
stress_period_data = stress_period_data.view(np.recarray)
print('stress_period_data: ', stress_period_data)
print('type is: ', type(stress_period_data))
stress_period_data[0] = (77, 10.7, 5000., -5.7)
stress_period_data[1] = (245, 10.7, 5000., -5.7)
stress_period_data[2] = (450034, 10.7, 5000., -5.7)
print(stress_period_data)
m = flopy.modflow.Modflow(modelname='test', model_ws=workspace)
riv = flopy.modflow.ModflowRiv(m, stress_period_data=stress_period_data, dtype=rivu_dtype)
m.write_input()
print(workspace)
# !head -n 10 'data/test.riv'
# ## Combining two boundary condition packages
ml = flopy.modflow.Modflow(modelname="test",model_ws=workspace)
dis = flopy.modflow.ModflowDis(ml,10,10,10,10)
sp_data1 = {3: [1, 1, 1, 1.0],5:[1,2,4,4.0]}
wel1 = flopy.modflow.ModflowWel(ml, stress_period_data=sp_data1)
ml.write_input()
# !head -n 10 'data/test.wel'
sp_data2 = {0: [1, 1, 3, 3.0],8:[9,2,4,4.0]}
wel2 = flopy.modflow.ModflowWel(ml, stress_period_data=sp_data2)
ml.write_input()
# !head -n 10 'data/test.wel'
# Now we create a third wel package, using the ```MfList.append()``` method:
wel3 = flopy.modflow.ModflowWel(ml,stress_period_data=\
wel2.stress_period_data.append(
wel1.stress_period_data))
ml.write_input()
# !head -n 10 'data/test.wel'
| examples/Notebooks/flopy3_modflow_boundaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: rabpro
# language: python
# name: rabpro
# ---
# # Multiple basins
# This example demonstrates the power of the `rapbro` interface to Google Earth Engine (GEE) to collect many subbasin attributes associated with multiple coordinate pairs.
# +
import os
import pandas as pd
from tqdm import tqdm
import geopandas as gpd
import contextily as cx # You may need to 'conda install contextily' or 'pip install contextily'
import matplotlib.pyplot as plt
import rabpro
from rabpro.basin_stats import Dataset
# os.environ['RABPRO_DATA'] = r'X:\Data' # Point this to your rabpro datapath if it different than the default
# -
# If you have not authenticated Google Earth Engine yet, this is a good time to do it. Running the following code should prompt you for an authorization token and instructions for getting it.
# ```python
# import ee
# ee.Authenticate()
# ```
# First let's create a list of coordinate pairs that we'll use to delineate our basins of interest. For this example, we'll retrieve the subbasins associated with dams in Sri Lanka. Data are from [VotE-Dams](https://data.ess-dive.lbl.gov/view/doi:10.15485/1843541).
dam_coords = [(9.299352, 80.324096), (6.9159649, 81.0100197), (8.1709424, 80.223766), (8.3519444, 80.3852778), (6.8577186, 80.5855038), (6.3098805, 80.8557684), (7.3017445, 80.700303), (7.8666667, 80.6166667), (5.9907437, 80.654368), (7.29212186, 80.63798077), (6.1064653, 80.6434438), (7.8002358, 80.5507896), (7.1644604, 81.6172083), (6.0966667, 80.5980556), (7.9926439, 80.9124781), (7.7713804, 80.4618547), (6.6663889, 81.1519444), (8.2166667, 80.7166667), (7.21031, 81.534525), (9.342165, 80.447647), (6.8531681, 80.1937833), (8.0116232, 80.5577336), (7.8749822, 80.7075809), (8.166120326, 80.92907567), (5.97662322, 80.59436068), (7.060356, 80.598767), (6.58, 80.326944), (6.8356819, 80.1798343), (8.5977778, 80.9497222), (8.3951818, 80.5521785), (8.4649153, 80.1930859), (7.276247, 81.0395821), (6.4406287, 80.9580932), (8.027173, 80.89291), (8.5850794, 81.0064442), (6.9811916, 81.5020328), (8.251949, 80.469926), (7.2667672, 81.1002985), (7.6801531, 80.6139585), (7.296071, 81.5130095), (7.4583255, 81.6007655), (6.913889, 80.521667), (8.347577, 80.419759), (8.8077739, 80.7574688), (7.9448769, 80.292059), (7.0793219, 81.6281681), (7.895393, 80.988042), (8.6907253, 80.4298704), (7.7185503, 81.1885293), (7.3219103, 80.645308), (6.9190503, 80.4894399), (8.1325, 80.2458333), (7.201942, 80.922261), (7.1996059, 80.9496288), (7.5082139, 81.0558007), (6.2057327, 80.9843446), (7.6461111, 81.4808333), (6.675005, 80.7865558), (8.0727324, 79.9556424), (6.428509, 80.838542), (7.618288, 81.5488207), (6.946745731, 80.65814325), (8.0608333, 80.3175), (9.090326112, 80.33821621), (7.241352, 80.783926), (8.7224936, 80.8350109), (6.5474919, 81.260143), (7.647458, 81.213197)]
dam_ids = [13331, 13332, 13333, 13334, 13335, 13336, 13337, 13338, 13339, 13340, 13341, 13342, 13343, 13344, 13345, 13346, 13347, 13348, 13349, 13350, 13351, 13352, 13353, 13354, 13355, 13356, 13357, 13358, 13359, 13360, 13361, 13362, 13363, 13364, 13365, 13366, 13367, 13368, 13369, 13370, 13371, 13372, 13373, 13374, 13375, 13376, 13377, 13378, 13379, 13380, 13381, 13382, 13383, 13384, 13385, 13386, 13387, 13388, 13389, 13390, 13391, 13392, 13393, 13394, 13395, 13396, 13397, 22927]
dam_das = [116.0, 1.6, 131.0, 4.0, 138.0, 176.0, 0.95, 155.0, 4.15, 2.75, 7.03, 26.0, 32.0, 4.9, 1.08, 58.0, 45.0, 0.79, 992.0, 555.0, 16.16, 834.0, 96.9, 88.0, 5.5, 572.0, 318.0, 5.7, 152.0, 326.0, 405.0, 2.79, 231.0, 213.0, 73.7, 69.2, 598.0, 72.6, 29.7, 30.9, 181.0, 173.6, 834.5, 552.0, 19.2, 26.3, 85.7, 297.25, 102.2, 1350.0, 170.7, 1616.0, 2350.0, 3109.0, 38.7, 26.8, 981.0, 340.0, 311.0, 1160.0, 239.0, 312.0, 183.0, 250.0, 1900.0, 72.0, 549.8, 470.5]
vdams_srilanka = pd.DataFrame(data={'coords':dam_coords, 'vote_id':dam_ids, 'da_km2':dam_das})
vdams_srilanka.head()
# We loop through each coordinate pair and delineate its watershed using MERIT (this is only possible if an estimated drainage area is known, which is the case here, else `rabpro` would use HydroBasins), then combine all watershed polygons into a GeoDataFrame. If you want to use MERIT, download tile 'n00e060' following the MERIT download [example](https://github.com/VeinsOfTheEarth/rabpro/blob/main/docs/source/examples/notebooks/downloading_data.ipynb).
if os.path.isfile("basins_sl.gpkg") is False:
basins_sl = []
for i, row in tqdm(vdams_srilanka.iterrows(), total=vdams_srilanka.shape[0]):
rpo = rabpro.profiler(row['coords'], da=row['da_km2'], verbose=False)
rpo.delineate_basin(force_merit=True)
rpo.watershed['vote_id'] = row['vote_id']
basins_sl.append(rpo.watershed)
basins_sl = pd.concat(basins_sl)
basins_sl.to_file("basins_sl.gpkg", driver="GPKG")
else:
basins_sl = gpd.read_file("basins_sl.gpkg")
# We need to upload our basin vector layer as a GEE asset because it too large to send as a json payload. This can be done manually through the GEE code editor or via `rabpro` automation. This automation requires that you have a writeable Google Cloud Platform (GCP) bucket and that you are authenticated via the command-line to call the `gsutil` and `earthengine` programs. These programs enable sending files to GCP and onward to GEE respectively.
#
# For now, you can skip this step as we've uploaded the sl_basins file as a public asset.
# ```python
# gcp_bucket = "your_gcp_bucket"
# gee_user = "your_gee_username"
# zip_path = rabpro.utils.build_gee_vector_asset(basins_sl, "basins_sl.zip")
# your_gee_asset = rabpro.utils.upload_gee_vector_asset(
# zip_path, gee_user, gcp_bucket, gcp_folder="rabpro"
# )
#
# ```
# We'd like to get zonal statistics over each of our delineated basins. We need to first define a list of Google Earth Engine (GEE) datasets we'd like to sample.
dataset_list = [
Dataset(
"JRC/GSW1_3/GlobalSurfaceWater",
"occurrence",
time_stats=["median"],
stats=["mean"],
),
Dataset(
"ECMWF/ERA5_LAND/MONTHLY",
"temperature_2m",
time_stats=["median"],
stats=["mean"],
),
Dataset(
"UCSB-CHG/CHIRPS/DAILY", "precipitation", time_stats=["median"], stats=["mean"]
),
]
# Now let's do the sampling with `rabpro.basin_stats.compute()`, format the results, and join it to our basins GeoDataFrame.
your_gee_asset = 'users/jstacompute/basins_sl'
# %%time
# This could take a few minutes to run, depending on how quickly GEE completes your tasks.
if not os.path.exists("res.gpkg"):
urls, tasks = rabpro.basin_stats.compute(
dataset_list, gee_feature_path=your_gee_asset, folder="rabpro"
)
tag_list = ["wateroccur", "temperature_2m", "precip"]
data = rabpro.basin_stats.fetch_gee(urls, tag_list)
res = gpd.GeoDataFrame(data.merge(basins_sl, on='vote_id'))
res = res.set_geometry("geometry")
res.to_file("res.gpkg", driver="GPKG")
else:
res = gpd.read_file("res.gpkg")
# Finally, we can visualize our results!
# +
def build_panel(col_name, col_label, axis):
axis.set_xlim([left-0.5, right+0.5])
axis.set_ylim([bot-0.2, top+0.2])
axis.set_axis_off()
res.plot(
ax=axis,
column=col_name,
legend=True,
legend_kwds={"label": col_label, "orientation": "horizontal", "pad": 0.01},
)
cx.add_basemap(axis, zoom=10, crs=res.crs, attribution="")
return axis
left, bot, right, top = res.total_bounds
fig, ax = plt.subplots(1, 3, figsize=(10, 10))
ax = [
build_panel(col_name, col_label, ax[i])
for col_name, col_label, i in zip(
[x for x in res.columns[1:4]],
["Water occurence %", "Temperature (K)", "Precipitation (mm/day)"],
range(0, 3),
)
]
| docs/source/examples/notebooks/multiple_basins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
data = pd.read_csv('water_potability.csv')
data.head()
# -
X = data.drop('Potability',axis=1)
y = data['Potability']
print(X.shape)
print(y)
# +
from sklearn.impute import SimpleImputer
impute = SimpleImputer()
X_impute = pd.DataFrame(impute.fit_transform(X))
# imputation removed columns. so put them back
X_impute.columns = X.columns
X= X_impute
# +
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,random_state=42)
# -
# KNN CLASSIFIER
# +
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=30)
knn.fit(X_train,y_train)
pred = knn.predict(X_train)
print("Training set score:{:.2f}".format(knn.score(X_train,y_train)*100))
print("Testing set score:{:.2f}".format(knn.score(X_test,y_test)*100))
# +
import matplotlib.pyplot as plt
training_accuracy = []
testing_accuracy = []
neighbors_settings = range(1,40)
for n_neighbors in neighbors_settings:
knn = KNeighborsClassifier(n_neighbors = n_neighbors)
knn.fit(X_train,y_train)
training_accuracy.append(knn.score(X_train,y_train))
testing_accuracy.append(knn.score(X_test,y_test))
#print("training accuracy of the model".format(training_accuracy))
#print("Tesing accuracy of the model:\n{}".format(testing_accuracy))
plt.plot(neighbors_settings,training_accuracy,label = "training accuracy")
plt.plot(neighbors_settings,testing_accuracy,label = "testing accuracy")
plt.ylabel("accuracy")
plt.xlabel("n_neighbors")
plt.legend()
# +
#FEATURE IMPORTANCES
from sklearn.inspection import permutation_importance
knn.fit(X,y)
results = permutation_importance(knn,X,y,scoring="accuracy") # performs permutation importances
imp = results.importances_mean # get importances
for i,v in enumerate(imp):
print("Feature: %0d, score: %.3f"%(i,v))
plt.bar([X for X in range(len(imp))],imp)
plt.show()
# -
# RANDOM FOREST
# +
#Random forest
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=10,random_state=42,max_depth=3,n_jobs=-1)
rfc.fit(X_train,y_train)
pred = rfc.predict(X_train)
print("Training set score:{:.2f}".format(rfc.score(X_train,y_train)*100))
print("Testing set score:{:.2f}".format(rfc.score(X_test,y_test)*100))
# +
training_accuracy = []
testing_accuracy = []
estimators_settings = range(1,50)
for n_estimators in estimators_settings:
rfc = RandomForestClassifier(n_estimators)
rfc.fit(X_train,y_train)
training_accuracy.append(rfc.score(X_train,y_train)*100)
testing_accuracy.append(rfc.score(X_test,y_test)*100)
plt.plot(estimators_settings,training_accuracy,label="training accuracy")
plt.plot(estimators_settings,testing_accuracy,label="testing accuracy")
plt.ylabel("accuracy")
plt.xlabel("n_estimators")
plt.legend()
# +
rfc.fit(X,y)
imp = rfc.feature_importances_
for i,v in enumerate(imp):
print("Feature:%0d , score: %.2f"%(i,v))
plt.bar([X for X in range(len(imp))],imp)
plt.show()
# -
# GRADIENT BOOSTER REGRESSION
# +
# Gradient Booster Regression
from sklearn.ensemble import GradientBoostingClassifier
gbrt = GradientBoostingClassifier(random_state=42)
gbrt.fit(X_train,y_train)
print("Training set score:{:.2f}".format(gbrt.score(X_train,y_train)*100))
print("Testing set score:{:.2f}".format(gbrt.score(X_test,y_test)*100))
# -
# LOGISTIC REGRESSION
# +
# logistic regression
# implementation with all the features
from sklearn.linear_model import LogisticRegression
lreg = LogisticRegression().fit(X_train,y_train)
pred = lreg.predict(X_train)
print("Training set score:{:.2f}".format(lreg.score(X_train,y_train)*100))
print("Testing set score:{:.2f}".format(lreg.score(X_test,y_test)*100))
lreg10 = LogisticRegression(C=10).fit(X_train,y_train)
print("\nTraining set score with C=10:\n{:.2f}".format(lreg10.score(X_train,y_train)*100))
print("Testing set score with C=10:\n{:.2f}".format(lreg10.score(X_test,y_test)*100))
lreg100 = LogisticRegression(C=100).fit(X_train,y_train)
print("\nTraining set score with C=100:\n{:.2f}".format(lreg100.score(X_train,y_train)*100))
print("Testing set score with C=100:\n{:.2f}".format(lreg100.score(X_test,y_test)*100))
lreg001 = LogisticRegression(C=0.01).fit(X_train,y_train)
print("\nTraining set score with C=0.01:\n{:.2f}".format(lreg001.score(X_train,y_train)*100))
print("Testing set score with C=0.01:\n{:.2f}".format(lreg001.score(X_test,y_test)*100))
# -
lreg.fit(X,y)
importance = lreg.coef_[0]
for i,v in enumerate(importance):
print('Feature: %0d, Score: %.5f' % (i,v))
# plot feature importance
plt.bar([x for x in range(len(importance))], importance)
plt.show()
# +
# since feature 6 i.e, organic_carbon is triggering a huge loss we will eliminate that feature
X1 = X.drop('Organic_carbon',axis=1)
X1
# +
X1_train,X1_test,y_train,y_test = train_test_split(X1,y,random_state=0)
lreg0 = LogisticRegression().fit(X1_train,y_train)
print("training score after removing feature 6:\n{:.2f}".format(lreg0.score(X1_train,y_train)*100))
print("testing score after removing feature 6:\n{:.2f}".format(lreg0.score(X1_test,y_test)*100))
# -
lreg.fit(X1,y)
importance = lreg.coef_[0]
for i,v in enumerate(importance):
print('Feature: %0d, Score: %.5f' % (i,v))
#plot feature importance
plt.bar([x for x in range(len(importance))], importance)
plt.show()
# +
X2 = X1.drop('Hardness',axis=1)
X2
X2_train,X2_test,y_train,y_test = train_test_split(X2,y,random_state=0)
lreg1 = LogisticRegression().fit(X2_train,y_train)
print("training score after removing feature 6 & 1:\n{:.2f}".format(lreg1.score(X2_train,y_train)*100))
print("testing score after removing feature 6 & 1:\n{:.2f}".format(lreg1.score(X2_test,y_test)*100))
# -
| Water Potability/water potability calssification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="qnyTxjK_GbOD" colab_type="text"
# # Beyond Hello World, A Computer Vision Example
# In the previous exercise you saw how to create a neural network that figured out the problem you were trying to solve. This gave an explicit example of learned behavior. Of course, in that instance, it was a bit of overkill because it would have been easier to write the function Y=2x-1 directly, instead of bothering with using Machine Learning to learn the relationship between X and Y for a fixed set of values, and extending that for all values.
#
# But what about a scenario where writing rules like that is much more difficult -- for example a computer vision problem? Let's take a look at a scenario where we can recognize different items of clothing, trained from a dataset containing 10 different types.
# + [markdown] id="H41FYgtlHPjW" colab_type="text"
# ## Start Coding
#
# Let's start with our import of TensorFlow
# + id="q3KzJyjv3rnA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7a1ceed7-66f6-4a43-b3f4-26d54bd95817" executionInfo={"status": "ok", "timestamp": 1550587660097, "user_tz": 480, "elapsed": 1577, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-RcxktLY-TBk/AAAAAAAAAAI/AAAAAAAAABY/b4V4dTIqmPI/s64/photo.jpg", "userId": "06401446828348966425"}}
import tensorflow as tf
print(tf.__version__)
# + [markdown] id="n_n1U5do3u_F" colab_type="text"
# The Fashion MNIST data is available directly in the tf.keras datasets API. You load it like this:
# + id="PmxkHFpt31bM" colab_type="code" colab={}
mnist = tf.keras.datasets.fashion_mnist
# + [markdown] id="GuoLQQBT4E-_" colab_type="text"
# Calling load_data on this object will give you two sets of two lists, these will be the training and testing values for the graphics that contain the clothing items and their labels.
#
# + id="BTdRgExe4TRB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="104aab14-aa09-4ed2-806c-38807fbac030" executionInfo={"status": "ok", "timestamp": 1550587667916, "user_tz": 480, "elapsed": 1381, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-RcxktLY-TBk/AAAAAAAAAAI/AAAAAAAAABY/b4V4dTIqmPI/s64/photo.jpg", "userId": "06401446828348966425"}}
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
# + [markdown] id="rw395ROx4f5Q" colab_type="text"
# What does these values look like? Let's print a training image, and a training label to see...Experiment with different indices in the array. For example, also take a look at index 42...that's a a different boot than the one at index 0
#
# + id="FPc9d3gJ3jWF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1238} outputId="5efae963-162d-4418-c001-2e8d6af83f40" executionInfo={"status": "ok", "timestamp": 1550587722332, "user_tz": 480, "elapsed": 357, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-RcxktLY-TBk/AAAAAAAAAAI/AAAAAAAAABY/b4V4dTIqmPI/s64/photo.jpg", "userId": "06401446828348966425"}}
import matplotlib.pyplot as plt
plt.imshow(training_images[0])
print(training_labels[0])
print(training_images[0])
# + [markdown] id="3cbrdH225_nH" colab_type="text"
# You'll notice that all of the values in the number are between 0 and 255. If we are training a neural network, for various reasons it's easier if we treat all values as between 0 and 1, a process called '**normalizing**'...and fortunately in Python it's easy to normalize a list like this without looping. You do it like this:
# + id="kRH19pWs6ZDn" colab_type="code" colab={}
training_images = training_images / 255.0
test_images = test_images / 255.0
# + [markdown] id="3DkO0As46lRn" colab_type="text"
# Now you might be wondering why there are 2 sets...training and testing -- remember we spoke about this in the intro? The idea is to have 1 set of data for training, and then another set of data...that the model hasn't yet seen...to see how good it would be at classifying values. After all, when you're done, you're going to want to try it out with data that it hadn't previously seen!
# + [markdown] id="dIn7S9gf62ie" colab_type="text"
# Let's now design the model. There's quite a few new concepts here, but don't worry, you'll get the hang of them.
# + id="7mAyndG3kVlK" colab_type="code" colab={}
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
# + [markdown] id="-lUcWaiX7MFj" colab_type="text"
# **Sequential**: That defines a SEQUENCE of layers in the neural network
#
# **Flatten**: Remember earlier where our images were a square, when you printed them out? Flatten just takes that square and turns it into a 1 dimensional set.
#
# **Dense**: Adds a layer of neurons
#
# Each layer of neurons need an **activation function** to tell them what to do. There's lots of options, but just use these for now.
#
# **Relu** effectively means "If X>0 return X, else return 0" -- so what it does it it only passes values 0 or greater to the next layer in the network.
#
# **Softmax** takes a set of values, and effectively picks the biggest one, so, for example, if the output of the last layer looks like [0.1, 0.1, 0.05, 0.1, 9.5, 0.1, 0.05, 0.05, 0.05], it saves you from fishing through it looking for the biggest value, and turns it into [0,0,0,0,1,0,0,0,0] -- The goal is to save a lot of coding!
#
# + [markdown] id="c8vbMCqb9Mh6" colab_type="text"
# The next thing to do, now the model is defined, is to actually build it. You do this by compiling it with an optimizer and loss function as before -- and then you train it by calling **model.fit ** asking it to fit your training data to your training labels -- i.e. have it figure out the relationship between the training data and its actual labels, so in future if you have data that looks like the training data, then it can make a prediction for what that data would look like.
# + id="BLMdl9aP8nQ0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="91a80606-f0a5-41f0-9d03-91d1ecdc8e17" executionInfo={"status": "ok", "timestamp": 1550587849991, "user_tz": 480, "elapsed": 29548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-RcxktLY-TBk/AAAAAAAAAAI/AAAAAAAAABY/b4V4dTIqmPI/s64/photo.jpg", "userId": "06401446828348966425"}}
model.compile(optimizer = tf.train.AdamOptimizer(),
loss = 'sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(training_images, training_labels, epochs=5)
# + [markdown] id="-JJMsvSB-1UY" colab_type="text"
# Once it's done training -- you should see an accuracy value at the end of the final epoch. It might look something like 0.9098. This tells you that your neural network is about 91% accurate in classifying the training data. I.E., it figured out a pattern match between the image and the labels that worked 91% of the time. Not great, but not bad considering it was only trained for 5 epochs and done quite quickly.
#
# But how would it work with unseen data? That's why we have the test images. We can call model.evaluate, and pass in the two sets, and it will report back the loss for each. Let's give it a try:
# + id="WzlqsEzX9s5P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="4dd8018a-f3ba-4e5b-e163-3d4f5dd12f19" executionInfo={"status": "ok", "timestamp": 1550587895553, "user_tz": 480, "elapsed": 832, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-RcxktLY-TBk/AAAAAAAAAAI/AAAAAAAAABY/b4V4dTIqmPI/s64/photo.jpg", "userId": "06401446828348966425"}}
model.evaluate(test_images, test_labels)
# + [markdown] id="6tki-Aro_Uax" colab_type="text"
# For me, that returned a accuracy of about .8838, which means it was about 88% accurate. As expected it probably would not do as well with *unseen* data as it did with data it was trained on! As you go through this course, you'll look at ways to improve this.
#
# To explore further, try the below exercises:
#
# + [markdown] id="htldZNWcIPSN" colab_type="text"
# # Exploration Exercises
# + [markdown] id="rquQqIx4AaGR" colab_type="text"
# ###Exercise 1:
# For this first exercise run the below code: It creates a set of classifications for each of the test images, and then prints the first entry in the classifications. The output, after you run it is a list of numbers. Why do you think this is, and what do those numbers represent?
# + id="RyEIki0z_hAD" colab_type="code" colab={}
classifications = model.predict(test_images)
print(classifications[0])
# + [markdown] id="MdzqbQhRArzm" colab_type="text"
# Hint: try running print(test_labels[0]) -- and you'll get a 9. Does that help you understand why this list looks the way it does?
# + id="WnBGOrMiA1n5" colab_type="code" colab={}
print(test_labels[0])
# + [markdown] id="uUs7eqr7uSvs" colab_type="text"
# ### What does this list represent?
#
#
# 1. It's 10 random meaningless values
# 2. It's the first 10 classifications that the computer made
# 3. It's the probability that this item is each of the 10 classes
#
#
# + [markdown] id="wAbr92RTA67u" colab_type="text"
# ####Answer:
# The correct answer is (3)
#
# The output of the model is a list of 10 numbers. These numbers are a probability that the value being classified is the corresponding value, i.e. the first value in the list is the probability that the handwriting is of a '0', the next is a '1' etc. Notice that they are all VERY LOW probabilities.
#
# For the 7, the probability was .999+, i.e. the neural network is telling us that it's almost certainly a 7.
# + [markdown] id="CD4kC6TBu-69" colab_type="text"
# ### How do you know that this list tells you that the item is an ankle boot?
#
#
# 1. There's not enough information to answer that question
# 2. The 10th element on the list is the biggest, and the ankle boot is labelled 9
# 2. The ankle boot is label 9, and there are 0->9 elements in the list
#
#
#
# + [markdown] id="I-haLncrva5L" colab_type="text"
# ####Answer
# The correct answer is (2). Both the list and the labels are 0 based, so the ankle boot having label 9 means that it is the 10th of the 10 classes. The list having the 10th element being the highest value means that the Neural Network has predicted that the item it is classifying is most likely an ankle boot
# + [markdown] id="OgQSIfDSOWv6" colab_type="text"
# ##Exercise 2:
# Let's now look at the layers in your model. Experiment with different values for the dense layer with 512 neurons. What different results do you get for loss, training time etc? Why do you think that's the case?
#
#
# + id="GSZSwV5UObQP" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels) , (test_images, test_labels) = mnist.load_data()
training_images = training_images/255.0
test_images = test_images/255.0
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(),
tf.keras.layers.Dense(1024, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
model.compile(optimizer = 'adam',
loss = 'sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
classifications = model.predict(test_images)
print(classifications[0])
print(test_labels[0])
# + [markdown] id="bOOEnHZFv5cS" colab_type="text"
# ###Question 1. Increase to 1024 Neurons -- What's the impact?
#
# 1. Training takes longer, but is more accurate
# 2. Training takes longer, but no impact on accuracy
# 3. Training takes the same time, but is more accurate
#
# + [markdown] id="U73MUP2lwrI2" colab_type="text"
# ####Answer
# The correct answer is (1) by adding more Neurons we have to do more calculations, slowing down the process, but in this case they have a good impact -- we do get more accurate. That doesn't mean it's always a case of 'more is better', you can hit the law of diminishing returns very quickly!
# + [markdown] id="WtWxK16hQxLN" colab_type="text"
# ##Exercise 3:
#
# What would happen if you remove the Flatten() layer. Why do you think that's the case?
#
# You get an error about the shape of the data. It may seem vague right now, but it reinforces the rule of thumb that the first layer in your network should be the same shape as your data. Right now our data is 28x28 images, and 28 layers of 28 neurons would be infeasible, so it makes more sense to 'flatten' that 28,28 into a 784x1. Instead of wriitng all the code to handle that ourselves, we add the Flatten() layer at the begining, and when the arrays are loaded into the model later, they'll automatically be flattened for us.
# + id="ExNxCwhcQ18S" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels) , (test_images, test_labels) = mnist.load_data()
training_images = training_images/255.0
test_images = test_images/255.0
model = tf.keras.models.Sequential([#tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)])
model.compile(optimizer = 'adam',
loss = 'sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
classifications = model.predict(test_images)
print(classifications[0])
print(test_labels[0])
# + [markdown] id="VqoCR-ieSGDg" colab_type="text"
# ##Exercise 4:
#
# Consider the final (output) layers. Why are there 10 of them? What would happen if you had a different amount than 10? For example, try training the network with 5
#
# You get an error as soon as it finds an unexpected value. Another rule of thumb -- the number of neurons in the last layer should match the number of classes you are classifying for. In this case it's the digits 0-9, so there are 10 of them, hence you should have 10 neurons in your final layer.
# + id="MMckVntcSPvo" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels) , (test_images, test_labels) = mnist.load_data()
training_images = training_images/255.0
test_images = test_images/255.0
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation=tf.nn.relu),
tf.keras.layers.Dense(5, activation=tf.nn.softmax)])
model.compile(optimizer = 'adam',
loss = 'sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
classifications = model.predict(test_images)
print(classifications[0])
print(test_labels[0])
# + [markdown] id="-0lF5MuvSuZF" colab_type="text"
# ##Exercise 5:
#
# Consider the effects of additional layers in the network. What will happen if you add another layer between the one with 512 and the final layer with 10.
#
# Ans: There isn't a significant impact -- because this is relatively simple data. For far more complex data (including color images to be classified as flowers that you'll see in the next lesson), extra layers are often necessary.
# + id="b1YPa6UhS8Es" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels) , (test_images, test_labels) = mnist.load_data()
training_images = training_images/255.0
test_images = test_images/255.0
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(256, activation=tf.nn.relu),
tf.keras.layers.Dense(5, activation=tf.nn.softmax)])
model.compile(optimizer = 'adam',
loss = 'sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
classifications = model.predict(test_images)
print(classifications[0])
print(test_labels[0])
# + id="sE7PDe6LWAHb" colab_type="code" colab={}
# + [markdown] id="Bql9fyaNUSFy" colab_type="text"
# #Exercise 6:
#
# Consider the impact of training for more or less epochs. Why do you think that would be the case?
#
# Try 15 epochs -- you'll probably get a model with a much better loss than the one with 5
# Try 30 epochs -- you might see the loss value stops decreasing, and sometimes increases. This is a side effect of something called 'overfitting' which you can learn about [somewhere] and it's something you need to keep an eye out for when training neural networks. There's no point in wasting your time training if you aren't improving your loss, right! :)
# + id="uE3esj9BURQe" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels) , (test_images, test_labels) = mnist.load_data()
training_images = training_images/255.0
test_images = test_images/255.0
model = tf.keras.models.Sequential([tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(5, activation=tf.nn.softmax)])
model.compile(optimizer = 'adam',
loss = 'sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=30)
model.evaluate(test_images, test_labels)
classifications = model.predict(test_images)
print(classifications[34])
print(test_labels[34])
# + [markdown] id="HS3vVkOgCDGZ" colab_type="text"
# #Exercise 7:
#
# Before you trained, you normalized the data, going from values that were 0-255 to values that were 0-1. What would be the impact of removing that? Here's the complete code to give it a try. Why do you think you get different results?
# + id="JDqNAqrpCNg0" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
mnist = tf.keras.datasets.mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images/255.0
test_images=test_images/255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=5)
model.evaluate(test_images, test_labels)
classifications = model.predict(test_images)
print(classifications[0])
print(test_labels[0])
# + [markdown] id="E7W2PT66ZBHQ" colab_type="text"
# #Exercise 8:
#
# Earlier when you trained for extra epochs you had an issue where your loss might change. It might have taken a bit of time for you to wait for the training to do that, and you might have thought 'wouldn't it be nice if I could stop the training when I reach a desired value?' -- i.e. 95% accuracy might be enough for you, and if you reach that after 3 epochs, why sit around waiting for it to finish a lot more epochs....So how would you fix that? Like any other program...you have callbacks! Let's see them in action...
# + id="pkaEHHgqZbYv" colab_type="code" colab={}
import tensorflow as tf
print(tf.__version__)
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if(logs.get('loss')<0.4):
print("\nReached 60% accuracy so cancelling training!")
self.model.stop_training = True
callbacks = myCallback()
mnist = tf.keras.datasets.fashion_mnist
(training_images, training_labels), (test_images, test_labels) = mnist.load_data()
training_images=training_images/255.0
test_images=test_images/255.0
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy')
model.fit(training_images, training_labels, epochs=5, callbacks=[callbacks])
| dlaicourse/Course 1 - Part 4 - Lesson 2 - Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A1: Data Curation
# ___
# ## Step 1: Data Acquisition
# Here we import all necessary Python libraries for this assignment
# +
import json
import requests
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
plt.rcParams["figure.figsize"] = "25,10"
plt.rcParams["legend.fontsize"] = 16
plt.rcParams["axes.labelsize"] = 16
# -
# These are the necessary endpoints for collecting page counts and page views. For each API, we have formatted it such that we can insert different values to gather different data based on access-site
# +
endpoint_pagecounts = 'https://wikimedia.org/api/rest_v1/metrics/legacy/pagecounts/aggregate/{project}/{access-site}/{granularity}/{start}/{end}'
endpoint_pageviews = 'https://wikimedia.org/api/rest_v1/metrics/pageviews/aggregate/{project}/{access-site}/{agent}/{granularity}/{start}/{end}'
# -
# Here, we make the actual API calls to collect the json data for each access type. This data is stored in the `data_raw` folder as json files
# +
params_template = {'project' : 'en.wikipedia.org',
'granularity':'monthly'}
headers = {
'User-Agent': 'https://github.com/sandeeptiwari6',
'From': '<EMAIL>'
}
def api_call(endpoint,parameters):
call = requests.get(endpoint.format(**parameters), headers=headers)
response = call.json()
return response
for access in ["desktop-site", "mobile-site", "all-sites"]:
params_template["access-site"] = access
params_template['start'] = '2007120100'
params_template['end'] = '2016080100'
json_data = api_call(endpoint_pagecounts, params_template)
outfile_path = f'data_raw/pagecounts_{access}_200712-201607.json'
with open(outfile_path, "w") as f:
json.dump(json_data, f)
for access in ["desktop", "mobile-app", "mobile-web"]:
params_template["access-site"] = access
params_template['start'] = '2015070100'
params_template['end'] = '2021080100'
params_template['agent'] = 'user'
json_data = api_call(endpoint_pageviews, params_template)
outfile_path = f'data_raw/pageviews_{access}_201507-202109.json'
with open(outfile_path, "w") as f:
json.dump(json_data, f)
# -
# ## Step 2: Data Preprocessing
# Here we begin the data preprocessing steps. Because the Pageview API splits the mobile views based on web and app, we need to consolidate these numbers into a sum total count of all mobile pageviews.
# +
mobile_total_views = []
with open('data_raw/pageviews_mobile-web_201507-202109.json') as f:
mobile_web_data = json.load(f)
with open('data_raw/pageviews_mobile-app_201507-202109.json') as f:
mobile_app_data = json.load(f)
nitems = len(mobile_web_data['items'])
for i in range(nitems):
dic = {}
d_web = mobile_web_data['items'][i]
d_app = mobile_app_data['items'][i]
year = d_web['timestamp'][:4]
month = d_web['timestamp'][4:6]
try:
total = d_web['count'] + d_app['count']
except:
total = d_web['views'] + d_app['views']
dic['month'] = month
dic['year'] = year
dic['pageview_mobile_views'] = total
mobile_total_views.append(dic)
mobile_total_df = pd.DataFrame(mobile_total_views)
mobile_total_df.head()
# -
# Here, we collect the pagecounts for each access type and store them into a singular dataframe that specifically contains page counts.
# +
counts_by_date = {}
with open('data_raw/pagecounts_all-sites_200712-201607.json') as f:
data = json.load(f)
items = data['items']
for dic in items:
date = dic['timestamp']
year = date[:4]
month = date[4:6]
if (month, year) not in counts_by_date:
counts_by_date[(month, year)] = {'month': month, 'year': year}
counts_by_date[(month, year)]['pagecount_all_views'] = dic['count']
with open('data_raw/pagecounts_desktop-site_200712-201607.json') as f:
data = json.load(f)
items = data['items']
for dic in items:
date = dic['timestamp']
year = date[:4]
month = date[4:6]
if (month, year) not in counts_by_date:
counts_by_date[(month, year)] = {'month': month, 'year': year}
counts_by_date[(month, year)]['pagecount_desktop_views'] = dic['count']
with open('data_raw/pagecounts_mobile-site_200712-201607.json') as f:
data = json.load(f)
items = data['items']
for dic in items:
date = dic['timestamp']
year = date[:4]
month = date[4:6]
if (month, year) not in counts_by_date:
counts_by_date[(month, year)] = {'month': month, 'year': year}
counts_by_date[(month, year)]['pagecount_mobile_views'] = dic['count']
pagecounts = pd.DataFrame(list(counts_by_date.values()))
# -
# We do the same, putting the views for desktop page views into a singular dataframe.
# +
views_by_date = {}
with open('data_raw/pageviews_desktop_201507-202109.json') as f:
data = json.load(f)
items = data['items']
for dic in items:
date = dic['timestamp']
year = date[:4]
month = date[4:6]
if (month, year) not in views_by_date:
views_by_date[(month, year)] = {'month': month, 'year': year}
views_by_date[(month, year)]['pageview_desktop_views'] = dic['views']
pageviews = pd.DataFrame(list(views_by_date.values()))
pageviews.head()
# -
# We then merge these dataframes together to create our final Pandas dataframe. At the end of this step, we have a single CSV-formatted data file that can be used in later analysis with no significant additional processing.
final_df = pagecounts.merge(pageviews, on=['month', 'year'], how='outer')
final_df = final_df.merge(mobile_total_df, on=['month', 'year'], how='outer')
final_df['pageview_all_views'] = final_df['pageview_desktop_views'] + final_df['pageview_mobile_views']
final_df = final_df[['year',
'month',
'pagecount_all_views',
'pagecount_desktop_views',
'pagecount_mobile_views',
'pageview_all_views',
'pageview_desktop_views',
'pageview_mobile_views']]
final_df.head()
# We save the dataframe to a CSV file.
final_csv_file = 'data_clean/en-wikipedia_traffic_200712-202108.csv'
final_df.to_csv(final_csv_file, sep=',')
# ## Step 3: Analysis
# This block of code generates the resulting time series graph using the dataframe from the previous step
# +
fig, ax = plt.subplots()
final_df['timestamp'] = final_df['year'] + final_df['month'] + '01'
final_df['timestamp'] = pd.to_datetime(final_df['timestamp'])
ax.plot_date(final_df['timestamp'], final_df['pagecount_all_views'], '--', color='black', label='total pagecount')
ax.plot_date(final_df['timestamp'], final_df['pagecount_mobile_views'], '--', color='blue', label='mobile site pagecount')
ax.plot_date(final_df['timestamp'], final_df['pagecount_desktop_views'], '--',color='green', label = 'main site pagecount')
ax.plot_date(final_df['timestamp'], final_df['pageview_all_views'], '-', color='black', label='total pageviews')
ax.plot_date(final_df['timestamp'], final_df['pageview_mobile_views'], '-', color='blue', label='mobile site pageviews' )
ax.plot_date(final_df['timestamp'], final_df['pageview_desktop_views'], '-',color='green', label='main site pageviews')
plt.title('Page Counts and Page Views on Wikipedia (2007-2021) ', fontsize=16);
plt.xlabel('Time');
plt.ylabel('Views');
ax.legend(loc='upper left');
plt.savefig('results/final_ts_graph.png')
| hcds-a1-data-curation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading an OBO ontology into a `networkx` graph
# Ontology are commonly represented as knowledge graphs, and the OBO language can express an ontology as an RDF graph, with terms as nodes and relationships as edges.
#
# As an example, here is how the *hexose biosynthetic process* and its superclasses in the [Gene Ontology](http://geneontology.org/docs/ontology-documentation/) can be represented as a graph, with edges showing the `rdfs:subClassOf` relationships:
#
# [](https://amigo.geneontology.org/amigo/term/GO:0019319)
#
#
# `fastobo` can be used to load the data from an OBO document and to build a graph using [NetworkX](https://networkx.org/), a graph library for Python. For this example, we will use the [Phenotype And Trait Ontology](http://www.obofoundry.org/ontology/pato.html). Let's start by creating an empty `DiGraph`:
import networkx
knowledge_graph = networkx.DiGraph()
# To load the latest version of PATO, we can use the Persistent URL form the OBO library which always resolves to the most up-to-date version:
# +
import urllib.request
import fastobo
with urllib.request.urlopen("http://purl.obolibrary.org/obo/pato.obo") as response:
pato = fastobo.load(response)
# -
# Now, we can populate the knowledge graph with all the terms that were just loaded, and add a link for every `is_a` clause appearing:
for frame in pato:
if isinstance(frame, fastobo.term.TermFrame):
knowledge_graph.add_node(str(frame.id))
for clause in frame:
if isinstance(clause, fastobo.term.IsAClause):
knowledge_graph.add_edge(str(frame.id), str(clause.term))
# Most OBO ontologies are DAGs, and PATO should be one as well:
networkx.is_directed_acyclic_graph(knowledge_graph)
# Now, we can have a look at only a portion of the graph, just like in the GO term example. Let's use `networkx.descendants` to extract the superclasses of an arbitrary PATO term:
superclass_nodes = networkx.descendants(knowledge_graph, "PATO:0000297")
superclass_nodes.add("PATO:0000297")
super_graph = knowledge_graph.subgraph(superclass_nodes)
# We can then draw the subgraph of `PATO:0000297` and its superclasses using the drawing capabilities of `networkx`:
from networkx.drawing.nx_agraph import graphviz_layout
networkx.draw(super_graph, pos=graphviz_layout(super_graph, prog="dot"), with_labels=True, arrows=True)
| docs/examples/graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Regression Modeling
# ## IMPORTS
from sklearn.linear_model import LinearRegression,Lasso,Ridge,ElasticNet
from sklearn.ensemble import GradientBoostingRegressor, RandomForestRegressor
from sklearn.cross_validation import train_test_split, cross_val_score
from sklearn.preprocessing import StandardScaler
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ## LOAD DATA
filename = 'insert file name here'
df = pd.read_csv(filename) # or read_json, read_sql, read_pickle, read_html, etc
df.head() # see head of data frame
# ## SPLIT INTO TRAIN/TEST
# - shuffle data at random state = 42
# - Train with 80% of data
# - Test with 20% of data
X = [] # features, should be matrix (or a vector if only 1 feature)
y = [] # target, should be vector
# X_std = StandardScaler().fit_transform(X) # scale if necessary
# y_std = StandardScaler().fit_transform(y) # scale if necessary
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.20,random_state=42)
# ## MACHINE LEARNING ALGORITHMS
models = {}
models['Ridge'] = Ridge()
models['Lasso'] = Lasso()
models['Elastic_Net'] = ElasticNet()
models['Linear_Regression'] = LinearRegression()
models['Random_Forest'] = RandomForestRegressor()
models['Gradient_Boost'] = GradientBoostingRegressor()
# ## TRAIN & SCORE MODELS
for name,model in models.items():
results = model.fit(X_train,y_train) #fit that model
y_pred = results.predict(X_test)
train_score = np.mean(cross_val_score(model,X_train,y_train, cv=4)) # 4 fold cross validation
test_score = results.score(X_test,y_test)
print 'MODEL:{} \t TRAIN_SCORE:{} \t TEST_SCORE:{}'.format(name,train_score,test_score)
# ## MODEL EVALUATION
def plot_analysis(name,y_true,y_pred):
figs,axs = plt.subplots(ncols=2,nrows=1)
figs.set_figwidth(15)
figs.set_figheight(10)
ax = axs[0]
residual = abs(y_true)-abs(y_pred)
ax.scatter(y_pred,residual)
ax.plot(np.linspace(min(residual),max(residual)),np.linspace(min(residual),max(residual))*0, '--r')
ax.set_title('Residual VS. Predicted (MODEL:{})'.format(name))
ax.set_xlabel('Predicted Yield')
ax.set_ylabel('Residual')
ax.grid(True)
ax = axs[1]
ax.scatter(y_true,y_pred)
ax.plot([min(y_true),max(true)],[min(y_true),max(y_true)],'--r')
ax.set_title('Predicted VS. True (MODEL:{})'.format(name))
ax.set_xlabel('True Yield')
ax.set_ylabel('Predicted Yield')
ax.grid(True)
| Python/regression_modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''ml3950'': conda)'
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.linear_model import SGDRegressor
import seaborn as sns
from sklearn.pipeline import Pipeline
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import Lasso
from sklearn.linear_model import Ridge
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import GridSearchCV
import linear_reg_demo_grad_desc
import time
# ## Linear Regression Redux and Cost
#
# We covered linear regression quite a bit in stats, it is one of the most simple and intuitive methods to make a predictive model - one that most of us did intuitively when doing a best fit line back in math class.
#
# The linear regression process we looked at used the algorithm of Linear Least Squares, which tries to create a model (the line) that has the lowest average squared distance from all the points. That is, the MSE between the model and the real values is a measurement of "badness" of the model - the smaller this value is, the better the model. We can also think of this MSE calculation as something called a Cost Function - the higher the cost, the worse the model is.
#
# The fitting part of a linear regression model is an attempt to minimize this cost function. The algorithm looks for the parameters (not hyperparameters) that minimize the cost; in a linear regression those parameters are the coefficients and the y-intercept.
#
# The linear least sqaures can calculate this directly, so the process of "looking" for the minimum is only one step. If you think back to logistic regression we can see a more dramatic example, the algorithm needs to test, try, and repeat. A decision tree is another example, the gini/entropy is the cost, and the algorithm searches through all potential splits until it finds the one that is best. That type of iteritive process is really common.
#
# ### Cost Functions
#
# The idea of cost is something that we will use throughout machine learning, it is critical for setting a goal that the algorithm can aim for when training. The cost function is just some function that measures the amount of error in a model, the lower the cost, the better a model we have. Usually this cost function is something that is a regular error metric that we have seen before - something like mean squared error for regression problems and accuracy or, more likely, cross-entropy for classifications. The cost to use is often something that we can specify as a hyperparameter when we create our models, such as the choice between gini and entropy in a tree.
#
# The cost function does not inherently need to be a regular error calculation, it could be almost any calculation at all. In certain weird situations the "goodness" of a model might not be measured accurately by a normal error calculation. Suppose you made a model to predict things on The Price is Right (https://priceisright.fandom.com/wiki/One_Bid); you want the model to make predictions that are close to the real price (probably well measured by MSE), however if the model predicts a price that is over, that's a tragedy and needs to be penalized severely. Maybe you'd want a model that calculated something more odd - MSE if the prediction is less than the real value, and residual^4 is the prediction is higher.
#
# #### Linear Regression Cost Function
#
# The cost function is just our old friend the Mean Squared Error:
#
# 
#Generate some random data
X1 = 2 * np.random.rand(1000, 1)
y1 = 4 + 3 * X1**3 + 3*np.random.randn(1000, 1)
# Generate a simple linear regression:
# +
lin_reg = LinearRegression()
lin_reg = lin_reg.fit(X1, y1)
preds1 = lin_reg.predict(X1)
d1 = pd.DataFrame(X1, columns={"X"})
d1["Y_pred"] = preds1
d1["Y_real"] = y1
sns.lineplot(data = d1, x="X", y="Y_pred", color="red")
sns.scatterplot(data=d1, x="X", y="Y_real")
# -
# #### Regression Results
#
# If we plot some data and generate a regression above, we have a solution that minimizes our cost function. There is no other model that we can create that has a lower mean squared residual between the model's predictions and the real values. Calculating the optimal model like we do here is great, however there are a couple of issues with that:
#
# <ul>
# <li>The computation cost when we have lots of data can become very large - growing exponentially. This can really matter when data grows massive.
# <li>Models that are not linear regression often can't be directly calculated (such as logistic regression). This is very common and we'll use gradient descent for things like neural networks later on.
# </ul>
#
# To deal with situations where we can't directly compute the optimal solution we need a different approach. Rather than determining the correct solution directly, we will make an attempt, evaluate the cost, make a slightly different attempt, and try to improve until we can't find a better cost score with subsequent attempts. This approach is called...
#
# ## Gradient Descent
#
# Gradient descent is a common approach to hunt for optimal solutions through an iteritive process. The process we can follow is:
#
# <ul>
# <li>Make an initial attempt to create a model, calculate the cost. This starting point is often random.
# <li>Compute the gradient - the derivitive or slope of the curve at that point. This indicates which direction to move.
# <li>Adjust the previous attempt, calculate the cost, compare to previous. The adjustment amount is determined by a value called the learning rate.
# <li>Repeat the previous step until moving does not improve the cost.
# </ul>
#
# We can visualize this process by looking at a curve of the cost function, and thinking about its derivitive or the slope. The gradient tells us two things:
# <ul>
# <li>Have we reached the best solution? If so, the gradient will be 0 indicating that we are at the minimum point on the curve. (i.e. there is no slope = we are at the bottom)
# <li>Which direction to go? We always want to go down the curve.
# </ul>
#
# 
#
# When we've "settled" at the bottom, that is the lowest amount of cost, so there are no moves to make to find a better model.
#
# #### Gradient Descent - From Scratch
#
# We can illustrate how gradient descent works. The file linear_reg_demo_grad_desc.py has an implementation of a linear regression that does gradient descent. I have modified it to return the set of predictions for each step of the training process. So what is happening in the background is:
# <ul>
# <li>Generate a linear regression.
# <li>Calculate the gradients.
# <li>Update the weights to move against the gradient (down the curve).
# <ul>
# <li>The size of the jump is defined by the learning rate. Big rate, big move!
# </ul>
# <li>Repeat until the trials are done. (In real implementations you'll also stop when improvement ends.)
# </ul>
#
# This example is what is called Batch Gradient Descent - at every step the entire process is recalculated.
#
# Play around with the learning rates and iterations and see how it progresses:
# +
#Play around with some options and see the results!
learn_rate = .05 #Learning rate - how large to move each update.
iterations = 100 #How many iterations to run.
num_show = 10 #How ofen to chart the line, i.e. 1 = every prediction, 10 = every 10th, etc..
train_rmses = []
test_rmses =[]
iters = []
plt.figure(figsize=(20,10))
X_train, X_test, y_train, y_test = train_test_split(X1, y1)
linreg = linear_reg_demo_grad_desc.LinearRegressionDemo(learning_rate=learn_rate, n_iters=iterations)
preds, test_preds = linreg.fit(X_train, y_train.ravel(), X_test)
sns.scatterplot(data=d1, x="X", y="Y_real")
for i in range(len(preds)):
d_tmp = pd.DataFrame(X_train, columns={"X"})
d_tmp["Y_pred"] = preds[i]
iters.append(i)
train_err = mean_squared_error(y_train, preds[i])
train_rmses.append(train_err)
test_err = mean_squared_error(y_test, test_preds[i])
test_rmses.append(test_err)
label = str(i) + " - RMSE:" + str(round(train_err, 3))
if (i%num_show) == 0:
sns.lineplot(data = d_tmp, x="X", y="Y_pred", label=label)
d_iters = pd.DataFrame(iters, columns={"Iteration"})
d_iters["Train"] = train_rmses
d_iters["Test"] = test_rmses
# -
# ### Key Code Bits
#
# We can look at the .py file and examine some of the key bits (you can open it directly for details and full code)
#
# <b>perform gradient descent for n iterations</b>
#
# ```python
# for _ in range(self.n_iters):
# # get y_prediction
# y_pred = self._get_prediction(X)
# # compute gradients
# dw, db = self._get_gradients(X, y, y_pred)
# # update weights & bias with gradients
# self._update_params(dw, db)
# ```
# Here the fit function is pretty simple, we make a prediction, take that prediction to compute the gradients (slopes), then make and update to the weights...
#
# ```python
# def _update_params(self, dw, db):
# self.weights -= self.lr * dw
# self.bias -= self.lr * db
# ```
# Updating the weights is just incrementing them by the amount of the learning rate.
#
# ```python
# def _get_gradients(self, X, y, y_pred):
# # get distance between y_pred and y_true
# error = y_pred - y
# # compute the gradients of weight & bias
# dw = (1 / self.n_samples) * np.dot(X.T, error)
# db = (1 / self.n_samples) * np.sum(error)
# return dw, db
# ```
# Calculating the gradients is just a recalculation of the gradients at our new point on the cost curve. The math uses vector math (dot products) that we can ignore for now.
#
# So the overall process, repeated for N iterations is to create a prediction (make a linear regression model), calculate the gradients on the cost curve at that point, update our position based on the position by the amount of the learning rate, and repeat. Eventually we will reach a point where the gradients, the slope of the location on the cost curve, is 0, then we won't be updating anymore - think about why by looking at the _update_params function.
# #### Errors
#
# We can visualize the errors by iteration, or epoch as it is often called. Here we can see how long it takes for us to narrow in on a solution.
#
# I've graphed both the test and train errors, here they tend to be extremely close and often flip-flop in terms of accuracy depending on random splits. This is not a constant pattern, it is due to the data.
plt.figure(figsize=(20,5))
sns.lineplot(data=d_iters, x="Iteration", y="Train", label="Train")
sns.lineplot(data=d_iters, x="Iteration", y="Test", label="Test")
# Our algorithm will work its way towards the solution.
#
# ### Errors and Gradients
#
# For a simple linear regression with one feature like we have here, the challenge is pretty simple. Our cost will be convex (shaped like a bowl), and we will always be able to find some solution, even if it takes a long time. With more complex datasets this isn't always true, we don't have a simple 2D curve, we have something that is in X dimension - maybe 100 or more at times. We can end up with curves that resemble this:
#
# 
#
# Here we have things like a local minima - a point at which the cost is minimized, but only locally, not overall. We don't want the algorithm to get 'stuck' in one of these holes, because we'll find a low cost, but not the lowest.
#
# Dealing with issues like this is there the learning rate comes in. By ensuring the learning rate is large enough, we have our attempts 'jump around' a little. This results in progress towards the minimum cost that is a little slower, but it also gives the function the ability to 'jump out' of problems like local minima. There's a balance with the learning rate, not too high, not too low.
# ## Other Types of Descents
#
# Batch gradient descent, like we did in the demo, suffers from the drawback that there is a lot of math involved in calculating the gradients, this can be slow with large amounts of data. In practice, there are a couple of things that we can do to speed this up.
#
# In the sample each time we iterated we calculated all of those gradients. For simple 2 varible datasets that's no big deal, but when the data gets massive, that's pretty slow. There are other implementations of gradient descent that apply the same concept, but take some shortcuts to lessen the amount of calculations needed. We can visualize how each 'hunts' the solution - the batch descent progresses steadily, the stochastic jumps around semi-randomly, and the mini-batch is between the two:
#
# 
#
# #### Stochastic Gradient Descent
#
# Stochastic (randomized) Gradient Descent, or SGD, speeds things up by just randomly selecting an instance and using that for the gradient calculation. The speed up comes from that massive reductions in the number of gradients calculated - instead of doing it for each record, SGD does it only once. Much faster.
#
# The downside to this is that there is much more randomness, and progress towards the solution isn't linear since you might randomly choose a point that is out of the way. This introduces some noise to the process, but the savings in computation time makes it generally worth it.
#
# #### Mini Batch Gradient Descent
#
# MBGD combines the ideas of batch and SGD - at each step a small random subset of the data is used for the gradient calculations.
#
# #### Gradients and Outcomes
#
# Note that the idea of all these algoritms is to generate a final model that is nearly the same, only the path there is different. Gradient descent isn't attempting to find a better model than linear regression (in a regular linear regression we can calculate the best model in a closed for solution), it is attempting to use a different method to reach that goal model. This idea of "narrowing in" on a solution can become even more useful when we don't have a solution that can be directly calculated - as long as we can set the cost function that defines accuracy, we can work our way towards an optimal solution.
#
#
#
# 
# ### SGD in Practice
#
# We do not need to implement this descent process by hand in practice, we can use the built in SKlearn modules, and the idea is also built into several other algorithms. For linear regression we can use SGDregressor, a generic implementation of SGD.
#
#
# We can do some gradient descent with actual data.
df = pd.read_csv("data/bodyfat.csv")
df.describe().T
#Create Datasets
y2 = np.array(df["BodyFat"]).reshape(-1,1)
X2 = np.array(df.drop(columns={"BodyFat"}))
X_train2, X_test2, y_train2, y_test2 = train_test_split(X2, y2, test_size=.3)
# ### Linear Regression Control Sample
#
# We can generate a standard linear regression model first, and see what the results are.
# +
#Closed Form Linear Regression
pipe_LR_steps = [('scale', MinMaxScaler()), ('model', LinearRegression())]
pipe_LR = Pipeline(pipe_LR_steps)
start = time.process_time()
pipe_LR.fit(X_train2, y_train2)
print(time.process_time() - start)
print('Training CrossVal Score:', cross_val_score(pipe_LR, X_train2, y_train2, cv=5))
print('Testing score:', pipe_LR.score(X_test2, y_test2))
# -
# ### SGD Regression
#
# Now we can create a model using gradient descent. There are a few options that we can specify in the SGD that are important:
# <ul>
# <li>eta0 - the initial learning rate.
# <li>learning_rate - how the learning rate is managed. In the examples we made by hand, the learning rate was constant. The SGD implementation provides for a way to adapt the learning rate - as you get closer to a solution, the learning rate slows. The defult is "invscaling", defined as: eta = eta0 / pow(t, power_t). t = number of updates, power_t = the exponent for inverse scaling learning rate.
# <ul>
# <li>This has the effect of using large learning rates to quickly narrow down to a close solution, hopefully taking advantage of both that speed increase and the ability to "jump" out of local minima. As the algorithm progresses, the rate slows to close in directly on a solution.
# </ul>
# <li>early_stopping - should the algorithm stop when it fails to improve. This will set aside a validation dataset, and if predictions for this set stop improving, end the training. If we look back to the demo, at some point the error flattens out and doesn't change much, early stopping stops at this point. We'll look at early stopping in more detail later on.
# <li>penalty - SGD applies regularization by default, which we'll discuss next time.
# </ul>
#
# <b>Note: Scaling values is very important in SGD algorithms. We'll probably get poor results if we forget.</b>
# +
#SGD Regressor.
pipe_SGD_steps = [('scale', MinMaxScaler()), ('model', SGDRegressor(max_iter=10000, eta0=.1))]
pipe_SGD = Pipeline(pipe_SGD_steps)
start = time.process_time()
pipe_SGD.fit(X_train2, y_train2.ravel())
print(time.process_time() - start)
#Print best model and test score
print('Training CrossVal Score:', cross_val_score(pipe_SGD, X_train2, y_train2.ravel(), cv=5))
print('Testing score:', pipe_SGD.score(X_test2, y_test2))
# -
# Note in the two examples above I added a timer. Play around with the learning rate and see what impact it has on the speed of SGD as compared to regular linear regression.
# ## Working Example
#
# Predict car prices using an SGD Regressor.
df3 = pd.read_csv("data/CarPrice_Assignment.csv")
df3.head()
#Describe/info data - look for data types as compared with dictionary
# <b>Note: There is probably a large amount of multicollinearity here. We'll try this example again with regularization soon, which is one way to deal with it in practice</b>
#
# Also, depending on randomness there may be a need to deal with rare categorical values. Keep this in mind, you should be able to find 2+ solutions by Googling the error.
# Start to preprocess and model.
# Use columnn transformer as there are mixed feature types
| 006_grad_dec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# We observed that time scaling matters *a lot* in convergence properties. Here we try and figure out why...
# +
from phimal_utilities.analysis import load_tensorboard
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import numpy as np
from natsort import natsorted
# #%matplotlib inline
# %config InlineBackend.figure_format='retina'
# -
# # With differentiable time scaling
# +
import pandas as pd
from tensorboard.backend.event_processing.event_accumulator import EventAccumulator
import os
def load_tensorboard(path):
'''Function to load tensorboard file from a folder.
Assumes one file per folder!'''
event_paths = [file for file in os.walk(path, topdown=True) if file[2][0][:len('events')] == 'events']
df = pd.DataFrame()
steps = None # steps are the same for all files
for event_idx, path in enumerate(event_paths):
summary_iterator = EventAccumulator(os.path.join(path[0], path[2][0])).Reload()
tags = summary_iterator.Tags()['scalars']
data = [[event.value for event in summary_iterator.Scalars(tag)] for tag in tags]
if steps is None:
steps = [event.step for event in summary_iterator.Scalars(tags[0])]
# Adding to dataframe
tags = [tag.replace('/', '_') for tag in tags] # for name consistency
if event_idx > 0: # We have one file in the top level, so after we need to use folder name
tags = [path[0].split('/')[-1]]
for idx, tag in enumerate(tags):
df[tag] = data[idx]
df.index = steps
return df, steps
# -
df, steps = load_tensorboard('runs/dt_differentiable/')
df.index = steps
df
class results:
def __init__(self, path):
self.df = load_tensorboard(path)
self.keys = self.get_keys()
def get_keys(self):
mse_keys = natsorted([key for key in self.df.keys() if key[:len('loss_mse')] == 'loss_mse'])
reg_keys = natsorted([key for key in self.df.keys() if key[:len('loss_reg')] == 'loss_reg'])
l1_keys = natsorted([key for key in self.df.keys() if key[:len('loss_l1')] == 'loss_l1'])
loss_key = 'loss_loss'
coeff_keys = natsorted([key for key in self.df.keys() if key[:len('coeffs')] == 'coeffs'])
unscaled_coeff_keys = natsorted([key for key in self.df.keys() if key[:len('unscaled')] == 'unscaled'])
estimator_coeff_keys = natsorted([key for key in self.df.keys() if key[:len('estimator')] == 'estimator'])
return {'mse': mse_keys, 'reg': reg_keys, 'l1': l1_keys, 'loss': loss_key,
'coeffs': coeff_keys, 'unscaled_coeffs': unscaled_coeff_keys, 'estimator_coeffs': estimator_coeff_keys}
def plot_losses(self):
fig, axes = plt.subplots(figsize=(12, 7), nrows=2, ncols=2, tight_layout=True)
ax = axes[0, 0]
ax.plot(self.df.index, self.df[self.keys['loss']])
ax.set_title('Loss')
ax = axes[0, 1]
for key in self.keys['mse']:
ax.semilogy(self.df.index, self.df[key], label=key[9:])
ax.set_title('MSE')
ax.legend()
ax.set_xlabel('Epoch')
ax = axes[1, 0]
for key in self.keys['reg']:
ax.semilogy(self.df.index, self.df[key], label=key[9:])
ax.set_title('MSE')
ax.legend()
ax.set_xlabel('Epoch')
ax = axes[1, 1]
for key in self.keys['l1']:
ax.semilogy(self.df.index, self.df[key], label=key[8:])
ax.set_title('MSE')
ax.legend()
ax.set_xlabel('Epoch')
def plot_coeffs(self):
fig, axes = plt.subplots(figsize=(15, 5), nrows=1, ncols=3, tight_layout=True)
ax = axes[0]
for key in self.keys['coeffs']:
ax.plot(self.df.index[1:], self.df[key][1:], label=key[7:])
ax.set_title('Coeffs')
ax.legend(ncol=2)
ax.set_xlabel('Epoch')
ax = axes[1]
for key in self.keys['unscaled_coeffs']:
ax.plot(self.df.index[1:], self.df[key][1:], label=key[16:])
ax.legend(ncol=2)
ax.set_title('Unscaled coeffs')
ax.set_xlabel('Epoch')
ax = axes[2]
for key in self.keys['estimator_coeffs']:
ax.plot(self.df.index[1:], self.df[key][1:], label=key[17:])
ax.legend(ncol=2)
ax.set_title('Estimator coeffs')
ax.set_xlabel('Epoch')
data = results('runs/dt_differentiable/')
data.plot_losses()
data.plot_coeffs()
for key in data.keys['coeffs']:
plt.plot(data.df[key], label=key)
#plt.plot(data.df[data.keys['coeffs'][3]], lw=3)
#plt.plot(data.df[data.keys['coeffs'][5]], lw=3)
plt.plot(np.arange(12), np.abs(data.df.tail(1)[data.keys['coeffs']].T), 'o')
for key in data.keys['unscaled_coeffs']:
plt.semilogy(np.abs(data.df[key]), label=key)
plt.semilogy(np.abs(data.df[data.keys['unscaled_coeffs'][3]]), lw=3)
plt.semilogy(np.abs(data.df[data.keys['unscaled_coeffs'][5]]), lw=3)
plt.plot(np.abs(data.df[data.keys['coeffs']]))
plt.plot(data.df[data.keys['unscaled_coeffs']])
# # Without differentiable time scaling
# # Without any scaling
| experiments/old/Timescaling/.ipynb_checkpoints/Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Network Models
#
# Probably the easiest kinds of statistical models for us to think about are the *network models*. These types of models (like the name imples) describe the random processes which you'd find when you're only looking at one network. We can have models which assume all of the nodes connect to each other essentially randomly, models which assume that the nodes are in distinct *communities*, and many more.
#
# The important realization to make about statistical models is that a model is *not* a network: it's the random process that *creates* a network. You can sample from a model a bunch of times, and because it's a random process, you'll end up with networks that look a little bit different each time -- but if you sampled a lot of networks and then averaged them, then you'd likely be able to get a reasonable ballpark estimation of what the model that they come from looks like.
#
# Let's pretend that we have a network, and the network is unweighted (meaning, we only have edges or not-edges) and undirected (meaning, edges connect nodes both ways). It'd have an adjacency matrix which consists of only 1's and 0's, because the only information we care about is whether there's an edge or not. The model that generated this network is pretty straightforward: there's just some universal probability that each node connects to each other node, and there are 10 nodes.
# + tags=["hide-input"]
import matplotlib.pyplot as plt
from graspologic.simulations import er_np
from graspologic.plot import binary_heatmap
# %config InlineBackend.figure_format = 'retina'
fig, ax = plt.subplots(figsize=(4,4))
n = 10
p = .5
A = er_np(n, p)
binary_heatmap(A, ax=ax, yticklabels=5, linewidths=1, linecolor="black", title="A small, simple network");
# -
# This small, simple network is one of many possible networks that we can generate with this model. Here are some more:
# + tags=["hide-input"]
fig, axs = plt.subplots(nrows=1, ncols=3, figsize=(12, 4))
for ax in axs.flat:
A = er_np(n, p)
hmap = binary_heatmap(A, ax=ax, yticklabels=5, linewidths=1, linecolor="black")
plt.suptitle("Three small, simple networks", fontsize=20)
# -
# One reasonable question to ask is how *many* possible networks we could make in this simple scenario? We've already made four, and it seems like there are more that this model could potentially generate.
#
# As it turns out, "more" is a pretty massive understatement. To actually figure out the number, think about the first node: there are two possibilities (weighted or not weighted), so you can generate two networks from a one-node model. Now, let's add an additional node. For each of the first two possibilities, there are two more -- so there are $2 \times 2 = 4$ total possible networks. Every node that we add doubles the number of networks - and since a network with $n$ nodes has $n \times n$ edges, the total number of possible networks ends up being $2^{n \times n} = 2^{n^2}$! So this ten-node model can generate $2^{10^2} = 2^{100}$ networks, which is, when you think carefully, an absurdly, ridiculously big number.
#
# Throughout many of the succeeding sections, we will attempt to make the content accessible to readers with, and without, a more technical background. To this end, we have added sections with trailing asterisks (\*). While we believe these sections build technical depth, we don't think they are critical to understanding many of the core ideas for network machine learning. In contrast with unstarred sections, these sections will assume familiarity with more advanced mathematical and probability concepts.
# ## Foundation*
# + active=""
# To understand network models, it is crucial to understand the concept of a network as a random quantity, taking a probability distribution. We have a realization $A$, and we think that this realization is random in some way. Stated another way, we think that there exists a network-valued random variable $\mathbf A$ that governs the realizations we get to see. Since $\mathbf A$ is a random variable, we can describe it using a probability distribution. The distribution of the random network $\mathbf A$ is the function $\mathbb P$ which assigns probabilities to every possible configuration that $\mathbf A$ could take. Notationally, we write that $\mathbf A \sim \mathbb P$, which is read in words as "the random network $\mathbf A$ is distributed according to $\mathbb P$."
#
# In the preceding description, we made a fairly substantial claim: $\mathbb P$ assigns probabilities to every possible configuration that realizations of $\mathbf A$, denoted by $A$, could take. How many possibilities are there for a network with $n$ nodes? Let's limit ourselves to simple networks: that is, $A$ takes values that are unweighted ($A$ is *binary*), undirected ($A$ is *symmetric*), and loopless ($A$ is *hollow*). In words, $\mathcal A_n$ is the set of all possible adjacency matrices $A$ that correspond to simple networks with $n$ nodes. Stated another way: every $A$ that is found in $\mathcal A$ is a *binary* $n \times n$ matrix ($A \in \{0, 1\}^{n \times n}$), $A$ is symmetric ($A = A^\top$), and $A$ is *hollow* ($diag(A) = 0$, or $A_{ii} = 0$ for all $i = 1,...,n$). Formally, we describe $\mathcal A_n$ as:
#
# \begin{align*}
# \mathcal A_n \triangleq \left\{A : A \textrm{ is an $n \times n$ matrix with $0$s and $1$s}, A\textrm{ is symmetric}, A\textrm{ is hollow}\right\}
# \end{align*}
#
# To summarize the statement that $\mathbb P$ assigns probabilities to every possible configuration that realizations of $\mathbf A$ can take, we write that $\mathbb P : \mathcal A_n \rightarrow [0, 1]$. This means that for any $A \in \mathcal A_n$ which is a possible realization of a random network $\mathbf A$, that $\mathbb P(\mathbf A = A)$ is a probability (it takes a value between $0$ and $1$). If it is completely unambiguous what the random variable $\mathbf A$ refers to, we might abbreviate $\mathbb P(\mathbf A = A)$ with $\mathbb P(A)$. This statement can alternatively be read that the probability that the random variable $\mathbf A$ takes the value $A$ is $\mathbb P(A)$. Finally, let's address that question we had in the previous paragraph. How many possible adjacency matrices are in $\mathcal A_n$?
#
# Let's imagine what just one $A \in \mathcal A_n$ can look like. Note that each matrix $A$ has $n \times n = n^2$ possible entries, in total, since $A$ is an $n \times n$ matrix. There are $n$ possible self-loops for a network, but since $\mathbf A$ is simple, it is loopless. This means that we can subtract $n$ possible edges from $n^2$, leaving us with $n^2 - n = n(n-1)$ possible edges that might not be unconnected. If we think in terms of a realization $A$, this means that we are ignoring the diagonal entries $a_{ii}$, for all $i \in [n]$. Remember that a simple network is also undirected. In terms of the realization $A$, this means that for every pair $i$ and $j$, that $a_{ij} = a_{ji}$. If we were to learn about an entry in the upper triangle of $A$ where $a_{ij}$ is such that $j > i$, note that we have also learned what $a_{ji}$ is, too. This symmetry of $A$ means that of the $n(n-1)$ entries that are not on the diagonal of $A$, we would, in fact, "double count" the possible number of unique values that $A$ could have. This means that $A$ has a total of $\frac{1}{2}n(n - 1)$ possible entries which are *free*, which is equal to the expression $\binom{n}{2}$. Finally, note that for each entry of $A$, that the adjacency can take one of two possible values: $0$ or $1$. To write this down formally, for every possible edge which is randomly determined, we have *two* possible values that edge could take. Let's think about building some intuition here:
# 1. If $A$ is $2 \times 2$, there are $\binom{2}{2} = 1$ unique entry of $A$, which takes one of $2$ values. There are $2$ possible ways that $A$ could look:
# \begin{align*}
# \begin{bmatrix}
# 0 & 1 \\
# 1 & 0
# \end{bmatrix}\textrm{ or }
# \begin{bmatrix}
# 0 & 0 \\
# 0 & 0
# \end{bmatrix}
# \end{align*}
# 2. If $A$ is $3 \times 3$, there are $\binom{3}{2} = \frac{3 \times 2}{2} = 3$ unique entries of $A$, each of which takes one of $2$ values. There are $8$ possible ways that $A$ could look:
# \begin{align*}
# &\begin{bmatrix}
# 0 & 1 & 1 \\
# 1 & 0 & 1 \\
# 1 & 1 & 0
# \end{bmatrix}\textrm{ or }
# \begin{bmatrix}
# 0 & 1 & 0 \\
# 1 & 0 & 1 \\
# 0 & 1 & 0
# \end{bmatrix}\textrm{ or }
# \begin{bmatrix}
# 0 & 0 & 1 \\
# 0 & 0 & 1 \\
# 1 & 1 & 0
# \end{bmatrix}
# \textrm{ or }\\
# &\begin{bmatrix}
# 0 & 1 & 1 \\
# 1 & 0 & 0 \\
# 1 & 0 & 0
# \end{bmatrix}\textrm{ or }
# \begin{bmatrix}
# 0 & 0 & 1 \\
# 0 & 0 & 0 \\
# 1 & 0 & 0
# \end{bmatrix}\textrm{ or }
# \begin{bmatrix}
# 0 & 0 & 0 \\
# 0 & 0 & 1 \\
# 0 & 1 & 0
# \end{bmatrix}\textrm{ or }\\
# &\begin{bmatrix}
# 0 & 1 & 0 \\
# 1 & 0 & 0 \\
# 0 & 0 & 0
# \end{bmatrix}\textrm{ or }
# \begin{bmatrix}
# 0 & 0 & 0 \\
# 0 & 0 & 0 \\
# 0 & 0 & 0
# \end{bmatrix}
# \end{align*}
#
# How do we generalize this to an arbitrary choice of $n$? The answer is to use *combinatorics*. Basically, the approach is to look at each entry of $A$ which can take different values, and multiply the total number of possibilities by $2$ for every element which can take different values. Stated another way, if there are $2$ choices for each one of $x$ possible items, we have $2^x$ possible ways in which we could select those $x$ items. But we already know how many different elements there are in $A$, so we are ready to come up with an expression for the number. In total, there are $2^{\binom n 2}$ unique adjacency matrices in $\mathcal A_n$. Stated another way, the *cardinality* of $\mathcal A_n$, described by the expression $|\mathcal A_n|$, is $2^{\binom n 2}$. The **cardinality** here just means the number of elements that the set $\mathcal A_n$ contains. When $n$ is just $15$, note that $\left|\mathcal A_{15}\right| = 2^{\binom{15}{2}} = 2^{105}$, which when expressed as a power of $10$, is more than $10^{30}$ possible networks that can be realized with just $15$ nodes! As $n$ increases, how many unique possible networks are there? In the below figure, look at the value of $|\mathcal A_n| = 2^{\binom n 2}$ as a function of $n$. As we can see, as $n$ gets big, $|\mathcal A_n|$ grows really really fast!
# + tags=["hide-input"]
import seaborn as sns
import numpy as np
from math import comb
n = np.arange(2, 51)
logAn = np.array([comb(ni, 2) for ni in n])*np.log10(2)
ax = sns.lineplot(x=n, y=logAn)
ax.set_title("")
ax.set_xlabel("Number of Nodes")
ax.set_ylabel("Number of Possible Graphs $|A_n|$ (log scale)")
ax.set_yticks([50, 100, 150, 200, 250, 300, 350])
ax.set_yticklabels(["$10^{{{pow:d}}}$".format(pow=d) for d in [50, 100, 150, 200, 250, 300, 350]])
ax;
# -
# So, now we know that we have probability distributions on networks, and a set $\mathcal A_n$ which defines all of the adjacency matrices that every probability distribution must assign a probability to. Now, just what is a network model? A **network model** is a set $\mathcal P$ of probability distributions on $\mathcal A_n$. Stated another way, we can describe $\mathcal P$ to be:
# \begin{align*}
# \mathcal P &\subseteq \{\mathbb P: \mathbb P\textrm{ is a probability distribution on }\mathcal A_n\}
# \end{align*}
#
# In general, we will simplify $\mathcal P$ through something called *parametrization*. We define $\Theta$ to be the set of all possible parameters of the random network model, and $\theta \in \Theta$ is a particular parameter choice that governs the parameters of a specific random network $\mathbf A$. In this case, we will write $\mathcal P$ as the set:
# \begin{align*}
# \mathcal P(\Theta) &\triangleq \left\{\mathbb P_\theta : \theta \in \Theta\right\}
# \end{align*}
# If $\mathbf A$ is a random network that follows a network model, we will write that $\mathbf A \sim \mathbb P_\theta$, for some choice $\theta$. We will often use the shorthand $\mathbf A \sim \mathbb P$.
#
# If you are used to traditional univariate or multivariate statistical modelling, an extremely natural choice for when you have a discrete sample space (like $\mathcal A_n$, which is discrete because we can count it) would be to use a categorical model. In the categorical model, we would have a single parameter for all possible configurations of an $n$-node network; that is, $|\theta| = \left|\mathcal A_n\right| = 2^{\binom n 2}$. What is wrong with this model? The limitations are two-fold:
# 1. As we explained previously, when $n$ is just $15$, we would need over $10^{30}$ bits of storage just to define $\theta$. This amounts to more than $10^{8}$ zetabytes, which exceeds the storage capacity of *the entire world*.
# 2. With a single network observed (or really, any number of networks we could collect in the real world) we would never be able to estimate $2^{\binom n 2}$ parameters for any reasonably non-trivial number of nodes $n$. For the case of one observed network $A$, an estimate of $\theta$ (referred to as $\hat\theta$) would simply be for $\hat\theta$ to have a $1$ in the entry corresponding to our observed network, and a $0$ everywhere else. Inferentially, this would imply that the network-valued random variable $\mathbf A$ which governs realizations $A$ is deterministic, even if this is not the case. Even if we collected potentially *many* observed networks, we would still (with very high probability) just get $\hat \theta$ as a series of point masses on the observed networks we see, and $0$s everywhere else. This would mean our parameter estimates $\hat\theta$ would not generalize to new observations at *all*, with high probability.
#
# So, what are some more reasonable descriptions of $\mathcal P$? We explore some choices below. Particularly, we will be most interested in the *independent-edge* networks. These are the families of networks in which the generative procedure which governs the random networks assume that the edges of the network are generated *independently*. **Statistical Independence** is a property which greatly simplifies many of the modelling assumptions which are crucial for proper estimation and rigorous statistical inference, which we will learn more about in the later chapters.
#
# ### Equivalence Classes*
#
# In all of the below models, we will explore the concept of the **likelihood equivalence class**, or an *equivalence class*, for short. The likelihood $\mathcal L$ is a function which in general, describes how effective a particular observation can be described by a random variable $\mathbf A$ with parameters $\theta$, written $\mathbf A \sim F(\theta)$. Formally, the likelihood is the function where $\mathcal L_\theta(A) \propto \mathbb P_\theta(A)$; that is, the likelihood is proportional to the probability of observing the realization $A$ if the underlying random variable $\mathbf A$ has parameters $\theta$. Why does this matter when it comes to equivalence classes? An equivalence class is a subset of the sample space $E \subseteq \mathcal A_n$, which has the following properties. Holding the parameters $\theta$ fixed:
#
# 1. If $A$ and $A'$ are members of the same equivalence class $E$ (written $A, A' \in E$), then $\mathcal L_\theta(A) = \mathcal L_\theta(A')$.
# 2. If $A$ and $A''$ are members of different equivalence classes; that is, $A \in E$ and $A'' \in E'$ where $E, E'$ are equivalence classes, then $\mathcal L_\theta(A) \neq \mathcal L_\theta(A'')$.
# 3. Using points 1 and 2, we can establish that if $E$ and $E'$ are two different equivalence classes, then $E \cap E' = \varnothing$. That is, the equivalence classes are **mutually disjoint**.
# 4. We can use the preceding properties to deduce that given the sample space $\mathcal A_n$ and a likelihood function $\mathcal L_\theta$, we can define a partition of the sample space into equivalence classes $E_i$, where $i \in \mathcal I$ is an arbitrary indexing set. A **partition** of $\mathcal A_n$ is a sequence of sets which are mutually disjoint, and whose union is the whole space. That is, $\bigcup_{i \in \mathcal I} E_i = \mathcal A_n$.
#
# We will see more below about how the equivalence classes come into play with network models, and in a later section, we will see their relevance to the estimation of the parameters $\theta$.
#
# ### Independent-Edge Random Networks*
#
# The below models are all special families of something called **independent-edge random networks**. An independent-edge random network is a network-valued random variable, in which the collection of edges are all independent. In words, this means that for every adjacency $\mathbf a_{ij}$ of the network-valued random variable $\mathbf A$, that $\mathbf a_{ij}$ is independent of $\mathbf a_{i'j'}$, any time that $(i,j) \neq (i',j')$. When the networks are simple, the easiest thing to do is to assume that each edge $(i,j)$ is connected with some probability (which may be different for each edge) $p_{ij}$. We use the $ij$ subscript to denote that this probability is not necessarily the same for each edge. This simple model can be described as $\mathbf a_{ij}$ has the distribution $Bern(p_{ij})$, for every $j > i$, and is independent of every other edge in $\mathbf A$. We only look at the entries $j > i$, since our networks are simple. This means that knowing a realization of $\mathbf a_{ij}$ also gives us the realizaaion of $\mathbf a_{ji}$ (and thus $\mathbf a_{ji}$ is a *deterministic* function of $\mathbf a_{ij}$). Further, we know that the random network is loopless, which means that every $\mathbf a_{ii} = 0$. We will call the matrix $P = (p_{ij})$ the **probability matrix** of the network-valued random variable $\mathbf A$. In general, we will see a common theme for the likelihoods of a realization $A$ of a network-valued random variable $\mathbf A$, which is that it will greatly simplify our computation. Remember that if $\mathbf x$ and $\mathbf y$ are binary variables which are independent, that $\mathbb P(\mathbf x = x, \mathbf y = y) = \mathbb P(\mathbf x = x) \mathbb P(\mathbf y = y)$. Using this fact:
#
# \begin{align*}
# \mathcal L_\theta(A) &= \mathbb P(\mathbf A = A) \\
# &= \mathbb P(\mathbf a_{11} = a_{11}, \mathbf a_{12} = a_{12}, ..., \mathbf a_{nn} = a_{nn}) \\
# &= \mathbb P(\mathbf a_{ij} = a_{ij} \text{ for all }j > i) \\
# &= \prod_{j > i}\mathbb P(\mathbf a_{ij} = a_{ij}), \;\;\;\;\textrm{Independence Assumption}
# \end{align*}
# Next, we will use the fact that if a random variable $\mathbf a_{ij}$ has the Bernoulli distribution with probability $p_{ij}$, that $\mathbb P(\mathbf a_{ij} = a_{ij}) = p_{ij}^{a_{ij}}(1 - p_{ij})^{1 - p_{ij}}$:
# \begin{align*}
# \mathcal L_\theta(A) &= \prod_{j > i}p_{ij}^{a_{ij}}(1 - p_{ij})^{1 - p_{ij}}
# \end{align*}
#
# Now that we've specified a likelihood and a very generalizable model, we've learned the full story behind network models and are ready to skip to estimating parameters, right? *Wrong!* Unfortunately, if we tried too estimate anything about each $p_{ij}$ individually, we would obtain that $p_{ij} = a_{ij}$ if we only have one realization $A$. Even if we had many realizations of $\mathbf A$, this still would not be very interesting, since we have a *lot* of $p_{ij}$s to estimate, and we've ignored any sort of structural model that might give us deeper insight into $\mathbf A$. In the below sections, we will learn successively less restrictive (and hence, *more expressive*) assumptions about $p_{ij}$s, which will allow us to convey fairly complex random networks, but *still* enable us with plenty of intteresting things to learn about later on.
# ## Erdös-Rényi (ER)
# + tags=["hide-input"]
from graspologic.plot import heatmap
from graspologic.simulations import er_np
n = 50 # network with 50 nodes
p = 0.3 # probability of an edge existing is .3
# sample a single simple adjacency mtx from ER(50, .3)
A = er_np(n=n, p=p, directed=False, loops=False)
# and plot it
binary_heatmap(A, title="$ER_{50}(0.3)$ Simulation")
plt.show()
# -
# The simplest random network model is called the Erdös Rényi (ER) model<sup>1</sup>. Consider a social network, where nodes represent students, and edges represent whether a pair of students arae friends. The simplest possible thing to do with our network would be to assume that a given pair of students within our network have the same chance of being friends as any other pair of people we select. The Erdös Rényi model formalizes this relatively simple model with a single parameter:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $p$ | $[0, 1]$ | Probability that an edge exists between a pair of nodes |
#
# In an Erdös Rényi network, each pair of nodes is connected with probability $p$, and therefore not connected with probability $1-p$. Statistically, we say that for each edge $\mathbf{a}_{ij}$, that $\mathbf{a}_{ij}$ is sampled independently and identically from a $Bern(p)$ distribution, whenever $j > i$. The word "independent" means that edges in the network occurring or not occurring do not affect one another. For instance, this means that if we knew a student named Alice was friends with Bob, and Alice was also friends with Chadwick, that we do not learn any information about whether Bob is friends with Chadwick. The word "identical" means that every edge in the network has the same probability $p$ of being connected. If Alice and Bob are friends with probability $p$, then Alice and Chadwick are friends with probability $p$, too. When $i > j$, we allow $\mathbf a_{ij} = \mathbf a_{ji}$. This means that the connections *across the diagonal* of the adjacency matrix are all equal, which means that we have built-in the property of undirectedness into our networks. Also, we let $\mathbf a_{ii} = 0$, which means that all self-loops are always unconnected. This means that all the networks are loopless, and consequently the adjacency matrices are hollow. If $\mathbf A$ is the adjacency matrix for an ER network with probability $p$, we write that $\mathbf A \sim ER_n(p)$.
# ### Practical Utility
#
# In practice, the ER model seems like it might be a little too simple to be useful. Why would it ever be useful to think that the best we can do to describe our network is to say that connections exist with some probability? Does this miss a *lot* of useful questions we might want to answer? Fortunately, there are a number of ways in which the simplicity of the ER model is useful. Given a probability and a number of nodes, we can easily describe the properties we would expect to see in a network if that network were ER. For instance, we know what the degree distribution of an ER network can look like. We can reverse this idea, too: given a network we think might *not* be ER, we could check whether it's different in some way from a network which is ER. For instance, if we see a half of the nodes have a very high degree, and the rest of the nodes with a much lower degree, we can reasonably conclude the network might be more complex than can be described by the ER model. If this is the case, we might look for other, more complex, models that could describe our network.
#
#
# ```{admonition} Working Out the Expected Degree in an Erdös-Rényi Network
# Suppose that $\mathbf A$ is a simple network which is random. The network has $n$ nodes $\mathcal V = (v_i)_{i = 1}^n$. Recall that the in a simple network, the node degree is $deg(v_i) = \sum_{j = 1}^n \mathbf a_{ij}$. What is the expected degree of a node $v_i$ of a random network $\mathbf A$ which is Erdös-Rényi?
#
# To describe this, we will compute the expectated value of the degree $deg(v_i)$, written $\mathbb E\left[deg(v_i)\right]$. Let's see what happens:
# \begin{align*}
# \mathbb E\left[deg(v_i)\right] &= \mathbb E\left[\sum_{j = 1}^n \mathbf a_{ij}\right] \\
# &= \sum_{j = 1}^n \mathbb E[\mathbf a_{ij}]
# \end{align*}
# We use the *linearity of expectation* in the line above, which means that the expectation of a sum with a finite number of terms being summed over ($n$, in this case) is the sum of the expectations. Finally, by definition, all of the edges $A_{ij}$ have the same distribution: $Bern(p)$. The expected value of a random quantity which takes a Bernoulli distribution is just the probability $p$. This means every term $\mathbb E[\mathbf a_{ij}] = p$. Therefore:
# \begin{align*}
# \mathbb E\left[deg(v_i)\right] &= \sum_{j = 1}^n p = n\cdot p
# \end{align*}
# Since all of the $n$ terms being summed have the same expected value. This holds for *every* node $v_i$, which means that the expected degree of all nodes is an undirected ER network is the same, $n \cdot p$.
# ```
#
# <!-- The ER model is also useful for the development of new computational techniques to use on random networks. This is because even if the "best" model for a network is something much more complex, we can still calculate an edge probability $p$ for the network without needing any information but the adjacency matrix. Consider, for instance, a case where we design a new algorithm for a social network, and we want to know how much more RAM we might need as the social network grows. We might want to investigate how the algorithm scales to networks with different numbers of people and different connection probabilities that might be realistic as our social network expands in popularity. Examining how the algorithm operates on ER networks with different values of $n$ and $p$ might be helpful. This is an especially common approach when people deal with networks that are said to be *sparse*. A **sparse network** is a network in which the number of edges is much less than the total possible number of edges. This contrasts with a **dense network**, which is a network in which the number of edges is close to the maximum number of possible edges. In the case of an $ER_{n}(p)$ network, the network is sparse when $p$ is small (closer to $0$), and dense when $p$ is large (closer to $1$). -->
#
# ### Code Examples
#
# In the next code block, we look to sample a single ER network with $50$ nodes and an edge probability $p$ of $0.3$:
# +
from graspologic.plot import heatmap
from graspologic.simulations import er_np
n = 50 # network with 50 nodes
p = 0.3 # probability of an edge existing is .3
# sample a single simple adjacency matrix from ER(50, .3)
A = er_np(n=n, p=p, directed=False, loops=False)
# and plot it
binary_heatmap(A, title="$ER_{50}(0.3)$ Simulation")
plt.show()
# -
# Above, we visualize the network using a heatmap. The dark red squares indicate that an edge exists between a pair of nodes, and white squares indicate that an edge does not exist between a pair of nodes.
# Next, let's see what happens when we use a higher edge probability, like $p=0.7$:
# +
p = 0.7 # network has an edge probability of 0.7
# sample a single adjacency matrix from ER(50, 0.7)
A = er_np(n=n, p=p, directed=False, loops=False)
# and plot it
binary_heatmap(A, title="$ER_{50}(0.7)$ Simulation")
plt.show()
# -
# As the edge probability increases, the sampled adjacency matrix tends to indicate that there are more connections in the network. This is because there is a higher chance of an edge existing when $p$ is larger.
#
# ### *Likelihood
#
# What is the likelihood for realizations of Erdös-Rényi networks? Remember that for Independent-edge graphs, that the likelihood can be written:
#
# \begin{align*}
# \mathcal L_{\theta}(A) &= \prod_{j > i} \mathbb P_\theta(\mathbf{a}_{ij} = a_{ij})
# \end{align*}
#
# Next, we recall that by assumption of the ER model, that the probability matrix $P = (p)$, or that $p_{ij} = p$ for all $i,j$. Therefore:
#
# \begin{align*}
# \mathcal L_\theta(A) &\propto \prod_{j > i} p^{a_{ij}}(1 - p)^{1 - a_{ij}} \\
# &= p^{\sum_{j > i} a_{ij}} \cdot (1 - p)^{\binom{n}{2} - \sum_{j > i}a_{ij}} \\
# &= p^{m} \cdot (1 - p)^{\binom{n}{2} - m}
# \end{align*}
#
# This means that the likelihood $\mathcal L_\theta(A)$ is a function *only* of the number of edges $m = \sum_{j > i}a_{ij}$ in the network represented by adjacency matrix $A$. The equivalence class on the Erdös-Rényi networks are the sets:
#
# \begin{align*}
# E_{i} &= \left\{A \in \mathcal A_n : m = i\right\}
# \end{align*}
#
# where $i$ index from $0$ (the minimum number of edges possible) all the way up to $n^2$ (the maximum number of edges possible). All of the relationships for equivalence classes discussed above apply to the sets $E_i$.
# ## Stochastic Block Model (SBM)
#
# Imagine that we are flipping a fair single coin. A *fair* coin is a coin in which the probability of seeing either a heads or a tails on a coin flip is $\frac{1}{2}$. Let's imagine we flip the coin $20$ times, and we see $10$ heads and $10$ tails.
#
# What would happen if we were to flip $2$ coins, which had a different probability of seeing heads or tails? Imagine that we flip each coin 10 times. The first 10 flips are with a fair coin, and we might see an outcome of five heads and five tails. On the other hand, the second ten flips are not with a fair coin, but with a coin that has a $\frac{4}{5}$ probability to land on heads, and a $\frac{1}{5}$ probability of landing on tails. In the second set of $10$ flips, we might see an outcome of nine heads and one tails.
#
# In the first set of 20 coin flips, all of the coin flips are performed with the same coin. Stated another way, we have a single *cluster*, or a set of coin flips which are similar. On the other hand, in the second set of twenty coin flips, twenty of the coin flips are performed with a fair coin, and ten of the coin flips are performed with a different coin which is not fair. Here, we have two *clusters* of coin flips, those that occur with the first coin, and those that occur with the second coin. Since the first cluster of coin flips are with a fair coin, we expect that coin flips from the first cluster will not necessarily have an identical number of heads and tails, but at least a similar number of heads and tails. On the other hand, coin flips from the second cluster will tend to have more heads than tails.
#
# What does this example have to do with networks? In the above examples, the two sets of coin flips differ in the number of coins with different probabilities that we use for the example. The first example has only one coin, whereas the second example has two coins with different probabilities of heads or tails. If we were to assume that the second example had been performed with only a single coin when in reality it was performed with two different coins, we would be unable to capture that the second 10 coin flips had a substantially different chance of landing on heads than the first ten coin flips. Just like coin flips can be performed with fundamentally different coins, the nodes of a network could also be fundamentally different. The way in which two nodes differ (or do not differ) sometimes holds value in determining the probability that an edge exists between them.
#
# To generalize this example to a network, let's imagine that we have $100$ students, each of whom can go to one of two possible schools: school $1$ or school $2$. Our network has $100$ nodes, and each node represents a single student. The edges of this network represent whether a pair of students are friends. Intuitively, if two students go to the same school, it might make sense to say that they have a higher chance of being friends than if they do not go to the same school. If we were to try to characterize this network using an ER network, we would run into a problem very similar to when we tried to capture the two cluster coin flip example with only a single coin. Intuitively, there must be a better way!
# The Stochastic Block Model, or SBM, captures this idea by assigning each of the $n$ nodes in the network to one of $K$ communities. A **community** is a group of nodes within the network. In our example case, the communities would represent the schools that students are able to attend in our network. In an SBM, instead of describing all pairs of nodes with a fixed probability like with the ER model, we instead describe properties that hold for edges between *pairs of communities*. In this sense, for a given school, we could think of the network that describes that school's students as ER. There are two types of SBMs: one in which the node-assignment vector is treated as *unknown* and one in which the node-assignment vector is treated as *known* (it is a *node attribute* for the network).
#
# ### *A Priori* Stochastic Block Model
#
# The *a priori* SBM is an SBM in which we know *a priori* (that is, ahead of time) which nodes are in which communities. Here, we will use the variable $K$ to denote the maximum number of different communities. The ordering of the communities does not matter; the community we call $1$ versus $2$ versus $K$ is largely a symbolic distinction (the only thing that matters is that they are *different*). The *a priori* SBM has the following parameter:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $B$ | [0,1]$^{K \times K}$ | The block matrix, which assigns edge probabilities for pairs of communities |
#
#
# To describe the *a priori* SBM, we will use a latent variable model. To do so, we will assume there is some vector-valued random variable, $\vec{\pmb \tau}$, which we will call the **node assignment vector**. This random variable takes values $\vec\tau$ which are in the space $\{1,...,K\}^n$. That means for each $\tau_i$ that is an element of a realization of $\vec{\pmb \tau}$, that $\tau_i$ takes on of $K$ possible values. Each node receives a community assignment, so we say that $i$ goes from $1$ to $n$. Stated another way, each node $i$ of our network receives an assignment $\tau_i$ to one of the $K$ communities. This model is called the *a priori* SBM because we use it when we have a realization $\vec\tau$ that we know ahead of time. In our social network example, for instance, $\tau_i$ would reflect that each student can attend one of two possible schools. For a single node $i$ that is in community $\ell$, where $\ell \in \{1, ..., K\}$, we write that $\tau_i = \ell$.
#
# Next, let's discuss the matrix $B$, which is known as the **block matrix** of the SBM. We write down that $B \in [0, 1]^{K \times K}$, which means that the block matrix is a matrix with $K$ rows and $K$ columns. If we have a pair of nodes and know which of the $K$ communities each node is from, the block matrix tells us the probability that those two nodes are connected. If our networks are simple, the matrix $B$ is also symmetric, which means that if $b_{kk'} = p$ where $p$ is a probability, that $b_{k'k} = p$, too. The requirement of $B$ to be symmetric exists *only* if we are dealing with simple networks, since they are undirected; if we relax the requirement of undirectedness (and allow directed networks) $B$ no longer need be symmetric.
#
# Finally, let's think about how to write down the generative model for the *a priori* SBM. Intuitionally what we want to reflect is, if we know that node $i$ is in community $\ell$ and node $j$ is in community $k$, that the $(\ell, k)$ entry of the block matrix is the probability that $i$ and $j$ are connected. We say that given $\tau_i = k'$ and $\tau_j = k$, $\mathbf a_{ij}$ is sampled independently from a $Bern(b_{k' k})$ distribution for all $j > i$. Note that the adjacencies $\mathbf a_{ij}$ are not *necessarily* identically distributed. Consider, for instance, another pair of nodes, $i'$ and $j'$, where $\tau_i = k'$ and $\tau_j = k'$. Then $\mathbf a_{i'j'}$ would have probability $b_{k'k'}$ instead of $b_{k'k}$, which specifies a different Bernoulli distribution (since the probabilities are different). If $\mathbf A$ is an *a priori* SBM network with parameter $B$, and $\vec{\tau}$ is a realization of the node-assignment vector, we write that $\mathbf A \sim SBM_{n,\vec \tau}(B)$.
#
# ### Code Examples
#
# We just covered a lot of intuition! This intuition will come in handy later, but let's take a break from the theory by working through an example. Say we have $300$ students, and we know that each student goes to one of two possible schools. We will begin by thinking about the *a priori* SBM, since it's a little more straightforward to generate samples. Remember the *a priori* SBM is the SBM where already have a realization of $\vec{\pmb \tau}$ ahead of time. We don't really care too much about the ordering of the students for now, so let's just assume that the first $150$ students all go to school $1$, and the second $150$ students all go to school $2$. Let's assume that the students from school $1$ are better friends in general than the students from school $2$, so we'll say that the probability of two students who both go to school $1$ being friends is $0.5$, and the probability of two students who both go to school $2$ being friends is $0.3$. Finally, let's assume that if one student goes to school $1$ and the other student goes to school $2$, that the probability that they are friends is $0.2$.
#
# ```{admonition} Thought Exercise
#
# Before you read on, try to think to yourself about what the node-assignment vector $\vec \tau$ and the block matrix $B$ look like.
# ```
#
# Next, let's plot what $\vec \tau$ and $B$ look like:
# + tags=["hide-input"]
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import matplotlib
def plot_tau(tau, title="", xlab="Node"):
cmap = matplotlib.colors.ListedColormap(["skyblue", 'blue'])
fig, ax = plt.subplots(figsize=(10,2))
with sns.plotting_context("talk", font_scale=1):
ax = sns.heatmap((tau - 1).reshape((1,tau.shape[0])), cmap=cmap,
ax=ax, cbar_kws=dict(shrink=1), yticklabels=False,
xticklabels=False)
ax.set_title(title)
cbar = ax.collections[0].colorbar
cbar.set_ticks([0.25, .75])
cbar.set_ticklabels(['School 1', 'School 2'])
ax.set(xlabel=xlab)
ax.set_xticks([.5,149.5,299.5])
ax.set_xticklabels(["1", "150", "300"])
cbar.ax.set_frame_on(True)
return
n = 300 # number of students
# tau is a column vector of 150 1s followed by 50 2s
# this vector gives the school each of the 300 students are from
tau = np.vstack((np.ones((int(n/2),1)), np.full((int(n/2),1), 2)))
plot_tau(tau, title="Tau, Node Assignment Vector",
xlab="Student")
# -
# So as we can see, the first $50$ students are from school $1$, and the second $50$ students are from school $2$. Next, let's look at the block matrix $B$:
# + tags=[]
K = 2 # 2 communities in total
# construct the block matrix B as described above
B = np.zeros((K, K))
B[0,0] = .5
B[0,1] = B[1,0] = .2
B[1,1] = .3
# + tags=["hide-input"]
def plot_block(X, title="", blockname="School", blocktix=[0.5, 1.5],
blocklabs=["School 1", "School 2"]):
fig, ax = plt.subplots(figsize=(8, 6))
with sns.plotting_context("talk", font_scale=1):
ax = sns.heatmap(X, cmap="Purples",
ax=ax, cbar_kws=dict(shrink=1), yticklabels=False,
xticklabels=False, vmin=0, vmax=1)
ax.set_title(title)
cbar = ax.collections[0].colorbar
ax.set(ylabel=blockname, xlabel=blockname)
ax.set_yticks(blocktix)
ax.set_yticklabels(blocklabs)
ax.set_xticks(blocktix)
ax.set_xticklabels(blocklabs)
cbar.ax.set_frame_on(True)
return
plot_block(B, title="Block Matrix")
plt.show()
# -
# As we can see, the matrix $B$ is a symmetric block matrix, since our network is undirected. Finally, let's sample a single network from the SBM with parameters $\vec \tau$ and $B$:
# +
from graspologic.simulations import sbm
from graspologic.plot import adjplot
import pandas as pd
# sample a graph from SBM_{300}(tau, B)
A = sbm(n=[int(n/2), int(n/2)], p=B, directed=False, loops=False)
meta = pd.DataFrame(
data = {"School": tau.reshape((n)).astype(int)}
)
ax=adjplot(A, meta=meta, color="School", palette="Blues")
# -
# The above network shows students, ordered by the school they are in (school 1 and school 2, respectively). As we can see in the above network, people from school $1$ are more connected than people from school $2$. We notice this from the fact that there are more connections between people from school $1$ than from school $2$. Also, the connections between people from different schools appear to be a bit *more sparse* (fewer edges) than connections betwen schools. The above heatmap can be described as **modular**: it has clear communities, which are the nodes that comprise the obvious "squares" in the above adjacency matrix.
#
# Something easy to mistake about the SBM is that the SBM will *not always* have the obvious modular structure defined above when we look at a heatmap. Rather, this modular structure is *only* made obvious because the students are ordered according to the school in which they are in. What do you think will happen if we look at the students in a random order? Do you think it will be obvious that the network will have a modular structure?
#
# The answer is: *No!* Let's see what happens when we use a reordering, called a *permutation* of the nodes, to reorder the nodes from the network into a random order:
# +
import numpy as np
# generate a permutation of the n nodes
vtx_perm = np.random.choice(n, size=n, replace=False)
meta = pd.DataFrame(
data = {"School": tau[vtx_perm].reshape((n)).astype(int)}
)
# same adjacency matrix (up to reorder of the nodes)
ax=adjplot(A[tuple([vtx_perm])] [:,vtx_perm], meta=meta, color="School", palette="Blues")
# -
# Notice that now, the students are *not* organized according to school. We can see this by looking at the school assignment vector, shown at the left and top, of the network. It becomes pretty tough to figure out whether there are communities in our network just by looking at an adjacency matrix, unless you are looking at a network in which the nodes are *already arranged* in an order which respects the community structure.
#
# In practice, this means that if you know ahead of time what natural groupings of the nodes might be (such knowing which school each student goes to) by way of your node attributes, you can visualize your data according to that grouping. If you don't know anything about natural groupings of nodes, however, we are left with the problem of *estimating community structure*. A later method, called the *spectral embedding*, will be paired with clustering techniques to allow us to estimate node assignment vectors.
# #### Likelihood*
#
# What does the likelihood for the *a priori* SBM look like? Fortunately, since $\vec \tau$ is a *parameter* of the *a priori* SBM, the likelihood is a bit simpler than for the *a posteriori* SBM. This is because the *a posteriori* SBM requires a marginalization over potential realizations of $\vec{\pmb \tau}$, whereas the *a priori* SBM does not, since we already know that $\vec{\pmb \tau}$ was realized as $\vec\tau$.
#
# Putting these steps together gives us that:
# \begin{align*}
# \mathcal L_\theta(A) &\propto \mathbb P_{\theta}(\mathbf A = A | \vec{\pmb \tau} = \vec\tau) \\
# &= \prod_{j > i} \mathbb P_\theta(\mathbf a_{ij} = a_{ij} | \vec{\pmb \tau} = \vec\tau),\;\;\;\;\textrm{Independence Assumption}
# \end{align*}
#
# Next, for the *a priori* SBM, we know that each edge $\mathbf a_{ij}$ only *actually* depends on the community assignments of nodes $i$ and $j$, so we know that $\mathbb P_{\theta}(\mathbf a_{ij} = a_{ij} | \vec{\pmb \tau} = \vec\tau) = \mathbb P(\mathbf a_{ij} = a_{ij} | \tau_i = k', \tau_j = k)$, where $k$ and $k'$ are any of the $K$ possible communities. This is because the community assignments of nodes that are not nodes $i$ and $j$ do not matter for edge $ij$, due to the independence assumption.
#
# Next, let's think about the probability matrix $P = (p_{ij})$ for the *a priori* SBM. We know that, given that $\tau_i = k'$ and $\tau_j = k$, each adjacency $\mathbf a_{ij}$ is sampled independently and identically from a $Bern(b_{k',k})$ distribution. This means that $p_{ij} = b_{k',k}$. Completing our analysis from above:
# \begin{align*}
# \mathcal L_\theta(A) &\propto \prod_{j > i} b_{k'k}^{a_{ij}}(1 - b_{k'k})^{1 - a_{ij}} \\
# &= \prod_{k,k' \in [K]}b_{k'k}^{m_{k'k}}(1 - b_{k'k})^{n_{k'k} - m_{k'k}}
# \end{align*}
#
# Where $n_{k' k}$ denotes the total number of edges possible between nodes assigned to community $k'$ and nodes assigned to community $k$. That is, $n_{k' k} = \sum_{j > i} \mathbb 1_{\tau_i = k'}\mathbb 1_{\tau_j = k}$. Further, we will use $m_{k' k}$ to denote the total number of edges observed between these two communities. That is, $m_{k' k} = \sum_{j > i}\mathbb 1_{\tau_i = k'}\mathbb 1_{\tau_j = k}a_{ij}$. Note that for a single $(k',k)$ community pair, that the likelihood is analogous to the likelihood of a realization of an ER random variable.
#
# <!--- We can formalize this a bit more explicitly. If we let $A^{\ell k}$ be defined as the subgraph *induced* by the edges incident nodes in community $\ell$ and those in community $k$, then we can say that $A^{\ell k}$ is a directed ER random network, --->
#
# Like the ER model, there are again equivalence classes of the sample space $\mathcal A_n$ in terms of their likelihood. For a two-community setting, with $\vec \tau$ and $B$ given, the equivalence classes are the sets:
# \begin{align*}
# E_{a,b,c}(\vec \tau, B) &= \left\{A \in \mathcal A_n : m_{11} = a, m_{21}=m_{12} = b, m_{22} = c\right\}
# \end{align*}
#
# The number of equivalence classes possible scales with the number of communities, and the manner in which nodes are assigned to communities (particularly, the number of nodes in each community).
#
#
# ### *A Posteriori* Stochastic Block Model
#
# In the *a posteriori* Stochastic Block Model (SBM), we consider that node assignment to one of $K$ communities is a random variable, that we *don't* know already like te *a priori* SBM. We're going to see a funky word come up, that you're probably not familiar with, the **$K$ probability simplex**. What the heck is a probability simplex?
#
# The intuition for a simplex is probably something you're very familiar with, but just haven't seen a word describe. Let's say I have a vector, $\vec\pi = (\pi_k)_{k \in [K]}$, which has a total of $K$ elements. $\vec\pi$ will be a vector, which indicates the *probability* that a given node is assigned to each of our $K$ communities, so we need to impose some additional constraints. Symbolically, we would say that, for all $i$, and for all $k$:
# \begin{align*}
# \pi_k = \mathbb P(\pmb\tau_i = k)
# \end{align*}
# The $\vec \pi$ we're going to use has a very special property: all of its elements are non-negative: for all $\pi_k$, $\pi_k \geq 0$. This makes sense since $\pi_k$ is being used to represent the probability of a node $i$ being in group $k$, so it certainly can't be negative. Further, there's another thing that we want our $\vec\pi$ to have: in order for each element $\pi_k$ to indicate the probability of something to be assigned to $k$, we need all of the $\pi_k$s to sum up to one. This is because of something called the Law of Total Probability. If we have $K$ total values that $\pmb \tau_i$ could take, then it is the case that:
# \begin{align*}
# \sum_{k=1}^K \mathbb P(\pmb \tau_i = k) = \sum_{k = 1}^K \pi_k = 1
# \end{align*}
# So, back to our question: how does a probability simplex fit in? Well, the $K$ probability simplex describes all of the possible values that our vector $\vec\pi$ could possibly take! In symbols, the $K$ probability simplex is:
# \begin{align*}
# \left\{\vec\pi : \text{for all $k$ }\pi_k \geq 0, \sum_{k = 1}^K \pi_k = 1 \right\}
# \end{align*}
# So the $K$ probability simplex is just the space for all possible vectors which could indicate assignment probabilities to one of $K$ communities.
#
# What does the probability simplex look like yy? Below, we take a look at the $2$-probability simplex (2-d $\vec\pi$s) and the $3$-probability simplex (3-dimensional $\vec\pi$s):
# + tags=["hide-input"]
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits.mplot3d.art3d import Poly3DCollection
import matplotlib.pyplot as plt
fig=plt.figure(figsize=plt.figaspect(.5))
fig.suptitle("Probability Simplexes")
ax=fig.add_subplot(1,2,1)
x=[1,0]
y=[0,1]
ax.plot(x,y)
ax.set_xticks([0,.5,1])
ax.set_yticks([0,.5,1])
ax.set_xlabel("$\pi_1$")
ax.set_ylabel("$\pi_2$")
ax.set_title("2-probability simplex")
ax=fig.add_subplot(1,2,2,projection='3d')
x = [1,0,0]
y = [0,1,0]
z = [0,0,1]
verts = [list(zip(x,y,z))]
ax.add_collection3d(Poly3DCollection(verts, alpha=.6))
ax.view_init(elev=20,azim=10)
ax.set_xticks([0,.5,1])
ax.set_yticks([0,.5,1])
ax.set_zticks([0,.5,1])
ax.set_xlabel("$\pi_1$")
ax.set_ylabel("$\pi_2$")
h=ax.set_zlabel("$\pi_3$", rotation=0)
ax.set_title("3-probability simplex")
plt.show()
# -
# The values of $\vec\pi = (\pi)$ that are in the $K$-probability simplex are indicated by the shaded region of each figure. This comprises the $(\pi_1, \pi_2)$ pairs that fall along a diagonal line from $(0,1)$ to $(1,0)$ for the $2$-simplex, and the $(\pi_1, \pi_2, \pi_3)$ tuples that fall on the surface of the triangular shape above with vertices at $(1,0,0)$, $(0,1,0)$, and $(0,0,1)$.
# This model has the following parameters:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $\vec \pi$ | the $K$ probability simplex | The probability of a node being assigned to community $K$ |
# | $B$ | [0,1]$^{K \times K}$ | The block matrix, which assigns edge probabilities for pairs of communities |
#
# The *a posteriori* SBM is a bit more complicated than the *a priori* SBM. We will think about the *a posteriori* SBM as a variation of the *a priori* SBM, where instead of the node-assignment vector being treated as a vector-valued random variable which takes a known fixed value, we will treat it as *unknown*. $\vec{\pmb \tau}$ is still a *latent variable* like it was before. In this case, $\vec{\pmb \tau}$ takes values in the space $\{1,...,K\}^n$. This means that for a given realization of $\vec{\pmb \tau}$, denoted by $\vec \tau$, that for each of the $n$ nodes in the network, we suppose that an integer value between $1$ and $K$ indicates which community a node is from. Statistically, we write that the node assignment for node $i$, denoted by $\pmb \tau_i$, is sampled independently and identically from $Categorical(\vec \pi)$. Stated another way, the vector $\vec\pi$ indicates the probability $\pi_k$ of assignment to each community $k$ in the network.
#
# The matrix $B$ behaves exactly the same as it did with the *a posteriori* SBM. Finally, let's think about how to write down the generative model in the *a posteriori* SBM. The generative model for the *a posteriori* SBM is, in fact, nearly the same as for the *a priori* SBM: we still say that given $\tau_i = k'$ and $\tau_j = k$, that $\mathbf a_{ij}$ are independent $Bern(b_{k'k})$. Here, however, we also describe that $\pmb \tau_i$ are sampled independent and identically from $Categorical(\vec\pi)$, as we learned above. If $\mathbf A$ is the adjacency matrix for an *a posteriori* SBM network with parameters $\vec \pi$ and $B$, we write that $\mathbf A \sim SBM_n(\vec \pi, B)$.
#
# #### Likelihood*
#
# What does the likelihood for the *a posteriori* SBM look like? In this case, $\theta = (\vec \pi, B)$ are the parameters for the model, so the likelihood for a realization $A$ of $\mathbf A$ is:
# \begin{align*}
# \mathcal L_\theta(A) &\propto \mathbb P_\theta(\mathbf A = A)
# \end{align*}
# Next, we use the fact that the probability that $\mathbf A = A$ is, in fact, the *marginalization* (over realizations of $\vec{\pmb \tau}$) of the joint $(\mathbf A, \vec{\pmb \tau})$. In this case, we will let $\mathcal T = \{1,...,K\}^n$ be the space of all possible realizations that $\vec{\pmb \tau}$ could take:
# \begin{align}
# \mathcal L_\theta(A)&\propto \sum_{\vec \tau \in \mathcal T} \mathbb P_\theta(\mathbf A = A, \vec{\pmb \tau} = \vec \tau)
# \end{align}
# Next, remember that by definition of a conditional probability for a random variable $\mathbf x$ taking value $x$ conditioned on random variable $\mathbf y$ taking the value $y$, that $\mathbb P(\mathbf x = x | \mathbf y = y) = \frac{\mathbb P(\mathbf x = x, \mathbf y = y)}{\mathbb P(\mathbf y = y)}$. Note that by multiplying through by $\mathbf P(\mathbf y = y)$, we can see that $\mathbb P(\mathbf x = x, \mathbf y = y) = \mathbb P(\mathbf x = x| \mathbf y = y)\mathbb P(\mathbf y = y)$. Using this logic for $\mathbf A$ and $\vec{\pmb \tau}$:
# \begin{align*}
# \mathcal L_\theta(A) &\propto\sum_{\vec \tau \in \mathcal T} \mathbb P_\theta(\mathbf A = A| \vec{\pmb \tau} = \vec \tau)\mathbb P(\vec{\pmb \tau} = \vec \tau)
# \end{align*}
# Intuitively, for each term in the sum, we are treating $\vec{\pmb \tau}$ as taking a fixed value, $\vec\tau$, to evaluate this probability statement.
#
# We will start by describing $\mathbb P(\vec{\pmb \tau} = \vec\tau)$. Remember that for $\vec{\pmb \tau}$, that each entry $\pmb \tau_i$ is sampled *independently and identically* from $Categorical(\vec \pi)$.The probability mass for a $Categorical(\vec \pi)$-valued random variable is $\mathbb P(\pmb \tau_i = \tau_i; \vec \pi) = \pi_{\tau_i}$. Finally, note that if we are taking the products of $n$ $\pi_{\tau_i}$ terms, that many of these values will end up being the same. Consider, for instance, if the vector $\tau = [1,2,1,2,1]$. We end up with three terms of $\pi_1$, and two terms of $\pi_2$, and it does not matter which order we multiply them in. Rather, all we need to keep track of are the counts of each $\pi$. term. Written another way, we can use the indicator that $\tau_i = k$, given by $\mathbb 1_{\tau_i = k}$, and a running counter over all of the community probability assignments $\pi_k$ to make this expression a little more sensible. We will use the symbol $n_k = \sum_{i = 1}^n \mathbb 1_{\tau_i = k}$ to denote this value, which is the number of nodes in community $k$:
# \begin{align*}
# \mathbb P_\theta(\vec{\pmb \tau} = \vec \tau) &= \prod_{i = 1}^n \mathbb P_\theta(\pmb \tau_i = \tau_i),\;\;\;\;\textrm{Independence Assumption} \\
# &= \prod_{i = 1}^n \pi_{\tau_i} ,\;\;\;\;\textrm{p.m.f. of a Categorical R.V.}\\
# &= \prod_{k = 1}^K \pi_{k}^{n_k},\;\;\;\;\textrm{Reorganizing what we are taking products of}
# \end{align*}
# Next, let's think about the conditional probability term, $\mathbb P_\theta(\mathbf A = A \big | \vec{\pmb \tau} = \vec \tau)$. Remember that the entries are all independent conditional on $\vec{\pmb \tau}$ taking the value $\vec\tau$. It turns out this is exactly the same result that we obtained for the *a priori* SBM:
# \begin{align*}
# \mathbb P_\theta(\mathbf A = A \big | \vec{\pmb \tau} = \vec \tau)
# &= \prod_{k',k} b_{\ell k}^{m_{k' k}}(1 - b_{k' k})^{n_{k' k} - m_{k' k}}
# \end{align*}
#
# Combining these into the integrand gives:
# \begin{align*}
# \mathcal L_\theta(A) &\propto \sum_{\vec \tau \in \mathcal T} \mathbb P_\theta(\mathbf A = A \big | \vec{\pmb \tau} = \vec \tau) \mathbb P_\theta(\vec{\pmb \tau} = \vec \tau) \\
# &= \sum_{\vec \tau \in \mathcal T} \prod_{k = 1}^K \left[\pi_k^{n_k}\cdot \prod_{k'=1}^K b_{k' k}^{m_{k' k}}(1 - b_{k' k})^{n_{k' k} - m_{k' k}}\right]
# \end{align*}
#
# Evaluating this sum explicitly proves to be relatively tedious and is a bit outside of the scope of this book, so we will omit it here.
# <!-- TODO: return to add equivalence classes -->
# ## Random Dot Product Graph (RDPG)
#
# Let's imagine that we have a network which follows the *a priori* Stochastic Block Model. To make this example a little bit more concrete, let's borrow the code example from above. The nodes of our network represent each of the $300$ students in our network. The node assignment vector represents which of the two schools eaach student attends, where the first $150$ students attend school $1$, and the second $150$ students attend school $2$. Remember that $\tau$ and $B$ look like:
# + tags=["hide-input"]
plot_tau(tau, title="Tau, Node Assignment Vector",
xlab="Student");
# + tags=["hide-input"]
plot_block(B, title="Block Matrix");
# -
# Are there any other ways to describe this scenario, other than using both $\tau$ and $B$?
#
# What if we were to look at the probabilities for *every* pair of edges? Remember, for a given $\tau$ and $B$, that a network which is SBM can be generated using the approach that, given that $\tau_i = \ell$ and $\tau_j = k$, that $\mathbf a_{ij} \sim Bern(b_{\ell k})$. That is, every entry is Bernoulli, with the probability indicated by appropriate entry of the block matrix corresponding to the pair of communities each node is in. However, there's another way we could write down this generative model. Suppose we had a $n \times n$ probability matrix, where for every $j > i$:
# \begin{align*}
# p_{ji} = p_{ij}, p_{ij} = \begin{cases}
# b_{11} & \tau_i = 1, \tau_j = 1 \\
# b_{12} & \tau_i = 1, \tau_j = 2 \\
# b_{22} & \tau_i = 2, \tau_j = 1
# \end{cases}
# \end{align*}
# We will call the matrix $P$ the *probability matrix* whose $i^{th}$ row and $j^{th}$ column is the entry $p_{ij}$, as defined above. If you've been following the advanced sections, you will already be familiar with this term. What does $P$ look like?
# + tags=["hide-input"]
def plot_prob(X, title="", nodename="Student", nodetix=None,
nodelabs=None):
fig, ax = plt.subplots(figsize=(8, 6))
with sns.plotting_context("talk", font_scale=1):
ax = sns.heatmap(X, cmap="Purples",
ax=ax, cbar_kws=dict(shrink=1), yticklabels=False,
xticklabels=False, vmin=0, vmax=1)
ax.set_title(title)
cbar = ax.collections[0].colorbar
ax.set(ylabel=nodename, xlabel=nodename)
if (nodetix is not None) and (nodelabs is not None):
ax.set_yticks(nodetix)
ax.set_yticklabels(nodelabs)
ax.set_xticks(nodetix)
ax.set_xticklabels(nodelabs)
cbar.ax.set_frame_on(True)
return
P = np.zeros((n,n))
P[0:150,0:150] = .5
P[150:300, 150:300] = .3
P[0:150,150:300] = .2
P[150:300,0:150] = .2
ax = plot_prob(P, title="Probablity Matrix", nodetix=[0,299],
nodelabs=["1", "300"])
plt.show()
# -
# As we can see, $P$ captures a similar modular structure to the actual adjacency matrix corresponding to the SBM network. Also, $P$ captures the probability of connections between each pair of students. Indeed, it is the case that $P$ contains the information of both $\vec\tau$ and $B$. This means that we can write down a generative model by specifying *only* $P$, and we no longer need to specify $\vec\tau$ and $B$ at all. To write down the generative model in this way, we say that for all $j > i$, that $\mathbf a_{ij} \sim Bern(p_{ij})$ independently, where $\mathbf a_{ji} = \mathbf a_{ij}$, and $\mathbf a_{ii} = 0$.
#
# What is so special about this formulation of the SBM problem? As it turns out, for a *positive semi-definite* probability matrix $P$, $P$ can be decomposed using a matrix $X$, where $P = X X^\top$. We will call a single row of $X$ the vector $\vec x_i$. Remember, using this expression, each entry $p_{ij}$ is the product $\vec x_i^\top \vec x_j$, for all $i, j$. Like $P$, $X$ has $n$ rows, each of which corresponds to a single node in our network. However, the special property of $X$ is that it doesn't *necessarily* have $n$ columns: rather, $X$ often will have many fewer columns than rows. For instance, with $P$ defined as above, there in fact exists an $X$ with just $2$ columns that can be used to describe $P$. This matrix $X$ will be called the *latent position matrix*, and each row $\vec x_i$ will be called the *latent position of a node*. Like previously, there are two types of RDPGs: one in which $X$ is treated as *known*, and another in which $X$ is treated as *unknown*.
#
# Now, your next thought might be that this requires a *lot* more space to represent an SBM network, and you'd be right: $\vec \tau$ has $n$ entries, and $B$ has $K \times K$ entries, where $K$ is typically much smaller than $n$. On the other hand, in this formulation, $P$ has $\binom{n}{2}$ entries, which is much bigger than $n + K \times K$ (since $K$ is usually much smaller than $n$). The advantage is that under this formulation, $P$ doesn't need to have this rigorous modular structure characteristic of SBM networks, and can look a *lot* more interesting. As we will see in later chapters, this network representation will prove extremely flexible for allowing us to capture networks that are fairly complex. Further, we can also perform analysis on the matrix $X$ itself, which will prove very useful for estimation of SBMs.
# ### *A Priori* RDPG
#
# The *a priori* Random Dot Product Graph is an RDPG in which we know *a priori* the latent position matrix $X$. The *a priori* RDPG has the following parameter:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $X$ | $ \mathbb R^{n \times d}$ | The matrix of latent positions for each node $n$. |
#
# $X$ is called the **latent position matrix** of the RDPG. We write that $X \in \mathbb R^{n \times d}$, which means that it is a matrix with real values, $n$ rows, and $d$ columns. We will use the notation $\vec x_i$ to refer to the $i^{th}$ row of $X$. $\vec x_i$ is referred to as the **latent position** of a node $i$. Visually, this looks something like this:
# \begin{align*}
# X = \begin{bmatrix}
# \vec x_{1}^\top \\
# \vdots \\
# \vec x_n^\top
# \end{bmatrix}
# \end{align*}
# Noting that $X$ has $d$ columns, this implies that $\vec x_i \in \mathbb R^d$, or that each node's latent position is a real-valued $d$-dimensional vector.
#
# What is the generative model for the *a priori* RDPG? As we discussed above, given $X$, for all $j > i$, $\mathbf a_{ij} \sim Bern(\vec x_i^\top \vec x_j)$ independently. If $i < j$, $\mathbf a_{ji} = \mathbf a_{ij}$ (the network is *undirected*), and $\mathbf a_{ii} = 0$ (the network is *loopless*). If $\mathbf A$ is an *a priori* RDPG with parameter $X$, we write that $\mathbf A \sim RDPG_n(X)$.
#
# #### Code Examples
#
# We will let $X$ be a little more complex than in our preceding example. Our $X$ will produce a $P$ that still *somewhat* has a modular structure, but not quite as much as before. Let's assume that we have $300$ people who live along a very long road that is $100$ miles long, and each person is $\frac{1}{3}$ of a mile apart. The nodes of our network represent the people who live along our assumed street. If two people are closer to one another, it might make sense to think that they have a higher probability of being friends. If two people are neighbors, we think that they will have a very high probability of being friends (almost $1$) and when people are very far apart, we think that they will have a very low probability of being friends (almost $0$). What could we use for $X$?
#
# One possible approach would be to let each $\vec x_i$ be defined as follows:
# \begin{align*}
# \vec x_i = \begin{bmatrix}
# \frac{300 - i}{300} \\
# \frac{i}{300}
# \end{bmatrix}
# \end{align*}
# For instance, $\vec x_1 = \begin{bmatrix}1 \\ 0\end{bmatrix}$, and $\vec x_{300} = \begin{bmatrix} 0 \\ 1\end{bmatrix}$. Note that:
# \begin{align*}
# p_{1,300} = \vec x_1^\top \vec x_j = 1 \cdot 0 + 0 \cdot 1 = 0
# \end{align*}
# What happens in between?
#
# Let's consider another person, person $100$. Note that person $100$ lives closer to person $1$ than to person $300$. Here, $\vec x_{100} = \begin{bmatrix} \frac{2}{3}\\ \frac{1}{3}\end{bmatrix}$. This gives us that:
# \begin{align*}
# p_{1,100} &= \vec x_1^\top \vec x_{100} = \frac{2}{3}\cdot 1 + 0 \cdot \frac{1}{3} = \frac{2}{3} \\
# p_{100, 300} &= \vec x_{100}^\top x_{300} = \frac{2}{3} \cdot 0 + \frac 1 3 \cdot 1 = \frac 1 3
# \end{align*}
# So this means that person $1$ and person $100$ have about a $67\%$ probability of being friends, but person $100$ and $300$ have about a $33\%$ probability of being friends.
#
# Let's consider another person, person $200$. Person $200$ lives closer to person $300$ than person $100$. With $\vec x_{200} = \begin{bmatrix}\frac{1}{3} \\ \frac{2}{3} \end{bmatrix}$, we obtain that:
# \begin{align*}
# p_{1,200} &= \vec x_1^\top \vec x_{200} = \frac{1}{3}\cdot 1 + 0 \cdot \frac{2}{3} = \frac{1}{3} \\
# p_{200, 300} &= \vec x_{100}^\top x_{300} = \frac{1}{3} \cdot 0 + \frac 2 3 \cdot 1 = \frac 2 3 \\
# p_{100,200} &= \vec x_{100}^\top x_{200} = \frac{2}{3} \cdot \frac 1 3 + \frac 1 3 \cdot \frac 2 3 = \frac 4 9
# \end{align*}
# Again, remember that these fractions capture the probability that two people will be friends. So, intuitively, it seems like our probability matrix $P$ will capture the intuitive idea we described above. First, we'll take a look at $X$, and then we'll look at $P$:
# +
n = 300 # the number of nodes in our network
# design the latent position matrix X according to
# the rules we laid out previously
X = np.zeros((n,2))
for i in range(0, n):
X[i,:] = [(n - i)/n, i/n]
# + tags=["hide-input"]
def plot_lp(X, title="", ylab="Student"):
fig, ax = plt.subplots(figsize=(4, 10))
with sns.plotting_context("talk", font_scale=1):
ax = sns.heatmap(X, cmap="Purples",
ax=ax, cbar_kws=dict(shrink=1), yticklabels=False,
xticklabels=False)
ax.set_title(title)
cbar = ax.collections[0].colorbar
ax.set(ylabel=ylab)
ax.set_yticks([0, 99, 199, 299])
ax.set_yticklabels(["1", "100", "200", "300"])
ax.set_xticks([.5, 1.5])
ax.set_xticklabels(["Dimension 1", "Dimension 2"])
cbar.ax.set_frame_on(True)
return
plot_lp(X, title="Latent Position Matrix, X")
# + [markdown] tags=["hide-input"]
# The latent position matrix $X$ that we plotted above is $n \times d$ dimensions. There are a number of approaches, other than looking at a heatmap of $X$, with which we can visualize $X$ to derive insights as to its structure. When $d=2$, another popular visualization is to look at the latent positions, $\vec x_i$, as individual points in $2$-dimensional space. This will give us a scatter plot of $n$ points, each of which has two coordinates. Each point is the latent position for a single node:
# + tags=["hide-input"]
def plot_latents(latent_positions, title=None, labels=None, **kwargs):
fig, ax = plt.subplots(figsize=(6, 6))
if ax is None:
ax = plt.gca()
ss = 6*np.arange(0, 50)
plot = sns.scatterplot(x=latent_positions[ss, 0], y=latent_positions[ss, 1], hue=labels,
s=10, ax=ax, palette="Set1", color='k', **kwargs)
ax.set_title(title)
ax.set(ylabel="Dimension 1", xlabel="Dimension 2")
ax.set_title(title)
return plot
# plot
plot_latents(X, title="Latent Position Matrix, X");
# + [markdown] tags=["hide-input"]
# The above scatter plot has been subsampled to show only every $6^{th}$ latent position, so that the individual $2$-dimensional latent positions are discernable. Due to the way we constructed $X$, the scatter plot would otherwise appear to be a line (due to points overlapping one another). The reason that the points fall along a vertical line when plotted as a vector is due to the method we used to construct entries of $X$, described above. Next, we will look at the probability matrix:
# + tags=["hide-input"]
plot_prob(X.dot(X.transpose()), title="Probability Matrix, P=$XX^T$",
nodelabs=["1", "100", "200", "300"], nodetix=[0,99,199,299])
# -
# Finally, we will sample an RDPG:
# +
from graspologic.simulations import rdpg
# sample an RDPG with the latent position matrix
# created above
A = rdpg(X, loops=False, directed=False)
# and plot it
ax = binary_heatmap(A, title="$RDPG_{300}(X)$ Simulation")
# -
# ### Likelihood*
#
# Given $X$, the likelihood for an RDPG is relatively straightforward, as an RDPG is another Independent-Edge Random Graph. The independence assumption vastly simplifies our resulting expression. We will also use many of the results we've identified above, such as the p.m.f. of a Bernoulli random variable. Finally, we'll note that the probability matrix $P = (\vec x_i^\top \vec x_j)$, so $p_{ij} = \vec x_i^\top \vec x_j$:
#
# \begin{align*}
# \mathcal L_\theta(A) &\propto \mathbb P_\theta(A) \\
# &= \prod_{j > i}\mathbb P(\mathbf a_{ij} = a_{ij}),\;\;\;\; \textrm{Independence Assumption} \\
# &= \prod_{j > i}(\vec x_i^\top \vec x_j)^{a_{ij}}(1 - \vec x_i^\top \vec x_j)^{1 - a_{ij}},\;\;\;\; a_{ij} \sim Bern(\vec x_i^\top \vec x_j)
# \end{align*}
#
# Unfortunately, the likelihood equivalence classes are a bit harder to understand intuitionally here compared to the ER and SBM examples so we won't write them down here, but they still exist!
# ### *A Posteriori* RDPG
#
# Like for the *a posteriori* SBM, the *a posteriori* RDPG introduces another strange set: the **intersection of the unit ball and the non-negative orthant**. Huh? This sounds like a real mouthful, but it turns out to be rather straightforward. You are probably already very familiar with a particular orthant: in two-dimensions, an orthant is called a quadrant. Basically, an orthant just extends the concept of a quadrant to spaces which might have more than $2$ dimensions. The non-negative orthant happens to be the orthant where all of the entries are non-negative. We call the **$K$-dimensional non-negative orthant** the set of points in $K$-dimensional real space, where:
# \begin{align*}
# \left\{\vec x \in \mathbb R^K : x_k \geq 0\text{ for all $k$}\right\}
# \end{align*}
# In two dimensions, this is the traditional upper-right portion of the standard coordinate axis. To give you a picture, the $2$-dimensional non-negative orthant is the blue region of the following figure:
# + tags=["hide-input"]
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.axisartist import SubplotZero
import matplotlib.patches as patch
class Axes():
def __init__(self, xlim=(-5,5), ylim=(-5,5), figsize=(6,6)):
self.xlim = xlim
self.ylim = ylim
self.figsize = figsize
self.__scale_arrows__()
def __drawArrow__(self, x, y, dx, dy, width, length):
plt.arrow(
x, y, dx, dy,
color = 'k',
clip_on = False,
head_width = self.head_width,
head_length = self.head_length
)
def __scale_arrows__(self):
""" Make the arrows look good regardless of the axis limits """
xrange = self.xlim[1] - self.xlim[0]
yrange = self.ylim[1] - self.ylim[0]
self.head_width = min(xrange/30, 0.25)
self.head_length = min(yrange/30, 0.3)
def __drawAxis__(self):
"""
Draws the 2D cartesian axis
"""
# A subplot with two additional axis, "xzero" and "yzero"
# corresponding to the cartesian axis
ax = SubplotZero(self.fig, 1, 1, 1)
self.fig.add_subplot(ax)
# make xzero axis (horizontal axis line through y=0) visible.
for axis in ["xzero","yzero"]:
ax.axis[axis].set_visible(True)
# make the other axis (left, bottom, top, right) invisible
for n in ["left", "right", "bottom", "top"]:
ax.axis[n].set_visible(False)
# Plot limits
plt.xlim(self.xlim)
plt.ylim(self.ylim)
ax.set_yticks([-1, 1, ])
ax.set_xticks([-2, -1, 0, 1, 2])
# Draw the arrows
self.__drawArrow__(self.xlim[1], 0, 0.01, 0, 0.3, 0.2) # x-axis arrow
self.__drawArrow__(0, self.ylim[1], 0, 0.01, 0.2, 0.3) # y-axis arrow
self.ax=ax
def draw(self):
# First draw the axis
self.fig = plt.figure(figsize=self.figsize)
self.__drawAxis__()
axes = Axes(xlim=(-2.5,2.5), ylim=(-2,2), figsize=(9,7))
axes.draw()
rectangle =patch.Rectangle((0,0), 3, 3, fc='blue',ec="blue", alpha=.2)
axes.ax.add_patch(rectangle)
plt.show()
# -
# Now, what is the unit ball? You are probably familiar with the idea of the unit ball, even if you haven't heard it called that specifically. Remember that the Euclidean norm for a point $\vec x$ which has coordinates $x_i$ for $i=1,...,K$ is given by the expression:
# \begin{align*}
# \left|\left|\vec x\right|\right|_2 \triangleq \sqrt{\sum_{i = 1}^K x_i^2}
# \end{align*}
# The Euclidean unit ball is just the set of points whose Euclidean norm is at most $1$. To be more specific, the **closed unit ball** with the Euclidean norm is the set of points:
# \begin{align*}
# \left\{\vec x \in \mathbb R^K :\left|\left|\vec x\right|\right|_2 \leq 1\right\}
# \end{align*}
#
# We draw the $2$-dimensional unit ball with the Euclidean norm below, where the points that make up the unit ball are shown in red:
# + tags=["hide-input"]
axes = Axes(xlim=(-2.5,2.5), ylim=(-2,2), figsize=(9,7))
axes.draw()
circle =patch.Circle((0,0), 1, fc='red',ec="red", alpha=.3)
axes.ax.add_patch(circle)
plt.show()
# -
# Now what is their intersection? Remember that the intersection of two sets $A$ and $B$ is the set:
# \begin{align*}
# A \cap B &= \{x : x \in A, x \in B\}
# \end{align*}
# That is, each element must be in *both* sets to be in the intersection. Formally, the interesction of the unit ball and the non-negative orthant will be the set:
#
# \begin{align*}
# \mathcal X_K \triangleq \left\{\vec x \in \mathbb R^K :\left|\left|\vec x\right|\right|_2 \leq 1, x_k \geq 0 \textrm{ for all $k$}\right\}
# \end{align*}
#
# visually, this will be the set of points in the *overlap* of the unit ball and the non-negative orthant, which we show below in purple:
# + tags=["hide-input"]
axes = Axes(xlim=(-2.5,2.5), ylim=(-2,2), figsize=(9,7))
axes.draw()
circle =patch.Circle((0,0), 1, fc='red',ec="red", alpha=.3)
axes.ax.add_patch(circle)
rectangle =patch.Rectangle((0,0), 3, 3, fc='blue',ec="blue", alpha=.2)
axes.ax.add_patch(rectangle)
plt.show()
# + [markdown] tags=["hide-input"]
# This space has an *incredibly* important corollary. It turns out that if $\vec x$ and $\vec y$ are both elements of $\mathcal X_K$, that $\left\langle \vec x, \vec y \right \rangle = \vec x^\top \vec y$, the **inner product**, is at most $1$, and at least $0$. Without getting too technical, this is because of something called the Cauchy-Schwartz inequality and the properties of $\mathcal X_K$. If you remember from linear algebra, the Cauchy-Schwartz inequality states that $\left\langle \vec x, \vec y \right \rangle$ can be at most the product of $\left|\left|\vec x\right|\right|_2$ and $\left|\left|\vec y\right|\right|_2$. Since $\vec x$ and $\vec y$ have norms both less than or equal to $1$ (since they are on the *unit ball*), their inner-product is at most $1$. Further, since $\vec x$ and $\vec y$ are in the non-negative orthant, their inner product can never be negative.
#
# -
# The *a posteriori* RDPG is to the *a priori* RDPG what the *a posteriori* SBM was to the *a priori* SBM. We instead suppose that we do *not* know the latent position matrix $X$, but instead know how we can characterize the individual latent positions. We have the following parameter:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | F | inner-product distributions | A distribution which governs each latent position. |
#
# The parameter $F$ is what is known as an **inner-product distribution**. In the simplest case, we will assume that $F$ is a distribution on a subset of the possible real vectors that have $d$-dimensions with an important caveat: for any two vectors within this subset, their inner product *must* be a probability. We will refer to the subset of the possible real vectors as $\mathcal X_K$, which we learned about above. This means that for any $\vec x_i, \vec x_j$ that are in $\mathcal X_K$, it is always the case that $\vec x_i^\top \vec x_j$ is between $0$ and $1$. This is essential because like previously, we will describe the distribution of each edge in the adjacency matrix using $\vec x_i^\top \vec x_j$ to represent a probability. Next, we will treat the latent position matrix as a matrix-valued random variable which is *latent* (remember, *latent* means that we don't get to see it in our real data). Like before, we will call $\vec{\mathbf x}_i$ the random latent positions for the nodes of our network. In this case, each $\vec {\mathbf x}_i$ is sampled independently and identically from the inner-product distribution $F$ described above. The latent-position matrix is the matrix-valued random variable $\mathbf X$ whose entries are the latent vectors $\vec {\mathbf x}_i$, for each of the $n$ nodes.
#
# The model for edges of the *a posteriori* RDPG can be described by conditioning on this unobserved latent-position matrix. We write down that, conditioned on $\vec {\mathbf x}_i = \vec x$ and $\vec {\mathbf x}_j = \vec y$, that if $j > i$, then $\mathbf a_{ij}$ is sampled independently from a $Bern(\vec x^\top \vec y)$ distribution. As before, if $i < j$, $\mathbf a_{ji} = \mathbf a_{ij}$ (the network is *undirected*), and $\mathbf a_{ii} = 0$ (the network is *loopless*). If $\mathbf A$ is the adjacency matrix for an *a posteriori* RDPG with parameter $F$, we write that $\mathbf A \sim RDPG_n(F)$.
#
# #### Likelihood*
#
# The likelihood for the *a posteriori* RDPG is fairly complicated. This is because, like the *a posteriori* SBM, we do not actually get to see the latent position matrix $\mathbf X$, so we need to use *marginalization* to obtain an expression for the likelihood. Here, we are concerned with realizations of $\mathbf X$. Remember that $\mathbf X$ is just a matrix whose rows are $\vec {\mathbf x}_i$, each of which individually have have the distribution $F$; e.g., $\vec{\mathbf x}_i \sim F$ independently. For simplicity, we will assume that $F$ is a disrete distribution on $\mathcal X_K$. This makes the logic of what is going on below much simpler since the notation gets less complicated, but does not detract from the generalizability of the result (the only difference is that sums would be replaced by multivariate integrals, and probability mass functions replaced by probability density functions).
#
# We will let $p$ denote the probability mass function (p.m.f.) of this discrete distribution function $F$. The strategy will be to use the independence assumption, followed by marginalization over the relevant rows of $\mathbf X$:
#
# \begin{align*}
# \mathcal L_\theta(A) &\propto \mathbb P_\theta(\mathbf A = A) \\
# &= \prod_{j > i} \mathbb P(\mathbf a_{ij} = a_{ij}), \;\;\;\;\textrm{Independence Assumption} \\
# \mathbb P(\mathbf a_{ij} = a_{ij})&= \sum_{\vec x \in \mathcal X_K}\sum_{\vec y \in \mathcal X_K}\mathbb P(\mathbf a_{ij} = a_{ij}, \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y),\;\;\;\;\textrm{Marginalization over }\vec {\mathbf x}_i \textrm{ and }\vec {\mathbf x}_j
# \end{align*}
# Next, we will simplify this expression a little bit more, using the definition of a conditional probability like we did before for the SBM:
#
# \begin{align*}
# \\
# \mathbb P(\mathbf a_{ij} = a_{ij}, \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y) &= \mathbb P(\mathbf a_{ij} = a_{ij}| \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y) \mathbb P(\vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y)
# \end{align*}
#
# Further, remember that if $\mathbf a$ and $\mathbf b$ are independent, then $\mathbb P(\mathbf a = a, \mathbf b = b) = \mathbb P(\mathbf a = a)\mathbb P(\mathbf b = b)$. Using that $\vec x_i$ and $\vec x_j$ are independent, by definition:
#
# \begin{align*}
# \mathbb P(\vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y) &= \mathbb P(\vec{\mathbf x}_i = \vec x) \mathbb P(\vec{\mathbf x}_j = \vec y)
# \end{align*}
#
# Which means that:
#
# \begin{align*}
# \mathbb P(\mathbf a_{ij} = a_{ij}, \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y) &= \mathbb P(\mathbf a_{ij} = a_{ij} | \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y)\mathbb P(\vec{\mathbf x}_i = \vec x) \mathbb P(\vec{\mathbf x}_j = \vec y)
# \end{align*}
# Finally, we that conditional on $\vec{\mathbf x}_i = \vec x_i$ and $\vec{\mathbf x}_j = \vec x_j$, $\mathbf a_{ij}$ is $Bern(\vec x_i^\top \vec x_j)$. This means that in terms of our probability matrix, each entry $p_{ij} = \vec x_i^\top \vec x_j$. Therefore:
#
# \begin{align*}
# \mathbb P(\mathbf a_{ij} = a_{ij}| \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y) &= (\vec x^\top \vec y)^{a_{ij}}(1 - \vec x^\top\vec y)^{1 - a_{ij}}
# \end{align*}
# This implies that:
# \begin{align*}
# \mathbb P(\mathbf a_{ij} = a_{ij}, \vec{\mathbf x}_i = \vec x, \vec{\mathbf x}_j = \vec y) &= (\vec x^\top \vec y)^{a_{ij}}(1 - \vec x^\top\vec y)^{1 - a_{ij}}\mathbb P(\vec{\mathbf x}_i = \vec x) \mathbb P(\vec{\mathbf x}_j = \vec y)
# \end{align*}
# Where $p(\vec x)$ is the p.m.f. $\mathbb
#
# So our complete expression for the likelihood is:
#
# \begin{align*}
# \mathcal L_\theta(A) &\propto \prod_{j > i}\sum_{\vec x \in \mathcal X_K}\sum_{\vec y \in \mathcal X_K} (\vec x^\top \vec y)^{a_{ij}}(1 - \vec x^\top\vec y)^{1 - a_{ij}}\mathbb P(\vec{\mathbf x}_i = \vec x) \mathbb P(\vec{\mathbf x}_j = \vec y)
# \end{align*}
# ## Inhomogeneous Erdös-Rényi (IER)
#
# In the preceding models, we typically made assumptions about how we could characterize the edge-existence probabilities using fewer than $\binom n 2$ unique probabilities (one for each edge). The reason for this is that in general, $n$ is usually relatively large, so attempting to actually learn $\binom n 2$ unique probabilities is not, in general, going to be very feasible (it is *never* feasible when we have a single network, since a single network only one observation for each independent edge). Further, it is relatively difficult to ask questions for which assuming edges share *nothing* in common (even if they don't share the same probabilities, there may be properties underlying the probabilities, such as the *latent positions* that we saw above with the RDPG, that we might still want to characterize) is actually favorable.
#
# Nonetheless, the most general model for an independent-edge random network is known as the Inhomogeneous Erdös-Rényi (IER) Random Network. An IER Random Network is characterized by the following parameters:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $P$ | [0,1]$^{n \times n}$ | The edge probability matrix. |
#
# The probability matrix $P$ is an $n \times n$ matrix, where each entry $p_{ij}$ is a probability (a value between $0$ and $1$). Further, if we restrict ourselves to the case of simple networks like we have done so far, $P$ will also be symmetric ($p_{ij} = p_{ji}$ for all $i$ and $j$). The generative model is similar to the preceding models we have seen: given the $(i, j)$ entry of $P$, denoted $p_{ij}$, the edges $\mathbf a_{ij}$ are independent $Bern(p_{ij})$, for any $j > i$. Further, $\mathbf a_{ii} = 0$ for all $i$ (the network is *loopless*), and $\mathbf a_{ji} = \mathbf a_{ij}$ (the network is *undirected*). If $\mathbf A$ is the adjacency maatrix for an IER network with probability matarix $P$, we write that $\mathbf A \sim IER_n(P)$.
#
# It is worth noting that *all* of the preceding models we have discussed so far are special cases of the IER model. This means that, for instance, if we were to consider only the probability matrices where all of the entries are the same, we could represent the ER models. Similarly, if we were to only to consider the probability matrices $P$ where $P = XX^\top$, we could represent any RDPG.
#
# ### Likelihood*
#
# The likelihood for a network which is IER is very straightforward. We use the independence assumption, and the p.m.f. of a Bernoulli-distributed random-variable $\mathbf a_{ij}$:
#
# \begin{align*}
# \mathcal L_\theta(A) &\propto \mathbb P(\mathbf A = A) \\
# &= \prod_{j > i}p_{ij}^{a_{ij}}(1 - p_{ij})^{1 - a_{ij}}
# \end{align*}
# ## Degree-Corrected Stochastic Block Model (DCSBM)
# Let's think back to our school example for the Stochastic Block Model. Remember, we had 100 students, each of whom could go to one of two possible schools: school one or school two. Our network had 100 nodes, representing each of the students. We said that the school for which each student attended was represented by their node assignment $\tau_i$ to one of two possible communities. The matrix $B$ was the block probaability matrix, where $b_{11}$ was the probability that students in school one were friends, $b_{22}$ was the probability that students in school two were friends, and $b_{12} = b_{21}$ was the probability that students were friends if they did not go to the same school. In this case, we said that $\mathbf A \sim SBM_n(\tau, B)$.
#
# When would this setup not make sense? Let's say that Alice and Bob both go to the same school, but Alice is more popular than Bob. If we were to look at a schoolmate Chadwick, it might not make sense to say that both Alice and Bob have the *same* probability of being friends with Chadwick. Rather, we might want to reflect that Alice has a higher probability of being friends with an arbitrary schoolmate than Bob. The problem here is that within a single community of an SBM, the SBM assumes that the **node degree** (the number of nodes each nodes is connected to) is the *same* for all nodes within a single community.
#
#
# ```{admonition} Degree Homogeneity in a Stochastic Block Model Network
# Suppose that $\mathbf A \sim SBM_{n, \vec\tau}(B)$, where $\mathbf A$ has $K=2$ communities. What is the node degree of each node in $\mathbf A$?
#
# For an arbitrary node $v_i$ which is in community $k$ (either $1$ or $2$), we will compute the expectated value of the degree $deg(v_i)$, written $\mathbb E\left[deg(v_i); \tau_i = k\right]$. We will let $n_k$ represent the number of nodes whose node assignments $\tau_i$ are to community $k$. Let's see what happens:
# \begin{align*}
# \mathbb E\left[deg(v_i); \tau_i = k\right] &= \mathbb E\left[\sum_{j = 1}^n \mathbf a_{ij}\right] \\
# &= \sum_{j = 1}^n \mathbb E[\mathbf a_{ij}]
# \end{align*}
# We use the *linearity of expectation* again to get from the top line to the second line. Next, instead of summing over all the nodes, we'll break the sum up into the nodes which are in the same community as node $i$, and the ones in the *other* community $k'$. We use the notation $k'$ to emphasize that $k$ and $k'$ are different values:
#
# \begin{align*}
# \mathbb E\left[deg(v_i); \tau_i = k\right] &= \sum_{j : i \neq j, \tau_j = k} \mathbb E\left[\mathbf a_{ij}\right] + \sum_{j : \tau_j =k'} \mathbb E[\mathbf a_{ij}]
# \end{align*}
# In the first sum, we have $n_k-1$ total edges (the number of nodes that aren't node $i$, but are in the same community), and in the second sum, we have $n_{k'}$ total edges (the number of nodes that are in the other community). Finally, we will use that the probability of an edge in the same community is $b_{kk}$, but the probability of an edge between the communities is $b_{k' k}$. Finally, we will use that the expected value of an adjacency $\mathbf a_{ij}$ which is Bernoulli distributed is its probability:
# \begin{align*}
# \mathbb E\left[deg(v_i); \tau_i = k\right] &= \sum_{j : i \neq j, \tau_j = k} b_{kk} + \sum_{j : \tau_j = \ell} b_{kk'},\;\;\;\;\mathbf a_{ij}\textrm{ are Bernoulli distributed} \\
# &= (n_k - 1)b_{kk} + n_{k'} b_{kk'}
# \end{align*}
# This holds for any node $i$ which is in community $k$. Therefore, the expected node degree is the same, or **homogeneous**, within a community of an SBM.
# ```
#
# To address this limitation, we turn to the Degree-Corrected Stochastic Block Model, or DCSBM. As with the Stochastic Block Model, there is both a *a priori* and *a posteriori* DCSBM.
# ### *A Priori* DCSBM
#
# Like the *a priori* SBM, the *a priori* DCSBM is where we know which nodes are in which node communities ahead of time. Here, we will use the variable $K$ to denote the maximum number of communities that nodes could be assigned to. The *a priori* DCSBM has the following two parameters:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $B$ | [0,1]$^{K \times K}$ | The block matrix, which assigns edge probabilities for pairs of communities |
# | $\vec\theta$ | $\mathbb R^n_+$ | The degree correction vector, which adjusts the degree for pairs of nodes |
#
# The latent community assignment vector $\vec{\pmb \tau}$ with a known *a priori* realization $\vec{\tau}$ and the block matrix $B$ are exactly the same for the *a priori* DCSBM as they were for the *a priori* SBM.
#
# The vector $\vec\theta$ is the degree correction vector. Each entry $\theta_i$ is a positive scalar. For every adjacency for a given node $i$, the degree correction $\theta_i$ will indicate the factor by which the probability for an adjacency which represents an edge incident node $i$ is adjusted.
#
# Finally, let's think about how to write down the generative model for the *a priori* DCSBM. We say that $\tau_i = k'$ and $\tau_j = k$, $\mathbf a_{ij}$ is sampled independently from a $Bern(\theta_i \theta_j b_{k'k})$ distribution for all $j > i$. As we can see, $\theta_i$ in a sense is "correcting" the probabilities of each adjacency to node $i$ to be higher, or lower, depending on the value of $\theta_i$ that that which is given by the block probabilities $b_{\ell k}$. If $\mathbf A$ is an *a priori* DCSBM network with parameters and $B$, we write that $\mathbf A \sim DCSBM_{n,\vec\tau}(\vec \theta, B)$.
# #### Likelihood*
#
# The derivation for the likelihood is the same as for the *a priori* SBM, with the change that $p_{ij} = \theta_i \theta_j b_{k'k}$ instead of just $b_{k'k}$. This gives that the likelihood turns out to be:
#
# \begin{align*}
# \mathcal L_\theta(A) &\propto \prod_{j > i} \left(\theta_i \theta_j b_{k'k}\right)^{a_{ij}}\left(1 - \theta_i \theta_j b_{k'k}\right)^{1 - a_{ij}}
# \end{align*}
# The expression doesn't simplify much more due to the fact that the probabilities are dependent on the particular $i$ and $j$, so we can't just reduce the statement in terms of $n_{k'k}$ and $m_{k'k}$ like for the SBM.
# ### *A Posteriori* DCSBM
# The *a posteriori* DCSBM is to the *a posteriori* SBM what the *a priori* DCSBM was to the *a priori* SBM. The changes are very minimal, so we will omit explicitly writing it all down here so we can get this section wrapped up, with the idea that the preceding section on the *a priori* DCSBM should tell you what needs to change.
#
# ## Network models for networks which aren't simple
# To make the discussions a little more easy to handle, in the above descriptions, we described network models for simple networks, which to recap, are binary networks which are both loopless and undirected. Stated another way, simple networks are networks whose adjacency matrices are only $0$s and $1$s, they are hollow, and symmetric. What happens our networks don't quite look this way?
#
# For now, we'll keep the assumption that the networks are binary, but we will discuss non-binary network models in a later chapter. We have three possibilities we can consider, and we will show how the "relaxations" of the assumptions change a description of a network model. We split these out so we can be as clear as possible about how the generative model changes.
#
# We will compare each relaxation to the statement about the generative model for the ER generative model. To recap, for a simple network, we wrote:
#
# Statistically, we say that for each edge $\mathbf{a}_{ij}$, that $\mathbf{a}_{ij}$ is sampled independently and identically from a $Bern(p)$ distribution, whenever $j > i$. When $i > j$, we allow $\mathbf a_{ij} = \mathbf a_{ji}$. Also, we let $\mathbf a_{ii} = 0$, which means that all self-loops are always unconnected.
#
# ### Binary network model which has loops, but is undirected
#
# Here, all we want to do is relax the assumption that the network is loopless. We simply ignore the statement that $\mathbf a_{ii} = 0$, and allow that the $\mathbf a_{ij}$ which follow a Bernoulli distribution (with some probability which depends on the network model choice) *now* applies to $j \geq i$, and not just $j > i$. We keep that $\mathbf a_{ji} = \mathbf a_{ij}$, which maintains the symmetry of $\mathbf A$ (and consequently, the undirectedness of the network).
#
# Our description of the ER network changes to:
#
# Statistically, we say that for each edge $\mathbf{a}_{ij}$, that $\mathbf{a}_{ij}$ is sampled independently and identically from a $Bern(p)$ distribution, whenever $j \geq i$. When $i > j$, we allow $\mathbf a_{ij} = \mathbf a_{ji}$.
#
# ### Binary network model which is loopless, but directed
#
# Like above, we simply ignore the statement that $\mathbf a_{ji} = \mathbf a_{ij}$, which removes the symmetry of $\mathbf A$ (and consequently, removes the undirectedness of the network). We allow that the $\mathbf a_{ij}$ which follows a Bernoulli distribution now apply to $j \neq i$, and not just $j > i$. We keep that $\mathbf a_{ii} = 0$, which maintains the hollowness of $\mathbf A$ (and consequently, the undirectedness of the network).
#
# Our description of the ER network changes to:
#
# Statistically, we say that for each edge $\mathbf{a}_{ij}$, that $\mathbf{a}_{ij}$ is sampled independently and identically from a $Bern(p)$ distribution, whenever $j \neq i$. Also, we let $\mathbf a_{ii} = 0$, which means that all self-loops are always unconnected.
#
#
# ### Binary network model which is has loops and is directed
#
# Finally, for a network which has loops and is directed, we combine the above two approaches. We ignore the statements that $\mathbf a_{ji} = \mathbf a_{ij}$, and the statement thhat $\mathbf a_{ii} = 0$.
#
# Our descriptiomn of the ER network changes to:
#
#
# Statistically, we say that for each edge $\mathbf{a}_{ij}$, that $\mathbf{a}_{ij}$ is sampled independently and identically from a $Bern(p)$ distribution, for all possible combinations of nodes $j$ and $i$.
# ## Generalized Random Dot Product Graph (GRDPG)
#
# The Generalized Random Dot Product Graph, or GRDPG, is the most general random network model we will consider in this book. Note that for the RDPG, the probability matrix $P$ had entries $p_{ij} = \vec x_i^\top \vec x_j$. What about $p_{ji}$? Well, $p_{ji} = \vec x_j^\top \vec x_i$, which is exactly the same as $p_{ij}$! This means that even if we were to consider a directed RDPG, the probabilities that can be captured are *always* going to be symmetric. The generalized random dot product graph, or GRDPG, relaxes this assumption. This is achieved by using *two* latent positin matrices, $X$ and $Y$, and letting $P = X Y^\top$. Now, the entries $p_{ij} = \vec x_i^\top \vec y_j$, but $p_{ji} = \vec x_j^\top \vec y_i$, which might be different.
#
# ### *A Priori* GRDPG
#
# The *a priori* GRDPG is a GRDPG in which we know *a priori* the latent position matrices $X$ and $Y$. The *a priori* GRDPG has the following parameters:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | $X$ | $ \mathbb R^{n \times d}$ | The matrix of left latent positions for each node $n$. |
# | $Y$ | $ \mathbb R^{n \times d}$ | The matrix of right latent positions for each node $n$. |
#
# $X$ and $Y$ behave nearly the same as the latent position matrix $X$ for the *a priori* RDPG, with the exception that they will be called the **left latent position matrix** and the **right latent position matrix** respectively. Further, the vectors $\vec x_i$ will be the left latent positions, and $\vec y_i$ will be the right latent positions, for a given node $i$, for each node $i=1,...,n$.
#
# What is the generative model for the *a priori* GRDPG? As we discussed above, given $X$ and $Y$, for all $j \neq i$, $\mathbf a_{ij} \sim Bern(\vec x_i^\top \vec y_j)$ independently. If we consider only loopless networks, $\mathbf a_{ij} = 0$. If $\mathbf A$ is an *a priori* GRDPG with left and right latent position matrices $X$ and $Y$, we write that $\mathbf A \sim GRDPG_n(X, Y)$.
#
# ### *A Posteriori* GRDPG
#
# The *A Posteriori* GRDPG is very similar to the *a posteriori* RDPG. We have two parameters:
#
# | Parameter | Space | Description |
# | --- | --- | --- |
# | F | inner-product distributions | A distribution for the left latent positions. |
# | G | inner-product distributions | A distribution for the right latent positions. |
#
# Here, we treat the left and right latent position matrices as latent variable matrices, like we did for *a posteriori* RDPG. That is, the left latent positions are sampled independently and identically from $F$, and the right latent positions $\vec y_i$ are sampled independently and identically from $G$.
#
# The model for edges of the *a posteriori* RDPG can be described by conditioning on the unobserved left and right latent-position matrices. We write down that, conditioned on $\vec {\mathbf x}_i = \vec x$ and $\vec {\mathbf y}_j = \vec y$, that if $j \neq i$, then $\mathbf a_{ij}$ is sampled independently from a $Bern(\vec x^\top \vec y)$ distribution. As before, assuming the network is loopless, $\mathbf a_{ii} = 0$. If $\mathbf A$ is the adjacency matrix for an *a posteriori* RDPG with parameter $F$, we write that $\mathbf A \sim GRDPG_n(F, G)$.
#
# # References
#
# [1] <NAME>. 1959. "On random graphs, I." Publ. Math. Debrecen 6:290–297.
#
| network_machine_learning_in_python/_build/jupyter_execute/representations/ch5/single-network-models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Model Training with TensorFlow 2.0
# ## Project Setup
# +
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
tf.__version__
# -
# ## Staging Data
# 
# **Pros and Cons of Preprocessing Data in Bigquery**
#
# - (+) Efficient transformations
# - (+) Easy to apply to new data within schema
# - (+) Scalability
# - (-) Limited Features
# - (-) Transfer over Network
# +
# %%bigquery flights_df --verbose
SELECT
-- We are binning the departure_delay to capture the data better 0, 2, 3, 6, 9, 13, 19, 28, 44, 76, 120
CASE WHEN (departure_delay < 2) THEN 1 ELSE 0 END AS departure_delay_bin_0,
CASE WHEN (2 <= departure_delay AND departure_delay < 3) THEN 1 ELSE 0 END AS departure_delay_bin_1,
CASE WHEN (3 <= departure_delay AND departure_delay < 6) THEN 1 ELSE 0 END AS departure_delay_bin_2,
CASE WHEN (6 <= departure_delay AND departure_delay < 9) THEN 1 ELSE 0 END AS departure_delay_bin_3,
CASE WHEN (9 <= departure_delay AND departure_delay < 13) THEN 1 ELSE 0 END AS departure_delay_bin_4,
CASE WHEN (13 <= departure_delay AND departure_delay < 19) THEN 1 ELSE 0 END AS departure_delay_bin_5,
CASE WHEN (19 <= departure_delay AND departure_delay < 28) THEN 1 ELSE 0 END AS departure_delay_bin_6,
CASE WHEN (28 <= departure_delay AND departure_delay < 44) THEN 1 ELSE 0 END AS departure_delay_bin_7,
CASE WHEN (44 <= departure_delay AND departure_delay < 76) THEN 1 ELSE 0 END AS departure_delay_bin_8,
CASE WHEN (76 <= departure_delay) THEN 1 ELSE 0 END AS departure_delay_bin_9,
departure_delay, -- just for demo purposes
-- We are binning the distance to capture the data better 600, 1200 km
CASE WHEN (distance < 600) THEN 1 ELSE 0 END AS distance_bin_0,
CASE WHEN (600 <= distance AND distance < 1200) THEN 1 ELSE 0 END AS distance_bin_1,
CASE WHEN (1200 <= distance) THEN 1 ELSE 0 END AS distance_bin_2,
-- Target column
CASE WHEN (arrival_delay >= 15) THEN 1 ELSE 0 END as delayed
FROM (
-- Inner Query
SELECT
ROUND(ST_DISTANCE(ST_GEOGPOINT(departure_lon, departure_lat), ST_GEOGPOINT(arrival_lon, arrival_lat))/1000) as distance,
departure_delay,
arrival_delay
FROM
`bigquery-samples.airline_ontime_data.flights`
WHERE date >= '2009-01-01'
AND date <= '2009-12-31'
AND departure_delay > 0
)
# -
flights_df.shape
flights_df.sample(n = 5)
# ## Data Preprocessing
# Why are we binning?
flights_df['departure_delay_bin'] = pd.cut(flights_df['departure_delay'], [0, 2, 3, 6, 9, 13, 19, 28, 44, 76, 120])
(flights_df
.filter(['departure_delay_bin', 'delayed'])
.groupby('departure_delay_bin')
.agg('mean')
.plot.bar(figsize = (16,5))
)
# Let's drop the departure_delay numeric variable because we are not needing it.
flights_df = flights_df.drop(columns = ["departure_delay_bin", "departure_delay"])
flights_df.dtypes
# ## Model Training using Keras API
# ### Training-Testing-Split
# +
train_df = flights_df.sample(frac=0.85,random_state=123)
test_df = flights_df.drop(train_df.index)
print(train_df.delayed.mean())
print(test_df.delayed.mean())
train_labels = train_df.pop('delayed')
test_labels = test_df.pop('delayed')
# +
# Initialize a Sequential model using TF.Keras API:
tf.keras.backend.clear_session()
model = tf.keras.models.Sequential([
tf.keras.layers.Dense(1, activation='sigmoid', input_shape = [len(train_df.keys())])])
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'],
)
# -
model.summary()
# The model itself is still very simple (logistic regression). The complexity comes through feature engineering and dummy variables. For the regression though this isn't a a problem because many of the computations will be 0 because it is essentially a weighted sum.
history = model.fit(train_df, train_labels,
epochs = 3,
callbacks = [tf.keras.callbacks.TensorBoard("logs/logistic-regression")])
# ## Testing Model Predictions
# ### Performance evaluation of logistic regression on test data
# +
test_predictions = model.predict(test_df).flatten()
test_predictions[:10]
# +
predicted_labels = (test_predictions > 0.5).astype(int)
predicted_labels[:10]
# -
results_df = pd.DataFrame({'Predicted': predicted_labels, 'Actual': test_labels})
results_df.head(10)
# ### Evaluate on test data
results = model.evaluate(test_df, test_labels)
print('test loss, test acc:', results)
# ### Create the confusion matrix
confusion_matrix = pd.crosstab(results_df['Actual'], results_df['Predicted'], margins = True)
confusion_matrix
# **Accuracy**: How many observations did we label correctly out of all the observations?
#
# **Precision**: How many of those that we labeled as positive were actually positive?
#
# **Recall**: Of all the positive class labels, how many of those did we correctly predict?
#
# **F1 Score**: F1-Score is a combination of both recall and precision. The F1 Score is low if one measure is improved at the expense of the other. For example, if Precision is 1 and Recall is 0, the F1 score is 0.
TP = confusion_matrix.loc[1, 1]
FP = confusion_matrix.loc[0, 1]
TN = confusion_matrix.loc[0, 0]
FN = confusion_matrix.loc[1, 0]
accuracy = (TP+TN)/(TP+FP+FN+TN)
accuracy
precision = TP/(TP+FP)
precision
recall = TP/(TP+FN)
recall
f1 = 2*(recall * precision) / (recall + precision)
f1
# ## Exporting the Model for Production
tf.saved_model.save(model, "model/1/")
# + language="bash"
# gsutil cp -r model/1/ gs://tensorflow-ml-course-blob/models
# -
# ## Deploy to AI Platform
# + language="bash"
# gcloud ai-platform models create flights_logistic_regression
# + language="bash"
#
# MODEL_DIR="gs://tensorflow-ml-course-blob/models/1"
# VERSION_NAME="v1"
# MODEL_NAME="flights_logistic_regression"
# FRAMEWORK="tensorflow"
#
# gcloud ai-platform versions create $VERSION_NAME \
# --model $MODEL_NAME \
# --origin $MODEL_DIR \
# --runtime-version=1.14 \
# --framework $FRAMEWORK \
# --python-version=3.5
# -
| 3.1 Lab Basic Model Training with Tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %%time
print('yes')
# +
# %matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# -
import numpy as np
import torch as t
t.autograd.set_detect_anomaly(True)
import tqdm
from torch.distributions.multivariate_normal import MultivariateNormal
import time
from csmc import *
def potential(x, _lambda, **kwargs):
"""
"""
return ((1. - _lambda)*(x**2) + _lambda * (0.5 * (x-1)**2)).sum()
def numpy_potential(x, _lambda):
return np.sum((1. - _lambda)*(x**2) + _lambda*(0.5 * (x-1.)**2))
# %%time
x = t.tensor([0.], requires_grad=True)
# %%time
x = t.tensor([0.], requires_grad=True)
pot = potential(x, 0.)
pot.backward()
# %%time
x = np.array([0.])
pot = numpy_potential(x, 0.)
np.finfo(float).eps
# %%time
from scipy import optimize
def func(x, c0, c1):
"Coordinate vector `x` should be an array of size two."
return c0 * x[0]**2 + c1*x[1]**2
x = np.ones(2)
c0, c1 = (1, 200)
eps = np.sqrt(np.finfo(float).eps)
optimize.approx_fprime(x, func, [eps,eps], c0, c1)
# %%time
pot.backward()
# let's plot the distributions at t=0,1
xs = t.tensor(np.linspace(-8, 8, 1000))
t0s = t.tensor(np.zeros(1000))
t1s = t.tensor(np.ones(1000))
ys_t0 = np.array([(-potential(x, t)).exp() for x, t in zip(xs, t0s)])
ys_t1 = np.array([(-potential(x, t)).exp() for x, t in zip(xs, t1s)])
plt.plot(xs, ys_t0, label = f"prior distribution")
plt.plot(xs, ys_t1, label = f"posterior distribution")
plt.legend()
# ok, now can we do AIS?
#
#define an initial position.
from torch.distributions.multivariate_normal import MultivariateNormal
mvn = MultivariateNormal(t.tensor([[0.]]), 0.5 * t.eye(1))
# +
num_particles = 1000
iterations = 10
incremental_works = t.zeros(num_particles, iterations)
positions = t.zeros(num_particles, iterations)
lambda_schedule = t.tensor(np.linspace(0., 1., iterations))
mvn = MultivariateNormal(t.tensor([[0.]]), 0.5 * t.eye(1))
dt=1e-2
import tqdm
for particle_idx in tqdm.trange(num_particles):
x = mvn.sample()
x.requires_grad=True
incremental_work_array, trajectory = AIS(x=x, potential=potential, lambda_schedule=lambda_schedule, propagator=MALA_propagator, dt=dt)
incremental_works[particle_idx,:] = incremental_work_array
positions[particle_idx] = t.flatten(trajectory)
# -
sns.distplot(positions[:,0])
sns.distplot(positions[:,-1])
incremental_works
cum_works = t.cumsum(incremental_works, 1)
plt.plot(incremental_works[0,:])
last_cum_works = cum_works[:,-1]
from pymbar import EXP
print(EXP(last_cum_works.numpy()))
sns.distplot(weights)
t.log(t.mean(weights))
plt.xlim(-1, 3)
# ok, so it looks like AIS works, more or less...can we make an ULA uncontrolled SMC proposal and assert that the logw returned is the same as a compute_ULA_logw?
x = MultivariateNormal(t.tensor([[0.]]), 0.5 * t.eye(1)).sample()
x.requires_grad=True
x_new, logw = ULA_uncontrolled_SMC(x, potential, t.tensor(0.), t.tensor(1.), dt=1e-2)
logw
x_new.requires_grad=True
# +
old_potential_old_x = potential(x, t.tensor(0.))
new_potential_old_x = potential(x, t.tensor(1.))
new_potential_new_x = potential(x_new, t.tensor(1.))
new_potential_old_x.backward()
new_potential_new_x.backward()
# -
compute_ULA_logw(log_gamma_old = -old_potential_old_x,
log_gamma_new = -new_potential_new_x,
x_old = x,
x_new = x_new,
grad_log_gamma_x_new = -x_new.grad,
grad_log_gamma_x_old = -x.grad,
dt = 1e-2)
# great, so we are recovering the correct log weight in the uncontrolled regime
# at this moment, i have to talk about the form of the controlled twisted initial distribution...
x = MultivariateNormal(t.tensor([0., 1.]), 0.5 * t.eye(2)).sample()
uncontrolled_alphas = t.tensor([0.5, 0.1, 0.7])
uncontrolled_mus = t.tensor([[0.1, 0.2], [0.3, 0.4], [0.5, 0.2]])
uncontrolled_sigmas = t.stack([t.eye(2) for _ in range(3)])
A_0 = t.eye(2)
b_0 = t.tensor([0., 0.])
x_0 = t.tensor([4.])
x.size()
# +
#x = MultivariateNormal(t.tensor([0., 0.]), 0.5 * t.eye(2)).sample()
uncontrolled_alphas = t.tensor([0.5, 0.0, 0.5])
uncontrolled_mus = t.tensor([[-5.0], [0.0], [5.0]])
uncontrolled_sigmas = t.stack([t.eye(1) for _ in range(3)])
A_0 = t.zeros(1)
b_0 = t.tensor([0.])
c_0 = t.tensor(0.)
twisted_gmm_proposal(
uncontrolled_alphas,
uncontrolled_mus,
uncontrolled_sigmas,
A_0,
b_0,
c_0)
# -
num_proposals = 1000
all_proposals = t.zeros(num_proposals)
lognorms = t.zeros(num_proposals)
for _ in range(num_proposals):
proposal, lognorm = twisted_gmm_proposal(
uncontrolled_alphas,
uncontrolled_mus,
uncontrolled_sigmas,
A_0,
b_0,
c_0)
all_proposals[_] = proposal
lognorms[_] = lognorm
sns.distplot(all_proposals, bins=50)
a = t.randn(3,3)
b = t.randn(3,3)
a>b
x= t.tensor(0.)
x.item()
list(x.size()) == []
# maybe we can build a twisted forward kernel now?
#
# can we play around with this a bit and see if it does what we want it to do?
# as a first sanity check, we can make sure that if the twisting is off, (i.e. uncontrolled), then there should be no
# twisting of a potential...right?
#
# if we define a potential, we can make a kernel density plot for uncontrolled smc, right?
def uncontrolled_kernel_density(potential, parameter, x, y, dt):
"""
report the log probability of transition from x to y under an Euler Maruyama discretized Unadjusted Langevin Algorithm
"""
mu, sigma = EL_mu_sigma(x,
potential(x, parameter),
dt)
mvg = MultivariateNormal(mu, sigma)
return mvg.log_prob(y)
# +
positionsx = [t.tensor([[i]], requires_grad=True) for i in np.linspace(-1,1,30)]
positionsy = [t.tensor([[i]], requires_grad=True) for i in np.linspace(-1,1,30)]
_data = np.zeros((30, 30))
for idx, x in enumerate(positionsx):
for idy, y in enumerate(positionsy):
z = t.exp(uncontrolled_kernel_density(potential, 0., x, y, 1e-2)).detach()
_data[idx, idy]=z
_data = np.array(_data)
# -
plt.contourf( np.linspace(-1,1,30), np.linspace(-1,1,30), _data)
plt.legend()
def A(x, time):
return t.eye(1)*0.
def b(x, time):
return t.tensor([0.])
def c(x, time):
return t.tensor(0.)
twisted_forward_kernel(x = t.tensor([0.], requires_grad=True),
potential = potential,
potential_parameters = t.tensor(0.),
dt = 1e-2,
A = A,
A_parameters = 0.,
b = b,
b_parameters = 0.,
c = c,
c_parameters = 0.,
d_t = t.tensor([0.]))
Theta = t.tensor([[1.]])
t.log((Theta.det().sqrt()) * t.exp(0.5 / t.tensor(1e-2)) * t.exp(-0.5 / t.tensor(1e-2) ))
twisted_gmm_proposal(uncontrolled_alphas = t.tensor([0.1, 0.2, 0.7]),
uncontrolled_mus = t.tensor([[0.], [1.], [2.]]),
uncontrolled_Sigmas = t.stack([t.eye(1), t.eye(1), t.eye(1)]),
A_0 = t.zeros(1,1),
b_0 = t.zeros(1),
c_0 = t.tensor(0.))
# ok, so it would seem at this point that we have an algorithm with components that are functioninig properly. CSMC says that the twisted weights look like:
# $$
# w_0^{\psi^{(i)}} \equiv \frac{\pi_0(\psi_0^{(i)})}{\psi_0^{(i)}(x_0)}
# $$
# and
# $$
# w_t^{\psi^{(i)}} \equiv \frac{w_t(x_{t-1}, x_t) K_t(\psi_t^{(i)})(x_{t-1})} {\psi_t^{(i)}(x_{t-1}, x_t)}
# $$
# +
def TwistedSMCSampler(potential,
uncontrolled_alphas,
uncontrolled_mus,
uncontrolled_sigmas,
A0,
b0,
c0,
A_t,
b_t,
c_t,
potential_parameter_sequence,
A_t_parameter_sequence,
b_t_parameter_sequence,
c_t_parameter_sequence,
dt,
uncontrolled=False):
"""
given a potential, A0, b0, c0, as well as functions A_t, b_t, c_t (coupled with a sequence of appropriate parameters), conduct a
forward pass of twisted SMC.
"""
#first thing to do is to make sure that the first dimension of all the sequences are the same length
sequence_first_dimensions = [entry.size()[0] for entry in [A_t_parameter_sequence, b_t_parameter_sequence, c_t_parameter_sequence]]
print(sequence_first_dimensions)
sequence_length = sequence_first_dimensions[0]
assert all(i ==sequence_first_dimensions[0] for i in sequence_first_dimensions)
#make an object that will store the trajectory and the twisted works...
trajectory = []
twisted_log_weights = []
#the potential parameter sequence is 1 greater than the rest since there is a parameterized _prior_ potential (index 0)
assert sequence_first_dimensions[0] + 1 == potential_parameter_sequence.size()[0]
#now we can conduct the pass of the 0th potential with a gaussian mixture model
x, log_pi0_psi0 = twisted_gmm_proposal(uncontrolled_alphas,
uncontrolled_mus,
uncontrolled_sigmas,
A0, b0, c0)
trajectory.append(x.numpy())
x.requires_grad=True #so we can compute potential gradient
#compute log_twisted_w0
log_phi0_x0 = quad_phi_0(x.detach(),A0.detach(), b0.detach(), c0.detach())
if uncontrolled:
assert np.isclose(log_phi0_x0, 0.)
assert np.isclose(log_pi0_psi0, 0.)
#print(log_psi0_x0)
#print(log_pi0_psi0)
initial_log_weight = log_pi0_psi0 + log_phi0_x0
#print(initial_log_weight)
twisted_log_weights.append(initial_log_weight)
#print(twisted_log_weights)
#now we can iteration from t=1 to T
for iteration in range(sequence_length):
"""
we have to make a proposal and then compute the twisted weight;
the twisted weight is the uncontrolled weight * K(\psi)(x_{t-1}) / \psi(x_{t-1}, x_t),
so we have to compute an uncontrolled weight first
"""
old_potential_parameters = potential_parameter_sequence[iteration] #previous (is current by indexing)
new_potential_parameters = potential_parameter_sequence[iteration+1] #current (is current+1 by indexing)
#make a proposal
d_t = t.tensor(0.)
# print(b_t)
# print(b_t_parameter_sequence[iteration])
x_new, logK_psi = twisted_forward_kernel(x = x,
potential = potential,
potential_parameters = new_potential_parameters,
dt = dt,
A = A_t,
A_parameters=A_t_parameter_sequence[iteration],
b = b_t,
b_parameters=b_t_parameter_sequence[iteration],
c = c_t,
c_parameters=c_t_parameter_sequence[iteration],
d_t = d_t
)
#print(f"logK_psi: {logK_psi}")
assert not x_new.requires_grad
x_new.requires_grad=True
log_gamma_old = -potential(x.detach(), old_potential_parameters) # \log \gamma_{t-1}(x_{t-1})
log_gamma_new = -potential(x_new, new_potential_parameters) # \log \gamma_{t}(x_t)
x_auxiliary = x.clone().detach().requires_grad_(True) # get a gradable detached clone of the auxiliary x variable
log_gamma_new_oldx = -potential(x_auxiliary, new_potential_parameters) # \log \gamma_{t}(x_{t-1})
log_gamma_new.backward()
log_gamma_new_oldx.backward()
grad_log_gamma_x_old = x_auxiliary.grad
grad_log_gamma_x_new = x_new.grad
uncontrolled_log_weight = compute_ULA_logw(log_gamma_old = log_gamma_old,
log_gamma_new = log_gamma_new,
x_old = x_auxiliary,
x_new = x_new,
grad_log_gamma_x_new = grad_log_gamma_x_new,
grad_log_gamma_x_old = grad_log_gamma_x_old,
dt=dt)
if uncontrolled:
"""
we are going to make a uncontrolled smc proposal and validate that the log weight is the same as the `uncontrolled
log weight`
"""
uncontrolled_log_weight_check = log_gamma_new - log_gamma_old
#forward
x_old_validate = x_auxiliary.clone().detach().requires_grad_(True)
#print(x_old_validate)
mu_forward, sigma_forward = EL_mu_sigma(x_old_validate, potential(x_old_validate, new_potential_parameters), dt)
forward_MVN = MultivariateNormal(mu_forward, sigma_forward)
logp_forward = forward_MVN.log_prob(x_new.detach())
#backward
x_new_validate = x_new.clone().detach().requires_grad_(True)
mu_backward, sigma_backward = EL_mu_sigma(x_new_validate, potential(x_new_validate, new_potential_parameters), dt)
backward_MVN = MultivariateNormal(mu_backward, sigma_backward)
logp_backward = backward_MVN.log_prob(x_old_validate.detach())
uncontrolled_log_weight_check += (logp_backward - logp_forward)
assert np.isclose(uncontrolled_log_weight_check.item(), uncontrolled_log_weight.item(), atol=1e-3), f"check: {uncontrolled_log_weight_check.item()}, computed: {uncontrolled_log_weight.item()} "
#print(old_potential_parameters)
#print(new_potential_parameters)
#compute phi_i
phi_i = (square_mahalanobis_distance(x_new.detach(), A_t(x.detach(), A_t_parameter_sequence[iteration]))
+x_new.detach().dot(b_t(x.detach(), b_t_parameter_sequence[iteration]))
+ c_t(c_t_parameter_sequence[iteration])
+ d_t
)
phi_t = quad_phi_t(x_tm1 = x.detach(),
x_t = x_new.detach(),
A_t = A_t,
b_t = b_t,
c_t = c_t,
d_t = d_t,
A_parameters = A_t_parameter_sequence[iteration],
b_parameters = b_t_parameter_sequence[iteration],
c_parameters = c_t_parameter_sequence[iteration]
)
# print(f"phi_i: {phi_i}")
# print(f"logK_psi: {logK_psi}")
# print(f"uncontrolled_log_weight: {uncontrolled_log_weight}")
twisted_log_weight = (uncontrolled_log_weight
+ logK_psi
+ phi_i
)
if uncontrolled:
assert np.isclose(logK_psi + phi_i, 0.)
twisted_log_weights.append(twisted_log_weight) #log the twisted log weights
trajectory.append(x_new.detach().numpy()) #log the new positions
x = x_new.clone().detach().requires_grad_(True) #reinitialize the new position
return trajectory, twisted_log_weights
# +
def dummy_A_t(x, param):
return t.tensor([[0.]])
def dummy_b_t(x, param):
return param[0]*(x-param[1])
def dummy_c_t(param):
return t.tensor(0.)
# +
import tqdm
#given the potential above, we are just shifting a gaussian...
iterations=100
sequence_length=20
trajs = []
twisted_log_weights = []
for iteration in tqdm.trange(iterations):
traj, twisted_log_weight = TwistedSMCSampler(potential = potential,
uncontrolled_alphas = t.tensor([1.]),
uncontrolled_mus = t.tensor([[0.]]),
uncontrolled_sigmas = t.tensor([[[0.5]]]), #this is the variance corresponding to the prior annealing protocol
A0 = t.tensor([[0.]]), #controlled gmm parameters are 0
b0 = t.tensor([0.]), #controlled gmm parameters are 0
c0 = t.tensor(0.), #controlled gmm parameters are 0
A_t = dummy_A_t,
b_t = dummy_b_t,
c_t = dummy_c_t,
potential_parameter_sequence = t.tensor(np.linspace(0,1,sequence_length)),
A_t_parameter_sequence = t.tensor([0. for _ in range(sequence_length-1)]), #this will always return 0
b_t_parameter_sequence = t.tensor([[0., 0.] for _ in range(sequence_length-1)]),
c_t_parameter_sequence = t.tensor(np.linspace(0,1,sequence_length)[1:]),
dt = 1e-2,
uncontrolled=True)
trajs.append(traj)
twisted_log_weights.append(twisted_log_weight)
# -
init_posits = [i[0] for i in trajs]
final_posits = [i[-1] for i in trajs]
sns.distplot(init_posits)
sns.distplot(final_posits)
twisted_log_weights = np.array(twisted_log_weights)
cumulatie_log_weights = np.array([np.cumsum(entry) for entry in twisted_log_weights])
final_log_weights = [entry[-1] for entry in cumulatie_log_weights]
sns.distplot(final_log_weights)
free_energy = EXP(-np.array(final_log_weights))
free_energy
np.exp(-free_energy[0])
# can we compare this to an ULA uncontrolled SMC?
# +
# trajs = []
# logws = []
# sequence_length=5
# iterations=1000
# for iteration in tqdm.trange(iterations):
# mvg = MultivariateNormal(t.tensor([[0.]]), t.tensor([[0.5]]))
# x = mvg.sample()
# x.requires_grad=True
# traj, logw = full_ULA_uncontrolled_SMC_sequence(x, potential, t.tensor(np.linspace(0,1,sequence_length)), dt=1e-3)
# #print(f"blaws: ", traj, logw)
# trajs.append(traj)
# logws.append(logw)
# -
np_logws = np.array(logws)
cum_logws = np.array([np.cumsum(entry) for entry in np_logws])
sns.distplot([entry[-1] for entry in cum_logws])
def ADP_value_training(x_sequences,
twisted_log_weights,
potential,
twisting_functions,
twisting_parameters,
minimization_iterations,
epsilon,
dt):
"""
given a set of twisting functions \psi \in \Psi(Q), and a set of i.i.d. samples {X_{0:T}^n}_{n-1}^N from Q^{\psi},
conduct approximate dynamic programming (ADP) for learning optimal sequences of potentials
arguments
x_sequences : t.tensor(R, T+1, M)
trajectory sequence where R is the number of samples, T is the number of annealing steps, and M is the
dimension of x
twisted_log_weights : t.tensor(R, T+1)
log incremental weights of the R annealing samples
twisting_functions : dict
dictionary of functions corresponding to A, b, c
twisting_initializers : dict
dictionary of A0, b0, c0
twisting_parameters : dict
dictionary of gradable parameters
contains:
A_t, b_t, c_t
minimization_iterations : int
number of iteration in minimization
epsilon : float
learning rate
dt : t.tensor(float)
timestep
"""
#initialize by setting K_{T+1}^{\psi}(\exp{-V_{T+1}})(X_T^n) = 1 for n = 1, ..., N
"""
iterate recursively from t=T, T-1, ..., 1
a. define V_{comp, t} = -log(w_t^{\psi})(X_{t-1}, X_t) - log(K_{t+1}^{\psi})
"""
num_samples, sequence_length = twisted_log_weights.size()[:2]
print(f"num_samples, sequence length: {num_samples, sequence_length}")
log_twisted_K = t.zeros(num_samples) #first log twisted Ks are 0.
print(f"the twisting parameters are: {twisting_parameters}")
#make holding variables for modified parameters:
returnable_twisting_parameters = {key: [] for key, val in twisting_parameters.items()}
#make a logger for the loss function
loss_functions = {}
for time in tqdm.tqdm(range(sequence_length)[::-1]): #iterate backward from T to 1
loss_logger = []
print(f"time: {time}")
#perform adp
A_params = twisting_parameters['A_t'][time]
b_params = twisting_parameters['b_t'][time]
c_params = twisting_parameters['c_t'][time]
#d_params = twisting_parameters['d_t'][t]
for iteration in range(minimization_iterations):
V_t_bar = -twisted_log_weights[:,time] - log_twisted_K # a.
if time==0: #then we do not compute quad_phi_t, but rather the 0th iteration equivalent
xis = [quad_phi_0(x_sequences[sample_idx, time],
A_params,
b_params,
c_params) for sample_idx in range(num_samples)
]
else:
xis = [quad_phi_t(x_tm1 = x_sequences[sample_idx, time-1],
x_t = x_sequences[sample_idx, time],
A_t = twisting_functions['A_t'],
b_t = twisting_functions['b_t'],
c_t = twisting_functions['c_t'],
d_t = twisting_functions['d_t'],
A_parameters = A_params,
b_parameters = b_params,
c_parameters = c_params) for sample_idx in range(num_samples)
]
loss_function = sum([(i-j)**2 for i, j in zip(xis, V_t_bar)])
loss_function.backward()
loss_logger.append(loss_function.item())
# print(f"loss: {loss_function.item()}")
# print(f"b params: {b_params}")
with t.no_grad():
try:
A_params -= epsilon * A_params.grad
A_params.grad.zero_()
except Exception as e:
#print(f"A_param optimizer: {e}")
pass
try:
b_params -= epsilon * b_params.grad
b_params.grad.zero_()
except Exception as e:
#print(f"b_params optimizer: {e}")
pass
try:
c_params -= epsilon * c_params.grad
c_params.grad.zero_()
except Exception as e:
#print(f"c_params optimizer: {e}")
pass
# try:
# d_params -= epsilon * d_params.grad
# except Exception as e:
# print(f"d_params optimizer: {e}")
#recompute the log_twisted_K
if time > 0:
log_twisted_K = t.tensor(
[twisted_forward_kernel(x = _x.clone().detach().requires_grad_(True),
potential = potential,
potential_parameters=twisting_parameters['potential'][time],
dt = dt,
A = twisting_functions['A_t'],
A_parameters = A_params.detach(),
b = twisting_functions['b_t'],
b_parameters = b_params.detach(),
c = twisting_functions['c_t'],
c_parameters = c_params.detach(),
d_t = t.tensor(0.))[1] for _x in x_sequences[:, time-1]
]
)
#for each of the parameters, update the returnable_twisting_parameters_dict...
for key, param in zip(['A_t', 'b_t', 'c_t'], [A_params, b_params, c_params]):
returnable_twisting_parameters[key].append(param.detach())
#add the loss logger to the loss dictionary
loss_functions[time] = np.array(loss_logger)
print(f"returnable twisting parameters: {returnable_twisting_parameters}")
flipped_returnable_twisting_parameters = {key: lst[::-1] for key, lst in returnable_twisting_parameters.items()}
return flipped_returnable_twisting_parameters, loss_functions
twisting_parameters, loss_functions = ADP_value_training(x_sequences = t.tensor(np.array(trajs)),
twisted_log_weights = t.tensor(twisted_log_weights),
potential = potential,
twisting_functions = {'A_t': dummy_A_t, 'b_t': dummy_b_t, 'c_t': dummy_c_t, 'd_t': t.tensor(0.)},
twisting_parameters = {'A_t': [t.tensor([[0.]]) for _ in range(sequence_length)],
'b_t': [t.tensor([0.], requires_grad=True)] + [t.tensor([0., 0.], requires_grad=True) for _ in range(sequence_length-1)],
'c_t': [t.tensor(0.) for _ in range(sequence_length)],
'potential': t.tensor(np.linspace(0,1,sequence_length))},
minimization_iterations=20,
epsilon = 1e-3,
dt = 1e-2)
len(loss_functions)
for i in range(100, 200):
plt.plot(loss_functions[i])
twisting_parameters
t.stack(twisting_parameters['b_t'][1:]).squeeze(0).size()
sequence_length
t.tensor([0. for _ in range(sequence_length-1)])
# +
import tqdm
#given the potential above, we are just shifting a gaussian...
iterations=50
sequence_length=200
trajs = []
twisted_log_weights = []
for iteration in tqdm.trange(iterations):
traj, twisted_log_weight = TwistedSMCSampler(potential = potential,
uncontrolled_alphas = t.tensor([1.]),
uncontrolled_mus = t.tensor([[0.]]),
uncontrolled_sigmas = t.tensor([[[0.5]]]), #this is the variance corresponding to the prior annealing protocol
A0 = t.tensor([[0.]]), #controlled gmm parameters are 0
b0 = twisting_parameters['b_t'][0], #controlled gmm parameters are 0
c0 = t.tensor(0.), #controlled gmm parameters are 0
A_t = dummy_A_t,
b_t = dummy_b_t,
c_t = dummy_c_t,
potential_parameter_sequence = t.tensor(np.linspace(0,1,sequence_length)),
A_t_parameter_sequence = t.tensor([0. for _ in range(sequence_length-1)]), #this will always return 0
c_t_parameter_sequence = t.tensor(np.linspace(0,1,sequence_length)[1:]),
b_t_parameter_sequence = t.stack(twisting_parameters['b_t'][1:]),
dt = 1e-2,
uncontrolled=False)
trajs.append(traj)
twisted_log_weights.append(twisted_log_weight)
# -
twisted_log_weights = np.array(twisted_log_weights)
cumulatie_log_weights = np.array([np.cumsum(entry) for entry in twisted_log_weights])
log_weights = np.array([entry[-1] for entry in cumulatie_log_weights])
sns.distplot(log_weights)
-np.log(np.sqrt(2))
from pymbar import EXP
free_energy = EXP(-log_weights)
free_energy
np.exp(-free_energy[0])
initial_posts = [tra[0] for tra in trajs]
final_posts = [tra[-1] for tra in trajs]
sns.distplot(initial_posts)
sns.distplot(final_posts)
# perhaps we should like to rewrite everything in numpy and use pytorch _only_ for optimization
def log_probability(potential):
"""
generate a log unnormalized probability
arguments
potential : np.float
-log unnormalized probability density
"""
return np.exp(-potential)
def EL_mu_sigma(x, func, dt, parameters, is_force_function=False, **kwargs):
"""
create mean vector and covariance marix for a multivariate gaussian proposal
"""
tau = dt/2.
if not is_force_function:
#we have to compute a numerical approximation of the gradient
force = compute_force(x, func, parameters, **kwargs)
else: #it is a force function, in which case, we just plug in the x, potential_params
force = func(x, parameters, **kwargs)
#now we can compute mu and Sigma
mu = x + tau * force
Sigma = 2*tau * np.eye(len(x))
return mu, Sigma
def compute_force(x, potential_function, potential_parameters, eps=None):
"""
given a potential function, compute an approximation of the Force (i.e. -grad(potential_function(x, parameters)))
"""
from scipy.optimize import approx_fprime
if not eps:
eps = np.sqrt(np.finfo(float).eps)
epsilons = [eps]*len(x)
grad_potential = approx_fprime(x, potential_function, epsilons, potential_parameters)
return -grad_potential
def potential(x, potential_parameters):
return np.sum(potential_parameters * x**2)
# +
def compute_ULA_logw():
"""
compute the unadjusted langevin algorithm log_weight
"""
logw = (
-potentialt_xt
+ 0.5 * x_tm1.dot(forcet_xt)
- 0.5 * xt.dot(forcet_xt)
- (dt/8.)*np.sum(forcet_xt**2)
+ potentialtm1_xtm1
- 0.5 * x_t.dot(forcet_xtm1)
+ 0.5 * xtm1.dot(forcet_xtm1)
+ (dt/8.)*np.sum(forcet_xtm1**2)
)
return logw
def compute_generalized_logw(log_gamma_old, log_gamma_new, log_forward_kernelt, log_backward_kernel_tm1):
"""
compute a generalized log incremental (unnormalized) weight
"""
logw = log_gamma_new + log_backward_kernel_tm1 - log_gamma_old - log_forward_kernelt
return logw
def twisted_gmm_components(uncontrolled_alphas, uncontrolled_mus, uncontrolled_Sigmas, A0, b0, c0):
"""
compute a twisted gaussian mixture model mixing components, twisted_mu, and twisted_Sigma
"""
assert len(uncontrolled_alphas) == len(uncontrolled_mus)
components, dimensions = uncontrolled_mus.shape
assert uncontrolled_Sigmas.shape == (components, dimension, dimension)
#compute mixture components
Sigma_tilde_js = np.linalg.inv(np.linalg.inv(uncontrolled_Sigmas) + 2.0*A0)
log_zetas = np.array([
gmm_log_zetas(sigma_tilde_j, sigma_j, mu_j, b0, c0)
for sigma_tilde_j, sigma_j in
zip(Sigma_tilde_js, uncontrolled_Sigmas)
])
log_alpha_tildes = np.log(uncontrolled_alphas) + log_zetas
return log_alpha_tildes, Sigma_tilde_js
def twisted_gmm_proposal(log_alpha_tildes, Sigma_tilde_js, b0):
"""
make a twisted gaussian mixture model proposal
"""
from scipy.special import logsumexp
from scipy.stats import multivariate_normal
normalized_alpha_tildes = np.exp(log_alpha_tildes - logsumexp(log_alpha_tildes))
#choose a component
component_index = np.random.choice(range(len(normalized_alpha_tildes)), p = normalized_alpha_tildes)
#then choose a position based on that gaussian
Sigma_tilde_j = Sigma_tilde_js[component_index]
mu_j = uncontrolled_mus[component_index]
Sigma_j = uncontrolled_Sigmas[component_index]
twisted_mean = np.matmul(Sigma_tilde_j, np.matmul(np.linalg.inv(Sigma_j), mu_j) - b0)
twisted_Sigma = Sigma_tilde_j
x = multivariate_normal.rvs(mean=twisted_mean, cov = twisted_Sigma)
logpdf = multivariate_normal.logpdf(x, mean=twisted_mean, cov = twisted_Sigma)
return x, logpdf
def compute_twisted_gmm_lognormalizer(log_alpha_tildes):
"""
compute the twisted gaussian mixture model log normalization constant with unnormalized log_alpha_tildes
"""
from scipy.special import logsumexp
return logsumexp(log_alpha_tildes)
# -
def gmm_log_zetas(Sigma_tilde_j, Sigma_j, mu_j, b0, c0):
"""
compute the logzeta_js mixture components
"""
from scipy.spatial.distance import mahalanobis
comp1 = -0.5 * np.log(np.linalg.det(Sigma_j))
comp2 = 0.5 * np.log(np.linalg.det(Sigma_tilde_j))
comp3 = 0.5 * mahalanobis(np.matmul(np.linalg.inv(Sigma_j), mu_j), b0, np.linalg.inv(Sigma_tilde_j))**2
comp4 = -0.5 * mahalanobis(mu_j, np.zeros(len(mu_j)), Sigma_j)**2
comp5 = -c0
return comp1 + comp2 + comp3 + comp4 + comp5
# +
def Theta_t(x_tm1, A_t, dt):
"""
compute Theta_t = (I_d _ 2*dt*A_t(x_tm1))^-1
"""
theta = np.linalg.inv(np.eye(len(x_tm1)) + 2. * dt * A_t)
return theta
def f_t(x_tm1, potential_function, parameters, dt, **kwargs):
"""
NOTE : parameters should reflect potential function at time t (not t-1)
compute f_t(x_tm1) = x_tm1 + 0.5 * dt * forcet(x_tm1) = mu
"""
mu, cov = EL_mu_sigma(x_tm1, potential_function, dt, parameters, is_force_function=False, **kwargs)
return mu
def twisted_forward_tools(x, potential_function, potential_parameters, A_function, A_parameters, dt, **kwargs):
"""
make a twisted forward
"""
dimension = len(x) #grab the dimension of x
f_t = f_t(x, potential_function, potential_parameters, dt)
A_t = A_function(x, A_parameters)
#theta and f
theta = Theta_t(x, A_t, dt)
f = f_t(x, potential, potential, dt)
return theta, f
def twisted_forward_proposal(theta, f, dt, b_t, **kwargs):
"""
make a forward twisted proposal
"""
twisted_mean = np.matmul(theta, f - dt*b_t)
twisted_covariance = dt * theta
x = multivariate_normal.rvs(mean=twisted_mean, cov = twisted_covariance)
return x
def twisted_forward_log_normalizer(theta, f, b_t, dt, c, d):
"""
perform one line computation to calculate the log normalization constant of the twisted forward proposal
"""
from scipy.spatial.distance import mahalanobis
#there are 3 components that will be computed separately and added.
comp1 = 0.5 * np.linalg.det(theta)
comp2 = (1./(2.*dt)) * mahalanobis(f, dt*b_t, np.linalg.inv(theta))**2
comp3 = -(1./(2.*dt)) * f.dot(f) - c - d
log_forward_normalizer = comp1 + comp2 + comp3
return log_forward_normalizer
# -
a = np.array([1, 2, 3, 4]).transpose()
b = np.array([0, 2, 3, 4])
a.dot(b)
a**2
a = np.array([[0., 1., 2.], [0., 1., 4.]]).shape
a
q = np.random.rand(2, 2)
np.matmul(q, np.array([1, 2]))
A0 = np.random.rand(2,2)
np.linalg.inv(np.linalg.inv(q) - A0)
np.array([np.linalg.inv(it) - A0 for it in q])
from scipy.stats import multivariate_normal
mv = multivariate_normal.rvs(mean = np.array([0., 1.]), cov = np.eye(2))
# %%time
mv = multivariate_normal.rvs(mean = np.random.randn(2000), cov = np.eye(2000))
| troubleshooting/csmc_stage.ipynb |