code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="P9bJCDjdlgG6" colab_type="text"
# # **Spit some [tensor] flow**
#
# Let's see some classifiers in action
#
# `Leggo`
# + id="aQwc0re5mFld" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + id="I-tpfuNjbj-q" colab_type="code" colab={}
# Visualising the results
def plot_model(classifier, X_set, y_set, y_test, y_pred, text):
from sklearn.metrics import accuracy_score
print("Accuracy Score :")
print(accuracy_score(y_test, y_pred))
return
# + id="qyw8HvOuBEZm" colab_type="code" colab={}
from sklearn.datasets import load_breast_cancer
data = load_breast_cancer()
# + id="EukWofARwuL9" colab_type="code" outputId="17659af5-4933-4385-9ceb-2b216d45c008" colab={"base_uri": "https://localhost:8080/", "height": 35}
data.keys()
# + id="9BjoFGYJwyS0" colab_type="code" colab={}
X = data.data
y = data.target
# + id="NbbCJdKu8l6w" colab_type="code" outputId="1d190fc9-a978-45e7-b9d7-6329b16e0644" colab={"base_uri": "https://localhost:8080/", "height": 35}
data.target_names
# + id="oUJyshkNW2YQ" colab_type="code" colab={}
# TRAIN TEST SPLIT
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="wKrVEKcmOH6I" colab_type="code" outputId="0f15d126-961e-43ff-bf80-ff9fcbb921ac" colab={"base_uri": "https://localhost:8080/", "height": 191}
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "Logistic Regression")
# + id="DG4OgIp1T4Nd" colab_type="code" colab={}
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="tFEMeP4FT7cb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="6c1e7e8a-6f74-4580-c763-57733172a552"
from sklearn.linear_model import LogisticRegression
classifier = LogisticRegression()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "Logistic Regression Feature Scaled")
# + id="9Thr6hcSUgNU" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="asjjB-OfP-V2" colab_type="code" outputId="b974c81c-b26b-4ee8-bbdf-d07d9d63767b" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.linear_model import RidgeClassifierCV
classifier = RidgeClassifierCV()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "RidgeClassifierCV")
# + id="NsqNNE4vUibr" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="QgZFqE-tUmG8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="b836f385-c847-417a-9a86-2d88185a9dc2"
from sklearn.linear_model import RidgeClassifierCV
classifier = RidgeClassifierCV()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "RidgeClassifierCV")
# + id="CUNrHU5RVHq7" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="j41ne3yuOLSm" colab_type="code" outputId="bfc56928-d7ed-4750-ce7f-61c2fc6ad1cf" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.svm import SVC
classifier = SVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SVC")
# + id="rb7GEKcpVLIL" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="jv5hbg4PVKxM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="9d522a13-976b-4738-9a61-139414f976dd"
from sklearn.svm import SVC
classifier = SVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SVC")
# + id="vwxUDZpcVyyW" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="G-H_ZGr5e1wk" colab_type="code" outputId="38bc1fdf-a60c-4ee2-932e-36f1d8fedc64" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.neural_network import MLPClassifier
classifier = MLPClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SVC")
# + id="kMy_BDIaV80k" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="IMzDo0DnVmRF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 107} outputId="6f5e3494-f8ff-463e-db1d-3f30784821c3"
from sklearn.neural_network import MLPClassifier
classifier = MLPClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SVC")
# + id="JgwnWyQfVzk1" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="BS6V1HSJad1f" colab_type="code" outputId="49d5b82b-5e4f-4fac-c3b7-6adef4031215" colab={"base_uri": "https://localhost:8080/", "height": 87}
from sklearn.svm import LinearSVC
classifier = LinearSVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "LinearSVC")
# + id="2ET4R3v4V9aU" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="z0kDhJKoVnbq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="9e59969a-d9a9-4171-d104-dba4a4767e10"
from sklearn.svm import LinearSVC
classifier = LinearSVC()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "LinearSVC")
# + id="2S5oHuofV0Kr" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="Cif5chCUbF3h" colab_type="code" outputId="d244a49d-b41e-45a8-e23e-d578750241ee" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "RandomForestClassifier")
# + id="HqgR8EiWV-Du" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="jP8Rm-TzVoXy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="155c79ab-9c6b-4297-ca12-53a6d5569b5c"
from sklearn.ensemble import RandomForestClassifier
classifier = RandomForestClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "RandomForestClassifier")
# + id="wdagMmauV0t-" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="vzRuKzGracLw" colab_type="code" outputId="34287811-52af-4621-b30e-075e1ab11e41" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.tree import DecisionTreeClassifier
classifier = DecisionTreeClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "DecisionTreeClassifier")
# + id="kQ7SKQimV-nf" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="Y0iG2CSvVpQq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="e36febd6-66cb-40eb-d1ec-f6480f375f6e"
from sklearn.tree import DecisionTreeClassifier
classifier = DecisionTreeClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "DecisionTreeClassifier")
# + id="w4HH6k1fV1K0" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="om8U4fa_bJop" colab_type="code" outputId="ae216722-68dd-4ce1-a9d6-3d458ae9ce10" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.ensemble import GradientBoostingClassifier
classifier = GradientBoostingClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "GradientBoostingClassifier")
# + id="-_UQDMl8V_GD" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="Ihmrh0aDVqCC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="208388db-157d-4456-faba-15650fcf7b27"
from sklearn.ensemble import GradientBoostingClassifier
classifier = GradientBoostingClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "GradientBoostingClassifier")
# + id="1pTXUOAyV2CW" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="gU2yGvvqbNaH" colab_type="code" outputId="62a7affe-ce70-4c3d-dbc6-4b7b98f3a67c" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.linear_model import SGDClassifier
classifier = SGDClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SGDClassifier")
# + id="C6c2pNSkV_nN" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="LdM17saDVq_T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="5f1bf97f-d0f0-4e0e-eaea-9af0d8253422"
from sklearn.linear_model import SGDClassifier
classifier = SGDClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "SGDClassifier")
# + id="hGwqgdIFV2ls" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="2Nr9fmGGcxjx" colab_type="code" outputId="2f152299-caf4-4601-eeb8-00e9ede218fb" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.linear_model import Perceptron
classifier = GradientBoostingClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "Perceptron")
# + id="jwKFPL3LWBYG" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="OLXVsjk1VsAa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="db4e2b67-8c67-43ae-dace-282bad70c7d0"
from sklearn.linear_model import Perceptron
classifier = GradientBoostingClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "Perceptron")
# + id="XtjnvnLUWCD7" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="bhfSkakWcx_g" colab_type="code" outputId="1c9dbb9e-0228-4d35-baaf-c6c92bdd68f9" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "GaussianNB")
# + id="JdqogsErWEM1" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="aaT1ig6hVsqE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="562d85be-603f-4dab-be74-20d98741519e"
from sklearn.naive_bayes import GaussianNB
classifier = GaussianNB()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "GaussianNB")
# + id="NGsgP_9DV4Tb" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.1)
# + id="JRMDGmgucyn4" colab_type="code" outputId="4f76ea2b-457c-4a2d-d086-2afd37e6b5a5" colab={"base_uri": "https://localhost:8080/", "height": 52}
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "KNeighborsClassifier")
# + id="obleHIUVWG2V" colab_type="code" colab={}
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# + id="8oWnP28zVtUr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="28b00025-1873-4513-de81-08dc4093d0b2"
from sklearn.neighbors import KNeighborsClassifier
classifier = KNeighborsClassifier()
classifier.fit(X_train,y_train)
y_pred = classifier.predict(X_test)
y_pred = np.round(y_pred).flatten()
plot_model(classifier, X_train, y_train, y_test, y_pred, "KNeighborsClassifier")
| MachineLearning_DataScience/Demo104_FeatureScaling_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import re
colitis_patt = r'\b(colitis|diarrhea|abdominal pains?|loose bowel movements?|(increase|increased) frequency of stools?|blood in stool|nausea|vomiting)\b'
tox_patts = [colitis_patt]
def match_tox_and_get_reduced_sent(text):
for tox_patt in tox_patts:
if re.search(tox_patt, text, re.I|re.M):
return text
return ""
# -
sentence = 'Does this actually work!.....abdominal pains...'
match_tox_and_get_reduced_sent(sentence)
| Data Preprocessing/SentenceCleaner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Autoregressive Moving Average (ARMA): Artificial data
# %matplotlib inline
import numpy as np
import statsmodels.api as sm
import pandas as pd
from statsmodels.tsa.arima_process import arma_generate_sample
np.random.seed(12345)
# Generate some data from an ARMA process:
arparams = np.array([.75, -.25])
maparams = np.array([.65, .35])
# The conventions of the arma_generate function require that we specify a 1 for the zero-lag of the AR and MA parameters and that the AR parameters be negated.
arparams = np.r_[1, -arparams]
maparams = np.r_[1, maparams]
nobs = 250
y = arma_generate_sample(arparams, maparams, nobs)
# Now, optionally, we can add some dates information. For this example, we'll use a pandas time series.
dates = sm.tsa.datetools.dates_from_range('1980m1', length=nobs)
y = pd.Series(y, index=dates)
arma_mod = sm.tsa.ARMA(y, order=(2,2))
arma_res = arma_mod.fit(trend='nc', disp=-1)
print(arma_res.summary())
y.tail()
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(10,8))
fig = arma_res.plot_predict(start='1999-06-30', end='2001-05-31', ax=ax)
legend = ax.legend(loc='upper left')
| examples/notebooks/tsa_arma_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="6RKbJM0frIDC"
#Principais importações
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import random
# + [markdown] id="cJzjRWSB8lxs"
# **Gráficos de linha**
# + id="CJlPSe109NPJ" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="d5dd49e3-427a-4b46-99e9-28f8c0ebb447"
# Exemplo 1: Gráfico de linha simples
plt.plot( [2,4,8,16,32] )
plt.title('Gráfico de linha')
plt.xlabel('Eixo X')
plt.ylabel('Eixo Y')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="Z3_IxUApLi81" outputId="790b2e66-43f6-4c38-ef40-aac0d5199d52"
# Exemplo 2: Gráfico de linha com formatação
x = np.linspace(1,5,5)
y = np.exp2(x)
plt.plot(x, y, '-p', color='gray',
markersize=15, linewidth=4,
markerfacecolor='white',
markeredgecolor='gray',
markeredgewidth=2)
plt.title('Gráfico de linha')
plt.xlabel('Eixo X')
plt.ylabel('Eixo Y')
plt.show()
# + id="2QblmUuvq5s-" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="93dfc41a-a011-4b5d-b801-589796ea34f2"
# Exemplo 3: Gráfico formatado com grade
fig, ax = plt.subplots()
t = np.arange(0.0, 2.0, 0.01)
s = 1 + np.sin(2 * np.pi * t)
ax.plot(t, s)
ax.set(xlabel='Tempo (s)', ylabel='Voltagem (mV)', title='Voltagem x Tempo')
ax.grid()
# Comando para salvar o gráfico em arquivo .png
# fig.savefig("Grafico_senoide.png")
plt.show()
# + [markdown] id="-zDsYf-w9oQ5"
# **Gráfico de barras**
# + colab={"base_uri": "https://localhost:8080/", "height": 452} id="PaSH0y1tB8fU" outputId="8c29b2cc-6569-4e60-8e4f-5df50cf52877"
# Fixando estado randômico
np.random.seed(1)
plt.rcdefaults()
fig, ax = plt.subplots()
# Dados
people = ('Device A', 'Device B', 'Device C', 'Device D', 'Device E')
y_pos = np.arange(len(people))
performance = 3 + 10 * np.random.rand(len(people))
# Gráfico
ax.bar(y_pos, performance, align='center')
ax.set_xticks(y_pos)
ax.set_xticklabels(people)
ax.set_ylabel('Performance')
ax.set_title('Medição de performance')
# Plot
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="5X5S4bp-EhmV" outputId="56d09c49-9914-47ca-d87d-edc8ea4371d6"
# Fixando estado randômico
np.random.seed(1)
plt.rcdefaults()
fig, ax = plt.subplots()
# Dados
people = ('Device A', 'Device B', 'Device C', 'Device D', 'Device E')
y_pos = np.arange(len(people))
performance = 3 + 10 * np.random.rand(len(people))
# Gráfico
ax.barh(y_pos, performance, align='center')
ax.set_yticks(y_pos)
ax.set_yticklabels(people)
ax.invert_yaxis()
ax.set_xlabel('Performance')
ax.set_title('Medição de performance')
# Plot
plt.show()
# + [markdown] id="6JpWmrMs9ssY"
# **Gráfico de pizza**
# + id="ykv4bIY-rFEj" colab={"base_uri": "https://localhost:8080/", "height": 406} outputId="3fb04eca-f560-4cd4-ce1c-65f80c350939"
labels = ['A', 'B', 'C', 'D']
sizes = [15, 30, 45, 10]
explode = (0, 0.1, 0, 0) # Destaca a fatia da segunda posição ( Ex: B )
fig1, ax1 = plt.subplots()
ax1.pie(sizes, explode=explode, labels=labels, autopct='%1.1f%%',
shadow=True, startangle=90)
ax1.axis('equal') # Assegura que a pizza será desenhada como um círculo
plt.show()
# + [markdown] id="ojt2S9HRqyaK"
# **Gráfico dispersão**
# + id="8eWzHnHEr57m" colab={"base_uri": "https://localhost:8080/", "height": 472} outputId="9df57de9-83a9-440b-a294-dcceb81a7c3e"
# Exemplo 1: Gráfico simples
x = np.linspace(0, 10, 30)
y = np.sin(x)
plt.plot(x, y, 'o', color='black')
plt.title('Gráfico de dispersão')
plt.xlabel('Eixo X')
plt.ylabel('Eixo Y')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 472} id="68VtfJ1XIgFR" outputId="9df7b66c-4f62-47b2-819c-c7cc66fb1993"
# Exemplo 2: Diferentes marcadores
for marker in ['o', '.', ',', 'x', '+', 'v', '^', '<', '>', 's', 'd']:
plt.plot(np.random.rand(3), np.random.rand(3), marker,
label=f"marker='{marker}'")
plt.legend(numpoints=1)
plt.xlim(0, 1.5)
plt.ylim(0, 1.5)
plt.title('Gráfico de dispersão')
plt.xlabel('Eixo X')
plt.ylabel('Eixo Y')
plt.show()
# + id="RZ0ez3Vdq5ee" colab={"base_uri": "https://localhost:8080/", "height": 472} outputId="a7dbed97-3296-4822-97e6-2d154d229c20"
# Exemplo 3: Dispersão com bolhas coloridas
N = 50
x = np.random.rand(N)
y = np.random.rand(N)
colors = np.random.rand(N)
area = (30 * np.random.rand(N))**2
plt.scatter(x, y, s=area, c=colors, alpha=0.35)
plt.title('Gráfico de dispersão')
plt.xlabel('Eixo X')
plt.ylabel('Eixo Y')
plt.show()
| Matplotlib.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Computation of atomic processes with JAC
#
# The following tutorials show how JAC can be utilized to compute the rates, cross sections and angular distributions for a (large) variety of atomic processes.
# ## 51-compute-Fe-X-spectrum.ipynb
# ### **Compute the spectrum of Fe X**
# We here consider the level structure of Fe$^{9+}$ ions which, in spectroscopic notation, give rise to the spectrum Fe X. Several medium-to-large scale computations have been carried out to calculate the excitation energies and transition probabilities of (and among) the levels from the [Ne] $3s^2 3p^5$ -> $3s^2 3p^4 3d + 3s 3p^6$ configuration. These three configurations comprise in total 31 fine-structure levels with total angular momenta $J = 1/2, ..., 9/2$ (cf. Dong et al., 1999). Although the $3s^2 3p^4 3d + 3s 3p^6$ levels mainly decay by electric-dipole (E1) transitions, also M1, E2 and M2 multipole transitions have been considered in the literature. Below, we show how this *complete* Fe X spectrum can be calculated approximately by just a single call within the JAC toolbox.
# ## 53-Ar-2s-2p-photoionization.ipynb
# ### **Photoionization of atomic argon: compute cross sections and angular parameters**
# Here, we show how one can calculate the cross sections and the $\beta_2$ angular parameters for the photoionzation of atomic argon by linearly-polarized synchrotron radiation. Since the ionization process depends on the energy and polarization of the incident radiatian, the **standard settings** (although available by default) are of little help, and at least the photon energies should normally be specified in advance.
# ## 54-compute-Ne-KLL-Auger-spectrum.ipynb
# ### **Compute the neon K-LL Auger spectrum**
# In this tutorial, we calculate the K-LL spectrum of atomic neon after $1s$ inner-shell ionization as it may occurs, for instance, following photon, electron or neutron impact. An autoionization of an atom or ion generally occurs if the initially-excited level is **energetically embedded into the continuum of the next higher charge state**, and then often also dominates the decay of the atom.
# ## 55-compute-Auger-rates-in-DH-plasma.ipynb
# ### **Compute K-LL Auger rates for neon-like iron in a Debye-Hückel potential**
# A plasma environment generally affects both, the level structure of atoms and ions as well their rates and cross sections. In this tutorial, we wish to compute the K-LL rates for neon-like iron in a Debye-Hückel potential following the work by Deprince *et al* (2019). --- A Debye-Hückel potential may affect the Auger rates of an atom or ion in three different ways: (i) the level structure due to the screened electron-nucleus and electron-electron interaction; (ii) the shape of the potential, seen by the emitted electrons, due to the plasma screening of the ionic core; as well as (iii) the screening of the electron-electron interaction within the Auger matrix. In the present implementation, only the (screened) Coulomb interaction can be taken into account for the computation of the plasma-modified Auger rates, i.e. **no** Breit interactions nor QED estimates are supported by the JAC program for such plasma computations.
# ## 56-compute-radiative-recombination-rates.ipynb
# ### **Radiative recombination of highly-charged lithium-like ions**
# We ...
# ## 58-compute-DR-resonance-strengths.ipynb
# ### **Dielectronic recombination of multiply-charged lithium-like ions**
# We ...
# ## 60-compute-two-photon-decay.ipynb
# ### **Bound-state two-photon decay of K-shell ionized ...**
# We ...
| tutorials/50-summarize-atomic-processes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# + deletable=false editable=false run_control={"frozen": true}
# **Prediction example:**
# ___
# In this example we will show how to:
# - Setup the required environment for accessing the ecosystem prediction server.
# - Upload data to ecosystem prediction server.
# - Load data into feature store and parse to frame.
# - Build and test a prediction model for prism scores.
# + deletable=false editable=false run_control={"frozen": true}
# ## Setup
# -
# **Setting up import path:**
# ___
# Add path of ecosystem notebook wrappers. It needs to point to the ecosystem notebook wrapper to allow access to the packages required for running the prediction server via python.
# - **notebook_path:** Path to notebook repository.
notebook_path = "/path of to ecosystem notebook repository"
# + deletable=false editable=false
# ---- Uneditible ----
import sys
sys.path.append(notebook_path)
# ---- Uneditible ----
# + deletable=false editable=false run_control={"frozen": true}
# **Import required packages:**
# ___
# Import and load all packages required for the following usecase.
# + deletable=false editable=false
# ---- Uneditible ----
import pymongo
from bson.son import SON
import pprint
import pandas as pd
import json
import numpy
import operator
import datetime
import time
import os
import matplotlib.pyplot as plt
from prediction import jwt_access
from prediction import notebook_functions
from prediction.apis import functions
from prediction.apis import data_munging_engine
from prediction.apis import worker_h2o
from prediction.apis import prediction_engine
from prediction.apis import worker_file_service
# %matplotlib inline
# ---- Uneditible ----
# + deletable=false editable=false run_control={"frozen": true}
# **Setup prediction server access:**
# ___
# Create access token for prediction server.
# - **url:** Url for the prediction server to access.
# - **username:** Username for prediction server.
# - **password:** Password for prediction server.
# -
url = "http://demo.ecosystem.ai:3001/api"
username = "<EMAIL>"
password = "password"
# + deletable=false editable=false
# ---- Uneditible ----
auth = jwt_access.Authenticate(url, username, password)
# ---- Uneditible ----
# + deletable=false editable=false heading_collapsed=true run_control={"frozen": true}
# ## Upload Data
# + deletable=false editable=false hidden=true run_control={"frozen": true}
# **List uploaded files:**
# ___
# List all files already uploaded.
# + deletable=false editable=false hidden=true
# ---- Uneditible ----
files = worker_file_service.get_files(auth, path="./", user=username)
files = files["item"]
for file in files:
file_name = file["name"]
fn_parts = file_name.split(".")
if len(fn_parts) > 1 and fn_parts[-1] != "log":
print(file_name)
# ---- Uneditible ----
# + deletable=false editable=false hidden=true run_control={"frozen": true}
# **List uploadable files:**
# ___
# List all files in path ready for upload to prediction server.
# + deletable=false editable=false hidden=true
# ---- Uneditible ----
path = "../example_data/"
upload_files = [f for f in os.listdir(path) if os.path.isfile(os.path.join(path, f))]
print(upload_files)
# ---- Uneditible ----
# + deletable=false editable=false hidden=true run_control={"frozen": true}
# **Upload file:**
# ___
# Select file to upload to prediction server.
# - **file_name:** file name of file to upload to prediction server. See list of available files for upload.
# + hidden=true
file_name = "multi_personality_tiny.csv"
# + deletable=false editable=false hidden=true
# ---- Uneditible ----
worker_file_service.upload_file(auth, path + file_name, "/data/")
# ---- Uneditible ----
# + deletable=false editable=false hidden=true run_control={"frozen": true}
# **List uploaded files:**
# ___
# List all files in path ready for upload to prediction server to compare with previous list to confirm that file was uploaded correctly.
# + deletable=false editable=false hidden=true
# ---- Uneditible ----
files = worker_file_service.get_files(auth, path="./", user=username)
files = files["item"]
for file in files:
file_name = file["name"]
fn_parts = file_name.split(".")
if len(fn_parts) > 1 and fn_parts[-1] != "log":
print(file_name)
# ---- Uneditible ----
# + hidden=true
j.RenderJSON(hexframename)
# + deletable=false editable=false run_control={"frozen": true}
# ## Build Model
# -
# **Train Model:**
# ___
# Set training parameters for model and train.
# - **predict_id:** Id for the prediction (for logging).
# - **description:** Description of model (for logging).
# - **model_id:** Id for the model (for logging).
# - **model_type:** Type of model to build (for logging).
# - **frame_name:** Name of frame used (for logging).
# - **frame_name_desc:** Description of frame used (for logging).
# - **model_purpose:** Purpose of model (for logging).
# - **version:** Model version (for logging).
#
# The following parameters are dependend on what is selected in the algo parameter.
#
# - **algo:** Algorithm to use to train model. (Availble algorithms: "H20-AUTOML", "PYTORCH")
# - **transformer:** Pytorch transformer to use. (Available transformers: "bert-base-uncase")
# - **model_name:** Output name of model being built.
# - **device:** Hardware on which to build model. (Available devices: "cpu")
# - **data_file_path:** Path to input data file.
# - **data_file_type:** Type of input data file. (Available types: "csv")
# - **model_path:** Path to output model.
# - **training_column:** Column in dataset containing training text.
# - **response_column:** Column in dataset containing predictor reponse.
# - **epochs:** Number of epochs for which to train the model.
# - **learning_rate:** (TODO)
# - **epsilon:** (TODO)
# - **seed:** Random seed with which to run training of model.
# - **model_checkpoint:** If set to True, will save model as a checkout of the base transformer, if false a whole model will be saved.
# - **train_test_split:** Percentage of data to use for validation.
# - **do_lower_case:** If True, for input text to lowercase.
# - **batch_size:** Number of rows in data to process concurrently.
# - **add_special_tokens:** If set to True, special tokens will be added to tokenized data.
# - **padding:** If input is less than max_length the fill rest of tokenized data with padding tokens.
# - **max_length:** Max length for tokenizers to allow.
# - **truncation:** If set to True, if input is more than max_length then data will be truncated to max_length. If set to False, if input is more than max_length model will not train.
#
version = "1010"
model_id = featurestore_name + version
model_purpose = "Prediction of personality based on text data."
description = "Automated features store generated for " + featurestore_name
model_params = {
"predict_id": featurestore_name,
"description": description,
"model_id": model_id,
"model_type": "PYTORCH",
"frame_name": hexframename,
"frame_name_desc": description,
"model_purpose": model_purpose,
"version": version,
"model_parms": {
"algo": "PYTORCH",
"transformer": "bert-base-uncased",
"transformer_configs": {
"model_name": "personality",
"device": "cpu",
"data_file_path": "data/multi_personality_tiny.csv",
"data_file_type": "csv",
"model_path": "modeling/personality.model",
"training_column": "text",
"response_column": "response",
"epochs": 1,
"learning_rate": 0.00001,
"epsilon": 1e-8,
"seed": 17,
"model_checkpoint": false,
"train_test_split": 0.15,
"do_lower_case": true,
"batch_size": 10,
"add_special_tokens": true,
"padding": true,
"max_length": 512,
"truncation": true
}
}
# + deletable=false editable=false
# ---- Uneditible ----
worker_h2o.train_model(auth, model_id, "pytorch", json.dumps(model_params["model_parms"]))
# ---- Uneditible ----
# + deletable=false editable=false run_control={"frozen": true}
# **View Model:**
# ___
# View autoML model to see which generated models are performing the best.
# -
# ---- Uneditible ----
model_data = worker_h2o.get_train_model(auth, model_id, "AUTOML")
notebook_functions.RenderJSON(model_data)
# ---- Uneditible ----
# +
sort_metric = model_data["leaderboard"]["sort_metric"]
model_names = []
for model in model_data["leaderboard"]["models"]:
model_names.append(model["name"])
model_metrics = model_data["leaderboard"]["sort_metrics"]
df = pd.DataFrame(
{
"model_names": model_names,
"model_metrics": model_metrics
}
)
df.sort_values("model_metrics", inplace=True, ascending=False)
ax = df.plot(y="model_metrics", x="model_names", kind="bar", align="center", alpha=0.5, legend=None)
plt.xticks(rotation=90)
ax.set_title("Performance of Models. Sorted Using Metric: {}".format(sort_metric))
ax.yaxis.grid(True)
# + deletable=false editable=false run_control={"frozen": true}
# **Save Model:**
# ___
# Save model for prediction.
# - **model_id:** Id for the model to save.
# -
best_model = df.iloc[0]["model_names"]
h2o_name = best_model
zip_name = h2o_name + ".zip"
# worker_h2o.download_model_mojo(auth, h2o_name)
high_level_mojo = worker_h2o.get_train_model(auth, h2o_name, "single")
model_to_save = high_level_mojo["models"][0]
model_to_save["model_identity"] = h2o_name
model_to_save["userid"] = "user"
model_to_save["timestamp"] = "time_stamp"
prediction_engine.save_model(auth, model_to_save)
# + deletable=false editable=false run_control={"frozen": true}
# **View Model Stats:**
# ___
# View stats of saved model.
# -
prediction_engine.get_user_model(auth, h2o_name)
stats = worker_h2o.get_model_stats(auth, h2o_name, "ecosystem", "variable_importances")
notebook_functions.RenderJSON(stats)
| data_science/00_From_Ingestion_to_Prediction_PyTorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p><font size="6"><b>03 - Pandas: Indexing and selecting data - Part I</b></font></p>
#
# > *© 2016, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
# + run_control={"frozen": false, "read_only": false}
import pandas as pd
# +
# redefining the example objects
# series
population = pd.Series({'Germany': 81.3, 'Belgium': 11.3, 'France': 64.3,
'United Kingdom': 64.9, 'Netherlands': 16.9})
population
# + run_control={"frozen": false, "read_only": false}
# dataframe
data = {'country': ['Belgium', 'France', 'Germany', 'Netherlands', 'United Kingdom'],
'population': [11.3, 64.3, 81.3, 16.9, 64.9],
'area': [30510, 671308, 357050, 41526, 244820],
'capital': ['Brussels', 'Paris', 'Berlin', 'Amsterdam', 'London']}
countries = pd.DataFrame(data)
countries
# -
# # Changing the DataFrame index
# We have mostly worked with DataFrames with the default *0, 1, 2, ... N* row labels. But, we can also set one of the columns as the index.
#
# Setting the index to the country names with [`set_index()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.set_index.html):
# + run_control={"frozen": false, "read_only": false}
countries = countries.set_index('country')
countries
# -
# Reversing this operation, is [`reset_index()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.reset_index.html):
# + run_control={"frozen": false, "read_only": false}
countries.reset_index('country')
# -
# # Selecting data based on the index
# ## Systematic indexing with `loc` and `iloc`
# We have used `[]` to obtain one column of a dataframe (e.g., `countries['area']`).
#
# Pandas has the attributes `.loc` and `.iloc` for indexing:
#
# * `loc`: selection by (row and column) labels
# * `iloc`: selection by position (like numpy)
#
# Both `loc` and `iloc` use the following pattern:
#
# `df.loc[ <selection of the rows> , <selection of the columns> ]`.
#
# This 'selection of the rows / columns' can be:
#
# * a single label
# * a list of labels
# * a slice or
# * a boolean mask.
#
# `loc` and `iloc` prevent any ambiguity when indexing and it is the preferred way to index in pandas. One of pandas' basic features is the labeling of rows and columns, but this makes indexing also a bit more complex compared to numpy.
#
# We now have to distuinguish between:
#
#
# - selection by **label** (using the row and column names)
# - selection by **position** (using integers)
#
# This is becomes very important if the row or column labels are numbers. For example, `countries` has the numbers 0 to n-1 as the index, so there is no ambiguity here:
countries
# However, if we changed the index to be different numbers:
countries.set_index(pd.Index([2, 3, 1, 6, 4]))
# If you use numbers to index here, e.g., `countries[2]` pandas does not know if you mean the 2nd row or if you want the row that is labeled `2`. These are different rows!
#
# This is why `loc` and `iloc` are important. Using `loc` indicates that we want to use the row label/index and `iloc` indicates that we want to use position.
# ## `loc`
#
# Selecting a single element:
countries
# + run_control={"frozen": false, "read_only": false}
countries.loc['Germany', 'area']
# -
# But the row or column indexer can also be a list, slice, boolean array (see next section), ..
# + run_control={"frozen": false, "read_only": false}
countries.loc['France':'Germany', ['area', 'population']]
# -
# <div class="alert alert-danger">
# <b>NOTE</b>:
#
# <ul>
# <li>Unlike slicing in numpy, the end label is **included**!</li>
# </ul>
# </div>
countries.loc[countries['area'] > 100000]
# ## Useful functions for boolean indexing:
#
# The [`isin`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.isin.html) method of Series is very useful to select rows that may contain certain values:
countries['capital'].isin(['Berlin', 'London'])
# This can then be used to filter the dataframe with boolean indexing:
countries['capital'].isin(['Berlin', 'London'])
countries.loc[countries['capital'].isin(['Berlin', 'London']),]
# Python [string methods](https://docs.python.org/3/library/stdtypes.html#string-methods) (such as [`startswith`](https://docs.python.org/3/library/stdtypes.html#str.startswith), [`replace`](https://docs.python.org/3/library/stdtypes.html#str.replace), etc.) have a pandas version: [`str.startswith`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.str.startswith.html), [`str.replace`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.str.replace.html), etc.
countries['capital'].str.startswith('B')
countries.loc[countries['capital'].str.startswith('B')]
# For an overview of all pandas string methods, see: [Working with text data](https://pandas.pydata.org/pandas-docs/stable/user_guide/text.html)
# # Exercises using the Titanic dataset
df = pd.read_csv("../data/titanic.csv")
df.head()
# <div class="alert alert-success">
#
# **EXERCISE:**
#
# - Select all the rows that contain male passengers and assign it to the variable `males`.
# - Calculate the mean age of the male passengers.
# - Do the same for the female passengers, but in a single statement.
#
# </div>
# +
# # %load _solutions/pandas_03a_selecting_data1.py
# +
# # %load _solutions/pandas_03a_selecting_data2.py
# +
# # %load _solutions/pandas_03a_selecting_data3.py
# -
# <div class="alert alert-success">
#
# **EXERCISE:**
#
# - Select the passengers that are between 30 and 40 years old?</li>
#
# Hint: You may find useful the section of the user guide that explains boolean operators: [Boolean indexing](https://pandas.pydata.org/pandas-docs/stable/user_guide/indexing.html#boolean-indexing).
# </div>
# +
# # %load _solutions/pandas_03a_selecting_data6.py
# -
# <div class="alert alert-success">
#
# **EXERCISE:**
#
# - Split the 'Name' column on the comma (`','`), take the first part (the surname), and add this as a new column named 'Surname'.
#
#
# Hint: Try first on a single string (and for this, check the [`split`](https://docs.python.org/3/library/stdtypes.html#str.split) method of a Python string), and then try to [`apply`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.apply.html) this on each row.
#
# </div>
# +
# # %load _solutions/pandas_03a_selecting_data7.py
# -
df.head()
# <div class="alert alert-success">
#
# **EXERCISE:**
#
# - Select all passenger that have a surname starting with 'Williams'.
#
# Hint: Use a pandas string method.
# </div>
# +
# # %load _solutions/pandas_03a_selecting_data8.py
# -
# <div class="alert alert-success">
#
# **EXERCISE:**
#
# - Select all rows for the passengers with a surname of more than 15 characters.
#
# Hint: Use a pandas string method.
# </div>
# +
# # %load _solutions/pandas_03a_selecting_data9.py
# -
# ---
#
# ## `iloc`
#
# Selecting by position with `iloc` works similar as **indexing numpy arrays**:
#
# Remember the syntax: `start:stop:step`
# + run_control={"frozen": false, "read_only": false}
countries.iloc[0:4:2, 1:3]
# -
# Next let's make a copy of our dataframe:
countries2 = countries.copy()
# The [`.copy()`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.copy.html) method makes a copy of the DataFrame. Simply assigning (e.g., `countries2 = countries`) gives you a view NOT a copy, like with numpy arrays.
# ---
#
# We can also use the different indexing methods can also be used to **assign data**:
# + run_control={"frozen": false, "read_only": false}
countries2.loc['Belgium':'Germany', 'population'] = 10
# + run_control={"frozen": false, "read_only": false}
countries2
# -
# <div class="alert alert-info">
# <b>REMEMBER</b>:
#
# Advanced indexing with **loc** and **iloc**
#
# - **loc**: select by label: `df.loc[row_indexer, column_indexer]`
# - **iloc**: select by position: `df.iloc[row_indexer, column_indexer]`
#
# </div>
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <p>
# <ul>
# <li>Add the population density as column to the DataFrame.</li>
# </ul>
# </p>
# Note: the population column is expressed in millions.
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_03b_indexing1.py
# -
countries
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>Select the capital and the population columns of those countries where the density is larger than 300.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_03b_indexing2.py
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Add a column 'density_ratio' with the ratio of the population density to the average population density for all countries.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_03b_indexing3.py
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Change the capital of the UK to Cambridge.</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_03b_indexing4.py
# -
# <div class="alert alert-success">
# <b>EXERCISE</b>:
#
# <ul>
# <li>Select all countries whose population density is between 100 and 300 people/km²</li>
# </ul>
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# # %load _solutions/pandas_03b_indexing5.py
# -
# # Alignment on the index
# <div class="alert alert-danger">
#
# **WARNING**: **Alignment!** (unlike numpy)
#
# - Pay attention to **alignment**: operations between series will align on the index:
#
# </div>
# + run_control={"frozen": false, "read_only": false}
population = countries['population']
s1 = population[['Belgium', 'France']]
s2 = population[['France', 'Germany']]
# + run_control={"frozen": false, "read_only": false}
s1
# + run_control={"frozen": false, "read_only": false}
s2
# + run_control={"frozen": false, "read_only": false}
s1 + s2
# -
# # Pitfall: chained indexing (and the 'SettingWithCopyWarning')
# + run_control={"frozen": false, "read_only": false}
countries.loc['Belgium', 'capital'] = 'Ghent'
# + run_control={"frozen": false, "read_only": false}
countries
# + run_control={"frozen": false, "read_only": false}
countries['capital']['Belgium'] = 'Antwerp'
# + run_control={"frozen": false, "read_only": false}
countries
# + run_control={"frozen": false, "read_only": false}
countries[countries['capital'] == 'Antwerp']['capital'] = 'Brussels'
# + run_control={"frozen": false, "read_only": false}
countries
# -
countries.loc[countries['capital'] == 'Antwerp', 'capital'] = 'Brussels'
countries
# <div class="alert alert-info">
#
# <b>REMEMBER!</b><br><br>
#
# The warning is given because chain indexing produces unpredictable results - it may return a copy or a view of the original data.
#
# What to do when encountering the *value is trying to be set on a copy of a slice from a DataFrame* error?
#
# - Use `loc` instead of chained indexing **if possible**!
# - Or `copy` explicitly if you don't want to change the original data.
#
# </div>
# # Exercises using the Titanic dataset
df.head()
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# - Select all rows for male passengers (using `.loc`) and calculate the mean age of those passengers.
# - Do the same for the female passengers.
#
# </div>
# + clear_cell=true
# # %load _solutions/pandas_03b_indexing6.py
# + clear_cell=true
# # %load _solutions/pandas_03b_indexing7.py
# -
# We will later see an easier way to calculate both averages at the same time with [`groupby`](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.groupby.html).
| Day_1_Scientific_Python/pandas/pandas_03a_indexing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finding Contours
# ### Import resources and display image
# +
import numpy as np
import matplotlib.pyplot as plt
import cv2
# %matplotlib inline
# Read in the image
image = cv2.imread('images/thumbs_up_down.jpg')
# Change color to RGB (from BGR)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
plt.imshow(image)
# -
# ### Produce a binary image for finding contours
# +
# Convert to grayscale
gray = cv2.cvtColor(image,cv2.COLOR_RGB2GRAY)
# Create a binary thresholded image
retval, binary = cv2.threshold(gray, 225, 255, cv2.THRESH_BINARY_INV)
plt.imshow(binary, cmap='gray')
# -
# ### Find and draw the contours
# +
# Find contours from thresholded, binary image
retval, contours, hierarchy = cv2.findContours(binary, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
# Draw all contours on a copy of the original image
contours_image = np.copy(image)
contours_image = cv2.drawContours(contours_image, contours, -1, (0,255,0), 3)
plt.imshow(contours_image)
# -
# ## Contour Features
#
# Every contour has a number of features that you can calculate, including the area of the contour, it's orientation (the direction that most of the contour is pointing in), it's perimeter, and many other properties outlined in [OpenCV documentation, here](http://opencv-python-tutroals.readthedocs.io/en/latest/py_tutorials/py_imgproc/py_contours/py_contour_properties/py_contour_properties.html).
#
# In the next cell, you'll be asked to identify the orientations of both the left and right hand contours. The orientation should give you an idea of which hand has its thumb up and which one has its thumb down!
# ### Orientation
#
# The orientation of an object is the angle at which an object is directed. To find the angle of a contour, you should first find an ellipse that fits the contour and then extract the `angle` from that shape.
#
# ```python
# # Fit an ellipse to a contour and extract the angle from that ellipse
# (x,y), (MA,ma), angle = cv2.fitEllipse(selected_contour)
# ```
#
# **Orientation values**
#
# These orientation values are in degrees measured from the x-axis. A value of zero means a flat line, and a value of 90 means that a contour is pointing straight up!
#
# So, the orientation angles that you calculated for each contour should be able to tell us something about the general position of the hand. The hand with it's thumb up, should have a higher (closer to 90 degrees) orientation than the hand with it's thumb down.
#
# ### TODO: Find the orientation of each contour
# +
## TODO: Complete this function so that
## it returns the orientations of a list of contours
## The list should be in the same order as the contours
## i.e. the first angle should be the orientation of the first contour
def orientations(contours):
"""
Orientation
:param contours: a list of contours
:return: angles, the orientations of the contours
"""
# Create an empty list to store the angles in
# Tip: Use angles.append(value) to add values to this list
angles = []
return angles
# ---------------------------------------------------------- #
# Print out the orientation values
angles = orientations(contours)
print('Angles of each contour (in degrees): ' + str(angles))
# -
# ### Bounding Rectangle
#
# In the next cell, you'll be asked to find the bounding rectangle around the *left* hand contour, which has its thumb up, then use that bounding rectangle to crop the image and better focus on that one hand!
#
# ```python
# # Find the bounding rectangle of a selected contour
# x,y,w,h = cv2.boundingRect(selected_contour)
#
# # Draw the bounding rectangle as a purple box
# box_image = cv2.rectangle(contours_image, (x,y), (x+w,y+h), (200,0,200),2)
# ```
#
# And to crop the image, select the correct width and height of the image to include.
#
# ```python
# # Crop using the dimensions of the bounding rectangle (x, y, w, h)
# cropped_image = image[y: y + h, x: x + w]
# ```
#
# ### TODO: Crop the image around a contour
# +
## TODO: Complete this function so that
## it returns a new, cropped version of the original image
def left_hand_crop(image, selected_contour):
"""
Left hand crop
:param image: the original image
:param selectec_contour: the contour that will be used for cropping
:return: cropped_image, the cropped image around the left hand
"""
## TODO: Detect the bounding rectangle of the left hand contour
## TODO: Crop the image using the dimensions of the bounding rectangle
# Make a copy of the image to crop
cropped_image = np.copy(image)
return cropped_image
## TODO: Select the left hand contour from the list
## Replace this value
selected_contour = None
# ---------------------------------------------------------- #
# If you've selected a contour
if(selected_contour is not None):
# Call the crop function with that contour passed in as a parameter
cropped_image = left_hand_crop(image, selected_contour)
plt.imshow(cropped_image)
| Lessons&CourseWorks/1. IntroToComputerVision/3.TypeOfFeatures&Segmentation/2.Contour&Features/Contour detection and features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="I08sFJYCxR0Z"
# 
# + [markdown] colab_type="text" id="VJbAaitZrXus"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/5.Spark_OCR.ipynb.ipynb)
# + [markdown] colab_type="text" id="Niy3mZAjoayg"
# # Spark OCR
#
# ### .. will be enriched ... work in progress ...
# -
from sparkocr.databricks import create_init_script_for_tesseract
create_init_script_for_tesseract()
from sparkocr.databricks import display_images, OCR_MODEL_DIR
# + colab={"base_uri": "https://localhost:8080/", "height": 216} colab_type="code" id="DFHbWwX4juxv" outputId="f3a5b571-f8c0-47cf-b540-c68cfa160c10"
import sparkocr
import sys
from pyspark.sql import SparkSession
from sparkocr import start
import os
spark = start(secret=os.environ['JSL_OCR_SECRET'])
spark
# + colab={"base_uri": "https://localhost:8080/", "height": 283} colab_type="code" id="Kj8tsYfljiBP" outputId="c29778e6-5696-4638-e279-d89f72c67521"
# %sh
wget 'http://www.asx.com.au/asxpdf/20171103/pdf/43nyyw9r820c6r.pdf'
# -
dbutils.fs.cp("file:/databricks/driver/43nyyw9r820c6r.pdf", "dbfs:/")
dbutils.fs.mv("dbfs:/43nyyw9r820c6r.pdf", "dbfs:/sample_pdf_ocr.pdf")
# + colab={} colab_type="code" id="FwAGTafIj4Bi"
import base64
from sparkocr.transformers import *
from pyspark.ml import PipelineModel
def pipeline():
# Transforrm PDF document to struct image format
pdf_to_image = PdfToImage()
pdf_to_image.setInputCol("content")
pdf_to_image.setOutputCol("image")
pdf_to_image.setResolution(200)
# Run tesseract OCR
ocr = TesseractOcr()
ocr.setInputCol("image")
ocr.setOutputCol("text")
ocr.setConfidenceThreshold(65)
ocr.setTessdata(OCR_MODEL_DIR)
pipeline = PipelineModel(stages=[
pdf_to_image,
ocr
])
return pipeline
# + colab={} colab_type="code" id="xNh4i1Woj6oo"
pdf = 'sample_pdf_ocr.pdf'
pdf_example_df = spark.read.format("binaryFile").load(pdf).cache().repartition(4)
pdf_example_df.show(3)
# -
display(pdf_example_df)
# + colab={} colab_type="code" id="Tp7a9yMqkCGu"
result = pipeline().transform(pdf_example_df).cache()
# + colab={"base_uri": "https://localhost:8080/", "height": 141} colab_type="code" id="VWND8q95kE47" outputId="08e20fb8-9792-4f12-8db7-6d8f4e78180a"
result.select("pagenum","text", "confidence").show()
# + colab={"base_uri": "https://localhost:8080/", "height": 141} colab_type="code" id="LJ4EB7lTkNIG" outputId="eea41119-c908-404a-a18a-6018b2f2efd3"
result.select("pagenum","text", "confidence").show()
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="rj7rv4b7kTNo" outputId="a46bfa6c-8a5c-4133-b6dd-0ec78a8f9790"
result.select("text").collect()
# + colab={"base_uri": "https://localhost:8080/", "height": 781} colab_type="code" id="bqbnotCfkK58" outputId="a3d08d14-9096-4d08-82dc-ba3b569ce33e"
print("\n".join([row.text for row in result.select("text").collect()]))
# -
display_images(PdfToImage().setOutputCol("image").transform(pdf_example_df), limit=3)
result = pipeline().transform(pdf_example_df).cache()
display(result.select("pagenum", "text", "confidence"))
# + colab={} colab_type="code" id="BAXZ3PFNjmgo"
# %sh
wget https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/ocr/datasets/news.2B/5/8205_003.2B.tif
# -
# ## Image parsing with Spark OCR
def pipeline():
# Transforrm binary data to struct image format
binary_to_image = BinaryToImage()
binary_to_image.setInputCol("content")
binary_to_image.setOutputCol("image")
# Run tesseract OCR
ocr = TesseractOcr()
ocr.setInputCol("image")
ocr.setOutputCol("text")
ocr.setConfidenceThreshold(65)
ocr.setTessdata(OCR_MODEL_DIR)
pipeline = PipelineModel(stages=[
binary_to_image,
ocr
])
return pipeline
dbutils.fs.mv("file:/databricks/driver/8205_003.2B.tif", "dbfs:/sample_image_OCR.tif")
# %sh
OCR_DIR=/dbfs/tmp/ocr
if [ ! -d "$OCR_DIR" ]; then
mkdir $OCR_DIR
cd $OCR_DIR
wget https://s3.amazonaws.com/auxdata.johnsnowlabs.com/public/ocr/datasets/images.zip
unzip images.zip
fi
display(dbutils.fs.ls("dbfs:/tmp/ocr/images/"))
# +
images_path = "/tmp/ocr/images/*.tif"
images_example_df = spark.read.format("binaryFile").load(images_path).cache()
display(images_example_df)
# +
# ACCESS_KEY = ""
# SECRET_KEY = ""
# sc._jsc.hadoopConfiguration().set("fs.s3n.awsAccessKeyId", ACCESS_KEY)
# sc._jsc.hadoopConfiguration().set("fs.s3n.awsSecretAccessKey", SECRET_KEY)
# imagesPath = "s3a://dev.johnsnowlabs.com/ocr/datasets/news.2B/0/*.tif"
# imagesExampleDf = spark.read.format("binaryFile").load(imagesPath).cache()
# display(imagesExampleDf)
# -
images_example_df.count()
display_images(BinaryToImage().transform(images_example_df), limit=3)
result = pipeline().transform(images_example_df).cache()
display(result.select("text", "confidence"))
| tutorials/Certification_Trainings/Healthcare/databricks_notebooks/2.4/5_Spark_OCR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# --- about
# this file starts with notebook data3 and adds legends to its plots
# notebooks left to do:
# bootstrapping
# kfold
# per spectrum plots
# +
# --- set up environment
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.artist as art
from sklearn.decomposition import FastICA
# for Set working folder, etc.
import os
import os.path
# for sampling
import random
# test code
# print(os.getcwd())
# +
# --- etl spectrum files
# turn files into lists and cut off first line
# put into database "spectra"
# turn deleted points (flagged as -1.23e+34) into 0's
# label spectra with datafile titles
# init spectra and class databases
datafolder = "C:/Users/Bonita/Documents/GitHub/spectra-analysis/data/data3"
num_samples = len([name for name in os.listdir(datafolder) if \
os.path.isfile(os.path.join(datafolder,name))])
# test code
print(num_samples)
spectrum_len = 480 # 480 >= the BECK spectrometer spectrum length
spectra = np.zeros((num_samples,spectrum_len))
spectrum_names = ["" for x in range(num_samples)]
y = np.zeros((num_samples, 1))
# fill databases
i = 0
# find spectra folder
os.chdir(datafolder)
# etl the data
# label data for plotting legend
for filename in os.listdir(datafolder):
file_object = open(filename, 'r').readlines()
# strip off header, add to matrix 'spectra'
spectra[i,:] = file_object[1:]
# label spectrum class, based on header
# actinolite: 0, alunite: 1, chlorite: 2
file_header = file_object[0]
spectrum_names[i] = file_header
# testcode
# print(material_name)
if file_header.find('Actinolite',)!= -1: #if material name contains actinolite
y[i,0] = 0
elif file_header.find('Alun',)!= -1:
y[i,0] = 1
else: #chlorite
y[i,0] = 2
# turn deleted points into 0
for j in range(spectrum_len):
if spectra[i,j] <0:
spectra[i,j]=0
i+=1
# test code
# print(y[:])
# print(spectra[0:2,0])
# +
# --- etl
# divide up data randomly
# 80% training data 20% test for this pilot
# eventually, 60% training set, 20% validation set, 20% test set
sample_indices = list(range(0, num_samples))
train_set_size = 4*(num_samples//5)
test_set_size= num_samples-train_set_size
train_set_indices = random.sample(sample_indices,train_set_size)
test_set_indices = np.setdiff1d(sample_indices,train_set_indices) #fixed bug: take remaining samples after making train set
# test code
# print(train_set_indices)
# print(test_set_indices)
# make train and test sets
train_set = spectra[train_set_indices, :]
train_labels = y[train_set_indices, :]
test_set = spectra[test_set_indices, :]
test_labels = y[test_set_indices, :]
# +
# --- plot the classes
# plot each class in a separate plot
# plot spectra names in legend
num0 = 0 #number of samples of class 0
num1 = 0
num2 = 0
# count the number of each class to make spectra0, spectra1, spectra2 databases
for i in range(num_samples):
if y[i,0]== 0:
num0 += 1
elif y[i,0]== 1:
num1 += 1
elif y[i,0]== 2:
num2 += 1
# make class-specific databases spectra0, ...1, ...2
# class-specific datasets for spectrum names too
spectra0 = np.zeros((num0,spectrum_len))
spectra1 = np.zeros((num1,spectrum_len))
spectra2 = np.zeros((num2,spectrum_len))
labels0 = ["" for x in range(num0)]
labels1 = ["" for x in range(num1)]
labels2 = ["" for x in range(num2)]
# make counters for each database to place spectra
i0 = 0
i1 = 0
i2 = 0
# populate class-specific databases spectra0, ...1, ...2
for i in range(num_samples):
if y[i,0]== 0:
spectra0[i0,:] = spectra[i,:]
labels0[i0] = spectrum_names[i]
i0 +=1
elif y[i,0]== 1:
spectra1[i1,:] = spectra[i,:]
labels1[i1] = spectrum_names[i]
i1 +=1
else:
spectra2[i2,:] = spectra[i,:]
labels2[i2] = spectrum_names[i]
i2 +=1
# plot each class-specific database separately
for i in range(i0):
plt.plot(range(1, spectrum_len+1), spectra0[i,:], label = labels0[i])
plt.legend(bbox_to_anchor=(1.1, 1.05))
plt.show()
for i in range(i1):
plt.plot(range(1, spectrum_len+1), spectra1[i,:], label = labels1[i])
plt.legend(bbox_to_anchor=(1.1, 1.05))
plt.show()
for i in range(i2):
plt.plot(range(1, spectrum_len+1), spectra2[i,:], label = labels2[i])
plt.legend(bbox_to_anchor=(1.1, 1.05))
plt.show()
# -
# --- kfold cross validation
# +
# --- Generate ICA spectra
# ICA Parameters
num_components = 25
# Create FastICA object
ica = FastICA(n_components=num_components)
# Fit ICA model
X = spectra.T
S_ = ica.fit_transform(X)
A_ = ica.mixing_ # Estimated independent components (mixing matrix)
print(X.shape)
print(S_.shape)
print(A_.shape)
#for i in range(A_.shape[1]):
# plt.title('Component {}'.format(i))
# plt.plot(A_[:,i])
# plt.figure()
# +
# --- make plain nn model
model = keras.Sequential([
keras.layers.Dense(300, activation=tf.nn.relu),
keras.layers.Dense(100, activation=tf.nn.relu),
keras.layers.Dense(3, activation=tf.nn.softmax)
])
# compile
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# train
model.fit(train_set, train_labels, epochs=15)
# +
# --- test plain nn
test_loss, test_acc = model.evaluate(test_set, test_labels)
print('NN Test accuracy:', test_acc)
# +
# --- transform train and test set into components
train_transformed = S_[train_set_indices, :]
test_transformed = S_[test_set_indices, :]
print(train_transformed.shape)
print(test_transformed.shape)
# +
# --- visualize transformed spectra
# --- graph the classes
num0 = 0 #number of samples of class 0
num1 = 0
num2 = 0
# count the number of each class to make spectra0, spectra1, spectra2 databases
for i in range(num_samples):
if y[i,0]== 0:
num0 += 1
elif y[i,0]== 1:
num1 += 1
elif y[i,0]== 2:
num2 += 1
# make class-specific databases spectra0, ...1, ...2
icaspectra0 = np.zeros((num0, num_components))
icaspectra1 = np.zeros((num1, num_components))
icaspectra2 = np.zeros((num2, num_components))
# make counters for each database to place spectra
i0 = 0
i1 = 0
i2 = 0
# populate class-specific databases spectra0, ...1, ...2
for i in range(num_samples):
if y[i,0]== 0:
icaspectra0[i0,:] = S_[i,:]
i0 +=1
elif y[i,0]== 1:
icaspectra1[i1,:] = S_[i,:]
i1 +=1
else:
icaspectra2[i2,:] = S_[i,:]
i2 +=1
# plot each class-specific database separately
for i in range(i0):
plt.plot(icaspectra0[i,:])
plt.figure()
for i in range(i1):
plt.plot(icaspectra1[i,:])
plt.figure()
for i in range(i2):
plt.plot(icaspectra2[i,:])
# +
# --- build NN on ICA model
model = keras.Sequential([
keras.layers.Dense(100, activation=tf.nn.relu),
keras.layers.Dense(3, activation=tf.nn.softmax)
])
# compile
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# train on ica
model.fit(train_transformed, train_labels, epochs=15)
# +
# --- test nn on ica
test_loss, test_acc = model.evaluate(test_transformed, test_labels)
print('NN Test accuracy:', test_acc)
| lab-notebook/bsong/2019-03-08 - from spectra to general data science notebook/2019-07-03-BPS - legends.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp distributed.models.lgb
# -
# # LGBMForecast
#
# > LightGBM forecaster
# Wrapper of `lightgbm.dask.DaskLGBMRegressor` that adds a `model_` property that contains the fitted booster and is sent to the workers to in the forecasting step.
# +
#export
import warnings
import lightgbm as lgb
# -
#export
class LGBMForecast(lgb.dask.DaskLGBMRegressor):
if lgb.__version__ <= '3.2.1':
warnings.warn(
"It is recommended to build LightGBM from source following the instructions here: "
"https://github.com/microsoft/LightGBM/tree/master/python-package#install-from-github, since "
"the current LightGBM version might be affected by https://github.com/microsoft/LightGBM/issues/4026, "
"which was fixed after 3.2.1."
)
@property
def model_(self):
return self.booster_
| nbs/distributed.models.lgb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Monte Carlo - Predicting Gross Profit - Part I
# *Suggested Answers follow (usually there are multiple ways to solve a problem in Python).*
# Imagine you are an experienced manager and you have forecasted revenues of \$200mln, with an expected deviation of $10mln. You are convinced Cogs will be near 40% of the revenues, and their expected deviation is 20% of its own value.
# Use NumPy’s random.random function to simulate the potential revenue stream for 250 iterations (which is the number of trading days in a year) and then the predicted Cogs value.
import numpy as np
import matplotlib.pyplot as plt
rev_m = 200
rev_stdev = 10
iterations = 250
rev = np.random.normal(rev_m, rev_stdev, iterations)
rev
# Plot the obtained data for revenues and Cogs on a graph and observe the behavior of the obtained values.
plt.figure(figsize=(15, 6))
plt.plot(rev)
plt.show()
# +
COGS = - (rev * np.random.normal(0.4,0.2))
plt.figure(figsize=(15, 6))
plt.plot(COGS)
plt.show()
# -
# Cogs mean:
COGS.mean()
# Cogs std:
COGS.std()
| Python for Finance - Code Files/100 Monte Carlo - Predicting Gross Profit - Part I/Python 3/MC - Predicting Gross Profit - Part I - Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from IPython.display import display
from bqplot import (Figure, Map, Mercator, Orthographic, ColorScale, ColorAxis,
AlbersUSA, topo_load, Tooltip)
# ## Basic Map ##
sc_geo = Mercator()
x = Map(scales={'projection': sc_geo})
fig = Figure(marks=[x], title='Basic Map Example')
display(fig)
# ## Advanced Map and Projection ##
sc_geo = Orthographic(scale_factor=375, center=[0, 25], rotate=(-50, 0))
x = Map(map_data=topo_load('WorldMapData.json'), scales={'projection': sc_geo},
colors={682: 'Green', 356: 'Red', 643: '#0000ff', 'default_color': 'DarkOrange'})
fig = Figure(marks=[x], fig_color='deepskyblue', title='Advanced Map Example')
display(fig)
sc_geo.scale = 350
# ## Choropleth ##
# +
sc_geo = Mercator()
sc_c1 = ColorScale(scheme='YlOrRd')
map_styles = {'color': {643: 105., 4: 21., 398: 23., 156: 42., 124:78., 76: 98.},
'scales': {'projection': sc_geo, 'color': sc_c1}, 'colors': {'default_color': 'Grey'}}
axis = ColorAxis(scale=sc_c1)
x = Map(map_data=topo_load('WorldMapData.json'), **map_styles)
fig = Figure(marks=[x], axes=[axis],title='Choropleth Example')
display(fig)
# -
# ## Alternate Map Data ##
sc_geo = AlbersUSA()
x = Map(map_data=topo_load('USMapData.json'),
scales={'projection': sc_geo})
fig = Figure(marks=[x], title='US States Map Example')
display(fig)
# ## Interactions ##
def_tt = Tooltip(fields=['id', 'name'])
map_mark = Map(scales={'projection': Mercator()}, tooltip=def_tt)
map_mark.interactions = {'click': 'select', 'hover': 'tooltip'}
fig = Figure(marks=[map_mark], title='Interactions Example')
display(fig)
| examples/Map.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_python3)
# language: python
# name: conda_python3
# ---
# # Bring Your Own R Algorithm
# _**Create a Docker container for training R algorithms and hosting R models**_
#
# ---
#
# ---
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Preparation](#Preparation)
# 1. [Code](#Code)
# 1. [Fit](#Fit)
# 1. [Serve](#Serve)
# 1. [Dockerfile](#Dockerfile)
# 1. [Publish](#Publish)
# 1. [Data](#Data)
# 1. [Train](#Train)
# 1. [Host](#Host)
# 1. [Predict](#Predict)
# 1. [Extensions](#Extensions)
#
# ---
# ## Background
#
# R is a popular open source statistical programming language, with a lengthy history in Data Science and Machine Learning. The breadth of algorithms available as an R package is impressive, which fuels a growing community of users. The R kernel can be installed into Amazon SageMaker Notebooks, and Docker containers which use R can be used to take advantage of Amazon SageMaker's flexible training and hosting functionality. This notebook illustrates a simple use case for creating an R container and then using it to train and host a model. In order to take advantage of boto, we'll use Python within the notebook, but this could be done 100% in R by invoking command line arguments.
#
# ---
# ## Preparation
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
# + isConfigCell=true
bucket = '<your_s3_bucket_name_here>'
prefix = 'sagemaker/r_byo'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
# -
# Now we'll import the libraries we'll need for the remainder of the notebook.
import time
import json
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# ### Permissions
#
# Running this notebook requires permissions in addition to the normal `SageMakerFullAccess` permissions. This is because we'll be creating a new repository in Amazon ECR. The easiest way to add these permissions is simply to add the managed policy `AmazonEC2ContainerRegistryFullAccess` to the role that you used to start your notebook instance. There's no need to restart your notebook instance when you do this, the new permissions will be available immediately.
#
# ---
# ## Code
#
# For this example, we'll need 3 supporting code files.
#
# ### Fit
#
# `mars.R` creates functions to fit and serve our model. The algorithm we've chosen to use is [Multivariate Adaptive Regression Splines](https://en.wikipedia.org/wiki/Multivariate_adaptive_regression_splines). This is a suitable example as it's a unique and powerful algorithm, but isn't as broadly used as Amazon SageMaker algorithms, and it isn't available in Python's scikit-learn library. R's repository of packages is filled with algorithms that share these same criteria.
# _The top of the code is devoted to setup. Bringing in the libraries we'll need and setting up the file paths as detailed in Amazon SageMaker documentation on bringing your own container._
#
# ```
# # Bring in library that contains multivariate adaptive regression splines (MARS)
# library(mda)
#
# # Bring in library that allows parsing of JSON training parameters
# library(jsonlite)
#
# # Bring in library for prediction server
# library(plumber)
#
#
# # Setup parameters
# # Container directories
# prefix <- '/opt/ml'
# input_path <- paste(prefix, 'input/data', sep='/')
# output_path <- paste(prefix, 'output', sep='/')
# model_path <- paste(prefix, 'model', sep='/')
# param_path <- paste(prefix, 'input/config/hyperparameters.json', sep='/')
#
# # Channel holding training data
# channel_name = 'train'
# training_path <- paste(input_path, channel_name, sep='/')
# ```
# _Next, we define a train function that actually fits the model to the data. For the most part this is idiomatic R, with a bit of maneuvering up front to take in parameters from a JSON file, and at the end to output a success indicator._
#
# ```
# # Setup training function
# train <- function() {
#
# # Read in hyperparameters
# training_params <- read_json(param_path)
#
# target <- training_params$target
#
# if (!is.null(training_params$degree)) {
# degree <- as.numeric(training_params$degree)}
# else {
# degree <- 2}
#
# # Bring in data
# training_files = list.files(path=training_path, full.names=TRUE)
# training_data = do.call(rbind, lapply(training_files, read.csv))
#
# # Convert to model matrix
# training_X <- model.matrix(~., training_data[, colnames(training_data) != target])
#
# # Save factor levels for scoring
# factor_levels <- lapply(training_data[, sapply(training_data, is.factor), drop=FALSE],
# function(x) {levels(x)})
#
# # Run multivariate adaptive regression splines algorithm
# model <- mars(x=training_X, y=training_data[, target], degree=degree)
#
# # Generate outputs
# mars_model <- model[!(names(model) %in% c('x', 'residuals', 'fitted.values'))]
# attributes(mars_model)$class <- 'mars'
# save(mars_model, factor_levels, file=paste(model_path, 'mars_model.RData', sep='/'))
# print(summary(mars_model))
#
# write.csv(model$fitted.values, paste(output_path, 'data/fitted_values.csv', sep='/'), row.names=FALSE)
# write('success', file=paste(output_path, 'success', sep='/'))}
# ```
# _Then, we setup the serving function (which is really just a short wrapper around our plumber.R file that we'll discuss [next](#Serve)._
#
# ```
# # Setup scoring function
# serve <- function() {
# app <- plumb(paste(prefix, 'plumber.R', sep='/'))
# app$run(host='0.0.0.0', port=8080)}
# ```
# _Finally, a bit of logic to determine if, based on the options passed when Amazon SageMaker Training or Hosting call this script, we are using the container to train an algorithm or host a model._
#
# ```
# # Run at start-up
# args <- commandArgs()
# if (any(grepl('train', args))) {
# train()}
# if (any(grepl('serve', args))) {
# serve()}
# ```
# ### Serve
# `plumber.R` uses the [plumber](https://www.rplumber.io/) package to create a lightweight HTTP server for processing requests in hosting. Note the specific syntax, and see the plumber help docs for additional detail on more specialized use cases.
# Per the Amazon SageMaker documentation, our service needs to accept post requests to ping and invocations. plumber specifies this with custom comments, followed by functions that take specific arguments.
#
# Here invocations does most of the work, ingesting our trained model, handling the HTTP request body, and producing a CSV output of predictions.
#
# ```
# # plumber.R
#
#
# #' Ping to show server is there
# #' @get /ping
# function() {
# return('')}
#
#
# #' Parse input and return the prediction from the model
# #' @param req The http request sent
# #' @post /invocations
# function(req) {
#
# # Setup locations
# prefix <- '/opt/ml'
# model_path <- paste(prefix, 'model', sep='/')
#
# # Bring in model file and factor levels
# load(paste(model_path, 'mars_model.RData', sep='/'))
#
# # Read in data
# conn <- textConnection(gsub('\\\\n', '\n', req$postBody))
# data <- read.csv(conn)
# close(conn)
#
# # Convert input to model matrix
# scoring_X <- model.matrix(~., data, xlev=factor_levels)
#
# # Return prediction
# return(paste(predict(mars_model, scoring_X, row.names=FALSE), collapse=','))}
# ```
# ### Dockerfile
#
# Smaller containers are preferred for Amazon SageMaker as they lead to faster spin up times in training and endpoint creation, so this container is kept minimal. It simply starts with Ubuntu, installs R, mda, and plumber libraries, then adds `mars.R` and `plumber.R`, and finally runs `mars.R` when the entrypoint is launched.
#
# ```Dockerfile
# FROM ubuntu:16.04
#
# MAINTAINER Amazon SageMaker Examples <<EMAIL>>
#
# RUN apt-get -y update && apt-get install -y --no-install-recommends \
# wget \
# r-base \
# r-base-dev \
# ca-certificates
#
# RUN R -e "install.packages(c('mda', 'plumber'), repos='https://cloud.r-project.org')"
#
# COPY mars.R /opt/ml/mars.R
# COPY plumber.R /opt/ml/plumber.R
#
# ENTRYPOINT ["/usr/bin/Rscript", "/opt/ml/mars.R", "--no-save"]
# ```
# ### Publish
# Now, to publish this container to ECR, we'll run the comands below.
#
# This command will take several minutes to run the first time.
# + language="sh"
#
# # The name of our algorithm
# algorithm_name=rmars
#
# #set -e # stop if anything fails
#
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
# region=${region:-us-west-2}
#
# fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
#
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login --region ${region} --no-include-email)
#
# # On a SageMaker Notebook Instance, the docker daemon may need to be restarted in order
# # to detect your network configuration correctly. (This is a known issue.)
# if [ -d "/home/ec2-user/SageMaker" ]; then
# sudo service docker restart
# fi
#
# # Build the docker image locally with the image name and then push it to ECR
# # with the full name.
# docker build -t ${algorithm_name} .
# docker tag ${algorithm_name} ${fullname}
#
# docker push ${fullname}
# -
# ---
# ## Data
# For this illustrative example, we'll simply use `iris`. This a classic, but small, dataset used to test supervised learning algorithms. Typically the goal is to predict one of three flower species based on various measurements of the flowers' attributes. Further detail can be found [here](https://en.wikipedia.org/wiki/Iris_flower_data_set).
#
# Then let's copy the data to S3.
train_file = 'iris.csv'
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train', train_file)).upload_file(train_file)
# _Note: Although we could, we'll avoid doing any preliminary transformations on the data, instead choosing to do those transformations inside the container. This is not typically the best practice for model efficiency, but provides some benefits in terms of flexibility._
# ---
# ## Train
#
# Now, let's setup the information needed to train a Multivariate Adaptive Regression Splines (MARS) model on iris data. In this case, we'll predict `Sepal.Length` rather than the more typical classification of `Species` to show how factors might be included in a model and limit the case to regression.
#
# First, we'll get our region and account information so that we can point to the ECR container we just created.
region = boto3.Session().region_name
account = boto3.client('sts').get_caller_identity().get('Account')
#
# - Specify the role to use
# - Give the training job a name
# - Point the algorithm to the container we created
# - Specify training instance resources (in this case our algorithm is only single-threaded so stick to 1 instance)
# - Point to the S3 location of our input data and the `train` channel expected by our algorithm
# - Point to the S3 location for output
# - Provide hyperparamters (keeping it simple)
# - Maximum run time
# +
r_job = 'r-byo-' + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print("Training job", r_job)
r_training_params = {
"RoleArn": role,
"TrainingJobName": r_job,
"AlgorithmSpecification": {
"TrainingImage": '{}.dkr.ecr.{}.amazonaws.com/rmars:latest'.format(account, region),
"TrainingInputMode": "File"
},
"ResourceConfig": {
"InstanceCount": 1,
"InstanceType": "ml.m4.xlarge",
"VolumeSizeInGB": 10
},
"InputDataConfig": [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": "s3://{}/{}/train".format(bucket, prefix),
"S3DataDistributionType": "FullyReplicated"
}
},
"CompressionType": "None",
"RecordWrapperType": "None"
}
],
"OutputDataConfig": {
"S3OutputPath": "s3://{}/{}/output".format(bucket, prefix)
},
"HyperParameters": {
"target": "Sepal.Length",
"degree": "2"
},
"StoppingCondition": {
"MaxRuntimeInSeconds": 60 * 60
}
}
# -
# Now let's kick off our training job on Amazon SageMaker Training, using the parameters we just created. Because training is managed (AWS takes care of spinning up and spinning down the hardware), we don't have to wait for our job to finish to continue, but for this case, let's setup a waiter so we can monitor the status of our training.
# +
# %%time
sm = boto3.client('sagemaker')
sm.create_training_job(**r_training_params)
status = sm.describe_training_job(TrainingJobName=r_job)['TrainingJobStatus']
print(status)
sm.get_waiter('training_job_completed_or_stopped').wait(TrainingJobName=r_job)
status = sm.describe_training_job(TrainingJobName=r_job)['TrainingJobStatus']
print("Training job ended with status: " + status)
if status == 'Failed':
message = sm.describe_training_job(TrainingJobName=r_job)['FailureReason']
print('Training failed with the following error: {}'.format(message))
raise Exception('Training job failed')
# -
# ---
# ## Host
#
# Hosting the model we just trained takes three steps in Amazon SageMaker. First, we define the model we want to host, pointing the service to the model artifact our training job just wrote to S3.
# +
r_hosting_container = {
'Image': '{}.dkr.ecr.{}.amazonaws.com/rmars:latest'.format(account, region),
'ModelDataUrl': sm.describe_training_job(TrainingJobName=r_job)['ModelArtifacts']['S3ModelArtifacts']
}
create_model_response = sm.create_model(
ModelName=r_job,
ExecutionRoleArn=role,
PrimaryContainer=r_hosting_container)
print(create_model_response['ModelArn'])
# -
# Next, let's create an endpoing configuration, passing in the model we just registered. In this case, we'll only use a few c4.xlarges.
# +
r_endpoint_config = 'r-endpoint-config-' + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print(r_endpoint_config)
create_endpoint_config_response = sm.create_endpoint_config(
EndpointConfigName=r_endpoint_config,
ProductionVariants=[{
'InstanceType': 'ml.m4.xlarge',
'InitialInstanceCount': 1,
'ModelName': r_job,
'VariantName': 'AllTraffic'}])
print("Endpoint Config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
# -
# Finally, we'll create the endpoints using our endpoint configuration from the last step.
# +
# %%time
r_endpoint = 'r-endpoint-' + time.strftime("%Y%m%d%H%M", time.gmtime())
print(r_endpoint)
create_endpoint_response = sm.create_endpoint(
EndpointName=r_endpoint,
EndpointConfigName=r_endpoint_config)
print(create_endpoint_response['EndpointArn'])
resp = sm.describe_endpoint(EndpointName=r_endpoint)
status = resp['EndpointStatus']
print("Status: " + status)
try:
sm.get_waiter('endpoint_in_service').wait(EndpointName=r_endpoint)
finally:
resp = sm.describe_endpoint(EndpointName=r_endpoint)
status = resp['EndpointStatus']
print("Arn: " + resp['EndpointArn'])
print("Status: " + status)
if status != 'InService':
raise Exception('Endpoint creation did not succeed')
# -
# ---
# ## Predict
# To confirm our endpoints are working properly, let's try to invoke the endpoint.
#
# _Note: The payload we're passing in the request is a CSV string with a header record, followed by multiple new lines. It also contains text columns, which the serving code converts to the set of indicator variables needed for our model predictions. Again, this is not a best practice for highly optimized code, however, it showcases the flexibility of bringing your own algorithm._
# +
iris = pd.read_csv('iris.csv')
runtime = boto3.Session().client('runtime.sagemaker')
payload = iris.drop(['Sepal.Length'], axis=1).to_csv(index=False)
response = runtime.invoke_endpoint(EndpointName=r_endpoint,
ContentType='text/csv',
Body=payload)
result = json.loads(response['Body'].read().decode())
result
# -
# We can see the result is a CSV of predictions for our target variable. Let's compare them to the actuals to see how our model did.
plt.scatter(iris['Sepal.Length'], np.fromstring(result[0], sep=','))
plt.show()
# ---
# ## Extensions
#
# This notebook showcases a straightforward example to train and host an R algorithm in Amazon SageMaker. As mentioned previously, this notebook could also be written in R. We could even train the algorithm entirely within a notebook and then simply use the serving portion of the container to host our model.
#
# Other extensions could include setting up the R algorithm to train in parallel. Although R is not the easiest language to build distributed applications on top of, this is possible. In addition, running multiple versions of training simultaneously would allow for parallelized grid (or random) search for optimal hyperparamter settings. This would more fully realize the benefits of managed training.
# ### (Optional) Clean-up
#
# If you're ready to be done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
sm.delete_endpoint(EndpointName=r_endpoint)
| advanced_functionality/r_bring_your_own/r_bring_your_own.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PySpark
# language: python
# name: pyspark
# ---
# # Yelp: Business Analysis
# !spark-shell --version
# <details>
# <summary><b>Table of Contents</b> (click to open)</summary>
# <!-- MarkdownTOC -->
#
# 1. [Top Businesses](#topbs)
# 1. [Impact of reviews on Businesses ](#revimpact)
# 1. [Top Restaurants](#toprest)
# 1. [Categories by Businesses](#catbs)
# 1. [Vegetarian Restuarants](#vegres)
# 1. [Top Category/Cuisine](#cuisine)
# <!-- /MarkdownTOC -->
# </details>
# ### Import Libraries
# +
import seaborn as sns
import folium
import numpy as np
import matplotlib.pyplot as plt
import pyspark.sql.functions as F
from pyspark.sql.functions import col
# +
font_size = 16
params = {"axes.titlesize": font_size,
"axes.labelsize": font_size*.7,
"xtick.labelsize": font_size*.5,
"ytick.labelsize": font_size*.5,
"legend.fontsize": font_size*.5,
"figure.figsize": (20, 10),
"axes.titlepad": 25}
plt.style.use('seaborn-darkgrid')
sns.set_context("notebook", rc=params)
# -
# ## Load Data
# Load the review data from google cloud storage to dataproc
file_review = "gs://proj-spark/yelp-dataset/yelp_academic_dataset_review.json"
file_business = "gs://proj-spark/yelp-dataset/yelp_academic_dataset_business.json"
file_users = "gs://proj-spark/yelp-dataset/yelp_academic_dataset_user.json"
file_checkin = "gs://proj-spark/yelp-dataset/yelp_academic_dataset_checkin.json"
file_tip = "gs://proj-spark/yelp-dataset/yelp_academic_dataset_tip.json"
# +
df_review = spark.read.format('json').load(file_review)
df_review.createOrReplaceTempView('reviewTable')
df_business = spark.read.format('json').load(file_business)
df_business.createOrReplaceTempView('businessTable')
df_users = spark.read.format('json').load(file_users)
df_users.createOrReplaceTempView('usersTable')
df_checkin = spark.read.format('json').load(file_checkin)
df_checkin.createOrReplaceTempView('checkinTable')
df_tip = spark.read.format('json').load(file_tip)
df_tip.createOrReplaceTempView('tipTable')
# -
# ## Top Businesses <a class="anchor" id="topbs"></a>
# +
# checkin count for each business
df_checkin_cnt = df_checkin.withColumn('checkin_count', F.size(F.split(col('date'),', ')))
# join the checkin info with businesses data
joinExpression = df_business['business_id'] == df_checkin_cnt['business_id']
df_bs_checkin = df_business.join(df_checkin_cnt, joinExpression)\
.drop(df_checkin_cnt['business_id'])
# -
top = 5
df_top_bs = df_bs_checkin.orderBy(F.desc('checkin_count')).limit(top)
# top 5 businesses on yelp
df_top_bs.select('business_id', 'name', 'city', 'state', 'stars', 'checkin_count', 'review_count').toPandas()
# ## Impact of reviews on Businesses <a class="anchor" id="revimpact"></a>
# Impact of reviews count on Business checkin in Oregon (State)
fig, ax = plt.subplots(figsize=(10, 5))
data = df_bs_checkin.filter("state = 'OR'").toPandas()
ax = sns.regplot(x='review_count', y='checkin_count', data=data, ax=ax,
marker="+",
scatter_kws={"s": 50})
ax.set_title('Impact of reviews on Business in Oregon')
# We can observed that business having high review count more likely to have high checkin count.
# ## Top Restaurants <a class="anchor" id="toprest"></a>
# Based on number of checkin find the top restaurants business in Oregon
top = 10
df_top_res = df_bs_checkin.filter(col('categories').contains('Res'))\
.filter("state = 'OR'")\
.orderBy(F.desc('checkin_count'))\
.limit(top)
fig, ax = plt.subplots(figsize=(10,5))
data = df_top_res.toPandas()
ax = sns.barplot(x='checkin_count', y='name', hue='stars', data=data, ax=ax,
dodge=False,
saturation=.7)
ax.set_title('Top Restaurants in Oregon')
# ## Categories by Businesses <a class="anchor" id="catbs"></a>
# Total number of businesses per category
# +
# Split the categories
df_business = df_business.withColumn('categories_splitted', F.split(col('categories'),', '))\
.withColumn('category', F.explode('categories_splitted'))
cat_grp = df_business.groupBy('category')\
.agg(F.count('business_id').alias('business_count'))\
.orderBy(F.desc('business_count'))
# -
#cat_grp.count()
cat_grp.show()
# Plot bar chart for top 15 categories
fig, ax = plt.subplots(figsize=(10, 5))
ax = sns.barplot(x='business_count', y='category', data=cat_grp.limit(15).toPandas(), ax=ax)
ax.set_title('Categories by Businesses')
# ## Vegetarian Restuarants <a class="anchor" id="vegres"></a>
# Yelp Business dataset is the collection of different business in different areas. Restaurants is most popular among them. Let's analyze the Vegetarian Restuarants.
# Filter Vegetarian Restaurants
df_veg_restuarant = df_business.withColumn('categories_splitted', F.split(col('categories'),', '))\
.filter(F.array_contains(col('categories_splitted'), 'Restaurants'))\
.filter(F.array_contains(col('categories_splitted'), 'Vegetarian'))
# Number of Vegetarian restaurants in each city
df_rest_city = df_veg_restuarant.groupBy('city').count().orderBy(F.desc('count'))
# Top-10 Cities for Vegetarian Restaurants
# Plot top-10 Cities for Vegetarian Restaurants
top = 10
data = df_rest_city.limit(top).toPandas()
fig, ax = plt.subplots(figsize=(10,5))
palette = np.array(sns.color_palette('Greens_d', top))[::-1]
ax = sns.barplot(x='count', y='city', data=data, palette=palette)
ax.set_title('Top Cities for Vegetarian Restaurants')
# Top Vegetarian Restaurants in Portland city which are open
df_portland = df_veg_restuarant.filter("city='Portland'")\
.filter("is_open=1")\
.withColumn('OutdoorSeating', col('attributes.OutdoorSeating'))\
.orderBy(F.desc('stars'))
# Plot top-10 Vegetarian Restaurants in Portland
top = 20
data = df_portland.limit(top).toPandas()
facetgrid = sns.catplot(x='review_count', y='name',
hue='stars',
col='OutdoorSeating',
kind='bar',
data=data)
facetgrid.fig.subplots_adjust(top=0.8)
facetgrid.fig.suptitle('Top Vegetarian Restaurants in Portland')
# +
#Geographical Map of Vegetarian restuarant in Portland
portland_map = folium.Map(location=[45.5051,-122.6750], tiles='OpenStreetMap', zoom_start=11)
for idx, row in data.iterrows():
text = f"""
<header>
<h6><b>Name: {row['name']}</b></h6>
</header>
<hr style='margin:10px;'>
<ul style='color: #444;list-style-type:circle;align-item:left;padding-left:20px'>
<li> Stars: {row['stars']} </li>
<li> Review Count: {row['review_count']} </li>
<li> OutdoorSeating: {row['OutdoorSeating']} </li>
</ul>
"""
popup = folium.Popup(folium.Html(text,script=True), max_width=2650)
folium.Marker([row['latitude'], row['longitude'] ], popup=popup, tooltip="Click!")\
.add_to(portland_map)
# -
portland_map
# ## Top Category/Cuisine <a class="anchor" id="cuisine"></a>
# +
def filter_business(city, category):
"""Filter business based on city and category"""
def _(df):
return df.filter(col('city')==city)\
.withColumn('categories_splitted', F.split(col('categories'),', '))\
.filter(F.array_contains(col('categories_splitted'), category))\
.withColumn('category', F.explode('categories_splitted'))
return _
def ntop_category(n:int=10):
"""Top n category"""
return lambda df: (
df.groupBy('category')\
.count()\
.orderBy(F.desc('count'))\
.limit(n)
)
# -
def ntop_cuisine(n, city):
""" """
def _(df):
return df.transform(filter_business(city, 'Restaurants'))\
.transform(ntop_category())\
.filter("category != 'Restaurants'")\
.filter("category != 'Food'")
return _
# +
# Top Category/Cuisine in Portland, Austin, Atlanta, Boston
top = 10
cities = ['Portland', 'Austin', 'Atlanta', 'Boston']
data = {}
for city in cities:
data[city] = df_business.transform(ntop_cuisine(top, city)).toPandas()
# +
# figure and grid for ploting
fig = plt.figure(figsize=(16, 8), constrained_layout=True)
grid = plt.GridSpec(2, 2, wspace=0.4, hspace=0.3, figure=fig)
fig.suptitle('Top Categories/Cuisine for Restaurants in City', fontsize=18)
palette = np.array(sns.color_palette("Blues_d", top))[::-1]
# Top categories for Portland Restaurants
portland_cat = data['Portland']
ax = plt.subplot(grid[0, 0])
ax = sns.barplot(x='count', y='category', data=portland_cat, ax=ax, palette=palette)
ax.set_title('Top categories for Portland Restaurants')
# Top categories for Austin Restaurants
austin_cat = data['Austin']
ax = plt.subplot(grid[0, 1])
ax = sns.barplot(x='count', y='category', data=austin_cat, ax=ax, palette=palette)
ax.set_title('Top categories for Austin Restaurants')
# Top categories for Austin Restaurants
alanta_cat = data['Atlanta']
ax = plt.subplot(grid[1, 0])
ax = sns.barplot(x='count', y='category', data=alanta_cat, ax=ax, palette=palette)
ax.set_title('Top categories for Atlanta Restaurants')
# Top categories for Boston Restaurants
boston_cat = data['Boston']
ax = plt.subplot(grid[1, 1])
ax = sns.barplot(x='count', y='category', data=boston_cat, ax=ax, palette=palette)
ax.set_title('Top categories for Boston Restaurants')
# -
| notebooks/business_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-Zl2SU5CqJsJ"
# <!--COURSE_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="https://user-images.githubusercontent.com/16768318/73986808-75b3ca00-4936-11ea-90f1-3a6c352766ce.png" width=10% >
# <img align="right" style="padding-left:10px;" src="https://user-images.githubusercontent.com/16768318/73986811-764c6080-4936-11ea-9653-a3eacc47caed.png" width=10% >
#
# **Bienvenidos!** Este *colab notebook* es parte del curso [**Introduccion a Google Earth Engine con Python**](https://github.com/csaybar/EarthEngineMasterGIS) desarrollado por el equipo [**MasterGIS**](https://www.mastergis.com/). Obten mas informacion del curso en este [**enlace**](https://www.mastergis.com/product/google-earth-engine/). El contenido del curso esta disponible en [**GitHub**](https://github.com/csaybar/EarthEngineMasterGIS) bajo licencia [**MIT**](https://opensource.org/licenses/MIT).
# + [markdown] id="4HfdNOuz37oi"
# ### **Ejercicio N°01: RUSLE a Nivel Mundial**
# <img src="https://user-images.githubusercontent.com/16768318/73690808-1604b700-46c9-11ea-8bdd-43e0e490a0a3.gif" align="right" width = 60%/>
#
# Genere una funcion para calcular la Ecuacion Universal de Perdida de Suelo (RUSLE) para cualquier parte del mundo. La funcion debe tener los siguientes parametros.**rusle(roi, prefix, folder, scale)**
#
# http://cybertesis.unmsm.edu.pe/handle/cybertesis/10078
#
# + id="tGfUCO7Vgv2Q"
#@title Credenciales Google Earth Engine
import os
credential = '{"refresh_token":"<PASSWORD>"}'
credential_file_path = os.path.expanduser("~/.config/earthengine/")
os.makedirs(credential_file_path,exist_ok=True)
with open(credential_file_path + 'credentials', 'w') as file:
file.write(credential)
# + id="bQZNW-c74xba" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1617022409487, "user_tz": 300, "elapsed": 31704, "user": {"displayName": "<NAME>\u00e1rdenas", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="79484139-b339-4ee2-8ca0-b4ad55a29126"
import ee
ee.Authenticate()
ee.Initialize()
# + id="EiCTu4NB4yAk" cellView="form"
#@title mapdisplay: Crea mapas interactivos usando folium
import folium
def mapdisplay(center, dicc, Tiles="OpensTreetMap",zoom_start=10):
'''
:param center: Center of the map (Latitude and Longitude).
:param dicc: Earth Engine Geometries or Tiles dictionary
:param Tiles: Mapbox Bright,Mapbox Control Room,Stamen Terrain,Stamen Toner,stamenwatercolor,cartodbpositron.
:zoom_start: Initial zoom level for the map.
:return: A folium.Map object.
'''
center = center[::-1]
mapViz = folium.Map(location=center,tiles=Tiles, zoom_start=zoom_start)
for k,v in dicc.items():
if ee.image.Image in [type(x) for x in v.values()]:
folium.TileLayer(
tiles = v["tile_fetcher"].url_format,
attr = 'Google Earth Engine',
overlay =True,
name = k
).add_to(mapViz)
else:
folium.GeoJson(
data = v,
name = k
).add_to(mapViz)
mapViz.add_child(folium.LayerControl())
return mapViz
# + [markdown] id="PKlMu7elDd-I"
# ### **1) Factor R**
#
# El **factor R** es el factor de erosividad de la lluvia. Este factor indica el potencial erosivo de la lluvia que afecta en el proceso de erosion del suelo. Haciendo una analogia, se podria decir que una lluvia fuerte un dia al año puede producir suficiente energia para erosionar el suelo que varias lluvias de mediana intensidad a lo largo de un ano.
#
# El factor erosividad (R) es definido como la sumatoria anual de los promedios de los valores individuales del indice de tormenta de erosion (EI30). Donde E es la energia cinetica por unidad de area e I30 es la maxima intensidad en 30 minutos de precipitacion. Esto se puede definir en la siguiente ecuacion:
#
# <img src="https://user-images.githubusercontent.com/16768318/73694650-67fd0b00-46d0-11ea-87f6-4ed9501cf964.png" width = 60%>
#
# Por tanto, la energia de la tormenta (EI o R) indica el volumen de lluvia y escurrimiento, pero una larga y suave lluvia puede tener el mismo valor de E que una lluvia de corta y mas alta intensidad. (Mannaerts, 1999). La energia se calcula a partir de la formula de Brown y Foster:
#
# <img src="https://user-images.githubusercontent.com/16768318/73694782-b3171e00-46d0-11ea-94fe-94f3f57941c5.png" width = 40%>
#
# A partir de la ecuación anterior, el calculo del factor R es un proceso complejo y requiere datos horarios o diarios de varios anos. Por lo que se han desarrollado diferentes ecuaciones que adaptan la erosividad local mediante una formula que solo requiera una data mensual o anual de precipitacion. A continuacion, se muestran algunas de las formulas adaptadas para una precipitacion media anual.
#
# <img src="https://user-images.githubusercontent.com/16768318/73694993-228d0d80-46d1-11ea-8bc4-9962963850b7.png">
#
#
# Si bien es cierto, se usa ampliamente una precipitacion media anual para estimar el **factor R** debido a la escasez de informacion, para este ejemplo se ha optado por utilizar la formula desarrollada por **(Wischmeier & Smith, 1978)** debido a que se cuenta con una serie historica de informacion de precipitacion mensual. La formula es:
#
#
# <img src="https://user-images.githubusercontent.com/16768318/73695488-2b321380-46d2-11ea-8033-0063f27698d8.png" width = 50%>
# + id="eImkby9KHNuf"
# Monthly precipitation in mm at 1 km resolution:
# https://zenodo.org/record/3256275#.XjibuDJKiM8
clim_rainmap = ee.Image("OpenLandMap/CLM/CLM_PRECIPITATION_SM2RAIN_M/v01")
year = clim_rainmap.reduce(ee.Reducer.sum())
R_monthly = ee.Image(10).pow(ee.Image(1.5).multiply(clim_rainmap.pow(2).divide(year).log10().subtract(-0.08188))).multiply(1.735)
factorR = R_monthly.reduce(ee.Reducer.sum())
# + id="erho-vrbLtYi" colab={"base_uri": "https://localhost:8080/", "height": 661} executionInfo={"status": "ok", "timestamp": 1617022424165, "user_tz": 300, "elapsed": 1684, "user": {"displayName": "<NAME>\u00e1rdenas", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="7f0a3f87-1166-4ff1-baef-37a36398e71b"
center_coordinate = [0,0]
palette_rain = ["#450155", "#3B528C", "#21918D", "#5DCA63","#FFE925"]
mapdisplay(center_coordinate, {'Factor_R':factorR.getMapId({'min':0,'max':6000,'palette':palette_rain})},zoom_start=3)
# + [markdown] id="TtGuoiN15n0R"
# ### **2) Factor K**
#
# A diferencia del factor R, el factor K muestra qué tan susceptible es el suelo a la erosion hidrica, esto es determinado por las propiedades fisicas y quimicas del suelo, que dependen de las caracteristicas de estos. Para determinar el factor K, existen una gran cantidad de formulas empiricas, adecuadas para diversos lugares del mundo y donde intervienen caracteristicas del suelo como porcentaje de arena, limo, arcilla; estructura del suelo; contenido de carbono organico o materia orgánica; entre otros.
#
# El factor K puede variar en una escala de 0 a 1, donde 0 indica suelos con la menor susceptibilidad a la erosion y 1 indica suelos altamente susceptibles a la erosion hidrica del suelo; cabe mencionar que esta escala fue hecha para el sistema de unidades americanas, y adaptandose al sistema internacional, la escala varia a normalmente entre 0 y 0.07.
#
# A continuacion, se muestran algunas ecuaciones para la estimación de este factor:
#
# <img src="https://user-images.githubusercontent.com/16768318/73704444-039b7500-46eb-11ea-9ccd-b7850bb17911.png" width = 50%>
# <img src="https://user-images.githubusercontent.com/16768318/73704442-039b7500-46eb-11ea-870c-a557ca50b777.png" width = 50%>
# <img src="https://user-images.githubusercontent.com/16768318/73704443-039b7500-46eb-11ea-9469-104f04983dfd.png" width = 50%>
#
#
# Para este ejemplo se ha optado por utilizar la formula desarrollada por **Williams (1975)**.
#
#
# + id="c2f4KFJY5rwQ"
# Cargamos toda la informacion necesaria para estimar el factor K
sand = ee.Image("OpenLandMap/SOL/SOL_CLAY-WFRACTION_USDA-3A1A1A_M/v02").select('b0')
silt = ee.Image('users/aschwantes/SLTPPT_I').divide(100)
clay = ee.Image("OpenLandMap/SOL/SOL_SAND-WFRACTION_USDA-3A1A1A_M/v02").select('b0')
morg = ee.Image("OpenLandMap/SOL/SOL_ORGANIC-CARBON_USDA-6A1C_M/v02").select('b0').multiply(0.58)
sn1 = sand.expression('1 - b0 / 100', {'b0': sand})
orgcar = ee.Image("OpenLandMap/SOL/SOL_ORGANIC-CARBON_USDA-6A1C_M/v02").select('b0')
# + id="uRHRfFcO3Q1O"
#Juntando todas las imagenes en una sola
soil = ee.Image([sand, silt, clay, morg, sn1, orgcar]).rename(['sand', 'silt', 'clay', 'morg', 'sn1', 'orgcar'] )
# + id="lB1wV7ECyKFh"
factorK = soil.expression(
'(0.2 + 0.3 * exp(-0.0256 * SAND * (1 - (SILT / 100)))) * (1 - (0.25 * CLAY / (CLAY + exp(3.72 - 2.95 * CLAY)))) * (1 - (0.7 * SN1 / (SN1 + exp(-5.51 + 22.9 * SN1))))',
{
'SAND': soil.select('sand'),
'SILT': soil.select('silt'),
'CLAY': soil.select('clay'),
'MORG': soil.select('morg'),
'SN1': soil.select('sn1'),
'CORG': soil.select('orgcar')
});
# + id="RzP4Ta2h7c8_" colab={"base_uri": "https://localhost:8080/", "height": 661} executionInfo={"status": "ok", "timestamp": 1617022438686, "user_tz": 300, "elapsed": 872, "user": {"displayName": "<NAME>\u00e1rdenas", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="db1a8047-75b3-421a-da95-aa0443dd47b8"
center_coordinate = [0,0]
palette_k = palette = [
'FFFFFF', 'CE7E45', 'DF923D', 'F1B555', 'FCD163', '99B718', '74A901',
'66A000', '529400', '3E8601', '207401', '056201', '004C00', '023B01',
'012E01', '011D01', '011301'
]
viz_param_k = {'min': 0.0, 'max': 0.5, 'palette': palette_k};
mapdisplay(center_coordinate, {'Factor_K':factorK.getMapId(viz_param_k)},zoom_start=3)
# + [markdown] id="Kknq-v_33f68"
# ### **3) Factor LS**
#
# El factor LS expresa el efecto de la topografia local sobre la tasa de erosion del suelo, combinando los efectos de la longitud de la pendiente (L) y la inclinación de la pendiente (S). A medida que mayor sea la longitud de la pendiente, mayor sera la cantidad de escorrentia acumulada y de la misma forma, mientras mas pronunciada sea la pendiente de la superficie, mayor sera la velocidad de la escorrentia, que influye directamente en la erosion. Existen diversas metodologias basadas en SIG para calcular estos factores, como se pueden mostrar a continuación:
#
# <img src="https://user-images.githubusercontent.com/16768318/73706484-7ce99680-46f0-11ea-8e0e-5fbb4a00731d.png" width = 50%>
# + id="Gtu7sQ2h9bur"
facc = ee.Image("WWF/HydroSHEDS/15ACC")
dem = ee.Image("WWF/HydroSHEDS/03CONDEM")
slope = ee.Terrain.slope(dem)
ls_factors = ee.Image([facc, slope]).rename(['facc','slope'])
# + id="Yv4kdbmZFcHf"
factorLS = ls_factors.expression(
'(FACC*270/22.13)**0.4*(SLOPE/0.0896)**1.3',
{
'FACC': ls_factors.select('facc'),
'SLOPE': ls_factors.select('slope')
});
# + id="2c98Zn_1GaOn" colab={"base_uri": "https://localhost:8080/", "height": 661} executionInfo={"status": "ok", "timestamp": 1617022448036, "user_tz": 300, "elapsed": 600, "user": {"displayName": "<NAME>00e1rdenas", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="145befee-e0f3-4d1f-b241-d1d7b65a727a"
center_coordinate = [0,0]
palette_ls = palette = [
'FFFFFF', 'CE7E45', 'DF923D', 'F1B555', 'FCD163', '99B718', '74A901',
'66A000', '529400', '3E8601', '207401', '056201', '004C00', '023B01',
'012E01', '011D01', '011301'
]
viz_param_k = {'min': 0, 'max': 100, 'palette': palette_ls};
mapdisplay(center_coordinate, {'Factor_LS':factorLS.getMapId(viz_param_k)},zoom_start=3)
# + [markdown] id="HV4zlFetUH0I"
# ### **4) Factor C**
#
# El factor C se utiliza para determinar la eficacia relativa de los sistemas de manejo del suelo y de los cultivos en terminos de prevencion o reduccion de la perdida de suelo. Este factor indica como la cobertura vegetal y los cultivos afectaran la perdida media anual de suelos y como se distribuira el potencial de perdida de suelos en el tiempo (Rahaman, 2015).
#
# El valor de C depende del tipo de vegetacion, la etapa de crecimiento y el porcentaje de cobertura. Valores mas altos del factor C indican que no hay efecto de cobertura y perdida de suelo, mientras que el menor valor de C significa un efecto de cobertura muy fuerte que no produce erosion.
#
#
# + id="ZdTMn78VUj7b"
ndvi_median = ee.ImageCollection("MODIS/006/MOD13A2").median().multiply(0.0001).select('NDVI')
# + id="CEgvwdUzVNV0" colab={"base_uri": "https://localhost:8080/", "height": 661} executionInfo={"status": "ok", "timestamp": 1617022456107, "user_tz": 300, "elapsed": 1134, "user": {"displayName": "<NAME>\u00e1rdenas", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="ce533ac8-6c0a-415e-eeee-a7e8d7d58c9a"
geo_ndvi = [
'FFFFFF', 'CE7E45', 'DF923D', 'F1B555', 'FCD163', '99B718', '74A901',
'66A000', '529400', '3E8601', '207401', '056201', '004C00', '023B01',
'012E01', '011D01', '011301'
]
l8_viz_params = {'palette':geo_ndvi,'min':0,'max': 0.8}
mapdisplay([0,0],{'composite_median':ndvi_median.getMapId(l8_viz_params)},zoom_start=3)
# + [markdown] id="OcMByLHSYSae"
# Otra forma de hallar este factor C, es haciendo una comparación entre el NDVI a partir de las fórmulas <NAME> (1999) [C1] y su adaptacion para paises asiaticos, que tambien se adecuan a la realidad de la costa peruana de Lin (2002) [C2]. Por ultimo se tiene la ecuacion formulada por De Jong(1994) [C3] adaptado a estudios de degradacion de suelos en un entorno mediterraneo.
# <center>
# <img src="https://user-images.githubusercontent.com/16768318/73713048-e6bf6b80-4703-11ea-80b1-1940e6b55707.png" width = 50%>
# </center>
# + id="bPj-eXMNZLfG"
factorC = ee.Image(0.805).multiply(ndvi_median).multiply(-1).add(0.431)
# + [markdown] id="Bp5B172zZJLy"
# ### **5) Calculo de la Erosion**
#
# **A = R\*K\*LS\*C\*1**
#
# <img src="https://user-images.githubusercontent.com/16768318/73690808-1604b700-46c9-11ea-8bdd-43e0e490a0a3.gif">
# + id="YDsavOPYYMWx"
erosion = factorC.multiply(factorR).multiply(factorLS).multiply(factorK)
# + id="iWXlpv_bbHRw" colab={"base_uri": "https://localhost:8080/", "height": 661} executionInfo={"status": "ok", "timestamp": 1617022484829, "user_tz": 300, "elapsed": 736, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="eb0c2ff9-2db1-4ea6-926f-da1a4e01aa4b"
geo_erosion = ["#00BFBF", "#00FF00", "#FFFF00", "#FF7F00", "#BF7F3F", "#141414"]
l8_viz_params = {'palette':geo_erosion,'min':0,'max': 6000}
mapdisplay([0,0],{'composite_median':erosion.getMapId(l8_viz_params)},zoom_start=3)
# + [markdown] id="uDrzlwocdUvp"
# ## **Ejercicio Propuesto 1**
#
# ### **Funcion para descargar RUSLE en cualquier parte del mundo**
# [Respuesta aqui](https://gist.github.com/csaybar/19a9db35f8c8044448d885b68e8c9eb8)
#
#
# #### Estimar la Erosión multianual para cualquier parte del mundo. El resultado final puede ser un vídeo ([ejemplo](https://drive.google.com/file/d/1u14mx7GfBj0TmDi6xTCk_vdxT0fBQN_B/view)) o enviar el código para revisarlo en formato .py o .txt.
# + id="TlnkZmAfdXSm" colab={"base_uri": "https://localhost:8080/", "height": 661} executionInfo={"status": "ok", "timestamp": 1617029624813, "user_tz": 300, "elapsed": 2712, "user": {"displayName": "<NAME>\u00e1rdenas", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GgyZb6d8eCK0c821mJyLUcgm5-sLGDNHqYpUgni=s64", "userId": "15908768188578917667"}} outputId="0d2905c9-d23b-4460-bcfa-aa0d81bc5962"
#### Ponga su funcion aqui (cree un snippet!)
# + id="2-Ol7GrqzPiB"
# Ambito de estudio aqui
geometry = ee.Geometry.Polygon([[[-81.9580078125,-5.659718554577273],
[-74.99267578125,-5.659718554577273],
[-74.99267578125,2.04302395742204],
[-81.9580078125,2.04302395742204],
[-81.9580078125,-5.659718554577273]]])
ec_erosion = rusle(geometry,'RUSLE_','RUSLE_MASTERGIS', scale = 100)
# + id="xfo5sr0U9aOj"
# Genere una vizualizacion de su ambito de estudio
geo_erosion = ["#00BFBF", "#00FF00", "#FFFF00", "#FF7F00", "#BF7F3F", "#141414"]
l8_viz_params = {'palette':geo_erosion,'min':0,'max': 6000}
center = geometry.centroid().coordinates().getInfo()
mapdisplay(center,{'composite_median':ec_erosion.select('A').getMapId(l8_viz_params)},zoom_start=6)
# + [markdown] id="v41ueZ40qCYa"
# ### **¿Dudas con este Jupyer-Notebook?**
#
# Estaremos felices de ayudarte!. Create una cuenta Github si es que no la tienes, luego detalla tu problema ampliamente en: https://github.com/csaybar/EarthEngineMasterGIS/issues
#
# **Tienes que dar clic en el boton verde!**
#
# <center>
# <img src="https://user-images.githubusercontent.com/16768318/79680748-d5511000-81d8-11ea-9f89-44bd010adf69.png" width = 70%>
# </center>
| module06/04_RUSLE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial 0: Overview of built-in examples
# +
# Standard imports
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# Insert path to mavenn beginning of path
import os
import sys
abs_path_to_mavenn = os.path.abspath('../../')
sys.path.insert(0, abs_path_to_mavenn)
# Load mavenn
import mavenn
# -
# MAVE-NN comes with built-in demos, tutorials, models, and datasets. Tutorials are provided in the form of Jupyter notebooks (like this one). To get a list of tutorials notebooks, as well as their locations on your local machine, import `mavenn` and execute the following:
mavenn.list_tutorials()
# A list of example datasets included within mavenn can be seen by execute the following:
mavenn.load_example_dataset()
# These string values represent different multiplex assays (see references) and entering any of these names into the `load_example_dataset` function returns a pandas dataframe containing sequences and their associated measurements. For example, using 'mpsa' as the input string loads the massively parallel splicing assay (MPSA) dataset reported by Wong et al., 2018.
mpsa_df = mavenn.load_example_dataset(name='mpsa')
mpsa_df.head()
# A value of `True` in the column `training_set` indicates rows that were used to train MAVE-NN models. All other rows indicate held-out test data.
#
# This mpsa dataframe show above consists of continuous measurement values $y$ and is appropriate for use with global epistasis regression in MAVE-NN. The values in the column d$y$ indicate the SEM values. Using 'name=gb1' as the string value in `load_example_dataset` yields a similar dataframe. Using 'sortseq' as the dataset name yields a dataset appropiate for measurement process agnostic (MPA) regression in MAVE-NN:
sortseq_df = mavenn.load_example_dataset('sortseq')
sortseq_df.head()
# In this dataframe, ct_i indicates the counts in bin i, where i indexes the bins.
#
# The `load_example_model` method allows for pretrained MAVE-NN models to be loaded and subsequently used on the above example datasets. To see a list of models that be loaded with MAVE-NN, execute the following:
mavenn.load_example_model()
# We now load the mpsa pairwise example model and use it in conjunction with the mpsa dataset loaded above.
mpsa_pairwise = mavenn.load_example_model(name='mpsa_ge_pairwise')
mpsa_pairwise.regression_type
# The `mpsa_pairwise` variable contain the MAVE-NN model objects, from which a number of helpful attributes and functions can be accessed. E.g., the function `x_to_yhat()` can be used to make predictions on input sequences. We first extract test sequences to evaluate various MAVE-NN model methods.
# +
# indices of training examples
i_training = mpsa_df['set']=='training'
# get test examples.
mpsa_test_df = mpsa_df[~i_training]
print('Test mpsa values:')
mpsa_test_df.head()
# +
x = mpsa_test_df['x'].values
y = mpsa_test_df['y'].values
mpsa_pairwise.set_data(x=mpsa_test_df['x'],
y=mpsa_test_df['y'])
fig, ax = plt.subplots(figsize=(4,4))
ax.scatter(mpsa_pairwise.x_to_yhat(x),y,s=5)
ax.set_title(mpsa_pairwise.regression_type+' '+mpsa_pairwise.gpmap_type)
ax.set_xlabel('predictions ($\hat{y}$)')
plt.show()
# -
# The function `x_to_phi` can be used to obtain the latent phenotype $\phi$ from sequences. Additionally, we can evaluate the function on a grid `phi_to_yhat` and plot it against $\phi$.
# +
phi_pairwise = mpsa_pairwise.x_to_phi(x)
phi_pairwise_lim = [min(phi_pairwise)-.5, max(phi_pairwise)+.5]
phi_pairwise_grid = np.linspace(phi_pairwise_lim[0], phi_pairwise_lim[1], 1000)
# Compute yhat each phi gridpoint
yhat_pairwise_grid = mpsa_pairwise.phi_to_yhat(phi_pairwise_grid)
# +
fig, ax = plt.subplots(figsize=(4,4))
ax.scatter(phi_pairwise,y,s=5,alpha=0.3)
ax.plot(phi_pairwise_grid,yhat_pairwise_grid,color='orange',lw=3.2,label='g($\phi$)')
ax.set_title(mpsa_pairwise.regression_type+' '+mpsa_pairwise.gpmap_type)
ax.set_ylabel('log PSI (y)')
ax.set_xlabel('latent phenotype ($\phi$)')
ax.legend()
plt.show()
# -
# Demos, on the other, hand are self-contained Python scripts that can be executed by calling `mavenn.run_demo()`. To get a list of demo names, execute:
mavenn.run_demo()
# The `"gb1_ge_evaluation"` demo illustrates an additive G-P map, along with a heteroskedastic Gaussian GE measurement process, fit to data from a deep mutational scanning (DMS) experiment on the protein GB1 performed by Olson et al., 2014. To see the code for this demo, set `print_code=True`.
mavenn.run_demo("gb1_ge_evaluation", print_code=False)
# The `"sortseq_mpa_visualization"` demo illustrates an additive G-P map, along with an MPA measurement process, fit to data from a sort-seq massively parallel reporter assay (MPRA) performed by Kinney et al., 2010.
mavenn.run_demo("sortseq_mpa_visualization", print_code=False)
# The `"mpsa_ge_training"` demo trains a pairwise G-P map, along with a homoskedastic Gaussian measurement process, on data from a massively parallel splicing assay (MPSA) reported by Wong et al., 2018. This training takes ~20 seconds on a standard laptop.
mavenn.run_demo("mpsa_ge_training", print_code=False)
# **References**
#
# 1. <NAME>., <NAME>., <NAME>. (2014). A comprehensive biophysical description of pairwise epistasis throughout an entire protein domain. Current biology : CB 24(22), 2643 - 2651. https://dx.doi.org/10.1016/j.cub.2014.09.072
#
# 1. <NAME>., <NAME>., <NAME>., <NAME>. (2010). Using deep sequencing to characterize the biophysical mechanism of a transcriptional regulatory sequence Proceedings of the National Academy of Sciences 107(20), 9158-9163. https://dx.doi.org/10.1073/pnas.1004290107
#
# 1. <NAME>., <NAME>., <NAME>. (2018). Quantitative Activity Profile and Context Dependence of All Human 5' Splice Sites. Molecular cell 71(6), 1012-1026.e3. https://dx.doi.org/10.1016/j.molcel.2018.07.033
| docs/tutorials/tutorial_0_built-in_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="S5DrIaIbDHKv"
# # Name:- <NAME>
# # Experiment No.:- 8
# # Roll No.- 34
# # UID:- 2019230071
# # Batch:- B
# + [markdown] id="fmywxCR8DWeX"
# <h2>Aim:- Understanding Support Vector Machine algorithm through building SVM algorithm in Python </h2>
#
# -
# <center> <h1> Support Vector Machine </h1><center>
# + [markdown] id="AykfHZgZDg5N"
# <h3>Importing the libraries </h3>
# + id="sTmr3VSzfP7h"
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# + [markdown] id="jKL35QIXDiWk"
# #### Linear Kernel
# + id="MOzOk5ijjfei"
from sklearn.metrics import confusion_matrix
# Function to draw a nice plot of an SVM
def plot_svc(svc, X, y, h=0.02, pad=0.25):
x_min, x_max = X[:, 0].min()-pad, X[:, 0].max()+pad
y_min, y_max = X[:, 1].min()-pad, X[:, 1].max()+pad
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max,h))
Z = svc.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.Paired, alpha=0.2)
plt.scatter(X[:,0], X[:,1], s=70, c=y, cmap=mpl.cm.Paired)
# Support vectors indicated in plot by vertical lines
sv = svc.support_vectors_
plt.scatter(sv[:,0], sv[:,1], c='k', marker='x', s=100, linewidths=1)
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xlabel('X1')
plt.ylabel('X2')
plt.show()
print('Number of support vectors: ', svc.support_.size)
# + id="0u01ooSNkMsp"
# Generating random data: 20 observations of 2 features and divide into two
np.random.seed(5)
X = np.random.randn(20,2)
y = np.repeat([1,-1], 10)
X[y == -1] = X[y == -1] + 1
# + colab={"base_uri": "https://localhost:8080/"} id="i21dY-9kkUux" outputId="7ba478d1-d841-4b38-e563-29f688d23b12"
X
# + colab={"base_uri": "https://localhost:8080/"} id="ibAsTTRekXAD" outputId="cd55a19a-8f80-4c3a-9bf3-b71387ddded0"
y
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="_iqH2MWXkXHZ" outputId="98c67150-b552-4c94-b2ac-dc8420b3f156"
# Scatter plot between the two features, where colour indicates the target class
plt.scatter(X[:,0], X[:,1], s=70, c=y, cmap=mpl.cm.Paired)
plt.xlabel('X1')
plt.ylabel('X2')
# + [markdown] id="8cGLwdYAD-kl"
# <h3>Linear or Non Linear? <br/>
# Not linearly separable </h3>
# + colab={"base_uri": "https://localhost:8080/"} id="ktSAXhn_mBe9" outputId="b537499a-fff2-4572-a011-717f4ba475ad"
# Fitting the Support Vector Classifier Model
from sklearn.svm import SVC
svc = SVC(C=1, kernel='linear')
svc.fit(X, y)
# + colab={"base_uri": "https://localhost:8080/"} id="ydizN60mmZhM" outputId="538fba03-1ece-49a9-b201-2376092d9240"
svc.support_ # Determining the identities of support vectors
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="NIhACSIpE7Ou" outputId="eecbd719-2dab-4760-b4da-794cad5660b1"
plot_svc(svc, X, y)
# + [markdown] id="RqNFbqkoBgWB"
# The region of feature space that will be assigned to the -1 class is shown in light blue, and the region that will be assigned to the +1 class is shown in brown.
#
# The decision boundary between the two classes is linear (because we used the argument kernel = ”linear”).
#
# The support vectors are plotted with crosses and the remaining observations are plotted as circles; we see here that there are 13 support vectors.
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="nXRB_G1A1EyR" outputId="7c528001-a937-4f8f-9c34-f1ae449809fe"
# Using a smaller value of C argument
svc2 = SVC(C=1, kernel='linear')
svc2.fit(X, y)
plot_svc(svc2, X, y)
# + colab={"base_uri": "https://localhost:8080/"} id="1Xm2UNfp1ISh" outputId="d2ed6e89-a608-4ecd-a415-7f7c72108211"
# Performing cross validation 10-fold
from sklearn.model_selection import GridSearchCV
# Select the optimal C parameter by cross-validation
tuned_parameters = [{'C': [0.001, 0.01, 0.1, 1, 5, 10, 100]}]
clf = GridSearchCV(
SVC(kernel='linear'),
tuned_parameters,
cv=10,
scoring='accuracy'
)
clf.fit(X, y)
# + colab={"base_uri": "https://localhost:8080/"} id="MgJtjYwO3eHL" outputId="d246c2b2-a3c2-494a-fbdf-918565488514"
clf.cv_results_ # Cross validation scores and errors for each split
# + colab={"base_uri": "https://localhost:8080/"} id="bPTkuB593hr6" outputId="081b798b-ed51-4515-9288-89d401afa817"
clf.best_params_
# + [markdown] id="TnsbgkQVBn25"
# C=0.001 is best parameter given by GridSearchCV
# + id="_G7jyrUO3nRL"
# Generating a test dataset
np.random.seed(1)
X_test = np.random.randn(20,2)
y_test = np.random.choice([-1,1], 20)
X_test[y_test == 1] = X_test[y_test == 1] -1
# + colab={"base_uri": "https://localhost:8080/"} id="Ozvfkhce4GuK" outputId="e6129fd7-df02-415c-b31b-d4e1caa1fdc5"
X_test
# + colab={"base_uri": "https://localhost:8080/"} id="wUH80zli4KlC" outputId="11e81a46-a1d9-4c65-bb25-275003d3c0e2"
y_test
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="r-tbjHqF4RcK" outputId="ba8b7e2d-9d88-4fb7-d39a-ef2b716ce762"
# Fitting SVC with best C value
svc2 = SVC(C=0.001, kernel='linear')
svc2.fit(X, y)
y_pred = svc2.predict(X_test)
pd.DataFrame(confusion_matrix(y_test, y_pred), index=svc2.classes_, columns=['-1','+1'])
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="6WSZXPxt4Syz" outputId="037931ba-8601-4416-d6e2-eb106f2d1e82"
pd.DataFrame(confusion_matrix(y_test, y_pred), index=['Actual -1','Actual +1'], columns=['Predicted -1','Predicted +1'])
# + colab={"base_uri": "https://localhost:8080/"} id="ae0KslWN4W4b" outputId="a54065ee-f76b-4829-d462-ca1b86f11b10"
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="-nQ-Zr1w4cSa" outputId="f852b8e3-282c-422b-9b9e-230f505aad14"
# Making nearly linearly separable data
X_test[y_test == 1] = X_test[y_test == 1] -1
plt.scatter(X_test[:,0], X_test[:,1], s=70, c=y_test, cmap=mpl.cm.Paired)
plt.xlabel('X1')
plt.ylabel('X2')
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="TNGIEYhh5FFb" outputId="660caba0-fa08-4023-fb00-e2074eaa4e27"
svc3 = SVC(C=1e5, kernel='linear')
svc3.fit(X_test, y_test)
plot_svc(svc3, X_test, y_test)
# + [markdown] id="Dj7AfKKKCPrE"
# No training errors were made and only three support vectors were used.
#
# However, we can see from the figure that the margin is very narrow (because the observations that are not support vectors, indicated as circles, are very close to the decision boundary).
#
# It seems likely that this model will perform poorly on test data
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="95SuyFKS5Szz" outputId="6f17eff9-b769-48ee-beba-da45042de8de"
# Trying a smaller value of C for this new dataset
svc4 = SVC(C=1, kernel='linear')
svc4.fit(X_test, y_test)
plot_svc(svc4, X_test, y_test)
# + [markdown] id="xJ5IYxLnCU6R"
# Using cost = 1, we misclassify a training observation, but we also obtain a much wider margin and make use of five support vectors.
#
# It seems likely that this model will perform better on test data than the model with cost = 1e5.
# + [markdown] id="7LB4v1EjCYhs"
# ### Non-linear Kernels
# + id="j102vIJk5X7r"
from sklearn.model_selection import train_test_split
np.random.seed(8)
X = np.random.randn(200,2)
X[:100] = X[:100] +2
X[101:150] = X[101:150] -2
y = np.concatenate([np.repeat(-1, 150), np.repeat(1,50)])
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="ZO7f2bf05mN7" outputId="5772d008-b8f2-4cdd-b955-a63e2fd24730"
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.5)
plt.scatter(X[:,0], X[:,1], s=70, c=y, cmap=mpl.cm.Paired)
plt.xlabel('X1')
plt.ylabel('X2')
# + [markdown] id="SVpzpnuVCh3h"
# One class is kind of stuck in the middle of another class. This suggests that we might want to use a radial kernel in our SVM.
# + [markdown] id="sbD-j64xDzTW"
# #### Rbf kernel
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="zdYqlGKM5q8j" outputId="8d174576-c075-4cb8-fac5-a661479a55a2"
# Using the SVC() function with a radial kernel and γ = 1
svm = SVC(C=1.0, kernel='rbf', gamma=1)
svm.fit(X_train, y_train)
plot_svc(svm, X_test, y_test)
# + [markdown] id="47aRfokuEA9x"
# The plot shows that the resulting SVM has a decidedly non-linear boundary.
#
# We can see from the figure that there are a fair number of training errors in this SVM fit.
#
# If we increase the value of cost, we can reduce the number of training error
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="Dg7BsgNC5yib" outputId="04708bc0-f654-47ba-b2bf-75a409f76d5c"
# Increasing C parameter, allowing more flexibility
svm2 = SVC(C=100, kernel='rbf', gamma=1.0)
svm2.fit(X_train, y_train)
plot_svc(svm2, X_test, y_test)
# + [markdown] id="f2W743cREE16"
# However, this comes at the price of a more irregular decision boundary that seems to be at risk of overfitting the data.
#
# We can perform cross-validation using GridSearchCV() to select the best choice of γ and cost for an SVM with a radial kernel
# + colab={"base_uri": "https://localhost:8080/"} id="ozzPCG3-6HCr" outputId="ad6e8f89-62ce-4bba-8283-d3d533595f8d"
tuned_parameters = [{'C': [0.01, 0.1, 1, 10, 100],'gamma': [0.5, 1,2,3,4]}]
clf = GridSearchCV(SVC(kernel='rbf'), tuned_parameters, cv=10, scoring='accuracy')
clf.fit(X_train, y_train)
clf.best_params_
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="j-mZ4lZc6M9c" outputId="e99bf6f6-33bf-4f62-d2ac-2628930681cb"
plot_svc(clf.best_estimator_, X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="YZQYJgH66W0c" outputId="d77f70bc-7d9c-4654-d94a-22444be6706c"
y_pred = clf.best_estimator_.predict(X_test)
pd.DataFrame(confusion_matrix(y_test, y_pred), index=['Actual -1','Actual +1'], columns=['Predicted -1','Predicted +1'])
# + colab={"base_uri": "https://localhost:8080/"} id="IugpSaAw6dyb" outputId="3dc5f0ef-12ec-4c53-ef47-074824fe143b"
print(clf.best_estimator_.score(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="kv44E7lA6m8G" outputId="f903099a-694b-4960-8280-69f6131cf40b"
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
# + [markdown] id="rbPsAmMEERmF"
# #### Polynomial kernel
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="17hzHmX66qos" outputId="66929e36-e35b-44c3-8092-a255c68aa4b2"
svm3 = SVC(C=100, kernel='poly', gamma=1.0)
svm3.fit(X_train, y_train)
plot_svc(svm2, X_test, y_test)
# + [markdown] id="TPq1TlsNEX2v"
# We can perform cross-validation using GridSearchCV() to select the best choice of γ and cost for an SVM with a polynomial kernel.
# + colab={"base_uri": "https://localhost:8080/"} id="yWenTBV16v2T" outputId="63a36280-68f3-4627-8c3b-036062b8a8ea"
tuned_parameters = [{'C': [0.01, 0.1, 1],'gamma': [0.5, 1, 2]}]
clf2 = GridSearchCV(SVC(kernel='poly'), tuned_parameters, cv=10, scoring='accuracy', n_jobs=-1)
clf2.fit(X_train, y_train)
clf2.best_params_
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="qj6b2wGt66oc" outputId="5cbb18c9-d875-480f-daa7-6eebf679a204"
plot_svc(clf2.best_estimator_, X_test, y_test)
# + colab={"base_uri": "https://localhost:8080/", "height": 112} id="I0aMiQNb7BX0" outputId="14032b76-bda8-471b-e729-78795e4007a9"
y_pred = clf2.best_estimator_.predict(X_test)
pd.DataFrame(confusion_matrix(y_test, y_pred), index=['Actual -1','Actual +1'], columns=['Predicted -1','Predicted +1'])
# + colab={"base_uri": "https://localhost:8080/"} id="3LLX6gCe7EJz" outputId="9cef5a30-28a0-4057-ccf8-3f86de2b5215"
print(clf2.best_estimator_.score(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="nYlVy_Lp7HRM" outputId="7a7ef015-5e67-4500-cc12-8fc0c295a90c"
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred))
# + [markdown] id="YxOWZwl2EctO"
# Rbf kernel performs better than polynomial kernel for the same data when non-linear SVM is used.
# + [markdown] id="3ZDXkEYHE66V"
# ### Working on Pulsar Dataset
# + [markdown] id="E8A4WoHBE_pG"
# #### Downloading the training set
# + colab={"base_uri": "https://localhost:8080/"} id="FFyGTaan6cpt" outputId="80691cdc-06a7-49c2-ab36-8d51429af53b"
# !gdown --id 1YSFW8DGpoU7iBB6TxnnltOFc1bkCBZC_
# + [markdown] id="XZx2gPvTFGt7"
# #### Downloading the testing set
# + colab={"base_uri": "https://localhost:8080/"} id="uZu6_ddh6xsP" outputId="1b2c4f73-d2d4-4434-c2c6-65428c4b4e81"
# !gdown --id 18suswE049ki9zxJreR45kGrvW14vgJ_N
# + [markdown] id="E6DdLhxFFJv7"
# #### Load training and testing set
# + id="qwFG_JMA62zy"
train = pd.read_csv('pulsar_data_train.csv')
# + id="pfkFI_8K6--0"
test = pd.read_csv('pulsar_data_test.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 302} id="vIJ8eq837Fg-" outputId="390602be-4496-4436-9165-b973befe9a2a"
train.head()
# + [markdown] id="EMe4H3Z5FO5d"
# #### Check for null values in dataset
# + colab={"base_uri": "https://localhost:8080/"} id="-Abvr7gi7KJM" outputId="4c46cd85-adc1-44a3-f62b-f080a1f59574"
train.isna().sum()
# + [markdown] id="WreMHtNpFgX5"
# #### Show the distribution of output class
# + colab={"base_uri": "https://localhost:8080/", "height": 288} id="nS7u8gBu7hR_" outputId="9197afb4-fa88-49e4-972e-9710448e33ab"
train['target_class'].value_counts().plot(kind='bar')
# + [markdown] id="wG-G4SdbGFkE"
# The dataset is imbalanced with class 1 as minority class
# + [markdown] id="DtFeKc-zFnIU"
# #### List the columns in dataset
# + colab={"base_uri": "https://localhost:8080/"} id="OtfkqxCV9Qe5" outputId="59216112-a5e6-43a9-e02a-0242ef74753f"
train.columns
# + [markdown] id="JYDSso4xFrWv"
# #### Use KNN Imputer to impute missing values
# + id="7Zyal9Nc7l_f"
from sklearn.impute import KNNImputer
# Imputer object using the mean strategy and
# missing_values type for imputation
imputer1 = KNNImputer(n_neighbors=2, weights="uniform")
traindata = imputer1.fit_transform(train)
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="i_Tni5ZQ9n-e" outputId="2a202ff5-0341-4a5e-b5a4-833e99d566af"
train
# + [markdown] id="D4JBSn_dF1sO"
# #### Create traindata as imputed dataset having no null values
# + id="2pfFcLVH-IWk"
traindata = pd.DataFrame(traindata, columns = train.columns)
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="NPdZ7uok-VdR" outputId="72f18d2a-a882-44f0-b0b7-9d5ef6033d5c"
traindata
# + colab={"base_uri": "https://localhost:8080/"} id="s6ltmvmg9qfb" outputId="70ac0c55-f024-4e96-daf6-317c0c42b70b"
traindata.isna().sum()
# + [markdown] id="0SED6e6aF-iL"
# #### Divide the dataset into X and y
# + id="QFE74YGZ-Zqu"
X = traindata.drop('target_class', axis = 1)
y = traindata['target_class']
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="DEVRLubd-4Le" outputId="23763e86-6031-4d41-bbed-e9a823326af0"
X
# + colab={"base_uri": "https://localhost:8080/"} id="L8_-sciC-6Wf" outputId="216f483d-7761-4994-9573-111522db9e74"
y
# + colab={"base_uri": "https://localhost:8080/"} id="cLKcCSku-7lJ" outputId="8b8c9c1a-0e8a-4afb-9b55-f9a83adb13fe"
# !pip install imbalanced-learn
# + [markdown] id="5Hy-M8C8GL2U"
# #### Import SMOTE to make the dataset balanced
# + id="1OPhNw2c_Aq5"
from imblearn.over_sampling import SMOTE
# + [markdown] id="dyy0m_R0GTYK"
# #### Oversampling the minority class
# + id="_r5tS3jV_DYg"
sm = SMOTE(random_state=42)
X_sampled, y_sampled = sm.fit_resample(X,y)
# + colab={"base_uri": "https://localhost:8080/", "height": 520} id="UbHrvlCX_o4y" outputId="8026d3cc-d65c-4359-8884-49df3598c585"
X_sampled
# + id="6jcbRXsB_rVl"
y_sampled_pd = pd.DataFrame(data=y_sampled)
# + [markdown] id="hC9eRFgNGXvo"
# #### Balanced dataset according to target_class
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="ClVIzO8v_NU4" outputId="843e0207-c8d6-462a-a60c-41e2008752b1"
y_sampled_pd['target_class'].value_counts().plot(kind='pie')
# + [markdown] id="8FgdMLsnGcO6"
# #### Split the dataset into 80% training set and 20% test set
# + id="TbvCWY8KAaXW"
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X_sampled,y_sampled_pd, test_size=0.20 , random_state=42)
# + [markdown] id="9wiuwFsaGijK"
# #### Use StandardScaler to apply transformation to training and test set
# + id="2JUWzwu0AqqV"
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
# + id="d3A5c88EAtdb"
X_train_scaled = sc.fit_transform(X_train)
X_test_scaled = sc.fit_transform(X_test)
# + [markdown] id="S0WBim4OGroA"
# #### Apply SVM with rbf kernel and note the accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="_VbYXiPFA463" outputId="768abae8-23a1-465f-f9a0-2802d4ba18e8"
from sklearn.metrics import accuracy_score
svm_rbf = SVC(kernel='rbf')
svm_rbf.fit(X_train_scaled, y_train)
y_pred_rbf = svm_rbf.predict(X_test_scaled)
print('Support Vector Machine RBF kernel accuracy= ', accuracy_score(y_test, y_pred_rbf))
# + [markdown] id="dOnN8E5QH__Y"
# #### Print the classification report
# + colab={"base_uri": "https://localhost:8080/"} id="X_PpqZvFA6Fw" outputId="fb541c74-3ff8-438b-f3e7-02537a547af1"
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred_rbf))
# + [markdown] id="neVkfIenIE4W"
# #### Apply SVM with linear kernel and note the accuracy
# + colab={"base_uri": "https://localhost:8080/"} id="P9u3WwwvBvCv" outputId="87bd16e1-9240-45c8-bb40-9b61c5cbc802"
svm_linear= SVC(kernel='linear')
svm_linear.fit(X_train_scaled, y_train)
y_pred_linear = svm_linear.predict(X_test_scaled)
print('Support Vector Machine Linear kernel accuracy= ', accuracy_score(y_test, y_pred_linear))
# + [markdown] id="6LCIoxEEILqJ"
# #### Print the classification report
# + colab={"base_uri": "https://localhost:8080/"} id="oS6mrBZ9A789" outputId="7cacd4d7-790e-47ed-a8f5-676f5ba605bd"
print(classification_report(y_test, y_pred_linear))
# + colab={"base_uri": "https://localhost:8080/"} id="C9tuVe1SIPBC" outputId="aa880dda-057d-46a8-ac7b-caf90ccf768d"
svm_poly= SVC(kernel='poly')
svm_poly.fit(X_train_scaled, y_train)
y_pred_poly = svm_poly.predict(X_test_scaled)
print('Support Vector Machine Linear kernel accuracy= ', accuracy_score(y_test, y_pred_poly))
# + [markdown] id="YYy9rrThIgnf"
# Inference:
#
#
# 1. The dataset was imbalanced, but it was balanced using SMOTE technique
# 2. The rbf kernel of SVM has more accuracy than linear and poly kernels.
#
#
# + [markdown] id="62Ug1Kn6I4Or"
# Conclusion: Hence, from this experiment, I understood the working of Support vector machine and its different kernels. Also, I applied SVM on Pulsar dataset. The dataset was balanced using SMOTE technique. Then, SVM with rbf,linear and poly kernels was applied and rbf kernel gave the highest accuracy of 94.37%.
| exp8/EXP8_Support_Vector_Machine.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lesson 12: GAN's
#
# <img src='https://snag.gy/lAKoVw.jpg' style='width:600px'/>
#
# ### Notes from last week - language model for medicine
#
# <img src='https://snag.gy/k7nY3X.jpg' style='width:600px' />
#
# - **Seeding a language model.** Once the language model is informed and updated, it can be seeded with some words to get the generative model working on generating text. In this case, a language model was fed a number of texts, and the "seed" or the prompt was a question.
#
# ### Competition on training CiFAR10
#
# We are going to be building some barebones models. And currently ( as of april) - how fast can one train a model to get to 94% accuracy?
#
# <img src='https://snag.gy/SnOx7W.jpg' style='width:600px'/>
| live_notes/dl2_050_GANs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example: D-optimal experiment design with random datasets
# Solve the D-Optimal experiment design problem
# $$
# \begin{array}{ll}
# \textrm{minimize} & F(x):=\log\left(\det\left(\sum_{i=1}^n x_i V_i V_i^T\right)\right) \\
# \textrm{subject to} & \sum_{i=1}^n x_i = 1, \\
# & x_i\geq 0, \quad i=1,\ldots,n
# \end{array}
# $$
# where $V_i\in R^m$ for $i=1,\ldots,n$.
# +
# #cd C:\\github\accbpg
# -
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
matplotlib.rcParams.update({'font.size': 16, 'legend.fontsize':14, 'font.family': 'serif'})
# matplotlib.rcParams.update({'text.usetex': True})
# ### Compare BPG and ABPG with different values of gamma
# +
import accbpg
# Generate a random instance of the D-optimal design problem of size m by n
m = 80
n = 200
f, h, L, x0 = accbpg.D_opt_design(m, n, randseed=10)
# Solve the problem using BPG and ABPG with different values of gamma (TSE)
x00, F00, G00, T00 = accbpg.BPG( f, h, L, x0, maxitrs=1000, linesearch=False, verbskip=100)
x10, F10, G10, T10 = accbpg.ABPG(f, h, L, x0, gamma=1.0, maxitrs=1000, theta_eq=True, verbskip=100)
x15, F15, G15, T15 = accbpg.ABPG(f, h, L, x0, gamma=1.5, maxitrs=1000, theta_eq=True, verbskip=100)
x20, F20, G20, T20 = accbpg.ABPG(f, h, L, x0, gamma=2.0, maxitrs=1000, theta_eq=True, verbskip=100)
x22, F22, G22, T22 = accbpg.ABPG(f, h, L, x0, gamma=2.2, maxitrs=2000, theta_eq=True, verbskip=100)
# +
# Plot the objective gap and estimated gains for triangle scaling
fig, _ = plt.subplots(1, 2, figsize=(11, 4))
labels = [r"BPG", r"ABPG $\gamma=1.0$", r"ABPG $\gamma=1.5$", r"ABPG $\gamma=2.0$", r"ABPG $\gamma=2.2$"]
styles = ['k:', 'g-', 'b-.', 'k-', 'r--']
dashes = [[1,2], [], [4,2,1,2], [], [4,2]]
ax1 = plt.subplot(1, 2, 1)
y_vals = [F00, F10, F15, F20, F22]
accbpg.plot_comparisons(ax1, y_vals, labels, x_vals=[], plotdiff=True, yscale="log", xlim=[-10, 1000], ylim=[1e-5, 2],
xlabel=r"Iteration number $k$", ylabel=r"$F(x_k)-F_\star$", legendloc="upper right",
linestyles=styles, linedash=dashes)
ax2 = plt.subplot(1, 2, 2)
y_vals = [G10, G15, G20, G22]
accbpg.plot_comparisons(ax2, y_vals, labels[1:], x_vals=[], plotdiff=False, yscale="log", xlim=[-10, 1000], ylim=[1e-3, 5],
xlabel=r"Iteration number $k$", ylabel=r'$\hat{G}_k$', legendloc="lower right",
linestyles=styles[1:], linedash=dashes[1:])
plt.tight_layout(w_pad=4)
plt.show()
# pad = 0.05
# extent = ax1.get_tightbbox(fig.canvas.renderer).expanded(1+pad, 1+pad).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('./figures/D_opt_m80n200gamma_semilog.pdf', format='pdf', bbox_inches=extent)
# extent = ax2.get_tightbbox(fig.canvas.renderer).expanded(1+pad, 1+pad).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('./figures/D_opt_m80n200gamma_gains.pdf', format='pdf', bbox_inches=extent)
# -
# ### Comparing BPG with line search and adaptive ABPG algorithms
# Solve the problem using BPG w/o line seach and adaptive ABPG with gamma=2 (TSE)
x00, F00, G00, T00 = accbpg.BPG( f, h, L, x0, maxitrs=1000, linesearch=False, verbskip=100)
xLS, FLS, GLS, TLS = accbpg.BPG( f, h, L, x0, maxitrs=1000, linesearch=True, verbskip=100)
x20, F20, G20, T20 = accbpg.ABPG(f, h, L, x0, gamma=2.0, maxitrs=1000, theta_eq=True, verbskip=100)
x2e, F2e, Gamma2e, G2e, T2e = accbpg.ABPG_expo(f, h, L, x0, gamma0=3, maxitrs=1000, theta_eq=True, verbskip=100)
x2g, F2g, G2g, Gdiv2g, Gavg2g, T2g = accbpg.ABPG_gain(f, h, L, x0, gamma=2, maxitrs=3000, G0=0.1, theta_eq=True, verbskip=100)
# +
# Plot the objective gap and estimated gains for triangle scaling
fig, _ = plt.subplots(1, 2, figsize=(11, 4))
labels = [r"BPG", r"BPG-LS", r"ABPG", r"ABPG-e", r"ABPG-g"]
styles = ['k:', 'g-', 'b-.', 'k-', 'r--']
dashes = [[1,2], [], [4,2,1,2], [], [4,2]]
ax1 = plt.subplot(1, 2, 1)
y_vals = [F00, FLS, F20, F2e, F2g]
accbpg.plot_comparisons(ax1, y_vals, labels, x_vals=[], plotdiff=True, yscale="log", xlim=[-10, 1000], ylim=[1e-5, 2],
xlabel=r"Iteration number $k$", ylabel=r"$F(x_k)-F_\star$", legendloc="upper right",
linestyles=styles, linedash=dashes)
ax2 = plt.subplot(1, 2, 2)
accbpg.plot_comparisons(ax2, y_vals, labels, x_vals=[], plotdiff=True, yscale="log", xscale='log', xlim=[1, 1000], ylim=[1e-5, 2],
xlabel=r"Iteration number $k$", ylabel=r"$F(x_k)-F_\star$", legendloc="lower left",
linestyles=styles, linedash=dashes)
plt.tight_layout(w_pad=4)
plt.show()
# pad = 0.05
# extent = ax1.get_tightbbox(fig.canvas.renderer).expanded(1+pad, 1+pad).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('./figures/D_opt_m80n200adapt_semilog.pdf', format='pdf', bbox_inches=extent)
# extent = ax2.get_tightbbox(fig.canvas.renderer).expanded(1+pad, 1+pad).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('./figures/D_opt_m80n200adapt_loglog.pdf', format='pdf', bbox_inches=extent)
# -
# ### Comparing ABPG and ABPG with restart for nearly strongly convex problems
# +
# Generate a random instance of the D-optimal design problem of size m by n
ms = 80
ns = 120
fs, hs, Ls, x0s = accbpg.D_opt_design(ms, ns, randseed=10)
# Solve the problem using BPG w/o line seach and adaptive ABPG with gamma=2 (TSE), also with restart
xs00, Fs00, Gs00, _ = accbpg.BPG( fs, hs, Ls, x0s, maxitrs=1000, linesearch=False, verbskip=100)
xsLS, FsLS, GsLS, _ = accbpg.BPG( fs, hs, Ls, x0s, maxitrs=1000, linesearch=True, verbskip=100)
xs20, Fs20, Gs20, _ = accbpg.ABPG(fs, hs, Ls, x0s, gamma=2.0, maxitrs=1000, theta_eq=True, restart=False, verbskip=100)
xs20rs, Fs20rs, Gs20rs, _ = accbpg.ABPG(fs, hs, Ls, x0s, gamma=2.0, maxitrs=1000, theta_eq=True, restart=True, verbskip=100)
xs2g, Fs2g, Gs2g, _, _, _ = accbpg.ABPG_gain(fs, hs, Ls, x0s, gamma=2, maxitrs=3000, G0=0.1, theta_eq=True, restart=False, verbskip=100)
xs2grs, Fs2grs, Gs2grs, _, _, _ = accbpg.ABPG_gain(fs, hs, Ls, x0s, gamma=2, maxitrs=3000, G0=0.1, theta_eq=True, restart=True, verbskip=100)
# +
# Plot the objective gap and estimated gains for triangle scaling
fig, _ = plt.subplots(1, 2, figsize=(11, 4))
labels = [r"BPG", r"BPG-LS", r"ABPG", r"ABPG RS", r"ABPG-g", r"ABPG-g RS"]
styles = ['k:', 'g-', 'b-.', 'm-', 'k-', 'r--']
dashes = [[1,2], [], [4,2,1,2], [4,2,1,2,1,2], [], [4,2]]
ax1 = plt.subplot(1, 2, 1)
y_vals = [Fs00, FsLS, Fs20, Fs20rs, Fs2g, Fs2grs]
accbpg.plot_comparisons(ax1, y_vals, labels, x_vals=[], plotdiff=True, yscale="log", xlim=[0, 50], ylim=[1e-10, 1],
xlabel=r"Iteration number $k$", ylabel=r"$F(x_k)-F_\star$", legendloc="upper right",
linestyles=styles, linedash=dashes)
ax2 = plt.subplot(1, 2, 2)
y_vals = [GsLS, Gs20, Gs20rs, Gs2g, Gs2grs]
accbpg.plot_comparisons(ax2, y_vals, labels[1:], x_vals=[], plotdiff=False, yscale="log", xlim=[0, 50], ylim=[1e-1, 1e2],
xlabel=r"Iteration number $k$", ylabel=r'$\hat{G}_k$', legendloc="upper left",
linestyles=styles[1:], linedash=dashes[1:])
plt.tight_layout(w_pad=4)
plt.show()
# pad = 0.05
# extent = ax1.get_tightbbox(fig.canvas.renderer).expanded(1+pad, 1+pad).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('./figures/D_opt_m80n120restart_semilog.pdf', format='pdf', bbox_inches=extent)
# extent = ax2.get_tightbbox(fig.canvas.renderer).expanded(1+pad, 1+pad).transformed(fig.dpi_scale_trans.inverted())
# fig.savefig('./figures/D_opt_m80n120restart_gains.pdf', format='pdf', bbox_inches=extent)
# -
| ipynb/ex_Dopt_random.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from keras.layers import Input, Dense, Dropout
from keras.models import Model
from keras.datasets import mnist
from keras.models import Sequential, load_model
from keras.optimizers import RMSprop
from keras.callbacks import TensorBoard
from __future__ import print_function
from keras.utils import plot_model
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
import keras
import matplotlib.pyplot as plt
import numpy as np
import math
import pydot
import graphviz
import pandas as pd
# -
# # Data Set Information
# This radar data was collected by a system in Goose Bay, Labrador. This system consists of a phased array of 16 high-frequency antennas with a total transmitted power on the order of 6.4 kilowatts. See the paper for more details. The targets were free electrons in the ionosphere. "Good" radar returns are those showing evidence of some type of structure in the ionosphere. "Bad" returns are those that do not; their signals pass through the ionosphere.
#
# Received signals were processed using an autocorrelation function whose arguments are the time of a pulse and the pulse number. There were 17 pulse numbers for the Goose Bay system. Instances in this databse are described by 2 attributes per pulse number, corresponding to the complex values returned by the function resulting from the complex electromagnetic signal.
# ## Attribute Information
# - All 34 are continuous
# - The 35th attribute is either "good" or "bad" according to the definition summarized above. This is a binary classification task.
# - https://archive.ics.uci.edu/ml/machine-learning-databases/ionosphere/ionosphere.names
# ## Data Import and preprocessing
data = np.genfromtxt("data/ionosphere.data")
data = pd.read_csv('data/ionosphere.data', sep=",", header=None)
data.describe()
data.head()
data.drop(data.columns[1], inplace=True, axis=1)
data[34] = [1 if e is "g" else 0 for e in data[34]]
data.head()
# sample the dataframe
data_train = data.sample(frac=0.9, random_state=42)
data_valid = data.drop(data_train.index)
# +
df_x_train = data_train.iloc[:,:-1]
df_y_train = data_train.iloc[:,-1]
df_x_valid = data_valid.iloc[:,:-1]
df_y_valid = data_valid.iloc[:,-1]
# -
df_y_train.sum()
df_y_train.sum()/len(df_y_train)
# ## preprocessing
# +
x_train = np.array(df_x_train.as_matrix())
y_train = np.array(pd.DataFrame(df_y_train).as_matrix())
x_val = np.array(df_x_valid.as_matrix())
y_val = np.array(pd.DataFrame(df_y_valid).as_matrix())
# -
input_dim = x_train.shape[1]
# ## Train Classifier
# The goal is to get a very accurate classifier.
result = []
for i in range(1,5):
model = Sequential()
model.add(Dense(15, input_dim=input_dim, activation='relu'))
model.add(Dropout(0.1))
model.add(Dense(15, activation='relu'))
model.add(Dropout(0.1))
model.add(Dense(15, activation='relu'))
model.add(Dropout(0.1))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='adadelta',
metrics=['accuracy'])
model.fit(x_train, y_train,
epochs=150,
shuffle=True,
batch_size=4,
verbose=0,
validation_split=0.1
)
score = model.evaluate(x_val, y_val)[1]
result.append(score)
print(np.mean(result))
# ## Single Hidden Layer Autoencoder
result = []
for i in range(1,5):
single_auto = Sequential()
single_auto.add(Dense(8, input_dim=input_dim, activation='relu'))
# model.add(Dropout(0.1))
single_auto.add(Dense(input_dim, activation='sigmoid'))
single_auto.compile(loss='mean_squared_error',
optimizer='adadelta')
single_auto.fit(x_train, x_train,
epochs=150,
shuffle=True,
batch_size=4,
verbose=0,
validation_split=0.1
)
score = single_auto.evaluate(x_val, x_val)
result.append(score)
print(np.mean(result))
# How well does a classifier after the vanilla autoencoder perform?
model.evaluate(single_auto.predict(x_val), y_val)
# So we lose about 12% by decoding
# # Stacked Autoencoder
######## constants for stacked autoencoder ############
encoding_dim1 = 16
encoding_dim2 = 8
decoding_dim1 = 16
decoding_dim2 = input_dim
result = []
for i in range(1,5):
stacked_auto = Sequential()
stacked_auto.add(Dense(encoding_dim1, input_dim=input_dim, activation='relu'))
# model.add(Dropout(0.1))
stacked_auto.add(Dense(encoding_dim2, activation='relu'))
# model.add(Dropout(0.05))
stacked_auto.add(Dense(decoding_dim1, activation='relu'))
# model.add(Dropout(0.05))
stacked_auto.add(Dense(decoding_dim2, activation='sigmoid'))
stacked_auto.compile(loss='mean_squared_error',
optimizer='adadelta')
stacked_auto.fit(x_train, x_train,
epochs=150,
shuffle=True,
batch_size=4,
verbose=0,
validation_split=0.1
)
score = stacked_auto.evaluate(x_val, x_val)
result.append(score)
print(np.mean(score))
| .ipynb_checkpoints/IONOSPHERE_Autoencoder-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Download MAST TESS Light Curves Within an FFI Footprint Using TAP
# <br> This notebook is a demo for accessing Transiting Exoplanet Survey Satellite (TESS) data in the Common Archive Observation Model (CAOM) catalog at MAST, using a Virtual Observatory standard Table Access Protocol (TAP) service.
#
# ***
# ### Table of Contents
#
# 1. [TAP Service Introduction](#TAP-Service-Introduction)
# 2. [Imports](#Imports)
# 3. [Service Specific Configuration](#Service-Specific-Configuration)
# 4. [Connecting to the TAP Service](#Connecting-to-the-TAP-Service)
# 5. [Use Case: Getting light curves from a sector, camera, and chip](#Use-Case:-Getting-images-from-a-sector,-camera,-and-chip)
# - [Step 1: Getting the footprint](#Step-1:-Getting-the-footprint)
# - [Step 2: Getting an inventory of TESS lightcurves within the footprint](#Step-2:-Getting-an-inventory-of-TESS-lightcurves-within-the-footprint)
# 5. [Additional Resources](#Additional-Resources)
# 6. [About This Notebook](#About-this-Notebook)
# ***
# ## TAP Service Introduction
# Table Access Protocol (TAP) services allow more direct and flexible access to astronomical data than the simpler types of IVOA standard data services. Queries are built with the SQL-like Astronomical Data Query Language (ADQL), and can include geographic / spatial queries as well as filtering on other characteristics of the data. This also allows the user fine-grained control over the returned columns, unlike the fixed set of coumns retunred from cone, image, and spectral services.
#
# For this example, we'll be using the astroquery TAP/TAP+ client, which was developed by the ESAC Space Data Centre for working with the GAIA catalog, but is interoperable with other valid TAP services, including those at MAST. As an astroquery project, TAP+ documentation is available at ReadTheDocs: http://astroquery.readthedocs.io/en/latest/utils/tap.html
#
# We'll be using TAP+ to call the CAOM Catalog TAP service at MAST and filter the results for TESS-related information. The schema for this catalog is an IVOA standard, and is also described within the service itself.
#
#
# ***
# ## Imports
# +
# Use the astroquery TapPlus library as our client to the data service.
from astroquery.utils.tap.core import TapPlus
# For handling ordinary astropy Tables in responses
from astropy.table import Table
# For displaying and manipulating some types of results
import requests
import astropy
import time
# To allow display tweaks for wider response tables
from IPython.core.display import display
from IPython.core.display import HTML
# -
# ## Service Specific Configuration
#
# Every TAP service has a "Base URL" plus associated endpoints for synchronous and asynchronous queries, as well as status and capability information, and sometimes service-provided sample queries. The endpoints are predefined in the TAP standard, so clients can infer them using the base. We therefore only have to provide astroquery that base.
TAP_URL = "http://vao.stsci.edu/caomtap/tapservice.aspx"
# ## Connecting to the TAP Service
#
# The TapPlus library is able to connect to any TAP service, given the "base" URL as noted in metadata registry resources describing the service. The CAOM TAP service at MAST has access to TESS FFI and time series, including file URLs for download.
TAP_service = TapPlus(url=TAP_URL)
# ## Use Case: Getting images from a sector, camera, and chip
# ### Step 1: Getting the footprint
#
# For our purposes, any one footprint related to a sector, camera, and chip combination is good enough. We are not currently accounting for small movements of the spacecraft to form a composite footprint. Observation IDs for this mission are constructed based on sector, camera, and chip combination, and we can use this to launch our footprint search:
sector = '1'
camera = '1'
chip = '2'
observationIDwildcard = 'tess%-s000{}-{}-{}-%-s'.format(sector, camera, chip)
# Here we query for a single ID and footprint to start with. For filters, we use the TESS mission, target types associated with light curves instead of full frame images, and the sector number. Note that the sector ID (sequence number) is in a different table in the CAOM database than most of the metadata we want, so we have to join these tables based on the shared observation ID.
#
# This query uses an asynchronous job with longer timeouts in case of connection issues.
# +
job = TAP_service.launch_job_async("""
SELECT top 1 obs_id, s_region
FROM dbo.caomobservation JOIN ivoa.obscore on dbo.caomobservation.observationID = ivoa.obscore.obs_id
WHERE collection = 'TESS' and trgType = 'FIELD' and
sequenceNumber = {} and
observationID like '{}'
""".format(sector, observationIDwildcard))
# Uncomment the line below to see the whole query with wildcards replaced
#print(job.parameters)
footprint_results = job.get_results()
footprint_results
# -
# ### Step 2: Getting an inventory of TESS lightcurves within the footprint
# Here we take the footprint from the above query and find all lightcurves intersecting with this footprint, in ALL sectors. Depending on where this is in the sky, there could be responses only in the original sector, or there could be overlaps with other sectors. There would be more sector overlap near the poles, for instance. By filtering on obs_collection = TESS, we filter based on the TESS mission and exclude High Level Science Products (including those based on TESS).
#
# The format must be reformatted for our next query. We separate the shape from the list of vertices, which themselves must be comma-separated.
# +
footprint = footprint_results['s_region'][0].decode('UTF-8')
footprintShape = footprint[0:footprint.find(' ')]
footprintVertices = footprint[footprint.find(' '):].strip().replace(' ', ', ')
print(footprintShape)
print(footprintVertices)
# -
# Once the footprint has been isolated and reformatted, we perform another query listing all lightcurves (minus the data validation timeseries, whose files end in _dvt) by their target name, sector, RA and Dec, as well as returning the access url for each FITS file and its estimated file size. We are doing this as an asynchronous query, which handles longer response times, just in case.
# +
job = TAP_service.launch_job_async("""
SELECT target_name, sequenceNumber as sector, s_ra, s_dec, access_url, access_estsize, obs_id
FROM dbo.caomobservation JOIN ivoa.obscore on dbo.caomobservation.observationID = ivoa.obscore.obs_id
WHERE
obs_collection = 'TESS' and dataproduct_type = 'timeseries'
and access_url like '%lc.fits' and
CONTAINS(POINT('ICRS',s_ra,s_dec),{}('ICRS', {}))=1
ORDER BY obs_id
""".format(footprintShape, footprintVertices))
# Uncomment the line below to see the whole query with wildcards replaced
#print(job.parameters)
TAP_results = job.get_results()
TAP_results
# -
# The returned data is in an astropy table; you can manipulate it to do more ordering or filtering. To download individual files or the whole set, you can use the <i>access_url</i> column, as below.
#
# Python's requests library lets you download files from a URL. The downloads will appear in the directory where your notebook is running.
# +
# Example: first result row:
single_url = TAP_results['access_url'][0].decode('UTF-8')
filename = TAP_results['obs_id'][0].decode('UTF-8') + "_lc.fits"
r = requests.get(single_url, allow_redirects=True)
open(filename, 'wb').write(r.content)
print('File downloaded: {} bytes'.format(r.headers['Content-length']))
# Uncomment the code below to download every file in a loop.
# Warning: this can take some time as there are ~900 2megabyte files listed from the notebook's original sample query
# (see "True length" of masked table above).
#for rows in TAP_results:
# single_url = rows['access_url'].decode('UTF-8')
# filename = rows['obs_id'].decode('UTF-8') + "_lc.fits"
# r = requests.get(single_url, allow_redirects=True)
# open(filename, 'wb').write(r.content)
#print('All files downloaded.')
# -
# If you have problems running requests or would rather save individual files through your browser, you can simply print clickable links, instead, or wrap them in curl or wget calls, which may be different based on your operating system.
# +
# Example: first result row:
single_url = TAP_results['access_url'][0].decode('UTF-8')
print(single_url)
# Uncomment the code below to display clickable links for every file in a loop.
#for rows in TAP_results:
# single_url = rows['access_url'].decode('UTF-8')
# print(single_url)
# -
# ***
# # Additional Resources
#
#
# ## Table Access Protocol
# * IVOA standard for RESTful web service access to tabular data
# * http://www.ivoa.net/documents/TAP/
#
#
# ## Astronomical Query Data Language (2.0)
# * IVOA standard for querying astronomical data in tabular format, with geometric search support
# * http://www.ivoa.net/documents/latest/ADQL.html
#
#
# ## Common Archive Observation Model (2)
# * IVOA standard data model whose relational representation this catalog follows
# * http://www.opencadc.org/caom2/
#
#
# ## TapPlus
# * Module created by ESAC Space Data Centre
# * http://astroquery.readthedocs.io/en/latest/utils/tap.html
# ***
# ## About this Notebook
# **Authors:** <NAME> & <NAME>, STScI Archive Scientists & Software Engineer
# **Updated On:** 07/09/2019
# ***
# <img style="float: right;" src="./stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="stsci_pri_combo_mark_horizonal_white_bkgd" width="200px"/>
| notebooks/MAST/TESS/beginner_tess_tap_search/beginner_tess_tap_search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
all = pd.read_csv('/mnt/Tsunami_HHD/Tang/all_feature_data/all_clusters_all_features_add_clusterID_add_cluster_2_new.csv')
# +
import pandas as pd
all = pd.read_csv('/home/tongwade780/pdb_website/python_code/all_cluster_feature_csv_new/cluster_1000_all_feature.csv')
# -
all
# +
import numpy as np
import pandas as pd
import h5py
f2 = h5py.File('/mnt/Tsunami_HHD/Tang/onebyone_clsuter/feature_matrix_GNM_profile/GNM_profile_3rnx_individual_feature.mat','r')
datas2= f2['nan_shift_GNM_co'].value.T
matlab_data = pd.DataFrame(datas2,columns=['GNM_profile'])
matlab_data['PDB_ID'] = ['3rnx']*len(matlab_data)
all = all.merge(matlab_data,how = 'left',on = 'PDB_ID')
# +
import sys
sys.path.append('/home/tongwade780/pdb_website/python_code/')
from charge_sasa_dssp_new import *
import os
import csv
import numpy as np
import pandas as pd
import h5py
import time
start = time.time()
pdbname_file = open ('/home/tongwade780/pdb_website/all_ca_number_all_code/onechain_100clust.csv').readlines ()
next_file = []
problem_pdb = []
frames = []
all_gnm = []
for line in pdbname_file[0:]:
next_file.append(line.rstrip('\r\n' + '').split(','))
for cluster in range(0,2792):
real_cluster_num = cluster+1
cluster_pdb_name = next_file[cluster]
cluster_pdb_name = np.asarray(cluster_pdb_name)
cluster_pdb_name = filter(None, cluster_pdb_name)
try:
all = pd.read_csv('/home/tongwade780/pdb_website/python_code/all_cluster_feature_csv_new/cluster_{0}_all_feature.csv'.format(real_cluster_num))
except:
problem_pdb.append((pdbname,'no_features'))
continue
frames = []
frames2 = []
for pdb_chain_name in cluster_pdb_name:
pdbname= pdb_chain_name[0:4].lower()
try:
f = h5py.File('/mnt/Tsunami_HHD/Tang/onebyone_clsuter/feature_matrix/{0}_individual_feature.mat'.format(pdbname),'r')
f2 = h5py.File('/mnt/Tsunami_HHD/Tang/onebyone_clsuter/feature_matrix_GNM_profile/GNM_profile_{0}_individual_feature.mat'.format(pdbname),'r')
except:
problem_pdb.append((pdbname,'no_gnm_profile'))
continue
datas = f['individual_feature'].value.T
datas2= f2['nan_shift_GNM_co'].value.T
matlab_data2 = pd.DataFrame(datas,index=datas[:,0],columns=['resID','eig_vv_1','eig_vv_2','vector_1','Rank_1','vector_2',
'Rank_2','GNM_co','co_rank','entropy','rmsf_overall','pca_feature',
'eig_first','eig_sec','shannon_entropy','shannon_entropy20',
'eig5_eig1','atomic_1','atomic_3','atomic_5','ssbond_matrix',
'Polarity','charge','stand_Dcom','side_chain_length',
'ca_len','hole_ss_bond','contact_per_res','gyradius',
'phobic_percent','RMSF','RMSF_0f_bfactor','shifted_RMSF',
'h_bond_group','intrinsicDisorder'])
matlab_data = pd.DataFrame(datas2,columns=['GNM_profile'])
matlab_data['PDB_ID'] = ['{0}'.format(pdbname)]*len(matlab_data)
frames2.append(matlab_data2)
frames.append(matlab_data)
result = pd.concat(frames)
result2 = pd.concat(frames2)
result['resID'] = result2['resID'].values
all_gnm.append(result)
result3 = pd.concat(all_gnm)
# all['GNM_profile'] = result['GNM_profile'].values
# all.to_csv('/home/tongwade780/pdb_website/python_code/all_cluster_feature_csv/cluster_{0}_all_feature.csv'.format(real_cluster_num),index = False)
# all = all.merge(result,how = 'left',on = 'PDB_ID')
# all.to_csv('/home/tongwade780/pdb_website/python_code/all_cluster_feature_csv/cluster_{0}_all_feature.csv'.format(real_cluster_num),index = False)
# +
all = all.merge(result3,how = 'left',on = ['PDB_ID','resID'])
change_order = ['PDB_ID','resID','atomic_1','atomic_3','atomic_5','ssbond_matrix','Polarity',
'eig_vv_1','eig_vv_2','vector_1','Rank_1','vector_2','Rank_2',
'GNM_co','co_rank','charge','stand_Dcom','side_chain_length','pka_charge',
'dssp_result','sasa','entropy','ca_len','hole_ss_bond','rmsf_overall',
'contact_per_res','gyradius','pca_feature','eig_first','eig_sec',
'shannon_entropy','shannon_entropy20','eig5_eig1','phobic_percent',
'loop_percent','h_bond_group','intrinsicDisorder','pca_feature_2','pca_feature_3','sheet_percent','helix_percent','RMSF','RMSF_0f_bfactor',
'shifted_RMSF','GNM_profile','clusterID']
all = all[change_order]
# -
all.to_csv('/mnt/Tsunami_HHD/Tang/all_feature_data/cluster_all_feature_add_gnm_profile.csv',index = False)
# +
#select pca 3 mode
pdbname_file = open ('/home/tongwade780/pdb_website/all_ca_number_all_code/onechain_100clust.csv').readlines ()
next_file = []
for line in pdbname_file[1673:]:
next_file.extend(line.strip().split(','))
import pandas as pd
next_file_new = filter(None, next_file)
next_file_new = map(lambda x: x[0:4].lower(),next_file_new)
pdbname_file = open ('/home/tongwade780/pdb_website/python_code/all_cluster_problem_pdb_add_feature/pca_3_mode_pdb_name_new.csv').readlines ()
next_file = []
for line in pdbname_file[0:]:
next_file.append(line.rstrip('\r\n')[0:4].lower())
next_file.extend(next_file_new)
all_new = all[all.PDB_ID.isin(next_file)]
all_new.to_csv('/mnt/Tsunami_HHD/Tang/all_feature_data/cluster_all_feature_add_gnm_profile_3_mode.csv',index = False)
# +
pdbname_file = open ('/home/tongwade780/pdb_website/python_code/close_meancoord_pdb_add_feature.csv').readlines ()
next_file = []
for line in pdbname_file[0:]:
next_file.append(line.rstrip('\r\n')[0:4].lower())
next_file
all_new_one = all[all.PDB_ID.isin(next_file)]
all_new_one.to_csv('/mnt/Tsunami_HHD/Tang/all_feature_data/cluster_all_feature_add_gnm_profile_1_mode.csv',index= False)
# -
all
| python_tornado/get_GNM_profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="K9q6exejeHE6"
# !pip3 install tensorflow==1.15
# + id="JI8wg9T8eKnu"
import tensorflow as tf
# + id="WFTBPv_Bd6z6"
hello = tf.constant("Hello, Tensorflow!")
sess = tf.Session()
print(sess.run(hello))
# + id="h36q0goeeumr"
# graph 정의하기
node1 = tf.constant(3.0, tf.float32)
node2 = tf.constant(4.0)
node3 = tf.add(node1, node2)
sess = tf.Session()
print("sess.run(node1, node2):", sess.run([node1, node2]))
print("sess.run(node3)]:", sess.run(node3))
# + id="tI6BQutPfIcm"
# graph를 미리 정의하고 값을 넘겨주는 방식
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
adder_node = a + b
print( sess.run(adder_node, feed_dict={a:3, b:4.5}))
print( sess.run(adder_node, feed_dict={a:[1,3], b:[2,4]}))
# + [markdown] id="pgCKD7CJgesr"
# # Linear Regreesion 구현하기
# + id="5Tf22FrKfkJt"
x_train = [1,2,3,4,5]
y_train = [2.1,3.1,4.1,5.1,6.1]
x = tf.placeholder(tf.float32)
y = tf.placeholder(tf.float32)
w = tf.Variable(tf.random_normal([1]), name='weight')
b = tf.Variable(tf.random_normal([1]), name='bias')
hypothesis = x_train * w + b
# loss function
# reduce_mean은 tensor를 평균 내주는 메소드
cost = tf.reduce_mean(tf.square(hypothesis - y_train))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
train = optimizer.minimize(cost)
sess = tf.Session()
# w, b를 초기화 해준다.
sess.run(tf.global_variables_initializer())
# 리스트를 이용해서 한번에 run할 수 있고, return을 받을 수도 있다.
for step in range(2001):
cost_val, w_val, b_val, _ = sess.run([cost, w, b, train], feed_dict={x:x_train, y:y_train})
if step%20==0:
print(step, cost_val, w_val, b_val)
# + id="lyyP8Otllgo3"
x_train = [1,2,3,4,5]
y_train = [2.1,3.1,4.1,5.1,6.1]
x = tf.placeholder(tf.float32)
y = tf.placeholder(tf.float32)
w = tf.Variable(tf.random_normal([1]), name='weight')
b = tf.Variable(tf.random_normal([1]), name='bias')
hypothesis = x_train * w + b
cost = tf.reduce_mean(tf.square(hypothesis - y_train))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.01)
# train = optimizer.minimize(cost) 이 코드 대신 아래 코드를 사용하면 gradient에 추가 작업을 진행할 수 있다.
gvs = optimizer.compute_gradients(cost)
train = optimizer.apply_gradients(gvs)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
for step in range(2001):
cost_val, w_val, b_val, _ = sess.run([cost, w, b, train], feed_dict={x:x_train, y:y_train})
if step%20==0:
print(step, cost_val, w_val, b_val)
# + [markdown] id="n8zki_EZmrbf"
# **multi variable**
# + id="hcVxzGAAmuze"
x_data = [[73., 50.,54.],[73., 50.,54.],[73., 50.,54.]]
y_data = [[73.],[50.],[54.]]
x = tf.placeholder(tf.float32, shape=[None,3])
y = tf.placeholder(tf.float32, shape=[None,1])
w = tf.Variable(tf.random_normal([3,1]), name='weight')
b = tf.Variable(tf.random_normal([1]), name='bias')
hypothesis = tf.matmul(x, w) + b
cost = tf.reduce_mean(tf.square(hypothesis-y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate=1e-5)
train = optimizer.minimize(cost)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
for step in range(2001):
cost_val, _ = sess.run([cost, train], feed_dict={x:x_data, y:y_data})
if step % 500 == 0:
print( step, cost_val )
# + [markdown] id="NrTln5QJqA57"
# # Tensor Manipulation
# + id="Rr3-OmycoKhl"
import numpy as np
t = tf.constant([1,2,3,4])
with tf.Session():
print(tf.shape(t).eval())
# + id="3u-uvkEUtRat"
t = tf.constant([[1,2],
[3,4]])
with tf.Session():
print(tf.shape(t).eval())
# + [markdown] id="igPJS3Rku3WS"
# **Broadcasting**
# + id="qVw7L_VAuMQR"
# broadcating에 의해 (1,2), (2,1)크기를 가지는 두 tensor의 크기가 모두 (2,2)로 확장되어 계산이 된다.
matrix1 = tf.constant([[1,2]])
matrix2 = tf.constant([[1],[2]])
with tf.Session():
print((matrix1+matrix2).eval())
# + [markdown] id="MqBmHuIQvck1"
# **Reduce mean**
# + id="Cg9uDpJOvCrX"
x=[[1.,10.],[3.,10.]]
with tf.Session():
print( tf.reduce_mean(x, axis=0).eval() )
print( tf.reduce_mean(x, axis=1).eval() )
print( tf.reduce_mean(x, axis=-1).eval()) # 가장 안쪽 축의 평균을 구하라
print( tf.reduce_mean(x).eval())
# + [markdown] id="iz_BsuTSwDeW"
# **Reduce sum**
# + id="3GwKoZz9vuwd"
x=[[1.,10.],[3.,10.]]
with tf.Session():
print( tf.reduce_sum(x, axis=0).eval() )
print( tf.reduce_sum(x, axis=1).eval() )
print( tf.reduce_sum(x, axis=-1).eval()) # 가장 안쪽 축의 합을 구하라
print( tf.reduce_sum(x).eval())
# + [markdown] id="pIuopgt-wUUP"
# Argmax
# + id="8fQF_ctiwID4"
x=[[1.,10.],[3.,10.]]
with tf.Session():
print( tf.argmax(x, axis=0).eval() )
print( tf.argmax(x, axis=1).eval() )
print( tf.argmax(x, axis=-1).eval()) # 가장 안쪽 축의 합을 구하라
# + [markdown] id="tgeWgVvQwdXk"
# **Reshape**
# + id="uP1GWSnpwYVr"
x=[[1.,10.],[3.,10.]]
with tf.Session():
print( tf.reshape(x, shape=[-1,4]).eval() )
# + id="bFdZABQvwqm7"
x=[[1.],[10.],[3.],[10.]]
with tf.Session():
print( tf.squeeze(x).eval() )
# + id="apCAkSQnw4_0"
x=[[1.,10.],[3.,10.]]
with tf.Session():
print( tf.expand_dims(x,1).eval() )
# + [markdown] id="qmTk50qBxND3"
# **One hot**
# + id="mEyrB-4JxE4n"
t = [[0],[1],[2],[0]]
with tf.Session():
t=tf.one_hot(t, depth=3) # depth는 class개수이다.
print( tf.reshape(t, shape=[-1,3]).eval() ) # one_hot을 쓸 경우 rank가 1증가하기 때문에 reshape
# + [markdown] id="bI4MXYRR4FPe"
# **Casting**
# + id="02lnwxH-xbOy"
with tf.Session():
print(tf.cast([1.5,2.2,2.4,5.6], tf.int32).eval())
print(tf.cast([True,False, 1==1, 0==1], tf.int32).eval())
# + [markdown] id="K0lkeyn94iKr"
# **stack**
# + id="3K_Kau6r4ND1"
x = [1,4]
y = [2,3]
z = [5,6]
with tf.Session():
print( tf.stack([x,y,z], axis=1).eval() )
# + [markdown] id="BJEQTTku4x9G"
# **Ones and Zeros like**
# + id="TW5yFCuv4ueG"
x=[[0,1,2],[3,4,5]]
with tf.Session():
print( tf.ones_like(x).eval())
print( tf.zeros_like(x).eval())
# + [markdown] id="Ow9suGfo5CT-"
# **Zip**
# + id="dj6V2TZq4_3H"
for x, y in zip([1,2,3],[4,5,6]):
print(x,y)
| tensorflow basic/tensorflow_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1 Write a Python Program to implement your own myreduce() function which works exactly like Python's built-in function reduce()
def myreduce(func, iterable):
"""
myreduce function is a user-built function version of Python built-in function reduce()
func : takes the function which needs to be applied on the iterable
iterable : list or any iterable object on which cumulative calculation can be performed
"""
it = iter(iterable)
result = next(it)
for element in it:
result = func(result, element)
return result
# +
import functools
l = [1,2,3,4,5]
print(functools.reduce(lambda a, b: a+b, l))
print(myreduce(lambda a, b: a+b, l))
# -
# 2 Write a Python program to implement your own myfilter() function which works exactly like Python's built-in function filter()
#
def myfilter(func, iterable):
"""
myfilter function is a user-built function version of Python built-in function filter()
func : takes the function which needs to be applied on the iterable
iterable : list or any iterable object on which cumulative calculation can be performed
"""
result = []
for i in iterable:
if func(i):
result.append(i)
return result
# +
seq = [0, 1, 2, 3, 5, 8, 13]
print(list(filter(lambda x: x % 2 != 0, seq)))
print(myfilter(lambda x: x % 2 != 0, seq))
# -
# 2. Implement List comprehensions to produce the following lists.
# Write List comprehensions to produce the following Lists :
#
# ['A', 'C', 'A', 'D', 'G', 'I', ’L’, ‘ D’]
#
# ['x', 'xx', 'xxx', 'xxxx', 'y', 'yy', 'yyy', 'yyyy', 'z', 'zz', 'zzz', 'zzzz']
#
# ['x', 'y', 'z', 'xx', 'yy', 'zz', 'xxx', 'yyy', 'zzz', 'xxxx', 'yyyy', 'zzzz']
#
# [[2], [3], [4], [3], [4], [5], [4], [5], [6]]
#
# [[2, 3, 4, 5], [3, 4, 5, 6], [4, 5, 6, 7], [5, 6, 7, 8]]
#
# [(1, 1), (2, 1), (3, 1), (1, 2), (2, 2), (3, 2), (1, 3), (2, 3), (3, 3)]
str1 = "ACADGILD"
str_list = [i for i in str1]
print(str_list)
str2 = ['x','y','z']
str_list = [i*n for i in str2 for n in range(1,5)]
print(str_list)
num_list = [2, 3, 4]
numsum_list = [[i+n] for i in num_list for n in range(0,3)]
print(numsum_list)
num_list = [2, 3, 4, 5]
numsum_list = [[i+n for i in num_list] for n in range(0,4)]
print(numsum_list)
pair_list = [1, 2, 3]
pairresult_list = [(b, a) for a in pair_list for b in pair_list]
print(pairresult_list)
| Python Assignment - 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Py3 research env
# language: python
# name: py3_research
# ---
# ## assignment 04: Decision Tree construction
# +
# If working in colab, uncomment the following line
# # ! wget https://raw.githubusercontent.com/girafe-ai/ml-mipt/basic_s20/homeworks_basic/assignment0_04_tree/tree.py -nc
# +
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
from sklearn.base import BaseEstimator
from sklearn.datasets import make_classification, make_regression, load_digits, load_boston
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.metrics import accuracy_score, mean_squared_error
import pandas as pd
# %load_ext autoreload
# %autoreload 2
# -
# Let's fix the `random_state` (a.k.a. random seed).
RANDOM_STATE = 42
# __Your ultimate task for today is to impement the `DecisionTree` class and use it to solve classification and regression problems.__
#
# __Specifications:__
# - The class inherits from `sklearn.BaseEstimator`;
# - Constructor is implemented for you. It has the following parameters:
# * `max_depth` - maximum depth of the tree; `np.inf` by default
# * `min_samples_split` - minimal number of samples in the leaf to make a split; `2` by default;
# * `criterion` - criterion to select the best split; in classification one of `['gini', 'entropy']`, default `gini`; in regression `variance`;
#
# - `fit` method takes `X` (`numpy.array` of type `float` shaped `(n_objects, n_features)`) and `y` (`numpy.array` of type float shaped `(n_objects, 1)` in regression; `numpy.array` of type int shaped `(n_objects, 1)` with class labels in classification). It works inplace and fits the `DecisionTree` class instance to the provided data from scratch.
#
# - `predict` method takes `X` (`numpy.array` of type `float` shaped `(n_objects, n_features)`) and returns the predicted $\hat{y}$ values. In classification it is a class label for every object (the most frequent in the leaf; if several classes meet this requirement select the one with the smallest class index). In regression it is the desired constant (e.g. mean value for `variance` criterion)
#
# - `predict_proba` method (works only for classification (`gini` or `entropy` criterion). It takes `X` (`numpy.array` of type `float` shaped `(n_objects, n_features)`) and returns the `numpy.array` of type `float` shaped `(n_objects, n_features)` with class probabilities for every object from `X`. Class $i$ probability equals the ratio of $i$ class objects that got in this node in the training set.
#
#
# __Small recap:__
#
# To find the optimal split the following functional is evaluated:
#
# $$G(j, t) = H(Q) - \dfrac{|L|}{|Q|} H(L) - \dfrac{|R|}{|Q|} H(R),$$
# where $Q$ is the dataset from the current node, $L$ and $R$ are left and right subsets defined by the split $x^{(j)} < t$.
#
#
#
# 1. Classification. Let $p_i$ be the probability of $i$ class in subset $X$ (ratio of the $i$ class objects in the dataset). The criterions are defined as:
#
# * `gini`: Gini impurity $$H(R) = 1 -\sum_{i = 1}^K p_i^2$$
#
# * `entropy`: Entropy $$H(R) = -\sum_{i = 1}^K p_i \log(p_i)$$ (One might use the natural logarithm).
#
# 2. Regression. Let $y_l$ be the target value for the $R$, $\mathbf{y} = (y_1, \dots, y_N)$ – all targets for the selected dataset $X$.
#
# * `variance`: $$H(R) = \dfrac{1}{|R|} \sum_{y_j \in R}(y_j - \text{mean}(\mathbf{y}))^2$$
#
# * `mad_median`: $$H(R) = \dfrac{1}{|R|} \sum_{y_j \in R}|y_j - \text{median}(\mathbf{y})|$$
#
#
# **Hints and comments**:
#
# * No need to deal with categorical features, they will not be present.
# * Siple greedy recursive procedure is enough. However, you can speed it up somehow (e.g. using percentiles).
# * Please, do not copy implementations available online. You are supposed to build very simple example of the Decision Tree.
# File `tree.py` is waiting for you. Implement all the needed methods in that file.
# ### Check yourself
from tree import entropy, gini, variance, mad_median, DecisionTree
# #### Simple check
# +
X = np.ones((4, 5), dtype=float) * np.arange(4)[:, None]
y = np.arange(4)[:, None] + np.asarray([0.2, -0.3, 0.1, 0.4])[:, None]
class_estimator = DecisionTree(max_depth=10, criterion_name='gini')
(X_l, y_l), (X_r, y_r) = class_estimator.make_split(1, 1., X, y)
assert np.array_equal(X[:1], X_l)
assert np.array_equal(X[1:], X_r)
assert np.array_equal(y[:1], y_l)
assert np.array_equal(y[1:], y_r)
# -
# #### Classification problem
digits_data = load_digits().data
digits_target = load_digits().target[:, None] # to make the targets consistent with our model interfaces
X_train, X_test, y_train, y_test = train_test_split(digits_data, digits_target, test_size=0.2, random_state=RANDOM_STATE)
assert len(y_train.shape) == 2 and y_train.shape[0] == len(X_train)
class_estimator = DecisionTree(max_depth=10, criterion_name='gini')
class_estimator.fit(X_train, y_train)
ans = class_estimator.predict(X_test)
accuracy_gini = accuracy_score(y_test, ans)
print(accuracy_gini)
reference = np.array([0.09027778, 0.09236111, 0.08333333, 0.09583333, 0.11944444,
0.13888889, 0.09930556, 0.09444444, 0.08055556, 0.10555556])
class_estimator = DecisionTree(max_depth=10, criterion_name='entropy')
class_estimator.fit(X_train, y_train)
ans = class_estimator.predict(X_test)
accuracy_entropy = accuracy_score(y_test, ans)
print(accuracy_entropy)
assert 0.84 < accuracy_gini < 0.9
assert 0.86 < accuracy_entropy < 0.9
assert np.sum(np.abs(class_estimator.predict_proba(X_test).mean(axis=0) - reference)) < 1e-4
# Let's use 5-fold cross validation (`GridSearchCV`) to find optimal values for `max_depth` and `criterion` hyperparameters.
param_grid = {'max_depth': range(3,11), 'criterion_name': ['gini', 'entropy']}
gs = GridSearchCV(DecisionTree(), param_grid=param_grid, cv=5, scoring='accuracy', n_jobs=-2)
# %%time
gs.fit(X_train, y_train)
gs.best_params_
assert gs.best_params_['criterion_name'] == 'entropy'
assert 6 < gs.best_params_['max_depth'] < 9
plt.figure(figsize=(10, 8))
plt.title("The dependence of quality on the depth of the tree")
plt.plot(np.arange(3,11), gs.cv_results_['mean_test_score'][:8], label='Gini')
plt.plot(np.arange(3,11), gs.cv_results_['mean_test_score'][8:], label='Entropy')
plt.legend(fontsize=11, loc=1)
plt.xlabel("max_depth")
plt.ylabel('accuracy')
plt.show()
# #### Regression problem
regr_data = load_boston().data
regr_target = load_boston().target[:, None] # to make the targets consistent with our model interfaces
RX_train, RX_test, Ry_train, Ry_test = train_test_split(regr_data, regr_target, test_size=0.2, random_state=RANDOM_STATE)
regressor = DecisionTree(max_depth=10, criterion_name='mad_median')
regressor.fit(RX_train, Ry_train)
predictions_mad = regressor.predict(RX_test)
mse_mad = mean_squared_error(Ry_test, predictions_mad)
print(mse_mad)
regressor = DecisionTree(max_depth=10, criterion_name='variance')
regressor.fit(RX_train, Ry_train)
predictions_mad = regressor.predict(RX_test)
mse_var = mean_squared_error(Ry_test, predictions_mad)
print(mse_var)
assert 9 < mse_mad < 20
assert 8 < mse_var < 12
param_grid_R = {'max_depth': range(2,9), 'criterion_name': ['variance', 'mad_median']}
gs_R = GridSearchCV(DecisionTree(), param_grid=param_grid_R, cv=5, scoring='neg_mean_squared_error', n_jobs=-2)
gs_R.fit(RX_train, Ry_train)
gs_R.best_params_
assert gs_R.best_params_['criterion_name'] == 'mad_median'
assert 3 < gs_R.best_params_['max_depth'] < 7
var_scores = gs_R.cv_results_['mean_test_score'][:7]
mad_scores = gs_R.cv_results_['mean_test_score'][7:]
plt.figure(figsize=(10, 8))
plt.title("The dependence of neg_mse on the depth of the tree")
plt.plot(np.arange(2,9), var_scores, label='variance')
plt.plot(np.arange(2,9), mad_scores, label='mad_median')
plt.legend(fontsize=11, loc=1)
plt.xlabel("max_depth")
plt.ylabel('neg_mse')
plt.show()
| homeworks_basic/assignment0_04_tree/assignment0_04_decision_tree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from functions_questions import add_to_file
from functions_variable_overview import *
# ## Create Table
# Read in description table
table = pd.read_csv("tables/covid_variable_description.csv", sep=";")
# Rename columns so they are easier to handle for the functions
table = table.rename(
columns={
"L_CoronavirusImpact_wave3_4p.dta": "wave#3",
"L_CoronavirusImpact_4.0p.dta": "wave#2",
"L_CoronavirusImpact_3p.dta": "wave#1",
"L_CoronavirusImpact_wave4_4p.dta": "wave#4",
"L_CoronavirusImpact_wave5_3p.dta": "wave#5",
"new_name": "variable",
"topic_english": "Topic",
"type": "Type",
"new_name": "Variable",
}
)
table.head()
# Select wave and language indicator.
# - `language` should be one lower-case letter.
# - `waves` should be a list of one or more values.
waves = [1,2,3,4,5]
language_short = "e"
language_long = "english"
# Create decription table and save to csv. If the table only includes one wave, it is saved to the respective language folder. If it is created for multiple waves, it is saved to "/docs/source/".
if len(waves) == 1:
variable_table = create_description_table(table, waves, language_short)
path = f"../docs/source/wave-{waves[0]}/"
else:
variable_table = create_description_table(
table, waves, language_short, routing=False
)
# Update path for multiple-wave overview:
path = f"../docs/source/"
variable_table
if language_short == "e":
variable_table.to_csv(
f"{path}variable_table-waves-{'-'.join(str(i) for i in waves)}-{language_short}.csv",
sep=",",
index=False,
)
# ## Create index file for questions sorted according to topics
if len(waves) == 1:
topic_table = create_description_table(
table, waves, language_short, return_links_as_symbols=False, routing=False
)
topic_dict = topic_table.groupby("Topic")[f"Links Wave {waves[0]}"].unique()
for key in ["skip", "Skip"]:
if key in topic_dict.keys():
topic_dict = topic_dict.drop(key)
create_overview_page(topic_dict, waves[0], language_long, path)
| python/create-variable-overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ijHoIPRArufd"
# #The History of Artificial Intelligence
# 1. Will quantum computing give AI the next big push it needs to achieve general artificial intelligence?
# 2. What is the environmental impact of achieving general artificial intelligence?
# 3. If computer speed and processing speed start to level out, what will be the next big driver of AI development toward general intelligence? (Moores law fails)
#
# #How Far are we from Achieving Artificial General Intelligence
# 1. Will production artificial general intelligence be our generation's "production flying car." Will we be saying "wheres my general artificial intelligence" at the end of our life like so many are saying "wheres my flying car" at the end of theirs.
# 2. Should artificial general intelligence have rights, and should the value of their life to comparable to a human?
# 3. What type of push back from governments and public citizens can researchers expect to see at the advent of general artificial intelligence?
#
# #Humans Fascination with Artificial General Intelligence
# 1. Does narrow artificial intelligence need to become more developed and researched (Comprehensive AI Services) before we can more clearly define a path toward general artificial intelligence?
# 2. When AGI can transfer knowledge from one domain to another as humans can, will AGI be capable of performing multiple positions simultaneously? In contrast, a human would still be limited to one, for example, an AGI that can perform hiring duties, but can act as a human resources personnel also?
#
# #Risks from Artificial Intelligence
# 1. In regards to safety, if an AGI were to hurt someone physically, who would be held responsible? Would humans have to waive their rights to prosecute if they worked in an environment where an AGI could cause possible harm to them?
# 2. In regards to security, if a country wanted to develop an AGI for military purposes, would there be the same barrier of entry that exists for nuclear weapons, and what country should have the most say? How significant is a first-mover advantage in the development of regulation around AGI?
#
# #The American Public is Already Worried About AI Catastrophe
# 1. Is AGI advancement and its ability to potentially take jobs being used as a red herring to distract from other topics?
# 2. Is the public afraid of AGI because it has not seen the jobs it will create?
| module3-autoencoders/U4_S3_M3_Reading_Questions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Memanggil Dependencies
import pandas as pd
import numpy as np
import random
import sklearn
from numpy import ndarray
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.neural_network import MLPClassifier
RANDOM_SEED = 27
# +
# Load dataset
df = pd.read_csv('input/diabetesPreProcessed.csv')
# Mengambil data pada kolom Outcome ke variabel target sebagai numpy Array
dftarget = df['Outcome']
target = dftarget.to_numpy()
# Menghapus kolom Outcome untuk mengambil data seluruh features
df = df.drop(columns=['Outcome'])
# Seluruh data features disimpan ke dalam variabel data sebagai numpy Array
data = df.to_numpy()
# -
# Standarisasi Data
from sklearn.preprocessing import StandardScaler
s = StandardScaler()
data = s.fit_transform(data)
# +
# Split Data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(data, target, test_size=0.2, random_state=RANDOM_SEED)
# Mengubah data target menjadi array 2D
# y_train = y_train.reshape(-1, 1)
print(X_train.shape, X_test.shape, y_train.shape, y_test.shape)
# +
# Training Neural Network
import time
start = time.perf_counter()
mlp = MLPClassifier(hidden_layer_sizes=(16), activation='tanh', solver='sgd', learning_rate_init=0.02,
early_stopping=True, batch_size=25, max_iter=2000, random_state=RANDOM_SEED)
mlp.fit(X_train,y_train)
elapsed = time.perf_counter() - start
print('Elapsed %.3f seconds.' % elapsed)
# -
# Testing Neural Network
import time
# %time
predict_train = mlp.predict(X_train)
predict_test = mlp.predict(X_test)
# Confusion Matrix untuk set training
print(confusion_matrix(y_train,predict_train))
print(classification_report(y_train,predict_train))
# Confusion Matrix untuk set testing
print(confusion_matrix(y_test,predict_test))
print(classification_report(y_test,predict_test))
| Main ANN Sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import cv2
import matplotlib
import matplotlib.pyplot as plt
import matplotlib as mpimg
import numpy as np
from IPython.display import HTML
import os, sys
import glob
import moviepy
from moviepy.editor import VideoFileClip
from moviepy.editor import *
from IPython import display
from IPython.core.display import display
from IPython.display import Image
import pylab
import scipy.misc
# +
def region_of_interest(img):
mask = np.zeros(img.shape, dtype=np.uint8) #mask image
roi_corners = np.array([[(200,675), (1200,675), (700,430),(500,430)]],
dtype=np.int32) # vertisies seted to form trapezoidal scene
channel_count = 1#img.shape[2] # image channels
ignore_mask_color = (255,)*channel_count
cv2.fillPoly(mask, roi_corners, ignore_mask_color)
masked_image = cv2.bitwise_and(img, mask)
return masked_image
# +
def ColorThreshold(img): # Threshold Yellow anf White Colos from RGB, HSV, HLS color spaces
HSV = cv2.cvtColor(img, cv2.COLOR_RGB2HSV)
# For yellow
yellow = cv2.inRange(HSV, (20, 100, 100), (50, 255, 255))
# For white
sensitivity_1 = 68
white = cv2.inRange(HSV, (0,0,255-sensitivity_1), (255,20,255))
sensitivity_2 = 60
HSL = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
white_2 = cv2.inRange(HSL, (0,255-sensitivity_2,0), (255,255,sensitivity_2))
white_3 = cv2.inRange(img, (200,200,200), (255,255,255))
bit_layer = yellow | white | white_2 | white_3
return bit_layer
# +
from skimage import morphology
def SobelThr(img): # Sobel edge detection extraction
gray=img
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0,ksize=15)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1,ksize=15)
abs_sobelx = np.absolute(sobelx)
abs_sobely = np.absolute(sobely)
scaled_sobelx = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
scaled_sobely = np.uint8(255*abs_sobely/np.max(abs_sobely))
binary_outputabsx = np.zeros_like(scaled_sobelx)
binary_outputabsx[(scaled_sobelx >= 70) & (scaled_sobelx <= 255)] = 1
binary_outputabsy = np.zeros_like(scaled_sobely)
binary_outputabsy[(scaled_sobely >= 100) & (scaled_sobely <= 150)] = 1
mag_thresh=(100, 200)
gradmag = np.sqrt(sobelx**2 + sobely**2)
scale_factor = np.max(gradmag)/255
gradmag = (gradmag/scale_factor).astype(np.uint8)
binary_outputmag = np.zeros_like(gradmag)
binary_outputmag[(gradmag >= mag_thresh[0]) & (gradmag <= mag_thresh[1])] = 1
combinedS = np.zeros_like(binary_outputabsx)
combinedS[(((binary_outputabsx == 1) | (binary_outputabsy == 1))|(binary_outputmag==1)) ] = 1
return combinedS
# -
def combinI(b1,b2): ##Combine color threshold + Sobel edge detection
combined = np.zeros_like(b1)
combined[((b1 == 1)|(b2 == 255)) ] = 1
return combined
def prespectI(img): # Calculate the prespective transform and warp the Image to the eye bird view
src=np.float32([[728,475],
[1058,690],
[242,690],
[565,475]])
dst=np.float32([[1058,20],
[1058,700],
[242,700],
[242,20]])
M = cv2.getPerspectiveTransform(src, dst)
warped = cv2.warpPerspective(img, M, (1280,720), flags=cv2.INTER_LINEAR)
return (warped, M)
# +
def undistorT(imgorg): # Calculate Undistortion coefficients
nx =9
ny = 6
objpoints = []
imgpoints = []
objp=np.zeros((6*9,3),np.float32)
objp[:,:2]=np.mgrid[0:6,0:9].T.reshape(-1,2)
images=glob.glob('./camera_cal/calibration*.jpg')
for fname in images: # find corner points and Make a list of calibration images
img = cv2.imread(fname)
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (6,9),None)
# If found, draw corners
if ret == True:
imgpoints.append(corners)
objpoints.append(objp)
# Draw and display the corners
#cv2.drawChessboardCorners(img, (nx, ny), corners, ret)
return cv2.calibrateCamera(objpoints,imgpoints,gray.shape[::-1],None,None)
# +
def undistresult(img, mtx,dist): # undistort frame
undist= cv2.undistort(img, mtx, dist, None, mtx)
return undist
# +
def LineFitting(wimgun): #Fit Lane Lines
# Set minimum number of pixels found to recenter window
minpix = 20
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
histogram = np.sum(wimgun[350:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((wimgun, wimgun, wimgun))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]/2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
nwindows = 9
# Set height of windows
window_height = np.int(wimgun.shape[0]/nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = wimgun.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated for each window
leftx_current = leftx_base
rightx_current = rightx_base
# Set the width of the windows +/- margin
margin =80
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = wimgun.shape[0] - (window+1)*window_height
win_y_high = wimgun.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) & (nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) & (nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
# Fit a second order polynomial to each
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, wimgun.shape[0]-1, wimgun.shape[0] )
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
# Create an image to draw on and an image to show the selection window
# out_img = np.dstack((wimgun, wimgun, wimgun))*255
window_img = np.zeros_like(out_img)
# Color in left and right line pixels
out_img[nonzeroy[left_lane_inds], nonzerox[left_lane_inds]] = [255, 0, 0]
out_img[nonzeroy[right_lane_inds], nonzerox[right_lane_inds]] = [0, 0, 255]
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
# plt.xlim(0, 1280)
# plt.ylim(720, 0)
# plt.imshow(out_img)
# # plt.savefig("./output_images/Window Image"+str(n)+".png")
# plt.show()
# Generate a polygon to illustrate the search window area
# And recast the x and y points into usable format for cv2.fillPoly()
left_line_window1 = np.array([np.transpose(np.vstack([left_fitx-margin, ploty]))])
left_line_window2 = np.array([np.flipud(np.transpose(np.vstack([left_fitx+margin, ploty])))])
left_line_pts = np.hstack((left_line_window1, left_line_window2))
right_line_window1 = np.array([np.transpose(np.vstack([right_fitx-margin, ploty]))])
right_line_window2 = np.array([np.flipud(np.transpose(np.vstack([right_fitx+margin, ploty])))])
right_line_pts = np.hstack((right_line_window1, right_line_window2))
# Draw the lane onto the warped blank image
cv2.fillPoly(window_img, np.int_([left_line_pts]), (0,255, 0))
cv2.fillPoly(window_img, np.int_([right_line_pts]), (0,255, 0))
result = cv2.addWeighted(out_img, 1, window_img, 0.3, 0)
# plt.title("r")
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
# plt.xlim(0, 1280)
# plt.ylim(720, 0)
# plt.imshow(result)
# # plt.savefig("./output_images/Line Image"+str(n)+".png")
# plt.show()
# Define y-value where we want radius of curvature
# I'll choose the maximum y-value, corresponding to the bottom of the image
y_eval = np.max(ploty)
left_curverad = ((1 + (2*left_fit[0]*y_eval + left_fit[1])**2)**1.5) / np.absolute(2*left_fit[0])
right_curverad = ((1 + (2*right_fit[0]*y_eval + right_fit[1])**2)**1.5) / np.absolute(2*right_fit[0])
#print(left_curverad, right_curverad)
ym_per_pix = 30/720 # meters per pixel in y dimension
xm_per_pix = 3.7/700 # meters per pixel in x dimension
# Fit new polynomials to x,y in world space
left_fit_cr = np.polyfit(ploty*ym_per_pix, left_fitx*xm_per_pix, 2)
right_fit_cr = np.polyfit(ploty*ym_per_pix, right_fitx*xm_per_pix, 2)
# y_eval = np.max(ploty)
# # Calculate the new radias of curvature
left_curverad = ((1 + (2*left_fit_cr[0]*y_eval*ym_per_pix + left_fit_cr[1])**2)**1.5) / np.absolute(2*left_fit_cr[0])
right_curverad = ((1 + (2*right_fit_cr[0]*y_eval*ym_per_pix + right_fit_cr[1])**2)**1.5) / np.absolute(2*right_fit_cr[0])
# # left_curverad = ((1 + (2*left_fit[0]*y_eval + left_fit[1])**2)**1.5) / np.absolute(2*left_fit[0])
# # right_curverad = ((1 + (2*right_fit[0]*y_eval + right_fit[1])**2)**1.5) / np.absolute(2*right_fit[0])
camera_center=wimgun.shape[0]/2
# #lane_center = (right_fitx[719] + left_fitx[719])/2
car_position = (camera_center - (left_fitx[-1]+right_fitx[-1])/2)*xm_per_pix
# print(left_curverad1, right_curverad1, lane_offset)
return (left_fit, ploty,right_fit,left_curverad, right_curverad,car_position)
# Create an image to draw the lines on
def unwrappedframe(img,pm, Minv, left_fit,ploty,right_fit):
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
nonzero = img.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
warp_zero = np.zeros_like(pm).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (img.shape[1], img.shape[0]))
# Combine the result with the original image
return cv2.addWeighted(img, 1, newwarp, 0.3, 0)
| LaneDetect.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
'''Code for fine-tuning Inception V3 for a new task.
Start with Inception V3 network, not including last fully connected layers.
Train a simple fully connected layer on top of these.
'''
import numpy as np
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential, Model
from keras.layers import Dense, Activation, Flatten, Dropout
import keras.applications.inception_v3 as inception
N_CLASSES = 2
IMSIZE = (299, 299)
# TO DO:: Replace these with paths to the downloaded data.
# Training directory
train_dir = '/mnt/e/data/catdog/train'
# Testing directory
test_dir = '/mnt/e/data/catdog/validation'
train_dir = r'E:\workshare\Mind\A3\data\catdog\train'
test_dir = r"E:\workshare\Mind\A3\data\catdog\validation"
# Start with an Inception V3 model, not including the final softmax layer.
base_model = inception.InceptionV3(weights='imagenet')
print ('Loaded Inception model')
# -
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.9
set_session(tf.Session(config=config))
for layer in base_model.layers:
print(layer.output)
for layer in base_model.layers:
print(layer.name)
# +
# Turn off training on base model layers
for layer in base_model.layers:
layer.trainable = True
# k=base_model.get_layer('flatten').output
# Add on new fully connected layers for the output classes.
x = Dense(32, activation='relu')(base_model.get_layer('avg_pool').output)
x = Dropout(0.5)(x)
predictions = Dense(N_CLASSES, activation='softmax', name='predictions')(x)
model = Model(inputs=base_model.input, outputs=[predictions])
model.compile(loss='categorical_crossentropy', optimizer='sgd', metrics=['accuracy'])
# Show some debug output
print (model.summary())
print ('Trainable weights')
print (model.trainable_weights)
# +
# Data generators for feeding training/testing images to the model.
model.load_weights('catdog_pretrain.h5')
train_datagen = ImageDataGenerator(rescale=1./255)
train_generator = train_datagen.flow_from_directory(
train_dir, # this is the target directory
target_size=IMSIZE, # all images will be resized to 299x299 Inception V3 input
batch_size=32,
class_mode='categorical')
test_datagen = ImageDataGenerator(rescale=1./255)
test_generator = test_datagen.flow_from_directory(
test_dir, # this is the target directory
target_size=IMSIZE, # all images will be resized to 299x299 Inception V3 input
batch_size=32,
class_mode='categorical')
model.fit_generator(
train_generator,
steps_per_epoch = 5,
epochs = 5000,
validation_data = test_generator,
verbose = 2,
validation_steps = 1)
model.save_weights('catdog_pretrain.h5') # always save your weights after training or during training
# img_path = '../data/sport3/validation/hockey/img_2997.jpg'
# img = image.load_img(img_path, target_size=IMSIZE)
# x = image.img_to_array(img)
# x = np.expand_dims(x, axis=0)
# x = inception.preprocess_input(x)
# preds = model.predict(x)
# print('Predicted:', preds)
# +
model.load_weights('catdog_pretrain.h5')
img_path = '../data/dog1.jpg'
img = image.load_img(img_path, target_size=IMSIZE)
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = inception.preprocess_input(x)
preds = model.predict(x)
print('Predicted:', preds)
| project/code/playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
from shutil import copyfile
path = '/home/mahmood1/matteo/pytorch-cnn-visualizations/results'
dest_path = '/home/mahmood1/matteo/pytorch-cnn-visualizations/jana_test_results'
for img in os.listdir(path):
layer = img.split('_')[1]
img_path = os.path.join(path, img)
current_dest_path = os.path.join(dest_path, 'layer_' + str(layer))
os.makedirs(current_dest_path, exist_ok = True)
current_dest_path = os.path.join(current_dest_path, img)
copyfile(img_path, current_dest_path)
os.remove(img_path)
| src/organizing_data_jana.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">5.3 Essential Functionality Part II
# <b>Function Application and Mapping
import pandas as pd
import numpy as np
# NumPy ufuncs (element-wise array methods) also work with pandas objects
frame = pd.DataFrame(np.random.randn(4, 3), columns=list('bde'),
index=['Utah', 'Ohio', 'Texas', 'Oregon'])
frame
np.abs(frame)
# Another frequent operation is applying a function on one-dimensional arrays to eachcolumn or row. DataFrame’s apply method does exactly this
f = lambda x: x.max() - x.min()
frame.apply(f)
frame.apply(f, axis=1)
# Element-wise Python functions can be used, too.
#
# Suppose you wanted to compute aformatted string from each floating-point value in frame. You can do this with applymap
format = lambda x: '%.2f' % x
frame.applymap(format)
# The reason for the name applymap is that Series has a map method for applying anelement-wise function
frame['e'].map(format)
# <b>Sorting and Ranking
# To sort lexicographically by row or column index, use the sort_index method, which returns a new sorted object
obj = pd.Series(range(4), index=['d', 'a', 'b', 'c'])
obj.sort_index()
frame = pd.DataFrame(np.arange(8).reshape((2, 4)),
index=['three', 'one'],
columns=['d', 'a', 'b', 'c'])
frame
frame.sort_index()
frame.sort_index(axis=1)
frame.sort_index(axis=1, ascending=False)
# To sort a Series by its values, use its sort_values method,NaN values are placed at last
obj.sort_values()
frame.sort_values(by='b',ascending=False)
frame.sort_values(by=['d','b'])
# Ranking assigns ranks from one through the number of valid data points in an array.
#
# The rank methods for Series and DataFrame are the place to look.
#
# By default rankbreaks ties by assigning each group the mean rank
obj_x=pd.Series([1,2,1,5,4,8,9,5,6])
obj_x.rank()
obj_x.rank(method='dense')
# DataFrame can compute ranks over the rows or the columns
frame
frame.rank(method='max',axis=0) #method does not work for dataframe they are only ranked by axis
# <b>Axis Indexes with Duplicate Labels
obj = pd.Series(range(5), index=['a', 'a', 'b', 'b', 'c'])
obj['a']
| Python-For-Data-Analysis/Chapter 5 Pandas Basics/5.3 Essential Functionality Part II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''nc2zarr'': conda)'
# name: python3
# ---
# +
#Notes to self: Run nc2zarr -c examples/nc2zarr-martin.yml on command line with conda nc2zarr activated first
#nc2zarr is taking care of the chunking - may be more efficient if we specify ourselves z = zarr.array(data, chunks=(1000, 1000), filters=filters, compressor=compressor)
#We are using directory store - GCSMap is supported: https://gcsfs.readthedocs.io/en/latest/api.html#gcsfs.mapping.GCSMap
#Questions
#1) Can we filter? - yes, like numpy filtering - would need to get indexs fro Lat,Lon and apply to melt array - must be a function for this?
#2) Understand access, and format? - done ish
#3) Lat/Lon change to hack? # easy to hack modify - see below
#4) Check with Gunner API - fine should work
#5) Do on Alex data in Unity
# -
import zarr
z1 = zarr.open('test_melt.zarr','r')
z1.info
print(z1.melt.chunks)
print(z1.melt)
# +
#View chunking split
z2 = z1.melt
for i in range(0,len(z2.chunks)):
print(z2.shape[i]/z2.chunks[i])
# -
#n5 store messing
store = zarr.N5Store('test_melt.n5')
root = zarr.group(store=store)
n1 = zarr.open('test_melt.n5','r')
# +
from_store = z1.store
to_store = zarr.ZipStore('test_melt.zip',mode='w')
#to_store = zarr.N5Store('test_melt.n5') #n5 doesn't work for some reason
zarr.copy_store(from_store,to_store)
# -
#Test updating values - can hack the lat, lon however we see fit
z1 = zarr.open('test_melt.zarr','a')
z1.lat[0,0]=-100
# +
zz1 = zarr.open('test_melt.zarr','r')
zz1.lat[0]
| data/explore.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Sample Classes in Python
class Car:
def __init__(self, color, model, year):
self.color = color
self.model = model
self.year = year
# ## Object Attributes
# +
my_car = Car("yellow", "beetle", 1967)
print(f"My car is {my_car.color}")
# -
# ## Add Attributes
# +
my_car.wheels = 5
print(f"Wheels: {my_car.wheels}")
# -
dir(my_car)
# In Python, when you declare a variable outside of a method, it’s treated as a class variable. Update the Car class as follows:
class Car:
wheels = 0
def __init__(self, color, model, year):
self.color = color
self.model = model
self.year = year
# This changes how you use the variable wheels. Instead of referring to it using an object, you refer to it using the class name:
# +
my_car = Car("yellow", "beetle", 1967)
print(f"My car is {my_car.color}")
print(f"It has {Car.wheels} wheels")
print(f"It has {my_car.wheels} wheels")
# -
# but be careful. Changing the value of the instance variable my_car.wheels will not change the value of the class variable Car.wheels:
# +
my_car = Car("yellow", "Beetle", "1966")
my_other_car = Car("red", "corvette", "1999")
print(f"My car is {my_car.color}")
print(f"It has {my_car.wheels} wheels")
print(f"My other car is {my_other_car.color}")
print(f"It has {my_other_car.wheels} wheels")
# +
# Change the class variable value
Car.wheels = 4
print(f"My car has {my_car.wheels} wheels")
print(f"My other car has {my_other_car.wheels} wheels")
# +
# Change the instance variable value for my_car
my_car.wheels = 5
print(f"My car has {my_car.wheels} wheels")
print(f"My other car has {my_other_car.wheels} wheels")
# -
# You define two Car objects on lines 2 and 3:
#
# my_car
# my_other_car
#
# At first, both of them have zero wheels. When you set the class variable using car.Car.wheels = 4 , both objects now have four wheels. However, when you set the instance variable using my_car.wheels = 5, only that object is affected.
#
# This means that there are now two different copies of the wheels attribute:
#
# A class variable that applies to all Car objects
# A specific instance variable applicable to the my_car object only
#
# Everything in Python is public. This code works with your existing Python class just fine:
# +
my_car = Car("blue", "Ford", 1972)
# Paint the car, No error
my_car.color = "red"
# -
# Instead of private, Python has a notion of a non-public instance variable. Any variable which starts with an underscore character is defined to be non-public. This naming convention makes it harder to access a variable, but it’s only a naming convention, and you can still access the variable directly.
#
# Add the following line to your Python Car class:
class Car:
wheels = 0
def __init__(self, color, model, year):
self.color = color
self.model = model
self.year = year
self._cupholders = 6
my_car = Car("yellow", "Beetle", "1969")
print(f"It was built in {my_car.year}")
# You can access the ._cupholders variable directly:
my_car.year = 1966
print(f"It was built in {my_car.year}")
print(f"It has {my_car._cupholders} cupholders.")
# Python further recognizes using double underscore characters in front of a variable to conceal an attribute in Python. When Python sees a double underscore variable, it changes the variable name internally to make it difficult to access directly. This mechanism avoids accidents but still doesn’t make data impossible to access.
#
# To show this mechanism in action, change the Python Car class again:
class Car:
wheels = 0
def __init__(self, color, model, year):
self.color = color
self.model = model
self.year = year
self.__cupholders = 6
# Now, when you try to access the .__cupholders variable, you see the following error:
my_car = Car("yellow", "Beetle", "1969")
print(f"It was built in {my_car.year}")
print(f"It has {my_car.__cupholders} cupholders.")
# So why doesn’t the .__cupholders attribute exist?
#
# When Python sees an attribute with double underscores, it changes the attribute by prefixing the original name of the attribute with an underscore, followed by the class name. To use the attribute directly, you need to change the name you use as well:
print(f"It has {my_car._Car__cupholders} cupholders")
# When you use double underscores to conceal an attribute from the user, Python changes the name in a well-documented manner. This means that a determined developer can still access the attribute directly.
# ## Access Control
# you access attributes directly in Python. Since everything is public, you can access anything at any time from anywhere. You set and get attribute values directly by referring to their names. You can even delete attributes in Python, which isn’t possible in Java:
# +
my_car = Car("yellow", "beetle", 1969)
print(f"My car was built in {my_car.year}")
my_car.year = 2003
print(f"It was built in {my_car.year}")
del my_car.year
print(f"It was built in {my_car.year}")
# -
# However, there are times you may want to control access to an attribute. In that case, you can use Python properties.
#
# In Python, properties provide controllable access to class attributes using Python decorator syntax.Properties allow functions to be declared in Python classes that are analogous to Java getter and setter methods, with the added bonus of allowing you to delete attributes as well.
#
# You can see how properties work by adding one to your Car class:
class Car:
def __init__(self, color, model, year):
self.color = color
self.model = model
self.year = year
self._voltage = 12
@property
def voltage(self):
return self._voltage
@voltage.setter
def voltage(self, volts):
print("Warning: this can cause problems!")
self._voltage = volts
@voltage.deleter
def voltage(self):
print("Warning: the radio will stop working!")
del self._voltage
# Here, you expand the notion of the Car to include electric vehicles. You declare the ._voltage attribute to hold the battery voltage
#
# To provide controlled access, you define a function called voltage() to return the private value. By using the @property decoration, you mark it as a getter that anyone can access directly
#
# Similarly, you define a setter function, also called voltage(). However, you decorate this function with @voltage.setter. Lastly, you use @voltage.deleter to decorate a third voltage(), which allows controlled deletion of the attribute.
#
# The names of the decorated functions are all the same, indicating they control access to the same attribute. The function names also become the name of the attribute you use to access the value. Here’s how these properties work in practice:
# +
my_car = Car("yellow", "beetle", 1969)
print(f"My car uses {my_car.voltage} volts")
my_car.voltage = 6
print(f"My car now uses {my_car.voltage} volts")
del my_car.voltage
# -
# Note that you use .voltage in the highlighted lines above, not ._voltage. This tells Python to use the property functions you defined:
#
# When you print the value of my_car.voltage , Python calls .voltage() decorated with @property.
# When you assign a value to my_car.voltage, Python calls .voltage() decorated with @voltage.setter.
# When you delete my_car.voltage, Python calls .voltage() decorated with @voltage.deleter.
#
# The @property, @.setter, and @.deleter decorations make it possible to control access to attributes without requiring users to use different methods. You can even make attributes appear to be read-only properties by omitting the @.setter and @.deleter decorated functions.
# ## Inheritance and Polymorphism
# Inheritance and polymorphism are two fundamental concepts in object-oriented programming.
#
# Inheritance allows objects to derive attributes and functionality from other objects, creating a hierarchy moving from more general objects to more specific. For example, a Car and a Boat are both specific types of Vehicles. Objects can inherit their behavior from a single parent object or multiple parent objects, and are referred to as child objects when they do.
#
# Polymorphism allows two or more objects to behave like one another, which allows them to be used interchangeably. For example, if a method or function knows how to paint a Vehicle object, then it can also paint a Car or Boat object, since they inherit their data and behavior from the Vehicle.
#
# These fundamental OOP concepts are implemented quite differently in Python vs Java.
# ## Inheritance
# Python supports multiple inheritance, or creating classes that inherit behavior from more than one parent class.
#
# To see how this works, update the Car class by breaking it into two categories, one for vehicles, and one for devices that use electricity:
# +
class Vehicle:
def __init__(self, color, model):
self.color = color
self.model = model
class Device:
def __init__(self):
self._voltage = 12
class Car(Vehicle, Device):
def __init__(self, color, model, year):
Vehicle.__init__(self, color, model)
Device.__init__(self)
self.year = year
@property
def voltage(self):
return self._voltage
@voltage.setter
def voltage(self, volts):
print("Warning: this can cause problems!")
self._voltage = volts
@voltage.deleter
def voltage(self):
print("Warning: the radio will stop working!")
del self._voltage
def __str__(self):
return f'Car {self.color} : {self.model} : {self.year}'
# -
# A Vehicle is defined as having .color and .model attributes. Then, a Device is defined to have a ._voltage attribute. Since the original Car object had these three attributes, it can be redefined to inherit both the Vehicle and Device classes. The color, model, and _voltage attributes will be part of the new Car class.
#
# In the .__init__() for Car, you call the .__init__() methods for both of the parent classes to make sure everything is initialized properly. Once done, you can add any other functionality you want to your Car. In this case, you add a .year attribute specific to Car objects, and getter and setter methods for .voltage.
#
# Functionally, the new Car class behaves as it always has. You create and use Car objects just as before:
# +
my_car = Car("yellow", "beetle", 1969)
print(f"My car is {my_car.color}")
print(f"My car uses {my_car.voltage} volts")
my_car.voltage = 6
print(f"My car now uses {my_car.voltage} volts")
# -
# ## Type Checking
# +
def charge(device):
if hasattr(device, '_voltage'):
print(f"Charging a {device._voltage} volt device")
else:
print(f"I can't charge a {device.__class__.__name__}")
class Phone(Device):
pass
class Rhino:
pass
my_car = Car("yellow", "Beetle", "1966")
my_phone = Phone()
my_rhino = Rhino()
charge(my_car)
charge(my_phone)
charge(my_rhino)
# -
# charge() must check for the existence of the ._voltage attribute in the object it’s passed. Since the Device class defines this attribute, any class that inherits from it (such as Car and Phone) will have this attribute, and will therefore show they are charging properly. Classes that do not inherit from Device (like Rhino) may not have this attribute, and will not be able to charge (which is good, since charging rhinos can be dangerous).
# ## Default Methods
# Python provides similar functionality with a set of common dunder (short for “double underscore”) methods. Every Python class inherits these methods, and you can override them to modify their behavior.
#
# For string representations of an object, Python provides __repr__() and __str__(), which you can learn about in Pythonic OOP String Conversion: __repr__ vs __str__. The unambiguous representation of an object is returned by __repr__(), while __str__() returns a human readable representation.
# Python provides default implementations of these dunder methods:
# +
my_car = Car("yellow", "Beetle", "1966")
print(repr(my_car))
print(str(my_car))
# -
# Overriding the dunder method gave us a more readable representation of your Car. You may want to override the .__repr__() as well, as it is often useful for debugging.
#
# Python offers a lot more dunder methods. Using dunder methods, you can define your object’s behavior during iteration, comparison, addition, or making an object callable directly, among other things.
# ## Operator Overloading
# Operator overloading refers to redefining how Python operators work when operating on user-defined objects. Python’s dunder methods allow you to implement operator overloading, something that Java doesn’t offer at all.
#
# Modify your Python Car class with the following additional dunder methods:
class Car:
def __init__(self, color, model, year):
self.color = color
self.model = model
self.year = year
def __str__(self):
return f'Car {self.color} : {self.model} : {self.year}'
def __eq__(self, other):
return self.year == other.year
def __lt__(self, other):
return self.year < other.year
def __add__(self, other):
return Car(self.color + other.color,
self.model + other.model,
int(self.year) + int(other.year))
# Dunder Method Operator Purpose
# - __eq__ == Do these Car objects have the same year?
# - __lt__ < Which Car is an earlier model?
# - __add__ + Add two Car objects in a nonsensical way
# When Python sees an expression containing objects, it calls any dunder methods defined that correspond to operators in the expression. The code below uses these new overloaded arithmetic operators on a couple of Car objects:
my_car = Car("yellow", "Beetle", "1966")
your_car = Car("red", "Corvette", "1967")
print (my_car < your_car)
print (my_car > your_car)
print (my_car == your_car)
print (my_car + your_car)
# There are many more operators you can overload using dunder methods. They offer a way to enrich your object’s behavior in a way that Java’s common base class default methods don’t.
# ## Reflection
# ### Examining an Object’s Type
# Both languages have ways to test or check an object’s type.
#
# In Python, you use type() to display the type of a variable, and isinstance() to determine if a given variable is an instance or child of a specific class:
# +
print(type(my_car))
print(isinstance(my_car, Car))
print(isinstance(my_car, Device))
# -
# ### Examining an Object’s Attributes
# In Python, you can view every attribute and function contained in any object (including all the dunder methods) using dir(). To get the specific details of a given attribute or function, use getattr():
print(dir(my_car))
print(getattr(my_car, "__init__"))
# ## Calling Methods Through Reflection
# Both Java and Python provide mechanisms to call methods through reflection.
# However, since Python doesn’t differentiate between functions and attributes, you have to look specifically for entries that are callable:
for method_name in dir(my_car):
if callable(getattr(my_car, method_name)):
print(method_name)
# The code below will find an object’s .__str__() and call it through reflection:
for method_name in dir(my_car):
attr = getattr(my_car, method_name)
if callable(attr):
if method_name == '__str__':
print(attr())
# Here, every attribute returned by dir() is checked. You get the actual attribute object using getattr(), and check if it’s a callable function using callable(). If so, you then check if its name is __str__(), and then call it.
| Class/OOP_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# language: python
# name: python3
# ---
# PROJETO - MÓDULO ESTATÍSTICA
#
# ALUNOS:
#
# <NAME>
#
# <NAME>
# ## CONTEXTUALIZAÇÃO
#
# A Autoridade De Estatística das Filipinas (PSA) lidera a realização da Pesquisa de Renda e Despesa Familiar (FIES) em todo o país. A pesquisa, realizada a cada 3 (três) anos, tem como objetivo fornecer dados sobre renda e despesas familiares, incluindo, entre outros, níveis de consumo por item de despesa, fontes de renda em dinheiro e informações relacionadas que afetam a renda e níveis de despesas e padrões nas Filipinas. Dessa forma, pede-se que seja elaborado um relatório prévio com insights e análise de dados por parte de um cientista de dados.
# +
# Importando as bibliotecas convencionais
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import plotly.express as px
import plotly.io as pio
# %matplotlib inline
# +
# importando bibliotecas específicas
from sklearn.linear_model import LinearRegression
import statsmodels.api as sm
from sklearn.metrics import r2_score, mean_squared_error
from scipy.stats import norm
from statsmodels.stats.weightstats import DescrStatsW, CompareMeans
from scipy.stats import pearsonr
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error as MSE
from sklearn.metrics import mean_squared_log_error
from sklearn.metrics import r2_score
# -
df = pd.read_csv('https://raw.githubusercontent.com/rodrigoviannini/Analises_Dados/main/Previsao%20de%20Renda%20com%20Regressao%20Linear%20-%20Filipinas/Dados_Renda_Despesa_Filipinas.csv')
df.head()
df.columns
df.info()
df.shape
# ## Milestone 1 - Amostra e Descrição
#
#
# 1. Calcule o tamanho de amostra ideal, considerando que você deseja estimar a média de dinheiro gasto com saúde (Medical Care Expenditure) com um erro máximo de 500, e um nível de confiança de 95%.
#
#
# +
## Calcular amostra ideal
# Média do $ gasto com saúde [Medical Care Expenditure]
# Erro Máximo = 500
# Confiança = 95%
sigma = df['Medical Care Expenditure'].std() #desvio padrão do meu gasto com saúde
maximum_error = 500
trust_level = 0.95
significance_level = (1 - trust_level)
# -
z_alpha_squared = norm.ppf(trust_level + (significance_level / 2))
z_alpha_squared
minimum_number = (z_alpha_squared * sigma / maximum_error) ** 2
minimum_number
ideal_sample = round(np.ceil(minimum_number))
ideal_sample
# 2. Obtenha uma amostra do dataset, considerando o tamanho da amostra que você calculou anteriormente e, em seguida, faça uma análise descritiva dos dados com, pelo menos, 10 variáveis quantitativas.
new_sample = df.sample(n=ideal_sample, random_state=101)
new_sample.head()
# +
## VARIÁVEIS QUANTITATIVAS:
# 1. renda familiar total, 2. despeza total de alimentos, 3. principal fonte de renda, 4. despesa com paes e cereais,
# 5. despeza total de arroz, 6. gastos com carne, 7. despesas de habitação e água, 8. despesas com frutas, 9. gastos com vegetais,
# 10. despesas com bebidas alcoolicas
df_ten_variables = df[['Total Household Income', 'Total Food Expenditure', 'Main Source of Income', 'Bread and Cereals Expenditure','Total Rice Expenditure', 'Meat Expenditure', 'Housing and water Expenditure', 'Vegetables Expenditure', 'Alcoholic Beverages Expenditure', ]]
df_ten_variables.head()
# -
# ## Milestone 2 - Inferência Estatística
#
#
# 1. Utilizando um nível de confiança de 95%, calcule o intervalo de confiança do dinheiro gasto com saúde e educação (colunas: Medical Care Expenditure e Education Expenditure).
# +
# Confiança = 95%
# Calcule o intervalo de confiança do $ gasto com saúde e educação
trust_level = 0.95
significance_level = (1 - trust_level) # alpha
average_health = new_sample['Medical Care Expenditure'].mean() # média de gastos com saúde
sample_health = len(new_sample['Medical Care Expenditure']) # Qtde de amostra de gastos com saúde
sigma_health = new_sample['Medical Care Expenditure'].std() # desvio padrão do meu gasto com saúde
average_education = new_sample['Education Expenditure'].mean() # média de gastos com saúde
sample_education = len(new_sample['Education Expenditure']) # Qtde de amostra de gastos com saúde
sigma_education = new_sample['Education Expenditure'].std() # desvio padrão do meu gasto com saúde
# +
# Intervalo de confiança de gastos com saúde
interval_trust_health = norm.interval(trust_level, loc=average_health, scale=(sigma_health / np.sqrt(sample_health)))
interval_trust_health
# +
# Intervalo de confiança de gastos com educação
interval_trust_education = norm.interval(trust_level, loc=average_education, scale=(sigma_education / np.sqrt(sample_education)))
interval_trust_education
# -
# 2. Calcule se há diferença de média de número total de membros da família pelo sexo do chefe de família (Household Head Sex e Total Number of Family members) com 95% de confiança.
#
#
# Há diferença de média de número total de membros da família pelo sexo do chefe de família?
new_sample['Household Head Sex'].value_counts()
# Chefe de família - Homem
male_householder = new_sample[new_sample['Household Head Sex'] == 'Male']
male_householder.shape[0]
# Chefe de família - Mulher
female_householder = new_sample[new_sample['Household Head Sex'] == 'Female']
female_householder.shape[0]
# Média e desvio padrão de homens como chefes de família
average_male = male_householder['Total Number of Family members'].mean()
sigma_male = male_householder['Total Number of Family members'].std()
average_male, sigma_male
# Média e desvio padrão de mulheres como chefes de família
average_female = female_householder['Total Number of Family members'].mean()
sigma_female = female_householder['Total Number of Family members'].std()
average_female, sigma_female
# +
# Calcular a diferença - confiança = 95%
trust_level = 0.95
significance_level = (1 - trust_level) # alpha
quantity_men = len(male_householder)
quantity_women = len(female_householder)
D0 = 0
# -
numerator = (average_male - average_female) - D0
denominator = np.sqrt((sigma_male ** 2 / quantity_men) + (sigma_female ** 2 / quantity_women))
Z = (numerator / denominator)
Z
p_value = norm.sf(Z) * 2
p_value
# Z de (alpha)²
z_alpha_squared = norm.ppf(trust_level + (significance_level / 2))
z_alpha_squared
# h0 -> Descartar: Z >= z_alpha_squared
Z >= z_alpha_squared
# h0 -> Descartar: p_value <= significance_level (alpha)
p_value <= significance_level
# H0 = m1 - m2 -> (average_male - average_female)
#
# ---
#
#
# H1 = m1 - m2! -> (average_male - average_female!)
# Testar
# Estatísticas descritivas e testes com pesos para pesos de caso
test_male = DescrStatsW(male_householder['Total Number of Family members'])
test_female = DescrStatsW(female_householder['Total Number of Family members'])
# Comparar
# classe para dois exemplos de comparação
to_compare_sex = CompareMeans(test_male, test_female)
# Z Test
# A função ztest.ind() testa a hipótese nula de que de que as duas distribuições possuem médias idênticas.
Z, p_value = to_compare_sex.ztest_ind()
Z, p_value
# - Z: Nas estatísticas, um escore-z (ou escore padrão) de uma observação é o número de desvios padrão acima ou abaixo da média da população. Para calcular um escore z, é necessário saber a média da população e o desvio padrão da população.
#
# ---
#
# - p-value: Na estatística clássica, o valor-p, é a probabilidade de se obter uma estatística de teste igual ou mais extrema que aquela observada em uma amostra, sob a hipótese nula. Por exemplo, em testes de hipótese, pode-se rejeitar a hipótese nula a 5% caso o valor-p seja menor que 5%
#
# - RESPOSTA: Há diferenças de média do número total de membros da família pelo sexo do householder
#
# +
# Calcular a diferença - confiança = 95%
trust_level = 0.95
significance_level = (1 - trust_level) # alpha
quantity_men = len(male_householder)
quantity_women = len(female_householder)
D0 = 0
# -
numerator = (average_male - average_female) - D0
denominator = np.sqrt((sigma_male ** 2 / quantity_men) + (sigma_female ** 2 / quantity_women))
Z = (numerator / denominator)
Z
p_value = norm.sf(Z) * 2
p_value
# Z de (alpha)²
z_alpha_squared = norm.ppf(trust_level + (significance_level / 2))
z_alpha_squared
# h0 -> Descartar: Z >= z_alpha_squared
Z >= z_alpha_squared
# h0 -> Descartar: p_value <= significance_level (alpha)
p_value <= significance_level
# 3. Verifique se há diferença de média total de renda da família (Total Household Income) de acordo com a fonte salarial (Main Source of Income).
# OBS: Considere “Enteroreneurial activities” e “other sources of income” como "outros".
# Visualizar todos os valores
new_sample['Main Source of Income'].value_counts() # Principal fonte de renda
# Substituir os nomes 'Other sources of Income', 'Enterpreneurial Activities' por 'other'
# Vizualizar modificação
new_sample['Main Source of Income'].replace(['Other sources of Income', 'Enterpreneurial Activities'], 'other', inplace = True)
new_sample['Main Source of Income'].value_counts()
wage_salaries = new_sample[new_sample['Main Source of Income'] == 'Wage/Salaries']
wage_salaries.head(2)
wage_salaries.shape[0]
other = new_sample[new_sample['Main Source of Income'] == 'other']
other.head(2)
other.shape[0]
test_income = DescrStatsW(wage_salaries['Total Household Income'])
test_total_income = DescrStatsW(other['Total Household Income'])
to_compare_income = CompareMeans(test_income, test_total_income)
Z, p_value = to_compare_income.ztest_ind()
Z, p_value
# - Z: Nas estatísticas, um escore-z (ou escore padrão) de uma observação é o número de desvios padrão acima ou abaixo da média da população. Para calcular um escore z, é necessário saber a média da população e o desvio padrão da população.
#
# ---
#
# - p-value: Na estatística clássica, o valor-p, é a probabilidade de se obter uma estatística de teste igual ou mais extrema que aquela observada em uma amostra, sob a hipótese nula. Por exemplo, em testes de hipótese, pode-se rejeitar a hipótese nula a 5% caso o valor-p seja menor que 5%
average_income_family_salaried = wage_salaries['Total Household Income'].mean()
sigma_income_family_salaried = wage_salaries['Total Household Income'].std()
average_income_family_salaried, sigma_income_family_salaried
average_total_family_income = other['Total Household Income'].mean()
sigma_total_family_income = other['Total Household Income'].std()
average_total_family_income, sigma_total_family_income
# H0 = m1 - m2 -> (average_income_family_salaried - average_total_family_income)
#
# ---
#
#
# H1 = m1 - m2! -> (average_income_family_salaried - average_total_family_income!)
# +
# Calcular a diferença - confiança = 95%
trust_level = 0.95
significance_level = (1 - trust_level) # alpha
quantity_wage_salaries = len(wage_salaries)
quantity_other = len(other)
D0 = 0
# -
numerator = (average_income_family_salaried - average_total_family_income) - D0
denominator = np.sqrt((sigma_income_family_salaried ** 2 / quantity_wage_salaries) + (sigma_total_family_income ** 2 / quantity_other))
Z = (numerator / denominator)
Z
p_value = norm.sf(Z) * 2
p_value
# Z de (alpha)²
z_alpha_squared = norm.ppf(trust_level + (significance_level / 2))
z_alpha_squared
# h0 -> Descartar: Z >= z_alpha_squared
Z >= z_alpha_squared
# h1 -> Descartar: p_value <= significance_level (alpha)
p_value <= significance_level
#
# - RESPOSTA: Há diferença de média entre rendas de famílias assalariadas e de famílias com outras rendas.
#
# ## Milestone 3 - Previsão de Renda Com Regressão Linear
#
# Usando Regressão Linear, crie um modelo de previsão de renda de forma a minimizar o erro. Considere 70% dos dados para treinar o modelo, e 30% para testá-lo. Por fim, utilize as métricas de avaliação do modelo que foram estudadas para validá-lo.
new_sample.info()
new_sample.shape
def correlations(dataframe, threshold):
""" Função que percorre um DataFrame filtrando os valores númericos, determinando o Coeficiencte de Correlação de Pearson de x (Renda familiar Total) e o y (colunas). Se este valor de Pearson for maior que o Limiar desejado adicionamos a Lista de Colunas à ser retornada """
features = []
for column in dataframe.columns:
if dataframe[column].dtype == 'int64':
pearson_correlation_coefficient, p_value = pearsonr(dataframe['Total Household Income'], dataframe[column])
if pearson_correlation_coefficient > threshold:
features.append(column)
return features
correlations(new_sample, 0.6)
# +
features = ['Total Food Expenditure',
'Clothing, Footwear and Other Wear Expenditure',
'Housing and water Expenditure',
'Imputed House Rental Value',
'Transportation Expenditure',
'Communication Expenditure',
'Miscellaneous Goods and Services Expenditure']
# Retirar: 'Imputed House Rental Value' (Valor imputado do aluguel da casa),
# Pois pode ser considerado valor duplicado, no item 'Housing and water Expenditure' (Despesas de habitação e água)
# Provavelmente o custo de aluguel já está incluso nos despesas com habitação e água
# -
X = new_sample[features]
X.shape
y = new_sample['Total Household Income']
y.shape
lr = LinearRegression().fit(X,y)
y_estimated = lr.predict(X)
lr.score(X,y)
r2_score(y,y_estimated)
x = sm.add_constant(X)
model = sm.OLS(y,x).fit()
model.summary()
# Média dos resíduos
model.resid.mean()
plt.figure(figsize=(8, 7))
sns.histplot(model.resid, kde=True);
plt.figure(figsize=(8, 7))
plt.scatter(y, model.resid);
plt.figure(figsize=(8, 7))
sns.heatmap(X.corr(), cmap='coolwarm', annot=True, vmin=-1, vmax=1);
plt.figure(figsize=(8, 10))
sns.jointplot(data=new_sample, x='Total Household Income', y='Imputed House Rental Value', kind='reg');
df.boxplot(column = features, figsize = (15,10));
plt. xticks(rotation=90);
# ## Retirando Outliers
def calc_min_and_max_range(data_features):
Q1 = data_features.quantile(q=0.25) # Primeiro quartil
Q3 = data_features.quantile(q=0.75) # Terceiro quartil
IQR = Q3 - Q1 # Intervalo interquartílico
low = Q1 - 1.5 * IQR
up = Q3 + 1.5 * IQR
return low, up
calc_min_and_max_range(new_sample)
# +
columns_to_remove_outliers = ['Total Household Income',
'Total Food Expenditure',
'Clothing, Footwear and Other Wear Expenditure',
'Housing and water Expenditure',
'Imputed House Rental Value',
'Transportation Expenditure',
'Communication Expenditure',
'Miscellaneous Goods and Services Expenditure']
for column in columns_to_remove_outliers:
low, up = calc_min_and_max_range(new_sample[features])
# Filtro para pegar apenas as linhas cujos dados estão dentro do intervalo aceitável (não outliers)
df = df[(new_sample[features] >= low) & (new_sample[features] <= up)]
# -
new_sample.shape
X_final = new_sample[['Total Food Expenditure',
'Clothing, Footwear and Other Wear Expenditure',
'Housing and water Expenditure',
'Imputed House Rental Value',
'Transportation Expenditure',
'Communication Expenditure',
'Miscellaneous Goods and Services Expenditure']]
X_final.head(2)
y_final = new_sample[['Total Household Income']]
y_final.head(2)
# Teste = 0.3
X_train, X_test, y_train, y_test = train_test_split(X_final, y_final, test_size=0.3, random_state=101)
X_train.shape
X_final.shape
X_test.shape
lr_final = LinearRegression().fit(X_train, y_train)
y_estimated_final = lr_final.predict(X_test)
r2_score(y_test, y_estimated_final)
lr.score(X_train, y_train)
# +
x = sm.add_constant(X_final)
model = sm.OLS(y_final, x).fit()
model.summary()
# -
# Média dos resíduos sem outliers
model.resid.mean()
plt.figure(figsize=(8, 7))
sns.histplot(model.resid, kde=True);
plt.figure(figsize=(8, 7))
plt.scatter(y_final, model.resid);
plt.figure(figsize=(8, 7))
sns.heatmap(X_final.corr(), cmap='coolwarm', annot=True, vmin=-1, vmax=1);
plt.figure(figsize=(8, 10))
sns.jointplot(data=new_sample, x='Total Household Income', y='Imputed House Rental Value', kind='reg');
r2_score(y_test, y_estimated_final)
mean_absolute_error(y_test, y_estimated_final)
MSE(y_test, y_estimated_final)
mean_squared_log_error(y_test, y_estimated_final)
| Previsao de Renda com Regressao Linear - Filipinas/Previsao_Renda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Simple acceptor
#
# - from http://pyfst.github.io/introduction.html
# - for more details see http://pyfst.github.io/
# +
import fst
a = fst.Acceptor()
a.add_arc(0, 1, 'py')
a[1].final = True
a
# +
b = fst.linear_chain(('fst'), syms=a.isyms)
b
# +
c = a.closure() + b
c
# -
# Determinization does not help in this case!
# +
c.determinize()
c
# -
# The error message shows unsupported operation when the FST is not deterministic.
# +
c.minimize()
c
| notebooks/simple_acceptor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MariaMalycha/dw_matrix/blob/master/Day5.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="gbKNd8PqbjGB" colab_type="code" outputId="19ad5f51-9452-47bc-824b-7843947a4476" colab={"base_uri": "https://localhost:8080/", "height": 228}
# !pip install eli5
# + id="x-mKRTvYbz1e" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import VarianceThreshold
import eli5
from eli5.sklearn import PermutationImportance
#dealing with dictionaries
from ast import literal_eval
from tqdm import tqdm_notebook
# + [markdown] id="amEbGvVRcF84" colab_type="text"
# # Modeling
# + id="iLT6NcjXd8sf" colab_type="code" outputId="01e160d3-8b14-4e39-ad83-36dce2e76a7b" colab={"base_uri": "https://localhost:8080/", "height": 35}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/"
# + id="B7c-qNkZeOnm" colab_type="code" outputId="b7e4f2eb-0726-42b6-a7cd-98291ec78dea" colab={"base_uri": "https://localhost:8080/", "height": 35}
# ls data
# + id="7Tps3KYkeRMv" colab_type="code" outputId="71c02a80-34ac-4430-d203-747cd8542d55" colab={"base_uri": "https://localhost:8080/", "height": 35}
df = pd.read_csv('data/men-shoes.csv', low_memory=False)
df.shape
# + id="dtCTpNr3fCWd" colab_type="code" outputId="0375e38c-2702-4357-e8bc-f96735f69ae8" colab={"base_uri": "https://localhost:8080/", "height": 225}
df.columns
# + id="EoOlkR_Kfyki" colab_type="code" colab={}
def run_model(feats, model = DecisionTreeRegressor(max_depth=5)):
X = df[ feats ].values
y = df.prices_amountmin.values
scores = cross_val_score(model, X, y, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="ZpyIrnARgkDu" colab_type="code" colab={}
df['brand_cat'] = df.brand.map(lambda x: str(x).lower()).factorize()[0]
# + id="sB_dIwdvhhvW" colab_type="code" outputId="a0cedb58-f4b0-4ea5-a080-dae6c53e456b" colab={"base_uri": "https://localhost:8080/", "height": 35}
run_model(['brand_cat'])
# + id="VY3JkafhhsvB" colab_type="code" outputId="69ab6cbd-e051-413a-f02b-d9b0d8e11b88" colab={"base_uri": "https://localhost:8080/", "height": 35}
model=RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
run_model(['brand_cat'], model)
# + id="fDvAYFvSnh0C" colab_type="code" outputId="227f1ddc-b802-491a-b9b7-034a48f9ea69" colab={"base_uri": "https://localhost:8080/", "height": 517}
df.head()
# + id="ZxpRwZ7uyVX2" colab_type="code" colab={}
def parse_features(x):
output_dict = {}
if str(x) == 'nan': return output_dict
features = literal_eval(x.replace('\\"', '"'))
#[{'key': 'Gender', 'value': ['Men']}
for item in features:
key = item['key'].lower().strip()
value = item['value'][0].lower().strip()
output_dict[key] = value
return output_dict
df['features_parsed'] = df['features'].map(parse_features)
# + id="mpq7Z9EM2XA5" colab_type="code" outputId="b9283c77-be0e-4ba5-c504-afe5bd7c8d1b" colab={"base_uri": "https://localhost:8080/", "height": 35}
keys = set()
df['features_parsed'].map(lambda x: keys.update(x.keys()))
len(keys)
# + id="xPnBBawH9KQx" colab_type="code" outputId="a6571873-84a2-4144-e57b-bd3d5129e3d4" colab={"base_uri": "https://localhost:8080/", "height": 67, "referenced_widgets": ["7102d15a03ad42d9ad9d1a67aadcb0ae", "1bfe8d51cb9f4e3cbb886831ad6d17f8", "<KEY>", "7c5cb8834e9943feb91178296e6109df", "5868c348de724d9bb5fe1611e9424ac8", "<KEY>", "23626699916c4ce3ab24bdabcc8a77fe", "<KEY>"]}
def get_name_feat(key):
return 'feat_' + key
for key in tqdm_notebook(keys):
df[get_name_feat(key)] = df.features_parsed.map(lambda feats: feats[key] if key in feats else np.nan)
# + id="2ZOF7MG8Cl6h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 139} outputId="4440dbe4-813f-4da7-80cb-a5a7014073c8"
df.columns
# + id="Dbwrbah48Okb" colab_type="code" colab={}
keys_stat = {}
for key in keys:
keys_stat[key] = df[ False == df[ get_name_feat(key) ].isnull() ].shape[0] / df.shape[0] * 100
# + id="4j6mQlUz8aUY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="2dc8b409-9473-4a88-d64f-543a31e04a0d"
{k:v for k,v in keys_stat.items() if v > 30}
# + id="pR9l-n7x81fp" colab_type="code" colab={}
'''
df['feat_brand_cat'] = df['feat_brand'].factorize()[0]
df['feat_color_cat'] = df['feat_color'].factorize()[0]
df['feat_gender_cat'] = df['feat_gender'].factorize()[0]
df['feat_manufacturer part number_cat'] = df['feat_manufacturer part number'].factorize()[0]
df['feat_material_cat'] = df['feat_material'].factorize()[0]
df['feat_sport_cat'] = df['feat_sport'].factorize()[0]
df['feat_style_cat'] = df['feat_style'].factorize()[0]
'''
for key in keys:
df[get_name_feat(key) + '_cat'] = df[get_name_feat(key)].factorize()[0]
# + id="7FLRigwDA4rG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="c9693c16-4f9c-4be0-aabc-e93bcd64de1f"
run_model(['brand_cat'])
# + id="QUIpkJV9BH7m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0ce22bec-cb16-405e-815d-b105e2a8de71"
df['brand'] = df['brand'].map( lambda x: str(x).lower() )
df[ df.brand != df.feat_brand ][ ['brand', 'feat_brand'] ].shape
# + id="xsi41t1OCqZq" colab_type="code" colab={}
feats = ['']
# + id="LTuVJ8JEC-6Y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d22edc1a-530b-4d45-b518-63d4dad1a92c"
model = RandomForestRegressor(max_depth=5, n_estimators=100)
feats = ['brand_cat']
run_model( feats, model)
# + id="mb0DBojvMcvW" colab_type="code" colab={}
#feats_cat = [x for x in df.columns if 'cat' in x]
feats_cat = [x for x in df.columns if '_cat' in x and x != 'feat_catalog']
# + id="FPGIbw6XFrTf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3d0ff8e2-cfba-49ee-9873-6f17c176a8c5"
feats = ['brand_cat'
,'feat_style_cat'
,'feat_sport_cat'
,'feat_brand_cat'
,'feat_gender_cat'
,'feat_material_cat']
#feats += feats_search
#feats = list(set(feats))
model = RandomForestRegressor(max_depth=5, n_estimators=100)
result_benchmark = run_model( feats, model)
result_benchmark
# + id="OaBYpKiDSsCV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="53a836bb-9048-4945-dd0e-d90edb263972"
print(len(feats_cat))
feats_search = feats_cat[301:400]
# + id="x7hktK82aEw4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="946c8c54-5bd6-4671-b396-4618b3df654f"
feats = ['brand_cat'
,'feat_age_cat'
,'feat_weight_cat'
,'feat_brand_cat'
,'feat_gender_cat'
,'feat_material_cat']
#feats += feats_search
feats = list(set(feats))
model = RandomForestRegressor(max_depth=5, n_estimators=100)
result = run_model( feats, model)
print(result_benchmark)
print(result)
# + id="QLzXabj1GcNY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="8d693866-abe7-49e0-c666-b53da98ee253"
X = df[ feats ].values
y = df['prices_amountmin'].values
m = RandomForestRegressor(max_depth=5, n_estimators=100, random_state=0)
m.fit(X,y)
print(result)
perm = PermutationImportance(m, random_state=1).fit(X, y);
eli5.show_weights(perm, feature_names=feats)
# + id="VR8MzYTwIs_e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="70e1a356-18a2-4538-e516-d02c08c11d09"
df['brand'].value_counts(normalize=True)
# + id="q1UhCq3kE8Wx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 141} outputId="59184822-d3fe-49f8-ffb3-72579d01e135"
df[ df['brand'] == 'nike'].features_parsed.sample(5).values
# + id="LUhfsEbeKxxe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="0a3ca8a4-1b15-4c28-85ed-7f73ad9b7d21"
df['feat_age group'].value_counts()
# + id="JabaYBh8JDkh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="7a292cb0-2505-4ffc-8474-4069d845e38e"
df.feat_age.value_counts()
# + id="c8jJYOpFrf18" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="6467d230-8f33-4598-cb62-40033d992511"
df.feat_material.value_counts()
# + id="q_L3a8r-s58w" colab_type="code" colab={}
'''
feats_search = ['brand_cat'
,'feat_shoe category_cat'
,'feat_style_cat'
,'feat_casual & dress shoe style_cat'
,'feat_gender_cat'
,'feat_brand_cat'
,'feat_inseam_cat'
, 'feat_material_cat'
,'feat_resizable_cat'
,'feat_shoe size_cat'
,'feat_condition_cat'
,'feat_weight_cat'
,'feat_bridge_cat'
,'feat_adjustable_cat'
,'feat_age_cat'
,'feat_color_cat'
,'feat_metal type_cat']
'''
| Day5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
data = pd.read_csv("./data/Visualization for company stakeholders.csv")
data.head()
# # Bar plot
loan_status = data['Loan_Status'].value_counts()
print(loan_status)
plt.xlabel('Loan Status')
plt.ylabel('Count for Loan Status')
plt.title('Bar plot for Loan Status')
loan_status.plot.bar(rot=0)
#
# # Bar plot(Unstack) with groupby
# +
property_and_loan = data.groupby(['Property_Area','Loan_Status'])
property_and_loan = property_and_loan.size().unstack()
property_and_loan.plot(kind='bar', stacked=False, rot=45)
plt.show()
# -
# # Bar plot(Stacked) with groupby
# +
education_and_loan = data.groupby(['Education','Loan_Status'])
education_and_loan = education_and_loan.size().unstack()
education_and_loan.plot(kind='bar',stacked=True, rot=45)
plt.xlabel("Education Status")
plt.ylabel("Loan Status")
# -
# # Density plot
# +
graduate = data[data['Education']=='Graduate']
not_graduate = data[data['Education']=='Not Graduate']
graduate['LoanAmount'].plot(kind='density', label='Graduate')
not_graduate['LoanAmount'].plot(kind='density', label='Not Graduate')
#For automatic legend display
plt.legend()
# -
# # Scatter plot
# +
fig,(ax_1,ax_2,ax_3) = plt.subplots(nrows=3, ncols=1, figsize = (8,10))
plt.subplots_adjust(hspace=1.35)
ax_1.scatter(data['LoanAmount'], data['ApplicantIncome'])
ax_1.set_title("Applicant Income")
ax_2.scatter(data['LoanAmount'] ,data['CoapplicantIncome'])
ax_2.set_title('Coapplicant Income')
data['TotalIncome'] = data['ApplicantIncome'] + data['CoapplicantIncome']
ax_3.scatter(data['LoanAmount'], data['TotalIncome'])
ax_3.set_title('Total Income')
# -
# This is an Analysis of the stake holders sentiment.
| Project - Visualization for company stakeholders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Method: Koopman Autoencoders
# # Dataset: Lorenz-96, F = 8
# # Purpose: Prediction
# # 1. Set-up
# +
# GPU
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "3"
# Package
import sys
sys.path.append("../..")
# +
from create_data import load_data
from utils import * # Number of testing samples
import numpy as np
import matplotlib.pyplot as plt
from time import time, localtime, strftime
from functools import partial
import jax
from jax import jit, value_and_grad
import jax.numpy as jnp
from jax import random
from jax.example_libraries import optimizers
import jax.example_libraries.stax as stax
from jax.example_libraries.stax import Dense, Relu, Tanh
from jax.nn.initializers import glorot_normal
# -
SEED = 42
train, test = load_data("Lorenz 96, F = 8", "../../data/lorenz8", 0.5)
train.data = train.data[:18000]
train.time = train.time[:18000]
print(f"Train size: {train.data.shape}")
print(f"Test size: {test.data.shape}")
# **Create test set**
L_forecast_test = 400 # steps to forecast forward (when testing)
# +
np.random.seed(1)
data_test = test.data
T_test, data_dim = data_test.shape
possible_idx = T_test - (L_forecast_test + 1) # minus number of steps forward, and the warm-up period
T_indices = np.random.randint(0, possible_idx, size = NUM_TEST)
t_past_batch = np.repeat(T_indices[:, None], WARM_UP_TEST, axis = 1).astype(int) # 200 warmup
t_pred_batch = (T_indices[:, None] + np.arange(1, 1 + L_forecast_test)[None, :].astype(int))
X_test = data_test[t_past_batch]
y_test = data_test[t_pred_batch]
# -
print(f"Test input size: {X_test.shape}") # Number of test points x input length x dim
print(f"Test output size: {y_test.shape}") # Number of test points x horizon x dim
# # 2. Koopman Autoencoder Implementation
def Dense_no_bias(out_dim, W_init = glorot_normal()):
"""Layer constructor function for a dense (fully-connected) layer."""
def init_fun(rng, input_shape):
output_shape = input_shape[:-1] + (out_dim,)
k1, k2 = random.split(rng)
W = W_init(k1, (input_shape[-1], out_dim))
return output_shape, W
def apply_fun(params, inputs, **kwargs):
W = params
return jnp.dot(inputs, W)
return init_fun, apply_fun
def get_params(hidden_state_list, max_lag, lambda_list, seed, batch_size, num_epoch, lr_schedule,
early_stopping = EARLY_STOPPING):
assert len(num_epoch) == len(lr_schedule)
def create_network():
encoder_init, encoder = stax.serial(
Dense(p1), Tanh,
Dense(p2), Tanh,
Dense(p3), Tanh,
Dense(kappa))
decoder_init, decoder = stax.serial(
Dense(p3), Tanh,
Dense(p2), Tanh,
Dense(p1), Tanh,
Dense(data_dim))
forward_init, forward = stax.serial(
Dense_no_bias(kappa))
backward_init, backward = stax.serial(
Dense_no_bias(kappa))
# initialization
enc_init_rng, dec_init_rng, forward_init_rng, backward_init_rng = random.split(key, num = 4)
enc_in_shape = (-1, data_dim)
dec_in_shape = (-1, kappa)
forward_in_shape = (-1, kappa)
backward_in_shape = (-1, kappa)
_, enc_params = encoder_init(enc_init_rng, enc_in_shape)
_, dec_params = decoder_init(dec_init_rng, dec_in_shape)
_, fwd_params = forward_init(forward_init_rng, forward_in_shape)
_, bwd_params = backward_init(backward_init_rng, backward_in_shape)
network = (encoder, decoder, forward, backward)
params = (enc_params, dec_params, fwd_params, bwd_params)
return network, params
def training(init_params):
@jit
def loss(params, batch):
"""
each batch has dimension Batch_size x (max_lag+1) x dim
"""
# unpack parameters
enc_params, dec_params, fwd_params, bwd_params = params
lambda_id, lambda_fwd, lambda_bwd, lambda_con = lambda_list
loss_id, loss_fwd, loss_bwd, loss_con = 0, 0, 0, 0
# AUTOENCODER
f_start = batch[:,0,:]
f_target = batch[:,0,:]
f_encoded = encoder(enc_params, f_start)
f_predict = decoder(dec_params, f_encoded)
loss_id = 0.5 * jnp.mean((f_predict - f_target)**2)
# FORWARD
f_start = batch[:,0,:]
f_encoded = encoder(enc_params, f_start)
for t_steps in range(1, max_lag+ 1):
# simulate forward
f_encoded = forward(fwd_params, f_encoded)
f_predict = decoder(dec_params, f_encoded)
f_target = batch[:,t_steps,:]
loss_fwd += 0.5 * jnp.mean((f_predict - f_target)**2)
loss_fwd = loss_fwd / max_lag
# BACKWARD
f_start = batch[:,-1,:]
f_encoded = encoder(enc_params, f_start)
for t_steps in range(1, max_lag + 1):
f_encoded = backward(bwd_params, f_encoded)
f_predict = decoder(dec_params, f_encoded)
f_target = batch[:,-t_steps-1,:]
loss_bwd += 0.5 * jnp.mean((f_predict - f_target)**2)
loss_bwd = loss_bwd / max_lag
# CONSISTENT
for k in range(1, kappa + 1):
C_upper = fwd_params[0][:k, :]
D_left = bwd_params[0][:, :k]
C_left = fwd_params[0][:, :k]
D_upper = bwd_params[0][:k, :]
I_k = jnp.identity(k)
loss_con += (jnp.sum((D_upper @ C_left - I_k)**2) + jnp.sum((C_upper @ D_left - I_k)**2)) / (2 * k)
return lambda_id * loss_id + lambda_fwd * loss_fwd + lambda_bwd * loss_bwd + lambda_con * loss_con
@jit
def step(i, opt_state, batch):
params = get_params(opt_state)
value, g = value_and_grad(loss)(params, batch)
opt_state = opt_update(i, g, opt_state)
return get_params(opt_state), opt_state, value
def make_forecast(params, batch, L_forecast):
enc_params, dec_params, fwd_params, bwd_params = params
preds = []
f_start = batch[:, 0, :]
f_encoded = encoder(enc_params, f_start)
for t in range(L_forecast):
f_encoded = forward(fwd_params, f_encoded)
f_predict = decoder(dec_params, f_encoded)
preds.append(f_predict)
return np.swapaxes(preds, 0, 1)
def mse(params, x_batch, y_truth):
"""
For each time-series in a batch, forecasts over a finite horizon
and compute the MSE. Primarily used for validation calculation
args:
====
params: neural parameters
x_batch: a batch of inputs with dimension (batch_size, T_past, dim_data)
y_truth: a batch of values to forecasts with dimension (batch_size, T_future, dim_data)
outputs:
=======
MSE: MSE between forecasts and targets
"""
# horizon of the forecast
L_forecast = y_truth.shape[1]
y_pred = make_forecast(params, x_batch, L_forecast_test)
#compute MSE
error = y_pred - y_truth
mu_loss = np.mean(error**2)
return mu_loss
start = time()
best_params = init_params
overall_best_mse = 99999999
# train/val split
t_size = int(0.9 * train_size)
v_size = train_size - t_size
T_indices_val = np.arange(t_size, train_size - (L_forecast_test + max_lag))
t_start_val = T_indices_val[::10]
t_past_batch_val = (t_start_val[:,None] + np.arange(max_lag)[None,:]).astype(int)
t_pred_batch_val = (t_start_val[:,None] + np.arange(max_lag, max_lag + L_forecast_test)[None,:]).astype(int)
x_val = x[t_past_batch_val]
y_val = y[t_pred_batch_val]
print("Backpropogation start", end = "\n\n")
for i, lr in enumerate(lr_schedule):
opt_init, opt_update, get_params = optimizers.adam(step_size = lr)
opt_state = opt_init(best_params)
counter = 0
best_mse = 999999999
for epoch in range(num_epoch[i]):
e_start = time()
# randomize the order of the data
T_indices = np.arange(t_size - max_lag - 1)
np.random.shuffle(T_indices)
# training
loss_epoch_train = []
for k in range(t_size // batch_size + 1):
# create a batch of data
t_start = T_indices[np.arange(k*batch_size, (k+1)*batch_size).astype(int) % len(T_indices)] # start of each time series in the batch
# create 2d array of dimension (batch_size, max_lag + 1) containing all the time indices
t_batch = (t_start[:,None] + np.arange(max_lag + 1)[None,:]).astype(int) # transposes data
#create batch of dimension (batch_size, max_lag + 1, data_dim)
x_batch = x[t_batch]
params, opt_state, loss_current = step(k, opt_state, x_batch)
loss_epoch_train.append(loss_current.item())
mse_train = np.mean(loss_epoch_train)
# validation
mse_val = mse(params, x_val, y_val)
if best_mse > mse_val: # Improvement
counter = 0
best_mse = mse_val
best_params = params
else:
counter += 1
e_end = time()
if (epoch + 1) % 10 == 0 or (counter == 0 and epoch >= 30):
print(f"Epoch {epoch + 1}: Time taken = {e_end - e_start:.2f} | Train loss = {mse_train:.7f} | Val loss = {mse_val: .7f}")
if counter >= early_stopping:
print(f"EARLY STOPPING. Epoch {epoch + 1}: Train loss = {mse_train:.7f} | Val loss = {mse_val: .7f}")
break
print(f"Best Validation MSE: {best_mse:.7f}")
if best_mse < overall_best_mse: # Best round so far
print("IMPROVED VALIDATION MSE")
overall_best_mse = best_mse
overall_best_params = best_params
print()
end = time()
print(f"Total time: {end - start:.2f}")
return overall_best_params
start = time()
x, y = train.data[:-1], train.data[1:]
train_size, data_dim = x.data.shape
p1, p2, p3, kappa = hidden_state_list # 4 layers (including bottleneck)
np.random.seed(seed)
key = jax.random.PRNGKey(seed)
(encoder, decoder, forward, backward), params = create_network()
final_params = training(params)
return final_params, (encoder, decoder, forward, backward)
def get_test_pred(data_test, params, network):
start = time()
(encoder, decoder, forward, backward) = network
enc_params, dec_params, fwd_params, bwd_params = params
num_data_test, L_past, data_dim = data_test.shape # testing ex, # steps used before, dim of data
preds = []
f_start = data_test[:, 0, :]
f_encoded = encoder(enc_params, f_start)
for t in range(L_forecast_test):
f_encoded = forward(fwd_params, f_encoded)
f_predict = decoder(dec_params, f_encoded)
preds.append(f_predict)
end = time()
print(f"Time taken: {end - start:.2f}")
return np.swapaxes(preds, 0, 1)
# # 3. Parameter tuning
#
# There are 3 parameters to tune
# - hidden_size - size of the hidden layer
# - max_lag - Number of steps calculated for forward and backward dynamics
# - loss_weights - [encoder, forward, backward, consistent] loss
# ## 3.1 hidden_size
# **Parameters**
# - hidden_size varied
# - max_lag = 8
# - loss_weights = [1, 1, 0.1, 0.01]
# +
max_lag = 8
loss_weights = [1, 1, .1, .01]
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
res_folder = os.path.join("results", "koopman")
# ### 3.1.1 hidden_size = [32, 16, 16, 8]
hidden_size = [32, 16, 16, 8]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.1.2 hidden_size = [16, 8, 8, 4]
hidden_size = [16, 8, 8, 4]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.1.3 hidden_size = [64, 128, 128, 256]
hidden_size = [64, 128, 128, 256]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.1.4 hidden_size = [32, 24, 24, 16]
hidden_size = [32, 24, 24, 16]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.1.5 hidden_size = [36, 32, 32, 24]
hidden_size = [36, 32, 32, 24]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.1.6 hidden_size = [40, 40, 40, 40]
hidden_size = [40, 40, 40, 40]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ## 3.2 max_lag
# +
loss_weights = [1, 1, .1, .01]
hidden_size = [64, 128, 128, 256]
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
# ### 3.2.1 max_lag = 4
max_lag = 4
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.2.2 max_lag = 8
# This is already done in 3.1.6
# ### 3.2.3 max_lag = 16
max_lag = 16
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.2.4 max_lag = 24
max_lag = 24
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.2.5 max_lag = 32
max_lag = 32
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ## 3.3 Loss coefficients
# +
hidden_size = [64, 128, 128, 256]
max_lag = 16
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
# ### 3.3.1 Base [1, 1, 0.1, 0.01]
# This is done in 3.2.3
# ### 3.3.2 Increased focus on forward loss [1, 2, 0.1, 0.01]
loss_weights = [1, 2, .1, .01]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.3.3 Only autoencoder and forward loss [1, 1, 0, 0]
# +
hidden_size = [40, 40, 40, 40]
max_lag = 8
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
loss_weights = [1, 1, 0, 0]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.3.4 Equal weightage to forward and backward dynamic [1, 0.5, 0.5, 0.01]
# +
hidden_size = [40, 40, 40, 40]
max_lag = 8
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
loss_weights = [1, 0.5, 0.5, 0.01]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# ### 3.3.5 Increased and Equal weightage to forward and backward dynamic [1, 1, 1, 0.01]
# +
hidden_size = [40, 40, 40, 40]
max_lag = 8
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
loss_weights = [1, 1, 1, 0.01]
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# # 4. Final Model
# +
hidden_size = [64, 128, 128, 256]
max_lag = 16
loss_weights = [1, 1, .1, .01]
batch_size = 128
epoch_list = [50, 50, 50]
lr_list = [1e-3, 1e-4, 1e-5]
# -
params, network = get_params(hidden_size, max_lag, loss_weights, SEED, batch_size, epoch_list, lr_list)
mean_pred = get_test_pred(X_test, params, network)
save_obj(params, res_folder, "best_params.pkl")
desc_name = "koopman_autoencoder"
res_single = PointExperimentResult(mean_pred - y_test, desc_name)
res_single.plot_rmse(error_thresh = 0.5)
print()
res_single.get_loss([0.2, 0.5, 1, 2, 3])
# # 5. Plot images based on Lyapunov Time
res_folder = os.path.join("results", "koopman")
def create_network(hidden_size):
p1, p2, p3, kappa = hidden_size
encoder_init, encoder = stax.serial(
Dense(p1), Tanh,
Dense(p2), Tanh,
Dense(p3), Tanh,
Dense(kappa))
decoder_init, decoder = stax.serial(
Dense(p3), Tanh,
Dense(p2), Tanh,
Dense(p1), Tanh,
Dense(data_dim))
forward_init, forward = stax.serial(
Dense_no_bias(kappa))
backward_init, backward = stax.serial(
Dense_no_bias(kappa))
network = (encoder, decoder, forward, backward)
return network
hidden_size = [64, 128, 128, 256]
network = create_network(hidden_size)
params = load_obj(os.path.join(res_folder, "best_params.pkl"))
mean_pred = get_test_pred(X_test, params, network)
save_obj(mean_pred, "results", "koopman_pred.pkl")
mean_pred = load_obj("results/koopman_pred.pkl")
total_lyapunov_time_pred = 4 / LORENZ_LT # Amount of Lyapunov Time in Test set
plot_predictions(mean_pred, y_test, total_lyapunov_time_pred, save_name = "lorenz_small_koop", pred_only = True)
res_single = PointExperimentResultLyapunov(mean_pred - y_test, "lorenz")
res_single.plot_rmse(save_name = "Lorenz_Small_Koopman_PH")
print()
res_single.get_loss()
| 2_small_size/1_lorenz8_20percent/4_koopman_lorenz8_small.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook compares the outputs from VESIcal to the Shishkina et al. (2014) Calibration dataset.
# - This notebook relies on the Excel spreadsheet entitled: "S7_Testing_Shishkina_et_al_2014.xlsx"
# - Test 1 compares the experimental pressures in the calibration dataset of Shishkina et al. (2014) for H$_2$O-only experiments to the saturation pressures obtained from VESIcal for the "ShishkinaWater" model.
# - Test 2 compares the experimental pressures in the calibration dataset of Shishkina et al. (2014) for CO$_2$-only experiments to the saturation pressures obtained from VESIcal for the "ShishkinaCarbon" model.
# - Test 3 compares the experimental pressures for mixed H$_2$O-CO$_2$ bearing fluids presented in Table 2 of the main text to the saturation pressures obtained from VESIcal for the "Shishkina" model.
# - Test 4 justifies the approach used in VESIcal, where cation fractions for their equation 9 are calculated ignoring H$_2$O and CO$_2$
#
import VESIcal as v
import matplotlib.pyplot as plt
import numpy as np
from IPython.display import display, HTML
import pandas as pd
import matplotlib as mpl
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
import statsmodels.api as sm
from statsmodels.sandbox.regression.predstd import wls_prediction_std
# %matplotlib inline
sns.set(style="ticks", context="poster",rc={"grid.linewidth": 1,"xtick.major.width": 1,"ytick.major.width": 1, 'patch.edgecolor': 'black'})
plt.style.use("seaborn-colorblind")
plt.rcParams["font.size"] =12
plt.rcParams["mathtext.default"] = "regular"
plt.rcParams["mathtext.fontset"] = "dejavusans"
plt.rcParams['patch.linewidth'] = 1
plt.rcParams['axes.linewidth'] = 1
plt.rcParams["xtick.direction"] = "in"
plt.rcParams["ytick.direction"] = "in"
plt.rcParams["ytick.direction"] = "in"
plt.rcParams["xtick.major.size"] = 6 # Sets length of ticks
plt.rcParams["ytick.major.size"] = 4 # Sets length of ticks
plt.rcParams["ytick.labelsize"] = 12 # Sets size of numbers on tick marks
plt.rcParams["xtick.labelsize"] = 12 # Sets size of numbers on tick marks
plt.rcParams["axes.titlesize"] = 14 # Overall title
plt.rcParams["axes.labelsize"] = 14 # Axes labels
plt.rcParams["legend.fontsize"]= 14
# ## Test 1 and 2 - comparing saturation pressures to experimental pressures
myfile_CO2 = v.BatchFile('S7_Testing_Shishkina_et_al_2014.xlsx', sheet_name='CO2') # Loading Carbon calibration dataset
satPs_wtemps_Shish_CO2= myfile_CO2.calculate_saturation_pressure(temperature="Temp", model='ShishkinaCarbon') # Calculating saturation pressures
myfile_H2O = v.BatchFile('S7_Testing_Shishkina_et_al_2014.xlsx', sheet_name='H2O') # Loading Water calibration dataset
satPs_wtemps_Shish_H2O= myfile_H2O.calculate_saturation_pressure(temperature="Temp", model='ShishkinaWater') # Calculating Saturation pressures
# +
######################## H2O only experiments
# This calculating a linear regression, and plots experimental pressures vs. saturation pressures for the Water calibration dataset
X_Test1=satPs_wtemps_Shish_H2O['Press']
Y_Test1=satPs_wtemps_Shish_H2O['SaturationP_bars_VESIcal']
mask_Test1 = (X_Test1>-1) & (Y_Test1>-1) # This gets rid of Nans
X_Test1noNan=X_Test1[mask_Test1].values.reshape(-1, 1)
Y_Test1noNan=Y_Test1[mask_Test1].values.reshape(-1, 1)
lr=LinearRegression()
lr.fit(X_Test1noNan,Y_Test1noNan)
Y_pred_Test1=lr.predict(X_Test1noNan)
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(12,5)) # adjust dimensions of figure here
ax1.plot(X_Test1noNan,Y_pred_Test1, color='red', linewidth=0.5, zorder=1) # This plots the best fit line
ax1.scatter(satPs_wtemps_Shish_H2O['Press'], satPs_wtemps_Shish_H2O['SaturationP_bars_VESIcal'], s=50, edgecolors='k', facecolors='silver', marker='o', zorder=5)
# This bit plots the regression parameters on the graph
I='Intercept= ' + str(np.round(lr.intercept_, 1))[1:-1]
G='Gradient= ' + str(np.round(lr.coef_, 3))[2:-2]
R='R$^2$= ' + str(np.round(r2_score(Y_Test1noNan, Y_pred_Test1), 3))
ax1.text(3000, 1500, R, fontsize=14)
ax1.text(3000, 1000, G, fontsize=14)
ax1.text(3000, 500, I, fontsize=14)
################### CO2 experiments
X_Test2=satPs_wtemps_Shish_CO2['Press']
Y_Test2=satPs_wtemps_Shish_CO2['SaturationP_bars_VESIcal']
mask_Test2 = (X_Test2>-1) & (Y_Test2>-1) # This gets rid of Nans
X_Test2noNan=X_Test2[mask_Test2].values.reshape(-1, 1)
Y_Test2noNan=Y_Test2[mask_Test2].values.reshape(-1, 1)
lr=LinearRegression()
lr.fit(X_Test2noNan,Y_Test2noNan)
Y_pred_Test2=lr.predict(X_Test2noNan)
ax2.plot(X_Test2noNan,Y_pred_Test2, color='red', linewidth=0.5, zorder=1) # This plots the best fit line
ax2.scatter(satPs_wtemps_Shish_CO2['Press'], satPs_wtemps_Shish_CO2['SaturationP_bars_VESIcal'], s=50, edgecolors='k', facecolors='silver', marker='o', zorder=5)
# This bit plots the regression parameters on the graph
I='Intercept= ' + str(np.round(lr.intercept_, 2))[1:-1]
G='Gradient= ' + str(np.round(lr.coef_, 3))[2:-2]
R='R$^2$= ' + str(np.round(r2_score(Y_Test2noNan, Y_pred_Test2), 2))
ax2.text(4000, 500, I, fontsize=14)
ax2.text(4000, 1000, G, fontsize=14)
ax2.text(4000, 1500, R, fontsize=14)
ax1.set_xlabel('Experimental Pressure (bar)', fontsize=14)
ax1.set_ylabel('P$_{Sat}$ VESIcal (bar)', fontsize=14)
ax2.set_xlabel('Experimental Pressure (bar)', fontsize=14)
ax2.set_ylabel('P$_{Sat}$ VESIcal (bar)', fontsize=14)
ax1.set_xticks([0, 2000, 4000, 6000, 8000, 10000])
ax1.set_yticks([0, 2000, 4000, 6000, 8000, 10000])
ax2.set_xticks([0, 2000, 4000, 6000, 8000, 10000])
ax2.set_yticks([0, 2000, 4000, 6000, 8000, 10000])
ax1.set_xlim([-200, 6500])
ax1.set_ylim([-200, 6500])
ax2.set_xlim([-200, 8000])
ax2.set_ylim([-200, 8000])
plt.subplots_adjust(left=0.125, bottom=None, right=0.9, top=None, wspace=0.3, hspace=None)
ax1.text(-150, 6200, 'a)', fontsize=14)
ax2.text(-150, 7600, 'b)', fontsize=14)
ax1.set_title('H$_{2}$O-only', fontsize=14)
ax2.set_title('CO$_2$-only', fontsize=14)
fig.savefig('Shishkina_Test1and2.png', transparent=True)
# -
# ## Test 3 - Mixed H$_2$O - CO$_2$ experiments from Table 2 in the text.
# - We show the regression for experimental pressure vs. saturation pressure calculated in VESIcal for all data, and data with experimental pressures <4000 bars (to remove the most scattered datapoints).
myfile_Comb = v.BatchFile('S7_Testing_Shishkina_et_al_2014.xlsx', sheet_name='Table2_Text') # Loads experimental data from Table 2
satPs_wtemps_Shish_Comb= myfile_Comb.calculate_saturation_pressure(temperature="Temp", model='ShishkinaIdealMixing') # Calculates saturation pressures for these compositions + tempts
# +
######################## H2O only experiments
X_Test3b=satPs_wtemps_Shish_Comb['Press']
Y_Test3b=satPs_wtemps_Shish_Comb['SaturationP_bars_VESIcal']
mask_Test3b = (X_Test3b>-1) & (Y_Test3b>-1) # This gets rid of Nans
X_Test3bnoNan=X_Test3b[mask_Test3b].values.reshape(-1, 1)
Y_Test3bnoNan=Y_Test3b[mask_Test3b].values.reshape(-1, 1)
lr=LinearRegression()
lr.fit(X_Test3bnoNan,Y_Test3bnoNan)
Y_pred_Test3b=lr.predict(X_Test3bnoNan)
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(12,5)) # adjust dimensions of figure here
ax1.plot(X_Test3bnoNan,Y_pred_Test3b, color='red', linewidth=0.5, zorder=1) # This plots the best fit line
ax1.scatter(satPs_wtemps_Shish_Comb['Press'], satPs_wtemps_Shish_Comb['SaturationP_bars_VESIcal'], s=50, edgecolors='k', facecolors='silver', marker='o', zorder=5)
# This bit plots the regression parameters on the graph
I='Intercept= ' + str(np.round(lr.intercept_, 1))[1:-1]
G='Gradient= ' + str(np.round(lr.coef_, 3))[2:-2]
R='R$^2$= ' + str(np.round(r2_score(Y_Test3bnoNan, Y_pred_Test3b), 3))
ax1.text(3000, 1500, R, fontsize=14)
ax1.text(3000, 1000, G, fontsize=14)
ax1.text(3000, 500, I, fontsize=14)
################### CO2 experiments
X_Test3=satPs_wtemps_Shish_Comb['Press']
Y_Test3=satPs_wtemps_Shish_Comb['SaturationP_bars_VESIcal']
mask_Test3 = (X_Test3>-1) & (Y_Test3>-1) &(X_Test3<4000) # This gets rid of Nans
X_Test3noNan=X_Test3[mask_Test3].values.reshape(-1, 1)
Y_Test3noNan=Y_Test3[mask_Test3].values.reshape(-1, 1)
lr=LinearRegression()
lr.fit(X_Test3noNan,Y_Test3noNan)
Y_pred_Test3=lr.predict(X_Test3noNan)
ax2.plot(X_Test3noNan,Y_pred_Test3, color='red', linewidth=0.5, zorder=1) # This plots the best fit line
ax2.scatter(satPs_wtemps_Shish_Comb['Press'], satPs_wtemps_Shish_Comb['SaturationP_bars_VESIcal'], s=50, edgecolors='k', facecolors='silver', marker='o', zorder=5)
# This bit plots the regression parameters on the graph
I='Intercept= ' + str(np.round(lr.intercept_, 2))[1:-1]
G='Gradient= ' + str(np.round(lr.coef_, 3))[2:-2]
R='R$^2$= ' + str(np.round(r2_score(Y_Test3noNan, Y_pred_Test3), 2))
ax2.text(2000, 100, I, fontsize=14)
ax2.text(2000, 400, G, fontsize=14)
ax2.text(2000, 700, R, fontsize=14)
ax1.set_xlabel('Experimental Pressure (bar)', fontsize=14)
ax1.set_ylabel('P$_{Sat}$ VESIcal (bar)', fontsize=14)
ax2.set_xlabel('Experimental Pressure (bar)', fontsize=14)
ax2.set_ylabel('P$_{Sat}$ VESIcal (bar)', fontsize=14)
ax1.set_xticks([0, 2000, 4000, 6000, 8000, 10000])
ax1.set_yticks([0, 2000, 4000, 6000, 8000, 10000])
ax2.set_xticks([0, 2000, 4000, 6000, 8000, 10000])
ax2.set_yticks([0, 2000, 4000, 6000, 8000, 10000])
ax1.set_xlim([-200, 8000])
ax1.set_ylim([-200, 8000])
ax2.set_xlim([-200, 4000])
ax2.set_ylim([-200, 4000])
plt.subplots_adjust(left=0.125, bottom=None, right=0.9, top=None, wspace=0.3, hspace=None)
ax1.text(-150, 7600, 'a)', fontsize=14)
ax2.text(-150, 3800, 'b)', fontsize=14)
ax1.set_title('All Experiments', fontsize=14)
ax2.set_title('Experimental Pressure < 4000 bars)', fontsize=14)
fig.savefig('Shishkina_Test3.png', transparent=True)
# -
# ## Test 4 - Intepretation of "atomic fractions of cations in Equation 9.
# - We can only recreate the chemical data for cation fractions shown in their Fig. 7a if the "atomic fractions of cations" are calculated excluding volatiles. Including atomic proportions including H$_2$O and CO$_2$ results in a significantly worse fit to experimental data for the ShishkinaWater model shown in test 2. The choice of normalization doesn't affect the results for the CO$_2$ model, where the compositional dependence is expressed as a fraction
# Removed CO2 and H2O
oxides = ['SiO2', 'TiO2', 'Al2O3', 'Fe2O3', 'Cr2O3', 'FeO', 'MnO', 'MgO', 'NiO', 'CoO', 'CaO', 'Na2O', 'K2O', 'P2O5']
oxideMass = {'SiO2': 28.085+32, 'MgO': 24.305+16, 'FeO': 55.845+16, 'CaO': 40.078+16, 'Al2O3': 2*26.982+16*3, 'Na2O': 22.99*2+16,
'K2O': 39.098*2+16, 'MnO': 54.938+16, 'TiO2': 47.867+32, 'P2O5': 2*30.974+5*16, 'Cr2O3': 51.996*2+3*16,
'NiO': 58.693+16, 'CoO': 28.01+16, 'Fe2O3': 55.845*2+16*3}
CationNum = {'SiO2': 1, 'MgO': 1, 'FeO': 1, 'CaO': 1, 'Al2O3': 2, 'Na2O': 2,
'K2O': 2, 'MnO': 1, 'TiO2': 1, 'P2O5': 2, 'Cr2O3': 2,
'NiO': 1, 'CoO': 1, 'Fe2O3': 2}
Normdata = myfile_H2O.get_data(normalization="additionalvolatiles")
for ind,row in Normdata.iterrows():
for ox in oxides:
Normdata.loc[ind, ox + 'molar']=((row[ox]*CationNum[ox])/oxideMass[ox]) # helps us get desired column name with its actual name, rather than its index. If by number, do by iloc.
#oxide_molar[ind, ox]=ox+'molar'
Normdata.loc[ind,'sum']=sum(Normdata.loc[ind, ox+'molar'] for ox in oxides)
for ox in oxides:
Normdata.loc[ind, ox + 'norm']=Normdata.loc[ind, ox+'molar']/Normdata.loc[ind, 'sum']
# helps us get desired column name with its actual name, rather than its index. If by number, do by iloc.
Normdata.head()
# +
### Comparison of these cation fractions to those shown in their Fig. 7a
# -
fig, ax1 = plt.subplots(figsize = (10,8)) # adjust dimensions of figure here
font = {'family': 'sans-serif',
'color': 'black',
'weight': 'normal',
'size': 20,
}
plt.xlim([0, 0.25])
plt.ylim([1, 13])
plt.title('Calculated using VESIcal')
plt.scatter(Normdata['Na2Onorm']+Normdata['K2Onorm'], Normdata['H2O'], edgecolor='k', facecolor='b', s=50, label='Normalized')
plt.xlabel('Na+K')
plt.ylabel('H$_2$O')
plt.legend()
# # Their graph below
# 
| manuscript/Supplement/JupyterNotebooks/Shishkina/S7_Testing_Shishkina_et_al_2014.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import numpy as np
import torch as th
import hypergrad as hg
TARGET_DEVICE = th.device('cuda') if th.cuda.is_available() else th.device('cpu')
# -
# Code for training a simple equilibrium network with "RNN-style" dynamics on a subset of the MNIST data.
#
# For more details refer to Section 3.2 of the paper
# "On the iteration complexity of hypergradient computation" (https://arxiv.org/abs/2006.16218)
# + pycharm={"name": "#%%\n"}
# --------------------------------------------
# UTILS
# --------------------------------------------
def to_numpy(tensor):
if isinstance(tensor, list):
return [to_numpy(v) for v in tensor]
else:
return tensor.detach().to(th.device('cpu')).numpy()
def set_requires_grad(lst): [l.requires_grad_(True) for l in lst]
def acc(preds, targets):
"""Computes the accuracy"""
return preds.argmax(dim=1).eq(targets).float().mean()
class NamedLists(list):
def __init__(self, lst, names) -> None:
super().__init__(lst)
assert len(lst) == len(names)
self.names = names
def __getitem__(self, i):
if isinstance(i, str):
return self.__getattribute__(i)
else:
return super().__getitem__(i)
class TVT(NamedLists): # train val & test
def __init__(self, lst) -> None:
super().__init__(lst, ['train', 'val', 'test'])
self.train, self.val, self.test = lst
class DT(NamedLists): # data & targets
def __init__(self, lst) -> None:
super().__init__(lst, ['data', 'targets'])
self.data, self.targets = lst
class LA(NamedLists): # loss and accuracy
def __init__(self, lst):
super().__init__(lst, ['loss', 'acc'])
self.loss, self.acc = lst
def load_mnist(seed=0, num_train=50000, num_valid=10000):
"""Load MNIST dataset with given number of training and validation examples"""
from torchvision import datasets
rnd = np.random.RandomState(seed)
mnist_train = datasets.MNIST('../data', download=True, train=True)
train_indices = rnd.permutation(list(range(60000)))
dta, targets = mnist_train.data, mnist_train.targets
# print(train_indices)
tr_inds = train_indices[:num_train]
mnist_tr1 = DT([dta[tr_inds], targets[tr_inds]])
val_inds = train_indices[num_train:num_train + num_valid]
mnist_valid = DT([dta[val_inds], targets[val_inds]])
mnist_test = datasets.MNIST('../data', download=True, train=False)
def _process_dataset(dts):
dt, tgt = np.array(dts.data.numpy(), dtype=np.float32), dts.targets.numpy()
return DT([th.from_numpy(
np.reshape(dt / 255., (-1, 28 * 28))).to(TARGET_DEVICE),
th.from_numpy(tgt).to(TARGET_DEVICE)])
return TVT([_process_dataset(dtt) for dtt in [mnist_tr1, mnist_valid, mnist_test]])
# + pycharm={"name": "#%%\n"}
i_sig = 0.01 # initialization
dw = 200 # dimensionality of the hidden state
lr = 0.5
th.manual_seed(0)
data = load_mnist(0, num_train=5000, num_valid=5000)
num_exp, dim_x = data.train.data.shape
# + pycharm={"name": "#%%\n"}
do_projection = True
T = K = 20 # number of iterations; T for forward iterations, K for backward;
# + pycharm={"name": "#%%\n"}
# choose between
# # rm (reverse-mode iterative differentiation),
# fp (fixed point implicit differentiation) and
# cg (conjugate gradient implicit differentiation)
hg_mode = 'rm'
# + pycharm={"name": "#%%\n"}
def matrix_projection_on_spectral_ball(a, radius=0.99, project=True):
A = a.detach()
if A.is_cuda: A = A.cpu()
A = A.numpy()
U, S, V = np.linalg.svd(A)
if project:
S1 = np.minimum(S, radius)
a = U @ np.diag(S1) @ V
else:
a = A
return th.from_numpy(a).type(th.FloatTensor).to(TARGET_DEVICE).requires_grad_(True), S
# + pycharm={"name": "#%%\n"}
initial_states = TVT([th.zeros(d.data.shape[0], dw, device=TARGET_DEVICE) for d in data])
if hg == 'rm': set_requires_grad(initial_states) # necessary only for reverse-mode with unrolling
# define model's parameters
parameters = [
i_sig * th.randn(dw, dw, device=TARGET_DEVICE),
i_sig * th.randn(dim_x, dw, device=TARGET_DEVICE),
i_sig * th.randn(dw, device=TARGET_DEVICE),
i_sig * th.randn(dw, 10, device=TARGET_DEVICE),
th.zeros(10, device=TARGET_DEVICE)
]
set_requires_grad(parameters)
# + pycharm={"name": "#%%\n"}
def get_fully_connected_dynamics(x):
def fully_connected_dynamics(state_list, params):
# RNNs like dynamics (the fp_map of the bi-level problem)
A, B, c = params[:3]
state = state_list[0]
return [th.tanh(state @ A + x @ B + c)]
return fully_connected_dynamics
get_dynamics = get_fully_connected_dynamics # change this line for changing type of dynamics
# obtain one dynamics per set (training, validation and test) which is a callable
tvt_dynamics = TVT([get_dynamics(dt.data) for dt in data])
# + pycharm={"name": "#%%\n"}
def linear(state_list, params):
return state_list[0] @ params[-2] + params[-1]
def get_loss(targets):
def loss(state_list, params):
# cross entropy loss (the outer loss of the bi-level problem)
outputs = linear(state_list, params)
criterion = th.nn.CrossEntropyLoss()
return th.mean(criterion(outputs, targets))
return loss
# obtain one loss per dataset (note: the losses remain callable as well as the dynamics!).
tvt_losses = TVT([get_loss(dt.targets) for dt in data])
# + pycharm={"name": "#%%\n"}
# forward pass
def get_forward(initial_state, dynamics):
def forward():
states = [[initial_state]]
for _ in range(T):
states.append(dynamics(states[-1], parameters))
return states
return forward
# one per dataset
tvt_forward = TVT([get_forward(s, dyna) for s, dyna in zip(initial_states, tvt_dynamics)])
def metric_after_fw(forward, metric):
def _f():
states = forward()
return metric(states[-1], parameters)
return _f
def accuracy(targets):
def _f(states, params):
return acc(linear(states, params), targets)
return _f
# obtain callables for loss and accuracy for each set (after executing the model's dynamics)
tvt_metrics = TVT([
LA([metric_after_fw(fww, lss), metric_after_fw(fww, accuracy(dt.targets))])
for fww, lss, dt in zip(tvt_forward, tvt_losses, data)
])
# + pycharm={"name": "#%%\n"}
# optimizer
opt = th.optim.SGD(parameters, lr, momentum=0.9)
# + pycharm={"name": "#%%\n"}
# training!
for t in range(1000):
opt.zero_grad()
states = tvt_forward.train()
# compute the hypergradient (with different methods)
if hg_mode == 'fp':
hg.fixed_point(states[-1], parameters, K, tvt_dynamics.train, tvt_losses.train)
elif hg_mode == 'cg':
hg.CG_normaleq(states[-1], parameters, K, tvt_dynamics.train, tvt_losses.train)
elif hg_mode == 'rm':
hg.reverse_unroll(states[-1], parameters, tvt_losses.train)
else:
raise NotImplementedError('{} not available!'.format(hg_mode))
opt.step()
try: # perform projection
A_proj, svl = matrix_projection_on_spectral_ball(parameters[0], project=do_projection)
parameters[0].data = A_proj.data
except (ValueError, np.linalg.LinAlgError) as e:
print('there were nans most probably: aborting all')
break
if t % 20 == 0:
valid_acc = to_numpy(tvt_metrics.val.acc())
hgs = to_numpy([l.grad for l in parameters])
print('Validation accuracy at iteration {}:'.format(t), valid_acc) # update early stopping
| examples/Equilibrium models (RNN-style model on MNIST).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h3><b>Introduction</b></h3>
# <p> The goal of this project is to take the publically available Beijing weather data from 2013 to 2017 and apply machine learning techniques to see if we can predict the amount of PM2.5 concentration in the air given other environmental features. This is a project I am working on during my free time applying some of the machine learning algorithms I have learned. I hope to come up with a predictive model with a high accuracy and a very low Root Mean Square Error (RMSE).</p>
# <br>
# <h5><b>Dataset Information</b></h5>
# <p>This data set includes hourly air pollutants data from 12 nationally-controlled air-quality monitoring sites. The air-quality data are from the Beijing Municipal Environmental Monitoring Center. The meteorological data in each air-quality site are matched with the nearest weather station from the China Meteorological Administration. The time period is from March 1st, 2013 to February 28th, 2017. Missing data are denoted as NA.</p>
# <br>
# <h5><b>Project workflow</b></h5>
# <p>This project utilizes the Supervised Machine Learning algorithms from python’s Scikit-learn library. The model we hope to succeed in training is a regression model and below are the steps we will go through in this jupyter notebook for this project:</p>
# <ul>
# <li>Import the neccessary libraries and loading the data</li>
# <li>Data preprocessing</li>
# <li>Exploratory Data Ananlysis</li>
# <li>Model training and Evaluation</li>
# <li>Saving the model</li>
# </ul>
# <br>
# <h5><b>Import libraries and loading data</b></h5>
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import warnings
warnings.filterwarnings(action='ignore')
from statsmodels.tsa.seasonal import seasonal_decompose
import statsmodels.formula.api as formula
from statsmodels.stats.outliers_influence import variance_inflation_factor
import statsmodels as sm
from sklearn import preprocessing
from sklearn.model_selection import train_test_split,GridSearchCV,RandomizedSearchCV
from sklearn.linear_model import LinearRegression,Lasso,Ridge
from sklearn.externals import joblib
from sklearn.metrics import r2_score,mean_squared_error
from sklearn.ensemble import RandomForestRegressor,GradientBoostingRegressor
from sklearn.tree import DecisionTreeRegressor
from IPython.display import display
# +
# get the name of the csv file
#file = 'https://raw.githubusercontent.com/prince381/air-pollution/master/data/PRSA_Data_Aotizhongxin_20130301-20170228.csv'
file = r'C:\Users\DELL\Desktop\EDUCATE\DATA CSV\PRSA_Data_20130301-20170228\data\PRSA_Data_Aotizhongxin_20130301-20170228.csv'
# read the csv file into a pandas DataFrame using the pd.read_csv()
data = pd.read_csv(file)
data.head()
# -
# <h3><b>Data Preprocessing</b></h3>
# drop the unwanted columns/features
cols_to_drop = ['No','station']
data = data.drop(cols_to_drop,axis=1)
# print out the info of the data
data.info()
# check for duplicated values and null values
print('Are there any duplicated values in our data ? : {}\n'.format(data.duplicated().any()))
print('The total number of null values in each colum:')
display(data.isnull().sum())
# find the most appearing wind direction value
data.wd.mode()
# fill in the missing values with the mean of the particular column
data.fillna(value=data.mean(),inplace=True)
# replace the missing values for the wind direction with the modal value
data.wd.fillna(value='NE',inplace=True)
# let's check the data again if there are any missing values
data.isnull().any()
# +
# create a datetime column using the year,month,day and hour columns.
years = data['year'].values
months = data['month'].values
days = data['day'].values
hours = data['hour'].values
full_date = []
for i in range(data.shape[0]):
date_time = str(years[i])+'-'+str(months[i])+'-'+str(days[i])+' '+str(hours[i])+':'+str(0)
full_date.append(date_time)
dates = pd.to_datetime(full_date)
dates = pd.DataFrame(dates,columns=['date'])
data = pd.concat([dates,data],axis=1)
data.head()
# -
# <h3><b>Exploratory Data Analysis</b></h3>
# <p>Before we start fitting a machine learning model on the data, we need to know much about the data by performing an Exploratory Data Analysis to gain insight from it. <abbr title="Exploratory Data Analysis">EDA</abbr> is simply describing the data by means of visualization. It involves asking questions about the data and answering them with the help of charts/graphs (graphical representation of the data). In this process, we will try to study the behavoir of the amount of pollutant (PM2.5 concentration) in the air and the relationship between other features. Below are some of the questions we will try to answer by analyzing the data, to know more about our dependent and independent variables:</p>
# <ul>
# <li>what pattern does the amount of PM2.5 concentration in the air recorded in an hour follow
# for a daily time period ?</li>
# <li>In which month does the amount of PM2.5 contained in the air rises ?</li>
# <li>At what time of the day do we expect the amount of PM2.5 concentration in the
# air to be high ?</li>
# <li>In which direction does polluted air/wind mostly move ?</li>
# <li>How do the other environmental factors affect the amount of PM2.5 concentration
# in the air ?</li>
# </ul>
# <p>We now have our questions so let's just dive into our data and start finding and interpreting some results. But since we are going to take averages of the dependent variable, we shoul know the distribtution of the data before we do take averages.</p>
plt.figure(figsize=(12,5))
sns.distplot(data['PM2.5'],bins=50)
plt.title('Distribution of the hourly recorded PM2.5 concetration in the air\nin Aotizhongxin-Beijin',
fontsize=16)
plt.show()
# <h5><b>what pattern does the amount of PM2.5 concentration in the air recorded in an hour follow
# for a daily time period ?</b></h5>
# +
# find the daily average of PM2.5 contained in the air in any given hour
daily_data = data[['date','PM2.5']]
daily_data = daily_data.set_index('date')
daily_data = daily_data.resample('D').median()
decomposition = seasonal_decompose(daily_data,model='addictive')
# plot the data
with plt.style.context('fivethirtyeight'):
decomposition.trend.plot(figsize=(12,5),style='k-',linewidth=.9,legend=False)
plt.xlabel('Date',fontsize=14)
plt.ylabel('PM2.5 concentration (ug/m^3)',fontsize=14)
plt.title('Daily trend in the hourly recorded PM2.5 concentration in\nthe air in Aotizhongxin-Beijin',fontsize=16)
plt.grid(axis='x')
plt.tight_layout()
plt.show()
# -
# <h5><b>In which month does the amount of PM2.5 contained in the air rises ?</b></h5>
# +
monthly_data = data[['month','PM2.5']]
months = ['January','February','March','April','May','June','July',
'August','September','October','November','December']
ordered_monthdf = pd.DataFrame(months,columns=['month'])
map_dict = {}
for i,j in enumerate(months):
map_dict.setdefault(i+1,j)
monthly_data.month = monthly_data.month.map(map_dict)
monthly_average = monthly_data.groupby('month').median()
monthly_average = pd.merge(ordered_monthdf,monthly_average,left_on='month',right_index=True)
monthly_average = np.round(monthly_average,1)
monthly_average = monthly_average.set_index('month')
# plot the data
with plt.style.context('ggplot'):
monthly_average.plot(figsize=(12,5),legend=False,kind='bar',linewidth=.9)
plt.xlabel('Month',fontsize=14)
plt.ylabel('PM2.5 concentration (ug/m^3)',fontsize=14)
plt.title('Monthly average of the hourly recorded PM2.5 concentration in\nthe air in Aotizhongxin-Beijin',fontsize=16)
plt.grid(axis='x')
plt.tight_layout()
plt.show()
# -
# <h5><b>At what time of the day do we expect the amount of PM2.5 concentration in the air to be high ?</b></h5>
# +
hourly_data = data[['hour','PM2.5']]
hrs = ['12 AM','1 AM','2 AM','3 AM','4 AM','5 AM','6 AM','7 AM','8 AM','9 AM','10 AM',
'11 AM','12 PM','1 PM','2 PM','3 PM','4 PM','5 PM','6 PM','7 PM',
'8 PM','9 PM','10 PM','11 PM']
hour_dict = {}
for i,j in enumerate(hrs):
hour_dict.setdefault(i,j)
hourly_data = hourly_data.groupby('hour').median().reset_index()
hourly_data.hour = hourly_data.hour.map(hour_dict)
hourly_data = hourly_data.set_index('hour')
# plot the data
with plt.style.context('ggplot'):
hourly_data.plot(figsize=(12,8),legend=False,kind='barh',linewidth=.9)
plt.ylabel('Hours',fontsize=14)
plt.xlabel('PM2.5 concentration (ug/m^3)',fontsize=14)
plt.title('Average recorded PM2.5 concentration in the air in Aotizhongxin-Beijin\nby the hour of the day',fontsize=16)
plt.grid(axis='y')
plt.tight_layout()
plt.show()
# -
# <h5><b>In which direction does polluted air/wind mostly move ?</b></h5>
# +
wind_dir = data[['wd','PM2.5']]
wind_dir = wind_dir.groupby('wd').median()
# plot the data
with plt.style.context('ggplot'):
wind_dir.plot(figsize=(12,5),legend=False,kind='bar',linewidth=.9)
plt.xlabel('Wind direction',fontsize=14)
plt.ylabel('PM2.5 concentration (ug/m^3)',fontsize=14)
plt.title('Average hourly recorded PM2.5 concentration in the air in Aotizhongxin-Beijin\ngrouped by wind direction',fontsize=16)
plt.grid(axis='x')
plt.tight_layout()
plt.show()
# -
# <h5><b>How do the other environmental factors affect the amount of PM2.5 concentration in the air ?</b></h5>
# let's try and visualize the relationships between the features of the data
plt.figure(figsize=(13,9))
correlation_data = data[['PM2.5', 'PM10', 'SO2', 'NO2',
'CO', 'O3', 'TEMP', 'PRES',
'DEWP', 'RAIN', 'WSPM']]
sns.heatmap(correlation_data.corr(),cmap=plt.cm.Reds,annot=True)
plt.title('Heatmap displaying the correlation matrix of the variables',fontsize=16)
plt.show()
# <h3><b>Model Training and Evaluation</b></h3>
# <br>
# <h5><b>check for multicollinearity among variables and fit a regression model using statsmodels</b></h5>
# +
cols_to_drop = ['date','year','month','day','hour','wd']
newdata = data.drop(cols_to_drop,axis=1)
# calculate the variance inflation factor of each feature and detect multicollinearity
cons_data = sm.tools.add_constant(newdata)
series_before = pd.Series([variance_inflation_factor(cons_data.values,i) for i in range(cons_data.shape[1])],
index=cons_data.columns)
series_before
# -
# we can see that TEMP (temperature) and DEWP (dewpoint) are highly correlated as the VIF value is
# greater than 5. As a result, we get rid of one of those features and probably the one that has the
# lowest correlation with the dependent variable.
newdata = newdata.drop('DEWP',axis=1)
cons_data2 = sm.tools.add_constant(newdata)
series_after = pd.Series([variance_inflation_factor(cons_data2.values,i) for i in range(cons_data2.shape[1])],
index=cons_data2.columns)
series_after
# +
newdata.columns = ['PM2_5','PM10','SO2','NO2','CO','O3','TEMP','PRES','RAIN','WSPM']
# PM2.5 is skewed to the right so we log transform the values to normalize the distribution
newdata['PM2_5'] = np.log(newdata['PM2_5'])
# -
# fit the regression model
mul_reg = formula.ols(formula='PM2_5 ~ PM10 + SO2 + NO2 + CO + O3 + TEMP + PRES + RAIN + WSPM',
data=newdata).fit()
mul_reg.summary()
# <p>The OLS model from statsmodels gives us an accuracy of 71% (0.712) which is not satisfactory for prediction. So we move on to fit a linear regression model from the scikit-learn library.</p>
# <br>
# <h5><b>fitting a linear regression model with sklearn.linear_model.LinearRegression()</b></h5>
# +
# we split the data into predictor variables and Outcome variable
X = newdata.drop('PM2_5',axis=1)
y = newdata['PM2_5']
# we need to scale or normalize the predictor variables since they are not on the same
# scale and some of their distributions are skewed.
X_scaled = preprocessing.scale(X)
X_scaled = pd.DataFrame(X_scaled,columns=X.columns)
X_scaled.dropna(inplace=True)
# print the scaled predictor variables.
X_scaled.head()
# +
# we now split out data into train and test data
X_train,X_test,y_train,y_test = train_test_split(X_scaled,y,test_size=.2,random_state=0)
# instantiate the linear regression model
lin_model = LinearRegression()
lin_model.fit(X_train,y_train) # fit the model
# -
# we now score the model
print('Score on train data: {}\n'.format(lin_model.score(X_train,y_train)))
print('Score on test data: {}'.format(lin_model.score(X_test,y_test)))
# +
prediction = lin_model.predict(X_test)
mse = mean_squared_error(y_test,prediction)
accuracy = r2_score(y_test,prediction)
print('Mean Squared Error: {}\n'.format(mse))
print('Overall model accuracy: {}'.format(accuracy))
# -
# <p>The model accuracy for the LinearRegression() is no better than that of the statsmodels. They all give the same accuracy is not better for making predictions. We now move on to fit other models by using the ensemble methods</p>
# <br>
# <br>
# <h3><b>Ensemble methods</b></h3>
# <p>For ensemble methods (DecisionTreeRegressor,RandomForestRegressor,and GradientBoostingRegressor),we include the pressure and rain features and we won't scale the predictor variables,neither would we log transform the outcome variable. At this part, model fitting and hyper-parameter tunning will be done at the same time. Instead of fitting the model with single parameters, we will straight away perform the grid search with multiple values for a parameter and find the best parameters for fitting the model on our data to get a satisfactory accuracy.</p>
# +
ensemble_data = data.drop(cols_to_drop,axis=1)
# we split the data into predictor variables and Outcome variable
X = ensemble_data.drop('PM2.5',axis=1)
y = ensemble_data['PM2.5']
# -
xtrain,xtest,ytrain,ytest = train_test_split(X,y,test_size=.2)
# ### DecisionTreeRegressor
# we will now fit a decision tree regression model on the data and tune some of its parameters to increase the accuracy.
# we go ahead to use the ensemble methods as the LinearRegression model has a low accuracy
# on both the test and train data.
decision_tree = DecisionTreeRegressor(max_depth=5,
max_features='auto',
min_samples_split=3,
min_samples_leaf=2)
decision_tree.fit(xtrain,ytrain)
# +
# we now score the model
print('Score on train data: {}\n'.format(decision_tree.score(xtrain,ytrain)))
print('Score on test data: {}\n'.format(decision_tree.score(xtest,ytest)))
tree_pred = decision_tree.predict(xtest)
tree_mse = mean_squared_error(ytest,tree_pred)
tree_accuracy = r2_score(ytest,tree_pred)
print('Root Mean Squared Error: {}\n'.format(np.sqrt(tree_mse)))
print('Overall model accuracy: {}'.format(tree_accuracy))
# +
# We now tune the parameters of the model to see if we can increase the accuracy
params = {'max_depth':[3,4,5,6,7],
'max_features':['auto','sqrt','log2'],
'min_samples_split':[2,3,4,5,6,7,8,9,10],
'min_samples_leaf':[2,3,4,5,6,7,8,9,10]}
tree = DecisionTreeRegressor()
# initialize the grid search for the best parameters
tree_search = GridSearchCV(tree,param_grid=params,
n_jobs=-1,cv=5)
tree_search.fit(xtrain,ytrain) # fit the model
# +
# we now score the model
print('Score on train data: {}\n'.format(tree_search.score(xtrain,ytrain)))
print('Score on test data: {}\n'.format(tree_search.score(xtest,ytest)))
print('Best parameters found:')
display(tree_search.best_params_)
tree_search_pred = tree_search.predict(xtest)
tree_search_mse = mean_squared_error(ytest,tree_search_pred)
tree_search_accuracy = r2_score(ytest,tree_search_pred)
print('Root Mean Squared Error: {}\n'.format(np.sqrt(tree_search_mse)))
print('Overall model accuracy: {}'.format(tree_search_accuracy))
# -
# ### RandomForestRegressor
#
# we now fit a random forest regression model on the data to see if we would get a better accuracy results than that of the decision tree regression model.
# +
# instantiate the RandomForestRegressor model and fit the model on the training data
forest = RandomForestRegressor(n_estimators=100,
max_depth=7,
max_features='auto',
min_samples_split=7,
min_samples_leaf=3)
forest.fit(xtrain,ytrain)
# +
# we now score the model
print('Score on train data: {}\n'.format(forest.score(xtrain,ytrain)))
print('Score on test data: {}\n'.format(forest.score(xtest,ytest)))
forest_pred = forest.predict(xtest)
forest_mse = mean_squared_error(ytest,forest_pred)
forest_accuracy = r2_score(ytest,forest_pred)
print('Root Mean Squared Error: {}\n'.format(np.sqrt(forest_mse)))
print('Overall model accuracy: {}'.format(forest_accuracy))
# +
# we now tune the parameters of the RandomForestRegressor model using RandomizedSearchCV to
# find the best parameters and increase the accuracy of the model
params['n_estimators'] = [100,200,300,400,500]
# instantiate the model
random_forest = RandomForestRegressor()
# perform the grid search for the best parameters
forest_search = RandomizedSearchCV(random_forest,params,n_jobs=-1,
cv=5,verbose=2)
forest_search.fit(xtrain,ytrain)
# +
# we now score the model
print('Score on train data: {}\n'.format(forest_search.score(xtrain,ytrain)))
print('Score on test data: {}\n'.format(forest_search.score(xtest,ytest)))
print('Best parameters found:')
display(forest_search.best_params_)
forest_search_pred = forest_search.predict(xtest)
forest_search_mse = mean_squared_error(ytest,forest_search_pred)
forest_search_accuracy = r2_score(ytest,forest_search_pred)
print('Root Mean Squared Error: {}\n'.format(np.sqrt(forest_search_mse)))
print('Overall model accuracy: {}'.format(forest_search_accuracy))
# -
# ### GradientBoostingRegressor
#
# we now fit a gradient boosting regression model on the data to see if we would get a better accuracy results than that of the decision tree and random forest regression model and also minimize the error.
# +
# instantiate the GradientBoostingRegressor model and fit the model on the training data
grad_boost = GradientBoostingRegressor(n_estimators=100,
max_depth=7,
max_features='auto',
min_samples_split=7,
min_samples_leaf=3,
learning_rate=0.1)
grad_boost.fit(xtrain,ytrain)
# +
# we now score the model
print('Score on train data: {}\n'.format(grad_boost.score(xtrain,ytrain)))
print('Score on test data: {}\n'.format(grad_boost.score(xtest,ytest)))
gboost_pred = grad_boost.predict(xtest)
gboost_mse = mean_squared_error(ytest,gboost_pred)
gboost_accuracy = r2_score(ytest,gboost_pred)
print('Root Mean Squared Error: {}\n'.format(np.sqrt(gboost_mse)))
print('Overall model accuracy: {}'.format(gboost_accuracy))
# +
# we now tune the parameters of the GradientBoostingRegressor model using RandomizedSearchCV to
# find the best parameters and increase the accuracy of the model
params['learning_rate'] = np.linspace(0.1,1,10)
# instantiate the model
gradient_boosting = GradientBoostingRegressor()
# perform the grid search for the best parameters
gboost_search = RandomizedSearchCV(gradient_boosting,params,n_jobs=-1,
cv=5,verbose=2)
gboost_search.fit(xtrain,ytrain)
# +
# we now score the model
print('Score on train data: {}\n'.format(gboost_search.score(xtrain,ytrain)))
print('Score on test data: {}\n'.format(gboost_search.score(xtest,ytest)))
print('Best parameters found:')
display(gboost_search.best_params_)
gboost_search_pred = gboost_search.predict(xtest)
gboost_search_mse = mean_squared_error(ytest,gboost_search_pred)
gboost_search_accuracy = r2_score(ytest,gboost_search_pred)
print('Root Mean Squared Error: {}\n'.format(np.sqrt(gboost_search_mse)))
print('Overall model accuracy: {}'.format(gboost_search_accuracy))
# +
# we now use the best model (GradientBoostingRegressor model) to predict the PM2.5
# concetration and compare it to the actual PM2.5 recorded in the data by means of
# visualization
compare_data = pd.DataFrame({'dates':data['date'],
'Actual PM2.5':y,
'Predicted PM2.5':gboost_search.predict(X.values)})
compare_data.set_index('dates',inplace=True)
compare_data['Predicted PM2.5'] = np.round(compare_data['Predicted PM2.5'],1)
# let's plot the daily averages of the Actual PM10 and the predicted PM2.5 concentration.
compare_data = compare_data.resample('D').mean()
with plt.style.context('fivethirtyeight'):
plt.figure(figsize=(12,5))
plt.scatter(compare_data.index,compare_data['Actual PM2.5'],s=15,label='Actual PM2.5',
alpha=.6)
plt.scatter(compare_data.index,compare_data['Predicted PM2.5'],s=15,label='Predicted PM2.5',
alpha=.6)
plt.legend()
plt.title('Evaluating the GradientBoostingRegressor model\n(model accuracy = 93%)',
fontsize=18)
plt.xlabel('period',fontsize=15)
plt.ylabel('PM2.5 concentration',fontsize=15)
plt.show()
# -
# <h5><b>Residuals analysis</b></h5>
# <p>Now that we have successfully trained a regression model that predicts the amount of PM2.5 concetration in the air with a 93% accuracy given other environmental features, we have to analyze the errors of prediction to see if the model satisfies the regression errors asumption. That is,the errors must be normally distributed and do not follow any pattern.</p>
# +
# calculate the errors
compare_data['Residuals'] = compare_data['Actual PM2.5'] - compare_data['Predicted PM2.5']
# make a scatter plot of the errors to see if they follow any pattern
with plt.style.context('ggplot'):
plt.figure(figsize=(12,5))
plt.scatter(compare_data.index,compare_data.Residuals,alpha=.7)
plt.title('Residual scatter plot',fontsize=16)
plt.ylabel('Errors',fontsize=15)
plt.grid(axis='x')
plt.show()
# -
# plot the histogram to see check the normality of the errors
plt.figure(figsize=(12,5))
sns.distplot(compare_data.Residuals,bins=50)
plt.title('Distribution of the residuals from the gradient boosting model',
fontsize=16)
plt.show()
# <p>As seen above, the residuals of the predictions follow no pattern and also have a normal distribution which satisfies the regression error assumptions. This proves that our model is accurate and good for further predictions.</p>
# <br>
# <h3><b>Saving and loading the model</b></h3>
# <p>Now that the model has been successfully trained, the next thing is to save and be able to load it anytime we want to use it. To do this, we have to save it as a pickle file using the joblib module from sklearn.externals. The cells below shows you how to save your trained machine learning model and also load it anytime you want.</p>
from sklearn.externals import joblib
# save the model to my desktop
joblib.dump(gboost_search,'.\\Desktop\\pm25_model.pkl')
# loading the model
regression_model = joblib.load('.\\Desktop\\pm25_model.pkl')
# <br>
# <br>
# <br>
# <h4><b>Project completed by: <NAME></b></h4>
# <p><a href="mailto:<EMAIL>" target="_blank">Email</a> || <a href="https://www.linkedin.com/in/prince-owusu-356914198?lipi=urn%3Ali%3Apage%3Ad_flagship3_profile_view_base_contact_details%3B2NYoXqMHQKOMp0yWSME5mQ%3D%3D" target="_blank">LinkedIn</a> || <a href="https://twitter.com/iam_kwekhu" target="_blank">Twitter</a></p>
| Beijin Air quality (predicting PM2.5).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Run the demo
#
# For FuxiCTR v1.0.x only.
#
# We provide [multiple demo scripts](https://github.com/xue-pai/FuxiCTR/tree/main/demo) to run a given model on the tiny dataset. Please follow these examples to get started. The code workflow is structured as follows:
#
# ```python
# # Set data params and model params
# params = {...}
#
# # Set the feature encoding specs
# feature_encoder = FeatureEncoder(feature_cols, label_col, ...) # define the feature encoder
# feature_encoder.fit(...) # fit and transfrom the data
#
# # Load data generators
# train_gen, valid_gen, test_gen = data_generator(feature_encoder, ...)
#
# # Define a model
# model = DeepFM(...)
#
# # Train the model
# model.fit_generator(train_gen, validation_data=valid_gen, ...)
#
# # Evaluation
# model.evaluate_generator(test_gen)
#
# ```
# .
#
# In the following, we show the demo `DeepFM_demo.py`.
import sys
import os
from fuxictr.datasets import data_generator
from fuxictr.datasets.taobao import FeatureEncoder
from datetime import datetime
from fuxictr.utils import set_logger, print_to_json
import logging
from fuxictr.pytorch.models import DeepFM
from fuxictr.pytorch.utils import seed_everything
# After importing the required packages, one needs to define the params dict for DeepFM.
# +
feature_cols = [{'name': ["userid","adgroup_id","pid","cate_id","campaign_id","customer","brand","cms_segid",
"cms_group_id","final_gender_code","age_level","pvalue_level","shopping_level","occupation"],
'active': True, 'dtype': 'str', 'type': 'categorical'}]
label_col = {'name': 'clk', 'dtype': float}
params = {'model_id': 'DeepFM_demo',
'dataset_id': 'tiny_data_demo',
'train_data': '../data/tiny_data/train_sample.csv',
'valid_data': '../data/tiny_data/valid_sample.csv',
'test_data': '../data/tiny_data/test_sample.csv',
'model_root': '../checkpoints/',
'data_root': '../data/',
'feature_cols': feature_cols,
'label_col': label_col,
'embedding_regularizer': 0,
'net_regularizer': 0,
'hidden_units': [64, 64],
'hidden_activations': "relu",
'learning_rate': 1e-3,
'net_dropout': 0,
'batch_norm': False,
'optimizer': 'adam',
'task': 'binary_classification',
'loss': 'binary_crossentropy',
'metrics': ['logloss', 'AUC'],
'min_categr_count': 1,
'embedding_dim': 10,
'batch_size': 16,
'epochs': 3,
'shuffle': True,
'seed': 2019,
'monitor': 'AUC',
'monitor_mode': 'max',
'use_hdf5': True,
'pickle_feature_encoder': True,
'save_best_only': True,
'every_x_epochs': 1,
'patience': 2,
'workers': 1,
'verbose': 0,
'version': 'pytorch',
'gpu': -1}
# Set the logger and random seed
set_logger(params)
logging.info('Start the demo...')
logging.info(print_to_json(params))
seed_everything(seed=params['seed'])
# -
# Then set the FeatureEncoder to fit the training data and encode the raw features (e.g., normalizing continious values and mapping/reindex categorical features) from csv files.
feature_encoder = FeatureEncoder(feature_cols,
label_col,
dataset_id=params['dataset_id'],
data_root=params["data_root"],
version=params['version'])
feature_encoder.fit(train_data=params['train_data'],
min_categr_count=params['min_categr_count'])
# Preprocess the csv files to h5 files and get the data generators ready for train/validation/test. Note that the h5 files can be reused for subsequent experiments directly.
train_gen, valid_gen, test_gen = data_generator(feature_encoder,
train_data=params['train_data'],
valid_data=params['valid_data'],
test_data=params['test_data'],
batch_size=params['batch_size'],
shuffle=params['shuffle'],
use_hdf5=params['use_hdf5'])
# Initialize a DeepFM model and fit the model with the training and validation data.
model = DeepFM(feature_encoder.feature_map, **params)
model.fit_generator(train_gen, validation_data=valid_gen, epochs=params['epochs'],
verbose=params['verbose'])
# Reload the saved best model checkpoint for testing.
logging.info('***** validation/test results *****')
model.load_weights(model.checkpoint)
model.evaluate_generator(valid_gen)
model.evaluate_generator(test_gen)
| tutorials/v1.0/1_run_the_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# ----------
# User Instructions:
#
# Write a function optimum_policy that returns
# a grid which shows the optimum policy for robot
# motion. This means there should be an optimum
# direction associated with each navigable cell from
# which the goal can be reached.
#
# Unnavigable cells as well as cells from which
# the goal cannot be reached should have a string
# containing a single space (' '), as shown in the
# previous video. The goal cell should have '*'.
# ----------
grid = [[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 0, 0, 0, 1, 0]]
init = [0, 0]
goal = [len(grid)-1, len(grid[0])-1]
cost = 1 # the cost associated with moving from a cell to an adjacent one
delta = [[-1, 0 ], # go up
[ 0, -1], # go left
[ 1, 0 ], # go down
[ 0, 1 ]] # go right
delta_name = ['^', '<', 'v', '>']
def optimum_policy(grid,goal,cost):
# ----------------------------------------
# modify code below
# ----------------------------------------
value = [[99 for row in range(len(grid[0]))] for col in range(len(grid))]
# Policy table for printing delta output
policy = [[' ' for col in range(len(grid[0]))] for row in range(len(grid))]
change = True
while change:
change = False
for x in range(len(grid)):
for y in range(len(grid[0])):
if goal[0] == x and goal[1] == y:
if value[x][y] > 0:
value[x][y] = 0
policy[x][y] = '*' # goal location
change = True
elif grid[x][y] == 0:
for a in range(len(delta)):
x2 = x + delta[a][0]
y2 = y + delta[a][1]
if x2 >= 0 and x2 < len(grid) and y2 >= 0 and y2 < len(grid[0]) and grid[x2][y2] == 0:
v2 = value[x2][y2] + cost
if v2 < value[x][y]:
value[x][y] = v2
policy[x][y] = delta_name[a] # memorize action
change = True
print(policy[0])
print(policy[1])
print(policy[2])
print(policy[3])
print(policy[4])
print("")
# # Show actionable path
# for i in range(len(policy)):
# print(policy[i])
return policy
optimum_policy(grid,goal,cost)
# +
##### Do Not Modify ######
import grader
from test import delta, delta_name
try:
response = grader.run_grader(search)
print(response)
except Exception as err:
print(str(err))
# -
| optimum-policy_quiz/optimum-policy_quiz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="vtp9xikALsD0" colab_type="code" colab={}
from keras.models import Model
from keras.layers import Flatten, Dense, Input, BatchNormalization
from keras.layers import Convolution2D, MaxPooling2D, AveragePooling2D
from keras.engine.topology import get_source_inputs
from keras.utils.layer_utils import convert_all_kernels_in_model
from keras.utils.data_utils import get_file
from keras import backend as K
from keras.layers import Dense, GlobalAveragePooling2D
from keras.layers.core import Dropout, Lambda
from keras.layers import merge
import warnings
from keras.layers.core import Activation
from keras.regularizers import l2
from keras.preprocessing.image import ImageDataGenerator
from keras.utils import np_utils
from keras.datasets import cifar10
import cv2
import numpy as np
# (x_train, y_train), (x_test, y_test) = cifar100.load_data()
# y_train = np_utils.to_categorical(y_train, 100)
# y_test = np_utils.to_categorical(y_test, 100)
# x_train = x_train.astype('float32')
# x_test = x_test.astype('float32')
# x_train /= 255
# x_test /= 255
# slim = tf.contrib.slim
# + id="wlMGah7N6UB7" colab_type="code" colab={}
import tensorflow as tf
slim = tf.contrib.slim
trunc_normal = lambda stddev: tf.truncated_normal_initializer(0.0, stddev)
# + id="rd6DJAuv_VHj" colab_type="code" colab={}
def inception_v3_base(inputs,
final_endpoint='Mixed_7c',
min_depth=16,
depth_multiplier=1.0,
scope=None):
end_points = {}
if depth_multiplier <= 0:
raise ValueError('depth_multiplier is not greater than zero.')
depth = lambda d: max(int(d * depth_multiplier), min_depth)
with tf.variable_scope(scope, 'InceptionV3', [inputs]):
with slim.arg_scope([slim.conv2d, slim.max_pool2d, slim.avg_pool2d],
stride=1, padding='VALID'):
# 299 x 299 x 3
end_point = 'Conv2d_1a_3x3'
net = slim.conv2d(inputs, depth(32), [3, 3], strides=2, scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 149 x 149 x 32
end_point = 'Conv2d_2a_3x3'
net = slim.conv2d(net, depth(32), [3, 3], scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 147 x 147 x 32
end_point = 'Conv2d_2b_3x3'
net = slim.conv2d(net, depth(64), [3, 3], padding='SAME', scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 147 x 147 x 64
end_point = 'MaxPool_3a_3x3'
net = slim.max_pool2d(net, [3, 3], stride=2, scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 73 x 73 x 64
end_point = 'Conv2d_3b_1x1'
net = slim.conv2d(net, depth(80), [1, 1], scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 73 x 73 x 80
end_point = 'Conv2d_4a_3x3'
net = slim.conv2d(net, depth(192), [3, 3], scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 71 x 71 x 192
end_point = 'MaxPool_5a_3x3'
net = slim.max_pool2d(net, [3, 3], stride=2, scope=end_point)
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# 35 x 35 x 192
with slim.arg_scope([slim.conv2d, slim.max_pool2d, slim.avg_pool2d], stride=1, padding='SAME'):
# mixed: 35 x 35 x 256
end_point = 'Mixed_5b'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(48), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(64), [5, 5], scope='Conv2d_0b_5x5')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(96), [3, 3], scope='Conv2d_0b_3x3')
branch_2 = slim.conv2d(branch_2, depth(96), [3, 3], scope='Conv2d_0c_3x3')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(32), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint:
return net, end_points
# mixed_1: 35 x 35 x 288
end_point = 'Mixed_5c'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(48), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(64), [5, 5], scope='Conv2d_0b_5x5')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(96), [3, 3], scope='Conv2d_0b_3x3')
branch_2 = slim.conv2d(branch_2, depth(96), [3, 3], scope='Conv2d_0c_3x3')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(64), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, value=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_2: 35 x 35 x 288
end_point = 'Mixed_5d'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(48), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(64), [5, 5], scope='Conv2d_0b_5x5')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(96), [3, 3], scope='Conv2d_0b_3x3')
branch_2 = slim.conv2d(branch_2, depth(96), [3, 3], scope='Conv2d_0c_3x3')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(64), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_3 : 17 x 17 x 768
end_point = 'Mixed_6a'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(384), [3, 3], stride=2, padding='VALID', scope='Conv2d_1a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(64), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(96), [3, 3], scope='Conv2d_0b_3x3')
branch_1 = slim.conv2d(branch_1, depth(96), [3, 3], stride=2, padding='VALID', scope='Conv2d_1a_1x1')
with tf.variable_scope('Branch_2'):
branch_2 = slim.max_pool2d(net, [3, 3], stride=2, padding='VALID', scope='MaxPool_1a_3x3')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_4 : 17 x 17 x 768
end_point = 'Mixed_6b'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(128), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(128), [1, 7], scope='Conv2d_0b_1x7')
branch_1 = slim.conv2d(branch_1, depth(192), [7, 1], scope='Conv2d_0c_7x1')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(128), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(128), [7, 1], scope='Conv2d_0b_7x1')
branch_2 = slim.conv2d(branch_2, depth(128), [1, 7], scope='Conv2d_0c_1x7')
branch_2 = slim.conv2d(branch_2, depth(128), [7, 1], scope='Conv2d_0d_7x1')
branch_2 = slim.conv2d(branch_2, depth(128), [1, 7], scope='Conv2d_0e_1x7')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(192), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_5 : 17 x 17 x 768
end_point = 'Mixed_6c'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(160), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(160), [1, 7], scope='Conv2d_0b_1x7')
branch_1 = slim.conv2d(branch_1, depth(192), [7, 1], scope='Conv2d_0c_7x1')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(160), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(160), [7, 1], scope='Conv2d_0b_7x1')
branch_2 = slim.conv2d(branch_2, depth(160), [1, 7], scope='Conv2d_0c_1x7')
branch_2 = slim.conv2d(branch_2, depth(160), [7, 1], scope='Conv2d_0d_7x1')
branch_2 = slim.conv2d(branch_2, depth(160), [1, 7], scope='Conv2d_0e_1x7')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(192), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_6 : 17 x 17 x 768
end_point = 'Mixed_6d'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(160), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(160), [1, 7], scope='Conv2d_0b_1x7')
branch_1 = slim.conv2d(branch_1, depth(160), [7, 1], scope='Conv2d_0c_7x1')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(160), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(160), [7, 1], scope='Conv2d_0b_7x1')
branch_2 = slim.conv2d(branch_2, depth(160), [1, 7], scope='Conv2d_0c_1x7')
branch_2 = slim.conv2d(branch_2, depth(160), [7, 1], scope='Conv2d_0d_7x1')
branch_2 = slim.conv2d(branch_2, depth(160), [1, 7], scope='Conv2d_0e_1x7')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(192), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_7 : 17 x 17 x 768
end_point = 'Mixed_6e'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(192), [1, 7], scope='Conv2d_0b_1x7')
branch_1 = slim.conv2d(branch_1, depth(192), [7, 1], scopee='Conv2d_0c_7x1')
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(192), [7, 1], scope='Conv2d_0b_7x1')
branch_2 = slim.conv2d(branch_2, depth(192), [1, 7], scope='Conv2d_0c_1x7')
branch_2 = slim.conv2d(branch_2, depth(192), [7, 1], scope='Conv2d_0d_7x1')
branch_2 = slim.conv2d(branch_2, depth(192), [1, 7], scope='Conv2d_0e_1x7')
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(192), [1, 1], scopee='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_8: 8 x 8 x 1280
end_point = 'Mixed_7a'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
branch_0 = slim.conv2d(branch_0, depth(320), [3, 3], stride=2, padding='VALID', scope='Conv2d_1a_3x3')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(192), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = slim.conv2d(branch_1, depth(192), [1, 7], scope='Conv2d_0b_1x7')
branch_1 = slim.conv2d(branch_1, depth(192), [7, 1], scope='Conv2d_0c_7x1')
branch_1 = slim.conv2d(branch_1, depth(192), [3, 3], stride=2, padding='VALID', scope='Conv2d_1a_3x3')
with tf.variable_scope('Branch_2'):
branch_2 = slim.max_pool2d(net, [3, 3], stride=2, padding='VALID', scope='MaxPool_1a_3x3')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_9: 8 x 8 x 2048
end_point = 'Mixed_7b'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(320), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(384), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = tf.concat(axis=3, values=[
slim.conv2d(branch_1, depth(384), [1, 3], scope='Conv2d_0b_1x3'),
slim.conv2d(branch_1, depth(384), [3, 1], scope='Conv2d_0b_3x1')
])
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(448), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(384), [3, 3], scope='Conv2d_0b_3x3')
branch_2 = tf.concat(axis=3, values=[
slim.conv2d(branch_2, depth(384), [1, 3], scope='Conv2d_0c_1x3'),
slim.conv2d(branch_2, deppth(384), [3, 1], scope='Conv2d_0d_3x1')
])
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(192), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
# mixed_10 : 8 x 8 x 2048
end_point = 'Mixed_7c'
with tf.variable_scope(end_point):
with tf.variable_scope('Branch_0'):
branch_0 = slim.conv2d(net, depth(320), [1, 1], scope='Conv2d_0a_1x1')
with tf.variable_scope('Branch_1'):
branch_1 = slim.conv2d(net, depth(384), [1, 1], scope='Conv2d_0a_1x1')
branch_1 = tf.concat(axis=3, values=[
slim.conv2d(branch_1, depth(384), [1, 3], scope='Conv2d_0a_1x3'),
slim.conv2d(branch_1, depth(384), [3, 1], scope='Conv2d_0b_3x1')
])
with tf.variable_scope('Branch_2'):
branch_2 = slim.conv2d(net, depth(448), [1, 1], scope='Conv2d_0a_1x1')
branch_2 = slim.conv2d(branch_2, depth(384), [3, 3], scope='Conv2d_0b_3x3')
branch_2 = tf.concat(axis=3, values=[
slim.conv2d(branch_2, depth(384), [1, 3], scope='Conv2d_0c_1x3'),
slim.conv2d(branch_2, depth(384), [3, 1], scope='Conv2d_0d_3x1')
])
with tf.variable_scope('Branch_3'):
branch_3 = slim.avg_pool2d(net, [3, 3], scope='AvgPool_0a_3x3')
branch_3 = slim.conv2d(branch_3, depth(192), [1, 1], scope='Conv2d_0b_1x1')
net = tf.concat(axis=3, values=[branch_0, branch_1, branch_2, branch_3])
end_points[end_point] = net
if end_point == final_endpoint: return net, end_points
raise ValueError('Unknown final endpoint %s' % final_endpoint)
def inception_v3(inputs,
num_classes=1000,
is_training=True,
dropout_keep_prob=0.8,
min_depth=16,
depth_multiplier=1.0,
prediction_fn=slim.softmax,
spatial_squeeze=True,
reuse=None,
create_aux_logits=True,
scope='InceptionV3',
global_pool=False):
if depth_multiplier <= 0:
raise ValueError('depth_multiplier is not greater than zero.')
depth = lambda d: max(int(d * depth_multiplier), min_depth)
with tf.variable_scope(scope, 'InceptionV3', [inputs], reuse=reuse) as scope:
with slim.arg_scope([slim.batch_norm, slim.dropout],
is_training=is_training):
net, end_points = inception_v3_base(
inputs, scope=scope, min_depth=min_depth,
depth_multiplier=depth_multiplier)
if create_aux_logits and num_classes:
with slim.arg_scope([slim.conv2d, slim.max_pool2d, slim.avg_pool2d],
stride=1, padding='SAME'):
aux_logits = end_points['Mixed_6e']
with tf.variable_scope('AuxLogits'):
aux_logits = slim.avg_pool2d(
aux_logits, [5, 5], stride=3, padding='VALID',
scope='AvgPool_1a_5x5')
aux_logits = slim.conv2d(aux_logits, depth(128), [1, 1],
scope='Conv2d_1b_1x1')
kernel_size = _reduced_kernel_size_for_small_input(
aux_logits, [5, 5])
aux_logits = slim.conv2d(
aux_logits, depth(768), kernel_size,
weights_initializer=trunc_normal(0.01),
padding='VALID', scope='Conv2d_2a_{}x{}'.format(*kernel_size))
aux_logits = slim.conv2d(
aux_logits, num_classes, [1, 1], activation_fn=None,
normalizer_fn=None, weights_initializer=trunc_normal(0.001),
scope='Conv2d_2b_1x1')
if spatial_squeeze:
aux_logits = tf.squeeze(aux_logits, [1, 2], name='SpatialSqueeze')
end_points['AuxLogits'] = aux_logits
with tf.variable_scope('Logits'):
if global_pool:
net = tf.reduce_mean(net, [1, 2], keep_dims=True, name='GlobalPool')
end_points['global_pool'] = net
else:
kernel_size = _reduced_kernel_size_for_small_input(net, [8, 8])
net = slim.avg_pool2d(net, kernel_size, padding='VALID',
scope='AvgPool_1a_{}x{}'.format(*kernel_size))
end_points['AvgPool_1a'] = net
if not num_classes:
return net, end_points
# 1 x 1 x 2048
net = slim.dropout(net, keep_prob=dropout_keep_prob, scope='Dropout_1b')
end_points['PreLogits'] = net
# 2048
logits = slim.conv2d(net, num_classes, [1, 1], activation_fn=None,
normalizer_fn=None, scope='Conv2d_1c_1x1')
if spatial_squeeze:
logits = tf.squeeze(logits, [1, 2], name='SpatialSqueeze')
# 1000
end_points['Logits'] = logits
end_points['Predictions'] = prediction_fn(logits, scope='Predictions')
return logits, end_points
inception_v3.default_image_size = 299
def inception_arg_scope(weight_decay=0.00004,
use_batch_norm=True,
batch_norm_decay=0.9997,
batch_norm_epsilon=0.001,
activation_fn=tf.nn.relu):
batch_norm_params = {
'decay': batch_norm_decay,
'epsilon': batch_norm_epsilon,
'updates_collections': tf.GraphKeys.UPDATE_OPS,
'fused': None,
}
if use_batch_norm:
normalizer_fn = slim.batch_norm
normalizer_params = batch_norm_params
else:
normalizer_fn = None
normalizer_params = {}
with slim.arg_scope([slim.conv2d, slim.fully_connected],
weights_regularizer=slim.l2_regularizer(weight_decay)):
with slim.arg_scope(
[slim.conv2d],
weights_initializer=slim.variance_scaling_initializer(),
activation_fn=activation_fn,
normalizer_fn=normalizer_fn,
normalizer_params=normalizer_params) as sc:
return sc
# + id="lycyMPH_rgjL" colab_type="code" colab={}
def conv2d_bn(x, nb_filter, nb_row, nb_col,
border_mode='same', subsample=(1, 1),
batch_norm=True, activation='relu',
weight_decay=0, name=None):
if name is not None:
bn_name = 'bn_' + name
conv_name = 'conv_' + name
else:
bn_name = None
conv_name = None
if K.image_dim_ordering() == 'th':
bn_axis = 1
else:
bn_axis = 3
if weight_decay and weight_decay > 0:
x = Convolution2D(nb_filter, nb_row, nb_col,
subsample=subsample,
activation='relu',
W_regularizer=l2(weight_decay),
border_mode=border_mode,
name=conv_name)(x)
else:
x = Convolution2D(nb_filter, nb_row, nb_col,
subsample=subsample,
activation='relu',
border_mode=border_mode,
name=conv_name)(x)
if batch_norm:
x = BatchNormalization(axis=bn_axis, name=bn_name)(x)
if activation:
x = Activation(activation)(x)
return x
# + id="1yybWobi_fyz" colab_type="code" colab={}
def Inception_V3(include_top=True, weights='imagenet',
input_tensor=None, input_shape=None,
weight_decay=0.00004, num_classes=1000,
dropout_prob=0., aux_include=True):
if input_shape is None:
input_shape = (299, 299)
if K.image_dim_ordering() == 'th':
input_shape = (3,) + input_shape
channel_axis = 1
else:
input_shape = input_shape + (3,)
channel_axis = 3
if input_tensor is None:
img_input = Input(shape=input_shape)
else:
img_input = input_tensor
# Using `tf` order
# 299 x 299 x 3
x = conv2d_bn(img_input, 32, 3, 3, subsample=(2, 2),
border_mode='valid', weight_decay=weight_decay,
name='0')
# 149 x 149 x 32
x = conv2d_bn(x, 32, 3, 3, border_mode='valid',
weight_decay=weight_decay, name='1')
# 147 x 147 x 32
x = conv2d_bn(x, 64, 3, 3, weight_decay=weight_decay, name='2')
# 147 x 147 x 64
x = MaxPooling2D(pool_size=(3, 3), strides=(2, 2), name='pool_1')(x)
# 73 x 73 x 64
x = conv2d_bn(x, 80, 1, 1, weight_decay=weight_decay, name='3')
# 73 x 73 x 80
x = conv2d_bn(x, 192, 3, 3, border_mode='valid',
weight_decay=weight_decay, name='4')
# 71 x 71 x 192
x = MaxPooling2D(pool_size=(3, 3), strides=(2, 2), name='pool_2')(x)
# 35 x 35 x 192
# mixed 0: 35 x 35 x 256
branch1x1 = conv2d_bn(x, 64, 1, 1, weight_decay=weight_decay)
branch5x5 = conv2d_bn(x, 48, 1, 1, weight_decay=weight_decay)
branch5x5 = conv2d_bn(branch5x5, 64, 5, 5, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(x, 64, 1, 1, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(branch3x3dbl, 96, 3, 3, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(branch3x3dbl, 96, 3, 3, weight_decay=weight_decay)
branch_pool = AveragePooling2D(
(3, 3), strides=(1, 1), border_mode='same')(x)
branch_pool = conv2d_bn(branch_pool, 32, 1, 1, weight_decay=weight_decay)
x = merge([branch1x1, branch5x5, branch3x3dbl, branch_pool],
mode='concat', concat_axis=channel_axis,
name='mixed_0')
for i in range(2):
branch1x1 = conv2d_bn(x, 64, 1, 1, weight_decay=weight_decay)
branch5x5 = conv2d_bn(x, 48, 1, 1, weight_decay=weight_decay)
branch5x5 = conv2d_bn(branch5x5, 64, 5, 5, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(x, 64, 1, 1, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(
branch3x3dbl, 96, 3, 3, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(
branch3x3dbl, 96, 3, 3, weight_decay=weight_decay)
branch_pool = AveragePooling2D(
(3, 3), strides=(1, 1), border_mode='same')(x)
branch_pool = conv2d_bn(
branch_pool, 64, 1, 1, weight_decay=weight_decay)
x = merge([branch1x1, branch5x5, branch3x3dbl, branch_pool],
mode='concat', concat_axis=channel_axis,
name='mixed_' + str(i + 1))
# mixed_3: 17 x 17 x 768
branch3x3 = conv2d_bn(
x, 384, 3, 3, subsample=(2, 2),
border_mode='valid', weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(x, 64, 1, 1, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(branch3x3dbl, 96, 3, 3, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(
branch3x3dbl, 96, 3, 3, subsample=(2, 2),
border_mode='valid', weight_decay=weight_decay)
branch_pool = MaxPooling2D((3, 3), strides=(2, 2), border_mode='valid')(x)
x = merge([branch3x3, branch3x3dbl, branch_pool], mode='concat',
concat_axis=channel_axis, name='mixed_3')
# mixed_4: 17 x 17 x 768
branch1x1 = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch7x7 = conv2d_bn(x, 128, 1, 1, weight_decay=weight_decay)
branch7x7 = conv2d_bn(branch7x7, 128, 1, 7, weight_decay=weight_decay)
branch7x7 = conv2d_bn(branch7x7, 192, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(x, 128, 1, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 128, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 128, 1, 7, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 128, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 192, 1, 7, weight_decay=weight_decay)
branch_pool = AveragePooling2D(
(3, 3), strides=(1, 1), border_mode='same')(x)
branch_pool = conv2d_bn(branch_pool, 192, 1, 1, weight_decay=weight_decay)
x = merge([branch1x1, branch7x7, branch7x7dbl, branch_pool],
mode='concat', concat_axis=channel_axis, name='mixed_4')
# mixed 5, 6: 17 x 17 x 768
for i in range(2):
branch1x1 = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch7x7 = conv2d_bn(x, 160, 1, 1, weight_decay=weight_decay)
branch7x7 = conv2d_bn(branch7x7, 160, 1, 7, weight_decay=weight_decay)
branch7x7 = conv2d_bn(branch7x7, 192, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(x, 160, 1, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 160, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 160, 1, 7, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 160, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 192, 1, 7, weight_decay=weight_decay)
branch_pool = AveragePooling2D(
(3, 3), strides=(1, 1), border_mode='same')(x)
branch_pool = conv2d_bn(
branch_pool, 192, 1, 1, weight_decay=weight_decay)
x = merge([branch1x1, branch7x7, branch7x7dbl, branch_pool],
mode='concat', concat_axis=channel_axis,
name='mixed_' + str(i + 5))
# mixed 7: 17 x 17 x 768
branch1x1 = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch7x7 = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch7x7 = conv2d_bn(branch7x7, 192, 1, 7, weight_decay=weight_decay)
branch7x7 = conv2d_bn(branch7x7, 192, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 192, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 192, 1, 7, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 192, 7, 1, weight_decay=weight_decay)
branch7x7dbl = conv2d_bn(
branch7x7dbl, 192, 1, 7, weight_decay=weight_decay)
branch_pool = AveragePooling2D(
(3, 3), strides=(1, 1), border_mode='same')(x)
branch_pool = conv2d_bn(branch_pool, 192, 1, 1, weight_decay=weight_decay)
x = merge([branch1x1, branch7x7, branch7x7dbl, branch_pool],
mode='concat', concat_axis=channel_axis,
name='mixed_7')
if aux_include:
aux_classifier = AveragePooling2D(
(5, 5), strides=(3, 3), border_mode='valid')(x)
aux_classifier = conv2d_bn(
aux_classifier, 128, 1, 1, weight_decay=weight_decay)
aux_classifier = conv2d_bn(aux_classifier, 768, 5, 5,
border_mode='valid',
weight_decay=weight_decay)
aux_classifier = Flatten()(aux_classifier)
if weight_decay and weight_decay > 0:
aux_classifier = Dense(num_classes, activation='softmax',
W_regularizer=l2(weight_decay),
name='aux_classifier')(aux_classifier)
else:
aux_classifier = Dense(
num_classes, activation='softmax',
name='aux_classifier')(aux_classifier)
# mixed 8: 8 x 8 x 1280.
branch3x3 = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch3x3 = conv2d_bn(branch3x3, 320, 3, 3,
subsample=(2, 2),
border_mode='valid',
weight_decay=weight_decay)
branch7x7x3 = conv2d_bn(x, 192, 1, 1, weight_decay=weight_decay)
branch7x7x3 = conv2d_bn(branch7x7x3, 192, 1, 7, weight_decay=weight_decay)
branch7x7x3 = conv2d_bn(branch7x7x3, 192, 7, 1, weight_decay=weight_decay)
branch7x7x3 = conv2d_bn(branch7x7x3, 192, 3, 3,
subsample=(2, 2),
border_mode='valid',
weight_decay=weight_decay)
branch_pool = MaxPooling2D((3, 3), strides=(2, 2), border_mode='valid')(x)
x = merge([branch3x3, branch7x7x3, branch_pool],
mode='concat', concat_axis=channel_axis,
name='mixed_8')
# mixed 9 10: 8 x 8 x 2048
for i in range(2):
branch1x1 = conv2d_bn(x, 320, 1, 1, weight_decay=weight_decay)
branch3x3 = conv2d_bn(x, 384, 1, 1, weight_decay=weight_decay)
branch3x3 = merge([conv2d_bn(branch3x3, 384, 1, 3,
weight_decay=weight_decay),
conv2d_bn(branch3x3, 384, 3, 1,
weight_decay=weight_decay)],
mode='concat', concat_axis=channel_axis)
branch3x3dbl = conv2d_bn(x, 448, 1, 1, weight_decay=weight_decay)
branch3x3dbl = conv2d_bn(
branch3x3dbl, 384, 3, 3, weight_decay=weight_decay)
branch3x3dbl = merge([conv2d_bn(branch3x3dbl, 384, 1, 3,
weight_decay=weight_decay),
conv2d_bn(branch3x3dbl, 384, 3, 1,
weight_decay=weight_decay)],
mode='concat', concat_axis=channel_axis)
branch_pool = AveragePooling2D(
(3, 3), strides=(1, 1), border_mode='same')(x)
branch_pool = conv2d_bn(
branch_pool, 192, 1, 1, weight_decay=weight_decay)
x = merge([branch1x1, branch3x3, branch3x3dbl, branch_pool],
mode='concat', concat_axis=channel_axis,
name='mixed_' + str(9 + i))
# 2048 x 8 x 8
x = conv2d_bn(x, 1024, 1, 1,
weight_decay=weight_decay)
# 1024 x 8 x 8
x = GlobalAveragePooling2D()(x)
x = Dropout(dropout_prob)(x)
# 1024
if weight_decay and weight_decay > 0:
predictions = Dense(num_classes,
activation='softmax',
W_regularizer=l2(weight_decay),
name='predictions')(x)
else:
predictions = Dense(num_classes,
activation='softmax',
name='predictions')(x)
if input_tensor is not None:
inputs = get_source_inputs(input_tensor)
else:
inputs = img_input
if aux_include:
model = Model(
inputs, [predictions, aux_classifier],
name='inception_v3_with_aux')
else:
model = Model(inputs, predictions, name='inception_v3')
return model
# + id="17JtCy3aoYWk" colab_type="code" colab={}
def load_data():
(_, _), (x_train, y_train) = cifar10.load_data()
x_train = x_train[:100]
y_train = y_train[:100]
print(x_train.shape)
data_upscaled = np.zeros((100, 3, 299, 299))
for i, img in enumerate(x_train):
#im = img.transpose((1, 2, 0))
large_img = cv2.resize(img, dsize=(299, 299), interpolation=cv2.INTER_CUBIC)
#data_upscaled[i] = large_img.transpose((2, 0, 1))
y_train = np_utils.to_categorical(y_train, 10)
return data_upscaled, y_train
# + id="CG3ZKg6fonGe" colab_type="code" outputId="cbae7c02-cd2f-4bed-bc2b-d50a1f50ea49" colab={"base_uri": "https://localhost:8080/", "height": 589}
x_train, y_train = load_data()
model = Inception_V3(num_classes=10)
model.compile(optimizer='rmsprop',
loss={'predictions': 'categorical_crossentropy',
'aux_classifier': 'categorical_crossentropy'},
loss_weights={'predictions': 1., 'aux_classifier': 0.2})
# + id="_NCZ_7QDRBfY" colab_type="code" colab={}
input_shape = x_train.shape[1:]
n_classes = 100
model = inception_v3(input_shape, n_classes)
sgd = optimizers.SGD(lr=0.045, decay=0.94, momentum=0.9, nesterov=True)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="TyDb3-FiRBuJ" colab_type="code" colab={}
datagen = ImageDataGenerator(horizontal_flip=True, width_shift_range=0.1, height_shift_range=0.1)
datagen.fit(x_train)
model.fit_generator(datagen.flow(x_train, y_train, batch_size=64),
steps_per_epoch=x_train.shape[0] // 64,
epochs=60,
verbose=1,
validation_data=(x_test, y_test))
model.evaluate(x_test, y_test)
| Implementation/Image_Classification/Inception/Inception_v3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/victorog17/soulcode_aulas_python/blob/main/Soulcode_Python_001_aula_introdutoria.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="IRkY_xkAQXr8"
#Declarar uma variavel em Python
idade = 27
nome = 'Maria'
nome2 = 'João'
altura = 1.72
# + colab={"base_uri": "https://localhost:8080/"} id="bPtEtUKgeFMA" outputId="2cb59bbd-301b-46da-887a-153eacf26a62"
type(altura)
# + id="mPZ1ZcxIeG3h"
def soma(n1, n2):
total = n1 + n2
return print(f'A soma entre {n1} e {n2} é igual a {total}')
# + colab={"base_uri": "https://localhost:8080/"} id="3B4X0cc6mqBg" outputId="538a4d73-1fa6-4db1-b34d-2eaeb4fb06df"
soma(3, 5)
# + id="324p42-omsJq"
def contas(salario, despesas):
x = (despesas / salario) * 100
return print(f'Sendo que seu salário é de R${salario:.2f} e suas despesas são R${despesas:.2f}\nSuas despesas correspondem a {x:.2f}% do seu salário')
# + colab={"base_uri": "https://localhost:8080/"} id="kLO2AzphzVQr" outputId="50e776ba-a42d-43b1-8b61-f126d4756dcb"
contas(2000, 1438)
# + colab={"base_uri": "https://localhost:8080/"} id="MesurIsX0rAB" outputId="c23b0fb3-7ae5-40e9-b917-cb760e6fe70f"
n1 = int(input('Digite o número a ser dividido: '))
n2 = int(input('Digite o divisor:' ))
n3 = n1 // n2
n4 = n1 % n2
print(f'A divisão inteira de {n1} por {n2} é igual a {n3} com resto de {n4}')
# + colab={"base_uri": "https://localhost:8080/"} id="IPGqiza--dLa" outputId="17879e91-e361-4900-850d-770392e4256c"
x1 = int(input('Digite o número que quer elevar ao cubo: '))
x2 = pow(x1, 3)
print(f'{x1} ao cubo é igual a {x2}')
# + id="60qJ5Jh1_HZg"
| Soulcode_Python_001_aula_introdutoria.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !ls -lh ../data
# !head ../data/supp_table_haplotype_panel.csv
import pandas as pd
df_haps = pd.read_csv('../data/supp_table_haplotype_panel.csv')
df_haps.head()
len(df_haps[df_haps.hierarchy_haplotype_group.str.startswith('F', na=False)])
len(df_haps[df_haps.core_haplotype.str.startswith('F', na=False)])
743 / 775
len(df_haps[df_haps.hierarchy_haplotype_group.str.startswith('S', na=False)])
len(df_haps[df_haps.core_haplotype.str.startswith('S', na=False)])
425 / 430
df_haps.hierarchy_haplotype_group.value_counts()
df_haps.core_haplotype.value_counts()
# +
# %run setup.ipynb
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import hapclust
from scipy.cluster.hierarchy import _convert_to_double
from scipy.spatial import distance
from scipy.cluster.hierarchy import _hierarchy
# -
region_vgsc = '2L', 2358158, 2431617
def load_data(chrom, start=None, stop=None, n_variants=None):
# load data
callset_haps = np.load('../data/haps_phase1.npz')
haps = allel.HaplotypeArray(callset_haps['haplotypes'])
pos = allel.SortedIndex(callset_haps['POS'])
ann = callset_haps['ANN']
# locate the region of interest
if start and stop:
loc = pos.locate_range(start, stop)
elif start and n_variants:
start_idx = bisect.bisect_left(pos, start)
stop_idx = start_idx + n_variants
loc = slice(start_idx, stop_idx)
elif stop and n_variants:
stop_idx = bisect.bisect_right(pos, stop)
start_idx = stop_idx - n_variants
loc = slice(start_idx, stop_idx)
else:
raise ValueError('bad args')
# obtain haplotypes for the region of interest
pos = pos[loc]
h = haps[loc]
#setup missense
tbl_variants_selected = etl.frompickle('../data/tbl_variants_missense_selected.pkl')
tbl_selected_redux = (
tbl_variants_selected
.cut('POS', 'REF', 'ALT', 'AGAP004707-RA')
.mergeduplicates(key=('POS'))
.convert('ALT', lambda v: ','.join(v) if len(v) > 1 else v)
.addfield('label', lambda rec: '%s:%s>%s %s' % (rec.POS, rec.REF, rec.ALT.ljust(3), rec['AGAP004707-RA'].rjust(6)))
.sort('POS')
.selectin('AGAP004707-RA', {'L995F', 'L995S'})
)
# extract positions for the missense variants
pos_missense = allel.SortedIndex(tbl_selected_redux['POS'])
# extract haplotypes for the missense variants
missense_bool = np.in1d(pos, pos_missense)
h_missense = h.compress(missense_bool)
missense_mutations = list(tbl_selected_redux['AGAP004707-RA'])
return pos, h, h_missense, missense_mutations
pos, h, h_missense, missense_mutations = load_data(*region_vgsc)
h.shape
missense_mutations
def plot_missense_haplotypes(ax, h, mut_labels=missense_mutations):
h = h.copy()
# colours for colormap
mycol = ['r', 'w', 'k']
# alter rows with kdr mutations for color map to pick up
# known_muts = ['L995S (2984T>C)', 'L995F (2985A>T)', 'N1570Y (4708A>T)']
known_muts = ['L995S', 'L995F', 'N1570Y']
for mut in known_muts:
if mut in mut_labels:
h[mut_labels.index(mut)] *= -1
# make colormap
cake = mpl.colors.ListedColormap(mycol, name='mymap', N=3)
# plot
ax.pcolormesh(np.asarray(h[::-1]), cmap=cake, vmin=-1, vmax=1, zorder=-10)
ax.set_yticks(np.arange(h.shape[0])+.5)
lbl = [l for l in mut_labels[::-1]]
# lbl = ['%s' % l for l in mut_labels[::-1]]
ax.set_yticklabels(lbl, family='monospace', fontsize=6)
for ytick in ax.get_yticklabels():
if ytick.get_text() in known_muts:
ytick.set_color('r')
ax.hlines(np.arange(h.shape[0]+1), 0, h.shape[1], color='k', lw=.5)
ax.set_xlim(0, h.shape[1])
ax.set_ylim(0, h.shape[0])
# ax.set_xticks([])
ax.yaxis.tick_left()
ax.set_xticks([])
# rasterize to avoid SVG antialiasing issues and reduce file size
ax.set_rasterization_zorder(-5)
# ax.set_xticks(list(range(0, h.shape[1], 200)) + [h.shape[1]])
# ax.xaxis.tick_bottom()
# ax.spines['top'].set_visible(False)
# ax.spines['bottom'].set_visible(False)
fig, ax = plt.subplots()
plot_missense_haplotypes(ax, h_missense)
# +
def plot_dendrogram(h, ax, method='complete', color_threshold=0, above_threshold_color='k',
max_dist=None):
# compute distance matrix
dist = allel.stats.pairwise_distance(h, 'hamming') * h.shape[0]
# HACKING SCIPY TO GET TO OLD CLUSTERING METHOD
# https://github.com/scipy/scipy/blob/v0.18.1/scipy/cluster/hierarchy.py#L470-L667
# 1. fiddle with format
y = _convert_to_double(np.asarray(dist, order='c'))
# 2. get n
n = int(distance.num_obs_y(dist))
# 3. do clustering
method = dict(single=0, complete=1)[method]
z = _hierarchy.linkage(y, n, method)
# plot dendrogram
sns.despine(ax=ax, offset=5, bottom=True, top=False)
r = scipy.cluster.hierarchy.dendrogram(z, no_labels=True, count_sort=True,
color_threshold=color_threshold,
above_threshold_color=above_threshold_color,
ax=ax)
xmin, xmax = ax.xaxis.get_data_interval()
xticklabels = np.array([0, h.shape[1]])
xticks = xticklabels / h.shape[1]
xticks = (xticks * (xmax - xmin)) + xmin
ax.set_xticks(xticks)
ax.set_xticklabels(xticklabels)
ax.set_xlabel('Haplotypes', va='top')
ax.xaxis.set_label_position('top')
ax.set_ylim(bottom=-10, top=max_dist)
# ax.set_xlim(left=-10)
ax.set_ylabel('No. SNPs')
ax.autoscale(axis='x', tight=True)
return z, r
fig, ax = plt.subplots(figsize=(10, 5))
z, r = plot_dendrogram(h, ax);
# -
plt.plot(z[:, 2])
populations = phase1_ar3.pop_ids
pop_colours = phase1_ar3.pop_colors
pop_labels = phase1_ar3.pop_labels
df_haplotypes = phase1_ar31.df_haplotypes.query('population != "colony"')
df_haplotypes.head()
import numpy as np
import seaborn as sns
# +
def plot_dist_hist(h, ax=None, pop=None, max_dist=320):
if ax is None:
fig, ax = plt.subplots(figsize=(7, 4))
sns.despine(ax=ax, offset=5)
color = 'k'
if pop:
hap_ix = df_haplotypes.query('population == "{}"'.format(pop)).index.values
h = h.take(hap_ix, axis=1)
color = pop_colours[pop]
# compute distance matrix
dist = allel.stats.pairwise_distance(h, 'hamming') * h.shape[0]
print(pop, np.median(dist), np.percentile(dist, q=25), np.percentile(dist, q=75),
np.min(dist), np.max(dist))
ax.hist(dist, bins=np.arange(0, np.max(dist), 3), color=color)
ax.set_xlim(0, max_dist)
plot_dist_hist(h)
# -
df_haplotypes.population.value_counts()
plot_dist_hist(h, pop='GWA')
plot_dist_hist(h, pop='BFS')
plot_dist_hist(h, pop='UGS')
plot_dist_hist(h, pop='GAS')
x = allel.HaplotypeArray(h)
x
allel.haplotype_diversity(h)
# +
def fig_hap_structure(h, h_display=None, pop=None, figsize=(8, 2.5),
fn=None, dpi=120,
height_ratios=(10, 3),
width_ratios=(1, 1),
legend=True, max_dist=320):
# create the figure
fig = plt.figure(figsize=figsize, facecolor='white', dpi=dpi)
# define subplot layout
gs_nrows = 2
gs_ncols = 2
gs = mpl.gridspec.GridSpec(gs_nrows, gs_ncols, hspace=0.04, wspace=0.04,
height_ratios=height_ratios,
width_ratios=width_ratios)
# setup data
if pop:
hap_ix = df_haplotypes.query('population == "{}"'.format(pop)).index.values
h = h.take(hap_ix, axis=1)
if h_display is not None:
h_display = h_display.take(hap_ix, axis=1)
hap_pops = df_haplotypes.query('population == "{}"'.format(pop)).population.values
else:
hap_pops = df_haplotypes.population.values
# dendrogram
ax_dend = fig.add_subplot(gs[0, 0])
z, r = plot_dendrogram(h, ax_dend, color_threshold=0, max_dist=max_dist)
ax_dend.set_ylim(bottom=-5, top=max_dist)
ax_dend.xaxis.set_tick_params(length=3, pad=2)
ax_dend.yaxis.set_tick_params(length=3, pad=2)
# # population colours
# ax_pops = fig.add_subplot(gs[1, 0])
# x = hap_pops.take(r['leaves'])
# hap_clrs = [pop_colours[p] for p in x]
# ax_pops.broken_barh(xranges=[(i, 1) for i in range(h.shape[1])], yrange=(0, 1), color=hap_clrs);
# sns.despine(ax=ax_pops, offset=5, left=True, bottom=True)
# ax_pops.set_xticks([])
# ax_pops.set_yticks([])
# ax_pops.set_xlim(0, h.shape[1])
# ax_pops.yaxis.set_label_position('left')
# ax_pops.set_ylabel('Population', rotation=0, ha='right', va='center')
# missense mutations
if h_display is not None:
ax_mut = fig.add_subplot(gs[1, 0])
plot_missense_haplotypes(ax_mut, h_display.take(r['leaves'], axis=1))
ax_mut.set_xticks([])
ax_mut.yaxis.set_tick_params(length=3, pad=2)
# histogram
ax_hist = fig.add_subplot(gs[0, 1])
sns.despine(ax=ax_hist, offset=5, left=True, right=False)
plot_dist_hist(h, ax=ax_hist, max_dist=max_dist)
ax_hist.yaxis.tick_right()
ax_hist.set_xlabel('No. SNPs')
ax_hist.set_ylabel('Frequency')
ax_hist.yaxis.set_label_position('right')
if pop:
txt = '{}'.format(pop_labels[pop])
fig.suptitle('Population = {}'.format(pop_labels[pop]), y=1.1, ha='right', x=.9)
# ax_hist.text(ax_hist.get_xlim()[1], ax_hist.get_ylim()[1], txt, ha='right', va='bottom')
if fn:
fig.savefig(fn, jpeg_quality=100, dpi=dpi, bbox_inches='tight')
return z, r
# -
fig_hap_structure(h, h_missense, pop='GWA');
fig_hap_structure(h, h_missense, pop='BFS');
z, r = fig_hap_structure(h, h_missense, pop='UGS');
plt.plot(z[:, 2])
z[-10:, 2]
fig_hap_structure(h, h_missense, pop='GAS', max_dist=200);
fig_hap_structure(h, h_missense, pop='BFM', max_dist=200);
fig_hap_structure(h, h_missense, pop='GNS', max_dist=200);
fig_hap_structure(h, h_missense, pop='AOM', max_dist=200);
| notebooks/chapter6/extra_metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import pymongo
from pymongo import InsertOne, DeleteMany, ReplaceOne, UpdateOne
from pymongo.errors import BulkWriteError
client = pymongo.MongoClient('192.168.127.12:27017')
db = client.tweet
import pandas as pd
from datetime import datetime
# + active=""
# records = []
# for i in db.test.find({'tweet.date':{'$lt':datetime.strptime('2017-10-03 00:00:00','%Y-%m-%d %H:%M:%S')}},{'tweet.text':1,'tweet.date':1}):
# records.append((i['_id'],i['tweet']['date'],i['tweet']['text']))
# + active=""
# data = pd.DataFrame.from_records(records,columns=['id','date','text'],index=['id'])
# ids = data.index.tolist()
# test_ids = pd.DataFrame(ids)
# test_ids.to_csv('test_ids.csv',header=False,index=False)
# -
# ## positive
records = []
for i in db.test.find({'class.1':{'$gte':0.9},'tweet.date':{'$lt':datetime.strptime('2017-10-03 00:00:00','%Y-%m-%d %H:%M:%S')}},{'tweet.text':1,'tweet.date':1}):
records.append((i['_id'],i['tweet']['date'],i['tweet']['text']))
data = pd.DataFrame.from_records(records,columns=['id','date','text'],index=['id'])
ids = data.index.tolist()
test_ids = pd.DataFrame(ids)
test_ids.to_csv('test_ids_pos.csv',header=False,index=False)
| experiment/test_data[10.1~10.2].ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
"""In progress.
<NAME>, Jan 2016.
"""
# %%
import tensorflow as tf
from libs.connections import conv2d, linear
from collections import namedtuple
from math import sqrt
# +
# %%
def residual_network(x, n_outputs,
activation=tf.nn.relu):
"""Builds a residual network.
Parameters
----------
x : Placeholder
Input to the network
n_outputs : TYPE
Number of outputs of final softmax
activation : Attribute, optional
Nonlinearity to apply after each convolution
Returns
-------
net : Tensor
Description
Raises
------
ValueError
If a 2D Tensor is input, the Tensor must be square or else
the network can't be converted to a 4D Tensor.
"""
# %%
LayerBlock = namedtuple(
'LayerBlock', ['num_layers', 'num_filters', 'bottleneck_size'])
blocks = [LayerBlock(3, 128, 32),
LayerBlock(3, 256, 64),
LayerBlock(3, 512, 128),
LayerBlock(3, 1024, 256)]
# %%
input_shape = x.get_shape().as_list()
if len(input_shape) == 2:
ndim = int(sqrt(input_shape[1]))
if ndim * ndim != input_shape[1]:
raise ValueError('input_shape should be square')
x = tf.reshape(x, [-1, ndim, ndim, 1])
# %%
# First convolution expands to 64 channels and downsamples
net = conv2d(x, 64, k_h=7, k_w=7,
batch_norm=True, name='conv1',
activation=activation)
# %%
# Max pool and downsampling
net = tf.nn.max_pool(
net, [1, 3, 3, 1], strides=[1, 2, 2, 1], padding='SAME')
# %%
# Setup first chain of resnets
net = conv2d(net, blocks[0].num_filters, k_h=1, k_w=1,
stride_h=1, stride_w=1, padding='VALID', name='conv2')
# %%
# Loop through all res blocks
for block_i, block in enumerate(blocks):
for layer_i in range(block.num_layers):
name = 'block_%d/layer_%d' % (block_i, layer_i)
conv = conv2d(net, block.bottleneck_size, k_h=1, k_w=1,
padding='VALID', stride_h=1, stride_w=1,
activation=activation, batch_norm=True,
name=name + '/conv_in')
conv = conv2d(conv, block.bottleneck_size, k_h=3, k_w=3,
padding='SAME', stride_h=1, stride_w=1,
activation=activation, batch_norm=True,
name=name + '/conv_bottleneck')
conv = conv2d(conv, block.num_filters, k_h=1, k_w=1,
padding='VALID', stride_h=1, stride_w=1,
activation=activation, batch_norm=True,
name=name + '/conv_out')
net = conv + net
try:
# upscale to the next block size
next_block = blocks[block_i + 1]
net = conv2d(net, next_block.num_filters, k_h=1, k_w=1,
padding='SAME', stride_h=1, stride_w=1, bias=False,
name='block_%d/conv_upscale' % block_i)
except IndexError:
pass
# %%
net = tf.nn.avg_pool(net,
ksize=[1, net.get_shape().as_list()[1],
net.get_shape().as_list()[2], 1],
strides=[1, 1, 1, 1], padding='VALID')
net = tf.reshape(
net,
[-1, net.get_shape().as_list()[1] *
net.get_shape().as_list()[2] *
net.get_shape().as_list()[3]])
net = linear(net, n_outputs, activation=tf.nn.softmax)
# %%
return net
def test_mnist():
"""Test the resnet on MNIST."""
import tensorflow.examples.tutorials.mnist.input_data as input_data
mnist = input_data.read_data_sets('MNIST_data/', one_hot=True)
x = tf.placeholder(tf.float32, [None, 784])
y = tf.placeholder(tf.float32, [None, 10])
y_pred = residual_network(x, 10)
# %% Define loss/eval/training functions
cross_entropy = -tf.reduce_sum(y * tf.log(y_pred))
optimizer = tf.train.AdamOptimizer().minimize(cross_entropy)
# %% Monitor accuracy
correct_prediction = tf.equal(tf.argmax(y_pred, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, 'float'))
# %% We now create a new session to actually perform the initialization the
# variables:
sess = tf.Session()
sess.run(tf.initialize_all_variables())
# %% We'll train in minibatches and report accuracy:
batch_size = 50
n_epochs = 5
for epoch_i in range(n_epochs):
# Training
train_accuracy = 0
for batch_i in range(mnist.train.num_examples // batch_size):
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
train_accuracy += sess.run([optimizer, accuracy], feed_dict={
x: batch_xs, y: batch_ys})[1]
train_accuracy /= (mnist.train.num_examples // batch_size)
# Validation
valid_accuracy = 0
for batch_i in range(mnist.validation.num_examples // batch_size):
batch_xs, batch_ys = mnist.validation.next_batch(batch_size)
valid_accuracy += sess.run(accuracy,
feed_dict={
x: batch_xs,
y: batch_ys
})
valid_accuracy /= (mnist.validation.num_examples // batch_size)
print('epoch:', epoch_i, ', train:',
train_accuracy, ', valid:', valid_accuracy)
if __name__ == '__main__':
test_mnist()
| files/nb_demo/tensorflow/tutorial/10_residual_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fully Convolutional Neural Networks
#
# Objectives:
# - Load a CNN model pre-trained on ImageNet
# - Transform the network into a Fully Convolutional Network
# - Apply the network perform weak segmentation on images
# +
# %matplotlib inline
import warnings
import numpy as np
from scipy.misc import imread as scipy_imread, imresize as scipy_imresize
import matplotlib.pyplot as plt
np.random.seed(1)
# +
# Wrapper functions to disable annoying warnings:
def imread(*args, **kwargs):
with warnings.catch_warnings():
warnings.simplefilter("ignore")
return scipy_imread(*args, **kwargs)
def imresize(*args, **kwargs):
with warnings.catch_warnings():
warnings.simplefilter("ignore")
return scipy_imresize(*args, **kwargs)
# +
# Load a pre-trained ResNet50
# We use include_top = False for now,
# as we'll import output Dense Layer later
from keras.applications.resnet50 import ResNet50
base_model = ResNet50(include_top=False)
print(base_model.output_shape)
# +
#print(base_model.summary())
# -
res5c = base_model.layers[-1]
type(res5c)
res5c.output_shape
# ### Fully convolutional ResNet
#
# - Out of the `res5c` residual block, the resnet outputs a tensor of shape $W \times H \times 2048$.
# - For the default ImageNet input, $224 \times 224$, the output size is $7 \times 7 \times 2048$
#
# #### Regular ResNet layers
#
# The regular ResNet head after the base model is as follows:
# ```py
# x = base_model.output
# x = Flatten()(x)
# x = Dense(1000)(x)
# x = Softmax()(x)
# ```
#
# Here is the full definition of the model: https://github.com/keras-team/keras-applications/blob/master/keras_applications/resnet50.py/resnet50.py
#
# #### Our Version
#
# - We want to retrieve the labels information, which is stored in the Dense layer. We will load these weights afterwards
# - We will change the Dense Layer to a Convolution2D layer to keep spatial information, to output a $W \times H \times 1000$.
# - We can use a kernel size of (1, 1) for that new Convolution2D layer to pass the spatial organization of the previous layer unchanged (it's called a _pointwise convolution_).
# - We want to apply a softmax only on the last dimension so as to preserve the $W \times H$ spatial information.
#
# #### A custom Softmax
#
# We build the following Custom Layer to apply a softmax only to the last dimension of a tensor:
# +
import keras
from keras.engine import Layer
import keras.backend as K
# A custom layer in Keras must implement the four following methods:
class SoftmaxMap(Layer):
# Init function
def __init__(self, axis=-1, **kwargs):
self.axis = axis
super(SoftmaxMap, self).__init__(**kwargs)
# There's no parameter, so we don't need this one
def build(self, input_shape):
pass
# This is the layer we're interested in:
# very similar to the regular softmax but note the additional
# that we accept x.shape == (batch_size, w, h, n_classes)
# which is not the case in Keras by default.
# Note that we substract the logits by their maximum to
# make the softmax more numerically stable.
def call(self, x, mask=None):
e = K.exp(x - K.max(x, axis=self.axis, keepdims=True))
s = K.sum(e, axis=self.axis, keepdims=True)
return e / s
# The output shape is the same as the input shape
def get_output_shape_for(self, input_shape):
return input_shape
# -
# Let's check that we can use this layer to normalize the classes probabilities of some random spatial predictions:
n_samples, w, h, n_classes = 10, 3, 4, 5
random_data = np.random.randn(n_samples, w, h, n_classes)
random_data.shape
# Because those predictions are random, if we some accross the classes dimensions we get random values instead of class probabilities that would need to some to 1:
random_data[0].sum(axis=-1)
# Let's wrap the `SoftmaxMap` class into a test model to process our test data:
# +
from keras.models import Sequential
model = Sequential([SoftmaxMap(input_shape=(w, h, n_classes))])
model.output_shape
# -
softmax_mapped_data = model.predict(random_data)
softmax_mapped_data.shape
# All the values are now in the [0, 1] range:
softmax_mapped_data[0]
# The last dimension now approximately sum to one, we can therefore be used as class probabilities (or parameters for a multinouli distribution):
softmax_mapped_data[0].sum(axis=-1)
# Note that the highest activated channel for each spatial location is still the same before and after the softmax map. The ranking of the activations is preserved as softmax is a monotonic function (when considered element-wise):
random_data[0].argmax(axis=-1)
softmax_mapped_data[0].argmax(axis=-1)
# #### Exercise
# - What is the shape of the convolution kernel we want to apply to replace the Dense ?
# - Build the fully convolutional model as described above. We want the output to preserve the spatial dimensions but output 1000 channels (one channel per class).
# - You may introspect the last elements of `base_model.layers` to find which layer to remove
# - You may use the Keras Convolution2D(output_channels, filter_w, filter_h) layer and our SotfmaxMap to normalize the result as per-class probabilities.
# - For now, ignore the weights of the new layer(s) (leave them initialized at random): just focus on making the right architecture with the right output shape.
# +
from keras.layers import Convolution2D
from keras.models import Model
input = base_model.layers[0].input
# TODO: compute per-area class probabilites
output = input
fully_conv_ResNet = Model(inputs=input, outputs=output)
# +
# # %load solutions/fully_conv.py
# -
# You can use the following random data to check that it's possible to run a forward pass on a random RGB image:
prediction_maps = fully_conv_ResNet.predict(np.random.randn(1, 200, 300, 3))
prediction_maps.shape
# How do you explain the resulting output shape?
#
# The class probabilities should sum to one in each area of the output map:
prediction_maps.sum(axis=-1)
# ### Loading Dense weights
#
# - We provide the weights and bias of the last Dense layer of ResNet50 in file `weights_dense.h5`
# - Our last layer is now a 1x1 convolutional layer instead of a fully connected layer
# +
import h5py
with h5py.File('weights_dense.h5', 'r') as h5f:
w = h5f['w'][:]
b = h5f['b'][:]
# +
last_layer = fully_conv_ResNet.layers[-2]
print("Loaded weight shape:", w.shape)
print("Last conv layer weights shape:", last_layer.get_weights()[0].shape)
# +
# reshape the weights
w_reshaped = w.reshape((1, 1, 2048, 1000))
# set the conv layer weights
last_layer.set_weights([w_reshaped, b])
# -
# ### A forward pass
#
# - We define the following function to test our new network.
# - It resizes the input to a given size, then uses `model.predict` to compute the output
# +
from keras.applications.imagenet_utils import preprocess_input
def forward_pass_resize(img_path, img_size):
img_raw = imread(img_path)
print("Image shape before resizing: %s" % (img_raw.shape,))
img = imresize(img_raw, size=img_size).astype("float32")
img = preprocess_input(img[np.newaxis])
print("Image batch size shape before forward pass:", img.shape)
z = fully_conv_ResNet.predict(img)
return z
# -
output = forward_pass_resize("dog.jpg", (800, 600))
print("prediction map shape", output.shape)
# ### Finding dog-related classes
# ImageNet uses an ontology of concepts, from which classes are derived. A synset corresponds to a node in the ontology.
#
# For example all species of dogs are children of the synset [n02084071](http://image-net.org/synset?wnid=n02084071) (Dog, domestic dog, Canis familiaris):
# Helper file for importing synsets from imagenet
import imagenet_tool
synset = "n02084071" # synset corresponding to dogs
ids = imagenet_tool.synset_to_dfs_ids(synset)
print("All dog classes ids (%d):" % len(ids))
print(ids)
for dog_id in ids[:10]:
print(imagenet_tool.id_to_words(dog_id))
print('...')
# ### Unsupervised heatmap of the class "dog"
#
# The following function builds a heatmap from a forward pass. It sums the representation for all ids corresponding to a synset
def build_heatmap(z, synset):
class_ids = imagenet_tool.synset_to_dfs_ids(synset)
class_ids = np.array([id_ for id_ in class_ids if id_ is not None])
x = z[0, :, :, class_ids].sum(axis=0)
print("size of heatmap: " + str(x.shape))
return x
def display_img_and_heatmap(img_path, heatmap):
dog = imread(img_path)
plt.figure(figsize=(12, 8))
plt.subplot(1, 2, 1)
plt.imshow(dog)
plt.axis('off')
plt.subplot(1, 2, 2)
plt.imshow(heatmap, interpolation='nearest', cmap="viridis")
plt.axis('off')
# **Exercise**
# - What is the size of the heatmap compared to the input image?
# - Build 3 dog heatmaps from `"dog.jpg"`, with the following sizes:
# - `(400, 640)`
# - `(800, 1280)`
# - `(1600, 2560)`
# - What do you observe?
#
# You may plot a heatmap using the above function `display_img_and_heatmap`. You might also want to reuse `forward_pass_resize` to compute the class maps them-selves
# +
# dog synset
s = "n02084071"
# TODO
# +
# # %load solutions/build_heatmaps.py
# -
# ### Combining the 3 heatmaps
# By combining the heatmaps at different scales, we obtain a much better information about the location of the dog.
#
# **Bonus**
# - Combine the three heatmap by resizing them to a similar shape, and averaging them
# - A geometric norm will work better than standard average!
# +
# # %load solutions/geom_avg.py
# -
# **Bonus**
#
# Experiment with Semantic segmentation. You may train on COCO dataset http://mscoco.org/dataset/#overview
#
# - Use the GPU to precompute the activations of a headless and convolutionalized ResNet50 or Xception model;
# - Initialize the weights of a new Convolution2D(n_classes, 1, 1) at random;
# - Train the top of the segmentation model on class label data extracted from the MS COCO 2016 dataset;
# - Start with a single low resolution model. Then add multi-scale and see the improvement.
#
# To go further, consider open source implementation of models rather than building your own from scratch. For instance, FAIR's detection lib (in Caffe2) provides a lot of state of the art models. https://github.com/facebookresearch/Detectron
| labs/05_conv_nets_2/Fully_Convolutional_Neural_Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Progress--Four detailed steps
# ## 1. Data cleaning
# * Smooth the time series data using a moving average
# * Reduce the dimensionality
#
# ## 2. Statistical validation
# * Motivation
# * Package
# * Scatterplot with user input
# * table of regression parameters)
#
# ## 3. Time series visualization
#
# ## 4. Spatial visualization
#
# # Deliverables for next period
# * Data(cleaned, reduced)
# * Package for statistical validation
# * Function for time series plotting
# * Maps in Earth Engine
#
# # Challenges
# * Earth Engine (Interface between EE and Python)
# * Function with user inputs from comobox
# * Space and time statistics
#
| presentations/Changming_2_22.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='toc'>Table of Contents</a>
# 1. <a href='#fgo'>Figure Object</a>
# 2. <a href='#axeso'>Axes Object</a>
# 3. <a href='#plts'>Plots</a>
# 1. <a href='#plotf'>Plotting Functions</a>
# 2. <a href='#plotmf'>Plotting Multiple Functions</a>
# 4. <a href='#seaborn'>Seaborn</a>
# The matplotlib plotting library provides powerful data visualization functionality. The recommended method to use matplotlib is via the pyplot interface as it provides an object-oriented access to the full matplotlib library. Since we will be focusing on writing Python programs, we will use the pyplot access method. The basic concept behind creating a visualization is to create a _figure_ upon which a _visualization_ will be made. The _figure_ is represented in matplotlib by a `Figure` object, while the _visualization_ is represented by an `Axes` object.
import matplotlib.pyplot as plt
import numpy as np
# <a id='fgo'></a>
# ### Figure Object
#
# Controls the global appearence of the visualization. Size, resolution etc.
fig = plt.figure()
# <a href='#toc'>Back to ToC</a>
# <a id='axeso'></a>
# ### Axes Object
#
# Makes the actual plot in the figure. A figure can have multiple subplots.
ax = fig.add_subplot()
ax
# <a href='#toc'>Back to ToC</a>
# <a id='plts'></a>
# ### Plots
# One-line
fig, ax = plt.subplots(figsize=(10,5))
plt.show()
# <a href='#toc'>Back to ToC</a>
# <a id='plotf'></a>
# ### Plotting Functions
# $y_i = m \cdot x_i + b$.
# Formula
m = -5
b = 10
x = np.linspace(0, 10)
y = m * x + b
# +
# Plotting the formulae
fig, ax = plt.subplots(figsize=(10,5))
# Generating the Plot
ax.plot(x, y)
# Setting Title, X and Y Labels
ax.set_xlabel("X Axis")
ax.set_ylabel("Y Axis")
ax.set_title("$y_i = m \cdot x_i + b$")
# Setting the axis limits
ax.set_xlim(-1, 11)
ax.set_ylim(-45, 11)
# Setting the x and y ticks
ax.set_xticks(np.arange(0,11,1))
ax.set_yticks(np.arange(-45,15,5))
# Show the plot
plt.show()
# -
# <a href='#toc'>Back to ToC</a>
# <a id='plotmf'></a>
# ### Plotting Multiple Functions
# +
# Creating Figure and Axes Objects
fig, ax = plt.subplots(figsize=(10,5))
# Now we generate something to plot. In this case, we will plot a straight line.
m = 2
b = 5
x1 = np.linspace(0,10)
y1 = m * x1 + b
x2 = x1
y2 = -1 * y1
# We can either plot each set of data separately as shown, or plot them all at
# once by calling ax.plot(x1, y1, x2, y2)
ax.plot(x1, y1, label='y = -2x+5')
ax.plot(x2, y2, label='y = 2x-5', color='green')
#display legend
plt.legend()
# Set our axis labels
ax.set_xlabel("X Axis")
ax.set_ylabel("Y Axis")
# Change the axis limits displayed in our plot
ax.set_xlim(-2, 12)
ax.set_ylim(-20, 20)
# Change the ticks on each axis and the corresponding numerical values that are displayed
ax.set_xticks(np.arange(0, 15, 5))
ax.set_yticks(np.arange(-15, 20, 5))
# Set the title
ax.set_title("Our Final Plot!")
plt.show()
# -
# $y = m x^2 + b$
# +
m = 4
b = 4
x = np.linspace(0, 10)
y = m * x **2 + b
fig, ax = plt.subplots(figsize=(10,15))
ax.set_xlabel("Height (cm)")
ax.set_ylabel("Mass (kg)")
# Set the title
ax.set_title("$y = m x^2 + b$")
plt.plot(x,y)
plt.show()
# -
# <a href='#toc'>Back to ToC</a>
import matplotlib.pylab as plt
x = np.linspace(-np.pi, np.pi, 201)
plt.plot(x, np.sin(x))
plt.xlabel('Angle [rad]')
plt.ylabel('sin(x)')
plt.axis('tight')
plt.show()
# <a href='#toc'>Back to ToC</a>
# <a id='seaborn'></a>
# ### Seaborn
# +
import seaborn as sns
sns.set()
# +
# Now we create our figure and axes for the plot we will make.
fig, ax = plt.subplots(figsize=(10,5))
# We can either plot each set of data separately as shown, or plot them all at
# once by calling ax.plot(x1, y1, x2, y2)
ax.plot(x1, y1)
ax.plot(x2, y2)
# Set our axis labels
ax.set_xlabel("X Axis")
ax.set_ylabel("Y Axis")
# Change the axis limits displayed in our plot
ax.set_xlim(-2, 12)
ax.set_ylim(-20, 20)
# Change the ticks on each axis and the corresponding numerical values that are displayed
ax.set_xticks(np.arange(0, 15, 5))
ax.set_yticks(np.arange(-15, 20, 5))
# Set the title
ax.set_title("Our Final Plot!")
plt.show()
# -
# In the following code cell, we demonstrate how to change the overall appearance of the plot by using a white style with tick marks and a specific seaborn context, which can take one of four predefined types:
#
# - `notebook`
# - `paper`
# - `talk`
# - `poster`
#
# We use the `despine` method, which can remove the box appearance of the plot, and we change the font size, resulting in a very different plot appearance.
#
# +
# Now seaborn specific modifications
sns.set_context("poster", font_scale=1.25)
sns.set_style("white")
sns.set_style("ticks")
# Now we create our figure and axes for the plot we will make.
fig, ax = plt.subplots(figsize=(10,5))
# We can either plot each set of data separately as shown, or plot them all at
# once by calling ax.plot(x1, y1, x2, y2)
ax.plot(x1, y1, label = 'Plot 1', ls = ':')
ax.plot(x2, y2, label = 'Plot 2', ls = '--')
# Set our axis labels
ax.set_xlabel("X Axis")
ax.set_ylabel("Y Axis")
plt.legend(loc='upper left')
# Change the axis limits displayed in our plot
ax.set_xlim(-2, 12)
ax.set_ylim(-20, 20)
# Set the title
ax.set_title("Our Final Plot!")
# Now seaborn specific modifications
sns.despine(offset=10, trim=True)
# -
# **Plotting** $y = m x^3 + b$
# +
m = 2
b = 5
x = np.linspace(0, 10)
y = m * x **3 + b
# Now seaborn specific modifications
sns.set_context("poster", font_scale=1.25)
sns.set_style("white")
sns.set_style("ticks")
# Now we create our figure and axes for the plot we will make.
fig, ax = plt.subplots(figsize=(10,5))
# We can either plot each set of data separately as shown, or plot them all at
# once by calling ax.plot(x1, y1, x2, y2)
#ax.plot(x1, y1, label = 'Plot 1', ls = ':')
#ax.plot(x2, y2, label = 'Plot 2', ls = '--')
ax.plot(x, y)
# Set our axis labels
ax.set_xlabel("Height (cm)")
ax.set_ylabel("Mass (kg)")
#plt.legend(loc='upper left')
# Change the axis limits displayed in our plot
ax.set_xlim(0, 3)
ax.set_ylim(0, 50)
# Set the title
ax.set_title("$y = 2x^3 + 5$")
# Now seaborn specific modifications
sns.despine(offset=10, trim=True)
plt.plot(x,y)
plt.show()
# -
# **plotting** $y = m x^3 + b$
# +
m = 6
b = 5
x = np.linspace(0, 10)
y = m * x **3 + b
# Now seaborn specific modifications
sns.set_context("poster", font_scale=1.25)
sns.set_style("white")
sns.set_style("ticks")
# Now we create our figure and axes for the plot we will make.
fig, ax = plt.subplots(figsize=(10,5))
# We can either plot each set of data separately as shown, or plot them all at
# once by calling ax.plot(x1, y1, x2, y2)
#ax.plot(x1, y1, label = 'Plot 1', ls = ':')
#ax.plot(x2, y2, label = 'Plot 2', ls = '--')
ax.plot(x, y)
# Set our axis labels
ax.set_xlabel("Height (cm)")
ax.set_ylabel("Mass (kg)")
#plt.legend(loc='upper left')
# Change the axis limits displayed in our plot
ax.set_xlim(0, 3)
ax.set_ylim(0, 50)
# Set the title
ax.set_title("$y = 6 x^3 + 5$")
# Now seaborn specific modifications
sns.despine(offset=10, trim=True)
plt.plot(x,y)
plt.show()
# -
# ---
# <a href='#toc'>Back to ToC</a>
# ---
| Matplotlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#import dependencies
from splinter import Browser
from bs4 import BeautifulSoup as bs
import pandas as pd
import requests
from webdriver_manager.chrome import ChromeDriverManager
#declare urls
url1 = 'https://redplanetscience.com/'
url2 = 'https://spaceimages-mars.com/'
url3 = 'https://galaxyfacts-mars.com/'
url4 = 'https://marshemispheres.com/'
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
# +
#scrape news
browser.visit(url1)
html = browser.html
soup = bs(html, 'html.parser')
news_title = soup.find('div', class_='content_title').text
news_p = soup.find('div', class_='article_teaser_body').text
print(news_title)
print(news_p)
# +
#scrape featured image
browser.visit(url2)
html = browser.html
soup = bs(html, 'html.parser')
featured_image_url = url2 + soup.find('img', class_='headerimage')['src']
print(featured_image_url)
# -
#scrape facts in pandas
tables = pd.read_html(url3)
tables
#select table for mars facts
mars_facts_df = tables[0]
# +
#rename columns
renamed_mars_facts_df = mars_facts_df.rename(columns={0:'Description', 1:'Mars', 2:'Earth'})
#render html table from pandas
mars_facts_table = renamed_mars_facts_df.to_html(index=False)
#clean
final_mars_facts_table = mars_facts_table.replace('\n', '')
final_mars_facts_table
# +
#scrape 4 hemispheres
browser.visit(url4)
html = browser.html
soup = bs(html, 'html.parser')
#get 4 hemispheres
hemispheres = soup.find_all('div', class_='description')
#hemispheres_link = hemispheres.find_all('a')
#print (hemispheres)
#set up hemi url list
hemisphere_image_urls = []
#loop through each hemisphere div
for hemi in hemispheres:
hemi_dict = {}
#extract hemisphere title
hemi_title = hemi.find('h3').text
#go to the link in div
browser.links.find_by_partial_text(hemi_title).click()
hemi_html = browser.html
hemi_soup = bs(hemi_html, 'html.parser')
#pull href to complete image urls
temp_div = hemi_soup.find('div', class_='downloads')
hemi_image = temp_div.find('a')['href']
hemi_image_link = url4 + hemi_image
#print(temp_div)
#add values to dictionary
hemi_dict['title'] = hemi_title
hemi_dict['img_url'] = hemi_image_link
#add dictionary to list hemisphere_image_urls
hemisphere_image_urls.append(hemi_dict)
#go back a page
browser.back()
# -
#display list
print (hemisphere_image_urls)
| Mission_to_Mars/mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit (conda)
# name: python3
# ---
# # Common Python Data Structures (Guide)
#
# [https://realpython.com/python-data-structures/](https://realpython.com/python-data-structures/)
# Neste tutorial, você aprenderá:
#
# - Quais tipos de dados abstratos comuns são integrados à biblioteca padrão do Python
# - Como os tipos de dados abstratos mais comuns são mapeados para o esquema de nomenclatura do Python
# - Como colocar tipos de dados abstratos em uso prático em vários algoritmos
# ## Dicionários, Mapas e Tabelas Hash
#
# [https://realpython.com/python-data-structures/#dictionaries-maps-and-hash-tables](https://realpython.com/python-data-structures/#dictionaries-maps-and-hash-tables)
# Dicionários (ou `dicts`) são estruturas de dados que armazenam um número arbitrário de objetos, cada um deles associado a um identificador único, uma chave (key).
#
# Os dicionários também são chamados de mapas, hashmaps, hookup tables ou associative arrays. Eles permitem a pesquisa, inserção e exclusão eficientes de qualquer objeto associado a uma determinada chave.
#
# Uma analogia com o mundo real são as agendas de telefone. Você não sai olhando a agenda inteira, do início, para achar um telefone. Você, a partir do nome (a chave) da pessoa ou empresa, vai direto no nome dela e obtém o número.
#
# Os dicionários, portanto, tem como característica uma performance de pesquisa, inserção e remoção muito maior que outras estruturas de dados como listas e vetores.
#
# Dicionáris são estruturas de dados muito importantes e usadas com bastante frequências.
# ### `dict`: Seu dicionário do dia a dia
# Dicionários são tão importantes que o Python mantém uma implementação padrão, bastante robusta, diretamente no núcleo da linguagem, o tipo de dado `dict`.
#
# Python define 3 maneiras de se criar um dicionário:
# - Usando a função `dict()`
# - Usando os literais de dicionários, com chaves `{}`
# - Usando dict comprehentions
# Criando um dicionário usando a função dict()
agenda_dict = dict()
agenda_dict['maria'] = '11-11111-1111'
agenda_dict['pedro'] = '22-22222-2222'
agenda_dict['joao'] = '33-33333-3333'
agenda_dict
# criando um dicionário usando um literal
agenda_literal = {
"alice": '44-44444-4444',
"mário": '55-55555-5555',
"joão": '66-66666-6666',
}
agenda_literal
# +
# Criando um dicionário usando um dict comprehension
nomes = ['ciro', 'mula', 'bozo']
numeros = ['77-77777-77', '88-88888-8888', '99-99999-9999']
agenda_comprehension = {chave: valor for chave, valor in zip(nomes, numeros)}
agenda_comprehension
# -
# Para acessar um dicionário, você usa a sintaxe índice, passando a chave buscada entre colchetes.
print(agenda_dict["maria"])
print(agenda_literal["alice"])
print(agenda_comprehension["ciro"])
# Existem algumas restrições sobre os objetos que podem ser usados como chaves válidas.
#
# Os dicionários do Python são indexados por chaves que podem ser de qualquer tipo *hashable*. Um objeto *hashable* tem um valor *hash* que nunca muda durante seu tempo de vida (consulte `__hash__`) e pode ser comparado a outros objetos (consulte `__eq__`).
#
# Tipos imutáveis, como strings e números, são *hashables* e, portanto, ótimos candidatos a chave. Também podemos usar tuplas como chaves, desde que elas contenham apenas objetos *hashables*.
# Python forcene uma implementação padrão bastante robusta para dicionários, mas você pode usar qualquer outra implementação de dicionário que você desejar. Além da implementação padrão, Python também oferece algums implementações especializadas, para dicionários mais específicos.
# ### `collections.OrderedDict`: Lembre-se da ordem de inserção das chaves
# O dicionário padrão mantém as ordens de inserção das chaves, mas isso é meramente um efeito colateral de sua implementação, não sendo definido na especificação. Desse moto, se precisamos garantir a ordem de de inserção das chaves, devemos usar o `collections.OrderedDict`.
#
# Como ele não é uma implementação built-in, você precisa importar a classe `OrderedDict`.
# +
import collections # noqa E402
d = collections.OrderedDict(one=1, two=2, three=3)
print(d)
# -
d["four"] = 4
d
d.keys()
# Até o Python 3.8, você não podia iterar sobre itens de dicionário em ordem inversa usando `reversed()`. Apenas as instâncias de `OrderedDict` ofereciam essa funcionalidade. Mesmo no Python 3.8, objetos `dict` e `OrderedDict` não são exatamente a mesma coisa.
#
# A classe `OrderedDict` possui um método `.move_to_end()`, que não está disponível na classe `dict`. Ela também tem um método `.popitem()` mais customizável que o `.popitem()` da classe `dict`.
# ### `collections.defaultdict`: Retornar valores padrão para chaves ausentes
# A classe `defaultdict` é outra subclasse de dicionário que aceita um `callable` em seu construtor cujo valor de retorno será usado se uma chave solicitada não puder ser encontrada.
#
# Isso pode economizar alguma digitação e tornar suas intenções mais claras em comparação com o método `get()` ou pegando uma exceção `KeyError` em dicionários regulares.
# +
from collections import defaultdict # noqa E402
dd = defaultdict(list)
# Acessar uma chave ausente a cria e inicializa usando a fábrica padrão,
# isto é, a list() neste exemplo:
dd["dogs"].append("Rufus")
dd["dogs"].append("Spike")
dd["cats"].append("Sophie")
dd["dogs"].append("Mr Sniffles")
print(dd)
print(dd["dogs"])
print(dd["cats"])
# -
# ### `collections.ChainMap`: Pesquise múltiplos dicionários como um único mapeamento
# A estrutura de dados `collections.ChainMap` agrupa múltiplos dicionários em um único mapeamento. A busca procura nos mapeamentos subjacentes, um por um, até que uma chave seja encontrada. Inserções, atualizações e exclusões afetam apenas o primeiro mapeamento adicionado à cadeia:
# +
from collections import ChainMap # noqa E402
dict1 = {"a": 1, "b": 2}
dict2 = {"c": 3, "d": 4}
chain = ChainMap(dict1, dict2)
chain
# -
chain["a"]
chain["c"]
# +
import traceback # noqa E402
try:
chain["e"]
except KeyError:
traceback.print_exc()
# -
# ### `types.MappingProxyType`: Um wrapper para fazer dicionários somente leitura
# O `MappingProxyType` é um wreapper ao redor do dicionário padrão, que provê uma interface somente para leitura aos dados do dicionário. Essa classe foi adicionada no Python 3.3 e pode ser usada pra criar uma versão proxy imutável de dicionários.
#
# `MappingProxyType` pode ser útil se, por exemplo, você quer retornar um dicionário que carrega o estado interno de uma classe ou modulo, enquanto desoncoraja acesssos de escrita a esse objeto. Usar o `MappingProxyType` é mais eficiente que usar uma cópia de um dicionário, e ainda permite que você possa colocar as restrições necessárias.
# +
from types import MappingProxyType # noqa E402
gravavel = {'a': 1, 'b': 2}
somente_leitura = MappingProxyType(gravavel)
somente_leitura['a']
# +
import traceback # noqa E402
try:
somente_leitura['a'] = 3
except TypeError:
traceback.print_exc()
# -
# alteraçoes no dicionário original se refletem no proxy
gravavel['a'] = 4
somente_leitura
# ### Dicionários em Python: Resumo
# Todas essas implementações estão presentes a biblioteca padrão do Python. Se possível, use o dicionário padrão. Além de ser bastante versátil e bem otimizado, seu código será mais limpo. Porém, em caso de requisitos especiais, não hesite em usar os dicionários especiais.
#
# Nesse capítulo, nós vemos:
#
# - dict: Dicionário padrão
# - collections.OrderedDict: Dicionário que mantém a ordem de inserção das chaves
# - collections.defaultdict: Dicionário que retorna um valor padrão para chaves ausentes
# - collections.ChainMap: Dicionário que agrupa múltiplos dicionários em um único mapeamento
# - types.MappingProxyType: Wrapper para dicionários somente leitura
| notebook/01-dicitionaries-maps-and-hash-tables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimización de funciones escalares diferenciables con `SymPy`
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/thumb/b/b6/Extrema_example_es.svg/1200px-Extrema_example_es.svg.png" width="300px" height="100px" />
#
# > - Mediante optimización se obtienen soluciones elegantes tanto en teoría como en ciertas aplicaciones.
# > - La teoría de optimización usa elementos comenzando con cálculo elemental y álgebra lineal básica, y luego se extiende con análisis funcional y convexo.
# > - Las aplicaciones en optimización involucran ciencia, ingeniería, economía, finanzas e industria.
# > - El amplio y creciente uso de la optimización lo hace escencial para estudiantes y profesionales de cualquier rama de la ciencia y la tecnología.
#
# **Referencia:**
# - http://www.math.uwaterloo.ca/~hwolkowi//henry/reports/talks.d/t06talks.d/06msribirs.d/optimportance.shtml
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/thumb/e/e7/OpenStreetMap_routing_service.png/300px-OpenStreetMap_routing_service.png" width="300px" height="100px" />
#
# Algunas aplicaciones son:
#
# 1. Ingeniería
# - Encontrar la composición de equilibrio de una mezcla de diferentes átomos.
# - Planeación de ruta para un robot (o vehículo aéreo no tripulado).
# - Planeación de la mano de obra óptima en una construcción o planta de producción.
# 2. Distribución óptima de recursos.
# - Distribución de rutas de vuelo.
# - Encontrar una dieta óptima.
# - Planeación de ruta óptima.
# 3. Optimización financiera
# - Administración de riesgos.
# - Portafolios de inversión.
# En esta clase veremos aspectos básicos de optimización. En específico, veremos cómo obtener máximos y mínimos de una función escalar de una variable (como en cálculo diferencial).
# ___
# ## 0. Librerías que usaremos
#
# Como ya dijimos en la primer clase `python` es el lenguaje de programación (el cual es de alto nivel). Sin embargo, `python` solo tiene unos pocos comandos primitivos y para hacer más fácil su uso en nuestras actividades de simulación en ingeniería, otras personas ya han escrito ciertas librerías por nosotros.
# ### 0.1 `NumPy`
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="data:image/png;base64,iVBORw0KGgoAAAANSUhEUgAAAOEAAADhCAMAAAAJbSJIAAAAkFBMVEX///9Nq89Nd89Iqc4+bsxFcs7R2vF6vdmVyuCWrOA7pczW3/Pg5vVAp82l0eSczeJ2u9jq9Pn1+vyEwtxCcM3e7vXU6fJktNRasNI5a8zN5fCZzOG12emNxt7y+Pvn8/jD4O3K1O9UfNHs8Pmqu+a7yOpoitV1k9h9mdq/zOydseLk6veGoNxykddcgdKywuhmzM3GAAAJhUlEQVR4nO2da1fiMBCGS9OuUq0googXBNz1fvn//25bEOllMjNJ06T05P22C+3Jw5ummZkkBoGXl5eXl5eXl5eXl5eXl5eXF6C7c9ctaFdnIk3ue8yY8Q0GA5H2lTHjE4ONRB99nGz926l/Ppb5eufj5Cyt8vWLEeb76asL161rLIl/vfGR4Dt4HyejhOI7aB8nI9K/g/Yx4xNMvh8fD4tRke/gfFzcKvNtGY8PhfGYNcAASpOl67YztbhNdExMxpeuW87XRL2jJrMr161Wk5qPIr2+cd1idS2O2T6mg6Xr1uqJyZiKO9ct1dfimOyrIj1z3UpSiyX24T3qY/YSnCAXm26qljKbkusl8oXze6mPIrlAJt1Xs+QWwbekn26ID4XnEh+Towf5RZfjJP8Jbt36ONk/ZgTjRd3HZIa84B92F4jUoY+VYSQhfCx9WaQz5NulH0QkIzeMwLQlwVq9tyVv9GCJ3Lk6ADvxUTJl4TGmA+QFCEZeIrXs40T+Ksenlzkj/gKsZ1YdMBLxH8F4hPW4u4E88LLGyAgbdMOE5TV+ayuMzLAoGasz3hB8Vhj5IZFQDWevZrzEQKuMamGtEuNmAsNU2hajemqCzfhwpHbrVuYAGmmJTVOOaEZoRkfJuI+aqSUOIxJ54IwmfUTe7xzEMdqSLETSvLk5H5e66c8Br5teKYwy5ZuniaH01aXiQLAXc6jJfNQCvEDCS1XGsQ6jwtRGnVEkBvk2jKo+4gFgU0bjfBtGFR+FRgb0hs3IegnpiM0o0ABQLh5jK/7txGPEM7x3WPOy2TfJ15J/O9GMKRrgZgEgXrZfooyt8+XCGbMAHnkNbwNAYsmXnNEKXy45o0BTuPsAkFh+AUfCxvmGb/JcLMyIl+LLASBRtq8ztuDfcB79RdpbY8Qz9PUAkCjbLwdFRorvUmfsHsbhNEYYyz4KPEMPzheIJSbLQqcm+MbJMUEDKSMMw2n4Lf9GwUd0AioPAPHKUzbw5hdSgXT+8wltwowxphnRCSgeAIoE7av5y4X0L/8VGhCGYUT4iNZk6NwAkbK/Q2fvu0elGWHO+E/+vaul/LPJGSd21k677IeCpoQZ49erxi1kGXrARw3G4lDXnDBjXK8Ub4Bl6Jszll9XJghzH1UYy+8zhpTSLtXXsRnCMJw/cRmpCoSEEZvTInzmCDPGZ04b6BioCWOdzyRhGD/RLdDKK/0yEn0V4jNKGM5P8MvOm/Dl8xf0/SfLrpokjJ+J6/RTvOT8TN47TBKG81PqSs0Urz6fYcL4hb5W41Ek59foLY0ShtE742pFRur5C4JjdHg2Sxh/sK5XYKT5gmCEV/iNEoZz5lIzJqNgZf+tEsZD7j04KV7mogarhGHMn0ASjDz/7BNOkfSNEqPCohS7hOFU6U43koUkhH83tw4JIySvATYW8JHimyUXxX9bJgzXqnerhhspxZcOhFPCSD2pcXO995H2L2+1U0J1E4O9jyTf9muOCSPVrM1GeeiPL7ItPLKOCUNpJPznCYsgl2Ocr/C4uiaURsIn8/kzGWDBKg+5rgnjT8l1J1EYzz81GKuVUdeE0kg4I8z4lX2sV36dE8qCqA2hKiNU2XZOGM4fMUIVRrhy755QEkT9EuaMLwxG2coE94RhBJpYIOQwIisv3BPGbyThhhHJ62DZ/w4QhhEUCVcIMUZidVCJ8MwJIRgJ1whljOT+kQ54GIY8QoiRrr51wcMwAqrfIGHOWJrmMZarKhCKRGe3NMtDwEQJYRj9KRHStQ12L1Xen6NCCETCLRFKPdTl43r4ZYtQ5qHOHjIlwnokbNXDRseF8AjrNWGLHhLpAjOE9UjYmocN+diEtZqwJQ8NHPfCJKxFwjY8FCl6ToVhwmok3D6hGT4+YTUSbruXmuJTIKxEwu16qLMZpzFhGJVqwm16qLsZpylhORI2R3hUvCDz0CyfCmEYWyAcpcaPk1IgLEXCrRFiAZLubgTAre8p2Pz2Ca+QtQOXR/q7EWrNfPwE/7sQCbdEiPI12o1Qaeb7yRxqfiGIsky4Pa3AIOFp8Az+/z6Iskq4O9PHKOEpYaJFwoeL331DJgmDF3B59G/7rREWzp0yTPgOmbgvJ5oiFAlKWN5MZZYw+ABN3AVRZghFgs5fqpupDBM+gibuFtaaIMziBxU+44QSE99NERLxwwLYDGea8BFM3H+YISTiP3izn2lC2XzHAKGGf60QLsAn8a0xIeGf/LxX44TBG/ThdNKQkOJDqqnGCRdQiLENorQJ9fnaIIRNjBsQUvmlEb6a2jxhAH76rUtI589sVUj3hH+hfrrWI+TkB+0TBiDFSoOQl/90QPgNkXwpE3LrKw4IQROzIEqJkF9fcUH4D0CJn1QIiTN5S8k1F4TBGjLxlE2IHwaWxX9j54SvkIkvpzxC+kzlWs7bPmHwBX3jnySTXCa8p8837wLhCrJrDQNWCEm+bhCCJsrEJHz4jY86QfgHzCw2ICzGt50gDMAcvzbhQym+7QYhnOPXI6z+jYhuECqYSBAC+bNuEPJNRAnPL+rxbUcI4UKNIqEkf9YRQrhQo0IoOwxMuJ+1bQUWaviE8sPOOkMIpoe5hNhhbp0hhHP8LELisLq2CVcxkImBCMFCDYOQOo29dcIgeItq7kCE3NXEZULZ34CSEpo/2yTTYlhlBAl5T2KZcMTYjWCBMGv8x7zECBIyl/WXCMeMKjefME0bLJd6fykywoQLdUJOHZ9L2PhPCJw87YcSmBBOD1siNPInElZfuydNQhg4I0wT5jl9lF7XEUrIMLENQvY5hBx9T6cI4YSxWdE4oTH/dvo7jaWEgWTNYouE+NnveloM5/Lj6Ohdw0YJjfbPgh5fpGdeQDn+1gjb4sO1tkbohg/O8bdB2MbzxxRhohnCVLjxbyMwx9+AcFAndNU/d8Jz/M0JM//sAtW0QkPhpoSJY/82emqT0LV/G6GFmoaEHRGWWewHIZbP6Achls/oCSEyAe8LofyNoT6nmbmCwCXNgKsSGt9uaExGPBSu52eY3iXRvgqh+/kZqtN6MUCJUDiMj5havEDDDZNQdNy/H63Cuo0sQtHd8aWqYc1GBuGB+Pejk69IkdD0dvv29V0ecQjCA+qfe5VHHJzw4Pz70Wo95RCKRvU/x9pXyqWEh/f8lfX4OUcJD/L5q+jn5QgSHrp/Ow3zQjlA2Be+TKdPUZ2wR3y5XuN5hbBffJkmH6XzB8eHP74QanycnJeXl5eXl5eXl5eXl5eXl1fP9B/O8eEv4cS03gAAAABJRU5ErkJggg==" width="300px" height="100px" />
#
# `NumPy` (Numerical Python) es la librería fundamental para computación científica (numérica) con `Python`. Contiene, entre otras cosas:
# - un objeto tipo arreglo N-dimensional muy poderoso
# - funciones sofisticadas
# - funciones de álgebra lineal, transformada de Fourier y números aleatorios.
#
# Por lo anterior, `NumPy` es de amplio uso entre la comunidad científica e ingenieril (por su manejo de cantidades vectoriales). De la misma manera, se usa para guardar datos. Para nuestros propósitos, se puede usar libremente.
#
# **Referencia:**
# - http://www.numpy.org/
# `NumPy` ya viene incluido en la instalación estándar de Anaconda por defecto. Para comenzar a usarlo, solo debemos de importarlo:
# importar la librería numpy
import numpy as np
# Creamos un vector
x = np.array([0, 7, 8.4, -3.5, 2])
x
# Función seno sobre los elementos de vector x
np.sin(x)
# $$
# \int e^x dx = e^x +C
# $$
# $$
# \int_{1}^{2} e^{x^2} dx
# $$
# ### 0.2 `SymPy`
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="http://www.sympy.org/static/images/logo.png" width="300px" height="100px" />
#
# `SymPy` (Symbolic Python) es una librería de `Python` para matemáticas simbólicas. Su objetivo es convertirse en un sistema de álgebra computacional con las mejores características, manteniendo el código lo más simple posible para que sea comprensible.
#
# **Referencia:**
# - http://www.sympy.org/en/index.html
# `SymPy` ya viene incluido en la instalación estándar de Anaconda por defecto. Para comenzar a usarlo, solo debemos de importarlo:
# importar la librería sympy
import sympy as sym
# Ayuda en la función init_printing
help(sym.init_printing)
# imprimir en formato latex
sym.init_printing(use_latex='mathjax')
# La funcionalidad de imprimir en formato LaTex que nos da `SymPy` mediante el proyecto `mathjax` hace de `SymPy` una herramienta muy atractiva...
# Notar que en `SymPy` y en `NumPy` existen funciones con el mismo nombre, pero reciben tipos de datos diferentes...
# Definimos la variable simbólica x
sym.var('x')
sym.sin(x)
np.sin(x)
y = np.array([0, 1, 3.14])
np.sin(y)
sym.sin(y)
# +
# from sympy import *
# from numpy import *
# Esta práctica no es para nada recomendable.
# -
sin(y)
log(y)
# Explicar el uso de la sintaxis `from numpy import *` y sus peligros (no recomendable).
# ### 0.3 `PyPlot` de `matplotlib`
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="https://matplotlib.org/_static/logo2.svg" width="300px" height="100px" />
#
# El módulo `PyPlot` de la librería `matplotlib` contiene funciones que nos permite generar una gran cantidad de gráficas rápidamente. Las funciones de este módulo están escritas con el mismo nombre que las funciones para graficar en `Matlab`.
#
# **Referencia:**
# - https://matplotlib.org/api/pyplot_summary.html
# importar matplotlib.pyplot
# import matplotlib.pyplot as plt
from matplotlib import pyplot as plt
# comando para que las gráficas salgan en la misma ventana
# %matplotlib inline
# Ya que revisamos todas las librerías que usaremos, empecemos con la clase como tal...
# ___
# Basamos todos los resultados en los siguientes teoremas:
# ## 1. Teorema de Fermat (análisis)
#
# Si una función $f(x)$ alcanza un máximo o mínimo local en $x=c$, y si la derivada $f'(c)$ existe en el punto $c$, entonces $f'(c) = 0$.
# ### Ejemplo
#
# Sabemos que la función $f(x)=x^2$ tiene un mínimo global en $x=0$, pues
#
# $$f(x)=x^2\geq0,\qquad\text{y}\qquad f(x)=x^2=0 \qquad\text{si y solo si}\qquad x=0.$$
# declarar la variable real x
sym.var('x')
# declarar ahora f=x^2 y mostrar
f = x**2 # En python x^2 es otra cosa
# Cuidado
4**2
4^2 # 4: 1 0 0
# 2: 0 1 0
# 1 1 0: 6
# derivar f respecto a x y mostrar
df = sym.diff(f, x) # df / dx
df
# resolver f'(x)=0 y mostrar soluciones
help(sym.solve)
sol = sym.solve(df, x)
sol
# Encuentra también soluciones complejas
sym.solve(x**2 + 1, x)
# Puedo resolver ecuaciones que involucren más de una variable
sym.solve(x**2 * y + 3 * y**3 * x + 1, x)
# Veamos la gráfica...
# convertir f e una función que se pueda evaluar numéricamente (función lambdify de la librería sympy)
f_num = sym.lambdify(x, f, 'numpy')
help(sym.lambdify)
f
# f es una expresión simbólica y no se puede evaluar
f(10)
# f_num sí se puede evaluar
f_num(10)
help(np.linspace)
# Coordenadas x (abscisas)
x_num = np.linspace(-10, 10, 100)
x_num
y_num = f_num(x_num)
y_num
# graficar
# Crear ventana de graficos y damos medidas de la ventana
plt.figure(figsize=(6, 4))
# Sirve para hacer el grafico y determinar sus caracteristicas
plt.plot(x_num, y_num, label='$f(x)=x^2$')
# Punto de mínimo
plt.plot(0, f_num(0), 'o', label='Mínimo de $f(x)$')
# Los signos de $ son para escribir expresiones matemáticas
# Nombre del eje x de la grafica
plt.xlabel('$x$')
# Nombre del eje y
plt.ylabel('$y$')
# Sirve para poner las etiquetas de las graficas
plt.legend()
# Sirve para poner la cuadricula
plt.grid()
# **Otra manera de hacer lo anterior**
#
# Concepto de función...
# Concepto de función
# def nombre_de_la_funcion(argumento[s]):
# --> Operación 1
# --> Operación 2
# --> ...
# --> Operación N
# return [lo que vaya a devolver]
def cuadrado(x):
return x**2
cuadrado(x)
cuadrado(x_num)
dcuadrado = sym.diff(cuadrado(x), x)
dcuadrado
sol = sym.solve(dcuadrado, x)
sol
# graficar
# Crear ventana de graficos y damos medidas de la ventana
plt.figure(figsize=(6, 4))
# Sirve para hacer el grafico y determinar sus caracteristicas
plt.plot(x_num, cuadrado(x_num), label='$f(x)=x^2$')
# Punto de mínimo
plt.plot(0, cuadrado(0), 'o', label='Mínimo de $f(x)$')
# Los signos de $ son para escribir expresiones matemáticas
# Nombre del eje x de la grafica
plt.xlabel('$x$')
# Nombre del eje y
plt.ylabel('$y$')
# Sirve para poner las etiquetas de las graficas
plt.legend()
# Sirve para poner la cuadricula
plt.grid()
# El converso del teorema anterior no es cierto.
#
# ### Actividad
# Considere $g(x)=x^3$.
# - Usando `sympy`, muestre que $g'(0)=0$.
# - Sin embargo, descartar que $x=0$ es un extremo de $g(x)$ viendo su **gráfica**.
# Declarar la variable simbolica x
sym.var('x')
# Definimos funcion g(x)
def g(x):
return x**3
# Derivamos g(x)
dg = sym.diff(g(x), x)
dg
# Puntos criticos
sol = sym.solve(dg, x)
sol
# graficar
x_num = np.linspace(-1, 1, 100)
y_num = g(x_num)
# Crear ventana de graficos y damos medidas de la ventana
plt.figure(figsize=(6, 4))
plt.axvline(x=0, color='k')
plt.axhline(y=0, color='k')
# Sirve para hacer el grafico y determinar sus caracteristicas
plt.plot(x_num, y_num, linewidth=3, label='$g(x)=x^3$')
plt.plot(0, 0, 'o', markersize=20, label='$(0, 0)$')
# Nombre del eje x de la grafica
plt.xlabel('$x$')
# Nombre del eje y
plt.ylabel('$y$')
# Sirve para poner las etiquetas de las graficas
plt.legend()
# Sirve para poner la cuadricula
plt.grid()
# ## 2. Criterio de la segunda derivada
#
# Sea $f(x)$ una función tal que $f’(c)=0$ y cuya segunda derivada existe en un intervalo abierto que contiene a $c$.
# - Si $f’’(c)>0$, entonces $f(c)$ es un mínimo relativo.
# - Si $f’’(c)<0$, entonces $f(c)$ es un máximo relativo.
# - Si $f’’(c)=0$, entonces el criterio no decide.
# ### Ejemplo
#
# Mostrar, usando `sympy`, que la función $f(x)=x^2$ tiene un mínimo relativo en $x=0$.
#
# Ya vimos que $f'(0)=0$. Notemos que:
# Método subs
# Sacamos la segunda derivada
# Por tanto, por el criterio de la segunda derivada, $f(0)=0$ es un mínimo relativo (en efecto, el mínimo global).
# ### Ejemplo
# ¿Qué pasa con $g(x)=x^3$ al intentar utilizar el criterio de la segunda derivada? (usar `sympy`).
# Definimos la función g(x)=x^3
# Sacamos la derivada de la función g respecto a x g'(x)=3x^2
# Observamos que g'(0)=0 (es un punto crítico)
# Obtenemos la segunda derivada de la función g respecto a x g''(x)=6x
# Evaluamos g''(0)
# Como $g''(0)=0$ entonces el criterio de la segunda derivada no concluye.
# ### Actividad
#
# ¿Qué pasa con $h(x)=x^4$ al intentar utilizar el criterio de la segunda derivada?.
#
# ## 3. Método para determinar extremos absolutos de una función continua y=f(x) en [a,b]
#
# - Determinar todos los valores críticos $c_1, c_2, c_3, \dots, c_n$ en $(a,b)$.
# - Evaluar $f$ en todos los valores críticos y en los extremos $x=a$ y $x=b$.
# - El más grande y el más pequeño de los valores de la lista $f(a), f(b), f(c_1), f(c_2), \dots, f(c_n)$ son el máximo absoluto y el mínimo absoluto, respectivamente, de f en el intervalo [a,b].
# ### Ejemplo
#
# Determinar los extremos absolutos de $f(x)=x^2-6x$ en $\left[0,5\right]$.
#
# Obtenemos los puntos críticos de $f$ en $\left[0,5\right]$:
# Definimos f
# Derivamos f
# Resolver la ecuación f'(x)=0 para encontrar puntos críticos
# Evaluamos $f$ en los extremos y en los puntos críticos:
# Concluimos que el máximo absoluto de $f$ en $\left[0,5\right]$ es $0$ y se alcanza en $x=0$, y que el mínimo absoluto es $-9$ y se alcanza en $x=3$.
# +
# graficar
# Crear ventana de graficos y damos medidas de la ventana
# Sirve para hacer el grafico y determinar sus caracteristicas
# Nombre del eje x de la grafica
# Nombre del eje y
# Sirve para poner las etiquetas de las graficas
# Sirve para poner la cuadricula
# -
# ### Actividad
# Determinar los valores extremos absolutos de $h(x)=x^3-3x$ en $\left[-2.2,1.8\right]$, usando `sympy`. Mostrar en una gráfica.
# ### En varias variables...
#
# El procedimiento es análogo.
#
# Si una función $f:\mathbb{R}^n\to\mathbb{R}$ alcanza un máximo o mínimo local en $\boldsymbol{x}=\boldsymbol{c}\in\mathbb{R}^n$, y $f$ es diferenciable en el punto $\boldsymbol{x}=\boldsymbol{c}$, entonces $\left.\frac{\partial f}{\partial \boldsymbol{x}}\right|_{\boldsymbol{x}=\boldsymbol{c}}=\boldsymbol{0}$ (todas las derivadas parciales en el punto $\boldsymbol{x}=\boldsymbol{c}$ son cero).
#
# **Criterio de la segunda derivada:** para ver si es máximo o mínimo, se toma la segunda derivada (matriz jacobiana) y se verifica definición negativa o positiva, respectivamente.
#
# Si se restringe a cierta región, hay ciertas técnicas. La más general, pero también la más compleja es la de **multiplicadores de Lagrange**.
# **Ejemplo:** hacer a mano a la vez para corroborar...
sym.var('x y')
x, y
def f(x, y):
return x**2 + y**2
dfx = sym.diff(f(x,y), x)
dfy = sym.diff(f(x,y), y)
dfx, dfy
xy_c = sym.solve([dfx, dfy], [x, y])
xy_c
x_c, y_c = xy_c[x], xy_c[y]
x_c, y_c
# +
d2fx = sym.diff(f(x,y), x, 2)
d2fy = sym.diff(f(x,y), y, 2)
dfxy = sym.diff(f(x,y), x, y)
Jf = sym.Matrix([[d2fx, dfxy], [dfxy, d2fy]])
Jf.eigenvals()
# -
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# +
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
x = np.linspace(-2, 2, 100)
y = x
X, Y = np.meshgrid(x, y)
ax.plot_surface(X, Y, f(X, Y))
ax.plot([x_c], [y_c], [f(x_c,y_c)], '*r')
# -
# # Anuncios parroquiales
#
# ## 1. [Curso gratis sugerido](https://www.kaggle.com/learn/python)
#
# ## 2. La próxima clase hay quiz (clases 1 y 2 - lo que hemos visto).
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>.
# </footer>
| Modulo1/Clase2_OptimizacionSympy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Create dataset from real bugs extracted from GitHub
# Here we go through the binary operations extracted from the commits of real bugs in GitHub. The goal is to creat
# a dataset in a format expected by DeepBugs.
# +
import os
from pathlib import Path
import codecs
import json
from typing import List, Dict, Any
import pandas as pd
from multiprocessing import Pool, cpu_count
from tqdm.notebook import trange, tqdm
benchmarks_dir = '../benchmarks'
real_bugs_dataset_file_path = os.path.join(benchmarks_dir, 'binops_real_bugs.pkl')
real_bugs_dataset_dir = os.path.join(benchmarks_dir, 'binops_real_bugs')
# + pycharm={"name": "#%%\n"}
def read_json_file(json_file_path)->Dict:
try:
obj_text = codecs.open(json_file_path, 'r', encoding='utf-8').read()
return json.loads(obj_text)
except FileNotFoundError:
print(
"Please provide a correct file p. Eg. ./results/validated-conflicts.json")
return {}
except Exception as e:
# Empty JSON file most likely due to abrupt killing of the process while writing
# print (e)
return {}
def read_dataset_given_files(extracted_data_files: List) -> pd.DataFrame:
d = []
with Pool(cpu_count()) as p:
with tqdm(total=len(extracted_data_files)) as pbar:
pbar.set_description_str(
desc="Reading dataset from files", refresh=False)
for i, each_vars in enumerate(
p.imap_unordered(read_json_file, extracted_data_files, 20)):
pbar.update()
d.extend(each_vars)
p.close()
p.join()
extracted_dataset = pd.DataFrame(d)
return extracted_dataset
def file_path_to_dataset(dataset_file_path, dir_path):
if not Path(dataset_file_path).is_file():
file_paths = list(Path(dir_path).rglob('*.json'))
print(f"Number of files={len(file_paths)}")
dataset = read_dataset_given_files(extracted_data_files=file_paths)
print(f"Saving {dataset_file_path}")
dataset.to_pickle(dataset_file_path,'gzip')
else:
print(f'Reading from {dataset_file_path}')
dataset = pd.read_pickle(dataset_file_path,'gzip')
print(f"Dataset contains {len(dataset)} examples")
return dataset
# + pycharm={"name": "#%%\n"}
def get_file_loc(row):
d = row.to_dict()
if 'benchmarks/real_bugs_github/buggy_' in d['src']:
file_name = d['src'].replace('benchmarks/real_bugs_github/buggy_','')
else:
file_name = d['src'].replace('benchmarks/real_bugs_github/correct_','')
range = str(d['range'])
return file_name+'_'+range
# + pycharm={"name": "#%%\n"}
dataset = file_path_to_dataset(dataset_file_path=real_bugs_dataset_file_path, dir_path=real_bugs_dataset_dir)
row_iter = [row for _, row in dataset.iterrows()]
locations = []
for row in tqdm(row_iter):
loc = get_file_loc(row)
locations.append(loc)
dataset['filename_loc'] = locations
# + pycharm={"name": "#%%\n"}
dataset
# + pycharm={"name": "#%%\n"}
correct_dataset = dataset[dataset['src'].apply(lambda x: 'correct_' in x)]
buggy_dataset = dataset[dataset['src'].apply(lambda x: 'buggy_' in x)]
# + pycharm={"name": "#%%\n"}
merged = correct_dataset.merge(buggy_dataset,left_on='filename_loc', right_on='filename_loc', suffixes=['_CORRECT','_BUGGY'])
merged
# + pycharm={"name": "#%%\n"}
def get_buggy_non_buggy_data(row):
d = row.to_dict()
correct = {k.replace('_CORRECT',''):v for k, v in d.items() if '_CORRECT' in k}
correct['probability_that_incorrect'] = 0
buggy = {k.replace('_BUGGY',''):v for k, v in d.items() if '_BUGGY' in k}
buggy['probability_that_incorrect'] = 1
if (correct['left'] != buggy['left'] or correct['right'] != buggy['right'] ) and correct['op'] == buggy['op'] :
return [correct, buggy]
else:
return []
# + pycharm={"name": "#%%\n"}
correct_bin_ops = []
buggy_bin_ops = []
x_y_pair_given = []
for _,row in tqdm(list(merged.iterrows()), desc='Get lines'):
r = get_buggy_non_buggy_data(row)
if len(r):
correct_bin_ops.append(r[0])
buggy_bin_ops.append(r[1])
x_y_pair_given.append(r)
print(f'Number of buggy/correct binOps extracted are {len(correct_bin_ops)}')
# + pycharm={"name": "#%%\n"}
print(len(x_y_pair_given))
filtered_x_y_pair = []
for pr in x_y_pair_given:
if pr[0]['parent'] =='AwaitExpression' or 'AwaitExpression' == pr[0]['grandParent']:
continue
if pr[1]['parent'] =='AwaitExpression'or 'AwaitExpression' == pr[1]['grandParent']:
continue
filtered_x_y_pair.append(pr)
x_y_pair_given = filtered_x_y_pair
print(len(x_y_pair_given))
# -
# We give the buggy lines as input to a trained model in DeepBugs and check how many are actually classified as buggy.
# Then we confirm them with the correct extracted binops.
# + pycharm={"name": "#%%\n"}
def write_json(content, out_file):
with open(out_file, 'w+') as f:
json.dump(content, f)
# write_json(correct_bin_ops, os.path.join(benchmarks_dir, 'correct_real_binops.json'))
# write_json(buggy_bin_ops, os.path.join(benchmarks_dir, 'buggy_real_binops.json'))
write_json(x_y_pair_given, os.path.join(benchmarks_dir, 'correct_buggy_real_binops.json'))
| compare_real_bug_finding_ability/create_dataset_from_real_bugs_binopnds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/katarinagresova/ia161/blob/main/IA161_Language_modeling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PEg9l2Z51sm0"
# This notebook contains practical part of Language modeling lesson from Advanced NLP course. Goal is to train simple neural network based on word pairs and use it to generate new text.
# + id="FkWFruXL9_90"
import numpy as np
from collections import defaultdict
import re
# + [markdown] id="7wrY-8VYA9dS"
# # Data
# Books in plain text from Project Gutenberg
#
# + colab={"base_uri": "https://localhost:8080/"} id="KSjUCa3dA946" outputId="ff598b6e-fd83-43a9-f6cc-c893942f04bd"
# !wget https://gutenberg.net.au/ebooks01/0100021.txt # en 1984
# + [markdown] id="YqzxYgyiqGUT"
# ## Tokenization
#
# + id="aT7Hj51iBJSL" colab={"base_uri": "https://localhost:8080/"} outputId="b0ef625a-e923-40d9-bf5a-410fa989c46e"
train_text = open("0100021.txt").read()
train_text = train_text.replace('\n\n','\n<p>\n')
print(train_text[3000:3300])
toks = train_text.split()
toks[1000:1020]
# + [markdown] id="usLfD8d9ptLK"
# # Neural Model
#
# [expit](https://docs.scipy.org/doc/scipy/reference/generated/scipy.special.expit.html) is the logistic sigmoid
#
# + id="jg9Aj_Zjqp_U"
from scipy.special import expit
dim = 30
neg_examples = 0
# + [markdown] id="6jMDw8ll3jnT"
# `vocab` maps word ID to string, `w2id` maps a word to its ID, `wfrq` contains frequences of all words in `vocab`, `prob` contains respective probabilities
#
# + colab={"base_uri": "https://localhost:8080/"} id="PyWX9yBUquFo" outputId="8f428dc4-c57f-4b8f-acd6-de78148e05e9"
vocab = list(set(toks))
w2id = {w:i for (i,w) in enumerate(vocab)}
wfrq = np.zeros(len(vocab))
tokIDs = [w2id[w] for w in toks]
for id in tokIDs:
wfrq[id] += 1
wprob = wfrq/sum(wfrq)
print(len(vocab), w2id['a'], wfrq[w2id['a']], vocab[:4], wfrq[:4])
print(len(toks), len(tokIDs), wprob)
# + [markdown] id="Ul_FW7si4CT4"
# `node_vec` and `ctx_vec` are matrices containding a word embedding vector for each word
#
# We train them on pairs of words *(w1, w2)*, *w2* follows *w1*, an embedding of *w1* in `ctx_vec` should be close to an embedding of *w2* in `node_vec`.
# + id="AWNbBzDjsXTL"
node_vec = np.random.rand(len(vocab), dim)
ctx_vec = np.zeros((len(vocab), dim))
# + colab={"base_uri": "https://localhost:8080/"} id="8C4A5pBuvQ21" outputId="3301f5cd-470f-48ea-c700-c77755c9ce89"
wfrq, len(wfrq)
# + id="jpGcUV-3tX0x"
def train_pair(nodeid, ctxid, alpha):
global node_vec, ctx_vec
L1 = node_vec[nodeid]
L2 = ctx_vec[ctxid]
corr = (1 - expit(np.dot(L2, L1)))* alpha
node_vec[nodeid] += corr * (L2 - L1)
ctx_vec[ctxid] += corr * (L1 - L2)
if neg_examples == 0:
return
negs = np.random.choice(len(vocab), neg_examples, p=wprob)
L2n = ctx_vec[negs]
corrn = expit(np.dot(L2n, L1))* alpha
#node_vec[nodeid] += corr * (L2 - L1)
L2n += corr * (L2n - L1)
def tranin_corpus(epochs=10, start_alpha=0.5):
parcnt = 0
last_parcnt = 0
parid = w2id['<p>']
total_parcnt = float(epochs * wfrq[parid])
alpha = start_alpha
for e in range(epochs):
print('epoch:', e, 'paragraphs:', parcnt, 'alpha:', alpha)
last = tokIDs[0]
for wid in tokIDs[1:]:
if wid == parid:
parcnt += 1
train_pair(wid, last, alpha)
last = wid
if parcnt >= last_parcnt + 200:
a = start_alpha * (1 - parcnt/total_parcnt)
alpha = max(a, start_alpha * 0.0001)
# + id="dWvc1IaPBEGP" colab={"base_uri": "https://localhost:8080/"} outputId="1067b469-2633-4bfb-fe79-686730323fb5"
tranin_corpus(100)
# + [markdown] id="rBAtQsF-135Y"
# ### Similarity function
# find most similar words for the given one, it finds the most probable following word with default `src` and `tar` parameters
#
# + id="NDX_1_ia30QA"
def sims(word, maxitems=5, src=None, tar=None):
if src is None:
src = ctx_vec
if tar is None:
tar = node_vec
wid = w2id[word]
norms = np.linalg.norm(tar, axis=1)
L1 = src[wid]
allsims = np.dot(tar, L1)
allsims /= norms
allsims /= np.linalg.norm(L1)
top = np.argpartition(allsims, len(allsims) - maxitems -1)[-maxitems -1:]
top = [i for i in top if i != wid]
top.sort(key=lambda i:allsims[i], reverse=True)
return [(vocab[i], round(allsims[i],3)) for i in top]
# + colab={"base_uri": "https://localhost:8080/"} id="bF5BDZrLBC1Z" outputId="37efc5b2-ee04-4794-80ee-ce9627cab0af"
# print following words
for w in 'Brother Big he she said is'.split():
print(w, sims(w))
# + id="8UiKKOEoTcOj" colab={"base_uri": "https://localhost:8080/"} outputId="cd13ebe4-e6bc-475c-b9e0-2ee5e303a466"
# print similar words
for w in 'she small years'.split():
print(w, sims(w, 5, node_vec, node_vec))
# + id="Dq0k5rFmDfxQ"
import random
def generate_text(seed='We', words=20):
text = seed
for _ in range(words):
next_words = sims(seed)
selected_word = random.choice(next_words)[0]
text += " " + selected_word
seed = selected_word
return text
# + colab={"base_uri": "https://localhost:8080/"} id="lzFZ1OM4DoIf" outputId="41de1e85-0383-4938-93c8-8b1ee73b47ac"
print(generate_text('We'))
print(generate_text('We'))
print(generate_text('We'))
| Language_modeling/IA161_Language_modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy arrays
# NumPy arrays, *ndarray*, are more efficient than Python native lists when it comes to numerical operations.
# Although *ndarrays* allows various types as elements, but, remember, every element has to be the same type.
# The calculations are over entire NumPy arrays.
#
# ## Create arrays
# Using numpy.arange() or numpy.array()
# +
import numpy as np
a1=np.arange(5) # arange() is similar like range(), but generates ndraary instead of list
a2=np.array([0,1,2,3,4])
a3=np.array([[0,1,2,3],[0,1,4,9]]) # Multi-dimentions is avialable through applying the function array()
# with list of list or list of ndarray.
print('object type of a1 is ', type(a1), "and it's content is ",a1)
print('object type of a2 is ', type(a2), "and it's content is ",a2)
print('content of a3 is ',a3,',and a3[1][3] uses to access that element equals to ', a3[1][3])
'''The outputs would be:
object type of a1 is <class 'numpy.ndarray'> and it's content is [0 1 2 3 4]
object type of a2 is <class 'numpy.ndarray'> and it's content is [0 1 2 3 4]
content of a3 is [[0 1 2 3]
[0 1 4 9]] ,and a3[1][3] uses to access that element equals to 9
'''
# -
# ## Information of arrays
# The following classes provide some properties for related information of arrays:
# - dtype
# - ndim
# - shape
# - size
a3=np.array([[0,1,2,3,4],[0,1,4,9,16]])
a3.ndim
| Chapter 1 - Essential packages/Essential pieces of NumPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''anaconda3'': virtualenv)'
# metadata:
# interpreter:
# hash: 4e4b1c0eba25b43fb0828087952fe1bfe39b80460d2a5f528a72e68cadaca174
# name: python3
# ---
# # Eco Score Pipeline for beautofuel
# +
# %load_ext autoreload
# %autoreload 2
from influxdb import DataFrameClient
from lib.utils.constants import INFLUXDB_HOST, INFLUXDB_PORT, INFLUXDB_USER, INFLUXDB_PASSWORD, INFLUXDB_DB
# -
# ## Pipeline configuration
# +
# Database client
grafanadb = DataFrameClient(
host=INFLUXDB_HOST,
port=INFLUXDB_PORT,
database=INFLUXDB_DB,
username=INFLUXDB_USER,
password=INFLUXDB_PASSWORD
)
user = "zlatka"
# -
# ## Helper functions
# +
def get_query_result_value(result, field):
return result['tracks'][field][0]
def get_query_end(phase):
return "FROM \"tracks\" WHERE (\"phase\"='{}' AND \"user\"='{}')".format(phase, user)
# -
# ## Setup Queries
# Consumption queries
stdddev_consumption_query = "SELECT stddev(\"consumption\") {};".format(get_query_end(1))
mean_consumption_query = "SELECT mean(\"consumption\") {};".format(get_query_end(1))
min_consumption_query = "SELECT min(\"consumption\") {};".format(get_query_end(1))
max_consumption_query = "SELECT max(\"consumption\") {};".format(get_query_end(1))
# ## Execute Queries
# Execute consumption queries
stddev_consumption = get_query_result_value(grafanadb.query(stdddev_consumption_query), 'stddev')
mean_consumption = get_query_result_value(grafanadb.query(mean_consumption_query), 'mean')
min_consumption = get_query_result_value(grafanadb.query(min_consumption_query), 'min')
max_consumption = get_query_result_value(grafanadb.query(max_consumption_query), 'max')
# Print consumption queries results
print("consumption_100_std:", stddev_consumption)
print("consumption_100_mean:", mean_consumption)
print("consumption_100_min:", min_consumption)
print("consumption_100_max:", max_consumption)
# ## Eco-score Calculation
# +
tracks_2_query = "SELECT \"consumption\" {};".format(get_query_end(2))
tracks_2 = grafanadb.query(tracks_2_query)
for _, track_row in tracks_2['tracks'].iterrows():
track_2_consumption = track_row['consumption']
part_50 = None
part_30 = None
part_20 = None
part_10 = None
lower_consumption_limit = mean_consumption - stddev_consumption
upper_consumption_limit = mean_consumption + stddev_consumption
if track_2_consumption >= upper_consumption_limit:
part_50 = 0
elif track_2_consumption <= lower_consumption_limit:
part_50 = 100
else:
track_2_consumption = track_2_consumption - lower_consumption_limit
upper_consumption_limit = upper_consumption_limit - lower_consumption_limit
part_50 = int((track_2_consumption / upper_consumption_limit) * 100)
print("50%:", part_50)
if track_2_consumption < min_consumption:
part_30 = 100
else:
part_30 = 0
print("30%:", part_30)
if track_2_consumption < mean_consumption:
part_20 = 100
else:
part_20 = 0
print("20%:", part_20)
if track_2_consumption < max_consumption:
part_10 = 100
else:
part_10 = 0
print("10%:", part_10)
eco_score = part_50 * 0.5 + part_30 * 0.3 + part_20 * 0.2 + part_10 * 0.1
print("eco_score:", int(eco_score))
print()
# -
| src/updater/app/eco_score.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/verypluming/JapaneseNLI/blob/master/JapaneseXLM_NLI.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="oidzllMksJWm"
# transformersライブラリのXLM: https://github.com/huggingface/transformers をファインチューニングして日本語テキスト推論を試すコード
# + id="SpoDfZd7TNNu"
# 必要なモジュールのインストール
# ! pip install transformers==2.6.0 mecab-python3==0.996.5 tensorflow scikit-learn pandas lxml
# %tensorflow_version 2.x.
# !mkdir data
# !mkdir models
# Google Driveのマウント
from google.colab import drive
drive.mount('/content/drive')
# + id="OnaqlPeLWrj0"
# 必要なモジュール・関数の読み込み
import codecs
import os
import re
import sys
import glob
from collections import Counter
import pandas as pd
import json
import numpy as np
import tensorflow as tf
from transformers import XLMConfig, TFXLMForSequenceClassification, XLMTokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
from tensorflow.keras.callbacks import EarlyStopping
class Vocab:
# 正解ラベルの設定(今回はcontradiction, entailment, neutralの3値を設定)
def __init__(self):
self.token_index = {label: i for i, label in enumerate(["contradiction", "entailment", "neutral"])}
self.index_token = {v: k for k, v in self.token_index.items()}
def encode(self, labels):
label_ids = [self.token_index.get(label) for label in labels]
return label_ids
def decode(self, label_ids):
labels = [self.index_token.get(label_id) for label_id in label_ids]
return labels
@property
def size(self):
return len(self.token_index)
def save(self, file_path):
with open(file_path, 'w') as f:
config = {
'token_index': self.token_index,
'index_token': self.index_token
}
f.write(json.dumps(config))
@classmethod
def load(cls, file_path):
with open(file_path) as f:
config = json.load(f)
vocab = cls()
vocab.token_index = config.token_index
vocab.index_token = config.index_token
return vocab
def convert_examples_to_features(x, y, vocab, max_seq_length, tokenizer):
features = {
'input_ids': [],
'attention_mask': [],
'token_type_ids': [],
'label_ids': np.asarray(vocab.encode(y))
}
for pairs in x:
tokens = [tokenizer.cls_token]
token_type_ids = []
for i, sent in enumerate(pairs):
word_tokens = tokenizer.tokenize(sent)
tokens.extend(word_tokens)
tokens += [tokenizer.sep_token]
len_sent = len(word_tokens) + 1
token_type_ids += [i] * len_sent
input_ids = tokenizer.convert_tokens_to_ids(tokens)
attention_mask = [1] * len(input_ids)
features['input_ids'].append(input_ids)
features['attention_mask'].append(attention_mask)
features['token_type_ids'].append(token_type_ids)
for name in ['input_ids', 'attention_mask', 'token_type_ids']:
features[name] = pad_sequences(features[name], padding='post', maxlen=max_seq_length)
x = [features['input_ids'], features['attention_mask'], features['token_type_ids']]
y = features['label_ids']
return x, y
def build_model(pretrained_model_name_or_path, num_labels):
config = XLMConfig.from_pretrained(
pretrained_model_name_or_path,
num_labels=num_labels
)
model = TFXLMForSequenceClassification.from_pretrained(
pretrained_model_name_or_path,
config=config
)
model.layers[-1].activation = tf.keras.activations.softmax
return model
def evaluate(model, target_vocab, features, labels):
label_ids = model.predict(features)
label_ids = np.argmax(label_ids, axis=-1)
y_pred = target_vocab.decode(label_ids)
y_true = target_vocab.decode(labels)
print(classification_report(y_true, y_pred, digits=4))
# + id="baN8z3fCfLjK"
# ハイパーパラメータの設定
batch_size = 100
epochs = 50
model_path = 'models/'
pretrained_model_name_or_path = 'xlm-mlm-100-1280'
tokenizer = XLMTokenizer.from_pretrained(pretrained_model_name_or_path)
maxlen = 250
target_vocab =Vocab()
# モデルの構築
model = build_model(pretrained_model_name_or_path, target_vocab.size)
model.compile(optimizer='sgd', loss='sparse_categorical_crossentropy')
# callbacksの設定
callbacks = [
EarlyStopping(patience=3),
]
# + id="jQ53YL45VaQm"
# ファインチューニングに用いるデータの読み込み
# 各行にタブ区切りでpremise(前提文)、hypothesis(仮説文)、gold_label(正解ラベル)が書かれたtrain.tsvファイルを用意し、Google Driveにアップロード
# (一行目はpremise, hypothesis, gold_labelと記述)
# データの例
# premise hypothesis gold_label
# 太郎は花子が山頂まで登っている間に、山頂まで登った。 太郎は花子が山頂まで登る前に、山頂まで登った。 entailment
# !cp /content/drive/My\ Drive/train.tsv data/.
df = pd.read_csv("data/train.tsv", sep="\t")
premises = list(df['premise'])
hypotheses = list(df['hypothesis'])
x = [(premise, hypothesis) for (premise, hypothesis) in zip(premises, hypotheses)]
y = list(df['gold_label'])
# + id="ci94_l0BsIVI"
# 全データをファインチューニングに使う場合
x_train = x
y_train = y
# train:testを9:1で分割して評価する場合
#x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.1, random_state=42)
features_train, labels_train = convert_examples_to_features(
x_train,
y_train,
target_vocab,
max_seq_length=maxlen,
tokenizer=tokenizer
)
# モデルのファインチューニング
model.fit(x=features_train,
y=labels_train,
batch_size=batch_size,
epochs=epochs,
validation_split=0.1,
callbacks=callbacks)
model.save_pretrained(model_path)
# + id="QWanq1vpW2Jy"
# ファインチューニングしたモデルをMyDriveに保存
# !cp models/tf_model.h5 /content/drive/My\ Drive/.
# !cp models/config.json /content/drive/My\ Drive/.
# + id="ywLq9ejNW4Dt"
# MyDriveに保存したモデルの読み込み(モデルがある場合に使用)
# !cp /content/drive/My\ Drive/tf_model.h5 models/.
# !cp /content/drive/My\ Drive/config.json models/.
config = XLMConfig.from_json_file('models/config.json')
model = TFXLMForSequenceClassification.from_pretrained('models/tf_model.h5', config=config)
# + id="uLcAGFXJdRQ5"
# 任意のデータでテスト
# x_test: 前提文と仮説文のペアをタプルとしたリスト, y_test: 正解ラベル(entailmentかneutralかcontradiction)のリスト
x_test = [('太郎は花子が山頂まで登っている間に、山頂まで登った。', '太郎は花子が山頂まで登る前に、山頂まで登った。')]
y_test = ['entailment']
features_test, labels_test = convert_examples_to_features(x_test, y_test, target_vocab, max_seq_length=maxlen, tokenizer=tokenizer)
# ラベルを予測
label_ids = model.predict(features_test)
label_ids = np.argmax(label_ids, axis=-1)
y_pred = target_vocab.decode(label_ids)
y_true = target_vocab.decode(labels_test)
print(y_pred, y_true)
# 混同行列の作成
evaluate(model, target_vocab, features_test, labels_test)
| JapaneseXLM_NLI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
#
# + active=""
#
# -
# ##### To communicate with SQL Databases from within a JupyterLab notebook, we can use the SQL "magic" provided by the [ipython-sql](https://github.com/catherinedevlin/ipython-sql) extension. "Magic" is JupyterLab's term for special commands that start with "%". Below, we'll use the _load_\__ext_ magic to load the ipython-sql extension. In the lab environemnt provided in the course the ipython-sql extension is already installed and so is the ibm_db_sa driver.
# %load_ext sql
# ##### Now we have access to SQL magic. With our first SQL magic command, we'll connect to a Db2 database. However, in order to do that, you'll first need to retrieve or create your credentials to access your Db2 database.
# + active=""
#
# +
# Enter your Db2 credentials in the connection string below
# Recall you created Service Credentials in Part III of the first lab of the course in Week 1
# i.e. from the uri field in the Service Credentials copy everything after db2:// (but remove the double quote at the end)
# for example, if your credentials are as in the screenshot above, you would write:
# # %sql ibm_db_sa://my-username:my-password@dashdb-txn-sbox-yp-dal09-03.services.dal.bluemix.net:50000/BLUDB
# Note the ibm_db_sa:// prefix instead of db2://
# This is because JupyterLab's ipython-sql extension uses sqlalchemy (a python SQL toolkit)
# which in turn uses IBM's sqlalchemy dialect: ibm_db_sa
# %sql ibm_db_sa://pzz71823:3kmfnjk22vf8fh%5E9@dashdb-txn-sbox-yp-lon02-07.services.eu-gb.bluemix.net:50000/BLUDB
# -
# ##### For convenience, we can use %%sql (two %'s instead of one) at the top of a cell to indicate we want the entire cell to be treated as SQL. Let's use this to create a table and fill it with some test data for experimenting.
# + language="sql"
#
# CREATE TABLE INTERNATIONAL_STUDENT_TEST_SCORES (
# country VARCHAR(50),
# first_name VARCHAR(50),
# last_name VARCHAR(50),
# test_score INT
# );
# INSERT INTO INTERNATIONAL_STUDENT_TEST_SCORES (country, first_name, last_name, test_score)
# VALUES
# ('United States', 'Marshall', 'Bernadot', 54),
# ('Ghana', 'Celinda', 'Malkin', 51),
# ('Ukraine', 'Guillermo', 'Furze', 53),
# ('Greece', 'Aharon', 'Tunnow', 48),
# ('Russia', 'Bail', 'Goodwin', 46),
# ('Poland', 'Cole', 'Winteringham', 49),
# ('Sweden', 'Emlyn', 'Erricker', 55),
# ('Russia', 'Cathee', 'Sivewright', 49),
# ('China', 'Barny', 'Ingerson', 57),
# ('Uganda', 'Sharla', 'Papaccio', 55),
# ('China', 'Stella', 'Youens', 51),
# ('Poland', 'Julio', 'Buesden', 48),
# ('United States', 'Tiffie', 'Cosely', 58),
# ('Poland', 'Auroora', 'Stiffell', 45),
# ('China', 'Clarita', 'Huet', 52),
# ('Poland', 'Shannon', 'Goulden', 45),
# ('Philippines', 'Emylee', 'Privost', 50),
# ('France', 'Madelina', 'Burk', 49),
# ('China', 'Saunderson', 'Root', 58),
# ('Indonesia', 'Bo', 'Waring', 55),
# ('China', 'Hollis', 'Domotor', 45),
# ('Russia', 'Robbie', 'Collip', 46),
# ('Philippines', 'Davon', 'Donisi', 46),
# ('China', 'Cristabel', 'Radeliffe', 48),
# ('China', 'Wallis', 'Bartleet', 58),
# ('Moldova', 'Arleen', 'Stailey', 38),
# ('Ireland', 'Mendel', 'Grumble', 58),
# ('China', 'Sallyann', 'Exley', 51),
# ('Mexico', 'Kain', 'Swaite', 46),
# ('Indonesia', 'Alonso', 'Bulteel', 45),
# ('Armenia', 'Anatol', 'Tankus', 51),
# ('Indonesia', 'Coralyn', 'Dawkins', 48),
# ('China', 'Deanne', 'Edwinson', 45),
# ('China', 'Georgiana', 'Epple', 51),
# ('Portugal', 'Bartlet', 'Breese', 56),
# ('Azerbaijan', 'Idalina', 'Lukash', 50),
# ('France', 'Livvie', 'Flory', 54),
# ('Malaysia', 'Nonie', 'Borit', 48),
# ('Indonesia', 'Clio', 'Mugg', 47),
# ('Brazil', 'Westley', 'Measor', 48),
# ('Philippines', 'Katrinka', 'Sibbert', 51),
# ('Poland', 'Valentia', 'Mounch', 50),
# ('Norway', 'Sheilah', 'Hedditch', 53),
# ('Papua New Guinea', 'Itch', 'Jubb', 50),
# ('Latvia', 'Stesha', 'Garnson', 53),
# ('Canada', 'Cristionna', 'Wadmore', 46),
# ('China', 'Lianna', 'Gatward', 43),
# ('Guatemala', 'Tanney', 'Vials', 48),
# ('France', 'Alma', 'Zavittieri', 44),
# ('China', 'Alvira', 'Tamas', 50),
# ('United States', 'Shanon', 'Peres', 45),
# ('Sweden', 'Maisey', 'Lynas', 53),
# ('Indonesia', 'Kip', 'Hothersall', 46),
# ('China', 'Cash', 'Landis', 48),
# ('Panama', 'Kennith', 'Digance', 45),
# ('China', 'Ulberto', 'Riggeard', 48),
# ('Switzerland', 'Judy', 'Gilligan', 49),
# ('Philippines', 'Tod', 'Trevaskus', 52),
# ('Brazil', 'Herold', 'Heggs', 44),
# ('Latvia', 'Verney', 'Note', 50),
# ('Poland', 'Temp', 'Ribey', 50),
# ('China', 'Conroy', 'Egdal', 48),
# ('Japan', 'Gabie', 'Alessandone', 47),
# ('Ukraine', 'Devlen', 'Chaperlin', 54),
# ('France', 'Babbette', 'Turner', 51),
# ('Czech Republic', 'Virgil', 'Scotney', 52),
# ('Tajikistan', 'Zorina', 'Bedow', 49),
# ('China', 'Aidan', 'Rudeyeard', 50),
# ('Ireland', 'Saunder', 'MacLice', 48),
# ('France', 'Waly', 'Brunstan', 53),
# ('China', 'Gisele', 'Enns', 52),
# ('Peru', 'Mina', 'Winchester', 48),
# ('Japan', 'Torie', 'MacShirrie', 50),
# ('Russia', 'Benjamen', 'Kenford', 51),
# ('China', 'Etan', 'Burn', 53),
# ('Russia', 'Merralee', 'Chaperlin', 38),
# ('Indonesia', 'Lanny', 'Malam', 49),
# ('Canada', 'Wilhelm', 'Deeprose', 54),
# ('Czech Republic', 'Lari', 'Hillhouse', 48),
# ('China', 'Ossie', 'Woodley', 52),
# ('Macedonia', 'April', 'Tyer', 50),
# ('Vietnam', 'Madelon', 'Dansey', 53),
# ('Ukraine', 'Korella', 'McNamee', 52),
# ('Jamaica', 'Linnea', 'Cannam', 43),
# ('China', 'Mart', 'Coling', 52),
# ('Indonesia', 'Marna', 'Causbey', 47),
# ('China', 'Berni', 'Daintier', 55),
# ('Poland', 'Cynthia', 'Hassell', 49),
# ('Canada', 'Carma', 'Schule', 49),
# ('Indonesia', 'Malia', 'Blight', 48),
# ('China', 'Paulo', 'Seivertsen', 47),
# ('Niger', 'Kaylee', 'Hearley', 54),
# ('Japan', 'Maure', 'Jandak', 46),
# ('Argentina', 'Foss', 'Feavers', 45),
# ('Venezuela', 'Ron', 'Leggitt', 60),
# ('Russia', 'Flint', 'Gokes', 40),
# ('China', 'Linet', 'Conelly', 52),
# ('Philippines', 'Nikolas', 'Birtwell', 57),
# ('Australia', 'Eduard', 'Leipelt', 53)
#
# -
# #### Using Python Variables in your SQL Statements
# ##### You can use python variables in your SQL statements by adding a ":" prefix to your python variable names.
# ##### For example, if I have a python variable `country` with a value of `"Canada"`, I can use this variable in a SQL query to find all the rows of students from Canada.
country = "Canada"
# %sql select * from INTERNATIONAL_STUDENT_TEST_SCORES where country = :country
# #### Assigning the Results of Queries to Python Variables
# ##### You can use the normal python assignment syntax to assign the results of your queries to python variables.
# ##### For example, I have a SQL query to retrieve the distribution of test scores (i.e. how many students got each score). I can assign the result of this query to the variable `test_score_distribution` using the `=` operator.
# test_score_distribution = %sql SELECT test_score as "Test Score", count(*) as "Frequency" from INTERNATIONAL_STUDENT_TEST_SCORES GROUP BY test_score;
test_score_distribution
# #### Converting Query Results to DataFrames
# ##### You can easily convert a SQL query result to a pandas dataframe using the `DataFrame()` method. Dataframe objects are much more versatile than SQL query result objects. For example, we can easily graph our test score distribution after converting to a dataframe.
# +
dataframe = test_score_distribution.DataFrame()
# %matplotlib inline
# uncomment the following line if you get an module error saying seaborn not found
# # !pip install seaborn
import seaborn
plot = seaborn.barplot(x='Test Score',y='Frequency', data=dataframe)
# -
# Now you know how to work with Db2 from within JupyterLab notebooks using SQL "magic"!
# + language="sql"
#
# -- Feel free to experiment with the data set provided in this notebook for practice:
# SELECT country, first_name, last_name, test_score FROM INTERNATIONAL_STUDENT_TEST_SCORES;
# + active=""
#
#
| sql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="OZmbd7WIjql5"
# # !pip install face_recognition
# + id="G5ZxzRBTkBCN"
import face_recognition
from pathlib import Path
from PIL import Image
# + id="HSmiEvPZkmFt"
# Load the image of the person we want to find similar people for
known_image = face_recognition.load_image_file("test_face7.jpg")
# + id="iVjmFf4Kko5K"
# Encode the known image
known_image_encoding = face_recognition.face_encodings(known_image)[0]
# + id="Q5LsX-bXkqhY"
# Variables to keep track of the most similar face match we've found
best_face_distance = 1.0
best_face_image = None
# + id="h7asH271kseT"
# Loop over all the images we want to check for similar people
for image_path in Path("people").glob("*.jpg"):
# Load an image to check
unknown_image = face_recognition.load_image_file(image_path)
# Get the location of faces and face encodings for the current image
face_encodings = face_recognition.face_encodings(unknown_image)
# Get the face distance between the known person and all the faces in this image
face_distance = face_recognition.face_distance(face_encodings, known_image_encoding)[0]
# If this face is more similar to our known image than we've seen so far, save it
if face_distance < best_face_distance:
# Save the new best face distance
best_face_distance = face_distance
# Extract a copy of the actual face image itself so we can display it
best_face_image = unknown_image
# + colab={"base_uri": "https://localhost:8080/", "height": 434} id="a-YMUSWukt_U" outputId="717ed190-83b9-46b4-a175-03b5c3cce8c5"
# Display the face image that we found to be the best match!
pil_image = Image.fromarray(best_face_image)
pil_image
| ML/Week10_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # StackOverflow, performance
#
# Main analysis.
import collabclass
import matplotlib.pyplot as plt
import numpy as np
import pickle
# +
with open("../_data/stackoverflow/categories-final.pkl", "rb") as f:
cats = pickle.load(f)
cls2idx = {cls: idx for idx, cls in enumerate(sorted(set(cats.values())))}
k = len(cls2idx)
# +
# %%time
with open("../_data/stackoverflow/workspace.pkl", "rb") as f:
data = pickle.load(f)
user_cnt = 0
item_cnt = 0
user2idx = dict()
item2idx = dict()
edges = list()
for uid, qid, _ in data["edges"]:
if qid not in cats:
# We dropped the question -> skip.
continue
if qid not in item2idx:
item2idx[qid] = item_cnt
item_cnt += 1
if uid not in user2idx:
user2idx[uid] = user_cnt
user_cnt += 1
edges.append((user2idx[uid], item2idx[qid]))
m = user_cnt
n = item_cnt
graph = collabclass.graph_from_edges(m, n, edges)
# -
print("Number of users: {:,}".format(m))
print("Number of items: {:,}".format(n))
print("Number of edges: {:,}".format(len(graph.user_edges)))
idx2item = {v: k for k, v in item2idx.items()}
vs = list()
for j in range(n):
cat = cats[idx2item[j]]
vs.append(cls2idx[cat])
vs = np.array(vs)
np.random.seed(0)
vs_hat = collabclass.symmetric_channel(vs, k, delta=0.1)
# +
vals, cnts = np.unique(vs, return_counts=True)
fig, ax = plt.subplots()
ax.bar(vals, cnts / cnts.sum());
ax.set_xticks(vals)
ax.set_xticklabels(sorted(set(cats.values())), rotation=30, ha="right");
# -
# ## CAVI
alpha = np.ones((m, k))
beta = collabclass.init_beta(k, vs_hat, delta=0.30)
# %%time
apost, bpost = collabclass.cavi(graph, alpha, beta, 3)
vs_bar = np.argmax(bpost, axis=1)
collabclass.print_accuracy(vs_bar, vs, vs_hat)
collabclass.degree_breakdown(vs_bar, vs, graph, qs=(50, 90, 98))
collabclass.confusion_matrix(vs_bar, vs);
# What about if we "loosen" the criterion a bit and look at the top-2 classes?
collabclass.degree_breakdown_topk(bpost, vs, graph, k=2)
# ## wvRN
# %%time
vs_bar = collabclass.wvrn(graph, vs_hat)
collabclass.print_accuracy(vs_bar, vs, vs_hat)
collabclass.degree_breakdown(vs_bar, vs, graph, qs=(50, 90, 95, 99))
collabclass.confusion_matrix(vs_bar, vs);
# ## Deep dive into errors of CAVI
# +
res = np.argsort(bpost, axis=1)[:,::-1]
mistakes = ((res[:,0] != vs) & (res[:,1] != vs))
not_corrupted = (vs_hat == vs)
high_degree = (graph.item_idx[:,1] >= 10)
idx2cls = {v: k for k, v in cls2idx.items()}
for idx in np.argwhere(mistakes & not_corrupted & high_degree).ravel()[:10]:
print("---")
print(f"Actual: {idx2cls[vs[idx]]}, predicted: {idx2cls[vs_bar[idx]]}")
print(f"URL: https://stackoverflow.com/questions/{idx2item[idx]}")
| notebooks/stackoverflow-performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K-Means Algorithm Demo
#
# _Source: 🤖[Homemade Machine Learning](https://github.com/trekhleb/homemade-machine-learning) repository_
#
# > ☝Before moving on with this demo you might want to take a look at:
# > - 📗[Math behind the K-Means Algorithm](https://github.com/trekhleb/homemade-machine-learning/tree/master/homemade/k_means)
# > - ⚙️[K-Means Algorithm Source Code](https://github.com/trekhleb/homemade-machine-learning/blob/master/homemade/k_means/k_means.py)
#
# **K-means clustering** aims to partition _n_ observations into _K_ clusters in which each observation belongs to the cluster with the nearest mean, serving as a prototype of the cluster.
#
# > **Demo Project:** In this example we will try to cluster Iris flowers into tree categories that we don't know in advance based on `petal_length` and `petal_width` parameters using K-Means unsupervised learning algorithm.
# +
# To make debugging of logistic_regression module easier we enable imported modules autoreloading feature.
# By doing this you may change the code of logistic_regression library and all these changes will be available here.
# %load_ext autoreload
# %autoreload 2
# Add project root folder to module loading paths.
import sys
sys.path.append('../..')
# -
# ### Import Dependencies
#
# - [pandas](https://pandas.pydata.org/) - library that we will use for loading and displaying the data in a table
# - [numpy](http://www.numpy.org/) - library that we will use for linear algebra operations
# - [matplotlib](https://matplotlib.org/) - library that we will use for plotting the data
# - [k_means](https://github.com/trekhleb/homemade-machine-learning/blob/master/homemade/k_means/k_means.py) - custom implementation of K-Means algorithm
# +
# Import 3rd party dependencies.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# Import custom k-means implementation.
from homemade.k_means import KMeans
# -
# ### Load the Data
#
# In this demo we will use [Iris data set](http://archive.ics.uci.edu/ml/datasets/Iris).
#
# The data set consists of several samples from each of three species of Iris (`Iris setosa`, `Iris virginica` and `Iris versicolor`). Four features were measured from each sample: the length and the width of the sepals and petals, in centimeters. Based on the combination of these four features, [Ronald Fisher](https://en.wikipedia.org/wiki/Iris_flower_data_set) developed a linear discriminant model to distinguish the species from each other.
# +
# Load the data.
data = pd.read_csv('../../data/iris.csv')
# Print the data table.
data.head(10)
# -
# ### Plot the Data
#
# Let's take two parameters `petal_length` and `petal_width` for each flower into consideration and plot the dependency of the Iris class on these two parameters.
#
# Since we have an advantage of knowing the actual flower labels (classes) let's illustrate the real-world classification on the plot. But K-Means algorithm is an example of unsuervised learning algorithm which means that this algorithm doesn't need to know about labels. Thus below in this demo we will try to split Iris flowers into unknown clusters and compare the result of such split with the actual flower classification.
# +
# List of suppported Iris classes.
iris_types = ['SETOSA', 'VERSICOLOR', 'VIRGINICA']
# Pick the Iris parameters for consideration.
x_axis = 'petal_length'
y_axis = 'petal_width'
# Make the plot a little bit bigger than default one.
plt.figure(figsize=(12, 5))
# Plot the scatter for every type of Iris.
# This is the case when we know flower labels in advance.
plt.subplot(1, 2, 1)
for iris_type in iris_types:
plt.scatter(
data[x_axis][data['class'] == iris_type],
data[y_axis][data['class'] == iris_type],
label=iris_type
)
plt.xlabel(x_axis + ' (cm)')
plt.ylabel(y_axis + ' (cm)')
plt.title('Iris Types (labels are known)')
plt.legend()
# Plot non-classified scatter of Iris flowers.
# This is the case when we don't know flower labels in advance.
# This is how K-Means sees the dataset.
plt.subplot(1, 2, 2)
plt.scatter(
data[x_axis][:],
data[y_axis][:],
)
plt.xlabel(x_axis + ' (cm)')
plt.ylabel(y_axis + ' (cm)')
plt.title('Iris Types (labels are NOT known)')
# Plot all subplots.
plt.show()
# -
# ### Prepara the Data for Training
#
# Let's extract `petal_length` and `petal_width` data and form a training feature set.
# +
# Get total number of Iris examples.
num_examples = data.shape[0]
# Get features.
x_train = data[[x_axis, y_axis]].values.reshape((num_examples, 2))
# -
# ### Init and Train Logistic Regression Model
#
# > ☝🏻This is the place where you might want to play with model configuration.
#
# - `num_clusters` - number of clusters into which we want to split our training dataset.
# - `max_iterations` - maximum number of training iterations.
# +
# Set K-Means parameters.
num_clusters = 3 # Number of clusters into which we want to split our training dataset.
max_iterations = 50 # maximum number of training iterations.
# Init K-Means instance.
k_means = KMeans(x_train, num_clusters)
# Train K-Means instance.
(centroids, closest_centroids_ids) = k_means.train(max_iterations)
# -
# # Plot the Clustering Results
#
# Now let's plot the original Iris flow classification along with our unsupervised K-Means clusters to see how the algorithm performed.
# +
# List of suppported Iris classes.
iris_types = ['SETOSA', 'VERSICOLOR', 'VIRGINICA']
# Pick the Iris parameters for consideration.
x_axis = 'petal_length'
y_axis = 'petal_width'
# Make the plot a little bit bigger than default one.
plt.figure(figsize=(12, 5))
# Plot ACTUAL Iris flower classification.
plt.subplot(1, 2, 1)
for iris_type in iris_types:
plt.scatter(
data[x_axis][data['class'] == iris_type],
data[y_axis][data['class'] == iris_type],
label=iris_type
)
plt.xlabel(x_axis + ' (cm)')
plt.ylabel(y_axis + ' (cm)')
plt.title('Iris Real-World Clusters')
plt.legend()
# Plot UNSUPERWISED Iris flower classification.
plt.subplot(1, 2, 2)
for centroid_id, centroid in enumerate(centroids):
current_examples_indices = (closest_centroids_ids == centroid_id).flatten()
plt.scatter(
data[x_axis][current_examples_indices],
data[y_axis][current_examples_indices],
label='Cluster #' + str(centroid_id)
)
# Plot clusters centroids.
for centroid_id, centroid in enumerate(centroids):
plt.scatter(centroid[0], centroid[1], c='black', marker='x')
plt.xlabel(x_axis + ' (cm)')
plt.ylabel(y_axis + ' (cm)')
plt.title('Iris K-Means Clusters')
plt.legend()
# Show all subplots.
plt.show()
| notebooks/k_means/k_means_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from os.path import join
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from sklearn.neighbors import NearestNeighbors
from sklearn.mixture import GaussianMixture
from sklearn.cluster import MeanShift, DBSCAN, estimate_bandwidth
# -
# ## Import preprocessed data
df = pd.read_csv(join('..', 'data', 'tugas_preprocessed.csv'))
df.head()
df.columns
# Splitting feature names into groups
non_metric_features = df.columns[df.columns.str.startswith('x')]
pc_features = df.columns[df.columns.str.startswith('PC')]
metric_features = df.columns[~df.columns.str.startswith('x') & ~df.columns.str.startswith('PC')]
def get_ss(df):
"""Computes the sum of squares for all variables given a dataset
"""
ss = np.sum(df.var() * (df.count() - 1))
return ss # return sum of sum of squares of each df variable
# # Density Based Clustering
# ## Mean Shift Clustering
# What is Mean-shift clustering? How does it work?
#
# Single seed | Multiple seeds
# :-------------------------:|:-------------------------:
#  | 
#
# ### Characteristics:
# - No need to define number of clusters apriori
# - Can detect clusters of any shape
# - Robust to outliers
# - Depends on the bandwidth hyperparameter (but there's a way to estimate it)
# - **Main drawback**: Poor scalability (on both the algorithm and in estimating the bandwidth)
# The following bandwidth can be automatically detected using (we need to set quantile though)
# Based on distance to nearest neighbors for all observations
bandwidth = estimate_bandwidth(df[metric_features], random_state=1, n_jobs=-1 , quantile=.055)
# TO-DO: manipulate the quantile value such that we obtain a small enough bandwidth
bandwidth
# +
# Perform mean-shift clustering with bandwidth set using estimate_bandwidth
# TO-DO: explore the MeanShift class and obtain the cluster labels
ms = MeanShift(bandwidth=bandwidth, bin_seeding=True, n_jobs=-1)
ms_labels = ms.fit_predict(df[metric_features])
ms_n_clusters = len(np.unique(ms_labels))
print("Number of estimated clusters : %d" % ms_n_clusters)
# -
from collections import Counter
Counter(ms_labels)
# Concatenating the labels to df
df_concat = pd.concat([df[metric_features], pd.Series(ms_labels, index=df.index, name="ms_labels")], axis=1)
df_concat.head()
# Computing the R^2 of the cluster solution
sst = get_ss(df[metric_features]) # get total sum of squares
ssw_labels = df_concat.groupby(by='ms_labels').apply(get_ss) # compute ssw for each cluster labels
ssb = sst - np.sum(ssw_labels) # remember: SST = SSW + SSB
r2 = ssb / sst
print("Cluster solution with R^2 of %0.4f" % r2)
# ## DBSCAN (Density-based spatial clustering of applications with noise)
# What is DBSCAN clustering? How does it work?
#
# DBSCAN animation | Core, border and noise
# :-------------------------:|:-------------------------:
#  | 
#
#
# ### Characteristics:
# - No need to define number of clusters apriori
# - Resistant to noise and outliers
# - Can identify outliers
# - Can handle clusters of different shapes and sizes
# - Depends highly on the epsilon hyperparameter and it can be hard to tune
# - Does not work well with clusters of varying densities
# +
# Perform DBSCAN clustering
# TO-DO: explore the DBSCAN class and obtain the cluster labels
dbscan = DBSCAN(eps=.5, min_samples=5,n_jobs=-1)
dbscan_labels = dbscan.fit_predict(df[metric_features])
dbscan_n_clusters = len(np.unique(dbscan_labels))
print("Number of estimated clusters : %d" % dbscan_n_clusters)
# +
#Counter(dbscan_labels)
# -
# ### Defining eps and min_samples:
# - **MinPts**: As a rule of thumb, **minPts = 2 x dim** can be used, but it may be necessary to choose larger values for very large data, for noisy data or for data that contains many duplicates.
#
# - **ε**: The value for ε can then be chosen by using a **k-distance graph**, plotting the distance to the kth (k = minPts - 1) nearest neighbor ordered from the largest to the smallest value. Good values of ε are where this plot shows an **"elbow"**: if ε is chosen much too small, a large part of the data will not be clustered; whereas for a too high value of ε, clusters will merge and the majority of objects will be in the same cluster. **The assumption is that for points in a cluster, their k nearest neighbors are at roughly the same distance**. Noise points have their k-th nearest neighbors at farther distance
len(metric_features)
#best practice is: min number of observations per feature - 1
# K-distance graph to find out the right eps value
neigh = NearestNeighbors(n_neighbors=19)
neigh.fit(df[metric_features])
distances, _ = neigh.kneighbors(df[metric_features])
distances = np.sort(distances[:, -1])
plt.plot(distances)
plt.show()
# +
# Perform DBSCAN clustering
# TO-DO: Re-cluster the data using the defined hyperparameters
dbscan = DBSCAN(eps=1.75, min_samples=20,n_jobs=-1)
dbscan_labels = dbscan.fit_predict(df[metric_features])
dbscan_n_clusters = len(np.unique(dbscan_labels))
print("Number of estimated clusters : %d" % dbscan_n_clusters)
# -
#its a good way to find outliers, but not a great way to find clusters
Counter(dbscan_labels)
# Concatenating the labels to df
df_concat = pd.concat([df[metric_features], pd.Series(dbscan_labels, index=df.index, name="dbscan_labels")], axis=1)
df_concat.head()
# Detecting noise (potential outliers)
# TO-DO: can we identify the noisy data?
df_concat[df_concat['dbscan_labels']==-1]
# Computing the R^2 of the cluster solution
df_nonoise = df_concat[df_concat['dbscan_labels']!=-1]# TO-DO: Remove the noisy data (we don't want to include it in the R² computation. Why?)
sst = get_ss(df_nonoise) # get total sum of squares
ssw_labels = df_nonoise.groupby(by='dbscan_labels').apply(get_ss) # compute ssw for each cluster labels
ssb = sst - np.sum(ssw_labels) # remember: SST = SSW + SSB
r2 = ssb / sst
print("Cluster solution with R^2 of %0.4f" % r2)
# - Why did the DBSCAN gave us just one cluster?
# - What can we do with the noisy data?
# ## GMM (Gaussian Mixture Model )
# What is GMM? How does it work?
# 
#
# ---
#
# $$\mathcal{p(\vec{x})} \ = \ \sum_{i=1}^K \phi_i \mathcal{N}(\vec{x}|\vec{\mu_i}, \Sigma_i) \tag{eq1}$$
# $$\mathcal{N}(\vec{x}|\vec{\mu_i}, \Sigma_i) \ = \ \frac{1}{\sqrt{{(2\pi)}^{K}|\Sigma_i|}}e^{-\frac{1}{2} (\vec{x} - \vec{\mu_i})^T \Sigma_i^{-1} (\vec{x} - \vec{\mu_i})} \tag{eq2}$$
# $$\sum_{i=1}^K \phi_i \ = \ 1 \tag{eq3}$$
#
# , where:
# - $\phi_i$ is the component weight (scalar) for Component $i$ (probability of an observation being generated by Component $i$)
# - $\vec{\mu_i}$ is the mean vector for Component $i$,
# - $\Sigma_i$ is the Covariance matrix for Component $i$
#
# ---
#
# - **(eq1)** gives the probability of a point $x$ given the estimated Gaussian mixture
# - **(eq2)** is the probability density function of a multivariate Gaussian with mean $\vec{\mu_i}$ and covariance $\Sigma_i$
# - **(eq3)** states that the sum of the component weights is 1, such that the total probability distribution normalizes to 1
#
# ### Characteristics:
# - Assumes the data is generated from a mixture of finite number of Gaussian distributions with unknown parameters
# - Use the EM (Expectation Maximization algorithm) to estimate the parameters
# - Provides a probability of each observation belonging to each cluster
# - Advantages over K-Means:
# - Can deal with spherical and elipsoid cluster shapes
# - Number of components needs to be defined apriori
# Performing GMM clustering
# TO-DO: explore the GaussianMixture class and obtain the cluster labels and the cluster probabilities
gmm = GaussianMixture(n_components=4,n_init=10,init_params='kmeans',random_state=9)
gmm_labels = gmm.fit_predict(df[metric_features])
labels_proba = gmm.predict_proba(df[metric_features])
gmm_labels, labels_proba
# **Let's look at the estimated parameters:**
# The estimated component weights
gmm.weights_
# The estimated mean vectors of the Components
gmm.means_
# The estimated covariance matrices of the Components
gmm.covariances_.shape
# ### Defining covariance_type:
# This hyperparameter controls the **degrees of freedom** in the shape of each cluster The more degrees of freedom we have the more complex shapes the model can fit and the more computationally expensive the model will be.
#
# 
#
# - `covariance_type="tied"` makes all components share the same general covariance matrix
# ### Defining n_components:
#
# **AIC**: estimates the relative amount of information lost by a model used to represent the data-generation process. The smaller the better.
#
# **BIC**: similar to AIC but penalizes more complex models (i.e. favors simpler models). The smaller the better.
# +
# Selecting number of components based on AIC and BIC
n_components = np.arange(1, 16)
models = [GaussianMixture(n, covariance_type='full', n_init=10, random_state=1).fit(df[metric_features])
for n in n_components]
# TO-DO: compute the aic and bic values for each cluster solution. Use the appropriate GMM methods.
#we want the lowest values here:
bic_values = [gmm.bic(df[metric_features]) for gmm in models]
aic_values = [gmm.aic(df[metric_features]) for gmm in models]
plt.plot(n_components, bic_values, label='BIC')
plt.plot(n_components, aic_values, label='AIC')
plt.legend(loc='best')
plt.xlabel('n_components')
plt.xticks(n_components)
plt.show()
# +
#number of componendts to keep: 3-5
# -
# **Note**: the AIC and BIC measures can also be used to select diferent hyperparameters such as the covariance_type
# Performing GMM clustering
# TO-DO: Re-cluster the data using the defined hyperparameters
gmm = GaussianMixture(5, covariance_type='full', n_init=10, random_state=1)
gmm_labels = gmm.fit_predict(df[metric_features])
Counter(gmm_labels)
# Concatenating the labels to df
df_concat = pd.concat([df[metric_features], pd.Series(gmm_labels, index=df.index, name="gmm_labels")], axis=1)
df_concat.groupby('gmm_labels').mean()
# Computing the R^2 of the cluster solution
sst = get_ss(df[metric_features]) # get total sum of squares
ssw_labels = df_concat.groupby(by='gmm_labels').apply(get_ss) # compute ssw for each cluster labels
ssb = sst - np.sum(ssw_labels) # remember: SST = SSW + SSB
r2 = ssb / sst
print("Cluster solution with R^2 of %0.4f" % r2)
# # Clustering by Perspectives
# - Demographic Perspective:
# - Value Perspective:
# - Product Perspective:
# ## Merging the Perspectives
# - How can we merge different cluster solutions?
| notebooks/lab12_density_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
# In this chapter, you will start the journey by laying out the foundation of modern mathematics – set algebra and discrete math. You will be exposed to the definitions and essential properties of sets and various operations associated with them including combinatorics. Moreover, you will also learn how to use built-in Python libraries to implement these mathematical concepts yourself.
#
# In the second half of the chapter, you will get familiarized with properties of numbers and series. We will examine definition and properties of various types of numbers and series that are encountered in data science and machine learning domain for analysis and modeling – prime numbers, real and complex domain, continued fraction, arithmetic and geometric series, Fibonacci series, etc.
# ## HEADING 1: Concept of set, definition of set relationships, and null set
# Set theory is a branch of mathematical logic that studies sets, which informally are collections of objects. Although any type of object can be collected into a set, set theory is applied most often to objects that are relevant to mathematics. The language of set theory can be used in the definitions of nearly all mathematical objects.
#
# **Set theory is commonly employed as a foundational system for modern mathematics**, particularly in the form of [**Zermelo–Fraenkel set theory**](https://en.wikipedia.org/wiki/Zermelo-Fraenkel_set_theory) with the [axiom of choice](https://en.wikipedia.org/wiki/Axiom_of_choice).
# ### Let's create a set using Python with the `set` and `{...}`
# Directly with curly braces
Set1 = {1,2}
print (Set1)
type(Set1)
# By calling the 'set' function i.e. typecasting
Set2 = set({2,3})
print(Set2)
# By typecasting from another Python object e.g. list
my_list=[1,2,3,4]
my_set_from_list = set(my_list)
print(my_set_from_list)
# #### Let's check the size of a set using `len` function
print("Size of the set created from the list with 4 elements:",len(my_set_from_list))
# ### Empty (Null) set is a special set
# Null set is a set which has no element.
#
# $$ \forall x, x \notin \varnothing $$
# #### Do not try to create the empty set by declaring an empty {}. That denotes an empty dictionary object
null_set = {}
print(type(null_set))
# #### Instead, use the `set()` function to create the empty (null) set from any empty data type e.g. dictionary or list
a = {}
print("Type of 'a':",type(a))
null_set = set(a)
print("Type after casting:",type(null_set))
b = []
print("Type of 'b':",type(b))
null_set_2 = set(b)
print("Type after casting:",type(null_set_2))
# #### Check the size of a Null set
print("Size of a null set:",len(null_set_2))
# ### Subset and Superset
# A set `A` is called a **subset** of another set `B` (or equivalently `B` is called the **superset** of `A`) if all the elements of `A` is also contained by `B`. Note, `B` may contain additional elements.
#
# Subset/superset relationship is shown as follows,
#
# $$ {\displaystyle A\subseteq B} $$
# $$ {\displaystyle B\supseteq A} $$
# #### Let us define two sets using the list comprehension technique of Python. One set will be all integeres from 1 to 10 and another will be only the odd integers between 1 and 10 (inclusive).
# +
set_A = set([i for i in range(1,11) if i%2==1])
set_B = set([i for i in range(1,11)])
print("set_A: ",set_A)
print("set_B: ",set_B)
# -
# #### Python's set objects provide built-in methods (functions) to check for subset or superset property. Use `issubset` or `issuperset` method for this.
set_A.issubset(set_B)
set_B.issubset(set_A)
set_B.issuperset(set_A)
# #### Alternatively, operators like `<=` or `>` can be used to check relations
set_A > set_B
set_B >= set_A
# #### Every set is both subset and superset of itself
set_A.issubset(set_A)
set_B.issuperset(set_B)
# #### Null set if subset to every other set (and every set is superset to the null set)
null_set.issubset(set_A)
set_B.issuperset(null_set_2)
# ### Membership testing using `in` and `not in`
# We can test whether an element belongs to a set or not using familiar Python keyword `in` or `not in`
2 in set_A
11 in set_B
5 in set_A
# ### Disjoint
# Two sets are said to be disjoint sets if they have **no element in common**. Equivalently, disjoint sets are sets whose **intersection** is the null set. We will study interesection little later but as the name implies it simply means the common cross-section between two sets. If there is no common element then the degree of commonality is zero and those two sets are disjoint.
set_C = set([i for i in range(1,11) if i%2==0])
print(set_C)
set_A.isdisjoint(set_C)
# ## HEADING 2: Properties of sets and basic set algebra
# In this section, we will study basic properties of set and the fundamental operations associated with them.
# ### Algebra of inclusion
# If ***`A`***, ***`B`*** and ***`C`*** are sets then the following hold:
#
# **Reflexivity:**
#
# $$ {\displaystyle A\subseteq A} $$
#
# **Antisymmetry:**
#
# $$ A\subseteq B\ \ and\ B\subseteq A\ \text if\ and\ only\ if\ A=B $$
#
# **Transitivity:**
#
# $$If\ {\displaystyle A\subseteq B}\ and \ {\displaystyle B\subseteq C}, then\ A ⊆ C $$
# ### Let's write a short Python program to verify the transivity rule
# +
A = {1,5}
B = {1,2,3,4,5}
C = set([i for i in range(1,11)])
print("A:",A)
print("B:",B)
print("C:",C)
# -
A.issubset(B)
B.issubset(C)
A.issubset(C)
# ### Equality
S1 = {1,2}
S2 = {2,2,1,1,2}
print ("S1 and S2 are equal because order or repetition of elements do not matter for sets\nS1==S2:", S1==S2)
S3 = {1,2,3,4,5,6}
S4 = {1,2,3,4,0,6}
print ("S3 and S4 are NOT equal because at least one element is different\nS3==S4:", S3==S4)
# ### Intersection of sets
# The intersection `A ∩ B` of two sets `A` and `B` is the set that contains all elements of `A` that also belong to `B` (or equivalently, all elements of `B` that also belong to `A`), but no other elements. Formally,
#
# $$ {\displaystyle A\cap B=\{x:x\in A{\text{ and }}x\in B\}.} $$
#
# 
# Define a set using list comprehension
S5 = set([x for x in range(1,11) if x%3==0])
print("S5:", S5)
S6 = set([x for x in range(1,5)])
print("S6:", S6)
# #### Using `intersection` method
S_intersection = S5.intersection(S6)
print("Intersection of S5 and S6:", S_intersection)
# #### Using `&` operator
S_intersection = S5 & S6
print("Intersection of S5 and S6:", S_intersection)
# #### We can chain the methods to get intersection between more than two sets
S7 = set([x for x in range(4,10)])
print("S7:", S7)
S5_S6_S7 = S5.intersection(S6).intersection(S7)
print("Intersection of S5, S6, and S7:", S5_S6_S7)
# #### Now modify `S7` to contain 3 and repeat the exercise above
S7 = set([x for x in range(3,10)])
S5_S6_S7 = S5.intersection(S6).intersection(S7)
print("Intersection of S5, S6, and S7:", S5_S6_S7)
# #### The symbol '&' can be used for intersection
A = {1, 2, 3}
B = {5,3,1}
print("Intersection of {} and {} is: {} with size {}".format(A,B,A&B,len(A&B)))
# ### Union of sets
# In set theory, the union (denoted by ∪) of a collection of sets is the set of all elements in the collection. It is one of the fundamental operations through which sets can be combined and related to each other. Formally,
#
# $$ {\displaystyle A\cup B=\{x:x\in A{\text{ or }}x\in B\}}$$
# #### Both `union` method or `|` can be used
# +
S1 = set([x for x in range(1,11) if x%3==0])
print("S1:", S1)
S2 = set([x for x in range(1,5)])
print("S2:", S2)
S_union = S1.union(S2)
print("Union of S1 and S2:", S_union)
S_union = S1 | S2
print("Union of S1 and S2:", S_union)
# -
# #### Just like intersection, chaining can be done to combine more than two sets
S3 = set([5*x for x in range(1,3)])
print("S3:", S3)
S4 = set ([7,8])
print("S4:", S4)
S1_S2_S3_S4 = S1.union(S2).union(S3).union(S4)
print("Union of S1, S2, S3, and S4:", S1_S2_S3_S4)
# ### Some algebraic identities (laws)
# For any three sets `A`, `B`, and `C`, following laws hold,
#
# **Commutative law:**
# $$ {\displaystyle A\cap B=B\cap A} $$
# $$ {\displaystyle A\cup B=B\cup A} $$
#
# **Associative law:**
# $$ {\displaystyle (A\cap B)\cap C=A\cap (B\cap C)} $$
# $$ {\displaystyle A\cup (B\cup C)=(A\cup B)\cup C} $$
#
# ** Distributive law:**
# $$ {\displaystyle A\cap (B\cup C)=(A\cap B)\cup (A\cap C)} $$
# $$ {\displaystyle A\cup (B\cap C)=(A\cup B)\cap (A\cup C)} $$
# ### More algebra of inclusion involving union and intersection
# If `A`, `B` and `C` are subsets of a set `S` then the following hold:
#
# **Existence of a least element and a greatest element:**
#
# $$ {\displaystyle \varnothing \subseteq A\subseteq S} $$
#
# **Existence of joins:**
#
# $$ {\displaystyle A\subseteq A\cup B} $$
#
# $$ If\ {\displaystyle A\subseteq C}\ and\ {\displaystyle B\subseteq C,}\ then\ {\displaystyle A\cup B\subseteq C} $$
#
# **Existence of meets:**
# $$ {\displaystyle A\cap B\subseteq A} $$
#
# $$ If\ {\displaystyle C\subseteq A}\ and\ {\displaystyle C\subseteq B,}\ then\ {\displaystyle C\subseteq A\cap B} $$
#
# ### Let's write a short Python program to verify the *existence of joins*
# +
A = {1,5}
B = {1,3,4}
S = set([i for i in range(1,11)])
print("A:",A)
print("B:",B)
print("S:",S)
# -
A_union_B=A.union(B)
print(A_union_B)
A_union_B.issubset(S)
# ### Venn diagrams
# A Venn diagram, named after English mathematician <NAME>, is an extremely useful and intuitive visual tool for representing sets and their inter-relationships.
import matplotlib.pyplot as plt
import matplotlib_venn as venn
S = {1, 2, 3}
T = {0, 2, -1, 5}
venn.venn2([S, T], set_labels=('S','T'))
plt.show()
venn.venn3(subsets = (1, 1, 1, 2, 1, 2, 2), set_labels = ('Set1', 'Set2', 'Set3'))
plt.show()
# #### Three sets' intersection shown in a Venn diagram
# 
# #### Union of three sets shown in this Venn diagram
# 
# ### Complement of a set
# If `A` is a set, then the absolute complement of `A` (or simply the complement of A) is the **set of all elements that are not in `A`**. In other words, if `U` is the universe that contains all the elements under study (and there is no need to mention it because it is obvious and unique), then the absolute complement of `A` is the relative complement of `A` in `U`. Formally,
#
# $$ {\displaystyle A^{\complement }=\{x\in U\mid x\notin A\}.} $$
S=set([x for x in range (21) if x%2==0])
print ("S is the set of even numbers between 0 and 20:", S)
S_complement = set([x for x in range (21) if x%2!=0])
print ("S_complement is the set of odd numbers between 0 and 20:", S_complement)
# You can take the union of two sets and if that is equal to the universal set (in the context of your problem), then you have found the right complement.
print ("Is the union of S and S_complement equal to all numbers between 0 and 20?",
S.union(S_complement)==set([x for x in range (21)]))
# ### Set algebra related to complements
#
# ** De Morgan's laws:**
#
# $$ {\displaystyle \left(A\cup B\right)^{\complement }=A^{\complement }\cap B^{\complement }.} $$
# $$ {\displaystyle \left(A\cap B\right)^{\complement }=A^{\complement }\cup B^{\complement }.} $$
#
# ** Complement laws **
#
# $$ {\displaystyle A\cup A^{\complement }=U.} $$
# $$ {\displaystyle A\cap A^{\complement }=\varnothing .} $$
# $$ {\displaystyle \varnothing ^{\complement }=U.} $$
# $$ {\displaystyle U^{\complement }=\varnothing .} $$
# $$ {\displaystyle {\text{If }}A\subset B{\text{, then }}B^{\complement }\subset A^{\complement }.} $$
# ### Verification of De Morgan's law using a simple Python program
A={-6,3,4,5}
B={-6,5,13}
U=A|B|{12,-2,-4}
print("U:",U)
# #### Function to compute complement of union
def complement_of_union(S1,S2,S3):
Su = S1|S2
S4 = set()
for item in S3:
if item not in Su:
S4.add(item)
return S4
# #### Function to compute intersection of the complements
def intersection_of_complement(S1,S2,S3):
S1C = set()
S2C = set()
for item in S3:
if item not in S1:
S1C.add(item)
for item in S3:
if item not in S2:
S2C.add(item)
return (S1C & S2C)
complement_of_union(A,B,U) == intersection_of_complement(A,B,U)
complement_of_union(A,B,U)
intersection_of_complement(A,B,U)
# ### Difference between sets
# If `A` and `B` are two sets, then the relative complement of `A` in `B`, also termed the **set-theoretic difference of B and A**, is the **set of elements in B but not in A**.
#
# $$ {\displaystyle B\setminus A=\{x\in B\mid x\notin A\}.} $$
#
# 
S1 = set([x for x in range(31) if x%3==0])
print ("Set S1:", S1)
S2 = set([x for x in range(31) if x%5==0])
print ("Set S2:", S2)
# #### Both the `difference` method or `-` operator can be used to compute set-theoretic difference in Python
# +
S2_difference_S1 = S2-S1
print("Difference of S2 and S1 i.e. S2\S1:", S2_difference_S1)
S1_difference_S2 = S1.difference(S2)
print("Difference of S1 and S2 i.e. S1\S2:", S1_difference_S2)
# -
# ### Set algebraic identities involving difference
# ** Following identities can be obtained with algebraic manipulation: **
#
# $$ {\displaystyle C\setminus (A\cap B)=(C\setminus A)\cup (C\setminus B)} $$
# $$ {\displaystyle C\setminus (A\cup B)=(C\setminus A)\cap (C\setminus B)} $$
# $$ {\displaystyle C\setminus (B\setminus A)=(C\cap A)\cup (C\setminus B)} $$
# $$ {\displaystyle C\setminus (C\setminus A)=(C\cap A)} $$
# $$ {\displaystyle (B\setminus A)\cap C=(B\cap C)\setminus A=B\cap (C\setminus A)} $$
# $$ {\displaystyle (B\setminus A)\cup C=(B\cup C)\setminus (A\setminus C)} $$
# $$ {\displaystyle A\setminus A=\emptyset} $$
# $$ {\displaystyle \emptyset \setminus A=\emptyset } $$
# $$ {\displaystyle A\setminus \emptyset =A} $$
# $$ {\displaystyle A\setminus U=\emptyset } $$
# ### Symmetric difference
# In set theory, the ***symmetric difference***, also known as the ***disjunctive union***, of two sets is the **set of elements which are in either of the sets and not in their intersection**.
# $$ {\displaystyle A\,\triangle \,B=\{x:(x\in A)\oplus (x\in B)\}}$$
#
# $$ {\displaystyle A\,\triangle \,B=(A\smallsetminus B)\cup (B\smallsetminus A)} $$
#
# $${\displaystyle A\,\triangle \,B=(A\cup B)\smallsetminus (A\cap B)} $$
#
# 
# #### Some identities related to symmetric difference,
# $$ {\displaystyle A\,\triangle \,B=B\,\triangle \,A,} $$
# $$ {\displaystyle (A\,\triangle \,B)\,\triangle \,C=A\,\triangle \,(B\,\triangle \,C).} $$
#
# **The empty set is neutral, and every set is its own inverse:**
#
# $$ {\displaystyle A\,\triangle \,\varnothing =A,} $$
# $$ {\displaystyle A\,\triangle \,A=\varnothing .} $$
# #### Symmetric difference can be computed by using `symmetric_difference` method or `^` operator
print("S1",S1)
print("S2",S2)
print("Symmetric difference", S1^S2)
print("Symmetric difference", S2.symmetric_difference(S1))
# ### Cartesian product
# In set theory, a Cartesian product is a mathematical operation that returns a set (or product set or simply product) from multiple sets. That is, for sets `A` and `B`, the Cartesian product `A × B` is the set of all ordered pairs (a, b) where a ∈ `A` and b ∈ `B`.
#
# $$ {\displaystyle A\times B=\{\,(a,b)\mid a\in A\ {\mbox{ and }}\ b\in B\,\}.} $$
#
# More generally, a Cartesian product of *n* sets, also known as an n-fold Cartesian product, can be represented by an array of n dimensions, where each element is an *n-tuple*. An ordered pair is a *2-tuple* or couple.
#
# The Cartesian product is named after [<NAME>](https://en.wikipedia.org/wiki/Ren%C3%A9_Descartes) whose formulation of analytic geometry gave rise to the concept
A = set(['a','b','c'])
B = {1,2,3}
# #### Let's write a simple function in Python to compute Cartesian product of two sets
def cartesian_product(S1,S2):
result = set()
for i in S1:
for j in S2:
result.add(tuple([i,j]))
return (result)
C = cartesian_product(A,B)
print("Cartesian product of A and B\n{} X {}: {}".format(A,B,C))
# #### The size of the Cartesian product is naturally the product of the individual sizes of the two sets
# +
A = set(['a','b','c','d'])
B = {1,2}
C = cartesian_product(A,B)
print("Size of A:", len(A))
print("Size of B:", len(B))
print("Size of C:", len(C))
# -
# #### Note that because these are ordered pairs, same element can be repeated inside the pair i.e. even if two sets contain some identical elements, they can be paired up in the Cartesian product
# +
A = {1,2,3,4}
B = {2,3,4}
print ("Cartesian product of {} and {} is:\n{}".format(A,B,cartesian_product(A,B)))
# -
# #### Instead of writing functions ourselves, we could use the `itertools` library of Python. Remember to turn the resulting product object into a list for viewing and subsequent processing
from itertools import product as prod
A = {'a','b','c','d'}
B = {2,3,4}
p=list(prod(A,B))
print (p)
# ### Cartesian Power
#
# The Cartesian square (or binary Cartesian product) of a set X is the Cartesian product $X^2 = X × X$. An example is the 2-dimensional plane $R^2 = R × R$ where _R_ is the set of real numbers: $R^2$ is the set of all points (_x_,_y_) where _x_ and _y_ are real numbers (see the [Cartesian coordinate system](https://en.wikipedia.org/wiki/Cartesian_coordinate_system)).
#
# The cartesian power of a set X can be defined as:
#
# ${\displaystyle X^{n}=\underbrace {X\times X\times \cdots \times X} _{n}=\{(x_{1},\ldots ,x_{n})\ |\ x_{i}\in X{\text{ for all }}i=1,\ldots ,n\}.} $
#
# The [cardinality of a set](https://en.wikipedia.org/wiki/Cardinality) is the number of elements of the set. Cardinality of a Cartesian power set is $|S|^{n}$ where |S| is the cardinality of the set _S_ and _n_ is the power.
#
# __We can easily use itertools again for calculating Cartesian power__. The _repeat_ parameter is used as power.
A = {1,2,3} # 3 element set
print("Set A:",A)
print("\n")
p2=list(prod(A,repeat=2)) # Power set of power 2
print("Cartesian power 2 with length {}: {}".format(len(p2),p2))
print("\n")
p3=list(prod(A,repeat=3)) # Power set of power 3
print("Cartesian power 3 with length {}: {}".format(len(p3),p3))
# ## HEADING 3: Factorials, permutation, combination
# In this section, we will discuss the concept of factorials and thereafter move to the topics of combinatorics such as permutation and combination.
# ### Factorial
# Factorials are normally defined for positive integers as following,
# $$n! = 1 \times 2 \times 3 \times \ ...\ \times (n-1) \times n$$
# Factorial of zero is defined as 1 and others follow naturally. For example,
# $$0!=1$$
# $$1!=1$$
# $$2!=1 \times 2 = 2$$
# $$3!=1 \times 2 \times 3 = 6$$
# #### Therefore, $n!$ can be expressed in terms of $(n-1)!$ easily,
# $$n! = n \times (n-1)!$$
# ### Let us write Python functions to compute factorial of a given number in a couple of different ways
def factorial_from_definition(n):
prod = 1
for i in range(1,n+1):
prod*=i
return prod
factorial_from_definition(5)
def factorial_recursive(n):
if n==0 or n==1:
return 1
else:
return (n*factorial_recursive(n-1))
factorial_recursive(5)
# #### The second function definition is short and elegant using recursion. But there is a cost to that. You can write another simple Python program to compare the computation time taken by these two approaches.
from time import time
t1 = time()
for i in range(1000000):
x = factorial_from_definition(20)
t2 = time()
print("Computing factorial of 20 one million times using the straight-forward function took {} seconds".format(t2-t1))
t1 = time()
for i in range(1000000):
x = factorial_recursive(20)
t2 = time()
print("Computing factorial of 20 one million times using the recursive function took {} seconds".format(t2-t1))
# ### Permutations
# In mathematics, the notion of permutation relates to the act of arranging all the members of a set into some sequence or order, or if the set is already ordered, rearranging (reordering) its elements, a process called __permuting__.
import itertools
A = {'a','b','c'}
permute_all = set(itertools.permutations(A))
print("Permutations of {}: ".format(A))
for i in permute_all:
print(i)
print ("\nNumber of permutations: ", len(permute_all))
# #### Finding all _k_-permutations of a set
A = {'a','b','c','d'}
k = 2
n = len(A)
permute_k = list(itertools.permutations(A, k))
print("{}-permutations of {}: ".format(k,A))
for i in permute_k:
print(i)
print ("\nSize = {}!/({}-{})! = {}".format(n,n,k, len(permute_k)))
# ### Combinations
#
# Combinatorics is an area of mathematics primarily concerned with counting, both as a means and an end in obtaining results, and certain properties of finite structures. It is closely related to many other areas of mathematics and has many applications ranging from logic to statistical physics, from evolutionary biology to computer science, etc.
#
# Combinatorics is well known for the breadth of the problems it tackles. Combinatorial problems arise in many areas of pure mathematics, notably in algebra, [probability theory](https://en.wikipedia.org/wiki/Probability_theory), [topology](https://en.wikipedia.org/wiki/Topology), and geometry, as well as in its many application areas.
#
# Many combinatorial questions have historically been considered in isolation, giving an _ad hoc_ solution to a problem arising in some mathematical context. In the later twentieth century, however, powerful and general theoretical methods were developed, making combinatorics into an independent branch of mathematics in its own right. One of the oldest and most accessible parts of combinatorics is [graph theory](https://en.wikipedia.org/wiki/Graph_theory), which by itself has numerous natural connections to other areas. Combinatorics is used frequently in computer science to obtain formulas and estimates in the [analysis of algorithms](https://en.wikipedia.org/wiki/Analysis_of_algorithms).
#
# We find the number of $k$-combinations of $A$, first by determining the set of combinations and then by simply calculating ${|A|}\choose{k}$.
A = {'a','b','c','d'}
k = 2
choose_k = list(itertools.combinations(A,k))
print("{}-combinations of {}: ".format(k,A))
for i in choose_k:
print(i)
print("\nNumber of combinations = {}!/({}!({}-{})!) = {}".format(n,k,n,k,len(choose_k)))
# ### Where in data science these concepts are used?
# Data science draws heavily from **probability and statistics** (as we will see in the next part of this book). One of the most intuitive application of combinatorics is in probability calculations. Let's think about the following problem,
#
# _"Suppose we have two dice which we throw together. <br>
# We want to know the probability of the event that the sum of the throws is divisible by 3."_
#
# Let us write a simple Python program to calculate this probability.
# #### First, compute all possible dice throw combinations. It is a combination of 2 samples drawn from a set of 6 elements.
D = {1,2,3,4,5,6}
all_events = list(itertools.permutations(D,2))
print("All possible dice throw permutations: ",all_events)
# #### How many sums of these pairs are divisible by 3?
div_by_3 = [x for x in all_events if (x[0]+x[1])%3==0]
print("Dice throw combinations where the sum is divisible by 3: ",div_by_3)
# #### So, the desired probability is simply the ratio of the number of events, where the sum is divisible by 3, to the total number of events
prob = len(div_by_3)/len(all_events)
print("Desired probability: ",prob)
# ## HEADING 4: Real and complex number
# In this section, we touch upon the topics of real and complex numbers. Although these may seem very basic concepts, they appear repetitively in all kind of discussion of functions and numerical analyses.
# ### Computing $\pi$ using _continued fraction_
# +
from fractions import Fraction
import itertools
def compute_pi(num_terms=4):
terms = list(itertools.islice(zip([6]*num_terms, [(2*i+3)**2 for i in range(num_terms)]), num_terms))
z = Fraction(1,1)
for a, b in reversed(terms):
z = a + b / z
return 3+(1/float(z))
# -
compute_pi(2)
compute_pi(10)
compute_pi(100)
# ### Functions for manipulating complex numbers
def complex_add(z1,z2):
z_real = z1[0]+z2[0]
z_imag = z1[1]+z2[1]
print("Sum of {}+{}j and {}+{}j is: {}+{}j".format(z1[0],z1[1],z2[0],z2[1],z_real,z_imag))
return ((z_real,z_imag))
# +
z1 = [4,3]
z2 = [-1,5]
complex_add(z1,z2)
# -
def complex_mult(z1,z2):
z_real = z1[0]*z2[0] - z1[1]*z2[1]
z_imag = z1[1]+z2[0] + z1[0]+z2[1]
print("Product of {}+{}j and {}+{}j is: {}+{}j".format(z1[0],z1[1],z2[0],z2[1],z_real,z_imag))
return ((z_real,z_imag))
# +
z1 = [4,3]
z2 = [-1,5]
complex_mult(z1,z2)
# -
# ### Python has a built-in `complex` type and a built-in `cmath` module to manipulate complex numbers easily
z = complex(3,4)
print(z)
print("Real part of {} is {}".format(z,z.real))
print("Imaginary part of {} is {}".format(z,z.imag))
print("Conjugate of {}: {}".format(z,z.conjugate()))
import cmath
cmath.polar(z)
cmath.phase(z)
cmath.exp(z)
cmath.sin(z)
# ## HEADING 5: Prime number
# ### Prime checking functions
def is_prime_naive(n):
for i in range(2,n-1):
if n%i==0:
return False
return True
is_prime_naive(25)
from math import sqrt
def is_prime_fast(n):
if n%2==0:
return False
for i in range(2,int(sqrt(n))+1):
if n%i==0:
return False
return True
is_prime_fast(23)
# ### Find all prime numbers between 1000000 and 1000500
# #### Using the naive (slow) function
primes=[]
t1=time()
for i in range(1000000,1000501):
if is_prime_naive(i):
primes.append(i)
t2=time()
print("List of primes between 1,000,000 and 1,000,500: ",primes)
print("\nTook {} seconds".format(t2-t1))
# #### Using the faster function
primes=[]
t1=time()
for i in range(1000000,1000501):
if is_prime_fast(i):
primes.append(i)
t2=time()
print("List of primes between 1,000,000 and 1,000,500: ",primes)
print("\nTook {} seconds".format(t2-t1))
# ### Function for prime factorization
def prime_factors(n):
i = 2
factors = []
while i * i <= n:
if n % i:
i += 1
else:
n //= i
factors.append(i)
if n > 1:
factors.append(n)
return factors
prime_factors(779)
# ### Show how the computational complexity of prime factorization rises with the size (order) of the number
# +
start = [1e3,1e4,1e5,1e6,1e7]
for i in start:
f = []
while len(f)<2:
i+=1
if is_prime_fast(i):
f.append(int(i))
prod = int(1)
for j in f:
prod*=j
t1=time()
factors = prime_factors(prod)
t2=time()
print("\nThe number to be factorized: ",prod)
print("Prime factors: ",factors)
print("Took {} microseconds".format((t2-t1)*1e6))
# -
# ## HEADING 6: Arithmetic, geometric, Fibonacci series
# ### Function to build an arithmatic series
def build_arithmatic_series(start,step,num_elements):
end = start + step*num_elements
return (list(range(start,end,step)))
build_arithmatic_series(2,5,10)
# ### Sum of an arithmatic series
s = sum(build_arithmatic_series(10,7,15))
print("Sum of the arithmatic series with starting element 10, step of 7, and 15 elements is: ",s)
# ### Function to build a geometric series
def build_geometric_series(start,mult,num_elements):
series = [start]
for i in range(1,num_elements):
term=start
for j in range(1,i+1):
term*=mult
series.append(term)
return series
build_geometric_series(2,1.5,4)
build_geometric_series(100,0.7,10)
# ### Sum of a geometric series
s = sum(build_geometric_series(10,0.8,10))
print("Sum of the geometric series with starting element 10, multiplier of 0.8, and 10 elements is: ",s)
s = sum(build_geometric_series(10,0.8,90))
print("Sum of the geometric series with starting element 10, multiplier of 0.8, and 90 elements is: ",s)
s = sum(build_geometric_series(10,0.8,100))
print("Sum of the geometric series with starting element 10, multiplier of 0.8, and 100 elements is: ",s)
# ### Fibonacci series
def fib_recursive(n):
if n==1 or n==2:
return 1
else:
return fib_recursive(n-1)+fib_recursive(n-2)
fib_recursive(2)
fib_recursive(6)
print("First 10 numbers in the Fibonacci sequence:")
for i in range(1,11):
print(fib_recursive(i), end=', ')
def fib_dynamic(n):
n1 = 1
n2 = 1
count = 2
nth=1
while count<n:
nth = n1+n2
n1=n2
n2=nth
count+=1
return (nth)
fib_dynamic(6)
print("First 10 numbers in the Fibonacci sequence:")
for i in range(1,11):
print(fib_dynamic(i), end=', ')
for i in range(21,35):
t1=time()
print("Number: ",fib_recursive(i))
t2=time()
print("Took {} milliseconds".format((t2-t1)*1e3))
for i in range(21,35):
t1=time()
print("Number: ",fib_dynamic(i))
t2=time()
print("Took {} milliseconds".format((t2-t1)*1e3))
| Packt-Maths-Data-Scientist/CHAPTER 1 - Set algebra, discrete math, and numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#get Iris test dataset using sklearn library
from sklearn import datasets
import numpy as np
iris = datasets.load_iris()
X = iris.data[:, [2, 3]] #take only petal lengh and petal width parameters from the Iris dataset
y = iris.target
#split test data into 30% testing and 70% training data
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
#feature scaling (standardize) using the StandardScaler
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
sc.fit(X_train)
X_train_std = sc.transform(X_train)
X_test_std = sc.transform(X_test)
X_combined = np.vstack((X_train, X_test))
y_combined = np.hstack((y_train, y_test))
from matplotlib.colors import ListedColormap
import matplotlib.pyplot as plt
def plot_decision_regions(X, y, classifier, test_idx=None, resolution=0.02):
#setup marker generator and color map
markers = ('s', 'x', 'o', '^', 'v')
colors = ('red', 'blue', 'lightgreen', 'gray', 'cyan')
cmap = ListedColormap(colors[:len(np.unique(y))])
#plot the decision surface
x1_min, x1_max = X[:, 0].min() - 1, X[:, 0].max() + 1
x2_min, x2_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx1, xx2 = np.meshgrid(np.arange(x1_min, x1_max, resolution), np.arange(x2_min, x2_max, resolution))
Z = classifier.predict(np.array([xx1.ravel(), xx2.ravel()]).T)
Z = Z.reshape(xx1.shape)
plt.contourf(xx1, xx2, Z, alpha=0.4, cmap=cmap)
plt.xlim(xx1.min(), xx1.max())
plt.ylim(xx2.min(), xx2.max())
#plot all samples
for idx, cl in enumerate(np.unique(y)):
plt.scatter(x=X[y == cl, 0], y=X[y == cl, 1], alpha=0.8, c=cmap(idx), marker=markers[idx], label=cl)
#highlight test samples
if test_idx:
X_test, y_test = X[test_idx, :], y[test_idx]
plt.scatter(X_test[:, 0], X_test[:, 1], facecolors='none', edgecolors='grey', alpha=1.0, linewidths=1, marker='o', s=55, label='test set')
# -
# random forest classifier
from sklearn.ensemble import RandomForestClassifier
forest = RandomForestClassifier(criterion='entropy', n_estimators=10, random_state=1, n_jobs=2)
forest.fit(X_train, y_train)
plot_decision_regions(X_combined, y_combined, classifier=forest, test_idx=range(105,150))
plt.xlabel('petal length [cm]')
plt.ylabel('petal width [cm]')
plt.legend(loc='upper left')
plt.show()
| ch03/07-random-forest-classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predictive modeling - Hyperparameter Tuning
# In this section we apply techniques for [hyperparameter tuning][1] on a real world data set, the _adult_ data set. The data set is available on the [UC Irvine Machine Learning Repository](https://archive.ics.uci.edu/ml/index.php) and can be assessed and downloaded [here](https://archive.ics.uci.edu/ml/datasets/Adult).
#
# For the purpose of this tutorial we already downloaded the data set. You may fin it in the `datasets` folder (`../datasets/adult.csv`).
#
# Please note that this tutorial bases on a talk given by [<NAME>](https://github.com/ogrisel) and [<NAME>](https://github.com/betatim) at [EuroScipy 2017](https://www.euroscipy.org/2017/). You can watch their tutorial on YouTube ([Part I](https://www.youtube.com/watch?v=Vs7tdobwj1k&index=3&list=PL55N1lsytpbekFTO5swVmbHPhw093wo0h) and [Part II](https://www.youtube.com/watch?v=0eYOhEF_aK0&list=PL55N1lsytpbekFTO5swVmbHPhw093wo0h&index=2)).
#
#
# [1]: https://en.wikipedia.org/wiki/Hyperparameter_(machine_learning)
# **Import libraries**
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# **Global setting**
pd.options.display.max_columns = 200
plt.rcParams["figure.figsize"] = [12,6]
# ## Load the data
filepath = "../datasets/adult_data.txt"
names = ("age, workclass, fnlwgt, education, education-num, "
"marital-status, occupation, relationship, race, sex, "
"capital-gain, capital-loss, hours-per-week, "
"native-country, income").split(', ')
data = pd.read_csv(filepath , names=names)
data = data.drop('fnlwgt', axis=1)
# We take a look at the first rows of the data set by calling the `head()` function.
data.head()
# > __The goal is to predict whether a person makes over 50K $ a year.__
# ## Training-Test Split
# Split the data set into `target` and `feature` data sets.
# +
target = data['income']
features_data = data.drop('income', axis=1)
features = pd.get_dummies(features_data)
print("Target variable: ", target.shape)
print("Features: ", features.shape)
# -
X = features.values.astype(np.float32)
y = (target.values == ' >50K').astype(np.int32)
X.shape
y
# +
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(
X, y, test_size=0.2, random_state=42)
print("Training set: ", X_train.shape)
print("Validation set: ", X_val.shape)
# -
# ## Learning Algorithm - Decision Trees
# [__Decision Trees__](https://en.wikipedia.org/wiki/Decision_tree_learning) are a non-parametric supervised learning method used for classification and regression. The goal is to create a model that predicts the value of a target variable by learning simple decision rules inferred from the data features.
#
#
# Some advantages of decision trees are:
#
# * Simple to understand and to interpret (white box model). Trees can be visualized.
# * Requires little data preparation.
# * The cost of using the tree (i.e., predicting data) is logarithmic in the number of data points used to train the tree.
# * Able to handle both numerical and categorical data. Other techniques are usually specialized in analyzing datasets that have only one type of variable. See algorithms for more information.
#
# The disadvantages of decision trees include:
#
# * Decision-tree learners can create over-complex trees that do not generalize the data well. This is called [overfitting](https://en.wikipedia.org/wiki/Overfitting).
# * Decision trees can be unstable because small variations in the data might result in a completely different tree being generated. This problem is mitigated by using decision trees within an ensemble.
#
#
#
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier(max_depth=8)
clf
# +
from sklearn.model_selection import cross_val_score
scores = cross_val_score(clf, X_train, y_train, cv=5, scoring='roc_auc')
print("ROC AUC Decision Tree: {:.4f} +/-{:.4f}".format(
np.mean(scores), np.std(scores)))
# -
# ## Tuning your estimator
#
# Hyperparameters are not directly learned by the classifier or regressor from the data. They need setting from the outside. An example of a hyper-parameter is `max_depth` for a decision tree classifier. In `scikit-learn` you can spot them as the parameters that are passed to the constructor of your estimator.
#
#
# The best value of a hyper-parameter depends on the kind of problem you are solving:
#
# * how many features and samples do you have?
# * mostly numerical or mostly categorical features?
# * is it a regression or classification task?
#
# Therefore you should optimize the hyper-parameters for each problem, otherwise the performance of your classifier will not be as good as it could be.
#
# ### Search over a grid of parameters
#
# This is the simplest strategy: you try every combination of values for each hyper-parameter.
# In scikit-learn __grid search__ is provided by [`GridSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html), which exhaustively generates candidates from a grid of parameter values specified with the `param_grid`.
# +
from sklearn.model_selection import GridSearchCV
param_grid = {"max_depth": [1, 2, 4, 8, 16, 32]}
grid_search = GridSearchCV(clf, param_grid=param_grid,
scoring='roc_auc', return_train_score=True)
# -
grid_search.fit(X_train, y_train)
type(grid_search)
# Once we have created a `sklearn.model_selection._search.GridSearchCV` object we can access its attributes using the `.`-notation. For instance, the results of the cross-validation are stored in the `cv_results_` attribute.
grid_search.cv_results_
# We print out the values of `max_depth` and the average train and test scores for each iteration.
for n, max_depth in enumerate(grid_search.cv_results_['param_max_depth']):
print("Max depth: {}, train score: {:.3f}, test score {:.3f}".format(max_depth,
grid_search.cv_results_['mean_train_score'][n],
grid_search.cv_results_['mean_test_score'][n],))
# For the purpose of model diagnostics we write a function, `plot_grid_scores`, which allows us to compare test and train performance at for each value of of a particular hyperparameter, such as `max_depth`.
def plot_grid_scores(param_name, cv_result):
# access the parameter
param_values = np.array(cv_result["param_{}".format(param_name)])
# plotting
fix, ax = plt.subplots()
ax.set_title("Scores for {}".format(param_name), size=18)
ax.grid()
ax.set_xlabel(param_name)
ax.set_ylabel("Score")
train_scores_mean = cv_result['mean_train_score']
test_scores_mean = cv_result['mean_test_score']
ax.scatter(param_values, train_scores_mean, s=80 ,marker='o', color="r",
label="Training scores")
ax.scatter(param_values, test_scores_mean, s=80, marker='o', color="g",
label="Cross-validation scores")
ax.legend(loc="best")
print("Best test score: {:.4f}".format(np.max(test_scores_mean)))
# Once implemented we can use the `plot_grid_scores` and apply it on the `grid_search.cv_results_` object.
plot_grid_scores("max_depth", grid_search.cv_results_)
# >**Challenge:** Extend the parameter grid to also search over different values for the `max_features` hyper-parameter. (Try: 3, 6, 12, 24, 48, and 96). Plot the results using the `plot_grid_scores` function from above.
# +
## your code here ...
# -
plot_grid_scores("max_features", grid_search.cv_results_)
# Another interesting information might be to lookt at the best three parameter combinations so far. We write a function called `report` to achieve tis task.
def report(results, n_top=3):
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_score'] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
results['mean_test_score'][candidate],
results['std_test_score'][candidate]))
print("Parameters: {0}\n".format(results['params'][candidate]))
report(grid_search.cv_results_)
# ### Random grid search
#
# An alternative to the exhaustive grid search is to sample parameter values at random. This has two main benefits over an exhaustive search:
# * A budget can be chosen independent of the number of parameters and possible values.
# * Adding parameters that do not influence the performance does not decrease efficiency.
#
# [`RandomizedSearchCV`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html#sklearn.model_selection.RandomizedSearchCV) implements a randomized search over parameters, where each setting is sampled from a distribution over possible parameter values. In contrast to `GridSearchCV`, not all parameter values are tried out, but rather a fixed number of parameter settings is sampled from the specified distributions. The number of parameter settings that are tried is given by `n_iter`.
#
# +
from scipy.stats import randint as sp_randint
from sklearn.model_selection import RandomizedSearchCV
param_grid = {"max_depth": sp_randint(1, 32),
"max_features": sp_randint(1, 96),
}
random_search = RandomizedSearchCV(clf, param_distributions=param_grid,
n_iter=36, scoring='roc_auc', return_train_score=True)
random_search.fit(X_train, y_train)
# -
plot_grid_scores("max_depth", random_search.cv_results_)
# For the same number of model evaluations you get a much better view of how the performance varies as a function of `max_depth`. This is a big advantage especially if one of the hyper-parameters does not influence the performance of the estimator. Though as you increase the number of dimensions making a projection into just one becomes more noisy.
param_grid = {"max_depth": sp_randint(1, 32),
"max_features": sp_randint(1, 96),
"min_samples_leaf": sp_randint(15, 40)
}
random_search = RandomizedSearchCV(clf, param_distributions=param_grid,
n_iter=36, scoring='roc_auc', return_train_score=True)
random_search.fit(X_train, y_train)
plot_grid_scores("max_depth", random_search.cv_results_)
plot_grid_scores("max_features", random_search.cv_results_)
plot_grid_scores("min_samples_leaf", random_search.cv_results_)
# You may assess the best performing parameter combination using the `best_params_` attribute.
random_search.best_params_
# ### Bayesian optimization
#
# Neither the exhaustive grid search nor random search adapt their search for the best hyper-parameter as they evaluate points. For the grid all points are chosen upfront, and for random search all of them are chosen at random.
#
# It makes sense to use the knowledge from the first few evaluations to decide what hyper-parameters to try next. This is what tools like [`scikit-optimize`](https://scikit-optimize.github.io/) try and do. The technique is known as Bayesian optimization or sequential model based optimization.
# .
#
# The basic algorithm goes like this:
# * evaluate a new set of hyper-parameters
# * fit a regression model to all sets of hyper-parameters
# * use the regression model to predict which set of hyper-parameters is the best
# * evaluate that set of hyper-parameters
# * repeat.
#
# `scikit-optimize` provides a drop-in replacement for `GridSearchCV` and `RandomSearchCV` that performs all this on the inside:
#
# _Note that if `scikit-optimize` is not yet installed on your machine type `conda install scikit-optimize` into your shell._
from skopt import BayesSearchCV
bayes_search = BayesSearchCV(
clf,
{"max_depth": (1, 32),
"max_features": (1, 96),
"min_samples_leaf": (15, 40)
},
n_iter=15,
scoring='roc_auc',
return_train_score=True
)
bayes_search.fit(X_train, y_train)
#
# Once the computation finished, we can access the results in the same fashion as we did before.
#
plot_grid_scores("max_depth", bayes_search.cv_results_)
bayes_search.best_params_
bayes_search.best_score_
np.mean(bayes_search.cv_results_["mean_test_score"])
# ## Using cross validation results for predictions
#
# Once we finished our hyperparameter search, we may actually use the best model for predictions. Note that so far we did not build a test set, hence for the purpose of demonstration we use the validation set as test set:
X_test = np.copy(X_val)
y_test = np.copy(y_val)
# We use accuracy as our model evaluation metric.
from sklearn.metrics import accuracy_score
# Now there is more than one way to make predictions for a hold out set (`X_test`). We may use the `best_estimator_` attribute to instantiate an estimator object, or use `predict` directly on the CV-object.
# variant 1
m = bayes_search.best_estimator_
y_pred_v1 = m.fit(X_train, y_train).predict(X_test)
print("Accuracy on the test set: ", accuracy_score(y_true=y_test, y_pred=y_pred_v1))
# variant 2
y_pred_v2 = bayes_search.predict(X_val)
print("Accuracy on the test set: ", accuracy_score(y_true=y_test, y_pred=y_pred_v2))
# The results should be the same.
| notebooks/04d - Predictive modeling - Hyperparameter Tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import display, HTML, IFrame, YouTubeVideo
from ipywidgets import interact,fixed
import pandas as pd
from numpy import cos,sin,pi,tan,log,exp,sqrt,array,linspace,arange
from mpl_toolkits import mplot3d
# from mpl_toolkits.mplot3d.art3d import Poly3DCollection
from ipywidgets import interact
plt.rcParams["figure.figsize"] = [7,7]
from numpy.linalg import norm
# %matplotlib inline
# Uncomment the one that corresponds to your Jupyter theme
plt.style.use('dark_background')
# plt.style.use('fivethirtyeight')
# plt.style.use('Solarize_Light2')
# -
# $\renewcommand{\vec}{\mathbf}$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Exercises
#
# 1. Wheat production $W$ in a given year depends on the average temperature $T$ and the annual rainfall $R$. Scientists estimate that the average temperature is rising at a rate of $0.15^\circ$C/year and rainfall is decreasing at a rate of $0.1$ cm/year. They also estimate that at current production levels, $\partial W/\partial T = -2$ and $\partial W/\partial R = 8$.
#
# 1. What is the significance of the signs of these partial derivatives?
#
# As temperature goes up, wheat production decreases. More rain, on the other hand, more wheat.
#
# 2. Estimate the current rate of change of wheat production, $dW/dt$.
#
# $$\frac{dW}{dt} = \frac{\partial W}{\partial T}\frac{dT}{dt} + \frac{\partial W}{\partial R}\frac{dR}{dt} = -2(0.15) + 8(-0.1) = -1.1 \text{ wheats} / \text{year}$$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# 2. Suppose
# \begin{align}
# z &= z(x,y) \\
# x &= x(u,v) \\
# y &= y(u,v) \\
# u &= u(s,t) \\
# v &= v(s,t) \\
# \end{align}
# are all differentiable. Find an expression for $\frac{\partial z}{\partial s}$.
# -
# $$ \frac{\partial z}{\partial s} = \frac{\partial z}{\partial x}\frac{\partial x}{\partial u}\frac{\partial u}{\partial s}
# + \frac{\partial z}{\partial x}\frac{\partial x}{\partial v}\frac{\partial v}{\partial s}
# + \frac{\partial z}{\partial y}\frac{\partial y}{\partial u}\frac{\partial u}{\partial s}
# + \frac{\partial z}{\partial y}\frac{\partial y}{\partial v}\frac{\partial v}{\partial s} $$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example
#
# If $g:\RR\to\RR$ is any smooth function, show that $f(x,y) = g(x^2+y^2)$ is radially symmetric. That is, $\frac{\partial f}{\partial \theta} =0$
# -
# $$\frac{\partial f }{\partial \theta} = \frac{\partial}{\partial x} (g(x^2 + y^2)) \frac{\partial x}{\partial \theta}
# + \frac{\partial}{\partial y} (g(x^2 + y^2)) \frac{\partial y}{\partial \theta} $$
#
# $$ = g'(x^2 + y^2)2x (-r \sin \theta) + g'(x^2 + y^2)2y (-r \cos \theta) $$
#
# $$ = g'(x^2 + y^2)( -2xy + 2yx) = 0 $$
# <p style="padding-bottom:40%;"> </p>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Example
#
# Find the slope of the tangent line to
#
# $$ x \sin(y) - \frac12 = \sqrt{2} - 2\cos(xy)$$ at the point $\left(\frac12,\frac\pi2\right)$.
# + hide_input=true jupyter={"source_hidden": true} slideshow={"slide_type": "fragment"}
x = y = np.linspace(-pi,pi,102)
x,y = np.meshgrid(x,y)
z = x*sin(y) + 2*cos(x*y) - sqrt(2) - 1/2
plt.figure(figsize=(7,7))
cp = plt.contour(x,y,z,levels=arange(-3,3.4,.5),alpha=.5,colors='y')
cp = plt.contour(x,y,z,levels=[0],colors='y')
# plt.clabel(cp,fmt="%d");
x = np.linspace(-2.5,3.5,102)
plt.plot(x,pi/2 + (x-1/2) * (sqrt(2) - pi),color='r');
plt.grid(True)
plt.scatter(1/2,pi/2)
plt.xlim(-pi,pi)
plt.ylim(-pi,pi);
# -
# $$F(x,y) = x \sin y + 2\cos(xy) = \frac12 + \sqrt 2 $$
# $$ \frac{dy}{dx} = \left.-\frac{F_x}{F_y} \right\rvert_{(1/2,\pi/2)} = \left.-\frac{\sin y -2\sin(xy)y}{x\cos y -2 \sin(xy)x}\right\rvert_{(1/2,\pi/2)} $$
# $$ = - \frac{1 - \frac{\pi}{\sqrt2}}{-\frac{1}{\sqrt2}} = \sqrt2 - \pi $$
# ### Example
# Differentiate the function $$f(t) = \int_0^t e^{-tx^2}dx.$$
# **Solution** This is a funny example as it is ostensibly a one-variable calculus problem. $x$ is just a dummy variable so the only variable to differentiate here is $t$, but you are not likely to find this example in a Calculus 1 text.
# + jupyter={"source_hidden": true}
@interact
def _(t = (0.,3.,0.05)):
x = np.linspace(0,3,200)
plt.plot(x,exp(-x**2),label = "$e^{-x^2}$")
plt.plot(x,exp(-t*x**2),label = "$e^{-tx^2}$")
y = np.array([0] + list(np.linspace(0,t,150)) + [t])
z = exp(-t*y**2)
z[0] = 0
z[-1] = 0
plt.fill(y,z)
plt.legend();
# -
# We cannot only apply the Fundamental Theorem of Calculus here directly as $t$ appears in both the limits and the integrand. So instead, we define
#
# $$F(a,b) = \int_0^a e^{-bx^2}dx$$
#
# to separate those roles and then realize $f(t) = F(t,t)$ so we apply the chain rule
#
# $$f'(t) = F_a(t,t) + F_b(t,t)$$ where of course here $\frac{da}{dt} = 1 = \frac{db}{dt}$. The first partial is computed via FTC and the second by differentiating under the integral sign. And thus,
#
# $$f'(t) = e^{-t^3} + \int_0^t (-x^2)e^{-tx^2}\,dx $$
#
#
# which is not beautiful but can be evaluated to arbitrary precision.
# +
from scipy.integrate import quad
def fprime(t):
val = quad(lambda x: (-x**2)*exp(-t*x**2),0,t)[0]
return exp(-t**3) + val
fprime(1)
# + jupyter={"source_hidden": true}
t = np.linspace(0,3,200)
plt.figure(figsize=(8,8))
plt.plot(t, [fprime(tt) for tt in t],label="$df/dt$")
plt.plot(t, [quad(lambda x: exp(-tt*x**2),0,tt)[0] for tt in t],label="$f$")
plt.legend();
plt.plot(t, 0*t);
# -
| exercises/L10-Exercises-Solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="19LYAr64HuOw"
#@ IMPORTING THE LIBRARIES:
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
# + id="XqRfocAmIHrV"
#@ IMPLEMENTATION OF LOGISTIC REGRESSION:
class LogisticRegression(): # Implementation of Logistic Regression.
def __init__(self, learning_rate, iterations): # Constructor Function.
self.learning_rate = learning_rate # Initializing the Learning rate.
self.iterations = iterations # Initializing the Iterations.
def fit(self, features, labels): # Function for Training.
self.features = features # Initializing Features.
self.labels = labels # Initializing Labels.
self.num_train = features.shape[0] # Number of Training examples.
self.num_features = features.shape[1] # Number of Features.
self.W, self.b = np.zeros(self.num_features), 0 # Initializing Weights.
for i in range(self.iterations):
self.update_weights()
return self
def update_weights(self): # Function for updating weights.
A = 1 / (1 + np.exp(-(self.features.dot(self.W) + self.b)))
pred = (A - self.labels.T)
pred = np.reshape(pred, self.num_train)
dW = np.dot(self.features.T, pred) / self.num_train
db = np.sum(pred) / self.num_train # Average of the Predictions.
self.W = self.W - self.learning_rate * dW # Updating Weights.
self.b = self.b - self.learning_rate * db # Updating Weights.
return self
def predict(self, X): # Function for Predictions.
Z = 1 / (1 + np.exp(-(X.dot(self.W) + self.b)))
Y = np.where(Z > 0.5, 1, 0)
return Y
# + colab={"base_uri": "https://localhost:8080/"} id="aEZlOSt4thQU" outputId="40a0db7f-067c-45c3-f753-252216bdcb79"
#@ IMPLEMENTATION:
def main():
data = pd.read_csv(PATH) # Reading the Dataset.
X = data.iloc[:, :-1].values
Y = data.iloc[:, -1:].values
X_train, X_test, y_train, y_test = train_test_split(X, Y, test_size=0.3,
random_state=11)
model = LogisticRegression(learning_rate=0.01, iterations=1000) # Initializing Logistic Regression.
model.fit(X_train, y_train) # Training the Model.
Y_pred = model.predict(X_test)
correctly_classified = 0
count = 0
for count in range(np.size(Y_pred)):
if y_test[count] == Y_pred[count]:
correctly_classified += 1
count += 1
print("Accuracy:", (correctly_classified / count) * 100)
if __name__ == "__main__":
main()
| LogisticRegression/LogisticRegression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MALARIA DETECTION
# ### Lets take a look at how we are gonna make our model
#
# #### Step 1: Loading and Splitting of the Dataset
#
# - The first step is to load the data and scaling the images to binary 0 and 1 from Parasitized and Uninfected.
# - Then we will resize the images to 50 x 50
# - After that suffling of the images before train-test-split and converting the images to a single numpy array
# - Splitting the data
# - Converting the type of X_train and X_valid to float32
# - Add then One Hot Encoding on y
#
#
# #### Step 2: Building the CNN model
# - The CNN model is one of the efficient nueral networks for images and performing classifications. We will use tf.keras to build the CNN model.
# - We will build a Sequential CNN model.
# - We will build a CNN Layer followed by MaxPooling layer which is later followed by BatchNormalisation to normalize the previous layer's output and implement the Dropout regularization. After that we will use Flatten to the outputs. Then the last layer that has function Softmax is the output layer.
# - Finally we have to compile the CNN model. We will use optimizer called Adam then will apply the loss function as categorical_crossentropy and an evaluation metric as accuracy.
# - Next step is to use the fit function, to train our convolutional neural network (CNN) with X_train and y_train. Lets set the total amounts of epochs as 25 epochs, which is essentially 25 cycles or iterations of the full dataset including a batch size of 120.
#
# #### Step 3 : Predictions and Testing of the Model
# - After this we will predict and do evaluation on the builded model.
# - The last step will be to test our model on the HOLDOUT DATASET and making predictions.
# ### Importing Libraries
# +
# importing the libraries for loading data and visualisation
import os
import cv2
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from PIL import Image
import seaborn as sns
# import for train-test-split
from sklearn.model_selection import train_test_split
# import for One Hot Encoding
from keras.utils import to_categorical
# importing libraries for Model
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, MaxPooling2D
from tensorflow.keras.layers import Dense, Flatten, Dropout, BatchNormalization
# importing libraries for evaluating the model
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
# -
# ## Loading Data and Train-Test-Split
# +
# loading the data of images and setting their labels
data = []
labels = []
Parasitized = os.listdir("input/cell_images/train/Parasitized/")
for a in Parasitized:
try:
image = cv2.imread("input/cell_images/train/Parasitized/" + a)
image_from_array = Image.fromarray(image, 'RGB')
size_image = image_from_array.resize((50, 50))
data.append(np.array(size_image))
labels.append(0)
except AttributeError:
print("Error0")
Uninfected = os.listdir("input/cell_images/train/Uninfected/")
for b in Uninfected:
try:
image = cv2.imread("input/cell_images/train/Uninfected/" + b)
image_from_array = Image.fromarray(image, 'RGB')
size_image = image_from_array.resize((50, 50))
data.append(np.array(size_image))
labels.append(1)
except AttributeError:
print("Error1")
# Creating single numpy array of all the images and labels
data = np.array(data)
labels = np.array(labels)
print('Cells : {} and labels : {}'.format(data.shape , labels.shape))
# lets shuffle the data and labels before splitting them into training and testing sets
n = np.arange(data.shape[0])
np.random.shuffle(n)
data = data[n]
labels = labels[n]
# +
### Splitting the dataset into the Training set and Test set
X_train, X_valid, y_train, y_valid = train_test_split(data, labels, test_size = 0.2, random_state = 0)
print('Train data shape {} ,Test data shape {} '.format(X_train.shape, X_valid.shape))
# -
X_train = X_train.astype('float32')
X_valid = X_valid.astype('float32')
# One Hot Encoding
y_train = to_categorical(y_train)
y_valid = to_categorical(y_valid)
y_train
# ## Building Model
# +
# Defining Model
classifier = Sequential()
# CNN layers
classifier.add(Conv2D(32, kernel_size=(3, 3), input_shape = (50, 50, 3), activation = 'relu'))
classifier.add(MaxPooling2D(pool_size = (2, 2)))
classifier.add(BatchNormalization(axis = -1))
classifier.add(Dropout(0.5)) # Dropout prevents overfitting
classifier.add(Conv2D(32, kernel_size=(3, 3), input_shape = (50, 50, 3), activation = 'relu'))
classifier.add(MaxPooling2D(pool_size = (2, 2)))
classifier.add(BatchNormalization(axis = -1))
classifier.add(Dropout(0.5))
classifier.add(Flatten())
classifier.add(Dense(units=128, activation='relu'))
classifier.add(BatchNormalization(axis = -1))
classifier.add(Dropout(0.5))
classifier.add(Dense(units=2, activation='softmax'))
# -
classifier.compile(optimizer = 'adam', loss = 'categorical_crossentropy', metrics = ['accuracy'])
history = classifier.fit(X_train, y_train, batch_size=120, epochs=15, verbose=1, validation_data=(X_valid, y_valid))
print("Test_Accuracy: {:.2f}%".format(classifier.evaluate(X_valid, y_valid)[1]*100))
# - Summary of the Model
classifier.summary()
loss_train = history.history['loss']
loss_val = history.history['val_loss']
epochs = range(1,16)
plt.plot(epochs, loss_train, 'g', label='Training loss')
plt.plot(epochs, loss_val, 'b', label='validation loss')
plt.title('Training and Validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
loss_train = history.history['accuracy']
loss_val = history.history['val_accuracy']
epochs = range(1,16)
plt.plot(epochs, loss_train, 'g', label='Training accuracy')
plt.plot(epochs, loss_val, 'b', label='validation accuracy')
plt.title('Training and Validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.show()
# ## Prediction, Evaluation and Testing of the Model
# - Lets do our first prediction using predict and predict X_valid and store in y_pred variable
y_pred = classifier.predict(X_valid)
# Convert back to categorical values
y_pred = np.argmax(y_pred, axis=1)
y_valid = np.argmax(y_valid, axis=1)
print('Accuracy Score: ', accuracy_score(y_valid, y_pred))
# - Evaluation of the CNN model by Plotting Confusion Matrix
# Plotting the Confusion Matrix
conf = confusion_matrix(y_valid, y_pred)
sns.heatmap(conf, annot=True)
classifier.save("malaria-model.h5")
| Notebooks/malariadetection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# Lambda School Data Science, Unit 2: Predictive Modeling
#
# # Kaggle Challenge, Module 1
#
# ## Assignment
# - [ ] Do train/validate/test split with the Tanzania Waterpumps data.
# - [ ] Define a function to wrangle train, validate, and test sets in the same way. Clean outliers and engineer features. (For example, [what other columns have zeros and shouldn't?](https://github.com/Quartz/bad-data-guide#zeros-replace-missing-values) What other columns are duplicates, or nearly duplicates? Can you extract the year from date_recorded? Can you engineer new features, such as the number of years from waterpump construction to waterpump inspection?)
# - [ ] Select features. Use a scikit-learn pipeline to encode categoricals, impute missing values, and fit a decision tree classifier.
# - [ ] Get your validation accuracy score.
# - [ ] Get and plot your feature importances.
# - [ ] Submit your predictions to our Kaggle competition. (Go to our Kaggle InClass competition webpage. Use the blue **Submit Predictions** button to upload your CSV file. Or you can use the Kaggle API to submit your predictions.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# ## Stretch Goals
#
# ### Reading
#
# - A Visual Introduction to Machine Learning
# - [Part 1: A Decision Tree](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/)
# - [Part 2: Bias and Variance](http://www.r2d3.us/visual-intro-to-machine-learning-part-2/)
# - [Decision Trees: Advantages & Disadvantages](https://christophm.github.io/interpretable-ml-book/tree.html#advantages-2)
# - [How a Russian mathematician constructed a decision tree — by hand — to solve a medical problem](http://fastml.com/how-a-russian-mathematician-constructed-a-decision-tree-by-hand-to-solve-a-medical-problem/)
# - [How decision trees work](https://brohrer.github.io/how_decision_trees_work.html)
# - [Let’s Write a Decision Tree Classifier from Scratch](https://www.youtube.com/watch?v=LDRbO9a6XPU) — _Don’t worry about understanding the code, just get introduced to the concepts. This 10 minute video has excellent diagrams and explanations._
# - [Random Forests for Complete Beginners: The definitive guide to Random Forests and Decision Trees](https://victorzhou.com/blog/intro-to-random-forests/)
#
#
# ### Doing
# - [ ] Add your own stretch goal(s) !
# - [ ] Try other [scikit-learn imputers](https://scikit-learn.org/stable/modules/impute.html).
# - [ ] Try other [scikit-learn scalers](https://scikit-learn.org/stable/modules/preprocessing.html).
# - [ ] Make exploratory visualizations and share on Slack.
#
#
# #### Exploratory visualizations
#
# Visualize the relationships between feature(s) and target. I recommend you do this with your training set, after splitting your data.
#
# For this problem, you may want to create a new column to represent the target as a number, 0 or 1. For example:
#
# ```python
# train['functional'] = (train['status_group']=='functional').astype(int)
# ```
#
#
#
# You can try [Seaborn "Categorical estimate" plots](https://seaborn.pydata.org/tutorial/categorical.html) for features with reasonably few unique values. (With too many unique values, the plot is unreadable.)
#
# - Categorical features. (If there are too many unique values, you can replace less frequent values with "OTHER.")
# - Numeric features. (If there are too many unique values, you can [bin with pandas cut / qcut functions](https://pandas.pydata.org/pandas-docs/stable/getting_started/basics.html?highlight=qcut#discretization-and-quantiling).)
#
# You can try [Seaborn linear model plots](https://seaborn.pydata.org/tutorial/regression.html) with numeric features. For this problem, you may want to use the parameter `logistic=True`
#
# You do _not_ need to use Seaborn, but it's nice because it includes confidence intervals to visualize uncertainty.
#
# #### High-cardinality categoricals
#
# This code from a previous assignment demonstrates how to replace less frequent values with 'OTHER'
#
# ```python
# # Reduce cardinality for NEIGHBORHOOD feature ...
#
# # Get a list of the top 10 neighborhoods
# top10 = train['NEIGHBORHOOD'].value_counts()[:10].index
#
# # At locations where the neighborhood is NOT in the top 10,
# # replace the neighborhood with 'OTHER'
# train.loc[~train['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
# test.loc[~test['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
# ```
#
#
# + colab={} colab_type="code" id="o9eSnDYhUGD7"
# If you're in Colab...
import os, sys
in_colab = 'google.colab' in sys.modules
if in_colab:
# Install required python packages:
# category_encoders, version >= 2.0
# pandas-profiling, version >= 2.0
# plotly, version >= 4.0
# !pip install --upgrade category_encoders pandas-profiling plotly
# Pull files from Github repo
os.chdir('/content')
# !git init .
# !git remote add origin https://github.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge.git
# !git pull origin master
# Change into directory for module
os.chdir('module1')
# + colab={} colab_type="code" id="QJBD4ruICm1m"
import pandas as pd
from sklearn.model_selection import train_test_split
train = pd.merge(pd.read_csv('../data/tanzania/train_features.csv'),
pd.read_csv('../data/tanzania/train_labels.csv'))
test = pd.read_csv('../data/tanzania/test_features.csv')
sample_submission = pd.read_csv('../data/tanzania/sample_submission.csv')
train.shape, test.shape
# + colab={} colab_type="code" id="2Amxyx3xphbb"
# Import block
pd.set_option('display.max_columns', None)
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
plt.style.use('dark_background')
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegressionCV
from sklearn.tree import DecisionTreeClassifier
from sklearn.pipeline import make_pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import accuracy_score
import category_encoders as ce
# +
# train/validation split
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify=train['status_group'], random_state=42)
train.shape, val.shape, test.shape
# -
train.columns
train.describe(include = 'number')
# construction year of zero doesn't make sense
# Nor does latitude/longitude of zero.
# population of zero is potentially meaningful, it's possible a region with a pump has no one in it
train.describe(exclude='number')
train[['waterpoint_type','waterpoint_type_group','source_class']].head(10)
def wrangler(X):
# Make a copy to avoid warning, prevent making changes from view.
X = X.copy()
# Replace near-zero latitudes with zero
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# Replace near-zero longitudes with zero
X['longitude'] = X['longitude'].replace(-2e-08, 0)
# Swap zeros with nulls
cols_with_zeros = ['longitude', 'latitude','construction_year']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
# All the following are duplicate drops
# quantity & quantity_group are duplicates, so drop one
X = X.drop(columns='quantity_group')
# extraction_type is a more specific version of extraction_type_group/class
X = X.drop(columns=['extraction_type_group','extraction_type_class'])
# Same for management and management_group
X = X.drop(columns = 'management_group')
# And payment / payment_type
X = X.drop(columns = 'payment')
# And water_quality/quality_group
X = X.drop(columns = 'quality_group')
# And water_quality/quality_group
X = X.drop(columns = 'waterpoint_type_group')
# Feature creation
# create a 'distance from Dodoma' feature
X['dodomadistance'] = (((X['latitude']-(6.1630))**2)+((X['longitude']-(35.7516))**2))**0.5
# create a pump age feature
X['pump_age'] = 2013 - X['construction_year']
# create 'year_recorded' - year from date_recorded
X['year_recorded'] = pd.to_datetime(X.date_recorded).dt.year
# create 'structspect_interval' - number of years between construction and date recorded
X['structspect_interval'] = X['year_recorded'] - X['construction_year']
# return the wrangled dataframe
return X
# Wrangle all datasets
train = wrangler(train)
val = wrangler(val)
test = wrangler(test)
# +
# Define target
target = 'status_group'
# Frame without target and id
train_features = train.drop(columns=[target, 'id'])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
# Get a series with the number of unique classes for each categorical variable
catswithcounts = train_features.select_dtypes(exclude='number').nunique()
# set upper bound on class count
categorical_features = catswithcounts[catswithcounts <= 50].index.tolist()
# Combine the lists
features = numeric_features + categorical_features
# Arrange train/val/test data
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
# map Ys to integers for the encoder
mapdict = {
'functional': 1,
'non functional': -1,
'functional needs repair': 0
}
y_train_mapped = y_train.map(mapdict)
y_val_mapped = y_val.map(mapdict)
# +
# Make decision tree pipeline
thelorax = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
DecisionTreeClassifier(max_depth=38, min_samples_leaf=1, random_state=42)
)
# -
# fit it and score it
thelorax.fit(X_train,y_train)
print ('train accuracy: ', thelorax.score(X_train, y_train))
print ('validation accuracy: ', thelorax.score(X_val, y_val))
# +
# Grab my gradient descent code, use it for tree depth and leaf samples
# tree length first
treemin = 1
treemax = 50
bounddict = {
'lowbound': treemin,
'midbound': int(round((treemin+treemax)/2)),
'upbound': treemax
}
for i in range(1,10):
boundaccuracy={}
# Set the central value according to the bounds
bounddict['midbound'] = int(round((bounddict['lowbound']+bounddict['upbound'])/2))
for key,value in bounddict.items():
# Make decision tree pipeline with the bound we're testing
thelorax = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
DecisionTreeClassifier(max_depth=value, min_samples_leaf=1, random_state=42)
)
# Fit the model, score it
thelorax.fit(X_train,y_train)
print ('train accuracy: ', thelorax.score(X_train, y_train))
print ('validation accuracy: ', thelorax.score(X_val, y_val))
# Get the error for the value
boundaccuracy[key] = thelorax.score(X_val, y_val)
#Eliminate whichever extremal bound is worse
if boundaccuracy['lowbound'] < boundaccuracy['upbound']:
bounddict['lowbound'] = bounddict['midbound']
else:
bounddict['upbound'] = bounddict['midbound']
# -
bounddict
# +
# Checking whether 26 or 27 produced the better value
thelorax = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
DecisionTreeClassifier(max_depth=28, min_samples_leaf=1, random_state=42)
)
# fit it and score it
thelorax.fit(X_train,y_train)
print ('train accuracy: ', thelorax.score(X_train, y_train))
print ('validation accuracy: ', thelorax.score(X_val, y_val))
# +
# Grab my gradient descent code, use it for tree depth and leaf samples
# now leaves
leafmin = 1
leafmax = 100
bounddict = {
'lowbound': leafmin,
'midbound': int(round((leafmin+leafmax)/2)),
'upbound': leafmax
}
for i in range(1,10):
boundaccuracy={}
# Set the central value according to the bounds
bounddict['midbound'] = int(round((bounddict['lowbound']+bounddict['upbound'])/2))
for key,value in bounddict.items():
# Make decision tree pipeline with the bound we're testing
thelorax = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
DecisionTreeClassifier(max_depth=28, min_samples_leaf=value, random_state=42)
)
# Fit the model, score it
thelorax.fit(X_train,y_train)
print ('train accuracy: ', thelorax.score(X_train, y_train))
print ('validation accuracy: ', thelorax.score(X_val, y_val))
# Get the error for the value
boundaccuracy[key] = thelorax.score(X_val, y_val)
#Eliminate whichever extremal bound is worse
if boundaccuracy['lowbound'] < boundaccuracy['upbound']:
bounddict['lowbound'] = bounddict['midbound']
else:
bounddict['upbound'] = bounddict['midbound']
# -
bounddict
# +
# Checking whether 17 or 18 produced the better value
thelorax = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
DecisionTreeClassifier(max_depth=28, min_samples_leaf=17, random_state=42)
)
# fit it and score it
thelorax.fit(X_train,y_train)
print ('train accuracy: ', thelorax.score(X_train, y_train))
print ('validation accuracy: ', thelorax.score(X_val, y_val))
# +
# feature importances
model = thelorax.named_steps['decisiontreeclassifier']
encoder = thelorax.named_steps['onehotencoder']
encoded_columns = encoder.transform(X_train).columns
importances = pd.Series(model.feature_importances_, encoded_columns)
importances.sort_values().plot.barh(color='grey', figsize = [8,20])
# +
# Predicting test data
y_test_pred = thelorax.predict(X_test)
# -
# Formatting submission
submission = sample_submission.copy()
submission['status_group'] = y_test_pred
submission.to_csv('submission-03.csv', index = False)
| module1/assignment_kaggle_challenge_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Cleaning Notebook - Example
# This is the more detailed notebook containing the data cleaning for several features. This contains only the "category" and "amount" features. I will leave the cleaning of the rest of the features as an exercise for you.
#
# ## Importing the data
# +
import json
with open("customer_data_example.json") as f_in:
data = json.load(f_in)
# taking a look at the JSON data
print(json.dumps(data, indent=2))
# +
import pandas as pd
# reading the JSON data in to a dataframe
df = pd.read_json("customer_data_example.json")
df
# -
# taking a look at the types for each feature
df.dtypes
# +
# parsing the XML data and putting it in to a dataframe
import xml.etree.ElementTree as ET
tree = ET.parse('location_data.xml')
root = tree.getroot()
# root tag
print(root.tag)
# child tag
print(root[0].tag)
# number of children elements
num_children = len(root.getchildren())
print(num_children)
# number of subchildren elements
num_subchildren = len(root[0].getchildren())
print(num_subchildren)
# pulling out all of the subchildren tags
tags = []
for subchild in root[0]:
tags.append(subchild.tag)
print(tags)
# creating an empty dictionary to store the data
d = {}
for tag in tags:
d[tag] = []
print(d)
# pulling out all of the data
for i in range(0, num_children):
for j in range(0, num_subchildren):
value = root[i][j].text
d[tags[j]].append(value)
#print(d)
# converting to a dataframe
df = pd.DataFrame(data=d)
print(df)
# -
# ## Cleaning the "category" feature
# summing the missing values
print(sum(df["category"].isna()))
# It doesn't look like there's any standard missing value types. Let's take a look at the unique values.
# unique values
print(df["category"].unique())
# There's a carrot "^" character in the "elec^tronics" feature. I'll need to remove that.
# +
# removing incorrect "^" character from the strings
bad_characters = ["^"]
cnt = 0
for row in df["category"]:
for character in bad_characters:
if character in row:
df.loc[cnt, "category"] = row.replace(character, "")
cnt+=1
print(df["category"].unique())
# -
# It looks like the features aren't all consistent. The "household" and "house" features are the same. I'll change "house" to "household" to make sure that my feature labels are consistent.
# +
consistent_format = ["house"]
cnt = 0
for row in df["category"]:
if row in consistent_format:
df.loc[cnt, "category"] = "household"
cnt+=1
print(df["category"].unique())
# -
# The data in the "category" column has now been cleaned.
# ## Cleaning the "amount" feature
# looking for missing values
print(sum(df["amount"].isna()))
# It doesn't look like there's any standard missing values.
#
# Rather than look at unique values, I'll take a look at the type first.
#
# I would expect the type to be a float. If the type is an object (string), there's probably some additional dirty data that needs to be cleaned, such as dollar signs "$" or unexpected characters.
# looking at unique values
df["amount"].dtype
# The type is "float", so it looks like we should be all set.
#
# # Exporting the clean data
#
# I'll finish by exporting the clean data to a CSV file called "customer_data_cleaned.csv".
df.to_csv("customer_data_cleaned.csv")
| Data Cleaning Notebook - Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Playing with Pandas and Numpy
#
#
# ---
# + [markdown] tags=["solution"]
# *Key questions:*
#
# - "How can I import data in Python ?"
# - "What is Pandas ?"
# - "Why should I use Pandas to work with data ?"
# -
import urllib.request
# You can also get this URL value by right-clicking the `survey.csv` link above and selecting "Copy Link Address"
url = 'https://github.com/aaneloy/DATA2010-Fall2021-Lab/blob/main/Dataset/survey.csv'
urllib.request.urlretrieve(url, 'surveys.csv')
# !pip install pandas matplotlib
import pandas as pd
import numpy as np
surveys_df = pd.read_csv("survey.csv")
# Notice when you assign the imported DataFrame to a variable, Python does not
# produce any output on the screen. We can view the value of the `surveys_df`
# object by typing its name into the cell.
surveys_df
# You can also select just a few rows, so it is easier to fit on one window, you can see that pandas has neatly formatted the data to fit our screen.
#
# Here, we will be using a function called **head**.
#
# The `head()` function displays the first several lines of a file. It is discussed below.
#
surveys_df.head(10)
# ## Exploring Our Species Survey Data
#
# Again, we can use the `type` function to see what kind of thing `surveys_df` is:
#
#
type(surveys_df)
#
# As expected, it's a DataFrame (or, to use the full name that Python uses to refer
# to it internally, a `pandas.core.frame.DataFrame`).
#
# What kind of things does `surveys_df` contain? DataFrames have an attribute
# called `dtypes` that answers this:
#
#
surveys_df.dtypes
# + [markdown] tags=["challenge"]
# ## Challenge - DataFrames
#
# Using our DataFrame `surveys_df`, try out the attributes & methods below to see
# what they return.
#
# 1. `surveys_df.columns`
# 2. `surveys_df.shape` Take note of the output of `shape` - what format does it
# return the shape of the DataFrame in? HINT: [More on tuples, here](https://docs.python.org/3/tutorial/datastructures.html#tuples-and-sequences).
# 3. `surveys_df.head()` Also, what does `surveys_df.head(15)` do?
# 4. `surveys_df.tail()`
#
#
#
# + [markdown] tags=["solution"]
# ## Solution - DataFrames
#
# ... try it yourself !
# -
# # Calculating Statistics From Data
#
# We've read our data into Python. Next, let's perform some quick summary
# statistics to learn more about the data that we're working with. We might want
# to know how many animals were collected in each plot, or how many of each
# species were caught. We can perform summary stats quickly using groups. But
# first we need to figure out what we want to group by.
#
# Let's begin by exploring our data:
#
#
# Look at the column names
surveys_df.columns
surveys_df['species_id']
# Let's get a list of all the species. The `pd.unique` function tells us all of
# the unique values in the `species_id` column.
pd.unique(surveys_df['species_id'])
len((surveys_df['site_id']))
surveys_df['site_id'].unique()
# # Interpreting missing data
#
# 1. Check if there is any missing data
# 2. Replace the missing data by `zero`
# 3. Copy the dataframe so that we can use later.
# +
# Check for missing values
surveys_df.isna().sum() # or s.isnull().sum() for older pandas versions
# -
surveys_df.isna()
# +
#Copy the dataframe and replace the Null values by zero
surveys_df_2 = surveys_df.copy()
# -
surveys_df_2['weight'].fillna(0, inplace=True)
surveys_df_2['hindfoot_length'].fillna(0, inplace=True)
surveys_df_2
surveys_df_2.isna().sum()
surveys_df_2.dropna()
surveys_df_2
# We can also simply drop the 'NaN' or 'NULL' value rows
surveys_df_3 = surveys_df.dropna()
surveys_df_3
# +
#Check if any null values
surveys_df_3.isna().sum()
# + [markdown] tags=["challenge"]
# ## Challenge - Statistics
#
# 1. Create a list of unique site ID's found in the surveys data. Call it
# `site_names`. How many unique sites are there in the data? How many unique
# species are in the data?
#
# 2. What is the difference between `len(site_names)` and `surveys_df['site_id'].nunique()`?
# + [markdown] tags=["solution"]
# ## Solution - Statistics
# + tags=["solution"]
site_names = pd.unique(surveys_df['site_id'])
print(len(site_names), surveys_df['site_id'].nunique())
# -
# # Groups in Pandas
#
# We often want to calculate summary statistics grouped by subsets or attributes
# within fields of our data. For example, we might want to calculate the average
# weight of all individuals per site.
#
# We can calculate basic statistics for all records in a single column using the
# syntax below:
surveys_df['weight'].describe()
#
# We can also extract one specific metric if we wish:
#
#
surveys_df['weight'].min()
surveys_df['weight'].max()
surveys_df['weight'].mean()
surveys_df['weight'].std()
# only the last command shows output below - you can try the others above in new cells
surveys_df['weight'].count()
#
# But if we want to summarize by one or more variables, for example sex, we can
# use **Pandas' `.groupby` method**. Once we've created a groupby DataFrame, we
# can quickly calculate summary statistics by a group of our choice.
#
#
# Group data by sex
grouped_data = surveys_df.groupby('sex')
grouped_data
#
# The **pandas function `describe`** will return descriptive stats including: mean,
# median, max, min, std and count for a particular column in the data. **Note** Pandas'
# `describe` function will only return summary values for columns containing
# numeric data.
#
#
# +
# Summary statistics for all numeric columns by sex
grouped_data.describe()
# Provide the mean for each numeric column by sex
# As above, only the last command shows output below - you can try the others above in new cells
grouped_data.mean()
# -
#
# The `groupby` command is powerful in that it allows us to quickly generate
# summary stats.
#
#
# + [markdown] tags=["challenge"]
# ## Challenge - Summary Data
#
# 1. How many recorded individuals are female `F` and how many male `M`
# - A) 17348 and 15690
# - B) 14894 and 16476
# - C) 15303 and 16879
# - D) 15690 and 17348
#
#
# 2. What happens when you group by two columns using the following syntax and
# then grab mean values:
# - `grouped_data2 = surveys_df.groupby(['site_id','sex'])`
# - `grouped_data2.mean()`
#
#
# 3. Summarize weight values for each site in your data. HINT: you can use the
# following syntax to only create summary statistics for one column in your data
# `by_site['weight'].describe()`
#
#
# + [markdown] tags=["solution"]
# ## Solution- Summary Data
# + tags=["solution"]
## Solution Challenge 1
grouped_data.count()
# + [markdown] tags=["solution"]
# ### Solution - Challenge 2
#
# The mean value for each combination of site and sex is calculated. Remark that the
# mean does not make sense for each variable, so you can specify this column-wise:
# e.g. I want to know the last survey year, median foot-length and mean weight for each site/sex combination:
# + tags=["solution"]
# Solution- Challenge 3
surveys_df.groupby(['site_id'])['weight'].describe()
# + [markdown] tags=["solution"]
# ## Did you get #3 right?
# **A Snippet of the Output from part 3 of the challenge looks like:**
#
# ```
# site_id
# 1 count 1903.000000
# mean 51.822911
# std 38.176670
# min 4.000000
# 25% 30.000000
# 50% 44.000000
# 75% 53.000000
# max 231.000000
# ...
# ```
#
#
# -
# ## Quickly Creating Summary Counts in Pandas
#
# Let's next count the number of samples for each species. We can do this in a few
# ways, but we'll use `groupby` combined with **a `count()` method**.
#
#
#
# Count the number of samples by species
species_counts = surveys_df.groupby('species_id')['record_id'].count()
print(species_counts)
#
# Or, we can also count just the rows that have the species "DO":
#
#
surveys_df.groupby('species_id')['record_id'].count()['DO']
# ## Basic Math Functions
#
# If we wanted to, we could perform math on an entire column of our data. For
# example let's multiply all weight values by 2. A more practical use of this might
# be to normalize the data according to a mean, area, or some other value
# calculated from our data.
#
#
# Multiply all weight values by 2 but does not change the original weight data, rather than create new column with "weighted value" variable
surveys_df['weighted_in_Kg'] = surveys_df['weight']*2
surveys_df
# ## Quick & Easy Plotting Data Using Pandas
#
# We can plot our summary stats using Pandas, too.
#
#
# +
import matplotlib.pyplot as plt
## To make sure figures appear inside Jupyter Notebook
# %matplotlib inline
# Create a quick bar chart
plt.figure(figsize=(12,6))
species_counts.plot(kind='bar')
# -
# #### Animals per site plot
#
# We can also look at how many animals were captured in each site.
total_count = surveys_df.groupby('site_id')['record_id'].nunique()
# Let's plot that too
plt.figure(figsize=(12,6))
total_count.plot(kind='bar')
# + [markdown] tags=["challenge"]
# ## _Extra Plotting Challenge_
#
# 1. Create a plot of average weight across all species per plot.
#
# 2. Create a plot of total males versus total females for the entire dataset.
#
# 3. Create a stacked bar plot, with weight on the Y axis, and the stacked variable being sex. The plot should show total weight by sex for each plot. Some tips are below to help you solve this challenge:
# [For more on Pandas plots, visit this link.](http://pandas.pydata.org/pandas-docs/stable/visualization.html#basic-plotting-plot)
#
#
#
#
# -
surveys_df.groupby('site_id').mean()["weight"].plot(kind='bar')
# + [markdown] tags=["solution"]
# ### _Solution to Extra Plotting Challenge 1_
# -
# 
# + [markdown] tags=["solution"]
# ### _Solution to Extra Plotting Challenge 2_
# + [markdown] tags=["solution"]
# # Solution Plotting Challenge 2
# ## Create plot of total males versus total females for the entire dataset.
#
# The output should look like this <img src="https://github.com/aaneloy/DATA2010-Fall2021-Lab/blob/main/plot2.png" width=400 height=400 />
# -
surveys_df.groupby('sex').count()["record_id"].plot(kind='bar')
# + [markdown] tags=["solution"]
# ### _Solution to Extra Plotting Challenge 3_
#
# First we group data by site and by sex, and then calculate a total for each site.
#
# 
# + [markdown] tags=["solution"]
#
# This calculates the sums of weights for each sex within each plot as a table
#
# ```
# site sex
# site_id sex
# 1 F 38253
# M 59979
# 2 F 50144
# M 57250
# 3 F 27251
# M 28253
# 4 F 39796
# M 49377
# <other sites removed for brevity>
# ```
#
# Below we'll use `.unstack()` on our grouped data to figure out the total weight that each sex contributed to each plot.
#
#
# + [markdown] tags=["solution"]
# Now, create a stacked bar plot with that data where the weights for each sex are stacked by plot.
#
# Rather than display it as a table, we can plot the above data by stacking the values of each sex as follows:
| Pandas_Numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
from torch import Tensor
import numpy as np
import matplotlib.pyplot as plt
# #### Modules
class Module(object):
def forward(self, *input):
raise NotImplementedError
def backward(self, *gradwrtoutput):
raise NotImplementedError
def param(self):
return []
def update(self, lr=None):
pass
class Linear(Module):
"""Implements the fully connected layer module
It requires the number of inputs and outputs.
Weights are initialized assuming that a ReLU module
will be used afterwards. If a Tanh module will be used
instead, it is recommended to set
std_w = 1 / np.sqrt(n_input)
It is possible to set a default learning rate that will be used
during backpropagation if no other learning rate is stated.
"""
def __init__(self, n_input, n_output, lr=1e-5,
std_w=None, bias=True, std_b=0):
if std_w is None:
# "Xavier" initialization
std_w = 1 / np.sqrt(.5 * n_input)
# Set parameters
self.lr = lr
self.w = Tensor(n_output, n_input).normal_(0, std_w)
self.dw = Tensor(self.w.size()).zero_()
self.cum_dw = Tensor(self.w.size()).zero_()
self.bias = bias
if bias:
if not std_b:
self.b = Tensor(n_output, 1).fill_(0)
else:
self.b = Tensor(n_output, 1).normal_(0, std_b)
self.db = Tensor(self.b.size()).zero_()
self.cum_db = Tensor(self.b.size()).zero_()
def forward(self, x):
"""Carries out the forward pass for backpropagation."""
if len(x.shape) < 2:
# reshape if required
x = x.view(-1,1)
self.x = x
self.s = self.w @ x
if self.bias:
self.s += self.b
return self.s
def backward(self, grad):
"""Carries out the backward pass for backpropagation.
It does not update the parameters.
"""
if len(grad.shape) < 2:
# reshape if required
grad = grad.view(1,-1)
out = grad @ self.w
self.dw = (self.x @ grad).t()
self.cum_dw.add_(self.dw)
if self.bias:
self.db = grad.t()
self.cum_db.add_(self.db)
return out
def param(self):
"""Returns the list of parameters and gradients."""
out = [(self.w, self.dw)]
if self.bias:
out.append((self.b, self.db))
return out
def update(self, lr=None):
"""Updates the parameters with the accumulated gradients.
It must be called explicitly. If no lr is stated, the
default lr of the module is used.
"""
if lr is None:
lr = self.lr
self.w.add_(-lr * self.cum_dw)
self.cum_dw = Tensor(self.w.size()).zero_()
if self.bias:
self.b.add_(-lr * self.cum_db)
self.cum_db = Tensor(self.b.size()).zero_()
class ReLU(Module):
"""Implements the Rectified Linear Unit activation layer"""
def forward(self, x):
"""Carries out the forward pass for backpropagation."""
self.x = x
return self.x.clamp(min=0)
def backward(self, grad):
"""Carries out the backward pass for backpropagation."""
return grad * Tensor(np.where(self.x <= 0,
0, 1)).view(grad.size())
class LeakyReLU(Module):
"""Implements the Leaky ReLU activation layer"""
def __init__(self, a=.001):
self.a = a
"""Carries out the forward pass for backpropagation."""
self.x = x
return Tensor(np.where(x >= 0, x, self.a * x ))
def backward(self, grad):
"""Carries out the backward pass for backpropagation."""
return grad * Tensor(np.where(self.x >= 0,
1, self.a)).view(grad.size())
class Tanh(Module):
"""Implements the Tanh activation layer"""
def forward(self, x):
"""Carries out the forward pass for backpropagation."""
self.x_tanh = x.tanh()
return self.x_tanh
def backward(self, grad):
"""Carries out the backward pass for backpropagation."""
return grad * (1 - self.x_tanh ** 2).view(grad.size())
class Sigmoid(Module):
"""Implements the Rectified Linear Unit activation layer
It is recommended to use the Tanh module instead.
"""
def forward(self, x):
"""Carries out the forward pass for backpropagation."""
self.sigmoid = (1 + (x / 2).tanh()) / 2
return self.sigmoid
def backward(self, grad):
"""Carries out the backward pass for backpropagation."""
return grad * (self.sigmoid * (1 - self.sigmoid)).view(grad.size())
class LossMSE(Module):
"""Implements the MSE loss computation"""
def forward(self, output, target):
"""Carries out the forward pass for backpropagation."""
self.diff = output.float() - target.float().view(output.size())
return (self.diff ** 2).sum()
def backward(self):
"""Carries out the backward pass for backpropagation."""
return self.diff * 2
class Sequential(Module):
"""Allows to combine several modules sequentially
It is possible to either include a loss module in the Sequential
module or to not include it and use a loss module defined outside
of the Sequential module instead.
"""
def __init__(self, layers, loss=None):
self.layers = layers
self.loss = loss
def forward(self, x, target=None):
"""Carries out the forward pass for backpropagation
To do it it calls the forward functions of each individual
module.
"""
if self.loss is not None:
assert target is not None, "Target required for loss module"
for l in self.layers:
x = l.forward(x)
if self.loss is not None:
x = self.loss.forward(x, target)
self.x = x
return x
def backward(self, grad=None):
"""Carries out the backward pass for backpropagation
To do it it calls the backward functions of each individual
module
"""
if self.loss is not None:
grad = self.loss.backward()
else:
assert grad is not None, "Initial gradient required when no loss module defined"
for l in reversed(self.layers):
grad = l.backward(grad)
def param(self):
return [p for l in layers for p in l.param()]
def update(self, lr=None):
for l in self.layers:
l.update(lr)
# #### Training
def adaptive_lr(kappa=0.75, eta0=1e-5):
"""Adaptive learning rate. After creating the lr with the
values for kappa and eta0, it yields the value for the learning
rate of the next iteration. Used for (Stochastic) Gradient Descent
methods.
"""
t = 1
while True:
yield eta0 * t ** -kappa
t += 1
def SGD(model, loss, train_input, train_target, n_epochs, mini_batch_size, a_lr):
"""Trains a model, which is usually defined as a Sequential module"""
for e in range(n_epochs):
L_tot = 0
errors_tot = 0
p = np.random.permutation(len(train_target))
train_zipped = zip(train_input[p], train_target[p])
predicted_labels = []
for i, (data, label) in enumerate(train_zipped):
d = Tensor(data)
l = Tensor(1).fill_(label)
# Forward pass
output = model.forward(d)
L = loss.forward(output, l)
predicted_labels.append(output)
# Backward pass
grad = loss.backward()
model.backward(grad)
# update gradients
if not (i + 1) % mini_batch_size:
next_a_lr = next(a_lr)
model.update(next_a_lr)
# Compute total loss and accuracy
L_tot += L
errors = (l != (output >= 0.5).float() ).sum()
errors_tot += errors
next_a_lr = next(a_lr)
model.update(next_a_lr)
accuracy = (len(train_target) - errors_tot) / len(train_target)
print('Epoch {:d} Loss {:.08f} Accuracy {:.02f} Errors {}'.format(
e, L_tot, accuracy, errors_tot))
return train_input[p], train_target[p], np.array(predicted_labels)
# #### Generate dataset
#
# Where is the disk centered? We assume (0,0)
# +
def convert_to_one_hot_labels(input, target):
tmp = input.new(target.size(0), target.max() + 1).fill_(0)
tmp.scatter_(1, target.view(-1, 1), 1.0)
return tmp.long()
def generate_disc_set(nb):
data = torch.Tensor(nb, 2).uniform_(0, 1)
label = ((data - .5) ** 2).sum(1) <= 1 / (2 * np.pi)
return data, convert_to_one_hot_labels(data,label.long())
# -
train_input, train_target = generate_disc_set(1000)
train_data_0 = train_input[train_target == 0]
train_data_1 = train_input[train_target == 1]
# Plot data points
import matplotlib.pyplot as plt
plt.scatter(train_data_0.transpose()[0], train_data_0.transpose()[1], c="red")
plt.scatter(train_data_1.transpose()[0], train_data_1.transpose()[1], c="blue")
plt.show()
# #### Test
# Why two output units?
def get_rand_params():
"""Generates random parameters for testing purposes"""
kappas = np.linspace(.5, .95, 30)
#kappas = np.linspace(.76, .92, 10)
etas = np.logspace(-7, -1, 30)
#etas = np.logspace(-4.32, -2, 10)
mbs = np.linspace(20, 160, 30).round()
return (np.random.choice(kappas),
np.random.choice(etas),
np.random.choice(mbs))
# + active=""
# # Iterate through several combinations of random parameters
# # This should be helpful for finding the learning rate and minibatch size
# for i in range(10):
# k, e0, mb = get_rand_params()
# a_lr = adaptive_lr(kappa=k, eta0=e0)
# print()
# print()
# print("-------------------")
# print("k:{}, e0:{}, mb:{}".format(k, e0, mb))
# print("-------------------")
#
# loss = LossMSE()
# hl1 = [Linear(2, 25, lr=0), ReLU()]
# hl2 = [Linear(25, 25, lr=0), ReLU()]
# hl3 = [Linear(25, 25, lr=0), ReLU()]
# out = [Linear(25, 1, lr=0), Sigmoid()]
#
# model = Sequential(hl1 + hl2 + hl3 + out)
#
# SGD(model, loss, train_input, train_target, 20, mb, a_lr)
# +
# Some good parameter combinations:
#k:0.593103448275862, e0:0.014873521072935119, mb:112.0
#k:0.5, e0:0.009236708571873866, mb:34.0
#k:0.6241379310344828, e0:0.03856620421163472, mb:92.0
k, e0, mb = (.6, 3.5e-2, 60)
a_lr = adaptive_lr(kappa=k, eta0=e0)
loss = LossMSE()
hl1 = [Linear(2, 25, lr=0), ReLU()]
hl2 = [Linear(25, 25, lr=0), ReLU()]
hl3 = [Linear(25, 25, lr=0), ReLU()]
out = [Linear(25, 1, lr=0), Sigmoid()]
model = Sequential(hl1 + hl2 + hl3 + out)
p_data, p_target, pred_labels = SGD(model, loss, train_input, train_target, 200, mb, a_lr)
pred_labels = (pred_labels >= 0.5)
# +
pred_data_errors = p_data[pred_labels != p_target]
# Plot predicted labels
plt.scatter(train_data_0.transpose()[0], train_data_0.transpose()[1], c="red", alpha=.3)
plt.scatter(train_data_1.transpose()[0], train_data_1.transpose()[1], c="blue", alpha=.3)
plt.scatter(pred_data_errors.transpose()[0], pred_data_errors.transpose()[1], c="yellow", alpha=.5)
plt.show()
# +
test_input, test_target = generate_disc_set(1000)
def testing(model, loss, test_input, test_target):
"""Tests a model, which is usually defined as a Sequential module"""
L_tot = 0
errors_tot = 0
test_zipped = zip(test_input, test_target)
predicted_labels = []
for data, label in test_zipped:
d = Tensor(data)
l = Tensor(1).fill_(label)
# Get predicted label and loss
output = model.forward(d)
predicted_labels.append(output)
L = loss.forward(output, l)
# Compute total loss and accuracy
L_tot += L
errors = (l != (output >= 0.5).float() ).sum()
errors_tot += errors
accuracy = (len(test_target) - errors_tot) / len(test_target)
print('Loss {:.08f} Accuracy {:.02f} Errors {}'.format(
L_tot, accuracy, errors_tot))
return np.array(predicted_labels)
pred_labels = testing(model, loss, test_input, test_target)
pred_labels = (pred_labels >= 0.5)
test_data_0 = test_input[test_target == 0]
test_data_1 = test_input[test_target == 1]
pred_data_errors = test_input[pred_labels != test_target]
# Plot data points
plt.scatter(test_data_0.transpose()[0], test_data_0.transpose()[1], c="gray", alpha=1)
plt.scatter(test_data_1.transpose()[0], test_data_1.transpose()[1], c="lightgray", alpha=1)
plt.scatter(pred_data_errors.transpose()[0], pred_data_errors.transpose()[1], c="red", alpha=1)
plt.show()
# -
| Projects/Project2/Notebooks and figs/Project2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Basics
# ### Let's Start
# - The main difference between tensors and NumPy arrays is that tensors can be used on GPUs (graphical processing units) and TPUs (tensor processing units).
# - The number of dimensions of a tensor is called its rank. A scalar has rank $0$, a vector has rank $1$, a matrix is rank $2$, a tensor has rank $n$.
# - There are $2$ ways of creating tensors. `tf.Variable()` and `tf.constant()` the difference being tensors created with `tf.constant()` are immutable, tensors created with `tf.Variable()` are mutable. `any_tensor[2].assign(7)` can be used to change a value of a specific element in the tensor, same would fail for `tf.constant()`.
# - There are other ways of creating tensors examples being `tf.zeros` or `tf.ones`. You can also convert numpy arrays into tensors.
# - Tensors can also be indexed just like Python lists.
# - You can an extra dimension by using `tf.newaxis` or `tf.expand_dims`
# - `tf.reshape()`,`tf.transpose()` allows us to reshape a tensor
# - Data type of a tesnor can be changed with `tf.cast(t1, dtype=tf.float16)`
# - You can squeeze a tensor to remove single-dimensions (dimensions with size 1) using `tf.squeeze()`.
# +
# Some common commands are as follows
import tensorflow as tf
print("Check TF version: ",tf.__version__)
t1 = tf.constant([[10., 7.],
[3., 2.],
[8., 9.]], dtype=tf.float16) # by default TF creates tensors with either an int32 or float32 datatype.
print("Access a specific feature of the tensor, in this case shape of t1: ",t1.shape)
print("Size of t1: ", tf.size(t1))
print("Datatype of every element:", t1.dtype)
print("Number of dimensions (rank):", t1.ndim)
print("Shape of tensor:", t1.shape)
print("Elements along axis 0 of tensor:", t1.shape[0])
print("Elements along last axis of tensor:", t1.shape[-1])
print("Total number of elements:", tf.size(t1).numpy()) # .numpy() converts to NumPy array
print("Details of the tensor: ",t1)
print("Index tensors: ", t1[:1,:])
# +
import tensorflow as tf
# Math operations
t1 = tf.constant([[10., 7.],
[3., 2.],
[8., 9.]], dtype=tf.float16) # by default TF creates tensors with either an int32 or float32 datatype.
print("Sum: ",t1+10)
print("Substraction: ",t1-10)
print("Multiplication: ",t1*10, tf.multiply(t1, 10))
print("Matrix Multiplication: ",t1 @ tf.transpose(t1)) # can also be done with tf.tensordot()
# Aggregation functions
print("Max: ", tf.reduce_max(t1)) # same or min, mean
print("Sum: ", tf.reduce_sum(t1))
print("Max Position: ", tf.argmax(t1)) # same or min
# -
# ### Random
# Randomness is often used in deep learning, be it initializing weights in a Neural Network or shuffling images while feeding data to the model.
random_1 = tf.random.Generator.from_seed(35) # setting seed ensures reproducibility
random_1 = random_1.normal(shape = (3,2))
print("Generating tensor from a normal distribution: ", random_1)
print("Shuffling the elements of the tesnor: ", tf.random.shuffle(random_1))
#
| _build/jupyter_execute/contents/Tensorflow/Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Excercises Electric Machinery Fundamentals
# ## Chapter 6
# ## Problem 6-2
# + slideshow={"slide_type": "skip"}
# %pylab inline
# -
# ### Description
# Answer the questions in Problem 6-1 for a 480-V three-phase two-pole 60-Hz induction motor running at a slip of 0.025.
fse = 60.0 # [Hz]
p = 2.0
s = 0.025
# ### SOLUTION
# #### (a)
# The speed of the magnetic fields is:
#
# $$n_\text{sync} = \frac{120f_{se}}{p}$$
n_sync = 120*fse / p
print('''
n_sync = {:.0f} r/min
==================='''.format(n_sync))
# #### (b)
# The speed of the rotor is:
#
# $$n_m = (1-s)n_\text{sync}$$
n_m = (1 - s) * n_sync
print('''
n_m = {:.0f} r/min
================'''.format(n_m))
# #### (c)
# The slip speed of the rotor is:
#
# $$n_\text{slip} = s\cdot n_\text{sync}$$
n_slip = s * n_sync
print('''
n_slip = {:.0f} r/min
================='''.format(n_slip))
# #### (d)
# The rotor frequency is:
#
# $$f_{re} = \frac{p\cdot n_\text{slip}}{120}$$
fre = p * n_slip / 120
print('''
fre = {:.1f} Hz
============'''.format(fre))
| Chapman/Ch6-Problem_6-02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''env'': venv)'
# name: python3
# ---
# +
base_transfer_set = ['01', '02', '04', '05', '08', '09', '12', '13', '16', '17', '18', '20']
target_transfer_set = ['03', '06', '07', '10', '11', '14', '15', '19']
import random
def random_combination(iterable, r):
"Random selection from itertools.combinations(iterable, r)"
pool = tuple(iterable)
n = len(pool)
indices = sorted(random.sample(range(n), r))
return tuple(pool[i] for i in indices)
transfers_size_6 = []
for i in range(4):
transfers_size_6.append(random_combination(target_transfer_set, 6))
print(transfers_size_6)
transfers_size_6 = [('03', '06', '07', '10', '11', '14'), ('03', '06', '07', '10', '14', '15'), ('03', '06', '07', '10', '14', '15'), ('03', '07', '10', '14', '15', '19')]
for i, tmp in enumerate(transfers_size_6):
transfers_size_6[i] = list(transfers_size_6[i])
print(transfers_size_6)
transfers_size_4 = []
for i in range(4):
transfers_size_4.append(random_combination(target_transfer_set, 4))
print(transfers_size_4)
transfers_size_4 = [('06', '10', '14', '15'), ('03', '10', '14', '19'), ('03', '06', '10', '15'), ('03', '07', '10', '15')]
for i, tmp in enumerate(transfers_size_4):
transfers_size_4[i] = list(transfers_size_4[i])
print(transfers_size_4)
transfers_size_3 = []
for i in range(4):
transfers_size_3.append(random_combination(target_transfer_set, 3))
print(transfers_size_3)
transfers_size_3 = [('07', '11', '14'), ('06', '07', '10'), ('03', '15', '19'), ('06', '14', '19')]
for i, tmp in enumerate(transfers_size_3):
transfers_size_3[i] = list(transfers_size_3[i])
print(transfers_size_3)
transfers_size_2 = []
for i in range(4):
transfers_size_2.append(random_combination(target_transfer_set, 2))
print(transfers_size_2)
transfers_size_2 = [('06', '10'), ('07', '11'), ('06', '15'), ('14', '15')]
for i, tmp in enumerate(transfers_size_2):
transfers_size_2[i] = list(transfers_size_2[i])
print(transfers_size_2)
# +
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
def make_confusion_matrix(cf,
group_names=None,
categories='auto',
count=True,
percent=True,
cbar=True,
xyticks=True,
xyplotlabels=True,
sum_stats=True,
figsize=None,
cmap='Blues',
title=None):
'''
This function will make a pretty plot of an sklearn Confusion Matrix cm using a Seaborn heatmap visualization.
Arguments
---------
cf: confusion matrix to be passed in
group_names: List of strings that represent the labels row by row to be shown in each square.
categories: List of strings containing the categories to be displayed on the x,y axis. Default is 'auto'
count: If True, show the raw number in the confusion matrix. Default is True.
normalize: If True, show the proportions for each category. Default is True.
cbar: If True, show the color bar. The cbar values are based off the values in the confusion matrix.
Default is True.
xyticks: If True, show x and y ticks. Default is True.
xyplotlabels: If True, show 'True Label' and 'Predicted Label' on the figure. Default is True.
sum_stats: If True, display summary statistics below the figure. Default is True.
figsize: Tuple representing the figure size. Default will be the matplotlib rcParams value.
cmap: Colormap of the values displayed from matplotlib.pyplot.cm. Default is 'Blues'
See http://matplotlib.org/examples/color/colormaps_reference.html
title: Title for the heatmap. Default is None.
'''
# CODE TO GENERATE TEXT INSIDE EACH SQUARE
blanks = ['' for i in range(cf.size)]
if group_names and len(group_names)==cf.size:
group_labels = ["{}\n".format(value) for value in group_names]
else:
group_labels = blanks
if count:
group_counts = ["{0:0.0f}\n".format(value) for value in cf.flatten()]
else:
group_counts = blanks
if percent:
group_percentages = ["{0:.2%}".format(value) for value in cf.flatten()/np.sum(cf)]
else:
group_percentages = blanks
box_labels = [f"{v1}{v2}{v3}".strip() for v1, v2, v3 in zip(group_labels,group_counts,group_percentages)]
box_labels = np.asarray(box_labels).reshape(cf.shape[0],cf.shape[1])
# CODE TO GENERATE SUMMARY STATISTICS & TEXT FOR SUMMARY STATS
if sum_stats:
#Accuracy is sum of diagonal divided by total observations
accuracy = np.trace(cf) / float(np.sum(cf))
#if it is a binary confusion matrix, show some more stats
if len(cf)==2:
#Metrics for Binary Confusion Matrices
precision = cf[1,1] / sum(cf[:,1])
recall = cf[1,1] / sum(cf[1,:])
f1_score = 2*precision*recall / (precision + recall)
stats_text = "\n\nAccuracy={:0.3f}\nPrecision={:0.3f}\nRecall={:0.3f}\nF1 Score={:0.3f}".format(
accuracy,precision,recall,f1_score)
else:
stats_text = "\n\nAccuracy={:0.3f}".format(accuracy)
else:
stats_text = ""
# SET FIGURE PARAMETERS ACCORDING TO OTHER ARGUMENTS
if figsize==None:
#Get default figure size if not set
figsize = plt.rcParams.get('figure.figsize')
if xyticks==False:
#Do not show categories if xyticks is False
categories=False
# MAKE THE HEATMAP VISUALIZATION
plt.figure(figsize=figsize)
sns.heatmap(cf,annot=box_labels,fmt="",cmap=cmap,cbar=cbar,xticklabels=categories,yticklabels=categories)
if xyplotlabels:
plt.ylabel('True label')
plt.xlabel('Predicted label' + stats_text)
else:
plt.xlabel(stats_text)
if title:
plt.title(title)
# +
import os
import pandas as pd
import warnings
warnings.filterwarnings("ignore")
def create_best_model(gesture_subset):
gesture_subset.sort()
print("Loadind Dataset for gestures: ", gesture_subset)
path = 'gestures-dataset'
dataset = None
samples = 0
for subject in os.listdir(path):
if os.path.isfile(os.path.join(path, subject)):
continue
if subject in ('U01', 'U02', 'U03', 'U04', 'U05', 'U06', 'U07', 'U08'):
for gesture in os.listdir(os.path.join(path, subject)):
if os.path.isfile(os.path.join(path, subject, gesture)):
continue
gesture = str(gesture)
if gesture not in gesture_subset:
continue
for samplefile in os.listdir(os.path.join(path, subject, gesture)):
if os.path.isfile(os.path.join(path, subject, gesture, samplefile)):
df = pd.read_csv(os.path.join(path, subject, gesture, samplefile), \
sep = ' ', \
names = ['System.currentTimeMillis()', \
'System.nanoTime()', \
'sample.timestamp', \
'X', \
'Y', \
'Z' \
])
df = df[["sample.timestamp", "X", "Y", "Z"]]
start = df["sample.timestamp"][0]
df["sample.timestamp"] -= start
df["sample.timestamp"] /= 10000000
df["subject"] = subject
df["gesture"] = gesture
df["sample"] = str(samplefile[:-4])
samples += 1
#print(df)
if dataset is None:
dataset = df.copy()
else:
dataset = pd.concat([dataset, df])
dataset = dataset.sort_values(by=['gesture','subject','sample','sample.timestamp'])
data = dataset
print(str(samples) + " samples loaded")
print("Scaling Dataset for gestures: ", gesture_subset)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
dataset_scaled = None
samples = 0
for i, gesture in enumerate(gesture_subset):
df_gesture=data[data['gesture']==gesture]
for j, subject in enumerate(df_gesture['subject'].unique()):
df_subject=df_gesture[df_gesture['subject']==subject]
for k, sample in enumerate(df_subject['sample'].unique()):
df_sample=df_subject[df_subject['sample']==sample].copy()
df_sample.sort_values(by=['sample.timestamp'])
sc = scaler
sc = sc.fit_transform(df_sample[["X", "Y", "Z"]])
sc = pd.DataFrame(data=sc, columns=["X", "Y", "Z"])
df_sample['X'] = sc['X']
df_sample['Y'] = sc['Y']
df_sample['Z'] = sc['Z']
if dataset_scaled is None:
dataset_scaled = df_sample.copy()
else:
dataset_scaled = pd.concat([dataset_scaled, df_sample])
samples += 1
print(str(samples) + " samples scaled")
data = dataset_scaled
print("Cleaning Dataset for gestures: ", gesture_subset)
dataset_outliers = None
dataset_cleaned = None
samples = 0
outliers = 0
for i, gesture in enumerate(gesture_subset):
df_gesture = data[data['gesture']==gesture]
for j, subject in enumerate(df_gesture['subject'].unique()):
df_subject = df_gesture[df_gesture['subject']==subject]
time_mean = df_subject.groupby(["gesture","subject", "sample"]).count().groupby(["gesture","subject"]).agg({'sample.timestamp': ['mean']})
time_std = df_subject.groupby(["gesture","subject", "sample"]).count().groupby(["gesture","subject"]).agg({'sample.timestamp': ['std']})
time_max = time_mean['sample.timestamp'].iloc[0]['mean'] + 1.0 * time_std['sample.timestamp'].iloc[0]['std']
time_min = time_mean['sample.timestamp'].iloc[0]['mean'] - 1.0 * time_std['sample.timestamp'].iloc[0]['std']
for k, sample in enumerate(df_subject['sample'].unique()):
df_sample=df_subject[df_subject['sample']==sample]
df_sample_count = df_sample.count()['sample.timestamp']
if df_sample_count < time_min or df_sample_count > time_max:
if dataset_outliers is None:
dataset_outliers = df_sample.copy()
else:
dataset_outliers = pd.concat([dataset_outliers, df_sample])
outliers += 1
else:
if dataset_cleaned is None:
dataset_cleaned = df_sample.copy()
else:
dataset_cleaned = pd.concat([dataset_cleaned, df_sample])
samples += 1
print(str(samples) + " samples cleaned")
print(str(outliers) + " samples outliers")
data = dataset_cleaned
print("Time slicing Cleaned Dataset for gestures: ", gesture_subset)
dataset_timecut = None
samples = 0
damaged = 0
for i, gesture in enumerate(data['gesture'].unique()):
df_gesture = data[data['gesture']==gesture]
for j, subject in enumerate(df_gesture['subject'].unique()):
df_subject = df_gesture[df_gesture['subject']==subject]
time_max = 19 # 18 * 11 = 198
for i, sample in enumerate(df_subject['sample'].unique()):
df_sample = df_subject[df_subject['sample']==sample]
df_sample_count = df_sample.count()['sample.timestamp']
#print(df_sample_count)
if df_sample_count >= time_max:
df_sample = df_sample[df_sample['sample.timestamp'] <= (11 * (time_max-1))]
df_sample_count = df_sample.count()['sample.timestamp']
#print(df_sample_count)
elif df_sample_count < time_max:
for tmp in range(df_sample_count * 11, (time_max) * 11, 11):
df = pd.DataFrame([[tmp, 0.0, 0.0, 0.0, gesture, subject, sample]], columns=['sample.timestamp', 'X', 'Y', 'Z', 'gesture', 'subject', 'sample'])
df_sample = df_sample.append(df, ignore_index=True)
#print(df_sample)
df_sample_count = df_sample.count()['sample.timestamp']
#print(df_sample_count)
if df_sample_count != time_max:
damaged += 1
continue
if dataset_timecut is None:
dataset_timecut = df_sample.copy()
else:
dataset_timecut = pd.concat([dataset_timecut, df_sample])
samples += 1
dataset_cleaned = dataset_timecut
print(str(samples) + " cleaned samples sliced")
print(str(damaged) + " cleaned samples damaged")
data = dataset_outliers
print("Time slicing Outliers Dataset for gestures: ", gesture_subset)
dataset_timecut = None
samples = 0
damaged = 0
for i, gesture in enumerate(data['gesture'].unique()):
df_gesture = data[data['gesture']==gesture]
for j, subject in enumerate(df_gesture['subject'].unique()):
df_subject = df_gesture[df_gesture['subject']==subject]
time_max = 19 # 18 * 11 = 198
for i, sample in enumerate(df_subject['sample'].unique()):
df_sample = df_subject[df_subject['sample']==sample]
df_sample_count = df_sample.count()['sample.timestamp']
#print(df_sample_count)
if df_sample_count >= time_max:
df_sample = df_sample[df_sample['sample.timestamp'] <= (11 * (time_max-1))]
df_sample_count = df_sample.count()['sample.timestamp']
#print(df_sample_count)
elif df_sample_count < time_max:
for tmp in range(df_sample_count * 11, (time_max) * 11, 11):
df = pd.DataFrame([[tmp, 0.0, 0.0, 0.0, gesture, subject, sample]], columns=['sample.timestamp', 'X', 'Y', 'Z', 'gesture', 'subject', 'sample'])
df_sample = df_sample.append(df, ignore_index=True)
#print(df_sample)
df_sample_count = df_sample.count()['sample.timestamp']
#print(df_sample_count)
if df_sample_count != time_max:
damaged += 1
continue
if dataset_timecut is None:
dataset_timecut = df_sample.copy()
else:
dataset_timecut = pd.concat([dataset_timecut, df_sample])
samples += 1
dataset_outliers = dataset_timecut
print(str(samples) + " outliers samples sliced")
print(str(damaged) + " outliers samples damaged")
data = dataset_cleaned
from keras.models import Sequential
from keras.layers import Bidirectional
from keras.layers import LSTM
from keras.layers import Dense
from keras.layers import Dropout
from keras.optimizers import adam_v2
from keras.wrappers.scikit_learn import KerasClassifier
# from scikeras.wrappers import KerasClassifier
from sklearn.model_selection import StratifiedGroupKFold
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import GridSearchCV
from keras.utils import np_utils
from sklearn.preprocessing import LabelEncoder
from sklearn.pipeline import Pipeline
import numpy as np
# fix random seed for reproducibility
seed = 1000
np.random.seed(seed)
# create the dataset
def get_dataset(data):
X_train = []
Y_train = []
groups = []
for i, gesture in enumerate(data['gesture'].unique()):
df_gesture = data[data['gesture']==gesture]
for j, subject in enumerate(df_gesture['subject'].unique()):
df_subject = df_gesture[df_gesture['subject']==subject]
for k, sample in enumerate(df_subject['sample'].unique()):
df_sample = df_subject[df_subject['sample']==sample]
accel_vector = []
for index, row in df_sample.sort_values(by='sample.timestamp').iterrows():
accel_vector.append([row['X'],row['Y'],row['Z']])
accel_vector = np.asarray(accel_vector)
X_train.append(accel_vector)
Y_train.append(gesture)
groups.append(subject)
X_train = np.asarray(X_train)
Y_train = LabelEncoder().fit_transform(Y_train)
#print(Y_train)
return X_train, Y_train, groups
# Function to create model, required for KerasClassifier
def create_model(dropout_rate=0.8, units=128, optimizer=adam_v2.Adam(learning_rate=0.001)):
model = Sequential()
model.add(
Bidirectional(
LSTM(
units=units,
input_shape=[19, 3]
)
)
)
model.add(Dropout(rate=dropout_rate))
model.add(Dense(units=units, activation='relu'))
model.add(Dense(len(gesture_subset), activation='softmax'))
model.compile(loss='sparse_categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
#print(model.summary())
return model
model = KerasClassifier(build_fn=create_model, verbose=0)
cv = StratifiedGroupKFold(n_splits=5, shuffle=True, random_state=1000)
# get the dataset
X, y, g = get_dataset(dataset_cleaned)
#cv = cv.split(X, y, g)
batch_size = [19]
epochs = [64, 128]
#epochs = [128]
units = [16, 32, 64, 128]
# units = [16]
dropout_rate = [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]
# dropout_rate = [0.5]
param_grid = dict(epochs=epochs, units=units, batch_size=batch_size, dropout_rate=dropout_rate)
print("Hyperparameter tunning started for Dataset for gestures: ", gesture_subset)
grid = GridSearchCV(estimator=model, param_grid=param_grid, n_jobs=1, cv=cv, verbose=1)
grid_result = grid.fit(X, y, groups=g)
# summarize results
print("Best: %f using %s" % (grid_result.best_score_, grid_result.best_params_))
means = grid_result.cv_results_['mean_test_score']
stds = grid_result.cv_results_['std_test_score']
train_mean = grid_result.cv_results_['mean_fit_time']
train_std = grid_result.cv_results_['std_fit_time']
score_mean = grid_result.cv_results_['mean_score_time']
score_std = grid_result.cv_results_['std_score_time']
params = grid_result.cv_results_['params']
for mean, stdev, train_mean, train_std, score_mean, score_std, param in zip(means, stds, train_mean, train_std, score_mean, score_std, params):
print("accuracy: %f (%f) train time: %f (%f) score time: %f (%f) with: %r" % (mean, stdev, train_mean, train_std, score_mean, score_std, param))
print("Hyperparameter tunning completed for Dataset: ", gesture_subset)
model = grid_result.best_estimator_
import pickle
def save_model(model, gesture_subset):
gesture_subset.sort()
name = '-'.join(gesture_subset)
# saving model
pickle.dump(model.classes_, open(name + '_model_classes.pkl','wb'))
model.model.save(name + '_lstm')
print("Saving model to disk started for Dataset gestures: ", gesture_subset)
save_model(model, gesture_subset)
print("Saving model to disk completed for Dataset gestures: ", gesture_subset)
import tensorflow as tf
def load_model(gesture_subset):
gesture_subset.sort()
name = '-'.join(gesture_subset)
# loading model
build_model = lambda: tf.keras.models.load_model(name + '_lstm')
classifier = KerasClassifier(build_fn=build_model, epochs=1, batch_size=10, verbose=0)
classifier.classes_ = pickle.load(open(name + '_model_classes.pkl','rb'))
classifier.model = build_model()
return classifier
print("Loading model to disk started for Dataset gestures: ", gesture_subset)
model = load_model(gesture_subset)
#print(model.model.summary())
print("Loading model to disk completed for Dataset gestures: ", gesture_subset)
print("Testing model against outliers for Dataset gestures: ", gesture_subset)
data = dataset_outliers
X, y, g = get_dataset(dataset_outliers)
y_pred = model.predict(X)
#print(y)
#print(y_pred)
from sklearn.metrics import classification_report
print(classification_report(y, y_pred, target_names=gesture_subset))
from sklearn.metrics import confusion_matrix
cf_matrix = confusion_matrix(y, y_pred)
make_confusion_matrix(cf_matrix, categories=gesture_subset)
return model
base_transfer_set = ['01', '02', '04', '05', '08', '09', '12', '13', '16', '17', '18', '20']
dataset = transfers_size_6[0]
model = create_best_model(dataset)
# -
from IPython.display import Image
Image('gestures-dataset/gestures.png')
| EDA/01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="rHxKzDGIdgev"
# **Nmed Entity Recognition**
#
# - NER is an NLP task of categorizing the words in a text to common categories such as the name of a person, date, organization, etc.
# - It can be used to extract information like products named in a complaint, location, companies in an article etc.
#
# + [markdown] id="BJNmaWh0fBhB"
# NER implimentation using NLTK
# + id="vj75MN2ndcgB"
# Downloading nltk requirements
import nltk
from nltk.tokenize import word_tokenize
from nltk.tag import pos_tag
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
# + id="sdMlkA-nfiP2"
Document = ['''Dr. <NAME> was named Canada’s Chief Public Health Officer (CPHO) on June 26, 2017.''', ''' She is a physician with expertise in immunization, infectious disease, emergency preparedness and global health security.''', '''
As the federal government’s lead health professional, Dr. Tam provides advice to the Minister of Health, supports and provides advice to the President of the Public Health Agency of Canada, and works in collaboration with the President in the leadership and management of the Agency.''',
'''The Public Health Agency of Canada Act empowers the CPHO to communicate with other levels of government, voluntary organizations, the private sector and Canadians on public health issues.''', '''Each year, the CPHO is required to submit a report to the Minister of Health on the state of public health in Canada.''']
# + id="qlQY2zUDgC8Z" colab={"base_uri": "https://localhost:8080/"} outputId="80df834e-5cda-4331-962d-70b2c3963ecd"
# Word tokenization
def pre_process(Doc):
Named = []
for sent in Doc:
sent = nltk.word_tokenize(sent)
sent = nltk.pos_tag(sent) # part of speech tag
Named.append(sent)
return Named
tokenized_sentences = pre_process(Document)
tokenized_sentences
# + [markdown] id="_kbvVNjTn4dP"
# Chunking
#
# - Use regular expressions to identity the named entities
# - For example the noun phrase chunking pattern below
# - The chunk pattern consists of one rule, that a noun phrase, NP, should be formed whenever the chunker finds an optional determiner, DT, followed by any number of adjectives, JJ, and then a noun, NN.
# + colab={"base_uri": "https://localhost:8080/"} id="YOSN3BOAtBjM" outputId="bccb607a-89e2-4c1c-a3b9-aca39302bf18"
# Pattern
pattern = 'NP: {<DT>?<JJ>*<NN>}'
# Chunk paser
cp = nltk.RegexpParser(pattern)
cs = cp.parse(tokenized_sentences[0])
print(cs)
# + [markdown] id="9tetE6nifrtR"
# Using IOB format to represent the chunk structures in files.
# - I - word in inside, eg. I-NP means word is inside a noun phrase
# - O - End of sentence
# - B - Beginning of a phrase, NP of VP
#
# + colab={"base_uri": "https://localhost:8080/"} id="Pma8uBigtZ0U" outputId="68bde166-1b9c-4e9e-d1ed-c02b1b361097"
from nltk import tree2conlltags
iob_tags = tree2conlltags(cs)
iob_tags
# Returns word, part of speech tag, IOBtag tuples.
# + [markdown] id="rLmn4U-qiwQN"
# nltk.ne_chunk can be used to indetify named entity and classify them using the classifier.
# + id="aTywF1YNkeiW"
from nltk import ne_chunk
nltk.download('maxent_ne_chunker')
nltk.download('words')
# + colab={"base_uri": "https://localhost:8080/"} id="V4p9n_0wEVgY" outputId="386399d1-1b28-4083-e356-b4740cbf3181"
ne_tree = [ne_chunk(pos_tag(word_tokenize(sent))) for sent in Document]
ne_tree
# + [markdown] id="A84fnMuHlfSf"
# Using **Spacy** for named entity recognition
# + id="1E8jQLBAlmoC"
import spacy
from spacy import displacy
from collections import Counter
import en_core_web_sm
nlp = en_core_web_sm.load()
# + [markdown] id="2FlzSQy09nM-"
# Entity level.
# + colab={"base_uri": "https://localhost:8080/"} id="zNh2KLg55Z7f" outputId="d232526b-13f3-491f-8052-dcefb911e64f"
# Tokenize and print entity level tags
Entities = nlp(Document[0])
[(x.text, x.label_) for x in Entities.ents]
# + [markdown] id="2ZWSRW5h9rVB"
# Token level entity annotation.
#
# - **B**egin - The first token of a multi_token entity
# - **I**n - An inner token of a multi-token entity
# - **L**ast - The final token of a multi-token entity
# - **U**nit - A single token entity
# - **O**ut - A non-entity token
# + colab={"base_uri": "https://localhost:8080/"} id="67LnsQLH9TmY" outputId="a331522d-25ee-491d-b81c-1104ef424858"
[(x, x.ent_iob_, x.ent_type_) for x in Entities]
# + [markdown] id="hTfTOW1GV243"
# **NER** extraction from an article.
# + id="LGGHz19_V9I8"
from bs4 import BeautifulSoup
import requests
import re
def url_to_string(URL):
res = requests.get(URL)
html = res.text
soup = BeautifulSoup(html, 'html5lib')
for script in soup(['script', 'style', 'aside']):
script.extract()
return " ".join(re.split(r'[\n\t]+', soup.get_text()))
Preprocessed = url_to_string('https://www.nytimes.com/2018/08/13/us/politics/peter-strzok-fired-fbi.html?hp&action=click&pgtype=Homepage&clickSource=story-heading&module=first-column-region®ion=top-news&WT.nav=top-news')
Article = nlp(Preprocessed)
# + colab={"base_uri": "https://localhost:8080/"} id="kEDTORqmaS6N" outputId="1dacc7f0-bc24-482d-8f1b-2498cd283721"
# Labels counter
Labels = [X.label_ for X in Article.ents]
Counter(Labels)
# + colab={"base_uri": "https://localhost:8080/"} id="eIfHy3tOchsE" outputId="9475783e-00ce-41f6-c97b-924a7caee8b4"
# Most frequently mentioned tokens
# To quickly find out what the article is about
items = [x.text for x in Article.ents]
Counter(items).most_common(3)
# + [markdown] id="q-f5I-BrhIxs"
# Naming the entities and extracting the lemma
# + colab={"base_uri": "https://localhost:8080/"} id="MffetQ8of1Qf" outputId="04ef12b2-5e24-457e-b2af-5498be408250"
#
sentences = [x for x in Article.sents]
[(X.orth_, X.pos_, X.lemma_) for X in [Y for Y in nlp(str(sentences[20]))
if not Y.is_stop and Y.pos_ != 'PUNCT']]
| Named_Entity_Recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="nm_n651NMm7M" colab_type="code" outputId="17d62d82-c386-4747-f320-783d2aac0d75" executionInfo={"status": "ok", "timestamp": 1583185729996, "user_tz": -60, "elapsed": 469, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "/content/drive/My Drive/Colab Notebooks/"
# + id="i1bunmOlM2CE" colab_type="code" outputId="6bcac896-29cb-4e29-9ef9-9f15d834d0ad" executionInfo={"status": "ok", "timestamp": 1583185363464, "user_tz": -60, "elapsed": 3767, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# ls
# + id="ay_xEgAoM3L9" colab_type="code" colab={}
# !mkdir -p "dw_matrix2"
# + id="_D47CfLiM7Cg" colab_type="code" outputId="dc118c6a-42a4-4a3c-e51a-e27758fc2b98" executionInfo={"status": "ok", "timestamp": 1583185380901, "user_tz": -60, "elapsed": 1905, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# ls
# + id="nlyf_r-8M75h" colab_type="code" outputId="caeff00f-dbc9-4adf-dff5-e6a93bc786d1" executionInfo={"status": "ok", "timestamp": 1583185742594, "user_tz": -60, "elapsed": 453, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd "dw_matrix2"
# + id="4IrGEj7YNEdk" colab_type="code" outputId="f2d17496-f3c1-4bda-807a-223a073bb39f" executionInfo={"status": "ok", "timestamp": 1583185746286, "user_tz": -60, "elapsed": 1558, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# ls
# + id="QGGBEKIqNG7U" colab_type="code" outputId="53200377-a971-415d-95fe-0990134e30a0" executionInfo={"status": "ok", "timestamp": 1583185543640, "user_tz": -60, "elapsed": 4263, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
# !curl -L http://bit.ly/dw_car_data -o car.h5
# + id="yzLRK9SQNjDc" colab_type="code" outputId="7db5729f-0d42-4a77-ab3c-94092ca0aa34" executionInfo={"status": "ok", "timestamp": 1583185721088, "user_tz": -60, "elapsed": 1978, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd
# + id="o2DhaYg-NwRq" colab_type="code" colab={}
import pandas as pd
# + id="cOVgYwFfN4YZ" colab_type="code" outputId="045c814a-d19a-47f7-facf-84e2a508d4a2" executionInfo={"status": "ok", "timestamp": 1583185770567, "user_tz": -60, "elapsed": 2009, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
df = pd.read_hdf("car.h5")
df.shape
# + id="al48hZK7N9ap" colab_type="code" outputId="e5f6bec2-8307-49de-87da-c149d12f3a07" executionInfo={"status": "ok", "timestamp": 1583185814247, "user_tz": -60, "elapsed": 445, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 593}
df.sample(5)
# + id="Ky2kxVS5OEGz" colab_type="code" outputId="93430910-264a-41ae-cc34-2aec447d38a5" executionInfo={"status": "ok", "timestamp": 1583187372913, "user_tz": -60, "elapsed": 489, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd dw_matrix2/
# + id="DyaGjbs_SmCL" colab_type="code" outputId="7a084e88-976f-4cd2-d6a0-fea64d1d0db0" executionInfo={"status": "ok", "timestamp": 1583187088688, "user_tz": -60, "elapsed": 1932, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "02416341188871661815"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# !git add day1_meta.ipynb
# + id="ZvqsmVDGYW_P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="a79b8e0c-8887-45ed-c9c9-577874b1f891" executionInfo={"status": "ok", "timestamp": 1583188532153, "user_tz": -60, "elapsed": 6851, "user": {"displayName": "kombin<NAME>", "photoUrl": "", "userId": "02416341188871661815"}}
# !git commit -m "Lets start the second stage"
# + [markdown] id="mybiFq1hXgtQ" colab_type="text"
#
| matrix_two/day1_meta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 1 Prediction Model: an application of Convolutional Neural Networks on Graphs
#
#
# This code was inspired by the [Graph Conv Net] library, whose algorithm was proposed in the [paper] *Convolutional Neural Networks on Graphs with Fast Localized Spectral Filtering*.
#
# [Graph Conv Net]: https://github.com/mdeff/cnn_graph,
# [paper]: https://arxiv.org/abs/1606.09375
# +
import os
import pandas as pd
pd.options.mode.chained_assignment = None
import numpy as np
import ast
from scipy import sparse
import seaborn as sns
import networkx as nx
import tensorflow as tf
import matplotlib
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
mycmap = ListedColormap(['blue','white','red'])
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore",category=matplotlib.cbook.mplDeprecation)
# %load_ext autoreload
# %autoreload
from lib import models_alt, graph
tf.logging.set_verbosity(tf.logging.ERROR)
# -
VOTES_PATH = '../data/votes/votes_{}.csv'
BILLS_PATH = '../data/bills/bills_{}.csv'
SEED = 123
np.random.seed(SEED)
# The CNN code as proposed in the reference was adapted for the regression problem, using Mean Squared Error as cost function, for an input tensor of and label tensor of dimension $N_{tr}\times N_{s}$, where $N_{tr}$ is the training set length and $N_{s}$ is the number of senators.
#
# Further details on the changes made can bee seen in `lib/models_alt.py`.
# +
params = dict()
params['dir_name'] = 'demo'
params['num_epochs'] = 200 # Number of training epochs
params['batch_size'] = 10 # Size of training batch
params['eval_frequency'] = 40 # Frequence of evaluation
# Building blocks.
params['filter'] = 'chebyshev5' # Type of filter
params['brelu'] = 'b1relu' # Type of activation function
params['pool'] = 'apool1' # Type of pooling
# Architecture.
params['F'] = [32, 64] # Number of graph convolutional filters.
params['p'] = [1, 1] # Pooling rate
params['K'] = [4, 4] # Polynomial degree
# Optimization.
params['regularization'] = 5e-4 # Regularization weight
params['dropout'] = 1 # 1- Dropout probability
params['decay_rate'] = 0.95 # Learning rate decay
params['momentum'] = 0 # Momentum
params['learning_rate'] = 0.1 # Learning rate
# +
def numeric_votes(vote_position):
'''
Converts vote positions to numeric values:
Yes -> 1
Not voting -> 0
No -> -1
'''
if vote_position == 'Yes':
return 1
elif vote_position == 'Not Voting' or pd.isnull(vote_position):
return 0
else:
return -1
def get_train_test(senate_id=115, ratio_train=0.5, ratio_val=0.1, shuffle=False, seed=123):
'''
Loads bills and votes and generates training, validation
and testing data. Inputs are cosponsor vote positions and
targets are the final vote outcomes.
'''
# Load votes and bills
VOTES_PATH = '../data/votes/votes_{}.csv'.format(senate_id)
BILLS_PATH = '../data/bills/bills_{}.csv'.format(senate_id)
bills = pd.read_csv(BILLS_PATH)
votes = pd.read_csv(VOTES_PATH)
# Keep only the bills sponsored by Senators
bills_sen = bills[bills.sponsor_title=='Sen.']
N_BILLS_SEN = bills_sen.shape[0]
# Get the members
members_aux = pd.DataFrame(list(votes['member_id'].unique())).rename(columns={0: 'member_id'})
members = members_aux.merge(votes.drop_duplicates(subset='member_id')[['member_id', 'party']], how='left').sort_values(['party', 'member_id'])
N_SENATORS = len(members)
for i,j in enumerate(bills_sen.cosponsors_sen):
bills_sen.cosponsors_sen.iloc[i] = ast.literal_eval(j)
# Create binary matrices indicating sponsorship
S = np.zeros((N_BILLS_SEN, N_SENATORS))
for i, j in enumerate(bills_sen.cosponsors_sen):
S[i, list(members['member_id']).index(bills_sen.sponsor_id.iloc[i])] = 1
# Create binary matrices indicating cosponsorship
CS = np.zeros((N_BILLS_SEN, N_SENATORS))
for i, j in enumerate(bills_sen.cosponsors_sen):
for k, l in enumerate(bills_sen.cosponsors_sen.iloc[i]):
CS[i, list(members['member_id']).index(bills_sen.cosponsors_sen.iloc[i][k])] = 1
# Sponsorship mask
Z = S + CS
# The target Y is the vote position of all senators for a given bill
Y = np.zeros((N_BILLS_SEN, N_SENATORS))
for i, l in enumerate(list(bills_sen.vote_uri)):
votes_i = (members.merge(votes[votes.vote_uri == l], how='left'))
feature = votes_i.vote_position.apply(numeric_votes).values
Y[i] = feature
# The input data X is the vote position of all (co)sponsors for a given bill
X = np.zeros((N_BILLS_SEN, N_SENATORS))
X[Z==1] = Y[Z==1]
n = X.shape[0]
d = X.shape[1]
n_train = int(n * ratio_train)
n_val = int(n * ratio_val)
if shuffle:
np.random.seed(seed)
perm_idx = np.random.permutation(n)
else:
perm_idx = np.arange(n)
X_train = X[perm_idx[:n_train]]
X_val = X[perm_idx[n_train:n_train+n_val]]
X_test = X[perm_idx[n_train+n_val:]]
y_train = Y[perm_idx[:n_train]]
y_val = Y[perm_idx[n_train:n_train+n_val]]
y_test = Y[perm_idx[n_train+n_val:]]
return X_train, X_val, X_test, y_train, y_val, y_test
def compute_adjacency(y, k=15):
'''
Computes an adjacency matrix from
'''
dist, idx = graph.distance_scipy_spatial(y.T, k=k, metric='euclidean')
A = graph.adjacency(dist, idx).astype(np.float32)
A = (A > 0.01).astype(np.float32)
return A
def compute_laplacian_list(A):
'''
Computes the normalized laplacian to use as input for the CNN.
'''
L = [graph.laplacian(A, normalized=True) for i in range(2)]
return L
def set_params_cnn(X_train, params):
'''
Sets parameters for the CNN.
'''
params['decay_steps'] = X_train.shape[0] / params['batch_size']
params['M'] = [X_train.shape[1]]
return params
def run_CNN(L, params, X_train, y_train, X_val, y_val, print_flag=False):
'''
Trains CNN model.
'''
model = models_alt.cgcnn(L, **params)
loss_train, loss_val, train_accs, val_accs, t_step = model.fit(X_train, y_train, X_val, y_val, print_flag)
return loss_train, loss_val, train_accs, val_accs, model
def rel_err(truth, preds):
'''
Calculates the relative error between the true labels and an estimate
Args:
truth: Ground truth signal
preds: Estimated signal
'''
voted_idxs = np.where(truth != 0)
return np.linalg.norm(truth[voted_idxs] - preds[voted_idxs], 2) / np.linalg.norm(truth[voted_idxs], 2)
def compute_acc(truth, preds):
'''
Computes the prediction accuracy.
Args:
truth: Ground truth signal
preds: Estimated signal
'''
valid_indices = np.where(np.any([truth==1, truth==-1], axis=0))
return (truth[valid_indices] == preds[valid_indices]).sum() / len(valid_indices)
def compute_set_accuracy(truth, preds):
'''
Computes the prediction accuracy for a set.
Args:
truth: Ground truth set
preds: Estimated set
'''
not_vot = np.not_equal(truth, 0)
a = np.equal(preds, truth)
return np.sum(a[not_vot])/np.size(truth[not_vot])
def predict_labels(X):
'''
Predicts labels for the input set X.
Args:
X: input dataset
'''
Pred = model.predict(X)
Pred = eval_prediction(Pred)
return Pred
def compute_prediction_err_acc(Truth, Pred):
'''
Computes the prediction error and accuracy for each bill.
Args:
Truth: Ground truth signal matrix
Preds: Estimated signal matrix
'''
err_v = []
acc_v = []
for i in range(Truth.shape[0]):
acc_v.append(compute_acc(Truth[i], Pred[i]))
err_v.append(rel_err(Truth[i], Pred[i]))
return err_v, acc_v
def eval_prediction(Pred):
'''
Binarize predictions into 1 and -1.
Args:
Pred: predicted labels set.
'''
Pred_bool = (Pred-Pred.mean(axis=1)[:,np.newaxis])>0
Pred[Pred_bool] = 1
Pred[~Pred_bool] = -1
return Pred
def plot_predictions(known_votes, true_votes, pred_votes, G, coords):
'''
Plot predicted labels over a network layout.
Args:
known_votes: training set.
true_votes: test set.
pred_votes: predicted labels.
G: graph.
coords: nodes coordinates in the layout.
'''
fig, ax = plt.subplots(ncols=3, figsize=(15,5))
nx.draw_networkx_nodes(G, coords, node_size=60, node_color=known_votes, cmap='bwr',
edgecolors='black', vmin=-1, vmax=1, ax=ax[0])
nx.draw_networkx_edges(G, coords, alpha=0.2, ax=ax[0])
ax[0].set_title('Known cosponsor votes')
nx.draw_networkx_nodes(G, coords, node_size=60, node_color=true_votes, cmap='bwr',
edgecolors='black', vmin=-1, vmax=1, ax=ax[1])
nx.draw_networkx_edges(G, coords, alpha=0.2, ax=ax[1])
ax[1].set_title('Final votes')
nx.draw_networkx_nodes(G, coords, node_size=60, node_color=pred_votes, cmap='bwr',
edgecolors='black', vmin=-1, vmax=1, ax=ax[2])
nx.draw_networkx_edges(G, coords, alpha=0.2, ax=ax[2])
ax[2].set_title('Predicted votes')
yes_patch = mpatches.Patch(label='Voted Yes', edgecolor='black', facecolor='red')
no_patch = mpatches.Patch(label='Voted No', edgecolor='black', facecolor='blue')
not_voted_patch = mpatches.Patch(label='Did not vote', edgecolor='black', facecolor='white')
fig.legend(handles=[yes_patch, no_patch, not_voted_patch], loc='lower center', ncol=3)
fig.subplots_adjust(bottom=0.12)
ax[0].tick_params(which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
ax[1].tick_params(which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
ax[2].tick_params(which='both', bottom=False, left=False, labelbottom=False, labelleft=False)
# -
# ### 2 Example: Congress 115
# In order to illustrate the use of the CNN applied to the senators graph, we consider the 115th Senate. As training input, half of the votes related to active bills are used, as well as graph (or its Laplacian) built from the training label vectors, i.e. from the actual votes outcome.
#
# In this case, the number of senators is $N_{s} = 105$ (5 of them replaced other during the term).
#
# The adjacency matrix is created from the similarity of votes between every two senators for the training set. This is done in function `compute_adjacency`, using the Euclidean distance measure.
X_train, X_val, X_test, y_train, y_val, y_test = get_train_test(senate_id = 115, shuffle = True)
A = compute_adjacency(y_train)
L = compute_laplacian_list(A)
params = set_params_cnn(X_train, params)
loss_train, loss_val, train_accs, val_accs, model = run_CNN(L, params, X_train, y_train, X_val, y_val, print_flag = False)
fig, ax1 = plt.subplots(figsize=(15, 5))
ax1.plot(loss_train, 'b.-')
ax1.set_ylabel('Loss', color='b')
ax1.plot(loss_val, 'b.:')
ax1.legend(['Train', 'Validation'])
ax2 = ax1.twinx()
ax2.plot(train_accs, 'r.-')
ax2.set_ylabel('Accuracy', color='r')
ax2.plot(val_accs, 'r.:')
ax2.legend(['Train', 'Validation'])
ax1.set_xlabel('Steps')
plt.show()
# We can see that the accuracy on the validation set reaches a score of $77$% by the end of the 200 training epochs.
#
# Now we perform the prediction over the test set, and illustrate it as:
# - Red: Vote yes;
# - Blue: Vote no;
# - White: Did not vote.
Pred_train = predict_labels(X_train)
Pred_test = predict_labels(X_test)
fig = plt.figure(figsize=(20,10))
plt.subplot(1,3,1)
plt.imshow(X_train, vmax = 1, vmin= -1, cmap = mycmap)
plt.xlabel('Senator #')
plt.ylabel('Bill-vote #')
plt.title('Training data')
plt.subplot(1,3,2)
plt.imshow(Pred_train, vmax = 1, vmin= -1, cmap = mycmap)
plt.xlabel('Senator #')
plt.ylabel('Bill-vote #')
plt.title('Training prediction')
plt.subplot(1,3,3)
plt.imshow(y_train, vmax = 1, vmin=-1, cmap = mycmap)
plt.xlabel('Senator #')
plt.ylabel('Bill-vote #')
plt.title('Training truth');
print('Accuracy train: {:.2f}%'.format(100*compute_set_accuracy(y_train, Pred_train)))
plt.figure(figsize=(20,10))
plt.subplot(1,3,1)
plt.imshow(X_test, vmax = 1, vmin= -1, cmap = mycmap)
plt.xlabel('Senator #')
plt.ylabel('Bill-vote #')
plt.title('Test data')
plt.subplot(1,3,2)
plt.imshow(Pred_test, vmax = 1, vmin= -1, cmap = mycmap)
plt.xlabel('Senator #')
plt.ylabel('Bill-vote #')
plt.title('Test prediction')
plt.subplot(1,3,3)
plt.imshow(y_test, vmax = 1, vmin= -1, cmap = mycmap)
plt.xlabel('Senator #')
plt.ylabel('Bill-vote #')
plt.title('Test truth');
print('Accuracy test: {:.2f} %'.format(100*compute_set_accuracy(y_test, Pred_test)))
# To better visualize the regression results, the following example is presented in network layout form, where one cosponsorship profile is selected among the validation set to emphasize the result predicted by the CNN and the respective groundtruth.
# +
G = nx.from_numpy_matrix(A.todense())
coords = nx.spring_layout(G)
Pred_val = predict_labels(X_val)
plot_predictions(X_val[2], y_val[2], Pred_val[2], G, coords)
# plt.savefig('figs/net_cnn.pdf', bbox_inches = 'tight')
# -
# Note that the CNN is quite successfull in identifying the final vote pattern using as input the initial cosponsorship profile and the graph structure, resulting in an accuracy of 100% (since we do not consider absences in the prediction evaluation).
# ## 3 Analysis of all congresses
# Next we extend the analysis to all US congresses, from Senate 105 to 115, where the same procedure detailed applied is for each of those Senates. The only exception is Senate 106, whose data provided by the ProPublica Congress API was inconsistent and faulty.
# +
Acc_tr = []
Acc_te = []
Err_tr = []
Err_te = []
for congress_nb in range(105, 116):
if congress_nb != 106:
X_train, X_val, X_test, y_train, y_val, y_test = get_train_test(senate_id = congress_nb, shuffle = True)
A = compute_adjacency(y_train)
L = compute_laplacian_list(A)
params = set_params_cnn(X_train, params)
loss_train, loss_val, train_accs, val_accs, model = run_CNN(L, params, X_train, y_train, X_val, y_val)
Pred_train = predict_labels(X_train)
Pred_test = predict_labels(X_test)
err_tr, acc_tr = compute_prediction_err_acc(y_train, Pred_train)
err_te, acc_te = compute_prediction_err_acc(y_test, Pred_test)
Err_tr.append(err_tr)
Err_te.append(err_te)
Acc_tr.append(acc_tr)
Acc_te.append(acc_te)
# -
# Build dataframe with error and accuracy statistics for train and test sets.
c=0
df=pd.DataFrame()
for i in range(105, 116):
if i != 106:
df = df.append(pd.DataFrame(data=[Err_tr[c], Acc_tr[c], Err_te[c], Acc_te[c], [i for j in range(len(Err_tr[c]))]]).transpose())
c+=1
df.columns = ['Err_tr', 'Acc_tr', 'Err_te', 'Acc_te', 'Congress']
# Next we show the distribution of error and accuracy for the test set for each Senate.
# +
fig, ax= plt.subplots(2,1, figsize=(8,6))
sns.set(style="whitegrid")
sns.boxplot(x='Congress',y='Err_te', data=df, palette="vlag", ax=ax[0])
sns.swarmplot(x='Congress',y='Err_te', data=df, size=2, color=".3", ax=ax[0])
sns.boxplot(x='Congress',y='Acc_te', data=df, palette="vlag", ax=ax[1])
sns.swarmplot(x='Congress',y='Acc_te', data=df, size=2, color=".3", ax=ax[1])
ax[0].set_ylabel('Test Error')
ax[1].set_ylabel('Test Accuracy (%)')
ax[0].set_xticks([])
ax[0].set_xlabel('')
plt.xlabel('Senate')
plt.xticks(np.arange(10),[i for i in range(105,116) if i !=106]);
plt.savefig('figs/test_err_cnn.pdf')
# -
# We can see from the image above that for later Senates, the prediction accuracy tends to increase. This can be due to the fact that as the training data gets scarcer for the older Senates, the amount of information available may be insufficient to correctly train all parameters in the CNN.
# The mean training and test errors and accuracy scores, as well as their standard deviations, can be seen below:
df.groupby('Congress').mean()
df.groupby('Congress').std()
| notebooks/cnn-congress-all-cosponsors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="aRK1drwdlJqv"
import tensorflow as tf
import matplotlib.image as img
# %matplotlib inline
import numpy as np
from collections import defaultdict
import collections
from shutil import copy
from shutil import copytree, rmtree
import tensorflow.keras.backend as K
from tensorflow.keras.models import load_model
from tensorflow.keras.preprocessing import image
import matplotlib.pyplot as plt
import numpy as np
import os
import random
import tensorflow as tf
import tensorflow.keras.backend as K
from tensorflow.keras import regularizers
from tensorflow.keras.applications.inception_v3 import InceptionV3
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.layers import Dense, Dropout, Activation, Flatten
from tensorflow.keras.layers import Convolution2D, MaxPooling2D, ZeroPadding2D, GlobalAveragePooling2D, AveragePooling2D
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.callbacks import ModelCheckpoint, CSVLogger
from tensorflow.keras.optimizers import SGD
from tensorflow.keras.regularizers import l2
from tensorflow import keras
from tensorflow.keras import models
import cv2
# + [markdown] id="1e24_39ecVsf"
# Dataset is stored on shared google drive.
# + colab={"base_uri": "https://localhost:8080/"} id="BCszFLCtcVsk" outputId="47d81335-abd2-488b-b7de-f0eb181fb7eb"
from google.colab import drive
drive.mount("/content/drive")
# + [markdown] id="jZjXuSQacVsn"
# Create .txt files with image filenames. Separate file for original and fake images in train and test datasets.
# + id="WWZVxY6mcVsn"
# !ls /content/drive/Shareddrives/datasets/animals/data/train/original/*.jpg > /content/drive/Shareddrives/datasets/animals/originals_train.txt
# !ls /content/drive/Shareddrives/datasets/animals/data/train/fake/*.jpg > /content/drive/Shareddrives/datasets/animals/fakes_train.txt
# !ls /content/drive/Shareddrives/datasets/animals/data/test/original/*.jpg > /content/drive/Shareddrives/datasets/animals/originals_test.txt
# !ls /content/drive/Shareddrives/datasets/animals/data/test/fake/*.jpg > /content/drive/Shareddrives/datasets/animals/fakes_test.txt
path = "/content/drive/Shareddrives/datasets/animals/"
original_train_filenames_path = path + "originals_train.txt"
fake_train_filenames_path = path + "fakes_train.txt"
original_test_filenames_path = path + "originals_test.txt"
fake_test_filenames_path = path + "fakes_test.txt"
# + [markdown] id="hrM428lIcVso"
# Paths to folders that include images. Current structure (after copying from previous notebook):
# data/
# -train/
# -original/
# -fake/
# -test/
# -original/
# -fake/
# + id="qQKTHXsNcVsq"
original_train_path = path + "data/train/original"
fake_train_path = path + "data/train/fake"
original_test_path = path + "data/test/original"
fake_test_path = path + "data/test/fake"
# + [markdown] id="4UG1O1p-cVsr"
# Function that creates array of filenames
# + id="i9INiDzYcVsr"
def get_array_of_filenames(original_train_filenames_path, original_test_filenames_path, fake_train_filenames_path, fake_test_filenames_path, main_path):
ori_train_paths = []
fake_train_paths = []
ori_test_paths = []
fake_test_paths = []
with open(original_train_filenames_path, 'r') as file:
ori_train_paths = [main_path + read.strip() for read in file.readlines()]
with open(fake_train_filenames_path, 'r') as file:
fake_train_paths = [main_path + read.strip() for read in file.readlines()]
with open(original_test_filenames_path, 'r') as file:
ori_test_paths = [main_path + read.strip() for read in file.readlines()]
with open(fake_test_filenames_path, 'r') as file:
fake_test_paths = [main_path + read.strip() for read in file.readlines()]
return ori_train_paths, fake_train_paths, ori_test_paths, fake_test_paths
# + id="4orLkB1NcVst"
ori_train, fakes_train, ori_test, fakes_test = get_array_of_filenames(original_train_filenames_path, fake_train_filenames_path, original_test_filenames_path, fake_test_filenames_path, "")
# + [markdown] id="5HN-NbcCcVsu"
#
# + id="gFwDbiuUcVsu"
def transformation_to_magnitude(image):
image = np.mean(image, axis=2)
dft = np.fft.fft2(image)
dft_shift = np.fft.fftshift(dft)
magnitude = np.abs(dft_shift)
magnitude = magnitude/np.max(magnitude)
result = np.zeros((magnitude.shape[0], magnitude.shape[1],3))
result[:,:,0] = magnitude
result[:,:,1] = magnitude
result[:,:,2] = magnitude
return result
# + id="i0gFOZV3nBG1"
def transformation_to_phase(image):
image = np.mean(image, axis=2)
dft = np.fft.fft2(image)
dft_shift = np.fft.fftshift(dft)
angle = np.angle(dft_shift)
angle = (angle+np.pi)/(2*np.pi)
result = np.zeros((angle.shape[0], angle.shape[1],3))
result[:,:,0] = angle
result[:,:,1] = angle
result[:,:,2] = angle
return result
# + id="qwab2bTwcVsw"
def transformation_to_everything(image):
image = np.mean(image, axis=2)
dft = np.fft.fft2(image)
dft_shift = np.fft.fftshift(dft)
magnitude = np.abs(dft_shift)
magnitude = magnitude/np.max(magnitude)
angle = np.angle(dft_shift)
angle = (angle+np.pi)/(2*np.pi)
result = np.zeros((magnitude.shape[0], magnitude.shape[1],3))
result[:,:,0] = image
result[:,:,1] = angle
result[:,:,2] = magnitude
return result
# + [markdown] id="BDB6jSEecVsy"
# Main analisys
# + colab={"base_uri": "https://localhost:8080/", "height": 575} id="T0JCjvZUcVsz" outputId="a4d34bca-3f9b-4c42-b63e-75e45db18193"
K.clear_session()
n_classes = 2
img_width, img_height = 512, 512
train_data_dir = '/content/drive/Shareddrives/datasets/animals/data/train'
validation_data_dir = '/content/drive/Shareddrives/datasets/animals/data/test'
nb_train_samples = 1414 #75750
nb_validation_samples = 142 #25250
batch_size = 16
transformation_function = transformation_to_phase
train_datagen = ImageDataGenerator(
rescale=1. / 255)
#preprocessing_function=transformation_function)
test_datagen = ImageDataGenerator(
rescale=1. / 255)
#preprocessing_function=transformation_function)
train_generator = train_datagen.flow_from_directory(
train_data_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='categorical')
validation_generator = test_datagen.flow_from_directory(
validation_data_dir,
target_size=(img_height, img_width),
batch_size=batch_size,
class_mode='categorical')
inception = InceptionV3(weights="imagenet", include_top=False, classes=2)
x = inception.output
x = GlobalAveragePooling2D()(x)
x = Dense(128,activation='relu')(x)
x = Dropout(0.2)(x)
predictions = Dense(2,kernel_regularizer=regularizers.l2(0.005), activation='softmax')(x)
model = Model(inputs=inception.input, outputs=predictions)
model.compile(optimizer=SGD(lr=0.0001, momentum=0.9), loss='categorical_crossentropy', metrics=['accuracy'])
checkpointer = ModelCheckpoint(filepath='/content/drive/Shareddrives/datasets/results/best_model_animals_batch_16_imagenet.hdf5', verbose=1, save_best_only=True)
csv_logger = CSVLogger('/content/drive/Shareddrives/datasets/results/history_animals_batch_16_imagenet.log')
history = model.fit_generator(train_generator,
steps_per_epoch = nb_train_samples // batch_size,
validation_data=validation_generator,
validation_steps=nb_validation_samples // batch_size,
epochs=30,
verbose=1,
callbacks=[csv_logger, checkpointer])
model.save('/content/drive/Shareddrives/datasets/results/model_trained_animals_batch_16_imagenet.hdf5')
# + [markdown] id="ZMkkRb6jcVsz"
# How to use model?
# First load model:
# ```
# trained_model = models.load_model('path to model')
# ```
# Use this function to load image:
#
# ```
# def load_file(path):
# img = cv2.imread(path)
# img = img.astype('float64')
# img = img/255.0
# img = img.reshape(-1, 512, 512, 3)
# return img
# ```
#
#
#
# Execute predict() method and print results:
# ```
# print(trained_model.predict(load_file(addr2)))
# ```
# Have fun :)
#
| notebooks/AO_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # sbpy.photometry: Disk-integrated phase function
#
# [sbpy.photometry](https://sbpy.readthedocs.io/en/latest/sbpy/photometry.html) defines the classes and functions to implement disk-integrated phase function models for (atmosphereless) solar system objects. The models currently implemented include a linear phase function model, IAU HG model, HG$_1$G$_2$ model, and HG$_{12}$ model.
# ## Disk-integrated phase function
#
# Disk-integrated phase function can be defined by the magnitude or by the average bidirectional reflectance over the whole cross-section of the object with respect to the observer (both illuminated and unilluminated), as a function of solar phase angle.
#
# The magnitude of an object is,
#
# $M(\alpha, r_h, \Delta) = \Phi(\alpha) + 5\log_{10}(r_h) + 5\log_{10}(\Delta)$,
#
# where $\alpha$ is phase angle, $r_h$ is heliocentric distance, and $\Delta$ is the distance to observer. $\Phi(\alpha)$ is the disk-integrated phase function defined in magnitude, or equivalently the reduced magnitude of an object (the magnitude at $r_h=\Delta=1 au$.
#
# Alternatively, the total flux of an object can be expressed by its average bidirectional reflectance over the whole cross-sectional area,
#
# $f(\alpha, r_h, \Delta) = \frac{\bar r(\alpha) \pi R^2 f_{Sun}}{r^2\Delta^2}$
#
# where $\bar r(\alpha)$ is the disk-integrated phaes function defined by the average bidirectional reflectance, and $R$ is the radius of an object such that $\pi R^2$ is its cross-sectional area. Note that the phase function $\bar r(\alpha)$ can be normalized to unity at opposition as $\bar r_n(\alpha)$ such that $\bar r_n(0)=1$. In this case, the geometric albedo of the object $p=\pi \bar r_n(0)$.
#
# The Bond albedo is expressed as,
#
# $A_B = p q$,
#
# where $q$ is phase integral,
#
# $q = 2\int_0^\pi \sin(\alpha) \bar r_n(\alpha)$
#
# ### Conversion between magnitude and average bidirectional reflectance
#
# The conversion between reduced magnitude $M(\alpha, 1, 1)$ and average bidirectional reflectance $\bar r(\alpha)$ is provided by functions `photometry.mag2ref` and `photometry.ref2mag`:
# +
import numpy as np
import astropy.units as u
from sbpy.photometry import ref2mag, mag2ref
# phys = Phys.from_dict({'r':2*u.km})
# for Ceres
radius = 480 * u.km # radius
mag = 3.34 # V-band absolute magnitude
rbar = mag2ref(mag, radius)
print('geometric albedo of Ceres is {0:.4f}'.format(np.pi*u.sr*rbar))
radius = 262 * u.km # radius
rbar = 0.38/np.pi # V-band geometric albedo 0.38
mag = ref2mag(rbar, radius)
print('absolute magnitude of Vesta is {0:.2f}'.format(mag))
# -
# ## Linear phase function model
#
# In this model, the total magnitude of an object is defined by its absolute magnitude $H = M(0, 1, 1)$, and a phase slope $-S$ in magnitude per unit phase angle change.
# +
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib notebook
from sbpy.photometry import LinearPhaseFunc
# Linear phase function with H = 5 and S = 0.04 mag/deg = 2.29 mag/rad
linear_phasefunc = LinearPhaseFunc(5, 2.29, radius=300)
pha = np.linspace(0, np.pi*0.75, 200)
print('Geometric albedo is {0:.3}'.format(linear_phasefunc.geoalb))
print('Bond albedo is {0:.3}'.format(linear_phasefunc.bondalb))
print('Phase integral is {0:.3}'.format(linear_phasefunc.phaseint))
# plot linear phase function
f, ax = plt.subplots(2, 1, sharex=True)
ax[0].plot(np.rad2deg(pha), linear_phasefunc.mag(pha))
plt.setp(ax[0], ylabel='Magnitude', ylim=(11, 4))
ax[1].plot(np.rad2deg(pha), linear_phasefunc.ref(pha))
plt.setp(ax[1], xlabel='Phase Angle (deg)', ylabel=r'$\bar r$')
plt.tight_layout()
# -
# ## IAU HG model
#
# IAU HG model ([Bowell et al. 1989](https://ui.adsabs.harvard.edu/#abs/1989aste.conf..524B/abstract)) describes the disk-integrated phase function of small bodies with two parameters, $H$, and $G$. The $H$ parameter is the absolute magnitude, and the $G$ parameter describe the slope of the phase function. Class `photometry.HG` implements this model.
#
# [Muinonen et al. (2010)](https://ui.adsabs.harvard.edu/#abs/2010Icar..209..542M/abstract) proposed a 3-parameter model that better describes the disk-integrated phase function of asteroids at phase angles up to 150$^\circ$. This model was subsequently adopted by the IAU as the new standard asteroid phase function model. In this model, parameter $H$ is the same as that in the HG model, and parameters $G_1$ and $G_2$ describe the slope of phase function. Class `photometry.HG1G2` implements the 3-parameter HG$_1$G$_2$ model.
#
# Based on the three-parameter HG$_1$G$_2$ model, [Muinonen et al. (2010)](https://ui.adsabs.harvard.edu/#abs/2010Icar..209..542M/abstract) also derived a two-parameter model, in which the phase slope parameter $G_{12}$ is a combination of G$_1$ and G$_2$ parameters that describe the phase slope. Class `photometry.HG12` implements this 2-parameter HG$_{12}$ model.
# +
import numpy as np
from astropy.table import Table
from sbpy.photometry import HG, HG1G2, HG12
# phase function models for (24) Themis
themis_phase = []
# HG model: H=7.08, G=0.19 (JPL Small Bodies Database)
themis_phase.append(HG(7.08, 0.19, radius=99))
# HG1G2 model: H=7.121, G1=0.67, G2=0.14 (Muinonen et al. 2010)
themis_phase.append(HG1G2(7.121, 0.67, 0.14, radius=99))
# HG12 model: H=7.121, G12=0.68 (Muinonen et al. 2010)
themis_phase.append(HG12(7.121, 0.68, radius=99))
# photometric properties of Themis based on all three models
geoalb = [m.geoalb for m in themis_phase] # geometric albedo
bondalb = [m.bondalb for m in themis_phase] # bond albedo
phaseint = [m.phaseint for m in themis_phase] # phase integral
oeamp = [m.oe_amp for m in themis_phase[1:]] # opposition-effect amplitude
oeamp.insert(0,None)
coeff = [m.phasecoeff for m in themis_phase[1:]] # phase coefficient (slope)
coeff.insert(0,None)
# table of properties
model_names = ['HG', 'HG1G2','HG12']
phopars = Table([model_names, geoalb, bondalb, phaseint, oeamp, coeff],
names=['Model', 'Geometric Albedo', 'Bond Albedo', 'Phase Integral',
'OE Amplitude', 'Phase Coeff'], masked=True)
phopars['OE Amplitude'].mask[0] = True
phopars['Phase Coeff'].mask[0] = True
for k in phopars.keys()[1:]:
phopars[k].format='%.4f'
phopars.show_in_notebook()
# +
from matplotlib import pyplot as plt
# %matplotlib notebook
import numpy as np
# plot phase function models for Themis constructed above
pha = np.linspace(0, np.pi*0.75, 200)
f, ax = plt.subplots(3, 1, sharex=True, figsize=(6,8))
sym = ['-','.','--']
for m, s in zip(themis_phase,sym):
ax[0].plot(np.rad2deg(pha), m.mag(pha), s)
ax[1].plot(np.rad2deg(pha), m.ref(pha), s)
ax[2].plot(np.rad2deg(pha), m.ref(pha, normalized=0), s)
plt.setp(ax[0], ylabel='Magnitude',ylim=[14,6])
plt.setp(ax[1], ylabel=r'$\bar r$')
plt.setp(ax[2], xlabel='Phase Angle (deg)', ylabel=r'$\bar r$ Normalized')
ax[2].legend(model_names)
plt.tight_layout()
# -
# ## Plot the predicted magnitude of (24) Themis based on various models
#
# In this example, we used the three different phase function model constructed above to calculate the predicted magnitude of Themis in year 2018, and compare with the prediction by JPL Horizons. The emphemerides of Themis is generated by JPL Horizons using `data.Ephem` class.
# +
from astropy.time import Time
from matplotlib import pyplot as plt
from sbpy.data import Ephem
# ephemerides of Themis
epochs = {'start':'2018-01-01', 'stop': '2018-12-31', 'step': '1d'}
eph = Ephem.from_horizons('themis', epochs=epochs)
# set up figure
fig = plt.figure()
fig.clear()
ax = fig.gca()
# plot JPL magnitude
ts = Time(eph['datetime_jd'], format='jd')
ax.plot_date(ts.plot_date, eph['V'], 'v', mfc='w')
# plot calculated magnitude
for m in themis_phase:
ax.plot_date(ts.plot_date, m.mag(eph), '-')
ax.legend(['JPL']+model_names)
plt.setp(ax, ylabel='Magnitude', ylim=(13.5,11))
# -
# ## Define a double-exponential phase function model
#
# In this example, we show how to define a phase function model with `photometry.DiskIntegratedModelClass`. This model is defined in average bidirectional reflectance with double-exponential function,
#
# $\bar r(\alpha) = \frac{p}{\pi} \frac{1}{1+c} [c\exp(-\alpha/a_1)+\exp(-\alpha/a_2)]$
#
# This model is defined by geometric albedo $p$, parameters $c$ and $a_1$ that characterize the opposition amplitude and width, respectively, and parameter $a_2$ that characterize the phase slope.
# +
import numpy as np
from astropy.modeling import Parameter
from matplotlib import pyplot as plt
# %matplotlib notebook
from sbpy.photometry import DiskIntegratedModelClass
from sbpy.data import Ephem
# model definition
class DoubleExpPhaseFunc(DiskIntegratedModelClass):
_unit = 'ref'
p = Parameter(description='Geometric albedo')
c = Parameter(description='Opposition effect amplitude')
a1 = Parameter(description='Opposition effect width')
a2 = Parameter(description='Phase slope')
@staticmethod
def evaluate(a, p, c, a1, a2):
return p / (np.pi * (1+c)) * (c*np.exp(-a/a1) + np.exp(-a/a2))
# define a double-exponential phase function and print the basic photometric properties
phasefunc = DoubleExpPhaseFunc(0.1, 0.3, 0.02, 0.5, radius=100)
print('geometric albedo = {0:.3f}'.format(phasefunc.geoalb))
print('bond albedo = {0:.3f}'.format(phasefunc.bondalb))
print('phase integral = {0:.3f}'.format(phasefunc.phaseint))
# plot phase function model
pha = np.linspace(0, np.pi*0.75, 200)
fig = plt.figure()
ax = fig.gca()
ax.plot(np.rad2deg(pha), phasefunc.ref(pha))
plt.setp(ax, xlabel='Phase Angle (deg)', ylabel=r'$\bar r$')
# assume this model for (24) Themis, plot its predicted magnitude for year 2018
epochs = {'start': '2018-01-01', 'stop': '2018-12-31', 'step': '1d'}
eph = Ephem.from_horizons('themis', epochs=epochs)
ts = Time(eph['datetime_jd'], format='jd')
fig = plt.figure()
fig.clear()
ax = fig.gca()
ax.plot_date(ts.plot_date, phasefunc.mag(eph), '-')
plt.setp(ax, ylabel='Mag', ylim=(13,10.5))
| notebooks/photometry/integrated_phasefunc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"".isspace()
" ".isspace()
" fdsf".isspace()
"sdfsd ".isspace()
'sdf'*3
"this should be capitalized each letter".title()
"First string"+" "+"Second string"
sen="i want ot partition right at this point as well okay?"
sen.partition(" point ")
car="mercedes"
truck="mercedes"
print(id(car))
print(id(truck))
#removing vowels
strng="i want to remove all the vowels from this string"
vowels={'a','e','i','o','u'}
''.join([k for k in strng if k not in vowels])
| string questions 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# # Iris flower classification with scikit-learn (run model explainer locally)
# 
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# Explain a model with the AML explain-model package
#
# 1. Train a SVM classification model using Scikit-learn
# 2. Run 'explain_model' with full data in local mode, which doesn't contact any Azure services
# 3. Run 'explain_model' with summarized data in local mode, which doesn't contact any Azure services
# 4. Visualize the global and local explanations with the visualization dashboard.
from sklearn.datasets import load_iris
from sklearn import svm
from azureml.explain.model.tabular_explainer import TabularExplainer
# # 1. Run model explainer locally with full data
# ## Load the breast cancer diagnosis data
iris = load_iris()
X = iris['data']
y = iris['target']
classes = iris['target_names']
feature_names = iris['feature_names']
# Split data into train and test
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# ## Train a SVM classification model, which you want to explain
clf = svm.SVC(gamma=0.001, C=100., probability=True)
model = clf.fit(x_train, y_train)
# ## Explain predictions on your local machine
tabular_explainer = TabularExplainer(model, x_train, features = feature_names, classes=classes)
# ## Explain overall model predictions (global explanation)
global_explanation = tabular_explainer.explain_global(x_test)
# Sorted SHAP values
print('ranked global importance values: {}'.format(global_explanation.get_ranked_global_values()))
# Corresponding feature names
print('ranked global importance names: {}'.format(global_explanation.get_ranked_global_names()))
# feature ranks (based on original order of features)
print('global importance rank: {}'.format(global_explanation.global_importance_rank))
# per class feature names
print('ranked per class feature names: {}'.format(global_explanation.get_ranked_per_class_names()))
# per class feature importance values
print('ranked per class feature values: {}'.format(global_explanation.get_ranked_per_class_values()))
dict(zip(global_explanation.get_ranked_global_names(), global_explanation.get_ranked_global_values()))
# ## Explain overall model predictions as a collection of local (instance-level) explanations
# feature shap values for all features and all data points in the training data
print('local importance values: {}'.format(global_explanation.local_importance_values))
# ## Explain local data points (individual instances)
# explain the first member of the test set
instance_num = 0
local_explanation = tabular_explainer.explain_local(x_test[instance_num,:])
# +
# get the prediction for the first member of the test set and explain why model made that prediction
prediction_value = clf.predict(x_test)[instance_num]
sorted_local_importance_values = local_explanation.get_ranked_local_values()[prediction_value]
sorted_local_importance_names = local_explanation.get_ranked_local_names()[prediction_value]
dict(zip(sorted_local_importance_names, sorted_local_importance_values))
# -
# ## Load visualization dashboard
# Note you will need to have extensions enabled prior to jupyter kernel starting
# !jupyter nbextension install --py --sys-prefix azureml.contrib.explain.model.visualize
# !jupyter nbextension enable --py --sys-prefix azureml.contrib.explain.model.visualize
# Or, in Jupyter Labs, uncomment below
# jupyter labextension install @jupyter-widgets/jupyterlab-manager
# jupyter labextension install microsoft-mli-widget
from azureml.contrib.explain.model.visualize import ExplanationDashboard
ExplanationDashboard(global_explanation, model, x_test)
| how-to-use-azureml/explain-model/explain-tabular-data-local/explain-local-sklearn-multiclass-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: U4-S2-NNF
# language: python
# name: u4-s2-nnf
# ---
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 4 Sprint 3 Assignment 1*
#
# # Recurrent Neural Networks and Long Short Term Memory (LSTM)
#
# 
#
# It is said that [infinite monkeys typing for an infinite amount of time](https://en.wikipedia.org/wiki/Infinite_monkey_theorem) will eventually type, among other things, the complete works of Wiliam Shakespeare. Let's see if we can get there a bit faster, with the power of Recurrent Neural Networks and LSTM.
#
# This text file contains the complete works of Shakespeare: https://www.gutenberg.org/files/100/100-0.txt
#
# Use it as training data for an RNN - you can keep it simple and train character level, and that is suggested as an initial approach.
#
# Then, use that trained RNN to generate Shakespearean-ish text. Your goal - a function that can take, as an argument, the size of text (e.g. number of characters or lines) to generate, and returns generated text of that size.
#
# Note - Shakespeare wrote an awful lot. It's OK, especially initially, to sample/use smaller data and parameters, so you can have a tighter feedback loop when you're trying to get things running. Then, once you've got a proof of concept - start pushing it more!
# + colab={} colab_type="code" id="Ltj1je1fp5rO"
import numpy as np
from tensorflow import keras
from tensorflow.keras.utils import get_file
url = "https://www.gutenberg.org/files/100/100-0.txt"
path = get_file("shakespeare.txt", url)
with open(path, encoding="utf8") as f:
big_text = f.read()
# -
len(big_text[10700:20810])
# +
text = big_text[10700:20810]
chars = list(set(text))
char_int = {c:i for i,c in enumerate(chars)}
int_char = {i:c for i,c in enumerate(chars)}
# -
chars
# + [markdown] colab_type="text" id="zE4a4O7Bp5x1"
# # Resources and Stretch Goals
# + [markdown] colab_type="text" id="uT3UV3gap9H6"
# ## Stretch goals:
# - Refine the training and generation of text to be able to ask for different genres/styles of Shakespearean text (e.g. plays versus sonnets)
# - Train a classification model that takes text and returns which work of Shakespeare it is most likely to be from
# - Make it more performant! Many possible routes here - lean on Keras, optimize the code, and/or use more resources (AWS, etc.)
# - Revisit the news example from class, and improve it - use categories or tags to refine the model/generation, or train a news classifier
# - Run on bigger, better data
#
# ## Resources:
# - [The Unreasonable Effectiveness of Recurrent Neural Networks](https://karpathy.github.io/2015/05/21/rnn-effectiveness/) - a seminal writeup demonstrating a simple but effective character-level NLP RNN
# - [Simple NumPy implementation of RNN](https://github.com/JY-Yoon/RNN-Implementation-using-NumPy/blob/master/RNN%20Implementation%20using%20NumPy.ipynb) - Python 3 version of the code from "Unreasonable Effectiveness"
# - [TensorFlow RNN Tutorial](https://github.com/tensorflow/models/tree/master/tutorials/rnn) - code for training a RNN on the Penn Tree Bank language dataset
# - [4 part tutorial on RNN](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/) - relates RNN to the vanishing gradient problem, and provides example implementation
# - [RNN training tips and tricks](https://github.com/karpathy/char-rnn#tips-and-tricks) - some rules of thumb for parameterizing and training your RNN
| module1-rnn-and-lstm/LS_DS_431_RNN_and_LSTM_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# formats: md,ipynb
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--TITLE:Data Augmentation-->
#
# # Introduction #
#
# Now that you've learned the fundamentals of convolutional classifiers, you're ready to move on to more advanced topics.
#
# In this lesson, you'll learn a trick that can give a boost to your image classifiers: it's called **data augmentation**.
# # The Usefulness of Fake Data #
#
# The best way to improve the performance of a machine learning model is to train it on more data. The more examples the model has to learn from, the better it will be able to recognize which differences in images matter and which do not. More data helps the model to *generalize* better.
#
# One easy way of getting more data is to use the data you already have. If we can transform the images in our dataset in ways that preserve the class, we can teach our classifier to ignore those kinds of transformations. For instance, whether a car is facing left or right in a photo doesn't change the fact that it is a *Car* and not a *Truck*. So, if we **augment** our training data with flipped images, our classifier will learn that "left or right" is a difference it should ignore.
#
# And that's the whole idea behind data augmentation: add in some extra fake data that looks reasonably like the real data and your classifier will improve.
#
# # Using Data Augmentation #
#
# Typically, many kinds of transformation are used when augmenting a dataset. These might include rotating the image, adjusting the color or contrast, warping the image, or many other things, usually applied in combination. Here is a sample of the different ways a single image might be transformed.
#
# <figure>
# <img src="https://i.imgur.com/UaOm0ms.png" width=400, alt="Sixteen transformations of a single image of a car.">
# </figure>
#
# Data augmentation is usually done *online*, meaning, as the images are being fed into the network for training. Recall that training is usually done on mini-batches of data. This is what a batch of 16 images might look like when data augmentation is used.
#
# <figure>
# <img src="https://i.imgur.com/MFviYoE.png" width=400, alt="A batch of 16 images with various random transformations applied.">
# </figure>
#
# Each time an image is used during training, a new random transformation is applied. This way, the model is always seeing something a little different than what it's seen before. This extra variance in the training data is what helps the model on new data.
#
# It's important to remember though that not every transformation will be useful on a given problem. Most importantly, whatever transformations you use should not mix up the classes. If you were training a [digit recognizer](https://www.kaggle.com/c/digit-recognizer), for instance, rotating images would mix up '9's and '6's. In the end, the best approach for finding good augmentations is the same as with most ML problems: try it and see!
#
# # Example - Training with Data Augmentation #
#
# Keras lets you augment your data in two ways. The first way is to include it in the data pipeline with a function like [`ImageDataGenerator`](https://www.tensorflow.org/api_docs/python/tf/keras/preprocessing/image/ImageDataGenerator). The second way is to include it in the model definition by using Keras's **preprocessing layers**. This is the approach that we'll take. The primary advantage for us is that the image transformations will be computed on the GPU instead of the CPU, potentially speeding up training.
#
# In this exercise, we'll learn how to improve the classifier from Lesson 1 through data augmentation. This next hidden cell sets up the data pipeline.
# +
#$HIDE_INPUT$
# Imports
import os, warnings
import matplotlib.pyplot as plt
from matplotlib import gridspec
import numpy as np
import tensorflow as tf
from tensorflow.keras.preprocessing import image_dataset_from_directory
# Reproducability
def set_seed(seed=31415):
np.random.seed(seed)
tf.random.set_seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
os.environ['TF_DETERMINISTIC_OPS'] = '1'
set_seed()
# Set Matplotlib defaults
plt.rc('figure', autolayout=True)
plt.rc('axes', labelweight='bold', labelsize='large',
titleweight='bold', titlesize=18, titlepad=10)
plt.rc('image', cmap='magma')
warnings.filterwarnings("ignore") # to clean up output cells
# Load training and validation sets
ds_train_ = image_dataset_from_directory(
'../input/car-or-truck/train',
labels='inferred',
label_mode='binary',
image_size=[128, 128],
interpolation='nearest',
batch_size=64,
shuffle=True,
)
ds_valid_ = image_dataset_from_directory(
'../input/car-or-truck/valid',
labels='inferred',
label_mode='binary',
image_size=[128, 128],
interpolation='nearest',
batch_size=64,
shuffle=False,
)
# Data Pipeline
def convert_to_float(image, label):
image = tf.image.convert_image_dtype(image, dtype=tf.float32)
return image, label
AUTOTUNE = tf.data.experimental.AUTOTUNE
ds_train = (
ds_train_
.map(convert_to_float)
.cache()
.prefetch(buffer_size=AUTOTUNE)
)
ds_valid = (
ds_valid_
.map(convert_to_float)
.cache()
.prefetch(buffer_size=AUTOTUNE)
)
# -
# ## Step 2 - Define Model ##
#
# To illustrate the effect of augmentation, we'll just add a couple of simple transformations to the model from Tutorial 1.
# +
from tensorflow import keras
from tensorflow.keras import layers
# these are a new feature in TF 2.2
from tensorflow.keras.layers.experimental import preprocessing
pretrained_base = tf.keras.models.load_model(
'../input/cv-course-models/cv-course-models/vgg16-pretrained-base',
)
pretrained_base.trainable = False
model = keras.Sequential([
# Preprocessing
preprocessing.RandomFlip('horizontal'), # flip left-to-right
preprocessing.RandomContrast(0.5), # contrast change by up to 50%
# Base
pretrained_base,
# Head
layers.Flatten(),
layers.Dense(6, activation='relu'),
layers.Dense(1, activation='sigmoid'),
])
# -
# ## Step 3 - Train and Evaluate ##
#
# And now we'll start the training!
# +
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['binary_accuracy'],
)
history = model.fit(
ds_train,
validation_data=ds_valid,
epochs=30,
verbose=0,
)
# +
import pandas as pd
history_frame = pd.DataFrame(history.history)
history_frame.loc[:, ['loss', 'val_loss']].plot()
history_frame.loc[:, ['binary_accuracy', 'val_binary_accuracy']].plot();
# -
# The training and validation curves in the model from Tutorial 1 diverged fairly quickly, suggesting that it could benefit from some regularization. The learning curves for this model were able to stay closer together, and we achieved some modest improvement in validation loss and accuracy. This suggests that the dataset did indeed benefit from the augmentation.
#
# # Your Turn #
#
# Move on to the [**Exercise**](#$NEXT_NOTEBOOK_URL$) to apply data augmentation to the custom convnet you built in Lesson 5. This will be your best model ever!
| notebooks/computer_vision/raw/tut6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3.7
# language: python
# name: py3.7
# ---
# # "Augmentations: Cutout, Mixup, Cutmix, Label Smoothing"
# > "Visual summary of modern augmentations technique"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [augmentations]
# Data augmentation is one of the major techniques used when we want to improve the model's performance on new data (i.e. generalization). It is especially useful in cases where only limited training data are available. In this post we'll discuss some of the augmentation techniques (besides affine transformation) that help the models to perform well (in computer vision).
# +
# install required libraries
# # !pip install opencv-python==4.5.1.48
# # !pip install numpy==1.21.0
# # !pip install matplotlib==3.4.2
# +
# import the required libraries
import cv2
import numpy as np
import matplotlib.pyplot as plt
from copy import deepcopy
# set random seed to reproduce results
np.random.seed(42)
# +
# define labels and read images
cat_label = np.array([[1, 0]])
dog_label = np.array([[0, 1]])
cat_img = cv2.imread("../assets/images/aug/cat.jpg")/255.0
dog_img = cv2.imread("../assets/images/aug/dog.jpg")/255.0
# +
# plot the original images
fig, ((ax1, ax2)) = plt.subplots(1, 2)
ax1.imshow(cat_img)
ax1.set_title("Cat")
ax2.imshow(dog_img)
ax2.set_title("Dog")
plt.suptitle("Original images",fontsize=15)
plt.show()
# -
# # Cutout
# [paper link](https://arxiv.org/abs/1708.04552)
#
# The main motivation of Cutout is to simulate the situation of object occlusion that is mostly encountered in tasks such as object recognition or human pose estimation. We occlude the part of image randomly. Instead of model seeing the same image everytime, it sees different parts of that image which helps it to perform well. The occluded part does not contain any information. In the example below, the randomly occluded part is replaced by all 0s.
class Cutout(object):
def __init__(self, cutout_width):
# width of the square to occlude
self.cutout_width = cutout_width
def __call__(self, img):
h = img.shape[0]
w = img.shape[1]
mask = np.ones(img.shape)
y = np.random.randint(h)
x = np.random.randint(w)
x1 = np.clip(x - self.cutout_width // 2, 0, w)
x2 = np.clip(x + self.cutout_width // 2, 0, w)
y1 = np.clip(y - self.cutout_width // 2, 0, h)
y2 = np.clip(y + self.cutout_width // 2, 0, h)
mask[y1: y2, x1: x2] = 0.0
# occlude the part of image using mask (zero out the part)
img = img * mask
return img
# +
cutout = Cutout(50)
plt.imshow(cutout(cat_img))
plt.suptitle("Cutout example",fontsize=15)
plt.show()
# -
# # Mixup
# [paper link](https://arxiv.org/abs/1710.09412)
#
# Mixup linearly mixes two images and their labels in the same proportion. With this approach the model will be able to see a different image with different label during training. It enables the model to make smooth decision boundaries while classifying the object since we are using linearly interpolated images and labels for classification decision intead of binary decision. As can be seen in the example below, with the mixup of images and labels, the new image will have the labels `[0.19, 0.81]` which means the class distribution is tilted more towards dog and the mixup visualization also proves this.
class Mixup:
def __init__(self, img1, img2, label1, label2):
self.img1 = img1
self.img2 = img2
self.label1 = label1
self.label2 = label2
def __call__(self):
alpha = 1
lam = np.random.beta(alpha, alpha)
# mix image and labels
mix_img = lam * self.img1 + (1 - lam) * self.img2
mix_label = lam * self.label1 + (1 - lam) * self.label2
return mix_img, mix_label
mix_img, mix_label = Mixup(cat_img, dog_img, cat_label, dog_label)()
# +
fig, ((ax1, ax2, ax3)) = plt.subplots(1, 3)
ax1.imshow(cat_img)
ax1.set_title("Cat")
ax2.imshow(dog_img)
ax2.set_title("Dog")
ax3.imshow(mix_img)
ax3.set_title("Cat/Dog Mixup")
plt.suptitle("Mixup example",fontsize=15)
plt.show()
# -
# # Cutmix
# [paper link](https://arxiv.org/abs/1905.04899)
#
# As we saw above, the occluded part in Cutout does not contain any information which in unwanted since it just increases resource usage without adding any value. Cutmix utilizes above two techniques to mix the images and utilize the occluded part. As can be seen in the example, the cutout portion in dog image is replaced by the same portion of cat image. It also linearly interpolates the label as in Mixup. As seen in the example below, the new labels are `[0.18, 0.82]` for cat and dog respectively and this can be verified from image as well.
class Cutmix:
def __init__(self, img1, img2, lbl1, lbl2):
self.img1 = img1
self.img2 = img2
self.lbl1 = lbl1
self.lbl2 = lbl2
def __call__(self):
# sample bounding box B
alpha = 1
lam = np.random.beta(alpha, alpha)
h, w, c = self.img1.shape
r_x = np.random.randint(0, w)
r_y = np.random.randint(0, h)
r_w = np.int32(w * np.sqrt(1 - lam))
r_h = np.int32(h * np.sqrt(1 - lam))
mid_x, mid_y = r_w//2, r_h//2
point1_x = np.clip(r_x - mid_x, 0, w)
point2_x = np.clip(r_x + mid_x, 0, w)
point1_y = np.clip(r_y - mid_y, 0, h)
point2_y = np.clip(r_y + mid_y, 0, h)
# refer to paper to see how M is cimputed, in short it is to make the ratio of mixing images and labels same
M = np.ones(self.img1.shape)
M[point1_x:point2_x, point1_y:point2_y, :] = 0
cut_mix_img = M * self.img1 + (1 - M) * self.img2
cut_mix_label = lam * self.lbl1 + (1 - lam) * self.lbl2
return cut_mix_img, cut_mix_label
cut_mix_img, cut_mix_lbl = Cutmix(cat_img, dog_img, cat_label, dog_label)()
# +
fig, ((ax1, ax2, ax3)) = plt.subplots(1, 3)
ax1.imshow(cat_img)
ax1.set_title("Cat")
ax2.imshow(dog_img)
ax2.set_title("Dog")
ax3.imshow(cut_mix_img)
ax3.set_title("Cat mixed onto Dog")
plt.suptitle("Cutmix example",fontsize=15)
plt.show()
# -
# # Label Smoothing
# [paper link](https://arxiv.org/abs/1512.00567)
#
# Label smoothing is used so that the model can be prevented from memorizing the training data and being over-confident. It adds noise to the labels without modifying the data itself. If we are using a 5-class classifier then instead of label being assigned only to one class, it is distributed among all the classes. As seen in example below, $\epsilon$ is used to control the smoothing.
# +
ϵ = 0.01
num_labels = 5
classes = ["cat", "dog", "horse", "bus", "carrot"]
original_label = [1, 0, 0, 0, 0]
new_neg_labels = ϵ/(num_labels-1)
# after label smoothing, cat gets 1 - ϵ and other classes get ϵ/(1-ϵ) probability
smooth_labels = [1 - ϵ, new_neg_labels, new_neg_labels, new_neg_labels, new_neg_labels]
smooth_labels
# -
# All the above implementations assume that both the images are of same resolution. There might be some minor differences while compared with the original paper. However the main motivation and results of these techniques are same to that of paper. Please feel free to post any comment, question or suggestion. I'll see you in the next one. :)
| _notebooks/2022-03-27-Augmentations-visually-explained.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="V9zNGvape2-I"
# # **CARE: Content-aware image restoration (2D)**
#
# ---
#
# <font size = 4>CARE is a neural network capable of image restoration from corrupted bio-images, first published in 2018 by [Weigert *et al.* in Nature Methods](https://www.nature.com/articles/s41592-018-0216-7). The CARE network uses a U-Net network architecture and allows image restoration and resolution improvement in 2D and 3D images, in a supervised manner, using noisy images as input and low-noise images as targets for training. The function of the network is essentially determined by the set of images provided in the training dataset. For instance, if noisy images are provided as input and high signal-to-noise ratio images are provided as targets, the network will perform denoising.
#
# <font size = 4> **This particular notebook enables restoration of 2D datasets. If you are interested in restoring a 3D dataset, you should use the CARE 3D notebook instead.**
#
# ---
#
# <font size = 4>*Disclaimer*:
#
# <font size = 4>This notebook is part of the *Zero-Cost Deep-Learning to Enhance Microscopy* project (https://github.com/HenriquesLab/DeepLearning_Collab/wiki). Jointly developed by the Jacquemet (link to https://cellmig.org/) and Henriques (https://henriqueslab.github.io/) laboratories.
#
# <font size = 4>This notebook is based on the following paper:
#
# <font size = 4>**Content-aware image restoration: pushing the limits of fluorescence microscopy**, by Weigert *et al.* published in Nature Methods in 2018 (https://www.nature.com/articles/s41592-018-0216-7)
#
# <font size = 4>And source code found in: https://github.com/csbdeep/csbdeep
#
# <font size = 4>For a more in-depth description of the features of the network, please refer to [this guide](http://csbdeep.bioimagecomputing.com/doc/) provided by the original authors of the work.
#
# <font size = 4>We provide a dataset for the training of this notebook as a way to test its functionalities but the training and test data of the restoration experiments is also available from the authors of the original paper [here](https://publications.mpi-cbg.de/publications-sites/7207/).
#
#
# <font size = 4>**Please also cite this original paper when using or developing this notebook.**
# + [markdown] id="jWAz2i7RdxUV"
# # **How to use this notebook?**
#
# ---
#
# <font size = 4>Video describing how to use our notebooks are available on youtube:
# - [**Video 1**](https://www.youtube.com/watch?v=GzD2gamVNHI&feature=youtu.be): Full run through of the workflow to obtain the notebooks and the provided test datasets as well as a common use of the notebook
# - [**Video 2**](https://www.youtube.com/watch?v=PUuQfP5SsqM&feature=youtu.be): Detailed description of the different sections of the notebook
#
#
# ---
# ###**Structure of a notebook**
#
# <font size = 4>The notebook contains two types of cell:
#
# <font size = 4>**Text cells** provide information and can be modified by douple-clicking the cell. You are currently reading the text cell. You can create a new text by clicking `+ Text`.
#
# <font size = 4>**Code cells** contain code and the code can be modfied by selecting the cell. To execute the cell, move your cursor on the `[ ]`-mark on the left side of the cell (play button appears). Click to execute the cell. After execution is done the animation of play button stops. You can create a new coding cell by clicking `+ Code`.
#
# ---
# ###**Table of contents, Code snippets** and **Files**
#
# <font size = 4>On the top left side of the notebook you find three tabs which contain from top to bottom:
#
# <font size = 4>*Table of contents* = contains structure of the notebook. Click the content to move quickly between sections.
#
# <font size = 4>*Code snippets* = contain examples how to code certain tasks. You can ignore this when using this notebook.
#
# <font size = 4>*Files* = contain all available files. After mounting your google drive (see section 1.) you will find your files and folders here.
#
# <font size = 4>**Remember that all uploaded files are purged after changing the runtime.** All files saved in Google Drive will remain. You do not need to use the Mount Drive-button; your Google Drive is connected in section 1.2.
#
# <font size = 4>**Note:** The "sample data" in "Files" contains default files. Do not upload anything in here!
#
# ---
# ###**Making changes to the notebook**
#
# <font size = 4>**You can make a copy** of the notebook and save it to your Google Drive. To do this click file -> save a copy in drive.
#
# <font size = 4>To **edit a cell**, double click on the text. This will show you either the source code (in code cells) or the source text (in text cells).
# You can use the `#`-mark in code cells to comment out parts of the code. This allows you to keep the original code piece in the cell as a comment.
# + [markdown] id="vNMDQHm0Ah-Z"
# #**0. Before getting started**
# ---
# <font size = 4> For CARE to train, **it needs to have access to a paired training dataset**. This means that the same image needs to be acquired in the two conditions (for instance, low signal-to-noise ratio and high signal-to-noise ratio) and provided with indication of correspondence.
#
# <font size = 4> Therefore, the data structure is important. It is necessary that all the input data are in the same folder and that all the output data is in a separate folder. The provided training dataset is already split in two folders called "Training - Low SNR images" (Training_source) and "Training - high SNR images" (Training_target). Information on how to generate a training dataset is available in our Wiki page: https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki
#
# <font size = 4>**We strongly recommend that you generate extra paired images. These images can be used to assess the quality of your trained model (Quality control dataset)**. The quality control assessment can be done directly in this notebook.
#
# <font size = 4> **Additionally, the corresponding input and output files need to have the same name**.
#
# <font size = 4> Please note that you currently can **only use .tif files!**
#
#
# <font size = 4>Here's a common data structure that can work:
# * Experiment A
# - **Training dataset**
# - Low SNR images (Training_source)
# - img_1.tif, img_2.tif, ...
# - High SNR images (Training_target)
# - img_1.tif, img_2.tif, ...
# - **Quality control dataset**
# - Low SNR images
# - img_1.tif, img_2.tif
# - High SNR images
# - img_1.tif, img_2.tif
# - **Data to be predicted**
# - **Results**
#
# ---
# <font size = 4>**Important note**
#
# <font size = 4>- If you wish to **Train a network from scratch** using your own dataset (and we encourage everyone to do that), you will need to run **sections 1 - 4**, then use **section 5** to assess the quality of your model and **section 6** to run predictions using the model that you trained.
#
# <font size = 4>- If you wish to **Evaluate your model** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 5** to assess the quality of your model.
#
# <font size = 4>- If you only wish to **run predictions** using a model previously generated and saved on your Google Drive, you will only need to run **sections 1 and 2** to set up the notebook, then use **section 6** to run the predictions on the desired model.
# ---
# -
# ## 0.1 Download example data
# + jupyter={"source_hidden": true}
data_import = "Download example data from Biostudies" #@param ["Download example data from Biostudies", "Use my own"]
if data_import:
# !wget -r ftp://ftp.ebi.ac.uk/biostudies/nfs/S-BSST/666/S-BSST666/Files/ZeroCostDl4Mic/Stardist_v2 --show-progress -q --cut-dirs=7 -nH -np
# + [markdown] id="b4-r1gE7Iamv"
# # **1. Initialise the Colab session**
# ---
# + [markdown] id="DMNHVZfHmbKb"
#
# ## **1.1. Check for GPU access**
# ---
#
# By default, the session should be using Python 3 and GPU acceleration, but it is possible to ensure that these are set properly by doing the following:
#
# <font size = 4>Go to **Runtime -> Change the Runtime type**
#
# <font size = 4>**Runtime type: Python 3** *(Python 3 is programming language in which this program is written)*
#
# <font size = 4>**Accelerator: GPU** *(Graphics processing unit)*
#
# + cellView="form" id="BDhmUgqCStlm" partialCollapse=true
#@markdown ##Run this cell to check if you have GPU access
# %tensorflow_version 1.x
import tensorflow as tf
if tf.test.gpu_device_name()=='':
print('You do not have GPU access.')
print('Did you change your runtime ?')
print('If the runtime setting is correct then Google did not allocate a GPU for your session')
print('Expect slow performance. To access GPU try reconnecting later')
else:
print('You have GPU access')
# !nvidia-smi
# + [markdown] id="-oqBTeLaImnU"
# ## **1.2. Mount your Google Drive**
# ---
# <font size = 4> To use this notebook on the data present in your Google Drive, you need to mount your Google Drive to this notebook.
#
# <font size = 4> Play the cell below to mount your Google Drive and follow the link. In the new browser window, select your drive and select 'Allow', copy the code, paste into the cell and press enter. This will give Colab access to the data on the drive.
#
# <font size = 4> Once this is done, your data are available in the **Files** tab on the top left of notebook.
# + cellView="form" id="01Djr8v-5pPk" jupyter={"source_hidden": true}
#@markdown ##Run this cell to connect your Google Drive to Colab
#@markdown * Click on the URL.
#@markdown * Sign in your Google Account.
#@markdown * Copy the authorization code.
#@markdown * Enter the authorization code.
#@markdown * Click on "Files" site on the right. Refresh the site. Your Google Drive folder should now be available here as "drive".
#mounts user's Google Drive to Google Colab.
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="n4yWFoJNnoin"
# # **2. Install CARE and dependencies**
# ---
#
# + [markdown] id="5d6BsNWn_bHL"
# ## **2.1. Install key dependencies**
# ---
# <font size = 4>
# + cellView="form" id="3u2mXn3XsWzd" partialCollapse=true
#@markdown ##Install CARE and dependencies
#Here, we install libraries which are not already included in Colab.
# !pip install tifffile # contains tools to operate tiff-files
# !pip install csbdeep # contains tools for restoration of fluorescence microcopy images (Content-aware Image Restoration, CARE). It uses Keras and Tensorflow.
# !pip install wget
# !pip install memory_profiler
# !pip install fpdf
#Force session restart
exit(0)
# + [markdown] id="3m8GnyWX-r0Z"
# ## **2.2. Restart your runtime**
# ---
# <font size = 4>
# + [markdown] id="bK6zwRkh-usk"
# **<font size = 4> Your Runtime has automatically restarted. This is normal.**
#
#
# + [markdown] id="eDrWDRP2_fRm"
# ## **2.3. Load key dependencies**
# ---
# <font size = 4>
# + cellView="form" id="aGxvAcGT-rTq" partialCollapse=true
#@markdown ##Load key dependencies
Notebook_version = ['1.12']
from builtins import any as b_any
def get_requirements_path():
# Store requirements file in 'contents' directory
current_dir = os.getcwd()
dir_count = current_dir.count('/') - 1
path = '../' * (dir_count) + 'requirements.txt'
return path
def filter_files(file_list, filter_list):
filtered_list = []
for fname in file_list:
if b_any(fname.split('==')[0] in s for s in filter_list):
filtered_list.append(fname)
return filtered_list
def build_requirements_file(before, after):
path = get_requirements_path()
# Exporting requirements.txt for local run
# !pip freeze > $path
# Get minimum requirements file
df = pd.read_csv(path, delimiter = "\n")
mod_list = [m.split('.')[0] for m in after if not m in before]
req_list_temp = df.values.tolist()
req_list = [x[0] for x in req_list_temp]
# Replace with package name and handle cases where import name is different to module name
mod_name_list = [['sklearn', 'scikit-learn'], ['skimage', 'scikit-image']]
mod_replace_list = [[x[1] for x in mod_name_list] if s in [x[0] for x in mod_name_list] else s for s in mod_list]
filtered_list = filter_files(req_list, mod_replace_list)
file=open(path,'w')
for item in filtered_list:
file.writelines(item + '\n')
file.close()
import sys
before = [str(m) for m in sys.modules]
# %load_ext memory_profiler
#Here, we import and enable Tensorflow 1 instead of Tensorflow 2.
# %tensorflow_version 1.x
import tensorflow
import tensorflow as tf
print(tensorflow.__version__)
print("Tensorflow enabled.")
# ------- Variable specific to CARE -------
from csbdeep.utils import download_and_extract_zip_file, plot_some, axes_dict, plot_history, Path, download_and_extract_zip_file
from csbdeep.data import RawData, create_patches
from csbdeep.io import load_training_data, save_tiff_imagej_compatible
from csbdeep.models import Config, CARE
from csbdeep import data
from __future__ import print_function, unicode_literals, absolute_import, division
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# ------- Common variable to all ZeroCostDL4Mic notebooks -------
import numpy as np
from matplotlib import pyplot as plt
import urllib
import os, random
import shutil
import zipfile
from tifffile import imread, imsave
import time
import sys
import wget
from pathlib import Path
import pandas as pd
import csv
from glob import glob
from scipy import signal
from scipy import ndimage
from skimage import io
from sklearn.linear_model import LinearRegression
from skimage.util import img_as_uint
import matplotlib as mpl
from skimage.metrics import structural_similarity
from skimage.metrics import peak_signal_noise_ratio as psnr
from astropy.visualization import simple_norm
from skimage import img_as_float32
from skimage.util import img_as_ubyte
from tqdm import tqdm
from fpdf import FPDF, HTMLMixin
from datetime import datetime
import subprocess
from pip._internal.operations.freeze import freeze
# Colors for the warning messages
class bcolors:
WARNING = '\033[31m'
W = '\033[0m' # white (normal)
R = '\033[31m' # red
#Disable some of the tensorflow warnings
import warnings
warnings.filterwarnings("ignore")
print("Libraries installed")
# Check if this is the latest version of the notebook
Latest_notebook_version = pd.read_csv("https://raw.githubusercontent.com/HenriquesLab/ZeroCostDL4Mic/master/Colab_notebooks/Latest_ZeroCostDL4Mic_Release.csv")
print('Notebook version: '+Notebook_version[0])
strlist = Notebook_version[0].split('.')
Notebook_version_main = strlist[0]+'.'+strlist[1]
if Notebook_version_main == Latest_notebook_version.columns:
print("This notebook is up-to-date.")
else:
print(bcolors.WARNING +"A new version of this notebook has been released. We recommend that you download it at https://github.com/HenriquesLab/ZeroCostDL4Mic/wiki")
# !pip freeze > requirements.txt
#Create a pdf document with training summary
def pdf_export(trained = False, augmentation = False, pretrained_model = False):
# save FPDF() class into a
# variable pdf
#from datetime import datetime
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'CARE 2D'
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Training report for '+Network+' model ('+model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
# add another cell
if trained:
training_time = "Training time: "+str(hour)+ "hour(s) "+str(mins)+"min(s) "+str(round(sec))+"sec(s)"
pdf.cell(190, 5, txt = training_time, ln = 1, align='L')
pdf.ln(1)
Header_2 = 'Information for your materials and methods:'
pdf.cell(190, 5, txt=Header_2, ln=1, align='L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
#print(all_packages)
#Main Packages
main_packages = ''
version_numbers = []
for name in ['tensorflow','numpy','Keras','csbdeep']:
find_name=all_packages.find(name)
main_packages = main_packages+all_packages[find_name:all_packages.find(',',find_name)]+', '
#Version numbers only here:
version_numbers.append(all_packages[find_name+len(name)+2:all_packages.find(',',find_name)])
cuda_version = subprocess.run('nvcc --version',stdout=subprocess.PIPE, shell=True)
cuda_version = cuda_version.stdout.decode('utf-8')
cuda_version = cuda_version[cuda_version.find(', V')+3:-1]
gpu_name = subprocess.run('nvidia-smi',stdout=subprocess.PIPE, shell=True)
gpu_name = gpu_name.stdout.decode('utf-8')
gpu_name = gpu_name[gpu_name.find('Tesla'):gpu_name.find('Tesla')+10]
#print(cuda_version[cuda_version.find(', V')+3:-1])
#print(gpu_name)
shape = io.imread(Training_source+'/'+os.listdir(Training_source)[1]).shape
dataset_size = len(os.listdir(Training_source))
text = 'The '+Network+' model was trained from scratch for '+str(number_of_epochs)+' epochs on '+str(dataset_size*number_of_patches)+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a '+config.train_loss+' loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). Key python packages used include tensorflow (v '+version_numbers[0]+'), Keras (v '+version_numbers[2]+'), csbdeep (v '+version_numbers[3]+'), numpy (v '+version_numbers[1]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
if pretrained_model:
text = 'The '+Network+' model was trained for '+str(number_of_epochs)+' epochs on '+str(dataset_size*number_of_patches)+' paired image patches (image dimensions: '+str(shape)+', patch size: ('+str(patch_size)+','+str(patch_size)+')) with a batch size of '+str(batch_size)+' and a '+config.train_loss+' loss function, using the '+Network+' ZeroCostDL4Mic notebook (v '+Notebook_version[0]+') (von Chamier & Laine et al., 2020). The model was re-trained from a pretrained model. Key python packages used include tensorflow (v '+version_numbers[0]+'), Keras (v '+version_numbers[2]+'), csbdeep (v '+version_numbers[3]+'), numpy (v '+version_numbers[1]+'), cuda (v '+cuda_version+'). The training was accelerated using a '+gpu_name+'GPU.'
pdf.set_font('')
pdf.set_font_size(10.)
pdf.multi_cell(190, 5, txt = text, align='L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(1)
pdf.cell(28, 5, txt='Augmentation: ', ln=0)
pdf.set_font('')
if augmentation:
aug_text = 'The dataset was augmented by a factor of '+str(Multiply_dataset_by)+' by'
if rotate_270_degrees != 0 or rotate_90_degrees != 0:
aug_text = aug_text+'\n- rotation'
if flip_left_right != 0 or flip_top_bottom != 0:
aug_text = aug_text+'\n- flipping'
if random_zoom_magnification != 0:
aug_text = aug_text+'\n- random zoom magnification'
if random_distortion != 0:
aug_text = aug_text+'\n- random distortion'
if image_shear != 0:
aug_text = aug_text+'\n- image shearing'
if skew_image != 0:
aug_text = aug_text+'\n- image skewing'
else:
aug_text = 'No augmentation was used for training.'
pdf.multi_cell(190, 5, txt=aug_text, align='L')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Parameters', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
if Use_Default_Advanced_Parameters:
pdf.cell(200, 5, txt='Default Advanced Parameters were enabled')
pdf.cell(200, 5, txt='The following parameters were used for training:')
pdf.ln(1)
html = """
<table width=40% style="margin-left:0px;">
<tr>
<th width = 50% align="left">Parameter</th>
<th width = 50% align="left">Value</th>
</tr>
<tr>
<td width = 50%>number_of_epochs</td>
<td width = 50%>{0}</td>
</tr>
<tr>
<td width = 50%>patch_size</td>
<td width = 50%>{1}</td>
</tr>
<tr>
<td width = 50%>number_of_patches</td>
<td width = 50%>{2}</td>
</tr>
<tr>
<td width = 50%>batch_size</td>
<td width = 50%>{3}</td>
</tr>
<tr>
<td width = 50%>number_of_steps</td>
<td width = 50%>{4}</td>
</tr>
<tr>
<td width = 50%>percentage_validation</td>
<td width = 50%>{5}</td>
</tr>
<tr>
<td width = 50%>initial_learning_rate</td>
<td width = 50%>{6}</td>
</tr>
</table>
""".format(number_of_epochs,str(patch_size)+'x'+str(patch_size),number_of_patches,batch_size,number_of_steps,percentage_validation,initial_learning_rate)
pdf.write_html(html)
#pdf.multi_cell(190, 5, txt = text_2, align='L')
pdf.set_font("Arial", size = 11, style='B')
pdf.ln(1)
pdf.cell(190, 5, txt = 'Training Dataset', align='L', ln=1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(29, 5, txt= 'Training_source:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_source, align = 'L')
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(27, 5, txt= 'Training_target:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = Training_target, align = 'L')
#pdf.cell(190, 5, txt=aug_text, align='L', ln=1)
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.cell(22, 5, txt= 'Model Path:', align = 'L', ln=0)
pdf.set_font('')
pdf.multi_cell(170, 5, txt = model_path+'/'+model_name, align = 'L')
pdf.ln(1)
pdf.cell(60, 5, txt = 'Example Training pair', ln=1)
pdf.ln(1)
exp_size = io.imread('/content/TrainingDataExample_CARE2D.png').shape
pdf.image('/content/TrainingDataExample_CARE2D.png', x = 11, y = None, w = round(exp_size[1]/8), h = round(exp_size[0]/8))
pdf.ln(1)
ref_1 = 'References:\n - ZeroCostDL4Mic: <NAME>, Lucas & Laine, Romain, et al. "Democratising deep learning for microscopy with ZeroCostDL4Mic." Nature Communications (2021).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- CARE: Weigert, Martin, et al. "Content-aware image restoration: pushing the limits of fluorescence microscopy." Nature methods 15.12 (2018): 1090-1097.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
if augmentation:
ref_3 = '- Augmentor: Bloice, <NAME>., <NAME>, and <NAME>. "Augmentor: an image augmentation library for machine learning." arXiv preprint arXiv:1708.04680 (2017).'
pdf.multi_cell(190, 5, txt = ref_3, align='L')
pdf.ln(3)
reminder = 'Important:\nRemember to perform the quality control step on all newly trained models\nPlease consider depositing your training dataset on Zenodo'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(model_path+'/'+model_name+'/'+model_name+"_training_report.pdf")
#Make a pdf summary of the QC results
def qc_pdf_export():
class MyFPDF(FPDF, HTMLMixin):
pass
pdf = MyFPDF()
pdf.add_page()
pdf.set_right_margin(-1)
pdf.set_font("Arial", size = 11, style='B')
Network = 'CARE 2D'
#model_name = os.path.basename(full_QC_model_path)
day = datetime.now()
datetime_str = str(day)[0:10]
Header = 'Quality Control report for '+Network+' model ('+QC_model_name+')\nDate: '+datetime_str
pdf.multi_cell(180, 5, txt = Header, align = 'L')
all_packages = ''
for requirement in freeze(local_only=True):
all_packages = all_packages+requirement+', '
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(2)
pdf.cell(190, 5, txt = 'Development of Training Losses', ln=1, align='L')
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'Quality Control/QC_example_data.png').shape
if os.path.exists(full_QC_model_path+'Quality Control/lossCurvePlots.png'):
pdf.image(full_QC_model_path+'Quality Control/lossCurvePlots.png', x = 11, y = None, w = round(exp_size[1]/10), h = round(exp_size[0]/13))
else:
pdf.set_font('')
pdf.set_font('Arial', size=10)
pdf.multi_cell(190, 5, txt='If you would like to see the evolution of the loss function during training please play the first cell of the QC section in the notebook.', align='L')
pdf.ln(2)
pdf.set_font('')
pdf.set_font('Arial', size = 10, style = 'B')
pdf.ln(3)
pdf.cell(80, 5, txt = 'Example Quality Control Visualisation', ln=1)
pdf.ln(1)
exp_size = io.imread(full_QC_model_path+'Quality Control/QC_example_data.png').shape
pdf.image(full_QC_model_path+'Quality Control/QC_example_data.png', x = 16, y = None, w = round(exp_size[1]/10), h = round(exp_size[0]/10))
pdf.ln(1)
pdf.set_font('')
pdf.set_font('Arial', size = 11, style = 'B')
pdf.ln(1)
pdf.cell(180, 5, txt = 'Quality Control Metrics', align='L', ln=1)
pdf.set_font('')
pdf.set_font_size(10.)
pdf.ln(1)
html = """
<body>
<font size="7" face="Courier New" >
<table width=94% style="margin-left:0px;">"""
with open(full_QC_model_path+'Quality Control/QC_metrics_'+QC_model_name+'.csv', 'r') as csvfile:
metrics = csv.reader(csvfile)
header = next(metrics)
image = header[0]
mSSIM_PvsGT = header[1]
mSSIM_SvsGT = header[2]
NRMSE_PvsGT = header[3]
NRMSE_SvsGT = header[4]
PSNR_PvsGT = header[5]
PSNR_SvsGT = header[6]
header = """
<tr>
<th width = 10% align="left">{0}</th>
<th width = 15% align="left">{1}</th>
<th width = 15% align="center">{2}</th>
<th width = 15% align="left">{3}</th>
<th width = 15% align="center">{4}</th>
<th width = 15% align="left">{5}</th>
<th width = 15% align="center">{6}</th>
</tr>""".format(image,mSSIM_PvsGT,mSSIM_SvsGT,NRMSE_PvsGT,NRMSE_SvsGT,PSNR_PvsGT,PSNR_SvsGT)
html = html+header
for row in metrics:
image = row[0]
mSSIM_PvsGT = row[1]
mSSIM_SvsGT = row[2]
NRMSE_PvsGT = row[3]
NRMSE_SvsGT = row[4]
PSNR_PvsGT = row[5]
PSNR_SvsGT = row[6]
cells = """
<tr>
<td width = 10% align="left">{0}</td>
<td width = 15% align="center">{1}</td>
<td width = 15% align="center">{2}</td>
<td width = 15% align="center">{3}</td>
<td width = 15% align="center">{4}</td>
<td width = 15% align="center">{5}</td>
<td width = 15% align="center">{6}</td>
</tr>""".format(image,str(round(float(mSSIM_PvsGT),3)),str(round(float(mSSIM_SvsGT),3)),str(round(float(NRMSE_PvsGT),3)),str(round(float(NRMSE_SvsGT),3)),str(round(float(PSNR_PvsGT),3)),str(round(float(PSNR_SvsGT),3)))
html = html+cells
html = html+"""</body></table>"""
pdf.write_html(html)
pdf.ln(1)
pdf.set_font('')
pdf.set_font_size(10.)
ref_1 = 'References:\n - ZeroCostDL4Mic: <NAME>, <NAME>, et al. "Democratising deep learning for microscopy with ZeroCostDL4Mic." Nature Communications (2021).'
pdf.multi_cell(190, 5, txt = ref_1, align='L')
ref_2 = '- CARE: <NAME>, et al. "Content-aware image restoration: pushing the limits of fluorescence microscopy." Nature methods 15.12 (2018): 1090-1097.'
pdf.multi_cell(190, 5, txt = ref_2, align='L')
pdf.ln(3)
reminder = 'To find the parameters and other information about how this model was trained, go to the training_report.pdf of this model which should be in the folder of the same name.'
pdf.set_font('Arial', size = 11, style='B')
pdf.multi_cell(190, 5, txt=reminder, align='C')
pdf.output(full_QC_model_path+'Quality Control/'+QC_model_name+'_QC_report.pdf')
# Build requirements file for local run
after = [str(m) for m in sys.modules]
build_requirements_file(before, after)
# + [markdown] id="Fw0kkTU6CsU4"
# # **3. Select your parameters and paths**
#
# ---
#
# + [markdown] id="WzYAA-MuaYrT"
# ## **3.1. Setting main training parameters**
# ---
# <font size = 4>
#
#
# + [markdown] id="CB6acvUFtWqd"
# <font size = 5> **Paths for training, predictions and results**
#
# <font size = 4>**`Training_source:`, `Training_target`:** These are the paths to your folders containing the Training_source (Low SNR images) and Training_target (High SNR images or ground truth) training data respecively. To find the paths of the folders containing the respective datasets, go to your Files on the left of the notebook, navigate to the folder containing your files and copy the path by right-clicking on the folder, **Copy path** and pasting it into the right box below.
#
# <font size = 4>**`model_name`:** Use only my_model -style, not my-model (Use "_" not "-"). Do not use spaces in the name. Avoid using the name of an existing model (saved in the same folder) as it will be overwritten.
#
# <font size = 4>**`model_path`**: Enter the path where your model will be saved once trained (for instance your result folder).
#
# <font size = 5>**Training Parameters**
#
# <font size = 4>**`number_of_epochs`:**Input how many epochs (rounds) the network will be trained. Preliminary results can already be observed after a few (10-30) epochs, but a full training should run for 100-300 epochs. Evaluate the performance after training (see 5). **Default value: 50**
#
# <font size = 4>**`patch_size`:** CARE divides the image into patches for training. Input the size of the patches (length of a side). The value should be smaller than the dimensions of the image and divisible by 8. **Default value: 128**
#
# <font size = 4>**When choosing the patch_size, the value should be i) large enough that it will enclose many instances, ii) small enough that the resulting patches fit into the RAM.**<font size = 4>
#
# <font size = 4>**`number_of_patches`:** Input the number of the patches per image. Increasing the number of patches allows for larger training datasets. **Default value: 50**
#
# <font size = 4>**Decreasing the patch size or increasing the number of patches may improve the training but may also increase the training time.**
#
# <font size = 5>**Advanced Parameters - experienced users only**
#
# <font size =4>**`batch_size:`** This parameter defines the number of patches seen in each training step. Reducing or increasing the **batch size** may slow or speed up your training, respectively, and can influence network performance. **Default value: 16**
#
# <font size = 4>**`number_of_steps`:** Define the number of training steps by epoch. By default or if set to zero this parameter is calculated so that each patch is seen at least once per epoch. **Default value: Number of patches / batch_size**
#
# <font size = 4>**`percentage_validation`:** Input the percentage of your training dataset you want to use to validate the network during training. **Default value: 10**
#
# <font size = 4>**`initial_learning_rate`:** Input the initial value to be used as learning rate. **Default value: 0.0004**
# + cellView="form" id="ewpNJ_I0Mv47" partialCollapse=true
#@markdown ###Path to training images:
Training_source = "" #@param {type:"string"}
InputFile = Training_source+"/*.tif"
Training_target = "" #@param {type:"string"}
OutputFile = Training_target+"/*.tif"
#Define where the patch file will be saved
base = "/content"
# model name and path
#@markdown ###Name of the model and path to model folder:
model_name = "" #@param {type:"string"}
model_path = "" #@param {type:"string"}
# other parameters for training.
#@markdown ###Training Parameters
#@markdown Number of epochs:
number_of_epochs = 50#@param {type:"number"}
#@markdown Patch size (pixels) and number
patch_size = 128#@param {type:"number"} # in pixels
number_of_patches = 50#@param {type:"number"}
#@markdown ###Advanced Parameters
Use_Default_Advanced_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please input:
batch_size = 16#@param {type:"number"}
number_of_steps = 0#@param {type:"number"}
percentage_validation = 10 #@param {type:"number"}
initial_learning_rate = 0.0004 #@param {type:"number"}
if (Use_Default_Advanced_Parameters):
print("Default advanced parameters enabled")
batch_size = 16
percentage_validation = 10
initial_learning_rate = 0.0004
#Here we define the percentage to use for validation
percentage = percentage_validation/100
#here we check that no model with the same name already exist, if so print a warning
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: "+model_name+" already exists and will be deleted in the following cell !!")
print(bcolors.WARNING +"To continue training "+model_name+", choose a new model_name here, and load "+model_name+" in section 3.3"+W)
# Here we disable pre-trained model by default (in case the cell is not ran)
Use_pretrained_model = False
# Here we disable data augmentation by default (in case the cell is not ran)
Use_Data_augmentation = False
print("Parameters initiated.")
# This will display a randomly chosen dataset input and output
random_choice = random.choice(os.listdir(Training_source))
x = imread(Training_source+"/"+random_choice)
# Here we check that the input images contains the expected dimensions
if len(x.shape) == 2:
print("Image dimensions (y,x)",x.shape)
if not len(x.shape) == 2:
print(bcolors.WARNING +"Your images appear to have the wrong dimensions. Image dimension",x.shape)
#Find image XY dimension
Image_Y = x.shape[0]
Image_X = x.shape[1]
#Hyperparameters failsafes
# Here we check that patch_size is smaller than the smallest xy dimension of the image
if patch_size > min(Image_Y, Image_X):
patch_size = min(Image_Y, Image_X)
print (bcolors.WARNING + " Your chosen patch_size is bigger than the xy dimension of your image; therefore the patch_size chosen is now:",patch_size)
# Here we check that patch_size is divisible by 8
if not patch_size % 8 == 0:
patch_size = ((int(patch_size / 8)-1) * 8)
print (bcolors.WARNING + " Your chosen patch_size is not divisible by 8; therefore the patch_size chosen is now:",patch_size)
os.chdir(Training_target)
y = imread(Training_target+"/"+random_choice)
f=plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.imshow(x, norm=simple_norm(x, percent = 99), interpolation='nearest')
plt.title('Training source')
plt.axis('off');
plt.subplot(1,2,2)
plt.imshow(y, norm=simple_norm(y, percent = 99), interpolation='nearest')
plt.title('Training target')
plt.axis('off');
plt.savefig('/content/TrainingDataExample_CARE2D.png',bbox_inches='tight',pad_inches=0)
# + [markdown] id="xGcl7WGP4WHt"
# ## **3.2. Data augmentation**
# ---
# + [markdown] id="5Lio8hpZ4PJ1"
# <font size = 4>Data augmentation can improve training progress by amplifying differences in the dataset. This can be useful if the available dataset is small since, in this case, it is possible that a network could quickly learn every example in the dataset (overfitting), without augmentation. Augmentation is not necessary for training and if your training dataset is large you should disable it.
#
# <font size = 4> **However, data augmentation is not a magic solution and may also introduce issues. Therefore, we recommend that you train your network with and without augmentation, and use the QC section to validate that it improves overall performances.**
#
# <font size = 4>Data augmentation is performed here by [Augmentor.](https://github.com/mdbloice/Augmentor)
#
# <font size = 4>[Augmentor](https://github.com/mdbloice/Augmentor) was described in the following article:
#
# <font size = 4><NAME>, <NAME>, <NAME>, Biomedical image augmentation using Augmentor, Bioinformatics, https://doi.org/10.1093/bioinformatics/btz259
#
# <font size = 4>**Please also cite this original paper when publishing results obtained using this notebook with augmentation enabled.**
# + cellView="form" id="htqjkJWt5J_8" partialCollapse=true
#Data augmentation
Use_Data_augmentation = False #@param {type:"boolean"}
if Use_Data_augmentation:
# !pip install Augmentor
import Augmentor
#@markdown ####Choose a factor by which you want to multiply your original dataset
Multiply_dataset_by = 30 #@param {type:"slider", min:1, max:30, step:1}
Save_augmented_images = False #@param {type:"boolean"}
Saving_path = "" #@param {type:"string"}
Use_Default_Augmentation_Parameters = True #@param {type:"boolean"}
#@markdown ###If not, please choose the probability of the following image manipulations to be used to augment your dataset (1 = always used; 0 = disabled ):
#@markdown ####Mirror and rotate images
rotate_90_degrees = 0 #@param {type:"slider", min:0, max:1, step:0.1}
rotate_270_degrees = 0 #@param {type:"slider", min:0, max:1, step:0.1}
flip_left_right = 0 #@param {type:"slider", min:0, max:1, step:0.1}
flip_top_bottom = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Random image Zoom
random_zoom = 0 #@param {type:"slider", min:0, max:1, step:0.1}
random_zoom_magnification = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Random image distortion
random_distortion = 0 #@param {type:"slider", min:0, max:1, step:0.1}
#@markdown ####Image shearing and skewing
image_shear = 0 #@param {type:"slider", min:0, max:1, step:0.1}
max_image_shear = 1 #@param {type:"slider", min:1, max:25, step:1}
skew_image = 0 #@param {type:"slider", min:0, max:1, step:0.1}
skew_image_magnitude = 0 #@param {type:"slider", min:0, max:1, step:0.1}
if Use_Default_Augmentation_Parameters:
rotate_90_degrees = 0.5
rotate_270_degrees = 0.5
flip_left_right = 0.5
flip_top_bottom = 0.5
if not Multiply_dataset_by >5:
random_zoom = 0
random_zoom_magnification = 0.9
random_distortion = 0
image_shear = 0
max_image_shear = 10
skew_image = 0
skew_image_magnitude = 0
if Multiply_dataset_by >5:
random_zoom = 0.1
random_zoom_magnification = 0.9
random_distortion = 0.5
image_shear = 0.2
max_image_shear = 5
if Multiply_dataset_by >25:
random_zoom = 0.5
random_zoom_magnification = 0.8
random_distortion = 0.5
image_shear = 0.5
max_image_shear = 20
list_files = os.listdir(Training_source)
Nb_files = len(list_files)
Nb_augmented_files = (Nb_files * Multiply_dataset_by)
if Use_Data_augmentation:
print("Data augmentation enabled")
# Here we set the path for the various folder were the augmented images will be loaded
# All images are first saved into the augmented folder
#Augmented_folder = "/content/Augmented_Folder"
if not Save_augmented_images:
Saving_path= "/content"
Augmented_folder = Saving_path+"/Augmented_Folder"
if os.path.exists(Augmented_folder):
shutil.rmtree(Augmented_folder)
os.makedirs(Augmented_folder)
#Training_source_augmented = "/content/Training_source_augmented"
Training_source_augmented = Saving_path+"/Training_source_augmented"
if os.path.exists(Training_source_augmented):
shutil.rmtree(Training_source_augmented)
os.makedirs(Training_source_augmented)
#Training_target_augmented = "/content/Training_target_augmented"
Training_target_augmented = Saving_path+"/Training_target_augmented"
if os.path.exists(Training_target_augmented):
shutil.rmtree(Training_target_augmented)
os.makedirs(Training_target_augmented)
# Here we generate the augmented images
#Load the images
p = Augmentor.Pipeline(Training_source, Augmented_folder)
#Define the matching images
p.ground_truth(Training_target)
#Define the augmentation possibilities
if not rotate_90_degrees == 0:
p.rotate90(probability=rotate_90_degrees)
if not rotate_270_degrees == 0:
p.rotate270(probability=rotate_270_degrees)
if not flip_left_right == 0:
p.flip_left_right(probability=flip_left_right)
if not flip_top_bottom == 0:
p.flip_top_bottom(probability=flip_top_bottom)
if not random_zoom == 0:
p.zoom_random(probability=random_zoom, percentage_area=random_zoom_magnification)
if not random_distortion == 0:
p.random_distortion(probability=random_distortion, grid_width=4, grid_height=4, magnitude=8)
if not image_shear == 0:
p.shear(probability=image_shear,max_shear_left=20,max_shear_right=20)
p.sample(int(Nb_augmented_files))
print(int(Nb_augmented_files),"matching images generated")
# Here we sort through the images and move them back to augmented trainning source and targets folders
augmented_files = os.listdir(Augmented_folder)
for f in augmented_files:
if (f.startswith("_groundtruth_(1)_")):
shortname_noprefix = f[17:]
shutil.copyfile(Augmented_folder+"/"+f, Training_target_augmented+"/"+shortname_noprefix)
if not (f.startswith("_groundtruth_(1)_")):
shutil.copyfile(Augmented_folder+"/"+f, Training_source_augmented+"/"+f)
for filename in os.listdir(Training_source_augmented):
os.chdir(Training_source_augmented)
os.rename(filename, filename.replace('_original', ''))
#Here we clean up the extra files
shutil.rmtree(Augmented_folder)
if not Use_Data_augmentation:
print(bcolors.WARNING+"Data augmentation disabled")
# + [markdown] id="bQDuybvyadKU"
#
# ## **3.3. Using weights from a pre-trained model as initial weights**
# ---
# <font size = 4> Here, you can set the the path to a pre-trained model from which the weights can be extracted and used as a starting point for this training session. **This pre-trained model needs to be a CARE 2D model**.
#
# <font size = 4> This option allows you to perform training over multiple Colab runtimes or to do transfer learning using models trained outside of ZeroCostDL4Mic. **You do not need to run this section if you want to train a network from scratch**.
#
# <font size = 4> In order to continue training from the point where the pre-trained model left off, it is adviseable to also **load the learning rate** that was used when the training ended. This is automatically saved for models trained with ZeroCostDL4Mic and will be loaded here. If no learning rate can be found in the model folder provided, the default learning rate will be used.
# + cellView="form" id="8vPkzEBNamE4" partialCollapse=true
# @markdown ##Loading weights from a pre-trained network
Use_pretrained_model = False #@param {type:"boolean"}
pretrained_model_choice = "Model_from_file" #@param ["Model_from_file"]
Weights_choice = "best" #@param ["last", "best"]
#@markdown ###If you chose "Model_from_file", please provide the path to the model folder:
pretrained_model_path = "" #@param {type:"string"}
# --------------------- Check if we load a previously trained model ------------------------
if Use_pretrained_model:
# --------------------- Load the model from the choosen path ------------------------
if pretrained_model_choice == "Model_from_file":
h5_file_path = os.path.join(pretrained_model_path, "weights_"+Weights_choice+".h5")
# --------------------- Download the a model provided in the XXX ------------------------
if pretrained_model_choice == "Model_name":
pretrained_model_name = "Model_name"
pretrained_model_path = "/content/"+pretrained_model_name
print("Downloading the 2D_Demo_Model_from_Stardist_2D_paper")
if os.path.exists(pretrained_model_path):
shutil.rmtree(pretrained_model_path)
os.makedirs(pretrained_model_path)
wget.download("", pretrained_model_path)
wget.download("", pretrained_model_path)
wget.download("", pretrained_model_path)
wget.download("", pretrained_model_path)
h5_file_path = os.path.join(pretrained_model_path, "weights_"+Weights_choice+".h5")
# --------------------- Add additional pre-trained models here ------------------------
# --------------------- Check the model exist ------------------------
# If the model path chosen does not contain a pretrain model then use_pretrained_model is disabled,
if not os.path.exists(h5_file_path):
print(bcolors.WARNING+'WARNING: weights_'+Weights_choice+'.h5 pretrained model does not exist')
Use_pretrained_model = False
# If the model path contains a pretrain model, we load the training rate,
if os.path.exists(h5_file_path):
#Here we check if the learning rate can be loaded from the quality control folder
if os.path.exists(os.path.join(pretrained_model_path, 'Quality Control', 'training_evaluation.csv')):
with open(os.path.join(pretrained_model_path, 'Quality Control', 'training_evaluation.csv'),'r') as csvfile:
csvRead = pd.read_csv(csvfile, sep=',')
#print(csvRead)
if "learning rate" in csvRead.columns: #Here we check that the learning rate column exist (compatibility with model trained un ZeroCostDL4Mic bellow 1.4)
print("pretrained network learning rate found")
#find the last learning rate
lastLearningRate = csvRead["learning rate"].iloc[-1]
#Find the learning rate corresponding to the lowest validation loss
min_val_loss = csvRead[csvRead['val_loss'] == min(csvRead['val_loss'])]
#print(min_val_loss)
bestLearningRate = min_val_loss['learning rate'].iloc[-1]
if Weights_choice == "last":
print('Last learning rate: '+str(lastLearningRate))
if Weights_choice == "best":
print('Learning rate of best validation loss: '+str(bestLearningRate))
if not "learning rate" in csvRead.columns: #if the column does not exist, then initial learning rate is used instead
bestLearningRate = initial_learning_rate
lastLearningRate = initial_learning_rate
print(bcolors.WARNING+'WARNING: The learning rate cannot be identified from the pretrained network. Default learning rate of '+str(bestLearningRate)+' will be used instead')
#Compatibility with models trained outside ZeroCostDL4Mic but default learning rate will be used
if not os.path.exists(os.path.join(pretrained_model_path, 'Quality Control', 'training_evaluation.csv')):
print(bcolors.WARNING+'WARNING: The learning rate cannot be identified from the pretrained network. Default learning rate of '+str(initial_learning_rate)+' will be used instead')
bestLearningRate = initial_learning_rate
lastLearningRate = initial_learning_rate
# Display info about the pretrained model to be loaded (or not)
if Use_pretrained_model:
print('Weights found in:')
print(h5_file_path)
print('will be loaded prior to training.')
else:
print(bcolors.WARNING+'No pretrained network will be used.')
# + [markdown] id="rQndJj70FzfL"
# # **4. Train the network**
# ---
# + [markdown] id="tGW2iaU6X5zi"
# ## **4.1. Prepare the training data and model for training**
# ---
# <font size = 4>Here, we use the information from 3. to build the model and convert the training data into a suitable format for training.
# + cellView="form" id="WMJnGJpCMa4y" partialCollapse=true
#@markdown ##Create the model and dataset objects
# --------------------- Here we delete the model folder if it already exist ------------------------
if os.path.exists(model_path+'/'+model_name):
print(bcolors.WARNING +"!! WARNING: Model folder already exists and has been removed !!"+W)
shutil.rmtree(model_path+'/'+model_name)
# --------------------- Here we load the augmented data or the raw data ------------------------
if Use_Data_augmentation:
Training_source_dir = Training_source_augmented
Training_target_dir = Training_target_augmented
if not Use_Data_augmentation:
Training_source_dir = Training_source
Training_target_dir = Training_target
# --------------------- ------------------------------------------------
# This object holds the image pairs (GT and low), ensuring that CARE compares corresponding images.
# This file is saved in .npz format and later called when loading the trainig data.
raw_data = data.RawData.from_folder(
basepath=base,
source_dirs=[Training_source_dir],
target_dir=Training_target_dir,
axes='CYX',
pattern='*.tif*')
X, Y, XY_axes = data.create_patches(
raw_data,
patch_filter=None,
patch_size=(patch_size,patch_size),
n_patches_per_image=number_of_patches)
print ('Creating 2D training dataset')
training_path = model_path+"/rawdata"
rawdata1 = training_path+".npz"
np.savez(training_path,X=X, Y=Y, axes=XY_axes)
# Load Training Data
(X,Y), (X_val,Y_val), axes = load_training_data(rawdata1, validation_split=percentage, verbose=True)
c = axes_dict(axes)['C']
n_channel_in, n_channel_out = X.shape[c], Y.shape[c]
# %memit
#plot of training patches.
plt.figure(figsize=(12,5))
plot_some(X[:5],Y[:5])
plt.suptitle('5 example training patches (top row: source, bottom row: target)');
#plot of validation patches
plt.figure(figsize=(12,5))
plot_some(X_val[:5],Y_val[:5])
plt.suptitle('5 example validation patches (top row: source, bottom row: target)');
#Here we automatically define number_of_step in function of training data and batch size
#if (Use_Default_Advanced_Parameters):
if (Use_Default_Advanced_Parameters) or (number_of_steps == 0):
number_of_steps = int(X.shape[0]/batch_size)+1
# --------------------- Using pretrained model ------------------------
#Here we ensure that the learning rate set correctly when using pre-trained models
if Use_pretrained_model:
if Weights_choice == "last":
initial_learning_rate = lastLearningRate
if Weights_choice == "best":
initial_learning_rate = bestLearningRate
# --------------------- ---------------------- ------------------------
#Here we create the configuration file
config = Config(axes, n_channel_in, n_channel_out, probabilistic=True, train_steps_per_epoch=number_of_steps, train_epochs=number_of_epochs, unet_kern_size=5, unet_n_depth=3, train_batch_size=batch_size, train_learning_rate=initial_learning_rate)
print(config)
vars(config)
# Compile the CARE model for network training
model_training= CARE(config, model_name, basedir=model_path)
# --------------------- Using pretrained model ------------------------
# Load the pretrained weights
if Use_pretrained_model:
model_training.load_weights(h5_file_path)
# --------------------- ---------------------- ------------------------
pdf_export(augmentation = Use_Data_augmentation, pretrained_model = Use_pretrained_model)
# + [markdown] id="wQPz0F6JlvJR"
# ## **4.2. Start Training**
# ---
# <font size = 4>When playing the cell below you should see updates after each epoch (round). Network training can take some time.
#
# <font size = 4>* **CRITICAL NOTE:** Google Colab has a time limit for processing (to prevent using GPU power for datamining). Training time must be less than 12 hours! If training takes longer than 12 hours, please decrease the number of epochs or number of patches.
#
# <font size = 4>Once training is complete, the trained model is automatically saved on your Google Drive, in the **model_path** folder that was selected in Section 3. It is however wise to download the folder from Google Drive as all data can be erased at the next training if using the same folder.
#
# <font size = 4>**Of Note:** At the end of the training, your model will be automatically exported so it can be used in the CSBDeep Fiji plugin (Run your Network). You can find it in your model folder (TF_SavedModel.zip). In Fiji, Make sure to choose the right version of tensorflow. You can check at: Edit-- Options-- Tensorflow. Choose the version 1.4 (CPU or GPU depending on your system).
# + cellView="form" id="j_Qm5JBmlvJg" partialCollapse=true
#@markdown ##Start training
start = time.time()
# Start Training
history = model_training.train(X,Y, validation_data=(X_val,Y_val))
print("Training, done.")
# # copy the .npz to the model's folder
shutil.copyfile(model_path+'/rawdata.npz',model_path+'/'+model_name+'/rawdata.npz')
# convert the history.history dict to a pandas DataFrame:
lossData = pd.DataFrame(history.history)
if os.path.exists(model_path+"/"+model_name+"/Quality Control"):
shutil.rmtree(model_path+"/"+model_name+"/Quality Control")
os.makedirs(model_path+"/"+model_name+"/Quality Control")
# The training evaluation.csv is saved (overwrites the Files if needed).
lossDataCSVpath = model_path+'/'+model_name+'/Quality Control/training_evaluation.csv'
with open(lossDataCSVpath, 'w') as f:
writer = csv.writer(f)
writer.writerow(['loss','val_loss', 'learning rate'])
for i in range(len(history.history['loss'])):
writer.writerow([history.history['loss'][i], history.history['val_loss'][i], history.history['lr'][i]])
# Displaying the time elapsed for training
dt = time.time() - start
mins, sec = divmod(dt, 60)
hour, mins = divmod(mins, 60)
print("Time elapsed:",hour, "hour(s)",mins,"min(s)",round(sec),"sec(s)")
model_training.export_TF()
print("Your model has been sucessfully exported and can now also be used in the CSBdeep Fiji plugin")
pdf_export(trained = True, augmentation = Use_Data_augmentation, pretrained_model = Use_pretrained_model)
# + [markdown] id="QYuIOWQ3imuU"
# # **5. Evaluate your model**
# ---
#
# <font size = 4>This section allows you to perform important quality checks on the validity and generalisability of the trained model.
#
# <font size = 4>**We highly recommend to perform quality control on all newly trained models.**
#
#
# + cellView="form" id="zazOZ3wDx0zQ" partialCollapse=true
# model name and path
#@markdown ###Do you want to assess the model you just trained ?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
QC_model_folder = "" #@param {type:"string"}
#Here we define the loaded model name and path
QC_model_name = os.path.basename(QC_model_folder)
QC_model_path = os.path.dirname(QC_model_folder)
if (Use_the_current_trained_model):
QC_model_name = model_name
QC_model_path = model_path
full_QC_model_path = QC_model_path+'/'+QC_model_name+'/'
if os.path.exists(full_QC_model_path):
print("The "+QC_model_name+" network will be evaluated")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
loss_displayed = False
# + [markdown] id="yDY9dtzdUTLh"
# ## **5.1. Inspection of the loss function**
# ---
#
# <font size = 4>First, it is good practice to evaluate the training progress by comparing the training loss with the validation loss. The latter is a metric which shows how well the network performs on a subset of unseen data which is set aside from the training dataset. For more information on this, see for example [this review](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6381354/) by Nichols *et al.*
#
# <font size = 4>**Training loss** describes an error value after each epoch for the difference between the model's prediction and its ground-truth target.
#
# <font size = 4>**Validation loss** describes the same error value between the model's prediction on a validation image and compared to it's target.
#
# <font size = 4>During training both values should decrease before reaching a minimal value which does not decrease further even after more training. Comparing the development of the validation loss with the training loss can give insights into the model's performance.
#
# <font size = 4>Decreasing **Training loss** and **Validation loss** indicates that training is still necessary and increasing the `number_of_epochs` is recommended. Note that the curves can look flat towards the right side, just because of the y-axis scaling. The network has reached convergence once the curves flatten out. After this point no further training is required. If the **Validation loss** suddenly increases again an the **Training loss** simultaneously goes towards zero, it means that the network is overfitting to the training data. In other words the network is remembering the exact patterns from the training data and no longer generalizes well to unseen data. In this case the training dataset has to be increased.
#
# <font size = 4>**Note: Plots of the losses will be shown in a linear and in a log scale. This can help visualise changes in the losses at different magnitudes. However, note that if the losses are negative the plot on the log scale will be empty. This is not an error.**
# + cellView="form" id="vMzSP50kMv5p"
#@markdown ##Play the cell to show a plot of training errors vs. epoch number
loss_displayed = True
lossDataFromCSV = []
vallossDataFromCSV = []
with open(QC_model_path+'/'+QC_model_name+'/Quality Control/training_evaluation.csv','r') as csvfile:
csvRead = csv.reader(csvfile, delimiter=',')
next(csvRead)
for row in csvRead:
lossDataFromCSV.append(float(row[0]))
vallossDataFromCSV.append(float(row[1]))
epochNumber = range(len(lossDataFromCSV))
plt.figure(figsize=(15,10))
plt.subplot(2,1,1)
plt.plot(epochNumber,lossDataFromCSV, label='Training loss')
plt.plot(epochNumber,vallossDataFromCSV, label='Validation loss')
plt.title('Training loss and validation loss vs. epoch number (linear scale)')
plt.ylabel('Loss')
plt.xlabel('Epoch number')
plt.legend()
plt.subplot(2,1,2)
plt.semilogy(epochNumber,lossDataFromCSV, label='Training loss')
plt.semilogy(epochNumber,vallossDataFromCSV, label='Validation loss')
plt.title('Training loss and validation loss vs. epoch number (log scale)')
plt.ylabel('Loss')
plt.xlabel('Epoch number')
plt.legend()
plt.savefig(QC_model_path+'/'+QC_model_name+'/Quality Control/lossCurvePlots.png',bbox_inches='tight',pad_inches=0)
plt.show()
# + [markdown] id="biT9FI9Ri77_"
# ## **5.2. Error mapping and quality metrics estimation**
# ---
#
# <font size = 4>This section will display SSIM maps and RSE maps as well as calculating total SSIM, NRMSE and PSNR metrics for all the images provided in the "Source_QC_folder" and "Target_QC_folder" !
#
# <font size = 4>**1. The SSIM (structural similarity) map**
#
# <font size = 4>The SSIM metric is used to evaluate whether two images contain the same structures. It is a normalized metric and an SSIM of 1 indicates a perfect similarity between two images. Therefore for SSIM, the closer to 1, the better. The SSIM maps are constructed by calculating the SSIM metric in each pixel by considering the surrounding structural similarity in the neighbourhood of that pixel (currently defined as window of 11 pixels and with Gaussian weighting of 1.5 pixel standard deviation, see our Wiki for more info).
#
# <font size=4>**mSSIM** is the SSIM value calculated across the entire window of both images.
#
# <font size=4>**The output below shows the SSIM maps with the mSSIM**
#
# <font size = 4>**2. The RSE (Root Squared Error) map**
#
# <font size = 4>This is a display of the root of the squared difference between the normalized predicted and target or the source and the target. In this case, a smaller RSE is better. A perfect agreement between target and prediction will lead to an RSE map showing zeros everywhere (dark).
#
#
# <font size =4>**NRMSE (normalised root mean squared error)** gives the average difference between all pixels in the images compared to each other. Good agreement yields low NRMSE scores.
#
# <font size = 4>**PSNR (Peak signal-to-noise ratio)** is a metric that gives the difference between the ground truth and prediction (or source input) in decibels, using the peak pixel values of the prediction and the MSE between the images. The higher the score the better the agreement.
#
# <font size=4>**The output below shows the RSE maps with the NRMSE and PSNR values.**
#
#
#
#
# + cellView="form" id="nAs4Wni7VYbq"
#@markdown ##Choose the folders that contain your Quality Control dataset
Source_QC_folder = "" #@param{type:"string"}
Target_QC_folder = "" #@param{type:"string"}
# Create a quality control/Prediction Folder
if os.path.exists(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction"):
shutil.rmtree(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction")
os.makedirs(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction")
# Activate the pretrained model.
model_training = CARE(config=None, name=QC_model_name, basedir=QC_model_path)
# List Tif images in Source_QC_folder
Source_QC_folder_tif = Source_QC_folder+"/*.tif"
Z = sorted(glob(Source_QC_folder_tif))
Z = list(map(imread,Z))
print('Number of test dataset found in the folder: '+str(len(Z)))
# Perform prediction on all datasets in the Source_QC folder
for filename in os.listdir(Source_QC_folder):
img = imread(os.path.join(Source_QC_folder, filename))
predicted = model_training.predict(img, axes='YX')
os.chdir(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction")
imsave(filename, predicted)
def ssim(img1, img2):
return structural_similarity(img1,img2,data_range=1.,full=True, gaussian_weights=True, use_sample_covariance=False, sigma=1.5)
def normalize(x, pmin=3, pmax=99.8, axis=None, clip=False, eps=1e-20, dtype=np.float32):
"""This function is adapted from Martin Weigert"""
"""Percentile-based image normalization."""
mi = np.percentile(x,pmin,axis=axis,keepdims=True)
ma = np.percentile(x,pmax,axis=axis,keepdims=True)
return normalize_mi_ma(x, mi, ma, clip=clip, eps=eps, dtype=dtype)
def normalize_mi_ma(x, mi, ma, clip=False, eps=1e-20, dtype=np.float32):#dtype=np.float32
"""This function is adapted from Martin Weigert"""
if dtype is not None:
x = x.astype(dtype,copy=False)
mi = dtype(mi) if np.isscalar(mi) else mi.astype(dtype,copy=False)
ma = dtype(ma) if np.isscalar(ma) else ma.astype(dtype,copy=False)
eps = dtype(eps)
try:
import numexpr
x = numexpr.evaluate("(x - mi) / ( ma - mi + eps )")
except ImportError:
x = (x - mi) / ( ma - mi + eps )
if clip:
x = np.clip(x,0,1)
return x
def norm_minmse(gt, x, normalize_gt=True):
"""This function is adapted from <NAME>"""
"""
normalizes and affinely scales an image pair such that the MSE is minimized
Parameters
----------
gt: ndarray
the ground truth image
x: ndarray
the image that will be affinely scaled
normalize_gt: bool
set to True of gt image should be normalized (default)
Returns
-------
gt_scaled, x_scaled
"""
if normalize_gt:
gt = normalize(gt, 0.1, 99.9, clip=False).astype(np.float32, copy = False)
x = x.astype(np.float32, copy=False) - np.mean(x)
#x = x - np.mean(x)
gt = gt.astype(np.float32, copy=False) - np.mean(gt)
#gt = gt - np.mean(gt)
scale = np.cov(x.flatten(), gt.flatten())[0, 1] / np.var(x.flatten())
return gt, scale * x
# Open and create the csv file that will contain all the QC metrics
with open(QC_model_path+"/"+QC_model_name+"/Quality Control/QC_metrics_"+QC_model_name+".csv", "w", newline='') as file:
writer = csv.writer(file)
# Write the header in the csv file
writer.writerow(["image #","Prediction v. GT mSSIM","Input v. GT mSSIM", "Prediction v. GT NRMSE", "Input v. GT NRMSE", "Prediction v. GT PSNR", "Input v. GT PSNR"])
# Let's loop through the provided dataset in the QC folders
for i in os.listdir(Source_QC_folder):
if not os.path.isdir(os.path.join(Source_QC_folder,i)):
print('Running QC on: '+i)
# -------------------------------- Target test data (Ground truth) --------------------------------
test_GT = io.imread(os.path.join(Target_QC_folder, i))
# -------------------------------- Source test data --------------------------------
test_source = io.imread(os.path.join(Source_QC_folder,i))
# Normalize the images wrt each other by minimizing the MSE between GT and Source image
test_GT_norm,test_source_norm = norm_minmse(test_GT, test_source, normalize_gt=True)
# -------------------------------- Prediction --------------------------------
test_prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction",i))
# Normalize the images wrt each other by minimizing the MSE between GT and prediction
test_GT_norm,test_prediction_norm = norm_minmse(test_GT, test_prediction, normalize_gt=True)
# -------------------------------- Calculate the metric maps and save them --------------------------------
# Calculate the SSIM maps
index_SSIM_GTvsPrediction, img_SSIM_GTvsPrediction = ssim(test_GT_norm, test_prediction_norm)
index_SSIM_GTvsSource, img_SSIM_GTvsSource = ssim(test_GT_norm, test_source_norm)
#Save ssim_maps
img_SSIM_GTvsPrediction_32bit = np.float32(img_SSIM_GTvsPrediction)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/SSIM_GTvsPrediction_'+i,img_SSIM_GTvsPrediction_32bit)
img_SSIM_GTvsSource_32bit = np.float32(img_SSIM_GTvsSource)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/SSIM_GTvsSource_'+i,img_SSIM_GTvsSource_32bit)
# Calculate the Root Squared Error (RSE) maps
img_RSE_GTvsPrediction = np.sqrt(np.square(test_GT_norm - test_prediction_norm))
img_RSE_GTvsSource = np.sqrt(np.square(test_GT_norm - test_source_norm))
# Save SE maps
img_RSE_GTvsPrediction_32bit = np.float32(img_RSE_GTvsPrediction)
img_RSE_GTvsSource_32bit = np.float32(img_RSE_GTvsSource)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/RSE_GTvsPrediction_'+i,img_RSE_GTvsPrediction_32bit)
io.imsave(QC_model_path+'/'+QC_model_name+'/Quality Control/RSE_GTvsSource_'+i,img_RSE_GTvsSource_32bit)
# -------------------------------- Calculate the RSE metrics and save them --------------------------------
# Normalised Root Mean Squared Error (here it's valid to take the mean of the image)
NRMSE_GTvsPrediction = np.sqrt(np.mean(img_RSE_GTvsPrediction))
NRMSE_GTvsSource = np.sqrt(np.mean(img_RSE_GTvsSource))
# We can also measure the peak signal to noise ratio between the images
PSNR_GTvsPrediction = psnr(test_GT_norm,test_prediction_norm,data_range=1.0)
PSNR_GTvsSource = psnr(test_GT_norm,test_source_norm,data_range=1.0)
writer.writerow([i,str(index_SSIM_GTvsPrediction),str(index_SSIM_GTvsSource),str(NRMSE_GTvsPrediction),str(NRMSE_GTvsSource),str(PSNR_GTvsPrediction),str(PSNR_GTvsSource)])
# All data is now processed saved
Test_FileList = os.listdir(Source_QC_folder) # this assumes, as it should, that both source and target are named the same
plt.figure(figsize=(20,20))
# Currently only displays the last computed set, from memory
# Target (Ground-truth)
plt.subplot(3,3,1)
plt.axis('off')
img_GT = io.imread(os.path.join(Target_QC_folder, Test_FileList[-1]))
plt.imshow(img_GT, norm=simple_norm(img_GT, percent = 99))
plt.title('Target',fontsize=15)
# Source
plt.subplot(3,3,2)
plt.axis('off')
img_Source = io.imread(os.path.join(Source_QC_folder, Test_FileList[-1]))
plt.imshow(img_Source, norm=simple_norm(img_Source, percent = 99))
plt.title('Source',fontsize=15)
#Prediction
plt.subplot(3,3,3)
plt.axis('off')
img_Prediction = io.imread(os.path.join(QC_model_path+"/"+QC_model_name+"/Quality Control/Prediction/", Test_FileList[-1]))
plt.imshow(img_Prediction, norm=simple_norm(img_Prediction, percent = 99))
plt.title('Prediction',fontsize=15)
#Setting up colours
cmap = plt.cm.CMRmap
#SSIM between GT and Source
plt.subplot(3,3,5)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imSSIM_GTvsSource = plt.imshow(img_SSIM_GTvsSource, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imSSIM_GTvsSource,fraction=0.046, pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsSource,3)),fontsize=14)
plt.ylabel('SSIM maps',fontsize=20, rotation=0, labelpad=75)
#SSIM between GT and Prediction
plt.subplot(3,3,6)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imSSIM_GTvsPrediction = plt.imshow(img_SSIM_GTvsPrediction, cmap = cmap, vmin=0,vmax=1)
plt.colorbar(imSSIM_GTvsPrediction,fraction=0.046, pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('mSSIM: '+str(round(index_SSIM_GTvsPrediction,3)),fontsize=14)
#Root Squared Error between GT and Source
plt.subplot(3,3,8)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imRSE_GTvsSource = plt.imshow(img_RSE_GTvsSource, cmap = cmap, vmin=0, vmax = 1)
plt.colorbar(imRSE_GTvsSource,fraction=0.046,pad=0.04)
plt.title('Target vs. Source',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsSource,3))+', PSNR: '+str(round(PSNR_GTvsSource,3)),fontsize=14)
#plt.title('Target vs. Source PSNR: '+str(round(PSNR_GTvsSource,3)))
plt.ylabel('RSE maps',fontsize=20, rotation=0, labelpad=75)
#Root Squared Error between GT and Prediction
plt.subplot(3,3,9)
#plt.axis('off')
plt.tick_params(
axis='both', # changes apply to the x-axis and y-axis
which='both', # both major and minor ticks are affected
bottom=False, # ticks along the bottom edge are off
top=False, # ticks along the top edge are off
left=False, # ticks along the left edge are off
right=False, # ticks along the right edge are off
labelbottom=False,
labelleft=False)
imRSE_GTvsPrediction = plt.imshow(img_RSE_GTvsPrediction, cmap = cmap, vmin=0, vmax=1)
plt.colorbar(imRSE_GTvsPrediction,fraction=0.046,pad=0.04)
plt.title('Target vs. Prediction',fontsize=15)
plt.xlabel('NRMSE: '+str(round(NRMSE_GTvsPrediction,3))+', PSNR: '+str(round(PSNR_GTvsPrediction,3)),fontsize=14)
plt.savefig(full_QC_model_path+'Quality Control/QC_example_data.png',bbox_inches='tight',pad_inches=0)
qc_pdf_export()
# + [markdown] id="69aJVFfsqXbY"
# # **6. Using the trained model**
#
# ---
#
# <font size = 4>In this section the unseen data is processed using the trained model (in section 4). First, your unseen images are uploaded and prepared for prediction. After that your trained model from section 4 is activated and finally saved into your Google Drive.
# + [markdown] id="tcPNRq1TrMPB"
# ## **6.1. Generate prediction(s) from unseen dataset**
# ---
#
# <font size = 4>The current trained model (from section 4.2) can now be used to process images. If you want to use an older model, untick the **Use_the_current_trained_model** box and enter the name and path of the model to use. Predicted output images are saved in your **Result_folder** folder as restored image stacks (ImageJ-compatible TIFF images).
#
# <font size = 4>**`Data_folder`:** This folder should contain the images that you want to use your trained network on for processing.
#
# <font size = 4>**`Result_folder`:** This folder will contain the predicted output images.
# + cellView="form" id="Am2JSmpC0frj"
#@markdown ### Provide the path to your dataset and to the folder where the predictions are saved, then play the cell to predict outputs from your unseen images.
Data_folder = "" #@param {type:"string"}
Result_folder = "" #@param {type:"string"}
# model name and path
#@markdown ###Do you want to use the current trained model?
Use_the_current_trained_model = True #@param {type:"boolean"}
#@markdown ###If not, please provide the path to the model folder:
Prediction_model_folder = "" #@param {type:"string"}
#Here we find the loaded model name and parent path
Prediction_model_name = os.path.basename(Prediction_model_folder)
Prediction_model_path = os.path.dirname(Prediction_model_folder)
if (Use_the_current_trained_model):
print("Using current trained network")
Prediction_model_name = model_name
Prediction_model_path = model_path
full_Prediction_model_path = os.path.join(Prediction_model_path, Prediction_model_name)
if os.path.exists(full_Prediction_model_path):
print("The "+Prediction_model_name+" network will be used.")
else:
W = '\033[0m' # white (normal)
R = '\033[31m' # red
print(R+'!! WARNING: The chosen model does not exist !!'+W)
print('Please make sure you provide a valid model path and model name before proceeding further.')
#Activate the pretrained model.
model_training = CARE(config=None, name=Prediction_model_name, basedir=Prediction_model_path)
# creates a loop, creating filenames and saving them
for filename in os.listdir(Data_folder):
img = imread(os.path.join(Data_folder,filename))
restored = model_training.predict(img, axes='YX')
os.chdir(Result_folder)
imsave(filename,restored)
print("Images saved into folder:", Result_folder)
# + [markdown] id="bShxBHY4vFFd"
# ## **6.2. Inspect the predicted output**
# ---
#
#
# + cellView="form" id="6b2t6SLQvIBO"
# @markdown ##Run this cell to display a randomly chosen input and its corresponding predicted output.
# This will display a randomly chosen dataset input and predicted output
random_choice = random.choice(os.listdir(Data_folder))
x = imread(Data_folder+"/"+random_choice)
os.chdir(Result_folder)
y = imread(Result_folder+"/"+random_choice)
plt.figure(figsize=(16,8))
plt.subplot(1,2,1)
plt.axis('off')
plt.imshow(x, norm=simple_norm(x, percent = 99), interpolation='nearest')
plt.title('Input')
plt.subplot(1,2,2)
plt.axis('off')
plt.imshow(y, norm=simple_norm(y, percent = 99), interpolation='nearest')
plt.title('Predicted output');
# + [markdown] id="hvkd66PldsXB"
# ## **6.3. Download your predictions**
# ---
#
# <font size = 4>**Store your data** and ALL its results elsewhere by downloading it from Google Drive and after that clean the original folder tree (datasets, results, trained model etc.) if you plan to train or use new networks. Please note that the notebook will otherwise **OVERWRITE** all files which have the same name.
# + [markdown] id="u4pcBe8Z3T2J"
# #**Thank you for using CARE 2D!**
| Colab_notebooks/CARE_2D_ZeroCostDL4Mic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
arr = np.array([[b'r', b'n', b'b', b'q', b'k', b'b', b'n', b'r'],
[b'p', b'p', b'p', b'p', b'p', b'p', b'p', b'p'],
[b'_', b'_', b'_', b'_', b'_', b'_', b'_', b'_'],
[b'_', b'_', b'_', b'_', b'_', b'_', b'_', b'_'],
[b'_', b'_', b'_', b'_', b'_', b'_', b'_', b'_'],
[b'_', b'_', b'_', b'_', b'_', b'_', b'_', b'_'],
[b'P', b'P', b'P', b'P', b'P', b'P', b'P', b'P'],
[b'R', b'N', b'B', b'Q', b'K', b'B', b'N', b'R']], dtype = np.object_)
arr
final = np.random.permutation(arr.ravel()).reshape(8, 8)
final
dc = {}
dc["P0"] = np.argwhere(final == b"P")[0]
dc["P1"] = np.argwhere(final == b"P")[1]
dc["P2"] = np.argwhere(final == b"P")[2]
np.unique(final, return_counts = True)
b"b".decode("utf-8")
for peca, counts in zip(*np.unique(final, return_counts = True)):
for i in range(counts):
dc[peca.decode("utf-8") + f"{i}"] = np.argwhere(final == peca)[i]
dc
for peca, pos in dc.items():
if peca[0]!= "_":
print(peca, pos)
poss = []
for peca, pos in dc.items():
if peca[0]!= "_" and peca[0].islower():
print(peca, pos)
poss.append(pos)
#cube = bpy.data.objects[peca]
#cube.location = mathutils.Vector((21.0 - pos[0] * 6.0 , 21.0 - pos[1] * 6.0 , 0.0))
len(np.unique(np.array(poss), axis = 1)) == len(np.array(poss))
np.random.normal(0, 1)
2 * np.pi * np.random.random()
| Blender/Blender Aux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="pLUMk3lRVx64"
# **Anamoly Detection using Isolation Forest Algorithm**
#
#
#
#
#
#
#
#
# + id="C6NSeNPqHorE"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 439} id="RN5MTrthRCXW" outputId="899bfe63-e7fe-4b56-84ab-87b6a73f0759"
df=pd.read_csv('Credit.csv')
df=df.drop(columns=['Class'])
df=df.dropna(how='any')
display(df)
# + id="rRvn0lTFR-CG"
X=df.iloc[:,:-1].values
Y=df.iloc[:,-1].values
# + colab={"base_uri": "https://localhost:8080/"} id="RCHkYjrJT2RF" outputId="32831c64-15b6-4eff-a794-29ea866afd0b"
Y=Y.reshape(len(Y),1)
Y
# + colab={"base_uri": "https://localhost:8080/"} id="hfLpDd_BRybm" outputId="d2ec7f33-eb2a-43f7-bc6f-eac33c746aeb"
from sklearn.ensemble import IsolationForest
forest_model=IsolationForest(max_features = 1.0,n_estimators=100, max_samples='auto',contamination=float(0.2))
forest_model.fit(Y)
# + colab={"base_uri": "https://localhost:8080/", "height": 379} id="V6nB0fl-SyHA" outputId="3c1ac34d-5c8a-4e5a-b08a-3cb4875a9a9e"
df['scores']=forest_model.decision_function(Y)
df['anomaly_Value']=forest_model.predict(Y)
df.head(10)
# + [markdown] id="cjduIKnFUhkI"
# Evaluation Of model
# + colab={"base_uri": "https://localhost:8080/"} id="oAm_AO0DUZfZ" outputId="c20f8ebf-aacb-4033-d13e-9d53568d8a65"
oc=len(df[df['Amount']>80])
oc
# + colab={"base_uri": "https://localhost:8080/"} id="S42taTmKVSVM" outputId="2eb8230f-f2f7-49a0-e1e2-f1cd4f73c924"
print("Accuracy percentage:", 100*list(df['anomaly_Value']).count(-1)/(oc))
| IFA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Programming Exercise 2 - Logistic Regression
#
# - [Logistic regression](#Logistic-regression)
# - [Regularized logistic regression](#Regularized-logistic-regression)
# +
# # %load ../../standard_import.txt
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
from scipy.optimize import minimize
from sklearn.preprocessing import PolynomialFeatures
pd.set_option('display.notebook_repr_html', False)
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', 150)
pd.set_option('display.max_seq_items', None)
# #%config InlineBackend.figure_formats = {'pdf',}
# %matplotlib inline
import seaborn as sns
sns.set_context('notebook')
sns.set_style('white')
# -
def loaddata(file, delimeter):
data = np.loadtxt(file, delimiter=delimeter)
print('Dimensions: ',data.shape)
print(data[1:6,:])
return(data)
def plotData(data, label_x, label_y, label_pos, label_neg, axes=None):
# Get indexes for class 0 and class 1
neg = data[:,2] == 0
pos = data[:,2] == 1
# If no specific axes object has been passed, get the current axes.
if axes == None:
axes = plt.gca()
axes.scatter(data[pos][:,0], data[pos][:,1], marker='+', c='k', s=60, linewidth=2, label=label_pos)
axes.scatter(data[neg][:,0], data[neg][:,1], c='y', s=60, label=label_neg)
axes.set_xlabel(label_x)
axes.set_ylabel(label_y)
axes.legend(frameon= True, fancybox = True);
# ### Logistic regression
data = loaddata('data/ex2data1.txt', ',')
X = np.c_[np.ones((data.shape[0],1)), data[:,0:2]]
y = np.c_[data[:,2]]
plotData(data, 'Exam 1 score', 'Exam 2 score', 'Admitted', 'Not admitted')
# #### Logistic regression hypothesis
# #### $$ h_{\theta}(x) = g(\theta^{T}x)$$
# #### $$ g(z)=\frac{1}{1+e^{−z}} $$
def sigmoid(z):
return(1 / (1 + np.exp(-z)))
# Scipy actually has a convenience function which does exactly the same:<BR>
# http://docs.scipy.org/doc/scipy/reference/generated/scipy.special.expit.html#scipy.special.expit
# #### Cost Function
# #### $$ J(\theta) = \frac{1}{m}\sum_{i=1}^{m}\big[-y^{(i)}\, log\,( h_\theta\,(x^{(i)}))-(1-y^{(i)})\,log\,(1-h_\theta(x^{(i)}))\big]$$
# #### Vectorized Cost Function
# #### $$ J(\theta) = \frac{1}{m}\big((\,log\,(g(X\theta))^Ty+(\,log\,(1-g(X\theta))^T(1-y)\big)$$
def costFunction(theta, X, y):
m = y.size
h = sigmoid(X.dot(theta))
J = -1*(1/m)*(np.log(h).T.dot(y)+np.log(1-h).T.dot(1-y))
if np.isnan(J[0]):
return(np.inf)
return(J[0])
# #### Partial derivative
#
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m}\sum_{i=1}^{m} ( h_\theta (x^{(i)})-y^{(i)})x^{(i)}_{j} $$
# #### Vectorized
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m} X^T(g(X\theta)-y)$$
#
def gradient(theta, X, y):
m = y.size
h = sigmoid(X.dot(theta.reshape(-1,1)))
grad =(1/m)*X.T.dot(h-y)
return(grad.flatten())
initial_theta = np.zeros(X.shape[1])
cost = costFunction(initial_theta, X, y)
grad = gradient(initial_theta, X, y)
print('Cost: \n', cost)
print('Grad: \n', grad)
# #### Optimize cost function
res = minimize(costFunction, initial_theta, args=(X,y), method=None, jac=gradient, options={'maxiter':400})
res
# #### Predict
def predict(theta, X, threshold=0.5):
p = sigmoid(X.dot(theta.T)) >= threshold
return(p.astype('int'))
# Student with Exam 1 score 45 and Exam 2 score 85
# Predict using the optimized Theta values from above (res.x)
sigmoid(np.array([1, 45, 85]).dot(res.x.T))
p = predict(res.x, X)
print('Train accuracy {}%'.format(100*sum(p == y.ravel())/p.size))
# #### Decision boundary
plt.scatter(45, 85, s=60, c='r', marker='v', label='(45, 85)')
plotData(data, 'Exam 1 score', 'Exam 2 score', 'Admitted', 'Not admitted')
x1_min, x1_max = X[:,1].min(), X[:,1].max(),
x2_min, x2_max = X[:,2].min(), X[:,2].max(),
xx1, xx2 = np.meshgrid(np.linspace(x1_min, x1_max), np.linspace(x2_min, x2_max))
h = sigmoid(np.c_[np.ones((xx1.ravel().shape[0],1)), xx1.ravel(), xx2.ravel()].dot(res.x))
h = h.reshape(xx1.shape)
plt.contour(xx1, xx2, h, [0.5], linewidths=1, colors='b');
# ### Regularized logistic regression
data2 = loaddata('data/ex2data2.txt', ',')
y = np.c_[data2[:,2]]
X = data2[:,0:2]
plotData(data2, 'Microchip Test 1', 'Microchip Test 2', 'y = 1', 'y = 0')
# #### Polynomials
# Note that this function inserts a column with 'ones' in the design matrix for the intercept.
poly = PolynomialFeatures(6)
XX = poly.fit_transform(data2[:,0:2])
XX.shape
# #### Regularized Cost Function
# #### $$ J(\theta) = \frac{1}{m}\sum_{i=1}^{m}\big[-y^{(i)}\, log\,( h_\theta\,(x^{(i)}))-(1-y^{(i)})\,log\,(1-h_\theta(x^{(i)}))\big] + \frac{\lambda}{2m}\sum_{j=1}^{n}\theta_{j}^{2}$$
# #### Vectorized Cost Function
# #### $$ J(\theta) = \frac{1}{m}\big((\,log\,(g(X\theta))^Ty+(\,log\,(1-g(X\theta))^T(1-y)\big) + \frac{\lambda}{2m}\sum_{j=1}^{n}\theta_{j}^{2}$$
def costFunctionReg(theta, reg, *args):
m = y.size
h = sigmoid(XX.dot(theta))
J = -1*(1/m)*(np.log(h).T.dot(y)+np.log(1-h).T.dot(1-y)) + (reg/(2*m))*np.sum(np.square(theta[1:]))
if np.isnan(J[0]):
return(np.inf)
return(J[0])
# #### Partial derivative
#
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m}\sum_{i=1}^{m} ( h_\theta (x^{(i)})-y^{(i)})x^{(i)}_{j} + \frac{\lambda}{m}\theta_{j}$$
# #### Vectorized
# #### $$ \frac{\delta J(\theta)}{\delta\theta_{j}} = \frac{1}{m} X^T(g(X\theta)-y) + \frac{\lambda}{m}\theta_{j}$$
# ##### $$\text{Note: intercept parameter } \theta_{0} \text{ is not to be regularized}$$
def gradientReg(theta, reg, *args):
m = y.size
h = sigmoid(XX.dot(theta.reshape(-1,1)))
grad = (1/m)*XX.T.dot(h-y) + (reg/m)*np.r_[[[0]],theta[1:].reshape(-1,1)]
return(grad.flatten())
initial_theta = np.zeros(XX.shape[1])
costFunctionReg(initial_theta, 1, XX, y)
# +
fig, axes = plt.subplots(1,3, sharey = True, figsize=(17,5))
# Decision boundaries
# Lambda = 0 : No regularization --> too flexible, overfitting the training data
# Lambda = 1 : Looks about right
# Lambda = 100 : Too much regularization --> high bias
for i, C in enumerate([0, 1, 100]):
# Optimize costFunctionReg
res2 = minimize(costFunctionReg, initial_theta, args=(C, XX, y), method=None, jac=gradientReg, options={'maxiter':3000})
# Accuracy
accuracy = 100*sum(predict(res2.x, XX) == y.ravel())/y.size
# Scatter plot of X,y
plotData(data2, 'Microchip Test 1', 'Microchip Test 2', 'y = 1', 'y = 0', axes.flatten()[i])
# Plot decisionboundary
x1_min, x1_max = X[:,0].min(), X[:,0].max(),
x2_min, x2_max = X[:,1].min(), X[:,1].max(),
xx1, xx2 = np.meshgrid(np.linspace(x1_min, x1_max), np.linspace(x2_min, x2_max))
h = sigmoid(poly.fit_transform(np.c_[xx1.ravel(), xx2.ravel()]).dot(res2.x))
h = h.reshape(xx1.shape)
axes.flatten()[i].contour(xx1, xx2, h, [0.5], linewidths=1, colors='g');
axes.flatten()[i].set_title('Train accuracy {}% with Lambda = {}'.format(np.round(accuracy, decimals=2), C))
# -
| Notebooks/.ipynb_checkpoints/Programming Exercise 2 - Logistic Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Main
# language: python
# name: main
# ---
# + slideshow={"slide_type": "skip"}
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
# + [markdown] slideshow={"slide_type": "slide"}
# # A fun introduction to programming using widgets
#
# A widget is an visual way to control a program
# + [markdown] slideshow={"slide_type": "slide"}
# Let's imagine, for example, that you wanted to write your name 516 times. And then 200. And then 1000. How would you do that?
#
# Here's one way to do it:
# + slideshow={"slide_type": "fragment"}
def repeat_name(name, times):
return print(name*times)
interact(repeat_name, name = 'alex', times = (0,1000));
# + slideshow={"slide_type": "slide"}
# Now imagine you want a perfect line
| Teaching_materials/Basics_presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import pandas as pd
import numpy as np
# import scipy as sp
import matplotlib.pyplot as plt
import seaborn as sns
from edgedroid.models.timings import EmpiricalExecutionTimeModel as Empirical, TheoreticalExecutionTimeModel as Theoretical
from collections import deque
neuros = [0.25, 0.5, 0.75]
fade_dist = 8
samples = 500
rng = np.random.default_rng()
# + pycharm={"name": "#%%\n"}
rows = deque()
for neuro in neuros:
emp = Empirical.from_default_data(neuroticism=neuro, transition_fade_distance=fade_dist)
theo = Theoretical.from_default_data(neuroticism=neuro, transition_fade_distance=fade_dist)
# get data and distributions
emp_data = emp._data_views
theo_dists = theo._dists
for imp_dur_trans in emp_data.keys():
emp_samples = rng.choice(emp_data[imp_dur_trans], replace=True, size=samples)
theo_samples = theo_dists[imp_dur_trans].rvs(size=samples)
impairment, duration, transition = imp_dur_trans
for sample in emp_samples:
rows.append({
"impairment": impairment,
"duration": duration,
"transition": transition,
"neuroticism": neuro,
"sample": sample,
"model": "Empirical"
})
for sample in theo_samples:
rows.append({
"impairment": impairment,
"duration": duration,
"transition": transition,
"neuroticism": neuro,
"sample": sample,
"model": "Theoretical"
})
sample_df = pd.DataFrame(rows)
sample_df
# + pycharm={"name": "#%%\n"}
for (imp, dur, trans), df in sample_df.groupby(["impairment", "duration", "transition"]):
print("Impairment:", imp, "| Duration:", dur, "| Transition:", trans)
fg = sns.displot(
kind="hist",
data=df,
x="sample",
hue="model",
stat="density",
col="neuroticism",
col_wrap=4,
# row="neuroticism",
multiple="dodge",
facet_kws=dict(margin_titles=True, legend_out=False,),
kde=True,
)
plt.show()
# df = sample_df.copy()
# df["state"] = df[["impairment", "duration", "transition", "neuroticism"]].apply(lambda row: str(row.to_dict()), axis=1)
#
# fg = sns.displot(
# kind="hist",
# data=df,
# x="sample",
# col="model",
# # col_wrap=4,
# hue="model",
# row="state",
# facet_kws=dict(margin_titles=True),
# # multiple="dodge",
# stat="density",
# )
# plt.show()
# del df
| analysis/state_distributions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sentinel-2 <img align="right" src="../Supplementary_data/DE_Africa_Logo_Stacked_RGB_small.jpg">
#
# * **Products used:**
# s2a_msil2a,
# s2b_msil2a
#
# ## Background
#
# Sentinel-2 is an Earth observation mission from the EU Copernicus Programme that systematically acquires optical imagery at high spatial resolution (up to 10 m for some bands).
# The mission is based on a constellation of two identical satellites in the same orbit, 180° apart for optimal coverage and data delivery.
# Together, they cover all Earth's land surfaces, large islands, inland and coastal waters every 3-5 days.
#
# Sentinel-2A was launched on 23 June 2015 and Sentinel-2B followed on 7 March 2017.
# Both of the Sentinel-2 satellites carry an innovative wide swath high-resolution multispectral imager with 13 spectral bands.
# For more information on the Sentinel-2 platforms and applications, check out the [European Space Agency website](http://www.esa.int/Applications/Observing_the_Earth/Copernicus/Overview4).
#
# Digital Earth Africa (DE Africa) provides [Sentinel 2, Level 2A](https://earth.esa.int/web/sentinel/technical-guides/sentinel-2-msi/level-2a-processing) surface reflectance data provided by ESA.
# Surface reflectance provides standardised optical datasets by using robust physical models to correct for variations in image radiance values due to atmospheric properties, as well as sun and sensor geometry.
# The resulting stack of surface reflectance grids are consistent over space and time, which is instrumental in identifying and quantifying environmental change.
#
# DE Africa provides one Sentinel-2 surface reflectance product:
#
# 1. **Sentinel-2** (i.e. `s2a_msil2a` & `s2b_msil2a`): These products represent the 'definitive' source of high quality Sentinel-2 surface reflectance data, and are available from the beginning of the Sentinel-2 archive.
# Sentinel-2, Level 2A surface reflectance products have 13 spectral channels and one pixel quality band ([scl](https://earth.esa.int/web/sentinel/technical-guides/sentinel-2-msi/level-2a/algorithm)):
#
# | Sentinel-2 bands | DEAfrica band name | Band number | Central wavelength (nm) | Resolution (m) | Bandwidth (nm) |
# | -----------------|---------------|-------------|-------------------------|----------------|----------------|
# | Coastal aerosol | `coastal_aerosol` | 1 | 443 | 60 | 20 |
# | Blue | `blue` | 2 | 490 | 10 | 65 |
# | Green | `green` | 3 | 560 | 10 | 35 |
# | Red | `red` | 4 | 665 | 10 | 30 |
# | Vegetation red edge | `red_edge_1` | 5 | 705 | 20 | 15 |
# | Vegetation red edge | `red_edge_2` | 6 | 740 | 20 | 15 |
# | Vegetation red edge | `red_edge_3` | 7 | 783 | 20 | 20 |
# | NIR | `nir_1` | 8 | 842 | 10 | 115 |
# | Narrow NIR | `nir_2` | 8A | 865 | 20 | 20 |
# | Water vapour | `water_vapour` | 9 | 945 | 60 | 20 |
# | SWIR | `swir_1` | 11 | 1610 | 20 | 90 |
# | SWIR | `swir_2` | 12 | 2190 | 20 | 180 |
# | SCL | `scl` | 13 | N/A
#
# These bands cover the visible, near-infrared and short-wave infrared wave lengths.
#
# 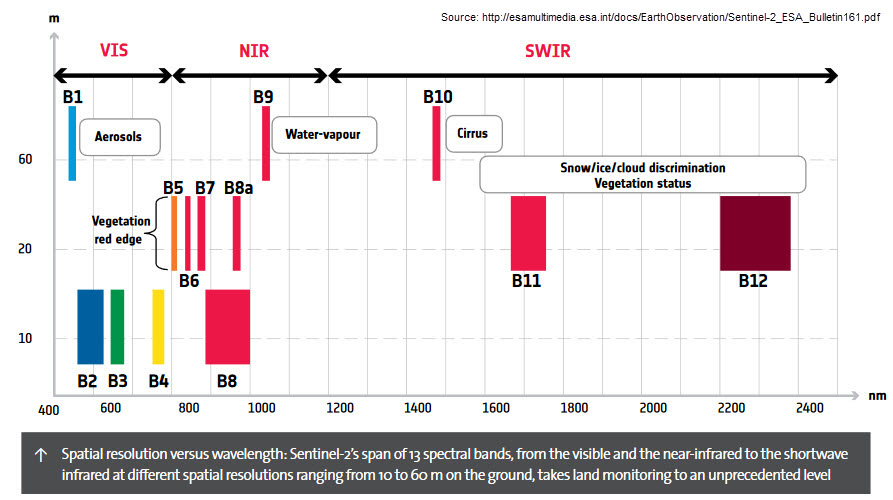
#
# ## Description
#
# This notebook will run through loading in Sentinel-2A and Sentinel-2B satellite images.
# Topics covered include:
#
# * Using the native `dc.load()` function to load in Sentinel-2 data from a single Sentinel-2 satellite
# * Using the `load_ard()` wrapper function to load in a concatenated, sorted, and cloud masked time series from both Sentinel-2A and 2B
#
# ***
# ## Getting started
#
# To run this analysis, run all the cells in the notebook, starting with the "Load packages" cell.
# ### Load packages
# +
import sys
import datacube
sys.path.append("../Scripts")
from deafrica_datahandling import load_ard, mostcommon_crs
from deafrica_plotting import rgb
# -
# ### Connect to the datacube
dc = datacube.Datacube(app="Sentinel_2")
# ## Load Sentinel-2 data from the datacube
#
# We will load **Sentinel-2** data from the Sentinel-2A and Sentinel-2B satellites using two methods.
# Firstly, we will use [dc.load()](../Beginners_guide/03_Loading_data.ipynb) to return a time series of satellite images from a single sensor.
# Secondly, we will load a time series using the [load_ard()](../Frequently_used_code/Using_load_ard.ipynb) function, which is a wrapper function around the dc.load module.
# This function will load all the images from both Sentinel-2A and Sentinel-2B, combine them, and then apply a cloud mask.
# The returned `xarray.Dataset` will contain analysis ready images with the cloudy and invalid pixels masked out.
#
# You can change any of the parameters in the `query` object below to adjust the location, time, projection, or spatial resolution of the returned datasets.
# To learn more about querying, refer to the Beginner's guide notebook on [loading data](../Beginners_guide/03_Loading_data.ipynb).
#
# Sentinel-2 data is stored on file with a range of different coordinate reference systems or CRS (i.e. multiple UTM zones).
# The different satellite bands also have different resolutions (10 m, 20 m and 60 m).
# Because of this, all Sentinel-2 queries need to include the following two query parameters:
#
# * `output_crs`: This sets a consistent CRS that all Sentinel-2 data will be reprojected to, regardless of the UTM zone the individual image is stored in.
# * `resolution`: This sets the resolution that all Sentinel-2 images will be resampled to.
#
# > **Note:** Be aware that setting `resolution` to the highest available resolution (i.e. `(-10, 10)`) will downsample the coarser resolution 20 m and 60 m bands, which may introduce unintended artefacts into your analysis.
# It is typically best practice to set `resolution` to match the lowest resolution band being analysed. For example, if your analysis uses both 10 m and 20 m resolution bands, set `"resolution": (-20, 20)`.
# Create a query object
query = {
'x': (-1.9, -2),
'y': (6.9,7),
"time": ("2018-01", "2018-02"),
"resolution": (-10, 10),
"group_by": "solar_day",
}
# ### Load Sentinel-2 using `dc.load()`
#
# The two **Sentinel-2** products are:
#
# * `s2a_msil2a`
# * `s2b_msil2a`
#
# Here we will load in a time-series of satellite images from only Sentinel-2A.
# To load in images from Sentinel-2B, change the `product` variable to `'s2b_msil2a'`.
# +
# Identify the most common projection system in the input query
output_crs = mostcommon_crs(dc=dc, product='s2a_msil2a', query=query)
#load data
ds = dc.load(product="s2a_msil2a",
output_crs = output_crs,
dask_chunks={},
**query)
print(ds)
# -
# The returned dataset contains all of the bands available for Sentinel-2.
# +
bands = ["blue", "green", "red"]
ds = dc.load(product="s2a_msil2a",
measurements=bands,
output_crs=output_crs,
dask_chunks={},
**query)
print(ds)
# -
# Once the load is complete, we can then analyse or plot the Sentinel-2 data:
rgb(ds, index=1)
# ### Load Sentinel-2 using `load_ard`
#
# This function will load images from both Sentinel-2A and Sentinel-2B, concatenate and sort the observations by time, and apply a cloud mask.
# The result is an analysis ready dataset.
# You can find more information on this function from the [Using load ard](../Frequently_used_code/Using_load_ard.ipynb) notebook.
# +
ds = load_ard(dc=dc,
products=["s2a_msil2a", "s2b_msil2a"],
measurements=bands,
output_crs=output_crs,
dask_chunks={},
**query)
print(ds)
# -
rgb(ds, index=[0,2,-4])
# ***
#
# ## Additional information
#
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
# Digital Earth Africa data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
#
# **Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
# If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/digitalearthafrica/deafrica-sandbox-notebooks).
#
# **Last modified:** Feb 2020
#
# **Compatible datacube version:**
print(datacube.__version__)
# ## Tags
# Browse all available tags on the DE Africa User Guide's [Tags Index]()
# + raw_mimetype="text/restructuredtext" active=""
# **Tags**: :index:`deafrica_datahandling`, :index:`dea_plotting`, :index:`load_ard`, :index:`mostcommon_crs`,:index:`rgb`, :index:`real world`, :Sentinel 2`
| Datasets/Sentinel_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 函数
#
# - 函数可以用来定义可重复代码,组织和简化
# - 一般来说一个函数在实际开发中为一个小功能
# - 一个类为一个大功能
# - 同样函数的长度不要超过一屏
# ## 定义一个函数
#
# def function_name(list of parameters):
#
# do something
# 
# - 以前使用的random 或者range 或者print.. 其实都是函数或者类
# ## 调用一个函数
# - functionName()
# - "()" 就代表调用
# 
# ## 带返回值和不带返回值的函数
# - return 返回的内容
# - return 返回多个值
# - 一般情况下,在多个函数协同完成一个功能的时候,那么将会有返回值
# 
#
# - 当然也可以自定义返回None
# ## EP:
# 
# ## 类型和关键字参数
# - 普通参数
# - 多个参数
# - 默认值参数
# - 不定长参数
# ## 普通参数
# ## 多个参数
# ## 默认值参数
# ## 不定长参数
# ## 变量的作用域
# - 局部变量 local
# - 全局变量 global
# - globals 函数返回一个全局变量的字典,包括所有导入的变量
# - locals() 函数会以字典类型返回当前位置的全部局部变量。
# ## 注意:
# - global :在进行赋值操作的时候需要声明
# - 官方解释:This is because when you make an assignment to a variable in a scope, that variable becomes local to that scope and shadows any similarly named variable in the outer scope.
# - 
x = 1
y = []
def test():
# x = x + 1
y.append(100)
print(x)
test()
print(x)
# ## EP:
# - 定义一个邮箱加密函数,使得输入邮箱后进行ASCII码加密
# - 定义一个判断其是否为闰年的函数
# - 函数的嵌套:定义两个函数A,B,B函数接受A函数的数值并判定是否为奇数或者偶数
# # Homework
# - 1
# 
def test(num):
for i in range(1,num):
print(i * (3*i -1) /2,end=' ')
if i % 10 == 0 and i !=0:
print()
test(100)
# - 2
# 
def sumDigits(n):
sum=0
while n!=0:
sum+=n%10
n=n//10
return sum
n = int(input("Enter an integer:"))
sum=sumDigits(234)
print(sum)
# - 3
# 
def displaySortedNumbers(num1,num2,num3):
i=0
list=[0,0,0]
list[0]=num1
list[1]=num2
list[2]=num3
list.sort()
i=0
while i<3:
print(list[i])
i=i+1
num1=eval(input())
num2=eval(input())
num3=eval(input())
displaySortedNumbers(num1,num2,num3)
# - 4
# 
# - 5
# 
def printChars(ch1,ch2,numberPerLine):
i=0
n=0
ch3=ord(ch1)+1
while ch3<ord(ch2):
print(chr(ch3),end="")
n+=1
if n==numberPerLine:
print("")
n=0
ch3=ch3+1;
ch1=input()
ch2=input()
numberPerLine=int(input())
printChars(ch1,ch2,numberPerLine)
# - 6
# 
def numberOfDaysInAYear(year):
days=365
if (year%4==0 and year%100!=0) or (year%400==0):
days+=1
print(year,"年有",days,"天")
year=2010
while year<=2020:
numberOfDaysInAYear(year)
year+=1
# - 7
# 
# - 8
# 
import math
def sushu(n):
i=2
while i<=math.sqrt(n):
if n%i==0:
break
i+=1
if i<math.sqrt(n):
return 0
else:
return 1
p=2
while p<=31:
if sushu(pow(2,p)-1):
print(pow(2,p)-1,"\t",p)
p+=1
# - 9
# 
# 
import time
time.time()
# - 10
# 
# - 11
# ### 去网上寻找如何用Python代码发送邮件
# +
#需要本机安装SMTP服务器
import smtplib
from email.mime.text import MIMEText
from email.header import Header
sender = '<EMAIL>'
receivers = ['12060<EMAIL>'] # 接收邮件,可设置为你的QQ邮箱或者其他邮箱
# 三个参数:第一个为文本内容,第二个 plain 设置文本格式,第三个 utf-8 设置编码
message = MIMEText('Python 邮件发送测试...', 'plain', 'utf-8')
message['From'] = Header("菜鸟教程", 'utf-8') # 发送者
message['To'] = Header("测试", 'utf-8') # 接收者
subject = 'Python SMTP 邮件测试'
message['Subject'] = Header(subject, 'utf-8')
try:
smtpObj = smtplib.SMTP('localhost')
smtpObj.sendmail(sender, receivers, message.as_string())
print "邮件发送成功"
except smtplib.SMTPException:
print "Error: 无法发送邮件"
| 7.20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
import re
import numpy as np
import pandas as pd
def gather_requests(url):
result = requests.get(url)
if result.status_code != 200:
print('Error')
else:
return result.content
# +
movie_Dict = {'Name':[],'Year':[], 'Level':[], 'Duration':[], 'Genre':[], 'Score':[], 'Director':[], 'Votes':[], 'Gross_in_Million':[]}
movie_List = []
def parse(text):
soup = BeautifulSoup(text, 'html.parser')
content_List = soup.find_all('div', class_ = 'lister-item-content')
for movie in content_List:
movies_Name = movie.find_all('a')[0].text
try:
movies_Year = movie.find_all('span')[1].text
movies_ReleaseYear = re.findall(r'\d{4}', movies_Year)[0]
except:
movies_ReleaseYear = ' '
try:
movies_R_Level = movie.find_all('span', class_ = 'certificate')[0].text.strip()
except:
movies_R_Level = 'NA'
try:
movies_Duration = movie.find_all('span', class_ = 'runtime')[0].text.strip()
except:
movies_Duration = ' '
try:
movies_Genre = movie.find_all('span', class_ = 'genre')[0].text.strip().replace(',','')
except:
movies_Genre = 'Not Classified '
try:
movies_Score = float(movie.find_all('div', class_ = 'inline-block ratings-imdb-rating')[0]['data-value'])
except:
movies_Score = ' '
try:
movies_Director = movie.find_all('p')[2].find_all('a')[0].text
except:
movies_Director = 'No Director Info'
try:
movies_Votes = int(movie.find_all('p')[3].find_all('span')[1]['data-value'])
except:
movies_Votes = ' '
try:
movies_Gross = int(movie.find_all('p')[3].find_all('span')[4]['data-value'].replace(',',''))/1000000
except:
movies_Gross = ' '
movie_Dict['Name'].append(movies_Name)
movie_Dict['Year'].append(movies_ReleaseYear)
movie_Dict['Level'].append(movies_R_Level)
movie_Dict['Duration'].append(movies_Duration)
movie_Dict['Genre'].append(movies_Genre)
movie_Dict['Score'].append(movies_Score)
movie_Dict['Director'].append(movies_Director)
movie_Dict['Votes'].append(movies_Votes)
movie_Dict['Gross_in_Million'].append(movies_Gross)
movie_List.append({'Name': movies_Name, 'Year': movies_ReleaseYear, 'Level':movies_R_Level,
'Duration':movies_Duration, 'Genre': movies_Genre, 'Score': movies_Score,
'Director':movies_Director, 'Votes': movies_Votes, 'Gross_in_Million':movies_Gross})
def main():
numbers = int(input("Please enter a number of page(s) you want to crawl (it should be an integer between 1 to 8048):"))
# Require an integer input between 1 to 8048
if numbers > 0 and numbers < 8049:
for i in range(numbers):
x = 1 + (i * 50)
url = f'https://www.imdb.com/search/title?title_type=feature&start={x}&ref_=adv_nxt'
text = gather_requests(url)
parse(text)
else:
raise ValueError('Number Invalid. Please enter again')
if __name__ == '__main__':
main()
# -
Dataframe = pd.DataFrame(movie_Dict)
Dataframe[[ 'Name','Year','Level', 'Duration', 'Genre','Score','Director','Votes','Gross_in_Million']].to_csv('movies.csv') #export data to csv
movies_table = pd.read_csv("movies.csv")
Dataframe = movies_table[['Name','Year','Level', 'Duration', 'Genre','Score','Director','Votes','Gross_in_Million']]#create table
Dataframe.head(100)
| scripts/418 IMDb final project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.10 ('AI_exam')
# language: python
# name: python3
# ---
# ### Konlpy Pos Tagging
# +
from konlpy.tag import Kkma, Okt
f = open("test_text.txt", 'r', encoding='UTF-8')
data = f.read()
# kkma
def kkmapos(data):
kkma = Kkma()
pos = kkma.pos(data)
return pos
# okt
def oktpos(data):
okt = Okt()
pos = okt.pos(data)
return pos
f.close()
print(kkmapos(data))
print(oktpos(data))
# -
# kkma options
kkma = Kkma()
# print(kkma.morphs(data)) # 형태소 추출
print(kkma.nouns(data)) # 명사 추출
print(kkma.pos(data)) # 품사 부착
print(kkma.sentences(data)) # 문장 추출
# okt options
okt = Okt()
# print(okt.morphs(data)) # 형태소 추출
print(okt.nouns(data)) # 명사 추출
print(okt.pos(data)) # 품사 부착
# ### Mecab Pos Tagging
# +
from konlpy.tag import Mecab
f = open("test_text.txt", 'r', encoding='UTF-8')
data = f.read()
def mecabpos(data):
mecab = Mecab(dicpath=r"C:\mecab\mecab-ko-dic")
pos = mecab.pos(data)
return pos
print(mecabpos(data))
| pos_tagging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pororo import Pororo
age = Pororo(task="age_suitability", lang="en")
age('''
[That stormy night, a tall house, on a rock island somewhere out at sea. The Dursleys, including Harry have moved there, due to the chaos with the letters. The family is sleeping, with Harry on the cold, dirt floor and Dudley sleeping on a sofa. Harry has drawn a birthday cake which reads, Happy Birthday Harry. Harry looks at Dudley's watch, which beeps 12:00.]
Harry: [to himself] Make a wish, Harry. [he blows the "candles" on the drawn birthday cake]
[Suddenly, the door thumps, in sync with the thunder flashing. Harry jumps. The door thumps again and Dudley and Harry jump up and back away. Harry hides behind a wall, and Dudley cowers on a windowsill. Petunia and Vernon appear, with Vernon holding a double barrel gun.]
Vernon: Who's there? Ahh!
[The door bangs again and then falls down, and a giant man appears. As he comes into the clear lighting, he is revealed to be Hagrid.]
Hagrid: Sorry 'bout that. [He puts the door back up]
Vernon: [aiming his gun at him] I demand that you leave at once, sir! You are breaking and entering!
[As Petunia quietly gasps in fear, Hagrid grabs the gun and bends it upwards.]
Hagrid: Dry up, Dursley, you great prune. [The gun fires upwards, blasting a hole in the ceiling, causing the two to shriek in fear. Hagrid notices Dudley.] Boy, I haven't seen you since you was a baby, Harry, but you're a bit more along than I would have expected. Particularly 'round the middle!
Dudley: I-I-I'm not Harry.
[Harry reveals himself]
Harry: I-I am.
Hagrid: Oh, well, of course you are! Got something for ya. 'Fraid I might have sat on it at some point! I imagine that it'll taste fine just the same. Ahh. Baked it myself. [Hands Harry the cake] Words and all. Heh.
[Harry opens cake, which reads "<NAME>" in green frosting.]
Harry: Thank you!
Hagrid: It's not every day that your young man turns eleven, now is it?
[Hagrid sits down on the sofa, takes out an umbrella and points it at the empty fire. Two sparks fly out and the fire starts. The family gasps.]
Harry: [puts cake down] Excuse me, who are you?
Hagrid: Rubeus Hagrid. Keeper of keys and grounds at Hogwarts. Course, you'll know all about Hogwarts.
Harry: Sorry, no.
Hagrid: No? Blimey, Harry, didn't you ever wonder where your mum and dad learned it all?
Harry: Learned what?
Hagrid: You're a wizard, Harry.
''')
| examples/age_suitability.ipynb |