code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# 
# # Linear Regression with Python
# !pip3 install pandas
# !pip3 install numpy
# !pip3 install seaborn
# !pip3 install matplotlib
# !pip3 install scikit-learn
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
# +
import numpy as np
X = 2 * np.random.rand(100, 1)
y = 4 + 3 * X + np.random.randn(100, 1)
# -
plt.plot(X, y, "b.")
plt.xlabel("$x_1$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.axis([0, 2, 0, 15])
plt.show()
lin_reg = LinearRegression()
lin_reg.fit(X, y)
lin_reg.intercept_, lin_reg.coef_
wine_data = pd.read_csv('../data/winequality-red.csv')
wine_data.head(100)
wine_data.describe()
correlation = wine_data.corr()
fig = plt.subplots(figsize=(10,10))
sns.heatmap(correlation,vmax=1,square=True,annot=True,cmap='Blues')
X = wine_data.drop('quality', axis=1)
y = wine_data['quality']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=101, train_size=0.8)
regressor = LinearRegression()
regressor.fit(X_train,y_train)
from sklearn.metrics import mean_absolute_error, mean_squared_error
y_pred = regressor.predict(X_test)
mse = mean_squared_error(y_test, y_pred)
rmse = np.sqrt(mse)
print(mse)
print(rmse)
mae = mean_absolute_error(y_test, y_pred)
print(mae)
y_pred_round = np.round(y_pred, 1)
mse = mean_squared_error(y_pred_round, y_test)
mse
np.sqrt(mse)
import pickle
with open ('../docker/model.pkl', 'wb') as file:
pickle.dump(regressor, file)
| Presentations/.ipynb_checkpoints/Linear Regression - Wine Quality-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## CSCS530 Winter 2015
# #### Complex Systems 530 - Computer Modeling of Complex Systems (Winter 2015)
#
# * Course ID: CMPLXSYS 530
# * Course Title: Computer Modeling of Complex Systems
# * Term: Winter 2015
# * Schedule: Wednesdays and Friday, 1:00-2:30PM ET
# * Location: 120 West Hall (http://www.lsa.umich.edu/cscs/research/computerlab)
# * Teachers: [<NAME>](https://www.linkedin.com/in/bommarito) and [<NAME>](https://www.linkedin.com/pub/sarah-cherng/35/1b7/316)
#
# #### [View this repository on NBViewer](http://nbviewer.ipython.org/github/mjbommar/cscs-530-w2016/tree/master/)
#
# # Buffon's Needle
#
# One of the first "modern" simulations ever performed was Buffon's needle (and subsequent correction by Laplace). This simulation was devised as a physical means of estimating $\pi$, and is reasonably accurate with simple measurement tools and a small number of samples.
#
# Many of the visual elements of this notebook were inspired by this wonderful book:
# https://github.com/unpingco/Python-for-Signal-Processing/blob/master/Buffons_Needle_Sim.ipynb
# +
# %matplotlib inline
# Imports
import numpy
import numpy.random
import matplotlib.pyplot as plt
import seaborn
seaborn.set_style("darkgrid")
# -
# Draw center, angle, calculate start/end points for length
def sample_line(length=0.5):
# Draw center
xC, yC = numpy.random.random(2)
# Draw angle
theta = numpy.random.uniform(0, numpy.pi)
# Get endpoints
x0 = xC - (0.5 * length) * numpy.cos(theta)
x1 = xC + (0.5 * length) * numpy.cos(theta)
y0 = yC - (0.5 * length) * numpy.sin(theta)
y1 = yC + (0.5 * length) * numpy.sin(theta)
return x0, x1, y0, y1
def is_unit(z):
return (0 < z < 1)
# +
N = 1000
# Setup plot
f, ax = plt.subplots()
ax.set_xlim((-0.25, 1.25))
ax.set_ylim((-0.25, 1.25))
ax.set_aspect('equal')
# Drop needles
count_in = 0
count_out = 0
for i in range(N):
x0, x1, y0, y1 = sample_line()
if all(map(is_unit, [x0, x1, y0, y1])):
ax.add_line(plt.Line2D([x0, x1], [y0, y1], alpha=0.1, marker="o", markersize=4, color="red"))
count_in += 1
else:
ax.add_line(plt.Line2D([x0, x1], [y0, y1], alpha=0.1, marker="o", markersize=4, color="blue"))
count_out += 1
# Draw box around unit square
ax.add_patch(plt.Rectangle((0, 0), 1, 1, alpha=0.25, color="green"))
# -
p_cut = float(count_out) / (count_in + count_out)
print("In/out: {0}/{1}".format(count_in, count_out))
print("P(out): {0}".format(p_cut))
# Theoretical cut
(1 - numpy.power(1 - 2 * length / numpy.pi, 2))
estimate = (2 * length) / (1 - numpy.sqrt(1 - p_cut))
print("Estimate: {0}".format(estimate))
# +
def run_simulation(samples=100, simulations=100):
# Run sims
length = 0.5
# Iterate over each sim
pi_estimates = []
for n in range(simulations):
# Drop needles
count_in = 0
count_out = 0
for i in range(samples):
x0, x1, y0, y1 = sample_line()
if all(map(is_unit, [x0, x1, y0, y1])):
count_in += 1
else:
count_out += 1
# Calculate cut probability
p_cut = float(count_out) / (count_in + count_out)
# Calculate estimate
estimate = (2 * length) / (1 - numpy.sqrt(1 - p_cut))
pi_estimates.append(estimate)
return pi_estimates
pi_estimates = run_simulation()
_ = plt.hist(pi_estimates)
plt.axvline(numpy.mean(pi_estimates), color="red", alpha=0.5)
print("Estimate: {0}".format(numpy.mean(pi_estimates)))
| notebooks/basic-random/004-buffons-needle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from matplotlib.pyplot import plot, figure, \
arrow, Circle, gca, text, bar, grid
import numpy as np
figure(figsize= (9, 9), dpi = 150)
x = 1
y = np.pi/4
plot(x, y, "go")
#arrow(0, 0, x, y, color = "red", linewidth = 1.5, linestyle = "dashed")
arrow(0, 0, x, y, head_width = 0.04, head_length = 0.04, color = "b")
# +
arrow(0,0,1.1,0,head_width=0.04,head_length=0.08)
arrow(0,0,-1.1,0,head_width=0.04,head_length=0.08)
arrow(0,0,0,-1.1,head_width=0.04,head_length=0.08)
arrow(0,0,0,1.1,head_width=0.04,head_length=0.08)
arrow(0, 0, np.cos(x), y, head_width = 0.04, head_length = 0.04, color = "b")
gca().add_patch(Circle((0, 0), 1, color = "black", fill = False))
text(np.cos(x), y + 0.1, "It is the point: (cos(1), π/4)", fontsize=12)
grid(True)
# -
bar(np.random.randint(0, 10, 100),np.random.randint(0, 10, 100))
# +
import sys
sys.path.insert(0, '../qworld/include/')
from drawing import draw_axes, draw_unit_circle, draw_quantum_state, draw_qubit, draw_qubit_grover, show_plt
from quantum_state import random_qstate_by_value, random_qstate_by_angle, angle_qstate
from grover import giant_oracle, giant_oracle2, giant_diffusion, Uf, Uf_8
# -
draw_quantum_state(2, 3, "test")
draw_qubit()
| practice /Drawing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:mp]
# language: python
# name: conda-env-mp-py
# ---
# # Materials API - Exercise 2: Using the MPRester and Pymatgen to Find Materials With Exotic Mechanical Properties
#
# The tetragonal SiO$_2$ polymorph $\alpha$-cristobalite is one of the very few crystalline materials known to have a negative average Poisson's ratio, which means that its cross-section expands under tensile strain rather than contracting. This property can be extremely useful in a variety of applications such as scratch-resistant coatings and high-toughness ceramics.
#
# Why does $\alpha$-cristobalite exhibit this property while other materials do not? The prevailing hypothesis is that $\alpha$-cristobalite's negative Poisson's ratio is a result of its crystal structure. If that's the case, then perhaps we can find other materials with this exotic property by looking for materials with similar structures and then calculating their Poisson's ratios.
# ## Step 1: Retrieve the structure of $\alpha$-cristobalite
#
# First, we should find the ground state structure for $\alpha$-cristobalite, which has the spacegroup $P4_12_12$ [92]. Remember, stable materials have *low* energy above hull (`e_above_hull`).
#
# _Hint: Query for the formula and spacegroup for the desired compound and then select the one with the lowest e_above_hull if there are multiple results._
# +
# Your code here
from pymatgen import MPRester
with MPRester("9LO7vrUFbVTMaxdK") as mpr:
ac = mpr.query({"spacegroup.number": 92,
"pretty_formula": "SiO2",
"e_above_hull": {"$lt": 0.1}},
["material_id", "structure"])[0]
# -
ac_structure = ac["structure"]
# ## Step 2: Build a structure comparison engine and test it out
#
# The code below creates a structure matcher object that can be used to compare if two structures are simlar (loose tolerances = similar, tight tolerances = identical). Please verify that the structure matcher works for identical structure by comparing the structure of AC with itself.
#
# _Hint: Replace <YOUR CODE HERE> with the two structures you want to compare, separated by a comma._
# +
##### Don't edit code below #####
from pymatgen.analysis.structure_matcher import StructureMatcher
from pymatgen.analysis.structure_matcher import FrameworkComparator
# This line initializes our comparator. Please don't play with its tolerances until after you are done
comparison_engine = StructureMatcher(ltol=.2, stol=.5, angle_tol=10, primitive_cell=True,
scale=True, attempt_supercell=True,
comparator=FrameworkComparator())
##### Don't edit code above #####
# -
print(comparison_engine.fit(ac_structure, ac_structure))
# We know that the high-temperature phase of cristobalite, $\beta$-cristobalite [(mp-546794)](https://materialsproject.org/materials/mp-546794/), has a very similar structure to $\alpha$-cristobalite. Let's see if the structure matcher agrees. Please retreive the structure for `mp-546794` and then compare it with our prototype structure.
#
# _Hint: No need to re-initialize the structure matcher. Just call it's `fit` function again._
# Your code here
with MPRester("9LO7vrUFbVTMaxdK") as mpr:
bc_structure = mpr.get_structure_by_material_id("mp-546794")
print(comparison_engine.fit(bc_structure, ac_structure))
# Just to make sure we haven't increased the tolerances too much, please try it against a random compound to make sure it's not matching dissimilar structures.
#
# _Hint: "mp-4991" is a good random MPID you can use_
# Your code here
# Your code here
with MPRester("9LO7vrUFbVTMaxdK") as mpr:
bc_structure = mpr.get_structure_by_material_id("mp-4991")
print(comparison_engine.fit(bc_structure, ac_structure))
# ## Step 3: Get a set of candidate structures to compare
#
# Now that we have our comparator, we need some candidates to screen!
#
# Imagine that we have an experimental colleague, <NAME>, who is an expert at synthesizing vanadate compounds. We have a hunch that some of the vanadates coming out of Dr. Tsarpinski's lab might have similar structures to $\alpha$-cristobalite and therefore might have negative Poisson's ratios. Let's see if we're right:
#
# For our search, we want to start with a set of structures that are:
# * Computationally tractable, so not too many sites (i.e `nsites` <50 is a good number)
# * Not too unlikely to be synthesizable (energy above hull <100 meV, i.e. `{e_above_hull: {"$lt": 0.1}`)
# * Have a "vanadate" composition, i.e. `"*V3O8"`
#
# Construct and execute a query to get the structure, MPID, and pretty_formula for all materials that match these criteria:
#
# _Aside; <NAME> is an anagram. There is a prize for the first workshop atendee that can guess the original phrase it comes from. (Submit your anwswers on Slack, please.)_
# Your code here
with MPRester("9LO7vrUFbVTMaxdK") as mpr:
vanadate_ids = [entry["material_id"] for entry in mpr.query('*V3O8', ["material_id"])]
vanadates = mpr.query({"material_id": {"$in": vanadate_ids},
"e_above_hull": {"$lt": 0.1},
"nsites":{"$lt": 50}}, ["material_id", "structure"])
# ## Step 4: Screen the vanadates for similar structures to $\alpha$-cristobalite and then check if our hypothesis is valid by querying for their Poisson's ratios.
#
# Now that we have a list of vanadates, let's screen it for similar structures. After we have the similar structures, make one final query to the Materials API to retrieve the `pretty_formula` and `elasticity.homogenious_poisson` property for each one.
#
# _Hint: Create an empty list for matches and then iterate through the vanadate entries with a for loop. If its structure is similar to $\alpha$-cristobalite, append its `material_id` to a list of matches. After you have a match list, query the Materials API for entries with a `material_id` that is in your matches list using the `$in` mongodb operator (e.g `{"material_id": {"$in$: matches}}`)_
# +
# Your code here
matches = []
for v in vanadates:
if comparison_engine.fit(ac_structure, v["structure"]):
matches.append(v["material_id"])
with MPRester("9LO7vrUFbVTMaxdK") as mpr:
elastic_data = mpr.query({"material_id": {"$in": matches}}, ['material_id', 'pretty_formula', 'elasticity.homogeneous_poisson'])
for e in elastic_data:
print(e)
# -
| workshop/lessons/04_materials_api/MAPI Exercise 2 Sloutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Analysis of combined data sets Cf072115 - Cf072215b
#
# <NAME>
# University of Michigan
# 2/9/2018
# We are expecting 8.5" of snow today. ..........
#
# I am combining four data sets:
#
# * Cf072115
# * Cf072115b
# * Cf072215a
# * Cf072215b
#
# I have combined the `sparse_bhm.npz`, `sparse_bhm_neg.npz`, and `singles_hist.npz` files on flux and downloaded to my local machine.
#
# Now I will revive those files and produce `bhp_nn` for positive and negative time ranges. This is so that I don't have to keep importing the entire `bhm` files each time because it takes forever and a ton of memory.
#
# I'm going to use 1 ns time binning for this to save 16x space in the time dimensions.
import os
import sys
import matplotlib.pyplot as plt
import matplotlib.colors
import numpy as np
import imageio
import scipy.io as sio
sys.path.append('../../scripts/')
import bicorr as bicorr
import bicorr_plot as bicorr_plot
# %load_ext autoreload
# %autoreload 2
# Use seaborn to make plots prettier
import seaborn as sns
sns.set(style='ticks')
# # Load `det_df`, channel lists
os.listdir('../../meas_info/')
det_df = bicorr.load_det_df('../../meas_info/det_df_pairs_angles.csv',plot_flag=True)
chList, fcList, detList, num_dets, num_det_pairs = bicorr.build_ch_lists(print_flag=True)
# # Calculate `num_fissions`
#
# Sometime it would be convenient to store all of the measurement in a database, and then load it according to which datasets are specified in `note`. For now, input the data manually.
#
# (Calculated in excel file `analysis_status.xlsx`)
num_fissions = int(int(sio.loadmat('datap/num_fissions.mat')['num_fissions'])*float(sio.loadmat('datap/fc_efficiency.mat')['fc_efficiency']))
num_fissions
# # Load data
# ## Load `sparse_bhm.npz`, revive `bhm`
os.listdir()
sparse_bhm, dt_bin_edges, note = bicorr.load_sparse_bhm(filepath='datap')
sparse_bhm.nbytes
bhm_pos = bicorr.revive_sparse_bhm(sparse_bhm, det_df, dt_bin_edges)
(bhm_pos.nbytes)/16 # .5 ns bins
# I'm going to perform the background subtraction, then store `bhp_nn_diff` for all 990 pairs to disk so I can reload it later.
# ## Load `sparse_bhm_neg.npz`, revive `bhm_neg`
sparse_bhm_neg, dt_bin_edges_neg, note_neg = bicorr.load_sparse_bhm(filename = 'sparse_bhm_neg.npz', filepath='datap')
bhm_neg = bicorr.revive_sparse_bhm(sparse_bhm_neg, det_df, dt_bin_edges_neg)
# ## Load `singles_hist.npz`
singles_hist, dt_bin_edges_sh, dict_det_to_index, dict_index_to_det = bicorr.load_singles_hist(filepath='datap')
help(bicorr.load_singles_hist)
plt.figure(figsize=(4,3))
dt_bin_centers_sh = (dt_bin_edges_sh[:-1]+dt_bin_edges_sh[1:])/2
plt.plot(dt_bin_centers_sh,np.sum(singles_hist,axis=(0,1)))
plt.xlabel('Time (ns)')
plt.ylabel('Number of events')
plt.title('TOF distribution, all events')
plt.yscale('log')
sns.despine(right=False)
bicorr_plot.save_fig_to_folder('singles_hist_allt_allp',extensions=['png','pdf'])
plt.show()
plt.figure(figsize=(4,3))
plt.plot(dt_bin_centers_sh,np.sum(singles_hist[0,:,:],axis=(0)))
plt.plot(dt_bin_centers_sh,np.sum(singles_hist[1,:,:],axis=(0)))
plt.xlabel('Time (ns)')
plt.ylabel('Number of events')
plt.title('TOF distribution, all detectors')
plt.legend(['N','G'])
plt.yscale('log')
sns.despine(right=False)
bicorr_plot.save_fig_to_folder('singles_hist_ng_allp',extensions=['png','pdf'])
plt.show()
plt.figure(figsize=(4,3))
plt.plot(dt_bin_centers_sh,singles_hist[0,dict_det_to_index[2],:])
plt.plot(dt_bin_centers_sh,singles_hist[1,dict_det_to_index[2],:])
plt.xlabel('Time (ns)')
plt.ylabel('Number of events')
plt.title('TOF distribution, channel 2')
plt.legend(['N','G'])
plt.yscale('log')
sns.despine(right=False)
bicorr_plot.save_fig_to_folder('singles_hist_ng_ch2',extensions=['png','pdf'])
plt.show()
# # Coarsen `bhm` to 1 ns. time binning.
print(bhm_pos.shape)
print(bhm_neg.shape)
bhm_pos, dt_bin_edges = bicorr.coarsen_bhm(bhm_pos,dt_bin_edges, 4,True)
bhm_neg, dt_bin_edges_neg = bicorr.coarsen_bhm(bhm_neg,dt_bin_edges_neg,4,True)
print(bhm_pos.shape)
print(bhm_neg.shape)
# # Produce `bhp` for $nn$ events
# One key piece of data that I am going to work with for producing multiple plots is the `bhp` for $nn$ events across all detector pairs. (Actually, only the pairs not next to the fission chambers)
#
# So I am going to produce that for future use. This will be copied into another notebook, but the process of loading all of the data is the same so I'm doing that here since all the data is loaded.
#
# I'm going to make this with 1 ns time binning to keep the file size manageable.
# ## Produce `pair_is` for pairs not next to fission chamber
pair_is = bicorr.generate_pair_is(det_df, ignore_fc_neighbors_flag=True)
len(pair_is)
# Look at this distribution.
plt.figure(figsize=(6,6))
plt.plot(det_df.iloc[pair_is]['d1'],det_df.iloc[pair_is]['d2'],'sk')
for i in [1,17,33]:
plt.axvline(i,c='r')
plt.axhline(i,c='r')
plt.xlabel('Detector 1 channel')
plt.ylabel('Detector 2 channel')
plt.title('Included detector pairs')
sns.despine(right=False)
bicorr_plot.save_fig_to_folder(fig_filename='pair_is_without_fc_neighbors',extensions=['png','pdf'])
plt.show()
# ## Create `bhp_nn_pos`, `bhp_nn_neg`, `bhp_nn_diff`
#
# Following instructions from `bicorr > methods > nn_sum_and_br_subtraction`.
#
# I'm going to create arrays with 1 ns time binning and save them to disk, so I can easily reload them in the future.
bhm_pos.shape
bhm_pos_shape = bhm_pos[pair_is,:,:,:].shape
print(bhm_pos_shape)
# The challenge here is that I want to preserve the dimension of `pair_is` (I don't want to sum across all pairs in `pair_is`). How can I do this without significantly modifying my code base?
#
# Set up arrays to fill
bhp_nn_pos = np.zeros((bhm_pos_shape[0],bhm_pos_shape[2],bhm_pos_shape[3]))
bhp_nn_neg = np.zeros((bhm_pos_shape[0],bhm_pos_shape[2],bhm_pos_shape[3]))
bhp_nn_neg.shape
# +
for i in np.arange(len(pair_is)):
pair_i = pair_is[i]
bhp_nn_pos[i,:,:] = bicorr.build_bhp(bhm_pos,dt_bin_edges,pair_is=[pair_i],type_is=[0])[0]
bhp_nn_neg[i,:,:] = bicorr.build_bhp(bhm_neg,dt_bin_edges_neg,pair_is=[pair_i],type_is=[0])[0]
print(bhp_nn_pos.shape)
print(bhp_nn_neg.shape)
# -
# Plot a few to make sure they look good.
i = 500
bicorr_plot.bhp_plot(bhp_nn_pos[i,:,:],dt_bin_edges,show_flag=True,title='bhp_nn_pos at i={}'.format(i))
bicorr_plot.bhp_plot(bhp_nn_neg[i,:,:],dt_bin_edges_neg,show_flag=True,title='bhp_nn_neg at i={}'.format(i))
# Plot them now as sums across all pairs.
bicorr_plot.bhp_plot(np.sum(bhp_nn_pos,axis=0),dt_bin_edges,show_flag=True,title='bhp_nn_pos')
bicorr_plot.bhp_plot(np.sum(bhp_nn_neg,axis=0),dt_bin_edges_neg,show_flag=True,title='bhp_nn_neg')
# Now create `bhp_nn_diff`.
#
# Question: Should I create `bhp_nn_diff` here, or work with `bhp_nn_pos` and `bhp_nn_neg`? The data is still pretty sparse, so `bhp_nn_diff` would end up with a lot of negative values in it. Mathematically, once I start taking sums, it would be the same. But I will always have to load `bhp_nn_pos` and `bhp_nn_neg` anyway, so I could just create `bhp_nn_diff` whenever I load them. Yeah. Do that.
bhp_nn_diff = np.subtract(bhp_nn_pos.astype(np.int32),bhp_nn_neg[:,::-1,::-1].astype(np.int32))
bicorr_plot.bhp_plot(np.sum(bhp_nn_diff,axis=0),dt_bin_edges,show_flag=True,title='bhp_nn_diff')
i = 4
bicorr_plot.bhp_plot(bhp_nn_diff[i,:,:],dt_bin_edges,show_flag=True,title='bhp_nn_diff')
# One thing to keep in mind is that `bicorr.bicorr_plot` does not show negative values, so the background subtraction makes it look "cleaner" than it is in reality.
# ## Verify `bhp_nn_diff`
# Calculate `bhp_nn_diff` pair by pair and make sure it matches what I've already done.
bhp_nn_diff_pair = np.zeros((861, 200, 200))
for i in np.arange(len(pair_is)):
pair_i = pair_is[i]
bhp_nn_diff_pair[i,:,:] = np.subtract(bhp_nn_pos[i,:,:].astype(np.int32),bhp_nn_neg[i,::-1,::-1].astype(np.int32))
bhp_nn_diff_pair.shape
np.array_equal(bhp_nn_diff,bhp_nn_diff_pair)
# # Store `bhp_nn_pos`, `bhp_nn_neg`
#
# * `bhp_nn_neg, bhp_nn_pos`
# * `dt_bin_edges`
# * `pair_is`
# * `note`
#
# Save these four arrays and the note to the same `.npz` file
note = 'Stored from Cf072115_to_Cf072215b with 1 ns time binning. Pairs are without fc neighbors. -PFS, 2/9/18'
save_filename = 'datap/bhp_nn_by_pair_1ns'
np.savez(save_filename, bhp_nn_neg = bhp_nn_neg, bhp_nn_pos = bhp_nn_pos,
dt_bin_edges = dt_bin_edges, pair_is = pair_is, note = note)
# # Reload instructions
#
# In my other analysis files, I'm going to load these variables from disk.
#
# Restart the notebook so I start fresh and start from here.
whos
# Go back and import all the packages.
whos
load_filename = 'datap/bhp_nn_by_pair_1ns.npz'
npzfile = np.load(load_filename)
print(npzfile.files)
print(npzfile['note'])
pair_is = npzfile['pair_is']
bhp_nn_pos = npzfile['bhp_nn_pos']
bhp_nn_neg = npzfile['bhp_nn_neg']
dt_bin_edges = npzfile['dt_bin_edges']
# Calculate `bhp_nn_diff`.
bhp_nn_diff = np.subtract(bhp_nn_pos.astype(np.int32),bhp_nn_neg[:,::-1,::-1].astype(np.int32))
bhp_nn_diff.shape
# Plot them to make sure they look good.
bicorr.bicorr_plot(np.sum(bhp_nn_diff,axis=0),dt_bin_edges,show_flag=True,title='bhp_nn_diff')
| analysis/Cf072115_to_Cf072215b/create_bhp_nn_1ns.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0
# language: julia
# name: julia-1.3
# ---
using GeometryTypes: HyperRectangle, FlexibleConvexHull, Vec, contains
using CoordinateTransformations: AffineMap
using StaticArrays
using EnhancedGJK
using MeshCat
using Polyhedra
using StaticArrays: SVector
using GeometryTypes: HyperSphere, Point
# +
geo1 = HyperRectangle{3,Float32}(Float32[-0.08559614, -0.125, 0.0], Float32[0.117596135, 0.25, 0.07049995])
geo2 = FlexibleConvexHull{Vec{3,Float32}}(Vec{3,Float32}[[-1.4459023, 0.8858041, 0.9851251], [-0.32686943, -0.23770937, -0.0762], [-0.32686943, -0.23770937, 0.204528], [-0.32686943, 0.23770937, -0.0762], [-0.32686943, 0.23770937, 0.204528], [0.32682246, -0.23770937, -0.0762], [0.32682246, -0.23770937, 0.204528], [0.32682246, 0.23770937, -0.0762], [0.32682246, 0.23770937, 0.204528]])
convex_hull_gg(vertices) = collect(points(vrep(polyhedron(hrep(polyhedron(vrep(vertices)))))))
geo2_ch = FlexibleConvexHull(map(Vec{3,Float32}, convex_hull_gg(map(Array, geo2.vertices))))
# geo2_ch = FlexibleConvexHull{Vec{3,Float32}}(Vec{3,Float32}[[-0.32686943, -0.23770937, 0.204528], [-0.32686943, -0.23770937, -0.0762], [-0.32686943, 0.23770937, -0.0762], [0.32682246, -0.23770937, 0.204528], [0.32682246, -0.23770937, -0.0762], [-1.4459023, 0.8858041, 0.9851251], [0.32682246, 0.23770937, -0.0762], [0.32682246, 0.23770937, 0.204528]])
definitely_inside__Fgeo2 = Vec{3, Float32}(-1.4459023, 0.8858041, 0.9851251)
tx__Fworld__Fgeo1 = AffineMap((@SArray Float32[1.0 -1.3267949e-6 1.7603859e-12; 1.3267949e-6 1.0 1.3267932e-6; -3.5207683e-12 -1.3267932e-6 1.0]), (@SArray Float32[0.28507775, 4.0943226e-7, 1.0193151]))
tx__Fworld__Fgeo2 = AffineMap((@SArray Float32[0.809017 -0.58778524 0.0; 0.58778524 0.809017 0.0; 0.0 0.0 1.0]), (@SArray Float32[2.0, 0.1, 0.0762]))
# Convex hull check
# result1 = EnhancedGJK.gjk(geo1, geo2, tx__Fworld__Fgeo1, tx__Fworld__Fgeo2)
# result2 = EnhancedGJK.gjk_original(geo1, geo2, tx__Fworld__Fgeo1, tx__Fworld__Fgeo2)
# Convex hull check
result1 = EnhancedGJK.gjk(geo1, geo2_ch, tx__Fworld__Fgeo1, tx__Fworld__Fgeo2)
result2 = EnhancedGJK.gjk_original(geo1, geo2_ch, tx__Fworld__Fgeo1, tx__Fworld__Fgeo2)
print("Final weights: ", result1.weights,"\n")
print("Final simplex: ", result1.simplex,"\n")
print("Iterations: ", result1.iterations,"\n") # terminates in 5 iterations
if result1.in_collision
@show simplex_penetration_distance(result1)
else
@show separation_distance(result1)
end
print("\n", "Final weights: ", result2.weights,"\n")
print("Final simplex: ", result2.simplex,"\n")
print("Iterations: ", result2.iterations,"\n") # continues to iterate
@show separation_distance(result2)
print("Collision Checks: ", "\n")
definitely_inside_Fgeo1 = inv(tx__Fworld__Fgeo1)(tx__Fworld__Fgeo2(definitely_inside__Fgeo2))
print(contains(geo1, definitely_inside_Fgeo1),"\n")
print(result1.in_collision,"\n") # pass
print(result2.in_collision,"\n") # fail
# -
function visualize_simplex(vis::Visualizer, simplex)
p = polyhedron(vrep(simplex))
setobject!(vis[:simplex], Polyhedra.Mesh(p))
for (i, point) in enumerate(simplex)
setobject!(vis["p$i"], HyperSphere(Point(point), convert(eltype(point), 0.03)))
end
end
# +
convex_mesh(points) = Polyhedra.Mesh(Polyhedra.polyhedron(Polyhedra.vrep(map(Array, points))))
vis = Visualizer()
# open(vis)
IJuliaCell(vis)
# +
setobject!(vis[:geo1], geo1)
settransform!(vis[:geo1], tx__Fworld__Fgeo1)
setobject!(vis[:geo2], convex_mesh(geo2.vertices))
settransform!(vis[:geo2], tx__Fworld__Fgeo2)
# visualize_simplex(vis[:simplex], result1.simplex)
# -
| test/notebooks/sv_degeneracy_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3.7Test
# language: python
# name: python3.7test
# ---
# # Notes
#
# Use state-level version of ensemble (once Xiao uploads them)
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="darkgrid")
# +
DEATH_DATE = '4/3/2020'
ENSEMBLE_FILE = "../predictions_state_level_04_01.csv"
IHME_FILE = "./ihme/2020-03-31.csv"
PROJECTION_DATE = '4/3/2020'
# -
# ## Read in
# +
## Import the Data and Predictions
deaths_true = pd.read_csv("../../data/usafacts/deaths.csv")
## Add transpose step
deaths_true = deaths_true.T
deaths_true = deaths_true.rename(columns=deaths_true.iloc[0]).drop(deaths_true.index[0])
## Getting the columns we want
deaths_true[DEATH_DATE] = deaths_true[DEATH_DATE].astype('float')
deaths_true = deaths_true[["State", DEATH_DATE]].groupby("State")\
.agg( {DEATH_DATE: 'sum'}).reset_index().rename(columns={"State": "state", DEATH_DATE: "deaths"})
# predictions
deaths_pred = pd.read_csv(ENSEMBLE_FILE)
deaths_pred = deaths_pred[deaths_pred["state"] != "US"]
# IHME predictions
deaths_ihme = pd.read_csv(IHME_FILE)
filtered_cols = ["Other Counties, WA", "US","King and Snohomish Counties (excluding Life Care Center), WA",
"Life Care Center, Kirkland, WA"]
deaths_ihme = \
deaths_ihme[~deaths_ihme["location"].isin(filtered_cols)]
deaths_ihme["state"] = ["AL", "AK", "AZ", "AR", "CA", "CO", "CT", "DE", "DC","FL", "GA",
"HI", "ID", "IL", "IN", "IA", "KS", "KY", "LA", "ME", "MD",
"MA", "MI", "MN", "MS", "MO", "MT", "NE", "NV", "NH", "NJ",
"NM", "NY", "NC", "ND", "OH", "OK", "OR", "PA", "RI", "SC",
"SD", "TN", "TX", "UT", "VT", "VA", "WA", "WV", "WI", "WY"]
deaths_ihme = deaths_ihme[["state", "3_day_cumul_mean"]].rename(columns={"3_day_cumul_mean": "3_day_mean"})
# -
### PROCESS THE MODEL WE WANT HERE
deaths_pred = deaths_pred[["state", "predicted_deaths_ensemble_3"]]
# deaths_pred = deaths_pred[["state", "predicted_deaths_simple_ensemble_3"]]\
# .rename(columns={"predicted_deaths_simple_ensemble_3": "predicted_deaths_ensemble_3"})
# Dataframe of Truth and Lies (Predictions)
df = deaths_true.merge(deaths_pred, how="left", on="state")\
.merge(deaths_ihme, how="left", on="state")
# ## Losses
# +
## Let's Compute some losses
# L1 Loss
df = df.assign(l1_ours=lambda x: np.abs(x.deaths - x.predicted_deaths_ensemble_3))
df = df.assign(l1_theirs = lambda x: np.abs(x.deaths - x["3_day_mean"]))
# L2 Loss
df = df.assign(l2_ours=lambda x: (x.deaths - x.predicted_deaths_ensemble_3)**2)
df = df.assign(l2_theirs = lambda x: (x.deaths - x["3_day_mean"])**2)
# Log L1 Loss
df = df.assign(l1_log_ours=lambda x: np.log(np.abs(x.deaths - x.predicted_deaths_ensemble_3) + 1))
df = df.assign(l1_log_theirs = lambda x: np.log(np.abs(x.deaths - x["3_day_mean"]) + 1))
# Log L2 Loss
df = df.assign(l2_log_ours=lambda x: np.log((x.deaths - x.predicted_deaths_ensemble_3)**2 + 1))
df = df.assign(l2_log_theirs = lambda x: np.log((x.deaths - x["3_day_mean"])**2 + 1))
## Compute the Results
res_cols = ["l1_ours", "l1_theirs",
"l2_ours", "l2_theirs",
"l1_log_ours", "l1_log_theirs",
"l2_log_ours", "l2_log_theirs"]
res = df[res_cols]
res.agg("sum")
# -
# ## Plots for log space
# +
## Scatter Plots
# Log Transform since idk seaborn
plt_df = np.log10(df[["deaths"]] + 1)
plt_df["predicted_deaths_ensemble_3"] = np.log10(df["predicted_deaths_ensemble_3"] + 1)
plt_df["3_day_mean"] = np.log10(df["3_day_mean"] + 1)
# Our Preds -- Their Preds
f, ax = plt.subplots()
sns.scatterplot(x = "predicted_deaths_ensemble_3", y="3_day_mean", data=plt_df)
# Our Preds -- Real Data
f, ax = plt.subplots()
ax.plot([0, 4],[0,4],
ls="--", c=".3")
sns.scatterplot(x = "deaths", y="predicted_deaths_ensemble_3", data=plt_df)
ax.set(ylabel="Ensemble 3 Day Prediction",
xlabel="(Log) Number of Deaths per State on " + PROJECTION_DATE)
# Their Preds Real Data
f, ax = plt.subplots()
ax.plot([0,4],[0, 4],
ls="--", c=".3")
sns.scatterplot(x = "deaths", y="3_day_mean", data=plt_df)
ax.set(ylabel="IHME Model 3 Day Mean",
xlabel="(Log) Number of Deaths per State on " + PROJECTION_DATE)
# +
## Histograms
f, ax = plt.subplots()
sns.distplot(plt_df['3_day_mean'], kde=False, rug=True, bins = 15);
ax.set(ylabel="IHME Counts",
xlabel="(Log) Number of Deaths per State on " + PROJECTION_DATE)
ax.set(xlim=(0, 3.5), ylim=(0,12))
f, ax = plt.subplots()
sns.distplot(plt_df['predicted_deaths_ensemble_3'], kde=False, rug=True, bins = 15);
ax.set(ylabel="Ensemble Counts",
xlabel="(Log) Number of Deaths per State on " + PROJECTION_DATE)
ax.set(xlim=(0, 3.5), ylim=(0,12))
f, ax = plt.subplots()
sns.distplot(plt_df['predicted_deaths_ensemble_3'], kde=False, rug=True, bins = 15);
ax.set(ylabel="Ensemble Counts",
xlabel="(Log) Number of Deaths per State on " + PROJECTION_DATE)
ax.set(xlim=(0, 3.5), ylim=(0,12))
sns.distplot(plt_df['3_day_mean'], kde=False, rug=True, bins = 15);
ax.set(ylabel="Counts",
xlabel="(Log) Number of Deaths per State on " + PROJECTION_DATE)
f.legend(loc='center left', labels=['ensemble','ihme'], bbox_to_anchor=(0.65, 0.8), ncol=1)
# -
# ## Plots for non-log
# +
## Scatter Plots
# plt_df = np.log10(df[["deaths"]] + 1)
# plt_df["predicted_deaths_ensemble_3"] = np.log10(df["predicted_deaths_ensemble_3"] + 1)
# plt_df["3_day_mean"] = np.log10(df["3_day_mean"] + 1)
## Scatter Plots
# Log Transform since idk seaborn
plt_df = df
# Our Preds -- Their Preds
f, ax = plt.subplots()
sns.scatterplot(x = "predicted_deaths_ensemble_3", y="3_day_mean", data=plt_df)
# Our Preds -- Real Data
f, ax = plt.subplots()
ax.plot([0, 3000],[0,3000],
ls="--", c=".3")
sns.scatterplot(x = "deaths", y="predicted_deaths_ensemble_3", data=plt_df)
ax.set(ylabel="Ensemble 3 Day Prediction",
xlabel="Number of Deaths per State on " + PROJECTION_DATE)
# Their Preds Real Data
f, ax = plt.subplots()
ax.plot([0,3000],[0, 3000],
ls="--", c=".3")
sns.scatterplot(x = "deaths", y="3_day_mean", data=plt_df)
ax.set(ylabel="IHME Model 3 Day Mean",
xlabel="Number of Deaths per State on " + PROJECTION_DATE)
| predictions/other_modeling/Model Comparisons.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # DeepPurpose Deep Dive
# ## Tutorial 2: Training a Drug Property Prediction Model from Scratch for Assay Data
# #### [@KexinHuang5](https://twitter.com/KexinHuang5)
#
# In this tutorial, we further extends the use cases of DeepPurpose to assay data where there are only drug information and its affinity score to the protein in the assay.
#
# Agenda:
#
# - Part I: Introduction to Assay Data
# - Part II: Drug Property Prediction
#
# Let's start!
from DeepPurpose import utils, models, dataset, property_pred
import warnings
warnings.filterwarnings("ignore")
# ## Part I: Introduction to Assay Data
#
# Drug-target interaction measures the binding of drug molecules to the protein targets. In drug discovery process, we usually already have a protein of interest and traditionally, high-throughput screening assay is performed on a potential drug library to get affinity scores. But it is expensive and time-consuming, and it cannot go through a huge amount of molecule candidates, which means it potentially can miss a drug candidate. Machine learning can provide an important role to facilitate this process. One way to do it is to narrow down the search space for HTS. Here are the steps:
#
# 1. For a target protein of interest, conduct an initial high-throughput screening assay on a set of drugs (ideally, structurally diverse), where the set size depends on the time and cost constraint.
#
# 2. Train a deep learning model using DeepPurpose and select a threshold to control the false positive rate (e.g., < 0.02).
#
# 3. Apply the trained model on a large set of drugs. Select the set of drugs that meet the threshold and send to HTS.
#
# Through the above three steps, it greatly cuts down the time of screening and also reduces the likelihood of missing a potential drug candidate. Now, we show the step 2: using DeepPurpose to train a deep learning model for screening the assay.
#
# (**Data**) DeepPurpose takes into an array of drug's SMILES strings (**d**) and an array of label (**y**), which can either be binary 0/1 indicating interaction outcome or a real number indicating affinity value. Note **y**\[0\] is the score for **d**\[0\].
#
# Besides transforming into numpy arrays through some data wrangling on your own, DeepPurpose also provides two ways to help data preparation.
#
# The first way is to read from local files. For example, to load drug assay data, we expect a file.txt where each line is a drug SMILES string, followed by an affinity score or 0/1 label:
#
# ```CC1=C...C4)N 7.365```
#
# Then, we use ```dataset.read_file_training_dataset_bioassay``` to load it.
X_drugs, X_targets, y = dataset.read_file_training_dataset_bioassay('./toy_data/AID1706.txt')
print('Drug 1: ' + X_drugs[0])
print('Score 1: ' + str(y[0]))
# DeepPurpose also provides data loaders to ease preprocessing. For example, in this tutorial, we will use the AID1706 screening data for SARS-CoV 3CL Protease. We can use ```dataset.load_AID1706_SARS_CoV_3CL```. It will download, preprocess to the designated data format. It supports label log-scale transformation for easier regression and also allows label binarization given a customized threshold. In this case, we use the binary label.
X_drugs, X_targets, y = dataset.load_AID1706_SARS_CoV_3CL(path = './data', binary = True, threshold = 15, balanced = True)
print('Drug 1: ' + X_drugs[0])
print('Score 1: ' + str(y[0]))
# For more detailed examples and tutorials of data loading, checkout this [tutorial](./DEMO/load_data_tutorial.ipynb).
#
# ## Part II: Drug Property Prediction Framework
#
# DeepPurpose provides a simple framework to do drug property prediction research using 8 encoders for drugs. It basically consists of the following steps, where each step corresponds to one line of code:
#
# - Encoder specification
# - Data encoding and split
# - Model configuration generation
# - Model initialization
# - Model Training
# - Model Prediction and Repuposing/Screening
# - Model Saving and Loading
#
# Let's start with data encoding!
#
# (**Encoder specification**) After we obtain the required data format from Part I, we need to prepare them for the encoders. Hence, we first specify the encoder to use for drug and protein. Here we try MPNN for drug.
#
# If you find MPNN is too large for the CPUs, you can try smaller encoders by uncommenting the last line:
drug_encoding = 'MPNN'
#drug_encoding = 'Morgan'
# Note that you can switch encoder just by changing the encoding name above. The full list of encoders are listed [here](https://github.com/kexinhuang12345/DeepPurpose#encodings).
#
# (**Data encoding and split**) Now, we encode the data into the specified format, using ```utils.data_process``` function. It specifies train/validation/test split fractions, and random seed to ensure same data splits for reproducibility. This function also support data splitting methods such as ```cold_drug```, which splits on drug for model robustness evaluation to test on unseen drug/proteins.
#
# The function outputs train, val, test pandas dataframes.
train, val, test = utils.data_process(X_drug = X_drugs, y = y, drug_encoding = drug_encoding,
split_method='random',frac=[0.7,0.1,0.2],
random_seed = 1)
train.head(1)
# (**Model configuration generation**) Now, we initialize a model with its configuration. You can modify almost any hyper-parameters (e.g., learning rate, epoch, batch size), model parameters (e.g. hidden dimensions, filter size) and etc in this function. The supported configurations are listed here in this [link](https://github.com/kexinhuang12345/DeepPurpose/blob/e169e2f550694145077bb2af95a4031abe400a77/DeepPurpose/utils.py#L486).
#
# For the sake of example, we specify the epoch size to be 3, and set the model parameters to be small so that you can run on both CPUs & GPUs quickly and can proceed to the next steps. For a reference parameters, checkout the notebooks in the DEMO folder.
config = utils.generate_config(drug_encoding = drug_encoding,
cls_hidden_dims = [1024,1024,512],
train_epoch = 5,
LR = 0.001,
batch_size = 128,
hidden_dim_drug = 128,
mpnn_hidden_size = 128,
mpnn_depth = 3
)
# (**Model initialization**) Next, we initialize a model using the above configuration.
model = property_pred.model_initialize(**config)
model
# (**Model Training**) Next, it is ready to train, using the ```model.train``` function! If you do not have test set, you can just use ```model.train(train, val)```.
model.train(train, val, test)
# We see that the model will automatically generate and plot the training process, along with the validation result and test result.
#
# (**Model Prediction and Repuposing/Screening**) Next, we see how we can predict affinity scores on new data. Suppose the new data is a new drug below.
X_drug = ['CC1=C2C=C(C=CC2=NN1)C3=CC(=CN=C3)OCC(CC4=CC=CC=C4)N']
X_pred = utils.data_process(X_drug = X_drug, y = y, drug_encoding = drug_encoding, split_method='no_split')
y_pred = model.predict(X_pred)
print('The predicted score is ' + str(y_pred))
# We can also do repurposing/screening using the trained model. Basically, for repurposing/screening a set of new drugs (**r**), we run the above prediction function. We wrap the operation into a ```models.repurpose```.
#
# For example, suppose we want to do repurposing from a set of antiviral drugs for the COVID-19 target 3CL protease. The corresponding data can be retrieved using ```dataset``` functions.
#
r, r_name, r_pubchem_cid = dataset.load_antiviral_drugs()
print('Repurposing Drug 1 Name: ' + r_name[0])
print('Repurposing Drug 1 SMILES: ' + r[0])
print('Repurposing Drug 1 Pubchem CID: ' + str(r_pubchem_cid[0]))
# Now, we can call the ```repurpose``` function. After feeding the necessary inputs, it will print a list of repurposed drugs ranked on its affinity to the target protein. The ```convert_y``` parameter should be set to be ```False``` when the ranking is ascending (i.e. lower value -> higher affinity) due to the log transformation, vice versus.
y_pred = property_pred.repurpose(X_repurpose = r, model = model, drug_names = r_name)
# Saving and loading models are also really easy. The loading function also automatically detects if the model is trained on multiple GPUs. To save a model:
model.save_model('./tutorial_model')
# To load a saved/pretrained model:
model = property_pred.model_pretrained(path_dir = './tutorial_model')
model
# We also provided more functionalities for DTI research purposes.
#
# For example, this [demo](https://github.com/kexinhuang12345/DeepPurpose/blob/master/DEMO/Drug_Property_Pred-Ax-Hyperparam-Tune.ipynb) shows how to use Ax platform to do some latest hyperparameter tuning methods such as Bayesian Optimization on DeepPurpose.
#
# They are described in details in tutorial 1 and in the github repository.
# That wraps up our tutorials on the main functionalities of DeepPurpose's Drug Property Prediction framework!
#
# Do checkout the previous & upcoming tutorials:
#
# Tutorial 1: Training a Drug-Target Interaction Model from Scratch
#
# Tutorial 3: Repurposing and Virtual Screening Using One Line of Code
#
# **Star & watch & contribute to DeepPurpose's [github repository](https://github.com/kexinhuang12345/DeepPurpose)!**
#
# Feedbacks would also be appreciated and you can send me an email (<EMAIL>)!
| Tutorial_2_Drug_Property_Pred_Assay_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="8QxfmwOzapfE"
# Convolutional Neural Networks (CNN) are used for classifying images and Computer Vision.
#
# To implement Neural Networks, we will use Tensorflow library.
#
# + [markdown] id="whrNq1W5a5zF"
# ## Description of the dataset
#
# **Build a CNN to recognize whether the input image is cat or dog**
#
# 1) Single Prediction - cat and dog image
#
# 2) Training - 4000 images of cat and 4000 images of dog
#
# 3) Test set - 1000 images of cat and 1000 images of dog
#
# Total 10,000 images
# + [markdown] id="p2yF0s-_gQL5"
# **As the dataset is very big, we will not implement all on Google Collab**
# + id="dNf7JO1LgvYC"
import numpy as np
import tensorflow as tf
from keras.preprocessing.image import ImageDataGenerator
# + colab={"base_uri": "https://localhost:8080/"} id="vwosvbwHARN7" outputId="9417c502-63d8-44e2-d0ee-82326cd124b2"
# from google.colab import drive
# drive.mount('/content/drive/')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="bWK3iHdugwDC" outputId="946503c3-5868-435f-f63d-5120fa00918e"
tf.__version__
# + [markdown] id="RkHiKj8zgevV"
# ## Part 1 - Data Preprocessing
# + [markdown] id="Rf1z-tRwhM7-"
# ### Preprocessing the Training Set
# + id="FxGnTZmFaqPq"
## Apply transformations on the images of the Training Set - to avoid overfitting
# these transformations are geometrical like zoom in/out, rotate, flips - image augmentation
# in this way our CNN wont be over trained on these images
# Keras is a deep learning library which is integrated into Tensorflow
# + colab={"base_uri": "https://localhost:8080/"} id="I7qPytmmhp4_" outputId="81235975-90fb-442f-e6a2-142e1894b56d"
# Create train_datagen object
train_datagen = ImageDataGenerator(rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
# rescale - feature scaling applied on the pixels (absolutetly necessary for neural networks)
# Import the training dataset
training_set = train_datagen.flow_from_directory('image_dataset/training_set', # change path of train set
target_size = (64, 64), # final size of image to be fed into CNN
batch_size = 32, # how many images in a batch to train at a time
class_mode = 'binary') # binary or categorical (multiclass) classification
# + [markdown] id="GUV7t6i2IEaG"
# ### Preprocessing the Training Set
# + colab={"base_uri": "https://localhost:8080/"} id="xORNGKFUB1Ad" outputId="e054f459-1df0-4924-d00e-1d49a8c2c3cc"
# New images - when deploying our model in production
# we have to the images of the test set intact --> so NO TRANSFORMATION!
test_datagen = ImageDataGenerator(rescale = 1./255) # test set must also be feature scaled
test_set = test_datagen.flow_from_directory('image_dataset/test_set', # change path of test set
target_size = (64, 64),
batch_size = 32,
class_mode = 'binary')
# + [markdown] id="NSnjV9FJIc6Y"
# ## Part 2 - Building the CNN
# + [markdown] id="KrSrzGMYIkXD"
# ### Initialising the CNN
# + id="OXgJNqhZIHnb"
# Sequential library is a part of Keras library. Keras library is integrated into Tenserflow
cnn = tf.keras.models.Sequential() # intialize the network, cnn object
# + [markdown] id="GIe5AxeotdyN"
# ### Step 1 - Convolution
# + id="5buqX4qOIHvE"
# Create the fully connected layer object.
# using Convolutional class from Keras (Tensorflow) - tf.keras.Conv2D
# filters --> no. of feature detectors to apply to our image
# kernel_size --> size of the filter (rows and cols) e.g. size = 3 -> means 3x3
# rectifier function - 'relu' (activation function)
cnn.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, activation='relu', input_shape=[64, 64, 3]))
# input_shape --> specifiy shape of inputs images (1st time)
# coloured image --> input_shape=[X, X, 3]
# B/W image --> input_shape=[X, X, 1]
### 1) Feature Map Created
# + [markdown] id="363EGfhrtjrS"
# ### Step 2 - Pooling (Max Pooling)
# + id="OB7JAYQ2IH1S"
# Apply MaxPool from Keras
# pool_size --> 2 (which means 2x2)
# strides -> shift pooling frame by 2 steps (2x2 pixels)
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
### 2) Pooled Feature Map Created
# + [markdown] id="D7kHFXl5tlIU"
# ### Adding a second convolutional layer
# + id="ugKoNBAHIH4I"
# Convolutional + Max Pooling for another layer
# remove input_shape parameter (it exists only in 1st layer)
cnn.add(tf.keras.layers.Conv2D(filters=32, kernel_size=3, activation='relu'))
cnn.add(tf.keras.layers.MaxPool2D(pool_size=2, strides=2))
# we can add more layers. More layers is better, but it increases computational costs
# also shape of layers must be same
# + [markdown] id="_OEaPMkEtos4"
# ### Step 3 - Flattening
# + id="XmRSWMT3IH6c"
# Create the Flattening layers (1D layers) by Flattern class from Keras
cnn.add(tf.keras.layers.Flatten())
# + [markdown] id="7VzD0a_EtpZc"
# ### Step 4 - Full Connection
# + id="w72c4ioHIIB_"
# Now create a full connected layer by Dense class
# units --> no. of hidden neurons (a larger number is preferred for image classfication)
# activation function --> rectifier function
cnn.add(tf.keras.layers.Dense(units=128, activation='relu'))
# + [markdown] id="9PjTFnF5ttbx"
# ### Step 5 - Output Layer
# + id="h-Nr-9jZtqF6"
# change units and activation function
# Binary classification - 0 or 1, so dimension of neuron is 1 (1 output neuron)
cnn.add(tf.keras.layers.Dense(units=1, activation='sigmoid')) # sigmoid activation function (gives probability)
# for multiclass classification, output layer dimensions will depend on no.of classes
# for binary classification - activation function is 'sigmoid'
# for multiclass classification - activation function is 'softmax'
# + [markdown] id="FlUqKeSvt4gZ"
# ## Part 3 - Training the CNN
# + [markdown] id="aqgKGajj6Ruj"
# ### Compiling the CNN
# + id="dqa2QMfjtqIk"
# using optimizer and loss function, with an evaluation metric
# best optimizer - stochastic gradient descent (adam optimizer) [reduces loss error]
# loss function, i.e. Cost function J theta - difference between prediction and real result
# for binary classification - loss function is 'binary_crossentropy'
# for multiclass classification - loss function is 'categorical_crossentropy'
cnn.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# + [markdown] id="utcUVOXK6epv"
# ### Training the CNN on the Training set and evaluating it on the Test set
# + id="BUv6L3emtqNP"
# Train and Test at the same time
# same fit method, but neural networks will have extra hyperparameters
# no. of epocs --> no. of iterations the neural network is trained (improves accuracy over time) (make it small as we have 10,000 images)
cnn.fit(x = training_set, validation_data = test_set, epochs = 25)
# -
# Therefore,
#
# **Training Accuracy** = 0.8864
#
# **Test Accuracy** = 0.8000
# + [markdown] id="U9is5KtZ6lJs"
# ## Part 4 - Making a single prediction
#
# + id="GbaBVM98tqPZ"
from keras.preprocessing import image
# load the images - give path of image
# image size must be the same as the one used in training the model
test_image = image.load_img('image_dataset/single_prediction/cat_or_dog_1.jpg', target_size = (64, 64))
# Conver the images from pixels to numpy array
test_image = image.img_to_array(test_image)
# CNN was not trained on single image, but on batch of images. This must also be specified for predict method
# Place the image in a batch (extra dimension)
test_image = np.expand_dims(test_image, axis = 0) # axis = 0 --> 1st dimension
result = cnn.predict(test_image)
# The way to figure out 0 or 1, is to call the class indices attritbute from our training set object
training_set.class_indices
# result also has batch dimension - result[0][0] - access batch, then single element within the batch
if result[0][0] == 1:
prediction = 'dog'
else:
prediction = 'cat'
# + id="AIpeCBIVtqRy"
print(prediction)
| Supervised Learning/Neural Networks/Convolutional Neural Networks (CNN)/Convolutional_Neural_Networks_(CNN).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib
matplotlib.rcParams["figure.figsize"] = (20,10)
df_test=pd.read_csv('Test.csv')
df_train=pd.read_csv('Train.csv')
df_train.head()
df_train.shape
df_train.groupby('area_type')['area_type'].agg('count')
df_2=df_train.drop(['area_type','availability','balcony','society'],axis='columns')
df_2.head()
b
df_2.isnull().sum()
df3=df_2.dropna()
df3.head()
df3.shape
df3['size'].unique()
df3['bhk']=df3['size'].apply(lambda x: int(x.split(' ')[0]))
df3.head()
df3['bhk'].unique()
df3.total_sqft.unique()
def is_float(x):
try:
float(x)
except:
return False
return True
is_float('1145 - 1340')
df3[~df3['total_sqft'].apply(is_float)].head(25)
def convert_sq_to_num(x):
tokens=x.split('-')
if len(tokens)== 2:
return(float(tokens[0])+float(tokens[1]))/2
try:
return float(x)
except:
return None
convert_sq_to_num('716Sq. Meter ')
df4=df3.copy()
df4['total_sqft']=df4['total_sqft'].apply(convert_sq_to_num)
df4.head()
df4.loc[1400]
df4.loc[410]
df4['total_sqft'].unique()
df4.shape
df5=df4.dropna()
df5.shape
# +
df5['price_per_sq']=df5['price']*100000/df5['total_sqft']
# -
df5.head()
len(df5.location.unique())
df5.location=df5.location.apply(lambda x:x.strip())
location_stats=df5.groupby('location')['location'].agg('count').sort_values(ascending=False)
location_stats
len(location_stats[location_stats<10])
location_stats_less_10=location_stats[location_stats<10]
location_stats_less_10
len(df5.location.unique())
df5.location=df5.location.apply(lambda x:'other' if x in location_stats_less_10 else x)
len(df5.location.unique())
df5.head(10)
df5[df5.total_sqft/df5.bhk<300].head()
df5.shape
df6=df5[~(df5.total_sqft/df5.bhk<300)]
df6.shape
df6.price_per_sq.describe()
def remove_pps_outliers(df):
df_out = pd.DataFrame()
for key, subdf in df.groupby('location'):
m = np.mean(subdf.price_per_sq)
st = np.std(subdf.price_per_sq)
reduced_df = subdf[(subdf.price_per_sq>(m-st)) & (subdf.price_per_sq<=(m+st))]
df_out = pd.concat([df_out,reduced_df],ignore_index=True)
return df_out
df7 = remove_pps_outliers(df6)
df7.shape
# +
def plot_scatter_chart(df,location):
bhk2 = df[(df.location==location) & (df.bhk==2)]
bhk3 = df[(df.location==location) & (df.bhk==3)]
matplotlib.rcParams['figure.figsize'] = (15,10)
plt.scatter(bhk2.total_sqft,bhk2.price,color='blue',label='2 BHK', s=50)
plt.scatter(bhk3.total_sqft,bhk3.price,marker='+', color='green',label='3 BHK', s=50)
plt.xlabel("Total Square Feet Area")
plt.ylabel("Price (Lakh Indian Rupees)")
plt.title(location)
plt.legend()
plot_scatter_chart(df7,"Rajaji Nagar")
# +
plot_scatter_chart(df7,"Hebbal")
# -
def remove_bhk_outliers(df):
exclude_indices = np.array([])
for location, location_df in df.groupby('location'):
bhk_stats = {}
for bhk, bhk_df in location_df.groupby('bhk'):
bhk_stats[bhk] = {
'mean': np.mean(bhk_df.price_per_sq),
'std': np.std(bhk_df.price_per_sq),
'count': bhk_df.shape[0]
}
for bhk, bhk_df in location_df.groupby('bhk'):
stats = bhk_stats.get(bhk-1)
if stats and stats['count']>5:
exclude_indices = np.append(exclude_indices, bhk_df[bhk_df.price_per_sq<(stats['mean'])].index.values)
return df.drop(exclude_indices,axis='index')
df8 = remove_bhk_outliers(df7)
# df8 = df7.copy()
df8.shape
plot_scatter_chart(df8,"Rajaji Nagar")
# +
plot_scatter_chart(df8,"Hebbal")
# -
import matplotlib
matplotlib.rcParams["figure.figsize"] = (20,10)
plt.hist(df8.price_per_sq,rwidth=0.8)
plt.xlabel("Price Per Square Feet")
plt.ylabel("Count")
# +
df8.bath.unique()
# +
plt.hist(df8.bath,rwidth=0.8)
plt.xlabel("Number of bathrooms")
plt.ylabel("Count")
# +
df8[df8.bath>10]
# +
df8[df8.bath>df8.bhk+2]
# -
df9 = df8[df8.bath<df8.bhk+2]
df9.shape
# +
df10 = df9.drop(['size','price_per_sq'],axis='columns')
df10.head(3)
# -
dummies = pd.get_dummies(df10.location)
dummies.head(3)
df11 = pd.concat([df10,dummies.drop('other',axis='columns')],axis='columns')
df11.head()
df12 = df11.drop('location',axis='columns')
df12.head(2)
# +
df12.shape
# -
X = df12.drop(['price'],axis='columns')
X.head(3)
X.shape
# +
y = df12.price
y.head(3)
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,y,test_size=0.2,random_state=10)
from sklearn.linear_model import LinearRegression
lr_clf = LinearRegression()
lr_clf.fit(X_train,y_train)
lr_clf.score(X_test,y_test)
# +
from sklearn.model_selection import ShuffleSplit
from sklearn.model_selection import cross_val_score
cv = ShuffleSplit(n_splits=5, test_size=0.2, random_state=0)
cross_val_score(LinearRegression(), X, y, cv=cv)
# +
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Lasso
from sklearn.tree import DecisionTreeRegressor
def find_best_model_using_gridsearchcv(X,y):
algos = {
'linear_regression' : {
'model': LinearRegression(),
'params': {
'normalize': [True, False]
}
},
'lasso': {
'model': Lasso(),
'params': {
'alpha': [1,2],
'selection': ['random', 'cyclic']
}
},
'decision_tree': {
'model': DecisionTreeRegressor(),
'params': {
'criterion' : ['mse','friedman_mse'],
'splitter': ['best','random']
}
}
}
scores = []
cv = ShuffleSplit(n_splits=5, test_size=0.2, random_state=0)
for algo_name, config in algos.items():
gs = GridSearchCV(config['model'], config['params'], cv=cv, return_train_score=False)
gs.fit(X,y)
scores.append({
'model': algo_name,
'best_score': gs.best_score_,
'best_params': gs.best_params_
})
return pd.DataFrame(scores,columns=['model','best_score','best_params'])
find_best_model_using_gridsearchcv(X,y)
# -
def predict_price(location,sqft,bath,bhk):
loc_index = np.where(X.columns==location)[0][0]
x = np.zeros(len(X.columns))
x[0] = sqft
x[1] = bath
x[2] = bhk
if loc_index >= 0:
x[loc_index] = 1
return lr_clf.predict([x])[0]
# +
predict_price('1st Phase JP Nagar',4000, 3, 2)
# -
predict_price('1st Phase JP Nagar',1000, 3, 3)
predict_price('Indira Nagar',1000, 3, 3)
import pickle
with open('banglore_home_prices_model.pickle','wb') as f:
pickle.dump(lr_clf,f)
import json
columns = {
'data_columns' : [col.lower() for col in X.columns]
}
with open("columns.json","w") as f:
f.write(json.dumps(columns))
| House price prediction in Banglore/House_price prediction .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=false editable=false
# Initialize OK
from client.api.notebook import Notebook
ok = Notebook('hw10.ok')
# -
# # Homework 10: Linear Regression
# **Reading**:
# * [Linear Regression](https://www.inferentialthinking.com/chapters/15/2/Regression_Line.html)
# * [Method of Least Squares](https://www.inferentialthinking.com/chapters/15/3/Method_of_Least_Squares.html)
# * [Least Squares Regression](https://www.inferentialthinking.com/chapters/15/4/Least_Squares_Regression.html)
# Please complete this notebook by filling in the cells provided. Before you begin, execute the following cell to load the provided tests. Each time you start your server, you will need to execute this cell again to load the tests.
#
# Homework 10 is due **Thursday, 4/16 at 11:59pm**. You will receive an early submission bonus point if you turn in your final submission by Wednesday, 4/15 at 11:59pm. Start early so that you can come to office hours if you're stuck. Check the website for the office hours schedule. Late work will not be accepted as per the [policies](http://data8.org/sp20/policies.html) of this course.
#
# Directly sharing answers is not okay, but discussing problems with the course staff or with other students is encouraged. Refer to the policies page to learn more about how to learn cooperatively.
#
# For all problems that you must write our explanations and sentences for, you **must** provide your answer in the designated space. Moreover, throughout this homework and all future ones, please be sure to not re-assign variables throughout the notebook! For example, if you use `max_temperature` in your answer to one question, do not reassign it later on.
# +
# Don't change this cell; just run it.
import numpy as np
from datascience import *
# These lines do some fancy plotting magic.
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
import warnings
warnings.simplefilter('ignore', FutureWarning)
from client.api.notebook import Notebook
ok = Notebook('hw10.ok')
# -
# ## Exploring the PTEN Gene with Linear Regression
# ## 1. PTEN Linear Regression
# This week's homework is about linear regression. The dataset we'll be using is from the Cancer Cell Line Encyclopedia -- you can read more about this database in this [paper](https://www.nature.com/articles/s41586-019-1186-3) and interact with the data yourself at the online portal [here](https://portals.broadinstitute.org/ccle).
#
# The specific dataset we'll be taking a look at is expression data for the PTEN gene in around 1000 cell lines. The PTEN gene is a tumor-suppressing gene, and mutations in the PTEN gene are associated with many types of cancer. A cell line is group of cells that are kept alive and replicate indefinitely in culture (grown in petri dishes, for example).
#
# Run the following cell to load the `pten` table. The `pten` table has four columns, a column for the specific `Cell Line`, a column for the `Copy Number`, which is how many times a copy of a portion of the PTEN gene is found in the DNA of that cell line, `mRNA Expression (Affy)`, and `mRNA Expression (RNAseq)`.
# Just run this cell
pten = Table().read_table("pten.csv")
pten.show(5)
# Just run this cell
pten.hist("Copy Number", bins = np.arange(-1, 1.5, 0.5))
# + [markdown] deletable=false editable=false
# **Question 1**
#
# Looking at the histogram above, we want to check whether or not `Copy Number` is in standard units. For this question, compute the mean and the standard deviation of the values in `Copy Number` and assign these values to `copy_number_mean` and `copy_number_sd` respectively. After you calculate these values, assign `is_su` to either `True` if you think that `Copy Numbers` is in standard units or `False` if you think otherwise.
#
# <!--
# BEGIN QUESTION
# name: q1_1
# manual: false
# -->
# -
copy_number = pten.column("Copy Number")
copy_number_mean = ...
copy_number_sd = ...
is_su = ...
print(f"Mean: {copy_number_mean}, SD: {copy_number_sd}, Is in standard units?: {is_su}")
# + deletable=false editable=false
ok.grade("q1_1");
# + [markdown] deletable=false editable=false
# **Question 2**
#
# Create the function `standard_units` so that it converts the values in the array `arr` to standard units. We'll then use `standard_units` to create a new table, `pten_su`, that converts all the values in the table `pten` to standard units.
#
# <!--
# BEGIN QUESTION
# name: q1_2
# manual: false
# -->
# +
def standard_units(arr):
...
# DON'T DELETE OR MODIFY ANY OF THE LINES OF CODE BELOW IN THIS CELL
pten_su = Table().with_columns("Cell Line", pten.column("Cell Line"),
"Copy Number SU", standard_units(pten.column("Copy Number")),
"mRNA Expression (Affy) SU", standard_units(pten.column("mRNA Expression (Affy)")),
"mRNA Expression (RNAseq) SU", standard_units(pten.column("mRNA Expression (RNAseq)"))
)
pten_su.show(5)
# + deletable=false editable=false
ok.grade("q1_2");
# -
# You should always visually inspect your data, before numerically analyzing any relationships in your dataset. Run the following cell in order to look at the relationship between the variables in our dataset.
# Just run this cell
pten_su.scatter("Copy Number SU", "mRNA Expression (Affy) SU")
pten_su.scatter("Copy Number SU", "mRNA Expression (RNAseq) SU")
pten_su.scatter("mRNA Expression (Affy) SU", "mRNA Expression (RNAseq) SU")
# + [markdown] deletable=false editable=false
# **Question 3**
#
# Which of the following relationships do you think has the highest correlation (i.e. highest absolute value of `r`)? Assign `highest_correlation` to the number corresponding to the relationship you think has the highest correlation.
#
# 1. Copy Number vs. mRNA Expression (Affy)
# 2. Copy Number vs. mRNA Expression (RNAseq)
# 3. mRNA Expression (Affy) vs. mRNA Expression (RNAseq)
#
# <!--
# BEGIN QUESTION
# name: q1_3
# manual: false
# -->
# -
highest_correlation = ...
# + deletable=false editable=false
ok.grade("q1_3");
# + [markdown] deletable=false editable=false
# **Question 4**
#
# Now, using the `standard units` function, define the function `correlation` which computes the correlation between `arr1` and `arr2`.
#
# <!--
# BEGIN QUESTION
# name: q1_4
# manual: false
# -->
# +
def correlation(arr1, arr2):
...
# This computes the correlation between the different variables in pten
copy_affy = correlation(pten.column("Copy Number"), pten.column("mRNA Expression (Affy)"))
copy_rnaseq = correlation(pten.column("Copy Number"), pten.column("mRNA Expression (RNAseq)"))
affy_rnaseq = correlation(pten.column("mRNA Expression (Affy)"), pten.column("mRNA Expression (RNAseq)"))
print(f" \
Copy Number vs. mRNA Expression (Affy) Correlation: {copy_affy}, \n \
Copy Number vs. mRNA Expression (RNAseq) Correlation: {copy_rnaseq}, \n \
mRNA Expression (Affy) vs. mRNA Expression (RNAseq) Correlation: {affy_rnaseq}")
# + deletable=false editable=false
ok.grade("q1_4");
# + [markdown] deletable=false editable=false
# **Question 5**
#
# If we switch what we input as arguments to `correlation`, i.e. found the correlation between `mRNA Expression (Affy)` vs. `Copy Number` instead of the other way around, would the correlation change? Assign `correlation_change` to either `True` if you think yes, or `False` if you think no.
#
# <!--
# BEGIN QUESTION
# name: q1_5
# manual: false
# -->
# -
correlation_change = ...
# + deletable=false editable=false
ok.grade("q1_5");
# + [markdown] deletable=false editable=false
# **Question 6**
#
# Looking at both the scatter plots after Question 2 and the correlations computed in Question 4, describe a pattern you see in the relationships between the variables.
#
# <!--
# BEGIN QUESTION
# name: q1_6
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] export_pdf=true
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# **Question 7**
#
# Let's look at the relationship between mRNA Expression (Affy) vs. mRNA Expression (RNAseq) only. Define a function called `regression_parameters` that returns the parameters of the regression line as a two-item array containing the slope and intercept of the regression line as the first and second elements respectively. The function `regression_line` takes in two arguments, an array of `x` values, and an array of `y` values.
#
# <!--
# BEGIN QUESTION
# name: q1_7
# manual: false
# -->
# +
def regression_parameters(x, y):
...
slope = ...
intercept = ...
return make_array(slope, intercept)
parameters = regression_parameters(pten.column("mRNA Expression (Affy)"), pten.column("mRNA Expression (RNAseq)"))
parameters
# + deletable=false editable=false
ok.grade("q1_7");
# + [markdown] deletable=false editable=false
# **Question 8**
#
# If we switch what we input as arguments to `regression_parameters`, i.e. found the parameters for the regression line for `mRNA Expression (RNAseq)` vs. `mRNA Expression (Affy)` instead of the other way around, would the regression parameters change (would the slope and/or intercept change)? Assign `parameters_change` to either `True` if you think yes, or `False` if you think no.
#
# <!--
# BEGIN QUESTION
# name: q1_8
# manual: false
# -->
# -
parameters_change = ...
# + deletable=false editable=false
ok.grade("q1_8");
# + [markdown] deletable=false editable=false
# **Question 9**
#
# Now, let's look at how the regression parameters look like in standard units. Use the table `pten_su` and the function `regression_parameters`, and assign `parameters_su` to a two-item array containing the slope and the intercept of the regression line for mRNA Expression (Affy) in standard units vs. mRNA Expression (RNAseq) in standard units.
#
#
# <!--
# BEGIN QUESTION
# name: q1_9
# manual: false
# -->
# -
parameters_su = ...
parameters_su
# + deletable=false editable=false
ok.grade("q1_9");
# + [markdown] deletable=false editable=false
# **Question 10**
#
# Looking at the array `parameters_su`, what do you notice about the slope and intercept values specifically? Relate them to another value we already calculated in a previous question, as well as relate them to an equation.
#
#
# <!--
# BEGIN QUESTION
# name: q1_10
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] export_pdf=true
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# **Question 11**
#
# The oldest and most commonly used cell line in Biology is the HeLa cell line, named after <NAME>, whose cervical cancer cells were taken without her consent in 1951 to create this cell line. The issue of data privacy and consent is very important to data science! You can read more about this topic [here](https://www.hopkinsmedicine.org/henriettalacks/).
#
# The HeLa cell line is missing from our dataset. If we know that the HeLa mRNA Expression (Affy) value is 8.2, what is the predicted mRNA Expression (RNAseq) value? Use the values in `parameters` that we derived in Question 1.7, and assign the result to `hela_rnaseq`.
#
# <!--
# BEGIN QUESTION
# name: q1_11
# manual: false
# -->
# -
hela_rnaseq = ...
hela_rnaseq
# + deletable=false editable=false
ok.grade("q1_11");
# + [markdown] deletable=false editable=false
# **Question 12**
#
# Compute the predicted mRNA Expression (RNAseq) values from the mRNA Expression (Affy) values in the `pten` table. Use the values in the `parameters` array from Question 1.7, and assign the result to `predicted_rnaseq`. We'll plot your computed regression line with the scatter plot from after question 1.2 of mRNA Expression (Affy) vs. mRNA Expression (RNAseq).
#
# <!--
# BEGIN QUESTION
# name: q1_12
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + export_pdf=true
predicted_rnaseq = ...
# DON'T CHANGE/DELETE ANY OF THE BELOW CODE IN THIS CELL
(pten.with_column("Predicted mRNA Expression (RNAseq)", predicted_rnaseq)
.select("mRNA Expression (Affy)", "mRNA Expression (RNAseq)", "Predicted mRNA Expression (RNAseq)")
.scatter("mRNA Expression (Affy)"))
plt.ylabel("mRNA Expression (RNAseq)");
# -
# ## Fitting a least-squares regression line
# Recall that the least-square regression line is the unique straight line that minimizes root mean squared error (RMSE) among all possible fit lines. Using this property, we can find the equation of the regression line by finding the pair of slope and intercept values that minimize root mean squared error.
# + [markdown] deletable=false editable=false
# **Question 13**
#
# Define a function called `RMSE`. It should take two arguments:
#
# 1. the slope of a line (a number)
# 2. the intercept of a line (a number).
#
# It should return a number that is the root mean squared error (RMSE) for a line defined with the arguments slope and intercept used to predict mRNA Expression (RNAseq) values from mRNA Expression (Affy) values for each row in the `pten` table.
#
# *Hint: Errors are defined as the difference between the actual `y` values and the predicted `y` values.*
#
# *Note: if you need a refresher on RMSE, here's the [link](https://www.inferentialthinking.com/chapters/15/3/Method_of_Least_Squares.html#Root-Mean-Squared-Error) from the textbook*
#
# <!--
# BEGIN QUESTION
# name: q1_13
# manual: false
# -->
# +
def RMSE(slope, intercept):
affy = pten.column("mRNA Expression (Affy)")
rnaseq = pten.column("mRNA Expression (RNAseq)")
predicted_rnaseq = ...
...
# DON'T CHANGE THE FOLLOWING LINES BELOW IN THIS CELL
rmse_example = RMSE(0.5, 6)
rmse_example
# + deletable=false editable=false
ok.grade("q1_13");
# + [markdown] deletable=false editable=false
# **Question 14**
#
# What is the RMSE of a line with slope 0 and intercept of the mean of `y` equal to?
#
# *Hint 1: The line with slope 0 and intercept of mean of `y` is just a straight horizontal line at the mean of `y`*
#
# *Hint 2: What does the formula for RMSE become if we input our predicted `y` values in the formula. Try writing it out on paper! It should be a familiar formula.*
#
# <!--
# BEGIN QUESTION
# name: q1_14
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] export_pdf=true
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# **Question 15**
#
# Find the parameters that minimizes RMSE of the regression line for mRNA Expression (Affy) vs. mRNA Expression (RNAseq). Assign the result to `minimized_parameters`.
#
# If you haven't tried to use the `minimize` [function](http://data8.org/sp20/python-reference.html) yet, now is a great time to practice. Here's an [example from the textbook](https://www.inferentialthinking.com/chapters/15/3/Method_of_Least_Squares.html#numerical-optimization).
#
# *Hint: Use the `RMSE` function in Question 1.13*
#
# **NOTE: When you use the minimize function, please pass in `smooth=True` as the second argument to this function. You'll need to do this, otherwise, your answer will be incorrect**
#
# <!--
# BEGIN QUESTION
# name: q1_15
# manual: false
# -->
# -
minimized_parameters = ...
minimized_parameters
# + deletable=false editable=false
ok.grade("q1_15");
# + [markdown] deletable=false editable=false
# **Question 16**
#
# The slope and intercept pair you found in Question 1.15 should be very similar to the values that you found in Question 1.7. Why were we able to minimize RMSE to find the same slope and intercept from the previous formulas?
#
#
# <!--
# BEGIN QUESTION
# name: q1_16
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] export_pdf=true
# *Write your answer here, replacing this text.*
# + [markdown] deletable=false editable=false
# **Question 17**
#
# If we had instead minimized mean squared error (MSE), would we have gotten the same slope and intercept of the minimized root mean squared error (RMSE) results? Assign `same_parameters` to either `True` if you think yes, or `False` if you think no.
#
#
# <!--
# BEGIN QUESTION
# name: q1_17
# manual: false
# -->
# -
same_parameters = ...
same_parameters
# + deletable=false editable=false
ok.grade("q1_17");
# -
# Let's look at the scatter plot of the relationship between mRNA Expression (Affy) and mRNA Expression (RNAseq) again:
pten.scatter("mRNA Expression (Affy)", "mRNA Expression (RNAseq)")
# + [markdown] deletable=false editable=false
# **Question 18**
#
# Using a linear regression model, would we be able to obtain accurate predictions for most of the points? Explain why or why not.
#
#
# <!--
# BEGIN QUESTION
# name: q1_18
# manual: true
# -->
# <!-- EXPORT TO PDF -->
# + [markdown] export_pdf=true
# *Write your answer here, replacing this text.*
# -
# ## 2. Properties of Binary Distributions
#
# Binary distributions arise in regular everyday life, and as data scientists you will encounter them constantly. A binary distribution is a distribution across two categories: such as voting in support of a proposition or voting against it on your local ballot, flipping heads or tails, having heart disease or not having heart disease. Generally we represent 'yes' or `True` as 1, and 'no' or `False` as 0. Binary distributions have some special properties that make working with them especially easy!
# The intent of this section of the homework is to walk you through these properties, so we decided to make all of the tests for this section public (i.e. there are no hidden tests to worry about for this section only).
# + [markdown] deletable=false editable=false
# #### Question 1
#
# Let's generate a random binary distribution of 0's and 1's. Assign `binary_options` to the correct array of possible values in a binary distribution (i.e. look at the previous sentence).
#
#
# <!--
# BEGIN QUESTION
# name: q2_1
# manual: false
# -->
# + deletable=false manual_problem_id="sample_with_proportions"
binary_options = ...
# DON'T DELETE/MODIFY ANY OF THE CODE IN THIS CELL BELOW
sample_size = 100
binary_sample = np.random.choice(binary_options, sample_size)
# Run this to see a histogram of this random distribution.
Table().with_columns("Value", make_array(1, 0), "Number in Sample", make_array(sum(binary_sample), sample_size - sum(binary_sample))).barh("Value")
# + deletable=false editable=false
ok.grade("q2_1");
# + [markdown] deletable=false editable=false
# #### Question 2
#
# The first property you should note is that the proportion of ones in a binary distribution is equal to the mean of the distribution. [Think about why this is true](https://www.inferentialthinking.com/chapters/14/1/Properties_of_the_Mean.html#Proportions-are-Means). Complete the following cell to show that this is the case for your `binary_sample`. Assign `number_of_ones` and `number_of_zeros` to the number of `1`'s and the number of `0`'s respectively from your `binary_sample`.
#
#
# <!--
# BEGIN QUESTION
# name: q2_2
# manual: false
# -->
# + deletable=false manual_problem_id="resample_yes_proportions"
number_of_ones = ...
number_of_zeros = ...
# DON'T DELETE/MODIFY ANY OF THE CODE BELOW IN THIS CELL
number_values = len(binary_sample)
sum_of_binary_sample = sum(binary_sample)
# Remember that the mean is equal to the sum divided by the number of items
mean_binary_sample = sum_of_binary_sample / number_values
# Don't change this!
print(f"In your binary sample there were {number_of_ones} ones and {number_of_zeros} zeros. 1*{number_of_ones} + 0*{number_of_zeros} = {number_of_ones}")
print(f"The sum of values in your sample was {sum_of_binary_sample}, divided by the number of items, {number_values}, gives us a mean of {mean_binary_sample}")
print(f"The proportion of ones in your sample was {number_of_ones} ones, divided by the number of items, {number_values}, gives us a value of {mean_binary_sample}" )
print('Those values are equal!')
# + deletable=false editable=false
ok.grade("q2_2");
# -
# Since the proportion of ones is the same as the mean, the Central Limit Theorem applies! That is, if we resample our sample a lot of times, the distribution of the proportion of ones in our resamples will be roughly normal, with a predictable center and spread!
# + deletable=false editable=false
# Just run this cell
resampled_proportion_of_ones = make_array()
for i in np.arange(5000):
resample = Table().with_column("Value", binary_sample).sample()
resample_proportion_ones = resample.where("Value", 1).num_rows / resample.num_rows
resampled_proportion_of_ones = np.append(resampled_proportion_of_ones, resample_proportion_ones)
Table().with_column('Resampled Proportions', resampled_proportion_of_ones).hist()
# -
# Let's generate a table where each row has a different number of ones and zeros that we'll use for the following parts.
# + deletable=false editable=false
# Just run this cell
possible_number_ones = np.arange(sample_size + 1)
possible_number_zeros = sample_size - possible_number_ones
possibilities_table = Table().with_columns("Values of One", possible_number_ones, "Values of Zero", possible_number_zeros)
possibilities_table.show(5)
# + [markdown] deletable=false editable=false
# #### Question 3
# The second important property of binary distributions is that the standard deviation of every binary distribution is equal to:
# $$\sqrt{\text{proportion_ones} *\text{proportion_zeros}}$$
#
# While this property is useful in some cases, a more useful extension of this property is that it tells us that the maximum standard deviation for a binary distribution is 0.5!
#
# Let's explore why that is the case!
#
# Complete the `binary_std_formula` function below so that it returns the standard deviation of a binary distribution according to the formula above.
#
#
# <!--
# BEGIN QUESTION
# name: q2_3
# manual: false
# -->
# + deletable=false manual_problem_id="polling_3"
def binary_std_formula(row):
num_ones = row.item("Values of One")
num_zeros = row.item("Values of Zero")
sum_ones_and_zeros = ...
prop_ones = ...
prop_zeros = ...
...
# DON'T DELETE/MODIFY ANY OF THE LINES BELOW IN THIS CELL
possibilities_table = possibilities_table.with_column("Formula SD", possibilities_table.apply(binary_std_formula))
possibilities_table.show(5)
# + deletable=false editable=false
ok.grade("q2_3");
# -
# Here's another function that takes in a row object from a table, generates a sample that has the same number of ones and zeros as the row specifies, and then returns the standard deviation of that table. You should be able to understand exactly what this function does! It also does the same thing as above, where we return the standard deviation, but we just use `np.std` for this function.
# + deletable=false editable=false
# Just run this cell
def binary_std(row):
values = make_array()
for i in np.arange(row.item("Values of One")):
values = np.append(values, 1)
for i in np.arange(row.item("Values of Zero")):
values = np.append(values, 0)
return np.std(values)
possibilities_table = possibilities_table.with_column("Empirical SD", possibilities_table.apply(binary_std))
possibilities_table.show(5)
# -
# All the values are the same! Let's see what this formula means!
# + deletable=false editable=false
# Just run this cell
possibilities_table.scatter("Values of One", "Formula SD")
# -
# What a beautiful curve!
#
# Looking at that curve, we can see that maximum value is $0.5$, which occurs in the middle of the distribution, when the two categories have equal proportions (proportion of ones = proportion of zeros = $\frac{1}{2}$).
# ## (OPTIONAL, NOT IN SCOPE) Logarithmic Plots
# A kind of visualization you will frequently encounter as a data scientist is a scatter plot or line plot that uses a logarithmic scale. This **Optional** section will cover how to read and generate logarithmic plots. Since this is optional, there is no autograded/free response questions for these sections. Just read, run cells, and explore.
# What is a logarithm? A logarithm helps us find the inverse of an equation that uses exponentials. Specifically, if
#
# $$a^y = x$$
#
# Then
#
# $$\log_a{x} = y$$
#
# The most commonly used $a$, which is known as the base of the logarithm, is $e$, which is equivalent to about 2.718, or 10 (for powers of 10).
#
# We can use `numpy` to take logs in Python! By default, np.log uses a base of e.
make_array(np.log(np.e), np.log(np.e**2), np.log(100))
# Back to the visualization: when we are plotting trends that grow exponentially, such as the line
#
# $$ y = e^x$$
#
# our y-axis needs to have a large range of values, which makes it difficult to understand.
#
# Let's see what this looks like:
# +
x = np.arange(0, 10, 1/100)
y = 10 ** x
Table().with_columns("X", x, "Y", y).scatter(0,1)
# -
# Note that since $10^{10}$ is so big, we can't really see what's happening at all to the y values when they have x values below 8.
#
# One solution to this to change our y and/or x axis so that instead of having even spaces between the tick marks, our marks grow by an uneven factor. We do this by making the tick marks go on a logarithmic scale, and we'll then be able to understand our data better!
Table().with_columns("X", x, "Y", y).scatter(0,1)
plt.yscale("log")
# Now we can tell what's happening to the y values for every x value!
#
# Note how the y values start at $10^0=1$, and increase by a *factor* of $10$ each mark - the next mark is $10^1 = 10$, then $10^2=100$.
#
# You still read this plot like a normal plot, so at a value of $x=5, y=10^5=10000$.
#
# How do you calculate intermediate values?
#
# At a value like $x = 2.5$ it looks like the y value is somewhere in-between $10^1$ and $10^3$. In this graph with a logarithmic scale, you would say that $y=10^{2.5} \approx 316$.
# When visualizing data about the spread of diseases, you will commonly run into plots with logarithmic scales, such as this example from the New York Times. Make sure to always know what the scales of the data are!
#
# <img src="virus-log-chart.jpg" width="650"/>
#
# Image is from https://www.nytimes.com/2020/03/20/health/coronavirus-data-logarithm-chart.html
# ## 3. Submission
#
# Once you're finished, select "Save and Checkpoint" in the File menu and then execute the `submit` cell below. The result will contain a link that you can use to check that your assignment has been submitted successfully. If you submit more than once before the deadline, we will only grade your final submission. If you mistakenly submit the wrong one, you can head to [okpy.org](https://okpy.org/) and flag the correct version. To do so, go to the website, click on this assignment, and find the version you would like to have graded. There should be an option to flag that submission for grading!
_ = ok.submit()
# For your convenience, you can run this cell to run all the tests at once!
import os
print("Running all tests...")
_ = [ok.grade(q[:-3]) for q in os.listdir("tests") if q.startswith('q') and len(q) <= 10]
print("Finished running all tests.")
| materials/sp20/hw/hw10/hw10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using the PyTorch JIT Compiler with Pyro
#
# This tutorial shows how to use the PyTorch [jit compiler](https://pytorch.org/docs/master/jit.html) in Pyro models.
#
# #### Summary:
# - You can use compiled functions in Pyro models.
# - You cannot use pyro primitives inside compiled functions.
# - If your model has static structure, you can use a `Jit*` version of an `ELBO` algorithm, e.g.
# ```diff
# - Trace_ELBO()
# + JitTrace_ELBO()
# ```
# - The [HMC](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.HMC) and [NUTS](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.NUTS) classes accept `jit_compile=True` kwarg.
# - Models should input all tensors as `*args` and all non-tensors as `**kwargs`.
# - Each different value of `**kwargs` triggers a separate compilation.
# - Use `**kwargs` to specify all variation in structure (e.g. time series length).
# - To ignore jit warnings in safe code blocks, use `with pyro.util.ignore_jit_warnings():`.
# - To ignore all jit warnings in `HMC` or `NUTS`, pass `ignore_jit_warnings=True`.
#
# #### Table of contents
# - [Introduction](#Introduction)
# - [A simple model](#A-simple-model)
# - [Varying structure](#Varying-structure)
# +
import os
import torch
import pyro
import pyro.distributions as dist
from torch.distributions import constraints
from pyro import poutine
from pyro.distributions.util import broadcast_shape
from pyro.infer import Trace_ELBO, JitTrace_ELBO, TraceEnum_ELBO, JitTraceEnum_ELBO, SVI
from pyro.infer.mcmc import MCMC, NUTS
from pyro.infer.autoguide import AutoDiagonalNormal
from pyro.optim import Adam
smoke_test = ('CI' in os.environ)
assert pyro.__version__.startswith('1.2.1')
pyro.enable_validation(True) # <---- This is always a good idea!
# -
#
# ## Introduction
#
# PyTorch 1.0 includes a [jit compiler](https://pytorch.org/docs/master/jit.html) to speed up models. You can think of compilation as a "static mode", whereas PyTorch usually operates in "eager mode".
#
# Pyro supports the jit compiler in two ways. First you can use compiled functions inside Pyro models (but those functions cannot contain Pyro primitives). Second, you can use Pyro's jit inference algorithms to compile entire inference steps; in static models this can reduce the Python overhead of Pyro models and speed up inference.
#
# The rest of this tutorial focuses on Pyro's jitted inference algorithms: [JitTrace_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.trace_elbo.JitTrace_ELBO), [JitTraceGraph_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.tracegraph_elbo.JitTraceGraph_ELBO), [JitTraceEnum_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.traceenum_elbo.JitTraceEnum_ELBO), [JitMeanField_ELBO](http://docs.pyro.ai/en/dev/inference_algos.html#pyro.infer.trace_mean_field_elbo.JitTraceMeanField_ELBO), [HMC(jit_compile=True)](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.HMC), and [NUTS(jit_compile=True)](http://docs.pyro.ai/en/dev/mcmc.html#pyro.infer.mcmc.NUTS). For further reading, see the [examples/](https://github.com/pyro-ppl/pyro/tree/dev/examples) directory, where most examples include a `--jit` option to run in compiled mode.
#
# ## A simple model
#
# Let's start with a simple Gaussian model and an [autoguide](http://docs.pyro.ai/en/dev/infer.autoguide.html).
# +
def model(data):
loc = pyro.sample("loc", dist.Normal(0., 10.))
scale = pyro.sample("scale", dist.LogNormal(0., 3.))
with pyro.plate("data", data.size(0)):
pyro.sample("obs", dist.Normal(loc, scale), obs=data)
guide = AutoDiagonalNormal(model)
data = dist.Normal(0.5, 2.).sample((100,))
# -
# First let's run as usual with an SVI object and `Trace_ELBO`.
# %%time
pyro.clear_param_store()
elbo = Trace_ELBO()
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(2 if smoke_test else 1000):
svi.step(data)
# Next to run with a jit compiled inference, we simply replace
# ```diff
# - elbo = Trace_ELBO()
# + elbo = JitTrace_ELBO()
# ```
# Also note that the `AutoDiagonalNormal` guide behaves a little differently on its first invocation (it runs the model to produce a prototype trace), and we don't want to record this warmup behavior when compiling. Thus we call the `guide(data)` once to initialize, then run the compiled SVI,
# +
# %%time
pyro.clear_param_store()
guide(data) # Do any lazy initialization before compiling.
elbo = JitTrace_ELBO()
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(2 if smoke_test else 1000):
svi.step(data)
# -
# Notice that we have a more than 2x speedup for this small model.
#
# Let us now use the same model, but we will instead use MCMC to generate samples from the model's posterior. We will use the No-U-Turn(NUTS) sampler.
# %%time
nuts_kernel = NUTS(model)
pyro.set_rng_seed(1)
mcmc_run = MCMC(nuts_kernel, num_samples=100).run(data)
# We can compile the potential energy computation in NUTS using the `jit_compile=True` argument to the NUTS kernel. We also silence JIT warnings due to the presence of tensor constants in the model by using `ignore_jit_warnings=True`.
# %%time
nuts_kernel = NUTS(model, jit_compile=True, ignore_jit_warnings=True)
pyro.set_rng_seed(1)
mcmc_run = MCMC(nuts_kernel, num_samples=100).run(data)
# We notice a significant increase in sampling throughput when JIT compilation is enabled.
# ## Varying structure
#
# Time series models often run on datasets of multiple time series with different lengths. To accomodate varying structure like this, Pyro requires models to separate all model inputs into tensors and non-tensors.$^\dagger$
#
# - Non-tensor inputs should be passed as `**kwargs` to the model and guide. These can determine model structure, so that a model is compiled for each value of the passed `**kwargs`.
# - Tensor inputs should be passed as `*args`. These must not determine model structure. However `len(args)` may determine model structure (as is used e.g. in semisupervised models).
#
# To illustrate this with a time series model, we will pass in a sequence of observations as a tensor `arg` and the sequence length as a non-tensor `kwarg`:
# +
def model(sequence, num_sequences, length, state_dim=16):
# This is a Gaussian HMM model.
with pyro.plate("states", state_dim):
trans = pyro.sample("trans", dist.Dirichlet(0.5 * torch.ones(state_dim)))
emit_loc = pyro.sample("emit_loc", dist.Normal(0., 10.))
emit_scale = pyro.sample("emit_scale", dist.LogNormal(0., 3.))
# We're doing manual data subsampling, so we need to scale to actual data size.
with poutine.scale(scale=num_sequences):
# We'll use enumeration inference over the hidden x.
x = 0
for t in pyro.markov(range(length)):
x = pyro.sample("x_{}".format(t), dist.Categorical(trans[x]),
infer={"enumerate": "parallel"})
pyro.sample("y_{}".format(t), dist.Normal(emit_loc[x], emit_scale),
obs=sequence[t])
guide = AutoDiagonalNormal(poutine.block(model, expose=["trans", "emit_scale", "emit_loc"]))
# This is fake data of different lengths.
lengths = [24] * 50 + [48] * 20 + [72] * 5
sequences = [torch.randn(length) for length in lengths]
# -
# Now lets' run SVI as usual.
# %%time
pyro.clear_param_store()
elbo = TraceEnum_ELBO(max_plate_nesting=1)
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(1 if smoke_test else 10):
for sequence in sequences:
svi.step(sequence, # tensor args
num_sequences=len(sequences), length=len(sequence)) # non-tensor args
# Again we'll simply swap in a `Jit*` implementation
# ```diff
# - elbo = TraceEnum_ELBO(max_plate_nesting=1)
# + elbo = JitTraceEnum_ELBO(max_plate_nesting=1)
# ```
# Note that we are manually specifying the `max_plate_nesting` arg. Usually Pyro can figure this out automatically by running the model once on the first invocation; however to avoid this extra work when we run the compiler on the first step, we pass this in manually.
# +
# %%time
pyro.clear_param_store()
# Do any lazy initialization before compiling.
guide(sequences[0], num_sequences=len(sequences), length=len(sequences[0]))
elbo = JitTraceEnum_ELBO(max_plate_nesting=1)
svi = SVI(model, guide, Adam({'lr': 0.01}), elbo)
for i in range(1 if smoke_test else 10):
for sequence in sequences:
svi.step(sequence, # tensor args
num_sequences=len(sequences), length=len(sequence)) # non-tensor args
# -
# Again we see more than 2x speedup. Note that since there were three different sequence lengths, compilation was triggered three times.
#
# $^\dagger$ Note this section is only valid for SVI, and HMC/NUTS assume fixed model arguments.
| tutorial/source/jit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="GJRp0mYP-UP3"
import os
import glob
import numpy as np
from tensorflow.keras import layers
from tensorflow import keras
import tensorflow as tf
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 72} id="Z6_0EXBHFAPD" outputId="8812d7eb-6652-4751-fddd-d0d9eef575d9"
from google.colab import files
uploaded = files.upload()
# + id="2-jCaomNEyWN"
f = open("mini_classes.txt","r")
# And for reading use
classes = f.readlines()
f.close()
# + id="eGcCE-7VFuEt"
classes = [c.replace('\n','').replace(' ','_') for c in classes]
# + id="O945ypInFyTn"
# !mkdir data
# + id="nnGmmpPTF1kF"
import urllib.request
def download():
base = 'https://storage.googleapis.com/quickdraw_dataset/full/numpy_bitmap/'
for c in classes:
cls_url = c.replace('_', '%20')
path = base+cls_url+'.npy'
print(path)
urllib.request.urlretrieve(path, 'data/'+c+'.npy')
# + colab={"base_uri": "https://localhost:8080/"} id="Psw9PatyF3dN" outputId="42c93704-cd35-4e54-9e2e-b43df346bfa6"
download()
# + id="FuGizPGs-m4x"
def load_data(root, vfold_ratio=0.2, max_items_per_class= 4000 ):
all_files = glob.glob(os.path.join(root, '*.npy'))
#initialize variables
x = np.empty([0, 784])
y = np.empty([0])
class_names = []
#load each data file
for idx, file in enumerate(all_files):
data = np.load(file)
data = data[0: max_items_per_class, :]
labels = np.full(data.shape[0], idx)
x = np.concatenate((x, data), axis=0)
y = np.append(y, labels)
class_name, ext = os.path.splitext(os.path.basename(file))
class_names.append(class_name)
data = None
labels = None
#randomize the dataset
permutation = np.random.permutation(y.shape[0])
x = x[permutation, :]
y = y[permutation]
#separate into training and testing
vfold_size = int(x.shape[0]/100*(vfold_ratio*100))
x_test = x[0:vfold_size, :]
y_test = y[0:vfold_size]
x_train = x[vfold_size:x.shape[0], :]
y_train = y[vfold_size:y.shape[0]]
return x_train, y_train, x_test, y_test, class_names
# + id="2VwxmWoXCS-p"
x_train, y_train, x_test, y_test, class_names = load_data('data')
num_classes = len(class_names)
image_size = 28
# + colab={"base_uri": "https://localhost:8080/"} id="rPu0DMKFEmA0" outputId="ffd03dc0-2461-468c-843a-87675f1bd800"
print(len(x_train))
# + id="sA9sMZs_Eoi4" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="ce67ea75-1b07-47e9-eab5-ec0bafe4f83d"
import matplotlib.pyplot as plt
from random import randint
# %matplotlib inline
idx = randint(0, len(x_train))
plt.imshow(x_train[idx].reshape(28,28))
print(class_names[int(y_train[idx].item())])
# + id="QUB1_FxSeAy_"
# Reshape and normalize
x_train = x_train.reshape(x_train.shape[0], image_size, image_size, 1).astype('float32')
x_test = x_test.reshape(x_test.shape[0], image_size, image_size, 1).astype('float32')
x_train /= 255.0
x_test /= 255.0
# Convert class vectors to class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + colab={"base_uri": "https://localhost:8080/"} id="m40rLrzEeBTX" outputId="8f475ff5-0a55-4662-ea74-94a95f6dc98e"
# Define model
model = keras.Sequential()
model.add(layers.Convolution2D(16, (3, 3),
padding='same',
input_shape=x_train.shape[1:], activation='relu'))
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Convolution2D(32, (3, 3), padding='same', activation= 'relu'))
model.add(layers.MaxPooling2D(pool_size=(2, 2)))
model.add(layers.Convolution2D(64, (3, 3), padding='same', activation= 'relu'))
model.add(layers.MaxPooling2D(pool_size =(2,2)))
model.add(layers.Flatten())
model.add(layers.Dense(128, activation='relu'))
model.add(layers.Dense(100, activation='softmax'))
# Train model
adam = tf.optimizers.Adam()
model.compile(loss='categorical_crossentropy',
optimizer=adam,
metrics=['top_k_categorical_accuracy'])
print(model.summary())
# + id="m83nEN0veEjZ"
# + colab={"base_uri": "https://localhost:8080/"} id="l3seg33JeaK8" outputId="ea5da514-ad22-46d0-965f-98f0ac7698b9"
model.fit(x = x_train, y = y_train, validation_split=0.1, batch_size = 256, verbose=2, epochs=5)
# + colab={"base_uri": "https://localhost:8080/"} id="SVtsAOd5ectC" outputId="ad37c418-0d09-42a8-9b5b-5ea615b6170b"
score = model.evaluate(x_test, y_test, verbose=0)
print('Test accuarcy: {:0.2f}%'.format(score[1] * 100))
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="04CP3JZMe8g7" outputId="738c9090-9733-4c01-ec8f-c957554938d9"
import matplotlib.pyplot as plt
from random import randint
# %matplotlib inline
idx = randint(0, len(x_test))
img = x_test[idx]
plt.imshow(img.squeeze())
pred = model.predict(np.expand_dims(img, axis=0))[0]
ind = (-pred).argsort()[:5]
latex = [class_names[x] for x in ind]
print(latex)
# + id="3GBSRMy8fAw3"
with open('class_names.txt', 'w') as file_handler:
for item in class_names:
file_handler.write("{}\n".format(item))
# + colab={"base_uri": "https://localhost:8080/"} id="iHN58YYJfD8X" outputId="9faf3b9e-1e33-41a6-8409-bb10bed3d804"
# !pip install tensorflowjs
# + id="aHi9n-UefGfz"
model.save('keras.h5')
# + colab={"base_uri": "https://localhost:8080/"} id="jsp0qnjTfJqW" outputId="f07bfe26-31f7-40c5-c5ad-875ec521d4eb"
# !mkdir model
# !tensorflowjs_converter --input_format keras keras.h5 model/
# + id="xxeQpQMefLyl"
# !cp class_names.txt model/class_names.txt
# + colab={"base_uri": "https://localhost:8080/"} id="fI-zQNmbfPkH" outputId="6aa6ceca-0f9b-4aba-83c9-0e00e19648e7"
# !zip -r model.zip model
# + colab={"base_uri": "https://localhost:8080/", "height": 17} id="RTUPC8b8fSP9" outputId="58adf5f5-4afc-43d0-808a-f399b09bdd2f"
from google.colab import files
files.download('model.zip')
# + id="_7ecs3QOfUbp"
| paint/Uni.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: waf_tutorial_part1
# language: python
# name: waf_tutorial_part1
# ---
# # Notebook 07: Complex ML Regression
#
# ### Goal: Training a ML using all features/predictors/inputs
#
# #### Reminder of Problem Statement
#
#
# Reminder of the ML task we want to accomplish in the paper.
#
# 1. Does this image contain a thunderstorm? <-- Classification
# 2. How many lightning flashes are in this image? <-- Regression
#
# #### Background
#
# Please make sure you already did Notebook 5, because this notebook extends what we did in Notebook 5 to now include many more input predictors. This is the same thing as Notebook 6, but for the regression task.
#
# #### Step 1 & 2: Import packages and load data for Classification
# In Notebook 5 we only wanted 1 feature, no we want all available inputs (36 total). So all we need to change is the ```features_to_keep``` keyword to include all indices. Remember in notebook 5 we wanted to drop the zeros to help. lets do that right away
# +
#needed packages
import xarray as xr
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
#plot parameters that I personally like, feel free to make these your own.
import matplotlib
matplotlib.rcParams['axes.facecolor'] = [0.9,0.9,0.9] #makes a grey background to the axis face
matplotlib.rcParams['axes.labelsize'] = 14 #fontsize in pts
matplotlib.rcParams['axes.titlesize'] = 14
matplotlib.rcParams['xtick.labelsize'] = 12
matplotlib.rcParams['ytick.labelsize'] = 12
matplotlib.rcParams['legend.fontsize'] = 12
matplotlib.rcParams['legend.facecolor'] = 'w'
matplotlib.rcParams['savefig.transparent'] = False
#make default resolution of figures much higher (i.e., High definition)
# %config InlineBackend.figure_format = 'retina'
#import some helper functions for our other directory.
import sys
sys.path.insert(1, '../scripts/')
from aux_functions import load_n_combine_df
(X_train,y_train),(X_validate,y_validate),(X_test,y_test) = load_n_combine_df(path_to_data='../datasets/sevir/',features_to_keep=np.arange(0,36,1),class_labels=False,dropzeros=True)
# -
# I have a habit of always checking shapes, so let's do that
print('X_train, y_train shapes: {},{}'.format(X_train.shape,y_train.shape))
print('X_val, y_val shapes: {},{}'.format(X_validate.shape,y_validate.shape))
print('X_test, y_test shapes: {},{}'.format(X_test.shape,y_test.shape))
# Good, these shapes are indeed less than the classification task (Notebook 6) because we are dropping the zeros.
#
# #### Change from Notebook 5
#
# Since we are using more than 1 input predictor it is important to normalize our predictors. Why is this important? because in reality each one of our inputs have a range of valid values associated with them, and that range could be large (e.g., -100 to 100) or it could be small (e.g., 0,1). The machine learning will weight these inputs quantitatively, so if we use the default scalings, it might be biased to use the larger magnitude predictors more than the small magnitude predictors. To prevent this we will scale the data to have mean 0, and variance 1. You are likely more familiar with the term *standard anomaly* which is the same thing:
#
# $$ z = \frac{x - \mu}{\sigma} $$
#
# where $\mu$ is the mean of that specific feature and $\sigma$ is the standard deviation, both calculated from the training dataset. We could implement this ourselves, but ```sklearn``` has this built for us: ```sklearn.preprocessing.StandardScaler``` and it works alot like how we fit the machine learning model before.
# +
from sklearn.preprocessing import StandardScaler
#create scaling object
scaler = StandardScaler()
#fit scaler to training data
scaler.fit(X_train)
#transform feature data into scaled space
X_train = scaler.transform(X_train)
X_validate = scaler.transform(X_validate)
X_test = scaler.transform(X_test)
#double check that mean is 0 and std is 1.
np.mean(X_train,axis=0),np.std(X_train,axis=0)
# -
# Note that e-16 means $\times 10^{-16}$ which is effectively 0. So it has successfully scaled our data to now have mean 0 and std 1. We are now ready to train our machine learning model again.
# #### Step 3: Initialize model
#
# To keep consistency with Notebook 5, we will continue using linear regression
# +
#load model from sklearn
from sklearn.linear_model import LinearRegression
#initialize
model = LinearRegression()
print(model)
# -
# #### Step 4: Train your ML model!
model = model.fit(X_train,y_train)
# #### Step 5: Evaluate your ML model
#
# As a sanity check, we will first look at the *one-to-one* plot where the x-axis is the predicted number of flashes, and the y-axis is the true number of flashes. A perfect prediction will be directly along the diagonal.
# +
#get predictions
yhat = model.predict(X_validate)
#color I like. The order of ratios is [Red,Green,Blue]
r = [255/255,127/255,127/255]
#make figure
fig = plt.figure(figsize=(5,5))
#set background color to white so we can copy paste out of the notebook if we want
fig.set_facecolor('w')
#get axis for drawing
ax = plt.gca()
#plot data
ax.scatter(yhat,y_validate,color=r,s=1,marker='+')
ax.plot([0,3500],[0,3500],'-k')
ax.set_xlabel('ML Prediction, [$number of flashes$]')
ax.set_xlabel('GLM measurement, [$number of flashes$]')
# -
# To me this looks better than the 1 input feature model we showed in Notebook 5. One troubling thing with these types of scatter plots though is that there are so many points, it is hard to see where the 'mode' of the distribution is. To look a bit deeper into this plot, and to match the plots we used in the paper (Figures 14 and 16). I have a function that will go ahead and bin up the data on this plot and count how many points are in each bin. This function is called ```boxbin``` and it is located in the ```aux_functions.py``` script
# +
from aux_functions import boxbin
#make figure
fig = plt.figure(figsize=(5,5))
#set background color to white so we can copy paste out of the notebook if we want
fig.set_facecolor('w')
#get axis for drawing
ax = plt.gca()
#build the bins we want to use for our data
n = 33
xbins = np.logspace(0,3.5,n) #this logspace logarithmically spaces bins from 10^0 to 10^3.5. The lightning data is better shown on a log-log scale
ybins = np.logspace(0,3.5,n) #we choose to keep the bins square
#Color I like
r = [255/255,127/255,127/255]
#make axes log-log
ax.semilogy()
ax.semilogx()
#same scatter as before
ax.scatter(yhat,y_validate,color=r,s=1,marker='+')
#use the boxbin function to bin data!
ax,cbar,C = boxbin(yhat,y_validate,xbins,ybins,ax=ax,mincnt=100,normed=True,cmap='Reds_r',vmin=0,vmax=2)
cbar.set_label('$\%$ of all points')
ax.set_xlim([1,4000])
ax.set_xticks([1,10,100,1000])
ax.set_yticks([1,10,100,1000])
ax.set_ylim([1,4000])
ax.plot([1,4000],[1,4000],'--k',alpha=0.5)
ax.set_ylabel('$y$, [# of flashes]')
ax.set_xlabel(r'$\hat{y}$, [# of flashes]')
plt.tight_layout()
# -
# Ah that’s better, we can now see where the highest density of points are. Note that only bins with more than 100 points in them will be colored in the boxes.
#
# Let's check on the quantitative metrics to see if it is indeed working better with more features. We will calculate these:
#
# $$ \mathrm{Bias} = \frac{1}{N} \sum_{j=1}^{N} (y_j - \hat{y}_j) $$
#
# $$ \mathrm{MAE} = \frac{1}{N} \sum_{j=1}^{N} |y_j - \hat{y}_j| $$
#
# $$ \mathrm{RMSE} = \sqrt{\frac{1}{N} \sum_{j=1}^{N} (y_j - \hat{y}_j)^{2}} $$
#
# $$ \mathrm{R^{2}} = 1 - \frac{\sum_{j=1}^{N} (y_j - \hat{y}_j)^{2}}{\sum_{j=1}^{N} (y_j - \bar{y})^{2}} $$
#
# All of these metrics again in the ```gewitter_functions.py``` script.
# +
from gewitter_functions import get_mae,get_rmse,get_bias,get_r2
yhat = model.predict(X_validate)
mae = get_mae(y_validate,yhat)
rmse = get_rmse(y_validate,yhat)
bias = get_bias(y_validate,yhat)
r2 = get_r2(y_validate,yhat)
#print them out so we can see them
print('MAE:{} flashes, RMSE:{} flashes, Bias:{} flashes, Rsquared:{}'.format(np.round(mae,2),np.round(rmse,2),np.round(bias,2),np.round(r2,2)))
# -
# The numbers from the single feature model are: *MAE:146.68 flashes, RMSE:235.46 flashes, Bias:30.98 flashes, Rsquared:0.2*. So things have indeed improved on the MAE, Bias and Rsquared metrics. But this is a linear model, so trying to use a linear model for a non-linear forecast (like the number of flashes in a satellite image) is not great, but does do ok.
| jupyter_notebooks/Notebook07_ComplexMLRegression.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Groovy
// language: groovy
// name: groovy
// ---
// ## Minimum Value
// +
//load ImageJ
// %classpath config resolver scijava.public https://maven.scijava.org/content/groups/public
// %classpath add mvn net.imagej imagej 2.0.0-rc-67
//create ImageJ object
ij = new net.imagej.ImageJ()
// -
// This `Op` finds the [Minimum Value](https://en.wikipedia.org/wiki/Maxima_and_minima) of any [`Iterable`](https://docs.oracle.com/javase/8/docs/api/java/lang/Iterable.html).
// +
sinusoid32 = ij.op().run("create.img", [150, 100])
formula = "63 * (Math.cos(0.3*p[0]) + Math.sin(0.3*p[1])) + 127"
ij.op().image().equation(sinusoid32, formula)
ij.notebook().display(sinusoid32)
// -
// All `Img`s are `Iterable`s, so we can just pass through the `Img` to `min()`:
// +
import net.imglib2.type.numeric.real.DoubleType
output = new DoubleType()
ij.op().stats().min(output, sinusoid32)
output
| notebooks/1-Using-ImageJ/Ops/stats/min.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from abc import ABC, abstractmethod
from pathlib import Path
from typing import List
from functools import partial, reduce
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
DATA_CLEANED_DIR = Path('../data/cleaned/')
# import files
df_country_codes = pd.read_csv(Path(DATA_CLEANED_DIR, 'country_codes.csv'))
df_life_exp = pd.read_csv(Path(DATA_CLEANED_DIR, 'life_expectancy.csv'))
df_mortality = pd.read_csv(Path(DATA_CLEANED_DIR, 'mortality.csv'))
df_population = pd.read_csv(Path(DATA_CLEANED_DIR, 'population.csv'))
# # Netherlands & Japan & Canada
# We first want to visualize the data of The Netherlands
df_country_codes.head(1)
df_country_codes[(df_country_codes['country'] == 'Netherlands') |
(df_country_codes['country'] == 'Japan') |
(df_country_codes['country'] == 'Canada')]
# As we can see, the country code of The Netherlands is `4210`. Therefore we will filter the *life expectancy*, *mortality* and *population datasets on this country code.
NL_CODE = 4210
JP_CODE = 3160
CA_CODE = 2090
# At the moment the **Life expectancy** dataset contains only data for the Netherlands. The data is obtained from https://www.who.int/data/maternal-newborn-child-adolescent-ageing/indicator-explorer-new/mca/life-expectancy-at-birth. Due to the fact that the *Export* button did not work at that time, I copied it manually in the dataset.
df_life_exp
# +
nl_life_exp = df_life_exp[(df_life_exp['country'] == 'Netherlands')]
jp_life_exp = df_life_exp[(df_life_exp['country'] == 'Japan')]
ca_life_exp = df_life_exp[(df_life_exp['country'] == 'Canada')]
nl_mortality = df_mortality[(df_mortality['country code'] == NL_CODE)]
jp_mortality = df_mortality[(df_mortality['country code'] == JP_CODE)]
ca_mortality = df_mortality[(df_mortality['country code'] == CA_CODE)]
nl_population = df_population[(df_population['country code'] == NL_CODE)]
jp_population = df_population[(df_population['country code'] == JP_CODE)]
ca_population = df_population[(df_population['country code'] == CA_CODE)]
# -
# So it is best to merge them into one single file. In this way we can standardize the values. You want to merge them on `year` and `sex`.
#
# Coding:
# `{1: male, 2: female, 3: both sexes}`
life_exp_set = [nl_life_exp, jp_life_exp, ca_life_exp]
mort_set = [nl_mortality, jp_mortality, ca_mortality]
pop_set = [nl_population, jp_population, ca_population]
for df in life_exp_set:
print(df.sex.unique())
for df in mort_set:
print(df.sex.unique())
for df in pop_set:
print(df.sex.unique())
ca_mortality[ca_mortality['sex'] == 9]
ca_population[ca_population['sex'] == 9]
# drop sex 9 as this is not usesful
ca_mortality = ca_mortality[~ca_mortality['sex'].isin([9])]
ca_population = ca_population[~ca_population['sex'].isin([9])]
# #### It is first important to create an aggregate of sexes for the *mortality* and *population* dataset.
# +
def generate_ICD_codes(lower, upper, symbol):
codes = []
for i in range(lower, upper + 1, 1):
if i < 10:
codes.append(f'{symbol}0{i}')
else:
codes.append(f'{symbol}{i}')
return np.array(codes)
def convert_format(series, n=3):
"""Only keep the n first characters of the column"""
return series.apply(lambda x: x[:n])
def filter_column(df: pd.DataFrame, column: str, elements):
"""
:param df:
:param column:
:param elements:
:return:
"""
target = df[column].unique()
found_elements = find_elements(target, elements)
dataset = df[df[column].isin(found_elements)]
return dataset
def groupby_sum(df, by, on):
"""Groups the dataframe by the index columns, and sums the target column and
returns the result as a dataframe."""
grouped = df.groupby(by, as_index=False)[on].sum()
return grouped
def find_elements(target, elements):
"""
Checks if the targets can be found in an arbitrary list of elements.
:param elements:
:param target:
:return:
"""
mask = np.isin(target, elements)
found = np.where(mask, target, '')
valid = [c for c in found if c != '']
return valid
# +
class Selector(ABC):
def __init__(self, file):
self.file = file
@abstractmethod
def get_selection(self):
pass
class DataFrameSelector(Selector):
def __init__(self, df):
if not isinstance(df, pd.DataFrame):
raise ValueError('Expects a pandas DataFrame.')
super().__init__(df)
self.selection = {}
def filter_column(self, column: str, elements):
target = self.file[column].unique()
found_elements = find_elements(target, elements)
dataset = self.file[self.file[column].isin(found_elements)]
return dataset
def split_dataframe(self, column: str, labels: List[str], selection: List[np.ndarray]):
"""Expects labels as keys and the selection to be the string to select the dataframe on.
if unique elements are given then it searches for elements found in the selection and the unique list
"""
for label, selector in zip(labels, selection):
dataset = self.filter_column(column, selector)
self.selection[label] = dataset
def rename_selection(self, column, mapping):
for k, df in self.selection.items():
self.selection[k] = df.rename(columns={column: mapping[k]})
def get_selection(self):
return self.selection
# -
class Aggregator:
def __init__(self, df):
self.df = df
self.aggregation = None
def handler(self, df):
if self.aggregation is None:
self.aggregation = df
else:
self.aggregation = self.aggregation.append(df)
def calc_aggr(self, by, on, column='', value=None):
# calculate aggregate and append it to the original
if not column and not value:
self.handler(groupby_sum(self.df, by, on))
return
aggr = groupby_sum(self.df, by, on)
aggr[column] = value
self.handler(aggr)
def get_aggregation(self, sort_by):
return self.aggregation.sort_values(sort_by).reset_index(drop=True)
# +
C_codes = generate_ICD_codes(0, 97, 'C')
I_codes = generate_ICD_codes(5, 99, 'I')
E_codes = generate_ICD_codes(10, 13, 'E')
J_codes = generate_ICD_codes(40, 47, 'J')
K_codes = generate_ICD_codes(0, 93, 'K')
code_to_name_map = {
'C': 'cancer [deaths]',
'I': 'cardiovascular disease [deaths]',
'E': 'diabetes mellitus [deaths]',
'J': 'chronic respiratory diseases [deaths]',
'K': 'diseases of digestive system [deaths]'
}
name_to_code_map = {
'cancer [deaths]': 'C',
'cardiovascular disease [deaths]': 'I',
'diabetes mellitus [deaths]': 'E',
'chronic respiratory diseases [deaths]': 'J',
'diseases of digestive system [deaths]': 'K',
}
code_map = {
'C': 'C',
'I': 'I',
'E': 'E',
'J': 'J',
'K': 'K'
}
codes = [C_codes, I_codes, E_codes, J_codes, K_codes]
labels = ['C', 'I', 'E', 'J', 'K']
# -
def convert_format(items: list, n=3):
"""Only keep the n first characters of the column"""
series = pd.Series(items)
return series.apply(lambda x: x[:n])
# +
nl_causes = convert_format(nl_mortality['cause'], 3)
jp_causes = convert_format(jp_mortality['cause'], 3)
ca_causes = convert_format(ca_mortality['cause'], 3)
nl_unique_causes = nl_causes.unique()
jp_unique_causes = jp_causes.unique()
ca_unique_causes = ca_causes.unique()
nl_mortality.loc[:, 'cause'] = nl_causes
jp_mortality.loc[:, 'cause'] = jp_causes
ca_mortality.loc[:, 'cause'] = ca_causes
# -
nl_mortality
# +
# Select correct data
nl_mortality_selector = DataFrameSelector(nl_mortality)
nl_mortality_selector.split_dataframe('cause', labels, codes)
nl_mortality_selector.rename_selection('deaths', code_to_name_map)
nl_mortality_sets = nl_mortality_selector.get_selection()
jp_mortality_selector = DataFrameSelector(jp_mortality)
jp_mortality_selector.split_dataframe('cause', labels, codes)
jp_mortality_selector.rename_selection('deaths', code_to_name_map)
jp_mortality_sets = jp_mortality_selector.get_selection()
ca_mortality_selector = DataFrameSelector(ca_mortality)
ca_mortality_selector.split_dataframe('cause', labels, codes)
ca_mortality_selector.rename_selection('deaths', code_to_name_map)
ca_mortality_sets = ca_mortality_selector.get_selection()
# create aggregates
for k, df in nl_mortality_sets.items():
aggr = Aggregator(df)
aggr.calc_aggr(by=['year', 'sex'], on=code_to_name_map[k])
aggr.calc_aggr(by=['year'], on=code_to_name_map[k], column='sex', value=3)
data = aggr.get_aggregation(sort_by='year')
data['country'] = 'Netherlands'
nl_mortality_sets[k] = data
for k, df in jp_mortality_sets.items():
aggr = Aggregator(df)
aggr.calc_aggr(by=['year', 'sex'], on=code_to_name_map[k])
aggr.calc_aggr(by=['year'], on=code_to_name_map[k], column='sex', value=3)
data = aggr.get_aggregation(sort_by='year')
data['country'] = 'Japan'
jp_mortality_sets[k] = data
for k, df in ca_mortality_sets.items():
aggr = Aggregator(df)
aggr.calc_aggr(by=['year', 'sex'], on=code_to_name_map[k])
aggr.calc_aggr(by=['year'], on=code_to_name_map[k], column='sex', value=3)
data = aggr.get_aggregation(sort_by='year')
data['country'] = 'Canada'
ca_mortality_sets[k] = data
# -
nl_mortality_sets['C'].head()
jp_mortality_sets['C'].head()
ca_mortality_sets['C'].head()
# ### Population
# Population is already properly formatted, thus no need to make a selection.
#
# +
aggr = Aggregator(nl_population)
aggr.calc_aggr(by=['year', 'sex'], on='population')
aggr.calc_aggr(by=['year'], on='population', column='sex', value=3)
data = aggr.get_aggregation(sort_by='year')
data['country'] = 'Netherlands'
nl_pop_agg = data
nl_pop_agg.isna().sum()
# +
aggr = Aggregator(jp_population)
aggr.calc_aggr(by=['year', 'sex'], on='population')
aggr.calc_aggr(by=['year'], on='population', column='sex', value=3)
data = aggr.get_aggregation(sort_by='year')
data['country'] = 'Japan'
jp_pop_agg = data
jp_pop_agg.isna().sum()
# +
aggr = Aggregator(ca_population)
aggr.calc_aggr(by=['year', 'sex'], on='population')
aggr.calc_aggr(by=['year'], on='population', column='sex', value=3)
data = aggr.get_aggregation(sort_by='year')
data['country'] = 'Canada'
ca_pop_agg = data
ca_pop_agg.isna().sum()
# -
# ## Combine all
# +
# nl_pop_agg = nl_pop_agg[nl_pop_agg['year'] >= 1996]
# nl_life_exp = nl_life_exp[(nl_life_exp['year'] >= 1996) & (nl_life_exp['year'] <= 2018)]
# jp_pop_agg = jp_pop_agg[jp_pop_agg['year'] >= 1996]
# jp_life_exp = jp_life_exp[(jp_life_exp['year'] >= 1996) & (jp_life_exp['year'] <= 2018)]
# ca_pop_agg = ca_pop_agg[ca_pop_agg['year'] >= 1996]
# ca_life_exp = ca_life_exp[(ca_life_exp['year'] >= 1996) & (ca_life_exp['year'] <= 2018)]
# -
nl_pop_agg.isna().sum()
jp_pop_agg.isna().sum()
ca_pop_agg.isna().sum()
# +
nl_dfs = []
jp_dfs = []
ca_dfs = []
nl_dfs.extend([df for df in nl_mortality_sets.values()])
jp_dfs.extend([df for df in jp_mortality_sets.values()])
ca_dfs.extend([df for df in ca_mortality_sets.values()])
nl_dfs.append(nl_pop_agg)
nl_dfs.append(nl_life_exp)
jp_dfs.append(jp_pop_agg)
jp_dfs.append(jp_life_exp)
ca_dfs.append(ca_pop_agg)
ca_dfs.append(ca_life_exp)
# -
def multi_merge(dfs, on):
merge = partial(pd.merge, on=on, how='outer')
dataset = reduce(merge, dfs)
return dataset
# +
nl_dataset = multi_merge(nl_dfs, on=['year', 'sex', 'country']).sort_values('year')
nl_dataset = dataset[(dataset['year'] > 1995) & (dataset['year'] < 2019)]
nl_dataset['non-communicable chronic disease [deaths]'] = 0
for disease in name_to_code_map.keys():
nl_dataset['non-communicable chronic disease [deaths]'] += dataset[disease]
nl_dataset = nl_dataset.sort_values(['year', 'sex']).reset_index(drop=True)
# +
jp_dataset = multi_merge(jp_dfs, on=['year', 'sex', 'country']).sort_values('year')
jp_dataset = dataset[(dataset['year'] > 1995) & (dataset['year'] < 2019)]
jp_dataset['non-communicable chronic disease [deaths]'] = 0
for disease in name_to_code_map.keys():
jp_dataset['non-communicable chronic disease [deaths]'] += dataset[disease]
jp_dataset = jp_dataset.sort_values(['year', 'sex']).reset_index(drop=True)
# +
ca_dataset = multi_merge(ca_dfs, on=['year', 'sex', 'country']).sort_values('year')
ca_dataset = dataset[(dataset['year'] > 1995) & (dataset['year'] < 2019)]
ca_dataset['non-communicable chronic disease [deaths]'] = 0
for disease in name_to_code_map.keys():
ca_dataset['non-communicable chronic disease [deaths]'] += dataset[disease]
ca_dataset = ca_dataset.sort_values(['year', 'sex']).reset_index(drop=True)
# -
full_dataset = pd.concat([nl_dataset, jp_dataset, ca_dataset]).reset_index(drop=True)
full_dataset
full_dataset.to_csv('../data/prepared/full_dataset.csv', index=False)
# +
cols = ['cancer [deaths]',
'cardiovascular disease [deaths]', 'diabetes mellitus [deaths]',
'chronic respiratory diseases [deaths]',
'diseases of digestive system [deaths]',
'non-communicable chronic disease [deaths]']
full_dataset_standardized = full_dataset
full_dataset_standardized[cols] = full_dataset[cols].divide(full_dataset['population'], axis=0) * 100000
full_dataset_standardized
# +
full_dataset_standardized.to_csv('../data/prepared/full_dataset_standardized.csv', index=False)
# -
| src/notebooks/Data selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Load Libraries
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import torchvision
# ### Defining Memnet Model
# +
"""MemNet"""
class MemNet(nn.Module):
def __init__(self, in_channels, channels, num_memblock, num_resblock):
super(MemNet, self).__init__()
self.feature_extractor = BNReLUConv(in_channels, channels)
self.reconstructor = BNReLUConv(channels, in_channels)
self.dense_memory = nn.ModuleList(
[MemoryBlock(channels, num_resblock, i+1) for i in range(num_memblock)]
)
def forward(self, x):
# x = x.contiguous()
residual = x
out = self.feature_extractor(x)
ys = [out]
for memory_block in self.dense_memory:
out = memory_block(out, ys)
out = self.reconstructor(out)
out = out + residual
return out
class MemoryBlock(nn.Module):
"""Note: num_memblock denotes the number of MemoryBlock currently"""
def __init__(self, channels, num_resblock, num_memblock):
super(MemoryBlock, self).__init__()
self.recursive_unit = nn.ModuleList(
[ResidualBlock(channels) for i in range(num_resblock)]
)
self.gate_unit = BNReLUConv((num_resblock+num_memblock) * channels, channels, 1, 1, 0)
def forward(self, x, ys):
"""ys is a list which contains long-term memory coming from previous memory block
xs denotes the short-term memory coming from recursive unit
"""
xs = []
residual = x
for layer in self.recursive_unit:
x = layer(x)
xs.append(x)
gate_out = self.gate_unit(torch.cat(xs+ys, 1))
ys.append(gate_out)
return gate_out
class ResidualBlock(torch.nn.Module):
"""ResidualBlock
introduced in: https://arxiv.org/abs/1512.03385
x - Relu - Conv - Relu - Conv - x
"""
def __init__(self, channels, k=3, s=1, p=1):
super(ResidualBlock, self).__init__()
self.relu_conv1 = BNReLUConv(channels, channels, k, s, p)
self.relu_conv2 = BNReLUConv(channels, channels, k, s, p)
def forward(self, x):
residual = x
out = self.relu_conv1(x)
out = self.relu_conv2(out)
out = out + residual
return out
class BNReLUConv(nn.Sequential):
def __init__(self, in_channels, channels, k=3, s=1, p=1, inplace=True):
super(BNReLUConv, self).__init__()
self.add_module('bn', nn.BatchNorm2d(in_channels))
self.add_module('relu', nn.ReLU(inplace=inplace))
self.add_module('conv', nn.Conv2d(in_channels, channels, k, s, p, bias=False))
# -
# ### Data
# +
imagesPath = './images/'
train_dataset = torchvision.datasets.ImageFolder(
root=imagesPath,
transform=torchvision.transforms.ToTensor()
)
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=64,
num_workers=0,
shuffle=True
)
# -
# ### Train Model
# +
def to_img(x):
x = 0.5 * (x + 1)
x = x.clamp(0, 1)
x = x.view(x.size(0), 1, 28, 28)
return x
num_epochs = 100
batch_size = 64
learning_rate = 1e-3
model = MemNet(2,2,2,2).cuda()
criterion = nn.MSELoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate, weight_decay=1e-5)
for epoch in range(num_epochs):
for data in train_loader:
img, = data
img = img.view(img.size(0), -1)
img = Variable(img).cuda()
# ===================forward=====================
output = model(img)
loss = criterion(output, img)
# ===================backward====================
optimizer.zero_grad()
loss.backward()
optimizer.step()
# ===================log========================
print('epoch [{}/{}], loss:{:.4f}'
.format(epoch + 1, num_epochs, loss.item()))
if epoch % 10 == 0:
pic = to_img(output.cpu().data)
save_image(pic, './mlp_img/image_{}.png'.format(epoch))
torch.save(model.state_dict(), './sim_autoencoder.pth')
# +
# #!/usr/bin/env python
from __future__ import print_function
import os
path = '.'
files = os.listdir(path)
for name in files:
print(name)
# -
| Main/MemNet/VanderMode Memnet/VanderMode Memnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Transformer spam classifier"
# > "Spam classifier using allennlp and pretrained transformer"
#
# - toc: true
# - badges: False
# - comments: true
# - categories: [allennlp]
# - hide: false
# This is a classifier which uses pretrained transformer as its embedding, implemented using allennlp 2.4.0.
#
# Dataset: https://www.kaggle.com/team-ai/spam-text-message-classification
#hide
# !pip install allennlp
# DatasetReader
# +
# DatasetReader
from allennlp.data import DatasetReader, Instance
from allennlp.data.fields import LabelField, TextField
class ClassificationTsvReader(DatasetReader):
def __init__(
self,
tokenizer=None,
token_indexers=None,
**kwargs
):
super().__init__(**kwargs)
self.tokenizer = tokenizer
self.token_indexers = token_indexers
def _read(self, file_path):
with open(file_path, "r") as lines:
for line in lines:
label, text = line.strip().split("\t")
tokens = self.tokenizer.tokenize(text)
text_field = TextField(tokens, self.token_indexers)
label_field = LabelField(label)
fields = {"tokens": text_field, "label": label_field}
yield Instance(fields)
def text_to_instance(self, text: str, label: str = None) -> Instance:
tokens = self.tokenizer.tokenize(text)
text_field = TextField(tokens, self.token_indexers)
fields = {'tokens': text_field}
if label:
fields['label'] = LabelField(label)
return Instance(fields)
# -
# Tokenizer
# Tokenizer
from allennlp.data.tokenizers import PretrainedTransformerTokenizer
tokenizer = PretrainedTransformerTokenizer("roberta-base")
# Indexer
# Indexer
from allennlp.data.token_indexers import PretrainedTransformerIndexer
token_indexers = {"tokens": PretrainedTransformerIndexer("roberta-base")}
# Reading instances
# +
# Reading instances
train_file='spam-classifier/data/spam-train.tsv'
dev_file='spam-classifier/data/spam-dev.tsv'
dataset_reader = ClassificationTsvReader(
tokenizer=tokenizer,
token_indexers=token_indexers)
train_instances = list(dataset_reader.read(train_file))
dev_instances = list(dataset_reader.read(dev_file))
# -
# Vocabulary
# +
# Vocabulary
from allennlp.data.vocabulary import Vocabulary
vocab = Vocabulary()
vocab = vocab.from_pretrained_transformer(model_name="roberta-base", namespace="tokens")
vocab.extend_from_instances(train_instances+dev_instances)
# -
# Token embedder
# Token embedder
from allennlp.modules.token_embedders import PretrainedTransformerEmbedder
token_embedder = PretrainedTransformerEmbedder("roberta-base")
# Text-field embedder
# Text-field embedder
from allennlp.modules.text_field_embedders import BasicTextFieldEmbedder
embedder = BasicTextFieldEmbedder({"tokens": token_embedder})
# Encoder
# Encoder
from allennlp.modules.seq2vec_encoders import ClsPooler
encoder = ClsPooler(embedding_dim=embedder.get_output_dim())
# Model
# +
# Model
from allennlp.models import BasicClassifier
model=BasicClassifier(
vocab=vocab,
text_field_embedder=embedder,
seq2vec_encoder=encoder,
namespace="tokens",
label_namespace="labels"
)
# -
# DataLoaders
# +
# DataLoaders
from allennlp.data.data_loaders import SimpleDataLoader
train_data_loader=SimpleDataLoader(
instances=train_instances,
batch_size=16,
shuffle=True,
vocab=vocab,
)
dev_data_loader=SimpleDataLoader(
instances=dev_instances,
batch_size=16,
shuffle=True,
vocab=vocab
)
# -
# Trainer
# +
# Trainer
from allennlp.training import GradientDescentTrainer
from torch.optim import Adam
trainer = GradientDescentTrainer(
model=model,
optimizer=Adam(model.parameters()),
data_loader=train_data_loader,
validation_data_loader=dev_data_loader,
patience=3,
num_epochs=100,
)
# -
# Preparing GPU training
# +
# Preparing GPU training
import torch
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
model.to(device)
train_data_loader.set_target_device(device)
dev_data_loader.set_target_device(device)
# -
# Train the model
# Train the model
# %time trainer.train()
# Predictor
# Predictor
from allennlp.predictors.text_classifier import TextClassifierPredictor
predictor = TextClassifierPredictor(model=model, dataset_reader=dataset_reader)
# Test predictor
# Test predictor
predictor.predict("hello world")
# Save model
# +
# Save model
model_dir = 'simple-classifier-spam/'
with open(model_dir + "model.th", 'wb') as f:
torch.save(model.state_dict(), f)
vocab.save_to_files(model_dir + "vocabulary")
| _notebooks/2021-06-04-transformer-classifer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
dataname="synthetic"
patch_size=256 #size of the tiles to put into DB
data_size=[10000,100]
balance=.5
classes=[0,1] #what classes we expect to have in the data, in this case data without boxes and data with boxes
max_circles=10
max_squares=1
diameter_min=10
diameter_max=50
phases=["train","val"]
# +
import random
import tables
import sys
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image, ImageDraw
seed = random.randrange(sys.maxsize) #get a random seed so that we can reproducibly do the cross validation setup
random.seed(seed) # set the seed
print(f"random seed (note down for reproducibility): {seed}")
# -
img_dtype = tables.UInt8Atom() # dtype in which the images will be saved, this indicates that images will be saved as unsigned int 8 bit, i.e., [0,255]
# +
# %matplotlib inline
storage={} #holder for future pytables
block_shape=np.array((patch_size,patch_size)) #block shape specifies what we'll be saving into the pytable array, here we assume that masks are 1d and images are 3d
filters=tables.Filters(complevel=6, complib='zlib') #we can also specify filters, such as compression, to improve storage speed
for phase,nimgs in zip(phases,data_size): #now for each of the phases, we'll loop through the files
print(phase)
totals=np.zeros(2) # we can to keep counts of all the classes in for in particular training, since we
hdf5_file = tables.open_file(f"./{dataname}_{phase}.pytable", mode='w') #open the respective pytable
storage["imgs"]= hdf5_file.create_earray(hdf5_file.root, "imgs", img_dtype,
shape=np.append([0],block_shape),
chunkshape=np.append([1],block_shape),
filters=filters)
storage["labels"]= hdf5_file.create_earray(hdf5_file.root, "labels", img_dtype,
shape=[0],
chunkshape=[1],
filters=filters)
for filei in range(nimgs): #now for each of the files
img=np.zeros((patch_size,patch_size))
img = Image.fromarray(img)
draw= ImageDraw.Draw(img)
for i in range(np.random.randint(0,high=max_circles)):
d=np.random.randint(diameter_min,diameter_max)
ul=np.random.randint(diameter_min,patch_size-diameter_max,2)
draw.ellipse(list(np.append(ul,ul+d)),fill=255)
label=np.random.random()>balance
if label:
for i in range(np.random.randint(1,high=max_squares+1)):
d=np.random.randint(diameter_min,diameter_max)
ul=np.random.randint(diameter_min,patch_size-diameter_max,2)
draw.rectangle(list(np.append(ul,ul+d)),fill=255)
totals[1]+=1
else:
totals[0]+=1
#add square
del draw
storage["imgs"].append(np.array(img)[None,::])
storage["labels"].append([np.uint8(label)]) #add the filename to the storage array
#lastely, we should store the number of pixels
npixels=hdf5_file.create_carray(hdf5_file.root, 'classsizes', tables.Atom.from_dtype(totals.dtype), totals.shape)
npixels[:]=totals
hdf5_file.close()
print("done")
# -
| visualization_densenet/make_hdf5_synthetic_circles_and_boxes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What's this TensorFlow business?
#
# You've written a lot of code in this assignment to provide a whole host of neural network functionality. Dropout, Batch Norm, and 2D convolutions are some of the workhorses of deep learning in computer vision. You've also worked hard to make your code efficient and vectorized.
#
# For the last part of this assignment, though, we're going to leave behind your beautiful codebase and instead migrate to one of two popular deep learning frameworks: in this instance, TensorFlow (or PyTorch, if you switch over to that notebook)
#
# #### What is it?
# TensorFlow is a system for executing computational graphs over Tensor objects, with native support for performing backpropogation for its Variables. In it, we work with Tensors which are n-dimensional arrays analogous to the numpy ndarray.
#
# #### Why?
#
# * Our code will now run on GPUs! Much faster training. Writing your own modules to run on GPUs is beyond the scope of this class, unfortunately.
# * We want you to be ready to use one of these frameworks for your project so you can experiment more efficiently than if you were writing every feature you want to use by hand.
# * We want you to stand on the shoulders of giants! TensorFlow and PyTorch are both excellent frameworks that will make your lives a lot easier, and now that you understand their guts, you are free to use them :)
# * We want you to be exposed to the sort of deep learning code you might run into in academia or industry.
# ## How will I learn TensorFlow?
#
# TensorFlow has many excellent tutorials available, including those from [Google themselves](https://www.tensorflow.org/get_started/get_started).
#
# Otherwise, this notebook will walk you through much of what you need to do to train models in TensorFlow. See the end of the notebook for some links to helpful tutorials if you want to learn more or need further clarification on topics that aren't fully explained here.
#
#
# # Table of Contents
#
# This notebook has 5 parts. We will walk through TensorFlow at three different levels of abstraction, which should help you better understand it and prepare you for working on your project.
#
# 1. Preparation: load the CIFAR-10 dataset.
# 2. Barebone TensorFlow: we will work directly with low-level TensorFlow graphs.
# 3. Keras Model API: we will use `tf.keras.Model` to define arbitrary neural network architecture.
# 4. Keras Sequential API: we will use `tf.keras.Sequential` to define a linear feed-forward network very conveniently.
# 5. CIFAR-10 open-ended challenge: please implement your own network to get as high accuracy as possible on CIFAR-10. You can experiment with any layer, optimizer, hyperparameters or other advanced features.
#
# Here is a table of comparison:
#
# | API | Flexibility | Convenience |
# |---------------|-------------|-------------|
# | Barebone | High | Low |
# | `tf.keras.Model` | High | Medium |
# | `tf.keras.Sequential` | Low | High |
# # Part I: Preparation
#
# First, we load the CIFAR-10 dataset. This might take a few minutes to download the first time you run it, but after that the files should be cached on disk and loading should be faster.
#
# In previous parts of the assignment we used CS231N-specific code to download and read the CIFAR-10 dataset; however the `tf.keras.datasets` package in TensorFlow provides prebuilt utility functions for loading many common datasets.
#
# For the purposes of this assignment we will still write our own code to preprocess the data and iterate through it in minibatches. The `tf.data` package in TensorFlow provides tools for automating this process, but working with this package adds extra complication and is beyond the scope of this notebook. However using `tf.data` can be much more efficient than the simple approach used in this notebook, so you should consider using it for your project.
# +
import os
import tensorflow as tf
import numpy as np
import math
import timeit
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def load_cifar10(num_training=49000, num_validation=1000, num_test=10000):
"""
Fetch the CIFAR-10 dataset from the web and perform preprocessing to prepare
it for the two-layer neural net classifier. These are the same steps as
we used for the SVM, but condensed to a single function.
"""
# Load the raw CIFAR-10 dataset and use appropriate data types and shapes
cifar10 = tf.keras.datasets.cifar10.load_data()
(X_train, y_train), (X_test, y_test) = cifar10
X_train = np.asarray(X_train, dtype=np.float32)
y_train = np.asarray(y_train, dtype=np.int32).flatten()
X_test = np.asarray(X_test, dtype=np.float32)
y_test = np.asarray(y_test, dtype=np.int32).flatten()
# Subsample the data
mask = range(num_training, num_training + num_validation)
X_val = X_train[mask]
y_val = y_train[mask]
mask = range(num_training)
X_train = X_train[mask]
y_train = y_train[mask]
mask = range(num_test)
X_test = X_test[mask]
y_test = y_test[mask]
# Normalize the data: subtract the mean pixel and divide by std
mean_pixel = X_train.mean(axis=(0, 1, 2), keepdims=True)
std_pixel = X_train.std(axis=(0, 1, 2), keepdims=True)
X_train = (X_train - mean_pixel) / std_pixel
X_val = (X_val - mean_pixel) / std_pixel
X_test = (X_test - mean_pixel) / std_pixel
return X_train, y_train, X_val, y_val, X_test, y_test
# Invoke the above function to get our data.
NHW = (0, 1, 2)
X_train, y_train, X_val, y_val, X_test, y_test = load_cifar10()
print('Train data shape: ', X_train.shape)
print('Train labels shape: ', y_train.shape, y_train.dtype)
print('Validation data shape: ', X_val.shape)
print('Validation labels shape: ', y_val.shape)
print('Test data shape: ', X_test.shape)
print('Test labels shape: ', y_test.shape)
# -
# ### Preparation: Dataset object
#
# For our own convenience we'll define a lightweight `Dataset` class which lets us iterate over data and labels. This is not the most flexible or most efficient way to iterate through data, but it will serve our purposes.
# +
class Dataset(object):
def __init__(self, X, y, batch_size, shuffle=False):
"""
Construct a Dataset object to iterate over data X and labels y
Inputs:
- X: Numpy array of data, of any shape
- y: Numpy array of labels, of any shape but with y.shape[0] == X.shape[0]
- batch_size: Integer giving number of elements per minibatch
- shuffle: (optional) Boolean, whether to shuffle the data on each epoch
"""
assert X.shape[0] == y.shape[0], 'Got different numbers of data and labels'
self.X, self.y = X, y
self.batch_size, self.shuffle = batch_size, shuffle
def __iter__(self):
N, B = self.X.shape[0], self.batch_size
idxs = np.arange(N)
if self.shuffle:
np.random.shuffle(idxs)
return iter((self.X[i:i+B], self.y[i:i+B]) for i in range(0, N, B))
train_dset = Dataset(X_train, y_train, batch_size=64, shuffle=True)
val_dset = Dataset(X_val, y_val, batch_size=64, shuffle=False)
test_dset = Dataset(X_test, y_test, batch_size=64)
# -
# We can iterate through a dataset like this:
for t, (x, y) in enumerate(train_dset):
print(t, x.shape, y.shape)
if t > 5: break
# You can optionally **use GPU by setting the flag to True below**. It's not neccessary to use a GPU for this assignment; if you are working on Google Cloud then we recommend that you do not use a GPU, as it will be significantly more expensive.
# +
# Set up some global variables
USE_GPU = True
if USE_GPU:
device = '/device:GPU:0'
else:
device = '/cpu:0'
# Constant to control how often we print when training models
print_every = 100
print('Using device: ', device)
# -
# # Part II: Barebone TensorFlow
# TensorFlow ships with various high-level APIs which make it very convenient to define and train neural networks; we will cover some of these constructs in Part III and Part IV of this notebook. In this section we will start by building a model with basic TensorFlow constructs to help you better understand what's going on under the hood of the higher-level APIs.
#
# TensorFlow is primarily a framework for working with **static computational graphs**. Nodes in the computational graph are Tensors which will hold n-dimensional arrays when the graph is run; edges in the graph represent functions that will operate on Tensors when the graph is run to actually perform useful computation.
#
# This means that a typical TensorFlow program is written in two distinct phases:
#
# 1. Build a computational graph that describes the computation that you want to perform. This stage doesn't actually perform any computation; it just builds up a symbolic representation of your computation. This stage will typically define one or more `placeholder` objects that represent inputs to the computational graph.
# 2. Run the computational graph many times. Each time the graph is run you will specify which parts of the graph you want to compute, and pass a `feed_dict` dictionary that will give concrete values to any `placeholder`s in the graph.
#
# ### TensorFlow warmup: Flatten Function
#
# We can see this in action by defining a simple `flatten` function that will reshape image data for use in a fully-connected network.
#
# In TensorFlow, data for convolutional feature maps is typically stored in a Tensor of shape N x H x W x C where:
#
# - N is the number of datapoints (minibatch size)
# - H is the height of the feature map
# - W is the width of the feature map
# - C is the number of channels in the feature map
#
# This is the right way to represent the data when we are doing something like a 2D convolution, that needs spatial understanding of where the intermediate features are relative to each other. When we use fully connected affine layers to process the image, however, we want each datapoint to be represented by a single vector -- it's no longer useful to segregate the different channels, rows, and columns of the data. So, we use a "flatten" operation to collapse the `H x W x C` values per representation into a single long vector. The flatten function below first reads in the value of N from a given batch of data, and then returns a "view" of that data. "View" is analogous to numpy's "reshape" method: it reshapes x's dimensions to be N x ??, where ?? is allowed to be anything (in this case, it will be H x W x C, but we don't need to specify that explicitly).
#
# **NOTE**: TensorFlow and PyTorch differ on the default Tensor layout; TensorFlow uses N x H x W x C but PyTorch uses N x C x H x W.
def flatten(x):
"""
Input:
- TensorFlow Tensor of shape (N, D1, ..., DM)
Output:
- TensorFlow Tensor of shape (N, D1 * ... * DM)
"""
N = tf.shape(x)[0]
return tf.reshape(x, (N, -1))
def test_flatten():
# Clear the current TensorFlow graph.
tf.reset_default_graph()
# Stage I: Define the TensorFlow graph describing our computation.
# In this case the computation is trivial: we just want to flatten
# a Tensor using the flatten function defined above.
# Our computation will have a single input, x. We don't know its
# value yet, so we define a placeholder which will hold the value
# when the graph is run. We then pass this placeholder Tensor to
# the flatten function; this gives us a new Tensor which will hold
# a flattened view of x when the graph is run. The tf.device
# context manager tells TensorFlow whether to place these Tensors
# on CPU or GPU.
with tf.device(device):
x = tf.placeholder(tf.float32)
x_flat = flatten(x)
# At this point we have just built the graph describing our computation,
# but we haven't actually computed anything yet. If we print x and x_flat
# we see that they don't hold any data; they are just TensorFlow Tensors
# representing values that will be computed when the graph is run.
print('x: ', type(x), x)
print('x_flat: ', type(x_flat), x_flat)
print()
# We need to use a TensorFlow Session object to actually run the graph.
with tf.Session() as sess:
# Construct concrete values of the input data x using numpy
x_np = np.arange(24).reshape((2, 3, 4))
print('x_np:\n', x_np, '\n')
# Run our computational graph to compute a concrete output value.
# The first argument to sess.run tells TensorFlow which Tensor
# we want it to compute the value of; the feed_dict specifies
# values to plug into all placeholder nodes in the graph. The
# resulting value of x_flat is returned from sess.run as a
# numpy array.
x_flat_np = sess.run(x_flat, feed_dict={x: x_np})
print('x_flat_np:\n', x_flat_np, '\n')
# We can reuse the same graph to perform the same computation
# with different input data
x_np = np.arange(12).reshape((2, 3, 2))
print('x_np:\n', x_np, '\n')
x_flat_np = sess.run(x_flat, feed_dict={x: x_np})
print('x_flat_np:\n', x_flat_np)
test_flatten()
# ### Barebones TensorFlow: Two-Layer Network
# We will now implement our first neural network with TensorFlow: a fully-connected ReLU network with two hidden layers and no biases on the CIFAR10 dataset. For now we will use only low-level TensorFlow operators to define the network; later we will see how to use the higher-level abstractions provided by `tf.keras` to simplify the process.
#
# We will define the forward pass of the network in the function `two_layer_fc`; this will accept TensorFlow Tensors for the inputs and weights of the network, and return a TensorFlow Tensor for the scores. It's important to keep in mind that calling the `two_layer_fc` function **does not** perform any computation; instead it just sets up the computational graph for the forward computation. To actually run the network we need to enter a TensorFlow Session and feed data to the computational graph.
#
# After defining the network architecture in the `two_layer_fc` function, we will test the implementation by setting up and running a computational graph, feeding zeros to the network and checking the shape of the output.
#
# It's important that you read and understand this implementation.
def two_layer_fc(x, params):
"""
A fully-connected neural network; the architecture is:
fully-connected layer -> ReLU -> fully connected layer.
Note that we only need to define the forward pass here; TensorFlow will take
care of computing the gradients for us.
The input to the network will be a minibatch of data, of shape
(N, d1, ..., dM) where d1 * ... * dM = D. The hidden layer will have H units,
and the output layer will produce scores for C classes.
Inputs:
- x: A TensorFlow Tensor of shape (N, d1, ..., dM) giving a minibatch of
input data.
- params: A list [w1, w2] of TensorFlow Tensors giving weights for the
network, where w1 has shape (D, H) and w2 has shape (H, C).
Returns:
- scores: A TensorFlow Tensor of shape (N, C) giving classification scores
for the input data x.
"""
w1, w2 = params # Unpack the parameters
x = flatten(x) # Flatten the input; now x has shape (N, D)
h = tf.nn.relu(tf.matmul(x, w1)) # Hidden layer: h has shape (N, H)
scores = tf.matmul(h, w2) # Compute scores of shape (N, C)
return scores
# +
def two_layer_fc_test():
# TensorFlow's default computational graph is essentially a hidden global
# variable. To avoid adding to this default graph when you rerun this cell,
# we clear the default graph before constructing the graph we care about.
tf.reset_default_graph()
hidden_layer_size = 42
# Scoping our computational graph setup code under a tf.device context
# manager lets us tell TensorFlow where we want these Tensors to be
# placed.
with tf.device(device):
# Set up a placehoder for the input of the network, and constant
# zero Tensors for the network weights. Here we declare w1 and w2
# using tf.zeros instead of tf.placeholder as we've seen before - this
# means that the values of w1 and w2 will be stored in the computational
# graph itself and will persist across multiple runs of the graph; in
# particular this means that we don't have to pass values for w1 and w2
# using a feed_dict when we eventually run the graph.
x = tf.placeholder(tf.float32)
w1 = tf.zeros((32 * 32 * 3, hidden_layer_size))
w2 = tf.zeros((hidden_layer_size, 10))
# Call our two_layer_fc function to set up the computational
# graph for the forward pass of the network.
scores = two_layer_fc(x, [w1, w2])
# Use numpy to create some concrete data that we will pass to the
# computational graph for the x placeholder.
x_np = np.zeros((64, 32, 32, 3))
with tf.Session() as sess:
# The calls to tf.zeros above do not actually instantiate the values
# for w1 and w2; the following line tells TensorFlow to instantiate
# the values of all Tensors (like w1 and w2) that live in the graph.
sess.run(tf.global_variables_initializer())
# Here we actually run the graph, using the feed_dict to pass the
# value to bind to the placeholder for x; we ask TensorFlow to compute
# the value of the scores Tensor, which it returns as a numpy array.
scores_np = sess.run(scores, feed_dict={x: x_np})
print(scores_np.shape)
two_layer_fc_test()
# -
# ### Barebones TensorFlow: Three-Layer ConvNet
# Here you will complete the implementation of the function `three_layer_convnet` which will perform the forward pass of a three-layer convolutional network. The network should have the following architecture:
#
# 1. A convolutional layer (with bias) with `channel_1` filters, each with shape `KW1 x KH1`, and zero-padding of two
# 2. ReLU nonlinearity
# 3. A convolutional layer (with bias) with `channel_2` filters, each with shape `KW2 x KH2`, and zero-padding of one
# 4. ReLU nonlinearity
# 5. Fully-connected layer with bias, producing scores for `C` classes.
#
# **HINT**: For convolutions: https://www.tensorflow.org/api_docs/python/tf/nn/conv2d; be careful with padding!
#
# **HINT**: For biases: https://www.tensorflow.org/performance/xla/broadcasting
def three_layer_convnet(x, params):
"""
A three-layer convolutional network with the architecture described above.
Inputs:
- x: A TensorFlow Tensor of shape (N, H, W, 3) giving a minibatch of images
- params: A list of TensorFlow Tensors giving the weights and biases for the
network; should contain the following:
- conv_w1: TensorFlow Tensor of shape (KH1, KW1, 3, channel_1) giving
weights for the first convolutional layer.
- conv_b1: TensorFlow Tensor of shape (channel_1,) giving biases for the
first convolutional layer.
- conv_w2: TensorFlow Tensor of shape (KH2, KW2, channel_1, channel_2)
giving weights for the second convolutional layer
- conv_b2: TensorFlow Tensor of shape (channel_2,) giving biases for the
second convolutional layer.
- fc_w: TensorFlow Tensor giving weights for the fully-connected layer.
Can you figure out what the shape should be?
- fc_b: TensorFlow Tensor giving biases for the fully-connected layer.
Can you figure out what the shape should be?
"""
conv_w1, conv_b1, conv_w2, conv_b2, fc_w, fc_b = params
scores = None
############################################################################
# TODO: Implement the forward pass for the three-layer ConvNet. #
############################################################################
o = tf.nn.relu(tf.nn.conv2d(x, conv_w1, strides=(1,1,1,1),padding='SAME') + conv_b1)
o = tf.nn.relu(tf.nn.conv2d(o, conv_w2, strides=(1,1,1,1),padding='SAME') + conv_b2)
o = flatten(o)
scores = tf.matmul(o,fc_w) + fc_b
############################################################################
# END OF YOUR CODE #
############################################################################
return scores
# After defing the forward pass of the three-layer ConvNet above, run the following cell to test your implementation. Like the two-layer network, we use the `three_layer_convnet` function to set up the computational graph, then run the graph on a batch of zeros just to make sure the function doesn't crash, and produces outputs of the correct shape.
#
# When you run this function, `scores_np` should have shape `(64, 10)`.
# +
def three_layer_convnet_test():
tf.reset_default_graph()
with tf.device(device):
x = tf.placeholder(tf.float32)
conv_w1 = tf.zeros((5, 5, 3, 6))
conv_b1 = tf.zeros((6,))
conv_w2 = tf.zeros((3, 3, 6, 9))
conv_b2 = tf.zeros((9,))
fc_w = tf.zeros((32 * 32 * 9, 10))
fc_b = tf.zeros((10,))
params = [conv_w1, conv_b1, conv_w2, conv_b2, fc_w, fc_b]
scores = three_layer_convnet(x, params)
# Inputs to convolutional layers are 4-dimensional arrays with shape
# [batch_size, height, width, channels]
x_np = np.zeros((64, 32, 32, 3))
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
scores_np = sess.run(scores, feed_dict={x: x_np})
print('scores_np has shape: ', scores_np.shape)
with tf.device('/cpu:0'):
three_layer_convnet_test()
# -
# ### Barebones TensorFlow: Training Step
# We now define the `training_step` function which sets up the part of the computational graph that performs a single training step. This will take three basic steps:
#
# 1. Compute the loss
# 2. Compute the gradient of the loss with respect to all network weights
# 3. Make a weight update step using (stochastic) gradient descent.
#
# Note that the step of updating the weights is itself an operation in the computational graph - the calls to `tf.assign_sub` in `training_step` return TensorFlow operations that mutate the weights when they are executed. There is an important bit of subtlety here - when we call `sess.run`, TensorFlow does not execute all operations in the computational graph; it only executes the minimal subset of the graph necessary to compute the outputs that we ask TensorFlow to produce. As a result, naively computing the loss would not cause the weight update operations to execute, since the operations needed to compute the loss do not depend on the output of the weight update. To fix this problem, we insert a **control dependency** into the graph, adding a duplicate `loss` node to the graph that does depend on the outputs of the weight update operations; this is the object that we actually return from the `training_step` function. As a result, asking TensorFlow to evaluate the value of the `loss` returned from `training_step` will also implicitly update the weights of the network using that minibatch of data.
#
# We need to use a few new TensorFlow functions to do all of this:
# - For computing the cross-entropy loss we'll use `tf.nn.sparse_softmax_cross_entropy_with_logits`: https://www.tensorflow.org/api_docs/python/tf/nn/sparse_softmax_cross_entropy_with_logits
# - For averaging the loss across a minibatch of data we'll use `tf.reduce_mean`:
# https://www.tensorflow.org/api_docs/python/tf/reduce_mean
# - For computing gradients of the loss with respect to the weights we'll use `tf.gradients`: https://www.tensorflow.org/api_docs/python/tf/gradients
# - We'll mutate the weight values stored in a TensorFlow Tensor using `tf.assign_sub`: https://www.tensorflow.org/api_docs/python/tf/assign_sub
# - We'll add a control dependency to the graph using `tf.control_dependencies`: https://www.tensorflow.org/api_docs/python/tf/control_dependencies
def training_step(scores, y, params, learning_rate):
"""
Set up the part of the computational graph which makes a training step.
Inputs:
- scores: TensorFlow Tensor of shape (N, C) giving classification scores for
the model.
- y: TensorFlow Tensor of shape (N,) giving ground-truth labels for scores;
y[i] == c means that c is the correct class for scores[i].
- params: List of TensorFlow Tensors giving the weights of the model
- learning_rate: Python scalar giving the learning rate to use for gradient
descent step.
Returns:
- loss: A TensorFlow Tensor of shape () (scalar) giving the loss for this
batch of data; evaluating the loss also performs a gradient descent step
on params (see above).
"""
# First compute the loss; the first line gives losses for each example in
# the minibatch, and the second averages the losses acros the batch
losses = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=scores)
loss = tf.reduce_mean(losses)
# Compute the gradient of the loss with respect to each parameter of the the
# network. This is a very magical function call: TensorFlow internally
# traverses the computational graph starting at loss backward to each element
# of params, and uses backpropagation to figure out how to compute gradients;
# it then adds new operations to the computational graph which compute the
# requested gradients, and returns a list of TensorFlow Tensors that will
# contain the requested gradients when evaluated.
grad_params = tf.gradients(loss, params)
# Make a gradient descent step on all of the model parameters.
new_weights = []
for w, grad_w in zip(params, grad_params):
new_w = tf.assign_sub(w, learning_rate * grad_w)
new_weights.append(new_w)
# Insert a control dependency so that evaluting the loss causes a weight
# update to happen; see the discussion above.
with tf.control_dependencies(new_weights):
return tf.identity(loss)
# ### Barebones TensorFlow: Training Loop
# Now we set up a basic training loop using low-level TensorFlow operations. We will train the model using stochastic gradient descent without momentum. The `training_step` function sets up the part of the computational graph that performs the training step, and the function `train_part2` iterates through the training data, making training steps on each minibatch, and periodically evaluates accuracy on the validation set.
def train_part2(model_fn, init_fn, learning_rate):
"""
Train a model on CIFAR-10.
Inputs:
- model_fn: A Python function that performs the forward pass of the model
using TensorFlow; it should have the following signature:
scores = model_fn(x, params) where x is a TensorFlow Tensor giving a
minibatch of image data, params is a list of TensorFlow Tensors holding
the model weights, and scores is a TensorFlow Tensor of shape (N, C)
giving scores for all elements of x.
- init_fn: A Python function that initializes the parameters of the model.
It should have the signature params = init_fn() where params is a list
of TensorFlow Tensors holding the (randomly initialized) weights of the
model.
- learning_rate: Python float giving the learning rate to use for SGD.
"""
# First clear the default graph
tf.reset_default_graph()
is_training = tf.placeholder(tf.bool, name='is_training')
# Set up the computational graph for performing forward and backward passes,
# and weight updates.
with tf.device(device):
# Set up placeholders for the data and labels
x = tf.placeholder(tf.float32, [None, 32, 32, 3])
y = tf.placeholder(tf.int32, [None])
params = init_fn() # Initialize the model parameters
scores = model_fn(x, params) # Forward pass of the model
loss = training_step(scores, y, params, learning_rate)
# Now we actually run the graph many times using the training data
with tf.Session() as sess:
# Initialize variables that will live in the graph
sess.run(tf.global_variables_initializer())
for t, (x_np, y_np) in enumerate(train_dset):
# Run the graph on a batch of training data; recall that asking
# TensorFlow to evaluate loss will cause an SGD step to happen.
feed_dict = {x: x_np, y: y_np}
loss_np = sess.run(loss, feed_dict=feed_dict)
# Periodically print the loss and check accuracy on the val set
if t % print_every == 0:
print('Iteration %d, loss = %.4f' % (t, loss_np))
check_accuracy(sess, val_dset, x, scores, is_training)
# ### Barebones TensorFlow: Check Accuracy
# When training the model we will use the following function to check the accuracy of our model on the training or validation sets. Note that this function accepts a TensorFlow Session object as one of its arguments; this is needed since the function must actually run the computational graph many times on the data that it loads from the dataset `dset`.
#
# Also note that we reuse the same computational graph both for taking training steps and for evaluating the model; however since the `check_accuracy` function never evalutes the `loss` value in the computational graph, the part of the graph that updates the weights of the graph do not execute on the validation data.
def check_accuracy(sess, dset, x, scores, is_training=None):
"""
Check accuracy on a classification model.
Inputs:
- sess: A TensorFlow Session that will be used to run the graph
- dset: A Dataset object on which to check accuracy
- x: A TensorFlow placeholder Tensor where input images should be fed
- scores: A TensorFlow Tensor representing the scores output from the
model; this is the Tensor we will ask TensorFlow to evaluate.
Returns: Nothing, but prints the accuracy of the model
"""
num_correct, num_samples = 0, 0
for x_batch, y_batch in dset:
feed_dict = {x: x_batch, is_training: 0}
scores_np = sess.run(scores, feed_dict=feed_dict)
y_pred = scores_np.argmax(axis=1)
num_samples += x_batch.shape[0]
num_correct += (y_pred == y_batch).sum()
acc = float(num_correct) / num_samples
print('Got %d / %d correct (%.2f%%)' % (num_correct, num_samples, 100 * acc))
# ### Barebones TensorFlow: Initialization
# We'll use the following utility method to initialize the weight matrices for our models using Kaiming's normalization method.
#
# [1] He et al, *Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification
# *, ICCV 2015, https://arxiv.org/abs/1502.01852
def kaiming_normal(shape):
if len(shape) == 2:
fan_in, fan_out = shape[0], shape[1]
elif len(shape) == 4:
fan_in, fan_out = np.prod(shape[:3]), shape[3]
return tf.random_normal(shape) * np.sqrt(2.0 / fan_in)
# ### Barebones TensorFlow: Train a Two-Layer Network
# We are finally ready to use all of the pieces defined above to train a two-layer fully-connected network on CIFAR-10.
#
# We just need to define a function to initialize the weights of the model, and call `train_part2`.
#
# Defining the weights of the network introduces another important piece of TensorFlow API: `tf.Variable`. A TensorFlow Variable is a Tensor whose value is stored in the graph and persists across runs of the computational graph; however unlike constants defined with `tf.zeros` or `tf.random_normal`, the values of a Variable can be mutated as the graph runs; these mutations will persist across graph runs. Learnable parameters of the network are usually stored in Variables.
#
# You don't need to tune any hyperparameters, but you should achieve accuracies above 40% after one epoch of training.
# +
def two_layer_fc_init():
"""
Initialize the weights of a two-layer network, for use with the
two_layer_network function defined above.
Inputs: None
Returns: A list of:
- w1: TensorFlow Variable giving the weights for the first layer
- w2: TensorFlow Variable giving the weights for the second layer
"""
hidden_layer_size = 4000
w1 = tf.Variable(kaiming_normal((3 * 32 * 32, 4000)))
w2 = tf.Variable(kaiming_normal((4000, 10)))
return [w1, w2]
learning_rate = 1e-2
train_part2(two_layer_fc, two_layer_fc_init, learning_rate)
# -
# ### Barebones TensorFlow: Train a three-layer ConvNet
# We will now use TensorFlow to train a three-layer ConvNet on CIFAR-10.
#
# You need to implement the `three_layer_convnet_init` function. Recall that the architecture of the network is:
#
# 1. Convolutional layer (with bias) with 32 5x5 filters, with zero-padding 2
# 2. ReLU
# 3. Convolutional layer (with bias) with 16 3x3 filters, with zero-padding 1
# 4. ReLU
# 5. Fully-connected layer (with bias) to compute scores for 10 classes
#
# You don't need to do any hyperparameter tuning, but you should see accuracies above 43% after one epoch of training.
# +
def three_layer_convnet_init():
"""
Initialize the weights of a Three-Layer ConvNet, for use with the
three_layer_convnet function defined above.
Inputs: None
Returns a list containing:
- conv_w1: TensorFlow Variable giving weights for the first conv layer
- conv_b1: TensorFlow Variable giving biases for the first conv layer
- conv_w2: TensorFlow Variable giving weights for the second conv layer
- conv_b2: TensorFlow Variable giving biases for the second conv layer
- fc_w: TensorFlow Variable giving weights for the fully-connected layer
- fc_b: TensorFlow Variable giving biases for the fully-connected layer
"""
params = None
############################################################################
# TODO: Initialize the parameters of the three-layer network. #
############################################################################
conv_w1 = tf.Variable(kaiming_normal((5,5,3,32)))
conv_b1 = tf.Variable(tf.zeros(32))
conv_w2 = tf.Variable(kaiming_normal((3,3,32,16)))
conv_b2 = tf.Variable(tf.zeros(16))
fc_w = tf.Variable(kaiming_normal((16*32*32,10)))
fc_b = tf.Variable(tf.zeros(10))
params = conv_w1, conv_b1, conv_w2, conv_b2, fc_w, fc_b
############################################################################
# END OF YOUR CODE #
############################################################################
return params
learning_rate = 3e-3
train_part2(three_layer_convnet, three_layer_convnet_init, learning_rate)
# -
# # Part III: Keras Model API
# Implementing a neural network using the low-level TensorFlow API is a good way to understand how TensorFlow works, but it's a little inconvenient - we had to manually keep track of all Tensors holding learnable parameters, and we had to use a control dependency to implement the gradient descent update step. This was fine for a small network, but could quickly become unweildy for a large complex model.
#
# Fortunately TensorFlow provides higher-level packages such as `tf.keras` and `tf.layers` which make it easy to build models out of modular, object-oriented layers; `tf.train` allows you to easily train these models using a variety of different optimization algorithms.
#
# In this part of the notebook we will define neural network models using the `tf.keras.Model` API. To implement your own model, you need to do the following:
#
# 1. Define a new class which subclasses `tf.keras.model`. Give your class an intuitive name that describes it, like `TwoLayerFC` or `ThreeLayerConvNet`.
# 2. In the initializer `__init__()` for your new class, define all the layers you need as class attributes. The `tf.layers` package provides many common neural-network layers, like `tf.layers.Dense` for fully-connected layers and `tf.layers.Conv2D` for convolutional layers. Under the hood, these layers will construct `Variable` Tensors for any learnable parameters. **Warning**: Don't forget to call `super().__init__()` as the first line in your initializer!
# 3. Implement the `call()` method for your class; this implements the forward pass of your model, and defines the *connectivity* of your network. Layers defined in `__init__()` implement `__call__()` so they can be used as function objects that transform input Tensors into output Tensors. Don't define any new layers in `call()`; any layers you want to use in the forward pass should be defined in `__init__()`.
#
# After you define your `tf.keras.Model` subclass, you can instantiate it and use it like the model functions from Part II.
#
# ### Module API: Two-Layer Network
#
# Here is a concrete example of using the `tf.keras.Model` API to define a two-layer network. There are a few new bits of API to be aware of here:
#
# We use an `Initializer` object to set up the initial values of the learnable parameters of the layers; in particular `tf.variance_scaling_initializer` gives behavior similar to the Kaiming initialization method we used in Part II. You can read more about it here: https://www.tensorflow.org/api_docs/python/tf/variance_scaling_initializer
#
# We construct `tf.layers.Dense` objects to represent the two fully-connected layers of the model. In addition to multiplying their input by a weight matrix and adding a bias vector, these layer can also apply a nonlinearity for you. For the first layer we specify a ReLU activation function by passing `activation=tf.nn.relu` to the constructor; the second layer does not apply any activation function.
#
# Unfortunately the `flatten` function we defined in Part II is not compatible with the `tf.keras.Model` API; fortunately we can use `tf.layers.flatten` to perform the same operation. The issue with our `flatten` function from Part II has to do with static vs dynamic shapes for Tensors, which is beyond the scope of this notebook; you can read more about the distinction [in the documentation](https://www.tensorflow.org/programmers_guide/faq#tensor_shapes).
# +
class TwoLayerFC(tf.keras.Model):
def __init__(self, hidden_size, num_classes):
super().__init__()
initializer = tf.variance_scaling_initializer(scale=2.0)
self.fc1 = tf.layers.Dense(hidden_size, activation=tf.nn.relu,
kernel_initializer=initializer)
self.fc2 = tf.layers.Dense(num_classes,
kernel_initializer=initializer)
def call(self, x, training=None):
x = tf.layers.flatten(x)
x = self.fc1(x)
x = self.fc2(x)
return x
def test_TwoLayerFC():
""" A small unit test to exercise the TwoLayerFC model above. """
tf.reset_default_graph()
input_size, hidden_size, num_classes = 50, 42, 10
# As usual in TensorFlow, we first need to define our computational graph.
# To this end we first construct a TwoLayerFC object, then use it to construct
# the scores Tensor.
model = TwoLayerFC(hidden_size, num_classes)
with tf.device(device):
x = tf.zeros((64, input_size))
scores = model(x)
# Now that our computational graph has been defined we can run the graph
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
scores_np = sess.run(scores)
print(scores_np.shape)
test_TwoLayerFC()
# -
# ### Funtional API: Two-Layer Network
# The `tf.layers` package provides two different higher-level APIs for defining neural network models. In the example above we used the **object-oriented API**, where each layer of the neural network is represented as a Python object (like `tf.layers.Dense`). Here we showcase the **functional API**, where each layer is a Python function (like `tf.layers.dense`) which inputs and outputs TensorFlow Tensors, and which internally sets up Tensors in the computational graph to hold any learnable weights.
#
# To construct a network, one needs to pass the input tensor to the first layer, and construct the subsequent layers sequentially. Here's an example of how to construct the same two-layer nework with the functional API.
# +
def two_layer_fc_functional(inputs, hidden_size, num_classes):
initializer = tf.variance_scaling_initializer(scale=2.0)
flattened_inputs = tf.layers.flatten(inputs)
fc1_output = tf.layers.dense(flattened_inputs, hidden_size, activation=tf.nn.relu,
kernel_initializer=initializer)
scores = tf.layers.dense(fc1_output, num_classes,
kernel_initializer=initializer)
return scores
def test_two_layer_fc_functional():
""" A small unit test to exercise the TwoLayerFC model above. """
tf.reset_default_graph()
input_size, hidden_size, num_classes = 50, 42, 10
# As usual in TensorFlow, we first need to define our computational graph.
# To this end we first construct a two layer network graph by calling the
# two_layer_network() function. This function constructs the computation
# graph and outputs the score tensor.
with tf.device(device):
x = tf.zeros((64, input_size))
scores = two_layer_fc_functional(x, hidden_size, num_classes)
# Now that our computational graph has been defined we can run the graph
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
scores_np = sess.run(scores)
print(scores_np.shape)
test_two_layer_fc_functional()
# -
# ### Keras Model API: Three-Layer ConvNet
# Now it's your turn to implement a three-layer ConvNet using the `tf.keras.Model` API. Your model should have the same architecture used in Part II:
#
# 1. Convolutional layer with 5 x 5 kernels, with zero-padding of 2
# 2. ReLU nonlinearity
# 3. Convolutional layer with 3 x 3 kernels, with zero-padding of 1
# 4. ReLU nonlinearity
# 5. Fully-connected layer to give class scores
#
# You should initialize the weights of your network using the same initialization method as was used in the two-layer network above.
#
# **Hint**: Refer to the documentation for `tf.layers.Conv2D` and `tf.layers.Dense`:
#
# https://www.tensorflow.org/api_docs/python/tf/layers/Conv2D
#
# https://www.tensorflow.org/api_docs/python/tf/layers/Dense
class ThreeLayerConvNet(tf.keras.Model):
def __init__(self, channel_1, channel_2, num_classes):
super().__init__()
########################################################################
# TODO: Implement the __init__ method for a three-layer ConvNet. You #
# should instantiate layer objects to be used in the forward pass. #
########################################################################
initializer = tf.variance_scaling_initializer(scale=2.0)
self.conv1 = tf.layers.Conv2D(channel_1, 5, padding='same', activation=tf.nn.relu, kernel_initializer=initializer)
self.conv2 = tf.layers.Conv2D(channel_2, 3, padding='same', activation=tf.nn.relu, kernel_initializer=initializer)
self.fc = tf.layers.Dense(num_classes)
########################################################################
# END OF YOUR CODE #
########################################################################
def call(self, x, training=None):
scores = None
########################################################################
# TODO: Implement the forward pass for a three-layer ConvNet. You #
# should use the layer objects defined in the __init__ method. #
########################################################################
o = self.conv1(x)
o = self.conv2(o)
o = tf.layers.flatten(o)
scores = self.fc(o)
########################################################################
# END OF YOUR CODE #
########################################################################
return scores
# Once you complete the implementation of the `ThreeLayerConvNet` above you can run the following to ensure that your implementation does not crash and produces outputs of the expected shape.
# +
def test_ThreeLayerConvNet():
tf.reset_default_graph()
channel_1, channel_2, num_classes = 12, 8, 10
model = ThreeLayerConvNet(channel_1, channel_2, num_classes)
with tf.device(device):
x = tf.zeros((64, 3, 32, 32))
scores = model(x)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
scores_np = sess.run(scores)
print(scores_np.shape)
test_ThreeLayerConvNet()
# -
# ### Keras Model API: Training Loop
# We need to implement a slightly different training loop when using the `tf.keras.Model` API. Instead of computing gradients and updating the weights of the model manually, we use an `Optimizer` object from the `tf.train` package which takes care of these details for us. You can read more about `Optimizer`s here: https://www.tensorflow.org/api_docs/python/tf/train/Optimizer
def train_part34(model_init_fn, optimizer_init_fn, num_epochs=1):
"""
Simple training loop for use with models defined using tf.keras. It trains
a model for one epoch on the CIFAR-10 training set and periodically checks
accuracy on the CIFAR-10 validation set.
Inputs:
- model_init_fn: A function that takes no parameters; when called it
constructs the model we want to train: model = model_init_fn()
- optimizer_init_fn: A function which takes no parameters; when called it
constructs the Optimizer object we will use to optimize the model:
optimizer = optimizer_init_fn()
- num_epochs: The number of epochs to train for
Returns: Nothing, but prints progress during trainingn
"""
tf.reset_default_graph()
with tf.device(device):
# Construct the computational graph we will use to train the model. We
# use the model_init_fn to construct the model, declare placeholders for
# the data and labels
x = tf.placeholder(tf.float32, [None, 32, 32, 3])
y = tf.placeholder(tf.int32, [None])
# We need a place holder to explicitly specify if the model is in the training
# phase or not. This is because a number of layers behaves differently in
# training and in testing, e.g., dropout and batch normalization.
# We pass this variable to the computation graph through feed_dict as shown below.
is_training = tf.placeholder(tf.bool, name='is_training')
# Use the model function to build the forward pass.
scores = model_init_fn(x, is_training)
# Compute the loss like we did in Part II
loss = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=scores)
loss = tf.reduce_mean(loss)
# Use the optimizer_fn to construct an Optimizer, then use the optimizer
# to set up the training step. Asking TensorFlow to evaluate the
# train_op returned by optimizer.minimize(loss) will cause us to make a
# single update step using the current minibatch of data.
# Note that we use tf.control_dependencies to force the model to run
# the tf.GraphKeys.UPDATE_OPS at each training step. tf.GraphKeys.UPDATE_OPS
# holds the operators that update the states of the network.
# For example, the tf.layers.batch_normalization function adds the running mean
# and variance update operators to tf.GraphKeys.UPDATE_OPS.
optimizer = optimizer_init_fn()
update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(update_ops):
train_op = optimizer.minimize(loss)
# Now we can run the computational graph many times to train the model.
# When we call sess.run we ask it to evaluate train_op, which causes the
# model to update.
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
t = 0
for epoch in range(num_epochs):
print('Starting epoch %d' % epoch)
for x_np, y_np in train_dset:
feed_dict = {x: x_np, y: y_np, is_training:1}
loss_np, _ = sess.run([loss, train_op], feed_dict=feed_dict)
if t % print_every == 0:
print('Iteration %d, loss = %.4f' % (t, loss_np))
check_accuracy(sess, val_dset, x, scores, is_training=is_training)
print()
t += 1
# ### Keras Model API: Train a Two-Layer Network
# We can now use the tools defined above to train a two-layer network on CIFAR-10. We define the `model_init_fn` and `optimizer_init_fn` that construct the model and optimizer respectively when called. Here we want to train the model using stochastic gradient descent with no momentum, so we construct a `tf.train.GradientDescentOptimizer` function; you can [read about it here](https://www.tensorflow.org/api_docs/python/tf/train/GradientDescentOptimizer).
#
# You don't need to tune any hyperparameters here, but you should achieve accuracies above 40% after one epoch of training.
# +
hidden_size, num_classes = 4000, 10
learning_rate = 1e-2
def model_init_fn(inputs, is_training):
return TwoLayerFC(hidden_size, num_classes)(inputs)
def optimizer_init_fn():
return tf.train.GradientDescentOptimizer(learning_rate)
train_part34(model_init_fn, optimizer_init_fn)
# -
# ### Keras Model API: Train a Two-Layer Network (functional API)
# Similarly, we train the two-layer network constructed using the functional API.
# +
hidden_size, num_classes = 4000, 10
learning_rate = 1e-2
def model_init_fn(inputs, is_training):
return two_layer_fc_functional(inputs, hidden_size, num_classes)
def optimizer_init_fn():
return tf.train.GradientDescentOptimizer(learning_rate)
train_part34(model_init_fn, optimizer_init_fn)
# -
# ### Keras Model API: Train a Three-Layer ConvNet
# Here you should use the tools we've defined above to train a three-layer ConvNet on CIFAR-10. Your ConvNet should use 32 filters in the first convolutional layer and 16 filters in the second layer.
#
# To train the model you should use gradient descent with Nesterov momentum 0.9.
#
# **HINT**: https://www.tensorflow.org/api_docs/python/tf/train/MomentumOptimizer
#
# You don't need to perform any hyperparameter tuning, but you should achieve accuracies above 45% after training for one epoch.
# +
learning_rate = 3e-3
channel_1, channel_2, num_classes = 32, 16, 10
def model_init_fn(inputs, is_training):
model = None
############################################################################
# TODO: Complete the implementation of model_fn. #
############################################################################
model = ThreeLayerConvNet(channel_1=channel_1, channel_2=channel_2,num_classes=num_classes)
########################t####################################################
# END OF YOUR CODE #
############################################################################
return model(inputs, is_training)
def optimizer_init_fn():
optimizer = None
############################################################################
# TODO: Complete the implementation of model_fn. #
############################################################################
optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
############################################################################
# END OF YOUR CODE #
############################################################################
return optimizer
train_part34(model_init_fn, optimizer_init_fn)
# -
# # Part IV: Keras Sequential API
# In Part III we introduced the `tf.keras.Model` API, which allows you to define models with any number of learnable layers and with arbitrary connectivity between layers.
#
# However for many models you don't need such flexibility - a lot of models can be expressed as a sequential stack of layers, with the output of each layer fed to the next layer as input. If your model fits this pattern, then there is an even easier way to define your model: using `tf.keras.Sequential`. You don't need to write any custom classes; you simply call the `tf.keras.Sequential` constructor with a list containing a sequence of layer objects.
#
# One complication with `tf.keras.Sequential` is that you must define the shape of the input to the model by passing a value to the `input_shape` of the first layer in your model.
#
# ### Keras Sequential API: Two-Layer Network
# Here we rewrite the two-layer fully-connected network using `tf.keras.Sequential`, and train it using the training loop defined above.
#
# You don't need to perform any hyperparameter tuning here, but you should see accuracies above 40% after training for one epoch.
# +
learning_rate = 1e-2
def model_init_fn(inputs, is_training):
input_shape = (32, 32, 3)
hidden_layer_size, num_classes = 4000, 10
initializer = tf.variance_scaling_initializer(scale=2.0)
layers = [
tf.layers.Flatten(input_shape=input_shape),
tf.layers.Dense(hidden_layer_size, activation=tf.nn.relu,
kernel_initializer=initializer),
tf.layers.Dense(num_classes, kernel_initializer=initializer),
]
model = tf.keras.Sequential(layers)
return model(inputs)
def optimizer_init_fn():
return tf.train.GradientDescentOptimizer(learning_rate)
train_part34(model_init_fn, optimizer_init_fn)
# -
# ### Keras Sequential API: Three-Layer ConvNet
# Here you should use `tf.keras.Sequential` to reimplement the same three-layer ConvNet architecture used in Part II and Part III. As a reminder, your model should have the following architecture:
#
# 1. Convolutional layer with 16 5x5 kernels, using zero padding of 2
# 2. ReLU nonlinearity
# 3. Convolutional layer with 32 3x3 kernels, using zero padding of 1
# 4. ReLU nonlinearity
# 5. Fully-connected layer giving class scores
#
# You should initialize the weights of the model using a `tf.variance_scaling_initializer` as above.
#
# You should train the model using Nesterov momentum 0.9.
#
# You don't need to perform any hyperparameter search, but you should achieve accuracy above 45% after training for one epoch.
# +
def model_init_fn(inputs, is_training):
model = None
############################################################################
# TODO: Construct a three-layer ConvNet using tf.keras.Sequential. #
############################################################################
input_shape = (32, 32, 3)
initializer = tf.variance_scaling_initializer(scale=2.0)
model = tf.keras.Sequential([tf.layers.Conv2D(filters=32, kernel_size=5, padding='same', activation=tf.nn.relu, kernel_initializer=initializer, input_shape=input_shape),
tf.layers.Conv2D(filters=16, kernel_size=3, padding='same', activation=tf.nn.relu, kernel_initializer=initializer),
tf.layers.Flatten(),
tf.layers.Dense(10, kernel_initializer=initializer)])
############################################################################
# END OF YOUR CODE #
############################################################################
return model(inputs)
learning_rate = 5e-4
def optimizer_init_fn():
optimizer = None
############################################################################
# TODO: Complete the implementation of model_fn. #
############################################################################
optimizer = tf.train.MomentumOptimizer(learning_rate=learning_rate, momentum=0.9)
############################################################################
# END OF YOUR CODE #
############################################################################
return optimizer
train_part34(model_init_fn, optimizer_init_fn)
# -
# # Part V: CIFAR-10 open-ended challenge
#
# In this section you can experiment with whatever ConvNet architecture you'd like on CIFAR-10.
#
# You should experiment with architectures, hyperparameters, loss functions, regularization, or anything else you can think of to train a model that achieves **at least 70%** accuracy on the **validation** set within 10 epochs. You can use the `check_accuracy` and `train` functions from above, or you can implement your own training loop.
#
# Describe what you did at the end of the notebook.
#
# ### Some things you can try:
# - **Filter size**: Above we used 5x5 and 3x3; is this optimal?
# - **Number of filters**: Above we used 16 and 32 filters. Would more or fewer do better?
# - **Pooling**: We didn't use any pooling above. Would this improve the model?
# - **Normalization**: Would your model be improved with batch normalization, layer normalization, group normalization, or some other normalization strategy?
# - **Network architecture**: The ConvNet above has only three layers of trainable parameters. Would a deeper model do better?
# - **Global average pooling**: Instead of flattening after the final convolutional layer, would global average pooling do better? This strategy is used for example in Google's Inception network and in Residual Networks.
# - **Regularization**: Would some kind of regularization improve performance? Maybe weight decay or dropout?
#
# ### WARNING: Batch Normalization / Dropout
# Batch Normalization and Dropout **WILL NOT WORK CORRECTLY** if you use the `train_part34()` function with the object-oriented `tf.keras.Model` or `tf.keras.Sequential` APIs; if you want to use these layers with this training loop then you **must use the tf.layers functional API**.
#
# We wrote `train_part34()` to explicitly demonstrate how TensorFlow works; however there are some subtleties that make it tough to handle the object-oriented batch normalization layer in a simple training loop. In practice both `tf.keras` and `tf` provide higher-level APIs which handle the training loop for you, such as [keras.fit](https://keras.io/models/sequential/) and [tf.Estimator](https://www.tensorflow.org/programmers_guide/estimators), both of which will properly handle batch normalization when using the object-oriented API.
#
# ### Tips for training
# For each network architecture that you try, you should tune the learning rate and other hyperparameters. When doing this there are a couple important things to keep in mind:
#
# - If the parameters are working well, you should see improvement within a few hundred iterations
# - Remember the coarse-to-fine approach for hyperparameter tuning: start by testing a large range of hyperparameters for just a few training iterations to find the combinations of parameters that are working at all.
# - Once you have found some sets of parameters that seem to work, search more finely around these parameters. You may need to train for more epochs.
# - You should use the validation set for hyperparameter search, and save your test set for evaluating your architecture on the best parameters as selected by the validation set.
#
# ### Going above and beyond
# If you are feeling adventurous there are many other features you can implement to try and improve your performance. You are **not required** to implement any of these, but don't miss the fun if you have time!
#
# - Alternative optimizers: you can try Adam, Adagrad, RMSprop, etc.
# - Alternative activation functions such as leaky ReLU, parametric ReLU, ELU, or MaxOut.
# - Model ensembles
# - Data augmentation
# - New Architectures
# - [ResNets](https://arxiv.org/abs/1512.03385) where the input from the previous layer is added to the output.
# - [DenseNets](https://arxiv.org/abs/1608.06993) where inputs into previous layers are concatenated together.
# - [This blog has an in-depth overview](https://chatbotslife.com/resnets-highwaynets-and-densenets-oh-my-9bb15918ee32)
#
# ### Have fun and happy training!
# +
import tensorflow.contrib.slim as slim
class ResUnit(tf.keras.Model):
def __init__(self):
super().__init__()
initializer=tf.variance_scaling_initializer(scale=2.0)
self.forward_unit = tf.keras.Sequential([tf.keras.layers.BatchNormalization(),
tf.keras.layers.ReLU(),
tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding='same', kernel_initializer=initializer),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.ReLU(),
tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding='same', kernel_initializer=initializer)])
def call(self, x, training=False):
f=self.forward_unit(x)
return f+x
total_layers = 20 #Specify how deep we want our network
resne_units = 4
units_between_stride = total_layers // resne_units
class SimpleResNet(tf.keras.Model):
def __init__(self, input_shape):
super().__init__()
initializer=tf.variance_scaling_initializer(scale=2.0)
self.conv1 = tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding='same', kernel_initializer=initializer, input_shape=input_shape)
self.res_layers=[]
# input_shape=[32,32,64]
for i in range(resne_units):
for j in range(units_between_stride):
self.res_layers.append(ResUnit())
self.res_layers.append(tf.keras.layers.Conv2D(filters=64, kernel_size=3, strides=2, kernel_initializer=initializer))
# input_shape[0],input_shape[1] = input_shape[0]//2,input_shape[1]//2
self.fc = tf.keras.layers.Dense(units=10, activation=None, kernel_initializer=initializer)
# self.top_conv = tf.keras.layers.Conv2D(filters=10, kernel_size=3, kernel_initializer=initializer, padding='same')
# self.flatten = tf.keras.layers.Flatten()
def call(self, x, training=False):
f = self.conv1(x)
for l in self.res_layers:
f=l(f)
f=tf.layers.flatten(f)
scores=self.fc(f)
# print(scores.shape)
# scores=tf.layers.flatten(f)
return scores
initializer=tf.variance_scaling_initializer(scale=2.0)
def bn_conv_maxpool_layer(out_channels=64, conv_kernel_size=3, conv_stride=1, pool_kernel_size=3,pool_stride=2):
padding=(conv_kernel_size-conv_stride)/2
return tf.keras.Sequential([tf.keras.layers.Conv2D(filters=out_channels, kernel_size=conv_kernel_size, strides=conv_stride, padding='same', kernel_initializer=initializer),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.ReLU(),
tf.keras.layers.MaxPool2D(pool_size=pool_kernel_size, strides=pool_stride)])
def model_init_fn(inputs, is_training):
model = None
############################################################################
# TODO: Construct a model that performs well on CIFAR-10 #
############################################################################
input_shape = [32,32,3]
model = tf.keras.Sequential([tf.keras.layers.Conv2D(filters=64, kernel_size=3, padding='same', input_shape=input_shape),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.ReLU(),
tf.keras.layers.MaxPool2D(pool_size=2, strides=2), # 16 x 16
bn_conv_maxpool_layer(out_channels=128), # 8 x 8
bn_conv_maxpool_layer(out_channels=256), # 4 x 4
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(2048,kernel_initializer=initializer),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(10, kernel_initializer=initializer)
])
net = model(inputs, is_training)
############################################################################
# END OF YOUR CODE #
############################################################################
return net
def optimizer_init_fn():
optimizer = None
############################################################################
# TODO: Construct an optimizer that performs well on CIFAR-10 #
############################################################################
optimizer = tf.train.AdamOptimizer()
############################################################################
# END OF YOUR CODE #
############################################################################
return optimizer
device = '/gpu:0'
print_every = 700
num_epochs = 10
train_part34(model_init_fn, optimizer_init_fn, num_epochs)
# -
# ## Describe what you did
#
# In the cell below you should write an explanation of what you did, any additional features that you implemented, and/or any graphs that you made in the process of training and evaluating your network.
# TODO: Tell us what you did
| spring1718_assignment2_v2/TensorFlow.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # Table
#
# These tables are from the [RST specification](http://docutils.sourceforge.net/docs/ref/rst/restructuredtext.html#grid-tables):
# ## Grid Tables
#
# A simple rst table with header
#
# |C1|C2|
# |:----:|:----:|
# |a|b|
# |c|d|
# **Note:** Tables without a header are currently not supported as markdown does
# not support tables without headers.
# ## Simple Tables
#
# |A|B|A and B|
# |:---:|:---:|:-----:|
# |False|False|False|
# |True|False|False|
# |False|True|False|
# |True|True|True|
# ## Directive Table Types
#
# These table types are provided by [sphinx docs](http://www.sphinx-doc.org/en/master/rest.html#directives)
# ### List Table directive
# ### Frozen Delights!
# |Treat|Quantity|Description|
# |:-------------:|:--------:|:----------------------------:|
# |Albatross|2.99|On a stick!|
# |Crunchy Frog|1.49|If we took the bones out, it wouldn’t be crunchy, now would it?|
# |Gannet Ripple|1.99|On a stick!|
# ### CSV Table Directive
# ### Frozen Delights!
# |Treat|Quantity|Description|
# |:-------------:|:--------:|:----------------------------:|
# |Albatross|2.99|On a stick!|
# |Crunchy Frog|1.49|If we took the bones out, it wouldn’t be crunchy, now would it?|
# |<NAME>|1.99|On a stick!|
# ## Complex Tables
#
# **MultiColumn and MultiRow** tables are currently **not** supported by this extension
| tests/base/ipynb/tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Module 4 - More Control Flow Tools
#
# Besides the while statement just introduced, Python knows the usual control flow statements known from other languages, with some twists.
# + [markdown] slideshow={"slide_type": "slide"}
# # `if` Statements
#
# Perhaps the most well-known statement type is the if statement. For example:
# + slideshow={"slide_type": "fragment"}
from __future__ import print_function
x = int(input("Please enter an integer: "))
if x < 0:
x = 0
print('Negative changed to zero')
elif x == 0:
print('Zero')
elif x == 1:
print('Single')
else:
print('More')
# + [markdown] slideshow={"slide_type": "subslide"}
# There can be zero or more `elif` parts, and the else part is optional.
#
# The keyword `elif` is short for `else if`, and is useful to avoid excessive indentation.
#
# An `if ... elif ... elif ...` sequence is a substitute for the switch or case statements found in other languages.
# + [markdown] slideshow={"slide_type": "slide"}
# # `for` Statements
#
# The for statement in Python differs a bit from what you may be used to in C or Pascal.
#
# Rather than
#
# - always iterating over an arithmetic progression of numbers (like in Pascal),
# - or giving the user the ability to define both the iteration step and halting condition (as C)
#
# Python’s for statement iterates over the items of any sequence (a list or a string), in the order that they appear in the sequence.
#
# For example (no pun intended):
# + slideshow={"slide_type": "fragment"}
# Measure some strings:
words = ['cat', 'window', 'defenestrate']
for w in words:
print(w, len(w))
# + [markdown] slideshow={"slide_type": "slide"}
# If you need to modify the sequence you are iterating over while inside the loop (for example to duplicate selected items), it is recommended that you first make a copy.
#
# Iterating over a sequence does not implicitly make a copy. The slice notation makes this especially convenient:
# + slideshow={"slide_type": "fragment"}
for w in words[:]: # Loop over a slice copy of the entire list.
if len(w) > 6:
words.insert(0, w)
words
# + [markdown] slideshow={"slide_type": "slide"}
# # The `range()` Function
#
# If you do need to iterate over a sequence of numbers, the built-in function range() comes in handy.
#
# It creates generators containing arithmetic progressions:
# + slideshow={"slide_type": "fragment"}
list(range(10))
# + [markdown] slideshow={"slide_type": "slide"}
# The given end point is never part of the generated list; `range(10)` generates a list of 10 values, the legal indices for items of a sequence of length 10.
#
# It is possible to let the range start at another number, or to specify a different increment (even negative; sometimes this is called the ‘step’):
#
# + slideshow={"slide_type": "fragment"}
list(range(5, 10))
# + slideshow={"slide_type": "fragment"}
list(range(0, 10, 3))
# + slideshow={"slide_type": "fragment"}
list(range(-10, -100, -30))
# + [markdown] slideshow={"slide_type": "slide"}
# To iterate over the indices of a sequence, you can combine `range()` and `len()` as follows:
# + slideshow={"slide_type": "fragment"}
a = ['Mary', 'had', 'a', 'little', 'lamb']
for i in range(len(a)):
print(i, a[i])
# + [markdown] slideshow={"slide_type": "fragment"}
# In most such cases, however, it is convenient to use the `enumerate()` function, see Looping Techniques.
# + [markdown] slideshow={"slide_type": "slide"}
# # `break` and `continue` Statements, and `else` Clauses on Loops
#
# The break statement, like in C, breaks out of the smallest enclosing for or while loop.
#
# Loop statements may have an `else` clause:
# - it is executed when the loop terminates through exhaustion of the list (with `for`)
# - or when the condition becomes false (with `while`)
# - but not when the loop is terminated by a `break` statement.
#
# This is exemplified by the following loop, which searches for prime numbers:
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Prime Number Search
# + slideshow={"slide_type": "fragment"}
for n in range(2, 10):
for x in range(2, n):
if n % x == 0:
print(n, 'equals', x, '*', n/x)
break
else:
# loop fell through without finding a factor
print(n, 'is a prime number')
# + [markdown] slideshow={"slide_type": "fragment"}
# (Yes, this is the correct code. Look closely: the else clause belongs to the for loop, not the if statement.)
# + [markdown] slideshow={"slide_type": "slide"}
# When used with a loop, the `else` clause has more in common with the `else` clause of a `try` statement than it does that of `if` statements:
# - a `try` statement’s else clause runs when no exception occur
# - a loop’s else clause runs when no break occurs.
#
# For more on the `try` statement and exceptions, see Handling Exceptions.
#
# The `continue` statement, also borrowed from C, continues with the next iteration of the loop:
# + slideshow={"slide_type": "fragment"}
for num in range(2, 10):
if num % 2 == 0:
print("Found an even number", num)
continue
print("Found a number", num)
# + [markdown] slideshow={"slide_type": "slide"}
# # `pass` Statements
#
# The pass statement does nothing.
#
# It can be used when a statement is required syntactically but the program requires no action.
#
# For example:
# + slideshow={"slide_type": "fragment"}
# while True:
# pass # Busy-wait for keyboard interrupt (Ctrl+C)
# + [markdown] slideshow={"slide_type": "slide"}
# This is commonly used for creating minimal classes:
# + slideshow={"slide_type": "fragment"}
class MyEmptyClass:
pass
# + [markdown] slideshow={"slide_type": "fragment"}
# Another place pass can be used is as a place-holder for a function or conditional body when you are working on new code, allowing you to keep thinking at a more abstract level.
#
# The pass is silently ignored:
# + slideshow={"slide_type": "fragment"}
def initlog(*args):
pass # Remember to implement this!
# + [markdown] slideshow={"slide_type": "slide"}
# # Defining Functions
#
# We can create a function that writes the Fibonacci series to an arbitrary boundary:
# + slideshow={"slide_type": "fragment"}
def fib(n): # write Fibonacci series up to n
"""Print a Fibonacci series up to n."""
a, b = 0, 1
while a < n:
print(a,end=' ')
a, b = b, a+b
# Now call the function we just defined:
fib(2000)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### `def`
#
# The keyword `def` introduces a function definition.
#
# It must be followed by the function name and the parenthesized list of formal parameters.
#
# The statements that form the body of the function start at the next line, and must be indented.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Document String
#
# The first statement of the function body can optionally be a string literal; this string literal is the function’s documentation string, or docstring. (More about docstrings can be found in the section Documentation Strings.)
#
# There are tools which use docstrings to automatically produce online or printed documentation, or to let the user interactively browse through code.
#
# It’s good practice to include docstrings in code that you write, so make a habit of it.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Function Scope
#
# The execution of a function introduces a new symbol table used for the local variables of the function.
#
# More precisely, all variable assignments in a function store the value in the local symbol table;
#
# whereas variable references first look in the local symbol table, then in the local symbol tables of enclosing functions, then in the global symbol table, and finally in the table of built-in names.
#
# Thus, global variables cannot be directly assigned a value within a function (unless named in a global statement), although they may be referenced.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Arguments
#
# The actual parameters (arguments) to a function call are introduced in the local symbol table of the called function when it is called;
#
# thus, arguments are passed using call by value (where the value is always an object reference, not the value of the object).
#
# When a function calls another function, a new local symbol table is created for that call.
# + [markdown] slideshow={"slide_type": "slide"}
# A function definition introduces the function name in the current symbol table.
#
# The value of the function name has a type that is recognized by the interpreter as a user-defined function.
#
# This value can be assigned to another name which can then also be used as a function. This serves as a general renaming mechanism:
# + slideshow={"slide_type": "fragment"}
fib
# + slideshow={"slide_type": "fragment"}
f = fib
f(100)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Default Return is `None`
#
# Coming from other languages, you might object that fib is not a function but a procedure since it doesn’t return a value.
#
# In fact, even functions without a return statement do return a value, albeit a rather boring one.
#
# This value is called `None` (it’s a built-in name).
#
# Writing the value None is normally suppressed by the interpreter if it would be the only value written.
#
# You can see it if you really want to using print:
#
# + slideshow={"slide_type": "fragment"}
fib(0) # Nothing prints
# + slideshow={"slide_type": "fragment"}
print(fib(0))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Return a List
#
# It is simple to write a function that returns a list of the numbers of the Fibonacci series, instead of printing it:
# + slideshow={"slide_type": "fragment"}
def fib2(n): # return Fibonacci series up to n
"""Return a list containing the Fibonacci series up to n."""
result = []
a, b = 0, 1
while a < n:
result.append(a) # see below
a, b = b, a+b
return result
f100 = fib2(100) # call it
f100 # write the result
# + [markdown] slideshow={"slide_type": "subslide"}
# This example, as usual, demonstrates some new Python features:
#
# - The `return` statement returns with a value from a function. `return` without an expression argument returns None. Falling off the end of a function also returns `None`.
#
# - The statement `result.append(a)` calls a method of the `list` object result.
# - A method is a function that ‘belongs’ to an object and is named `obj.methodname`, where `obj` is some object (this may be an expression), and `methodname` is the name of a method that is defined by the object’s type.
# - Different types define different methods. Methods of different types may have the same name without causing ambiguity. (It is possible to define your own object types and methods, using classes, see Classes)
# - The method `append()` shown in the example is defined for list objects; it adds a new element at the end of the list. In this example it is equivalent to `result = result + [a]`, but more efficient.
# + [markdown] slideshow={"slide_type": "slide"}
# # More on Defining Functions
#
# It is also possible to define functions with a variable number of arguments. There are three forms, which can be combined.
# + [markdown] slideshow={"slide_type": "slide"}
# # Default Argument Values
#
# The most useful form is to specify a default value for one or more arguments. This creates a function that can be called with fewer arguments than it is defined to allow. For example:
# + slideshow={"slide_type": "fragment"}
def ask_ok(prompt, retries=4, complaint='Yes or no, please!'):
while True:
ok = raw_input(prompt)
if ok in ('y', 'ye', 'yes'):
return True
if ok in ('n', 'no', 'nop', 'nope'):
return False
retries = retries - 1
if retries < 0:
raise IOError('refusenik user')
print(complaint)
# + [markdown] slideshow={"slide_type": "fragment"}
# This function can be called in several ways:
#
# - giving only the mandatory argument: ask_ok('Do you really want to quit?')
# - giving one of the optional arguments: ask_ok('OK to overwrite the file?', 2)
# - or even giving all arguments: ask_ok('OK to overwrite the file?', 2, 'Come on, only yes or no!')
# + [markdown] slideshow={"slide_type": "subslide"}
# This example also introduces the in keyword. This tests whether or not a sequence contains a certain value.
# + [markdown] slideshow={"slide_type": "slide"}
# The default values are evaluated at the point of function definition in the defining scope, so that
# + slideshow={"slide_type": "fragment"}
i = 5
def f(arg=i):
print(arg
)
i = 6
f()
# + [markdown] slideshow={"slide_type": "slide"}
# ### Important warning:
#
# The default value is evaluated only once.
#
# This makes a difference when the default is a mutable object such as a list, dictionary, or instances of most classes.
#
# For example, the following function accumulates the arguments passed to it on subsequent calls:
# + slideshow={"slide_type": "fragment"}
def f(a, L=[]):
L.append(a)
return L
print(f(1))
print(f(2))
print(f(3))
# + [markdown] slideshow={"slide_type": "subslide"}
# If you don’t want the default to be shared between subsequent calls, you can write the function like this instead:
# + slideshow={"slide_type": "fragment"}
def f(a, L=None):
if L is None:
L = []
L.append(a)
return L
print(f(1))
print(f(2))
print(f(3))
# + [markdown] slideshow={"slide_type": "slide"}
# # Keyword Arguments
#
# Functions can also be called using keyword arguments of the form `kwarg=value`. For instance, the following function:
# + slideshow={"slide_type": "fragment"}
def parrot(voltage, state='a stiff', action='voom', type='Norwegian Blue'):
print("-- This parrot wouldn't", action,)
print("if you put", voltage, "volts through it.")
print("-- Lovely plumage, the", type)
print("-- It's", state, "!")
# + [markdown] slideshow={"slide_type": "fragment"}
#
# accepts one required argument (voltage) and three optional arguments (state, action, and type). This function can be called in any of the following ways:
# + slideshow={"slide_type": "subslide"}
parrot(1000) # 1 positional argument
parrot(voltage=1000) # 1 keyword argument
parrot(voltage=1000000, action='VOOOOOM') # 2 keyword arguments
parrot(action='VOOOOOM', voltage=1000000) # 2 keyword arguments
parrot('a million', 'bereft of life', 'jump') # 3 positional arguments
parrot('a thousand', state='pushing up the daisies') # 1 positional, 1 keyword
# + [markdown] slideshow={"slide_type": "slide"}
# # Lambda Expressions
#
# Small anonymous functions can be created with the lambda keyword.
#
# This function returns the sum of its two arguments: `lambda a, b: a+b`.
#
# Lambda functions can be used wherever function objects are required.
#
# They are syntactically restricted to a single expression.
#
# Semantically, they are just syntactic sugar for a normal function definition.
#
# Like nested function definitions, lambda functions can reference variables from the containing scope:
# + slideshow={"slide_type": "fragment"}
def make_incrementor(n):
return lambda x: x + n
f = make_incrementor(42)
f(0), f(1)
# + [markdown] slideshow={"slide_type": "subslide"}
# The above example uses a lambda expression to return a function. Another use is to pass a small function as an argument:
# + slideshow={"slide_type": "fragment"}
pairs = [(1, 'one'), (2, 'two'), (3, 'three'), (4, 'four')]
pairs.sort(key=lambda pair: pair[1])
pairs
# + [markdown] slideshow={"slide_type": "slide"}
# # Document Strings
#
# There are emerging conventions about the content and formatting of documentation strings.
#
# The first line should always be a short, concise summary of the object’s purpose.
#
# For brevity, it should not explicitly state the object’s name or type, since these are available by other means (except if the name happens to be a verb describing a function’s operation).
#
# This line should begin with a capital letter and end with a period.
#
# If there are more lines in the documentation string, the second line should be blank, visually separating the summary from the rest of the description.
#
# The following lines should be one or more paragraphs describing the object’s calling conventions, its side effects, etc.
# + slideshow={"slide_type": "subslide"}
def my_function():
"""Do nothing, but document it.
No, really, it doesn't do anything.
"""
pass
print(my_function.__doc__)
# + [markdown] slideshow={"slide_type": "slide"}
# # Intermezzo: Coding Style
#
# Now that you are about to write longer, more complex pieces of Python, it is a good time to talk about coding style.
#
# Most languages can be written (or more concise, formatted) in different styles; some are more readable than others.
#
# Making it easy for others to read your code is always a good idea, and adopting a nice coding style helps tremendously for that.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### PEP 8
#
# For Python, PEP 8 has emerged as the style guide that most projects adhere to; it promotes a very readable and eye-pleasing coding style. Every Python developer should read it at some point; here are the most important points extracted for you:
#
# - Use 4-space indentation, and no tabs.
#
# - 4 spaces are a good compromise between small indentation (allows greater nesting depth) and large indentation (easier to read). Tabs introduce confusion, and are best left out.
#
# - Wrap lines so that they don’t exceed 79 characters.
#
# - This helps users with small displays and makes it possible to have several code files side-by-side on larger displays.
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# - Use blank lines to separate functions and classes, and larger blocks of code inside functions.
#
# - When possible, put comments on a line of their own.
#
# - Use docstrings.
#
# - Use spaces around operators and after commas, but not directly inside bracketing constructs: a = f(1, 2) + g(3, 4).
#
# - Name your classes and functions consistently; the convention is to use CamelCase for classes and lower_case_with_underscores for functions and methods. Always use self as the name for the first method argument (see A First Look at Classes for more on classes and methods).
#
# - Don’t use fancy encodings if your code is meant to be used in international environments. Plain ASCII works best in any case.
| python/Module-04-More_Control_Flow_Tools.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solving problems by Searching
#
# This notebook serves as supporting material for topics covered in **Chapter 3 - Solving Problems by Searching** and **Chapter 4 - Beyond Classical Search** from the book *Artificial Intelligence: A Modern Approach.* This notebook uses implementations from [search.py](https://github.com/aimacode/aima-python/blob/master/search.py) module. Let's start by importing everything from search module.
# +
from search import *
from notebook import psource, heatmap, gaussian_kernel, show_map, final_path_colors, display_visual, plot_NQueens
# Needed to hide warnings in the matplotlib sections
import warnings
warnings.filterwarnings("ignore")
# -
# ## CONTENTS
#
# * Overview
# * Problem
# * Node
# * Simple Problem Solving Agent
# * Search Algorithms Visualization
# * Breadth-First Tree Search
# * Breadth-First Search
# * Best First Search
# * Uniform Cost Search
# * Greedy Best First Search
# * A\* Search
# * Hill Climbing
# * Simulated Annealing
# * Genetic Algorithm
# * AND-OR Graph Search
# * Online DFS Agent
# * LRTA* Agent
# ## OVERVIEW
#
# Here, we learn about a specific kind of problem solving - building goal-based agents that can plan ahead to solve problems. In particular, we examine navigation problem/route finding problem. We must begin by precisely defining **problems** and their **solutions**. We will look at several general-purpose search algorithms.
#
# Search algorithms can be classified into two types:
#
# * **Uninformed search algorithms**: Search algorithms which explore the search space without having any information about the problem other than its definition.
# * Examples:
# 1. Breadth First Search
# 2. Depth First Search
# 3. Depth Limited Search
# 4. Iterative Deepening Search
#
#
# * **Informed search algorithms**: These type of algorithms leverage any information (heuristics, path cost) on the problem to search through the search space to find the solution efficiently.
# * Examples:
# 1. Best First Search
# 2. Uniform Cost Search
# 3. A\* Search
# 4. Recursive Best First Search
#
# *Don't miss the visualisations of these algorithms solving the route-finding problem defined on Romania map at the end of this notebook.*
# For visualisations, we use networkx and matplotlib to show the map in the notebook and we use ipywidgets to interact with the map to see how the searching algorithm works. These are imported as required in `notebook.py`.
# +
# %matplotlib inline
import networkx as nx
import matplotlib.pyplot as plt
from matplotlib import lines
from ipywidgets import interact
import ipywidgets as widgets
from IPython.display import display
import time
# -
# ## PROBLEM
#
# Let's see how we define a Problem. Run the next cell to see how abstract class `Problem` is defined in the search module.
psource(Problem)
# The `Problem` class has six methods.
#
# * `__init__(self, initial, goal)` : This is what is called a `constructor`. It is the first method called when you create an instance of the class as `Problem(initial, goal)`. The variable `initial` specifies the initial state $s_0$ of the search problem. It represents the beginning state. From here, our agent begins its task of exploration to find the goal state(s) which is given in the `goal` parameter.
#
#
# * `actions(self, state)` : This method returns all the possible actions agent can execute in the given state `state`.
#
#
# * `result(self, state, action)` : This returns the resulting state if action `action` is taken in the state `state`. This `Problem` class only deals with deterministic outcomes. So we know for sure what every action in a state would result to.
#
#
# * `goal_test(self, state)` : Return a boolean for a given state - `True` if it is a goal state, else `False`.
#
#
# * `path_cost(self, c, state1, action, state2)` : Return the cost of the path that arrives at `state2` as a result of taking `action` from `state1`, assuming total cost of `c` to get up to `state1`.
#
#
# * `value(self, state)` : This acts as a bit of extra information in problems where we try to optimise a value when we cannot do a goal test.
# ## NODE
#
# Let's see how we define a Node. Run the next cell to see how abstract class `Node` is defined in the search module.
psource(Node)
# The `Node` class has nine methods. The first is the `__init__` method.
#
# * `__init__(self, state, parent, action, path_cost)` : This method creates a node. `parent` represents the node that this is a successor of and `action` is the action required to get from the parent node to this node. `path_cost` is the cost to reach current node from parent node.
#
# The next 4 methods are specific `Node`-related functions.
#
# * `expand(self, problem)` : This method lists all the neighbouring(reachable in one step) nodes of current node.
#
# * `child_node(self, problem, action)` : Given an `action`, this method returns the immediate neighbour that can be reached with that `action`.
#
# * `solution(self)` : This returns the sequence of actions required to reach this node from the root node.
#
# * `path(self)` : This returns a list of all the nodes that lies in the path from the root to this node.
#
# The remaining 4 methods override standards Python functionality for representing an object as a string, the less-than ($<$) operator, the equal-to ($=$) operator, and the `hash` function.
#
# * `__repr__(self)` : This returns the state of this node.
#
# * `__lt__(self, node)` : Given a `node`, this method returns `True` if the state of current node is less than the state of the `node`. Otherwise it returns `False`.
#
# * `__eq__(self, other)` : This method returns `True` if the state of current node is equal to the other node. Else it returns `False`.
#
# * `__hash__(self)` : This returns the hash of the state of current node.
# We will use the abstract class `Problem` to define our real **problem** named `GraphProblem`. You can see how we define `GraphProblem` by running the next cell.
psource(GraphProblem)
# Have a look at our romania_map, which is an Undirected Graph containing a dict of nodes as keys and neighbours as values.
# +
romania_map = UndirectedGraph(dict(
Arad=dict(Zerind=75, Sibiu=140, Timisoara=118),
Bucharest=dict(Urziceni=85, Pitesti=101, Giurgiu=90, Fagaras=211),
Craiova=dict(Drobeta=120, Rimnicu=146, Pitesti=138),
Drobeta=dict(Mehadia=75),
Eforie=dict(Hirsova=86),
Fagaras=dict(Sibiu=99),
Hirsova=dict(Urziceni=98),
Iasi=dict(Vaslui=92, Neamt=87),
Lugoj=dict(Timisoara=111, Mehadia=70),
Oradea=dict(Zerind=71, Sibiu=151),
Pitesti=dict(Rimnicu=97),
Rimnicu=dict(Sibiu=80),
Urziceni=dict(Vaslui=142)))
romania_map.locations = dict(
Arad=(91, 492), Bucharest=(400, 327), Craiova=(253, 288),
Drobeta=(165, 299), Eforie=(562, 293), Fagaras=(305, 449),
Giurgiu=(375, 270), Hirsova=(534, 350), Iasi=(473, 506),
Lugoj=(165, 379), Mehadia=(168, 339), Neamt=(406, 537),
Oradea=(131, 571), Pitesti=(320, 368), Rimnicu=(233, 410),
Sibiu=(207, 457), Timisoara=(94, 410), Urziceni=(456, 350),
Vaslui=(509, 444), Zerind=(108, 531))
# -
# It is pretty straightforward to understand this `romania_map`. The first node **Arad** has three neighbours named **Zerind**, **Sibiu**, **Timisoara**. Each of these nodes are 75, 140, 118 units apart from **Arad** respectively. And the same goes with other nodes.
#
# And `romania_map.locations` contains the positions of each of the nodes. We will use the straight line distance (which is different from the one provided in `romania_map`) between two cities in algorithms like A\*-search and Recursive Best First Search.
#
# **Define a problem:**
# Now it's time to define our problem. We will define it by passing `initial`, `goal`, `graph` to `GraphProblem`. So, our problem is to find the goal state starting from the given initial state on the provided graph.
#
# Say we want to start exploring from **Arad** and try to find **Bucharest** in our romania_map. So, this is how we do it.
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
# ### Romania Map Visualisation
#
# Let's have a visualisation of Romania map [Figure 3.2] from the book and see how different searching algorithms perform / how frontier expands in each search algorithm for a simple problem named `romania_problem`.
# Have a look at `romania_locations`. It is a dictionary defined in search module. We will use these location values to draw the romania graph using **networkx**.
romania_locations = romania_map.locations
print(romania_locations)
# Let's get started by initializing an empty graph. We will add nodes, place the nodes in their location as shown in the book, add edges to the graph.
# +
# node colors, node positions and node label positions
node_colors = {node: 'white' for node in romania_map.locations.keys()}
node_positions = romania_map.locations
node_label_pos = { k:[v[0],v[1]-10] for k,v in romania_map.locations.items() }
edge_weights = {(k, k2) : v2 for k, v in romania_map.graph_dict.items() for k2, v2 in v.items()}
romania_graph_data = { 'graph_dict' : romania_map.graph_dict,
'node_colors': node_colors,
'node_positions': node_positions,
'node_label_positions': node_label_pos,
'edge_weights': edge_weights
}
# -
# We have completed building our graph based on romania_map and its locations. It's time to display it here in the notebook. This function `show_map(node_colors)` helps us do that. We will be calling this function later on to display the map at each and every interval step while searching, using variety of algorithms from the book.
# We can simply call the function with node_colors dictionary object to display it.
show_map(romania_graph_data)
# Voila! You see, the romania map as shown in the Figure[3.2] in the book. Now, see how different searching algorithms perform with our problem statements.
# ## SIMPLE PROBLEM SOLVING AGENT PROGRAM
#
# Let us now define a Simple Problem Solving Agent Program. Run the next cell to see how the abstract class `SimpleProblemSolvingAgentProgram` is defined in the search module.
psource(SimpleProblemSolvingAgentProgram)
# The SimpleProblemSolvingAgentProgram class has six methods:
#
# * `__init__(self, intial_state=None)`: This is the `contructor` of the class and is the first method to be called when the class is instantiated. It takes in a keyword argument, `initial_state` which is initially `None`. The argument `initial_state` represents the state from which the agent starts.
#
# * `__call__(self, percept)`: This method updates the `state` of the agent based on its `percept` using the `update_state` method. It then formulates a `goal` with the help of `formulate_goal` method and a `problem` using the `formulate_problem` method and returns a sequence of actions to solve it (using the `search` method).
#
# * `update_state(self, percept)`: This method updates the `state` of the agent based on its `percept`.
#
# * `formulate_goal(self, state)`: Given a `state` of the agent, this method formulates the `goal` for it.
#
# * `formulate_problem(self, state, goal)`: It is used in problem formulation given a `state` and a `goal` for the `agent`.
#
# * `search(self, problem)`: This method is used to search a sequence of `actions` to solve a `problem`.
# Let us now define a Simple Problem Solving Agent Program. We will create a simple `vacuumAgent` class which will inherit from the abstract class `SimpleProblemSolvingAgentProgram` and overrides its methods. We will create a simple intelligent vacuum agent which can be in any one of the following states. It will move to any other state depending upon the current state as shown in the picture by arrows:
#
# 
class vacuumAgent(SimpleProblemSolvingAgentProgram):
def update_state(self, state, percept):
return percept
def formulate_goal(self, state):
goal = [state7, state8]
return goal
def formulate_problem(self, state, goal):
problem = state
return problem
def search(self, problem):
if problem == state1:
seq = ["Suck", "Right", "Suck"]
elif problem == state2:
seq = ["Suck", "Left", "Suck"]
elif problem == state3:
seq = ["Right", "Suck"]
elif problem == state4:
seq = ["Suck"]
elif problem == state5:
seq = ["Suck"]
elif problem == state6:
seq = ["Left", "Suck"]
return seq
# Now, we will define all the 8 states and create an object of the above class. Then, we will pass it different states and check the output:
# +
state1 = [(0, 0), [(0, 0), "Dirty"], [(1, 0), ["Dirty"]]]
state2 = [(1, 0), [(0, 0), "Dirty"], [(1, 0), ["Dirty"]]]
state3 = [(0, 0), [(0, 0), "Clean"], [(1, 0), ["Dirty"]]]
state4 = [(1, 0), [(0, 0), "Clean"], [(1, 0), ["Dirty"]]]
state5 = [(0, 0), [(0, 0), "Dirty"], [(1, 0), ["Clean"]]]
state6 = [(1, 0), [(0, 0), "Dirty"], [(1, 0), ["Clean"]]]
state7 = [(0, 0), [(0, 0), "Clean"], [(1, 0), ["Clean"]]]
state8 = [(1, 0), [(0, 0), "Clean"], [(1, 0), ["Clean"]]]
a = vacuumAgent(state1)
print(a(state6))
print(a(state1))
print(a(state3))
# -
# ## SEARCHING ALGORITHMS VISUALIZATION
#
# In this section, we have visualizations of the following searching algorithms:
#
# 1. Breadth First Tree Search
# 2. Depth First Tree Search
# 3. Breadth First Search
# 4. Depth First Graph Search
# 5. Best First Graph Search
# 6. Uniform Cost Search
# 7. Depth Limited Search
# 8. Iterative Deepening Search
# 9. Greedy Best First Search
# 9. A\*-Search
# 10. Recursive Best First Search
#
# We add the colors to the nodes to have a nice visualisation when displaying. So, these are the different colors we are using in these visuals:
# * Un-explored nodes - <font color='black'>white</font>
# * Frontier nodes - <font color='orange'>orange</font>
# * Currently exploring node - <font color='red'>red</font>
# * Already explored nodes - <font color='gray'>gray</font>
# ## 1. BREADTH-FIRST TREE SEARCH
#
# We have a working implementation in search module. But as we want to interact with the graph while it is searching, we need to modify the implementation. Here's the modified breadth first tree search.
# +
def tree_breadth_search_for_vis(problem):
"""Search through the successors of a problem to find a goal.
The argument frontier should be an empty queue.
Don't worry about repeated paths to a state. [Figure 3.7]"""
# we use these two variables at the time of visualisations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
#Adding first node to the queue
frontier = deque([Node(problem.initial)])
node_colors[Node(problem.initial).state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
while frontier:
#Popping first node of queue
node = frontier.popleft()
# modify the currently searching node to red
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
# modify goal node to green after reaching the goal
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
frontier.extend(node.expand(problem))
for n in node.expand(problem):
node_colors[n.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
# modify the color of explored nodes to gray
node_colors[node.state] = "gray"
iterations += 1
all_node_colors.append(dict(node_colors))
return None
def breadth_first_tree_search(problem):
"Search the shallowest nodes in the search tree first."
iterations, all_node_colors, node = tree_breadth_search_for_vis(problem)
return(iterations, all_node_colors, node)
# -
# Now, we use `ipywidgets` to display a slider, a button and our romania map. By sliding the slider we can have a look at all the intermediate steps of a particular search algorithm. By pressing the button **Visualize**, you can see all the steps without interacting with the slider. These two helper functions are the callback functions which are called when we interact with the slider and the button.
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
a, b, c = breadth_first_tree_search(romania_problem)
display_visual(romania_graph_data, user_input=False,
algorithm=breadth_first_tree_search,
problem=romania_problem)
# ## 2. DEPTH-FIRST TREE SEARCH
# Now let's discuss another searching algorithm, Depth-First Tree Search.
# +
def tree_depth_search_for_vis(problem):
"""Search through the successors of a problem to find a goal.
The argument frontier should be an empty queue.
Don't worry about repeated paths to a state. [Figure 3.7]"""
# we use these two variables at the time of visualisations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
#Adding first node to the stack
frontier = [Node(problem.initial)]
node_colors[Node(problem.initial).state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
while frontier:
#Popping first node of stack
node = frontier.pop()
# modify the currently searching node to red
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
# modify goal node to green after reaching the goal
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
frontier.extend(node.expand(problem))
for n in node.expand(problem):
node_colors[n.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
# modify the color of explored nodes to gray
node_colors[node.state] = "gray"
iterations += 1
all_node_colors.append(dict(node_colors))
return None
def depth_first_tree_search(problem):
"Search the deepest nodes in the search tree first."
iterations, all_node_colors, node = tree_depth_search_for_vis(problem)
return(iterations, all_node_colors, node)
# -
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=depth_first_tree_search,
problem=romania_problem)
# ## 3. BREADTH-FIRST GRAPH SEARCH
#
# Let's change all the `node_colors` to starting position and define a different problem statement.
def breadth_first_search_graph(problem):
"[Figure 3.11]"
# we use these two variables at the time of visualisations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
node = Node(problem.initial)
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
frontier = deque([node])
# modify the color of frontier nodes to blue
node_colors[node.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
explored = set()
while frontier:
node = frontier.popleft()
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
explored.add(node.state)
for child in node.expand(problem):
if child.state not in explored and child not in frontier:
if problem.goal_test(child.state):
node_colors[child.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, child)
frontier.append(child)
node_colors[child.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
node_colors[node.state] = "gray"
iterations += 1
all_node_colors.append(dict(node_colors))
return None
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=breadth_first_search_graph,
problem=romania_problem)
# ## 4. DEPTH-FIRST GRAPH SEARCH
# Although we have a working implementation in search module, we have to make a few changes in the algorithm to make it suitable for visualization.
# +
def graph_search_for_vis(problem):
"""Search through the successors of a problem to find a goal.
The argument frontier should be an empty queue.
If two paths reach a state, only use the first one. [Figure 3.7]"""
# we use these two variables at the time of visualisations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
frontier = [(Node(problem.initial))]
explored = set()
# modify the color of frontier nodes to orange
node_colors[Node(problem.initial).state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
while frontier:
# Popping first node of stack
node = frontier.pop()
# modify the currently searching node to red
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
# modify goal node to green after reaching the goal
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
explored.add(node.state)
frontier.extend(child for child in node.expand(problem)
if child.state not in explored and
child not in frontier)
for n in frontier:
# modify the color of frontier nodes to orange
node_colors[n.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
# modify the color of explored nodes to gray
node_colors[node.state] = "gray"
iterations += 1
all_node_colors.append(dict(node_colors))
return None
def depth_first_graph_search(problem):
"""Search the deepest nodes in the search tree first."""
iterations, all_node_colors, node = graph_search_for_vis(problem)
return(iterations, all_node_colors, node)
# -
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=depth_first_graph_search,
problem=romania_problem)
# ## 5. BEST FIRST SEARCH
#
# Let's change all the `node_colors` to starting position and define a different problem statement.
def best_first_graph_search_for_vis(problem, f):
"""Search the nodes with the lowest f scores first.
You specify the function f(node) that you want to minimize; for example,
if f is a heuristic estimate to the goal, then we have greedy best
first search; if f is node.depth then we have breadth-first search.
There is a subtlety: the line "f = memoize(f, 'f')" means that the f
values will be cached on the nodes as they are computed. So after doing
a best first search you can examine the f values of the path returned."""
# we use these two variables at the time of visualisations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
f = memoize(f, 'f')
node = Node(problem.initial)
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
frontier = PriorityQueue('min', f)
frontier.append(node)
node_colors[node.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
explored = set()
while frontier:
node = frontier.pop()
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
explored.add(node.state)
for child in node.expand(problem):
if child.state not in explored and child not in frontier:
frontier.append(child)
node_colors[child.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
elif child in frontier:
incumbent = frontier[child]
if f(child) < f(incumbent):
del frontier[incumbent]
frontier.append(child)
node_colors[child.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
node_colors[node.state] = "gray"
iterations += 1
all_node_colors.append(dict(node_colors))
return None
# ## 6. UNIFORM COST SEARCH
#
# Let's change all the `node_colors` to starting position and define a different problem statement.
def uniform_cost_search_graph(problem):
"[Figure 3.14]"
#Uniform Cost Search uses Best First Search algorithm with f(n) = g(n)
iterations, all_node_colors, node = best_first_graph_search_for_vis(problem, lambda node: node.path_cost)
return(iterations, all_node_colors, node)
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=uniform_cost_search_graph,
problem=romania_problem)
# ## 7. DEPTH LIMITED SEARCH
#
# Let's change all the 'node_colors' to starting position and define a different problem statement.
# Although we have a working implementation, but we need to make changes.
# +
def depth_limited_search_graph(problem, limit = -1):
'''
Perform depth first search of graph g.
if limit >= 0, that is the maximum depth of the search.
'''
# we use these two variables at the time of visualisations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
frontier = [Node(problem.initial)]
explored = set()
cutoff_occurred = False
node_colors[Node(problem.initial).state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
while frontier:
# Popping first node of queue
node = frontier.pop()
# modify the currently searching node to red
node_colors[node.state] = "red"
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
# modify goal node to green after reaching the goal
node_colors[node.state] = "green"
iterations += 1
all_node_colors.append(dict(node_colors))
return(iterations, all_node_colors, node)
elif limit >= 0:
cutoff_occurred = True
limit += 1
all_node_color.pop()
iterations -= 1
node_colors[node.state] = "gray"
explored.add(node.state)
frontier.extend(child for child in node.expand(problem)
if child.state not in explored and
child not in frontier)
for n in frontier:
limit -= 1
# modify the color of frontier nodes to orange
node_colors[n.state] = "orange"
iterations += 1
all_node_colors.append(dict(node_colors))
# modify the color of explored nodes to gray
node_colors[node.state] = "gray"
iterations += 1
all_node_colors.append(dict(node_colors))
return 'cutoff' if cutoff_occurred else None
def depth_limited_search_for_vis(problem):
"""Search the deepest nodes in the search tree first."""
iterations, all_node_colors, node = depth_limited_search_graph(problem)
return(iterations, all_node_colors, node)
# -
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=depth_limited_search_for_vis,
problem=romania_problem)
# ## 8. ITERATIVE DEEPENING SEARCH
#
# Let's change all the 'node_colors' to starting position and define a different problem statement.
def iterative_deepening_search_for_vis(problem):
for depth in range(sys.maxsize):
iterations, all_node_colors, node=depth_limited_search_for_vis(problem)
if iterations:
return (iterations, all_node_colors, node)
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=iterative_deepening_search_for_vis,
problem=romania_problem)
# ## 9. GREEDY BEST FIRST SEARCH
# Let's change all the node_colors to starting position and define a different problem statement.
# +
def greedy_best_first_search(problem, h=None):
"""Greedy Best-first graph search is an informative searching algorithm with f(n) = h(n).
You need to specify the h function when you call best_first_search, or
else in your Problem subclass."""
h = memoize(h or problem.h, 'h')
iterations, all_node_colors, node = best_first_graph_search_for_vis(problem, lambda n: h(n))
return(iterations, all_node_colors, node)
psource(GraphProblem)
# -
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=greedy_best_first_search,
problem=romania_problem)
# ## 10. A\* SEARCH
#
# Let's change all the `node_colors` to starting position and define a different problem statement.
def astar_search_graph(problem, h=None):
"""A* search is best-first graph search with f(n) = g(n)+h(n).
You need to specify the h function when you call astar_search, or
else in your Problem subclass."""
h = memoize(h or problem.h, 'h')
iterations, all_node_colors, node = best_first_graph_search_for_vis(problem,
lambda n: n.path_cost + h(n))
return(iterations, all_node_colors, node)
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=astar_search_graph,
problem=romania_problem)
# ## 11. RECURSIVE BEST FIRST SEARCH
# Let's change all the `node_colors` to starting position and define a different problem statement.
def recursive_best_first_search_for_vis(problem, h=None):
"""[Figure 3.26] Recursive best-first search"""
# we use these two variables at the time of visualizations
iterations = 0
all_node_colors = []
node_colors = {k : 'white' for k in problem.graph.nodes()}
h = memoize(h or problem.h, 'h')
def RBFS(problem, node, flimit):
nonlocal iterations
def color_city_and_update_map(node, color):
node_colors[node.state] = color
nonlocal iterations
iterations += 1
all_node_colors.append(dict(node_colors))
if problem.goal_test(node.state):
color_city_and_update_map(node, 'green')
return (iterations, all_node_colors, node), 0 # the second value is immaterial
successors = node.expand(problem)
if len(successors) == 0:
color_city_and_update_map(node, 'gray')
return (iterations, all_node_colors, None), infinity
for s in successors:
color_city_and_update_map(s, 'orange')
s.f = max(s.path_cost + h(s), node.f)
while True:
# Order by lowest f value
successors.sort(key=lambda x: x.f)
best = successors[0]
if best.f > flimit:
color_city_and_update_map(node, 'gray')
return (iterations, all_node_colors, None), best.f
if len(successors) > 1:
alternative = successors[1].f
else:
alternative = infinity
node_colors[node.state] = 'gray'
node_colors[best.state] = 'red'
iterations += 1
all_node_colors.append(dict(node_colors))
result, best.f = RBFS(problem, best, min(flimit, alternative))
if result[2] is not None:
color_city_and_update_map(node, 'green')
return result, best.f
else:
color_city_and_update_map(node, 'red')
node = Node(problem.initial)
node.f = h(node)
node_colors[node.state] = 'red'
iterations += 1
all_node_colors.append(dict(node_colors))
result, bestf = RBFS(problem, node, infinity)
return result
all_node_colors = []
romania_problem = GraphProblem('Arad', 'Bucharest', romania_map)
display_visual(romania_graph_data, user_input=False,
algorithm=recursive_best_first_search_for_vis,
problem=romania_problem)
all_node_colors = []
# display_visual(romania_graph_data, user_input=True, algorithm=breadth_first_tree_search)
algorithms = { "Breadth First Tree Search": tree_breadth_search_for_vis,
"Depth First Tree Search": tree_depth_search_for_vis,
"Breadth First Search": breadth_first_search_graph,
"Depth First Graph Search": graph_search_for_vis,
"Best First Graph Search": best_first_graph_search_for_vis,
"Uniform Cost Search": uniform_cost_search_graph,
"Depth Limited Search": depth_limited_search_for_vis,
"Iterative Deepening Search": iterative_deepening_search_for_vis,
"Greedy Best First Search": greedy_best_first_search,
"A-star Search": astar_search_graph,
"Recursive Best First Search": recursive_best_first_search_for_vis}
display_visual(romania_graph_data, algorithm=algorithms, user_input=True)
# ## RECURSIVE BEST-FIRST SEARCH
# Recursive best-first search is a simple recursive algorithm that improves upon heuristic search by reducing the memory requirement.
# RBFS uses only linear space and it attempts to mimic the operation of standard best-first search.
# Its structure is similar to recursive depth-first search but it doesn't continue indefinitely down the current path, the `f_limit` variable is used to keep track of the f-value of the best _alternative_ path available from any ancestor of the current node.
# RBFS remembers the f-value of the best leaf in the forgotten subtree and can decide whether it is worth re-expanding the tree later.
# <br>
# However, RBFS still suffers from excessive node regeneration.
# <br>
# Let's have a look at the implementation.
psource(recursive_best_first_search)
# This is how `recursive_best_first_search` can solve the `romania_problem`
recursive_best_first_search(romania_problem).solution()
# `recursive_best_first_search` can be used to solve the 8 puzzle problem too, as discussed later.
puzzle = EightPuzzle((2, 4, 3, 1, 5, 6, 7, 8, 0))
assert puzzle.check_solvability((2, 4, 3, 1, 5, 6, 7, 8, 0))
recursive_best_first_search(puzzle).solution()
# ## A* HEURISTICS
#
# Different heuristics provide different efficiency in solving A* problems which are generally defined by the number of explored nodes as well as the branching factor. With the classic 8 puzzle we can show the efficiency of different heuristics through the number of explored nodes.
#
# ### 8 Puzzle Problem
#
# The *8 Puzzle Problem* consists of a 3x3 tray in which the goal is to get the initial configuration to the goal state by shifting the numbered tiles into the blank space.
#
# example:-
#
# Initial State Goal State
# | 7 | 2 | 4 | | 1 | 2 | 3 |
# | 5 | 0 | 6 | | 4 | 5 | 6 |
# | 8 | 3 | 1 | | 7 | 8 | 0 |
#
# We have a total of 9 blank tiles giving us a total of 9! initial configuration but not all of these are solvable. The solvability of a configuration can be checked by calculating the Inversion Permutation. If the total Inversion Permutation is even then the initial configuration is solvable else the initial configuration is not solvable which means that only 9!/2 initial states lead to a solution.
# <br>
# Let's define our goal state.
goal = [1, 2, 3, 4, 5, 6, 7, 8, 0]
# #### Heuristics :-
#
# 1) Manhattan Distance:- For the 8 puzzle problem Manhattan distance is defined as the distance of a tile from its goal state( for the tile numbered '1' in the initial configuration Manhattan distance is 4 "2 for left and 2 for upward displacement").
#
# 2) No. of Misplaced Tiles:- The heuristic calculates the number of misplaced tiles between the current state and goal state.
#
# 3) Sqrt of Manhattan Distance:- It calculates the square root of Manhattan distance.
#
# 4) Max Heuristic:- It assign the score as the maximum between "Manhattan Distance" and "No. of Misplaced Tiles".
# +
# Heuristics for 8 Puzzle Problem
def linear(node):
return sum([1 if node.state[i] != goal[i] else 0 for i in range(8)])
def manhattan(node):
state = node.state
index_goal = {0:[2,2], 1:[0,0], 2:[0,1], 3:[0,2], 4:[1,0], 5:[1,1], 6:[1,2], 7:[2,0], 8:[2,1]}
index_state = {}
index = [[0,0], [0,1], [0,2], [1,0], [1,1], [1,2], [2,0], [2,1], [2,2]]
x, y = 0, 0
for i in range(len(state)):
index_state[state[i]] = index[i]
mhd = 0
for i in range(8):
for j in range(2):
mhd = abs(index_goal[i][j] - index_state[i][j]) + mhd
return mhd
def sqrt_manhattan(node):
state = node.state
index_goal = {0:[2,2], 1:[0,0], 2:[0,1], 3:[0,2], 4:[1,0], 5:[1,1], 6:[1,2], 7:[2,0], 8:[2,1]}
index_state = {}
index = [[0,0], [0,1], [0,2], [1,0], [1,1], [1,2], [2,0], [2,1], [2,2]]
x, y = 0, 0
for i in range(len(state)):
index_state[state[i]] = index[i]
mhd = 0
for i in range(8):
for j in range(2):
mhd = (index_goal[i][j] - index_state[i][j])**2 + mhd
return math.sqrt(mhd)
def max_heuristic(node):
score1 = manhattan(node)
score2 = linear(node)
return max(score1, score2)
# -
# We can solve the puzzle using the `astar_search` method.
# Solving the puzzle
puzzle = EightPuzzle((2, 4, 3, 1, 5, 6, 7, 8, 0))
puzzle.check_solvability((2, 4, 3, 1, 5, 6, 7, 8, 0)) # checks whether the initialized configuration is solvable or not
# This case is solvable, let's proceed.
# <br>
# The default heuristic function returns the number of misplaced tiles.
astar_search(puzzle).solution()
# In the following cells, we use different heuristic functions.
# <br>
astar_search(puzzle, linear).solution()
astar_search(puzzle, manhattan).solution()
astar_search(puzzle, sqrt_manhattan).solution()
astar_search(puzzle, max_heuristic).solution()
# And here's how `recursive_best_first_search` can be used to solve this problem too.
recursive_best_first_search(puzzle, manhattan).solution()
# Even though all the heuristic functions give the same solution, the difference lies in the computation time.
# <br>
# This might make all the difference in a scenario where high computational efficiency is required.
# <br>
# Let's define a few puzzle states and time `astar_search` for every heuristic function.
# We will use the %%timeit magic for this.
puzzle_1 = EightPuzzle((2, 4, 3, 1, 5, 6, 7, 8, 0))
puzzle_2 = EightPuzzle((1, 2, 3, 4, 5, 6, 0, 7, 8))
puzzle_3 = EightPuzzle((1, 2, 3, 4, 5, 7, 8, 6, 0))
# The default heuristic function is the same as the `linear` heuristic function, but we'll still check both.
# %%timeit
astar_search(puzzle_1)
astar_search(puzzle_2)
astar_search(puzzle_3)
# %%timeit
astar_search(puzzle_1, linear)
astar_search(puzzle_2, linear)
astar_search(puzzle_3, linear)
# %%timeit
astar_search(puzzle_1, manhattan)
astar_search(puzzle_2, manhattan)
astar_search(puzzle_3, manhattan)
# %%timeit
astar_search(puzzle_1, sqrt_manhattan)
astar_search(puzzle_2, sqrt_manhattan)
astar_search(puzzle_3, sqrt_manhattan)
# %%timeit
astar_search(puzzle_1, max_heuristic)
astar_search(puzzle_2, max_heuristic)
astar_search(puzzle_3, max_heuristic)
# We can infer that the `manhattan` heuristic function works the fastest.
# <br>
# `sqrt_manhattan` has an extra `sqrt` operation which makes it quite a lot slower than the others.
# <br>
# `max_heuristic` should have been a bit slower as it calls two functions, but in this case, those values were already calculated which saved some time.
# Feel free to play around with these functions.
# For comparison, this is how RBFS performs on this problem.
# %%timeit
recursive_best_first_search(puzzle_1, linear)
recursive_best_first_search(puzzle_2, linear)
recursive_best_first_search(puzzle_3, linear)
# It is quite a lot slower than `astar_search` as we can see.
# ## <NAME>
#
# <NAME> is a heuristic search used for optimization problems.
# Given a large set of inputs and a good heuristic function, it tries to find a sufficiently good solution to the problem.
# This solution may or may not be the global optimum.
# The algorithm is a variant of generate and test algorithm.
# <br>
# As a whole, the algorithm works as follows:
# - Evaluate the initial state.
# - If it is equal to the goal state, return.
# - Find a neighboring state (one which is heuristically similar to the current state)
# - Evaluate this state. If it is closer to the goal state than before, replace the initial state with this state and repeat these steps.
# <br>
psource(hill_climbing)
# We will find an approximate solution to the traveling salespersons problem using this algorithm.
# <br>
# We need to define a class for this problem.
# <br>
# `Problem` will be used as a base class.
class TSP_problem(Problem):
""" subclass of Problem to define various functions """
def two_opt(self, state):
""" Neighbour generating function for Traveling Salesman Problem """
neighbour_state = state[:]
left = random.randint(0, len(neighbour_state) - 1)
right = random.randint(0, len(neighbour_state) - 1)
if left > right:
left, right = right, left
neighbour_state[left: right + 1] = reversed(neighbour_state[left: right + 1])
return neighbour_state
def actions(self, state):
""" action that can be excuted in given state """
return [self.two_opt]
def result(self, state, action):
""" result after applying the given action on the given state """
return action(state)
def path_cost(self, c, state1, action, state2):
""" total distance for the Traveling Salesman to be covered if in state2 """
cost = 0
for i in range(len(state2) - 1):
cost += distances[state2[i]][state2[i + 1]]
cost += distances[state2[0]][state2[-1]]
return cost
def value(self, state):
""" value of path cost given negative for the given state """
return -1 * self.path_cost(None, None, None, state)
# We will use cities from the Romania map as our cities for this problem.
# <br>
# A list of all cities and a dictionary storing distances between them will be populated.
# +
distances = {}
all_cities = []
for city in romania_map.locations.keys():
distances[city] = {}
all_cities.append(city)
all_cities.sort()
print(all_cities)
# -
# Next, we need to populate the individual lists inside the dictionary with the manhattan distance between the cities.
import numpy as np
for name_1, coordinates_1 in romania_map.locations.items():
for name_2, coordinates_2 in romania_map.locations.items():
distances[name_1][name_2] = np.linalg.norm(
[coordinates_1[0] - coordinates_2[0], coordinates_1[1] - coordinates_2[1]])
distances[name_2][name_1] = np.linalg.norm(
[coordinates_1[0] - coordinates_2[0], coordinates_1[1] - coordinates_2[1]])
# The way neighbours are chosen currently isn't suitable for the travelling salespersons problem.
# We need a neighboring state that is similar in total path distance to the current state.
# <br>
# We need to change the function that finds neighbors.
def hill_climbing(problem):
"""From the initial node, keep choosing the neighbor with highest value,
stopping when no neighbor is better. [Figure 4.2]"""
def find_neighbors(state, number_of_neighbors=100):
""" finds neighbors using two_opt method """
neighbors = []
for i in range(number_of_neighbors):
new_state = problem.two_opt(state)
neighbors.append(Node(new_state))
state = new_state
return neighbors
# as this is a stochastic algorithm, we will set a cap on the number of iterations
iterations = 10000
current = Node(problem.initial)
while iterations:
neighbors = find_neighbors(current.state)
if not neighbors:
break
neighbor = argmax_random_tie(neighbors,
key=lambda node: problem.value(node.state))
if problem.value(neighbor.state) <= problem.value(current.state):
current.state = neighbor.state
iterations -= 1
return current.state
# An instance of the TSP_problem class will be created.
tsp = TSP_problem(all_cities)
# We can now generate an approximate solution to the problem by calling `hill_climbing`.
# The results will vary a bit each time you run it.
hill_climbing(tsp)
# The solution looks like this.
# It is not difficult to see why this might be a good solution.
# <br>
# 
# ## SIMULATED ANNEALING
#
# The intuition behind Hill Climbing was developed from the metaphor of climbing up the graph of a function to find its peak.
# There is a fundamental problem in the implementation of the algorithm however.
# To find the highest hill, we take one step at a time, always uphill, hoping to find the highest point,
# but if we are unlucky to start from the shoulder of the second-highest hill, there is no way we can find the highest one.
# The algorithm will always converge to the local optimum.
# Hill Climbing is also bad at dealing with functions that flatline in certain regions.
# If all neighboring states have the same value, we cannot find the global optimum using this algorithm.
# <br>
# <br>
# Let's now look at an algorithm that can deal with these situations.
# <br>
# Simulated Annealing is quite similar to Hill Climbing,
# but instead of picking the _best_ move every iteration, it picks a _random_ move.
# If this random move brings us closer to the global optimum, it will be accepted,
# but if it doesn't, the algorithm may accept or reject the move based on a probability dictated by the _temperature_.
# When the `temperature` is high, the algorithm is more likely to accept a random move even if it is bad.
# At low temperatures, only good moves are accepted, with the occasional exception.
# This allows exploration of the state space and prevents the algorithm from getting stuck at the local optimum.
#
psource(simulated_annealing)
# The temperature is gradually decreased over the course of the iteration.
# This is done by a scheduling routine.
# The current implementation uses exponential decay of temperature, but we can use a different scheduling routine instead.
#
psource(exp_schedule)
# Next, we'll define a peak-finding problem and try to solve it using Simulated Annealing.
# Let's define the grid and the initial state first.
#
initial = (0, 0)
grid = [[3, 7, 2, 8], [5, 2, 9, 1], [5, 3, 3, 1]]
# We want to allow only four directions, namely `N`, `S`, `E` and `W`.
# Let's use the predefined `directions4` dictionary.
directions4
# Define a problem with these parameters.
problem = PeakFindingProblem(initial, grid, directions4)
# We'll run `simulated_annealing` a few times and store the solutions in a set.
solutions = {problem.value(simulated_annealing(problem)) for i in range(100)}
max(solutions)
# Hence, the maximum value is 9.
# Let's find the peak of a two-dimensional gaussian distribution.
# We'll use the `gaussian_kernel` function from notebook.py to get the distribution.
grid = gaussian_kernel()
# Let's use the `heatmap` function from notebook.py to plot this.
heatmap(grid, cmap='jet', interpolation='spline16')
# Let's define the problem.
# This time, we will allow movement in eight directions as defined in `directions8`.
directions8
# We'll solve the problem just like we did last time.
# <br>
# Let's also time it.
problem = PeakFindingProblem(initial, grid, directions8)
# %%timeit
solutions = {problem.value(simulated_annealing(problem)) for i in range(100)}
max(solutions)
# The peak is at 1.0 which is how gaussian distributions are defined.
# <br>
# This could also be solved by Hill Climbing as follows.
# %%timeit
solution = problem.value(hill_climbing(problem))
solution = problem.value(hill_climbing(problem))
solution
# As you can see, Hill-Climbing is about 24 times faster than Simulated Annealing.
# (Notice that we ran Simulated Annealing for 100 iterations whereas we ran Hill Climbing only once.)
# <br>
# Simulated Annealing makes up for its tardiness by its ability to be applicable in a larger number of scenarios than Hill Climbing as illustrated by the example below.
# <br>
# Let's define a 2D surface as a matrix.
grid = [[0, 0, 0, 1, 4],
[0, 0, 2, 8, 10],
[0, 0, 2, 4, 12],
[0, 2, 4, 8, 16],
[1, 4, 8, 16, 32]]
heatmap(grid, cmap='jet', interpolation='spline16')
# The peak value is 32 at the lower right corner.
# <br>
# The region at the upper left corner is planar.
# Let's instantiate `PeakFindingProblem` one last time.
problem = PeakFindingProblem(initial, grid, directions8)
# Solution by <NAME>
solution = problem.value(hill_climbing(problem))
solution
# Solution by Simulated Annealing
solutions = {problem.value(simulated_annealing(problem)) for i in range(100)}
max(solutions)
# Notice that even though both algorithms started at the same initial state,
# Hill Climbing could never escape from the planar region and gave a locally optimum solution of **0**,
# whereas Simulated Annealing could reach the peak at **32**.
# <br>
# A very similar situation arises when there are two peaks of different heights.
# One should carefully consider the possible search space before choosing the algorithm for the task.
# ## GENETIC ALGORITHM
#
# Genetic algorithms (or GA) are inspired by natural evolution and are particularly useful in optimization and search problems with large state spaces.
#
# Given a problem, algorithms in the domain make use of a *population* of solutions (also called *states*), where each solution/state represents a feasible solution. At each iteration (often called *generation*), the population gets updated using methods inspired by biology and evolution, like *crossover*, *mutation* and *natural selection*.
# ### Overview
#
# A genetic algorithm works in the following way:
#
# 1) Initialize random population.
#
# 2) Calculate population fitness.
#
# 3) Select individuals for mating.
#
# 4) Mate selected individuals to produce new population.
#
# * Random chance to mutate individuals.
#
# 5) Repeat from step 2) until an individual is fit enough or the maximum number of iterations was reached.
# ### Glossary
#
# Before we continue, we will lay the basic terminology of the algorithm.
#
# * Individual/State: A list of elements (called *genes*) that represent possible solutions.
#
# * Population: The list of all the individuals/states.
#
# * Gene pool: The alphabet of possible values for an individual's genes.
#
# * Generation/Iteration: The number of times the population will be updated.
#
# * Fitness: An individual's score, calculated by a function specific to the problem.
# ### Crossover
#
# Two individuals/states can "mate" and produce one child. This offspring bears characteristics from both of its parents. There are many ways we can implement this crossover. Here we will take a look at the most common ones. Most other methods are variations of those below.
#
# * Point Crossover: The crossover occurs around one (or more) point. The parents get "split" at the chosen point or points and then get merged. In the example below we see two parents get split and merged at the 3rd digit, producing the following offspring after the crossover.
#
# 
#
# * Uniform Crossover: This type of crossover chooses randomly the genes to get merged. Here the genes 1, 2 and 5 were chosen from the first parent, so the genes 3, 4 were added by the second parent.
#
# 
# ### Mutation
#
# When an offspring is produced, there is a chance it will mutate, having one (or more, depending on the implementation) of its genes altered.
#
# For example, let's say the new individual to undergo mutation is "abcde". Randomly we pick to change its third gene to 'z'. The individual now becomes "abzde" and is added to the population.
# ### Selection
#
# At each iteration, the fittest individuals are picked randomly to mate and produce offsprings. We measure an individual's fitness with a *fitness function*. That function depends on the given problem and it is used to score an individual. Usually the higher the better.
#
# The selection process is this:
#
# 1) Individuals are scored by the fitness function.
#
# 2) Individuals are picked randomly, according to their score (higher score means higher chance to get picked). Usually the formula to calculate the chance to pick an individual is the following (for population *P* and individual *i*):
#
# $$ chance(i) = \dfrac{fitness(i)}{\sum_{k \, in \, P}{fitness(k)}} $$
# ### Implementation
#
# Below we look over the implementation of the algorithm in the `search` module.
#
# First the implementation of the main core of the algorithm:
psource(genetic_algorithm)
# The algorithm takes the following input:
#
# * `population`: The initial population.
#
# * `fitness_fn`: The problem's fitness function.
#
# * `gene_pool`: The gene pool of the states/individuals. By default 0 and 1.
#
# * `f_thres`: The fitness threshold. If an individual reaches that score, iteration stops. By default 'None', which means the algorithm will not halt until the generations are ran.
#
# * `ngen`: The number of iterations/generations.
#
# * `pmut`: The probability of mutation.
#
# The algorithm gives as output the state with the largest score.
# For each generation, the algorithm updates the population. First it calculates the fitnesses of the individuals, then it selects the most fit ones and finally crosses them over to produce offsprings. There is a chance that the offspring will be mutated, given by `pmut`. If at the end of the generation an individual meets the fitness threshold, the algorithm halts and returns that individual.
#
# The function of mating is accomplished by the method `recombine`:
psource(recombine)
# The method picks at random a point and merges the parents (`x` and `y`) around it.
#
# The mutation is done in the method `mutate`:
psource(mutate)
# We pick a gene in `x` to mutate and a gene from the gene pool to replace it with.
#
# To help initializing the population we have the helper function `init_population`":
psource(init_population)
# The function takes as input the number of individuals in the population, the gene pool and the length of each individual/state. It creates individuals with random genes and returns the population when done.
# ### Explanation
#
# Before we solve problems using the genetic algorithm, we will explain how to intuitively understand the algorithm using a trivial example.
#
# #### Generating Phrases
#
# In this problem, we use a genetic algorithm to generate a particular target phrase from a population of random strings. This is a classic example that helps build intuition about how to use this algorithm in other problems as well. Before we break the problem down, let us try to brute force the solution. Let us say that we want to generate the phrase "genetic algorithm". The phrase is 17 characters long. We can use any character from the 26 lowercase characters and the space character. To generate a random phrase of length 17, each space can be filled in 27 ways. So the total number of possible phrases is
#
# $$ 27^{17} = 2153693963075557766310747 $$
#
# which is a massive number. If we wanted to generate the phrase "Genetic Algorithm", we would also have to include all the 26 uppercase characters into consideration thereby increasing the sample space from 27 characters to 53 characters and the total number of possible phrases then would be
#
# $$ 53^{17} = 205442259656281392806087233013 $$
#
# If we wanted to include punctuations and numerals into the sample space, we would have further complicated an already impossible problem. Hence, brute forcing is not an option. Now we'll apply the genetic algorithm and see how it significantly reduces the search space. We essentially want to *evolve* our population of random strings so that they better approximate the target phrase as the number of generations increase. Genetic algorithms work on the principle of Darwinian Natural Selection according to which, there are three key concepts that need to be in place for evolution to happen. They are:
#
# * **Heredity**: There must be a process in place by which children receive the properties of their parents. <br>
# For this particular problem, two strings from the population will be chosen as parents and will be split at a random index and recombined as described in the `recombine` function to create a child. This child string will then be added to the new generation.
#
#
# * **Variation**: There must be a variety of traits present in the population or a means with which to introduce variation. <br>If there is no variation in the sample space, we might never reach the global optimum. To ensure that there is enough variation, we can initialize a large population, but this gets computationally expensive as the population gets larger. Hence, we often use another method called mutation. In this method, we randomly change one or more characters of some strings in the population based on a predefined probability value called the mutation rate or mutation probability as described in the `mutate` function. The mutation rate is usually kept quite low. A mutation rate of zero fails to introduce variation in the population and a high mutation rate (say 50%) is as good as a coin flip and the population fails to benefit from the previous recombinations. An optimum balance has to be maintained between population size and mutation rate so as to reduce the computational cost as well as have sufficient variation in the population.
#
#
# * **Selection**: There must be some mechanism by which some members of the population have the opportunity to be parents and pass down their genetic information and some do not. This is typically referred to as "survival of the fittest". <br>
# There has to be some way of determining which phrases in our population have a better chance of eventually evolving into the target phrase. This is done by introducing a fitness function that calculates how close the generated phrase is to the target phrase. The function will simply return a scalar value corresponding to the number of matching characters between the generated phrase and the target phrase.
# Before solving the problem, we first need to define our target phrase.
target = 'Genetic Algorithm'
# We then need to define our gene pool, i.e the elements which an individual from the population might comprise of. Here, the gene pool contains all uppercase and lowercase letters of the English alphabet and the space character.
# +
# The ASCII values of uppercase characters ranges from 65 to 91
u_case = [chr(x) for x in range(65, 91)]
# The ASCII values of lowercase characters ranges from 97 to 123
l_case = [chr(x) for x in range(97, 123)]
gene_pool = []
gene_pool.extend(u_case) # adds the uppercase list to the gene pool
gene_pool.extend(l_case) # adds the lowercase list to the gene pool
gene_pool.append(' ') # adds the space character to the gene pool
# -
# We now need to define the maximum size of each population. Larger populations have more variation but are computationally more expensive to run algorithms on.
max_population = 100
# As our population is not very large, we can afford to keep a relatively large mutation rate.
mutation_rate = 0.07 # 7%
# Great! Now, we need to define the most important metric for the genetic algorithm, i.e the fitness function. This will simply return the number of matching characters between the generated sample and the target phrase.
def fitness_fn(sample):
# initialize fitness to 0
fitness = 0
for i in range(len(sample)):
# increment fitness by 1 for every matching character
if sample[i] == target[i]:
fitness += 1
return fitness
# Before we run our genetic algorithm, we need to initialize a random population. We will use the `init_population` function to do this. We need to pass in the maximum population size, the gene pool and the length of each individual, which in this case will be the same as the length of the target phrase.
population = init_population(max_population, gene_pool, len(target))
# We will now define how the individuals in the population should change as the number of generations increases. First, the `select` function will be run on the population to select *two* individuals with high fitness values. These will be the parents which will then be recombined using the `recombine` function to generate the child.
parents = select(2, population, fitness_fn)
# The recombine function takes two parents as arguments, so we need to unpack the previous variable
child = recombine(*parents)
# Next, we need to apply a mutation according to the mutation rate. We call the `mutate` function on the child with the gene pool and mutation rate as the additional arguments.
child = mutate(child, gene_pool, mutation_rate)
# The above lines can be condensed into
#
# `child = mutate(recombine(*select(2, population, fitness_fn)), gene_pool, mutation_rate)`
#
# And, we need to do this `for` every individual in the current population to generate the new population.
population = [mutate(recombine(*select(2, population, fitness_fn)), gene_pool, mutation_rate) for i in range(len(population))]
# The individual with the highest fitness can then be found using the `max` function.
current_best = max(population, key=fitness_fn)
# Let's print this out
print(current_best)
# We see that this is a list of characters. This can be converted to a string using the join function
current_best_string = ''.join(current_best)
print(current_best_string)
# We now need to define the conditions to terminate the algorithm. This can happen in two ways
# 1. Termination after a predefined number of generations
# 2. Termination when the fitness of the best individual of the current generation reaches a predefined threshold value.
#
# We define these variables below
ngen = 1200 # maximum number of generations
# we set the threshold fitness equal to the length of the target phrase
# i.e the algorithm only terminates whne it has got all the characters correct
# or it has completed 'ngen' number of generations
f_thres = len(target)
# To generate `ngen` number of generations, we run a `for` loop `ngen` number of times. After each generation, we calculate the fitness of the best individual of the generation and compare it to the value of `f_thres` using the `fitness_threshold` function. After every generation, we print out the best individual of the generation and the corresponding fitness value. Lets now write a function to do this.
def genetic_algorithm_stepwise(population, fitness_fn, gene_pool=[0, 1], f_thres=None, ngen=1200, pmut=0.1):
for generation in range(ngen):
population = [mutate(recombine(*select(2, population, fitness_fn)), gene_pool, pmut) for i in range(len(population))]
# stores the individual genome with the highest fitness in the current population
current_best = ''.join(max(population, key=fitness_fn))
print(f'Current best: {current_best}\t\tGeneration: {str(generation)}\t\tFitness: {fitness_fn(current_best)}\r', end='')
# compare the fitness of the current best individual to f_thres
fittest_individual = fitness_threshold(fitness_fn, f_thres, population)
# if fitness is greater than or equal to f_thres, we terminate the algorithm
if fittest_individual:
return fittest_individual, generation
return max(population, key=fitness_fn) , generation
# The function defined above is essentially the same as the one defined in `search.py` with the added functionality of printing out the data of each generation.
psource(genetic_algorithm)
# We have defined all the required functions and variables. Let's now create a new population and test the function we wrote above.
population = init_population(max_population, gene_pool, len(target))
solution, generations = genetic_algorithm_stepwise(population, fitness_fn, gene_pool, f_thres, ngen, mutation_rate)
# The genetic algorithm was able to converge!
# We implore you to rerun the above cell and play around with `target, max_population, f_thres, ngen` etc parameters to get a better intuition of how the algorithm works. To summarize, if we can define the problem states in simple array format and if we can create a fitness function to gauge how good or bad our approximate solutions are, there is a high chance that we can get a satisfactory solution using a genetic algorithm.
# - There is also a better GUI version of this program `genetic_algorithm_example.py` in the GUI folder for you to play around with.
# ### Usage
#
# Below we give two example usages for the genetic algorithm, for a graph coloring problem and the 8 queens problem.
#
# #### Graph Coloring
#
# First we will take on the simpler problem of coloring a small graph with two colors. Before we do anything, let's imagine how a solution might look. First, we have to represent our colors. Say, 'R' for red and 'G' for green. These make up our gene pool. What of the individual solutions though? For that, we will look at our problem. We stated we have a graph. A graph has nodes and edges, and we want to color the nodes. Naturally, we want to store each node's color. If we have four nodes, we can store their colors in a list of genes, one for each node. A possible solution will then look like this: ['R', 'R', 'G', 'R']. In the general case, we will represent each solution with a list of chars ('R' and 'G'), with length the number of nodes.
#
# Next we need to come up with a fitness function that appropriately scores individuals. Again, we will look at the problem definition at hand. We want to color a graph. For a solution to be optimal, no edge should connect two nodes of the same color. How can we use this information to score a solution? A naive (and ineffective) approach would be to count the different colors in the string. So ['R', 'R', 'R', 'R'] has a score of 1 and ['R', 'R', 'G', 'G'] has a score of 2. Why that fitness function is not ideal though? Why, we forgot the information about the edges! The edges are pivotal to the problem and the above function only deals with node colors. We didn't use all the information at hand and ended up with an ineffective answer. How, then, can we use that information to our advantage?
#
# We said that the optimal solution will have all the edges connecting nodes of different color. So, to score a solution we can count how many edges are valid (aka connecting nodes of different color). That is a great fitness function!
#
# Let's jump into solving this problem using the `genetic_algorithm` function.
# First we need to represent the graph. Since we mostly need information about edges, we will just store the edges. We will denote edges with capital letters and nodes with integers:
edges = {
'A': [0, 1],
'B': [0, 3],
'C': [1, 2],
'D': [2, 3]
}
# Edge 'A' connects nodes 0 and 1, edge 'B' connects nodes 0 and 3 etc.
#
# We already said our gene pool is 'R' and 'G', so we can jump right into initializing our population. Since we have only four nodes, `state_length` should be 4. For the number of individuals, we will try 8. We can increase this number if we need higher accuracy, but be careful! Larger populations need more computating power and take longer. You need to strike that sweet balance between accuracy and cost (the ultimate dilemma of the programmer!).
population = init_population(8, ['R', 'G'], 4)
print(population)
# We created and printed the population. You can see that the genes in the individuals are random and there are 8 individuals each with 4 genes.
#
# Next we need to write our fitness function. We previously said we want the function to count how many edges are valid. So, given a coloring/individual `c`, we will do just that:
def fitness(c):
return sum(c[n1] != c[n2] for (n1, n2) in edges.values())
# Great! Now we will run the genetic algorithm and see what solution it gives.
solution = genetic_algorithm(population, fitness, gene_pool=['R', 'G'])
print(solution)
# The algorithm converged to a solution. Let's check its score:
print(fitness(solution))
# The solution has a score of 4. Which means it is optimal, since we have exactly 4 edges in our graph, meaning all are valid!
#
# *NOTE: Because the algorithm is non-deterministic, there is a chance a different solution is given. It might even be wrong, if we are very unlucky!*
# #### Eight Queens
#
# Let's take a look at a more complicated problem.
#
# In the *Eight Queens* problem, we are tasked with placing eight queens on an 8x8 chessboard without any queen threatening the others (aka queens should not be in the same row, column or diagonal). In its general form the problem is defined as placing *N* queens in an NxN chessboard without any conflicts.
#
# First we need to think about the representation of each solution. We can go the naive route of representing the whole chessboard with the queens' placements on it. That is definitely one way to go about it, but for the purpose of this tutorial we will do something different. We have eight queens, so we will have a gene for each of them. The gene pool will be numbers from 0 to 7, for the different columns. The *position* of the gene in the state will denote the row the particular queen is placed in.
#
# For example, we can have the state "03304577". Here the first gene with a value of 0 means "the queen at row 0 is placed at column 0", for the second gene "the queen at row 1 is placed at column 3" and so forth.
#
# We now need to think about the fitness function. On the graph coloring problem we counted the valid edges. The same thought process can be applied here. Instead of edges though, we have positioning between queens. If two queens are not threatening each other, we say they are at a "non-attacking" positioning. We can, therefore, count how many such positionings are there.
#
# Let's dive right in and initialize our population:
population = init_population(100, range(8), 8)
print(population[:5])
# We have a population of 100 and each individual has 8 genes. The gene pool is the integers from 0 to 7, in string form. Above you can see the first five individuals.
#
# Next we need to write our fitness function. Remember, queens threaten each other if they are at the same row, column or diagonal.
#
# Since positionings are mutual, we must take care not to count them twice. Therefore for each queen, we will only check for conflicts for the queens after her.
#
# A gene's value in an individual `q` denotes the queen's column, and the position of the gene denotes its row. We can check if the aforementioned values between two genes are the same. We also need to check for diagonals. A queen *a* is in the diagonal of another queen, *b*, if the difference of the rows between them is equal to either their difference in columns (for the diagonal on the right of *a*) or equal to the negative difference of their columns (for the left diagonal of *a*). Below is given the fitness function.
def fitness(q):
non_attacking = 0
for row1 in range(len(q)):
for row2 in range(row1+1, len(q)):
col1 = int(q[row1])
col2 = int(q[row2])
row_diff = row1 - row2
col_diff = col1 - col2
if col1 != col2 and row_diff != col_diff and row_diff != -col_diff:
non_attacking += 1
return non_attacking
# Note that the best score achievable is 28. That is because for each queen we only check for the queens after her. For the first queen we check 7 other queens, for the second queen 6 others and so on. In short, the number of checks we make is the sum 7+6+5+...+1. Which is equal to 7\*(7+1)/2 = 28.
#
# Because it is very hard and will take long to find a perfect solution, we will set the fitness threshold at 25. If we find an individual with a score greater or equal to that, we will halt. Let's see how the genetic algorithm will fare.
solution = genetic_algorithm(population, fitness, f_thres=25, gene_pool=range(8))
print(solution)
print(fitness(solution))
# Above you can see the solution and its fitness score, which should be no less than 25.
# This is where we conclude Genetic Algorithms.
# ### N-Queens Problem
# Here, we will look at the generalized cae of the Eight Queens problem.
# <br>
# We are given a `N` x `N` chessboard, with `N` queens, and we need to place them in such a way that no two queens can attack each other.
# <br>
# We will solve this problem using search algorithms.
# To do this, we already have a `NQueensProblem` class in `search.py`.
psource(NQueensProblem)
# In [`csp.ipynb`](https://github.com/aimacode/aima-python/blob/master/csp.ipynb) we have seen that the N-Queens problem can be formulated as a CSP and can be solved by
# the `min_conflicts` algorithm in a way similar to Hill-Climbing.
# Here, we want to solve it using heuristic search algorithms and even some classical search algorithms.
# The `NQueensProblem` class derives from the `Problem` class and is implemented in such a way that the search algorithms we already have, can solve it.
# <br>
# Let's instantiate the class.
nqp = NQueensProblem(8)
# Let's use `depth_first_tree_search` first.
# <br>
# We will also use the %%timeit magic with each algorithm to see how much time they take.
# %%timeit
depth_first_tree_search(nqp)
dfts = depth_first_tree_search(nqp).solution()
plot_NQueens(dfts)
# `breadth_first_tree_search`
# %%timeit
breadth_first_tree_search(nqp)
bfts = breadth_first_tree_search(nqp).solution()
plot_NQueens(bfts)
# `uniform_cost_search`
# %%timeit
uniform_cost_search(nqp)
ucs = uniform_cost_search(nqp).solution()
plot_NQueens(ucs)
# `depth_first_tree_search` is almost 20 times faster than `breadth_first_tree_search` and more than 200 times faster than `uniform_cost_search`.
# We can also solve this problem using `astar_search` with a suitable heuristic function.
# <br>
# The best heuristic function for this scenario will be one that returns the number of conflicts in the current state.
psource(NQueensProblem.h)
# %%timeit
astar_search(nqp)
# `astar_search` is faster than both `uniform_cost_search` and `breadth_first_tree_search`.
astar = astar_search(nqp).solution()
plot_NQueens(astar)
# ## AND-OR GRAPH SEARCH
# An _AND-OR_ graph is a graphical representation of the reduction of goals to _conjunctions_ and _disjunctions_ of subgoals.
# <br>
# An _AND-OR_ graph can be seen as a generalization of a directed graph.
# It contains a number of vertices and generalized edges that connect the vertices.
# <br>
# Each connector in an _AND-OR_ graph connects a set of vertices $V$ to a single vertex, $v_0$.
# A connector can be an _AND_ connector or an _OR_ connector.
# An __AND__ connector connects two edges having a logical _AND_ relationship,
# while and __OR__ connector connects two edges having a logical _OR_ relationship.
# <br>
# A vertex can have more than one _AND_ or _OR_ connector.
# This is why _AND-OR_ graphs can be expressed as logical statements.
# <br>
# <br>
# _AND-OR_ graphs also provide a computational model for executing logic programs and you will come across this data-structure in the `logic` module as well.
# _AND-OR_ graphs can be searched in depth-first, breadth-first or best-first ways searching the state sapce linearly or parallely.
# <br>
# Our implementation of _AND-OR_ search searches over graphs generated by non-deterministic environments and returns a conditional plan that reaches a goal state in all circumstances.
# Let's have a look at the implementation of `and_or_graph_search`.
psource(and_or_graph_search)
# The search is carried out by two functions `and_search` and `or_search` that recursively call each other, traversing nodes sequentially.
# It is a recursive depth-first algorithm for searching an _AND-OR_ graph.
# <br>
# A very similar algorithm `fol_bc_ask` can be found in the `logic` module, which carries out inference on first-order logic knowledge bases using _AND-OR_ graph-derived data-structures.
# <br>
# _AND-OR_ trees can also be used to represent the search spaces for two-player games, where a vertex of the tree represents the problem of one of the players winning the game, starting from the initial state of the game.
# <br>
# Problems involving _MIN-MAX_ trees can be reformulated as _AND-OR_ trees by representing _MAX_ nodes as _OR_ nodes and _MIN_ nodes as _AND_ nodes.
# `and_or_graph_search` can then be used to find the optimal solution.
# Standard algorithms like `minimax` and `expectiminimax` (for belief states) can also be applied on it with a few modifications.
# Here's how `and_or_graph_search` can be applied to a simple vacuum-world example.
vacuum_world = GraphProblemStochastic('State_1', ['State_7', 'State_8'], vacuum_world)
plan = and_or_graph_search(vacuum_world)
plan
def run_plan(state, problem, plan):
if problem.goal_test(state):
return True
if len(plan) is not 2:
return False
predicate = lambda x: run_plan(x, problem, plan[1][x])
return all(predicate(r) for r in problem.result(state, plan[0]))
run_plan('State_1', vacuum_world, plan)
# ## ONLINE DFS AGENT
# So far, we have seen agents that use __offline search__ algorithms,
# which is a class of algorithms that compute a complete solution before executing it.
# In contrast, an __online search__ agent interleaves computation and action.
# Online search is better for most dynamic environments and necessary for unknown environments.
# <br>
# Online search problems are solved by an agent executing actions, rather than just by pure computation.
# For a fully observable environment, an online agent cycles through three steps: taking an action, computing the step cost and checking if the goal has been reached.
# <br>
# For online algorithms in partially-observable environments, there is usually a tradeoff between exploration and exploitation to be taken care of.
# <br>
# <br>
# Whenever an online agent takes an action, it receives a _percept_ or an observation that tells it something about its immediate environment.
# Using this percept, the agent can augment its map of the current environment.
# For a partially observable environment, this is called the belief state.
# <br>
# Online algorithms expand nodes in a _local_ order, just like _depth-first search_ as it does not have the option of observing farther nodes like _A* search_.
# Whenever an action from the current state has not been explored, the agent tries that action.
# <br>
# Difficulty arises when the agent has tried all actions in a particular state.
# An offline search algorithm would simply drop the state from the queue in this scenario whereas an online search agent has to physically move back to the previous state.
# To do this, the agent needs to maintain a table where it stores the order of nodes it has been to.
# This is how our implementation of _Online DFS-Agent_ works.
# This agent works only in state spaces where the action is reversible, because of the use of backtracking.
# <br>
# Let's have a look at the `OnlineDFSAgent` class.
psource(OnlineDFSAgent)
# It maintains two dictionaries `untried` and `unbacktracked`.
# `untried` contains nodes that have not been visited yet.
# `unbacktracked` contains the sequence of nodes that the agent has visited so it can backtrack to it later, if required.
# `s` and `a` store the state and the action respectively and `result` stores the final path or solution of the problem.
# <br>
# Let's look at another online search algorithm.
# ## LRTA* AGENT
# We can infer now that hill-climbing is an online search algorithm, but it is not very useful natively because for complicated search spaces, it might converge to the local minima and indefinitely stay there.
# In such a case, we can choose to randomly restart it a few times with different starting conditions and return the result with the lowest total cost.
# Sometimes, it is better to use random walks instead of random restarts depending on the problem, but progress can still be very slow.
# <br>
# A better improvement would be to give hill-climbing a memory element.
# We store the current best heuristic estimate and it is updated as the agent gains experience in the state space.
# The estimated optimal cost is made more and more accurate as time passes and each time the the local minima is "flattened out" until we escape it.
# <br>
# This learning scheme is a simple improvement upon traditional hill-climbing and is called _learning real-time A*_ or __LRTA*__.
# Similar to _Online DFS-Agent_, it builds a map of the environment and chooses the best possible move according to its current heuristic estimates.
# <br>
# Actions that haven't been tried yet are assumed to lead immediately to the goal with the least possible cost.
# This is called __optimism under uncertainty__ and encourages the agent to explore new promising paths.
# This algorithm might not terminate if the state space is infinite, unlike A* search.
# <br>
# Let's have a look at the `LRTAStarAgent` class.
psource(LRTAStarAgent)
# `H` stores the heuristic cost of the paths the agent may travel to.
# <br>
# `s` and `a` store the state and the action respectively.
# <br>
# `problem` stores the problem definition and the current map of the environment is stored in `problem.result`.
# <br>
# The `LRTA_cost` method computes the cost of a new path given the current state `s`, the action `a`, the next state `s1` and the estimated cost to get from `s` to `s1` is extracted from `H`.
# Let's use `LRTAStarAgent` to solve a simple problem.
# We'll define a new `LRTA_problem` instance based on our `one_dim_state_space`.
one_dim_state_space
# Let's define an instance of `OnlineSearchProblem`.
LRTA_problem = OnlineSearchProblem('State_3', 'State_5', one_dim_state_space)
# Now we initialize a `LRTAStarAgent` object for the problem we just defined.
lrta_agent = LRTAStarAgent(LRTA_problem)
# We'll pass the percepts `[State_3, State_4, State_3, State_4, State_5]` one-by-one to our agent to see what action it comes up with at each timestep.
lrta_agent('State_3')
lrta_agent('State_4')
lrta_agent('State_3')
lrta_agent('State_4')
# If you manually try to see what the optimal action should be at each step, the outputs of the `lrta_agent` will start to make sense if it doesn't already.
lrta_agent('State_5')
# There is no possible action for this state.
# <br>
# This concludes the notebook.
# Hope you learned something new!
| search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pymongo import MongoClient
from bs4 import BeautifulSoup
import pandas as pd
we_eat_client = MongoClient()
we_eat_db = we_eat_client['we_eat']
website_collection = we_eat_db['websites']
more_business_info_collection = we_eat_db['more_business_info']
# -
website_collection.count_documents({})
a = website_collection.find()[:3]
def parse_bizinfo(bizinfos, alias):
biz = {}
biz['alias'] = alias
for bizinfo in bizinfos:
dt = bizinfo.select_one('dt')
cat_name = dt.text.strip()
dd = bizinfo.select_one('dd')
response = dd.text.strip()
biz[cat_name] = response
return biz
def save_bizinfo(bizinfo_data):
more_business_info_collection.delete_many({
'alias' : bizinfo_data['alias']
})
more_business_info_collection.insert_one(bizinfo_data)
def collect_bizinfo(website_collection):
for website in website_collection.find():
url = website['url']
alias = url.rpartition('/')[2]
html = website['html']
soup = BeautifulSoup(html, 'html.parser')
bizinfos = soup.select('div.short-def-list dl')
print(alias)
data = parse_bizinfo(bizinfos, alias)
save_bizinfo(data)
for website in a:
html = website['html']
soup = BeautifulSoup(html, 'html.parser')
bizinfo = soup.select('div.short-def-list dl')
data = get_biz_info(bizinfo)
website = website_collection.find()[2:3]
html = website[0]['html']
soup = BeautifulSoup(html, 'html.parser')
bizinfo = soup.select('div.short-def-list dl')
bizinfo[0].find_all('dt')
len(bizinfo)
get_biz_info(bizinfo)
# %pdb
# +
#collect_bizinfo(website_collection)
# -
more_business_info_collection.count_documents({})
more_business_info_collection.find_one({'alias' : 'nachyo-average-seattle'})
more_business_info_collection.delete_many({'alias': {'$exists': False}})
biz_dicts = pd.DataFrame(list(more_business_info_collection.find()))
cols = list(biz_dicts.columns)
sorted_cols = cols[-2:] + cols[:-2]
biz_dicts = biz_dicts[sorted_cols]
biz_dicts.columns
bizinfo_df = biz_dicts.drop(['_id'], axis=1).set_index('alias')
bizinfo_df.to_pickle('bizinfo_df.pkl')
bizinfo_df.head()
| notebooks/extract_more_business_info.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <a href="https://cocl.us/NotebooksPython101">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/TopAd.png" width="750" align="center">
# </a>
# </div>
# <a href="https://cognitiveclass.ai/">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/CCLog.png" width="200" align="center">
# </a>
# <h1>2D <code>Numpy</code> in Python</h1>
# <p><strong>Welcome!</strong> This notebook will teach you about using <code>Numpy</code> in the Python Programming Language. By the end of this lab, you'll know what <code>Numpy</code> is and the <code>Numpy</code> operations.</p>
# <h2>Table of Contents</h2>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <ul>
# <li><a href="create">Create a 2D Numpy Array</a></li>
# <li><a href="access">Accessing different elements of a Numpy Array</a></li>
# <li><a href="op">Basic Operations</a></li>
# </ul>
# <p>
# Estimated time needed: <strong>20 min</strong>
# </p>
# </div>
#
# <hr>
# <h2 id="create">Create a 2D Numpy Array</h2>
# +
# Import the libraries
import numpy as np
import matplotlib.pyplot as plt
# -
# Consider the list <code>a</code>, the list contains three nested lists **each of equal size**.
# +
# Create a list
a = [[11, 12, 13], [21, 22, 23], [31, 32, 33]]
a
# -
# We can cast the list to a Numpy Array as follow
# +
# Convert list to Numpy Array
# Every element is the same type
A = np.array(a)
A
# -
# We can use the attribute <code>ndim</code> to obtain the number of axes or dimensions referred to as the rank.
# +
# Show the numpy array dimensions
A.ndim
# -
# Attribute <code>shape</code> returns a tuple corresponding to the size or number of each dimension.
# +
# Show the numpy array shape
A.shape
# -
# The total number of elements in the array is given by the attribute <code>size</code>.
# +
# Show the numpy array size
A.size
# -
# <hr>
# <h2 id="access">Accessing different elements of a Numpy Array</h2>
# We can use rectangular brackets to access the different elements of the array. The correspondence between the rectangular brackets and the list and the rectangular representation is shown in the following figure for a 3x3 array:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoEg.png" width="500" />
# We can access the 2nd-row 3rd column as shown in the following figure:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoFT.png" width="400" />
# We simply use the square brackets and the indices corresponding to the element we would like:
# +
# Access the element on the second row and third column
A[1, 2]
# -
# We can also use the following notation to obtain the elements:
# +
# Access the element on the second row and third column
A[1][2]
# -
# Consider the elements shown in the following figure
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoFF.png" width="400" />
# We can access the element as follows
# +
# Access the element on the first row and first column
A[0][0]
# -
# We can also use slicing in numpy arrays. Consider the following figure. We would like to obtain the first two columns in the first row
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoFSF.png" width="400" />
# This can be done with the following syntax
# +
# Access the element on the first row and first and second columns
A[0][0:2]
# -
# Similarly, we can obtain the first two rows of the 3rd column as follows:
# +
# Access the element on the first and second rows and third column
A[0:2, 2]
# -
# Corresponding to the following figure:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoTST.png" width="400" />
# <hr>
# <h2 id="op">Basic Operations</h2>
# We can also add arrays. The process is identical to matrix addition. Matrix addition of <code>X</code> and <code>Y</code> is shown in the following figure:
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoAdd.png" width="500" />
# The numpy array is given by <code>X</code> and <code>Y</code>
# +
# Create a numpy array X
X = np.array([[1, 0], [0, 1]])
X
# +
# Create a numpy array Y
Y = np.array([[2, 1], [1, 2]])
Y
# -
# We can add the numpy arrays as follows.
# +
# Add X and Y
Z = X + Y
Z
# -
# Multiplying a numpy array by a scaler is identical to multiplying a matrix by a scaler. If we multiply the matrix <code>Y</code> by the scaler 2, we simply multiply every element in the matrix by 2 as shown in the figure.
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoDb.png" width="500" />
# We can perform the same operation in numpy as follows
# +
# Create a numpy array Y
Y = np.array([[2, 1], [1, 2]])
Y
# +
# Multiply Y with 2
Z = 2 * Y
Z
# -
# Multiplication of two arrays corresponds to an element-wise product or Hadamard product. Consider matrix <code>X</code> and <code>Y</code>. The Hadamard product corresponds to multiplying each of the elements in the same position, i.e. multiplying elements contained in the same color boxes together. The result is a new matrix that is the same size as matrix <code>Y</code> or <code>X</code>, as shown in the following figure.
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Chapter%205/Images/NumTwoMul.png" width="500" />
# We can perform element-wise product of the array <code>X</code> and <code>Y</code> as follows:
# +
# Create a numpy array Y
Y = np.array([[2, 1], [1, 2]])
Y
# +
# Create a numpy array X
X = np.array([[1, 0], [0, 1]])
X
# +
# Multiply X with Y
Z = X * Y
Z
# -
# We can also perform matrix multiplication with the numpy arrays <code>A</code> and <code>B</code> as follows:
# First, we define matrix <code>A</code> and <code>B</code>:
# +
# Create a matrix A
A = np.array([[0, 1, 1], [1, 0, 1]])
A
# +
# Create a matrix B
B = np.array([[1, 1], [1, 1], [-1, 1]])
B
# -
# We use the numpy function <code>dot</code> to multiply the arrays together.
# +
# Calculate the dot product
Z = np.dot(A,B)
Z
# +
# Calculate the sine of Z
np.sin(Z)
# -
# We use the numpy attribute <code>T</code> to calculate the transposed matrix
# +
# Create a matrix C
C = np.array([[1,1],[2,2],[3,3]])
C
# +
# Get the transposed of C
C.T
# -
# <hr>
# <h2>The last exercise!</h2>
# <p>Congratulations, you have completed your first lesson and hands-on lab in Python. However, there is one more thing you need to do. The Data Science community encourages sharing work. The best way to share and showcase your work is to share it on GitHub. By sharing your notebook on GitHub you are not only building your reputation with fellow data scientists, but you can also show it off when applying for a job. Even though this was your first piece of work, it is never too early to start building good habits. So, please read and follow <a href="https://cognitiveclass.ai/blog/data-scientists-stand-out-by-sharing-your-notebooks/" target="_blank">this article</a> to learn how to share your work.
# <hr>
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <h2>Get IBM Watson Studio free of charge!</h2>
# <p><a href="https://cocl.us/NotebooksPython101bottom"><img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/PY0101EN/Ad/BottomAd.png" width="750" align="center"></a></p>
# </div>
# <h3>About the Authors:</h3>
# <p><a href="https://www.linkedin.com/in/joseph-s-50398b136/" target="_blank"><NAME></a> is a Data Scientist at IBM, and holds a PhD in Electrical Engineering. His research focused on using Machine Learning, Signal Processing, and Computer Vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.</p>
# Other contributors: <a href="www.linkedin.com/in/jiahui-mavis-zhou-a4537814a"><NAME></a>
# <hr>
# <p>Copyright © 2018 IBM Developer Skills Network. This notebook and its source code are released under the terms of the <a href="https://cognitiveclass.ai/mit-license/">MIT License</a>.</p>
| 5.2-Numpy2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple RNN Encode-Decoder for Translation
#
# **Learning Objectives**
# 1. Learn how to create a tf.data.Dataset for seq2seq problems
# 1. Learn how to train an encoder-decoder model in Keras
# 1. Learn how to save the encoder and the decoder as separate models
# 1. Learn how to piece together the trained encoder and decoder into a translation function
# 1. Learn how to use the BLUE score to evaluate a translation model
#
# ## Introduction
#
# In this lab we'll build a translation model from Spanish to English using a RNN encoder-decoder model architecture.
# We will start by creating train and eval datasets (using the `tf.data.Dataset` API) that are typical for seq2seq problems. Then we will use the Keras functional API to train an RNN encoder-decoder model, which will save as two separate models, the encoder and decoder model. Using these two separate pieces we will implement the translation function.
# At last, we'll benchmark our results using the industry standard BLEU score.
# +
import os
import pickle
import sys
import nltk
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow.keras.layers import (
Dense,
Embedding,
GRU,
Input,
)
from tensorflow.keras.models import (
load_model,
Model,
)
import utils_preproc
print(tf.__version__)
# -
SEED = 0
MODEL_PATH = 'translate_models/baseline'
DATA_URL = 'http://storage.googleapis.com/download.tensorflow.org/data/spa-eng.zip'
LOAD_CHECKPOINT = False
tf.random.set_seed(SEED)
# ## Downloading the Data
# We'll use a language dataset provided by http://www.manythings.org/anki/. The dataset contains Spanish-English translation pairs in the format:
#
# ```
# May I borrow this book? ¿Puedo tomar prestado este libro?
# ```
#
# The dataset is a curated list of 120K translation pairs from http://tatoeba.org/, a platform for community contributed translations by native speakers.
# +
path_to_zip = tf.keras.utils.get_file(
'spa-eng.zip', origin=DATA_URL, extract=True)
path_to_file = os.path.join(
os.path.dirname(path_to_zip),
"spa-eng/spa.txt"
)
print("Translation data stored at:", path_to_file)
# -
data = pd.read_csv(
path_to_file, sep='\t', header=None, names=['english', 'spanish'])
data.sample(3)
# From the `utils_preproc` package we have written for you,
# we will use the following functions to pre-process our dataset of sentence pairs.
# ## Sentence Preprocessing
# The `utils_preproc.preprocess_sentence()` method does the following:
# 1. Converts sentence to lower case
# 2. Adds a space between punctuation and words
# 3. Replaces tokens that aren't a-z or punctuation with space
# 4. Adds `<start>` and `<end>` tokens
#
# For example:
raw = [
"No estamos comiendo.",
"Está llegando el invierno.",
"El invierno se acerca.",
"Tom no comio nada.",
"Su pierna mala le impidió ganar la carrera.",
"Su respuesta es erronea.",
"¿Qué tal si damos un paseo después del almuerzo?"
]
processed = [utils_preproc.preprocess_sentence(s) for s in raw]
processed
# ## Sentence Integerizing
# The `utils_preproc.tokenize()` method does the following:
#
# 1. Splits each sentence into a token list
# 1. Maps each token to an integer
# 1. Pads to length of longest sentence
#
# It returns an instance of a [Keras Tokenizer](https://keras.io/preprocessing/text/)
# containing the token-integer mapping along with the integerized sentences:
integerized, tokenizer = utils_preproc.tokenize(processed)
integerized
# The outputted tokenizer can be used to get back the actual works
# from the integers representing them:
tokenizer.sequences_to_texts(integerized)
# ## Creating the tf.data.Dataset
# ### `load_and_preprocess`
# Let's first implement a function that will read the raw sentence-pair file
# and preprocess the sentences with `utils_preproc.preprocess_sentence`.
#
# The `load_and_preprocess` function takes as input
# - the path where the sentence-pair file is located
# - the number of examples one wants to read in
#
# It returns a tuple whose first component contains the english
# preprocessed sentences, while the second component contains the
# spanish ones:
def load_and_preprocess(path, num_examples):
with open(path_to_file, 'r') as fp:
lines = fp.read().strip().split('\n')
# TODO 1a
sentence_pairs = [
[utils_preproc.preprocess_sentence(sent) for sent in line.split('\t')]
for line in lines[:num_examples]
]
return zip(*sentence_pairs)
# +
en, sp = load_and_preprocess(path_to_file, num_examples=10)
print(en[-1])
print(sp[-1])
# -
# ### `load_and_integerize`
# Using `utils_preproc.tokenize`, let us now implement the function `load_and_integerize` that takes as input the data path along with the number of examples we want to read in and returns the following tuple:
#
# ```python
# (input_tensor, target_tensor, inp_lang_tokenizer, targ_lang_tokenizer)
# ```
#
# where
#
#
# * `input_tensor` is an integer tensor of shape `(num_examples, max_length_inp)` containing the integerized versions of the source language sentences
# * `target_tensor` is an integer tensor of shape `(num_examples, max_length_targ)` containing the integerized versions of the target language sentences
# * `inp_lang_tokenizer` is the source language tokenizer
# * `targ_lang_tokenizer` is the target language tokenizer
def load_and_integerize(path, num_examples=None):
targ_lang, inp_lang = load_and_preprocess(path, num_examples)
# TODO 1b
input_tensor, inp_lang_tokenizer = utils_preproc.tokenize(inp_lang)
target_tensor, targ_lang_tokenizer = utils_preproc.tokenize(targ_lang)
return input_tensor, target_tensor, inp_lang_tokenizer, targ_lang_tokenizer
# ### Train and eval splits
# We'll split this data 80/20 into train and validation, and we'll use only the first 30K examples, since we'll be training on a single GPU.
#
# Let us set variable for that:
TEST_PROP = 0.2
NUM_EXAMPLES = 30000
# Now let's load and integerize the sentence paris and store the tokenizer for the source and the target language into the `int_lang` and `targ_lang` variable respectively:
input_tensor, target_tensor, inp_lang, targ_lang = load_and_integerize(
path_to_file, NUM_EXAMPLES)
# Let us store the maximal sentence length of both languages into two variables:
max_length_targ = target_tensor.shape[1]
max_length_inp = input_tensor.shape[1]
# We are now using scikit-learn `train_test_split` to create our splits:
# +
splits = train_test_split(
input_tensor, target_tensor, test_size=TEST_PROP, random_state=SEED)
input_tensor_train = splits[0]
input_tensor_val = splits[1]
target_tensor_train = splits[2]
target_tensor_val = splits[3]
# -
# Let's make sure the number of example in each split looks good:
(len(input_tensor_train), len(target_tensor_train),
len(input_tensor_val), len(target_tensor_val))
# The `utils_preproc.int2word` function allows you to transform back the integerized sentences into words. Note that the `<start>` token is alwasy encoded as `1`, while the `<end>` token is always encoded as `0`:
# +
print("Input Language; int to word mapping")
print(input_tensor_train[0])
print(utils_preproc.int2word(inp_lang, input_tensor_train[0]), '\n')
print("Target Language; int to word mapping")
print(target_tensor_train[0])
print(utils_preproc.int2word(targ_lang, target_tensor_train[0]))
# -
# ### Create tf.data dataset for train and eval
# Below we implement the `create_dataset` function that takes as input
# * `encoder_input` which is an integer tensor of shape `(num_examples, max_length_inp)` containing the integerized versions of the source language sentences
# * `decoder_input` which is an integer tensor of shape `(num_examples, max_length_targ)`containing the integerized versions of the target language sentences
#
# It returns a `tf.data.Dataset` containing examples for the form
#
# ```python
# ((source_sentence, target_sentence), shifted_target_sentence)
# ```
#
# where `source_sentence` and `target_setence` are the integer version of source-target language pairs and `shifted_target` is the same as `target_sentence` but with indices shifted by 1.
#
# **Remark:** In the training code, `source_sentence` (resp. `target_sentence`) will be fed as the encoder (resp. decoder) input, while `shifted_target` will be used to compute the cross-entropy loss by comparing the decoder output with the shifted target sentences.
def create_dataset(encoder_input, decoder_input):
# TODO 1c
# shift ahead by 1
target = tf.roll(decoder_input, -1, 1)
# replace last column with 0s
zeros = tf.zeros([target.shape[0], 1], dtype=tf.int32)
target = tf.concat((target[:, :-1], zeros), axis=-1)
dataset = tf.data.Dataset.from_tensor_slices(
((encoder_input, decoder_input), target))
return dataset
# Let's now create the actual train and eval dataset using the function above:
BUFFER_SIZE = len(input_tensor_train)
BATCH_SIZE = 64
# +
train_dataset = create_dataset(
input_tensor_train, target_tensor_train).shuffle(
BUFFER_SIZE).repeat().batch(BATCH_SIZE, drop_remainder=True)
eval_dataset = create_dataset(
input_tensor_val, target_tensor_val).batch(
BATCH_SIZE, drop_remainder=True)
# -
# ## Training the RNN encoder-decoder model
# We use an encoder-decoder architecture, however we embed our words into a latent space prior to feeding them into the RNN.
# +
EMBEDDING_DIM = 256
HIDDEN_UNITS = 1024
INPUT_VOCAB_SIZE = len(inp_lang.word_index) + 1
TARGET_VOCAB_SIZE = len(targ_lang.word_index) + 1
# -
# Let's implement the encoder network with Keras functional API. It will
# * start with an `Input` layer that will consume the source language integerized sentences
# * then feed them to an `Embedding` layer of `EMBEDDING_DIM` dimensions
# * which in turn will pass the embeddings to a `GRU` recurrent layer with `HIDDEN_UNITS`
#
# The output of the encoder will be the `encoder_outputs` and the `encoder_state`.
# +
encoder_inputs = Input(shape=(None,), name="encoder_input")
# TODO 2a
encoder_inputs_embedded = Embedding(
input_dim=INPUT_VOCAB_SIZE,
output_dim=EMBEDDING_DIM,
input_length=max_length_inp)(encoder_inputs)
encoder_rnn = GRU(
units=HIDDEN_UNITS,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
encoder_outputs, encoder_state = encoder_rnn(encoder_inputs_embedded)
# -
# We now implement the decoder network, which is very similar to the encoder network.
#
# It will
# * start with an `Input` layer that will consume the source language integerized sentences
# * then feed that input to an `Embedding` layer of `EMBEDDING_DIM` dimensions
# * which in turn will pass the embeddings to a `GRU` recurrent layer with `HIDDEN_UNITS`
#
# **Important:** The main difference with the encoder, is that the recurrent `GRU` layer will take as input not only the decoder input embeddings, but also the `encoder_state` as outputted by the encoder above. This is where the two networks are linked!
#
# The output of the encoder will be the `decoder_outputs` and the `decoder_state`.
# +
decoder_inputs = Input(shape=(None,), name="decoder_input")
# TODO 2b
decoder_inputs_embedded = Embedding(
input_dim=TARGET_VOCAB_SIZE,
output_dim=EMBEDDING_DIM,
input_length=max_length_targ)(decoder_inputs)
decoder_rnn = GRU(
units=HIDDEN_UNITS,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
decoder_outputs, decoder_state = decoder_rnn(
decoder_inputs_embedded, initial_state=encoder_state)
# -
# The last part of the encoder-decoder architecture is a softmax `Dense` layer that will create the next word probability vector or next word `predictions` from the `decoder_output`:
# +
decoder_dense = Dense(TARGET_VOCAB_SIZE, activation='softmax')
predictions = decoder_dense(decoder_outputs)
# -
# To be able to train the encoder-decoder network defined above, we now need to create a trainable Keras `Model` by specifying which are the `inputs` and the `outputs` of our problem. They should correspond exactly to what the type of input/output in our train and eval `tf.data.Dataset` since that's what will be fed to the `inputs` and `outputs` we declare while instantiating the Keras `Model`.
#
# While compiling our model, we should make sure that the loss is the `sparse_categorical_crossentropy` so that we can compare the true word indices for the target language as outputted by our train `tf.data.Dataset` with the next word `predictions` vector as outputted by the decoder:
# +
# TODO 2c
model = Model(inputs=[encoder_inputs, decoder_inputs], outputs=predictions)
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy')
model.summary()
# -
# Let's now train the model!
# +
STEPS_PER_EPOCH = len(input_tensor_train)//BATCH_SIZE
EPOCHS = 1
history = model.fit(
train_dataset,
steps_per_epoch=STEPS_PER_EPOCH,
validation_data=eval_dataset,
epochs=EPOCHS
)
# -
# ## Implementing the translation (or decoding) function
#
# We can't just use model.predict(), because we don't know all the inputs we used during training. We only know the encoder_input (source language) but not the decoder_input (target language), which is what we want to predict (i.e., the translation of the source language)!
#
# We do however know the first token of the decoder input, which is the `<start>` token. So using this plus the state of the encoder RNN, we can predict the next token. We will then use that token to be the second token of decoder input, and continue like this until we predict the `<end>` token, or we reach some defined max length.
#
# So, the strategy now is to split our trained network into two independent Keras models:
#
# * an **encoder model** with signature `encoder_inputs -> encoder_state`
# * a **decoder model** with signature `[decoder_inputs, decoder_state_input] -> [predictions, decoder_state]`
#
# This way, we will be able to encode the source language sentence into the vector `encoder_state` using the encoder and feed it to the decoder model along with the `<start>` token at step 1.
#
# Given that input, the decoder will produce the first word of the translation, by sampling from the `predictions` vector (for simplicity, our sampling strategy here will be to take the next word to be the one whose index has the maximum probability in the `predictions` vector) along with a new state vector, the `decoder_state`.
#
# At this point, we can feed again to the decoder the predicted first word and as well as the new `decoder_state` to predict the translation second word.
#
# This process can be continued until the decoder produces the token `<stop>`.
#
# This is how we will implement our translation (or decoding) function, but let us first extract a separate encoder and a separate decoder from our trained encoder-decoder model.
#
#
# **Remark:** If we have already trained and saved the models (i.e, `LOAD_CHECKPOINT` is `True`) we will just load the models, otherwise, we extract them from the trained network above by explicitly creating the encoder and decoder Keras `Model`s with the signature we want.
#
#
# +
if LOAD_CHECKPOINT:
encoder_model = load_model(os.path.join(MODEL_PATH, 'encoder_model.h5'))
decoder_model = load_model(os.path.join(MODEL_PATH, 'decoder_model.h5'))
else:
# TODO 3a
encoder_model = Model(inputs=encoder_inputs, outputs=encoder_state)
decoder_state_input = Input(shape=(HIDDEN_UNITS,), name="decoder_state_input")
# Reuses weights from the decoder_rnn layer
decoder_outputs, decoder_state = decoder_rnn(
decoder_inputs_embedded, initial_state=decoder_state_input)
# Reuses weights from the decoder_dense layer
predictions = decoder_dense(decoder_outputs)
decoder_model = Model(
inputs=[decoder_inputs, decoder_state_input],
outputs=[predictions, decoder_state]
)
# -
# Now that we have a separate encoder and a separate decoder, let's implement a translation function, to which we will give the generic name of `decode_sequences` (to stress that this procedure is general to all seq2seq problems).
#
# `decode_sequences` will take as input
# * `input_seqs` which is the integerized source language sentence tensor that the encoder can consume
# * `output_tokenizer` which is the target languague tokenizer we will need to extract back words from predicted word integers
# * `max_decode_length` which is the length after which we stop decoding if the `<stop>` token has not been predicted
#
#
# **Note**: Now that the encoder and decoder have been turned into Keras models, to feed them their input, we need to use the `.predict` method.
def decode_sequences(input_seqs, output_tokenizer, max_decode_length=50):
"""
Arguments:
input_seqs: int tensor of shape (BATCH_SIZE, SEQ_LEN)
output_tokenizer: Tokenizer used to conver from int to words
Returns translated sentences
"""
# Encode the input as state vectors.
states_value = encoder_model.predict(input_seqs)
# Populate the first character of target sequence with the start character.
batch_size = input_seqs.shape[0]
target_seq = tf.ones([batch_size, 1])
decoded_sentences = [[] for _ in range(batch_size)]
# TODO 4: Sampling loop
for i in range(max_decode_length):
output_tokens, decoder_state = decoder_model.predict(
[target_seq, states_value])
# Sample a token
sampled_token_index = np.argmax(output_tokens[:, -1, :], axis=-1)
tokens = utils_preproc.int2word(output_tokenizer, sampled_token_index)
for j in range(batch_size):
decoded_sentences[j].append(tokens[j])
# Update the target sequence (of length 1).
target_seq = tf.expand_dims(tf.constant(sampled_token_index), axis=-1)
# Update states
states_value = decoder_state
return decoded_sentences
# Now we're ready to predict!
# +
sentences = [
"No estamos comiendo.",
"Está llegando el invierno.",
"El invierno se acerca.",
"Tom no comio nada.",
"Su pierna mala le impidió ganar la carrera.",
"Su respuesta es erronea.",
"¿Qué tal si damos un paseo después del almuerzo?"
]
reference_translations = [
"We're not eating.",
"Winter is coming.",
"Winter is coming.",
"Tom ate nothing.",
"His bad leg prevented him from winning the race.",
"Your answer is wrong.",
"How about going for a walk after lunch?"
]
machine_translations = decode_sequences(
utils_preproc.preprocess(sentences, inp_lang),
targ_lang,
max_length_targ
)
for i in range(len(sentences)):
print('-')
print('INPUT:')
print(sentences[i])
print('REFERENCE TRANSLATION:')
print(reference_translations[i])
print('MACHINE TRANSLATION:')
print(machine_translations[i])
# -
# ### Checkpoint Model
# Now let's us save the full training encoder-decoder model, as well as the separate encoder and decoder model to disk for latter reuse:
if not LOAD_CHECKPOINT:
os.makedirs(MODEL_PATH, exist_ok=True)
# TODO 3b
model.save(os.path.join(MODEL_PATH, 'model.h5'))
encoder_model.save(os.path.join(MODEL_PATH, 'encoder_model.h5'))
decoder_model.save(os.path.join(MODEL_PATH, 'decoder_model.h5'))
with open(os.path.join(MODEL_PATH, 'encoder_tokenizer.pkl'), 'wb') as fp:
pickle.dump(inp_lang, fp)
with open(os.path.join(MODEL_PATH, 'decoder_tokenizer.pkl'), 'wb') as fp:
pickle.dump(targ_lang, fp)
# ## Evaluation Metric (BLEU)
#
# Unlike say, image classification, there is no one right answer for a machine translation. However our current loss metric, cross entropy, only gives credit when the machine translation matches the exact same word in the same order as the reference translation.
#
# Many attempts have been made to develop a better metric for natural language evaluation. The most popular currently is Bilingual Evaluation Understudy (BLEU).
#
# - It is quick and inexpensive to calculate.
# - It allows flexibility for the ordering of words and phrases.
# - It is easy to understand.
# - It is language independent.
# - It correlates highly with human evaluation.
# - It has been widely adopted.
#
# The score is from 0 to 1, where 1 is an exact match.
#
# It works by counting matching n-grams between the machine and reference texts, regardless of order. BLUE-4 counts matching n grams from 1-4 (1-gram, 2-gram, 3-gram and 4-gram). It is common to report both BLUE-1 and BLUE-4
#
# It still is imperfect, since it gives no credit to synonyms and so human evaluation is still best when feasible. However BLEU is commonly considered the best among bad options for an automated metric.
#
# The NLTK framework has an implementation that we will use.
#
# We can't run calculate BLEU during training, because at that time the correct decoder input is used. Instead we'll calculate it now.
#
# For more info: https://machinelearningmastery.com/calculate-bleu-score-for-text-python/
def bleu_1(reference, candidate):
reference = list(filter(lambda x: x != '', reference)) # remove padding
candidate = list(filter(lambda x: x != '', candidate)) # remove padding
smoothing_function = nltk.translate.bleu_score.SmoothingFunction().method1
return nltk.translate.bleu_score.sentence_bleu(
reference, candidate, (1,), smoothing_function)
def bleu_4(reference, candidate):
reference = list(filter(lambda x: x != '', reference)) # remove padding
candidate = list(filter(lambda x: x != '', candidate)) # remove padding
smoothing_function = nltk.translate.bleu_score.SmoothingFunction().method1
return nltk.translate.bleu_score.sentence_bleu(
reference, candidate, (.25, .25, .25, .25), smoothing_function)
# Let's now average the `bleu_1` and `bleu_4` scores for all the sentence pairs in the eval set. The next cell takes some time to run, the bulk of which is decoding the 6000 sentences in the validation set. Please wait unitl completes.
# +
# %%time
num_examples = len(input_tensor_val)
bleu_1_total = 0
bleu_4_total = 0
for idx in range(num_examples):
# TODO 5
reference_sentence = utils_preproc.int2word(
targ_lang, target_tensor_val[idx][1:])
decoded_sentence = decode_sequences(
input_tensor_val[idx:idx+1], targ_lang, max_length_targ)[0]
bleu_1_total += bleu_1(reference_sentence, decoded_sentence)
bleu_4_total += bleu_4(reference_sentence, decoded_sentence)
print('BLEU 1: {}'.format(bleu_1_total/num_examples))
print('BLEU 4: {}'.format(bleu_4_total/num_examples))
# -
# ## Results
#
# **Hyperparameters**
#
# - Batch_Size: 64
# - Optimizer: adam
# - Embed_dim: 256
# - GRU Units: 1024
# - Train Examples: 24,000
# - Epochs: 10
# - Hardware: P100 GPU
#
# **Performance**
# - Training Time: 5min
# - Cross-entropy loss: train: 0.0722 - val: 0.9062
# - BLEU 1: 0.2519574312515255
# - BLEU 4: 0.04589972764144636
# ### References
#
# - <NAME>: https://github.com/keras-team/keras/blob/master/examples/lstm_seq2seq.py
#
| notebooks/text_classification/solutions/5_rnn_encoder_decoder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Apache Toree - Scala
# language: scala
# name: apache_toree_scala
# ---
# # Plongeur
#
# A *topological data analysis* library.
#
# > Core algorithm written in [Scala](http://www.scala-lang.org/), using Apache [Spark](http://spark.apache.org/).
# >
# > Executed in a [Jupyter](http://jupyter.org/) notebook, using the Apache [Toree](https://github.com/apache/incubator-toree) kernel and [declarative widgets](http://jupyter-incubator.github.io/declarativewidgets/docs.html).
# >
# > Graphs rendered with [Sigma](http://sigmajs.org/)/[Linkurious](https://github.com/Linkurious/linkurious.js), wrapped in a [Polymer](https://www.polymer-project.org/1.0/) component.
# >
# > Reactive machinery powered by [Rx](http://reactivex.io/) [RxScala](https://github.com/ReactiveX/RxScala).
#
# + [markdown] nbpresent={"id": "f5ac5e2a-9a1a-46ab-8882-2a492fbebc19"}
# #### Maven dependencies
# + nbpresent={"id": "c3144ed8-e62e-4316-917e-26b8904edc72"}
// %AddDeps org.apache.spark spark-mllib_2.10 1.6.2 --repository file:/Users/tmo/.m2/repository
// %AddDeps org.scalanlp breeze_2.10 0.11.2 --transitive
// %AddDeps org.scalanlp breeze-natives_2.10 0.11.2
// %AddDeps org.scalanlp breeze-macros_2.10 0.11.2
// %AddDeps com.github.haifengl smile-core 1.2.0 --transitive
// %AddDeps com.github.karlhigley spark-neighbors_2.10 0.3.6-FORK --repository file:/Users/tmo/.m2/repository
// %AddDeps io.reactivex rxscala_2.10 0.26.1 --transitive --repository file:/Users/tmo/.m2/repository
// %AddDeps com.softwaremill.quicklens quicklens_2.10 1.4.4 --repository file:/Users/tmo/.m2/repository
// %AddDeps org.tmoerman plongeur-spark_2.10 0.3.51 --repository file:/Users/tmo/.m2/repository
# + nbpresent={"id": "70465288-c658-4f45-a709-4e6dd0f9a390"}
// %addjar http://localhost:8888/nbextensions/declarativewidgets/declarativewidgets.jar
# + [markdown] nbpresent={"id": "ff4512c4-eb02-43ff-8911-26c511ea96b4"}
# #### Import classes
# + nbpresent={"id": "800b012e-aa8e-4989-a156-a0b1f1e34391"}
import rx.lang.scala.{Observer, Subscription, Observable}
import rx.lang.scala.subjects.PublishSubject
import rx.lang.scala.subjects._
import org.apache.commons.lang.StringUtils.trim
import org.apache.spark.SparkContext
import org.apache.spark.mllib.linalg.Vectors.dense
import org.apache.spark.rdd.RDD
import org.joda.time.DateTime
import org.tmoerman.plongeur.tda.TDAMachine
import org.tmoerman.plongeur.tda.Distances._
import org.tmoerman.plongeur.tda.Model._
import org.tmoerman.plongeur.tda.Filters._
import org.tmoerman.plongeur.tda.cluster.Clustering._
import org.tmoerman.plongeur.tda.cluster.Scale._
import org.tmoerman.plongeur.tda.Colour._
import org.tmoerman.plongeur.tda.Brewer
import org.tmoerman.plongeur.tda.LSH.LSHParams
import org.tmoerman.plongeur.tda.Model.{DataPoint, TDAContext, dp}
import org.tmoerman.plongeur.tda.knn.FastKNN.FastKNNParams
import org.tmoerman.plongeur.tda.knn.SampledKNN.SampledKNNParams
import org.tmoerman.plongeur.tda.knn.{FastKNN, SampledKNN, _}
import org.tmoerman.plongeur.util.RDDFunctions._
import org.tmoerman.plongeur.util.TimeUtils.time
import org.tmoerman.plongeur.tda.geometry.Laplacian._
import breeze.stats.distributions._
import org.apache.spark.mllib.linalg.SparseMatrix
# +
import declarativewidgets._
initWidgets
import declarativewidgets.WidgetChannels.channel
# +
import java.util.concurrent.atomic.AtomicReference
case class SubRef(val ref: AtomicReference[Option[Subscription]] = new AtomicReference[Option[Subscription]](None)) extends Serializable {
def update(sub: Subscription): Unit = ref.getAndSet(Option(sub)).foreach(old => old.unsubscribe())
def reset(): Unit = update(null)
}
# + [markdown] nbpresent={"id": "17926a43-50f9-444a-97a5-84eaf41caa6a"}
# #### Import polymer elements
#
# These cells triggers Bower installations of the specified web components.
#
# If it doesn't work, check whether Bower has sufficient permissions to install in the jupyter `/nbextensions` folder.
# + nbpresent={"id": "282de503-679a-43a3-af20-3002bd068851"} language="html"
# <link rel='import' href='urth_components/paper-slider/paper-slider.html'
# is='urth-core-import' package='PolymerElements/paper-slider'>
# <link rel='import' href='urth_components/paper-button/paper-button.html'
# is='urth-core-import' package='PolymerElements/paper-button'>
# <link rel='import' href='urth_components/plongeur-graph/plongeur-graph.html'
# is='urth-core-import' package='tmoerman/plongeur-graph'>
# <link rel='import' href='urth_components/urth-viz-scatter/urth-viz-scatter.html' is='urth-core-import'>
# + [markdown] nbpresent={"id": "3ee8048e-d1e3-4c46-99cc-5a3bc5c7f865"}
# #### Reactive TDA Machine
# -
# Keep references to Rx subscriptions apart.
val in$_subRef = SubRef()
# + [markdown] nbpresent={"id": "40c9258a-bbab-40d7-95ca-84b622b5951e"}
# Instantiate a `PublishSubject`. This stream of `TDAParams` instances represents the input of a `TDAMachine`. The `PublishSubject` listens to changes and sets these to the channel `"ch_TDA_1"` under the `"params"` key.
#
# *TODO: unsubscribe previous on re-evaluation*
# + nbpresent={"id": "53be8369-a4fb-4105-8e5b-808067019bb2"}
val in$ = PublishSubject[TDAParams]
in$_subRef.update(in$.subscribe(p => channel("ch_TDA_1").set("params", p.toString)))
# + [markdown] nbpresent={"id": "0a444806-7064-47e9-89f3-ed1313a47c74"}
# Create an initial `TDAParams` instance. In the same cell, we submit the instance to the `PublishSubject`.
# + [markdown] nbpresent={"id": "d23779c0-a7e1-4dbf-865b-2609f499a45f"}
# For the sake of illustration, we create an html snippet that listens to changes on the `"ch_TDA_1"` channel and displays the value of the `"params"` key.
# + nbpresent={"id": "6c896ce2-45c9-44fb-896e-d2144d14e08e"} language="html"
# <template is='urth-core-bind' channel='ch_TDA_1'>
# <div style='background: #FFB; padding: 10px;'>
# <span style='font-family: "Courier"'>[[params]]</span>
# </div>
# </template>
# + [markdown] nbpresent={"id": "b58f5933-8e2b-445c-92f6-27589989e0ee"}
# Notice that when we evaluate the `TDAParams` instantiation cells, the output of the yellow box changes.
# + [markdown] nbpresent={"id": "407e489d-0988-457e-a68b-5b13abe99c0d"}
# #### Inititalize rdd
#
# In this example, we are using a synthetic torus-shaped 2D data set.
# + nbpresent={"id": "f95081ee-5623-4d18-ac6f-24ee227e225e"}
import org.apache.spark.rdd.RDD
import org.apache.commons.lang.StringUtils.trim
import org.apache.spark.mllib.linalg.Vectors.dense
def readMixture(file: String): RDD[DataPoint] = {
sc
.textFile(file)
.zipWithIndex
.map{ case (line, idx) =>
val columns = line.split(",").map(trim)
val category = columns.head
val features = columns.tail.map(_.toDouble)
dp(idx, dense(features), Map("cat" -> category)) }
}
# + nbpresent={"id": "44093436-255d-4f4e-8a87-eef04e88a076"}
val data_path = "/Users/tmo/Work/batiskav/projects/plongeur/scala/plongeur-spark/src/test/resources/data/"
val mixture_path = data_path + "mixture.1000.2.csv"
val rdd = readMixture(mixture_path).cache
val ctx = TDAContext(sc, rdd)
# -
rdd.count
# + [markdown] nbpresent={"id": "5f060023-6ac5-4d84-9e36-8bcf7e8a4710"}
# Turn a TDAResult into a data structure.
# + nbpresent={"id": "4ee5b503-b74a-47c4-8f48-452d1f42e746"}
val r = scala.util.Random
def format(result: TDAResult) = Map(
"nodes" -> result.clusters.map(c =>
Map(
"id" -> c.id.toString,
"size" -> c.dataPoints.size,
"color" -> c.colours.headOption.getOrElse("#000000"),
"x" -> r.nextInt(100),
"y" -> r.nextInt(100))),
"edges" -> result.edges.map(e => {
val (from, to) = e.toArray match {case Array(f, t) => (f, t)}
Map(
"id" -> s"$from--$to",
"source" -> from.toString,
"target" -> to.toString)}))
# -
# Run the machine, obtaining an `Observable` of `TDAResult` instances
# + nbpresent={"id": "b59014a4-e289-4011-b1ab-8667dd1c893f"}
val out$: Observable[TDAResult] = TDAMachine.run(ctx, in$)
# -
val out$_subRef = SubRef()
# + nbpresent={"id": "8b43b0a5-55ee-4fdb-bcfc-4cca3df27a04"}
out$_subRef.update(
out$.subscribe(
onNext = (r) => channel("ch_TDA_1").set("result", format(r)),
onError = (e) => println("Error in TDA machine: ", e)))
# + [markdown] nbpresent={"id": "f329dd44-6902-4dc9-8a6a-77b00475a612"}
# #### Reactive inputs
#
# First, we set up a stream of updates to BASE TDAParams instance.
# -
val pipe$_subRef = SubRef()
import org.tmoerman.plongeur.ui.Controls._
kernel.magics.html(controlsCSS)
# +
import TDAParams._
val den = Filter(Density(sigma=1.0), 30, 0.30)
val pc0 = Filter(Feature(0), 20, 0.3)
val pc1 = Filter(Feature(1), 20, 0.3)
val selector = (d: DataPoint) => d.meta.get("cat")
val maxFq = ClusterMaxFrequency(Array("#F00", "#00F", "#999"), selector)
val avgFilterValue = AverageFilterValue(Brewer.palettes("PuOr")(9), den)
val BASE =
TDAParams(
lens = TDALens(pc0, pc1),
clusteringParams = ClusteringParams(),
scaleSelection = firstGap(5),
collapseDuplicateClusters = false,
colouring = Nop())
in$.onNext(BASE)
# + language="html"
# <template is='urth-core-bind' channel='ch_TDA_1'>
# <plongeur-graph height="600" data="{{result}}"></plongeur-graph>
# </template>
# -
val (sub, html) = BASE.makeControls(channel("ch_TDA_1"), in$)
pipe$_subRef.update(sub)
kernel.magics.html(html)
# +
val rawData = rdd.
map(dp => dp.features.toArray.toList).collect.toList
rawData.take(1)
# -
channel("data").set("raw", rawData)
# + language="html"
# <template is='urth-core-bind' channel='data'>
# <urth-viz-scatter
# datarows='[[raw]]'
# primary='0'
# secondary='1'
# />
# </template>
# -
| scala/plongeur-spark/plongeur-nb/Plongeur Mixture.2 - Density 0.3.51.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import os
import netCDF4
import numpy as np
import math
from scipy.interpolate import griddata
import matplotlib.pyplot as plt
import gc
import re
from collections import OrderedDict
from geophys_utils import NetCDFGridUtils
from geophys_utils import NetCDFLineUtils
from geophys_utils import get_gdal_wcs_dataset, get_gdal_grid_values
from geophys_utils import get_spatial_ref_from_wkt, get_coordinate_transformation, get_utm_wkt, transform_coords
from geophys_utils._transect_utils import line_length, point_along_line, utm_coords, coords2distance, sample_transect
# +
# Setup proxy as required
GA_STAFF_WIFI = False
if GA_STAFF_WIFI:
os.environ['http_proxy'] = 'http://proxy.inno.lan:3128'
os.environ['https_proxy'] = 'http://proxy.inno.lan:3128'
# -
###### AEM links will be updated when available ######
aem_nc_path = '/g/data2/uc0/rr2_dev/rcb547/AEM_examples/AUS_10008_WestK_LCI.nc'
if not os.path.isfile(aem_nc_path):
aem_nc_path = 'http://dapds00.nci.org.au/thredds/dodsC/uc0/rr2_dev/rcb547/AEM_examples/AUS_10008_WestK_LCI.nc'
aem_nc_dataset = netCDF4.Dataset(aem_nc_path + '#fillmismatch') # Note work-around for bad _FillValue: https://github.com/Unidata/netcdf-c/issues/1299
# The CRS definition in the file is INCORRECT in the test file! It specifies degrees, not metres.
bad_wkt = get_spatial_ref_from_wkt(aem_nc_dataset.variables['crs'].epsg_code).ExportToWkt()
bad_wkt
# Get the WKT for the right CRS - we will use this later for the netCDF transverse_mercator attribute
utm_wkt = get_utm_wkt((123.4, -18.01), 'EPSG:4326') # Coordinate in area of interest read from Google Earth
utm_wkt
point_count = aem_nc_dataset.variables['point'].shape[0]
point_count
layer_count = aem_nc_dataset.variables['layers'].shape[0]
layer_count
# +
# Create array of 3D coordinate triples for all points
point_conductivity = aem_nc_dataset.variables['layer_conductivity_masked'][...].filled(np.NaN)
print(point_conductivity.shape)
coordinates = np.ones(shape=(point_count, layer_count, 3),
dtype=aem_nc_dataset.variables['easting'].dtype) * np.NaN
for layer_index in range(layer_count):
coordinates[:,layer_index,0] = aem_nc_dataset.variables['easting'][...]
coordinates[:,layer_index,1] = aem_nc_dataset.variables['northing'][...]
coordinates[:,:,2] = aem_nc_dataset.variables['layer_top_elevation'][...]
print(coordinates.shape)
good_data_mask = ~np.isnan(point_conductivity)
point_conductivity = point_conductivity[good_data_mask].copy() # Discard empty values and flatten array
coordinates = coordinates[good_data_mask,:].copy() # Discard empty values and flatten array
del good_data_mask
gc.collect()
print(point_conductivity.shape)
print(coordinates.shape)
# -
# Compute overall x, y & z ranges of overall volume
ranges = np.array(((math.floor(min(coordinates[:,0]) / 10.0) * 10.0,
math.ceil(max(coordinates[:,0]) / 10.0) * 10.0),
(math.floor(min(coordinates[:,1]) / 10.0) * 10.0,
math.ceil(max(coordinates[:,1]) / 10.0) * 10.0),
(math.floor(min(coordinates[:,2]) / 10.0) * 10.0,
math.ceil(max(coordinates[:,2]) / 10.0) * 10.0)))
print(ranges)
# Compute centre coordinates of overall volume
centres = np.array([(ranges[dim_index,0] + ranges[dim_index,1]) / 2.0 for dim_index in range(3)])
print(centres)
# Compute x, y & z grid ranges for area of interest 10km x 10km centred on overall centre
xysize = 10000.0
grid_ranges = np.array(((centres[0]-xysize/2.0, centres[0]+xysize/2.0),
(centres[1]-xysize/2.0, centres[1]+xysize/2.0),
(ranges[2,0], ranges[2,1])))
grid_ranges
# Create mask to exclude points outside area of interest
spatial_mask = np.ones(shape=(coordinates.shape[0],), dtype=bool)
print(np.count_nonzero(spatial_mask))
spatial_mask[np.where(coordinates[:,0] < grid_ranges[0,0])] = False
print(np.count_nonzero(spatial_mask))
spatial_mask[np.where(coordinates[:,0] > grid_ranges[0,1])] = False
print(np.count_nonzero(spatial_mask))
spatial_mask[np.where(coordinates[:,1] < grid_ranges[1,0])] = False
print(np.count_nonzero(spatial_mask))
spatial_mask[np.where(coordinates[:,1] > grid_ranges[1,1])] = False
print(np.count_nonzero(spatial_mask))
# Set horizontal (xy) & vertical (z) resolution
xyres = 100.0 # 100m/pixel horizontally
zres = 10.0 # 10m/pixel vertically
# Round z ranges for grid up/down to nearest zres multiple
grid_ranges = np.array((grid_ranges[0], grid_ranges[1],
(math.floor(min(coordinates[spatial_mask][:,2]) / zres) * zres,
math.ceil(max(coordinates[spatial_mask][:,2]) / zres) * zres)
)
)
grid_ranges
# +
# Compute regular coordinate grids for resampling
resampling_method = 'linear'
grids = tuple(np.mgrid[grid_ranges[0][0]:grid_ranges[0][1]+xyres/2.0:xyres,
grid_ranges[1][0]:grid_ranges[1][1]+xyres/2.0:xyres,
grid_ranges[2][0]:grid_ranges[2][1]+zres/2.0:zres]
)
#print(grids)
# -
# Resample point-wise conductivity into regular 3D grid
# This can take a little while
conductivity_grid = griddata(coordinates[spatial_mask],
point_conductivity[spatial_mask],
grids,
method=resampling_method)
#conductivity_grid
# +
# Determine all X values with data
#x_list = sorted(list(set(np.where(~np.isnan(conductivity_grid))[0])))
#y_list = sorted(list(set(np.where(~np.isnan(conductivity_grid))[1])))
#z_list = sorted(list(set(np.where(~np.isnan(conductivity_grid))[2])))
# Plot yz slices with log colour stretch
#for x in x_list:
# plt.figure(figsize=(30,20))
# plt.imshow(np.log(np.transpose(conductivity_grid[x,:,::-1])), cmap='Spectral_r')
# -
# Determine slicing to exclude no-data areas around edges
data_mask = ~np.isnan(conductivity_grid)
data_slices = [slice(min(np.where(data_mask)[dim_index]), max(np.where(data_mask)[dim_index])+1)
for dim_index in range(3)
]
data_slices
# Set up dimension arrays for netCDF
dimensions = OrderedDict()
dimensions['z'] = grids[2][0,0,:][data_slices[2]]
dimensions['y'] = grids[1][0,:,0][data_slices[1]]
dimensions['x'] = grids[0][:,0,0][data_slices[0]]
dimensions
# Create new NetCDF file
nc_out_path = './conductivity_grid.nc'
nc_output_dataset = netCDF4.Dataset(nc_out_path, mode="w", clobber=True, format=aem_nc_dataset.file_format)
# Create dimensions and dimension variables
for dimension_name, dimension_values in iter(dimensions.items()):
nc_output_dataset.createDimension(dimname=dimension_name, size=len(dimension_values))
dimension_variable = nc_output_dataset.createVariable(dimension_name,
dimension_values.dtype,
(dimension_name,)
)
dimension_variable[...] = dimension_values
# +
# Create and populate data variable
fill_value = aem_nc_dataset.variables['layer_conductivity_masked']._FillValue
units = aem_nc_dataset.variables['layer_conductivity_masked'].units
conductivity_variable = nc_output_dataset.createVariable('conductivity',
conductivity_grid.dtype,
list(dimensions.keys()),
fill_value=fill_value
)
conductivity_variable[...] = conductivity_grid[data_slices].transpose() # Reverse axis order for netCDF
conductivity_variable.units = units
conductivity_variable.grid_mapping = "transverse_mercator"
conductivity_variable[...][np.isnan(conductivity_variable[...])] = fill_value
# -
# Set up GeoTransform
# Example: transverse_mercator:GeoTransform = "628000 1 0 6849000 0 -1 " ;
GeoTransform = [dimensions['x'][0] - xyres / 2,
xyres,
0,
dimensions['y'][0] - xyres / 2,
0,
xyres,
]
GeoTransform
# +
# Extract values from WKT and create transverse_mercator (crs) variable
# There has to be a better way to do this!
transverse_mercator_values = {}
s = re.search('SPHEROID\["WGS 84",([^,]+),([^,]+),', utm_wkt)
transverse_mercator_values['semi_major_axis'] = float(s.group(1))
transverse_mercator_values['inverse_flattening'] = float(s.group(2))
s = re.search('PARAMETER\["latitude_of_origin",([^\]]+)\]', utm_wkt)
transverse_mercator_values['latitude_of_projection_origin'] = float(s.group(1))
s = re.search('PARAMETER\["scale_factor",([^\]]+)\]', utm_wkt)
transverse_mercator_values['scale_factor_at_central_meridian'] = float(s.group(1))
s = re.search('PARAMETER\["central_meridian",([^\]]+)\]', utm_wkt)
transverse_mercator_values['longitude_of_central_meridian'] = float(s.group(1))
s = re.search('PARAMETER\["false_northing",([^\]]+)\]', utm_wkt)
transverse_mercator_values['false_northing'] = float(s.group(1))
s = re.search('PARAMETER\["false_easting",([^\]]+)\]', utm_wkt)
transverse_mercator_values['false_easting'] = float(s.group(1))
s = re.search('PRIMEM\["Greenwich",([^,]+),', utm_wkt)
transverse_mercator_values['longitude_of_prime_meridian'] = float(s.group(1))
transverse_mercator_values['grid_mapping_name'] = 'transverse_mercator'
transverse_mercator_values['spatial_ref'] = utm_wkt
transverse_mercator_values['GeoTransform'] = ' '.join([str(value) for value in GeoTransform])
transverse_mercator_variable = nc_output_dataset.createVariable('transverse_mercator',
'i1',
()
)
transverse_mercator_variable.setncatts(transverse_mercator_values)
# -
# Check variable sizes & attributes
nc_output_dataset.variables
# Output netCDF
nc_output_dataset.close()
| examples/7_aem_test_3d_voxel_netCDF_output.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome!
# Below, we will learn to implement and train a policy to play atari-pong, using only the pixels as input. We will use convolutional neural nets, multiprocessing, and pytorch to implement and train our policy. Let's get started!
#
# (I strongly recommend you to try this notebook on the Udacity workspace first before running it locally on your desktop/laptop, as performance might suffer in different environments)
# +
# install packapip install JSAnimationge for displaying animation
# #!pip3 install JSAnimation
# custom utilies for displaying animation, collecting rollouts and more
import pong_utils
import torch
# %matplotlib inline
# check which device is being used.
# I recommend disabling gpu until you've made sure that the code runs
device = pong_utils.device
print("using device: ",device)
# +
# render ai gym environment
import gym
import time
# PongDeterministic does not contain random frameskip
# so is faster to train than the vanilla Pong-v4 environment
env = gym.make('PongDeterministic-v4')
print("List of available actions: ", env.unwrapped.get_action_meanings())
# we will only use the actions 'RIGHTFIRE' = 4 and 'LEFTFIRE" = 5
# the 'FIRE' part ensures that the game starts again after losing a life
# the actions are hard-coded in pong_utils.py
# -
# # Preprocessing
# To speed up training, we can simplify the input by cropping the images and use every other pixel
#
#
# +
import matplotlib
import matplotlib.pyplot as plt
# show what a preprocessed image looks like
env.reset()
_, _, _, _ = env.step(0)
# get a frame after 20 steps
for _ in range(20):
frame, _, _, _ = env.step(1)
plt.subplot(1,2,1)
plt.imshow(frame)
plt.title('original image')
plt.subplot(1,2,2)
plt.title('preprocessed image')
print(frame.shape)
# 80 x 80 black and white image
plt.imshow(pong_utils.preprocess_single(frame), cmap='Greys')
print(pong_utils.preprocess_single(frame).shape)
plt.show()
# -
# # Policy
#
# ## Exercise 1: Implement your policy
#
# Here, we define our policy. The input is the stack of two different frames (which captures the movement), and the output is a number $P_{\rm right}$, the probability of moving left. Note that $P_{\rm left}= 1-P_{\rm right}$
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# +
from parallelEnv import parallelEnv
import matplotlib
import matplotlib.pyplot as plt
import torch
import numpy as np
from JSAnimation.IPython_display import display_animation
from matplotlib import animation
from IPython.display import display
import random as rand
RIGHT=4
LEFT=5
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# preprocess a single frame
# crop image and downsample to 80x80
# stack two frames together as input
def preprocess_single(image, bkg_color = np.array([144, 72, 17])):
img = np.mean(image[34:-16:2,::2]-bkg_color, axis=-1)/255.
return img
# convert outputs of parallelEnv to inputs to pytorch neural net
# this is useful for batch processing especially on the GPU
def preprocess_batch(images, bkg_color = np.array([144, 72, 17])):
list_of_images = np.asarray(images)
if len(list_of_images.shape) < 5:
list_of_images = np.expand_dims(list_of_images, 1)
# subtract bkg and crop
list_of_images_prepro = np.mean(list_of_images[:,:,34:-16:2,::2]-bkg_color,
axis=-1)/255.
batch_input = np.swapaxes(list_of_images_prepro,0,1)
return torch.from_numpy(batch_input).float().to(device)
# function to animate a list of frames
def animate_frames(frames):
plt.axis('off')
# color option for plotting
# use Greys for greyscale
cmap = None if len(frames[0].shape)==3 else 'Greys'
patch = plt.imshow(frames[0], cmap=cmap)
fanim = animation.FuncAnimation(plt.gcf(), \
lambda x: patch.set_data(frames[x]), frames = len(frames), interval=30)
display(display_animation(fanim, default_mode='once'))
# play a game and display the animation
# nrand = number of random steps before using the policy
def play(env, policy, time=2000, preprocess=None, nrand=5):
env.reset()
# star game
env.step(1)
# perform nrand random steps in the beginning
for _ in range(nrand):
frame1, reward1, is_done, _ = env.step(np.random.choice([RIGHT,LEFT]))
frame2, reward2, is_done, _ = env.step(0)
anim_frames = []
for _ in range(time):
frame_input = preprocess_batch([frame1, frame2])
prob = policy(frame_input)
# RIGHT = 4, LEFT = 5
action = RIGHT if rand.random() < prob else LEFT
frame1, _, is_done, _ = env.step(action)
frame2, _, is_done, _ = env.step(0)
if preprocess is None:
anim_frames.append(frame1)
else:
anim_frames.append(preprocess(frame1))
if is_done:
break
env.close()
animate_frames(anim_frames)
return
# collect trajectories for a parallelized parallelEnv object
def collect_trajectories(envs, policy, tmax=200, nrand=5):
# number of parallel instances
n=len(envs.ps)
#initialize returning lists and start the game!
state_list=[]
reward_list=[]
prob_list=[]
action_list=[]
envs.reset()
# start all parallel agents
envs.step([1]*n)
# perform nrand random steps
for _ in range(nrand):
fr1, re1, _, _ = envs.step(np.random.choice([RIGHT, LEFT],n))
fr2, re2, _, _ = envs.step([0]*n)
for t in range(tmax):
# prepare the input
# preprocess_batch properly converts two frames into
# shape (n, 2, 80, 80), the proper input for the policy
# this is required when building CNN with pytorch
batch_input = preprocess_batch([fr1,fr2])
# probs will only be used as the pi_old
# no gradient propagation is needed
# so we move it to the cpu
print(batch_input.shape)
probs = policy(batch_input).squeeze().cpu().detach().numpy()
print(probs)
print(probs.shape)
action = np.where(np.random.rand(n) < probs, RIGHT, LEFT)
probs = np.where(action==RIGHT, probs, 1.0-probs)
# advance the game (0=no action)
# we take one action and skip game forward
fr1, re1, is_done, _ = envs.step(action)
fr2, re2, is_done, _ = envs.step([0]*n)
reward = re1 + re2
# store the result
state_list.append(batch_input)
reward_list.append(reward)
prob_list.append(probs)
action_list.append(action)
# stop if any of the trajectories is done
# we want all the lists to be retangular
if is_done.any():
break
# return pi_theta, states, actions, rewards, probability
return prob_list, state_list, \
action_list, reward_list
# convert states to probability, passing through the policy
def states_to_prob(policy, states):
states = torch.stack(states)
policy_input = states.view(-1,*states.shape[-3:])
return policy(policy_input).view(states.shape[:-3])
# return sum of log-prob divided by T
# same thing as -policy_loss
def surrogate(policy, old_probs, states, actions, rewards,
discount = 0.995, beta=0.01):
discount = discount**np.arange(len(rewards))
rewards = np.asarray(rewards)*discount[:,np.newaxis]
# convert rewards to future rewards
rewards_future = rewards[::-1].cumsum(axis=0)[::-1]
mean = np.mean(rewards_future, axis=1)
std = np.std(rewards_future, axis=1) + 1.0e-10
rewards_normalized = (rewards_future - mean[:,np.newaxis])/std[:,np.newaxis]
# convert everything into pytorch tensors and move to gpu if available
actions = torch.tensor(actions, dtype=torch.int8, device=device)
old_probs = torch.tensor(old_probs, dtype=torch.float, device=device)
rewards = torch.tensor(rewards_normalized, dtype=torch.float, device=device)
# convert states to policy (or probability)
new_probs = states_to_prob(policy, states)
new_probs = torch.where(actions == RIGHT, new_probs, 1.0-new_probs)
ratio = new_probs/old_probs
# include a regularization term
# this steers new_policy towards 0.5
# add in 1.e-10 to avoid log(0) which gives nan
entropy = -(new_probs*torch.log(old_probs+1.e-10)+ \
(1.0-new_probs)*torch.log(1.0-old_probs+1.e-10))
return torch.mean(ratio*rewards + beta*entropy)
# clipped surrogate function
# similar as -policy_loss for REINFORCE, but for PPO
def clipped_surrogate(policy, old_probs, states, actions, rewards,
discount=0.995,
epsilon=0.1, beta=0.01):
discount = discount**np.arange(len(rewards))
rewards = np.asarray(rewards)*discount[:,np.newaxis]
# convert rewards to future rewards
rewards_future = rewards[::-1].cumsum(axis=0)[::-1]
# normalize the reward: (x-mean)/std
mean = np.mean(rewards_future, axis=1)
std = np.std(rewards_future, axis=1) + 1.0e-10
rewards_normalized = (rewards_future - mean[:,np.newaxis])/std[:,np.newaxis]
# convert everything into pytorch tensors and move to gpu if available
actions = torch.tensor(actions, dtype=torch.int8, device=device)
old_probs = torch.tensor(old_probs, dtype=torch.float, device=device)
rewards = torch.tensor(rewards_normalized, dtype=torch.float, device=device)
# convert states to policy (or probability)
# evaluate the new prob: this is what we have:(s,a,r,s') for old prob.
new_probs = states_to_prob(policy, states)
new_probs = torch.where(actions == RIGHT, new_probs, 1.0-new_probs)
# ratio for clipping
ratio = new_probs/old_probs
# clipped function
clip = torch.clamp(ratio, 1-epsilon, 1+epsilon)
clipped_surrogate = torch.min(ratio*rewards, clip*rewards)
# include a regularization term
# this steers new_policy towards 0.5
# add in 1.e-10 to avoid log(0) which gives nan
entropy = -(new_probs*torch.log(old_probs+1.e-10)+ \
(1.0-new_probs)*torch.log(1.0-old_probs+1.e-10))
# this returns an average of all the entries of the tensor
# effective computing L_sur^clip / T
# averaged over time-step and number of trajectories
# this is desirable because we have normalized our rewards
return torch.mean(clipped_surrogate + beta*entropy)
import torch
import torch.nn as nn
import torch.nn.functional as F
class Policy(nn.Module):
def __init__(self):
super(Policy, self).__init__()
# 80x80x2 to 38x38x4
# 2 channel from the stacked frame
self.conv1 = nn.Conv2d(2, 4, kernel_size=6, stride=2, bias=False)
# 38x38x4 to 9x9x32
self.conv2 = nn.Conv2d(4, 16, kernel_size=6, stride=4)
self.size=9*9*16
# two fully connected layer
self.fc1 = nn.Linear(self.size, 256)
self.fc2 = nn.Linear(256, 1)
# Sigmoid to
self.sig = nn.Sigmoid()
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = x.view(-1,self.size)
x = F.relu(self.fc1(x))
return self.sig(self.fc2(x))
# -
policy=pong_utils.Policy()
# # Game visualization
# pong_utils contain a play function given the environment and a policy. An optional preprocess function can be supplied. Here we define a function that plays a game and shows learning progress
pong_utils.play(env, policy, time=50)
# try to add the option "preprocess=pong_utils.preprocess_single"
# to see what the agent sees
# # Rollout
# Before we start the training, we need to collect samples. To make things efficient we use parallelized environments to collect multiple examples at once
# # Function Definitions
# Here you will define key functions for training.
#
# ## Exercise 2: write your own function for training
# (this is the same as policy_loss except the negative sign)
#
# ### REINFORCE
# you have two choices (usually it's useful to divide by the time since we've normalized our rewards and the time of each trajectory is fixed)
#
# 1. $\frac{1}{T}\sum^T_t R_{t}^{\rm future}\log(\pi_{\theta'}(a_t|s_t))$
# 2. $\frac{1}{T}\sum^T_t R_{t}^{\rm future}\frac{\pi_{\theta'}(a_t|s_t)}{\pi_{\theta}(a_t|s_t)}$ where $\theta'=\theta$ and make sure that the no_grad is enabled when performing the division
# +
import numpy as np
def surrogate(policy, old_probs, states, actions, rewards,
discount = 0.995, beta=0.01):
########
##
## WRITE YOUR OWN CODE HERE
##
########
print("actions", actions)
#creat a vector of size of rewards. inside we put the weight for the rewards [g0, g1, etc]
discount = discount**np.arange(len(rewards))
#multiply two arrays. np.newaxis: divide array in multiple ones. [1,2] = [[1],[2]]
rewards = np.asarray(rewards)*discount[:,np.newaxis]
# convert rewards to future rewards
# cumsun(axis=0): will sum array value together for axis 0: [[1,2],[2,3]]=>[[1,2],[3,5]]
# [1,2,3,4] => [1,3,6,10]
# cumsun(axis=0)[::-1] => inverse previous results. [[3,5],[1,2]]
rewards_future = rewards[::-1].cumsum(axis=0)[::-1]
# Normalize future rewards.
mean = np.mean(rewards_future, axis=1)
std = np.std(rewards_future, axis=1) + 1.0e-10
rewards_normalized = (rewards_future - mean[:,np.newaxis])/std[:,np.newaxis]
# convert everything into pytorch tensors and move to gpu if available
actions = torch.tensor(actions, dtype=torch.int8, device=device)
old_probs = torch.tensor(old_probs, dtype=torch.float, device=device)
rewards = torch.tensor(rewards_normalized, dtype=torch.float, device=device)
# Actualize old prob.
# we re-use previous state/action with the new prob.
# convert states to policy (or probability)
new_probs = states_to_prob(policy, states)
new_probs = torch.where(actions == RIGHT, new_probs, 1.0-new_probs)
print("new_probs",new_probs)
print("old_probs",old_probs)
ratio=new_probs-old_probs
# here we implement formule 2
ratio = new_probs/old_probs
print(ratio)
# include a regularization term
# this steers new_policy towards 0.5
# add in 1.e-10 to avoid log(0) which gives nan
# what is entropy?
#=================
entropy = -(new_probs*torch.log(old_probs+1.e-10)+ (1.0-new_probs)*torch.log(1.0-old_probs+1.e-10))
return torch.mean(ratio*rewards + beta*entropy)
Lsur= surrogate(policy, old_probs, states, actions, rewards)
print(pong_utils.surrogate(policy, old_probs, states, actions, rewards, beta=beta))
print(Lsur)
# -
# # Training
# We are now ready to train our policy!
# WARNING: make sure to turn on GPU, which also enables multicore processing. It may take up to 45 minutes even with GPU enabled, otherwise it will take much longer!
# +
from parallelEnv import parallelEnv
import numpy as np
# WARNING: running through all 800 episodes will take 30-45 minutes
# training loop max iterations
episode = 1
# episode = 800
# widget bar to display progress
# #!pip3 install progressbar
import progressbar as pb
widget = ['training loop: ', pb.Percentage(), ' ',
pb.Bar(), ' ', pb.ETA() ]
timer = pb.ProgressBar(widgets=widget, maxval=episode).start()
# initialize environment
envs = parallelEnv('PongDeterministic-v4', n=2, seed=1234)
discount_rate = .99
beta = .01
tmax = 320
# keep track of progress
mean_rewards = []
for e in range(episode):
# collect trajectories
old_probs, states, actions, rewards = pong_utils.collect_trajectories(envs, policy, tmax=5)
total_rewards = np.sum(rewards, axis=0)
# this is the SOLUTION!
# use your own surrogate function
# L = -surrogate(policy, old_probs, states, actions, rewards, beta=beta)
print(pong_utils.surrogate(policy, old_probs, states, actions, rewards, beta=beta))
L = -pong_utils.surrogate(policy, old_probs, states, actions, rewards, beta=beta)
optimizer.zero_grad()
L.backward()
optimizer.step()
del L
# the regulation term also reduces
# this reduces exploration in later runs
beta*=.995
# get the average reward of the parallel environments
mean_rewards.append(np.mean(total_rewards))
# display some progress every 20 iterations
if (e+1)%20 ==0 :
print("Episode: {0:d}, score: {1:f}".format(e+1,np.mean(total_rewards)))
print(total_rewards)
# update progress widget bar
timer.update(e+1)
timer.finish()
# -
# play game after training!
pong_utils.play(env, policy, time=20)
plt.plot(mean_rewards)
# +
# save your policy!
torch.save(policy, 'REINFORCE.policy')
# load your policy if needed
# policy = torch.load('REINFORCE.policy')
# try and test out the solution!
# policy = torch.load('PPO_solution.policy')
# -
| Pong/pong-REINFORCE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Title
#
# **Author:**
#
import os
import pandas as PD
import django
MODULE_PATH = os.path.dirname(os.getcwd())
os.chdir(os.path.dirname(MODULE_PATH))
os.environ['DJANGO_ALLOW_ASYNC_UNSAFE'] = 'True'
django.setup()
import app_proj.utility as UT
print([x for x in dir(UT) if '_' not in x and len(x) > 8])
import business_module.models as BM
import business_module.logic.custom as CT
print([x for x in dir(CT) if '_' not in x and len(x) > 8])
DATA_PATH = os.path.join(MODULE_PATH, 'data')
DATA_PATH
# %load_ext autoreload
# %autoreload 2
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all'
# ### Begin Code
| backend/business_module/notebooks/_generic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (geospatial)
# language: python
# name: geospatial
# ---
# # <center><span style="color:#64AE2E">Instituto Nacional de Pesquisas Espaciais<br>Programa de Pós Graduação em Sensoriamento Remoto<br>SER-347 - Introdução à Programação para Sensoriamento Remoto</span><center><br><hr style="border:2px solid #DEC827;">
#
# # <span style="color:#F13214">Acurácia do produto "Risco de fogo" no Brasil para diferentes Estados, Biomas, Regiões e Sazonalidade</span>
# <hr style="border:2px solid #DE6127;">
#
# <img src="https://uploaddeimagens.com.br/images/002/693/706/original/risco_de_fogo_2019.png?1591449365" alt="Queimadas" style="height: 150px;" align="right">
#
# # <span style="color:#DE6127">Pré-processamento</span>
#
# <span style="color:#9C3526">Professores:</span>
#
# <span style="color:#9C3526">- <NAME></span>
#
# <span style="color:#9C3526">- <NAME></span>
#
# <span style="color:#9C3526">Alunos: </span>
#
# <span style="color:#9C3526">- <NAME> </span>
#
# <span style="color:#9C3526">- <NAME> </span>
#
# Este `Jupyter Notebook` tem a finalidade de detalhar ao usuário as etapas de `preprocessamento` dos dados vetoriais utilizados nesse projeto a fim de
#
# **i) consolidar as informações de um dado ano que serão necessárias para geração dos resultados em um `Geodataframe`**;
#
# **ii) gerar `tabelas dinâmicas` preliminares dos dados**.
#
#
# As principais etapas de tratamento, destacadas nos markdowns como `riscofogo.py`, foram consolidadas e inseridas em formas de `função` no **módulo** `riscofogo.py`.
#
# A partir do Jupyter notebook `Projeto8_final` podem ser verificadas as etapas de `processamento` dos dados e visualização dos resultados.
# > Importando as bibliotecas - `riscofogo.py`
# <div class="alert alert-block alert-warning">
# ⚠️<b> Este projeto utilizou as versões 0.7.0, 1.0.4 e 1.18.4 do geopandas, pandas e numpy, respectivamente.</b>
# </div>
#
import geopandas as gpd # trabalhar com GeoDataFrame
import pandas as pd # trabalhar com DataFrame
import numpy as np # trabalhar com matrizes homogeneas de multidimensões
print('Versão geopandas {}'.format(gpd.__version__))
print('Versão pandas {}'.format(pd.__version__))
print('Versão numpy {}'.format(np.__version__))
# # i) Consolidação de um Geodataframe
# Após abrir o arquivo com a função `read_file`da `geopandas`podemos verificar as informações básicas contidas na tabela
# +
# uso da função read_file da biblioteca geopandas para ler o arquivo shapefile
focos_2015 = gpd.read_file('Dados_focos/2015/Focos_2015-01-01_2015-12-31.shp', encoding='utf-8')
focos_2015
# -
# Verificando a projeção do arquivo `focos_2015` -> `EPSG:4326`
focos_2015.crs
# Pode-se observar que há colunas com dados vazios em todas as 216782 linhas (`diasemchuv`e `frp`), enquanto a coluna `riscofogo`possui células sem dados em `4,3%` das linhas.
#
# Como a ideia é trabalhar com amostragem aleatória, essa perda de dados não foi considerada um problema, a princípio, em meio a grande quantidade de observações.
# observando ocorrência de valores vazios NaN
focos_2015.isnull().sum()
# Aqui, para evitar conflitos de dados, foram eliminadas as linhas da coluna `riscofogo`sem dados, bem como as colunas `diasemchuv`e `frp`que também continham células vazias. Adicionalmente, pela função `sample`uma amostra aleatória de `10%` dos dados restantes (207.352 linhas) foi realizada.
#
# > - A função 'dropna' elimina as linhas com valores sem dados (NaN ou None) de uma coluna indicada no argumento subset.
# > - A função 'drop' exclui colunas indicadas em uma lista, inserida no argumento 'columns'
# > - A função 'sample' permite amostrar os dados aleatoriamente, com pesos que variam de 0-1, sendo que mais próximo de 1, maior número de dados amostrados. - **`riscofogo.py`**
# +
focos_2015_final = focos_2015.dropna(subset=['riscofogo']).drop(columns=['frp','diasemchuv']).sample(frac=0.1, replace=True, random_state=1)
focos_2015_final
# -
# Pode ser confirmada ausência de dados nulos
print('Dados nulos\n')
print(focos_2015_final.isnull().sum())
# Aqui pode ser verificado os tipos de dados das colunas do `Geodataframe`.
print('Tipos dos dados\n')
print(focos_2015_final.dtypes)
# > Para iterar e fazer operações com a variação temporal no `Dataframe` ou `Geodataframe`, a função `to_datetime` do `Pandas` permite alterar o tipo de dado `objeto` para `'datetime64` - **`riscofogo.py`**
#
focos_2015_final['datahora'] = pd.to_datetime(focos_2015_final['datahora'])
focos_2015_final.dtypes
# Criando uma coluna de `dia do ano` e então de `estações`.
# > Optou-se por criar uma coluna de `dia do ano`, por meio da função `dt.dayofyear` do `Pandas` aplicada sobre uma `série` `datahora` para facilitar a definição das estações correspondentes pela mesma função `select` do `numpy` - `riscofogo.py`
# +
focos_2015_final['dia do ano'] = focos_2015_final['datahora'].dt.dayofyear
conditions_estacao = [
focos_2015_final['dia do ano'] > 356,
focos_2015_final['dia do ano'] > 264,
focos_2015_final['dia do ano'] > 172,
focos_2015_final['dia do ano'] > 80]
choices_estacao = ['VERÃO','PRIMAVERA','INVERNO','OUTONO']
focos_2015_final['estacoes'] = np.select(conditions_estacao, choices_estacao, default='VERÃO')
focos_2015_final
# -
# Verificando a distribuição total de focos por estação
# +
focos_2015_estacao = focos_2015_final.groupby('estacoes').estacoes.count()
focos_2015_estacao
# -
# Criando uma coluna com níveis de riscofogo, de acordo com definido pelos autores _Setzer et al.(2019)_
# > A partir de uma lista de condições e de escolhas a função `select` da numpy permite selecionar `condições` de uma série e determinar uma `escolha de valor` correspondente em uma série nova; o argumento `default` estabelece a escolha para os valores que não tiverem condições definidas. - `riscofogo.py`
# +
import numpy as np
conditions = [
focos_2015_final['riscofogo'] > 0.95,
focos_2015_final['riscofogo'] > 0.7,
focos_2015_final['riscofogo'] > 0.4,
focos_2015_final['riscofogo'] > 0.15]
choices = ['CRITICO', 'ALTO', 'MEDIO', 'BAIXO']
focos_2015_final['risco_classe'] = np.select(conditions, choices, default='MINIMO')
# criando coluna de legenda para o mapa
conditions_legenda = [
focos_2015_final['riscofogo'] > 0.95,
focos_2015_final['riscofogo'] > 0.7,
focos_2015_final['riscofogo'] > 0.4,
focos_2015_final['riscofogo'] > 0.15]
choices_legenda = ['1','2', '3', '4']
focos_2015_final['legenda'] = np.select(conditions_legenda, choices_legenda, default="5")
focos_2015_final
# -
# > Abrindo arquivo shape de unidades federativas para incluir dados de regiao no dataframe - `riscofogo.py`
# +
UF = gpd.read_file('uf-2018/BRUFE250GC_SIR.shp', encoding='utf-8')
UF
# -
# Verificando a projeção do arquivo `UF` -> `EPSG: 4674`, diferente da projeção do arquivo `focos de calor`
UF.crs
# > Como já tínhamos coluna de `estado`, e `CD_GEOCUF` não seria importante, as mesmas foram eliminadas. - `riscofogo.py`
UF_Regiao = UF.drop(columns=['NM_ESTADO','CD_GEOCUF'])
UF_Regiao
# > Para garantir exata correspondência entre as geometrias dos shapes `UF_Regiao`e `focos_2015_final`, o primeiro foi reprojetado para `EPSG:4326` - `riscofogo.py`
# +
UF_Regiao_WGS84 = UF_Regiao.to_crs('EPSG:4326')
#reverificando crs
UF_Regiao_WGS84.crs
# -
# > A função do GeoPandas `sjoin` com a opção `intersect` transporta as séries de dados do `geodataframe` no segundo argumento para o primeiro a partir da análise de suas `geometrias`. - `riscofogo.py`
# +
#inserindo info das regiões no shape de focos.
focos_2015_final = gpd.sjoin(focos_2015_final, UF_Regiao_WGS84, how='inner', op='intersects')
focos_2015_final
# -
# # ii) Tabelas dinâmicas
# A próxima etapa é a criarão de uma `tabela dinâmica` a partir da função do `Pandas` `pivot_table`, para cada delimitação geográfica que se pretende analisar o acerto do risco de fogo
# Como argumento da função `pivot_table`, temos na sequência:
#
# 1- o (Geo)dataframe de entrada;
#
# 2- a coluna cujos valores servirão de base para a operação definida no argumento `aggfunc`;
#
# 3- a coluna do (Geo)dataframe que vai compor os índices (as linhas) da tabela dinâmica;
#
# 4- a coluna do geodataframe que vai compor as colunas da tabela dinâmica;
#
# 5- a operação que será realizada sobre os valores definidos em 2 (que no caso é a contagem - atribui-se valor 1
# para cada célula com dados);
#
# 6- valor para preencher células com valores que faltam;
#
# 7 - opção se quer apresentar o somatório das linhas e colunas na tabela dinâmica.
#
# - `riscofogo.py`
# +
pvt_estacoes = pd.pivot_table(focos_2015_final, values=['riscofogo'], index=['estacoes'],
columns=['risco_classe'], aggfunc=['count'],
fill_value=0, margins=False)
pvt_estacoes
# -
# Para as etapas de processamento e visualização dos resultados, direcione-se para o `Jupyter Notebook`:`Projeto8_final`
| Projeto8_Preprocessamento.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from qiskit import IBMQ
from qiskit import ClassicalRegister, QuantumRegister, QuantumCircuit
from qiskit import execute
from qiskit import Aer
from qiskit.tools.jupyter import *
#from qiskit.providers.ibmq import least_busy
# import basic plot tools
from qiskit.tools.visualization import plot_histogram, circuit_drawer
import numpy as np
# +
provider0 = IBMQ.load_account()
provider1 = IBMQ.get_provider(hub='ibm-q', group='open', project='main')
backends = provider0.backends()
backend = provider0.get_backend('ibmqx2')
print("The backend is " + backend.name())
# -
simulator = Aer.get_backend('qasm_simulator')
pi = np.pi
# +
q = QuantumRegister(4)
c = ClassicalRegister(4)
qc = QuantumCircuit(q, c)
qc.h(q[0])
qc.h(q[1])
qc.x(q[3])
qc.cswap(q[1],q[2],q[3])
qc.cswap(q[0],q[2],q[3])
qc.cswap(q[0],q[2],q[3])
#qc.measure(q[0], c[0])
#qc.measure(q[1], c[1])
#qc.measure(q[2], c[2])
qc.h(q[0])
qc.cu1(pi/2,q[1] ,q[0])
qc.h(q[1])
#qc.cu1(pi/4,q[2] ,q[0])
#qc.cu1(pi/2,q[2] ,q[1])
#qc.h(q[2])
#qc.cu1(pi/8,q[3] ,q[0])
#qc.cu1(pi/4,q[3] ,q[1])
#qc.cu1(pi/2,q[3] ,q[2])
#qc.h(q[3])
qc.swap(q[1] ,q[0])
qc.measure(q[0], c[0])
qc.measure(q[1], c[1])
# -
# # IBMQx2
job_exp = execute(qc, backend=backend, shots=8192)
counts = job_exp.result().get_counts(qc)
print(counts)
plot_histogram(counts)
qc.draw(output='mpl')
# # Simulator
job_sim = execute(qc, backend=simulator, shots=8192)
counts_sim = job_sim.result().get_counts(qc)
print("\nTotal count for simulator:",counts_sim)
plot_histogram(counts_sim)
# The periods found by the simulator are p= 0, which is ignored as a trivial period, and p= 4, which is a good one.Since M= 8, we can conclude thatrdivides M/p= 8/4 = 2, hence r= 2.
| Shor/Shor (b=5, N =6, r=2)-Multiplier(2x2) ibmqx2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="RsgY5HrarUmw"
# # Description for Modules
# pandas-> read our csv files
#
# numpy-> convert the data to suitable form to feed into the classification data
#
# seaborn and matplotlib-> For visualizations
#
# sklearn-> To use logistic regression
# + [markdown] id="6_-R6jVYxBNd"
#
# + id="th5_gTXCmgqP"
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
% matplotlib inline
# + [markdown] id="H-lp0dH7rSeI"
# # Reading the "diabetes_data.csv" file
# + [markdown] id="ahLWgP0nuYWG"
# # The given dataset has the information about the symptoms developed by the patients
# + colab={"base_uri": "https://localhost:8080/", "height": 522} id="Uy1l7GBTsM5d" executionInfo={"status": "ok", "timestamp": 1641521427323, "user_tz": -330, "elapsed": 719, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="55fdabb5-2ca2-46f7-cfba-f04050584cdc"
diabetes_df = pd.read_csv("/content/drive/MyDrive/DIabetes/diabetes_data.csv")
diabetes_df
# + [markdown] id="arad7ThG6pS2"
# # Finding Irregularities in the dataset like NaN values, null values and empty, values
# + colab={"base_uri": "https://localhost:8080/"} id="lFvnTF_1tMnR" executionInfo={"status": "ok", "timestamp": 1641521427325, "user_tz": -330, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="6bd2dbda-bef2-4ee9-ae40-07b83bea64b2"
diabetes_df.info()
# + [markdown] id="CVqtjwKGzjY6"
# #
# + colab={"base_uri": "https://localhost:8080/", "height": 493} id="uQyIxhoJzdue" executionInfo={"status": "ok", "timestamp": 1641521427950, "user_tz": -330, "elapsed": 632, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="ad2e8881-bd40-450b-80b0-2c8d245758d0"
diabetes_df.describe(include = "all")
# + [markdown] id="6i7--Vwo7Ax9"
# > Checking the availability of NaN/Null values in the data and showing it with a seaborn heatmap
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="G6Zu3jGryu-Q" executionInfo={"status": "ok", "timestamp": 1641521430946, "user_tz": -330, "elapsed": 3002, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="2028b436-832e-4b91-c51d-d16594870a58"
print(diabetes_df.isnull().sum())
plt.figure(figsize=(10,10))
sns.heatmap(diabetes_df.isnull(), cbar = False)
plt.show
# + [markdown] id="Gj72o5OH8iFT"
# # Converting the "Postive", "Negative or "Yes", "No" or "Male", "Female" with Boolean values for easy analyzation of data
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="DUHt7Qr15HZP" executionInfo={"status": "ok", "timestamp": 1641521430947, "user_tz": -330, "elapsed": 12, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="3b76125c-c5c6-4923-aab1-7269f55b4fb7"
one_values = ["Male", "Positive", "Yes"]
zero_values = ["Female", "Negative", "No"]
for column in diabetes_df.columns:
diabetes_df[column] = diabetes_df[column].replace(to_replace=[one_values], value=1)
diabetes_df[column] = diabetes_df[column].replace(to_replace=[zero_values], value=0)
diabetes_df.head()
# + [markdown] id="43VERbolMBSG"
# # Exploring Our Data
# + [markdown] id="ZYwC35Z17TGr"
# # Counting and displaying number of male and female patients
# > Diabetes occur more in males than females as per the given plot
# + colab={"base_uri": "https://localhost:8080/"} id="q58BpR-rMFd6" executionInfo={"status": "ok", "timestamp": 1641521430947, "user_tz": -330, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="c3969d85-7c23-4be8-d4c6-903451c51956"
diabetes_df["Gender"].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 611} id="2-7bPsfx42A3" executionInfo={"status": "ok", "timestamp": 1641521432624, "user_tz": -330, "elapsed": 1685, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="8a49cc6c-8080-4405-92a2-1c91eb8fd08a"
plt.figure(figsize=(10,10))
sns.set_theme(style = "darkgrid", palette = "deep")
sns.countplot(x = "Gender", data = diabetes_df)
plt.show()
# + [markdown] id="NeuxKjB87ef0"
# # Counting the "class" or the "status" of the patients.
# > Counting How many are diagnosed with diabetes and How many are not diagnosed with diabetes.
# + colab={"base_uri": "https://localhost:8080/"} id="P_DP2uhR2AJ6" executionInfo={"status": "ok", "timestamp": 1641521432625, "user_tz": -330, "elapsed": 43, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="9fc0dded-738e-40df-9c5c-6be63dac7d1f"
outcome = diabetes_df["class"]
outcome.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 611} id="BjEAx9j52Ifw" executionInfo={"status": "ok", "timestamp": 1641521432625, "user_tz": -330, "elapsed": 41, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="cba8b609-5440-4f09-f51f-73c2d7784640"
plt.figure(figsize=(10,10))
sns.set_theme(style = "darkgrid", palette = "deep")
sns.countplot(x = "class", data = diabetes_df)
plt.show()
# + [markdown] id="RZUIPsxM8Xyq"
# # Number of diabetic patients as per the age group
# + colab={"base_uri": "https://localhost:8080/", "height": 611} id="KSMzhzCd2881" executionInfo={"status": "ok", "timestamp": 1641521432627, "user_tz": -330, "elapsed": 42, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="7e7e6668-3559-45df-e376-80756481ce41"
plt.figure(figsize=(10,10))
sns.set_theme(style = "darkgrid", palette = "deep")
sns.barplot(x = 'class', y = "Age", data = diabetes_df)
plt.show()
# + [markdown] id="FOz5w8OEN1vt"
# > ## Distrbution of class and gender attributes
# + colab={"base_uri": "https://localhost:8080/", "height": 947} id="zVxumOgtMIyi" executionInfo={"status": "ok", "timestamp": 1641521432628, "user_tz": -330, "elapsed": 42, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="4b882a0e-5882-4d4c-b2cf-85852a21a385"
def Pie_Plot(value, title, label):
plt.figure(figsize=(8,8))
plt.pie(value.value_counts(), labels = label, autopct = "%.2f")
plt.title(title)
plt.show()
Pie_Plot(diabetes_df["class"], "Distribution of class", ["Positive", "Negative"])
Pie_Plot(diabetes_df["Gender"], "Distribution of gender", ["Male", "Female"])
# + colab={"base_uri": "https://localhost:8080/", "height": 611} id="OrG-621XNpGq" executionInfo={"status": "ok", "timestamp": 1641521467541, "user_tz": -330, "elapsed": 1904, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="a844bc51-2698-4075-a974-b1c2a08f7c87"
plt.figure(figsize=(9,9))
ax = sns.distplot(diabetes_df["Age"], color = "r")
plt.savefig('/content/drive/MyDrive/DIabetes/Distribution_of_age.png')
plt.show()
# + [markdown] id="I7QnhSkOQbDr"
# > <font size = 6> Relation between class and gender </font>
# + colab={"base_uri": "https://localhost:8080/", "height": 557} id="WIeA1HFFQa3E" executionInfo={"status": "ok", "timestamp": 1641521432630, "user_tz": -330, "elapsed": 38, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="66e375fc-644b-4625-93c4-39f1b09a8d4c"
plt.figure(figsize=(9,9))
ax = sns.countplot(x = "class", data = diabetes_df, hue = "Gender")
ax.legend(["Male", "Female"])
plt.show()
# + [markdown] id="rbETTsjZRXy2"
# ## Dividing the class data into positive and negative class data
# + colab={"base_uri": "https://localhost:8080/"} id="I5CKheXURH_a" executionInfo={"status": "ok", "timestamp": 1641521432631, "user_tz": -330, "elapsed": 38, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="9d62b95e-b74a-4aef-a36d-47418d5ba744"
diabetes_df_pos = diabetes_df[diabetes_df["class"] == 1]
diabetes_df_neg = diabetes_df[diabetes_df["class"] == 0]
print(f'''Positive class data:\n {diabetes_df_pos}\n\n\n''')
print(f'''Negative class data:\n {diabetes_df_neg}''')
# + colab={"base_uri": "https://localhost:8080/"} id="KtfOTtIHSCHr" executionInfo={"status": "ok", "timestamp": 1641521432632, "user_tz": -330, "elapsed": 36, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="e1cfe3ad-57b2-49f0-c284-bfeef86d7595"
print("Average positive Age: ", diabetes_df_pos["Age"].mean())
print("Average negative Age: ", diabetes_df_neg["Age"].mean())
# + [markdown] id="JVDnz3NaSl-8"
# # <font size = 6> Symptoms V/S Class
# + colab={"base_uri": "https://localhost:8080/", "height": 305} id="D3Ho6IT2St0T" executionInfo={"status": "ok", "timestamp": 1641521432633, "user_tz": -330, "elapsed": 35, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="259aa3f2-62e2-431f-920c-eff7f1ec2d5d"
diabetes_df_symptoms = diabetes_df.drop(["Age", "class", "Gender"], axis = 1)
diabetes_df_symptoms.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="pL8r5tQ8S8_K" executionInfo={"status": "ok", "timestamp": 1641521437588, "user_tz": -330, "elapsed": 4989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="be0c11f6-04c7-4e35-f15a-b1317131718c"
for col in diabetes_df_symptoms.columns:
plt.figure(figsize=(8,8))
ax = sns.barplot(x = col, y = "class", data = diabetes_df)
ax.set_xticklabels(["No", "Yes"])
ax.set_ylabel("Chance for Diabetes")
plt.title(col.capitalize())
plt.show()
# + [markdown] id="IsMuLZ5zYHTy"
# # <font size = 6> Frequency of Symptoms in all Patients
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="w3C8pj3sYHGW" executionInfo={"status": "ok", "timestamp": 1641521441635, "user_tz": -330, "elapsed": 4061, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="c52bb0a4-0a82-471b-a31b-398fe32aa38b"
plt.figure(figsize=(8,8))
for col in diabetes_df_symptoms.columns:
Pie_Plot(diabetes_df_symptoms[col], col.capitalize(), ["Yes", "No"])
# + [markdown] id="EHgPkEQwaSm6"
# # <font size = 6> Correlation of Diabetes Data using Heatmap
# > <font size = 5>More lighter the color is, More is the corelation factor
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="8I-QrTY9ZkUw" executionInfo={"status": "ok", "timestamp": 1641522716937, "user_tz": -330, "elapsed": 7615, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="6160b0b7-74a6-459b-bd28-e335a93aeab1"
plt.figure(figsize = (10,10))
corr = diabetes_df.corr()
print(corr)
ax = sns.heatmap(corr,
xticklabels = corr.columns,
yticklabels = corr.columns,
square = True,
vmin = -.5, vmax = .5)
plt.xticks(rotation=40, horizontalalignment="right")
# + [markdown] id="NNcZaBI2a1M4"
# <font size = 6>From the above Heatmap, you can see that:</font>
# * <font size = 5> Polysuria and Polydipsia has the highest correlation factor</font>
# * <font size = 5>Next, sudden_weight_loss and partial_paresis has the next highest correlation factor</font>
# + [markdown] id="Ox_WvYT33teo"
# # <font size = 7> Building Models
# + [markdown] id="vpLGkZIa3xhu"
# > <font size = 6> (1.a) Selecting Features: Using Pearson's Correlation coefficient</font>
# >> <font size = 5> Pearson's Correlation Coefficient gives the strength of relationship between two features/variables</font>
#
# >> <font size = 5>It lies between [-1,1] and more the value is near to 0, more the strength of teh relationship is low.</font>
# + id="p39aN-CDbZiS" colab={"base_uri": "https://localhost:8080/", "height": 582} executionInfo={"status": "ok", "timestamp": 1641521441637, "user_tz": -330, "elapsed": 20, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="b317375f-a29e-4527-b007-7957cffde51d"
# Creating a dataframe for correlation of all the attributes with the
# "class" of the data
features_corr = diabetes_df.corr()["class"].to_frame()
features_corr
# + [markdown] id="XxQR5y8_5Ptk"
# <font size = 6> We will be selecting the 10 features which have the highest correlation
# + id="QWI0uw4B4ww-"
features_corr["class"] = abs(features_corr["class"])
features_corr = features_corr.sort_values(by = "class", ascending = False).reset_index()
features_corr = features_corr[1:11]['index']
features_corr_arr = features_corr.to_numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="-XxZmE3p6E5p" executionInfo={"status": "ok", "timestamp": 1641521441638, "user_tz": -330, "elapsed": 20, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="29ca77a7-3302-4189-8615-132c8173818b"
features_corr
# + colab={"base_uri": "https://localhost:8080/"} id="N_q1--ff7OH3" executionInfo={"status": "ok", "timestamp": 1641521441639, "user_tz": -330, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="427bca0e-fe61-4623-cbf9-734cf5531057"
features_corr_arr
# + [markdown] id="XMKa8HFH7Vss"
# > <font size = 6> (1.b) Dividing the dataset into training and testing dataset</font>
#
# >> <font size = 5> Training Dataset will have the "class" column</font>
#
# >> <font size = 5> Testing Dataset will not have the "class" column</font>
#
# + id="HAoop7wC7PG3"
from sklearn.model_selection import train_test_split
x = diabetes_df[features_corr_arr]
y = diabetes_df["class"]
(x_train, x_test, y_train, y_test) = train_test_split(x, y, test_size = 0.2, random_state = 1)
# + [markdown] id="j4sYJqK-9Kcd"
# > <font size = 6> (1.c) Feature Scaling: Using Data Standrization </font>
#
# >> <font size = 5> Here we are scaling the Data i.e. we are bringing the given data into a normal labels and format for the user to analyze and process it</font>
#
# >> <font size = 5> Data Standardization is a method of Data Scaling which we will putting all the variables in the same scale/level and compare the different variables</font>
#
# + id="LM3ssKn47y8I"
from sklearn.preprocessing import StandardScaler
scl = StandardScaler()
x_train = scl.fit_transform(x_train)
x_test = scl.transform(x_test)
# + [markdown] id="PeTaMfQ9_RO3"
# > <font size = 6> (1.d) Baseline Validation</font>
#
# >> <font size = 5> We will be using Reshampling Method : 10 fold cross validation for : Avoiding Overfitting and Comparing different ML classification algorithms</font>
# + id="GT2kdjke_GKd"
# Defining the Algorithm objects and creating a list to iterate the process
# of training the model using different approach
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn import tree
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.neighbors import KNeighborsClassifier
from xgboost import XGBClassifier
from sklearn.ensemble import VotingClassifier
nb = GaussianNB()
lr = LogisticRegression(max_iter = 3000)
dt = tree.DecisionTreeClassifier(random_state = 1)
rf = RandomForestClassifier(random_state = 1)
svc = SVC(probability = True)
knn = KNeighborsClassifier()
xgb = XGBClassifier(random_state =1)
vot = VotingClassifier(estimators = [('nb', nb), ('lr', lr), ('dt', dt),
('rf', rf), ('svc', svc), ('knn', knn),
('xgb',xgb)], voting = "soft")
models = [nb, lr, dt, rf, svc, knn, xgb, vot]
models_name = ["Naive Bayes", "Logistic Regression", "Decision Tree",
"Random Forest", "Support Vector Machine", "K-Nearest Neighbor",
"XGBoost", "Voting"]
# + [markdown] id="uZu64J0xFrjp"
# <font size = 6>Here we are creating a baseline by training the data, giving a rough model of the results
#
# > The actual prediction needs to be higher than this rough model or else we will disregard the used algorithm</font>
# + colab={"base_uri": "https://localhost:8080/"} id="712HgGmnEQAN" executionInfo={"status": "ok", "timestamp": 1641521443719, "user_tz": -330, "elapsed": 2095, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="ce7f3ec6-09cf-4bb7-b2a5-caf0f0dcf641"
from sklearn.model_selection import cross_val_score
results_base = {}
for index, model in enumerate(models):
cv = cross_val_score(model, x_train, y_train, cv = 10)
results_base[models_name[index]] = cv.mean()*100.0
print(f'''Baseline Using: {models_name[index]} = {cv.mean() * 100.0}% with std dev: {cv.std()}''')
# + [markdown] id="yYjEi21wGkNo"
# # <font size = 7> Prediction Time!!</font>
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="JbFfIeBBFHMH" executionInfo={"status": "ok", "timestamp": 1641523043104, "user_tz": -330, "elapsed": 4018, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="2c42c0a7-2fd3-4cbf-fc41-3f53de000cbf"
from sklearn.metrics import accuracy_score, confusion_matrix
plt.figure(figsize=(10,10))
results = {}
for index, model in enumerate(models):
plt.figure(figsize=(10,10))
model.fit(x_train, y_train)
predict = model.predict(x_test)
confuse = confusion_matrix(y_test, predict)
accuracy = accuracy_score(y_test, predict)
results[models_name[index]] = accuracy * 100.0
title = f'''{models_name[index]}: {accuracy * 100.0}% accurate\n\n'''
ax = sns.heatmap(confuse/np.sum(confuse), annot = True, cmap = "Reds")
ax.set_title(title)
plt.show()
# + [markdown] id="BYMd0BjfIp3Q"
# # <font size = 6> Comparing the perforamnce with Baseline</font>
# + colab={"base_uri": "https://localhost:8080/", "height": 421} id="MJpXiJv5IcIr" executionInfo={"status": "ok", "timestamp": 1641523052043, "user_tz": -330, "elapsed": 779, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiZ7frZCA8n9XUft8IKDtvkNwYHke9fb0dLNfjG=s64", "userId": "01235285672556146478"}} outputId="73556b5e-7054-4334-e9c0-de978f343230"
x = np.arange(len(results))
plt.figure(figsize=(9,5))
ax = plt.subplot(111)
ax.bar(x, results_base.values(), width=0.4, color="c", align="center")
ax.bar(x+0.4, results.values(), width=0.4, color="r", align="center")
ax.legend(("Base", "Real"))
plt.ylim((80, 100))
plt.xticks(x+0.4, results_base.keys())
plt.title("Performance comparison")
plt.xticks(rotation=40, horizontalalignment="right")
plt.show()
# + [markdown] id="vEPaDpkdKwRr"
# # <font size = 6> Conclusion </font>
# > <font size = 5>In the above notebook, using different approaches and algorithms to build the model. It turns out Voting, KNN, SVM, Descision Trre, Random Forest are the best models for prediction
#
# > The best accuracy I can get is with Voting with 90% accuracy and Random Forst with 89% accuracy uisng the Pearson's Correaltion Method for Feature Selection</font>
| DIabetes-20220107T024304Z-001/DIabetes/Diabetes_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #İNSTANCE VE ATRİBUTE ler ve
# # Sınıf içindeki fonksiyonlara METHOT DENİR.
class Superkahraman():
def __init__(self): #inislaser dan gelir başlatma fonksiyonu anlamında
print("init çalıştı. sınıf oluştu.") #her bir nesne oluştugunda init çalışır.
ali =Superkahraman()
metin = Superkahraman()
# #Şimdi metin ve ali isimli ik adet superkahraman sınııfnda nesne ürettik. Bu nesnelerin özelliklerinin olmasını
# isteyebiliriz. atribute denir.
class Superkahraman():
ozelGuc = "DAYANKILI"
def __init__(self, isim, yas, meslek):
print("init çalıştı. sınıf oluştu.")
self.isim = isim
self.yas=yas
self.meslek = meslek
cenk = Superkahraman("cenk",32,"Gazeteci")
cenk.meslek = "yazar"
cenk.meslek
cenk.yas
cenk.ozelGuc
cenk.ozelGuc = "<NAME>"
cenk.ozelGuc
class Superkahraman():
ozelGuc = "DAYANKILI"
def __init__(self, isim, yas, meslek):
print("init çalıştı. sınıf oluştu.")
self.isim = isim
self.yas=yas
self.meslek = meslek
def ornekguc(self):
print("Ben dürüst bir kahramanım.")
ali = Superkahraman("Ali",44,"TEKSTİL")
ali.ornekguc()
| OOP 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Active Qubit Reset on QCS
# In this notebook, we will walk through how to use **Active Qubit Reset** to drastically decrease the amount of time it takes to run a job on the QPU. Although we use a toy example in this notebook, the principles here can be extended to rapidly iterate through real-world applications such as optimizing a variational quantum algorithm (as we will do in the **Max-Cut QAOA** notebook).
#
# **NOTE**: This notebook depends on `pyquil >= 2.3.0`, which comes preinstalled on all new QMIs.
# +
import itertools
import time
from typing import List
from pyquil import get_qc, Program
from pyquil.gates import CNOT, H, MEASURE, RESET
# -
# ## Get Started
#
# Before running on the QPU, users must book a block of time on a lattice using the QCS command-line interface. To determine the lattices that are available for booking, you can run `qcs lattices` from within the QMI. For our [Aspen QPUs](https://www.rigetti.com/qpu), one of the currently available 8-qubit lattices is `Aspen-1-8Q-B`. Once your QPU time has started (which we call being "QPU-engaged"), you must then set up the `QuantumComputer` object associated with the booked lattice, which we do in the following cell.
#
# **NOTE**: When running this notebook, you will need to edit the `lattice` and `qubits` entries in the following cell to match whatever QPU lattice you end up booking. And remember that this code will only work from within the QMI!
lattice = 'Aspen-1-8Q-B' # edit as necessary
qpu = get_qc(lattice)
qubits = qpu.device.qubits()[-3:] # edit as necessary
print(f'All qubits on {lattice}: {qpu.device.qubits()}')
print(f'\nSelected qubits: {qubits}')
# ## Build GHZ Program
#
# We begin by putting the first qubit in the superposition state |+⟩ by using the [Hadamard](https://en.wikipedia.org/wiki/Quantum_logic_gate) gate. Then, we produce our [Greenberger–Horne–Zeilinger](https://en.wikipedia.org/wiki/Greenberger%E2%80%93Horne%E2%80%93Zeilinger_state) (GHZ) state (which looks like |000⟩ + |111⟩ for 3 qubits), by entangling all the qubits successively using [Controlled-NOT](https://en.wikipedia.org/wiki/Controlled_NOT_gate) (`CNOT`) gates. As in the **Parametric Compilation** notebook, we also declare our readout memory "ro", and measure each qubit into a readout register.
def ghz_program(qubits: List[int]) -> Program:
program = Program()
program.inst(H(qubits[0]))
for i in range(len(qubits) - 1):
program.inst(CNOT(qubits[i], qubits[i + 1]))
ro = program.declare('ro', 'BIT', len(qubits))
program.inst([MEASURE(qubit, ro[idx]) for idx, qubit in enumerate(qubits)])
return program
# ## Enable Active Reset
#
# We create two GHZ state programs, one with an initial `RESET` command, and one without. The `RESET` directive enables active qubit reset for all the measured qubits in the program. We then set the number of shots to take for each Quil program, and compile each into instrument binaries.
# +
program = Program()
program.inst(ghz_program(qubits))
program.wrap_in_numshots_loop(10_000)
binary = qpu.compile(program)
program_reset = Program(RESET())
program_reset.inst(ghz_program(qubits))
program_reset.wrap_in_numshots_loop(10_000)
binary_reset = qpu.compile(program_reset)
# -
# ## Compare Execution Time
#
# We run each binary on the QPU, comparing the total run time, to see a drastic speed increase with active qubit reset. For our Aspen-1 system, we are able to achieve an order of magnitude reduction in reset time between passive and active reset (~100μs vs. ~10μs). However, there are additional components beyond reset time that make up the total execution time, which is why we only see a ~5x overall improvement.
# +
start = time.time()
results = qpu.run(binary)
total = time.time() - start
print(f'Execution time without active reset: {total:.3f} s')
start_reset = time.time()
results_reset = qpu.run(binary_reset)
total_reset = time.time() - start_reset
print(f'\nExecution time with active reset: {total_reset:.3f} s')
# -
# ## Compare Execution Quality
#
# We compare the bitstring counts to see that there is no material performance hit for using active qubit reset. As we see in both count dictionaries, the `000` and `111` bitstrings are the most prevalent, as expected. However, we can see that the bitstring counts aren't perfect, which we can attribute to gate infidelity and decoherence.
# +
counts = {bit_tuple: 0 for bit_tuple in itertools.product((0, 1), repeat=3)}
for shot_result in results:
bit_tuple = tuple(shot_result)
counts[bit_tuple] += 1
print(f'Measurement results without active reset:')
for bit_tuple, count in counts.items():
print(bit_tuple, count)
counts_reset = {bit_tuple: 0 for bit_tuple in itertools.product((0, 1), repeat=3)}
for shot_result in results_reset:
bit_tuple = tuple(shot_result)
counts_reset[bit_tuple] += 1
print(f'\nMeasurement results with active reset:')
for bit_tuple, count in counts_reset.items():
print(bit_tuple, count)
| notebooks/qcs-only/ActiveQubitReset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
datos = pd.read_csv('events.csv')
datos_event = datos[datos["event"] == "ad campaign hit"]
datos_event[["person"]].dropna().drop_duplicates()
# +
datos_ = datos[["timestamp","person","event","campaign_source"]].dropna().groupby("campaign_source")["event"]
#datos_.apply(lambda x: 100 * x/float(x.sum()))
p = datos_.count().plot.barh(width = 0.5, figsize=(20,10))
p.axes.set_xscale("log")
p.set_xlabel("Cantidad de ads", size = 16)
p.set_ylabel("Origen del ad", size = 16)
p.set_title("Ads por donde se entra a la página", size = 22)
for v in p.patches:
p.annotate(str(int(v.get_width())), (v.get_x() + v.get_width()*1.5, v.get_y()), xytext=(-2, 4), textcoords='offset points', horizontalalignment='right')
| Ads mas efectivos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../')
# +
# %load_ext autoreload
# %autoreload 2
import sklearn
import copy
import numpy as np
import seaborn as sns
sns.set()
import scipy as sp
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import seaborn as sns
# from viz import viz
from bokeh.plotting import figure, show, output_notebook, output_file, save
from functions import merge_data
from sklearn.model_selection import RandomizedSearchCV
import load_data
from sklearn.linear_model import LinearRegression
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from fit_and_predict import fit_and_predict
# -
#
# ## Params:
aggregate_by_state = False
outcome_type = 'cases'
# ## Basic Data Visualization
# +
# Just something to quickly summarize the number of cases and distributions each day
# -
# 'deaths' and 'cases' contain the time-series of the outbreak
df = load_data.load_county_level(data_dir = '../data/')
df = df.sort_values('#Deaths_3/30/2020', ascending=False)
# outcome_cases = load_data.outcome_cases # most recent day
# outcome_deaths = load_data.outcome_deaths
important_vars = load_data.important_keys(df)
very_important_vars = ['PopulationDensityperSqMile2010',
# 'MedicareEnrollment,AgedTot2017',
'PopulationEstimate2018',
'#ICU_beds',
'MedianAge2010',
'Smokers_Percentage',
'DiabetesPercentage',
'HeartDiseaseMortality',
'#Hospitals'
# 'PopMale60-642010',
# 'PopFmle60-642010',
# 'PopMale65-742010',
# 'PopFmle65-742010',
# 'PopMale75-842010',
# 'PopFmle75-842010',
# 'PopMale>842010',
# 'PopFmle>842010'
]
def sum_lists(list_of_lists):
arr = np.array(list(list_of_lists))
sum_arr = np.sum(arr,0)
return list(sum_arr)
if aggregate_by_state:
# Aggregate by State
state_deaths_df = df.groupby('StateNameAbbreviation').deaths.agg(sum_lists).to_frame()
state_cases_df = df.groupby('StateNameAbbreviation').cases.agg(sum_lists).to_frame()
df = pd.concat([state_cases_df,state_deaths_df],axis =1 )
# +
# Distribution of the maximum number of cases
_cases = list(df['cases'])
max_cases = []
for i in range(len(df)):
max_cases.append(max(_cases[i]))
print('Number of counties with non-zero cases')
print(sum([v >0 for v in max_cases]))
# cases truncated below 20 and above 1000 for plot readability
plt.hist([v for v in max_cases if v > 20 and v < 1000],bins = 100)
# -
sum(max_cases)
print(sum([v > 50 for v in max_cases]))
np.quantile(max_cases,.5)
# +
# Distribution of the maximum number of cases
_deaths = list(df['deaths'])
max_deaths = []
for i in range(len(df)):
max_deaths.append(max(_deaths[i]))
print('Number of counties with non-zero deaths')
print(sum([v > 0 for v in max_deaths]))
# plt.hist(max_cases)
# print(sum([v >0 for v in max_cases]))
plt.hist([v for v in max_deaths if v > 5],bins=30)
# -
sum(max_deaths)
max(max_deaths)
np.quantile(max_deaths,.7)
# ### Clean data
# Remove counties with zero cases
max_cases = [max(v) for v in df['cases']]
df['max_cases'] = max_cases
max_deaths = [max(v) for v in df['deaths']]
df['max_deaths'] = max_deaths
df = df[df['max_cases'] > 0]
#
# ## Predict data from model:
method_keys = []
# clear predictions
for m in method_keys:
del df[m]
# +
# target_day = np.array([1])
# # Trains model on train_df and produces predictions for the final day for test_df and writes prediction
# # to a new column for test_df
# # fit_and_predict(df, method='exponential', outcome=outcome_type, mode='eval_mode',target_day=target_day)
# # fit_and_predict(df,method='shared_exponential', outcome=outcome_type, mode='eval_mode',target_day=target_day)
# # fit_and_predict(train_df, test_df,'shared_exponential', mode='eval_mode',demographic_vars=important_vars)
# # fit_and_predict(df,method='shared_exponential', outcome=outcome_type, mode='eval_mode',demographic_vars=very_important_vars,target_day=target_day)
# fit_and_predict(df, outcome=outcome_type, mode='eval_mode',demographic_vars=[],
# method='ensemble',target_day=target_day)
# fit_and_predict(df, outcome=outcome_type, mode='eval_mode',demographic_vars=[],
# method='ensemble',target_day=np.array([1,2,3]))
# # fit_and_predict(train_df, test_d f,method='exponential',mode='eval_mode',target_day = np.array([1,2]))
# # Finds the names of all the methods
# method_keys = [c for c in df if 'predicted' in c]
# method_keys
# +
# for days_ahead in [1, 2, 3]:
# for method in ['exponential', 'shared_exponential', 'ensemble']:
# fit_and_predict(df, method=method, outcome=outcome_type, mode='eval_mode',target_day=np.array([days_ahead]))
# if method == 'shared_exponential':
# fit_and_predict(df,method='shared_exponential',
# outcome=outcome_type,
# mode='eval_mode',
# demographic_vars=very_important_vars,
# target_day=np.array([days_ahead]))
# method_keys = [c for c in df if 'predicted' in c]
# geo = ['countyFIPS', 'CountyNamew/StateAbbrev']
# +
# method_keys = [c for c in df if 'predicted' in c]
# df_preds = df[method_keys + geo + ['deaths']]
# df_preds.to_pickle("multi_day_6.pkl")
# -
# ## Ensemble predictions
exponential = {'model_type':'exponential'}
shared_exponential = {'model_type':'shared_exponential'}
demographics = {'model_type':'shared_exponential', 'demographic_vars':very_important_vars}
linear = {'model_type':'linear'}
# +
# import fit_and_predict
# for d in [1, 2, 3]:
# df = fit_and_predict.fit_and_predict_ensemble(df,
# target_day=np.array([d]),
# mode='eval_mode',
# outcome=outcome_type,
# output_key=f'predicted_{outcome_type}_ensemble_{d}'
# )
# -
import fit_and_predict
for d in [1, 3, 5, 7]:
df = fit_and_predict.fit_and_predict_ensemble(df,
target_day=np.array(range(1, d+1)),
mode='eval_mode',
outcome=outcome_type,
methods=[exponential,
shared_exponential,
demographics,
linear
],
output_key=f'predicted_{outcome_type}_ensemble_{d}_with_exponential'
)
method_keys = [c for c in df if 'predicted' in c]
# +
# df = fit_and_predict.fit_and_predict_ensemble(df)
# -
method_keys
# ## Evaluate and visualize models
# ### Compute MSE and log MSE on relevant cases
# +
# TODO: add average rank as metric
# +
# Computes the mse in log space and non-log space for all columns
# -
def l1(arr1,arr2,norm=True):
"""
arr2 ground truth
arr1 predictions
"""
if norm:
sum_percent_dif = 0
for i in range(len(arr1)):
sum_percent_dif += np.abs(arr2[i]-arr1[i])/arr1[i]
return sum_percent_dif/len(arr1)
return sum([np.abs(a1-a2) for (a1,a2) in zip(arr1,arr2)])/len(arr1)
mse = sklearn.metrics.mean_squared_error
# Only evaluate points that exceed this number of deaths
# lower_threshold, upper_threshold = 10, 100000
lower_threshold, upper_threshold = 10, np.inf
# +
# Log scaled
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [np.log(p[-1] + 1) for p in df[key][(outcome > lower_threshold)]] # * (outcome < upper_threshold)]]
print('Log scale MSE for '+key)
print(mse(np.log(outcome[(outcome > lower_threshold) * (outcome < upper_threshold)] + 1),preds))
# -
# Log scaled
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [np.log(p[-1] + 1) for p in df[key][outcome > lower_threshold]]
print('Log scale l1 for '+key)
print(l1(np.log(outcome[outcome > lower_threshold] + 1),preds))
# No log scale
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [p[-1] for p in df[key][outcome > lower_threshold]]
print('Raw MSE for '+key)
print(mse(outcome[outcome > lower_threshold],preds))
# No log scale
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [p[-1] for p in df[key][outcome > lower_threshold]]
print('Raw l1 for '+key)
print(l1(outcome[outcome > lower_threshold],preds))
# No log scale
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [p[-1] for p in df[key][outcome > lower_threshold]]
print('Raw l1 for '+key)
print(l1(outcome[outcome > lower_threshold],preds,norm=False))
# ### Plot residuals
# +
# TODO: Create bounds automatically, create a plot function and call it instead of copying code, figure out way
# to plot more than two things at once cleanly
# Creates residual plots log scaled and raw
# We only look at cases with number of deaths greater than 5
# -
def method_name_to_pretty_name(key):
# TODO: hacky, fix
words = key.split('_')
words2 = []
for w in words:
if not w.isnumeric():
words2.append(w)
else:
num = w
model_name = ' '.join(words2[2:])
# model_name = 'model'
if num == '1':
model_name += ' predicting 1 day ahead'
else:
model_name += ' predicting ' +w+' days ahead'
return model_name
# Make log plots:
bounds = [1.5, 7]
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [np.log(p[-1]) for p in df[key][outcome > 5]]
plt.scatter(np.log(outcome[outcome > 5]),preds,label=method_name_to_pretty_name(key))
plt.xlabel('actual '+outcome_type)
plt.ylabel('predicted '+outcome_type)
plt.xlim(bounds)
plt.ylim(bounds)
plt.legend()
plt.plot(bounds, bounds, ls="--", c=".3")
plt.show()
# Make log plots zoomed in for the counties that have a fewer number of deaths
bounds = [1.5, 4]
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [np.log(p[-1]) for p in df[key][outcome > 5]]
plt.scatter(np.log(outcome[outcome > 5]),preds,label=method_name_to_pretty_name(key))
plt.xlabel('actual '+outcome_type)
plt.ylabel('predicted '+outcome_type)
plt.xlim(bounds)
plt.ylim(bounds)
plt.legend()
plt.plot(bounds, bounds, ls="--", c=".3")
plt.show()
# Make non-log plots zoomed in for the counties that have a fewer number of deaths# We set bounds
bounds = [10,400]
outcome = np.array([df[outcome_type].values[i][-1] for i in range(len(df))])
for key in method_keys:
preds = [p[-1] for p in df[key][outcome > 5]]
plt.scatter(outcome[outcome > 5],preds,label=method_name_to_pretty_name(key))
plt.xlabel('actual '+outcome_type)
plt.ylabel('predicted '+outcome_type)
plt.xlim(bounds)
plt.ylim(bounds)
plt.legend()
plt.plot(bounds, bounds, ls="--", c=".3")
plt.show()
# ### Graph Visualizations
# +
# Here we visualize predictions on a per county level.
# The blue lines are the true number of deaths, and the dots are our predictions for each model for those days.
# -
# +
def plot_prediction(row):
"""
Plots model predictions vs actual
row: dataframe row
window: autoregressive window size
"""
gold_key = outcome_type
for i,val in enumerate(row[gold_key]):
if val > 0:
start_point = i
break
# plt.plot(row[gold_key][start_point:], label=gold_key)
if len(row[gold_key][start_point:]) < 3:
return
sns.lineplot(list(range(len(row[gold_key][start_point:]))),row[gold_key][start_point:], label=gold_key)
for key in method_keys:
preds = row[key]
sns.scatterplot(list(range(len(row[gold_key][start_point:])))[-len(preds):],preds,label=method_name_to_pretty_name(key))
# plt.scatter(list(range(len(row[gold_key][start_point:])))[-len(preds):],preds,label=key)
# plt.legend()
# plt.show()
# sns.legend()
plt.title(row['CountyName']+' in '+row['StateNameAbbreviation'])
plt.ylabel(outcome_type)
plt.xlabel('Days since first death')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.figure(dpi=500)
plt.show()
# +
# feature_vals = {
# 'PopulationDensityperSqMile2010' : 1.1525491065255939e-05,
# "MedicareEnrollment,AgedTot2017" : -2.119520577282583e-06,
# 'PopulationEstimate2018' : 2.8898343032154275e-07,
# '#ICU_beds' : -0.000647030727828718,
# 'MedianAge2010' : 0.05032666600339253,
# 'Smokers_Percentage' : -0.013410742818946319,
# 'DiabetesPercentage' : 0.04395318355581005,
# 'HeartDiseaseMortality' : 0.0015473771787186525,
# '#Hospitals': 0.019248102357644396,
# 'log(deaths)' : 0.8805209010821442,
# 'bias' : -1.871552103871495
# }
# -
df = df.sort_values(by='max_deaths',ascending=False)
for i in range(len(df)):
row = df.iloc[i]
# If number of deaths greater than 10
if max(row['deaths']) > 10:
print(row['CountyName']+' in '+row['StateNameAbbreviation'])
plot_prediction(row)
for v in very_important_vars:
print(v+ ': '+str(row[v])) #+';\t contrib: '+ str(feature_vals[v]*float(row[v])))
print('\n')
| modeling/eda/basic_model_framework.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf_rl
# language: python
# name: tf_rl
# ---
# # 02. Neuralnet Q Learning example
#
# Neuralnet Q Learning을 실습해봅니다.
# - 신경망의 parameter(weight)를 업데이트 함에 따라 state에 대한 Q value가 변화합니다.
# - Q Learning의 TD error를 loss function으로 하여 학습합니다.
# ## Colab 용 package 설치 코드
# !pip install gym
# ### Package import
# +
import tensorflow as tf
import numpy as np
import random
import gym
import matplotlib.pyplot as plt
# %matplotlib inline
np.random.seed(777)
tf.set_random_seed(777)
print("tensorflow version: ", tf.__version__)
print("gym version: ", gym.__version__)
# -
# ## Frozen Lake
#
# **[state]**
#
# SFFF
# FHFH
# FFFH
# HFFG
#
# S : starting point, safe
# F : frozen surface, safe
# H : hole, fall to your doom
# G : goal, where the frisbee is located
#
# **[action]**
#
# LEFT = 0
# DOWN = 1
# RIGHT = 2
# UP = 3
# ### Frozen Lake (not Slippery)
# +
def register_frozen_lake_not_slippery(name):
from gym.envs.registration import register
register(
id=name,
entry_point='gym.envs.toy_text:FrozenLakeEnv',
kwargs={'map_name' : '4x4', 'is_slippery': False},
max_episode_steps=100,
reward_threshold=0.78, # optimum = .8196
)
register_frozen_lake_not_slippery('FrozenLakeNotSlippery-v0')
# -
# Load Environment
env = gym.make('FrozenLakeNotSlippery-v0')
state_size = env.observation_space.n
action_size = env.action_space.n
print("State_size : ", state_size)
print("Action_size: ",action_size)
# ## Q-Learning using Neural Network
# **Update 식**
#
# $J(w) = \mathbb{E}_{\pi}[(target - \hat q(S,A,w))^2]$
#
# $ \begin{align} \Delta w & = - \frac{1}{2} \alpha \nabla_w J(w)
# \\ & = \alpha (R_{t+1} + \gamma max(\hat q(S_{t+1},a ,w)) - \hat q(S_{t},A_{t},w))\nabla_w \hat q(S_{t},A_{t},w) \end{align}$
#
# ### 학습 순서
# 1. 초기 state 받음 (env.reset())
# 2. action 선택 (e - greedy policy)
# 3. 선택한 action으로 다음 state로 이동 (env.step())
# 4. 다음 state와 reward를 이용해 update식 작성
# 5. 신경망 업데이트
# 6. 반복
# ## Tensorflow 코드 흐름
# 1. 각 연산자에 대한 그래프를 구성한다.
# 2. 실제 데이터를 그래프에 할당하면서 전체 그래프를 실행한다.
# ### Build graph
# +
# placeholder 선언
# state
inputs = tf.placeholder(shape=[1], dtype=tf.int64)
# state에 대한 action
input_action = tf.placeholder(shape=[1], dtype=tf.int64)
# Loss 식의 target
target = tf.placeholder(shape=[1], dtype=tf.float32)
layers = tf.contrib.layers
# 신경망 구성 함수
# one-hot vector : 입력 1을 단순한 숫자로 받는 것보다 [1, 0, 0, 0] 처럼 encoding된 값으로 바꾸어 받는 것이
# 학습에 유리하다. 모든 입력이 크기에 관계없이 동등해지게 된다.
# tf.one_hot( 입력, one-hot size )
def build_network(inputs):
with tf.variable_scope('q_net'):
# 빈칸 {} 을 지우고 채워주세요.
input_onehot = tf.one_hot({}, {}, dtype=tf.float32)
fc1 = layers.fully_connected(inputs={},
num_outputs={},
activation_fn=None)
return fc1
# 신경망 구성
q_value = build_network(inputs)
# 현재 action에 대한 Q_value 구하는 연산
# q_value = [1, 2, 3, 4] curr_action = [0, 1, 0, 0] --(원소 곱)--> [0, 2, 0, 0] --(sum)--> [2]
curr_action = tf.one_hot({}, {})
curr_q_value = tf.reduce_sum(tf.multiply({}, {}))
# Loss 함수 구성
# 직접 구현해보세요. ( learning_rate = 0.1 )
# 참고) 제곱 : tf.square()
# optimizer : tf.train.GradientDescentOptimizer( learning_rate )
loss_op =
opt = tf.train.GradientDescentOptimizer(learning_rate=0.1)
train_op = opt.minimize(loss_op)
# -
# ### Executing a graph in a tf.Session
# +
# Session 열기
tf.reset_default_graph()
sess = tf.Session()
# 변수 초기화
sess.run(tf.global_variables_initializer())
# 변수(파라미터) 확인
for var in tf.trainable_variables('q_net'):
print(var)
print(sess.run(var))
# -
# ### Action select using Q value
# +
# 초기 state
state = env.reset()
state = np.reshape(state, [1])
print("Current state:", state)
# 현재 state에 대한 Q-value
# 빈칸 {}을 채워보세요.
# 참고) sess.run( "Q-value를 구하는 신경망 그래프", feed_dict={inputs: "신경망 입력"} )
curr_q = sess.run({}, feed_dict={inputs: {} })
print("Q value of current state:", curr_q)
# +
# action 선택 ( greedy )
action = sess.run({}, feed_dict={})
# 직접 구현해보세요. (hint. np.argmax)
print("Choose action: ", action)
# -
# ### 선택한 Action으로 다음 State, Reward 받기
# action을 이용해 env.step하기
# 빈칸 {} 을 채워보세요.
next_state, reward, done, _ = env.step({})
next_state = np.reshape(next_state, [1])
print("next_state", next_state)
# ### update를 위한 (미래)보상 값(target) 계산
gamma = 0.9
if done: # terminal state 이면 reward가 곧 next_q_value
next_q_value =
else:
# 직접 작성해보세요.
# 위 수식 참고.
# 참고) R + gamma * next state의 q-value 중 max
next_q_value =
print("next_q_value", next_q_value)
# ### Update Neural Net
# +
action = np.reshape(action, [1])
# train_op를 sess.run 하여 학습 실행.
# 빈칸 {} 을 채워보세요.
loss, _ = sess.run([loss_op, train_op], feed_dict={inputs: {}, target: {}, input_action: {}})
print("loss", loss)
for var in tf.trainable_variables('q_net'):
print(var)
print(sess.run(var))
# -
# ### 학습 시작
# +
rlist = []
slist = []
epsilon_list = []
EPISODE = 2000
gamma = 0.99
update_count = 0
loss_list = []
# Episode 수만큼 반복
for step in range(EPISODE):
# step마다 epsilon 감소
epsilon = 1. / ((step/50)+10)
epsilon_list.append(epsilon)
# 초기 state
state = env.reset()
state = np.reshape(state, [1])
print("[Episode {}]".format(step))
total_reward = 0
limit = 0
done = False
while not done and limit < 99:
# 위에서 했던 코드를 참조하여 아래 학습 코드를 작성해보세요.
# e-greedy policy로 action 선택
if epsilon > np.random.random():
# random
action = env.action_space.sample()
else:
# greedy
# 현재 state의 Q value의 argmax값을 sess.run().
curr_q =
action =
# 선택한 action으로 env.step 하기
next_state, reward, done, _ =
next_state = np.reshape(next_state, [1])
if reward == 1.0:
print("GOAL")
# 업데이트를 위한 (미래)보상값 반환
# episode가 끝났다면
if done:
next_q_value =
# 끝나지 않았다면
else:
next_q_value =
# Q update
action = np.reshape(action, [1])
loss, _ =
loss_list.append(loss)
update_count += 1
slist.append(state.item())
state = next_state
total_reward += reward
limit += 1
print(slist)
slist = []
print("total reward: ", total_reward)
rlist.append(total_reward)
print("Success Prob" + str(sum(rlist) / EPISODE) + "%")
# -
for var in tf.trainable_variables('q_net'):
print(var)
print(sess.run(var))
# epsilon 변화 그래프
steps = np.arange(EPISODE)
plt.title('Epsilon values')
plt.xlabel('Timestep')
plt.ylabel('$\\epsilon$')
plt.plot(steps, epsilon_list)
# loss 변화 그래프
update_count = np.arange(update_count)
plt.title('Loss values')
plt.xlabel('Update Count')
plt.ylabel('Loss')
plt.plot(update_count, loss_list)
# ### Test agent
# +
state = env.reset()
state = np.reshape(state, [1])
done = False
limit = 0
epsilon = 0.0
while not done and limit < 30:
# 학습된 신경망을 테스트하는 코드를 작성해보세요.
curr_q =
action =
next_state, reward, done, _ =
next_state = np.reshape(next_state, [1])
env.render()
state = next_state
limit += 1
# -
| 02_Neural_net_Q_Learning/02_Practice_Neural_net_Q_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import libraries
import numpy as np
import os
import pandas as pd
from sklearn import preprocessing
# environment settings
os.chdir("../data")
train_data = 'A2trainData_MMAI.xlsx'
test_data = 'A2testData_MMAI.xlsx'
# read in the train data
df_train = pd.read_excel(train_data)
df_test = pd.read_excel(test_data)
# get the shape of the train data
df_train.shape
# get the shape of the test data
df_test.shape
# get the head of the train data
df_train.head()
# get the head of the test data
df_test.head()
# get the tail of the train data
df_train.tail()
# get the tail of the test data
df_test.tail()
# rename all the columns to be without spaces and camel cases
df_train.columns = ['year', 'eps', 'liquidity', 'profitability', 'productivity', 'leverageRatio', 'marketBookRatio', 'tobinsQ', 'bUmd', 'excessReturnFromRiskModel', 'bHml', 'alpha', 'bSmb', 'returns', 'bMkt', 'totalVolatility', 'outputReturn']
df_test.columns = ['year', 'eps', 'liquidity', 'profitability', 'productivity', 'leverageRatio', 'marketBookRatio', 'tobinsQ', 'bUmd', 'excessReturnFromRiskModel', 'bHml', 'alpha', 'bSmb', 'returns', 'bMkt', 'totalVolatility', 'outputReturn']
# get all the train data columns
list(df_train.columns)
# get all the test data columns
list(df_test.columns)
# get the description of the train data
df_train.describe().transpose()
# get the description of the test data
df_test.describe().transpose()
# check for any null values in train data
df_train.isnull().any()
# check for any null values in train data
df_test.isnull().any()
# get list of unique years in the train data
np.unique(df_train['year'])
# get list of unique years in the test data
np.unique(df_test['year'])
# get correlation of train data
corr_train = df_train.corr()
corr_train.style.background_gradient()
# get correlation of test data
corr_test = df_test.corr()
corr_test.style.background_gradient()
# use >0.8 cut off for highly correlated features, and drop them (either returns or excessReturnFromRiskModel)
# no highly anti-correlated features, so we leave everything else as is
df_train = df_train.drop('returns', axis=1)
# drop year column
df_train = df_train.drop('year', axis=1)
# get all the train data columns
list(df_train.columns)
# use >0.8 cut off for highly correlated features, and drop them (either returns or excessReturnFromRiskModel)
# no highly anti-correlated features, so we leave everything else as is
df_test = df_test.drop('returns', axis=1)
# drop year column
df_test = df_test.drop('year', axis=1)
# get all the test data columns
list(df_test.columns)
# preprocess the data
scaler = preprocessing.MinMaxScaler()
# scale all the feature columns and disregard the target column using minmax scaler
df_train[df_train.columns] = scaler.fit_transform(df_train[df_train.columns])
df_test[df_test.columns] = scaler.transform(df_test[df_test.columns])
# get the head after scaling
df_train.head()
# get the head after scaling
df_test.head()
# get the tail after scaling
df_train.tail()
# get the tail after scaling
df_test.tail()
# describe the scaled data
df_train.describe().transpose()
# describe the scaled data
df_test.describe().transpose()
# write back the complete dataframe into one csv
df_train.to_csv(os.path.join(cwd, 'data', '2.0-sh-train-data.csv'), index=False)
df_test.to_csv(os.path.join(cwd, 'data', '2.0-sh-test-data.csv'), index=False)
| A2/cleansing/2.0-sh-data-cleansed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install PyPDF2
import PyPDF2
myfile = open('US_Declaration.pdf','rb')
pdf_reader = PyPDF2.PdfFileReader(myfile)
pdf_reader.numPages
page_one = pdf_reader.getPage(0)
page_one.extractText()
mytext = page_one.extractText()
print(mytext)
myfile.close()
# can't directly write python string directly into pdf, append
# # copy first page into the new filw
f = open('US_Declaration.pdf','rb')
pdf_reader = PyPDF2.PdfFileReader(f)
first_page = pdf_reader.getPage(0)
pdf_writer = PyPDF2.PdfFileWriter()
pdf_writer.addPage(first_page)
pdf_output = open('My_Brand_New.pdf','wb')
pdf_writer.write(pdf_output)
pdf_output.close()
f.close()
f = open('US_Declaration.pdf','rb')
pdf_reader = PyPDF2.PdfFileReader(f)
pdf_reader.numPages
f.close()
brand_new = open('My_Brand_New.pdf','rb')
pdf_reader = PyPDF2.PdfFileReader(brand_new)
pdf_reader.numPages
mytext = page_one.extractText()
print(mytext)
# whole pdf
f = open('US_Declaration.pdf','rb')
pdf_text = [0]
pdf_reader = PyPDF2.PdfFileReader(f)
for p in range(pdf_reader.numPages):
page = pdf_reader.getPage(p)
pdf_text.append(page.extractText())
f.close()
pdf_text
for page in pdf_text:
print(page)
print('\n')
print('\n')
print('\n')
print('\n')
| NLP_COURSE/00-Python-Text-Basics/PDF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Transpose of a Matrix
#
# In this set of exercises, you will work with the transpose of a matrix.
#
# Your first task is to write a function that takes the transpose of a matrix. Think about how to use nested for loops efficiently.
#
# The second task will be to write a new matrix multiplication function that takes advantage of your matrix transposition function.
# Takes in a matrix and outputs the transpose of the matrix
def transpose(matrix):
matrix_transpose = []
# Loop through columns on outside loop
for c in range(len(matrix[0])):
new_row = []
# Loop through columns on inner loop
for r in range(len(matrix)):
# Column values will be filled by what were each row before
new_row.append(matrix[r][c])
matrix_transpose.append(new_row)
return matrix_transpose
# +
### TODO: Run the code in the cell below. If there is no
### output, then your answers were as expected
assert transpose([[5, 4, 1, 7], [2, 1, 3, 5]]) == [[5, 2], [4, 1], [1, 3], [7, 5]]
assert transpose([[5]]) == [[5]]
assert transpose([[5, 3, 2], [7, 1, 4], [1, 1, 2], [8, 9, 1]]) == [[5, 7, 1, 8], [3, 1, 1, 9], [2, 4, 2, 1]]
# -
# ### Matrix Multiplication
#
# Now that you have your transpose function working, write a matrix multiplication function that takes advantage of the transpose.
#
# As part of the matrix multiplication code, you might want to re-use your dot product function from the matrix multiplication exercises. But you won't need your get_row and get_column functions anymore because the tranpose essentially takes care of turning columns into row vectors.
#
# Remember that if matrix A is mxn and matrix B is nxp, then the resulting product will be mxp.
# +
### TODO: Write a function called matrix_multiplication() that
### takes in two matrices and outputs the product of the two
### matrices
### TODO: Copy your dot_product() function here so that you can
### use it in your matrix_multiplication function
# Takes in two matrices and outputs the product of the two matrices
def dot_product(vectorA, vectorB):
result = 0
for i in range(len(vectorA)):
result += vectorA[i] * vectorB[i]
return result
def matrix_multiplication(matrixA, matrixB):
product = []
# Take the transpose of matrixB and store the result
transposeB = transpose(matrixB)
# Use a nested for loop to iterate through the rows
# of matrix A and the rows of the tranpose of matrix B
for r1 in range(len(matrixA)):
new_row = []
for r2 in range(len(transposeB)):
# Calculate the dot product between each row of matrix A
# with each row in the transpose of matrix B
dp = dot_product(matrixA[r1], transposeB[r2])
new_row.append(dp)
# Store the results in the product variable
product.append(new_row)
return product
# +
### TODO: Run the code in the cell below. If there is no
### output, then your answers were as expected
assert matrix_multiplication([[5, 3, 1],
[6, 2, 7]],
[[4, 2],
[8, 1],
[7, 4]]) == [[51, 17],
[89, 42]]
assert matrix_multiplication([[5]], [[4]]) == [[20]]
assert matrix_multiplication([[2, 8, 1, 2, 9],
[7, 9, 1, 10, 5],
[8, 4, 11, 98, 2],
[5, 5, 4, 4, 1]],
[[4],
[2],
[17],
[80],
[2]]) == [[219], [873], [8071], [420]]
assert matrix_multiplication([[2, 8, 1, 2, 9],
[7, 9, 1, 10, 5],
[8, 4, 11, 98, 2],
[5, 5, 4, 4, 1]],
[[4, 1, 2],
[2, 3, 1],
[17, 8, 1],
[1, 3, 0],
[2, 1, 4]]) == [[61, 49, 49], [83, 77, 44], [329, 404, 39], [104, 65, 23]]
# -
| Object-Tracking-4/23-Matrix-Transpose.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0
# language: julia
# name: julia-0.5
# ---
# # [DFA Preliminaries](./DFASlide.pdf)
logis(x) = 1 ./ (1 .+ exp(-x))
dlogis(x) = logis(x) .* (1 .- logis(x))
inp = [1 1 ; 0 1 ; 1 0 ; 0 0]'
out = [0 1 1 0]
y=zeros(size(out))
n_hidden = 10; # Number of hidden units
num_iterations = 1000; #Number of learning steps
trials = 4000;
w1 = randn(n_hidden,size(inp,1))
w2 = randn(n_hidden,n_hidden);
w3 = randn(size(out,1),n_hidden)
logis(w3 * (logis(w2 * (logis(w1 * inp)))))
bp_w1=w1
bp_w2=w2
bp_w3=w3
B1=rand(n_hidden,1)
B2=rand(n_hidden,1)
a1 = w1 * inp
z1 = logis(a1)
a2 = w2 * z1
z2 = logis(a2)
ay = w3 * z2
y .= logis(ay)
err=y-out
d_a1 = (B1.*err) .* dlogis(a1)
d_a2 = (B2*err) .* dlogis(a2)
dw1 = -d_a1 * inp'
dw2 = -d_a2 * z1'
dw3 = -err * z2'
w1 = w1 + dw1
w2 = w2 + dw2
w3 = w3 + dw3
bp_a1 = bp_w1*inp
bp_z1 = logis(bp_a1)
bp_a2 = bp_w2 * bp_z1
bp_z2 = logis(bp_a2)
bp_ay = bp_w3 * bp_z2
bp_y = logis(bp_ay)
bp_err = bp_y - out
bp_d3 = (bp_err) .* dlogis(bp_ay)
bp_d2 = (bp_w3' * bp_d3) .* dlogis(bp_a2)
bp_d1 = (bp_w2' * bp_d2) .* dlogis(bp_a1)
bp_w1 .-= (inp * bp_d1')'
bp_w2 .-= (z1 * bp_d2')'
bp_w3 .-= (z2 * bp_d3')'
e_store(ii,jj) = sum(abs(err))
bp_e_store(ii,jj) = sum(abs(bp_error))
# +
nprocs()==1 && addprocs()
@everywhere logis(x) = 1 ./ (1 .+ exp(-x))
@everywhere dlogis(x) = logis(x) .* (1 .- logis(x))
@everywhere function dfa!(w1,w2,w3,B1,B2,inp,out)
a1 = w1 * inp
z1 = logis(a1)
a2 = w2 * z1
z2 = logis(a2)
ay = w3 * z2
y = logis(ay)
err = y - out
d_a1 = (B1.*err) .* dlogis(a1)
d_a2 = (B2.*err) .* dlogis(a2)
w1 .-= d_a1 * inp'
w2 .-= d_a2 * z1'
w3 .-= err * z2'
return Dict("output"=>y,"err"=>err)
end
@everywhere function bp!(bp_w1,bp_w2,bp_w3,inp,out)
bp_a1 = bp_w1*inp
bp_z1 = logis(bp_a1)
bp_a2 = bp_w2 * bp_z1
bp_z2 = logis(bp_a2)
bp_ay = bp_w3 * bp_z2
bp_y = logis(bp_ay)
bp_err = bp_y - out
bp_d_a3 = (bp_err) .* dlogis(bp_ay)
bp_d_a2 = (bp_w3' * bp_d_a3) .* dlogis(bp_a2)
bp_d_a1 = (bp_w2' * bp_d_a2) .* dlogis(bp_a1)
bp_w1 .-= bp_d_a1 * inp'
bp_w2 .-= bp_d_a2 * bp_z1'
bp_w3 .-= bp_d_a3 * bp_z2'
return Dict("output"=>bp_y,"err"=>bp_err)
end
# +
inp = [1 1 ; 0 1 ; 1 0 ; 0 0]'
out = [0 1 1 0]
y=zeros(size(out))
n_hidden = 15; # Number of hidden units
num_iterations = 1000; #Number of learning steps
trials = 50;
e_store = SharedArray(Float64,(num_iterations,trials))
bp_e_store = SharedArray(Float64,(num_iterations,trials))
y_store = SharedArray(Float64,(size(out,2),num_iterations))
bp_y_store = SharedArray(Float64,(size(out,2),num_iterations))
@elapsed @sync @parallel for jj = 1:trials
w1 = randn(n_hidden,size(inp,1))
w2 = randn(n_hidden,n_hidden)
w3 = randn(size(out,1),n_hidden)
B1=rand(n_hidden,1)
B2=rand(n_hidden,1)
bp_w1 = randn(n_hidden,size(inp,1))
bp_w2 = randn(n_hidden,n_hidden)
bp_w3 = randn(size(out,1),n_hidden)
for ii = 1:num_iterations
dfa_res = dfa!(w1,w2,w3,B1,B2,inp,out)
bp_res = bp!(bp_w1,bp_w2,bp_w3,inp,out)
if jj == 1
y_store[:,ii] = dfa_res["output"]'
bp_y_store[:,ii] = bp_res["output"]'
end
e_store[ii,jj] = sum(abs(dfa_res["err"]))
bp_e_store[ii,jj] = sum(abs(bp_res["err"]))
end
end
# +
using Plots
l = @layout [
a b
c d
]
gr()
p1=plot(y_store',label=["1,1" "0,1" "1,0" "0,0"],title="DFA")
p2=plot(bp_y_store',label=["1,1" "0,1" "1,0" "0,0"],title="BP")
p3=plot([e_store[:,1] bp_e_store[:,1]],label=["DFA" "BP"],title="First Trial")
p4=plot([mean(e_store,2) mean(bp_e_store,2)],label=["DFA" "BP"],title="Mean Error")
plot(p1,p2,p3,p4,layout=l)
| DFA_OLD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from muzero import MuZero
mz = MuZero('anytrading')
mz.load_model(
checkpoint_path= 'results/anytrading/2021-01-15--05-57-26/model.checkpoint',
replay_buffer_path='results/anytrading/2021-01-15--05-57-26/replay_buffer.pkl',
)
mz.test(render=True, opponent="self", muzero_player=None)
| eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ep38]
# language: python
# name: conda-env-ep38-py
# ---
# # Open converted files
# ## Open a converted netCDF or Zarr dataset
# Converted netCDF files can be opened with the open_converted function that returns a lazy loaded EchoData object (only metadata are read during opening):
# ```python
# import echopype as ep
# file_path = "./converted_files/file.nc" # path to a converted nc file
# ed = ep.open_converted(file_path) # create an EchoData object
# ```
# Likewise, specify the path to open a Zarr dataset. To open such a dataset from cloud storage, use the same `storage_options` parameter as with [open_raw](convert.html#aws-s3-access). For example:
# ```python
# s3_path = "s3://s3bucketname/directory_path/dataset.zarr" # S3 dataset path
# ed = ep.open_converted(s3_path, storage_options={"anon": True})
# ```
# ## Combine EchoData objects
# Converted data found in multiple files corresponding to the same instrument deployment can be combined into a single `EchoData` object. First assemble a list of `EchoData` objects from the converted files (netCDF or Zarr). Then apply `combine_echodata` on this list to combine all the data into a single `EchoData` object in memory:
# ```python
# ed_list = []
# for converted_file in ["convertedfile1.nc", "convertedfile2.nc"]:
# ed_list.append(ep.open_converted(converted_file))
#
# combined_ed = ep.combine_echodata(ed_list)
# ```
# ## EchoData object
# `EchoData` is an object that conveniently handles raw converted data from either raw instrument files (via `open_raw`) or previously converted and standardized raw files (via `open_converted`). It is essentially a container for multiple `xarray Dataset` objects, where each such object corresponds to one of the netCDF4 groups specified in the SONAR-netCDF4 convention followed by echopype. `EchoData` objects are used for conveniently accessing and exploring the echosounder data, for calibration and other processing, and for [serializing into netCDF4 or Zarr file formats](convert.html#file-export).
#
# A sample `EchoData` object is presented below using the `Dataset` HTML browser generated by `xarray`, collected into SONAR-netCDF4 groups. Select each group and drill down to variables and attributes to examine the structure and representative content of an `EchoData` object.
# + tags=["remove-cell"]
from pathlib import Path
import echopype as ep
bucket = "ncei-wcsd-archive"
rawdirpath = "data/raw/Bell_M._Shimada/SH1707/EK60/Summer2017-D20170728-T181619.raw"
s3raw_fpath = f"s3://{bucket}/{rawdirpath}"
ed = ep.open_raw(s3raw_fpath, sonar_model='EK60', storage_options={'anon': True})
# Manually populate additional metadata about the dataset and the platform
# -- SONAR-netCDF4 Top-level Group attributes
ed.top.attrs['title'] = "2017 Pacific Hake Acoustic Trawl Survey"
ed.top.attrs['summary'] = (
f"EK60 raw file {s3raw_fpath} from the {ed.top.attrs['title']}, "
"converted to a SONAR-netCDF4 file using echopype."
)
# -- SONAR-netCDF4 Platform Group attributes
ed.platform.attrs['platform_type'] = "Research vessel"
ed.platform.attrs['platform_name'] = "Bell M. Shimada"
ed.platform.attrs['platform_code_ICES'] = "315"
# + tags=["remove-input"]
ed
| docs/source/open-converted.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
temp = 212.0
def toCelsius(fahrenheit):
return (fahrenheit - 32) * 5.0 / 9.0
print toCelsius(temp)
# +
import pandas as pd
from pandas.io import gbq
print "Imports run."
# +
projectId = "YOUR-PROJECT-ID-HERE"
sql = "SELECT year, month, day, weight_pounds FROM [publicdata:samples.natality] LIMIT 50"
print 'Running query...'
data = gbq.read_gbq(sql, project_id=projectId)
data[:5]
# -
| courses/unstructured/BigQuery-test-solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tunable Coupler (MIT)
# +
# %load_ext autoreload
# %autoreload 2
import qiskit_metal as metal
from qiskit_metal import designs, draw
from qiskit_metal import MetalGUI, Dict, Headings
import pyEPR as epr
# -
# ### Create the design in Metal
# Setup a design of a given dimension. Dimensions will be respected in the design rendering. <br>
# Note that the design size extends from the origin into the first quadrant.
# +
design = designs.DesignPlanar({}, True)
design.chips.main.size['size_x'] = '3mm'
design.chips.main.size['size_y'] = '3mm'
gui = MetalGUI(design)
# -
# Create two crossmons coupled together with an interdigitated tunable coupler.
from qiskit_metal.qlibrary.qubits.transmon_cross_fl import TransmonCrossFL
from qiskit_metal.qlibrary.qubits.tunable_coupler_01 import TunableCoupler01
TransmonCrossFL.get_template_options(design)
TunableCoupler01.get_template_options(design)
# +
Q1 = TransmonCrossFL(design, 'Q1', options = dict(pos_x = '0', pos_y='-0.3mm',
connection_pads = dict(
bus_01 = dict(connector_location = '180',claw_length ='95um'),
readout = dict(connector_location = '0')),
fl_options = dict()))
Q2 = TransmonCrossFL(design, 'Q2', options = dict(pos_x = '0', pos_y='0.3mm', orientation = '180',
connection_pads = dict(
bus_02 = dict(connector_location = '0',claw_length ='95um'),
readout = dict(connector_location = '180')),
fl_options = dict()))
tune_c_Q12 = TunableCoupler01(design,'Tune_C_Q12', options = dict(pos_x = '-0.06mm', pos_y = '0',
orientation=90, c_width='500um'))
gui.rebuild()
gui.autoscale()
# +
# Get a list of all the qcomponents in QDesign and then zoom on them.
all_component_names = design.components.keys()
gui.zoom_on_components(all_component_names)
# -
#Save screenshot as a .png formatted file.
gui.screenshot()
# + tags=["nbsphinx-thumbnail"]
# Screenshot the canvas only as a .png formatted file.
gui.figure.savefig('shot.png')
from IPython.display import Image, display
_disp_ops = dict(width=500)
display(Image('shot.png', **_disp_ops))
# -
# Closing the Qiskit Metal GUI
gui.main_window.close()
| docs/circuit-examples/qubit-couplers/TwoCrossmonsTunableCoupler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# Forest fires are a natural disaster that have significant environmental ramifications, and have increased in frequency which coincides with the issue of climate change. In the United States alone, over 70,000 forest fires burn an average of 7 million acres every year (1). Wildfires are often directly influenced by human interaction, with close to 90% having some link to a man-made ignition point (1). Another major factor in the spread of forest fires is the topography of the land. Fire travels much faster uphill than downhill, as it is able to preheat the fuel above the flames, meaning that the steeper the gradient of the hill, the faster the fire will spread (2). Finally, the type of fuel the fire is burning through contributes to the speed, flame size, and temperature of the fire. Larger fuels, such as trees and heavy slash (fallen branches, brush, etc.), burns hotter, with larger flame sizes than smaller fuels such as grass and thin brush (spread out over a larger area) (3). Historically, forests have contained roughly 50-70 trees per acre, while today, in some parts of the United States, forests contain anywhere from 500-1000 trees per acre (3). This overcrowding of forests has led to both high mortality rates among trees, and explosive forest fires that devastate entire tree populations in record times.
#
# The forest fire model involves a multitude of physical factors, yet all of them cannot be evaluated in this project. This research will consider three physical attributes that can influence the spread of a fire: the varying slopes of hills and valleys, the type of fuel the fire is burning through, and the concentration of trees in the simulated forest. With lower probabilties for a contribution to fire spread, ground vegetation will have less of a chance of spreading from cell to cell than larger fuels like the trees in this scenario. In order to mimic a forest that has sub-optimum conditions for a forest fire, the concentration of trees for this project will be above 5000 trees per 10000 cells. Finally, in an effort to mimic terrain that doesn't rise and fall in an easily modeled fashion, the slopes that trees are on will be generated in a fashion that will be described in further detail in the model section below. The larger the slope value, the more likely the transition from one cell to the next.
#
# While forest fires can have conditions that change rapidly, it is necessary to set up some constants that will set the boundaries for the model below. The size of the forest will be predetermined for each experiment, but the simulation will end once the probability for fire has been calculated for every cell within the "forest". The goal of this simulation is to experiment with ways to reduce the transition rate of fire from one cell to the next, in an effort to determine actual physical measures that humans can take to limit the impact of forest fires. Therefore, multiple simulations will be run with different possible values for slope size, and different values of ground vegetation fire transition. By the end of this project, the research should convey data on how the aforementioned factors relate to the spread of fire, as well as which preventitive measures (changing the variables) achieved effective results.
# ## Model
# In order to simulate a forest, this project will generate a two-dimensional grid that assigns each cell a value of either empty, a tree, or an active burn designation. The constants in this project are the probabilities for a lightning strike, new tree growth, vegetation, as well as the fraction of the forest that is already occupied by trees. Using an initialized forest grid, the cell values are assigned based on whether the numerical values in the initialized grid add up to the determined numerical values for the cell designation, or are within the probability constants. The hill was determined using the two-dimensional Gaussian distribution function:
# $ Aexp(−((x−xo)^2/(2σ_X^2)+(y−yo)^2/(2σ_Y^2)) $. The derivative of this function will also be used to determine the slope of each calculated cell. The forest simulation will involve a 100x100 grid, and the simulation will terminate once all of the cells have been evaluated for cell designation. The generated forest cell is shown below. The green cells are trees, the black cells are empty vegetation cells, and the orange cells are active fire cells.
# ## Numerical Methods
# In order to solve the Gaussian function for this project, the x0 and y0 values will be established through the mean value of the hill coordinates to the nearest integer. The spread of the distribution will be calculated by subtracting the minimum coordinate value from the maximum coordinate value for both the x and y values. In order to find the slope at each point within the coordinate field, a point will be generated on each side of the selected coordinate and the generated point with the smaller coordinate value of the two will be subtracted from the larger point in order to determine the slope for the desired coordinate. This will be repeated for each coordinate point within the Gaussian distribution. The slope will then be measured against burn probabilities in order to assign a burn or empty designation. The forest generated outside of the hill will receive cell designations based on whether the coordinate values from the initialized forest grid add up to a numerical designation for cell assignment. If the cell designation is empty, a random number will be generated and if that number fits within the probability for new tree growth, it will be reassigned a tree designation. Finally, the ground vegetation will be determined through a random number generation and will be given an equal probability to burn or remain intact. These methods are limited by accuracy, as the slope will be a generalization, due to the both the method, and the nature of hills to not be exactly alligned with a Gaussian distribution. Also, the use of random numbers and probabilities limit the project, as these are merely estimates, and can not exactly replicate the varying behavior of forest fires.
# ## Code
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import animation
from matplotlib import colors
rng = np.random.default_rng() # Create a default random number Generator
print(rng.uniform(1, 3)) # Test case
# +
X1 = 1 #defining an initial X1 so the code runs
def ground_vegetation(X,iy, ix):
"""Determines whether a space is empty vegetation or on fire.
Given the determination that ground fuel doesn't burn consistently, this function
assigns a burn designation for cells determined to have ground vegetation, based on a
random generation of a number. Only possibilities are empty cells, which indicates
non-burning ground vegetation, or fire designation, indicating vegetation on fire.
x = randomly generated number which is the basis for the burn designation."""
x = np.random.random()
if x < .5:
X1[iy,ix] = FIRE
else:
X1[iy,ix] = EMPTY
return X1[iy, ix]
def Gaussian(X, hill_min, hill_max, A):
"""Determines burn status for fuel on a determined hill.
Using the dervative of the Two Dimensional Gaussian distribution, this function
evaluates the slope at each point within the range, and proceeds to assign a burn
probability for that point. The burn probability is then used to evaluate whether
each point is on fire or not.
y0 = y-coordinate center of hill
x0 = x-coordinate center of hill
range_max = last point evaluated within the hill
hill_min = x,y coordinate of the beginning of the hill
hill_max = x,y coordinate of the end of the hill
x_spread = spread of distribution in x axis
y_spread = spread of distribution in y axis
A = variable for amplitude of hill
X2 = list of burn assignments for cells on the hill."""
X2 = [] #initializes burn list
y0 = (hill_max - hill_min)//2 #determines center point/peak of the hill
x0 = (hill_max - hill_min)//2 #in both x and y coordinates
range_max = (hill_max - hill_min) + 1 #sets range for point calculation
x_spread = hill_max - hill_min #sets the spread of the distribution
y_spread = hill_max - hill_min
for x in range(1, range_max):
for y in range(1, range_max):
#value for larger generated coordinate point
t = A * np.exp(-((((x+.5)-x0)**2/(2*(x_spread**2))) + (((y+.5)-y0)**2/(2*(y_spread**2)))))
#value for smaller generated coordinate point
q = A * np.exp(-((((x-.5)-x0)**2/(2*(x_spread**2))) + (((y-.5)-y0)**2/(2*(y_spread**2)))))
slope = t - q #determines slope value
if slope > 1: #if slope is steep, FIRE designation is guaranteed
X2.append(2)
elif -1 < slope < 1: #at top of hill, probability of FIRE decreases linearly
x = np.random.random()
if x > .25:
X2.append(2)
else:
X2.append(1)
else:
x = np.random.random #going downhill, probability of FIRE decreases further linearly
if x > .5:
X2.append(2)
else:
X2.append(1)
return X2
# the neighbourhood is the nearest 8 cells to the point in consideration
neighbourhood = ((-1,-1), (-1,0), (-1,1), (0,-1), (0, 1), (1,-1), (1,0), (1,1))
EMPTY, TREE, FIRE = 0, 1, 2 #number values for cell designations
# Colours for visualization: brown for EMPTY, dark green for TREE and orange
# for FIRE.
colors_list = [(0.2,0,0), (0,0.5,0), (1,0,0), 'orange']
cmap = colors.ListedColormap(colors_list) #sets up colors for simulation
bounds = [0,1,2,3]
norm = colors.BoundaryNorm(bounds, cmap.N)
def iterate(X, hill_min, hill_max, A):
"""Iterate the forest according to the forest-fire rules.
Given the number values for cell designation, this function generates a forest of
empty cells of ground vegetation, cells with trees that aren't on fire, and cells
that are actively burning. It also generates a hill within the forest.
X = initialized forest grid
hill_min = beginning of hill coordinates
hill_max = end of hill coordinates
X1 = coordinate list with the cell designations
A = variable for amplitude of hill
"""
Empty = []
Tree = []
Fire = []
X1 = np.zeros((ny, nx)) #initializes cells in the forest
for ix in range(1, hill_min):
for iy in range(1,hill_min): #values for cells before the hill
if X[iy,ix] == EMPTY and np.random.random() <= p: #assigns tree values based on new-tree growth
X1[iy,ix] = TREE #probability
Tree.append(1)
if X[iy,ix] == TREE:
X1[iy,ix] = TREE
Tree.append(1)
if X[iy, ix] == EMPTY and p < np.random.random() < b:
x = np.random.random()
if x < .5:
X1[iy,ix] = FIRE
Fire.append(2)
else:
X1[iy,ix] = EMPTY #generates vegetation burn designations
Empty.append(0)
for dx,dy in neighbourhood:
if X[iy+dy,ix+dx] == FIRE: #assigns a FIRE designation for cells within the neighborhood
X1[iy, ix] == FIRE #that have a numerical value equal to the FIRE value
Fire.append(2)
break
else:
if np.random.random() <= f: #assigns a FIRE designation for cells that are struck
X1[iy,ix] = FIRE #by lightning based on the lightning probability
Fire.append(2)
for ix in range(hill_min,hill_max): #assigns cell designations for cells within the hill
for iy in range(hill_min,hill_max):
if X[iy,ix] == EMPTY and np.random.random() <= p:
X1[iy,ix] = TREE
Tree.append(1)
if X[iy,ix] == TREE: #determines tree and empty cells for the hill
X1[iy,ix] = TREE
Tree.append(1)
if X[iy, ix] == EMPTY and p < np.random.random() < b:
x = np.random.random()
if x < .5:
X1[iy,ix] = FIRE
Fire.append(2)
else:
X1[iy,ix] = EMPTY #determines ground vegetation cells
Empty.append(0)
for iy in range(hill_min, hill_max): #assigns burn designations for the hill
X2 = Gaussian(X, hill_min, hill_max, A)
length = len(X2) #determines length of list
for i in range(0, length-1):
w = X2[i] #determines value of coordinate point
if w == 2:
X1[iy, ix] == FIRE #assigns cell designation for hill coordinates
Fire.append(2)
else:
X1[iy, ix] == TREE
Tree.append(1)
for ix in range(hill_max,nx-1):
for iy in range(hill_max,ny-1): #values for cells after the hill
if X[iy,ix] == EMPTY and np.random.random() <= p:
X1[iy,ix] = TREE
Tree.append(1)
#determines tree and empty cells
if X[iy,ix] == TREE:
X1[iy,ix] = TREE
Tree.append(1)
if X[iy, ix] == EMPTY and p < np.random.random() < b:
x = np.random.random()
if x < .5:
X1[iy,ix] = FIRE
Fire.append(2)
else:
X1[iy,ix] = EMPTY #determines ground vegetation cells
Empty.append(0)
for dx,dy in neighbourhood:
if X[iy+dy,ix+dx] == FIRE: #assigns a FIRE designation for cells within the neighborhood
X1[iy, ix] == FIRE #that have a numerical value equal to the FIRE value
Fire.append(2)
break
else:
if np.random.random() <= f: #assigns a FIRE designation for cells that are struck
X1[iy,ix] = FIRE #by lightning based on the lightning probability
Fire.append(2)
return X1, Empty, Tree, Fire
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.05, .85, .001
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Base Case")
X1, empty, tree, fire = iterate(X, 71, 80, 3)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
def animate(i):
"""Creates an animation progression for each generation of iteration function.
Uses the iteration function to display the results of the fire simulation.
X = the initialized forest grid."""
im.set_data(animate.X)
animate.X = iterate(animate.X, 71, 80, 3)
#animate.X = X
# Interval between frames
#interval = 100
#anim = animation.FuncAnimation(fig, animate, interval=interval)
#plt.show()
# +
# Second test case with higher lightning, lower vegetation, and lower new-growth probabilties. Gaussian hill is also
#steeper
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.02, .50, .003
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 2")
X1, empty, tree, fire = iterate(X, 71, 80, 5)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Third test case with lower lightning, higher vegetation, and higher new-growth probabilties. Gaussian hill is also
#less steep
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.10, .9, .0005
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 3")
X1, empty, tree, fire = iterate(X, 71, 80, .05)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Fourth test case with higher initial forest fraction, higher new-growth probabilties, and high vegetation probablilites
#Gaussian hill is also less steep
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.4
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.15, .9, .0005
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 4")
X1, empty, tree, fire = iterate(X, 71, 80, .05)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Fifth test case with lower lightning, low vegetation, and low new-growth probabilties. Gaussian hill is also
#very steep
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.2, .65, .0005
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 5")
X1, empty, tree, fire = iterate(X, 71, 80, 10)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Sixth test case with lower lightning, higher vegetation, and low new-growth probabilties. Gaussian hill is also
#less steep
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.02, .95, .0005
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 6")
X1, empty, tree, fire = iterate(X, 71, 80, 1)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Seventh test case simply changed the location of the Gaussian hill
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.10, .9, .0005
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 7")
X1, empty, tree, fire = iterate(X, 15, 24, 3)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Eight test case displays how a larger hill impacts the output numbers
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.10, .9, .0005
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 8")
X1, empty, tree, fire = iterate(X, 50, 80, .05)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# +
#Ninth test case with extremely high lightning, low tree, and low vegetation probabilities, Gaussian hill is not steep
# The initial fraction of the forest occupied by trees.
forest_fraction = 0.2
# Probability of new tree growth per empty cell, vegetation per empty cell, and of lightning strike.
p, b, f = 0.05, .65, .1
# Forest size (number of cells in x and y directions).
nx, ny = 100, 100
# Creates the forest grid used in the iteration function
X = np.zeros((ny, nx))
X[1:ny-1, 1:nx-1] = np.random.randint(0, 2, size=(ny-2, nx-2))
X[1:ny-1, 1:nx-1] = np.random.random(size=(ny-2, nx-2)) < forest_fraction
#creates the plot
fig = plt.figure(figsize=(25/3, 6.25))
ax = fig.add_subplot(111)
ax.set_axis_off()
im = ax.imshow(X, cmap=cmap, norm=norm)
ax.set_title("Fig. 9")
X1, empty, tree, fire = iterate(X, 71, 80, 3)
print("Empty =", len(empty))
print("Tree =", len(tree))
print("Fire =", len(fire))
# -
# ## Results
# Most of these results are reasonable, as the output numbers are close to the total cell value within the grid. In order to evaluate the impacts of forest density, lighting, and the hills on the simulation, I ran a variety of trials while varying those variables. Larger forest density and significant slope size on the hills were the two biggest factors in large fire cell outputs.
# ## Analysis
# Our results seem reasonable, but only when the hill is of a certain size value. I verified this because the output numbers added up to be very close to 10,000 which is the total value of the grid. However, when the hill size was made larger, the output numbers added up to well above the total grid value, which leads to the conclusion that the code adds mutliple cell designations to cells when the simulation runs a large hill. However, with specified boundaries for the hill size, I was able to evaluate the impacts of the other variables on the spread of the fire. The probability of lightning strike was not a significant factor, but rather the slope of the hill and the density of the forest were the most significant factors in the generation of fire cells. With a large slope or a dense vegetation and tree population, the fire cell output numbers were larger, and with a less dense forest and low vegetation the fire cell output dropped significantly. When we created a forest with a larger number of initial trees the fire cell output was also significantly larger.
# ## Summary
# Scientific Summary: My project attempted to model the spread of a forest fire with varying physical factors. I achieved this with the use of a Gaussian distribution equation, and a variance of probabilities for new tree growth, vegetation and the start of a fire. I showed that the slope of a the hill impacts the rate at which the fire grows, and that density of the forest is key to how devastating a fire will be. This means that more dense forests will have a higher burn probability, and a fire will most likely decimate a tree population when moving uphill. These findings are very important for the fight against forest fires.
#
# Personal Summary: This project was meaningful for me because I live so close to areas that have been ravaged by forest fires. I enjoyed being able to rewrite the same code over and over again until it finally did what I wanted it to do. This project was a great example of how perserverance in computational physics can be vital to the success of a project, and it was exciting to finally achieve a model that was close to a real-life simulation.
# ## References
# Appendix B Fire and Fuels, National Geographic Area Coordination Centers, Fire Behavior Fuel Model Descriptions, (2001).
# Influence of slope on fire spread rate, <NAME>, <NAME>, <NAME>, The Fire Environment-Innovations, Managementm and Policy: Conference Proceedings (P), (2007).
# Wildfire Statistics, <NAME>, <NAME>, Congressional Research Service (34), (2019).
| Final Project (4).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit (conda)
# language: python
# name: python38364bitcondaefb26bb7c6fc42ada05cffadde411c01
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Python Data Science Excercise
# ## Part 1
# #### First Testing
# + pycharm={"name": "#%%\n"}
print("Test?!")
# + [markdown] pycharm={"name": "#%% md\n"}
# Next test
#
# + pycharm={"name": "#%%\n"}
# -
| 01_mm_my_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# (2mins)=
# # A 2-minute primer
#
# ## Self-managing experiments
# With Mandala, the way you compose experiments out of function calls and
# collection creation/indexing automatically induces the organization of storage.
# For example, below are line-by-line analogous code blocks to
# - save results of an imagined clustering experiment (**left**);
# - query results from this and all analogous experiments in the storage (**right**):
#
# ```python
# with run(storage, lazy=True): with query(storage) as q:
# X, y = get_data(preprocess=True) X, y = get_data(preprocess=True)
# for n_clusters in (2, 4, 8): n_clusters = Query(int)
# clusters = get_clusters(X, y, n_clusters) clusters = get_clusters(X, y, n_clusters)
# score = eval_clusters(clusters, y) score = eval_clusters(clusters, y)
# if score > 0.95: score > 0.95
# for cluster in clusters: cluster = clusters[IndexQuery()]
# centroid = get_centroid(X, cluster) centroid = get_centroid(X, cluster)
# df = q.get_table(n_clusters,
# score, centroid)
# ```
# These blocks look a lot like code to just run computations, without data
# management concerns -- but in fact support various natural ways to store and
# interact with results.
#
# ### The `run` context: use code to traverse storage directly
# If you re-run `run`-wrapped code -- *or any sub-program of it* -- function calls
# you've already computed will load their results as needed to allow control flow
# to proceed. Consequences include
# - **flexible queries**: you can get to the results you want by directly
# retracing the steps that created them, however complicated they may be (an
# **imperative** query interface)
# - **easy iteration**: you can organically grow a piece of code with new
# parameters and functionality, without re-doing expensive work
# - **resilience**: recovering from failure is as simple as running the same
# code again -- which will retrace the steps that completed successfully, up to
# the failed computations.
#
# ### The `query` context: search in storage by pattern-matching to code
# With the `query` block, you
# - interpret function calls and collection operations as building a structure of
# dependencies between variables (such as `n_clusters` and `score`);
# - point to a sequence of variables to get a table where each row is a
# matching of values to these variables satisfying the dependencies (a
# **declarative** query interface)
#
# ## Next steps: abstraction and refactoring
# Read on to see how
# - these patterns can be composed with **abstraction** and **refactoring** to
# scale them up to evolving projects with many components
# - you can use imperative/declarative patterns similar to the above for deletions
| docs/_build/html/_sources/intros/two_mins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FantAPPalla server
#
# Documenting each of the functions in the server. So far I have completed up to
# +
#Assigning this notebook as the server for the app
#import anvil.server
#from anvil import Image, XYPanel
#anvil.server.connect("N2RJZAZY3RKJ65AUOA5JKOLC-GFGPDCRRTFT46NHQ")
# +
# Importing all the utilities
from utilities_stats import *
import copy
from anvil import URLMedia
import anvil.media
import datetime
from datetime import date
import time
#from joblib import Parallel, delayed
#Setting the total number of matchdays in the season
tot_giornate = 35
# setting useful parameters
Teams, Logos, parameters, Results_0, goal_marks = set_par(fasce = 2)
giornate = current_matchday()
# filling individual dataframes and conditional display
Results = fill_dataframe_partita(Results_0, giornate, parameters, goal_marks, Teams, Print = False)
# saving cumulative statistical data
pf_med, pf_std, ps_med, ps_std, gf_med, gf_std = cumulative_data(Results, giornate, Print = False)
# Building cumulative dataframe
Total = make_Total_df(Results, giornate, goal_marks)
Tot_per_round = partial_totals(Results, giornate, tot_giornate, goal_marks)
#Total.head(10)
# +
#####################################################
### TEAM STATS
#####################################################
def score_label(df, giornata):
return '%d-%d' % (df.at[giornata,'GF'], df.at[giornata,'GS'])
def best_worst_games(df):
'''Returns two dictionaries: best win and worst loss in Campionato'''
df['scarto_goal'] = df['GF'] - df['GS']
df['scarto_fp'] = df['punti_fatti'] - df['punti_subiti']
giornata_best_win = df['scarto_goal'].idxmax()
if df.at[giornata_best_win,'esito'] == 'V':
best_win = get_matchday_dict(df, giornata_best_win)
else: best_win = None
giornata_worst_loss = df['scarto_goal'].idxmin()
if df.at[giornata_worst_loss,'esito'] == 'S':
worst_loss = get_matchday_dict(df, giornata_worst_loss)
else: worst_loss = None
return best_win, worst_loss
def scontri_diretti(df, avversario):
'''Returns a disctionary with results against an opponent'''
df = df[df['avversario'] == avversario]
punti = df['pti'].sum()
media_punti = punti/len(df)
wins = len(df[df['esito'] == 'V'])
draws = len(df[df['esito'] == 'P'])
losses = len(df[df['esito'] == 'S'])
dic = {'avversario': avversario, 'vittorie': wins, 'pareggi': draws, 'sconfitte': losses}
best_win, worst_loss = best_worst_games(df)
dic['best_win'] = best_win
dic['worst_loss'] = worst_loss
dic['media_punti'] = media_punti
return dic
def get_matchday_dict(df, giornata):
'''Returns a disctionary with info about a matchday of a given team'''
dic = {}
dic['giornata'] = giornata
dic['avversario'] = df.at[giornata, 'avversario']
dic['score'] = score_label(df, giornata)
dic['GF'] = df.at[giornata, 'GF']
dic['GS'] = df.at[giornata, 'GS']
dic['fantapunti_fatti'] = df.at[giornata, 'punti_fatti']
dic['fantapunti_subiti'] = df.at[giornata, 'punti_subiti']
dic['esito'] = df.at[giornata, 'esito']
#--- ignobel dict
dic['ignobel'] = {}
dic['ignobel']['goal_subiti_por'] = df.at[giornata, 'goal_subiti_por']
dic['ignobel']['cartellini'] = df.at[giornata, 'cartellini']
dic['ignobel']['bonus_panchina'] = df.at[giornata, 'bonus_panchina']
dic['ignobel']['infortunati'] = df.at[giornata, 'infortunati']
dic['ignobel']['mod_difesa'] = df.at[giornata, 'mod_difesa']
return dic
def results_arr(df):
'''Returns a disctionary with results of a team'''
results_array = []
for giornata in range(1, len(df)+1):
dic = get_matchday_dict(df, giornata)
results_array.append(dic)
return results_array
def get_close_games_dict(df, threshold = 2, verbose=False):
'''Returns a disctionary with results of a team of close_games'''
giornate = len(df)
close_games_dict = {}
close_games_arr = []
pti = 0
n_close_games = 0
for gg in df.index:
if (np.abs(df['punti_fatti'][gg] - df['punti_subiti'][gg])) < threshold:
matchday_dict = get_matchday_dict(df, gg)
close_games_arr.append(matchday_dict)
res = esito(df.GF[gg], df.GS[gg])
if verbose:
print('G', gg, '| punti fatti:', df.punti_fatti[gg], ' ( subiti:', df.punti_subiti[gg],') |', res, '(', df.GF[gg], '-', df.GS[gg], ')')
pti = pti+punti(res)
n_close_games = n_close_games + 1
if n_close_games > 0:
if verbose: print('---> %d punti in %d giornate \n (media: %.2f)' % (pti, n_close_games, pti/n_close_games))
close_games_dict['punti'] = pti
close_games_dict['n_close_games'] = n_close_games
close_games_dict['media_punti'] = float(pti) / n_close_games
else:
if verbose: print('---> No close games found')
close_games_dict['punti'] = pti
close_games_dict['n_close_games'] = n_close_games
close_games_dict['media_punti'] = float(pti) / n_close_games
close_games_dict['games_list'] = close_games_arr
return close_games_dict
def get_esito_repetitions(Result, owner, esito):
'''Returns number of wins/draws/losses'''
df = Results[owner]
df = df[df['esito'] == esito]
return len(df)
#---------------------------------------------------------------------
# TRENDING TEAMS FUNCTIONS
#---------------------------------------------------------------------
from itertools import groupby
def get_longest_streak_dict(Results, owner):
'''Returns max number of consecutive wins/draws/losses'''
dic = {}
for esito in ['V', 'P', 'S']:
esito_list = list(Results[owner]['esito']) # lista di esito
cons_results = [(x[0], len(list(x[1]))) for x in groupby(esito_list)] #risultati consecutivi raggruppati
cons_results_esito = [x for x in cons_results if x[0]==esito] #solo risultati consecuti di uno specifico esito
try:
max_cons_esito = max(cons_results_esito, key=lambda x:x[1])
n_cons_games = max_cons_esito[1]
dic[esito] = n_cons_games
except: dic[esito] = 0
return dic
def get_current_trend(Results, owner, last_games = 5):
dic = {}
esito_list = list(Results[owner]['esito'])
giornate = min(last_games, len(esito_list))
esito_list = esito_list[-giornate:]
dic['results'] = esito_list
media_punti = np.sum(list(Results[owner]['pti'])[-giornate:])/giornate
dic['media_punti'] = media_punti
return dic
#---------------------------------------------------------------------
#################################################################
### FUNCTION BUILDING THE COMPLETE STATS DICTIONARY BY TEAM
#################################################################
def get_team_stats(Results, owner):
'''Returns a dictionary with all the season info by team'''
giornate = current_matchday()
db = {}
#--- TOP LEVEL KEYS ---
db['owner'] = owner
db['total'] = {} # cumulated raw data
db['season_data'] = {} # elaborated season statistics
db['calendar'] = [] # list of mathcday data
db['close_games'] = {} # games with scarto fantapunti < fascia gol
db['scontri_diretti'] = [] # list of cumulated data against each opponent
#----------------------
#--- total dictionary
db['total']['punti'] = Results[owner]['pti'].sum() # punti in calssifica
db['total']['vittorie'] = get_esito_repetitions(Results, owner, 'V')
db['total']['pareggi'] = get_esito_repetitions(Results, owner, 'P')
db['total']['sconfitte'] = get_esito_repetitions(Results, owner, 'S')
db['total']['fantapunti_fatti'] = Results[owner]['punti_fatti'].sum() # fantapunti fatti
db['total']['fantapunti_subiti'] = Results[owner]['punti_subiti'].sum() # fantapunti subiti
db['total']['GF'] = Results[owner]['GF'].sum() # gol fatti
db['total']['GS'] = Results[owner]['GS'].sum() # gol subiti
#--- total dictionary --- ignobel dictionary
db['total']['ignobel'] = {}
db['total']['ignobel']['goal_subiti_por'] = Results[owner]['goal_subiti_por'].sum() # gol subiti dal portiere schierato
db['total']['ignobel']['cartellini'] = Results[owner]['cartellini'].sum() # cartellini giocatori schierati
db['total']['ignobel']['bonus_panchina'] = Results[owner]['bonus_panchina'].sum() # bonus giocatori in panchina
db['total']['ignobel']['infortunati'] = Results[owner]['infortunati'].sum() # n. giocatori infortunati in rosa
db['total']['ignobel']['mod_difesa'] = Results[owner]['mod_difesa'].sum() # modificatoree difesa
#--- season data
db['season_data']['media_punti'] = db['total']['punti']/giornate
#--- season data --- fantapunti_fatti dictionary
db['season_data']['fantapunti_fatti'] = {}
db['season_data']['fantapunti_fatti']['mean'] = np.mean(Results[owner]['punti_fatti'])
db['season_data']['fantapunti_fatti']['std'] = np.std(Results[owner]['punti_fatti'])
db['season_data']['fantapunti_fatti']['max'] = max(Results[owner]['punti_fatti'])
db['season_data']['fantapunti_fatti']['min'] = min(Results[owner]['punti_fatti'])
#--- season data --- fantapunti_subiti dictionary
db['season_data']['fantapunti_subiti'] = {}
db['season_data']['fantapunti_subiti']['mean'] = np.mean(Results[owner]['punti_subiti'])
db['season_data']['fantapunti_subiti']['std'] = np.std(Results[owner]['punti_subiti'])
db['season_data']['fantapunti_subiti']['max'] = max(Results[owner]['punti_subiti'])
db['season_data']['fantapunti_subiti']['min'] = min(Results[owner]['punti_subiti'])
#--- season data --- gol fatti dictionary
db['season_data']['GF'] = {}
db['season_data']['GF']['mean'] = np.mean(Results[owner]['GF'])
db['season_data']['GF']['std'] = np.std(Results[owner]['GF'])
db['season_data']['GF']['max'] = max(Results[owner]['GF'])
db['season_data']['GF']['min'] = min(Results[owner]['GF'])
#--- season data --- gol subiti dictionary
db['season_data']['GS'] = {}
db['season_data']['GS']['mean'] = np.mean(Results[owner]['GS'])
db['season_data']['GS']['std'] = np.std(Results[owner]['GS'])
db['season_data']['GS']['max'] = max(Results[owner]['GS'])
db['season_data']['GS']['min'] = min(Results[owner]['GS'])
best_win, worst_loss = best_worst_games(Results[owner])
db['season_data']['best_win'] = best_win # matchday dictionary of best win (highest GF-GS)
db['season_data']['worst_loss'] = worst_loss # matchday dictionary of worst loss (highest GS-GF)
#--- season data --- trends
db['season_data']['longest_streaks'] = get_longest_streak_dict(Results, owner) # dictionary of longest stresks by esito
db['season_data']['current_trend'] = get_current_trend(Results, owner, last_games = 5) #dictionary with current results and media punti
#--- calendar
db['calendar'] = results_arr(Results[owner]) # array of matchday dictionaries (see get_matchday_dict())
#--- close games
db['close_games'] = get_close_games_dict(Results[owner]) # dictionary with overall info about close games and with an array of matchday info for those games
#--- scontri diretti
for avv in Results[owner]['avversario'].unique():
db['scontri_diretti'].append(scontri_diretti(Results[owner], avv)) # array of overall stats collected against a specific opponent
return db
###################################################################################################################
# +
#####################################################
### DICTIONARY FOR CAMPIONATO STATS DB - CURRENT SEASON
#####################################################
def get_team_totals(owner, Results = Results, Total = Total, Tot_per_round = Tot_per_round):
'''Returns a dictionary with total stats by team'''
df_prev = Tot_per_round[-2]
df_prev = df_prev[df_prev['Team'] == owner]
df = Total[Total['Team'] == owner]
dic = {}
dic['owner'] = owner
dic['posizione_precedente'] = int(df_prev['pos'].iat[0]) # posizione giornata precedente
dic['posizione'] = int(df['pos'].iat[0])
dic['punti'] = df['pti'].iat[0]
dic['vittorie'] = get_esito_repetitions(Results, owner, 'V')
dic['pareggi'] = get_esito_repetitions(Results, owner, 'P')
dic['sconfitte'] = get_esito_repetitions(Results, owner, 'S')
dic['fantapunti_fatti'] = df['punti_fatti'].iat[0]
dic['fantapunti_subiti'] = df['punti_subiti'].iat[0]
dic['expected_fantapunti_subiti'] = df['x_punti_subiti'].iat[0]
dic['GF'] = df['GF'].iat[0]
dic['GS'] = df['GS'].iat[0]
dic['expected_GS'] = df['x_GS'].iat[0]
#--- ignobel totals
dic['ignobel'] = {}
dic['ignobel']['goal_subiti_por'] = df['goal_subiti_por'].iat[0] # gol subiti dal portiere schierato
dic['ignobel']['cartellini'] = df['cartellini'].iat[0] # cartellini giocatori schierati
dic['ignobel']['bonus_panchina'] = df['bonus_panchina'].iat[0] # bonus giocatori in panchina
dic['ignobel']['infortunati'] = df['infortunati'].iat[0] # n. giocatori infortunati in rosa
dic['ignobel']['mod_difesa'] = df['mod_difesa'].iat[0] # modificatoree difesa
#--- ignobel --- previous game totals
dic['ignobel']['previous_goal_subiti_por'] = df_prev['goal_subiti_por'].iat[0] # gol subiti dal portiere schierato
dic['ignobel']['previous_cartellini'] = df_prev['cartellini'].iat[0] # cartellini giocatori schierati
dic['ignobel']['previous_bonus_panchina'] = df_prev['bonus_panchina'].iat[0] # bonus giocatori in panchina
dic['ignobel']['previous_infortunati'] = df_prev['infortunati'].iat[0] # n. giocatori infortunati in rosa
dic['ignobel']['previous_mod_difesa'] = df_prev['mod_difesa'].iat[0] # modificatoree difesa
return dic
def get_season_max_min(season_data_variable, team_stats_dict):
'''Returns max and min values and owner across teams results'''
arr_max = []
arr_min = []
owners_list = list(Results.keys())
for owner in owners_list:
arr_max.append(team_stats_dict[owner]['season_data'][season_data_variable]['max'])
arr_min.append(team_stats_dict[owner]['season_data'][season_data_variable]['min'])
max_val = max(arr_max)
max_owner = owners_list[arr_max.index(max_val)]
dict_max = {'value': max_val, 'owner': max_owner}
min_val = min(arr_min)
min_owner = owners_list[arr_min.index(min_val)]
dict_min = {'value': min_val, 'owner': min_owner}
return dict_max, dict_min
from random import sample
def make_calendar_array_univoque():
'''
Returns array of dictionaries with 4 items
owner: opponent
'''
calendar_list = make_calendar_array()
owners_found = []
matchday_dic = {}
calendar_matchups = []
owners_list = list(Results.keys())[:]
for dic in calendar_list:
for owner in sample(owners_list, len(owners_list)):
if dic[owner] not in owners_found:
matchday_dic[owner] = dic[owner]
owners_found.append(owner)
calendar_matchups.append(matchday_dic)
owners_found = []
matchday_dic = {}
return calendar_matchups
def get_campionato_calendar_array(team_stats_dict):
'''Returns an array of matchday dictionaries with matches info'''
matchups_list = make_calendar_array_univoque()
match_id = 0
games_array = []
for i, matchups in enumerate(matchups_list):
giornata = i + 1
for owner in matchups:
home_owner = owner
away_owner = matchups[owner]
match_tmp = team_stats_dict[home_owner]['calendar'][i]
single_match_dict = {}
single_match_dict['match_id'] = match_id
single_match_dict['giornata'] = giornata
match_id = match_id + 1
single_match_dict['home'] = {}
single_match_dict['home']['owner'] = home_owner
single_match_dict['home']['fantapunti'] = match_tmp['fantapunti_fatti']
single_match_dict['home']['GF'] = match_tmp['GF']
single_match_dict['home']['ignobel'] = match_tmp['ignobel']
single_match_dict['away'] = {}
single_match_dict['away']['owner'] = away_owner
single_match_dict['away']['fantapunti'] = match_tmp['fantapunti_subiti']
single_match_dict['away']['GF'] = match_tmp['GS']
match_tmp = team_stats_dict[away_owner]['calendar'][i]
single_match_dict['away']['ignobel'] = match_tmp['ignobel']
games_array.append(single_match_dict)
return games_array
#---------------------------------------------------------------------
# NOTABLE MATCHES FUNCTIONS
#---------------------------------------------------------------------
def get_highest_scoring_match(calendar_array):
'''
Returns game dictionary for the game with
max fantapunti
'''
parameter_arr = []
for game in calendar_array:
game['tot_fantapunti'] = game['home']['fantapunti'] + game['away']['fantapunti']
parameter_arr.append(game['tot_fantapunti'])
val_of_interest = max(parameter_arr)
index_of_interest = parameter_arr.index(val_of_interest)
game_of_interest = calendar_array[index_of_interest]
return game_of_interest
def get_lowest_scoring_match(calendar_array):
'''
Returns game dictionary for the game with
min fantapunti
'''
parameter_arr = []
for game in calendar_array:
game['tot_fantapunti'] = game['home']['fantapunti'] + game['away']['fantapunti']
parameter_arr.append(game['tot_fantapunti'])
val_of_interest = min(parameter_arr)
index_of_interest = parameter_arr.index(val_of_interest)
game_of_interest = calendar_array[index_of_interest]
return game_of_interest
def get_highest_differential_match(calendar_array):
'''
Returns game dictionary for the game with
max differenza goal
'''
parameter_arr = []
for game in calendar_array:
game['scarto_gol'] = np.abs(game['home']['GF'] - game['away']['GF'])
parameter_arr.append(game['scarto_gol'])
val_of_interest = max(parameter_arr)
index_of_interest = parameter_arr.index(val_of_interest)
game_of_interest = calendar_array[index_of_interest]
return game_of_interest
def get_highest_scoring_draw_match(calendar_array):
'''
Returns game dictionary for the game with
max fantapunti in un pareggio
'''
parameter_arr = []
games_arr = []
for game in calendar_array:
game['tot_fantapunti'] = game['home']['fantapunti'] + game['away']['fantapunti']
if (game['home']['GF'] == game['away']['GF']):
parameter_arr.append(game['tot_fantapunti'])
games_arr.append(game)
val_of_interest = max(parameter_arr)
index_of_interest = parameter_arr.index(val_of_interest)
game_of_interest = games_arr[index_of_interest]
return game_of_interest
def get_perfect_ties(calendar_array):
'''
Returns array of games dictionary for the games with
fantapunti uguali
'''
parameter_arr = []
games_arr = []
for game in calendar_array:
if (game['home']['fantapunti'] == game['away']['fantapunti']):
games_arr.append(game)
return games_arr
def get_zero_goal_matches(calendar_array):
'''
Returns array of games dictionary for the games with
0-0
'''
parameter_arr = []
games_arr = []
for game in calendar_array:
if (game['home']['GF'] == 0) & (game['away']['GF'] == 0):
games_arr.append(game)
return games_arr
def get_max_min_ignobel_matches(calendar_array, ig_par):
'''
Returns max and min games dictionary for the games with
max and min ignobel parameter total
'''
parameter_arr = []
games_arr = []
for game in calendar_array:
value = game['home']['ignobel'][ig_par] + game['away']['ignobel'][ig_par]
parameter_arr.append(value)
games_arr.append(game)
val_max = max(parameter_arr)
val_min = min(parameter_arr)
index_max = parameter_arr.index(val_max)
index_min = parameter_arr.index(val_min)
game_max = games_arr[index_max]
game_min = games_arr[index_min]
return game_max, game_min
#----------------------------------------------------------------------------
#----------------------------------------------------------------------------
# TRENDING TEAMS
#----------------------------------------------------------------------------
def longest_streaks_campionato(team_stats_dict):
'''Returns dictionary with longest V/P/S streaks'''
dic = {}
owners_list = list(Results.keys())
for esito in ['V', 'P', 'S']:
arr = []
for owner in owners_list:
arr.append(team_stats_dict[owner]['season_data']['longest_streaks'][esito])
max_val = max(arr)
index = arr.index(max_val)
owner_max = owners_list[index]
dic[esito] = {'owner': owner_max, 'length': max_val}
return dic
def get_hot_cold_teams(team_stats_dict):
'''Returns team with better results in last 5 games'''
owners_list = list(Results.keys())
arr = []
for owner in owners_list:
arr.append(team_stats_dict[owner]['season_data']['current_trend']['media_punti'])
max_val = max(arr)
index_max = arr.index(max_val)
owner_max = owners_list[index_max]
min_val = min(arr)
index_min = arr.index(min_val)
owner_min = owners_list[index_min]
dic_max = {'owner': owner_max, 'current_trend': team_stats_dict[owner_max]['season_data']['current_trend']}
dic_min = {'owner': owner_min, 'current_trend': team_stats_dict[owner_min]['season_data']['current_trend']}
return dic_max, dic_min
def get_pareggite_team(standings):
arr = []
owners_list = []
for team_totals in standings:
arr.append(team_totals['pareggi'])
owners_list.append(team_totals['owner'])
max_val = max(arr)
index_max = arr.index(max_val)
owner_max = owners_list[index_max]
return {'owner': owner_max, 'pareggi': max_val}
#----------------------------------------------------------------------------
##########################################################################
# MAIN FUNCTION BUILDING THE COMPLETE CAMPIONATO DICTIONARY
##########################################################################
def get_campionato_stats(team_stats_dict, Results = Results, Total = Total):
'''Returns a dictionary with cumulative info of Campionato'''
db = {}
db['calendar'] = [] # array with matches per matchday
db['standings'] = [] # array with teams info dictionary
db['season_data'] = {} # dictionary of campionanto stats data
db['notable_matches'] = {} # dictionary of specific matches to remember
db['trending_teams'] = {} # dictionary of teams status
#--- calendar
db['calendar'] = get_campionato_calendar_array(team_stats_dict) # calendar array
#--- standings
for owner in Results.keys():
db['standings'].append(get_team_totals(owner))
#--- season data
db['season_data']['media_punti'] = Total['pti'].sum()/len(Total)/giornate
#--- season data --- fantapunti_fatti dictionary
db['season_data']['fantapunti_fatti'] = {}
db['season_data']['fantapunti_fatti']['median'] = np.median(Total['punti_fatti']/giornate)
db['season_data']['fantapunti_fatti']['std'] = np.std(Total['punti_fatti']/giornate)
max_d, min_d = get_season_max_min('fantapunti_fatti', team_stats_dict)
db['season_data']['fantapunti_fatti']['max'] = max_d
db['season_data']['fantapunti_fatti']['min'] = min_d
#--- season data --- gol fatti dictionary
db['season_data']['GF'] = {}
db['season_data']['GF']['median'] = np.median(Total['GF']/giornate)
db['season_data']['GF']['std'] = np.std(Total['GF']/giornate)
max_d, min_d = get_season_max_min('GF', team_stats_dict)
db['season_data']['GF']['max'] = max_d
db['season_data']['GF']['min'] = min_d
#--- notable matches
db['notable_matches']['highest_scoring'] = get_highest_scoring_match(db['calendar']) # partita con piu' fantapunti
db['notable_matches']['lowest_scoring'] = get_lowest_scoring_match(db['calendar']) # partita con meno fantapunti
db['notable_matches']['highest_scoring_draw'] = get_highest_scoring_draw_match(db['calendar']) # pareggio ad alta quota (max fantapunti)
db['notable_matches']['highest_differential'] = get_highest_differential_match(db['calendar']) # partita con maggiore scarto di gol
db['notable_matches']['perfect_ties'] = get_perfect_ties(db['calendar']) # ARRAY di partite in perfetta parita' (uguali fantapunti)
db['notable_matches']['zero_goal'] = get_zero_goal_matches(db['calendar']) # ARRAY di partite 0-0
#--- notable matches --- ignobel stats
for ig_par in ['goal_subiti_por', 'cartellini', 'infortunati', 'bonus_panchina', 'mod_difesa']:
game_max, game_min = get_max_min_ignobel_matches(db['calendar'], ig_par)
db['notable_matches']['max_' + ig_par] = game_max
db['notable_matches']['min_' + ig_par] = game_min
#--- trending teams
db['trending_teams']['longest_streaks'] = longest_streaks_campionato(team_stats_dict) # dictionary of longest V/P/S streaks
hot_team, cold_team = get_hot_cold_teams(team_stats_dict)
db['trending_teams']['pareggite_team'] = get_pareggite_team(db['standings']) # dictionary with team with highest number of pareggi
db['trending_teams']['hot_team'] = hot_team # dictionary with team with highest media_punti in last 5 games
db['trending_teams']['cold_team'] = cold_team # dictionary with team with lowest media_punti in last 5 games
return db
#########################################################################################
# +
##########################################################################
# BUILDING THE COMPLETE DICTIONARY OF THE SEASON WITH
# SEASON ID, TEAM STATS, CAMPIONATO STATS
##########################################################################
def get_season_stats(Results = Results, Total = Total, Tot_per_round = Tot_per_round, season = '1900/01'):
'''Returns a dictionary with the season info of all teams'''
db = {}
db['season'] = season # season string
db['team_stats'] = {}
for owner in Results.keys():
db['team_stats'][owner] = get_team_stats(Results, owner)
db['campionato_stats'] = get_campionato_stats(team_stats_dict = db['team_stats'])
return db
season_dict = get_season_stats(season = '2020/21')
# +
############# SERVER FUNCTIONS ###################
#--- TEAM STATS
@anvil.server.callable
def team_stats_db(owner):
"""
Returns a dictionary with several info about team stats in Campionato
"""
dic = season_dict['team_stats'][owner]
return dic
#--- CAMPIONATO STATS
@anvil.server.callable
def campionato_stats_db(owner):
"""
Returns a dictionary with several info about team stats in Campionato
"""
dic = season_dict['campionato_stats']
return dic
# +
# ^
# |
# |
# |
########### NEW SERVER FUNCTIONS #################
########### TO BE DEPRECATED #################
# |
# |
# |
# v
######################################################################
# Dictionary for form: Squadra.Stats
######################################################################
def old_team_stats_dict(owner):
'''Builds a nested dictionary with stats in Campionato per team'''
#stats_dict = {}
df = Results[owner]
team_stats_dict = {}
team_stats_dict['media_fantapunti_fatti'] = np.mean(df['punti_fatti'])
team_stats_dict['media_fantapunti_subiti'] = np.mean(df['punti_subiti'])
team_stats_dict['media_GF'] = np.mean(df['GF'])
team_stats_dict['media_GS'] = np.mean(df['GS'])
team_stats_dict['media_punti'] = np.mean(df['pti'])
team_stats_dict['max_fantapunti'] = max(df['punti_fatti'])
team_stats_dict['min_fantapunti'] = min(df['punti_fatti'])
best_win, worst_loss = best_worst_games(df)
team_stats_dict['best_win'] = best_win
team_stats_dict['worst_loss'] = worst_loss
team_stats_dict['results'] = results_arr(df)
team_stats_dict['close_games'] = get_close_games_dict(df)
team_stats_dict['scontri_diretti'] = []
for avv in df['avversario'].unique():
team_stats_dict['scontri_diretti'].append(scontri_diretti(df, avv))
return team_stats_dict
# Anvil server function
@anvil.server.callable
def team_stats(owner):
"""
Returns a dictionary with several info about team stats in Campionato
"""
dic = old_team_stats_dict(owner)
return dic
# -
<EMAIL>
def current_matchDay():
"""
Returns the current matchday, based on the local database of the matches data in the utilities python script.
"""
return giornate
<EMAIL>
def generate_plots():
"""
Generates all the plots for the IGnobel section and for the Cfactor section and saves them in a local folder.
"""
giornate = current_matchday()
Results = fill_dataframe_partita(Results_0, giornate, parameters, goal_marks, Teams, Print = False)
for premio in ['Caduti','Porta Violata','Catenaccio','Panchina Oro','Cartellino Facile']:
_ = premio_plot(Results, giornate, Teams, Logos, premio)
fortuna_evo(Results, Teams, Tot_per_round)
C_factor_logos_2(Total, giornate, Teams, tot_giornate, Logos)
# +
#generate_plots()
# +
# Imports all the libraries needed to manage the database
import json
from pymongo import MongoClient
from pprint import pprint
import pymongo
from datetime import datetime
with open('credential.json','r') as f:
cred = json.load(f)
#creates the variables needed to manage the database
cluster = MongoClient(cred['cred'])
# choosing database
db = cluster["Game"]
# choosing collection
collection = db["Players"]
collection_man = db['Managers']
collection_tr = db['Transfers']
collection_temp = db["tempPlayers"]
collection_man_temp = db['tempManagers']
collection_tr_temp = db['tempTransfers']
# -
<EMAIL>
def man_team_name(owner):
"""
Given the owner of the team, fetches the name of the team from mongodb and returns it
"""
owner = owner.lower()
dic = collection_man.find_one({'owner': owner})
return dic['team_name']
# +
<EMAIL>
def rose_funct(owner, squad):
"""
This is one of the most fundamental functions about a manager's lineup, it returns all possible information
about the lineup, both contract players, including on loan, and loanee. Age, value, cost info etc.
It is not called directly but it is used in another function to merge info about main and primavera.
"""
flip_squad={
'main':'primavera',
'primavera':'main'
}
owner = owner.lower()
squad = squad.lower()
players = []
value_init = 0
value_now = 0
mean_age = 0
tot_cost = 0
p_num_dict={
'a contratto':0,
'dentro in prestito':0,
'fuori in prestito':0
}
posts = collection.find({'info.contract.owner': owner,'info.current_team.squad': squad})
for player in posts:
#check if loanee player comes from owner's squad
temp = ''
if player['info']['current_team']['on_loan']:
if squad not in player['info']['current_team']['previous_team']:
continue
else:
p_num_dict['fuori in prestito'] +=1
temp = '**'
p_num_dict['a contratto'] +=1
name_url = player['name']
name_url = name_url.replace(' ','-')
name_url = name_url.replace('.','')
dag = ''
cost_eff = player['info']['contract']['cost']
if player['info']['personal_info']['team_real'] is None:
stats_link = ''
dag = '\u2020'
else:
stats_link = 'https://www.fantacalcio.it/squadre/'+player['info']['personal_info']['team_real']+'/'+name_url+'/'+str(player['_id'])
age = int(np.floor((datetime.today()-datetime.strptime(player['info']['personal_info']['birthdate'], "%d/%m/%Y")).days/365.4))
players.append({'role': player['info']['personal_info']['FC_role']
, 'name': player['name']
, 'age': age
, 'quotation': player['info']['stats']['Qt_A']
, 'quotation_initial': player['info']['contract']['quotation_initial']
, 'difference': int(player['info']['stats']['Qt_A']) - int(player['info']['current_team']['quotation_initial'])
, 'loan': temp+dag,
'link': stats_link,
'owner':owner,
'cost':cost_eff,
'complete_db': player})
value_init += int(player['info']['contract']['quotation_initial'])
value_now += int(player['info']['stats']['Qt_A'])
mean_age += int(age)
tot_cost += int(player['info']['contract']['cost'])
posts = collection.find({'info.contract.owner': owner,'info.current_team.on_loan': True,'info.current_team.squad': flip_squad[squad]})
for player in posts:
#check if loanee player comes from owner's squad
temp = ''
if player['info']['current_team']['on_loan']:
if squad not in player['info']['current_team']['previous_team']:
continue
else:
p_num_dict['fuori in prestito'] +=1
temp = '**'
p_num_dict['a contratto'] +=1
name_url = player['name']
name_url = name_url.replace(' ','-')
name_url = name_url.replace('.','')
dag = ''
cost_eff = player['info']['contract']['cost']
if player['info']['personal_info']['team_real'] is None:
stats_link = ''
dag = '\u2020'
else:
stats_link = 'https://www.fantacalcio.it/squadre/'+player['info']['personal_info']['team_real']+'/'+name_url+'/'+str(player['_id'])
age = int(np.floor((datetime.today()-datetime.strptime(player['info']['personal_info']['birthdate'], "%d/%m/%Y")).days/365.4))
players.append({'role': player['info']['personal_info']['FC_role']
, 'name': player['name']
, 'age': age
, 'quotation': player['info']['stats']['Qt_A']
, 'quotation_initial': player['info']['contract']['quotation_initial']
, 'difference': int(player['info']['stats']['Qt_A']) - int(player['info']['current_team']['quotation_initial'])
, 'loan': temp+dag,
'link': stats_link,
'owner':owner,
'cost':cost_eff,
'complete_db': player})
value_init += int(player['info']['contract']['quotation_initial'])
value_now += int(player['info']['stats']['Qt_A'])
mean_age += int(age)
tot_cost += int(player['info']['contract']['cost'])
posts = collection.find({'info.current_team.owner': owner,'info.current_team.on_loan': True,'info.current_team.squad': squad})
for player in posts:
p_num_dict['dentro in prestito'] +=1
temp = '*'
name_url = player['name']
name_url = name_url.replace(' ','-')
name_url = name_url.replace('.','')
dag = ''
cost_eff = player['info']['current_team']['loan_info']['cost']
if player['info']['personal_info']['team_real'] is None:
stats_link = ''
dag = '\u2020'
else:
stats_link = 'https://www.fantacalcio.it/squadre/'+player['info']['personal_info']['team_real']+'/'+name_url+'/'+str(player['_id'])
age = int(np.floor((datetime.today()-datetime.strptime(player['info']['personal_info']['birthdate'], "%d/%m/%Y")).days/365.4))
players.append({'role': player['info']['personal_info']['FC_role']
, 'name': player['name']
, 'age': age
, 'quotation': player['info']['stats']['Qt_A']
, 'quotation_initial': player['info']['current_team']['quotation_initial']
, 'difference': int(player['info']['stats']['Qt_A']) - int(player['info']['current_team']['quotation_initial'])
, 'loan': temp+dag,
'link': stats_link,
'owner':owner,
'cost':cost_eff,
'complete_db': player})
value_init += int(player['info']['current_team']['quotation_initial'])
value_now += int(player['info']['stats']['Qt_A'])
mean_age += int(age)
tot_cost += int(player['info']['contract']['cost'])
mean_age = mean_age/len(players)
return players, value_init, value_now, round(mean_age, 1), man_team_name(owner), tot_cost, p_num_dict
# -
@anvil.server.callable
def rose_funct_all(owner):
"""
It uses the specific function rose_funct(owner, squad) to return information about the main and primavera
squad of a given owner.
Used by Form:
- Squadra.Rosa
"""
return rose_funct(owner, 'main'), rose_funct(owner, 'primavera')
def count_prizes(palmares):
"""
This is a support function used to count the prizes of each kind. It is given the palmares as it is in the
managers database on mongodb and it returns the overall number of trophies of each kind in a dictionary
"""
sc=ch=cop=sup=tot=ig=pv=cf=po=ca=0
for prize in palmares:
if prize['Type'] == 'Coppa di Lega':
cop+=1
elif prize['Type'] == 'Scudetto':
sc+=1
elif prize['Type'] == 'Champions':
ch+=1
elif prize['Type'] == 'Supercoppa':
sup+=1
elif prize['Type'] == 'Porta Violata':
pv +=1
elif prize['Type'] == 'Cartellino Facile':
cf +=1
elif prize['Type'] == 'Panchina D\'Oro':
po +=1
elif prize['Type'] == 'Caduti':
ca +=1
ig=pv+cf+po+ca
tot=cop+sc+ch+sup
return {'tot': tot,'sc': sc, 'ch': ch, 'cop': cop, 'sup': sup, 'tot_ig': ig,'pv':pv, 'cf':cf, 'po':po, 'ca':ca}
# +
@anvil.server.callable
def man_data_tot(owner):
"""
It returns a string with team_name and a dictionary with the details of a single owner.
Specifically it gives info about budgets, historic wins,
historic fines, current players overall cost (owned players, NOT received on loan). It also uses the function
count_prizes to extract the total number of each trophy which was won in the history of the league. This
function is called by several other functions in the server, as well directly by some forms.
Used by Forms:
- Squadra
"""
owner = owner.lower()
dic = collection_man.find_one({'owner': owner})
team_name = dic['team_name']
dic_out = {}
dic_out['budget'] = dic['budget']
dic_out['tot_wins'] = dic['total_wins']
if len(dic['fines']):
dic_out['tot_fines'] = pd.DataFrame(dic['fines']).Fine_eur.sum()
else:
dic_out['tot_fines'] = 0
dic_out['tot_value'] = 0
dic_out['tot_cost'] = 0
for pl in collection.find({'info.contract.owner': owner}):
dic_out['tot_value'] += int(pl['info']['stats']['Qt_A'])
dic_out['tot_cost'] += int(pl['info']['contract']['cost'])
dic_out['prizes'] = count_prizes(dic['palmares'])
#card = anvil.media.from_file('Logos/'+owner+dic_res[res]+'.png','image/png')
return dic_out, team_name
# -
@anvil.server.callable
def man_data_all():
"""
It returns two dictionaries, with the number of trophies of each possible kind won by each owner in the history
of the league. First dictionary if for league trophies, second id for ignobels.
It uses the other function man_data_tot('owner') to fetch the individual owner's prize history.
Used in the Forms:
- Albo
"""
dic_trophies = []
dic_ig = []
for owner in Teams.keys():
dic_in, team_name = man_data_tot(owner)
dic = {'team_logo_rep':owner,
'team_name': team_name,
'sc': dic_in['prizes']['sc'],
'ch': dic_in['prizes']['ch'],
'cop': dic_in['prizes']['cop'],
'sup': dic_in['prizes']['sup'],
'tot':dic_in['prizes']['tot']
}
dic_2 ={'team_logo_rep':owner,
'team_name': team_name,
'pv': dic_in['prizes']['pv'],
'cf': dic_in['prizes']['cf'],
'po': dic_in['prizes']['po'],
'ca': dic_in['prizes']['ca'],
'tot':dic_in['prizes']['tot_ig']
}
dic_trophies.append(dic)
dic_ig.append(dic_2)
return dic_trophies, dic_ig
# +
@anvil.server.callable
#cost is yet to fix
def finance_managers_data(SC = 360, LTL = 460): #SC and LTL will be determined from previous seasons, saved on mongoDB
"""
This returns two list of dictionaries, used to fill repeating panels. It uses salary cap (SC) and luxury tax limit (LTL) as inputs. The dictionaries contain financial
information of all the owners on the database. The reason for the two lists is because of the form that calls this
function. It calls the function general_standing() to get the general standing, and inverts it to get the
draft order, and the functino man_data_tot() for specific manager's details.
Used by Forms:
- Finanze
"""
table_filler_1 = []
table_filler_2 = []
dict_standing = general_standing()
for owner in Teams.keys():
dict_out_1 = {}
dict_out_2 = {}
dic, team_name = man_data_tot(owner)
_, value_init_main, value_now_main, mean_age_main ,_, tot_cost_main,_ = rose_funct(owner, 'main')
_, value_init_prima, value_now_prima, mean_age_prima ,_, tot_cost_main,_ = rose_funct(owner, 'primavera')
dict_out_1['team'] = owner
dict_out_2['team'] = owner
dict_out_1['extra_budget'] = max(0, 100 + SC - int(value_now_main))
dict_out_1['luxury_tax'] = max(0, int(value_now_main) - LTL)
dict_out_1['budget'] = dic['budget']
dict_out_1['draft_pick'] = 9 - int(dict_standing[owner])
dict_out_2['trophies'] = dic['prizes']['tot']
dict_out_2['trophies_ig'] = dic['prizes']['tot_ig']
dict_out_2['tot_wins'] = dic['tot_wins']
dict_out_2['tot_fines'] = dic['tot_fines']
table_filler_1.append(dict_out_1)
table_filler_2.append(dict_out_2)
return table_filler_1, table_filler_2
# +
@anvil.server.callable
#cost is yet to fix
def finance_one_manager_data(owner, SC = 360, LTL = 460): #SC and LTL will be determined from previous seasons, saved on mongoDB
"""
Uses owner, SC and LTL as inputs to return financial information about a given owner. It returns a dictionary with all
the specific information.
Used by Forms:
- Squadra.Finanze
"""
table_filler_1 = []
dict_standing = general_standing()
owner = owner.lower()
dic = collection_man.find_one({'owner': owner})
dic_2, team_name = man_data_tot(owner)
_, value_init_main, value_now_main, mean_age_main ,_, tot_cost_main,_= rose_funct(owner, 'main')
_, value_init_prima, value_now_prima, mean_age_prima ,_, tot_cost_prima,_= rose_funct(owner, 'primavera')
dic['extra_budget'] = max(0, 100 + SC - int(value_now_main))
dic['luxury_tax'] = max(0, int(value_now_main) - LTL)
dic['draft_pick'] = 9 - int(dict_standing[owner])
dic['tot_fines'] = dic_2['tot_fines']
dic['main_value'] = value_now_main
dic['main_cost'] = value_now_main
dic['prima_cost'] = value_now_main
return dic
# +
@anvil.server.callable
#cost is yet to fix
def rose_managers_data(SC = 360, LTL = 460):
"""
It takes salary cap (SC) and luxury tax limit (LTL) as input. It returns a list of dicionaries each with manager's
information about the lineup. It also returns a dictionary with the league medians.
Used in Forms:
- Rose
"""
table_filler = []
median_dict ={'Val_M':[],
'Val_P':[],
'Cost_M':[],
'Cost_P':[],
'Age_M':[],
'Age_P':[]}
for owner in Teams.keys():
dict_out = {}
dic, team_name = man_data_tot(owner)
_, value_init_main, value_now_main, mean_age_main ,_,tot_cost_main, main_pl_num = rose_funct(owner, 'main')
_, value_init_prima, value_now_prima, mean_age_prima ,_,tot_cost_prima, prima_pl_num= rose_funct(owner, 'primavera')
dict_out['team'] = owner
dict_out['extra_budget'] = max(0, 100 + SC - int(value_now_main))
dict_out['luxury_tax'] = max(0, int(value_now_main) - LTL)
dict_out['main_value'] = float(value_now_main)
dict_out['prima_value'] = float(value_now_prima)
dict_out['main_cost'] = float(tot_cost_main)
dict_out['prima_cost'] = float(tot_cost_prima)
dict_out['mean_age_main'] = float(mean_age_main)
dict_out['mean_age_prima'] = float(mean_age_prima)
#dict_out['draft_pick'] = 9 - int(dict_standing[owner])
dict_out['trophies'] = int(dic['prizes']['tot'])
dict_out['trophies_ig'] = int(dic['prizes']['tot_ig'])
dict_out['tot_wins'] = int(dic['tot_wins'])
dict_out['tot_fines'] = int(dic['tot_fines'])
table_filler.append(dict_out)
median_dict['Val_M'].append(float(value_now_main))
median_dict['Val_P'].append(float(value_now_prima))
median_dict['Age_M'].append(float(mean_age_main))
median_dict['Age_P'].append(float(mean_age_prima))
median_dict['Cost_M'].append(float(tot_cost_main))
median_dict['Cost_P'].append(float(tot_cost_prima))
median_dict['Val_M'] = float(np.median(median_dict['Val_M']))
median_dict['Val_P'] = float(np.median(median_dict['Val_P']))
median_dict['Age_M'] = float(np.median(median_dict['Age_M']))
median_dict['Age_P'] = float(np.median(median_dict['Age_P']))
median_dict['Cost_M'] = float(np.median(median_dict['Cost_M']))
median_dict['Cost_P'] = float(np.median(median_dict['Cost_P']))
return table_filler, median_dict
# -
@anvil.server.callable
def load_plot_C_all(regen = False):
"""
It returns the current matchday, as well as two plots for Cfactor uploaded from the local folder 'Plots'.
If regen = True, the function generate_plots() is called, that recreates these plots and saves them in the
same local folred, before uploading.
Used in Forms:
- Fortuna
"""
if regen:
generate_plots()
card_hist = anvil.media.from_file('Plots/C_fact_'+ 'Historic' +'.png','image/png')
card_tot = anvil.media.from_file('Plots/C_fact_'+ 'Total' +'.png','image/png')
return giornate, card_tot, card_hist
@anvil.server.callable
def load_IGNOBEL_db(plot):
#tot = premio_plot(Results, giornate, Teams, Logos, plot)
list_IG = sorted(dict_out[plot], key = lambda i: i['points'], reverse = True)
i=1
prize = 7 #to be determined via mongodb
for el in list_IG:
el['position'] = i
el['team'] = el['team'].capitalize()
el['prize'] = prize
i+=1
card = anvil.media.from_file('Plots/plot_'+ plot +'.png','image/png')
#out_plot = load_plot_IG(plot)
montepremi = str(4*prize)
return card, list_IG, giornate, montepremi
def tot_fines():
"""
Using the info extracted from man_data_owner(), it returns an updated dictionary with the total fines
accumulated by each owner.
It is called by other functions.
"""
fines = 0
for owner in Teams.keys():
dic_man, _ = man_data_tot(owner)
fines += dic_man['tot_fines']
return fines
@anvil.server.callable
def fetch_standing(standing):
"""
For a given 'standing' parameter (points, goal etc), it returns the standing based on that parameter as a
list of dictionaries.
It is called by other functions.
"""
fines = tot_fines()
posts = []
for owner in Teams.keys():
dic = Results[owner]
points = dic[standing].sum() #without the sum, dic[standing] contains the progress, just in case one needs it for plotting live
points_last = dic[standing][len(dic)]
if standing in ['infortunati', 'cartellini', 'goal_subiti_por','bonus_panchina']:
prize = 4 + fines/4
else:
prize = None
team = owner.capitalize()
dic_0 = {
'points':points,
'points_last':points_last,
'prize':prize,
'team':team,
'team_name': Teams[owner][0]
}
posts.append(dic_0)
posts = sorted(posts, key = lambda i: i['points'], reverse = True)
i=1
for el in posts:
el['position'] = i
i+=1
return posts, prize
<EMAIL>
def fetch_ALL_standings():
"""
This returns the complete set of all the standings in the league, namely IGnobels and general/points. It is used
by other functions
"""
dic_stand = {
'Caduti': 'infortunati',
'Cartellino Facile': 'cartellini',
'Porta Violata': 'goal_subiti_por',
'Catenaccio': 'mod_difesa',
'Panchina Oro': 'bonus_panchina',
'Generale': 'pti',
'Avulsa': 'punti_fatti'
}
dic_out = {}
for key, arg in dic_stand.items():
posts, temp_prize = fetch_standing(arg)
if temp_prize is not None:
prize = temp_prize
dic_out[key] = posts
return dic_out, prize
@anvil.server.callable
def fetch_Points_standings():
"""
This returns the points-based standings in a two-keys dictionary.
Used by Forms:
- Stats
"""
dic_stand = {
'Generale': 'pti',
'Avulsa': 'punti_fatti'
}
dic_out = {}
for key, arg in dic_stand.items():
posts, temp_prize = fetch_standing(arg)
dic_out[key] = posts
return dic_out
<EMAIL>
def fetch_IG_standings():
"""
This returns the complete set of the IGnobel standings. It is called by other functions.
"""
dic_stand = {
'Caduti': 'infortunati',
'Cartellino Facile': 'cartellini',
'Porta Violata': 'goal_subiti_por',
'Panchina Oro': 'bonus_panchina'
}
dic_out = {}
for key, arg in dic_stand.items():
posts, temp_prize = fetch_standing(arg)
if temp_prize is not None:
prize = temp_prize
dic_out[key] = posts
return dic_out, prize
def general_standing():
"""
It gives the general standing based on the current matchday. Note that it depends on the parameters that are
imported at the beginning of the notebook, specifically Results, hence in order to refresh it needs to be run
after Results is created from the utilities script.
This is called by other functions.
"""
posts,_ = fetch_standing('pti')
dict_out={}
for dic in posts:
dict_out[dic['team'].lower()] = dic['position']
return dict_out
@anvil.server.callable
def load_IGNOBEL_db_all():
"""
This returns a dictionary with the 4 lists corresponting to the standings for the ignobel prizes.
also the current matchday and the overall money prize for the competitions. It uses the function
fetch_IG_standings() to fetch the stats.
Used by Forms:
- Ignobel
"""
list_IG, prize = fetch_IG_standings()
montepremi_ig = 4*prize
return list_IG, giornate, montepremi_ig
@anvil.server.callable
def load_IGNOBEL_plots():
"""
Returns dictionaries of the ignobel plots in anvil media format. It calls the function generate_plots() to
generate the plots and save them in the local folder Plot. Then it uploads them from there.
Used by Forms:
- Ignobel
"""
plot_dict = {}
generate_plots()
time.sleep(3)
for plot in ['Porta Violata', 'Cartellino Facile', 'Panchina Oro', 'Caduti']:
plot_dict[plot] = anvil.media.from_file('Plots/plot_'+ plot +'.png','image/png')
return plot_dict
@anvil.server.callable
def all_players():
"""
Returns the complete list of the names of all the players in the database.
Used by the forms:
- Admin_Transfers
- Giocatori
- Trasferimenti
"""
down = list(collection.find({}))
name_list = list(pd.DataFrame(down).name)
return name_list
def transfer_list(name):
"""
Fetches all the transfers involving the player's name.
"""
posts = list(collection_tr.find({'name':name}))
return posts
@anvil.server.callable
def full_pl_info(name):
"""
Returns the player's info, including the full dictionary, the URL media from the link from fc.it, the link to the
statistics from FC and the list of all the transfers in which the player was involved.
To fetch the transfers the function transfer_list(name) is used.
Used by Forms:
- Giocatori_lower
"""
dic = collection.find_one({'name': name})
name_url = dic['name']
name_url = name_url.replace(' ','-')
name_url = name_url.replace('.','')
age = date.today().year - int(dic['info']['personal_info']['birthdate'][6:10])
dic['info']['personal_info']['age']= ' ('+ str(age)+')'
card = URLMedia('https://content.fantacalcio.it/web/campioncini/card/'+name_url+'.jpg')
if dic['info']['personal_info']['team_real'] is None:
stats_link = ''
else:
stats_link = 'https://www.fantacalcio.it/squadre/'+dic['info']['personal_info']['team_real']+'/'+name_url+'/'+str(dic['_id'])
return dic, card, stats_link, transfer_list(name)
# +
@anvil.server.callable
def save_transfer_mongo(dic, player = False, loan_info = False):
"""
It is used to make modifications directly to the databases of transfers and players. It takes all the parameters that are
inserted from the app and directly creates/overwrites the entries in the database.
Used by the Forms:
- Admin_Transfers
"""
now = datetime.today()
Id = now.strftime("%Y%m%d%H%M%S")
res = dic
res['_id'] = Id
if loan_info:
date_now = date.today()
m, y = date_now.month, date_now.year
if m > 7:
y = y + 1
loan_info['expire_date'] = str(y)+'/07/31'
res['loan_info'] = loan_info
collection_tr.insert_one(res)
if player:
if dic['operation'] in ['Asta', 'Draft', 'Acquisto', 'Scambio', 'Algoritmo', 'Svincolo']:
cost_exch = 0
if dic['operation'] == 'Scambio':
dic_exch = collection.find_one({'name': dic['exchange_player']})
cost_exch = int(dic_exch['info']['contract']['cost'])
collection.update_one({'name':dic['name']},{'$set':{'info.contract.start_date':dic['date']}})
collection.update_one({'name':dic['name']},{'$set':{'info.contract.cost':int(dic['cost']) + cost_exch}})
collection.update_one({'name':dic['name']},{'$set':{'info.contract.acquisition_mode':dic['operation']}})
collection.update_one({'name':dic['name']},{'$set':{'info.contract.previous_owner':dic['previous_owner']}})
collection.update_one({'name':dic['name']},{'$set':{'info.contract.quotation_initial':int(dic['quotation_to_date'])}})
if loan_info:
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.on_loan':True}})
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.loan_info':loan_info}})
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.start_date':dic['date']}})
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.owner':dic['new_owner']}})
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.squad':dic['squad']}})
if dic['previous_owner'] is None:
previous_team = None
else:
previous_team = dic['previous_owner']+', '+dic['previous_squad']
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.previous_team':previous_team}})
collection.update_one({'name':dic['name']},{'$set':{'info.current_team.quotation_initial':int(dic['quotation_to_date'])}})
dic_tr = collection_tr.find_one({'_id':Id})
dic_pl = 'Non Aggiornato'
if player:
dic_pl = collection.find_one({'name':dic['name']})
return str(dic_tr), str(dic_pl)
# -
@anvil.server.callable
def all_flags_list():
"""
It returns the flags of all the nationalities of the players on mongodb in anvil media format. It would give error if
some file is missing, in that case it needs to be downloaded and saved in the folder 'Bandiere' in png format.
Used in the Forms:
- Admin_area
"""
posts = collection.find({})
all_pl = []
for pl in posts:
if 'personal_info' in pl['info'].keys():
all_pl.append(pl['info']['personal_info']['nation'])
nations = []
for nat in all_pl:
if ',' in nat:
nat_2 = nat.split(', ')
nations.append(nat_2[0].lower())
nations.append(nat_2[1].lower())
else:
nations.append(nat.lower())
dict_flag = {}#[{'res':'high'}]
for nat in nations:
dict_flag[nat] = anvil.media.from_file('../Bandiere/'+ nat +'.png','image/png')
return dict_flag
@anvil.server.callable
def all_team_logos_list():
"""
It returns the logos of all the real teams of the players on mongodb in anvil media format. It would give error if
some file is missing, in that case it needs to be downloaded and saved in the folder 'Scudetti' in png format.
Used in the Forms:
- Admin_area
"""
posts = collection.find({})
all_pl = []
for pl in posts:
if 'personal_info' in pl['info'].keys():
if pl['info']['personal_info']['team_real'] is None:
all_pl.append('svincolato')
else:
all_pl.append(pl['info']['personal_info']['team_real'])
teams = []
for team in all_pl:
teams.append(team.lower())
dict_logos = {}#[{'res':'high'}]
for team in teams:
dict_logos[team] = anvil.media.from_file('../Scudetti/'+ team +'.png','image/png')
return dict_logos
@anvil.server.callable
def fetch_transfers(dic):
"""
This returns a list of dictionaries straight from the transfer database that match the filters included
in the dictionary dic used as input.
Used by Forms:
- Trasferimenti
"""
filt = {}
from_owners = []
for el in dic['from_list']:
if el['checked']:
from_owners.append(el['owner'])
filt['previous_owner'] = {'$in': from_owners}
to_owners = []
for el in dic['to_list']:
if el['checked']:
to_owners.append(el['owner'])
filt['new_owner'] = {'$in': to_owners}
operations = []
for el in dic['operations']:
if el['checked']:
operations.append(el['operation'])
filt['operation'] = {'$in': operations}
if dic['from_squad']['main']:
filt['previous_squad'] = 'main'
elif dic['from_squad']['primavera']:
filt['previous_squad'] = 'primavera'
if dic['to_squad']['main']:
filt['squad'] = 'main'
elif dic['to_squad']['primavera']:
filt['squad'] = 'primavera'
if dic['name'] is not None:
filt['name'] = dic['name']
Min = -100
Max = 1000
if dic['cost']['min'] is not None:
Min = dic['cost']['min']
if dic['cost']['max'] is not None:
Max = dic['cost']['max']
filt['cost'] = {'$gte':int(Min), '$lte':int(Max)}
date_Min = 0
date_Max = 10**10
if dic['dates']['from'] is not None:
date_Min = int(dic['dates']['from'].strftime('%Y%m%d'))
if dic['dates']['to'] is not None:
date_Max = int(dic['dates']['to'].strftime('%Y%m%d'))
filt['date_num'] = {'$gte':int(date_Min), '$lte':int(date_Max)}
#dic['dates']
print(filt)
output = list(collection_tr.find(filt))
return output
@anvil.server.callable
def fetch_teams_real(checked = True):
"""
It returns a list of dictionaries with all the real teams' names in the database.
Used by Forms:
- Giocatori
"""
posts = collection.find()
teams_dict = []
teams = []
for pl in posts:
team_real = filter_team = pl['info']['personal_info']['team_real']
if team_real is None:
continue #team_real = 'Non in Serie A'
if team_real not in teams:
teams_dict.append({'filter_team': filter_team,'team_name': team_real, 'checked': checked})
teams.append(team_real)
return teams_dict + [{'filter_team': None,'team_name': 'Non in Serie A', 'checked': True}]
@anvil.server.callable
def fetch_players_database(dic):
"""
Given an input from the appropriate filters as dictionary, it returns the list of dictionaries of the corresponding
players from mongodb.
Used in Forms:
- Giocatori
"""
filt = {}
owners = []
for el in dic['owners']:
if el['checked']:
owners.append(el['owner'])
filt['info.contract.owner'] = {'$in': owners}
if dic['squad']['main'] ^ dic['squad']['primavera']:
if dic['squad']['main']:
filt['info.current_team.squad'] = 'main'
elif dic['squad']['primavera']:
filt['info.current_team.squad'] = 'primavera'
if dic['squad']['loan']:
filt['info.current_team.on_loan'] = True
roles = []
for el in dic['roles']:
if el['checked']:
roles.append(el['role'])
filt['info.personal_info.FC_role'] = {'$in': roles}
Min_q = dic['quot']['min']
Max_q = dic['quot']['max']
filt['info.stats.Qt_A'] = {'$gte':int(Min_q), '$lte':int(Max_q)}
Min_birth = int(date.today().strftime('%Y%m%d'))-dic['age']['min']*10**4
Max_birth = int(date.today().strftime('%Y%m%d'))-dic['age']['max']*10**4
filt['info.personal_info.birthdate_num'] = {'$lte':int(Min_birth), '$gte':int(Max_birth)}#inversed of course
teams_real = []
for el in dic['teams_real']:
if el['checked']:
teams_real.append(el['filter_team'])
filt['info.personal_info.team_real'] = {'$in': teams_real}
posts = list(collection.find(filt))
return posts
| Database/App_server_pietro-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.015432, "end_time": "2021-09-09T04:01:59.577367", "exception": false, "start_time": "2021-09-09T04:01:59.561935", "status": "completed"} tags=[]
# # 1. Parameters
# + papermill={"duration": 0.027908, "end_time": "2021-09-09T04:01:59.614200", "exception": false, "start_time": "2021-09-09T04:01:59.586292", "status": "completed"} tags=["parameters"]
# Defaults
cases_dir = 'cases/unset'
reference_file = 'references/NC_045512.gbk.gz'
input_files_all = 'input/input-files.tsv'
iterations = 3
mincov = 10
ncores = 32
number_samples = 10
build_tree = False
# + papermill={"duration": 0.018136, "end_time": "2021-09-09T04:01:59.646264", "exception": false, "start_time": "2021-09-09T04:01:59.628128", "status": "completed"} tags=["injected-parameters"]
# Parameters
cases_dir = "cases/case-20000"
iterations = 3
number_samples = 20000
build_tree = False
# + papermill={"duration": 2.50566, "end_time": "2021-09-09T04:02:02.164495", "exception": false, "start_time": "2021-09-09T04:01:59.658835", "status": "completed"} tags=[]
from pathlib import Path
from shutil import rmtree
from os import makedirs
import imp
fp, pathname, description = imp.find_module('gdi_benchmark', ['../../lib'])
gdi_benchmark = imp.load_module('gdi_benchmark', fp, pathname, description)
cases_dir_path = Path(cases_dir)
if cases_dir_path.exists():
rmtree(cases_dir_path)
if not cases_dir_path.exists():
makedirs(cases_dir_path)
input_files_all = Path(input_files_all)
reference_file = Path(reference_file)
case_name = str(cases_dir_path.name)
reference_name = reference_file.name.split('.')[0]
cases_input = cases_dir_path / 'input-files-case.tsv'
index_path = cases_dir_path / 'index'
benchmark_path = cases_dir_path / 'index-info.tsv'
output_tree = cases_dir_path / 'tree.tre'
# + [markdown] papermill={"duration": 0.009054, "end_time": "2021-09-09T04:02:02.188013", "exception": false, "start_time": "2021-09-09T04:02:02.178959", "status": "completed"} tags=[]
# # 2. Create subset input
# + papermill={"duration": 0.363399, "end_time": "2021-09-09T04:02:02.558000", "exception": false, "start_time": "2021-09-09T04:02:02.194601", "status": "completed"} tags=[]
import pandas as pd
all_input_df = pd.read_csv(input_files_all, sep='\t')
all_input_total = len(all_input_df)
subset_input_df = all_input_df.head(number_samples)
subset_input_total = len(subset_input_df)
subset_input_df.to_csv(cases_input, sep='\t', index=False)
print(f'Wrote {subset_input_total}/{all_input_total} samples to {cases_input}')
# + [markdown] papermill={"duration": 0.007979, "end_time": "2021-09-09T04:02:02.579449", "exception": false, "start_time": "2021-09-09T04:02:02.571470", "status": "completed"} tags=[]
# # 2. Index genomes
# + papermill={"duration": 2.958601, "end_time": "2021-09-09T04:02:05.544450", "exception": false, "start_time": "2021-09-09T04:02:02.585849", "status": "completed"} tags=[]
# !gdi --version
# + [markdown] papermill={"duration": 0.00916, "end_time": "2021-09-09T04:02:05.569079", "exception": false, "start_time": "2021-09-09T04:02:05.559919", "status": "completed"} tags=[]
# ## 2.1. Index reads
# + papermill={"duration": 27594.924, "end_time": "2021-09-09T11:42:00.500317", "exception": false, "start_time": "2021-09-09T04:02:05.576317", "status": "completed"} tags=[]
results_handler = gdi_benchmark.BenchmarkResultsHandler(name=case_name)
benchmarker = gdi_benchmark.IndexBenchmarker(benchmark_results_handler=results_handler,
index_path=index_path, input_files_file=cases_input,
reference_file=reference_file, mincov=mincov,
build_tree=build_tree,
ncores=ncores)
benchmark_df = benchmarker.benchmark(iterations=iterations)
# + papermill={"duration": 0.055867, "end_time": "2021-09-09T11:42:00.577739", "exception": false, "start_time": "2021-09-09T11:42:00.521872", "status": "completed"} tags=[]
benchmark_df
# + papermill={"duration": 0.023262, "end_time": "2021-09-09T11:42:00.623071", "exception": false, "start_time": "2021-09-09T11:42:00.599809", "status": "completed"} tags=[]
benchmark_df.to_csv(benchmark_path, sep='\t', index=False)
# + [markdown] papermill={"duration": 0.011061, "end_time": "2021-09-09T11:42:00.647973", "exception": false, "start_time": "2021-09-09T11:42:00.636912", "status": "completed"} tags=[]
# # 3. Export trees
# + papermill={"duration": 0.021258, "end_time": "2021-09-09T11:42:00.679313", "exception": false, "start_time": "2021-09-09T11:42:00.658055", "status": "completed"} tags=[]
if build_tree:
# !gdi --project-dir {index_path} export tree {reference_name} > {output_tree}
print(f'Wrote tree to {output_tree}')
else:
print(f'build_tree={build_tree} so no tree to export')
| evaluations/sars-cov-2/3-index-genomes.case-20000.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Try It Yourself
#
# With all you've learned, you can start writing much more interesting programs. See if you can solve the problems below.
#
# As always, run the setup code below before working on the questions.
from learntools.core import binder; binder.bind(globals())
from learntools.python.ex5 import *
print('Setup complete.')
# # Exercises
# ## 1.
#
# Have you ever felt debugging involved a bit of luck? The following program has a bug. Try to identify the bug and fix it.
def has_lucky_number(nums):
"""Return whether the given list of numbers is lucky. A lucky list contains
at least one number divisible by 7.
"""
for num in nums:
if num % 7 == 0:
return True
else:
return False
# Try to identify the bug and fix it in the cell below:
# +
def has_lucky_number(nums):
"""Return whether the given list of numbers is lucky. A lucky list contains
at least one number divisible by 7.
"""
for num in nums:
if num % 7 == 0:
return True
else:
return False
# Check your answer
q1.check()
# -
#_COMMENT_IF(PROD)_
q1.hint()
#_COMMENT_IF(PROD)_
q1.solution()
# ## 2.
#
# ### a.
# Look at the Python expression below. What do you think we'll get when we run it? When you've made your prediction, uncomment the code and run the cell to see if you were right.
# +
#[1, 2, 3, 4] > 2
# -
# ### b
# R and Python have some libraries (like numpy and pandas) compare each element of the list to 2 (i.e. do an 'element-wise' comparison) and give us a list of booleans like `[False, False, True, True]`.
#
# Implement a function that reproduces this behaviour, returning a list of booleans corresponding to whether the corresponding element is greater than n.
#
# +
def elementwise_greater_than(L, thresh):
"""Return a list with the same length as L, where the value at index i is
True if L[i] is greater than thresh, and False otherwise.
>>> elementwise_greater_than([1, 2, 3, 4], 2)
[False, False, True, True]
"""
pass
# Check your answer
q2.check()
# -
#_COMMENT_IF(PROD)_
q2.solution()
# ## 3.
#
# Complete the body of the function below according to its docstring.
# +
def menu_is_boring(meals):
"""Given a list of meals served over some period of time, return True if the
same meal has ever been served two days in a row, and False otherwise.
"""
pass
# Check your answer
q3.check()
# -
#_COMMENT_IF(PROD)_
q3.hint()
#_COMMENT_IF(PROD)_
q3.solution()
# ## 4. <span title="A bit spicy" style="color: darkgreen ">🌶️</span>
#
# Next to the Blackjack table, the Python Challenge Casino has a slot machine. You can get a result from the slot machine by calling `play_slot_machine()`. The number it returns is your winnings in dollars. Usually it returns 0. But sometimes you'll get lucky and get a big payday. Try running it below:
play_slot_machine()
# By the way, did we mention that each play costs $1? Don't worry, we'll send you the bill later.
#
# On average, how much money can you expect to gain (or lose) every time you play the machine? The casino keeps it a secret, but you can estimate the average value of each pull using a technique called the **Monte Carlo method**. To estimate the average outcome, we simulate the scenario many times, and return the average result.
#
# Complete the following function to calculate the average value per play of the slot machine.
def estimate_average_slot_payout(n_runs):
"""Run the slot machine n_runs times and return the average net profit per run.
Example calls (note that return value is nondeterministic!):
>>> estimate_average_slot_payout(1)
-1
>>> estimate_average_slot_payout(1)
0.5
"""
pass
# When you think you know the expected value per spin, run the code cell below to view the solution and get credit for answering the question.
# Check your answer (Run this code cell to receive credit!)
q4.solution()
# # Keep Going
#
# Many programmers report that dictionaries are their favorite data structure. You'll get to **[learn about them](#$NEXT_NOTEBOOK_URL$)** (as well as strings) in the next lesson.
| notebooks/python/raw/ex_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.3
# language: julia
# name: julia-1.6
# ---
# # Handling missing values
# A singelton type Missing allows us to deal with missing values.
using DataFrames
missing, typeof(missing)
x = [1, 2, missing, 3]
ismissing(1), ismissing(missing), ismissing(x), ismissing.(x)
# We can extract the type combined with Missing from a Union via nonmissingtype
eltype(x), nonmissingtype(eltype(x))
missing == missing, missing != missing, missing < missing
1 == missing, 1 != missing, 1 < missing
# isequal, isless, and === produce results of type Bool. Notice that missing is considered greater than any numeric value.
isequal(missing, missing), missing === missing, isequal(1, missing), isless(1, missing)
map(x -> x(missing), [sin, cos, zero, sqrt]) # part 1
map(x -> x(missing, 1), [+, - , *, /, div]) # part 2
using Statistics # needed for mean
map(x -> x([1,2,missing]), [minimum, maximum, extrema, mean, float]) # part 3
collect(skipmissing([1, missing, 2, missing]))
# Here we use replace to create a new array that replaces all missing values with some value (NaN in this case).
replace([1.0, missing, 2.0, missing], missing=>NaN)
# Another way to do this:
coalesce.([1.0, missing, 2.0, missing], NaN)
# +
# You can also use recode from CategoricalArrays.jl if you have a default output value.
using CategoricalArrays
recode([1.0, missing, 2.0, missing], false, missing=>true)
# -
df = DataFrame(a=[1,2,missing], b=["a", "b", missing])
replace!(df.a, missing=>100)
df.b = coalesce.(df.b, 100)
unique([1, missing, 2, missing]), levels([1, missing, 2, missing])
x = [1,2,3]
y = allowmissing(x)
z = disallowmissing(y)
x,y,z
df = allowmissing(DataFrame(ones(2,3), :auto))
df[1,1] = missing
df
disallowmissing(df) # an error is thrown
disallowmissing(df, error=false) # column :x1 is left untouched as it contains missing
x = DataFrame(rand(Int, 2,3), :auto)
println("Before: ", eltype.(eachcol(x)))
allowmissing!(x, 1) # make first column accept missings
allowmissing!(x, :x3) # make :x3 column accept missings
println("After: ", eltype.(eachcol(x)))
x = DataFrame(A=[1, missing, 3, 4], B=["A", "B", missing, "C"])
println("Complete cases:\n", completecases(x))
y = dropmissing(x)
dropmissing!(x)
describe(x)
dropmissing!(x, disallowmissing=false)
| Day 10/missing_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="h2q27gKz1H20"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="TUfAcER1oUS6"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="Gb7qyhNL1yWt"
# # Text classification with TensorFlow Lite Model Maker with TensorFlow 2.0
# + [markdown] colab_type="text" id="Fw5Y7snSuG51"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/lite/tutorials/model_maker_text_classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/tutorials/model_maker_text_classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorflow/blob/master/tensorflow/lite/g3doc/tutorials/model_maker_text_classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/tensorflow/tensorflow/lite/g3doc/tutorials/model_maker_text_classification.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="sr3q-gvm3cI8"
# This notebook has been moved [here](https://www.tensorflow.org/lite/tutorials/model_maker_text_classification).
| tensorflow_examples/lite/model_maker/demo/text_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Using inaFaceAnalyzer API: image analysis quick-start tutorial
# In this tutorial, we use inaFaceAnalyzer with default analysis parameters on image files. We export results to CSV and display intermediate processing steps.
# ## Install inaFaceAnalyzer
# try to import inaFaceAnalyzer and import it from Pypi's
# if it is not available
try:
import inaFaceAnalyzer
except:
# install inaFaceAnalyzer Pypi's distribution
# !pip install inaFaceAnalyzer
# ## Download and display a sample images
# +
from IPython.display import Image, display
from tensorflow.keras.utils import get_file
urlprefix = 'https://raw.githubusercontent.com/ina-foss/inaFaceAnalyzer/master/media/'
# download Donald Knuth remote image
donald = get_file('donald.jpg', urlprefix + 'dknuth.jpg')
# print local path to image
print(donald)
# display image in a notebook
display(Image(donald))
# download a familily picture
family = get_file('family.jpg', urlprefix + '1328360239_e56cc26d26_o.jpg')
# print local path to image
print(family)
# display image full path
display(Image(family))
# -
# ## Analyse a single image with default parameters
# Single image analysis requires to use the ImageAnalyzer engine. ImageAnalyzer constructor may require several seconds and should be done a single time when processing several image files. ImageAnalyzer constructor may accept several parameters that will be covered in more advanced tutorials.
# Import ImageAnalyzer class
from inaFaceAnalyzer.inaFaceAnalyzer import ImageAnalyzer
# create an image analyzer engine instance
ia = ImageAnalyzer()
# Process an image
df = ia(donald)
# Analysis results are returned as pandas DataFrames
# see https://pandas.pydata.org/docs/
# Results contain one line per detected faces and several columns :
#
# frame: the full path to the imaged used
# bbox: (left, top, right, bottom) the bounding box of the face in the image frame
# detect_conf: the face detection confidence estimate (dependent on the face detection method used)
# sex_decfunc: raw gender classifier output : positive values are used for men and negative values for women
# sex_label: gender classifer prediction: 'm' for men and 'w' for 'women'
# age_decfunc: raw age regression output based on FairFace age categories.
# 0 for (0-3 years old), 1 for (4-9) years, 2 for (10-19) years, 3 for (20-29) years, etc...
# sex_label : "human-readable" age prediction
#
# For Donald Knuth picture, we found a single face, labelled as 62 years old male
df
# export results to CSV
df.to_csv('./donald.csv', index=False)
# for the family picture, we found 7 male and female faces from 2.4 to 60.3 years old
ia(family)
# ## Displaying results and intermediate processing steps
# Setting named argument `verbose=True` in analysis engine constructor allow to display intermediate processing steps and final results, in a more human-friendly way than CSV or dataframes. This may be usefull for debugging and integrating new components, or having insights on the quality of the results obtained for a given material.
# The information displayed for ImageAnalyzer are :
# * raw image
# * image with incrusted bounding boxes
# * original faces corresponding to detection bounding boxes
# * preprocessed faces (after bounding box scaling and eyes alignment)
# * classification results
#
#
# Intermediate image display is costly and should be avoided in production
# setting named argument verbose=True in ImageAnalyzer constructor allows to
# display intermediate processings. Image display is costly and should be avoided in production.
ia = ImageAnalyzer(verbose=True)
ia(donald)
ia(family)
# ## Faster analyses with image lists
# `inaFaceAnalyzer` back-end provide batches of 32 preprocess faces to face classifiers in order to speed-up computation time. Consequently, faster processing times can be obtained using list of images instead of isolated images.
# process image list with verbose=True in previously construted analyzer
df = ia([donald, family])
# display result
# a single line is used for <NAME>'s picture, and the 7 remaining lines are for the family picture
df
| tutorial_API_notebooks/quick_start-image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Challenge Part 2: Have Customers Narrow Their Travel Searches Based on Temperature and Precipitation
# +
import pandas as pd
import requests
import gmaps
from config import g_key
| Vacation_Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # lab
#
# Laboratory tests that have have been mapped to a standard set of measurements. Unmapped measurements are recorded in the customLab table. The lab table is fairly well populated by hospitals. It is possible some rarely obtained lab measurements are not interfaced into the system and therefore will not be available in the database. Absence of a rare lab measurement, such as serum lidocaine concentrations, would not indicate the lab was not drawn. However, absence of a platelet count would likely indicate the value was not obtained.
# +
# Import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import psycopg2
import getpass
import pdvega
# for configuring connection
from configobj import ConfigObj
import os
# %matplotlib inline
# +
# Create a database connection using settings from config file
config='../db/config.ini'
# connection info
conn_info = dict()
if os.path.isfile(config):
config = ConfigObj(config)
conn_info["sqluser"] = config['username']
conn_info["sqlpass"] = config['password']
conn_info["sqlhost"] = config['host']
conn_info["sqlport"] = config['port']
conn_info["dbname"] = config['dbname']
conn_info["schema_name"] = config['schema_name']
else:
conn_info["sqluser"] = 'postgres'
conn_info["sqlpass"] = ''
conn_info["sqlhost"] = 'localhost'
conn_info["sqlport"] = 5432
conn_info["dbname"] = 'eicu'
conn_info["schema_name"] = 'public,eicu_crd'
# Connect to the eICU database
print('Database: {}'.format(conn_info['dbname']))
print('Username: {}'.format(conn_info["sqluser"]))
if conn_info["sqlpass"] == '':
# try connecting without password, i.e. peer or OS authentication
try:
if (conn_info["sqlhost"] == 'localhost') & (conn_info["sqlport"]=='5432'):
con = psycopg2.connect(dbname=conn_info["dbname"],
user=conn_info["sqluser"])
else:
con = psycopg2.connect(dbname=conn_info["dbname"],
host=conn_info["sqlhost"],
port=conn_info["sqlport"],
user=conn_info["sqluser"])
except:
conn_info["sqlpass"] = getpass.getpass('Password: ')
con = psycopg2.connect(dbname=conn_info["dbname"],
host=conn_info["sqlhost"],
port=conn_info["sqlport"],
user=conn_info["sqluser"],
password=conn_info["sql<PASSWORD>"])
query_schema = 'set search_path to ' + conn_info['schema_name'] + ';'
# -
# ## Examine a single patient
patientunitstayid = 2704494
# +
query = query_schema + """
select *
from lab
where patientunitstayid = {}
order by labresultoffset
""".format(patientunitstayid)
df = pd.read_sql_query(query, con)
df.head()
# +
query = query_schema + """
select *
from patient
where patientunitstayid = {}
""".format(patientunitstayid)
pt = pd.read_sql_query(query, con)
pt[['patientunitstayid', 'apacheadmissiondx', 'hospitaladmitoffset']]
# -
# Immediately we can note the very large negative `labresultoffset`. This likely means we have some lab values pre-ICU. In some cases this will be a lab measured in another hospital location such as the emergency department or hospital floor. In this case, the large value (-99620 minutes, or ~70 days) is surprising, but we can see from the patient table that the patient was admitted to the hospital -99779 minutes before their ICU stay (`hospitaladmitoffset`). This patient was admitted to the ICU with thrombocytopenia (`apacheadmissiondx`), and inspection of the diagnosis table indicates they have a form of cancer, so likely this is a long hospital stay where labs were taken on hospital admission.
# ## Available labs
#
# We can group the lab table to summarize all available labs.
# +
query = query_schema + """
select labname, count(*) as n
from lab
group by labname
order by n desc
""".format(patientunitstayid)
lab = pd.read_sql_query(query, con)
print('{} total vlues for {} distinct labs.'.format(lab['n'].sum(), lab.shape[0]))
print('\nTop 5 labs by frequency:')
lab.head()
# -
# The lab table is a large table with over 39 million observations. The most frequent observation is bedside glucose which accounts for almost 10% of the lab table, followed by potassium and sodium.
# ## Hospitals with data available
# +
query = query_schema + """
with t as
(
select distinct patientunitstayid
from lab
)
select
pt.hospitalid
, count(distinct pt.patientunitstayid) as number_of_patients
, count(distinct t.patientunitstayid) as number_of_patients_with_tbl
from patient pt
left join t
on pt.patientunitstayid = t.patientunitstayid
group by pt.hospitalid
""".format(patientunitstayid)
df = pd.read_sql_query(query, con)
df['data completion'] = df['number_of_patients_with_tbl'] / df['number_of_patients'] * 100.0
df.sort_values('number_of_patients_with_tbl', ascending=False, inplace=True)
df.head(n=10)
# -
df.tail(n=10)
df[['data completion']].vgplot.hist(bins=10,
var_name='Number of hospitals',
value_name='Percent of patients with data')
# Above we can see that very few hospitals are missing lab data. Most of the data at < 90% data completion is driven by a few hospitals with very few patients. The majority of hospitals have 90-100% of patients with data in the lab table (right side of histogram, 0-90% bin).
| notebooks/lab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
import pandas as pd
file = "data/access-export/tblTexts.xlsx"
df = pd.read_excel(file)
# create necessary concept-schemes
ecce_genre,_ = SkosConceptScheme.objects.get_or_create(dc_title="ecce-genre")
ecce_dialect,_ = SkosConceptScheme.objects.get_or_create(dc_title="ecce-dialect")
for i, row in df.iterrows():
temp_genre,_ = SkosConcept.objects.get_or_create(pref_label=row['Genre'])
temp_genre.save()
temp_genre.scheme = [ecce_genre]
temp_dialect,_ = SkosConcept.objects.get_or_create(pref_label=row['Dialect'])
temp_dialect.save()
temp_dialect.scheme = [ecce_dialect]
for i, row in df.iterrows():
temp_corpus,_ = Corpus.objects.get_or_create(name=row['Corpus'])
temp_corpus.save()
temp_date,_ = Date.objects.get_or_create(dates=row['MeanDate'])
temp_date.save()
temp_text,_ = Text.objects.get_or_create(
text=row['Text'],
date=row['Date'],
genre=SkosConcept.objects.get(pref_label=row['Genre']),
corpus = temp_corpus,
mean_date = temp_date,
lower=int(row['Lower']),
size = float(row['Size']),
dialect=SkosConcept.objects.get(pref_label=row['Dialect'])
)
temp_text.save()
SkosConceptScheme.objects.all().delete()
#for x in Text.objects.all():
# x.delete()
| import_texts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from TT_utilities import Case, NL_METHODS
from scipy.stats import stats
from pathlib import Path
import pandas as pd
import numpy as np
import re
RECORD_DIRS = list(Path("./Data").glob("*p00*"))
CASES = list()
for record_dir in RECORD_DIRS:
record_name = re.search("p[0-9]{6}", str(record_dir))[0]
c = Case(record_dir.joinpath(record_name))
c.process()
CASES.append(c)
for c in CASES:
try:
c.process()
except ValueError:
pass
# +
# for mean (_m) and variance (_v)
columns = ["case", "record", "cond"]
for m in NL_METHODS:
columns.extend([m["tag"]+"_m", m["tag"]+"_v"])
print(columns)
CSV_DATA = pd.DataFrame(columns=columns)
CSV_DATA
# -
for n, c in enumerate(CASES):
for r in c:
vals = list()
for k, v in r.N_LINEAR.items():
s = stats.describe(v)
vals.extend([s[2], s[3]])
row = [c._case_name, r.name, c.pathology] + vals
CSV_DATA = CSV_DATA.append(pd.Series(data=row, index=columns), ignore_index=True)
CSV_DATA
CSV_DATA.to_csv("worksample_data.csv")
| Notebooks/csv_tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
directory = '/datasets/Our_dataset'
selected_cat = 'TED'
# +
# Read file names in videos directory
video_names = []
for video_file in os.listdir(os.path.join(directory, selected_cat)):
if video_file.endswith(".mp4"):
video_names.append(video_file)
num_files = len(video_names)
print('found', num_files, 'files')
#print(video_names)
# +
# Read spreadsheet
df = pd.read_excel(os.path.join(directory, selected_cat +'.xlsx'))
data = df[df['Video'].str.contains(video_names[0][:-4])==True]
print(data.shape)
print(data.iloc[0]['Link'])
print(data.iloc[0]['Video'])
# -
text_to_id("¿Cuál es tu mejor canción <NAME> TEDxMatamoros.mp4")
# +
filepath = 'prueba_lines.txt'
with open(filepath) as fp:
lines = fp.read().splitlines()
print(lines)
lines.append('another one')
with open(filepath, "w") as fp:
for line in lines:
print(line, file=fp)
# -
| .ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #Correlation Functions
#
# ###Contents
#
# - [Two-Time Correlation Functions](#twotime)
# - [Steady State Correlation Functions](#steady)
# - [Emission Spectrum](#emission)
# - [Non-Steady State Correlation Function](#nonsteady)
# %matplotlib inline
import numpy as np
from pylab import *
from qutip import *
# <a id='twotime'></a>
# ##Two-Time Correlation Functions
#
# With the QuTiP time-evolution functions (for example `mesolve` and `mcsolve`), a state vector or density matrix can be evolved from an initial state at :math:`t_0` to an arbitrary time $t$, $\rho(t)=V(t, t_0)\left\{\rho(t_0)\right\}$, where $V(t, t_0)$ is the propagator defined by the equation of motion. The resulting density matrix can then be used to evaluate the expectation values of arbitrary combinations of *same-time* operators.
#
# To calculate *two-time* correlation functions on the form $\left<A(t+\tau)B(t)\right>$, we can use the quantum regression theorem to write
#
# $$
# \left<A(t+\tau)B(t)\right> = {\rm Tr}\left[A V(t+\tau, t)\left\{B\rho(t)\right\}\right]
# = {\rm Tr}\left[A V(t+\tau, t)\left\{BV(t, 0)\left\{\rho(0)\right\}\right\}\right]
# $$
#
# We therefore first calculate $\rho(t)=V(t, 0)\left\{\rho(0)\right\}$ using one of the QuTiP evolution solvers with $\rho(0)$ as initial state, and then again use the same solver to calculate $V(t+\tau, t)\left\{B\rho(t)\right\}$ using $B\rho(t)$ as the initial state. Note that if the intial state is the steady state, then $\rho(t)=V(t, 0)\left\{\rho_{\rm ss}\right\}=\rho_{\rm ss}$ and
#
# $$
# \left<A(t+\tau)B(t)\right> = {\rm Tr}\left[A V(t+\tau, t)\left\{B\rho_{\rm ss}\right\}\right]
# = {\rm Tr}\left[A V(\tau, 0)\left\{B\rho_{\rm ss}\right\}\right] = \left<A(\tau)B(0)\right>,
# $$
#
# which is independent of $t$, so that we only have one time coordinate $\tau$.
#
#
# QuTiP provides a family of functions that assists in the process of calculating two-time correlation functions. The available functions and their usage is show in the table below. Each of these functions can use one of the following evolution solvers: Master-equation, Exponential series and the Monte-Carlo. The choice of solver is defined by the optional argument ``solver``.
#
#
# <table>
# <tr>
# <th>QuTiP Function</th>
# <th>Correlation Function Type</th>
# </tr>
# <tr>
# <td>`correlation` or `correlation_2op_2t`</td>
# <td>$\left<A(t+\tau)B(t)\right>$ or $\left<A(t)B(t+\tau)\right>$. </td>
# </tr>
# <tr>
# <td>`correlation_ss` or `correlation_2op_1t`</td>
# <td>$\left<A(\tau)B(0)\right>$ or $\left<A(0)B(\tau)\right>$.</td>
# </tr>
# <tr>
# <td>`correlation_3op_1t`</td>
# <td>$\left<A(0)B(\tau)C(0)\right>$.</td>
# </tr>
# <tr>
# <td>`correlation_3op_2t`</td>
# <td>$\left<A(t)B(t+\tau)C(t)\right>$.</td>
# </tr>
# <tr>
# <td>`correlation_4op_1t` <font color='red'>(Depreciated)</font></td>
# <td>$\left<A(0)B(\tau)C(\tau)D(0)\right>$</td>
# </tr>
# <tr>
# <td>`correlation_4op_2t` <font color='red'>(Depreciated)</font></td>
# <td style='min-width:200px'>$\left<A(t)B(t+\tau)C(t+\tau)D(t)\right>$ </td>
# </tr>
# </table>
#
# The most common use-case is to calculate correlation functions of the kind $\left<A(\tau)B(0)\right>$, in which case we use the correlation function solvers that start from the steady state, e.g., the `correlation_2op_1t` function. These correlation function solvers return a vector or matrix (in general complex) with the correlations as a function of the delay times.
# <a id='steady'></a>
# ##Steady State Correlation Function
#
# The following code demonstrates how to calculate the $\left<x(t)x(0)\right>$ correlation for a leaky cavity with three different relaxation rates.
# +
times = np.linspace(0,10.0,200)
a = destroy(10)
x = a.dag() + a
H = a.dag() * a
corr1 = correlation_2op_1t(H, None, times, [np.sqrt(0.5) * a], x, x)
corr2 = correlation_2op_1t(H, None, times, [np.sqrt(1.0) * a], x, x)
corr3 = correlation_2op_1t(H, None, times, [np.sqrt(2.0) * a], x, x)
plot(times, np.real(corr1), times, np.real(corr2), times, np.real(corr3))
legend(['0.5','1.0','2.0'])
xlabel(r'Time $t$')
ylabel(r'Correlation $\left<x(t)x(0)\right>$')
show()
# -
# <a id='emission'></a>
# ##Emission Spectrum
#
# Given a correlation function $\left<A(\tau)B(0)\right>$ we can define the corresponding power spectrum as
#
# $$
# S(\omega) = \int_{-\infty}^{\infty} \left<A(\tau)B(0)\right> e^{-i\omega\tau} d\tau.
# $$
#
# In QuTiP, we can calculate $S(\omega)$ using either `spectrum`, which first calculates the correlation function using the `essolve` solver and then performs the Fourier transform semi-analytically, or we can use the function `spectrum_correlation_fft` to numerically calculate the Fourier transform of a given correlation data using FFT.
#
# The following example demonstrates how these two functions can be used to obtain the emission power spectrum.
# +
N = 4 # number of cavity fock states
wc = wa = 1.0 * 2 * np.pi # cavity and atom frequency
g = 0.1 * 2 * np.pi # coupling strength
kappa = 0.75 # cavity dissipation rate
gamma = 0.25 # atom dissipation rate
# Jaynes-Cummings Hamiltonian
a = tensor(destroy(N), qeye(2))
sm = tensor(qeye(N), destroy(2))
H = wc * a.dag() * a + wa * sm.dag() * sm + g * (a.dag() * sm + a * sm.dag())
# collapse operators
n_th = 0.25
c_ops = [np.sqrt(kappa * (1 + n_th)) * a,
np.sqrt(kappa * n_th) * a.dag(), np.sqrt(gamma) * sm]
# calculate the correlation function using the mesolve solver, and then fft to
# obtain the spectrum. Here we need to make sure to evaluate the correlation
# function for a sufficient long time and sufficiently high sampling rate so
# that the discrete Fourier transform (FFT) captures all the features in the
# resulting spectrum.
tlist = np.linspace(0, 100, 5000)
corr = correlation_2op_1t(H, None, tlist, c_ops, a.dag(), a)
wlist1, spec1 = spectrum_correlation_fft(tlist, corr)
# calculate the power spectrum using spectrum, which internally uses essolve
# to solve for the dynamics (by default)
wlist2 = np.linspace(0.25, 1.75, 200) * 2 * np.pi
spec2 = spectrum(H, wlist2, c_ops, a.dag(), a)
# plot the spectra
fig, ax = subplots(1, 1)
ax.plot(wlist1 / (2 * np.pi), spec1, 'b', lw=2, label='eseries method')
ax.plot(wlist2 / (2 * np.pi), spec2, 'r--', lw=2, label='me+fft method')
ax.legend()
ax.set_xlabel('Frequency')
ax.set_ylabel('Power spectrum')
ax.set_title('Vacuum Rabi splitting')
ax.set_xlim(wlist2[0]/(2*np.pi), wlist2[-1]/(2*np.pi))
show()
# -
# <a id='nonsteady'></a>
# ##Non-Steady State Correlation Function
#
# More generally, we can also calculate correlation functions of the kind $\left<A(t_1+t_2)B(t_1)\right>$, i.e., the correlation function of a system that is not in its steadystate. In QuTiP, we can evoluate such correlation functions using the function `correlation_2op_2t`. The default behavior of this function is to return a matrix with the correlations as a function of the two time coordinates ($t_1$ and $t_2$).
# +
times = np.linspace(0, 10.0, 200)
a = destroy(10)
x = a.dag() + a
H = a.dag() * a
alpha = 2.5
rho0 = coherent_dm(10, alpha)
corr = correlation_2op_2t(H, rho0, times, times, [np.sqrt(0.25) * a], x, x)
pcolor(np.real(corr))
colorbar()
xlabel(r'Time $t_2$')
ylabel(r'Time $t_1$')
title(r'Correlation $\left<x(t)x(0)\right>$')
show()
# -
# However, in some cases we might be interested in the correlation functions on the form $\left<A(t_1+t_2)B(t_1)\right>$, but only as a function of time coordinate $t_2$. In this case we can also use the `correlation_2op_2t` function, if we pass the density matrix at time $t_1$ as second argument, and `None` as third argument. The `correlation_2op_2t` function then returns a vector with the correlation values corresponding to the times in `taulist` (the fourth argument).
#
# ###Ex: First-Order Optical Coherence Function
#
# This example demonstrates how to calculate a correlation function on the form $\left<A(\tau)B(0)\right>$ for a non-steady initial state. Consider an oscillator that is interacting with a thermal environment. If the oscillator initially is in a coherent state, it will gradually decay to a thermal (incoherent) state. The amount of coherence can be quantified using the first-order optical coherence function
# $$
# g^{(1)}(\tau) = \frac{\left<a^\dagger(\tau)a(0)\right>}{\sqrt{\left<a^\dagger(\tau)a(\tau)\right>\left<a^\dagger(0)a(0)\right>}}.
# $$
# For a coherent state $|g^{(1)}(\tau)| = 1$, and for a completely incoherent (thermal) state $g^{(1)}(\tau) = 0$. The following code calculates and plots $g^{(1)}(\tau)$ as a function of $\tau$.
# +
N = 15
taus = np.linspace(0,10.0,200)
a = destroy(N)
H = 2 * np.pi * a.dag() * a
# collapse operator
G1 = 0.75
n_th = 2.00 # bath temperature in terms of excitation number
c_ops = [np.sqrt(G1 * (1 + n_th)) * a, np.sqrt(G1 * n_th) * a.dag()]
# start with a coherent state
rho0 = coherent_dm(N, 2.0)
# first calculate the occupation number as a function of time
n = mesolve(H, rho0, taus, c_ops, [a.dag() * a]).expect[0]
# calculate the correlation function G1 and normalize with n to obtain g1
G1 = correlation_2op_2t(H, rho0, None, taus, c_ops, a.dag(), a)
g1 = G1 / np.sqrt(n[0] * n)
plot(taus, np.real(g1), 'b')
plot(taus, n, 'r')
title('Decay of a coherent state to an incoherent (thermal) state')
xlabel(r'$\tau$')
legend((r'First-order coherence function $g^{(1)}(\tau)$',
r'occupation number $n(\tau)$'))
show()
# -
# For convenience, the steps for calculating the first-order coherence function have been collected in the function `coherence_function_g1`.
#
#
# ###Example: Second-Order Optical Coherence Function
#
# The second-order optical coherence function, with time-delay $\tau$, is defined as
#
# $$
# \displaystyle g^{(2)}(\tau) = \frac{\langle a^\dagger(0)a^\dagger(\tau)a(\tau)a(0)\rangle}{\langle a^\dagger(0)a(0)\rangle^2}
# $$
#
# For a coherent state $g^{(2)}(\tau) = 1$, for a thermal state $g^{(2)}(\tau=0) = 2$ and it decreases as a function of time (bunched photons, they tend to appear together), and for a Fock state with $n$ photons $g^{(2)}(\tau = 0) = n(n - 1)/n^2 < 1$ and it increases with time (anti-bunched photons, more likely to arrive separated in time).
#
# To calculate this type of correlation function with QuTiP, we could use `correlation_4op_1t`, which computes a correlation function of the form $\left<A(0)B(\tau)C(\tau)D(0)\right>$ (four operators, one delay-time vector). However, the middle pair of operators are evaluated at the same time $\tau$, and thus can be simplified to a single operator $E(\tau)=B(\tau)C(\tau)$, and we can instead call the `correlation_3op_1t` function to compute $\left<A(0)E(\tau)D(0)\right>$. This simplification is done automatically inside the depreciated `correlation_4op_1t` function that calls `correlation_3op_1t` internally.
#
# The following code calculates and plots $g^{(2)}(\tau)$ as a function of $\tau$ for coherent, thermal and fock states.
# +
N = 25
taus = np.linspace(0, 25.0, 200)
a = destroy(N)
H = 2 * np.pi * a.dag() * a
kappa = 0.25
n_th = 2.0 # bath temperature in terms of excitation number
c_ops = [np.sqrt(kappa * (1 + n_th)) * a, np.sqrt(kappa * n_th) * a.dag()]
states = [{'state': coherent_dm(N, np.sqrt(2)), 'label': "coherent state"},
{'state': thermal_dm(N, 2), 'label': "thermal state"},
{'state': fock_dm(N, 2), 'label': "Fock state"}]
fig, ax = subplots(1, 1)
for state in states:
rho0 = state['state']
# first calculate the occupation number as a function of time
n = mesolve(H, rho0, taus, c_ops, [a.dag() * a]).expect[0]
# calculate the correlation function G2 and normalize with n(0)n(t) to
# obtain g2
G2 = correlation_3op_1t(H, rho0, taus, c_ops, a.dag(), a.dag() * a, a)
g2 = G2 / (n[0] * n)
ax.plot(taus, np.real(g2), label=state['label'], lw=2)
ax.legend(loc=0)
ax.set_xlabel(r'$\tau$')
ax.set_ylabel(r'$g^{(2)}(\tau)$')
show()
# -
# For convenience, the steps for calculating the second-order coherence function have been collected in the function `coherence_function_g2`.
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/guide.css", "r").read()
return HTML(styles)
css_styling()
| qutip-notebooks-master/docs/guide/CorrelationFunctions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from six.moves import cPickle as pickle
valid_data = pd.read_csv('data/full_train_wo_appts_cleaned_encoded.csv')
valid_data = valid_data.tail()
valid_data[['patient_id','Bucket','Revenue']]
valid_data = valid_data.drop_duplicates()
valid_pat_ids = valid_data['patient_id']
# Removing unnecessary columns
X_valid = valid_data.drop(columns=['patient_id','Bucket','Revenue'])
Y_valid = valid_data['Bucket']
Y_valid_reg = valid_data['Revenue']
Y_valid
Y_valid_reg
try:
# Loading the classifier
#with open('data/train_svc_clf_.pickle', 'rb') as f:
with open('data/train_svc_clf_wo_pat_id.pickle', 'rb') as f:
clf = pickle.load(f)
except Exception as e:
print('Exception ', e)
try:
# Loading the classifier
#with open('data/train_svc_clf_.pickle', 'rb') as f:
with open('data/train_svr_reg.pickle', 'rb') as f:
reg = pickle.load(f)
except Exception as e:
print('Exception ', e)
Y_predict = clf.predict(X_valid)
Y_predict_reg = reg.predict(X_valid)
Y_predict
Y_predict_reg
# Seems we got the same output for Classifier, the predictions are good.
# But it is not the same case for Regressor, it failed miserably
| 03_18_portea_challenge/portea_ml_train_validate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Auto Correct in Python
#
# Given a word w, find the most likely correction `c = correct(w)`.
#
# Approach: Try all candidate words `c` that are known words that are near `w`. Choose the most likely one.
#
# Our main data from where we'll be pulling out words is from [this location](http://www.openbookproject.net/courses/python4fun/_static/spellcheck/spell.words)
# ## 1. Download the data programmatically or via CLI (choose your poison)
#
# ### 1.1. Programmatically
# ### 1.2. Via CLI
# ***
#
# ## 2. Look around to read the file
#
# ### Choose the option that you are familiar with covered during the course
# ***
# ## 3. Make sure that the words don't contain any special characters. If you find any try to clean up the files
# ***
# ## 4. What does the theory say?
#
# ### 4.1 Generally, if we want to check if the word in contention is correct or not we need to calculate the edit distance:
#
# > Edit distance is a way of quantifying how dissimilar two strings (e.g., words) are to one another by counting the minimum number of operations required to transform one string into the other.
#
# > Edit distances find applications in natural language processing, where automatic spelling correction can determine candidate corrections for a misspelled word by selecting words from a dictionary that have a low distance to the word in question
# ### 4.2 Most commonly taught one is Levenstein's distance:
#
# #### Example:
#
# The Levenshtein distance between "kitten" and "sitting" is 3. A minimal edit script that transforms the former into the latter is:
#
# 
# ### 4.3. But we won't be going that way, instead we'll be generating encodings first
#
# In generic terms what are encodings:
#
# 
# ## The encoding algorithm that you'll have to write is: _Soundex Encoding_ (albeit it's variant)
# ***
# ## 5. Implementing Soundex Algorithm
#
# What are the steps in Soundex Algorithm
#
# - Retain the first letter of the name
# - Replace consonants with digits as follows (after the first letter):
# - a, e, i, o, u, y, h, w = 0
# - b, f, p, v = 1
# - c, g, j, k, q, s, x, z = 2
# - d, t = 3
# - l = 4
# - m, n = 5
# - r = 6
# - If two or more letters with the same number are adjacent in the original name (before step 1), only retain the first letter
# - (__We won't be coding this part__) If you have too few letters in your word that you can't assign three numbers, append with zeros until there are three numbers. If you have more than 3 letters, just retain the first 3 numbers.
# ### 5.1. Retaining the first letter of the name
#
# Example: Suppose your word is `comittee`. You need to retain: `c` from `comittee` separately. The next steps will be run on `omittee`
# ### 5.2. Replace consonants with digits (__refer above__):
#
# Example: `omittee` would be eventually --> `0503300`
# ### 5.3. If two or more letters with the same number are adjacent in the original name (before step 1), only retain the first letter
#
# Example: `0503300` would be --> `05030`, i.e. the adjacent `33` would be just replaced by `3`. Similar with `00`
# ***
# ## 6. Above are the building blocks for the algorithm. Now create a function `generate_soundex_encodings`
#
# #### `generate_soundex_encodings` would take in a list of the words (remember `words` from task two?) and return: `soundex_encodings`
# ***
# ## 7. Now create another function to calculate the encodings for the entire list `words` against this and save this as a `pickle` file. Read about pickle [here](https://stackoverflow.com/questions/4530611/saving-and-loading-objects-and-using-pickle)
# ***
# ## 8. Once we have the soundex encodings generated, next task is to find out a way to calculate distances comparing the encoding of the incoming word and the already generated soundex encodings
#
# ### 8.1. Below are the steps for the Jaro distance algorithm
#
# > The Jaro distance is a measure of similarity between two strings.
#
# > The higher the Jaro distance for two strings is, the more similar the strings are.
#
# > The score is normalized such that 0 equates to no similarity and 1 is an exact match.
#
# 
#
# ### 8.2. Example:
#
# 
# ### 8.3 Find the number of matching characters
# ### 8.4 Find the lengths of the two given strings
# ### 8.5 Find `t` i.e. half number of transpositions
#
# `t` is the number of characters that are shared but are in different positions, divided by `2`.
#
# For __MARTHA__ and __MARHTA__ , 2 characters (H and T) are shared but are in different positions so `t = 2/2 = 1`.
#
# But for __DWAYNE__ and __DUANE__ it is `t = 0/2 = 0`.
# ### 8.6 Write a function, `get_jaro_distance()` to calculate Jaro distance for both the strings
# ***
#
# ## 9. Next step will be to write a function: `autocorrect()` that will take in: Soundex encodings for all the words and the `target_word` that needs to be autocorrected
#
# Example: autocorrect(words_encodings, target_word="comittee") --> output would be top five choices of words for which the jaro distance (__see above__) is highest
#
# Output expected: `['committee', 'commit', 'comet', 'committed', 'com']` (__NOTE: Not a representative example__)
# ### Finally, you done folks with your own auto-correct
| notebooks/autocorrect/Creating our own auto-correct-Problem Statement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-mh4fYe9vvuk"
# # Model Evaluation
#
# + [markdown] id="NrP6gOUsaVsy"
# ## Classifier Evaluation
#
# ### True Positive (TP)
#
# The predicted value matches the actual value
# The actual value was positive and the model predicted a positive value
#
# ### True Negative (TN)
#
# The predicted value matches the actual value
# The actual value was negative and the model predicted a negative value
#
# ### False Positive (FP) – Type 1 error
#
# The predicted value was falsely predicted
# The actual value was negative but the model predicted a positive value
# Also known as the Type 1 error
#
# ### False Negative (FN) – Type 2 error
#
# The predicted value was falsely predicted
# The actual value was positive but the model predicted a negative value
# Also known as the Type 2 error
#
# ### Precision
# Precision is a good measure to determine, when the costs of False Positive is high. For instance, email spam detection. In email spam detection, a false positive means that an email that is non-spam (actual negative) has been identified as spam (predicted spam). The email user might lose important emails if the precision is not high for the spam detection model.
#
# $$Precision = \dfrac{TP}{TP + FP}$$
#
# ### Recall
# Recall shall be the model metric we use to select our best model when there is a high cost associated with False Negative.
# in sick patient detection. If a sick patient (Actual Positive) goes through the test and predicted as not sick (Predicted Negative). The cost associated with False Negative will be extremely high if the sickness is contagious.
#
# $$Recall = \dfrac{TP}{TP + FN}$$
#
# ### F1 score
# F1 Score might be a better measure to use if we need to seek a balance between Precision and Recall AND there is an uneven class distribution (large number of Actual Negatives).
#
# $$F1 = 2 \cdot \dfrac{Precision \cdot Recall}{Precision + Recall}$$
#
# ### Confusion Matrix
# A Confusion matrix is an N x N matrix used for evaluating the performance of a classification model, where N is the number of target classes. The matrix compares the actual target values with those predicted by the machine learning model. This gives us a holistic view of how well our classification model is performing and what kinds of errors it is making.
# + id="WVfD0etJidZq"
from sklearn import linear_model, metrics, datasets, model_selection
# + id="1LqNVHlaikt4"
x, y = datasets.load_breast_cancer(return_X_y=True)
x_train, x_test, y_train, y_test = model_selection.train_test_split(x, y, test_size = 0.1, random_state=42, stratify=y)
# + colab={"base_uri": "https://localhost:8080/"} id="ti3WlBgD40JL" outputId="4eeff02c-2a57-4677-bc0a-bcf6ff385736"
lgr = linear_model.LogisticRegression(solver='liblinear',random_state=42)
lgr.fit(x_train, y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 453} id="nevs86s-i-t0" outputId="5bce484c-8289-4652-d4fd-5eff58025937"
y_pred = lgr.predict(x_test)
print("Accuracy Score:", metrics.accuracy_score(y_test, y_pred))
cf_matrix = metrics.confusion_matrix(y_test, y_pred)
tp, tn, fp, fn = cf_matrix[1, 1], cf_matrix[0, 0], cf_matrix[0, 1], cf_matrix[1, 0]
print("True Positive:", tp)
print("True Negative:", tn)
print("False Positive:", fp)
print("False Negative:", fn)
precision1 = metrics.precision_score(y_test, y_pred)
precision2 = tp / (tp + fp)
print("Precision Score:", precision1)
print("Precision Score Manual:", precision2)
recall1 = metrics.recall_score(y_test, y_pred)
recall2 = tp / (tp + fn)
print("Recall Score:", recall1)
print("Recall Score Manual:", recall2)
metrics.plot_confusion_matrix(lgr, x_test, y_test)
# + [markdown] id="49Zi7e-89fwb"
# ## Regression Metrics
#
# ### Mean Squared Error
#
# MSE is the average of the squared error that is used as the loss function for least squares regression: It is the sum, over all the data points, of the square of the difference between the predicted and actual target variables, divided by the number of data points.
#
# $$mse = 1/m \sum_{i=0}^{m} (\hat{y_i} - y_i)^2$$
#
# Where m is the total number of examples being tested, and $\hat{y}$ is the predicted label while $y$ is the actual label
#
# ### Mean Absolute Error
#
# MAE is the average of the absolue errors that is used as the loss function for regression: It is the sum, over all the data points, of the absolute of the difference between the predicted and actual target variables, divided by the number of data points.
#
# $$mae = 1/m \sum_{i=0}^{m} |\hat{y_i} - y_i|$$
#
# Where m is the total number of examples being tested, and $\hat{y}$ is the predicted label while $y$ is the actual label
#
#
# ### $R^2$ score
# the $R^2$ score varies between 0 and 100%. It is closely related to the MSE .
#
# if it is 100%, the two variables are perfectly correlated, i.e., with no variance at all. A low value would show a low level of correlation, meaning a regression model that is not valid, but not in all cases.
#
# + id="92KHMKxH6n72"
x, y = datasets.load_boston(return_X_y=True)
x_train, x_test, y_train, y_test = model_selection.train_test_split(x, y, test_size = 0.1, random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="GOOfCGA3nH_e" outputId="5a1e0ef1-e21e-4c5e-b5ad-d05059ea95e6"
lnr = linear_model.LinearRegression()
lnr.fit(x_train, y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="PxpRftWOnMoV" outputId="9ef31b05-0a80-4240-c858-e56d07006ce5"
y_pred = lnr.predict(x_test)
print("Mean Squared Error:", metrics.mean_squared_error(y_test, y_pred))
print("Mean Absolute Error:", metrics.mean_absolute_error(y_test, y_pred))
print("R2 Score:", metrics.r2_score(y_test, y_pred))
| W3/Evaluation_Metrics (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stellargraph example: Graph Attention Network (GAT) on the CORA citation dataset
# + [markdown] tags=["CloudRunner"]
# <table><tr><td>Run the master version of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/node-classification/gat/gat-cora-node-classification-example.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/node-classification/gat/gat-cora-node-classification-example.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
# -
# Import NetworkX and stellar:
# + tags=["CloudRunner"]
# install StellarGraph if running on Google Colab
import sys
if 'google.colab' in sys.modules:
# %pip install -q stellargraph[demos]
# + tags=["VersionCheck"]
# verify that we're using the correct version of StellarGraph for this notebook
import stellargraph as sg
try:
sg.utils.validate_notebook_version("1.0.0b")
except AttributeError:
raise ValueError(
f"This notebook requires StellarGraph version 1.0.0b, but a different version {sg.__version__} is installed. Please see <https://github.com/stellargraph/stellargraph/issues/1172>."
) from None
# +
import networkx as nx
import pandas as pd
import os
import stellargraph as sg
from stellargraph.mapper import FullBatchNodeGenerator
from stellargraph.layer import GAT
from tensorflow.keras import layers, optimizers, losses, metrics, Model
from sklearn import preprocessing, feature_extraction, model_selection
from stellargraph import datasets
from IPython.display import display, HTML
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### Loading the CORA network
dataset = datasets.Cora()
display(HTML(dataset.description))
G, node_subjects = dataset.load()
print(G.info())
# We aim to train a graph-ML model that will predict the "subject" attribute on the nodes. These subjects are one of 7 categories:
set(node_subjects)
# ### Splitting the data
# For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to do this.
#
# Here we're taking 140 node labels for training, 500 for validation, and the rest for testing.
train_subjects, test_subjects = model_selection.train_test_split(
node_subjects, train_size=140, test_size=None, stratify=node_subjects
)
val_subjects, test_subjects = model_selection.train_test_split(
test_subjects, train_size=500, test_size=None, stratify=test_subjects
)
# Note using stratified sampling gives the following counts:
# +
from collections import Counter
Counter(train_subjects)
# -
# The training set has class imbalance that might need to be compensated, e.g., via using a weighted cross-entropy loss in model training, with class weights inversely proportional to class support. However, we will ignore the class imbalance in this example, for simplicity.
# ### Converting to numeric arrays
# For our categorical target, we will use one-hot vectors that will be fed into a soft-max Keras layer during training. To do this conversion ...
# +
target_encoding = preprocessing.LabelBinarizer()
train_targets = target_encoding.fit_transform(train_subjects)
val_targets = target_encoding.transform(val_subjects)
test_targets = target_encoding.transform(test_subjects)
# -
# We now do the same for the node attributes we want to use to predict the subject. These are the feature vectors that the Keras model will use as input. The CORA dataset contains attributes 'w_x' that correspond to words found in that publication. If a word occurs more than once in a publication the relevant attribute will be set to one, otherwise it will be zero.
# ## Creating the GAT model in Keras
# To feed data from the graph to the Keras model we need a generator. Since GAT is a full-batch model, we use the `FullBatchNodeGenerator` class to feed node features and graph adjacency matrix to the model.
generator = FullBatchNodeGenerator(G, method="gat")
# For training we map only the training nodes returned from our splitter and the target values.
train_gen = generator.flow(train_subjects.index, train_targets)
# Now we can specify our machine learning model, we need a few more parameters for this:
#
# * the `layer_sizes` is a list of hidden feature sizes of each layer in the model. In this example we use two GAT layers with 8-dimensional hidden node features for the first layer and the 7 class classification output for the second layer.
# * `attn_heads` is the number of attention heads in all but the last GAT layer in the model
# * `activations` is a list of activations applied to each layer's output
# * Arguments such as `bias`, `in_dropout`, `attn_dropout` are internal parameters of the model, execute `?GAT` for details.
# To follow the GAT model architecture used for Cora dataset in the original paper [Graph Attention Networks. <NAME>ic et al. ICLR 2018 https://arxiv.org/abs/1710.10903], let's build a 2-layer GAT model, with the 2nd layer being the classifier that predicts paper subject: it thus should have the output size of `train_targets.shape[1]` (7 subjects) and a softmax activation.
gat = GAT(
layer_sizes=[8, train_targets.shape[1]],
activations=["elu", "softmax"],
attn_heads=8,
generator=generator,
in_dropout=0.5,
attn_dropout=0.5,
normalize=None,
)
# Expose the input and output tensors of the GAT model for node prediction, via GAT.in_out_tensors() method:
x_inp, predictions = gat.in_out_tensors()
# ### Training the model
# Now let's create the actual Keras model with the input tensors `x_inp` and output tensors being the predictions `predictions` from the final dense layer
model = Model(inputs=x_inp, outputs=predictions)
model.compile(
optimizer=optimizers.Adam(lr=0.005),
loss=losses.categorical_crossentropy,
metrics=["acc"],
)
# Train the model, keeping track of its loss and accuracy on the training set, and its generalisation performance on the validation set (we need to create another generator over the validation data for this)
val_gen = generator.flow(val_subjects.index, val_targets)
# Create callbacks for early stopping (if validation accuracy stops improving) and best model checkpoint saving:
# +
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
if not os.path.isdir("logs"):
os.makedirs("logs")
es_callback = EarlyStopping(
monitor="val_acc", patience=20
) # patience is the number of epochs to wait before early stopping in case of no further improvement
mc_callback = ModelCheckpoint(
"logs/best_model.h5", monitor="val_acc", save_best_only=True, save_weights_only=True
)
# -
# Train the model
history = model.fit(
train_gen,
epochs=50,
validation_data=val_gen,
verbose=2,
shuffle=False, # this should be False, since shuffling data means shuffling the whole graph
callbacks=[es_callback, mc_callback],
)
# Plot the training history:
sg.utils.plot_history(history)
# Reload the saved weights of the best model found during the training (according to validation accuracy)
model.load_weights("logs/best_model.h5")
# Evaluate the best model on the test set
test_gen = generator.flow(test_subjects.index, test_targets)
test_metrics = model.evaluate(test_gen)
print("\nTest Set Metrics:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
# ### Making predictions with the model
# Now let's get the predictions for all nodes:
all_nodes = node_subjects.index
all_gen = generator.flow(all_nodes)
all_predictions = model.predict(all_gen)
# These predictions will be the output of the softmax layer, so to get final categories we'll use the `inverse_transform` method of our target attribute specifcation to turn these values back to the original categories
# Note that for full-batch methods the batch size is 1 and the predictions have shape $(1, N_{nodes}, N_{classes})$ so we we remove the batch dimension to obtain predictions of shape $(N_{nodes}, N_{classes})$.
node_predictions = target_encoding.inverse_transform(all_predictions.squeeze())
# Let's have a look at a few predictions after training the model:
df = pd.DataFrame({"Predicted": node_predictions, "True": node_subjects})
df.head(20)
# ## Node embeddings
# Evaluate node embeddings as activations of the output of the 1st GraphAttention layer in GAT layer stack (the one before the top classification layer predicting paper subjects), and visualise them, coloring nodes by their true subject label. We expect to see nice clusters of papers in the node embedding space, with papers of the same subject belonging to the same cluster.
#
# Let's create a new model with the same inputs as we used previously `x_inp` but now the output is the embeddings rather than the predicted class. We find the embedding layer by taking the first graph attention layer in the stack of Keras layers. Additionally note that the weights trained previously are kept in the new model.
emb_layer = next(l for l in model.layers if l.name.startswith("graph_attention"))
print(
"Embedding layer: {}, output shape {}".format(emb_layer.name, emb_layer.output_shape)
)
embedding_model = Model(inputs=x_inp, outputs=emb_layer.output)
# The embeddings can now be calculated using the predict function. Note that the embeddings returned are 64 dimensional features (8 dimensions for each of the 8 attention heads) for all nodes.
emb = embedding_model.predict(all_gen)
emb.shape
# Project the embeddings to 2d using either TSNE or PCA transform, and visualise, coloring nodes by their true subject label
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import pandas as pd
import numpy as np
# Note that the embeddings from the GAT model have a batch dimension of 1 so we `squeeze` this to get a matrix of $N_{nodes} \times N_{emb}$.
X = emb.squeeze()
y = np.argmax(target_encoding.transform(node_subjects), axis=1)
if X.shape[1] > 2:
transform = TSNE # PCA
trans = transform(n_components=2)
emb_transformed = pd.DataFrame(trans.fit_transform(X), index=list(G.nodes()))
emb_transformed["label"] = y
else:
emb_transformed = pd.DataFrame(X, index=list(G.nodes()))
emb_transformed = emb_transformed.rename(columns={"0": 0, "1": 1})
emb_transformed["label"] = y
# +
alpha = 0.7
fig, ax = plt.subplots(figsize=(7, 7))
ax.scatter(
emb_transformed[0],
emb_transformed[1],
c=emb_transformed["label"].astype("category"),
cmap="jet",
alpha=alpha,
)
ax.set(aspect="equal", xlabel="$X_1$", ylabel="$X_2$")
plt.title(
"{} visualization of GAT embeddings for cora dataset".format(transform.__name__)
)
plt.show()
# + [markdown] tags=["CloudRunner"]
# <table><tr><td>Run the master version of this notebook:</td><td><a href="https://mybinder.org/v2/gh/stellargraph/stellargraph/master?urlpath=lab/tree/demos/node-classification/gat/gat-cora-node-classification-example.ipynb" alt="Open In Binder" target="_parent"><img src="https://mybinder.org/badge_logo.svg"/></a></td><td><a href="https://colab.research.google.com/github/stellargraph/stellargraph/blob/master/demos/node-classification/gat/gat-cora-node-classification-example.ipynb" alt="Open In Colab" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg"/></a></td></tr></table>
| demos/node-classification/gat/gat-cora-node-classification-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercises for "Hands-on with Pydata: How to Build a Minimal Recommendation Engine"
# # Systems check: imports and files
import numpy as np
import pandas as pd
# # Pandas questions: Series and DataFrames
# ## 1. Adding a column in a DataFrame
# given the following DataFrame, add a new column to it
df = pd.DataFrame({'col1': [1,2,3,4]})
df['col2']=[5,6,7,8]
df
# ## 2. Deleting a row in a DataFrame
# given the following DataFrame, delete row 'd' from it
df = pd.DataFrame({'col1': [1,2,3,4]}, index = ['a','b','c','d'])
df.drop('a',axis=0)
# ## 3. Creating a DataFrame from a few Series
# given the following three Series, create a DataFrame such that it holds them as its columns
ser_1 = pd.Series(np.random.randn(6))
ser_2 = pd.Series(np.random.randn(6))
ser_3 = pd.Series(np.random.randn(6))
df=pd.DataFrame({'1':ser_1,'2':ser_2, '3':ser_3})
df
# # Pandas questions: Indexing
# ## 1. Indexing into a specific column
# given the following DataFrame, try to index into the 'col_2' column
df = pd.DataFrame(data={'col_1': [0.12, 7, 45, 10], 'col_2': [0.9, 9, 34, 11]},
columns=['col_1', 'col_2', 'col_3'],
index=['obs1', 'obs2', 'obs3', 'obs4'])
df.col_1
# ## 2. Label-based indexing
# using the same DataFrame, index into the row whose index is 'obs3'
df.loc['obs3']
# ## 2. Position-based indexing
# using the same DataFrame, index into into its first row
df.iloc[0,0]
# # Mini-Challenge prep: data loading
#
# ## 1. How to load the `users` and `movies` portions of MovieLens
# +
import pandas as pd
users = pd.read_table('data/ml-1m/users.dat',
sep='::', header=None,
names=['user_id', 'gender', 'age', 'occupation', 'zip'],engine='python')
movies = pd.read_table('data/ml-1m/movies.dat',
sep='::', header=None,
names=['movie_id', 'title', 'genres'],engine='python')
# -
# ## 2. How to load the training and testing subsets
# subset version (hosted notebook)
movielens_train = pd.read_csv('C:/Users/niharika/Desktop/data sciences/Springboard/Machine Learning/Recommender systems/1489104021_pycon2015_tutorial322/pycon2015_tutorial322/data/movielens_train.csv', index_col=0, encoding='latin')
movielens_test = pd.read_csv('C:/Users/niharika/Desktop/data sciences/Springboard/Machine Learning/Recommender systems/1489104021_pycon2015_tutorial322/pycon2015_tutorial322/data/movielens_test.csv', index_col=0, encoding='latin')
movielens_train.head()
movielens_test.head()
# # Mini-Challenge prep: evaluation functions
#
# These are the two functions that you will need to test your `estimate` method.
def compute_rmse(y_pred, y_true):
""" Compute Root Mean Squared Error. """
return np.sqrt(np.mean(np.power(y_pred - y_true, 2)))
def evaluate(estimate_f):
""" RMSE-based predictive performance evaluation with pandas. """
ids_to_estimate = zip(movielens_test.user_id, movielens_test.movie_id)
estimated = np.array([estimate_f(u,i) for (u,i) in ids_to_estimate])
real = movielens_test.rating.values
return compute_rmse(estimated, real)
# Test a dummy solution!
def my_estimate_func(user_id, movie_id):
return 3.0
# You can test for performance with the following line, which assumes that your function is called `my_estimate_func`:
print ('RMSE for my estimate function: %s' % evaluate(my_estimate_func))
# # Reco systems questions: Minimal reco engine v1.0
# ## 1. Simple collaborative filtering using mean ratings
# +
# write an 'estimate' function that computes the mean rating of a particular user
def collab_mean(user_id, movie_id):
# first, index into all ratings of this movie
# second, compute the mean of those ratings
user_condition = movielens_train.user_id != user_id
movie_condition = movielens_train.movie_id == movie_id
if movielens_train.loc[user_condition & movie_condition].empty:
return(3)
else:
a= movielens_train.loc[user_condition & movie_condition]
b= a.rating
return b.mean()
# try it out for a user_id, movie_id pair
collab_mean(4653, 2648)
# -
# # Mini-Challenge: first round
# Implement an `estimate` function of your own using other similarity notions, eg.:
#
# - collaborative filter based on age similarities
# - collaborative filter based on zip code similarities
# - collaborative filter based on occupation similarities
# - content filter based on movie genre
user_info = users.set_index('user_id')
user_info.head(5)
user_id = 3
user_info.loc[user_id, 'age']
# +
class CollabAgeReco:
""" Collaborative filtering using an implicit sim(u,u'). """
def learn(self):
""" Prepare datastructures for estimation. """
self.means_by_age = movielens_train.pivot_table('rating', index='movie_id', columns='age')
def estimate(self, user_id, movie_id):
""" Mean ratings by other users of the same gender. """
if movie_id not in self.means_by_age.index:
return 3.0
user_age = user_info.ix[user_id, 'age']
if ~np.isnan(self.means_by_age.ix[movie_id, user_age]):
return self.means_by_age.ix[movie_id, user_age]
else:
return self.means_by_age.ix[movie_id].mean()
reco = CollabAgeReco()
reco.learn()
print('RMSE for CollabGenderReco: %s' % evaluate(reco.estimate))
# -
# # Mini-Challenge: second round
# Implement an `estimate` function of your own using other custom similarity notions, eg.:
#
# - euclidean
# - cosine
def euclidean(s1, s2):
"""Take two pd.Series objects and return their euclidean 'similarity'."""
diff = s1 - s2
return 1 / (1 + np.sqrt(np.sum(diff ** 2)))
# +
users = pd.read_table('data/ml-1m/users.dat',
sep='::', header=None,
names=['user_id', 'gender', 'age', 'occupation', 'zip'], engine='python')
ratings = pd.read_table('data/ml-1m/ratings.dat',
sep='::', header=None,
names=['user_id', 'movie_id', 'rating', 'timestamp'], engine='python')
movies = pd.read_table('data/ml-1m/movies.dat',
sep='::', header=None,
names=['movie_id', 'title', 'genres'], engine='python')
# +
movielens = pd.merge(pd.merge(ratings, users), movies)
class CollabeuclideanReco:
""" Collaborative filtering using a custom sim(u,u'). """
def learn(self):
""" Prepare datastructures for estimation. """
self.all_user_profiles = movielens.pivot_table('rating', index='movie_id', columns='user_id')
def estimate(self, user_id, movie_id):
""" Ratings weighted by correlation similarity. """
user_condition = movielens_train.user_id != user_id
movie_condition = movielens_train.movie_id == movie_id
ratings_by_others = movielens_train.loc[user_condition & movie_condition]
if ratings_by_others.empty:
return 3.0
ratings_by_others.set_index('user_id', inplace=True)
their_ids = ratings_by_others.index
their_ratings = ratings_by_others.rating
their_profiles = self.all_user_profiles[their_ids]
user_profile = self.all_user_profiles[user_id]
sims = their_profiles.apply(lambda profile: euclidean(profile, user_profile), axis=0)
ratings_sims = pd.DataFrame({'sim': sims, 'rating': their_ratings})
ratings_sims = ratings_sims[ratings_sims.sim > 0]
if ratings_sims.empty:
return their_ratings.mean()
else:
return np.average(ratings_sims.rating, weights=ratings_sims.sim)
reco = CollabeuclideanReco()
reco.learn()
print('RMSE for CollabPearsonReco: %s' % evaluate(reco.estimate))
# -
| RecommenderSystem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:ap-northeast-1:102112518831:image/datascience-1.0
# ---
# # データセットの準備を行う
#
# 下記、Amazon Forecastのdocにある、電力消費量データを用いる。
# https://docs.aws.amazon.com/forecast/latest/dg/getting-started.html
#
# 48時間後までを予測するため、48時間前の情報から予測する必要があります。
# # 0.データセットのダウンロード
# !wget -P ../input https://docs.aws.amazon.com/forecast/latest/dg/samples/electricityusagedata.zip
# ## 1.データセット読み込み
import pandas as pd
df = pd.read_csv('../input/electricityusagedata.zip', names=['timestamp','demand','client'])
print(df.shape)
print(df['timestamp'].min())
print(df['timestamp'].max())
df.head()
df['client'] = df['client'].str.replace('client_','').astype(int)
df.head()
# ## 2.特徴量生成
# ### 1)時刻型に変換し、特徴量(時間帯、曜日)を抽出
df['timestamp'] = pd.to_datetime(df['timestamp'], format='%Y-%m-%d %H:%M:%S')
df['num_of_week'] = df['timestamp'].dt.dayofweek
df['hour'] = df['timestamp'].dt.hour
# ### 2)clientごとに時間差の値を特徴量にもつ(36hより前である必要がある)
df["before_36h"] = df.groupby(['client']).shift(36)['demand'].reset_index()['demand']
df["before_37h"] = df.groupby(['client']).shift(37)['demand'].reset_index()['demand']
df["before_38h"] = df.groupby(['client']).shift(38)['demand'].reset_index()['demand']
df["before_48h"] = df.groupby(['client']).shift(48)['demand'].reset_index()['demand']
df["before_72h"] = df.groupby(['client']).shift(72)['demand'].reset_index()['demand']
df["before_96h"] = df.groupby(['client']).shift(96)['demand'].reset_index()['demand']
# ### 3)clientごとに期間で集約する
df = df.sort_values(["client","timestamp"]).reset_index()
df['mean_24h'] = df.groupby(['client']).rolling(24)['demand'].mean().reset_index()['demand']
df['var_24h'] = df.groupby(['client']).rolling(24)['demand'].var().reset_index()['demand']
# +
#df = df.sort_values(["client","timestamp"]).reset_index()
#df['mean_24h_before_36h'] = df.groupby(['client']).rolling(2)['demand'].mean().reset_index()['demand']
# -
df["mean_24h_before_36h"] = df.groupby(['client']).shift(36)['mean_24h'].reset_index()['mean_24h']
df["var_24h_before_36h"] = df.groupby(['client']).shift(36)['var_24h'].reset_index()['var_24h']
df[df.client==21].head()
df[df.client==21].tail()
feature_col = [
'client',
'num_of_week',
'hour',
'before_36h',
'before_37h',
'before_38h',
'before_48h',
'before_72h',
'before_96h',
'mean_24h_before_36h',
'var_24h_before_36h'
]
#
# ## 3.データ分割
# all:2014-01-01 01:00:00 〜 2015-01-01 00:00:00
# train:2014-01-01 01:00:00 〜 2014-12-29 00:00:00
# valid:2014-12-29 01:00:00 〜 2014-12-30 12:00:00(36h)
# test:2014-12-30 13:00:00 〜 2015-01-01 00:00:00(36h)
df_train = df[df.timestamp <= '2014-12-29 00:00:00']
df_valid = df[(df.timestamp >= '2014-12-29 01:00:00') & (df.timestamp <= '2014-12-30 12:00:00')]
df_test = df[df.timestamp >= '2014-12-30 13:00:00']
df_train[df_train.client==21].shape
df_valid[df_valid.client==21].shape
df_test[df_test.client==21].shape
tr_x = df_train[feature_col]
tr_y = df_train['demand']
va_x = df_valid[feature_col]
va_y = df_valid['demand']
test_x = df_test[feature_col]
test_y = df_test[['timestamp','client','demand']]
# # XGBoost
# !pip install xgboost
import xgboost as xgb
dtrain = xgb.DMatrix(tr_x, label=tr_y)
dvalid = xgb.DMatrix(va_x, label=va_y)
dtest = xgb.DMatrix(test_x)
# ## 4.モデル学習
# 誤差はRMSEを用いる
#
# https://docs.aws.amazon.com/ja_jp/forecast/latest/dg/metrics.html
import xgboost as xgb
from sklearn.metrics import mean_squared_error
#fit by best params
regressor = xgb.XGBRegressor(n_estimators=50)
regressor.fit(tr_x, tr_y, eval_metric="rmse", eval_set=[(va_x, va_y)])
# ## 5.モデル評価
# ### testデータでのRMSE
import numpy as np
from sklearn.metrics import mean_squared_error
test_y
### RMSEを出力
np.sqrt(mean_squared_error(test_y['demand'], regressor.predict(test_x)))
# ### 重要度の可視化
xgb.plot_importance(regressor)
# ## 6.推論(予測)
# https://github.com/dmlc/xgboost/blob/master/demo/guide-python/sklearn_examples.py
regressor.predict(test_x)
# ## 7.推論結果の可視化
# client_21について
# 正解データ
# Amazon Foreast
# xgboost
df_xgb = pd.DataFrame(regressor.predict(test_x[test_x.client==21]), columns=['xgb'])
df_xgb
df_y = pd.DataFrame(test_y[test_y.client==21]['demand'], columns=['demand']).reset_index(drop=True)
df_y
# ### Foreastの結果
# +
#df_forecast = pd.read_csv('../amazon_forecast_official_dev_guide/export_forecast_drop1week/my_forecast_export_drop1week_2021-03-03T07-51-36Z_part1.csv')
df_forecast = pd.read_csv('../input/my_forecast_export_drop1week_2021-03-03T07-51-36Z_part1.csv')
# -
df_forecast['item_id'].unique()
df_result = df_forecast[df_forecast.item_id=='client_21'].reset_index(drop=True)
df_result
pd.concat([df_result, df_xgb, df_y], axis=1).plot(x='date',figsize=(20,5), grid=True)
# # XGBoostの利点
# 特徴量の重要度がわかる
#
# 【デメリット】
# コードのデバッグが大変
# 特徴量を作り込む必要あり
# ハイパーパラメータのチューニング
#
| AmazonForecast_vs_XGBoost/notebook/01_separate_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import networkx as nx
import matplotlib.pyplot as plt
import plotly.express as px
from tqdm import tqdm
import random
import pickle
with open("data/matrix.p", "rb") as f:
matrix = pickle.load(f)
with open("data/train-mix.p", "rb") as f:
data = pickle.load(f)
data
# +
id_to_pair = {}
for i in data:
id_to_pair[i[0]] = (i[1], i[2])
# -
df = pd.DataFrame()
link = [[i[1], i[2]] for i in data]
label = [i[3] for i in data]
df['Source'] = [i[1] for i in data]
df['Sink'] = [i[2] for i in data]
df['Label'] = [i[3] for i in data]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(link, label, test_size=0.1)
from features_generator import *
train_features = pd.DataFrame()
test_features = pd.DataFrame()
temp_train = []
for i in tqdm(X_train):
temp.append(common_neighbour(i[0], i[1], matrix))
train_features['c_nei'] = temp
# !open .
# +
from multiprocessing import Pool
def feature_generator(datum):
id, a, b, l = datum
# node feature
a_in = indegree(a, matrix)
a_out = outdegree(a, matrix)
b_in = indegree(b, matrix)
b_out = indegree(b, matrix)
# neighbouring feature
neighbour = common_neighbour(a, b, matrix)
jac = jaccard(neighbour, a, b, matrix)
dice = dice_idx(neighbour, a, b, matrix)
p_a = pref_attach(a, b, matrix)
cos = cosine_sim(neighbour, p_a)
lhn = LHN(neighbour, p_a)
adar = adamic_adar(a, b, matrix)
ra = resource_allocation(a, b, matrix)
reverse = reverse_link(a, b, matrix)
hp = hub_promoted(neighbour, a, b, matrix)
hd = hub_depressed(neighbour, a, b, matrix)
# path feature
#sim_r = sim_rank(a, b, matrix, 0)
flow2, flow3 = propflow3(a, b, matrix)
#print(flow)
#return flow
return [id,a_in,a_out,b_in,b_out,neighbour,jac,dice,p_a,cos,lhn,adar,reverse,hp,hd,flow2,flow3,l]
def logger(res):
train_test.append(res)
if len(train_test) % (len(data)//100) == 0:
print("{:.2%} done".format(len(train_test)/len(data)))
train_test = []
print("start")
pool = Pool(processes=4)
for item in data:
pool.apply_async(feature_generator, args=[item], callback=logger)
pool.close()
pool.join()
print("end")
train_test = np.array(train_test)
print(train_test.shape)
# -
len(sub_data)
# +
from multiprocessing import Pool
from features_generator import propflow3
def feature_generator(pair):
a, b = pair
flow2, flow3 = propflow3(a, b, matrix)
#print(flow)
#return flow
return [a, b, flow2,flow3]
def logger(res):
random_walk_sub[(res[0], res[1])] = (res[2], res[3])
if len(random_walk_sub) % (len(sub_data)//100) == 0:
print("{:.2%} done".format(len(random_walk_sub)/(len(sub_data))))
random_walk_sub = {}
print("start")
pool = Pool(processes=6)
for pair in sub_data.to_numpy():
pool.apply_async(feature_generator, args=[pair], callback=logger)
pool.close()
pool.join()
print("end")
print(len(random_walk_sub))
# -
with open("data/random_walk_sub", "wb") as f:
pickle.dump(random_walk_sub, f)
# +
# train_test.dump("data/data.npy")
# -
train_test = np.load("data/data.npy", allow_pickle=True)
random_walk_train = {}
for i in train_test[1:]:
pair = id_to_pair[i[0]]
# flow2, flow3
random_walk_train[pair] = [i[-2], i[-3]]
with open('data/random_walk_train.p', "wb") as f:
pickle.dump(random_walk_train, f)
len(data)
len(random_walk_train)
train_test_reduced = [i[5:] for i in train_test]
len(train_test_reduced)
X_train, X_test, y_train, y_test = train_test_split(train_test_reduced, label, test_size=0.1)
from sklearn.linear_model import LogisticRegression
lr_clf = LogisticRegression()
lr_clf.fit(X_train, y_train)
lr_clf.score(X_test, y_test)
# +
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import roc_auc_score
base = RandomForestClassifier(n_estimators=100)
parameters = {
"max_depth":[9,11,13,15,17],
"min_samples_leaf":[1,3,5],
"min_samples_split":[2,4,6,8,10],
}
model = GridSearchCV(base, parameters, n_jobs=-1)
model.fit(X_train, y_train)
print(model.score(X_test, y_test))
# +
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=200, learning_rate=0.5)
ada.fit(X_train, y_train)
print(roc_auc_score(y_test, np.squeeze(ada.predict_proba(X_test)[:,1])))
# +
from sklearn.preprocessing import Normalizer
scaler = Normalizer(norm='l1')
scaler.fit(X_train)
X_train = scaler.transform(X_train)
scaler.fit(X_test)
X_test = scaler.transform(X_test)
# -
# # Generate Rec Sim
# +
import numpy as np
from tqdm import tqdm
from sklearn.metrics.pairwise import cosine_similarity
import json
import pickle
train = {} # key: src value: [dest1, dest2, ...]
vector_count = {} # indegree count
print ("reading train set")
with open("train.txt") as trainfile:
for i, line in tqdm(enumerate(trainfile), position=0, leave=True):
line_list = [int(k) for k in line[:-1].split("\t")]
a = line_list[0]
train[a] = []
for b in line_list[1:]:
train[a].append(b)
vector_count[b] = vector_count.get(b,0)+1
train[a] = list(set(train[a]))
print ("--------complete")
print ("generating dictionary")
# -
import pandas as pd
def read_sub():
with open('test-public.txt', 'r') as f:
# skip the header
f.readline()
data = {'Source':[], 'Sink':[]}
for raw_line in f:
line = raw_line.strip().split("\t")
data['Source'].append(int(line[1]))
data['Sink'].append(int(line[2]))
return pd.DataFrame(data=data)
# generate new node set
# filter by indegree threshold
threshold = 10
new_set = set()
for i in vector_count:
if vector_count[i] > threshold:
new_set.add(i)
# add all source node
for i in train:
new_set.add(i)
with open("data/train-mix.p","rb") as f:
test = pickle.load(f)
# add all the node in testing to the new set
for _, j, k, _ in test:
new_set.add(j)
new_set.add(k)
sub_data = read_sub()
for source, sink in tqdm(sub_data.to_numpy(), position=0, leave=True):
new_set.add(source)
new_set.add(sink)
# +
sources_vector = list(train.keys())
source_to_index = {}
for i, j in enumerate(sources_vector):
source_to_index[j] = i
# -
from copy import deepcopy
new_train = deepcopy(train)
# remvoe existing link to prevent overfitting
for _, j, k, label in tqdm(test, position=0, leave=True):
try:
if label == 1:
new_train[j].remove(k)
except:
print(j, k)
# +
sink_source = {}
for source in tqdm(sources_vector, position=0, leave=True):
sinks = new_train[source]
for sink in sinks:
if sink in new_set:
value = sink_source.get(sink, set())
value.add(source_to_index[source])
sink_source[sink] = value
# -
from scipy.sparse import csr_matrix
# +
sink_to_vec = {}
for sink in tqdm(sink_source.keys(), position=0, leave=True):
vec = np.zeros(20000)
vec[list(sink_source[sink])] = 1/len(sink_source[sink])
sink_to_vec[sink] = csr_matrix(vec)
# +
# source_to_vec = {}
# for source in tqdm(sources_vector, position=0, leave=True):
# vec = csr_matrix(np.zeros(20000))
# counter = 0
# for sink in new_train[source]:
# if sink not in new_set:
# continue
# vec += sink_to_vec[sink]
# counter += 1
# source_to_vec[source] = vec / max(counter, 1)
# -
test_source = set()
for _, source, _, _ in test:
test_source.add(source)
# +
test_source_to_vec = {}
for source in tqdm(test_source, position=0, leave=True):
vec = csr_matrix(np.zeros(20000))
counter = 0
for sink in new_train[source]:
if sink not in new_set:
continue
vec += sink_to_vec[sink]
counter += 1
# add self reference
if source in sink_to_vec:
vec += sink_to_vec[source]
counter += 1
test_source_to_vec[source] = vec / max(counter, 1)
# -
with open("data/test_source_to_vec_add_self_t10.p", "wb") as f:
pickle.dump(test_source_to_vec, f)
# +
cosin_dict = {}
for _, source, sink, _ in tqdm(test, position=0, leave=True):
source_vec = test_source_to_vec[source]
sink_vec = sink_to_vec.get(sink, np.array([np.zeros(20000)]))
sim = cosine_similarity(X=source_vec, Y=sink_vec)[0][0]
cosin_dict[(source, sink)] = sim
# -
with open("data/cosin_dict_add_self_t10.p", "wb") as f:
pickle.dump(cosin_dict, f)
# ### Creating features for submission data
# +
# TODO : add whole training graph source here
sub_sink_source = {}
for source in tqdm(train.keys(), position=0, leave=True):
sinks = train[source]
for sink in sinks:
if sink in new_set:
value = sub_sink_source.get(sink, set())
value.add(source_to_index[source])
sub_sink_source[sink] = value
# +
# sub_sink_to_vec = {}
# for sink in tqdm(sub_sink_source.keys(), position=0, leave=True):
# vec = np.zeros(20000)
# vec[list(sub_sink_source[sink])] = 1/len(sub_sink_source[sink])
# sub_sink_to_vec[sink] = csr_matrix(vec)
# +
sub_sink_to_vec = {}
def sub_sink_to_vec_func(sink):
if sink in sub_sink_to_vec:
return sub_sink_to_vec[sink]
vec = np.zeros(20000)
if sink in sub_sink_source:
vec[list(sub_sink_source[sink])] = 1/len(sub_sink_source[sink])
else:
print(sink, "not in sink source")
res = csr_matrix(vec)
sub_sink_to_vec[sink] = res
return res
# +
sub_source_to_vec = {}
for source in tqdm(sub_data['Source'], position=0, leave=True):
vec = csr_matrix(np.zeros(20000))
counter = 0
for sink in train[source]:
if sink not in new_set:
continue
vec += sub_sink_to_vec_func(sink)
counter += 1
# add self reference
if source in sub_sink_source:
vec += sub_sink_to_vec_func(source)
counter += 1
sub_source_to_vec[source] = vec / max(counter, 1)
# +
sub_cosin_dict = {}
for source, sink in tqdm(sub_data.to_numpy(), position=0, leave=True):
source_vec = sub_source_to_vec[source]
# sink_vec = sub_sink_to_vec.get(sink, np.array([np.zeros(20000)]))
sink_vec = sub_sink_to_vec_func(sink)
sim = cosine_similarity(X=source_vec, Y=sink_vec)[0][0]
sub_cosin_dict[(source, sink)] = sim
# -
sub_cosin_dict
with open("data/sub_cosin_dict_add_sm_t10.p", "wb") as f:
pickle.dump(sub_cosin_dict, f)
# !open .
labels = np.array([i[-1] for i in test])
labels.reshape(-1,1)
# +
from sklearn.ensemble import AdaBoostClassifier
ada = AdaBoostClassifier(n_estimators=200, learning_rate=0.5)
ada.fit(,)
# -
print(roc_auc_score(labels[-2000:], np.squeeze(ada.predict_proba([[i] for i in test_with_cosin_feature[-2000:]])[:,1])))
# +
from sklearn.model_selection import GridSearchCV
base = RandomForestClassifier(n_estimators=100)
parameters = {
"max_depth":[5,7,9,11,13],
"min_samples_leaf":[1,3,5],
"min_samples_split":[2,4,6,8,10],
"max_features":["sqrt", "log2"]
}
rf_grid = GridSearchCV(base, parameters, n_jobs=-1)
rf_grid.fit([[i] for i in test_with_cosin_feature[:-2000]], labels[:-2000])
print(rf_grid.score([[i] for i in test_with_cosin_feature[-2000:]], labels[-2000:]))
# -
# # Sink-wise modelling
# +
id2v = list(new_set) # [v1, v2, ...]
v2id = {} # key: node value: index of the node in id2v
for i, j in enumerate(id2v):
v2id[j] = i
print ("length of new set:")
print (len(new_set))
# -
test[0]
# +
# generate new node id dictionary
new_train = {} # key: index value: set of connected nodes after filtering
for i in train:
# i is source node, train[i] is sink node from i-th source node
# select sink node that in our new_set
new_train[v2id[i]] = set([v2id[j] for j in train[i] if j in new_set])
new_test = {} # key: training sample id (old id) value: [new id for source, id for sink]
for i, j, k,_ in test:
# i is id, j is source, k is sink
new_test[i] = [v2id[j], v2id[k]]
# remove true edge
if v2id[k] in new_train[v2id[j]]:
new_train[v2id[j]].remove(v2id[k])
# +
tA = new_train.copy()
tB = {}
for i in new_train:
if i not in tA[i]:
# add self connection for each node
tA[i].add(i)
for j in new_train[i]:
tB[j] = tB.get(j,set([]))
tB[j].add(i)
print ("now processing...")
# -
len(tA.keys())
def sim(pair, tA, tB, l):
vi, vj = pair
tempA = np.zeros(l)
tempB = np.zeros(l)
tempA[list(tA[vi])] = 1/len(tA[vi])
if vj in tB:
for i in tB[vj]:
tempB[list(tA[i])] += 1/len(tB[vj])/len(tA[i])
return cosine_similarity([tempA, tempB])[0][1]
#return tempA,tempB
len(new_test.keys())
# +
l = len(new_set)
res = {}
for i in tqdm(new_test, ascii=True):
source, sink = new_test[i]
res[i] = []
res[i].append(sim([source, sink], tA, tB, l))
# res[i].append(sim([vj,vi], tB, tA, l))
with open("data/rec_sim.pickle","wb") as f:
pickle.dump(res, f)
# -
| Rec Cosin Sim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rationale
# EDA of ATC ontology to other ontology mappings. Main conclusions are: there are lot's of exact duplicates(same source, same classes mapped, probably we are missing some annotation of the mappings here), and quite some different-source duplicates (different source, same classes mapped, this is sort of expected). There are only 2 relevant sources (CUI & LOOM).
# +
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from zib_uploader.fetch_data import ClassMappingsFetcher
# -
sns.set_style()
# ## Fetch data or load from cache
mappings_filepath = 'mappings.csv'
try:
mappings = pd.read_csv(mappings_filepath)
except FileNotFoundError:
fetcher = ClassMappingsFetcher()
data = fetcher.fetch()
mappings = pd.DataFrame({'source': mapping.source,
'atc_class': mapping.classes[0],
'mapped_class': mapping.classes[1]}
for mapping in data)
mappings.to_csv('mappings.csv', index=False)
mappings.head()
# ## Deduct mapped ontology
def get_ontology_name(uri):
return '/'.join(uri.split('/')[:-1])
mappings['mapped_ontology_name'] = mappings['mapped_class'].apply(get_ontology_name)
mappings.head()
# ## Poop out some statistics
mappings_no_duplicates = mappings.drop_duplicates()
# ### Number of mappings per ATC class
print(f'{len(mappings)} mappings for {len(mappings["atc_class"].unique())} ATC classes')
print(f'{len(mappings)-len(mappings_no_duplicates)} duplicate mappings (this is kind of weird)')
sns.distplot(mappings['atc_class'].value_counts(), hist=True, kde=False)
plt.xlabel('Number of mappings per class')
plt.title('With duplicates')
y = plt.ylabel('Frequency')
sns.distplot(mappings_no_duplicates['atc_class'].value_counts(), hist=True, kde=False)
plt.xlabel('Number of mappings per class')
plt.title('Without duplicates')
y = plt.ylabel('Frequency')
# ### Number of mappings per source
mappings_no_duplicates['source'].value_counts()
# ### Number of mappings per ontology
def source_counts(df):
result_df = pd.DataFrame(df['mapped_ontology_name'].value_counts())
result_df.columns = ['total_count']
for source in df['source'].unique():
source_df = pd.DataFrame(df[df['source'] == source]['mapped_ontology_name'].value_counts())
source_df.columns = [f'{source}_count']
result_df = result_df.merge(source_df, how='left',left_index=True, right_index=True)
return result_df.fillna(0).astype(int)
source_counts(mappings_no_duplicates).head(50)
# ### Note that all mappings from source SAME_URI are for the http://purl.bioontology.org/ontology/STY ontology. These are semantic types, not medicines.
mappings_no_duplicates[mappings_no_duplicates['mapped_ontology_name'] == 'http://purl.bioontology.org/ontology/STY']
# ### For some mappings both CUI and LOOM are stated as a source
multiple_sources = mappings_no_duplicates[mappings_no_duplicates.duplicated(subset=['atc_class', 'mapped_class'], keep=False)]
multiple_sources.sort_values(['atc_class', 'mapped_class'])
# ### Some classes in the ATC ontology are mapped to multiple classes in other ontologies
# An example is 'silicones' in ATC which is mapped to 'Silicone' and 'Silicone-containing product' in Snowmed
# Below are some more examples for snowmed
snomed_df = mappings_no_duplicates[mappings_no_duplicates['mapped_ontology_name'] == 'http://purl.bioontology.org/ontology/SNOMEDCT']
snomed_df = snomed_df.drop(columns=['mapped_ontology_name'])
pd.set_option('display.max_colwidth', None)
snomed_df[snomed_df.duplicated(subset=['atc_class', 'source'], keep=False)].head(30)
| notebooks/eda-atc-ontology-mappings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'> <img src='../Pierian_Data_Logo.png' /></a>
# ___
# # NumPy Exercises
#
# Now that we've learned about NumPy let's test your knowledge. We'll start off with a few simple tasks, and then you'll be asked some more complicated questions.
# ## <NAME> worked exercises, Udemy.com PyDSML, 3/27/2019.
# #### Import NumPy as np
import numpy as np
# #### Create an array of 10 zeros
np.zeros(10)
# #### Create an array of 10 ones
np.ones(10)
# #### Create an array of 10 fives
np.ones(10)*5
# #### Create an array of the integers from 10 to 50
np.arange(10, 51)
# #### Create an array of all the even integers from 10 to 50
np.arange(10, 51, 2)
# #### Create a 3x3 matrix with values ranging from 0 to 8
mat = np.arange(0, 9)
mat.reshape(3, 3)
# #### Create a 3x3 identity matrix
np.eye(3)
np.eye(3, 3)
# #### Use NumPy to generate a random number between 0 and 1
np.random.rand(1)
# np.random.rand(d0, d1, d2, ...) selects a random number from [0, 1) range. Paramters are dimensions.]
help(np.random.rand)
# #### Use NumPy to generate an array of 25 random numbers sampled from a standard normal distribution
np.random.rand(25)
np.random.rand(25)
# #### Create the following matrix:
np.arange(0.01, 1.01, .01).reshape(10, 10)
# #### Create an array of 20 linearly spaced points between 0 and 1:
np.linspace(0, 1, 20)
np.linspace(0, 1, 20, endpoint=True)
# linspace returns an array of evenly spaced numbers. (start, spop, numbers, default include endpoint=True)
# ## Numpy Indexing and Selection
#
# Now you will be given a few matrices, and be asked to replicate the resulting matrix outputs:
mat = np.arange(1,26).reshape(5,5)
mat
# +
# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW
# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T
# BE ABLE TO SEE THE OUTPUT ANY MORE
# -
mat[2:6, 1:6]
# Indexing is same as list indexing.
# Try negative indexing.
mat[-3:, -4:]
# Negative indexing. Always counts from top-left. [Row start:end, Col start:end]
mat[:-3, :-4]
# +
# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW
# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T
# BE ABLE TO SEE THE OUTPUT ANY MORE
# -
mat[3, -1]
# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW
# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T
# BE ABLE TO SEE THE OUTPUT ANY MORE
mat
# mat 2nd column, 0 to 2 rows.
mat[:3, 1].reshape(3, 1)
# +
# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW
# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T
# BE ABLE TO SEE THE OUTPUT ANY MORE
# -
mat[-1, :]
# +
# WRITE CODE HERE THAT REPRODUCES THE OUTPUT OF THE CELL BELOW
# BE CAREFUL NOT TO RUN THE CELL BELOW, OTHERWISE YOU WON'T
# BE ABLE TO SEE THE OUTPUT ANY MORE
# -
mat[-2:, :] # Last two rows, all columns.
# ### Now do the following
# #### Get the sum of all the values in mat
mat.sum() # Method applied to mat object?
# mat is an ndarray object, a built-in numpy class.
# This class object has a method .sum() and .std().
# #### Get the standard deviation of the values in mat
mat.std()
# #### Get the sum of all the columns in mat
sum(mat)
# sum() function applied to mat ndarray object produce column sums by default.
sum(mat[:3, -2:])
# [row 0 to 2, col 4 to 5]
## mat.sum(axis=0)
# Columns axis=0, not 1.
x = mat.sum(axis=0)
y = mat.sum(axis=1)
print(x, y)
# # Great Job!
| udemy-ds-bc/Py_DS_ML_bootcamp/00-my-practice/02-Python-for-Data-Analysis-NumPy/numpy_ex1_JY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="YUV3eJoaHC-T"
# 
# + [markdown] id="e1ch_tb5HIOM"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/18.Chunk_Sentence_Splitter.ipynb)
# + [markdown] id="gNLwhcTeHT01"
# # Chunk Sentence Splitter
# We are releasing `ChunkSentenceSplitter` annotator that splits documents or sentences by chunks provided. Splitted parts can be named with the splitting chunks. <br/>
# By using this annotator, you can do some tasks like splitting clinical documents according into sections in accordance with CDA (Clinical Document Architecture).
# + [markdown] id="sdg3wYL6HXZj"
# ## Colab Setup
# + colab={"base_uri": "https://localhost:8080/", "height": 73, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": ""}}} executionInfo={"elapsed": 25965, "status": "ok", "timestamp": 1639491635600, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="Gf8bIwegE7lb" outputId="e378d6dc-09f6-4efe-c087-bdebab3551ef"
import json
import os
from google.colab import files
license_keys = files.upload()
with open(list(license_keys.keys())[0]) as f:
license_keys = json.load(f)
# Defining license key-value pairs as local variables
locals().update(license_keys)
# Adding license key-value pairs to environment variables
os.environ.update(license_keys)
# + executionInfo={"elapsed": 8513, "status": "ok", "timestamp": 1639491648012, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="J2fSZPxNJJKt"
# %%capture
# Installing pyspark and spark-nlp
# ! pip install --upgrade -q pyspark==3.1.2 spark-nlp==$PUBLIC_VERSION
# Installing Spark NLP Healthcare
# ! pip install --upgrade -q spark-nlp-jsl==$JSL_VERSION --extra-index-url https://pypi.johnsnowlabs.com/$SECRET
# + colab={"base_uri": "https://localhost:8080/", "height": 267} executionInfo={"elapsed": 248, "status": "ok", "timestamp": 1639491651717, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="fqL744xQHlwf" outputId="a124da15-27c1-4403-d7ff-78ccfa81b7e6"
import json
import os
from pyspark.ml import Pipeline, PipelineModel
from pyspark.sql import SparkSession
from pyspark.sql import functions as F
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
from sparknlp.base import *
import sparknlp_jsl
import sparknlp
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
params = {"spark.driver.memory":"16G",
"spark.kryoserializer.buffer.max":"2000M",
"spark.driver.maxResultSize":"2000M"}
print ("Spark NLP Version :", sparknlp.version())
print ("Spark NLP_JSL Version :", sparknlp_jsl.version())
spark = sparknlp_jsl.start(license_keys['SECRET'],params=params)
spark
# + [markdown] id="WyFoYZ25Hjwr"
# ## How It Works
#
# + id="OEzQVvAWFl9A"
#giving input chunks to the ChunkSentenceSplitter model by using regex
regex = """Reporting Template,title1
SPECIMEN,title2
RESULTS,title3"""
with open("title_regex.txt", 'w') as f:
f.write(regex)
# + id="mS62MMfgFl6o"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
regexMatcher = RegexMatcher()\
.setExternalRules("/content/title_regex.txt", ",")\
.setInputCols("document")\
.setOutputCol("chunks")
pipeline = Pipeline().setStages([
documentAssembler,
regexMatcher])
text_list = ["""
This is the header that have not title
Reporting Template
Writers write descriptive paragraphs because their purpose is to describe something. Their point is that something
is beautiful or disgusting or strangely intriguing.
Writers write persuasive and argument paragraphs because their purpose is to persuade or convince someone. T
SPECIMEN
+Adequacy of Sample for Testing
___ Adequate
+Estimated % Tumor Cellularity
___ Suboptimal (explain): _________________
RESULTS
+Mutational Analysis
___ Mutation detected
___ Mutation no identified
___ EGFR
"""]
data_chunk = spark.createDataFrame([["text"]]).toDF("text")
pipeline_model = pipeline.fit(data_chunk)
chunk_df = pipeline_model.transform(spark.createDataFrame(pd.DataFrame({'text': text_list})))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 4474, "status": "ok", "timestamp": 1639488346008, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="xNIZ4_dHFl3_" outputId="4b11c6cc-6413-43cc-eaae-178301006906"
chunk_df.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1353, "status": "ok", "timestamp": 1639488348624, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="mNp3BvxaO_LN" outputId="18bd3bd7-9fdd-4d3c-d505-0e4bb4ad8430"
chunk_df.selectExpr('explode(chunks)').show(truncate=False)
# + [markdown] id="feW0o5ta5XNI"
# Applying `ChunkSentenceSplitter()`
# + id="rIIg7qqsNpTS"
chunkSentenceSplitter = ChunkSentenceSplitter()\
.setInputCols("chunks","document")\
.setOutputCol("paragraphs")
paragraphs = chunkSentenceSplitter.transform(chunk_df)
# + colab={"base_uri": "https://localhost:8080/", "height": 173} executionInfo={"elapsed": 1384, "status": "ok", "timestamp": 1639488354955, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="Wi3PJLe4NuZs" outputId="1215021c-8aed-4ab2-fbd4-9db4060f99d5"
paragraphs.selectExpr("explode(paragraphs) as result").selectExpr("result.result","result.metadata.entity").toPandas()
# + [markdown] id="y2ukghZzH5lr"
# ### Ner Pipeline with Sentence Splitting
# + executionInfo={"elapsed": 247, "status": "ok", "timestamp": 1639491663923, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="_9vSrHwVM-bE"
#input data
input_list = ["""Sample Name: Mesothelioma - Pleural Biopsy
Description: Right pleural effusion and suspected malignant mesothelioma. (Medical Transcription Sample Report)
PREOPERATIVE DIAGNOSIS: Right pleural effusion and suspected malignant mesothelioma.
POSTOPERATIVE DIAGNOSIS: Right pleural effusion, suspected malignant mesothelioma.
ANESTHESIA: General double-lumen endotracheal.
DESCRIPTION OF FINDINGS: Right pleural effusion, firm nodules, diffuse scattered throughout the right pleura and diaphragmatic surface.
SPECIMEN: Pleural biopsies for pathology and microbiology.
INDICATIONS: Briefly, this is a 66-year-old gentleman who has been transferred from an outside hospital after a pleural effusion had been drained and biopsies taken from the right chest that were thought to be consistent with mesothelioma. Upon transfer, he had a right pleural effusion demonstrated on x-ray as well as some shortness of breath and dyspnea on exertion. The risks, benefits, and alternatives to right VATS pleurodesis and pleural biopsy were discussed with the patient and his family and they wished to proceed.
Dr. X was present for the entire procedure which was right VATS pleurodesis and pleural biopsies.The counts were correct x2 at the end of the case."""]
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 663, "status": "ok", "timestamp": 1639491665544, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="fIFfrkyVJ6Ba" outputId="22f331f0-b873-4608-90d8-cbe411afb3c7"
files = [f"{i}.txt" for i in (range(1, len(input_list)+1))]
df = spark.createDataFrame(pd.DataFrame({'text': input_list, 'file' : files}))
df.show()
# + [markdown] id="AQ0B-fo_KEYV"
# Now, creating NER pipeline for extracting chunks
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 8781, "status": "ok", "timestamp": 1639491685337, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="zRqO5n4uNuXA" outputId="051bb62c-113c-4e40-e78b-cafadc4be4ee"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentenceDetector = SentenceDetectorDLModel.pretrained("sentence_detector_dl_healthcare","en","clinical/models")\
.setInputCols(["document"])\
.setOutputCol("sentence")
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")\
word_embeddings = WordEmbeddingsModel.pretrained("embeddings_clinical", "en", "clinical/models")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
clinical_ner = MedicalNerModel.pretrained("ner_jsl_slim", "en", "clinical/models") \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "ner"]) \
.setOutputCol("ner_chunk")\
.setWhiteList(["Header"])
pipeline_sentence = Pipeline(
stages = [
documentAssembler,
sentenceDetector,
tokenizer,
word_embeddings,
clinical_ner,
ner_converter
])
empty_df = spark.createDataFrame([[""]]).toDF('text')
pipeline_model_sentence = pipeline_sentence.fit(empty_df)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2356, "status": "ok", "timestamp": 1639491704187, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="Yx8ARe0zNNEe" outputId="5e6fa79b-d69d-48e5-cb06-5a74f2ef4cd3"
result = pipeline_model_sentence.transform(df)
result.selectExpr('explode(ner_chunk)').show(truncate=False)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 260, "status": "ok", "timestamp": 1639491706039, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="poV6TdY0J3n4" outputId="2f48a220-985a-4370-efc1-e55613fa08df"
result.columns
# + executionInfo={"elapsed": 260, "status": "ok", "timestamp": 1639491709608, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="8ybHSZaQJ3k-"
#applying ChunkSentenceSplitter
chunkSentenceSplitter = ChunkSentenceSplitter()\
.setInputCols("document","ner_chunk")\
.setOutputCol("paragraphs")\
.setGroupBySentences(False)
paragraphs = chunkSentenceSplitter.transform(result)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1646, "status": "ok", "timestamp": 1639491712203, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="2mcZx6LJJ3iL" outputId="5c26724f-f983-45f6-e749-9d3d5ad5c66a"
paragraphs.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2328, "status": "ok", "timestamp": 1639491721281, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="VR3wH1kLO0pU" outputId="bc6cbc11-d231-43bb-991e-c205bbba594a"
paragraphs.select("paragraphs.result").show(truncate=100)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"elapsed": 2029, "status": "ok", "timestamp": 1639491728694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="7YEy6236PkpI" outputId="7260a9c2-e9cc-4f2d-abb7-c3567eadbe99"
pd.set_option('display.max_colwidth', None)
result_df = paragraphs.selectExpr("explode(paragraphs) as result").selectExpr("result.result","result.metadata.entity").toPandas()
result_df.head()
# + [markdown] id="mfPboKgfRkRy"
# ### Ner Pipeline without Sentence Splitter
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3293, "status": "ok", "timestamp": 1639491735404, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="vIINLRRUPkdH" outputId="77db0b44-f28e-478a-9c8c-fbd9d22a72e8"
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
#sentenceDetector = SentenceDetectorDLModel.pretrained("sentence_detector_dl_healthcare","en","clinical/models")\
# .setInputCols(["document"])\
# .setOutputCol("sentence")
tokenizer = Tokenizer()\
.setInputCols(["document"])\
.setOutputCol("token")\
tokenClassifier = BertForTokenClassification.pretrained("bert_token_classifier_ner_jsl_slim", "en", "clinical/models")\
.setInputCols("token", "document")\
.setOutputCol("ner")\
.setCaseSensitive(True)
ner_converter = NerConverter() \
.setInputCols(["document", "token", "ner"]) \
.setOutputCol("ner_chunk")\
.setWhiteList(["Header"])
pipeline = Pipeline(
stages = [
documentAssembler,
tokenizer,
tokenClassifier,
ner_converter
])
empty_df = spark.createDataFrame([[""]]).toDF('text')
pipeline_model = pipeline.fit(empty_df)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2111, "status": "ok", "timestamp": 1639491742634, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="YnfxFzC0zazx" outputId="f3ee4134-f726-496a-e961-cec1a36fe5a2"
result = pipeline_model.transform(df)
result.selectExpr('explode(ner_chunk)').show(truncate=False)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 251, "status": "ok", "timestamp": 1639491744119, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="u7KlfLavTBut" outputId="e0558aa3-8690-4eb9-a2cc-6fe6549f030d"
result.columns #no sentence column
# + executionInfo={"elapsed": 243, "status": "ok", "timestamp": 1639491747550, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="14Ms3D27O0j4"
#applying ChunkSentenceSplitter
chunkSentenceSplitter = ChunkSentenceSplitter()\
.setInputCols("ner_chunk","document")\
.setOutputCol("paragraphs")\
paragraphs = chunkSentenceSplitter.transform(result)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 1868, "status": "ok", "timestamp": 1639491753394, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="GlMF_Q8_NnNb" outputId="f2ba3afd-7825-4f73-9952-c8ea8f9661da"
paragraphs.show()
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2681, "status": "ok", "timestamp": 1639491763463, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="Bmo5ghIlPavC" outputId="bc5d06bb-9d79-48eb-c0d8-e06d299a2636"
paragraphs.select("paragraphs.result").show(truncate=100)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"elapsed": 2630, "status": "ok", "timestamp": 1639491770945, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="34Y2bJ94Pape" outputId="411d20ea-d8ab-4c82-a8cc-d9b6748fbc85"
result_df = paragraphs.selectExpr("explode(paragraphs) as result").selectExpr("result.result","result.metadata.entity", "result.metadata.splitter_chunk").toPandas()
result_df.head()
# + [markdown] id="bl3WOfMnT1D2"
# `.setInsertChunk()` parameter to set whether remove chunks from splitted parts or not.
# + executionInfo={"elapsed": 250, "status": "ok", "timestamp": 1639491772759, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="NMDiBa3PK57R"
chunkSentenceSplitter = ChunkSentenceSplitter()\
.setInputCols("ner_chunk","document")\
.setOutputCol("paragraphs")\
.setInsertChunk(False)
paragraphs = chunkSentenceSplitter.transform(result)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 2556, "status": "ok", "timestamp": 1639491781901, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="VlPzTyevK52L" outputId="72c5e63a-1751-4195-c997-578d446f43ca"
paragraphs.select("paragraphs.result").show(truncate=100)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"elapsed": 2728, "status": "ok", "timestamp": 1639491787596, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="-rpSK8WtK5xQ" outputId="b250c88d-7c1d-42bd-94a1-fe2c91bcb7af"
result_insert = paragraphs.selectExpr("explode(paragraphs) as result").selectExpr("result.result","result.metadata.entity", "result.metadata.splitter_chunk").toPandas()
result_insert.head()
# + [markdown] id="0r16mxmy3yon"
# Check how `.setInsertChunk(True)` affects the result
# + colab={"base_uri": "https://localhost:8080/", "height": 204} executionInfo={"elapsed": 2791, "status": "ok", "timestamp": 1639491794114, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "14855809472179427810"}, "user_tz": -180} id="V3hx_MQs3ezD" outputId="228f35fd-9081-4b73-98a2-dc76e1d10b5e"
chunkSentenceSplitter_2 = ChunkSentenceSplitter()\
.setInputCols("ner_chunk","document")\
.setOutputCol("paragraphs")\
.setInsertChunk(True)\
.setDefaultEntity("Intro") #to set the name of the introduction entity
paragraphs = chunkSentenceSplitter_2.transform(result)
result = paragraphs.selectExpr("explode(paragraphs) as result").selectExpr("result.result","result.metadata.entity", "result.metadata.splitter_chunk").toPandas()
result.head()
# + id="4bAP9GGQz2SU"
| tutorials/Certification_Trainings/Healthcare/18.Chunk_Sentence_Splitter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2021 June Embed
# ## Imports
# +
import sys
import delphi.plotter as dp
from delphi.cpp.DelphiPython import AnalysisGraph, InitialBeta, InitialDerivative
import pandas as pd
import numpy as np
#import graphviz
#from IPython.display import display
from IPython.display import Image
import warnings
warnings.filterwarnings('ignore')
# -
# ## Read Data
# +
df_temp = pd.read_csv('../../data/mini_use_case/TerraClimateOromiaMonthlyMaxTemp.csv')
temperature = df_temp['(deg C) Max Temperature (TerraClimate) at State, 1958-01-01 to 2019-12-31'].tolist()[:72]
df_rain = pd.read_csv('../../data/mini_use_case/TerraClimateOromiaMontlhyPrecip.csv')
rain = df_rain['(mm) Precipitation (TerraClimate) at State, 1958-01-01 to 2019-12-31'].tolist()[:72]
# -
# ## Create Base CAG
# +
statements = [
(
("", 1, "rain"),
("", -1, "temperature"),
)
]
data = {"rain": ("Monthly Precipitation (mm)", rain),
"temperature": ("Monthly Max Temperature (F)", temperature)
}
G = AnalysisGraph.from_causal_fragments_with_data((statements, data), kde_kernels=1000)
G.to_png('2021_june_embed.png', rankdir="TB", simplified_labels=False)
Image('2021_june_embed.png')
# -
# ## Train Model
# ### Head node modeling related parameteres
# <ol>
# <li>concept_periods</li>
# <ul>
# <li>Period for each head concept.</li>
# <li>Default is 1.</li>
# <li>A dictionary: {'concept name': period}</li>
# </ul>
# <li>concept_center_measures</li>
# <ul>
# <li>How to calculate the central tendency for each head concept.</li>
# <li>mean or median.</li>
# <li>Default is median.</li>
# </ul>
#
# <li>concept_models</li>
# <ul>
# <li>The model to be used for the head concept.</li>
# <li>The choises are:</li>
# <ol>
# <li>center - Always predicts the center value for each partition.</li>
# <--li>absolute_change - Predicts based on the absolute change between adjacent partiton centers. $ac_{i} = center_{i+1} - center_i$</li-->
# <--li>relative_change - Predicts based on the absolute change between adjacent partiton centers.</li-->
# </ol>
# </ul>
#
# </ol>
def train_predict_plot(G, period, center, model, constraints={}, concept_min_vals=-10000, concept_max_vals=1000):
# Training
G.run_train_model(res=70,
burn=0,
initial_beta=InitialBeta.ZERO,
initial_derivative=InitialDerivative.DERI_ZERO,
use_heuristic=False,
use_continuous=False,
train_start_timestep=0,
train_timesteps=48,
concept_periods={'rain': period},
concept_center_measures={'rain': center}, # mean, median
concept_models={'rain': model}, # center, absolute_change, relative_change
concept_min_vals={'rain': concept_min_vals},
concept_max_vals={'rain': concept_max_vals}
)
# Predicting
G.generate_prediction(49, 23, constraints=constraints)
# Plotting
model_state = G.get_complete_state()
dp.delphi_plotter(model_state, num_bins=400, rotation=45,
out_dir='plots', file_name_prefix='', save_csv=False)
# #### Using median
train_predict_plot(G, period=12, center='median', model='center')
Image('plots/8_Predictions_Median_and_CI_Monthly Precipitation (mm).png')
# #### Using mean
train_predict_plot(G, period=12, center='mean', model='center')
Image('plots/8_Predictions_Median_and_CI_Monthly Precipitation (mm).png')
# #### Let us try a different period
train_predict_plot(G, period=24, center='median', model='center')
Image('plots/8_Predictions_Median_and_CI_Monthly Precipitation (mm).png')
# ### Constraints
#
# <p>Three model options, center, absolute_change and relative_change behave differently upon constraints.</p>
#
# #### center with constraints
train_predict_plot(G, period=12, center='median', model='center', constraints={12: [('rain', 'Monthly Precipitation (mm)', 120)]})
Image('plots/8_Predictions_Median_and_CI_Monthly Precipitation (mm).png')
# ### Guiding the modeler with bounds
train_predict_plot(G, period=12, center='median', model='absolute_change', constraints={7: [('rain', 'Monthly Precipitation (mm)', 15)]})
Image('plots/8_Predictions_Median_and_CI_Monthly Precipitation (mm).png')
train_predict_plot(G, period=12, center='median', model='absolute_change', constraints={7: [('rain', 'Monthly Precipitation (mm)', 15)]}, concept_min_vals=0)
Image('plots/8_Predictions_Median_and_CI_Monthly Precipitation (mm).png')
| notebooks/2021_june_embed/2021_june_embed.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Today concepts**
# * Dimensionality Reduction
# * Applying PCA
# * manipulating categorical data with
# * LabelEncoder
# * OneHotEncoder
# * get_dummies
# * Saving/Reading ML model for future use
# * For Awareness we connecting ML model with Web application(Flask)
# **Dimensionality Reduction**
#
# Reduction of the dimension of the features space is called **Dimensionality Reduction**
# **In this dimensionality Reduction**
#
# 1. Feature Elimination
# 2. Feature Extraction
# **Feature Elimination**
#
# means Removing less important columns/features and taking only remaining featurres is called the feature elimination
# **Feature Extraction**
#
# we create new independent features/variable,where each new variable is a combination of each of the old columns**
# **simple example for feature extraction**
# if we have **number of production** and **price of product** as features then we can tranform these features into single feature as **total_price**
# **Advantage**
#
# by combining the features you can not loose information
#
#
# by reducing the features your model able to work well
# **Principal Component Analysis**
# it is the technique for feature extraction
#
#
#
# each of the new variable after PCA are all independent to each other
# **Now we apply PCA for breast cancer data**
import pandas as pd
import matplotlib.pyplot as plt
# **Get the data**
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
cancer.keys()
df = pd.DataFrame(cancer['data'])
df.columns = cancer['feature_names']
df['target'] = cancer['target']
df.head()
df.shape # 569 rows and 31 columns
df.isna().sum()
# **To apply PCA before that we have to standardize the data**
# Example salary range 10000 to 90000
# purchase price 1000 5000 to convert into same scale
#
# **step-1** import standardscaler
from sklearn.preprocessing import StandardScaler
# **step-2** create object
scaler = StandardScaler()
# **step-3** transform the data
#
# scaler.fit_transform(Features)
X = df.drop('target',axis=1)
X.head()
X_transformed = scaler.fit_transform(X)
# **we apply PCA**
from sklearn.decomposition import PCA
# **we have to tell number of principal components/new columns required**
pcaObj = PCA(n_components=2)
newFeatures = pcaObj.fit_transform(X_transformed)
pca_df = pd.DataFrame(newFeatures,columns=['PCA1','PCA2'])
pca_df.head()
# **How much information carried from actual features(30 features) to new features(2 pca components)**
# **explained_variance_ration**
pcaObj.explained_variance_ratio_*100
# **Visualisation**
pca_df.head(2)
pca_df['target'] = df['target']
pca_df.head(2)
# **separate the cancer and non cancer data**
mask1 = pca_df['target'] == 1
mask2 = pca_df['target'] == 0
hasCancer = pca_df[mask1]
hasNoCancer = pca_df[mask2]
hasCancer.head()
# +
plt.figure()
plt.xlabel('PCA1')
plt.ylabel('PCA2')
plt.title('visualisation of breast cancer with PCA')
plt.scatter(hasCancer['PCA1'] ,hasCancer['PCA2'],c='red',label='cancer')
plt.scatter(hasNoCancer['PCA1'] ,hasNoCancer['PCA2'],c='green',label='Not harm')
plt.legend()
plt.show()
# -
# **We can observe how much accuracy without PCA**
y = df['target']
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X,y)
print('score',model.score(X,y)*100)
# **we can observe how much accuracy with PCA**
pca_df.head()
X = pca_df.drop('target',axis=1)
y = pca_df['target']
model = LogisticRegression()
model.fit(X,y)
print('Score',model.score(X,y)* 100)
# **We are getting approximatly same accuracy even we use 2 new column for model**
# **manipulating the categorical data**
df = pd.read_csv('fruit_data_with_colours.csv')
df.drop('fruit_label',axis=1,inplace=True)
df.head(2)
# **unique items in fruit_name target**
df['fruit_name'].unique()
# **apple -> 0,mandrain-1,orange-2 lemon->3**
# **LabelEncoder**
# **step-1** import the label encoder
from sklearn.preprocessing import LabelEncoder
# **step-2** create object
le = LabelEncoder()
# **step-3** specifiy the unique items
# by fit()
le.fit(['apple','mandarin','orange','lemon'])
# **step-4** which value assigned for each data
le.classes_
# **step-4** give actual values
le.transform(df['fruit_name'])
# **or**
le = LabelEncoder()
le.fit_transform(df['fruit_name'])
# ## OneHotEncoder
from sklearn.preprocessing import OneHotEncoder
enc = OneHotEncoder()
enc.fit(df[['fruit_name']])
enc.transform(df[['fruit_name']]).toarray()
# #### get_dummies()
pd.get_dummies(df['fruit_name'])
# ### save the model/read model
df = pd.read_csv('fruit_data_with_colours.csv')
df.head(2)
df.sample(5)
X = df[['mass','width','height']]
y = df['fruit_label']
# **we train the model with Knn**
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=3)
model.fit(X,y)
print(model.score(X,y)*100)
# **to save the model for future use**
import pickle
pickle.dump(model, open('fruitModel.pkl','wb') )
# **how to reuse /read model**
model = pickle.load(open('fruitModel.pkl','rb'))
model.predict([[192,8.3,7.3]])
# **1. Kaggle website**
# **2. towardsdatascience**
# **3. analyticsvidya**
# **It will take 15 to 20 days to get the certificate**
| Day-9/Day9_24sep2020.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# language: python
# name: python37364bit31dfbc0e68ab47c2be432e09052b515f
# ---
# + jupyter={"outputs_hidden": true} tags=[]
# # !pip install yfinance
# # !pip install yahoofinancials
import time
import pandas as pd
import yfinance as yf
from yahoofinancials import YahooFinancials
# list of symbols from https://stockmarketmba.com/listofstocksforanexchange.php?s=GY
xetra_stocks = pd.read_csv('xetra_stocks.csv')
symbol_list_pre = xetra_stocks['Symbol'].values
symbol_list = []
for s in symbol_list_pre:
s = s.replace('GY:','')
s = s + '.DE'
symbol_list.append(s)
print(symbol_list)
data_list = []
start_date = '2018-12-22'
end_date = '2021-12-24'
i = 0
for s in symbol_list:
i = i + 1
print(f'Downloading {s} - {i} of {len(symbol_list)}')
df = yf.download(s,
start=start_date,
end=end_date,
progress=False)
data_list.append(df)
# niceness sleep.
time.sleep(2)
print(f'Downloaded {len(data_list)} histories.')
# + jupyter={"outputs_hidden": true} tags=[]
import pickle
with open('history.pkl', 'wb') as f:
pickle.dump(data_list, f)
| xetra_eod_download.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Scattered data interpolation via unregularized or regularized linear deconvolution
#
# This notebook describes and compares several methods of scattered data interpolation.
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import matplotlib
import numpy as np
import scipy.optimize as optimize
import pygsvd
import crr.scattered.sampling as sampling
import crr.scattered.roll_zeropad as roll_zeropad
matplotlib.rcParams['font.size'] = 16
matplotlib.rcParams['figure.figsize'] = [9., 7.]
dpi = 100
# The plot below shows the sampling (with the size of the points proportional to the width of the kernel.
samples = sampling.Sampling(nsamples=1000)
samples.set_flux(total_flux=1000., noise=1.e-0)
samples.imshow()
# We will reconstruct on a 21 by 21 grid, with the $x$ and $y$ coordinates set below (with coordinate $(0,0)$ at pixel center $(10, 10)$), and pixels space each unit of distance in the space.
# ### Modelling the image
# Relevant to some, though not all, methods of interpolation of scattered data is the idea of a model image that explains our sampled fluxes. To do so, we will define:
#
# * The vector of samples $\vec{f}$, of length $N$,
# * A model image $\vec{F}$, which represents a regular grid of delta functions, with a total of $M$ grid points, and
# * The kernel transformation ${\mathbf A}$, an $N\times M$ matrix.
#
# The kernel transformation defines how each sample responds to the delta function in the model image. We can build it with successive calls to the code that generates the fluxes. Under these definitions a model $\vec{m}$ can be written as:
#
# $$\vec{m} = {\mathbf A} \cdot \vec{f}$$
#
# Once we have ${\mathbf A}$ we can fit the parameters in the model image $\vec{F}$. We minimize:
#
# $$\chi^2 = \left(\vec{m} - \vec{f}\right)\cdot {\mathbf N}^{-1} \cdot \left(\vec{m} - \vec{f}\right)$$
#
# To find the values of $\vec{F}$ minimize $\chi^2$, we apply singular value decomposition as follows:
#
# $${\mathbf N}^{1/2} {\mathbf A} = {\mathbf U} \cdot \Sigma \cdot {\mathbf V}^T$$
#
# That makes the inversion of the problem easy so it is:
#
# $$\vec{F} = {\mathbf V}\cdot\Sigma \cdot {\mathbf U}^T \cdot {\mathbf N}^{-1/2} \cdot \vec{f} $$
#
# The covariance of this image can be calculated as:
#
# $${\mathbf C}_F^{-1} = {\mathbf A}^T\cdot{\mathbf N}^{-1}\cdot{\mathbf A}$$
#
# We can create a noiseless image and perform the fit. This works great. It is very closely equivalent to a full image deconvolution.
(U, S, VT) = np.linalg.svd(samples.A, full_matrices=False)
Sinv = np.zeros(len(S))
Sinv[S > 0] = 1. / S[S > 0]
W_F = VT.T.dot(np.diag(Sinv)).dot(U.T)
S_F = W_F.dot(samples.flux_nonoise)
S_F = S_F.reshape((samples.nx, samples.ny))
samples.imshow(S_F)
# But if we add even a tiny amount of noise this runs into terrible trouble. Even $10^{-6}$ level noise leads to a very troublesome covariance matrix, which leads to strong fluctuations.
samples.set_flux(total_flux=1000., noise=1.e-0)
S_F = W_F.dot(samples.flux)
S_F = S_F.reshape((samples.nx, samples.ny))
samples.imshow(S_F)
# The covariance matrix of this result can be calculated and clearly shows these issues.
C_F = W_F.dot(W_F.T)
myargs = {'interpolation': 'nearest', 'origin': 'lower', 'cmap': cm.Greys, 'vmin': -1., 'vmax': 1}
CC_F = 0. * C_F
for i in np.arange(samples.nx * samples.ny):
for j in np.arange(samples.nx * samples.ny):
CC_F[i, j] = C_F[i, j] / np.sqrt(C_F[i, i] * C_F[j, j])
plt.imshow(CC_F, **myargs)
nmid = (samples.nx * samples.ny) // 2
plt.xlim([nmid - 30, nmid + 30])
plt.ylim([nmid - 30, nmid + 30])
plt.colorbar()
plt.xlabel('pixel $i$')
plt.ylabel('pixel $j$')
# A very common approach to handling these issues is to regularize the fit. The simplest form of regularization is Tikhonov regularization. In its simplest form, it is designed to quadratically favor reconstruction values near zero. This case simply reduces to a conversion of the singular values that causes their inverse to smoothly go to zero below a value of $\Sigma \sim \lambda$. We can ask what this does, in the case of noise and no noise. The noiseless case is very enlightening. The PSF of this method is extremely irregular even for small values of $\lambda$.
llambda = 3.e-1
(U, S, VT) = np.linalg.svd(samples.A, full_matrices=False)
Sinv = np.zeros(len(S))
Sinv = S / (S**2 + llambda**2)
W_Ts = VT.T.dot(np.diag(Sinv)).dot(U.T)
S_Ts = W_Ts.dot(samples.flux)
S_Ts = S_Ts.reshape((samples.nx, samples.ny))
samples.imshow(S_Ts)
S_Ts = W_Ts.dot(samples.flux_nonoise)
S_Ts = S_Ts.reshape((samples.nx, samples.ny))
samples.imshow(S_Ts)
C_Ts = W_Ts.dot(W_Ts.T)
myargs = {'interpolation': 'nearest', 'origin': 'lower', 'cmap': cm.Greys, 'vmin': -1., 'vmax': 1}
CC_Ts = 0. * C_Ts
for i in np.arange(samples.nx * samples.ny):
for j in np.arange(samples.nx * samples.ny):
CC_Ts[i, j] = C_Ts[i, j] / np.sqrt(C_Ts[i, i] * C_Ts[j, j])
plt.imshow(CC_Ts, **myargs)
nmid = (samples.nx * samples.ny) // 2
plt.xlim([nmid - 30, nmid + 30])
plt.ylim([nmid - 30, nmid + 30])
plt.colorbar()
plt.xlabel('pixel $i$')
plt.ylabel('pixel $j$')
# There are more complicated ways of regularizing even in the Tikhonov case. In particular, one can construct $\Gamma$ to favor minimizing the squared differences of neighboring pixels. We can solve this problem using generalized SVD. First we will test that it works on the simplest Tikhonov case. It matches perfectly.
N = samples.nx * samples.ny
Gamma = np.diag(np.ones(N))
(sigma1_gs, sigma2_gs, X_gs, U_gs, V_gs) = pygsvd.gsvd(samples.A, Gamma, extras='uv')
XTinv_gs = np.linalg.inv(X_gs.T)
llambda = 3.e-3
sigmat_gs = np.diag(1. / (sigma1_gs * sigma1_gs + llambda**2 * sigma2_gs * sigma2_gs))
W_Tgs = XTinv_gs.dot(sigmat_gs).dot(np.diag(sigma1_gs)).dot(U_gs.T)
S_Tgs = W_Tgs.dot(samples.flux_nonoise)
S_Tgs = S_Tgs.reshape((samples.nx, samples.ny))
samples.imshow(S_Tgs)
# Now we construct $\Gamma$ that minimizes the squared error between neighboring pixels. This is somewhat different but also shows that the PSF for this case is a complicated beast with a lot of ringing.
nx = samples.nx
ny = samples.ny
Gamma = np.zeros((nx * ny, nx * ny))
ident = np.diag(np.ones(nx * ny)).reshape(nx, ny, nx, ny)
sident = - roll_zeropad.roll_zeropad(ident, -1, axis=2)
Gamma = Gamma + sident.reshape(nx * ny, nx * ny)
sident = - roll_zeropad.roll_zeropad(ident, 1, axis=2)
Gamma = Gamma + sident.reshape(nx * ny, nx * ny)
sident = - roll_zeropad.roll_zeropad(ident, -1, axis=3)
Gamma = Gamma + sident.reshape(nx * ny, nx * ny)
sident = - roll_zeropad.roll_zeropad(ident, 1, axis=3)
Gamma = Gamma + sident.reshape(nx * ny, nx * ny)
for indx in np.arange(nx * ny):
Gammasum = - Gamma[indx, :].sum()
Gamma[indx, :] = Gamma[indx, :] / Gammasum
Gamma[indx, indx] = 1.
(sigma1_g, sigma2_g, X_g, U_g, V_g) = pygsvd.gsvd(samples.A, Gamma, extras='uv')
XTinv_g = np.linalg.inv(X_g.T)
llambda = 3.e-2
sigmat_g = np.diag(1. / (sigma1_g * sigma1_g + llambda**2 * sigma2_g * sigma2_g))
W_Tg = XTinv_g.dot(sigmat_g).dot(np.diag(sigma1_g)).dot(U_g.T)
S_Tg = W_Tg.dot(samples.flux_nonoise)
S_Tg = S_Tg.reshape((nx, ny))
samples.imshow(S_Tg)
samples.set_flux(total_flux=1000., noise=1.e-0)
S_Tg = W_Tg.dot(samples.flux)
S_Tg = S_Tg.reshape((nx, ny))
samples.imshow(S_Tg)
C_Tg = W_Tg.dot(W_Tg.T)
myargs = {'interpolation': 'nearest', 'origin': 'lower', 'cmap': cm.Greys, 'vmin': -1., 'vmax': 1}
CC_Tg = 0. * C_Tg
for i in np.arange(nx * ny):
for j in np.arange(nx * ny):
CC_Tg[i, j] = C_Tg[i, j] / np.sqrt(C_Tg[i, i] * C_Tg[j, j])
plt.imshow(CC_Tg, **myargs)
nmid = (nx * ny) // 2
plt.xlim([nmid - 30, nmid + 30])
plt.ylim([nmid - 30, nmid + 30])
plt.colorbar()
plt.xlabel('pixel $i$')
plt.ylabel('pixel $j$')
| notebooks/scattered-linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Two runs for both the UMLfit algorithms
# +
import pickle
import json
from collections import Counter
import pandas as pd
import pickle
import re
import numpy as np
from collections import Counter, defaultdict, OrderedDict
from nltk import word_tokenize, pos_tag
import editdistance
import csv
from sklearn.metrics import f1_score
import numpy as np
import scipy.stats
from nltk.corpus import names
from sklearn.model_selection import train_test_split
from weighted_levenshtein import lev, osa, dam_lev
import langid
from nltk.tokenize.treebank import TreebankWordDetokenizer
from fastai.text import *
import os
# +
#import the test data
path = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/testDataST4_participants.txt'
test_data = pd.read_csv (path, sep = '\t', header = None)
test_data.columns = ['tweet_id', 'text']
print(test_data)
# -
txt = list(test_data['text'])
# +
#preprocessing
class Normalizer ():
def __init__(self):
pass
#to use this function the files need to be sorted in the same folder as the script under /obj_lex/
def load_obj(self, name):
with open('/home/dirksonar/Scripts/Project1_lexnorm/preprocessing_pipeline/obj_lex/' + name + '.pkl', 'rb') as f:
return pickle.load(f, encoding='latin1')
def load_files(self):
self.ext_vocab2 = self.load_obj('vocabulary_spelling_unique')
self.abbr_dict = self.load_obj ('abbreviations_dict')
self.celex_freq_dict = self.load_obj ('celex_lwrd_frequencies')
self.celex_list = list(self.celex_freq_dict.keys())
self.celex_set = set (self.celex_list)
self.drug_norm_dict = self.load_obj ('drug_normalize_dict')
def change_tup_to_list(self, tup):
thelist = list(tup)
return thelist
def change_list_to_tup(self,thelist):
tup = tuple(thelist)
return tup
#---------Remove URls, email addresses and personal pronouns ------------------
def replace_urls(self,list_of_msgs):
list_of_msgs2 = []
for msg in list_of_msgs:
nw_msg = re.sub(
r'\b' + r'((\(<{0,1}https|\(<{0,1}http|\[<{0,1}https|\[<{0,1}http|<{0,1}https|<{0,1}http)(:|;| |: )\/\/|www.)[\w\.\/#\?\=\+\;\,\&\%_\n-]+(\.[a-z]{2,4}\]{0,1}\){0,1}|\.html\]{0,1}\){0,1}|\/[\w\.\?\=#\+\;\,\&\%_-]+|[\w\/\.\?\=#\+\;\,\&\%_-]+|[0-9]+#m[0-9]+)+(\n|\b|\s|\/|\]|\)|>)',
' ', msg)
list_of_msgs2.append(nw_msg)
return list_of_msgs2
def replace_email(self,list_of_msgs):
list_of_msgs2 = []
for msg in list_of_msgs:
nw_msg = re.sub (r"([a-zA-Z0-9_.+-]+@[a-zA-Z0-9-]+\.[a-zA-Z0-9-.]+[. ])", ' ', msg) #remove email
nw_msg2 = re.sub (r"(@[a-zA-Z0-9]+[. ])", ' ', nw_msg) #remove usernames
# nw_msg3 = re.sub(r"(@ [a-zA-Z0-9]+[. ])", ' ', nw_msg2) #remove usernames
list_of_msgs2.append(nw_msg2)
return list_of_msgs2
def remove_empty (self,list_of_msgs):
empty = []
check_msgs3 =[]
for a, i in enumerate (list_of_msgs):
if len(i) == 0:
print('empty')
else:
check_msgs3.append(i)
return check_msgs3
def create_names_list (self):
male_names = names.words('male.txt')
female_names = names.words('female.txt')
male_set = set (male_names)
female_set = set (female_names)
names_set = male_set.union(female_set)
names_list = []
for word in names_set:
if (word != 'ned') & (word != 'Ned'): #ned means no evidence and is an important medical term
word1 = str.lower (word)
names_list.append(word1) #add the lowered words
names_list.append(word) #add the capitalized words
self.names_list = names_list
def remove_propernoun_names(self,msg):
try:
nw_msg = [self.change_tup_to_list(token) for token in msg]
for a, token in enumerate (nw_msg):
if (token[0] in self.names_list) and ((token[1] == 'NNP') or (token[1]== 'NNPS')):
new_token = token[0].replace (token[0], "-NAME-")
nw_msg[a] = [new_token, token[1]]
# nw_msg2 = [self.change_list_to_tup(token) for token in nw_msg]
return nw_msg
except TypeError:
pass
def remove_registered_icon (self, msg):
nw_msg = re.sub ('\u00AE', '', msg)
nw_msg2 = re.sub ('\u00E9', 'e', nw_msg)
return nw_msg2
#this function has been altered because we do not wnat to remove personal pronouns
def anonymize (self, posts):
posts2 = self.replace_urls (posts)
posts3 = self.replace_email (posts2)
posts4 = self.remove_empty(posts3)
posts5 = [self.remove_registered_icon(p) for p in posts4]
# posts5 = [p.encode('latin-1', errors = 'ignore').decode() for p in posts4]
posts6 = [word_tokenize (sent) for sent in posts5]
# posts6 = [pos_tag(sent) for sent in posts5]
# self.create_names_list()
# posts7 = [self.remove_propernoun_names (m) for m in posts6]
# posts8 = []
# for post in posts7:
# tg = [m[0] for m in post]
# posts8.append(tg)
return posts6
#---------Convert to lowercase ----------------------------------------------------
def lowercase (self, post):
post1 = []
for word in post:
word1 = word.lower()
post1.append (word1)
return post1
#---------Remove non_English posts -------------------------------------------------
def language_identify_basic (self, posts):
nw = []
tally = 0
list_removed = []
for post in posts:
out = langid.classify (post)
out2 = list(out)
if out2[0]=='en':
nw.append(post)
else:
tally += 1
list_removed.append(tuple ([post, out2[0], out2[1]]))
return nw, tally, list_removed
def language_identify_thres (self, msgs, lang_list, thres):
nw = []
tally = 0
list_removed = []
for post in msgs:
langid.set_languages(lang_list)
out = langid.classify (post)
out2 = list(out)
if out2[0]=='en':
nw.append(post)
elif out2[1] > thres:
nw.append(post)
else:
tally += 1
list_removed.append(tuple ([post, out2[0], out2[1]]))
return nw, tally, list_removed
def remove_non_english(self, posts):
d = TreebankWordDetokenizer()
posts2 = [d.detokenize(m) for m in posts]
posts_temp, tally, list_removed = self.language_identify_basic(posts2)
lang = []
for itm in list_removed:
lang.append(itm[1])
c = Counter(lang)
lang_list = ['en']
for itm in c.most_common(10):
z = list(itm)
lang_list.append(z[0])
print("Most common 10 languages in the data are:" + str(lang_list))
posts3, tally_nw, list_removed_nw = self.language_identify_thres(posts2, lang_list, thres = -100)
return posts3
#---------Lexical normalization pipeline (Sarker, 2017) -------------------------------
def loadItems(self):
'''
This is the primary load function.. calls other loader functions as required..
'''
global english_to_american
global noslang_dict
global IGNORE_LIST_TRAIN
global IGNORE_LIST
english_to_american = {}
lexnorm_oovs = []
IGNORE_LIST_TRAIN = []
IGNORE_LIST = []
english_to_american = self.loadEnglishToAmericanDict()
noslang_dict = self.loadDictionaryData()
for key, value in noslang_dict.items ():
value2 = value.lower ()
value3 = word_tokenize (value2)
noslang_dict[key] = value3
return None
def loadEnglishToAmericanDict(self):
etoa = {}
english = open('/home/dirksonar/Scripts/Project1_lexnorm/preprocessing_pipeline/obj_lex/englishspellings.txt')
american = open('/home/dirksonar/Scripts/Project1_lexnorm/preprocessing_pipeline/obj_lex/americanspellings.txt')
for line in english:
etoa[line.strip()] = american.readline().strip()
return etoa
def loadDictionaryData(self):
'''
this function loads the various dictionaries which can be used for mapping from oov to iv
'''
n_dict = {}
infile = open('/home/dirksonar/Scripts/Project1_lexnorm/preprocessing_pipeline/obj_lex/noslang_mod.txt')
for line in infile:
items = line.split(' - ')
if len(items[0]) > 0 and len(items) > 1:
n_dict[items[0].strip()] = items[1].strip()
return n_dict
#this has been changed becuase we are dealing with twitter data
def preprocessText(self, tokens, IGNORE_LIST, ignore_username=False, ignore_hashtag=True, ignore_repeated_chars=True, eng_to_am=True, ignore_urls=False):
'''
Note the reason it ignores hashtags, @ etc. is because there is a preprocessing technique that is
designed to remove them
'''
normalized_tokens =[]
#print tokens
text_string = ''
# NOTE: if nesting if/else statements, be careful about execution sequence...
for t in tokens:
t_lower = t.strip().lower()
# if the token is not in the IGNORE_LIST, do various transformations (e.g., ignore usernames and hashtags, english to american conversion
# and others..
if t_lower not in IGNORE_LIST:
# ignore usernames '@'
if re.match('@', t) and ignore_username:
IGNORE_LIST.append(t_lower)
text_string += t_lower + ' '
#ignore hashtags
elif re.match('#', t_lower) and ignore_hashtag:
IGNORE_LIST.append(t_lower)
text_string += t_lower + ' '
#convert english spelling to american spelling
elif t.strip().lower() in english_to_american.keys() and eng_to_am:
text_string += english_to_american[t.strip().lower()] + ' '
#URLS
elif re.search('http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', t_lower) and ignore_urls:
IGNORE_LIST.append(t_lower)
text_string += t_lower + ' '
elif not ignore_repeated_chars and not re.search(r'[^a-zA-Z]', t_lower):
# if t_lower only contains alphabetic characters
t_lower = re.sub(r'([a-z])\1+', r'\1\1', t_lower)
text_string += t_lower + ' '
# print t_lower
# if none of the conditions match, just add the token without any changes..
else:
text_string += t_lower + ' '
else: # i.e., if the token is in the ignorelist..
text_string += t_lower + ' '
normalized_tokens = text_string.split()
# print normalized_tokens
return normalized_tokens, IGNORE_LIST
def dictionaryBasedNormalization(self, tokens, I_LIST, M_LIST):
tokens2 =[]
for t in (tokens):
t_lower = t.strip().lower()
if t_lower in noslang_dict.keys() and len(t_lower)>2:
nt = noslang_dict[t_lower]
[tokens2.append(m) for m in nt]
if not t_lower in M_LIST:
M_LIST.append(t_lower)
if not nt in M_LIST:
M_LIST.append(nt)
else:
tokens2.append (t)
return tokens2, I_LIST, M_LIST
#----Using the Sarker normalization functions ----------------------------
#Step 1 is the English normalization and step 2 is the abbreviation normalization
def normalize_step1(self, tokens, oovoutfile=None):
global IGNORE_LIST
global il
MOD_LIST = []
# Step 1: preprocess the text
normalized_tokens, il = self.preprocessText(tokens, IGNORE_LIST)
normalized_minus_ignorelist = [t for t in normalized_tokens if t not in IGNORE_LIST]
return normalized_minus_ignorelist
def normalize_step2(self, normalized_tokens, oovoutfile=None):
global IGNORE_LIST
global il
MOD_LIST = []
ml = MOD_LIST
normalized_tokens, il, ml = self.dictionaryBasedNormalization(normalized_tokens, il, ml)
return normalized_tokens
def sarker_normalize (self,list_of_msgs):
self.loadItems()
msgs_normalized = [self.normalize_step1(m) for m in list_of_msgs]
msgs_normalized2 = [self.normalize_step2(m) for m in msgs_normalized]
return msgs_normalized2
#-------Domain specific abreviation expansion ----------------------------
# The list of abbreviations is input as a dictionary with tokenized output
def domain_specific_abbr (self, tokens, abbr):
post2 = []
for t in tokens:
if t in abbr.keys():
nt = abbr[t]
[post2.append(m) for m in nt]
else:
post2.append(t)
return post2
def expand_abbr (self, data, abbr):
data2 = []
for post in data:
post2 = self.domain_specific_abbr (tokens = post, abbr= abbr)
data2.append(post2)
return data2
#-------Spelling correction -------------------------------------------------
def load_files2 (self):
#load the edit matrices
#transpositions
self.edits_trans = self.load_obj ('weighted_edits_transpositions')
#deletions
self.edits_del = self.load_obj('weighted_edits_deletions')
#insertions
self.edits_ins = self.load_obj('weighted_edits_insertions')
#substitutions
self.edits_sub = self.load_obj('weighted_edits_substitutions')
#load the generic dictionary - CHANGE PATH!
self.celex_freq_dict = self.load_obj ('celex_lwrd_frequencies')
def initialize_weighted_matrices(self):
#initialize the cost matrixes for deletions and insertions
insert_costs = np.ones(128, dtype=np.float64) # make an array of all 1's of size 128, the number of ASCII characters
delete_costs = np.ones (128, dtype=np.float64)
for index,row in self.edits_ins.iterrows():
insert_costs[ord(index)] = row['transformed_frequency']
for index,row in self.edits_del.iterrows():
delete_costs[ord(index)] = row['transformed_frequency']
#substitution
substitute_costs = np.ones((128, 128), dtype=np.float64)
lst = []
for index,row in self.edits_sub.iterrows():
z = tuple([row['edit_from'], row['edit_to'], row['transformed_frequency']])
lst.append (z)
for itm in lst:
itm2 = list(itm)
try:
substitute_costs[ord(itm2[0]), ord(itm2[1])] = itm2[2]
except IndexError:
pass
#transposition
transpose_costs = np.ones((128, 128), dtype=np.float64)
lst = []
for index,row in self.edits_trans.iterrows():
z = tuple([row['first_letter'], row['second_letter'], row['transformed_frequency']])
lst.append (z)
for itm in lst:
itm2 = list(itm)
try:
transpose_costs[ord(itm2[0]), ord(itm2[1])] = itm2[2]
except IndexError:
print(itm2)
return insert_costs, delete_costs, substitute_costs, transpose_costs
def weighted_ed_rel (self, cand, token, del_costs, ins_costs, sub_costs, trans_costs):
try:
w_editdist = dam_lev(token, cand, delete_costs = del_costs, insert_costs = ins_costs, substitute_costs = sub_costs, transpose_costs = trans_costs)
rel_w_editdist = w_editdist/len(token)
return rel_w_editdist
except UnicodeEncodeError:
# print(token)
IGNORE_LIST.append(token)
rel_w_editdist = 100
return rel_w_editdist
def run_low (self, word, voc, func, del_costs, ins_costs, sub_costs, trans_costs):
replacement = [' ',100]
for token in voc:
sim = func(word, token, del_costs, ins_costs, sub_costs, trans_costs)
if sim < replacement[1]:
replacement[1] = sim
replacement[0] = token
return replacement
def spelling_correction (self, post, token_freq_dict, token_freq_ordered, min_rel_freq = 2, max_rel_edit_dist = 0.08):
post2 = []
cnt = 0
for a, token in enumerate (post):
if self.TRUE_WORD.fullmatch(token):
if token in self.spelling_corrections:
correct = self.spelling_corrections[token]
post2.append(correct)
cnt +=1
self.replaced.append(token)
self.replaced_with.append(correct)
elif token in self.celex_freq_dict:
post2.append(token)
else:
# make the subset of possible candidates
freq_word = token_freq_dict[token]
limit = freq_word * min_rel_freq
subset = [t[0] for t in token_freq_ordered if t[1]>= limit]
#compare these candidates with the word
candidate = self.run_low (token, subset, self.weighted_ed_rel, self.delete_costs_nw, self.insert_costs_nw,
self.substitute_costs_nw, self.transpose_costs_nw)
#if low enough RE - candidate is deemed good
if candidate[1] <= max_rel_edit_dist:
post2.append(candidate[0])
cnt +=1
self.replaced.append(token)
self.replaced_with.append(candidate[0])
self.spelling_corrections [token] = candidate[0]
else:
post2.append(token)
else: post2.append(token)
self.total_cnt.append (cnt)
return post2
def initialize_files_for_spelling(self):
total_cnt = []
replaced = []
replaced_with = []
spelling_corrections= {}
return total_cnt, replaced, replaced_with, spelling_corrections
def change_tup_to_list (self, tup):
thelist = list(tup)
return thelist
def create_token_freq (self, data):
flat_data = [item for sublist in data for item in sublist]
self.token_freq = Counter(flat_data)
token_freq_ordered = self.token_freq.most_common ()
self.token_freq_ordered2 = [self.change_tup_to_list(m) for m in token_freq_ordered]
def correct_spelling_mistakes(self, data):
# data= self.load_obj ('/data/dirksonar/Project1_lexnorm/spelling_correction/output/', 'gistdata_lemmatised')
self.load_files2()
self.insert_costs_nw, self.delete_costs_nw, self.substitute_costs_nw, self.transpose_costs_nw = self.initialize_weighted_matrices()
self.total_cnt, self.replaced, self.replaced_with, self.spelling_corrections= self.initialize_files_for_spelling()
self.TRUE_WORD = re.compile('[-a-z]+') # Only letters and dashes
# data2 = [word_tokenize(m) for m in data]
self.create_token_freq(data)
out = [self.spelling_correction (m, self.token_freq, self.token_freq_ordered2) for m in data]
return out, self.total_cnt, self.replaced, self.replaced_with, self.spelling_corrections
#--------Overall normalization function--------------------------------------
def normalize (self, posts):
self.load_files ()
posts1 = self.anonymize(posts)
posts2 = [self.lowercase (m) for m in posts1]
# posts3 = self.remove_non_english (posts2)
# posts3 = [word_tokenize(m) for m in posts2]
posts4 = [self.sarker_normalize(posts2)]
posts5 = [self.expand_abbr(posts4[0], self.abbr_dict)]
# posts6, total_cnt, replaced, replaced_with, spelling_corrections = self.correct_spelling_mistakes(posts5[0])
return posts5[0]
def normalize_extra(self, posts):
self.load_files()
self.loadItems()
posts2, total_cnt, replaced, replaced_with, spelling_corrections_nw = self.correct_spelling_mistakes(posts)
posts_ignored = []
for post in posts2:
p2 = [t for t in post if t not in IGNORE_LIST]
posts_ignored.append(p2)
return posts_ignored, total_cnt, replaced, replaced_with, spelling_corrections_nw
# -
txt_norm1 = Normalizer().normalize(txt)
txt_norm2, total_cnt, replaced, replaced_with, spelling_corrections_nw = Normalizer().normalize_extra(txt_norm1)
# +
# print(spelling_corrections_nw)
# +
# [print(m) for m in txt_norm2]
# +
def remove_punc (post):
temp = []
for word in post:
if re.fullmatch (r'[^\w\s]', word) == None:
temp.append (word)
else:
pass
return temp
# (u"\2026", '')
def post_filter_char (msg):
final1 = msg.replace('Â', '')
final2= final1.replace('’', '')
final3 = final2.replace('“', '')
final4 = final3.replace('–', '')
final5 = final4.replace('…', '')
final6 = final5.replace('â€', '')
final7 = final6.replace('...', '')
final8 = final7.replace ('`', '')
final9 = final8.replace ('ðÿ˜', '')
final10 = final9.replace ('¡', '')
final11 = final10.replace ('©', '')
final12 = re.sub(r'(@ ?[a-zA-Z0-9-_]+[\.: ]?)', '', final11)
return final12
txt_norm3 = [remove_punc(m) for m in txt_norm2]
d = TreebankWordDetokenizer ()
txt_norm4 = [d.detokenize(m) for m in txt_norm3]
txt_norm5 = [post_filter_char(m) for m in txt_norm4]
# -
# # Prediction time
# +
# os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
# os.environ["CUDA_VISIBLE_DEVICES"]="4"
# -
@dataclass
class Fbeta_binary(Callback):
"Computes the fbeta between preds and targets for single-label classification"
beta2: int = 2
eps: float = 1e-9
clas:int=1
def on_epoch_begin(self, **kwargs):
self.TP = 0
self.total_y_pred = 0
self.total_y_true = 0
def on_batch_end(self, last_output, last_target, **kwargs):
y_pred = last_output.argmax(dim=1)
y_true = last_target.float()
self.TP += ((y_pred==self.clas) * (y_true==self.clas)).float().sum()
self.total_y_pred += (y_pred==self.clas).float().sum()
self.total_y_true += (y_true==self.clas).float().sum()
def on_epoch_end(self, last_metrics, **kwargs):
beta2=self.beta2**2
prec = self.TP/(self.total_y_pred+self.eps)
rec = self.TP/(self.total_y_true+self.eps)
res = (prec*rec)/(prec*beta2+rec+self.eps)*(1+beta2)
self.metric = res
return add_metrics(last_metrics, self.metric)
# print(txt_norm5)
#run ULMfit algorithm1
path = '/data/dirksonar/Project3_sharedtasks_SMM4H/Task4/fastai/'
learn = load_learner (path, 'classifier_phm_1.pkl')
# +
def extract_label (output):
out = []
for i in output:
lst = list(i)
z = str(lst[0])
label = z[-1]
out.append(float(label))
return out
predicted = [learn.predict (i) for i in txt_norm5]
pred_labels = extract_label(predicted)
# -
print(pred_labels)
#run UMlfit algorithm2
path = '/data/dirksonar/Project3_sharedtasks_SMM4H/umlfit_languagemodel/'
learn = load_learner (path, 'classifier_phm_2.pkl')
# +
def extract_label (output):
out = []
for i in output:
lst = list(i)
z = str(lst[0])
label = z[-1]
out.append(float(label))
return out
predicted2 = [learn.predict (i) for i in txt_norm5]
pred_labels2 = extract_label(predicted2)
# +
from sklearn.metrics import accuracy_score
print(accuracy_score(pred_labels, pred_labels2))
# +
#save output in correct manner
output_test_task4_run1 = pd.concat([test_data, pd.Series(pred_labels)], axis = 1)
output_test_task4_run2 = pd.concat([test_data, pd.Series(pred_labels2)], axis = 1)
output_test_task4_run1.head()
# +
def save_obj(obj, name):
with open(name + '.pkl', 'wb') as f:
pickle.dump(obj, f, pickle.HIGHEST_PROTOCOL)
path = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/output_test_task4_run1'
path2 = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/output_test_task4_run2'
save_obj(output_test_task4_run1, path)
save_obj(output_test_task4_run2, path2)
# -
output_test_task4_run1_notext = output_test_task4_run1.drop('text', axis =1)
output_test_task4_run1_notext.columns =['Tweet ID', 'Label']
output_test_task4_run1_notext
output_test_task4_run2_notext = output_test_task4_run2.drop('text', axis =1)
output_test_task4_run2_notext.columns =['Tweet ID', 'Label']
# +
path = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/output_test_task4_run1_notext'
path2 = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/output_test_task4_run2_notext'
save_obj(output_test_task4_run1_notext, path)
save_obj(output_test_task4_run2_notext, path2)
# +
path = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/output_test_task4_run1_notext.txt'
path2 = '/data/dirksonar/Project3_sharedtasks_SMM4H/testdata/output_test_task4_run2_notext.txt'
output_test_task4_run1_notext.to_csv (path, index = False, sep= '\t', header = False)
output_test_task4_run2_notext.to_csv (path2, index = False, sep= '\t', header = False)
# -
| Task4_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Geo-Distribution of Tourism #
#
# The tourism industry has been masively affected by the Covid-19 situation. One of the indicators for travelling is the number of Airbnb reviews, treated here as demand.
#
# This notebook extracts the data from InsideAirbnb, aggregates it and displays the number of reviews for each neighbourhood per month.
#
# ### Inputs: ###
# <br>Bristol_reviews.csv - This dataset is the reviews file downloaded from http://insideairbnb.com/get-the-data.html. It contains all the reviews up to the last scraped date. <br/>
# <br> Bristol_listings.csv - This dataset is the listings file downloaded from http://insideairbnb.com/get-the-data.html. It contains the listings of that specific month. <br/>
# <br>neighbourhoods.geojson - This dataset is the geojson file downloaded from http://insideairbnb.com/get-the-data.html. It contains the geometry of the neighbourhoods.
#
#
# ### Steps ###
#
# 1. Get the number of reviews per day per city.
# 2. Get the number of reviews per day per district.
# 3. Normalize teh number of reviews for each month.
# 4. Plot each neighbourhood for a given month.
# +
# Install all the dependencies
# !pip install shapely
# !pip install geopandas
# !pip install geojsonio
# !pip install descartes
# !pip install wget
# !pip install pandas
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
# %matplotlib inline
import requests
from bs4 import BeautifulSoup
from tqdm import tqdm
import io
import wget
from os.path import isfile
from os import remove
# +
# Scrap the airbnb website and fetch the name and links to all available data files
r = requests.get("http://insideairbnb.com/get-the-data.html")
soup = BeautifulSoup(r.text)
alldata = {}
for table in tqdm(soup.findAll("table", {"class":"table-hover"})):
df = pd.read_html(io.StringIO(str(table)))[0]
df["downloadlink"] = ""
index = 0
for link in table.findAll("a",href=True):
df.at[index,"downloadlink"] = link["href"]
index += 1
city = df["Country/City"].unique()[0]
alldata[city] = df
# -
# ## Get reviews per day per city
def get_reviews_for_city(city_name):
city_name = city_name.replace(" ", "_")
filename = f"{city_name}_reviews.csv"
if not isfile(filename):
return None
df = pd.read_csv(filename)
df["number_of_reviews"] = 1
df = df[["date", "number_of_reviews"]]\
.groupby("date")\
.sum()\
.reset_index(drop=False)\
.rename(columns={
"number_of_reviews": city_name}
)
df = df[df["date"] != "0"]
df.date = pd.to_datetime(df.date)
df.set_index("date", drop=True, inplace=True)
df = df.astype(float)
df = df.resample('D').mean()
t_index = pd.DatetimeIndex(start='2014-01-01', end='2020-06-30', freq='D')
df = df.interpolate(method='linear', limit_area="inside")
return df
# +
# Get all the reviews for all the cities available on insideairbnb in one DataFrame
city_names = list(alldata.keys())
df_final = pd.DataFrame()
for idx, city_name in enumerate(city_names):
print("City: ", city_name)
if idx > 1000000:
break
df = get_reviews_for_city(city_name)
if df is not None:
df_final = pd.concat([df_final, df], axis=1)
# +
# Plot the number of reviews over time for Bristol and London
fig = df_final[["Bristol", "London"]].plot(figsize=(20, 10),fontsize= 16)
fig.legend(fontsize =16)
# -
# ## Get reviews per day per district
# +
# Define utility functions
def get_geojson_url(city_name):
df_city_info = alldata[city_name]
df_city_info = df_city_info[df_city_info["File Name"] == "neighbourhoods.geojson"]
return df_city_info["downloadlink"].iloc[0]
def download_geojson(geojson_url):
if isfile("neighbourhoods.geojson"):
remove("neighbourhoods.geojson")
wget.download(geojson_url)
# +
# Get a dataframe with all the reviews of a given city, per district.
def get_reviews_by_district(city_name):
geojson_url = get_geojson_url(city_name)
download_geojson(geojson_url)
if not isfile('neighbourhoods.geojson'):
return None
df_places = gpd.read_file('neighbourhoods.geojson')
df_places["neighbourhood"] = df_places["neighbourhood"].astype(str)
city_name = city_name.replace(" ", "_")
filename = f"{city_name}_listings.csv"
if not isfile(filename):
return None
df = pd.read_csv(filename)
df = df[["id", "neighbourhood_cleansed"]]
df.drop_duplicates(subset=["id"], keep="first", inplace=True)
df["neighbourhood_cleansed"] = df["neighbourhood_cleansed"].astype(str)
df = pd.merge(df, df_places, left_on="neighbourhood_cleansed", right_on="neighbourhood", how="left")
df.fillna(0, inplace=True)
df = df[["id", "neighbourhood", "geometry"]]
filename = f"{city_name}_reviews.csv"
if not isfile(filename):
return None
df_reviews = pd.read_csv(filename)
df = pd.merge(df, df_reviews, left_on="id", right_on="listing_id", how="right")
df["yearmonth"] = df["date"].apply(lambda x: str(x)[:7])
print(df)
df.drop(["listing_id", "date"], axis=1, inplace=True)
df["n_reviews"] = 1
df = df.groupby(["neighbourhood", "yearmonth"]).agg({
"geometry": lambda x: x.iloc[0],
"n_reviews": sum
}).reset_index(drop=False)
return df
# -
df = get_reviews_by_district("Bristol")
# ## Normalize the number of reviews for each month##
for yearmonth in df.yearmonth.unique():
idx = df[df["yearmonth"] == yearmonth].index
_sum = df[df["yearmonth"] == yearmonth]["n_reviews"].sum()
df.iloc[idx, 3] = 100 * (df.iloc[idx, 3] / _sum)
# ## Plot each neighbourhood for a given month##
# +
# Choose a month and plot the map of the chosen city to see how many bookings were made in each district
MONTH = "2020-04"
gdf = gpd.GeoDataFrame(df[df["yearmonth"]==MONTH],
geometry="geometry")
gdf.plot(column="n_reviews",
figsize=(20, 20),
cmap="Blues",
linewidth=0.8,
edgecolor="0.8")
| Mobility and Tourism - WS2/airbnb_analysis/Geo_Distribution_Tourism.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Interdisciplinary Communication Lesson
#
#
# Welcome to the Hour of Cyberinfrastructure (also known as the Hour of CI). This is the Interdisciplinary Communication Lesson. Interdisciplinary Communication involves discussing, listening, and speaking across levels and types of expertise. In this lesson, you will learn about some key components of communicating to people of different experiences, backgrounds, and expertise. These practices are essential for working in diverse teams and with diverse backgrounds.
#
# <br>
#
# Lesson Developer:
# * <NAME> (<EMAIL>),
# + hide_input=false init_cell=true slideshow={"slide_type": "skip"} tags=["Hide"]
# This code cell starts the necessary setup for Hour of CI lesson notebooks.
# First, it enables users to hide and unhide code by producing a 'Toggle raw code' button below.
# Second, it imports the hourofci package, which is necessary for lessons and interactive Jupyter Widgets.
# Third, it helps hide/control other aspects of Jupyter Notebooks to improve the user experience
# This is an initialization cell
# It is not displayed because the Slide Type is 'Skip'
from IPython.display import HTML, IFrame, Javascript, display
from ipywidgets import interactive
import ipywidgets as widgets
from ipywidgets import Layout
import getpass # This library allows us to get the username (User agent string)
# import package for hourofci project
import sys
sys.path.append('../../supplementary') # relative path (may change depending on the location of the lesson notebook)
import hourofci
# Retreive the user agent string, it will be passed to the hourofci submit button
agent_js = """
IPython.notebook.kernel.execute("user_agent = " + "'" + navigator.userAgent + "'");
"""
Javascript(agent_js)
# load javascript to initialize/hide cells, get user agent string, and hide output indicator
# hide code by introducing a toggle button "Toggle raw code"
HTML('''
<script type="text/javascript" src=\"../../supplementary/js/custom.js\"></script>
<style>
.output_prompt{opacity:0;}
</style>
<input id="toggle_code" type="button" value="Toggle raw code">
''')
# + [markdown] slideshow={"slide_type": "slide"}
# ## Before we get started
#
# You should know that the Hour of CI project is a research study and your participation is voluntary. Please read the following invitation to take part in the study and participate in the Hour of CI project.
# + [markdown] rise={"scroll": true} slideshow={"slide_type": "slide"}
# #### PERMISSION TO TAKE PART IN A RESEARCH STUDY
#
# <font size="+1">
#
# You are invited to be in a research study for the Hour of Cyberinfrastructure: Developing Cyber Literacy for GIScience project. We ask that you read this form and ask any questions you may have before agreeing to be in the study. This study is being conducted by: <NAME>, Department of Geography, Environment, and Society at the University of Minnesota.
#
# ##### Procedures:
# If you agree to be in this study, you will be learning about cyberinfrastructure and related concepts using a web-based platform that will take approximately one hour per lesson. Each lesson will include readings, short assessments such as multiple choice questions and short answers, and interactive components all to enhance and gauge learning. Your answers to questions will be recorded in a secure database. Participation in this study is voluntary. If you are not interested in participating please exit the browser or navigate to this website: <a href="http://www.umn.edu" target="http://www.umn.edu">http://www.umn.edu</a>.
#
# ##### Confidentiality:
# The individual records of this study will be kept private. In any sort of report we might publish, we will not include any information that will make it possible to identify a subject. Research records will be stored securely and only researchers will have access to the records. All information will be stored in a secure database for current and future research purposes.
#
# ##### Voluntary Nature of the Study:
# Participation in this study is voluntary. Your decision whether or not to participate will not affect your current or future relations with the University of Minnesota, University of Southern California, University of Illinois at Urbana-Champaign, or University of Massachusetts at Amherst.
#
# ##### Contacts and Questions:
# The principal researcher conducting this study is: <NAME>.
#
# If you have questions now or later, you are encouraged to contact him at:
# * Address: 267 – 19 th Avenue South, 414 Social Sciences Building, Minneapolis, MN, 55455,
# * Phone: 612-625-5970,
# * Email: <EMAIL>
#
# This research has been reviewed and approved by an IRB within the Human Research Protections Program (HRPP) - STUDY00007588. To share feedback privately with the HRPP about your research experience, call the Research Participants’ Advocate Line at 612-625-1650 (Toll Free: 1-888-224-8636) or go to z.umn.edu/participants. You are encouraged to contact the HRPP if:
# * Your questions, concerns, or complaints are not being answered by the research team.
# * You cannot reach the research team.
# * You want to talk to someone besides the research team.
#
# HRP-587 Template Version: 2/28/2019
# * You have questions about your rights as a research participant.
# * You want to get information or provide input about this research.
#
#
# ##### Permission:
# Participants in this research must be 18 years or older. If you are under the age of 18 then please exit this webpage or navigate to another website such as the Hour of Code at https://hourofcode.com, which is designed for K-12 students.
#
#
# By clicking on the "Agree" button below you are granting your permission to take part in this research and start an Hour of CI lesson. If you are not interested in participating please exit the browser or navigate to this website: http://www.umn.edu. Your participation is voluntary and you are free to stop the lesson at any time.
#
# Please print a copy of this information to keep for your records.
#
# </font>
#
# <a href="ic-2.ipynb" style="background-color:blue;color:white;padding:10px;margin:2px;font-weight:bold;">I am 18 years or older, and I agree to participate in the study</a>
#
#
| beginner-lessons/interdisciplinary-communication/ic-1.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # 변수,자료형,기본 함수, 작업공간
# - 변수는 분석에 필요한 자료를 일시적으로 저장하거나 처리결과를 담을 수 있는 기억장소를 지정해주는 역할을 한다. 즉 변수의 이름은 값을 저장하는 메모리 영역의 이름으로 할당된다. 또한 R은 모든 변수가 객체(object) 형태로 생성되기 때문에 하나의 변수에 자료와 함수 그리고 차트와 같은 이미지까지 모든 형식을 저장할 수 있다.
# - 변수 이름 작성 규칙
# - 첫 자는 영문자로 시작
# - 두 번째 단어는 숫자와 밑줄 문자 그리고 점(.)을 이용할 수 있다.
# - 대문자와 소문자는 서로 다른 변수로 인식한다.
# - 변수 이름은 의미를 파악할 수 있는 이름으로 지정하는 것이 좋다.
# - 두 단어를 포함하여 변수 이름을 지정할 경우 두 번째 단어의 첫 자는 대문자로 표기한다.
# - 한 번 정의된 변수는 재사용이 가능하고 가장 최근에 할당된 값으로 수정된다.
var1 <- 0
var1 = 9
var1
# - `=` 나 `<-` 둘 다 사용가능하다
# ---
# - 스칼라 변수 : 한 개의 값만 갖는 변수를 의미한다.
# - 벡터 변수 : 두 개 이상의 값을 갖는 변수를 의미한다.
# - 여러 개의 자료를 저장할 수 있는 1차원의 선형 자료구조이다.
age<-35 # 정수
name<-'홍길동' # 문자열
age;name
name<-c('홍길동','이순신','곽도성')
# 이건 두 개 이상의 값을 갖는 변수, 여러 개의 자료를 저장할 수 있는 1차원의 선형 자료구조
# 즉 벡터변수
# ---
# - R은 변수를 선언할 때 별도의 자료형(type)을 선언하지 않는다. 즉 변수에 저장하는 자료의 유형에 의해서 변수의 타입이 결정된다!
# - 숫자형 : 정수, 실수
# - 문자형 : 문자, 문자열
# - 논리형 : 참, 거짓
# - 결측 데이터 : 결측치, 비숫자(NA,NaN)
int<-20
string<-'홍길동'
boolean<-TRUE # True파이썬에서는 이렇게 작성해야 함
sum(10,20,30)
# 3개의 숫자형 값의 합계 연산
sum(10,20,30,NA)
# sum 안 됨
sum(10,20,30,NA,na.rm=TRUE)
# na.rm=TRUE를 통해 NA 결측치 제거 후 합계 연산
# 이건 현재 사용 중인 변수(객체) 보기
ls()
# ---
# - 자료형 확인
# - 변수에 저장된 자료형을 확인하는 함수를 이용하여 반환되는 TRUE 또는 FALSE의 결과를 통해서 해당 변수의 자료형을 확인할 수 있다.
# - is.numeric(x) - 수치형 여부
# - is.logical(x)
# - is.character(x) - 문자형 여부
# - is.data.frame(x) - 데이터프레임 여부
# - is.na(x)
# - is.integer(x) - 정수형 여부
# - is.double(x) - 실수형 여부
# - is.complex(x)
# - is.factor(x) - 범주형 여부
# - is.nan(x)
is.character(string)
# ---
# - 자료형 변환
# - 변수에 저장된 자료형을 다른 자료형으로 변환하자
# - as.numeric(x) - 수치형 변환
# - as.logical(x)
# - as.character(x) - 문자형 변환
# - as.data.frame(x) - 데이터프레임 변환
# - as.list(x) - 리스트형 변환
# - as.array(x) - 다차원 배열 변환
# - as.integer(x) - 정수형 변환
# - as.double(x) - 실수형 변환
# - as.complex(x)
# - as.factor(x) - 요인형 변환
# - as.Data(x) - 날짜형 변환
x<-c(1,2,'3') # 3개의 원소를 갖는 벡터 생성
# x*3
# 이렇게 작성하면 이항연산자에 수치가 아닌 인수가 있다는 error가 발생하게 됨
result<-as.numeric(x)*3
result2<-as.integer(x)*3
result
result2
# - c() 함수를 이용하여 벡터를 생성할 경우 원소 중 한 개라도 문자이면 모든 원소를 문자로 하여 객체가 생성된다. 따라서 이항연산자에 수치가 아닌 인수가 있다는 error가 발생하게 된다.
# ---
# - 복소수
z<-5.3-3i
Re(z) # 실수
Im(z) # 허수
is.complex(z)
as.complex(123)
# ---
# - 자료형과 자료구조 보기
# - 자료형은 변수에 저장된 자료의 성격(숫자형, 문자형, 논리형)을 의미하고, 자료구조는 변수에 저장된 자료의 메모리 구조(배열, 리스트, 테이블)를 의미한다. 메모리 구조는 객체가 생성될 때 만들어지기 때문에 자료구조를 객체형(Object Type)이라고도 한다. R에서는 mode()를 이용하여 자료형을 확인할 수 있고, class()를 이용하여 자료 구조, 즉 메모리 구조를 확인할 수 있다.
int<-3
mode(int) # 자료의 성격을 알려준다.
class(int) # 자료구조의 성격을 알려준다.
# - 이처럼 벡터 변수가 아닌 스칼라 변수일 땐 자료의 성격을 알려주는 mode()와 자료 구조의 성격을 알려주는 class() 함수의 결과는 같은 유형으로 나타난다.
# ---
# - 요인(Factor)형 변환
# - 요인(Factor)은 같은 성격인 값의 목록을 범주로 갖는 벡터 자료를 의미한다. 범주는 변수가 가질 수 있는 값의 범위로 예를 들면 성별 변수의 범주는 남자와 여자가 되낟. 요인형은 순서에 의미가 없는 Nominal 유형과 순서에 의미가 있는 Ordinal 유형으로 구분된다.
# - Nominal : 범주의 순서는 알파벳 순서로 정렬
# - Ordinal : 범주의 순서는 사용자가 지정한 순서대로 정렬
gender<- c('man','woman','woman','man','man')
# plot(gender)
# error 발생, 차트는 수치 데이터만 가능하다.
# - Factor Nominal : 벡터 원소를 요인형으로 변환한 경우 범주의 순서가 알파벳 순서로 정렬되는 요인형의 기본 유형이다.
Ngender<-as.factor(gender) # Factor형 변환
Ngender
# - 여기서 Levels가 의미하는 것이 범주이다. 여기서 범주의 수준(Levels)은 값의 목록을 알파벳 순서로 정렬한다.
table(Ngender)
# 빈도수 구하기
# - gender 원소 중에서 같은 값의 수량을 수치화한 빈도수를 확인할 수 있다. 여기서 man과 woman은 범주가 된다. 범주가 된다는 의미는 gender 변수가 값을 가질 수 잇는 범위를 의미한다. 또한 빈도수는 해당 범주의 발생 수를 의미한다.
plot(Ngender)
mode(Ngender)
class(Ngender)
is.factor(Ngender)
# ---
# - Factor Ordinal : 범주의 순서를 사용자가 지정한 순서대로 정렬하는 기능으로 factor() 함수의 형식은 다음과 같다.
# -
# ```r
# factor(x,levels,ordered)
# ```
# factor() 함수의 매개변수 보기
args(factor)
Ogender<-factor(gender,levels=c('woman','man'),ordered = TRUE)
Ogender
# factor 함수에서 사용할 수 있는 매개변수를 확인하고, 해당 변수를 이용하여 순서 잇는 요인형으로 변환한 후 Levels:에서 범주의 순서를 확인할 수 있다.
# 전에는 man이 먼저 나왔는데 이번엔 woman이 먼저 나온 것을 확인할 수 있다.
par(mfrow=c(1,2))
# 두 개의 그래프를 Plots 영역에 나타낼 수 있다.
# (2,2)라고 바꿔주면 4개의 그래프를 그릴 수 있다.
plot(Ngender)
plot(Ogender)
# ---
# - 날짜형 변환 : 인터넷 또는 로컬 파일로부터 가져온 자료 중에서 날짜형 칼럼은 요인형 또는 문자형으로 인식되기 때문에 정확한 날짜형으로 변환할 필요가 있다.
a=as.Date('20/02/28','%y/%m/%d')
a
class(a)
dates=c('15/02/24','10/02/2','16/08/14')
as.Date(dates,'%y/%m/%d')
dates=c('15/22/24','98/02/2','16/08/14')
as.Date(dates,'%y/%m/%d')
# - 22월이라는 날짜는 없으므로 NA처리 되었다.
# - 년도 4자리로 작성 : %Y
# - 년도 2자리로 작성 : %y
# - 24시간 : %H
# - 12시간 : %I
# - 분 : %M
# - 초 : %S
# ---
# - 현재 날짜와 시간 확인
a=Sys.time()
mode(a)
class(a)
# - strptime()를 이용한 날짜형 변환
sdate<-'2019-11-11 12:17:6'
class(sdate)
today<-strptime(sdate,format='%Y-%m-%d %H:%M:%S')
today
class(today)
# - 반면 as.Date()는 날짜 자료만 형 변환이 가능하다.
# ---
# - 기본 함수와 작업공간
# - R 패키지에서 제공되는 수많은 함수 사용법을 머릿속에 기억하기는 불가능한 일이다. 따라서 해당 함수의 사용법을 제공하는 도움말 기능을 이용할 수 있어야 한다.
# - 'help(함수명)' 또는 '?함수명'형식으로 볼 수 있다.
# - 파이썬에서는 '함수명?'이다.
# - google에서는 '함수명() in r' 형식으로 검색한 뒤 확인할 수 있다.
# - arg(함수명) : 특정 함수를 대상으로 사용 가능한 함수 파라미터를 보여준다.
# - 예를 들어보자
args(max)
# na.rm은 FALSE가 default이다.
# - example() 함수는 R에서 제공되는 기본 함수들을 사용하는 예제를 제공해준다.
example(seq)
# - seq() 함수를 사용하는 예로 다양한 방법으로 벡터 원소를 생성하는 과정을 보여준다.
# - mean의 예제도 알아보자
example(mean)
| _notebooks/2022-02-04-R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import Interpolation
# %matplotlib notebook
# # %matplotlib widget
import matplotlib.pyplot as plt
plt.rcParams['font.size']=16
plt.rcParams['font.family']='dejavu sans'
plt.rcParams['mathtext.fontset']='stix'
plt.rcParams['mathtext.rm']='custom'
plt.rcParams['mathtext.it']='stix:italic'
plt.rcParams['mathtext.bf']='stix:bold'
# -
_=np.loadtxt('example/eos2020.dat')
T,h_tab,g_tab=_[:,0],_[:,1],_[:,2]
# +
h0=Interpolation.Interpolation(T,h_tab)
h1=Interpolation.linearSpline(T,h_tab)
h3=Interpolation.cubicSpline(T,h_tab)
dh0=lambda T:1+1/3*T/h0(T)*h0.derivative_1(T)
dh1=lambda T:1+1/3*T/h1(T)*h1.derivative_1(T)
dh3=lambda T:1+1/3*T/h3(T)*h3.derivative_1(T)
g0=Interpolation.Interpolation(T,g_tab)
g1=Interpolation.linearSpline(T,g_tab)
g3=Interpolation.cubicSpline(T,g_tab)
# +
fig=plt.figure(figsize=(9,5))
fig.subplots_adjust(bottom=0.15, left=0.15, top = 0.9, right=0.9,wspace=0.0,hspace=0.0)
sub = fig.add_subplot(1,1,1)
T_int=np.logspace(np.log10(min(T)),np.log10(max(T)),len(T)*10)
Y_0=[h0(_) for _ in T_int]
sub.plot(T_int,Y_0,linestyle=':',linewidth=1,alpha=0.2,c='xkcd:blue',label='interpolation-0')
Y_1=[h1(_) for _ in T_int]
sub.plot(T_int,Y_1,linestyle='--',linewidth=1,alpha=0.2,c='xkcd:black',label='interpolation-1')
Y_3=[h3(_) for _ in T_int]
sub.plot(T_int,Y_3,linestyle='-',linewidth=1.5,alpha=1,c='xkcd:red',label='interpolation-3')
sub.scatter(T,h_tab,alpha=0.3,c='xkcd:gray',marker='+',label='data')
sub.legend(bbox_to_anchor=(0.01, 0.99),borderaxespad=0., columnspacing=1,labelspacing=-0.1,
borderpad=0,ncol=1,loc='upper left' ,framealpha=0)
sub.set_xlabel(r'$T ~ [{\rm GeV}]$')
sub.xaxis.set_label_coords(0.5, -0.09)
sub.set_ylabel(r'$h_{\rm eff}$')
sub.yaxis.set_label_coords(-0.09,0.5)
sub.set_xscale('log')
sub.set_yscale('linear')
fig.show()
# +
fig=plt.figure(figsize=(9,5))
fig.subplots_adjust(bottom=0.15, left=0.15, top = 0.9, right=0.9,wspace=0.0,hspace=0.0)
sub = fig.add_subplot(1,1,1)
T_int=np.logspace(np.log10(min(T)),np.log10(max(T)),len(T)*10)
Y_0=[g0(_) for _ in T_int]
sub.plot(T_int,Y_0,linestyle=':',linewidth=1,alpha=0.2,c='xkcd:blue',label='interpolation-0')
Y_1=[g1(_) for _ in T_int]
sub.plot(T_int,Y_1,linestyle='--',linewidth=1,alpha=0.2,c='xkcd:black',label='interpolation-1')
Y_3=[g3(_) for _ in T_int]
sub.plot(T_int,Y_3,linestyle='-',linewidth=1.5,alpha=1,c='xkcd:red',label='interpolation-3')
sub.scatter(T,g_tab,alpha=0.3,c='xkcd:gray',marker='+',label='data')
sub.legend(bbox_to_anchor=(0.01, 0.99),borderaxespad=0., columnspacing=1,labelspacing=-0.1,
borderpad=0,ncol=1,loc='upper left' ,framealpha=0)
sub.set_xlabel(r'$T ~ [{\rm GeV}]$')
sub.xaxis.set_label_coords(0.5, -0.09)
sub.set_ylabel(r'$g_{\rm *}$')
sub.yaxis.set_label_coords(-0.09,0.5)
sub.set_xscale('log')
sub.set_yscale('linear')
fig.show()
# +
fig=plt.figure(figsize=(9,6))
fig.subplots_adjust(bottom=0.15, left=0.15, top = 0.9, right=0.9,wspace=0.0,hspace=0.03)
fig.suptitle('')
sub = fig.add_subplot(2,1, 1)
T_int=np.logspace(np.log10(min(T)),np.log10(max(T)),len(T)*10)
gt=[g3(i) for i in T]
ht=[h3(i) for i in T]
dht0=[dh0(i) for i in T]
dht1=[dh1(i) for i in T]
dht3=[dh3(i) for i in T]
sub.plot(T,gt,linestyle='-',c='xkcd:red',label=r"$g_{\rm *} (T)$")
sub.plot(T,ht,linestyle='-',c='xkcd:black',label=r"$h_{\rm eff} (T)$")
sub.legend(bbox_to_anchor=(1, 0.0),borderaxespad=0.,
borderpad=0.05,ncol=1,loc='lower right',fontsize=14,framealpha=0)
sub.set_yscale('log')
sub.set_xscale('log')
sub.get_xaxis().set_visible(False)
sub.set_ylim(3,1.2e2)
sub = fig.add_subplot(2,1,2)
sub.plot(T,dht0,linestyle=':',c='xkcd:blue',label=r"Interpolation-0")
sub.plot(T,dht1,linestyle='--',c='xkcd:red',label=r"Interpolation-1")
sub.plot(T,dht3,linestyle='-',c='xkcd:black',label=r"Interpolation-3")
sub.legend(bbox_to_anchor=(0.99, 0.99),borderaxespad=0.,
borderpad=0.05,ncol=1,loc='upper right',fontsize=14,framealpha=0)
sub.set_ylabel(r'$\delta_h=1+\dfrac{1}{3} \dfrac{d \log h_{\rm eff}}{d \log T}$')
sub.yaxis.set_label_coords(-0.09,0.5)
sub.set_yscale('linear')
sub.set_xscale('log')
sub.set_xlabel(r'$T \; [{\rm GeV}]$')
sub.xaxis.set_label_coords(0.5, -0.17)
fig.show()
# -
| Interpolation/python/Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Geometric operations
#
# ## Overlay analysis
#
# In this tutorial, the aim is to make an overlay analysis where we create a new layer based on geometries from a dataset that `intersect` with geometries of another layer. As our test case, we will select Polygon grid cells from `TravelTimes_to_5975375_RailwayStation_Helsinki.shp` that intersects with municipality borders of Helsinki found in `Helsinki_borders.shp`.
#
# Typical overlay operations are (source: [QGIS docs](https://docs.qgis.org/2.8/en/docs/gentle_gis_introduction/vector_spatial_analysis_buffers.html#more-spatial-analysis-tools)):
# 
#
# ## Download data
#
# For this lesson, you should [download a data package](https://github.com/AutoGIS/data/raw/master/L4_data.zip) that includes 3 files:
#
# 1. Helsinki_borders.shp
# 2. Travel_times_to_5975375_RailwayStation.shp
# 3. Amazon_river.shp
#
# ```
# $ cd /home/jovyan/notebooks/L4
# $ wget https://github.com/AutoGIS/data/raw/master/L4_data.zip
# $ unzip L4_data.zip
# ```
#
# Let's first read the data and see how they look like.
#
# - Import required packages and read in the input data:
# +
import geopandas as gpd
import matplotlib.pyplot as plt
import shapely.speedups
# %matplotlib inline
# File paths
border_fp = "data/Helsinki_borders.shp"
grid_fp = "data/TravelTimes_to_5975375_RailwayStation.shp"
# Read files
grid = gpd.read_file(grid_fp)
hel = gpd.read_file(border_fp)
# -
# - Visualize the layers:
# Plot the layers
ax = grid.plot(facecolor='gray')
hel.plot(ax=ax, facecolor='None', edgecolor='blue')
# Here the grey area is the Travel Time Matrix grid (13231 grid squares) that covers the Helsinki region, and the blue area represents the municipality of Helsinki. Our goal is to conduct an overlay analysis and select the geometries from the grid polygon layer that intersect with the Helsinki municipality polygon.
#
# When conducting overlay analysis, it is important to check that the CRS of the layers match!
#
# - Check if Helsinki polygon and the grid polygon are in the same crs:
# Ensure that the CRS matches, if not raise an AssertionError
assert hel.crs == grid.crs, "CRS differs between layers!"
# Indeed, they do. Hence, the pre-requisite to conduct spatial operations between the layers is fullfilled (also the map we plotted indicated this).
#
# - Let's do an overlay analysis and create a new layer from polygons of the grid that `intersect` with our Helsinki layer. We can use a function called `overlay()` to conduct the overlay analysis that takes as an input 1) the GeoDataFrame where the selection is taken, 2) the GeoDataFrame used for making the selection, and 3) parameter `how` that can be used to control how the overlay analysis is conducted (possible values are `'intersection'`, `'union'`, `'symmetric_difference'`, `'difference'`, and `'identity'`):
intersection = gpd.overlay(grid, hel, how='intersection')
# - Let's plot our data and see what we have:
intersection.plot(color="b")
# As a result, we now have only those grid cells that intersect with the Helsinki borders. As we can see **the grid cells are clipped based on the boundary.**
#
# - Whatabout the data attributes? Let's see what we have:
#
print(intersection.head())
# As we can see, due to the overlay analysis, the dataset contains the attributes from both input layers.
#
# - Let's save our result grid as a GeoJSON file that is commonly used file format nowadays for storing spatial data.
#
# +
# Output filepath
outfp = "data/TravelTimes_to_5975375_RailwayStation_Helsinki.geojson"
# Use GeoJSON driver
intersection.to_file(outfp, driver="GeoJSON")
# -
# There are many more examples for different types of overlay analysis in [Geopandas documentation](http://geopandas.org/set_operations.html) where you can go and learn more.
# ## Aggregating data
#
# Data aggregation refers to a process where we combine data into groups. When doing spatial data aggregation, we merge the geometries together into coarser units (based on some attribute), and can also calculate summary statistics for these combined geometries from the original, more detailed values. For example, suppose that we are interested in studying continents, but we only have country-level data like the country dataset. If we aggregate the data by continent, we would convert the country-level data into a continent-level dataset.
#
# In this tutorial, we will aggregate our travel time data by car travel times (column `car_r_t`), i.e. the grid cells that have the same travel time to Railway Station will be merged together.
#
# - For doing the aggregation we will use a function called `dissolve()` that takes as input the column that will be used for conducting the aggregation:
#
# +
# Conduct the aggregation
dissolved = intersection.dissolve(by="car_r_t")
# What did we get
print(dissolved.head())
# -
# - Let's compare the number of cells in the layers before and after the aggregation:
print('Rows in original intersection GeoDataFrame:', len(intersection))
print('Rows in dissolved layer:', len(dissolved))
# Indeed the number of rows in our data has decreased and the Polygons were merged together.
#
# What actually happened here? Let's take a closer look.
#
# - Let's see what columns we have now in our GeoDataFrame:
print(dissolved.columns)
# As we can see, the column that we used for conducting the aggregation (`car_r_t`) can not be found from the columns list anymore. What happened to it?
#
# - Let's take a look at the indices of our GeoDataFrame:
print(dissolved.index)
# Aha! Well now we understand where our column went. It is now used as index in our `dissolved` GeoDataFrame.
#
# - Now, we can for example select only such geometries from the layer that are for example exactly 15 minutes away from the Helsinki Railway Station:
# Select only geometries that are within 15 minutes away
dissolved.iloc[15]
# See the data type
print(type(dissolved.iloc[15]))
# See the data
print(dissolved.iloc[15].head())
# As we can see, as a result, we have now a Pandas `Series` object containing basically one row from our original aggregated GeoDataFrame.
#
# Let's also visualize those 15 minute grid cells.
#
# - First, we need to convert the selected row back to a GeoDataFrame:
# Create a GeoDataFrame
selection = gpd.GeoDataFrame([dissolved.iloc[15]], crs=dissolved.crs)
# - Plot the selection on top of the entire grid:
# Plot all the grid cells, and the grid cells that are 15 minutes a way from the Railway Station
ax = dissolved.plot(facecolor='gray')
selection.plot(ax=ax, facecolor='red')
# ## Simplifying geometries
# Sometimes it might be useful to be able to simplify geometries. This could be something to consider for example when you have very detailed spatial features that cover the whole world. If you make a map that covers the whole world, it is unnecessary to have really detailed geometries because it is simply impossible to see those small details from your map. Furthermore, it takes a long time to actually render a large quantity of features into a map. Here, we will see how it is possible to simplify geometric features in Python.
#
# As an example we will use data representing the Amazon river in South America, and simplify it's geometries.
#
# - Let's first read the data and see how the river looks like:
# +
import geopandas as gpd
# File path
fp = "data/Amazon_river.shp"
data = gpd.read_file(fp)
# Print crs
print(data.crs)
# Plot the river
data.plot();
# -
# The LineString that is presented here is quite detailed, so let's see how we can generalize them a bit. As we can see from the coordinate reference system, the data is projected in a metric system using [Mercator projection based on SIRGAS datum](http://spatialreference.org/ref/sr-org/7868/).
#
# - Generalization can be done easily by using a Shapely function called `.simplify()`. The `tolerance` parameter can be used to adjusts how much geometries should be generalized. **The tolerance value is tied to the coordinate system of the geometries**. Hence, the value we pass here is 20 000 **meters** (20 kilometers).
#
#
# +
# Generalize geometry
data2 = data.copy()
data2['geom_gen'] = data2.simplify(tolerance=20000)
# Set geometry to be our new simlified geometry
data2 = data2.set_geometry('geom_gen')
# Plot
data2.plot()
# +
# plot them side-by-side
# %matplotlib inline
import matplotlib.pyplot as plt
#basic config
fig, (ax1,ax2) = plt.subplots(nrows=1, ncols=2, figsize=(20, 16))
#ax1, ax2 = axes
#1st plot
ax1 = data.plot(ax=ax1, color='red', alpha=0.5)
ax1.set_title('Original')
#2nd plot
ax2 = data2.plot(ax=ax2, color='orange', alpha=0.5)
ax2.set_title('Generalize')
fig.tight_layout()
# -
# Nice! As a result, now we have simplified our LineString quite significantly as we can see from the map.
| geometric-operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + deletable=true editable=true
from IPython.display import Image
import sympy as sp
import math
import numpy as np
import datetime
# + deletable=true editable=true
Image(filename='/Users/wy/Desktop/beales_function.png')
# + deletable=true editable=true
class GoldSearch(object):
def __init__(self):
self.l = 10**-5
self.alpha = (math.sqrt(5)-1)/2.
def g_lambda(self, a, b):
return a+(1-self.alpha)*(b-a)
def g_mu(self, a, b):
return a+self.alpha*(b-a)
def goldSearch(self, a, b,lambda_k,mu_k,function,k = 1):
# step1
if (b - a) < self.l:
return (a+b)/2.
if function(lambda_k) > function(mu_k):
# step2
a = lambda_k
b = b
lambda_k = mu_k
mu_k = self.g_mu(a,b)
k = k+1
return self.goldSearch(a,b,lambda_k,mu_k,function,k)
elif function(lambda_k) <= function(mu_k):
# step3
a = a
b = mu_k
mu_k = lambda_k
lambda_k = self.g_lambda(a,b)
k = k+1
return self.goldSearch(a,b,lambda_k,mu_k,function,k)
GoldSearch = GoldSearch()
# + deletable=true editable=true
def gradient(f):
return [sp.lambdify((x1,x2), f.diff(x, 1), 'numpy') for x in [x1,x2]]
# + [markdown] deletable=true editable=true
# # Fletcher_Reeves
# 初始點 (1,1)
# GoldSearch interval -5 ~ 5
# e = 10**-5
# number of iterations : 24
# run time : 0.91s
# + deletable=true editable=true
def Fletcher_Reeves(f,xj):
lambda_j = sp.symbols('lambda_j')
e = 10**-5
sj = np.array(map(lambda fun : fun( xj[0],xj[1] ),gradient(f)))*(-1)
i = 1
while np.linalg.norm(sj) > e:
i = i+1
tmp = xj+lambda_j*sj
new_f = f.subs([(x1,tmp[0]),(x2,tmp[1])])
lambdaJ = GoldSearch.goldSearch(a,b,GoldSearch.g_lambda(a,b),GoldSearch.g_mu(a,b),sp.lambdify(lambda_j , new_f))
xj_1 = xj+lambdaJ*sj
sj_1 = np.array(map(lambda fun : fun( xj_1[0],xj_1[1] ),gradient(f)))*(-1)
beta_j = np.dot(sj_1.T,sj_1)/np.dot(sj.T,sj)
sj_1 = sj_1+beta_j*sj
sj = sj_1
xj = xj_1
return xj_1,i
# + deletable=true editable=true
a = -5
b = 5
x1,x2 = sp.symbols('x1,x2')
f = (1.5-x1*(1-x2))**2 + (2.25-x1*(1-x2**2))**2 + (2.625-x1*(1-x2**3))**2
# 初始點
xj = np.array([1,1])
start = datetime.datetime.now()
xj_1,i = Fletcher_Reeves(f,xj)
end = datetime.datetime.now()
print xj_1
print i
print end - start
# + [markdown] deletable=true editable=true
# # DFP
# 初始點 (1,1)
# GoldSearch interval -5 ~ 5
# e = 10**-5
# number of iterations : 8
# run time : 0.34s
# + deletable=true editable=true
def DFP(f,xi):
lambda_i = sp.symbols('lambda_i')
e = 10**-3
gradient_f = (np.array(map(lambda fun : fun( xi[0],xi[1] ),gradient(f)))).reshape(2,1)
Bi = np.identity(2)
i = 0
while abs(np.linalg.norm(gradient_f)) > e:
i = i+1
si = (np.dot(Bi,gradient_f)*(-1)).reshape(1,2)[0]
tmp = xi+lambda_i*si
new_f = f.subs([(x1,tmp[0]),(x2,tmp[1])])
lambdaI = GoldSearch.goldSearch(a,b,GoldSearch.g_lambda(a,b),GoldSearch.g_mu(a,b),sp.lambdify(lambda_i , new_f))
xi_1 = xi+lambdaI*si
gradient_f_1 = (np.array(map(lambda fun : fun( xi_1[0],xi_1[1] ),gradient(f)))).reshape(2,1)
if abs(np.linalg.norm(gradient_f_1)) > e:
gi = (gradient_f_1 - gradient_f).reshape(1,2)[0]
Mi = (np.dot(si.reshape(2,1),si.reshape(2,1).T))*lambdaI/np.dot(si.T,gi)
Ni = np.dot(np.dot(Bi,gi).reshape(2,1),np.dot(Bi,gi).T.reshape(1,2))*(-1)/np.dot(np.dot(gi.T,Bi),gi)
Bi = Bi+Mi+Ni
xi = xi_1
gradient_f = (np.array(map(lambda fun : fun( xi[0],xi[1] ),gradient(f)))).reshape(2,1)
else:
return xi_1,i
# + deletable=true editable=true
a = -5
b = 5
x1,x2 = sp.symbols('x1,x2')
f = (1.5-x1*(1-x2))**2 + (2.25-x1*(1-x2**2))**2 + (2.625-x1*(1-x2**3))**2
xi = np.array([1,1])
start = datetime.datetime.now()
xi_1,i = DFP(f,xi)
end = datetime.datetime.now()
print xi_1
print i
print end - start
# + [markdown] deletable=true editable=true
# # BFGS
# 初始點 (1,1)
# GoldSearch interval -5 ~ 5
# e = 10**-5
# number of iterations : 8
# run time : 0.38s
# + deletable=true editable=true
def BFGS(f,xi):
lambda_i = sp.symbols('lambda_i')
e = 10**-3
gradient_f = (np.array(map(lambda fun : fun( xi[0],xi[1] ),gradient(f)))).reshape(2,1)
Bi = np.identity(2)
i = 0
while abs(np.linalg.norm(gradient_f)) > e:
i = i+1
si = (np.dot(Bi,gradient_f)*(-1)).reshape(1,2)[0]
tmp = xi+lambda_i*si
new_f = f.subs([(x1,tmp[0]),(x2,tmp[1])])
lambdaI = GoldSearch.goldSearch(a,b,GoldSearch.g_lambda(a,b),GoldSearch.g_mu(a,b),sp.lambdify(lambda_i , new_f))
xi_1 = xi+lambdaI*si
gradient_f_1 = (np.array(map(lambda fun : fun( xi_1[0],xi_1[1] ),gradient(f)))).reshape(2,1)
if abs(np.linalg.norm(gradient_f_1)) > e:
gi = (gradient_f_1 - gradient_f).reshape(1,2)[0]
di = xi_1-xi
Mi = ((1 + np.dot(np.dot(gi.T,Bi),gi)/np.dot(di.T,gi))*np.dot(di.reshape(2,1),di.reshape(1,2)))/np.dot(di.T,gi)
Ni = np.dot(np.dot(di.reshape(2,1),gi.reshape(1,2)),Bi)*(-1)/np.dot(di.T,gi)
Qi = np.dot(np.dot(Bi,gi).reshape(2,1),di.reshape(1,2))*(-1)/np.dot(di.T,gi)
Bi = Bi+Mi+Ni+Qi
xi = xi_1
gradient_f = (np.array(map(lambda fun : fun( xi[0],xi[1] ),gradient(f)))).reshape(2,1)
else:
return xi_1,i
# + deletable=true editable=true
a = -5
b = 5
x1,x2 = sp.symbols('x1,x2')
f = (1.5-x1*(1-x2))**2 + (2.25-x1*(1-x2**2))**2 + (2.625-x1*(1-x2**3))**2
xi = np.array([1,1])
start = datetime.datetime.now()
xi_1,i = BFGS(f,xi)
end = datetime.datetime.now()
print xi_1
print i
print end - start
# + deletable=true editable=true
from scipy.optimize import fmin
def fun(X):
return (1.5-X[0]*(1-X[1]))**2 + (2.25-X[0]*(1-X[1]**2))**2 + (2.625-X[0]*(1-X[1]**3))**2
fmin(fun,np.array([1,1]))
# + [markdown] deletable=true editable=true
# # scipy python做科學計算的lib
# 出處 : http://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.fmin.html#scipy.optimize.fmin
# Minimize a function using the downhill simplex algorithm.
# This algorithm only uses function values, not derivatives or second derivatives.
| Fletcher_Reeves, DFP, BFGS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creation of a contact sequence
#
# In this second notebook, we will manually create a contact sequence from a predefined gait. Then, we will add some centroidal data to the contact sequence and export it.
#
# ## Contact plan
#
# In this section we will create a contact sequence for a bipedal robot, with a gait alterning double support phases and single support phases, for a simple walking motion.
#
# First, we need to create the first contact phase: a phase with both feet in contact with a flat floor at `z=0`. All the values here are taken for the robot Talos.
# +
# Import the required lib
import numpy as np
from pinocchio import SE3
from multicontact_api import ContactType, ContactModel, ContactPatch, ContactPhase, ContactSequence
#Define the name of the contacts.
# As explained in the previous notebook, a good practice is to use the names of the frames as defined in the urdf
rf_name = 'leg_right_sole_fix_joint'
lf_name = 'leg_left_sole_fix_joint'
OFFSET_Y = 0.085 # the position along the y axis of the feet in the reference configuration
# Create a contact phase:
p0 = ContactPhase()
# Define the placement of each contact:
placement_rf = SE3.Identity()
placement_lf = SE3.Identity()
translation_rf = np.array([0, -OFFSET_Y, 0])
translation_lf = np.array([0, OFFSET_Y, 0])
placement_rf.translation = translation_rf
placement_lf.translation = translation_lf
# Add both contacts to the contact phase:
p0.addContact(rf_name, ContactPatch(placement_rf))
p0.addContact(lf_name, ContactPatch(placement_lf))
print("First phase: \n", p0)
# -
# As you can see in the print, a lot of data in this contact phase are undefined, we will fill this data.
# Now we can create an empty contact sequence and set this phase as the first one of the sequence:
cs = ContactSequence()
cs.append(p0)
print("Current size of the sequence : ", cs.size())
# Now we can add more phases to define the walking motion. The final contact plan will consist of 5 steps for each leg of 20cm forward (with the first and last step only 10cm), thus moving the robot of 1m forward. Let's create the first step, remenber that in our formulation there should only be one contact variation (creation OR removing) between each adjacent phases, one step is thus two contact phases: single support and double support.
# +
# First, create a new phase where we break the right feet contact:
p1 = ContactPhase(p0) # copy the previous double support phase
p1.removeContact(rf_name)
# Now, add it to the sequence:
cs.append(p1)
# Then, create the second double support phase by creating a new contact with the right foot:
placement_rf = SE3.Identity()
translation_rf[0] = 0.1 # move 10cm along the x axis
placement_rf.translation = translation_rf
p2 = ContactPhase(p1) # copy the previous phase
p2.addContact(rf_name, ContactPatch(placement_rf))
# Now, add it to the sequence:
cs.append(p2)
# Lets print the result:
print("number of contact phases in the contact sequence : ", cs.size())
#first phase:
print("# Right feet contact of phase 0: ", cs.contactPhases[0].isEffectorInContact(rf_name))
print(cs.contactPhases[0].contactPatch(rf_name).placement)
print("# Left feet contact of phase 0: ", cs.contactPhases[0].isEffectorInContact(lf_name))
print(cs.contactPhases[0].contactPatch(lf_name).placement)
#second phase:
print("# Right feet contact of phase 1: ", cs.contactPhases[1].isEffectorInContact(rf_name))
print("# Left feet contact of phase 1: ", cs.contactPhases[1].isEffectorInContact(lf_name))
print(cs.contactPhases[1].contactPatch(lf_name).placement)
#Third phase:
print("# Right feet contact of phase 2: ", cs.contactPhases[2].isEffectorInContact(rf_name))
print(cs.contactPhases[2].contactPatch(rf_name).placement)
print("# Left feet contact of phase 2: ", cs.contactPhases[2].isEffectorInContact(lf_name))
print(cs.contactPhases[2].contactPatch(lf_name).placement)
# -
# As expected we now have a contact sequence with 3 phases: a double support, a single support and a double support. The code above is quite verbose, fortunately several helper methods exist to achieve the same result easier. Lets create the second step with the left foot with this helpers:
# +
# This method add a new contact phase to the sequence,
# # copy the contacts from the previous phase exept the one specified
cs.breakContact(lf_name)
translation_lf[0] = 0.2 # move 20cm forward
placement_lf.translation = translation_lf
# This method add a new contact phase to the sequence,
# # copy the contacts from the previous phase and add the one specified
cs.createContact(lf_name, ContactPatch(placement_lf))
print("Current contact sequence size: ", cs.size())
# -
# For the next steps, we will use another helper usefull for gaited motion. This method is used to "reposition" a contact and automatically add the intermediate contact phase with the broken contact.
# +
# First define the step length:
displacement = SE3.Identity()
displacement.translation = np.array([0.2, 0, 0]) # 20cm forward
for _ in range(4):
cs.moveEffectorOf(rf_name, displacement)
cs.moveEffectorOf(lf_name, displacement)
# add the last step of only 10cm to end the motion with both feet side by side:
displacement.translation = np.array([0.1, 0, 0])
cs.moveEffectorOf(rf_name, displacement)
print("Final contact sequence size: ", cs.size())
print("Right foot position at the end of the motion: \n",
cs.contactPhases[-1].contactPatch(rf_name).placement.translation)
print("Left foot position at the end of the motion: \n",
cs.contactPhases[-1].contactPatch(lf_name).placement.translation)
# -
# At this point the contact sequence define a consistent contact plan, we can check this with the following method:
cs.haveConsistentContacts()
# This method check that there is no discontinuities between the phases, and that there is always 1 contact variation between each phases.
#
#
# ### Additionnal data
#
# #### Contact model
#
# The Contact phases created do not specify any contact model (see the previous notebook for more information about this). We can check that the contact model are indeed not defined with the following code:
#
print("Friction coefficient defined for all contacts: ", cs.haveFriction())
print("Contact models defined for all contacts: ", cs.haveContactModelDefined())
# We can create a contact model specific to the robot feet and assign it to all the contact patches of all phases with this code:
# +
# Create a contact model with a friction coefficient of 0.5 and the PLANAR type
contact_model = ContactModel(0.5, ContactType.CONTACT_PLANAR)
# Define 4 contacts points at the corners of a rectangle:
contact_model.num_contact_points = 4
lx = 0.2 / 2. # half size of the feet along x axis
ly = 0.13 / 2. # half size of the feet along y axis
contact_points = np.zeros([3,4])
contact_points[0, :] = [-lx, -lx, lx, lx]
contact_points[1, :] = [-ly, ly, -ly, ly]
contact_model.contact_points_positions = contact_points
# Now, add this model to all patches of all phases:
for phase in cs.contactPhases:
for ee_name in phase.effectorsInContact():
phase.contactPatch(ee_name).contact_model = contact_model
print("Friction coefficient defined for all contacts: ", cs.haveFriction())
print("Contact models defined for all contacts: ", cs.haveContactModelDefined())
# -
# #### Phase duration
#
# As explained in the previous notebook, a contact phase may be defined on a specific time interval. We can check if this is correctly defined for all the phases and that the timings are consistent with the following method:
cs.haveTimings()
# The code below set the duration for all the phases, depending on the number of contacts of this phase. We are going to define long phase duration here because in the next section we are going to generate a quasi-static centroidal reference.
# +
DURATION_SS = 2. # duration of the single support phases
DURATION_DS = 4. # duration of the double support phases
for i, phase in enumerate(cs.contactPhases):
if i == 0:
phase.timeInitial = 0.
else:
# Set the initial time as the final time of the previous phase
phase.timeInitial = cs.contactPhases[i-1].timeFinal
# set the duration of the phase based on the number of contacts
if phase.numContacts() == 1:
phase.duration = DURATION_SS
elif phase.numContacts() == 2:
phase.duration = DURATION_DS
else:
raise RuntimeError("Incorrect number of contacts for the phase " + str(i))
# Check that the timings are correctly set :
print("Contact sequence have consistent timings: ", cs.haveTimings())
# -
# ## Centroidal data
#
# Now that the contact sequence correctly define a contact plan, we are going to store centroidal data to the sequence: the center of mass position, velocity and acceleration trajectory.
# As this notebook is about the multicontact_api package and not about centroidal trajectory optimization, we are going to use really simple and quasi-static trajectory:
#
# During the single support phases, the CoM is fixed above the center of the support polygon. During the double support phases, the CoM will go from the previous support polygon to the next one in a straight line (starting and ending with a null velocity and acceleration).
#
# First, we need to compute the initial and final CoM position for each phases, the CoM velocity and acceleration are initialized to 0 by defaut so we do not need to modify it here.
# +
# Define the CoM height:
COM_HEIGHT = 0.85
for i, phase in enumerate(cs.contactPhases):
if i == 0:
# Define the initial CoM position:
cs.contactPhases[0].c_init = np.array([0, 0, COM_HEIGHT])
elif phase.numContacts() == 1:
# Single support phase: set the CoM position above the feet in contact:
com = phase.contactPatch(phase.effectorsInContact()[0]).placement.translation
com[2] += COM_HEIGHT
phase.c_init = com
# The CoM is not moving during single support phase, so we set the same position as the final point
phase.c_final = com
# Set the final point of the previous double support phase to be the same as this position
cs.contactPhases[i-1].c_final = com
elif phase.numContacts() == 2:
# Double support phase:
# set the initial CoM position to be equal to the final position of the previous phase
phase.c_init = cs.contactPhases[i-1].c_final
else:
raise RuntimeError("Incorrect number of contacts for the phase " + str(i))
# For the final phase: set the final position between the feets:
com = (phase.contactPatch(rf_name).placement.translation + phase.contactPatch(lf_name).placement.translation) / 2.
com[2] += COM_HEIGHT
phase.c_final = com
print("Final CoM position: ",cs.contactPhases[-1].c_final)
# -
# We can check that the position for all the phases have been set and that the initial position of each phase match the final position of the previous phase like this:
cs.haveCentroidalValues()
# Now we can generate trajectories for each phases. For the single support phases we will use Constant trajectories. For the double support phases we will use fifth order polynomials that connect the initial and final position and have a null initial and final velocity and acceleration.
#
# As explained in the previous notebook, this trajectories must be represented with objects from the `NDCurves` package.
#
# +
from ndcurves import polynomial
for phase in cs.contactPhases:
if phase.numContacts() == 1:
# Single support phase: build a constant trajectory at the CoM position:
phase.c_t = polynomial(phase.c_init, phase.timeInitial, phase.timeFinal) # Build a degree 0 polynomial curve
# Compute the derivate of this curve and store it in the phase
phase.dc_t = phase.c_t.compute_derivate(1)
phase.ddc_t = phase.dc_t.compute_derivate(1)
elif phase.numContacts() == 2:
# Double support phase: build a minJerk trajectory (5th degree) between the initial and final CoM position
phase.c_t = polynomial.MinimumJerk(phase.c_init, phase.c_final, phase.timeInitial, phase.timeFinal)
phase.dc_t = phase.c_t.compute_derivate(1)
phase.ddc_t = phase.dc_t.compute_derivate(1)
else:
raise RuntimeError("Incorrect number of contacts.")
# Check that all the phases have CoM trajectories:
print("Contact sequence have consistent CoM trajectories: ", cs.haveCOMtrajectories())
# -
# ## Conclusion
#
# In this notebook we saw how to manually build a `ContactSequence` from scratch. Adding several phases with contact placement and time interval. Then adding CoM trajectories for each phases.
#
# In the next notebook we will se how to use the data inside a contact sequence, plot it and display it on a 3D viewer.
#
# The last code below show how to export the ContactSequence build in this notebook to a file.
# Serialize the contact sequence
cs.saveAsBinary("notebook2.cs")
| notebooks/2_contact_sequence_creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # File Input Output
# Persistent Data Storage
# Types of Files: text file, binary file
#
# Text File: stores data as a stream of characters, each line ends with a new line character. typical extension of
# plain text file is .txt
# CSV File: A type of text file that stores tabular data.
# Each line is one observation of multiple values separated by comma.
# CSV stands for comma separated values. extension: .csv
# Binary File: Can store characters, numeric data as well as objects of all types.
# Typical extensions:.bin,.dat
#
# 
#
# <center>Figure:On the left is a text file opened in notepad while on the right is a binary file opened in notepad</center>
#
# ## Sequence of file operations:
# 1. Open the file: open(file,mode)
# Arguments
# file: filepath (relative: from working directory or absolute)
# mode: described in table below.
# Returns
# file object
# 
#
# 2. Write data (from main memory/RAM) to file or read from file
# 3. Close the file : fileobj.close() frees up the resources
outfile=open("util_files/test.txt","w")
outfile.write("test data\ntest2\ntest3")
outfile.close()
infile=open("util_files/test.txt")#default mode is "r"
print(infile.readline())
infile.close()
# ## Closing a file, exceptions, with statement
# When a File is closed, the memory taken up by it is released
infile=open("util_files/test.txt")#default mode is "r"
print(infile.fileno())
infile.close()
print(infile.fileno())
# However if an exception occurs after file is opened and before it is closed, the object keeps on taking up the memory
infile=open("util_files/test.txt")#default mode is "r"
print(infile.readline())
x=10/0
infile.close()
print(infile.fileno())
# Solution: <b>with </b> statement
# +
with open("util_files/test.txt") as infile:
print(infile.fileno())
print(infile.fileno())
# -
with open("util_files/test.txt") as infile:
print(infile.fileno())
7/0
print(infile.fileno()) #Resource freed despite exception
# ## Four methods to read data:
# 1. loop through each line of file
# or use following methods of file object:
# 2. read(): reads entire file into a string
# 3. readline(): reads next line as a string
# 4. readlines(): reads all the lines and returns them as a list
# Word of caution: use readline or loop so that if the file is too large, you don't run out of memory reading it all at once.
with open("util_files/test.txt") as infile:
x=infile.read()
print(type(x))
print(x)
with open("util_files/test.txt") as infile:
x=infile.readlines()
print(type(x))
print(x)
print(x[0])
with open("util_files/test.txt") as infile:
x=infile.readline()
print(type(x))
print(x)
x=infile.readline()
print(type(x))
print(x)
with open("util_files/test.txt") as infile:
for line in infile:
print(type(line))
print(line)
# ## Special example: read and write to file via lists
#
# #### 1. Always convert non-string data to string before writing to text file
years=[1995, 2000, 2019]
with open("util_files/years.txt","w") as outfile:
for year in years:
outfile.write(year)
years=[1995, 2000, 2019]
with open("util_files/years.txt","w") as outfile:
for year in years:
outfile.write(str(year)+"\n")
# #### 2. Convert string read from file to whichever datatype is expected, eg: int
years=[]
with open("util_files/years.txt","r") as infile:
for line in infile:
years.append(int(line))
print(years)
# #### 3. If reading data as list of strings, take care of \n character
#\n may pose a problem:
lines=[]
with open("util_files/test.txt","r") as infile:
for line in infile:
lines.append(line)
print(lines)
#Solution:
lines=[]
with open("util_files/test.txt","r") as infile:
for line in infile:
line=line.replace("\n","")
lines.append(line)
print(lines)
# ## CSV Files
# Each line is a record of multiple columns (fields) separated by comma
# +
import csv
fields = []
rows = []
with open("util_files/mycsv.csv", 'r') as csvfile:
csvreader = csv.reader(csvfile)
# fields = csvreader.next()
for row in csvreader:
rows.append(row)
print("Total no. of rows: %d"%(csvreader.line_num))
print('Field names are:' + ', '.join(field for field in fields))
print('\nThe rows are:\n')
for row in rows[:5]:
for col in row:
print("%10s"%col),
print('\n')
# -
import csv
days=[["Mon,",1],["Tue",2],["Wed",3],["Thu",4],["Fri",5],["Sat",6],["Sun",0]]
with open("days.csv","w",newline="") as file: # setting newline argument to ""
#enables universal newline mode so r/w works correctly on all OS
writer=csv.writer(file) #CSV writer object converts data to comma separated values
writer.writerows(days) #writes the rows to file
days=[]
with open("days.csv",newline="") as file:
reader=csv.reader(file)
for day,id in reader:
days.append([day,int(id)])
days
# +
# Or
# for row in reader:
# days.append([row[0],int(row[1])])
# -
# <br> If the data itself contains commas, newline CSV module automatically handles this by adding quotes around these special characters before writing to file
# If the data contains quotes, or tab is used to separate fields in csv file we want to read from, we can specify optional arguments while reading/writing
# 
| 6. Files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from run_pplm_regress_train import Discriminator
from transformers import GPT2Tokenizer
from tqdm import tqdm_notebook as tqdm
pretrained_model = 'gpt2-medium'
discriminator = Discriminator(
pretrained_model=pretrained_model,
cached_mode=False,
device='cpu',
reg_type=1
).to('cpu')
discriminator.regressor_head.load_state_dict(sd)
tokenizer = GPT2Tokenizer.from_pretrained(pretrained_model)
tokenized_ = tokenizer.encode("Hello, how are you doing today? I hadn't heard from you in a while!")
test_sentences = list()
test_labels = list()
with open('../../data/div/test.tsv') as f:
for line in f:
split = line.split('\t')
test_labels.append(float(split[0].strip()))
test_sentences.append(split[1])
predicted_lengths = list()
for sentence in tqdm(test_sentences):
tokenized = tokenizer.encode("A man")
predicted = discriminator(torch.tensor([tokenized]))
predicted_lengths.append(predicted)
predicted_lengths
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Introduction to Nilearn and image manipulation
#
# The goal of this notebook is to help get you comfortable with manipulating functional and anatomical images using nilearn. We'll be using the techniques we learned here in our final analysis...
#
# #### Content:
# 1. Basic Image Operations and Masking
# 2. Resampling data to work across modalities (T1/FUNC)
# Notice that we imported two things:
# 1. `image as nimg` - allows us to load NIFTI images using nibabel under the hood
# 2. `plotting as nplot`- allows us to using Nilearn's plotting library for easy visualization
#
# First let's grab some data from where we downloaded our **FMRIPREP** outputs. Note that we're using the argument <code>return_type='file'</code> so that pyBIDS gives us file paths directly rather than the standard BIDSFile objects
#
#Base directory for fmriprep output
# Here we used pyBIDS (as introduced in earlier sections) to pull a single participant. Specifically, we pulled all preprocessed (MNI152NLin2009cAsym, and T1w) anatomical files as well as their respective masks. Let's quickly view these files for review:
#
#Display preprocessed files inside of anatomy folder
# ## Basic Image Operations
#
# In this section we're going to deal with the following files:
#
# 1. <code>sub-10171_desc-preproc_T1w.nii.gz</code> - the T1 image in native space
# 2. <code>sub-10171_desc-brain_mask.nii.gz</code> - a mask with 1's representing the brain and 0's elsewhere.
# Using the <code>plotting</code> module (which we've aliased as <code>nplot</code>), we can view our MR image:
#
# This gives just a still image of the brain. We can also view the brain more interactively using the `view_img` function. It will require some additional settings however:
nplot.view_img(t1_img,
bg_img=False, # Disable using a standard image as the background
cmap='Greys_r', # Set color scale so white matter appears lighter than grey
symmetric_cmap=False, # We don't have negative values
threshold="auto", # Clears out the background
)
# Try clicking and dragging the image in each of the views that are generated!
# Try viewing the mask as well!
#View the mask image
# ### Arithmetic Operations
#
# Let’s start performing some image operations. The simplest operations we can perform is element-wise, what this means is that we want to perform some sort of mathematical operation on each voxel of the MR image. Since voxels are represented in a 3D array, this is equivalent to performing an operation on each element (i,j,k) of a 3D array. Let’s try inverting the image, that is, flip the colour scale such that all blacks appear white and vice-versa. To do this, we’ll use the method
#
# <code>nimg.math_img(formula, **imgs)</code> Where:
#
# - <code>formula</code> is a mathematical expression such as 'a+1'
# - </code>**imgs</code> is a set of key-value pairs linking variable names to images. For example a=T1
#
# In order to invert the image, we can simply flip the sign which will set the most positive elements (white) to the most negative elements (black), and the least positives elements (black) to the least negative elements (white). This effectively flips the colour-scale:
# Alternatively we don't need to first load in our <code>t1_img</code> using <code>img.load_img</code>. Instead we can feed in a path to <code>img.math_img</code>:
#
# ~~~
# invert_img = nimg.math_img('-a', a=t1)
# nplot.plot_anat(invert_img)
# ~~~
#
# This will yield the same result!
# ### Applying a Mask
# Let’s extend this idea of applying operations to each element of an image to multiple images. Instead of specifying just one image like the following:
#
# <code>nimg.math_img('a+1',a=img_a)</code>
#
# We can specify multiple images by tacking on additional variables:
#
# <code>nimg.math_img('a+b', a=img_a, b=img_b)</code>
#
# The key requirement here is that when dealing with multiple images, that the size of the images must be the same. The reason being is that we’re deaing with element-wise operations. That means that some voxel (i,j,k) in img_a is being paired with some voxel (i,j,k) in <code>img_b</code> when performing operations. So every voxel in <code>img_a</code> must have some pair with a voxel in <code>img_b</code>; sizes must be the same.
#
# We can take advantage of this property when masking our data using multiplication. Masking works by multipling a raw image (our <code>T1</code>), with some mask image (our <code>bm</code>). Whichever voxel (i,j,k) has a value of 0 in the mask multiplies with voxel (i,j,k) in the raw image resulting in a product of 0. Conversely, any voxel (i,j,k) in the mask with a value of 1 multiplies with voxel (i,j,k) in the raw image resulting in the same value. Let’s try this out in practice and see what the result is:
# As you can see areas where the mask image had a value of 1 were retained, everything else was set to 0
# #### Exercise!
# Try applying the mask such that the brain is removed, but the rest of the head is intact!
#
# *Hint*:
#
# Remember that a mask is composed of 0's and 1's, where parts of the data labelled 1 are regions to keep, and parts of the data that are 0, are to throw away.
#
# You can do this in 2 steps:
#
# 1. Switch the 0's and 1's using an equation (simple addition/substraction) or condition (like x == 0).
# 2. Apply the mask
# Step 1: Switch the 0's and 1's
inverted_mask = img.math_img(??, x=bm)
plot.plot_anat(inverted_mask)
# Step 2: Apply the mask
inverted_mask_t1 = img.math_img(??, a=t1, b=inverted_mask)
plot.plot_anat(inverted_mask_t1)
# ### Slicing
#
# Recall that our data matrix is organized in the following manner:
#
# <img src="./static/images/numpy_arrays.png" alt="Drawing" align="middle" width="500px"/>
# Slicing does exactly what it seems to imply. Given our 3D volume, we can pull out a 2D subset (called a "slice"). Here's an example of slicing moving from left to right via an animation:
#
# <img src="https://upload.wikimedia.org/wikipedia/commons/5/56/Parasagittal_MRI_of_human_head_in_patient_with_benign_familial_macrocephaly_prior_to_brain_injury_%28ANIMATED%29.gif"/>
#
# What you see here is a series of 2D images that start from the left, and move toward the right. Each frame of this GIF is a slice - a 2D subset of a 3D volume. Slicing can be useful for cases in which you'd want to loop through each MR slice and perform a computation; importantly in functional imaging data slicing is useful for pulling out timepoints as we'll see later!
#
# ***
# Sourced from: https://en.wikipedia.org/wiki/Neuroimaging#/media/File:Parasagittal_MRI_of_human_head_in_patient_with_benign_familial_macrocephaly_prior_to_brain_injury_(ANIMATED).gif
#
# ***
# Slicing is done easily on an image file using the attribute <code>.slicer</code> of a Nilearn <code>image</code> object. For example we can grab the $10^{\text{th}}$ slice along the x axis as follows:
# The statement $10:11$ is intentional and is required by <code>.slicer</code>. Alternatively we can slice along the x-axis using the data matrix itself:
# This will yield the same result as above. Notice that when using the <code>t1_data</code> array we can just specify which slice to grab instead of using <code>:</code>.
#
#
# We can also use slicing in order to modify visualizations. For example, when viewing the T1 image, we may want to specify at which slice we'd like to view the image. This can be done by specifying which coordinates to *cut* the image at:
# The <code>cut_coords</code> option specifies 3 numbers:
# - The first number says cut the X coordinate at slice 50 and display (sagittal view in this case!)
# - The second number says cut the Y coordinate at slice 30 and display (coronal view)
# - The third number says cut the Z coordinate at slice 20 and display (axial view)
#
# Remember <code>plot.plot_anat</code> yields 3 images, therefore <code>cut_coords</code> allows you to display where to take cross-sections of the brain from different perspectives (axial, sagittal, coronal)
# ***
#
# This covers the basics of image manipulation using T1 images. To review in this section we covered:
#
# - Basic image arithmetic
# - Visualization
# - Slicing
#
# In the next section we will cover how to integrate additional modalities (functional data) to what we've done so far using <code>Nilearn</code>. Then we can start using what we've learned in order to perform analysis and visualization!
| code/03-basic_image_manipulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="N2_J4Rw2r0SQ" outputId="6d69d167-9ba2-438a-d7b2-64414a7d06a6"
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
# %matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
# + id="ZTJPyL45DiJ3"
m = 5 # 5, 10, 20, 100, 500, 1000
# + id="g-do5dZWK5L6"
desired_num = 4000
# + colab={"base_uri": "https://localhost:8080/"} id="XTHbSeGAK7k7" outputId="9e72c9e8-7d3f-48b5-9f59-6982ec7c8fc4"
tr_i = 0
tr_j = int(desired_num/2)
tr_k = desired_num
tr_i, tr_j, tr_k
# + [markdown] id="F6fjud_Fr0Sa"
# # Generate dataset
# + colab={"base_uri": "https://localhost:8080/"} id="CqdXHO0Cr0Sd" outputId="1c8d48d2-dfc5-4de8-e41f-3940ee9b0bb1"
np.random.seed(12)
y = np.random.randint(0,10,5000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
# + id="ddhXyODwr0Sk"
x = np.zeros((5000,2))
# + id="DyV3N2DIr0Sp"
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [4,6.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [5.5,6],cov=[[0.01,0],[0,0.01]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [4.5,4.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [3,3.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [2.5,5.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [3.5,8],cov=[[0.01,0],[0,0.01]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [5.5,8],cov=[[0.01,0],[0,0.01]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [7,6.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [6.5,4.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [5,3],cov=[[0.01,0],[0,0.01]],size=sum(idx[9]))
# + colab={"base_uri": "https://localhost:8080/"} id="qh1mDScsU07I" outputId="c2fdec60-754b-4f2e-9e37-6cff3a77e67d"
x[idx[0]][0], x[idx[5]][5]
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="hJ8Jm7YUr0St" outputId="bdd82ff5-160e-474a-acb2-4b13eed02fc5"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + colab={"base_uri": "https://localhost:8080/"} id="3lMBZEHNBlF2" outputId="9ea33ef5-05cd-4591-8b24-259c2341945f"
bg_idx = [ np.where(idx[3] == True)[0],
np.where(idx[4] == True)[0],
np.where(idx[5] == True)[0],
np.where(idx[6] == True)[0],
np.where(idx[7] == True)[0],
np.where(idx[8] == True)[0],
np.where(idx[9] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
# + colab={"base_uri": "https://localhost:8080/"} id="blRbGZHeCwXU" outputId="0de8c30a-6106-41cb-d695-d9dfd2db47dc"
np.unique(bg_idx).shape
# + id="Y43sWeX7C15F"
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/"} id="ooII7N6UDWe0" outputId="d9870039-c1d6-453d-bb5a-c29173d4d818"
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
# + id="g21bvPRYDL9k"
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/"} id="GtFvIeHsDZJk" outputId="c7579856-51f1-4bed-e338-ac2a324ee89d"
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="8-VLhUfDDeHt" outputId="d3f5a0ec-7641-4164-bc27-b7cf1db37a73"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="UfFHcZJOr0Sz"
foreground_classes = {'class_0','class_1', 'class_2'}
background_classes = {'class_3','class_4', 'class_5', 'class_6','class_7', 'class_8', 'class_9'}
# + colab={"base_uri": "https://localhost:8080/"} id="OplNpNQVr0S2" outputId="0d1ea896-e669-4cfe-d321-fa967aac0a67"
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
# + colab={"base_uri": "https://localhost:8080/"} id="OoxzYI-ur0S_" outputId="4698b427-b7d6-48a8-e6ff-97a71760e98e"
np.reshape(a,(2*m,1))
# + id="jqbvfbwVr0TN"
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
for j in range(desired_num):
np.random.seed(j)
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,m)
a = []
for i in range(m):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(np.reshape(a,(2*m,1)))
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
# + colab={"base_uri": "https://localhost:8080/"} id="YzJPNP2mFwAG" outputId="29af77ec-68dd-4be3-a9fb-13ac0659c097"
mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
mosaic_list_of_images.shape
# + colab={"base_uri": "https://localhost:8080/"} id="2aIPMgLXNiXW" outputId="e10855e3-dc6f-481d-e9c6-542b463a0513"
mosaic_list_of_images.shape, mosaic_list_of_images[0]
# + colab={"base_uri": "https://localhost:8080/"} id="A3qcsbbzPfRG" outputId="5580b13d-8917-4376-cebb-43306a38dc98"
for j in range(m):
print(mosaic_list_of_images[0][2*j:2*j+2])
# + id="iPoIwbMHx44n"
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number, m):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
cnt = 0
counter = np.zeros(m) #np.array([0,0,0,0,0,0,0,0,0])
for i in range(len(mosaic_dataset)):
img = torch.zeros([2], dtype=torch.float64)
np.random.seed(int(dataset_number*10000 + i))
give_pref = foreground_index[i] #np.random.randint(0,9)
# print("outside", give_pref,foreground_index[i])
for j in range(m):
if j == give_pref:
img = img + mosaic_dataset[i][2*j:2*j+2]*dataset_number/m #2 is data dim
else :
img = img + mosaic_dataset[i][2*j:2*j+2]*(m-dataset_number)/((m-1)*m)
if give_pref == foreground_index[i] :
# print("equal are", give_pref,foreground_index[i])
cnt += 1
counter[give_pref] += 1
else :
counter[give_pref] += 1
avg_image_dataset.append(img)
print("number of correct averaging happened for dataset "+str(dataset_number)+" is "+str(cnt))
print("the averaging are done as ", counter)
return avg_image_dataset , labels , foreground_index
# + colab={"base_uri": "https://localhost:8080/"} id="30ZAjix3x8CM" outputId="cbd5439d-abb1-4041-d9de-0934e39ff148"
avg_image_dataset_1 , labels_1, fg_index_1 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:tr_j], mosaic_label[0:tr_j], fore_idx[0:tr_j] , 1, m)
test_dataset , labels , fg_index = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[tr_j : tr_k], mosaic_label[tr_j : tr_k], fore_idx[tr_j : tr_k] , m, m)
# + colab={"base_uri": "https://localhost:8080/"} id="0dYXnywAD-4l" outputId="c44b7e26-d31f-47e3-b008-7148c670e59e"
avg_image_dataset_1 = torch.stack(avg_image_dataset_1, axis = 0)
# avg_image_dataset_1 = (avg - torch.mean(avg, keepdims= True, axis = 0)) / torch.std(avg, keepdims= True, axis = 0)
# print(torch.mean(avg_image_dataset_1, keepdims= True, axis = 0))
# print(torch.std(avg_image_dataset_1, keepdims= True, axis = 0))
print("=="*40)
test_dataset = torch.stack(test_dataset, axis = 0)
# test_dataset = (avg - torch.mean(avg, keepdims= True, axis = 0)) / torch.std(avg, keepdims= True, axis = 0)
# print(torch.mean(test_dataset, keepdims= True, axis = 0))
# print(torch.std(test_dataset, keepdims= True, axis = 0))
print("=="*40)
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="bT9-kEI7NAnR" outputId="4f0c80c4-cf8f-4f86-8cd8-30f2fdc4895b"
x1 = (avg_image_dataset_1).numpy()
y1 = np.array(labels_1)
plt.scatter(x1[y1==0,0], x1[y1==0,1], label='class 0')
plt.scatter(x1[y1==1,0], x1[y1==1,1], label='class 1')
plt.scatter(x1[y1==2,0], x1[y1==2,1], label='class 2')
plt.legend()
plt.title("dataset4 CIN with alpha = 1/"+str(m))
# + colab={"base_uri": "https://localhost:8080/", "height": 299} id="Sc8W2N7LQQ_l" outputId="cdb8569a-00aa-4ab1-deb6-461bb8a9d477"
x1 = (test_dataset).numpy() / m
y1 = np.array(labels)
plt.scatter(x1[y1==0,0], x1[y1==0,1], label='class 0')
plt.scatter(x1[y1==1,0], x1[y1==1,1], label='class 1')
plt.scatter(x1[y1==2,0], x1[y1==2,1], label='class 2')
plt.legend()
plt.title("test dataset4")
# + colab={"base_uri": "https://localhost:8080/"} id="pWVkaBZGXCRN" outputId="919023f1-d272-47bf-b3c4-8ae42168c4e6"
test_dataset[0:10]/m
# + colab={"base_uri": "https://localhost:8080/"} id="JDZLkEGbXGqg" outputId="1b82f913-44a9-480c-a0a4-fa587dab212b"
test_dataset = test_dataset/m
test_dataset[0:10]
# + id="yL0BRf8er0TX"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
# + colab={"base_uri": "https://localhost:8080/"} id="4KsrW9qL9xgS" outputId="2f147be7-cb10-413d-b4b1-2e137797409e"
avg_image_dataset_1[0].shape
avg_image_dataset_1[0]
# + id="EY2l62APygaV"
batch = 200
traindata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
# + id="9suUslCj8YZK"
testdata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
# + id="Nh3mBQHZ8bEj"
testdata_11 = MosaicDataset(test_dataset, labels )
testloader_11 = DataLoader( testdata_11 , batch_size= batch ,shuffle=False)
# + id="5_XeIUk0r0Tl"
class Whatnet(nn.Module):
def __init__(self):
super(Whatnet,self).__init__()
self.linear1 = nn.Linear(2,3)
# self.linear2 = nn.Linear(50,10)
# self.linear3 = nn.Linear(10,3)
torch.nn.init.xavier_normal_(self.linear1.weight)
torch.nn.init.zeros_(self.linear1.bias)
def forward(self,x):
# x = F.relu(self.linear1(x))
# x = F.relu(self.linear2(x))
x = (self.linear1(x))
return x
# + id="pjD2VZuV9Ed4"
def calculate_loss(dataloader,model,criter):
model.eval()
r_loss = 0
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
outputs = model(inputs)
loss = criter(outputs, labels)
r_loss += loss.item()
return r_loss/(i+1)
# + id="uALi25pmzQHV"
def test_all(number, testloader,net):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
pred = np.concatenate(pred, axis = 0)
out = np.concatenate(out, axis = 0)
print("unique out: ", np.unique(out), "unique pred: ", np.unique(pred) )
print("correct: ", correct, "total ", total)
print('Accuracy of the network on the %d test dataset %d: %.2f %%' % (total, number , 100 * correct / total))
# + id="4vmNprlPzTjP"
def train_all(trainloader, ds_number, testloader_list):
print("--"*40)
print("training on data set ", ds_number)
torch.manual_seed(12)
net = Whatnet().double()
net = net.to("cuda")
criterion_net = nn.CrossEntropyLoss()
optimizer_net = optim.Adam(net.parameters(), lr=0.001 ) #, momentum=0.9)
acti = []
loss_curi = []
epochs = 1000
running_loss = calculate_loss(trainloader,net,criterion_net)
loss_curi.append(running_loss)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
net.train()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_net.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion_net(outputs, labels)
# print statistics
running_loss += loss.item()
loss.backward()
optimizer_net.step()
running_loss = calculate_loss(trainloader,net,criterion_net)
if(epoch%200 == 0):
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.05:
print('epoch: [%d] loss: %.3f' %(epoch + 1,running_loss))
break
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %.2f %%' % (total, 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,net)
print("--"*40)
return loss_curi
# + id="Yl41sE8vFERk"
train_loss_all=[]
testloader_list= [ testloader_1, testloader_11]
# + colab={"base_uri": "https://localhost:8080/"} id="5gQoPST5zW2t" outputId="3905f9a6-fed0-4527-8cde-94918128fdc0"
train_loss_all.append(train_all(trainloader_1, 1, testloader_list))
# + id="In76SYH_zZHV"
# %matplotlib inline
# + id="BS4HtOHEzZ0E" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="8144536e-9503-43b1-992d-a192424c172d"
for i,j in enumerate(train_loss_all):
plt.plot(j,label ="dataset "+str(i+1))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="1UbTkfLUINTI"
| AAAI/Learnability/CIN/Linear/ds4/size_2000/synthetic_type4_Linear_m_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 키보드 단축키
#
# notebook을 사용하는 데 유용한 몇 가지의 단축키를 소개하고자 합니다.
#
# 가장 첫번째로 수정 모드(edit mode)와 실행 모드(command mode)를 변경하는 것입니다. 수정모드에서는 셀 내에서 코드를 입력할 수 있게 해 주고, 실행 모드는 새로운 셀을 만들거나 커맨드 팔레트를 열 수 있게 도와줍니다.
#
# 셀을 선택할 때, 우리는 박스의 색을 보고 어떤 셀이 선택되어 있는지를 확인할 수 있습니다. 수정 모드에서는 박스의 왼쪽 부분이 **녹색**으로 표시되고, 커서가 깜빡이는 것을 확인할 수 있습니다. 실행 모드에서는 **푸른색**으로 변합니다.
#
# 새로운 셀을 만들고 다음 셀로 넘어가면 기본적으로 실행 모드에 들어가게 됩니다. 수정 모드에 들어가기 위해서는 Enter/Return를 눌어야 합니다. 수정모드에서 실행모드로 이동하기 위해서는 ESC를 눌러야 합니다.
#
# > **실습하기:** 셀을 클릭하고 **Enter + Shift**를 눌러 다음 셀에 이동해 봅시다. 수정모드와 실행모드를 전환해 보세요.
# +
# 연습하기
# -
# ## 단축키 살펴보기
#
# 단축키를 살펴보고 싶다면, 실행 모드에서 **h**를 눌러보세요. 툴박스에서 'Help'를 눌러도 해결 가능합니다.
# ## 새로운 셀 만들기
#
# 가장 많이 쓰는 명령어는 새로운 셀을 만드는 것입니다. 실행 모드에서 **a**를 눌러 현재 셀 위에 새로운 셀을 만들 수 있습니다. **b**를 누른다면 현재 셀 바로 밑에 새로운 셀을 만들어 줍니다.
# > **실습하기:** 이 셀 위에 새로운 셀을 만들어보세요.
# > **실습하기:** 이 셀 아래에 새로운 셀을 만들어보세요.
# ## 마크다운과 코드 전환하기
#
# 키보드 단축키를 이용해 Markdown 과 code cells을 전환하는 것도 가능합니다. Markdown을 code cell로 바꾸려면 **y**를, 반대의 경우에는 **m**을 누르면 됩니다.
#
# > **실습하기:** 아래의 코드를 Markdown과 code cell로 전환시켜 보세요.
# +
## 실습하기
def fibo(n): # 피보나치 수열 (재귀함수)
if n == 0:
return 0
elif n == 1:
return 1
return fibo(n-1) + fibo(n-2)
# -
# ## 행 숫자 표시하기
#
# 코드셀에서 몇 번쨰 행인지 찾아보는 것이 필요하는 것은 매우 유용할 수 있습니다. 이때는 간단히 **L**을 눌러 표시해 볼 수 있습니다.
#
# > **실습하기:** 위의 코드에서 L을 눌러 라인을 표시해 보세요.
# ## Deleting cells
#
# 셀을 삭제할 때는 'd'를 두번 누르면 됩니다.
#
# > **실습하기:** 아래의 셀을 삭제해 봅시다.
# +
# 삭제해 주세요
# -
# ## notebook 저장하기
#
# notebook은 스스로 자동저장하나, 직접 저장하는 것이 현재의 상태를 가장 잘 보존할 수 있습니다. 저장하기 위해서는 **s**를 누르면 됩니다.
# ## 커맨드 팔레트
#
# **Shift + Control/Command + p** 를 눌러 팔레트 모드로 이동할 수 있습니다.
#
# > **Note:** 이 명령어는 파이어폭스에서는 실행되지 않습니다. 크롬과 사파리에서만 실행 가능합니다.
#
# 커맨드 팔레트를 불러옴으로서 우리는 필요한 단축키를 찾아볼 수 있습니다. 예를 들어 셀을 위, 아래로 옮기고 싶을 수 있지만, 이를 제공하는 단축키는 없습니다. 대신에 커맨드 팔레트의 'move'를 이용해 해당 기능을 실행해 볼 수 있습니다.
#
# > **실습하기:** 커맨드 팔레트를 활용해 아래 셀의 위치를 이동해 봅시다.
# +
# 이 셀을 이동시켜 봅시다.
# +
# 이 셀 밑으로 이동시켜 주세요.
# -
# ## 마치며
#
# notebook에는 복사, 잘라내기, 붙여넣기 등의 다양한 기능이 있습니다. Jupyter를 사용하면서 더 필요한 기능들을 살펴봅시다. 나중에는 프로 프로그래머처럼 키보드에서 손을 떼지 않아도 될지도 모릅니다.
| assets/tutorial/shortcuts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Production Engineering**
# # *Reservoir Inflow Behaviour*
# 
# # Import Python Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %config Completer.use_jedi = False
# # **Important Functions**
# 
# ## **Productivity Index Taking into account Petrophysical and Fluid Properties**
#
# $$J=\frac{K_{o}\,h}{141.2\,B_{o}\,u_{o}(\ln{\frac{r_{e}}{r_{w}}}-0.75+s)}$$
#
# **Where:** \
# *$J$:* Productivity Index (bpd/psi)\
# *$K_{o}$:* Effective permeablity (md)\
# *$h$:* Thickness (ft)\
# *$B_{o}$:* Oil Formation Volume Factor (rb/stb)\
# *$u_{o}$:* Oil Viscosity (cp)\
# *$r_{e}$:* Drainage ratio (ft)\
# *$r_{w}$:* Well ratio (ft)\
# *$s$:* Skin
# Productivity Index (darcy law)
def J_darcy(ko, h, bo, uo, re, rw, s, flow_regime = 'seudocontinuo'):
if flow_regime == 'seudocontinuo':
J = ko * h / (141.2 * bo * uo * (np.log(re / rw) - 0.75 + s))
elif flow_regime == 'continuo':
J = ko * h / (141.2 * bo * uo * (np.log(re / rw) + s))
return J
# ## **Productivity Index with productivity test data**
#
# $$J=\frac{Q_{o}}{P_{r} - Pwf}$$
#
# **Where:** \
# *$J$:* Productivity Index (bpd/psi)\
# *$Q_{o}$:* Oil Flow Rate (bpd)\
# *$P_{r}$:* Reservoir Pressure (psia)\
# *$Pwf$:* Pressure Well Flowing (psia)
# Productivity Index
def J(q_test, pwf_test, pr, pb):
if pwf_test > pb:
J = q_test / (pr - pwf_test)
else:
J = q_test / ((pr - pb) + (pb / 1.8) * (1 - 0.2 * (pwf_test / pb) - 0.8 * (pwf_test / pb)**2))
return J
# ## **Oil Flow Rate at Bubble Point**
#
# $$Q_{b}=J(P_{r} - P_{b})$$
#
# **Where:** \
# *$Q_{b}$:* Oil Flow Rate at Bubble Point (bpd) \
# *$J$:* Productivity Index (bpd/psi) \
# *$P_{r}$:* Reservoir Pressure (psia) \
# *$P_{b}$:* Bubble Point Pressue
# Q(bpd) @ Pb
def Qb(q_test, pwf_test, pr, pb):
qb = J(q_test, pwf_test, pr, pb) * (pr - pb)
return qb
# ## **AOF at different conditions**
#
# **If $P_{r}$ > $P_{b}$** -> The oil reservoir is **UNDERSATURATED:**
#
# At this case, there are 2 conditions:
#
# *- If $Pwf$ >= $P_{b}$*:
# $$AOF=JP_{r}$$
#
# *Otherwise*,
# $$AOF=Q_{b} + \frac{JP_{b}}{1.8}$$
#
# On the other hand, if $P_{r}$ <= $P_{b}$ -> The oil reservoir is **SATURATED:**
#
# At this situation:
#
# $$AOF=\frac{Qo_{test}}{1 - 0.2 (\frac{Pwf_{test}}{P_{r}}) - 0.8 (\frac{Pwf_{test}}{P_{r}})^2}$$
#
# **Where:** \
# *$AOF$:* Absolute Open Flow (bpd)\
# *$Q_{b}$:* Oil Flow Rate at Bubble Point (bpd)\
# *$J$:* Productivity Index (bpd/psi)\
# *$P_{r}$:* Reservoir Pressure (psia)\
# *$P_{b}$:* Bubble Point Pressue\
# *$Pwf$:* Pressure Well Flowing (psia)\
# *$Pwf_{test}$:* Pressure Well Flowing of productity test (psia)\
# *$Qo_{test}$:* Oil Flow Rate of productivity test (bpd)
# AOF(bpd)
def aof(q_test, pwf_test, pr, pb, pwf):
if pr > pb: # Yac. subsaturado
if pwf >= pb:
aof = J(q_test, pwf_test, pr, pb) * pr
elif pwf < pb:
aof = Qb(q_test, pwf_test, pr, pb) + ((J(q_test, pwf_test, pr, pb) * pb) / (1.8))
else: # Yac. Saturado
aof = q_test / (1 - 0.2 * (pwf_test / pr) - 0.8 * (pwf_test / pr)**2)
return aof
# ## **$Q_{o}$ at Different Conditions**
#
# Here, it is assumed that FE (Flow Efficiency) = 100% by using Vogel's statements.
#
# **If $P_{r}$ > $P_{b}$** -> The oil reservoir is **UNDERSATURATED:**
#
# At this case, there are 2 conditions:
#
# *- If $Pwf$ >= $P_{b}$*:
# $$Q_{o}=J(P_{r} - Pwf)$$
#
# *Otherwise*,
# $$Q_{o}=Q_{b} + \frac{JP_{b}}{1.8}(1-0.2 (\frac{Pwf}{P_{b}})-0.8 (\frac{Pwf}{P_{b}})^2)$$
#
# On the other hand, if $P_{r}$ <= $P_{b}$ -> The oil reservoir is **SATURATED:**
#
# At this situation:
# $$Q_{o}=AOF(1-0.2 (\frac{Pwf}{P_{b}})-0.8 (\frac{Pwf}{P_{b}})^2)$$
#
# **Where:** \
# *$Q_{o}$:* Oil Flow Rate of productivity test (bpd)\
# *$AOF$:* Absolute Open Flow (bpd)\
# *$Q_{b}$:* Oil Flow Rate at Bubble Point (bpd)\
# *$J$:* Productivity Index (bpd/psi)\
# *$P_{r}$:* Reservoir Pressure (psia)\
# *$P_{b}$:* Bubble Point Pressue\
# *$Pwf$:* Pressure Well Flowing (psia)
#Qo(bpd) @ vogel conditions
def qo_vogel(q_test, pwf_test, pr, pwf, pb):
if pr > pb: # Yac. subsaturado
if pwf >= pb:
qo = J(q_test, pwf_test, pr, pb) * (pr - pwf)
elif pwf < pb:
qo = Qb(q_test, pwf_test, pr, pb) + ((J(q_test, pwf_test, pr, pb) * pb) / (1.8)) * \
(1 - 0.2 * (pwf / pb) - 0.8 * ( pwf / pb)**2)
elif pr <= pb: # Yac. Saturado
qo = aof(q_test, pwf_test, pr, pb, pwf) * (1 - 0.2 * (pwf / pr) - 0.8 * ( pwf / pr)**2)
return qo
# ## **Pwf at Different Conditions**
#
# Here, it is assumed that FE (Flow Efficiency) = 100% by using Vogel's statements.
#
# **If $P_{r}$ > $P_{b}$** -> The oil reservoir is **UNDERSATURATED:**
#
# At this case, there are 2 conditions:
#
# *- If $Q_{o}$ <= $Q_{b}$*:
# $$Pwf=P_{r} - \frac{Q_{o}}{J}$$
#
# *Otherwise*,
# $$Pwf=0.125P_{r}(-1+ \sqrt{81-\frac{80Q_{o}}{Qo_{max}}})$$ ; $$Qo_{max}=Q_{b} + \frac{JP_{b}}{1.8}$$
#
# On the other hand, if $P_{r}$ <= $P_{b}$ -> The oil reservoir is **SATURATED:**
#
# At this situation:
# $$Pwf=0.125P_{r}(-1+ \sqrt{81-\frac{80Q_{o}}{Qo_{max}}})$$ ; $$Qo_{max}=\frac{Qo_{test}}{1 - 0.2 (\frac{Pwf_{test}}{P_{r}}) - 0.8 (\frac{Pwf_{test}}{P_{r}})^2}$$
#
# **Where:** \
# *$Q_{o}$:* Oil Flow Rate of productivity test (bpd)\
# *$AOF = Qo_{max}$:* Absolute Open Flow (bpd)\
# *$Q_{b}$:* Oil Flow Rate at Bubble Point (bpd)\
# *$J$:* Productivity Index (bpd/psi)\
# *$P_{r}$:* Reservoir Pressure (psia)\
# *$P_{b}$:* Bubble Point Pressue\
# *$Pwf$:* Pressure Well Flowing (psia)\
# *$Pwf_{test}$:* Pressure Well Flowing of productity test (psia)\
# *$Qo_{test}$:* Oil Flow Rate of productivity test (bpd)
# Pwf @ vogel conditions
def pwf_vogel(q_test, pwf_test, pr, qo, pb):
if pr > pb:
if qo <= Qb(q_test, pwf_test, pr, pb):
pwf = pr - qo / J(q_test, pwf_test, pr, pb)
elif qo > Qb(q_test, pwf_test, pr, pb):
Qmax = Qb(q_test, pwf_test, pr, pb) + ((J(q_test, pwf_test, pr, pb) * pb) / (1.8))
pwf = 0.125*pr*(-1 + np.sqrt(81 - 80*qo/Qmax))
elif pr <= pb:
Qmax = q_test / (1 - 0.2 * (pwf_test / pr) - 0.8 * (pwf_test / pr)**2)
pwf = 0.125*pr*(-1 + np.sqrt(81 - 80*qo/Qmax))
return pwf
# ## *Ejercicio 1*
# Data
ko = 8.2 #md
h = 53 #ft
bo = 1.2 #rb/stb
uo = 1.2 #cp
re = 2978.4 # ft
rw = 0.328 # ft
s = 0
pr = 5651 #psia
# ### a) J
J_darcy = J_darcy(ko, h, bo, uo, re, rw, s)
print(f"J -> {J_darcy} bpd/psia")
# ### b) AOF
AOF = J_darcy * pr
print(f"AOF -> {AOF} bpd")
# ## *Ejercicio 2*
# Data
pr = 2400 #psia
pb = 2500 #psia
pwf = 1000 #psia
q_test = 100 #stb/d
pwf_test = 1800 #psia
# ### a) AOF
AOF = aof(q_test, pwf_test, pr, pb, pwf)
print(f"AOF -> {AOF} bpd")
# ### b) IPR Curve
# +
# Creating Dataframe
df = pd.DataFrame()
df['Pwf(psia)'] = np.array([2400, 2000, 1500, 1000, 500, 0])
df['Qo(bpd)'] = df['Pwf(psia)'].apply(lambda x: qo_vogel(q_test, pwf_test, pr, x, pb))
# -
df
# +
# Plot
fig, ax = plt.subplots(figsize=(18, 8))
ax.plot(df['Qo(bpd)'], df['Pwf(psia)'], c='red')
ax.set_xlabel('Qo(bpd)')
ax.set_ylabel('Pwf(psia)')
ax.set_title('IPR')
ax.set(xlim=(0, df['Qo(bpd)'].max() + 10), ylim=(0, df['Pwf(psia)'][0] + 100))
ax.grid()
plt.show()
# -
# ## *Ejercicio 3*
# Data
pr = 120 #bar
pb = 65 #bar
q_test = 400 #m3/d
pwf_test = 100 #bar
pwf = 40 #bar
# ### Qo @ Pwf=40 bar
# ## *Ejercicio 4*
# Data
pr = 4000 #psi
pb = 3000 #psi
qo_test = 600 #bpd
pwf_test = 2000 #bpd
# ### a) Qmax
# ### b) Qo @ Pwf = 3500 psi
# ### c) Qo @ Pwf = 1000 psi
# ### d) pwf @ Q = 1110 bpd
# ### e) IPR Curve
| Reservoir Inflow Behaviour.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NN tension prediction for SUPERball
# ### <NAME>
#
# This neural network uses the torque values of the 24 motors of SUPERball as features to predict the tension on one of the cables. The learning process is supervised and data is acquired by logging the motor status for a long period of time. To measure the tension of the cable, another motor was attached to said cable and used as a force sensor.
#
# Dataset used for training: 2017-10-24_BiggerBetterData_1kHz.mat
#
#
# Import libraries
import tensorflow as tf
import numpy as np
import collections
import os
import collections
import matplotlib.pyplot as plt
import scipy.io
# +
D=24 #number of features recorded at each time step
TRAIN_SET_SIZE=200000 # Size of training dataset
TEST_SET_SIZE=100000 # SIze of test dataset
#Importing dataset
dataset=scipy.io.loadmat('2017-10-24_BiggerBetterData_1kHz.mat')
#Selecting subset of the entire dataset that contains the features meaningful for this prediction
dataset_effort=dataset.get('effort')
#Dividing dataset into train and target subsets
train_target=[]
for i in range(0, TRAIN_SET_SIZE):
train_target.append(dataset_effort[i,D])
train_target=np.asarray(train_target)
train_target=np.reshape(train_target,[len(train_target),1])
print(train_target.shape)
test_target=[]
for i in range(TRAIN_SET_SIZE, TRAIN_SET_SIZE+TEST_SET_SIZE):
test_target.append(dataset_effort[i,D])
test_target=np.asarray(test_target)
test_target=np.reshape(test_target,[len(test_target),1])
print(test_target.shape)
train_features=[]
for i in range(0, TRAIN_SET_SIZE):
train_features.append(dataset_effort[i,0:D])
train_features=np.asarray(train_features)
train_features=np.reshape(train_features,[len(train_features),D])
print(train_features.shape)
test_features=[]
for i in range(TRAIN_SET_SIZE, TRAIN_SET_SIZE+TEST_SET_SIZE):
test_features.append(dataset_effort[i,0:D])
test_features=np.asarray(test_features)
test_features=np.reshape(test_features,[len(test_features),D])
print(test_features.shape)
# +
batch_size = 100 #size of the batch
batch_len =len(train_features)//batch_size
H = 5*D # size of hidden state
print('train_data_len=',len(train_features),' batch_size=',batch_size,' batch_len=',
batch_len,' D=',D,'H=',H)
# +
# Create placceholders for the models. In the training and test phases real data will be fed to the NN and will take the
# place of these placeholders
#Input shape: (batch_size,number of features)
#Output shape: (batch_size, number of outputs)
Xin= tf.placeholder(tf.float32,shape=[batch_size,D])
Ytarget = tf.placeholder(tf.float32,shape=[batch_size,1])
#Xavier initialization for weights
#http://andyljones.tumblr.com/post/110998971763/an-explanation-of-xavier-initialization
Wx = tf.get_variable("Wx", shape=[D, H],initializer=tf.contrib.layers.xavier_initializer()); print('Wx=',Wx.get_shape())
Wy = tf.get_variable("Wy", shape=[H, 1],initializer=tf.contrib.layers.xavier_initializer()); print('Wy=',Wy.get_shape())
#Bias initialized to 0
bh = tf.Variable(tf.zeros([1,H])); print('bh=',bh.get_shape())
by = tf.Variable(tf.zeros([1,1])); print('by=',by.get_shape())
# -
# ## NN Implementation
#
# The neural network only has one hidden layer and is using the ReLU function as activation function:
#
# $$
# \begin{aligned}
# h_t &= \textrm{ReLU}(W_x x_{in} + b_h)\\
# y_t &= W_y h_t + b_y
# \end{aligned}
# $$
#
#
# +
# NN implementation with ReLU function and one hidden layer
h_t=tf.nn.relu(tf.matmul(Xin,Wx)+bh)
y_=tf.matmul(h_t,Wy)+ by;
print('Ypredicted=',y_.get_shape())
print('Ytarget=',Ytarget.get_shape())
#Mean Squared Error cost function (worse performance)
#cost = tf.reduce_mean(tf.square(Ytarget-y_))
#Mean Absolute Error cost function
cost=tf.reduce_mean(tf.abs(tf.subtract(y_, Ytarget)))
#Optimizer used to implement backpropagation
optimizer = tf.train.AdamOptimizer(learning_rate=0.0002).minimize(cost)
# -
#Defining an interval for the accuracy
margin=0.1
lower_bound =tf.greater_equal(y_,tf.subtract(Ytarget,margin))
upper_bound= tf.less_equal(y_,tf.add(Ytarget,margin))
correct=tf.equal(lower_bound,upper_bound)
accuracy = tf.reduce_mean(tf.cast(correct,tf.float32))
print('Accuracy interval set')
# +
#Initializing the variables and the session
init_op = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init_op)
costs = []
# TRAINING
# For every epoch, feed the whole training set in batches and let the NN learn the weights and the biases
for epoch in range(300):
total_batch=int(len(train_features)/batch_size)
for i in range(total_batch):
inp_data= train_features[i*batch_size:(i+1)*batch_size,:]
out_data= train_target[i*batch_size:(i+1)*batch_size,:]
_,acc,c = sess.run([optimizer,accuracy,cost],feed_dict={Xin: inp_data, Ytarget: out_data})
print("Epoch: {}, Cost: {}, Accuracy: {}".format(epoch,c,acc))
costs.append(c)
print ("\nTraining complete!")
#TESTING
#Feed the test set in batches and compare the prediction with the actual output from the output test set
predicted=[]
test_batch=int(len(test_features)/batch_size)
for j in range (test_batch):
inp_data = test_features[j*batch_size:(j+1)*batch_size,:]
pred = sess.run(y_, feed_dict={Xin: inp_data})
predicted=np.append(predicted,pred)
# Plot predicted values, real output values and measurement from the motor connected to the cable whose tension
# we are trying to predict
x=np.arange(0,len(test_target))
y1= predicted
y2= test_target
y3=-test_features[:,21]*0.0175/0.008
fig= plt.figure(figsize=(20,10))
ax1=plt.subplot(211)
ax1.plot(x,y1,'b',label='Predictions')
ax1.plot(x,y2,'r',label='Targets')
ax1.plot(x,y3,'g',label='Motor 22')
ax1.legend(loc="upper right")
plt.title('Prediction using 24 motor torques as features')
plt.show()
sess.close()
# -
| NN_tension_pred_relu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression Project
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
#
# * Avg. Session Length: Average session of in-store style advice sessions.
# * Time on App: Average time spent on App in minutes
# * Time on Website: Average time spent on Website in minutes
# * Length of Membership: How many years the customer has been a member.
customers = pd.read_csv("Ecommerce Customers")
customers.head()
customers.info()
customers.describe()
sns.set_palette("GnBu_d")
sns.set_style("whitegrid")
customers.columns
sns.jointplot(customers['Time on Website'],customers['Yearly Amount Spent'])
sns.jointplot(customers['Time on App'],customers['Yearly Amount Spent'])
sns.jointplot(customers['Time on App'],customers['Length of Membership'],kind="hex")
sns.pairplot(customers)
sns.lmplot('Yearly Amount Spent','Length of Membership',customers)
customers.columns
y = customers['Yearly Amount Spent']
X = customers[['Avg. Session Length', 'Time on App','Time on Website', 'Length of Membership']]
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3,random_state = 101)
from sklearn.linear_model import LinearRegression
# **Create an instance of a LinearRegression() model named lm.**
lm = LinearRegression()
lm.fit(X_train,y_train)
# **Print out the coefficients of the model**
print("Coefficients:\n",lm.coef_)
predictions = lm.predict(X_test)
# ** Create a scatterplot of the real test values versus the predicted values. **
plt.scatter(y_test,predictions)
plt.xlabel("Y Test")
plt.ylabel("Predicted Y")
# ## Evaluating the Model
#
# Let's evaluate our model performance by calculating the residual sum of squares and the explained variance score (R^2).
# +
from sklearn import metrics
print("MAE: ",metrics.mean_absolute_error(y_test,predictions))
print("MSE: ",metrics.mean_squared_error(y_test,predictions))
print("RMSE: ",np.sqrt(metrics.mean_squared_error(y_test,predictions)))
# -
sns.distplot(y_test-predictions)
coefficients = pd.DataFrame(lm.coef_,X.columns)
coefficients.columns = ['Coefficients']
coefficients
# Interpreting the coefficients:
#
# - Holding all other features fixed, a 1 unit increase in **Avg. Session Length** is associated with an **increase of 25.98 total dollars spent**.
# - Holding all other features fixed, a 1 unit increase in **Time on App** is associated with an **increase of 38.59 total dollars spent**.
# - Holding all other features fixed, a 1 unit increase in **Time on Website** is associated with an **increase of 0.19 total dollars spent**.
# - Holding all other features fixed, a 1 unit increase in **Length of Membership** is associated with an **increase of 61.27 total dollars spent**.
# **The company should focus more on their mobile app or on their website?**
# This is tricky, there are two ways to think about this: Develop the Website to catch up to the performance of the mobile app, or develop the app more since that is what is working better. This sort of answer really depends on the other factors going on at the company, you would probably want to explore the relationship between Length of Membership and the App or the Website before coming to a conclusion!
#
| LINEAR REGRESSION - COMPANY SALES.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="aklJxkHBD5aR"
# # LAB 2: AutoML Tables Babyweight Training.
#
# **Learning Objectives**
#
# 1. Setup AutoML Tables
# 1. Create and import AutoML Tables dataset from BigQuery
# 1. Analyze AutoML Tables dataset
# 1. Train AutoML Tables model
# 1. Check evaluation metrics
# 1. Deploy model
# 1. Make batch predictions
# 1. Make online predictions
#
#
# ## Introduction
# In this notebook, we will use AutoML Tables to train a model to predict the weight of a baby before it is born. We will use the AutoML Tables UI to create a training dataset from BigQuery and will then train, evaluate, and predict with a Auto ML Tables model.
#
# In this lab, we will setup AutoML Tables, create and import an AutoML Tables dataset from BigQuery, analyze AutoML Tables dataset, train an AutoML Tables model, check evaluation metrics of trained model, deploy trained model, and then finally make both batch and online predictions using the trained model.
#
# Each learning objective will correspond to a series of steps to complete in this student lab notebook.
# -
# ## Verify tables exist
#
# Run the following cells to verify that we previously created the dataset and data tables. If not, go back to lab [1b_prepare_data_babyweight](../solutions/1b_prepare_data_babyweight.ipynb) to create them.
# %%bigquery
-- LIMIT 0 is a free query; this allows us to check that the table exists.
SELECT * FROM babyweight.babyweight_augmented_data
LIMIT 0
# ## Lab Task #1: Setup AutoML Tables
# ### Step 1: Open AutoML Tables
# Go the GCP console and open the console menu in the upper left corner. Then scroll down to the bottom to get to the `Artificial Intelligence` section. Click on `Tables` to open AutoML Tables.
# <img src="../assets/1_automl_tables_hamburger_dropdown.png" width='70%'>
# ### Step 2: Enable API
# If you haven't already enabled the AutoML Tables API, then you'll see the screen below. Make sure to click the `ENABLE API` button.
# <img src="../assets/2_automl_tables_enable_api.png" width='70%'>
# ### Step 3: Get started
# If this is your first time using AutoML Tables, then you'll see the screen below. Make sure to click the `GET STARTED` button.
# <img src="../assets/3_automl_tables_get_started.png" width='70%'>
# ## Lab Task #2: Create and import AutoML Tables dataset from BigQuery
# ### Step 4: Datasets
# You should now be on AutoML Table's Datasets page. This is where all imported datasets are shown. We'll want to add our babyweight dataset, so click the `+ NEW DATASET` button.
# <img src="../assets/4_automl_tables_click_create_dataset.png" width='70%'>
# ### Step 5: Create new dataset
# We need to give our new dataset a unique name. I named mine `babyweight_automl` but feel free to name yours whatever you want. When you are done choosing a unique name, click the `CREATE DATASET` button.
# <img src="../assets/5_automl_tables_create_new_dataset.png" width='70%'>
# ### Step 6: Import your data
# Now that we've created a dataset, let's import our data so that AutoML Tables can use it for training. Our data is currently already in BigQuery, so we will select the radio button `Import data from BigQuery`. This will give us some text boxes to fill in with our data's `BigQuery Project ID`, `BigQuery Dataset ID`, and `BigQuery Table or View ID`. Once you are done entering those in, click the `IMPORT` button.
# <img src="../assets/6_automl_tables_import_data.png" width='70%'>
# ### Step 7: Wait for your data to be imported
# AutoML Tables should now be importing your data from BigQuery. Depending on the size of your dataset, this could take a while, so this step is about just waiting and being patient.
# <img src="../assets/7_automl_tables_importing_data.png" width='70%'>
# ### Step 8: Select target column
# Awesome! Our dataset has been successfully imported! You can now look at the dataset's schema which will show for each column the column name, the data type, and its nullability. Out of these columns we need to select which column is we want to be our target or label column. Click the drop down for `Target column` and choose `weight_pounds`.
# <img src="../assets/8_automl_tables_schema_target_column.png" width='70%'>
# ### Step 9: Approve target and schema
# When you successfully choose your target column you will see a green checkmark and the target tag added to the column row on the right. It will also disable its nullability since machine learning doesn't do too well with null labels.
# <img src="../assets/9_automl_tables_schema_continue.png" width='70%'>
# ## Lab Task #3: Analyze AutoML Tables dataset
# ### Step 10: Analyze
# We can analyze some basic statistics here. We can see that we have 6 features, 4 of which are numeric and 2 of which are categorical. We can also see that there are 0% missing and 0 invalid values across all of our columns, which is great! We can also see the number of distinct values which we can compare with our expectations. Additionally, the linear correlation with the target column, `weight_pounds` in this instance, is shown as well as the mean and standard deviation for each column. Once you are satisfied with the analysis, then click the **TRAIN MODEL**.
# <img src="../assets/10_automl_tables_analyze.png" width='70%'>
# ## Lab Task #4: Train AutoML Tables model
# ### Step 11: Setup training
# We are almost ready to train our model. It took a lot of steps to get here but those were mainly to import the data and make sure the data is alright. As we all know, data is extremely important for ML and if it is not what we expect then our model will also not perfom as we expect. Garbage in, garbage out. We need to set the `Training budget` which is the maxmimum number of node hours to spend training our model. Thankfully, if improvement stops before that, then the training will stop and you'll only be charged for the actual node hours you used. For this dataset, I got decent results with a budget of just 1 to 3 node hours. We also need to select which features we want to use in our model out of the superset of features by selecting the `Input feature selection` dropdown where we will see details in the next step below. Once all of that is set the click the `TRAIN MODEL` button.
# <img src="../assets/11_automl_tables_train.png" width='70%'>
# ### Step 12: Input feature selection
# We imported six columns, one of which, `weight_pounds`, we have set aside to be our target or label column. This leaves five columns leftover. Clicking the `Input feature selection` dropdown provides you with a list of all of the remaining columns. We want `is_male`, `mother_age`, `plurality`, and `gestation_weeks` as our four features. `hashmonth` is leftover from when we did our repeatable splitting in the [2_prepare_babyweight](../solutions/2_prepare_babyweight.ipynb) lab. Whatever is selected will be trained with, so please click the checkbox to de-select it.
# <img src="../assets/12_automl_tables_feature_selection.png" width='70%'>
# ### Step 13: Wait for model to train
# Woohoo! Our model is training! We are going to have an awesome model when it finishes! And now we wait. Depending on the size of your dataset, your training budget, and other factors, this could take a while, anywhere from a couple hours to over a day, so this step is about just waiting and being patient. A good thing to do while you are waiting is to keep going through the next labs in this series and then come back to this once lab training completes.
# <img src="../assets/13_automl_tables_training.png" width='70%'>
# ## Lab Task #5: Check evaluation metrics
# ### Step 14: Evaluate model
# Yay! Our model is done training! Now we can check the `EVALUATE` tab and see how well we did. It reminds you what the target was, `weight_pounds`, what the training was optimized for, `RMSE`, and then many evaluation metrics like MAE, RMSE, etc. My training run did great with an RMSE of 1.030 after only an hour of training! It really shows you the amazing power of AutoML! Below you can see a feature importance bar chart. `gestation_weeks` is by far the most important which makes sense because usually the longer someone has been pregnant, the longer the baby has had time to grow, and therefore the heavier the baby weighs.
# <img src="../assets/14_automl_tables_evaluate.png" width='70%'>
# ## Lab Task #6: Deploy model
# ### Step 15: Deploy model for predictions
# So if you are satisified with how well our brand new AutoML Tables model trained and evaluated, then you'll probably want to do next what ML is all about; making great predictions! To do that, we'll have to deploy our trained model. If you go to the main `Models` page for AutoML Tables you'll see your trained model listed. It gives the model name, the dataset used, the problem type, the time of creation, the model size, and whether the model is deployed or not. Since we just finished training our model, `Deployed` should say `No`. Click the three vertical dots to the right and then click `Deploy model`.
# <img src="../assets/15_automl_tables_deploy.png" width='70%'>
# ### Step 16: Deploy model confirmation
# You should now see a confirmation box pop up on your screen. This is just a confirmation making sure you really want to deploy your model because then there will be charges depending on the model size and the number of machines used. Please click the `DEPLOY` button.
# <img src="../assets/16_automl_tables_deploy_confirmation.png" width='70%'>
# ## Lab Task # 7: Make batch predictions
# ### Step 17: Create batch prediction job
# Great! Once it is done deploying, `Deployed` should say `Yes` and you can now click your model name and then the `TEST & USE` tab. You'll start out with batch prediction. To make these easy, we can for now just predict on the BigQuery table that we used to train and evaluate on. To do that, select the radio button `Data from BigQuery` and then enter your `BigQuery Project Id`, `BigQuery Dataset Id`, and `BigQuery Table or View Id`. We could have also used CSVs from Google Cloud Storage. Then we need to select where we want to put our `Result`. Let's select the radio button `BigQuery project` and then enter our `BigQuery Project Id`. We also could have written the results to Google Cloud Storage. Once all of that is set, please click `SEND BATCH PREDICTION` which will submit a batch prediction job using our trained AutoML Tables model and the data at the location we chose above.
# <img src="../assets/17_automl_tables_batch_predict.png" width='70%'>
# ### Step 18: Batch prediction job finished
# After just a little bit of waiting, your batch predictions should be done. For me with my dataset it took just over 15 minutes. At the bottom of the `BATCH PREDICTION` page you should see a section labeled `Recent Predictions`. It shows the data input, where the results are stored, when it was created, and how long it took to process. Let's now move to the [BigQuery Console UI](https://console.cloud.google.com/bigquery) to have a look.
# <img src="../assets/18_automl_tables_batch_predict_results.png" width='70%'>
# ### Step 19: Batch prediction dataset
# On your list of projects on the far left, you will see the project you have been working in. Click the arrow to expand the dropdown list of all of the BigQuery datasets within the project. You'll see a new dataset there which is the same as what was shown for the `Results directory` from the last step. Expanding that dataset dropdown list you will see two BigQuery tables that have been created: `predictions` and `errors`. Let's first look at the `predictions` table.
# <img src="../assets/19_automl_tables_batch_predict_dataset.png" width='70%'>
# ### Step 20: Batch prediction predictions
# The `predictions` BigQuery table has essentially taken your input data to the batch prediction job and appended three new columns to it. Notice even columns that you did not use as features in your model are still here such as `hashmonth`. You should see the two `prediction_inteval` columns for `start` and `end`. The last column is the prediction `value` which for us is our predicted `weight_pounds` that was calculated by our trained AutoML Tables model uses the corresponding features in the row.
# <img src="../assets/20_automl_tables_batch_predict_prediction_table.png" width='70%'>
# ### Step 21: Batch prediction errors
# We can also look at the `errors` table for any possible errors. When I ran my batch prediction job, thankfully I didn't have any errors, but this is definitely the place to check in case you did. Since my `errors` table was empty, below you'll see the schema. Once again it has essentially taken your input data to the batch prediction job and appended three new columns to it. There is a record stored as well as an error `code` and `error` message. These could be helpful in debugging any unwanted behavior.
# <img src="../assets/21_automl_tables_batch_predict_errors_table_schema.png" width='70%'>
# ## Lab Task #8: Make online predictions
# ### Step 22: Online prediction setup
# We can also perform online prediction with our trained AutoML Tables model. To do that, in the `TEST & USE` tab, click `ONLINE PREDICTION`. You'll see on your screen something similar to below with a table our model's features. Each feature has the column name, the column ID, the data type, the status (whether it is required or not), and a prepopulated value. You can leave those values as is or enter values. For `Categorical` features, make sure to use valid values or else they will just end up in the OOV (out of vocabulary) spill-over and not take full advantage of the training. When you're done setting your values, click the `PREDICT` button.
# <img src="../assets/22_automl_tables_before_online_predict.png" width='70%'>
# ### Step 23: Online prediction result
# After just a moment, you should see your online predictions appear on your screen. There will be a `Prediction result` as well as a `95% prediction interval` returned. You can try other values for each feature and see what predictions they result in!
# <img src="../assets/23_automl_tables_after_online_predict.png" width='70%'>
# ## Lab Summary:
# In this lab, we setup AutoML Tables, created and imported an AutoML Tables dataset from BigQuery, analyzed AutoML Tables dataset, trained an AutoML Tables model, checked evaluation metrics of trained model, deployed trained model, and then finally made both batch and online predictions using the trained model.
# + [markdown] colab_type="text" id="PK_-WNGUD5bX"
# Copyright 2021 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# https://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
| notebooks/end-to-end-structured/labs/2_automl_tables_babyweight.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/iamfaith/DeepLearning/blob/master/hw7_Architecture_Design.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="8odNXcMV_wI8" colab_type="text"
# # Homework 7 - Network Compression (Architecuture Design)
#
# > Author: <NAME> (<EMAIL>)
#
# 若有任何問題,歡迎來信至助教信箱 <EMAIL>
# + [markdown] id="7YIYiHkT4CLp" colab_type="text"
# # Readme
#
# HW7的任務是模型壓縮 - Neural Network Compression。
#
# Compression有很多種門派,在這裡我們會介紹上課出現過的其中四種,分別是:
#
# * 知識蒸餾 Knowledge Distillation
# * 網路剪枝 Network Pruning
# * 用少量參數來做CNN Architecture Design
# * 參數量化 Weight Quantization
#
# 在這個notebook中我們會介紹MobileNet v1的Architecture Design。
# + [markdown] id="BTz5r-Zy4UDf" colab_type="text"
# # Architecture Design
#
# ## Depthwise & Pointwise Convolution
# 
# > 藍色為上下層Channel的關係,綠色則為該Receptive Field的擴張。
# > (圖片引用自arxiv:1810.04231)
#
# (a) 就是一般的Convolution Layer,所以他的Weight連接方式會跟Fully Connected一樣,只差在原本在FC是用數字相乘後相加,Convolution Layer是圖片卷積後相加。
#
# (b) DW(Depthwise Convolution Layer)你可以想像成一張feature map各自過**一個filter**處理後,再用PW(Pointwise Convolution Layer)把所有feature map的單個pixel資訊合在一起(就是1個pixel的Fully Connected Layer)。
#
# (c) GC(Group Convolution Layer)就是把feature map分組,讓他們自己過Convolution Layer後再重新Concat起來。算是一般的Convolution和Depthwise Convolution的折衷版。**所以說,Group Convolution的Group=Input Feautures數就會是Depthwise Convolution(因為每個Channel都各自獨立),Group=1就會是一般的Convolution(因為就等於沒有Group)。**
#
# <img src="https://i.imgur.com/Hqhg0Q9.png" width="500px">
#
#
# ## 實作細節
# ```python
# # 一般的Convolution, weight大小 = in_chs * out_chs * kernel_size^2
# nn.Conv2d(in_chs, out_chs, kernel_size, stride, padding)
#
# # Group Convolution, Group數目可以自行控制,表示要分成幾群。其中in_chs和out_chs必須要可以被groups整除。(不然沒辦法分群。)
# nn.Conv2d(in_chs, out_chs, kernel_size, stride, padding, groups=groups)
#
# # Depthwise Convolution, 輸入chs=輸出chs=Groups數目, weight大小 = in_chs * kernel_size^2
# nn.Conv2d(in_chs, out_chs=in_chs, kernel_size, stride, padding, groups=in_chs)
#
# # Pointwise Convolution, 也就是1 by 1 convolution, weight大小 = in_chs * out_chs
# nn.Conv2d(in_chs, out_chs, 1)
# ```
#
#
# + [markdown] id="SRnRXK3zQzVO" colab_type="text"
# # Model
#
# * training的部分請參考Network Pruning、Knowledge Distillation,或直接只用Hw3的手把手即可。
#
# > 註記: 這邊把各個Block多用一層Sequential包起來是因為Network Pruning的時候抓Layer比較方便。
# + id="nrBEYCCC7JQP" colab_type="code" colab={}
import torch.nn as nn
import torch.nn.functional as F
import torch
class StudentNet(nn.Module):
'''
在這個Net裡面,我們會使用Depthwise & Pointwise Convolution Layer來疊model。
你會發現,將原本的Convolution Layer換成Dw & Pw後,Accuracy通常不會降很多。
另外,取名為StudentNet是因為這個Model等會要做Knowledge Distillation。
'''
def __init__(self, base=16, width_mult=1):
'''
Args:
base: 這個model一開始的ch數量,每過一層都會*2,直到base*16為止。
width_mult: 為了之後的Network Pruning使用,在base*8 chs的Layer上會 * width_mult代表剪枝後的ch數量。
'''
super(StudentNet, self).__init__()
multiplier = [1, 2, 4, 8, 16, 16, 16, 16]
# bandwidth: 每一層Layer所使用的ch數量
bandwidth = [ base * m for m in multiplier]
# 我們只Pruning第三層以後的Layer
for i in range(3, 7):
bandwidth[i] = int(bandwidth[i] * width_mult)
self.cnn = nn.Sequential(
# 第一層我們通常不會拆解Convolution Layer。
nn.Sequential(
nn.Conv2d(3, bandwidth[0], 3, 1, 1),
nn.BatchNorm2d(bandwidth[0]),
nn.ReLU6(),
nn.MaxPool2d(2, 2, 0),
),
# 接下來每一個Sequential Block都一樣,所以我們只講一個Block
nn.Sequential(
# Depthwise Convolution
nn.Conv2d(bandwidth[0], bandwidth[0], 3, 1, 1, groups=bandwidth[0]),
# Batch Normalization
nn.BatchNorm2d(bandwidth[0]),
# ReLU6 是限制Neuron最小只會到0,最大只會到6。 MobileNet系列都是使用ReLU6。
# 使用ReLU6的原因是因為如果數字太大,會不好壓到float16 / or further qunatization,因此才給個限制。
nn.ReLU6(),
# Pointwise Convolution
nn.Conv2d(bandwidth[0], bandwidth[1], 1),
# 過完Pointwise Convolution不需要再做ReLU,經驗上Pointwise + ReLU效果都會變差。
nn.MaxPool2d(2, 2, 0),
# 每過完一個Block就Down Sampling
),
nn.Sequential(
nn.Conv2d(bandwidth[1], bandwidth[1], 3, 1, 1, groups=bandwidth[1]),
nn.BatchNorm2d(bandwidth[1]),
nn.ReLU6(),
nn.Conv2d(bandwidth[1], bandwidth[2], 1),
nn.MaxPool2d(2, 2, 0),
),
nn.Sequential(
nn.Conv2d(bandwidth[2], bandwidth[2], 3, 1, 1, groups=bandwidth[2]),
nn.BatchNorm2d(bandwidth[2]),
nn.ReLU6(),
nn.Conv2d(bandwidth[2], bandwidth[3], 1),
nn.MaxPool2d(2, 2, 0),
),
# 到這邊為止因為圖片已經被Down Sample很多次了,所以就不做MaxPool
nn.Sequential(
nn.Conv2d(bandwidth[3], bandwidth[3], 3, 1, 1, groups=bandwidth[3]),
nn.BatchNorm2d(bandwidth[3]),
nn.ReLU6(),
nn.Conv2d(bandwidth[3], bandwidth[4], 1),
),
nn.Sequential(
nn.Conv2d(bandwidth[4], bandwidth[4], 3, 1, 1, groups=bandwidth[4]),
nn.BatchNorm2d(bandwidth[4]),
nn.ReLU6(),
nn.Conv2d(bandwidth[4], bandwidth[5], 1),
),
nn.Sequential(
nn.Conv2d(bandwidth[5], bandwidth[5], 3, 1, 1, groups=bandwidth[5]),
nn.BatchNorm2d(bandwidth[5]),
nn.ReLU6(),
nn.Conv2d(bandwidth[5], bandwidth[6], 1),
),
nn.Sequential(
nn.Conv2d(bandwidth[6], bandwidth[6], 3, 1, 1, groups=bandwidth[6]),
nn.BatchNorm2d(bandwidth[6]),
nn.ReLU6(),
nn.Conv2d(bandwidth[6], bandwidth[7], 1),
),
# 這邊我們採用Global Average Pooling。
# 如果輸入圖片大小不一樣的話,就會因為Global Average Pooling壓成一樣的形狀,這樣子接下來做FC就不會對不起來。
nn.AdaptiveAvgPool2d((1, 1)),
)
self.fc = nn.Sequential(
# 這邊我們直接Project到11維輸出答案。
nn.Linear(bandwidth[7], 11),
)
def forward(self, x):
out = self.cnn(x)
out = out.view(out.size()[0], -1)
return self.fc(out)
# + [markdown] id="zPTYk9w-B_yt" colab_type="text"
# # Q&A
#
# 有任何問題Network Compression的問題可以寄信到<EMAIL>。
#
# 我有空的話會更新在這裡。
| hw7_Architecture_Design.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/anitayadav3/EmotionRecognitionInConversation/blob/master/BERT_on_IEMOCAP_for_ERC.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="irbKXIVqMGym" outputId="b748361b-96d8-4293-d8a9-6ced6940008d" colab={"base_uri": "https://localhost:8080/"}
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import numpy as np
import re
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
nltk.download('punkt')
from nltk.tokenize import word_tokenize
import pandas as pd
import tensorflow as tf
from tensorflow.keras.layers import Dense, Input
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import ModelCheckpoint
import tensorflow_hub as hub
import pickle
from sklearn.metrics import f1_score
from sklearn.metrics import accuracy_score
from keras.utils.np_utils import to_categorical
# + id="lNTNsf6SMN0W" outputId="50f9b287-2023-4be3-b54b-886802776c33" colab={"base_uri": "https://localhost:8080/"}
# !wget --quiet https://raw.githubusercontent.com/tensorflow/models/master/official/nlp/bert/tokenization.py
# !pip install sentencepiece
# + id="r72yzfDkMSVU"
import tokenization
# + id="gAuiS6t4MW_4"
def bert_encode(texts, tokenizer, max_len=512):
all_tokens = []
all_masks = []
all_segments = []
for text in texts:
text = tokenizer.tokenize(text)
text = text[:max_len-2]
input_sequence = ["[CLS]"] + text + ["[SEP]"]
pad_len = max_len - len(input_sequence)
tokens = tokenizer.convert_tokens_to_ids(input_sequence)
tokens += [0] * pad_len
pad_masks = [1] * len(input_sequence) + [0] * pad_len
segment_ids = [0] * max_len
all_tokens.append(tokens)
all_masks.append(pad_masks)
all_segments.append(segment_ids)
return np.array(all_tokens), np.array(all_masks), np.array(all_segments)
# + id="2R5uSnu5MZQ3"
def build_model(bert_layer, max_len=512):
input_word_ids = Input(shape=(max_len,), dtype=tf.int32, name="input_word_ids")
input_mask = Input(shape=(max_len,), dtype=tf.int32, name="input_mask")
segment_ids = Input(shape=(max_len,), dtype=tf.int32, name="segment_ids")
_, sequence_output = bert_layer([input_word_ids, input_mask, segment_ids])
clf_output = sequence_output[:, 0, :]
out = Dense(6, activation='softmax')(clf_output)
model = Model(inputs=[input_word_ids, input_mask, segment_ids], outputs=out)
model.compile(Adam(lr=2e-6), loss='binary_crossentropy', metrics=['accuracy'])
return model
# + id="SPBiw3jdQqXo" outputId="b0818ffb-a972-4fbe-b3ad-f9ea83bef7cc" colab={"base_uri": "https://localhost:8080/"}
# %%time
module_url = "https://tfhub.dev/tensorflow/bert_en_uncased_L-24_H-1024_A-16/1"
bert_layer = hub.KerasLayer(module_url, trainable=True)
# + id="T0lDyjgKQrbW" outputId="de8fc4d2-a888-4a6d-9f94-17680a5d2490" colab={"base_uri": "https://localhost:8080/"}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="v-g0HXbfQuZa"
with open('/content/gdrive/My Drive/iemocap/train/sentences.pkl', 'rb') as f:
data = pickle.load(f)
with open('/content/gdrive/My Drive/iemocap/train/labels.pkl', 'rb') as f:
labels = pickle.load(f)
with open('/content/gdrive/My Drive/iemocap/test/sentences.pkl', 'rb') as f:
test_data = pickle.load(f)
with open('/content/gdrive/My Drive/iemocap/test/labels.pkl', 'rb') as f:
test_labels = pickle.load(f)
# + id="VTVssce7QwPd"
def preprocessing(data,labels):
processed_data=[]
processed_label=[]
for i in range(0,len(data)):
for j in range(0,len(data[i])):
intermediate_data=[]
intermediate_label=[]
for k in range(0,len(data[i][j])):
text=data[i][j][k]
if text != '<eos>'and text!='<pad>':
intermediate_data.append(text)
processed_data.append(intermediate_data)
for i in labels:
for j in i:
processed_label.append(j)
return processed_data,processed_label
# + id="fKUpRX1cQw1y"
processed_data,processed_label = preprocessing(data,labels)
test_processed_data,test_processed_label = preprocessing(test_data,test_labels)
# + id="2Eib7uuMRLsc"
for i in range(0,len(processed_data)):
processed_data[i]= ' '.join(processed_data[i])
for i in range(0,len(test_processed_data)):
test_processed_data[i]=' '.join(test_processed_data[i])
# + id="Bf4C0QIsRPiS"
processed_data=np.asarray(processed_data)
test_processed_data=np.asarray(test_processed_data)
Y=to_categorical(processed_label, num_classes=6)
test_Y=to_categorical(test_processed_label, num_classes=6)
# + id="kiTdbVgdRTAi"
vocab_file = bert_layer.resolved_object.vocab_file.asset_path.numpy()
do_lower_case = bert_layer.resolved_object.do_lower_case.numpy()
tokenizer = tokenization.FullTokenizer(vocab_file, do_lower_case)
# + id="K8sAlpdtRW6n"
train_input = bert_encode(processed_data, tokenizer, max_len=160)
test_input = bert_encode(test_processed_data, tokenizer, max_len=160)
train_labels = Y
test_labels = test_Y
# + id="bYSK8AjpRYPx" outputId="835044ba-03a7-4bb2-eeec-5a92414ea20f" colab={"base_uri": "https://localhost:8080/"}
model = build_model(bert_layer, max_len=160)
model.summary()
# + id="99O-dUk_Ra5y" outputId="2944145f-93db-48d1-d56e-4aa820c194fd" colab={"base_uri": "https://localhost:8080/"}
train_history = model.fit(
train_input, train_labels,
epochs=6,
batch_size=1
)
model.save('model.h5')
# + id="i1yXEXJpRcdF"
y_pred=model.predict(test_input)
# + id="ifq6GS_ylKXA" outputId="4de43891-30e4-4a9c-a1ef-1e41ce8d360b" colab={"base_uri": "https://localhost:8080/"}
y_pred=np.argmax(y_pred,axis=1)
test_processed_label=np.asarray(test_processed_label)
print("Accuracy : " + str(accuracy_score(test_processed_label, y_pred1)))
print("Weighted F1-score : " + str(f1_score(test_processed_label, y_pred1, average='weighted')))
| BERT_on_IEMOCAP_for_ERC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visulalization
import matplotlib.pyplot as plt
import mpld3
# # Repesent matplotlib in 3 ways
# 1. random/numeric/str
# 2. image
# 3. live/vedio
x=range(10)
x
# +
x1=[2,3,4,10]
y1=[5,6,7,8]
x2=[10,24,12,13]
y2=[12,23,16,14]
# -
# # Graphs
# 1. dots
# 2. bar
# 3. stackplot
# 4. pichart
# 5. expo
# 6. live
# 7. guage
#
# +
plt.xlabel('hum',c='red')
plt.ylabel('tum',c='green')
plt.grid(c='blue') #line aa jayengi under we can also give color to it as (c='green')
plt.plot(x1,y1,label="cars",c='magenta')
plt.plot(x2,y2,label="bikes")
plt.legend() # to show the label on the screen if we not type this then label "cars" and "bikes" will not been shown
# -
#
# # Bar Plot
plt.xlabel('chalo')
plt.ylabel('chalta hai')
plt.bar(x1,y1,label="apple",)
plt.bar(x2,y2,label="MICOSOFT")
plt.plot(x2,y2,label="amazon",c='green')
plt.grid(c='red')
plt.legend()
# # Cricket Score
players=['virat','dhoni','sehwag','tendulkar']
runs=[15,10,9,18]
plt.bar(players,runs)
plt.xlabel("players")
plt.ylabel("run")
plt.grid(c='b')
# # Scatter/Dots PLOT
plt.scatter(x1,y1,marker="x",s=100)
plt.scatter(x2,y2,marker="^",s=100)
plt.plot(x1,y1)
plt.grid(c='y')
import numpy as np
x=np.array([3,5,6,8,1,9])
x
y=x**2
z=y**2
plt.scatter(x,y)
plt.scatter(x,z)
| Visulalization-matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Call expressions* invoke [functions](functions), which are named operations.
# The name of the function appears first, followed by expressions in
# parentheses.
#
# For example, `abs` is a function that returns the absolute value of the input
# argument:
abs(-12)
# `round` is a function that returns the input argument rounded to the nearest integer (counting number).
round(5 - 1.3)
max(2, 5, 4)
# In this last example, the `max` function is *called* on three *arguments*: 2,
# 5, and 4. The value of each expression within parentheses is passed to the
# function, and the function *returns* the final value of the full call
# expression. You separate the expressions with commas: `,`. The `max` function
# can take any number of arguments and returns the maximum.
# Many functions, like `max` can accept a variable number of arguments.
#
# `round` is an example. If you call `round` with one argument, it returns the number rounded to the nearest integer, as you have already seen:
round(3.3333)
# You can also call round with two arguments, where the first argument is the number you want to round, and the second argument is the number of decimal places you want to round to. If you don't pass this second argument, `round` assumes you mean 0, corresponding to no decimal places, and rounding to the nearest integer:
# The same as above, rounding to 0 decimal places.
round(3.3333, 0)
# You can also round to - say - 2 decimal places, like this:
# Rounding to 2 decimal places.
round(3.3333, 2)
# A few functions are available by default, such as `abs` and `round`, but most
# functions that are built into the Python language are stored in a collection
# of functions called a *module*. An *import statement* is used to provide
# access to a module, such as `math`.
import math
math.sqrt(5)
# Operators and call expressions can be used together in an expression. The
# *percent difference* between two values is used to compare values for which
# neither one is obviously `initial` or `changed`. For example, in 2014 Florida
# farms produced 2.72 billion eggs while Iowa farms produced 16.25 billion eggs
# [^eggs]. The percent difference is 100 times the absolute value of the
# difference between the values, divided by their average. In this case, the
# difference is larger than the average, and so the percent difference is
# greater than 100.
#
# [^eggs]: <http://quickstats.nass.usda.gov>
florida = 2.72
iowa = 16.25
100*abs(florida-iowa)/((florida+iowa)/2)
# Learning how different functions behave is an important part of learning a
# programming language. A Jupyter notebook can assist in remembering the names
# and effects of different functions. When editing a code cell, press the *tab*
# key after typing the beginning of a name to bring up a list of ways to
# complete that name. For example, press *tab* after `math.` to see all of the
# functions available in the `math` module. Typing will narrow down the list of
# options. To learn more about a function, place a `?` after its name. For
# example, typing `math.sin?` will bring up a description of the `sin`
# function in the `math` module. Try it now. You should get something like
# this:
#
# ```
# sqrt(x)
#
# Return the square root of x.
# ```
#
# The list of [Python's built-in
# functions](https://docs.python.org/3/library/functions.html) is quite long and
# includes many functions that are never needed in data science applications.
# The list of [mathematical functions in the `math`
# module](https://docs.python.org/3/library/math.html) is similarly long. This
# text will introduce the most important functions in context, rather than
# expecting the reader to memorize or understand these lists.
# ### Example ###
#
# In 1869, a French civil engineer named <NAME> created what is
# still considered one of the greatest graphs of all time. It shows the
# decimation of Napoleon's army during its retreat from Moscow. In 1812,
# Napoleon had set out to conquer Russia, with over 350,000 men in his army.
# They did reach Moscow but were plagued by losses along the way. The Russian
# army kept retreating farther and farther into Russia, deliberately burning
# fields and destroying villages as it retreated. This left the French army
# without food or shelter as the brutal Russian winter began to set in. The
# French army turned back without a decisive victory in Moscow. The weather got
# colder and more men died. Fewer than 10,000 returned.
#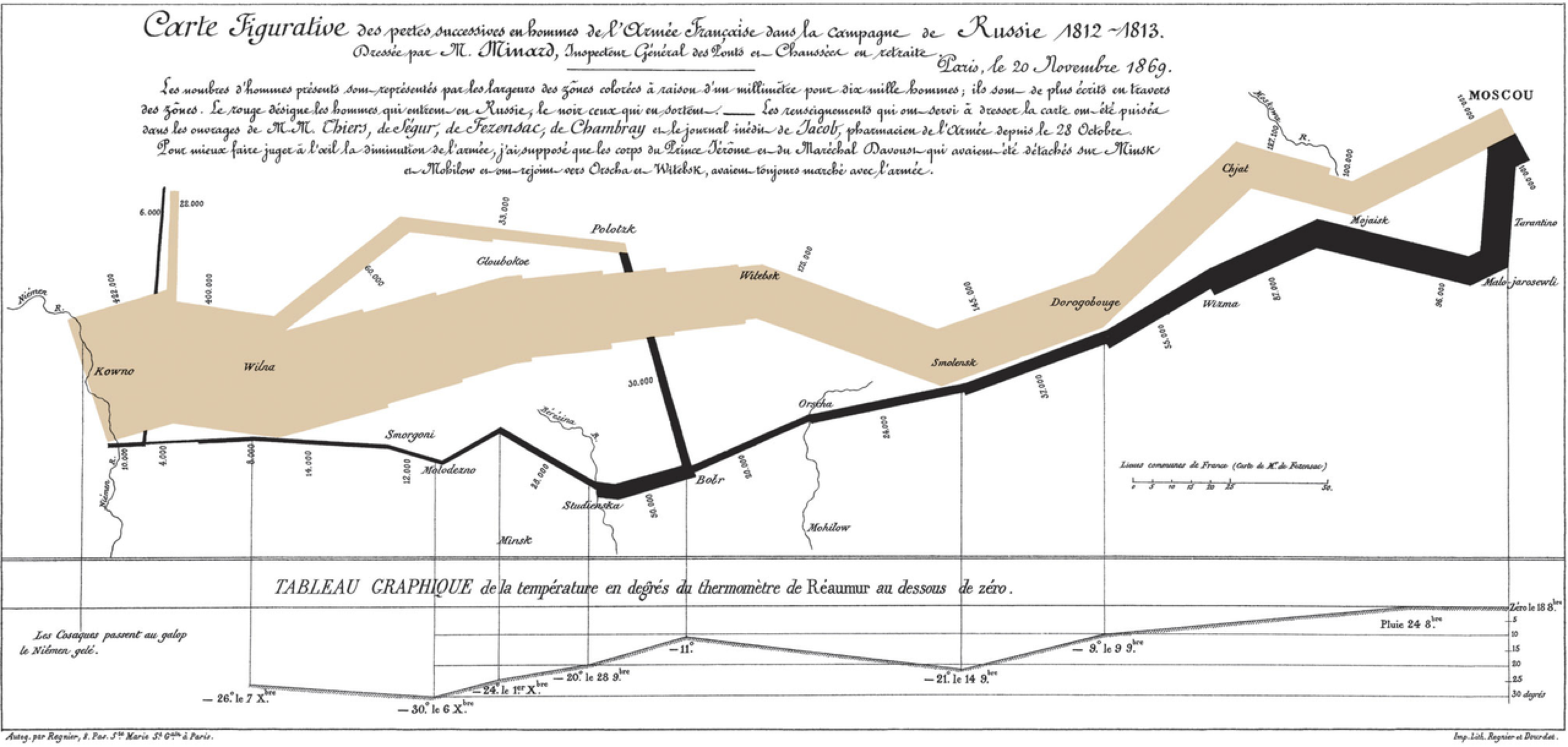 
# The graph is drawn over a map of eastern Europe. It starts at the
# Polish-Russian border at the left end. The light brown band represents
# Napoleon's army marching towards Moscow, and the black band represents the
# army returning. At each point of the graph, the width of the band is
# proportional to the number of soldiers in the army. At the bottom of the
# graph, Minard includes the temperatures on the return journey.
#
# Notice how narrow the black band becomes as the army heads back. The crossing
# of the Berezina river was particularly devastating; can you spot it on the
# graph?
#
# The graph is remarkable for its simplicity and power. In a single graph,
# Minard shows six variables:
#
# - the number of soldiers
# - the direction of the march
# - the latitude and longitude of location
# - the temperature on the return journey
# - the location on specific dates in November and December
#
# Tufte says that Minard's graph is "probably the best statistical graphic ever
# drawn."
# Here is a subset of Minard's data, adapted from *The Grammar of Graphics* by
# <NAME>.
#
# 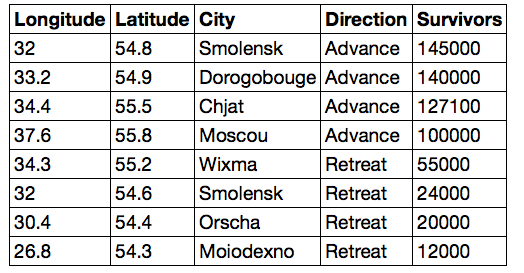
# Each row of the column represents the state of the army in a particular
# location. The columns show the longitude and latitude in degrees, the name of
# the location, whether the army was advancing or in retreat, and an estimate of
# the number of men.
#
# In this table the biggest change in the number of men between two consecutive
# locations is when the retreat begins at Moscow, as is the biggest percentage
# change.
moscou = 100000
wixma = 55000
wixma - moscou
(wixma - moscou)/moscou
# That's a 45% drop in the number of men in the fighting at Moscow. In other
# words, almost half of Napoleon's men who made it into Moscow didn't get very
# much farther.
#
# As you can see in the graph, Moiodexno is pretty close to Kowno where the army
# started out. Fewer than 10% of the men who marched into Smolensk during the
# advance made it as far as Moiodexno on the way back.
smolensk_A = 145000
moiodexno = 12000
(moiodexno - smolensk_A)/smolensk_A
# Yes, you could do these calculations by just using the numbers without names.
# But the names make it much easier to read the code and interpret the results.
# It is worth noting that bigger absolute changes don't always correspond to
# bigger percentage changes.
#
# The absolute loss from Smolensk to Dorogobouge during the advance was 5,000
# men, whereas the corresponding loss from Smolensk to Orscha during the retreat
# was smaller, at 4,000 men.
#
# However, the percent change was much larger between Smolensk and Orscha
# because the total number of men in Smolensk was much smaller during the
# retreat.
dorogobouge = 140000
smolensk_R = 24000
orscha = 20000
abs(dorogobouge - smolensk_A)
abs(dorogobouge - smolensk_A)/smolensk_A
abs(orscha - smolensk_R)
abs(orscha - smolensk_R)/smolensk_R
#
| ipynb/02/Calls.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from random import random
import matplotlib.pyplot as plt
from li_rn_network import li_rnn_networks, task_3back
input_data = np.array([[[0.8,0.2],[0.8,0.2]],[[0.2,0.8],[0.2,0.8]],[[0.8,0.2],[0.2,0.8]],[[0.2,0.8],[0.8,0.2]]]) # 80%-20%
#input_data = np.array([[[0.6,0.4],[0.6,0.4]],[[0.4,0.6],[0.4,0.6]],[[0.6,0.4],[0.4,0.6]],[[0.4,0.6],[0.6,0.4]]]) # 60%-40%
target_data = np.array([[1,0],[1,0],[0,1],[0,1]])
category_num = 2 #カテゴリー数
steps_num = 2 #チャンク数
# +
#ネットワークモデルの定義 difine network model
I = category_num # 入力層のニューロン数 number of input layer's neuron
H = 200 #リカレント結合をもつ中間層のニューロン数 number of hidden layer's neuron(reccurent)
O = 2 #出力層のニューロン数
Wih_size = 2 #入力層から中間層への結合の初期値の値 initial size of bond strength I to H
Whh_size = 2 #層内結合の結合の初期値の値 initial size of bond strength H to H
Who_size = 3 #中間層から出力層への結合の初期値の値 initial size of bond strength I to H
lr = 0.01 #学習率 learning rate
optimizer_model = 0 #オプティマイザーの選択 chose optimizer_model
# -
#ニューロンの生理学的パラメータの定義
Tau_H = 150 #ニューロンの時定数(10~200)
Tau_O = 10 #ニューロンの時定数(10~100)
dt = 0.005 #オイラー法の最小時間(0.1~0.005)
step_length = 200 #同じ入力が続く時間ms time length of one step
size_nois = 1
#ネットワークモデルの生成
li_rnn = li_rnn_networks(I,H,O,Wih_size,Whh_size,Who_size,steps_num)
#学習に関するパラメータの設定 set parameter of leaning
li_rnn.setlr(lr,optimizer_model)
epoch = 200000 #学習回数の定義 difine leaning epoch
#学習の実行 taraning
loss_memo = li_rnn.traning(input_data,target_data,epoch)#出力は誤差のリスト output is list of loss
#学種回数ごとの誤差の出力 visualize loss on taraning epochs
plt.plot(loss_memo)
ix = input_data[2]
li_rnn.def_parameter(Tau_H, Tau_O, dt, size_nois, step_length)
memo_out1, memo_out2 = li_rnn.li_forward(ix, target_data[1])
for i in range(20):
plt.plot(memo_out1.T[i])
#リカレント層(LI)の出力に対する判断層の再学習 relearning woth new output of Reccurebnt layer(LI)
li_rnn.def_parameter(Tau_H, Tau_O, dt, size_nois, step_length)
loss_memo = li_rnn.to_li_para(50000)
plt.plot(loss_memo)
def visu_dic(i, li_rnn, color):
input = input_data[i]
memo_out1, memo_out2 = li_rnn.li_forward(input, target_data[i])
print(target_data[i])
plt.plot(memo_out2.T[0],linewidth=3,color=color, label='much')
plt.plot(memo_out2.T[1],linewidth=3, linestyle="dotted", color=color, alpha=0.7, label='non-much')
num=0
color="#000000"
visu_dic(num, li_rnn, color)
num=1
color="#000000"
visu_dic(num, li_rnn, color)
num=2
color="#000000"
visu_dic(num, li_rnn, color)
num=3
color="#000000"
visu_dic(num, li_rnn, color)
| firerate_vs_time/firerate_vs_time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
df = pd.read_csv("../output_data/benchmark_datasize_sensitivity_lymph.csv")
df.head(10)
labels = {
"DLS": "IDS - DLS",
"SLS": "IDS - SLS",
"DUSM": "IDS - DUSM",
"RUSM": "IDS - RUSM",
"pyARC - M1": "pyARC - M1",
"pyARC - M2": "pyARC - M2"
}
# +
algorithm_dict = {
"DLS": "blue",
"SLS": "green",
"DUSM": "orange",
"RUSM": "red",
"pyARC - M1": "black",
"pyARC - M2": "cyan"
}
for algorithm, color in algorithm_dict.items():
df_alg = df[df["algorithm"] == algorithm]
label = labels[algorithm]
plt.plot(df_alg["data_count"].copy(), df_alg["duration"].copy(), label=label, color=color)
plt.legend()
plt.xlabel("Počet vstupních řádků dat")
plt.ylabel("Čas optimalizace [s]")
# +
algorithm_dict = {
"DLS": "blue",
"DUSM": "orange",
"RUSM": "red",
"pyARC - M1": "black",
"pyARC - M2": "cyan"
}
for algorithm, color in algorithm_dict.items():
df_alg = df[df["algorithm"] == algorithm]
label = labels[algorithm]
plt.plot(df_alg["data_count"], df_alg["duration"], label=label, color=color)
plt.legend()
plt.xlabel("Počet vstupních řádků dat")
plt.ylabel("Čas optimalizace [s]")
# +
algorithm_dict = {
"pyARC - M1": "black",
"pyARC - M2": "cyan"
}
for algorithm, color in algorithm_dict.items():
df_alg = df[df["algorithm"] == algorithm]
label = labels[algorithm]
plt.plot(df_alg["data_count"], df_alg["duration"], label=label, color=color)
plt.legend()
plt.xlabel("Počet vstupních řádků dat")
plt.ylabel("Čas optimalizace [s]")
# -
| jupyter_notebooks/plot_datasize_sensitivity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''ml4nlp'': conda)'
# name: python3
# ---
# # Introduction
#
# The goal of this assignment is to create a basic program that provides an overview of basic evaluation metrics (in particular, precision, recall, f-score and a confusion matrix) from documents provided in the conll format.
# You will need to implement the calculations for precision, recall and f-score yourself (i.e. do not use an existing module that spits them out). Make sure that your code can handle the situation where there are no true positives for a specific class.
#
# This notebook provides functions for reading in conll structures with pandas and proposes a structure for calculating your evaluation metrics and producing the confusion matrix. Feel free to adjust the proposed structure if you see fit.
import sys
import re
import pandas as pd
# see tips & tricks on using defaultdict (remove when you do not use it)
from collections import defaultdict, Counter
# module for verifying output
from nose.tools import assert_equal
# # A note Pandas
#
# Pandas is a module that provides data structures and is widely used for dealing with data representations in machine learning. It is a bit more advanced than the csv module we saw in the preprocessing notebook.
# Working with pandas data structures can be tricky, but it will generally work well if you follow online tutorials and examples closely. If your code is slow before you even started training your models, it is likely to be a problem with the way you are using Pandas (it will still work in most cases, you will just have to wait a bit longer). Once you are more used to working with modules and complex objects, it will also become easier to work with Pandas.
#
# In the examples below, we assume that the data representations that are used have headers (i.e. specific titles that indicate what information can be found in each column of the conll file). You can look at the mini- sample files in data to get an idea of how this works.
def remove_BIO(tag):
if tag == 'NN|SYM':
return 'NN'
return re.sub(r"[A-Z]-", "", tag)
def matching_tokens(conll1, conll2):
'''
Check whether the tokens of two conll files are aligned
:param conll1: tokens (or full annotations) from the first conll file
:param conll2: tokens (or full annotations) from the second conll file
:returns boolean indicating whether tokens match or not
'''
for row in conll1:
row2 = next(conll2)
if row[0] != row2[0]:
return False
return True
def extract_annotations(inputfile, annotationcolumn, delimiter='\t'):
'''
This function extracts annotations represented in the conll format from a file
:param inputfile: the path to the conll file
:param annotationcolumn: the name of the column in which the target annotation is provided
:param delimiter: optional parameter to overwrite the default delimiter (tab)
:type inputfile: string
:type annotationcolumn: string
:type delimiter: string
:returns: the annotations as a list
'''
#https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html
if type(annotationcolumn) == str:
conll_input = pd.read_csv(inputfile, sep=delimiter, quotechar=delimiter)
annotations = conll_input[annotationcolumn].tolist()
annotations = [remove_BIO(tag) for tag in annotations]
if type(annotationcolumn) == int:
conll_input = pd.read_csv(inputfile, sep=delimiter, header=None, quotechar=delimiter)
annotations = conll_input.iloc[:, annotationcolumn ].tolist()
annotations = [remove_BIO(tag) for tag in annotations]
return annotations
# +
def obtain_counts(goldannotations, machineannotations):
'''
This function compares the gold annotations to machine output
:param goldannotations: the gold annotations
:param machineannotations: the output annotations of the system in question
:type goldannotations: the type of the object created in extract_annotations
:type machineannotations: the type of the object created in extract_annotations
:returns: a countainer providing the counts for each predicted and gold class pair
'''
# TIP on how to get the counts for each class
# https://stackoverflow.com/questions/49393683/how-to-count-items-in-a-nested-dictionary, last accessed 22.10.2020
evaluation_counts = defaultdict(Counter)
if len(goldannotations) != len(machineannotations):
print("Error")
return None
labels = set(list(goldannotations))
for l1 in labels:
for l2 in labels:
evaluation_counts[l1][l2] = 0
for true, pred in zip(goldannotations, machineannotations):
evaluation_counts[true][pred] += 1
return evaluation_counts
def calculate_precision_recall_fscore(evaluation_counts):
'''
Calculate precision recall and fscore for each class and return them in a dictionary
:param evaluation_counts: a container from which you can obtain the true positives, false positives and false negatives for each class
:type evaluation_counts: type of object returned by obtain_counts
:returns the precision, recall and f-score of each class in a container
'''
# TIP: you may want to write a separate function that provides an overview of true positives, false positives and false negatives
# for each class based on the outcome of obtain counts
scores = defaultdict(dict)
total_predicted = 0
total_correct = 0
for c, cntr1 in evaluation_counts.items():
total_true_c = sum(cntr1.values()) # true total of gold class
total_predicted += total_true_c
total_correct += cntr1[c]
total_pred_c = 0 # pred total of gold class
for cl, cntr2 in evaluation_counts.items():
for name, value in cntr2.items():
if name == c:
total_pred_c += value
# calculate precision per class
# prevent division by 0
if total_pred_c == 0:
precision = 0
else:
precision = cntr1[c] / total_pred_c
# calculate recall per class
if total_true_c == 0:
recall = 0
else:
recall = cntr1[c] / total_true_c
# calculate f-score per class
if precision == 0 and recall == 0:
fscore = 0
else:
fscore = 2 * ((precision * recall)/(precision + recall))
scores[c]['precision'] = round(precision, 5)
scores[c]['recall'] = round(recall, 5)
scores[c]['f-score'] = round(fscore, 5)
# calculate accuracy
accuracy = total_correct / total_predicted
scores['accuracy'] = accuracy
# calculate weighed precision
weighed_p = 0
for c, cntr3 in evaluation_counts.items():
total_true_c = sum(cntr3.values()) # true total of gold class
weighed_p += scores[c]['precision'] * (total_true_c/total_predicted)
scores['weighed_p'] = weighed_p
# calculate weighed recall
weighed_r = 0
for c, cntr3 in evaluation_counts.items():
total_true_c = sum(cntr3.values()) # true total of gold class
weighed_r += scores[c]['recall'] * (total_true_c/total_predicted)
scores['weighed_r'] = weighed_r
# calculate weighed f-score
weighed_f = 0
for c, cntr3 in evaluation_counts.items():
total_true_c = sum(cntr3.values()) # true total of gold class
weighed_f += scores[c]['f-score'] * (total_true_c/total_predicted)
scores['weighed_f'] = weighed_f
return scores
def provide_confusion_matrix(evaluation_counts):
'''
Read in the evaluation counts and provide a confusion matrix for each class
:param evaluation_counts: a container from which you can obtain the true positives, false positives and false negatives for each class
:type evaluation_counts: type of object returned by obtain_counts
:prints out a confusion matrix
'''
# TIP: provide_output_tables does something similar, but those tables are assuming one additional nested layer
# your solution can thus be a simpler version of the one provided in provide_output_tables below
# YOUR CODE HERE (and remove statement below)
classes = list(evaluation_counts.keys())
firstrow = list(list(evaluation_counts.items())[0][1].values())
df = pd.DataFrame([firstrow], columns=classes, index=[classes[0]])
for i, cl in enumerate(classes[1:]):
data = list(list(evaluation_counts.items())[i+1][1].values())
newrow = pd.DataFrame([data], columns=classes, index=[cl])
df = df.append(newrow)
print("Confusion Matrix:")
print(df)
print(df.to_latex())
# -
def carry_out_evaluation(gold_annotations, systemfile, systemcolumn, delimiter='\t'):
'''
Carries out the evaluation process (from input file to calculating relevant scores)
:param gold_annotations: list of gold annotations
:param systemfile: path to file with system output
:param systemcolumn: indication of column with relevant information
:param delimiter: specification of formatting of file (default delimiter set to '\t')
returns evaluation information for this specific system
'''
system_annotations = extract_annotations(systemfile, systemcolumn, delimiter)
evaluation_counts = obtain_counts(gold_annotations, system_annotations)
provide_confusion_matrix(evaluation_counts)
evaluation_outcome = calculate_precision_recall_fscore(evaluation_counts)
return evaluation_outcome
def provide_output_tables(evaluations):
'''
Create tables based on the evaluation of various systems
:param evaluations: the outcome of evaluating one or more systems
'''
accuracy = evaluations['system1']['accuracy']
evaluations['system1'].pop('accuracy', None)
weighed_p = evaluations['system1']['weighed_p']
evaluations['system1'].pop('weighed_p', None)
weighed_r = evaluations['system1']['weighed_r']
evaluations['system1'].pop('weighed_r', None)
weighed_f = evaluations['system1']['weighed_f']
evaluations['system1'].pop('weighed_f', None)
# https:stackoverflow.com/questions/13575090/construct-pandas-dataframe-from-items-in-nested-dictionary
evaluations_pddf = pd.DataFrame.from_dict({(i,j): evaluations[i][j]
for i in evaluations.keys()
for j in evaluations[i].keys()},
orient='index')
print(evaluations_pddf)
print(evaluations_pddf.to_latex())
print("Accuracy:", round(accuracy,5))
print("Mean f-score:", evaluations_pddf['f-score'].mean())
print("Weighed precision:", round(weighed_p,5))
print("Weighed recall:", round(weighed_r,5))
print("Weighed f-score:", round(weighed_f,5))
def run_evaluations(goldfile, goldcolumn, systems):
'''
Carry out standard evaluation for one or more system outputs
:param goldfile: path to file with goldstandard
:param goldcolumn: indicator of column in gold file where gold labels can be found
:param systems: required information to find and process system output
:type goldfile: string
:type goldcolumn: integer
:type systems: list (providing file name, information on tab with system output and system name for each element)
:returns the evaluations for all systems
'''
evaluations = {}
#not specifying delimiters here, since it corresponds to the default ('\t')
gold_annotations = extract_annotations(goldfile, goldcolumn)
for system in systems:
sys_evaluation = carry_out_evaluation(gold_annotations, system[0], system[1])
evaluations[system[2]] = sys_evaluation
return evaluations
# # Checking the overall set-up
#
# The functions below illustrate how to run the setup as outlined above using a main function and, later, commandline arguments. This setup will facilitate the transformation to an experimental setup that no longer makes use of notebooks, that you will submit later on. There are also some functions that can be used to test your implementation You can carry out a few small tests yourself with the data provided in the data/ folder.
def identify_evaluation_value(system, class_label, value_name, evaluations):
'''
Return the outcome of a specific value of the evaluation
:param system: the name of the system
:param class_label: the name of the class for which the value should be returned
:param value_name: the name of the score that is returned
:param evaluations: the overview of evaluations
:returns the requested value
'''
return evaluations[system][class_label][value_name]
def create_system_information(system_information):
'''
Takes system information in the form that it is passed on through sys.argv or via a settingsfile
and returns a list of elements specifying all the needed information on each system output file to carry out the evaluation.
:param system_information is the input as from a commandline or an input file
'''
# https://stackoverflow.com/questions/312443/how-do-you-split-a-list-into-evenly-sized-chunks
systems_list = [system_information[i:i + 3] for i in range(0, len(system_information), 3)]
return systems_list
# +
def main(my_args=None):
'''
A main function. This does not make sense for a notebook, but it is here as an example.
sys.argv is a very lightweight way of passing arguments from the commandline to a script.
'''
if my_args is None:
my_args = sys.argv
system_info = create_system_information(my_args[2:])
evaluations = run_evaluations(my_args[0], my_args[1], system_info)
provide_output_tables(evaluations)
check_eval = identify_evaluation_value('system1', 'O', 'f-score', evaluations)
#if it does not work correctly, this assert statement will indicate that
# assert_equal("%.3f" % check_eval,"0.889")
# these can come from the commandline using sys.argv for instance
my_args = ['../../data/minigold.csv','gold','../../data/miniout1.csv','NER','system1']
main(my_args)
# -
#some additional tests
test_args = ['../../data/minigold.csv','gold','../../data/miniout2.csv','NER','system2']
system_info = create_system_information(test_args[2:])
evaluations = run_evaluations(test_args[0], test_args[1], system_info)
test_eval = identify_evaluation_value('system2', 'I-ORG', 'f-score', evaluations)
assert_equal("%.3f" % test_eval,"0.571")
test_eval2 = identify_evaluation_value('system2', 'I-PER', 'precision', evaluations)
assert_equal("%.3f" % test_eval2,"0.500")
test_eval3 = identify_evaluation_value('system2', 'I-ORG', 'recall', evaluations)
assert_equal("%.3f" % test_eval3,"0.667")
# Spacy evaluation
my_args = ['../../data/conll2003.dev_prep.conll', 3,'../../data/spacy_out.dev_prep.conll', 2,'system1']
main(my_args)
# Stanford evaluation
my_args = ['../../data/conll2003.dev_prep.conll', 3,'../../data/stanford_out.dev_prep.conll', 3,'system1']
main(my_args)
| code/assignment1/basic_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Description
# In this notebook, I use an image (manually generated in GIMP) to generate a simulated MSI dataset with overlapping isotopic envelopes.
# I'll create two images, one with lipids and one with synthetic polymers (spectra generated in IsoSpec).
# Each image also has a certain level of noise signal - random peaks, as well as contaminant molecules.
# Simulation steps:
# 1. Generate reference spectra for different regions of the image using manually selected proportions
# 2. For each pixel, distort the reference proportions of the spectra (depending on the corresponding region); Distort each spectrum by sampling from the multinomial process with the number of molecules given by the corresponding distorted proportion; Combine the distorted spectra; Add a handful of noise peaks amounting for 5% of the total signal to simulate contaminants; Apply a Gaussian filter to the spectra to lower the resolution; Apply a Gaussian distortion to each intensity measurement to simulate electronic noise; Centroid the spectra.
#
# The result is stored in imzML files.
# ## Module & data imports
# %matplotlib notebook
import imageio
import numpy as np
import numpy.random as rd
from masserstein import Spectrum, estimate_proportions
from matplotlib import pyplot as plt
from pyimzml.ImzMLWriter import ImzMLWriter
mask = imageio.imread('SimulationMask.png')
plt.figure()
plt.title('The reference image')
plt.imshow(mask)
plt.tight_layout()
plt.show()
np.unique(mask[:,:,1])
region1 = mask[:,:,1] == 128
region2 = mask[:,:,1] == 130
region3 = mask[:,:,1] == 46
region4 = mask[:,:,1] == 53
region_mapping = np.zeros(mask.shape[:2], dtype='int')
for reg_id, reg_value in enumerate([0, 128, 130, 46, 53]):
region_mapping[mask[:,:,1] == reg_value] = reg_id
plt.figure(figsize=(6,3))
plt.subplot(141)
plt.title('Region 1')
plt.imshow(region1)
plt.subplot(142)
plt.title('Region 2')
plt.imshow(region2)
plt.subplot(143)
plt.title('Region 3')
plt.imshow(region3)
plt.subplot(144)
plt.title('Region 4')
plt.imshow(region4)
plt.tight_layout()
plt.show()
plt.figure()
plt.imshow(region_mapping)
plt.show()
# ## Lipid dataset
# +
# PC_38_0 = 'C46H92NO8P'
# PC_38_1 = 'C46H90NO8P'
# PC_38_2 = 'C46H88NO8P'
# PC_38_3 = 'C46H86NO8P'
# PS_38_1 = 'C44H84NO10P'
# PS_38_3 = 'C44H80NO10P'
# PS_38_4 = 'C44H78NO10P'
# PS_38_5 = 'C44H76NO10P'
# PS_38_6 = 'C44H74NO10P'
# formulas = [PC_38_0, PC_38_1, PC_38_2, PC_38_3, PS_38_3, PS_38_4, PS_38_5, PS_38_6]
# formulas = [PC_38_0, PC_38_1, PC_38_2, PC_38_3, PS_38_4]
# -
TG_50_8 = 'C53H86O6' # 857.5
PC_38_0 = 'C46H92NO8P' # 856.6
PC_38_1 = 'C46H90NO8P' # 854.5
PS_38_0 = 'C44H86NO10P' # 858.5
PA_44_0 = 'C47H93O8P' # 855.6
formulas = [PC_38_1, PA_44_0, PC_38_0, TG_50_8, PS_38_0]
names = ['PC(38:1)', 'PA(44:0)', 'PC(38:0)', 'TG(50:8)', 'PS(38:0)']
spectra = [Spectrum(f, adduct='K', threshold=0.001, label=n) for f, n in zip(formulas, names)]
for s in spectra:
s.normalize()
# Monoisotopic peaks:
for s in spectra:
print(s.confs[0][0])
# Save the lipid table:
with open('lipid_table.tsv', 'w') as h:
h.write('\t'.join(['Name', 'Formula', 'Monoisotopic mass']) + '\n')
for s in spectra:
h.write(s.label + '\t' + s.formula + '\t' + str(s.confs[0][0]) + '\n')
# +
# for s in spectra:
# s.coarse_bin(1)
# print(s.confs)
# -
plt.figure()
plt.title('All input spectra')
Spectrum.plot_all(spectra)
plt.tight_layout()
plt.show()
# Set the proportions for the regions.
# We should probably make some lipids disappear in some regions to make them distinguishable.
# Strategy:
# Regions 3,4 enriched in PC(38:1).
# Regions 1,2 enriched in PA(44:0) with monoisotopic peak at 855.6, but intensity of 855.6 depleted due to interference with PC(38:1) in 3,4.
# Regions 2,4 enriched in PC(38:0) with monoisotopic peak at 856.6, but intensity of 856.6 shows multiple regions due to overlap from PA(44:0) in 1,2.
#
# +
# region_intensities = np.array([[0.00, 0.00, 0.00, 0.01, 0.02],
# [2.50, 0.00, 0.00, 0.02, 0.01],
# [2.50, 0.00, 0.50, 0.02, 0.01],
# [0.00, 1.00, 0.50, 0.02, 0.01],
# [0.00, 1.00, 0.00, 0.02, 0.01]])
region_intensities = np.array([[0.00, 0.00, 0.00, 0.1, 0.2], # rows = regions (including background as 0)
[0.00, 2.00, 1.00, 0.2, 0.1], # columns = lipids
[0.00, 2.00, 2.00, 0.2, 0.1],
[4.00, 1.00, 1.00, 0.2, 0.1],
[4.00, 1.00, 2.00, 0.2, 0.1]])
# -
# Save the reference intensity image:
region_intensities[region_mapping, 0].shape
plt.figure(figsize=(3, 2*len(spectra)))
for lipid_id, lipid_name in enumerate(names):
plt.subplot(511 + lipid_id)
spatial_intensity = region_intensities[region_mapping, lipid_id]
plt.imshow(spatial_intensity)
plt.title(lipid_name)
plt.tight_layout()
plt.savefig('lipid_enrichment_plot.pdf')
plt.figure()
for i in range(1, region_intensities.shape[0]):
plt.subplot(220 + i)
plt.title('Region %i reference spectrum' % i)
Spectrum.plot_all([s*p for s,p in zip(spectra, region_intensities[i])])
plt.tight_layout()
plt.show()
# Generate distorted reference spectra:
# +
noise_proportion = 0.05
nb_of_noise_peaks = 10
nb_of_molecules = 1e04
peak_sd = 0.05
mass_axis = np.arange(854, 865, 0.01)
# -
reference_spectra = [Spectrum() for _ in range(region_intensities.shape[0])]
for i in range(region_intensities.shape[0]):
for j in range(region_intensities.shape[1]):
reference_spectra[i] += Spectrum.sample_multinomial(spectra[j],
region_intensities[i,j]*nb_of_molecules,
1, 0.001)
reference_spectra[i].add_chemical_noise(nb_of_noise_peaks, noise_proportion)
reference_spectra[i].fuzzify_peaks(peak_sd, 0.01)
reference_spectra[i] = reference_spectra[i].resample(mass_axis)
#reference_spectra[i].add_gaussian_noise(20)
reference_spectra[i].confs = [c for c in reference_spectra[i].confs if c[1] >= 0]
max_intensity = max(y for s in reference_spectra for x,y in s.confs)
plt.figure()
for i in range(1, region_intensities.shape[0]):
plt.subplot(220 + i)
plt.title('Region %i reference spectrum' % i)
reference_spectra[i].plot(profile=True)
plt.ylim(-0.01*max_intensity, 1.1*max_intensity)
plt.tight_layout()
plt.show()
# Centroid the spectrum for estimation:
region = 3
example_spectrum = Spectrum(confs=reference_spectra[region].centroid(0.2, 0.4)[0])
plt.figure()
plt.subplot(121)
Spectrum.plot_all([s*p for s,p in zip(spectra, region_intensities[region])])
plt.subplot(122)
example_spectrum.plot(profile=False)
plt.show()
# Estimate the proportions based on monoisotopic peaks:
mono_peaks = [s.confs[0][0] for s in spectra]
mono_intensities = np.zeros(len(mono_peaks))
for i, p in enumerate(mono_peaks):
match = np.argmin([np.abs(p - c[0]) for c in example_spectrum.confs])
if np.abs(p-example_spectrum.confs[match][0]) < 0.05:
mono_intensities[i] = example_spectrum.confs[match][1]
mono_intensities /= np.sum(mono_intensities)
mono_intensities
# Test the regression:
example_spectrum.normalize()
# reg = estimate_proportions(example_spectrum, spectra, MTD=0.05) # for centroided
reg = estimate_proportions(example_spectrum, spectra, MTD=0.4) # for profile
print('True', 'Regressed', 'Monoisotopic')
for t, e1, e2 in zip(region_intensities[region]/sum(region_intensities[region]), reg['proportions'], mono_intensities):
print(round(t, 3), round(e1, 3), round(e2, 3))
# Now, go over each pixel and simulate the spectra:
image = np.zeros((mask.shape[0], mask.shape[1], len(mass_axis)))
for i in range(mask.shape[0]):
if not i % 50:
print(i)
for j in range(mask.shape[1]):
region_id = region_mapping[i, j]
proportions = region_intensities[region_id]
pixel_spectrum = Spectrum()
for s,p in zip(spectra, proportions):
s = Spectrum.sample_multinomial(s, p*nb_of_molecules, 1, 0.001)
pixel_spectrum += s*p
pixel_spectrum.add_chemical_noise(nb_of_noise_peaks, noise_proportion)
pixel_spectrum.fuzzify_peaks(peak_sd, 0.01)
pixel_spectrum = pixel_spectrum.resample(mass_axis)
# pixel_spectrum.add_gaussian_noise(100)
pixel_spectrum.set_confs([c for c in pixel_spectrum.confs if c[1] >= 0])
image[i,j,:] = [y for x,y in pixel_spectrum.confs]
# Save the image in profile mode in imzML:
with ImzMLWriter('lipid_MSI_profile_mode.imzML') as writer:
for i in range(mask.shape[0]):
if not i % 50:
print(i)
for j in range(mask.shape[1]):
writer.addSpectrum(mass_axis, image[i,j,:], (i, j))
# Centroid the image and save it in imzML:
with ImzMLWriter('lipid_MSI_centroid_mode.imzML') as writer:
for i in range(mask.shape[0]):
if not i % 50:
print(i)
for j in range(mask.shape[1]):
S = Spectrum(confs=list(zip(mass_axis, image[i,j,:])))
peaks, _ = S.centroid(0.25, 0.4)
mzs = [p[0] for p in peaks]
intsys = [p[1] for p in peaks]
writer.addSpectrum(mzs, intsys, (i, j))
# ## Data validation
from pyimzml.ImzMLParser import ImzMLParser
profile_image = ImzMLParser('lipid_MSI_profile_mode.imzML')
centroid_image = ImzMLParser('lipid_MSI_centroid_mode.imzML')
profile_ion_image = np.zeros(mask.shape[:2])
centroid_ion_image = np.zeros(mask.shape[:2])
# peak_to_plot = 856.626 # PC_38_0
peak_to_plot = 855.68 # PA_44_0
# peak_to_plot = 852.525 # PC_38_2 and PS_38_3
# peak_to_plot = 850.53 # PC_38_3 and PS_38_4
for i, (x,y,z) in enumerate(profile_image.coordinates):
mz, intsy = profile_image.getspectrum(i)
mz = np.array(mz)
peak_id = np.argmin(np.abs(mz-peak_to_plot))
profile_ion_image[x, y] = intsy[peak_id]
mz, intsy = centroid_image.getspectrum(i)
mz = np.array(mz)
peak_id = np.argmin(np.abs(mz-peak_to_plot))
centroid_ion_image[x, y] = intsy[peak_id]
plt.figure(figsize=(8,3))
plt.subplot(121)
plt.title('Profile mode')
plt.imshow(profile_ion_image)
plt.subplot(122)
plt.title('Centroid mode')
plt.imshow(centroid_ion_image)
plt.tight_layout()
plt.show()
# ## Estimation of optimal MTD based on selected pixels
px = (20, 20)
print('pixel', px, 'from region', region_mapping[px])
plt.figure()
plt.subplot(121)
plt.plot(mass_axis, image[20, 20,:])
plt.subplot(122)
plt.plot(mass_axis, image[20, 10,:])
plt.show()
S1 = Spectrum(confs=list(zip(mass_axis, image[20,10,:])))
T1 = np.trapz(image[20,10,:], mass_axis)
S1.normalize()
S2 = Spectrum(confs=list(zip(mass_axis, image[20,20,:])))
T2 = np.trapz(image[20,20,:], mass_axis)
S2.normalize()
C1 = Spectrum(confs=S1.centroid(0.25, 0.4)[0])
C1.normalize()
C2 = Spectrum(confs=S2.centroid(0.25, 0.4)[0])
C2.normalize()
R1 = estimate_proportions(S1, spectra, MTD=0.2)
R2 = estimate_proportions(S2, spectra, MTD=0.2)
for ptrue, pestim in zip(region_intensities[1], R1['proportions']):
print(ptrue, round(pestim*sum(region_intensities[1]), 2))
for ptrue, pestim in zip(region_intensities[2], R2['proportions']):
print(ptrue, round(pestim*sum(region_intensities[2]), 2))
# Estimate MTD based on Region 2:
mtd_array = np.linspace(0.01, 1., num=400)
correlations_P = np.zeros(mtd_array.shape)
l1_P = np.zeros(mtd_array.shape) # profile
correlations_C = np.zeros(mtd_array.shape)
l1_C = np.zeros(mtd_array.shape) # centroid
for i, mtd in enumerate(mtd_array):
R2 = estimate_proportions(S2, spectra, MTD=mtd)
p2 = np.array(R2['proportions'])*sum(region_intensities[2])
l1_P[i] = sum(np.abs(p2 - region_intensities[2]))
correlations_P[i] = np.corrcoef(region_intensities[2], p2)[0,1]
R2 = estimate_proportions(C2, spectra, MTD=mtd)
p2 = np.array(R2['proportions'])*sum(region_intensities[2])
l1_C[i] = sum(np.abs(p2 - region_intensities[2]))
correlations_C[i] = np.corrcoef(region_intensities[2], p2)[0,1]
plt.figure(figsize=(6,3))
plt.subplot(121)
plt.title('Correlations')
plt.plot(mtd_array, correlations_P)
plt.plot(mtd_array, correlations_C)
plt.legend(['profile', 'centroid'])
plt.subplot(122)
plt.title('L1')
plt.plot(mtd_array, l1_P)
plt.plot(mtd_array, l1_C)
plt.legend(['profile', 'centroid'])
plt.tight_layout()
plt.show()
region_intensities[2]/sum(region_intensities[2])
np.corrcoef(R2['proportions'], region_intensities[2])
| Simulations/0. Image simulation.ipynb |