code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia (faststart)
# language: julia
# name: julia-fast
# ---
# # Adaptive integration
using FundamentalsNumericalComputation
# +
f = x -> (x+1)^2*cos((2*x+1)/(x-4.3));
@show exact,errest = quadgk(f,0,4,atol=1e-14,rtol=1e-14); # 'exact' value
# -
# We perform the integration and show the nodes selected underneath the curve.
# +
Q,t = FNC.intadapt(f,0,4,0.001)
@show num_nodes = length(t);
plot(f,0,4,color=:black,legend=:none,
xlabel="x", ylabel="f(x)", title="Adaptive node selection")
plot!(t,f.(t),seriestype=:sticks,m=(:o,2))
# -
# The error turns out to be a bit more than we requested. It's only an estimate, not a guarantee.
@show err = exact - Q;
# Let's see how the number of integrand evaluations and the error vary with the requested tolerance.
# +
table = (tol=[],err=[],n=[])
for tol in 10.0.^(-4:-1:-14)
Q,t = FNC.intadapt(f,0,4,tol)
push!(table.tol,tol)
push!(table.err,exact-Q)
push!(table.n,length(t))
end
pretty_table(table,["tolerance","error","f-evaluations"],backend=:html)
# -
# As you can see, even though the errors are not less than the estimates, the two columns decrease in tandem. If we consider now the convergence not in $h$ (which is poorly defined) but in the number of nodes actually chosen, we come close to the fourth order accuracy of the underlying Simpson scheme.
# +
n = table.n
plot(n,abs.(table.err),m=:o,label="results",
xaxis=(:log10,"number of nodes"), yaxis=(:log10,"error"),
title="Convergence of adaptive quadrature")
order4 = @. 0.01*(n/n[1])^(-4)
plot!(n,order4,l=:dash,label="\$O(n^{-4})\$")
| book/localapprox/demos/adapt-usage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # User Interfaces
# Dask supports several user interfaces [docs](https://docs.dask.org/en/latest/user-interfaces.html):
#
# **High-Level**
# - Arrays: Parallel NumPy
# - Bags: Parallel lists
# - DataFrames: Parallel Pandas
# - Machine Learning : Parallel Scikit-Learn
# - Others from external projects, like XArray
#
# **Low-Level**
# - Delayed: Parallel function evaluation
# - Futures: Real-time parallel function evaluation
#
# Each of these user interfaces employs the same underlying parallel computing machinery, and so has the same scaling,
# diagnostics, resilience, and so on, but each provides a different set of parallel algorithms and programming style.
#
# This document helps you to decide which user interface best suits your needs, and gives some general information that
# applies to all interfaces. The pages linked above give more information about each interface in greater depth.
#
# -
# ## High-Level Collections
# Many people who start using Dask are explicitly looking for a scalable version of NumPy, Pandas, or Scikit-Learn.
# For these situations, the starting point within Dask is usually fairly clear. If you want scalable NumPy arrays,
# then start with Dask array; if you want scalable Pandas DataFrames, then start with Dask DataFrame, and so on.
#
# These high-level interfaces copy the standard interface with slight variations. These interfaces automatically
# parallelize over larger datasets for you for a large subset of the API from the original project.
#
# + pycharm={"name": "#%%\n"}
| User Interfaces.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import copy
import os
import time
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torchvision
from torchvision import datasets, transforms, models
import torch.optim as optim
from torch.optim import lr_scheduler
from torch.autograd import Variable
from torch.utils.data import DataLoader
#device
device=torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
print()
#version
print("PyTorch version: {} Torchvision version: {}".format(torch.__version__, torchvision.__version__))
# +
#load data
data_dir="/home/nsoma/AISuccess/DeepLearning/DLProjects/Diabetic Retnopahty/Original Images/"
input_size=224
#transform my data
data_transform={
"train":transforms.Compose([
transforms.ToTensor(),
transforms.RandomResizedCrop(input_size),
transforms.RandomHorizontalFlip(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])]),
"val":transforms.Compose([
transforms.Resize(input_size),
transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])])}
datasets={x: datasets.ImageFolder(root=os.path.join(data_dir, x), transform=data_transform[x]) for x in ["train", "val"]}
#make my data iteratible
shuffle=True
dataloader_dict={x: DataLoader(datasets[x], shuffle=shuffle, batch_size=4, num_workers=3) for x in ["train", "val"]}
# +
#looking my data
def show(img, title=None):
np_img=img.numpy().transpose(1, 2, 0)
mean=[0.485, 0.456, 0.406]
std=[0.229, 0.224, 0.225]
imgs=np_img*std + mean
imgs=np.clip(imgs, 0, 1)
plt.imshow(imgs)
plt.title(title)
i, o= next(iter(dataloader_dict["val"]))
imgs=torchvision.utils.make_grid(i)
class_name=datasets["val"].classes
show(imgs, title=[class_name[x] for x in o])
# +
#load my model
model_ft=models.resnet101(pretrained=True)
#freeze all parameters
for params in model_ft.parameters():
params.requires_grad=False
"""#handle the auxiliary net
num_ftrs=model_ft.AuxLogits.fc.in_features
model_ft.AuxLogits.fc=nn.Linear(num_ftrs, 5)"""
#handle the primary net
num_ftrs=model_ft.fc.in_features
model_ft.fc=nn.Linear(num_ftrs, 5)
#create my optimizer
params_to_update=model_ft.parameters()
for names, params in model_ft.named_parameters():
if params.requires_grad==True:
# params_to_update.append(params)
print("\t",names)
optimizer_ft=optim.SGD(params_to_update, lr=0.001, momentum=0.9)
scheduler_ft=lr_scheduler.StepLR(optimizer_ft, step_size=3, gamma=0.1)
#create my criterion
criterion_ft=nn.CrossEntropyLoss()
# -
print(model_ft)
def train_model(model, criterion, scheduler, optimizer, dataloader, num_epoch, is_inception=False):
since=time.time()
best_model=copy.deepcopy(model.state_dict())
best_acc=0.0
val_acc_hist, val_loss_hist=[], []
trn_acc_hist, trn_loss_hist=[],[]
for epoch in range(num_epoch):
print("{}/{}".format(epoch, num_epoch-1))
print(" - - "*10)
#every epoch have training and validation
for phase in ["train", "val"]:
if phase=="train":
model.train()
else:
model.eval()
running_loss=0.0
running_acc=0
#iterate over datasets
for inputs, labels in (dataloader[phase]):
inputs=Variable(inputs)
labels=Variable(labels)
inputs=inputs.to(device)
labels=labels.to(device)
#zero parameters
optimizer.zero_grad()
#forward
#track the history
with torch.set_grad_enabled(phase=="train"):
if is_inception and phase=="train":
outputs, aux_outputs=model(inputs)
loss1=criterion(outputs, labels)
loss2=criterion(aux_outputs, labels)
loss=loss1 + loss2*0.4
else:
outputs=model(inputs)
loss=criterion(outputs, labels)
_,pred=torch.max(outputs, 1)
#backwards
if phase=="train":
loss.backward()
optimizer.step()
#statistics
running_loss += loss.item()*inputs.size(0)
running_acc += torch.sum(pred==labels.data).item()
if phase=="val":
scheduler.step()
epoch_loss=running_loss/len(dataloader[phase].dataset)
epoch_acc=running_acc/len(dataloader[phase].dataset)
print("{} Loss: {:.4f} Acc: {:.4f}".format(phase, epoch_loss, epoch_acc))
if phase=="val" and epoch_acc > best_acc:
best_acc=epoch_acc
best_model=copy.deepcopy(model.state_dict())
if phase == "val":
val_acc_hist.append(epoch_acc)
val_loss_hist.append(epoch_loss)
else:
trn_acc_hist.append(epoch_acc)
trn_loss_hist.append(epoch_loss)
acc_loss_hist={
"val_acc_hist":val_acc_hist,
"val_loss_hist":val_loss_hist,
"trn_acc_hist":trn_acc_hist,
"trn_loss_hist":trn_loss_hist
}
print()
time_elipsed=time.time() - since
print("Training complete in {:.0f}m and {:.0f}s".format(time_elipsed//60 , time_elipsed%60))
print("Best val acc: {:.4f}".format(best_acc))
#load the best model
model.load_state_dict(best_model)
return acc_loss_hist , model
model_name="resnet101"
acc_loss_hist , model=train_model(model_ft, criterion_ft, scheduler_ft, optimizer_ft, dataloader_dict, num_epoch=24, is_inception=(model_name=="inception"))
path1="/home/isack/Desktop/isack/Diabetic Retnopathy/saved_models/model_resnet101.pth"
path2="/home/isack/Desktop/isack/Diabetic Retnopathy/saved_models/model_wight_resnet101.pth"
path3="/home/isack/Desktop/isack/Diabetic Retnopathy/saved_models/acc_loss_hist_resnet101.pth"
model_inception_v3=torch.save(model,path1)
model_wight_inception_v3=torch.save(model.state_dict(), path2)
acc_loss_hist_inception_v3=torch.save(acc_loss_hist, path3)
acc_loss_hist=torch.load(path3)
# +
plt.plot(range(1, len(acc_loss_hist["trn_acc_hist"])+1), acc_loss_hist["trn_acc_hist"], 'b', label='training accuracy')
plt.plot(range(1, len(acc_loss_hist["val_acc_hist"])+1), acc_loss_hist["val_acc_hist"], 'r', label='validation accuracy')
plt.legend()
plt.title("TRAINING & VALIDATION ACCURACY")
# +
plt.plot(range(1, len(acc_loss_hist["trn_loss_hist"])+1), acc_loss_hist["trn_loss_hist"], 'b', label='training loss')
plt.plot(range(1, len(acc_loss_hist["val_loss_hist"])+1), acc_loss_hist["val_loss_hist"], 'r', label='validation loss')
plt.legend()
plt.title("TRAINING & VALIDATION LOSSES")
| DiabeticRetnopathy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Models
# #!pip install seaborn
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
pd.set_option('display.width', 500)
pd.set_option('display.max_columns', 100)
df = pd.read_csv("local-olives-cleaned.csv")
df.head()
acidlist=['palmitic', 'palmitoleic', 'stearic', 'oleic', 'linoleic', 'linolenic', 'arachidic', 'eicosenoic']
dfsouth=df[df.regionstring=='South']
dfsouth.head()
# ## Predicting via SVM
dfnew=df[['eicosenoic', 'region', 'regionstring']]
dfnew['linoarch']=(0.969/1022.0)*df.linoleic + (0.245/105.0)*df.arachidic
dfnew.head()
dfnosouth=df[df.regionstring!='South']
dfnosouth.head()
plt.scatter(dfnosouth.linoleic, dfnosouth.arachidic, c=dfnosouth.region, s=50);
# +
from sklearn.cross_validation import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.svm import SVC # "Support Vector Classifier"
def plot_svc_decision_function(clf, ax=None):
"""Plot the decision function for a 2D SVC"""
if ax is None:
ax = plt.gca()
x = np.linspace(plt.xlim()[0], plt.xlim()[1], 30)
y = np.linspace(plt.ylim()[0], plt.ylim()[1], 30)
Y, X = np.meshgrid(y, x)
P = np.zeros_like(X)
for i, xi in enumerate(x):
for j, yj in enumerate(y):
P[i, j] = clf.decision_function([[xi, yj]])
return ax.contour(X, Y, P, colors='k',
levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# -
X = dfnosouth[['linoleic', 'arachidic']]
y = (dfnosouth.regionstring.values=='Sardinia')*1
Xtrain, Xtest, ytrain, ytest = train_test_split(X.values ,y)
clf = SVC(kernel="linear")
clf.fit(Xtrain, ytrain)
plt.scatter(Xtrain[:, 0], Xtrain[:, 1], c=ytrain, s=50, cmap='spring', alpha=0.3)
plot_svc_decision_function(clf, plt.gca())
plt.scatter(clf.support_vectors_[:, 0], clf.support_vectors_[:, 1],
s=200, facecolors='none')
plt.scatter(Xtest[:, 0], Xtest[:, 1], c=ytest, s=50, marker="s", cmap='spring', alpha=0.5);
clf.score(Xtest, ytest)
confusion_matrix(clf.predict(Xtest), ytest)
# ## Allowing for crossovers
# +
from sklearn.model_selection import GridSearchCV
def cv_optimize_svm(X, y, n_folds=10, num_p=50):
#clf = SVC()
#parameters = {"C": np.logspace(-4, 3, num=num_p), "gamma": np.logspace(-4, 3, num=10)}
clf = SVC(kernel="linear", probability=True)
parameters = {"C": np.logspace(-4, 3, num=num_p)}
gs = GridSearchCV(clf, param_grid=parameters, cv=n_folds)
gs.fit(X, y)
return gs
def get_optim_classifier_svm(indf, inacidlist, clon, clonval):
subdf=indf[inacidlist]
subdfstd=(subdf - subdf.mean())/subdf.std()
X=subdfstd.values
y=(indf[clon].values==clonval)*1
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, train_size=0.8)
#Xtrain, Xtest, ytrain, ytest=X,X,y,y
fitted=cv_optimize_svm(Xtrain, ytrain)
return fitted, Xtrain, ytrain, Xtest, ytest
# -
thesvcfit, Xtr, ytr, Xte, yte = get_optim_classifier_svm(dfnosouth, ['linoleic','arachidic'],'regionstring', "Sardinia")
#thesvcfit, Xtr, ytr, Xte, yte = get_optim_classifier_binary(dfsouthns, ['palmitic','palmitoleic'],'area', 3)
thesvcfit.best_estimator_, thesvcfit.best_params_, thesvcfit.best_score_
def plot_svm_new(clf,Xtr,ytr,Xte,yte):
plt.scatter(Xtr[:, 0], Xtr[:, 1], c=ytr, s=50, cmap='spring', alpha=0.5)
plt.scatter(Xte[:, 0], Xte[:, 1], marker='s', c=yte, s=50, cmap='spring', alpha=0.5)
#plt.xlim(-1, 4)
#plt.ylim(-1, 6)
plot_svc_decision_function(clf, plt.gca())
plt.scatter(clf.support_vectors_[:, 0], clf.support_vectors_[:, 1],
s=100, facecolors=None , lw=2, alpha=0.4)
print(dict(kernel="linear",**thesvcfit.best_params_))
clsvc=SVC(**dict(kernel="linear",**thesvcfit.best_params_)).fit(Xtr, ytr)
plot_svm_new(clsvc, Xtr, ytr, Xte, yte)
# The best fit allows for a bigger margin by allowing some inbetween penalization. If we use the standard C=1 in scikit-learn you see that we are allowing for less penalization.
| notebooks/olives-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (reco_full)
# language: python
# name: reco_full
# ---
# <i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
#
# <i>Licensed under the MIT License.</i>
# # Benchmark with Movielens dataset
#
# This illustrative comparison applies to collaborative filtering algorithms available in this repository such as Spark ALS, Surprise SVD, SAR and others using the Movielens dataset. These algorithms are usable in a variety of recommendation tasks, including product or news recommendations.
#
# The main purpose of this notebook is not to produce comprehensive benchmarking results on multiple datasets. Rather, it is intended to illustrate on how one could evaluate different recommender algorithms using tools in this repository.
#
# ## Experimentation setup:
#
# * Objective
# * To compare how each collaborative filtering algorithm perform in predicting ratings and recommending relevant items.
#
# * Environment
# * The comparison is run on a [Azure Data Science Virtual Machine](https://azure.microsoft.com/en-us/services/virtual-machines/data-science-virtual-machines/).
# * The virtual machine size is Standard NC6s_v2 (6 vcpus, 112 GB memory, 1P100 GPU).
# * It should be noted that the single node DSVM is not supposed to run scalable benchmarking analysis. Either scaling up or out the computing instances is necessary to run the benchmarking in an run-time efficient way without any memory issue.
# * **NOTE ABOUT THE DEPENDENCIES TO INSTALL**: This notebook uses CPU, GPU and PySpark algorithms, so make sure you install the `full environment` as detailed in the [SETUP.md](../SETUP.md).
#
# * Datasets
# * [Movielens 100K](https://grouplens.org/datasets/movielens/100k/).
# * [Movielens 1M](https://grouplens.org/datasets/movielens/1m/).
#
# * Data split
# * The data is split into train and test sets.
# * The split ratios are 75-25 for train and test datasets.
# * The splitting is stratified based on items.
#
# * Model training
# * A recommendation model is trained by using each of the collaborative filtering algorithms.
# * Empirical parameter values reported [here](http://mymedialite.net/examples/datasets.html) are used in this notebook. More exhaustive hyper parameter tuning would be required to further optimize results.
#
# * Evaluation metrics
# * Ranking metrics:
# * Precision@k.
# * Recall@k.
# * Normalized discounted cumulative gain@k (NDCG@k).
# * Mean-average-precision (MAP).
# * In the evaluation metrics above, k = 10.
# * Rating metrics:
# * Root mean squared error (RMSE).
# * Mean average error (MAE).
# * R squared.
# * Explained variance.
# * Run time performance
# * Elapsed for training a model and using a model for predicting/recommending k items.
# * The time may vary across different machines.
# ## 0 Globals settings
# +
import sys
sys.path.append("../")
import os
import json
import pandas as pd
import numpy as np
import seaborn as sns
import pyspark
import torch
import fastai
import tensorflow as tf
import surprise
from reco_utils.common.general_utils import get_number_processors
from reco_utils.common.gpu_utils import get_cuda_version, get_cudnn_version
from reco_utils.dataset import movielens
from reco_utils.dataset.python_splitters import python_stratified_split
from benchmark_utils import *
print("System version: {}".format(sys.version))
print("Pandas version: {}".format(pd.__version__))
print("PySpark version: {}".format(pyspark.__version__))
print("Surprise version: {}".format(surprise.__version__))
print("PyTorch version: {}".format(torch.__version__))
print("Fast AI version: {}".format(fastai.__version__))
print("Tensorflow version: {}".format(tf.__version__))
print("CUDA version: {}".format(get_cuda_version()))
print("CuDNN version: {}".format(get_cudnn_version()))
n_cores = get_number_processors()
print("Number of cores: {}".format(n_cores))
# %load_ext autoreload
# %autoreload 2
# -
# ## Parameters
# Run parameters
EPOCHS = 15
# Hide fastai progress bar
hide_fastai_progress_bar()
# fix random seeds to make sure out runs are reproducible
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed_all(SEED)
# +
environments = {
"als": "pyspark",
"sar": "python_cpu",
"svd": "python_cpu",
"fastai": "python_gpu",
"ncf": "python_gpu",
}
metrics = {
"als": ["rating", "ranking"],
"sar": ["ranking"],
"svd": ["rating", "ranking"],
"fastai": ["rating", "ranking"],
"ncf": ["ranking"],
}
# -
# Algorithm parameters
# +
als_params = {
"rank": 10,
"maxIter": EPOCHS,
"implicitPrefs": False,
"alpha": 0.1,
"regParam": 0.05,
"coldStartStrategy": "drop",
"nonnegative": False,
"userCol": DEFAULT_USER_COL,
"itemCol": DEFAULT_ITEM_COL,
"ratingCol": DEFAULT_RATING_COL,
}
sar_params = {
"remove_seen": True,
"similarity_type": "jaccard",
"time_decay_coefficient": 30,
"time_now": None,
"timedecay_formula": True,
"col_user": DEFAULT_USER_COL,
"col_item": DEFAULT_ITEM_COL,
"col_rating": DEFAULT_RATING_COL,
"col_timestamp": DEFAULT_TIMESTAMP_COL,
}
svd_params = {
"n_factors": 200,
"n_epochs": EPOCHS,
"lr_all": 0.005,
"reg_all": 0.02,
"random_state": SEED,
"verbose": False
}
fastai_params = {
"n_factors": 40,
"y_range": [0,5.5],
"wd": 1e-1,
"max_lr": 5e-3,
"epochs": EPOCHS
}
ncf_params = {
"model_type": "NeuMF",
"n_factors": 4,
"layer_sizes": [16, 8, 4],
"n_epochs": EPOCHS,
"batch_size": 1024,
"learning_rate": 1e-3,
"verbose": 10
}
params = {
"als": als_params,
"sar": sar_params,
"svd": svd_params,
"fastai": fastai_params,
"ncf": ncf_params,
}
# -
prepare_training_data = {
"als": prepare_training_als,
"svd": prepare_training_svd,
"fastai": prepare_training_fastai,
"ncf": prepare_training_ncf,
}
prepare_metrics_data = {
"als": lambda train, test: prepare_metrics_als(train, test),
"fastai": lambda train, test: prepare_metrics_fastai(train, test),
}
trainer = {
"als": lambda params, data: train_als(params, data),
"svd": lambda params, data: train_svd(params, data),
"sar": lambda params, data: train_sar(params, data),
"fastai": lambda params, data: train_fastai(params, data),
"ncf": lambda params, data: train_ncf(params, data),
}
rating_predictor = {
"als": lambda model, test: predict_als(model, test),
"svd": lambda model, test: predict_svd(model, test),
"fastai": lambda model, test: predict_fastai(model, test),
}
ranking_predictor = {
"als": lambda model, test, train: recommend_k_als(model, test, train),
"sar": lambda model, test, train: recommend_k_sar(model, test, train),
"svd": lambda model, test, train: recommend_k_svd(model, test, train),
"fastai": lambda model, test, train: recommend_k_fastai(model, test, train),
"ncf": lambda model, test, train: recommend_k_ncf(model, test, train),
}
# +
rating_evaluator = {
"als": lambda test, predictions: rating_metrics_pyspark(test, predictions),
"svd": lambda test, predictions: rating_metrics_python(test, predictions),
"fastai": lambda test, predictions: rating_metrics_python(test, predictions)
}
ranking_evaluator = {
"als": lambda test, predictions, k: ranking_metrics_pyspark(test, predictions, k),
"sar": lambda test, predictions, k: ranking_metrics_python(test, predictions, k),
"svd": lambda test, predictions, k: ranking_metrics_python(test, predictions, k),
"fastai": lambda test, predictions, k: ranking_metrics_python(test, predictions, k),
"ncf": lambda test, predictions, k: ranking_metrics_python(test, predictions, k),
}
# -
def generate_summary(data, algo, k, train_time, time_rating, rating_metrics, time_ranking, ranking_metrics):
summary = {"Data": data, "Algo": algo, "K": k, "Train time (s)": train_time, "Predicting time (s)": time_rating, "Recommending time (s)": time_ranking}
if rating_metrics is None:
rating_metrics = {
"RMSE": np.nan,
"MAE": np.nan,
"R2": np.nan,
"Explained Variance": np.nan,
}
if ranking_metrics is None:
ranking_metrics = {
"MAP": np.nan,
"nDCG@k": np.nan,
"Precision@k": np.nan,
"Recall@k": np.nan,
}
summary.update(rating_metrics)
summary.update(ranking_metrics)
return summary
# ## Benchmark loop
data_sizes = ["100k", "1m"] # Movielens data size: 100k, 1m, 10m, or 20m
algorithms = ["als", "svd", "sar", "ncf", "fastai"]
# +
# %%time
# For each data size and each algorithm, a recommender is evaluated.
cols = ["Data", "Algo", "K", "Train time (s)", "Predicting time (s)", "RMSE", "MAE", "R2", "Explained Variance", "Recommending time (s)", "MAP", "nDCG@k", "Precision@k", "Recall@k"]
df_results = pd.DataFrame(columns=cols)
for data_size in data_sizes:
# Load the dataset
df = movielens.load_pandas_df(
size=data_size,
header=[DEFAULT_USER_COL, DEFAULT_ITEM_COL, DEFAULT_RATING_COL, DEFAULT_TIMESTAMP_COL]
)
print("Size of Movielens {}: {}".format(data_size, df.shape))
# Split the dataset
df_train, df_test = python_stratified_split(df,
ratio=0.75,
min_rating=1,
filter_by="item",
col_user=DEFAULT_USER_COL,
col_item=DEFAULT_ITEM_COL
)
# print("Train set size: {}".format(df_train.shape))
# print("Test set size: {}".format(df_test.shape))
# Loop through the algos
for algo in algorithms:
print("\nComputing {} algorithm on Movielens {}".format(algo, data_size))
# Data prep for training set
train = prepare_training_data.get(algo, lambda x:x)(df_train)
# Get model parameters
model_params = params[algo]
# Train the model
model, time_train = trainer[algo](model_params, train)
# print("Training time: {}".format(time_train))
# Predict and evaluate
# print("\nEvaluating with {}".format(algo))
train, test = prepare_metrics_data.get(algo, lambda x,y:(x,y))(df_train, df_test)
if "rating" in metrics[algo]:
# Predict for rating
preds, time_rating = rating_predictor[algo](model, test)
# print("Rating prediction time: {}".format(time_rating))
# Evaluate for rating
ratings = rating_evaluator[algo](test, preds)
# print("Rating metrics: \n{}".format(json.dumps(ratings, indent=4, sort_keys=True)))
else:
ratings = None
time_rating = np.nan
if "ranking" in metrics[algo]:
# Predict for ranking
top_k_scores, time_ranking = ranking_predictor[algo](model, test, train)
# print("Ranking prediction time: {}".format(time_ranking))
# Evaluate for rating
rankings = ranking_evaluator[algo](test, top_k_scores, DEFAULT_K)
# print("Ranking metrics: \n{}".format(json.dumps(rankings, indent=4, sort_keys=True)))
else:
rankings = None
time_ranking = np.nan
# Record results
summary = generate_summary(data_size, algo, DEFAULT_K, time_train, time_rating, ratings, time_ranking, rankings)
df_results.loc[df_results.shape[0] + 1] = summary
# -
# ## Results
df_results
| benchmarks/movielens.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Raw Voltage Visualization
# <a id="plot"></a>
# ## Raw voltage plot
# 
from simpl_eeg import raw_voltage, eeg_objects
import warnings
warnings.filterwarnings('ignore')
# ```{note}
# Please include the line below in your IDE so that the changes would be simultaneously reflected when you make a change to the python scripts.
# ```
# %load_ext autoreload
# %autoreload 2
# <br>
# ### Define parameters
# A detailed description of all parameters can be found in the `raw_voltage.plot_voltage` docstring:
help(raw_voltage.plot_voltage)
# +
# change values below to values of interest
experiment_path = "../../data/927"
nth_epoch = 0
# -
# <br>
# ### Create epoched data
# For additional options see [Creating EEG Objects](eeg_objects.html#intro) section.
epochs = eeg_objects.Epochs(experiment_path)
epoch = epochs.get_epoch(nth_epoch)
# </br>
# ### Create the raw voltage plot
# #### Generating the plot
# You can create a plot for one epoch or multiple epochs by customizing the epoch object you pass to the function.
# multiple epochs
raw_voltage.plot_voltage(epochs.all_epochs, n_epochs=2);
# single epoch
voltage_plot = raw_voltage.plot_voltage(epoch)
voltage_plot;
# #### Saving the plot
# You can change the file to different formats by changing the format argument in the function. It supports `png`, `pdf`, `svg`.
# ```python
# # change the file path to where you would like to save the file
# file_path = "../../exports/examples/voltage_plot.svg"
#
# voltage_plot.savefig(file_path, format="svg")
# ```
# <br>
| docs/simpl_instructions/_build/html/_sources/raw_voltage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Api Manager Example
from onesaitplatform.apimanager import ApiManagerClient
# ## Create ApiManager
HOST = "development.onesaitplatform.com"
PORT = 443
TOKEN = "b<PASSWORD>"
#client = ApiManagerClient(host=HOST, port=PORT)
client = ApiManagerClient(host=HOST)
# ## Set token
client.setToken(TOKEN)
# ## Find APIs
ok_find, res_find = client.find("RestaurantsAPI", "Created", "analytics")
print("API finded: {}".format(ok_find))
print("Api info:")
print(res_find)
# ## List APIs
ok_list, res_list = client.list("analytics")
print("APIs listed {}".format(ok_list))
print("Apis info:")
for api in res_list:
print(api)
print("*")
# ## Make API request
ok_request, res_request = client.request(method="GET", name="RestaurantsAPI/", version=1, body=None)
print("API request: {}".format(ok_request))
print("Api request:")
print(res_request)
| python-client/examples/ApiManagerClient.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from utils import *
watcha_data = read_data('../reviews/watcha/contents_watcha.txt')
watcha_data[:3]
# %%time
watcha_docs = [tokenize(row[0]) for row in watcha_data]
watcha_docs_no_pos = [remove_pos(docs) for docs in watcha_docs]
# +
from pprint import pprint
pprint(watcha_docs[20])
# +
import nltk
tokens = [t for d in watcha_docs for t in d]
text = nltk.Text(tokens, name="WATCHA")
print(text)
# -
print(len(text.tokens))
print(len(set(text.tokens)))
pprint(text.vocab().most_common(10))
# %%time
from gensim.models import word2vec
model = word2vec.Word2Vec(watcha_docs, size=300, workers=4, min_count=10, iter=100)
save_pickle('../watcha_docs_0427.pickle', watcha_docs)
save_pickle('../watcha_docs_no_pos_0427.pickle', watcha_docs_no_pos)
save_pickle('../nltk_text_watcha_0427.pickle', text)
model.save('../model/review_word2vec_watcha_20180427')
emotion_pair = {'joy': '기쁘다', 'anger': '화나다',
'disgust': '역겹다', 'sadness': '슬프다', 'fear': '무섭다'}
emotion_ko_list = list(emotion_pair.values())
emotion_ko_dic = {
'기쁘다': 0,
'화나다': 1,
'역겹다': 2,
'슬프다': 3,
'무섭다': 4,
}
emotion_ko_list = [tokenize(row) for row in emotion_ko_list]
emotion_ko_list = [row[0] for row in emotion_ko_list]
emotion_ko_list
model.wv.most_similar('깜놀/Noun', topn=100)
file = read_data('../reviews/watcha/watcha_review_2018-04-26.csv')
# %%time
docs_2 = [tokenize(row[0]) for row in file]
# %%time
model_2 = word2vec.Word2Vec(docs_2, size=300, workers=4, min_count=10, iter=100)
save_pickle('../watcha_docs_0427.pickle', watcha_docs)
save_pickle('../watcha_docs_no_pos_0427.pickle', watcha_docs_no_pos)
save_pickle('../nltk_text_watcha_0427.pickle', text)
model.save('../model/review_word2vec_watcha_20180427')
save_pickle('../watcha_docs_0427_02.pickle', docs_2)
model_2.save('../model/review_word2vec_watcha_2_2018427')
| model/watcha_corpus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
# Import API key
weather_api_key = "Your_API_key_here"
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(low=-90.000, high=90.000, size=1500)
lngs = np.random.uniform(low=-180.000, high=180.000, size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# +
# OpenWeatherMap API Key
weather_api_key = "f2a2d5f8e3e39ff0829bcca2e60f5ead"
# Starting URL for Weather Map API Call
url = "http://api.openweathermap.org/data/2.5/weather?units=Imperial&APPID=" + weather_api_key
# +
# Create empty lists to append the API data into lists
city_name = []
cloudiness = []
country = []
date = []
humidity = []
lat = []
lng = []
max_temp = []
wind_speed = []
# Start the call counter
record = 1
# Set print statements
print(f"Beginning Data Retrieval")
print(f"-------------------------------")
# Loop through the cities in the city list
for city in cities:
# Try statement to append calls where value is found
# Not all calls return data as OpenWeatherMap will not have have records in all the cities generated by CityPy module
try:
response = requests.get(f"{url}&q={city}").json()
city_name.append(response["name"])
cloudiness.append(response["clouds"]["all"])
country.append(response["sys"]["country"])
date.append(response["dt"])
humidity.append(response["main"]["humidity"])
max_temp.append(response["main"]["temp_max"])
lat.append(response["coord"]["lat"])
lng.append(response["coord"]["lon"])
wind_speed.append(response["wind"]["speed"])
city_record = response["name"]
print(f"Processing Record {record} | {city_record}")
print(f"{url}&q={city}")
# Increase counter by one
record= record + 1
# Wait a second in loop to not over exceed rate limit of API
time.sleep(1.01)
# If no record found "skip" to next call
except:
print("City not found. Skipping...")
continue
# +
# Create a dictonary with the lists generated
weatherpy_dict = {
"City": city_name,
"Cloudiness":cloudiness,
"Country":country,
"Date":date,
"Humidity": humidity,
"Lat":lat,
"Lng":lng,
"Max Temp": max_temp,
"Wind Speed":wind_speed
}
# Create a dataframe from dictionary
weather_data = pd.DataFrame(weatherpy_dict)
# Display count of weather data values
weather_data.count()
# +
# Save dataframe to a CSV file
weather_data.to_csv('city_weather_data.csv')
# Display dataframe head
weather_data.head()
# +
# Create a scatter plot for each data type
plt.scatter(weather_data["Lat"], weather_data["Max Temp"], marker="o", s=10)
# Label the other graph properties
plt.title("City Latitude vs. Max Temperature (04/10/20)")
plt.ylabel("Max. Temperature (F)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("Plot-Max_Temp_vs_Latitude.png")
# Show plot
plt.show()
# -
# This plot is showing the data points for city latitude and maximum temperature to show if there is a noticeable relationship visually.
# +
# Create a scatter plot for each data type
plt.scatter(weather_data["Lat"], weather_data["Humidity"], marker="o", s=10)
# Label the other graph properties
plt.title("City Latitude vs. Humidity (04/10/20)")
plt.ylabel("Humidity (%)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("Plot-Humidity_vs_Latitude.png")
# Show plot
plt.show()
# -
# This plot is showing the data points for city latitude and humidity to show if there is a noticeable relationship visually.
# +
# Create a scatter plot for each data type
plt.scatter(weather_data["Lat"], weather_data["Cloudiness"], marker="o", s=10)
# Label the other graph properties
plt.title("City Latitude vs. Cloudiness (04/10/20)")
plt.ylabel("Cloudiness (%)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("Plot-Cloudiness_vs_Latitude.png")
# Show plot
plt.show()
# -
# This plot is showing the data points for city latitude and cloudiness to show if there is a noticeable relationship visually.
# +
# Create a scatter plot for each data type
plt.scatter(weather_data["Lat"], weather_data["Wind Speed"], marker="o", s=10)
# Label the other graph properties
plt.title("City Latitude vs. Wind Speed (04/10/20)")
plt.ylabel("Wind Speed (mph)")
plt.xlabel("Latitude")
plt.grid(True)
# Save the figure
plt.savefig("Plot-Wind_Speed_vs_Latitude.png")
# Show plot
plt.show()
# -
# This plot is showing the data points for city latitude and wind speed to show if there is a noticeable relationship visually.
# +
# Create Northern and Southern Hemisphere DataFrames
# Store the boolean criteria in a variable to pass to the dataframe indexing function
crit_north = weather_data.Lat >= 0
crit_south = weather_data.Lat < 0
# Create the northern and southern hemisphere dataframes using boolean indexing from the criteria from above
northern_weather = weather_data[crit_north]
southern_weather = weather_data[crit_south]
# The indexes will not be continuous so they need to be reset with the drop=True argument so we don't make
# the prior index as a column
northern_weather = northern_weather.reset_index(drop=True)
southern_weather = southern_weather.reset_index(drop=True)
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = northern_weather['Lat'].astype('float')
y_values = northern_weather['Max Temp'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Northern Hemisphere - Temperature (F) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Temperature(F)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = southern_weather['Lat'].astype('float')
y_values = southern_weather['Max Temp'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Southern Hemisphere - Temperature (F) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Temperature(F)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# -
# In the northern hemisphere plot there is a strong positive correlation between temperature and latitude as denoted by the r-squared value of 0.83. In the southern hemisphere plot there is a weak positive correlation between temperature and latitude as denoted by the r-squared value of 0.35.
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = northern_weather['Lat'].astype('float')
y_values = northern_weather['Humidity'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Northern Hemisphere - Humidity (%) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = southern_weather['Lat'].astype('float')
y_values = southern_weather['Humidity'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Southern Hemisphere - Humidity (%) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Humidity (%)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# -
# In the northern hemisphere plot there is a very weak positive correlation between humidity and latitude as denoted by the r-squared value of 0.062. In the southern hemisphere plot there is a very weak positive correlation between humidity and latitude as denoted by the r-squared value of 0.13.
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = northern_weather['Lat'].astype('float')
y_values = northern_weather['Cloudiness'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Northern Hemisphere - Cloudiness (%) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = southern_weather['Lat'].astype('float')
y_values = southern_weather['Cloudiness'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Southern Hemisphere - Cloudiness (%) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Cloudiness (%)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# -
# In the northern hemisphere plot there is a very weak positive correlation between cloudiness and latitude as denoted by the r-squared value of 0.0165. In the southern hemisphere plot there is a very weak positive correlation between cloudiness and latitude as denoted by the r-squared value of 0.102.
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = northern_weather['Lat'].astype('float')
y_values = northern_weather['Wind Speed'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Northern Hemisphere - Wind Speed (mph) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# +
# Convert to floats and store latitude and maximum temperature as x and y values
x_values = southern_weather['Lat'].astype('float')
y_values = southern_weather['Wind Speed'].astype('float')
# Run linear regression
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
# Plot scatter plot
plt.scatter(x_values,y_values)
# Plot regression line
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(6,10),fontsize=15,color="red")
# Label plot
plt.title('Southern Hemisphere - Wind Speed (mph) vs. Latitude (04/10/20)')
plt.xlabel('Latitude')
plt.ylabel('Wind Speed (mph)')
# Print r square value
print(f"R squared: {rvalue**2}")
# Show plot
plt.show()
# -
# In the northern hemisphere plot there is a very weak positive correlation between wind speed and latitude as denoted by the r-squared value of 0.046. In the southern hemisphere plot there is a very weak positive correlation between wind speed and latitude as denoted by the r-squared value of 0.069.
| Weather_Py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A Recurrent Network Mechanism of Time Integration in Perceptual Decisions
# Wong, Wang
# Journal of Neuroscience, January 2006, 26(4):1212-1328
# ## Material and Methods
# ### The two-variable network model
# ### Calling Library Fuctions
# +
# LIBRARY
import numpy as np # vector manipulation
import math # math functions
import sys
# THIS IS FOR PLOTTING
# %matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import warnings
warnings.filterwarnings("ignore")
# -
# ### The Reduced Network Model
def H(x):
a=270 # Hz/nA
b=108 # Hz
d=.154 # seconds
f=(a*x-b)/(1-np.exp(-d*(a*x-b)))
return f
x=np.arange(-1,1,0.01)
# ### Neural Circuit
# $$ x_{1}=J_{11}S_1-J_{12}S_2+I_{0}+I_{1}+I_{noise,1}$$
# $$ x_{2}=J_{22}S_2-J_{21}S_1+I_{0}+I_{2}+I_{noise,1}$$
#
# where the synaptic couplings are $J_{11}=0.2609$, $J_{22}=0.2609$, $J_{12}=0.0497$ and $J_{21}=0.0497$.
# $I_{0}=0.3255 nA$ represents external input
#
def total_synaptic_current(S_1,S_2,I_1,I_2,I_noise_1,I_noise_2):
# Synaptic coupling
J_11=0.2609 # nA
J_22=0.2609 # nA
J_12=0.0497 # nA
J_21=0.0497 # nA
I_0=0.3255 # nA
x_1=J_11*S_1-J_12*S_2+I_0+I_1+I_noise_1
x_2=J_22*S_2-J_21*S_1+I_0+I_2+I_noise_2
return x_1, x_2
# ### Background activity
# $$ \tau_{AMPA}\frac{d I_{noise,i}(t)}{dt} =-I_{noise,i}(t)+\eta_i(t)\sqrt{\tau_{AMPA}}\sigma_{noise}$$
def Background_Activity(I_noise):
h=0.1
sigma_noise=0.02 # nA
tau_AMPA=2 #ms
eta_noise=np.random.normal(0,1,1)
k=0#(-(I_noise)+eta_noise*np.sqrt(tau_AMPA)*sigma_noise)
I_noise_new=I_noise+h/tau_AMPA*(-(I_noise+h/2*k)+eta_noise
*np.sqrt(tau_AMPA)*sigma_noise)
return I_noise_new
# ### Network Dynamics
# $$ \frac{d S_{i}}{dt} =-\frac{S_{i}}{\tau_S}+(1-S_{i})\gamma H_{i}$$
# +
def Network_Dynamics(S,x):
h=0.1/1000 #ms
gamma=0.641
tau_S=.100 #s
k=(-S/tau_S+(1-S)*gamma*H(x)/1)
S_new=S+h*(-(S+h/2*k)/tau_S+(1-S+h/2*k)*gamma*H(x))
return S_new
def Network_Dynamics_nuncill(x):
h=0.1/1000 #ms
gamma=0.641
tau_S=.100 #s
S_new=tau_S*gamma*H(x)/(1+tau_S*gamma*H(x))
return S_new
# -
# ### Input Current Target
#
# $$ I_i=J_{A,ext}\mu_0\left(1+ \frac{c'}{100} \right) $$
#
def I_input_1(c_dash):
J_A_ext=5.2/10000 # nA/Hz
mu_0=30 # Hz
I_motion=J_A_ext*mu_0*(1+(c_dash)/100)
return I_motion
# $$ I_2=J_{A,ext}\mu_0\left(1- \frac{c'}{100} \right) $$
def I_input_2(c_dash):
J_A_ext=0.00052 # nA/Hz
mu_0=30 # Hz
I_motion=J_A_ext*mu_0*(1-(c_dash)/100)
return I_motion
# +
h=0.1
time=np.arange(0,100000,h)
J_A_ext=0.00052 # nA/Hz
mu_0=30 # Hz
Vector_coherence=[0.0]
Threshold=15
N=1
RT_coh_hit=np.zeros(len(Vector_coherence))
RT_coh_miss=np.zeros(len(Vector_coherence))
Prob=np.zeros(len(Vector_coherence))
count=0
#fig = plt.figure(figsize=(12,8))
ALL_F_1=0.2*np.ones((N,len(time)))
ALL_F_2=0.2*np.ones((N,len(time)))
I_1=0.0*np.ones(len(time)) # np.zeros((1,len(time)))
I_2=0.0*np.ones(len(time)) # np.zeros((1,len(time)))
Firing_target_1=0*time # np.zeros((1,len(time)))
Firing_target_2=0*time # np.zeros((1,len(time)))
I_noise_1=0.001*np.random.normal(0,1,len(time))
I_noise_2=0.001*np.random.normal(0,1,len(time))
x_1=J_A_ext*mu_0*np.random.uniform(0,1,len(time))
x_2=J_A_ext*mu_0*np.random.uniform(0,1,len(time))
S_1=np.random.uniform(0,1,len(time))#0.1*np.random.uniform(0,1,len(time))# np.random.normal(0,1,len(time))*0.0004
S_2=np.random.uniform(0,1,len(time)) #0.1*np.random.uniform(0,1,len(time)) #np.random.normal(0,1,len(time))*0.0004
S_1[0]=0.02
S_2[0]=0.5
count=0
for i in range (0,len(time)-1):
I_noise_1[i+1]=0*Background_Activity(I_noise_1[i])
I_noise_2[i+1]=0*Background_Activity(I_noise_2[i])
I_1[i+1]=0*I_input_1(c_dash)
I_2[i+1]=0*I_input_1(-c_dash)
x_1[i+1],x_2[i+1]=total_synaptic_current(S_1[i],S_2[i],I_1[i],I_2[i],I_noise_1[i],I_noise_2[i])
S_1[i+1]=Network_Dynamics_nuncill(x_1[i+1])
S_2[i+1]=Network_Dynamics(S_2[i],x_2[i+1])
# -
# ### Plotting
# #### Input
# +
fig = plt.figure(figsize=(8,4))
plt.plot(S_1,S_2,'-',color='blue',label="Right")
#plt.plot(time,Firing_target_2,'-',color='red',label="Left")
plt.grid()
plt.xlabel('time(ms)')
plt.ylabel('Firing Rate (Hz)')
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.savefig('Example_Firing_Pattern.png', bbox_inches='tight',dpi=300)
plt.xlim(0,1)
plt.ylim(0,1)
# plt.
#plt.ylim((30,50))
plt.show()
| OLD VERSIONS/A Network Mechanism of Perceptual Decision Time-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import svgwrite
from svgpathtools import Path, Line, svg2paths, wsvg, path_encloses_pt
random.seed(a = "THIS IS THE GENERATOR STRING", version = 2)
paths, attributes = svg2paths('./bitcoin_coin_plain.svg')
out = 0 + 0j
print([a['id'] for a in attributes])
inner_circle = next(p for p,a in zip(paths, attributes) if a['id'] == 'path4569')
outer_b = next(p for p,a in zip(paths, attributes) if a['id'] == 'path4600')
xmin, xmax, ymin, ymax = inner_circle.bbox()
print(xmin, xmax, ymin, ymax)
pt = ((xmin+xmax)/2) + ((ymin+ymax)/2)*1j
nodes = []
SCALE = 20
SIZE = 22
CONST = (ymax - ymin) / (SIZE / 1.75)
OFFSET = SIZE / 3
svg_document = svgwrite.Drawing(filename = "out.svg", size=(SCALE*SIZE, SCALE*SIZE))
TOP = 1
RIGHT = 2
BOTTOM = 4
LEFT = 8
VISITED = 16
ALL = TOP | RIGHT | BOTTOM | LEFT
matrix = [[(TOP | RIGHT | BOTTOM | LEFT) for _ in range(SIZE)] for _ in range(SIZE)]
def get_coords(x, y, direction):
if direction == TOP and y > 0:
return (x, y - 1)
if direction == BOTTOM and y < SIZE - 1:
return (x, y + 1)
if direction == LEFT and x > 0:
return (x - 1, y)
if direction == RIGHT and x < SIZE - 1:
return (x + 1, y)
return None
def is_visited(x, y):
return matrix[y][x] & VISITED
def opposite(direction):
if direction == TOP:
return BOTTOM
if direction == BOTTOM:
return TOP
if direction == LEFT:
return RIGHT
if direction == RIGHT:
return LEFT
def turn_left(direction):
return LEFT if direction == TOP else (direction >> 1)
def turn_right(direction):
return max(TOP, (direction << 1))
def run(x, y, path, prev_pt, drawing=False):
if matrix[y][x] & VISITED:
return False
matrix[y][x] |= VISITED
options = [TOP, RIGHT, RIGHT, RIGHT, BOTTOM, LEFT, LEFT, LEFT]
random.shuffle(options)
this_pt = xmin + (x - OFFSET)*CONST + 1j * ((y - OFFSET) * CONST + ymin)
this_line = Line(prev_pt, this_pt)
draw_this = path_encloses_pt(this_pt, out, inner_circle) and not path_encloses_pt(this_pt, out, outer_b)
if not draw_this and drawing:
# close the line before us
(t1, t2) = inner_circle.intersect(this_line, justonemode=True) or outer_b.intersect(this_line, justonemode=True)
paths.append(Path(Line(prev_pt, this_line.point(t2[0]))))
elif not drawing and draw_this:
# new line
(t1, t2) = inner_circle.intersect(this_line, justonemode=True) or outer_b.intersect(this_line, justonemode=True)
paths.append(Path(Line(this_pt, this_line.point(t2[0]))))
elif drawing:
paths.append(this_line)
path.push('L{},{}'.format(x * SCALE, y * SCALE))
for opt in options:
# already no wall
if (matrix[y][x] & opt) == 0:
continue
coords = get_coords(x, y, opt)
if coords is None:
continue
x1, y1 = coords
if is_visited(x1, y1):
continue
matrix[y][x] ^= opt
matrix[y1][x1] ^= opposite(opt)
path.push('M{},{}'.format(x * SCALE, y * SCALE))
run(x1, y1, path, this_pt, draw_this)
path.push('Z')
path = svg_document.path('M0,0 ', stroke_width=1, stroke='black', fill='none')
run(0, 0, path, 0+0j)
path.push('Z')
svg_document.add(path)
svg_document.save()
wsvg(paths, nodes=nodes, filename='out-masked.svg')
from IPython.display import SVG
SVG(data=svg_document.tostring())
| generators/b-coin/generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSJeong-me/KOSA-Python_Algorithm/blob/main/Tuesday/math2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="2QX7uijxYp65"
def addition(aa, bb=20):
return aa+bb
# + id="drKmEll0Y12f"
def multi(cc, dd):
return cc*dd
# + id="pxjce8QvZXZ5"
def divided(ee, ff):
y1 = ee//ff
y2 = ee%ff
return y1, y2
# + id="ebMQU0MzY-Qm"
r_val = addition(10)
# + id="fIdPHUTrZJjG"
r_val
# + id="wio4RtJ0ZI3B"
x1, x2 = divided(15, 5)
# + id="L3hbi0kmaBGY"
x1
# + id="jeGWIqibZ9S-"
x2
# + id="2pNkSMjcaC3I"
| Tuesday/math2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import tensorflow as tf
import numpy as np
# +
def ascii2char(list_):
list_ += 97
return [chr(item) for item in list_]
tf.reset_default_graph()
# Variables:
seq_len = 3
cell_size = 10
vocab_size = 26 - 2
embedding_size = vocab_size
# Placeholders:
enc_inp = [tf.placeholder(tf.int32,[None] ) ]
labels = [tf.placeholder(tf.int32,[None] ) ]
weights = [ tf.ones_like(enc_inp, tf.int32) ]#weight of each prediction in the sequence during loss calc.
dec_inp = ([tf.zeros_like(enc_inp[0], dtype=tf.int32, name="GO")] )
# Build Graph:
cell = tf.contrib.rnn.BasicLSTMCell(cell_size)
dec_outputs, states = tf.contrib.legacy_seq2seq.embedding_rnn_seq2seq(enc_inp, dec_inp, cell, vocab_size, vocab_size,
embedding_size, feed_previous=True)
# Optimisation:
loss = tf.reduce_mean( tf.nn.softmax_cross_entropy_with_logits(
labels=tf.one_hot(labels[0], depth=vocab_size, dtype=tf.float32),
logits=dec_outputs,
))
learning_rate = 00.1
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
# Start Session:
sess = tf.Session()
sess.run(tf.global_variables_initializer() )
for epoch_i in range(1000):
X = np.random.choice(vocab_size, size=(seq_len)) #97 is ascii of 'a'
Y = X + 2 #next to next ascii value
feed_dict = {enc_inp[0]:X , labels[0]:Y}
dec_outputs_, dec_inp_ = sess.run([dec_outputs, dec_inp],feed_dict)
loss_,_ = sess.run([loss, optimizer],feed_dict)
if epoch_i%100==0:
print "Iteration: ",epoch_i
print "X: ",ascii2char(X)
print "Prediction: ",ascii2char([logits_t.argmax(axis=1) for logits_t in dec_outputs_][0] )
print "Label: ",ascii2char(Y)
print "loss: ",loss_
print
| Simple seq2seq/predict_next2next_char.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="aibYvylFpXAZ"
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
import numpy as np
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
# + id="RoQJChFGpXAm"
class AdvDetector(nn.Module):
def __init__(self, in_channels, pooling1=False, pooling2=False):
super(AdvDetector, self).__init__()
self.pooling1=pooling1
self.pooling2=pooling2
self.conv1=nn.Conv2d(in_channels=in_channels, out_channels=96,kernel_size=3,stride=1, padding=1)
self.layer2=nn.BatchNorm2d(96)
self.layer3=nn.ReLU()
self.layer4=nn.MaxPool2d(2,2) #1st pooling layer
self.layer5=nn.Conv2d(in_channels=96, out_channels=192,kernel_size=3,stride=1,padding=1)
self.layer6=nn.BatchNorm2d(192)
self.layer7=nn.ReLU()
self.layer8=nn.MaxPool2d(2,2) #2nd pooling layer
self.layer9=nn.Conv2d(in_channels=192, out_channels=192,kernel_size=3,stride=1, padding=1)
self.layer10=nn.BatchNorm2d(192)
self.layer11=nn.ReLU()
self.layer12=nn.Conv2d(in_channels=192,out_channels=2,kernel_size=1,stride=1)
self.adaptive_avg_pool = nn.AdaptiveAvgPool2d((1, 1))
def forward(self,x):
x = self.conv1(x)
x = self.layer2(x)
x = self.layer3(x)
if self.pooling1==True:
x = self.layer4(x)
x = self.layer5(x)
x = self.layer6(x)
x = self.layer7(x)
if self.pooling2==True:
x = self.layer8(x)
x = self.layer9(x)
x = self.layer10(x)
x = self.layer11(x)
x = self.layer12(x)
x = self.adaptive_avg_pool(x)
return x
# + id="q9yaxY3FpXAt" outputId="e369e1d3-f376-4bbb-c638-1d64e23c2290" colab={"base_uri": "https://localhost:8080/"}
net = AdvDetector(64,False,False)
print(net)
net.train()
# + id="bptQ92lupXA0" outputId="8fb65f51-3b24-4521-cefc-047c55ea6e0a" colab={"base_uri": "https://localhost:8080/"}
temp_data = np.random.randn(128, 64, 8, 8)
temp_data = torch.Tensor(temp_data)
out = net(temp_data)
out
# + id="xTudelClpXA6" outputId="518f1bc3-2a50-47cc-93ec-4e0f5c5f93c5" colab={"base_uri": "https://localhost:8080/"}
print(out.shape)
out[1]
# + id="_e2edmY-pXBA" outputId="b05e541a-2d4d-423c-b084-5bd59f20868d" colab={"base_uri": "https://localhost:8080/"}
target = torch.ones((128))
out = out.reshape(128, 2)
print(target.shape, out.shape)
# + id="yctZsBqZpXBG" outputId="03b2c31d-bec7-47d9-daf0-92fe72c09824" colab={"base_uri": "https://localhost:8080/"}
criterion = nn.CrossEntropyLoss()
out = out.float()
loss = criterion(out, target.long())
# loss.backward()
print(loss)
# + id="BOiigL_ApXBN"
file=np.load('c1s_train.npy')
# + id="tVTbD7VepXBT" outputId="d3b38e5d-a468-495d-ab60-a9107aef330b"
print(file)
print(file.shape)
# + id="qZXHObwApXBZ"
#device = torch.device("cuda" if torch.cuda.is_available()
# else "cpu")
device="cpu"
# + id="Rj2Z8K92pXBf"
epochs = 1
steps = 0
running_loss = 0
print_every = 10
train_losses, test_losses = [], []
# + id="sMKaQy9OpXBq"
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters(), lr=0.001)
loss_function = nn.MSELoss()
# + id="Kehcx9AGqLv4"
X_train=temp_data
y_train=target
# + id="NE3k1J3hqMBU"
# + id="n93nZ7ibpXB1"
## train data
class trainData(Dataset):
def __init__(self, X_data, y_data):
self.X_data = X_data
self.y_data = y_data
def __getitem__(self, index):
return self.X_data[index], self.y_data[index]
def __len__ (self):
return len(self.X_data)
train_data = trainData(torch.FloatTensor(X_train),
torch.FloatTensor(y_train))
## test data
class testData(Dataset):
def __init__(self, X_data):
self.X_data = X_data
def __getitem__(self, index):
return self.X_data[index]
def __len__ (self):
return len(self.X_data)
#test_data = testData(torch.FloatTensor(X_test))
# + id="ynzSSaohqzTs"
# + id="4NUj1umuqqXg"
train_loader = DataLoader(dataset=train_data, batch_size=4, shuffle=True)
#test_loader = DataLoader(dataset=test_data, batch_size=1)
# + id="Q9s-sagyp54x"
def accuracy(y_pred, y_test):
y_pred_tag = torch.round(torch.sigmoid(y_pred))
correct_results_sum = (y_pred_tag == y_test).sum().float()
acc = correct_results_sum/y_test.shape[0]
acc = torch.round(acc * 100)
return acc
# + id="NSW_mac1rX2a"
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(net.parameters(), lr=0.001)
loss_function = nn.MSELoss()
# + id="JlF_d8mfrFtt" outputId="dd1a30fa-2e80-45f2-f491-2031eae27fb5" colab={"base_uri": "https://localhost:8080/", "height": 385}
# train
EPOCHS=20
net.train()
for e in range(1, EPOCHS+1):
epoch_loss = 0
epoch_acc = 0
for X_batch, y_batch in train_loader:
X_batch, y_batch = X_batch.to(device), y_batch.to(device)
optimizer.zero_grad()
y_pred = net(X_batch)
loss = criterion(y_pred, y_batch.unsqueeze(1))
acc = accuracy(y_pred, y_batch.unsqueeze(1))
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
print(f'Epoch {e+0:03}: | Loss: {epoch_loss/len(train_loader):.5f} | Acc: {epoch_acc/len(train_loader):.3f}')
# + id="75HK-EyHukNG"
y_pred_list = []
net.eval()
with torch.no_grad():
for X_batch in test_loader:
X_batch = X_batch.to(device)
y_test_pred = net(X_batch)
y_test_pred = torch.sigmoid(y_test_pred)
y_pred_tag = torch.round(y_test_pred)
y_pred_list.append(y_pred_tag.cpu().numpy())
y_pred_list = [a.squeeze().tolist() for a in y_pred_list]
| detector/other_notebooks/adversary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Building your own TensorFlow container
#
# With Amazon SageMaker, you can package your own algorithms that can then be trained and deployed in the SageMaker environment. This notebook guides you through an example using TensorFlow that shows you how to build a Docker container for SageMaker and use it for training and inference.
#
# By packaging an algorithm in a container, you can bring almost any code to the Amazon SageMaker environment, regardless of programming language, environment, framework, or dependencies.
#
# 1. [Building your own TensorFlow container](#Building-your-own-tensorflow-container)
# 1. [When should I build my own algorithm container?](#When-should-I-build-my-own-algorithm-container?)
# 1. [Permissions](#Permissions)
# 1. [The example](#The-example)
# 1. [The presentation](#The-presentation)
# 1. [Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker](#Part-1:-Packaging-and-Uploading-your-Algorithm-for-use-with-Amazon-SageMaker)
# 1. [An overview of Docker](#An-overview-of-Docker)
# 1. [How Amazon SageMaker runs your Docker container](#How-Amazon-SageMaker-runs-your-Docker-container)
# 1. [Running your container during training](#Running-your-container-during-training)
# 1. [The input](#The-input)
# 1. [The output](#The-output)
# 1. [Running your container during hosting](#Running-your-container-during-hosting)
# 1. [The parts of the sample container](#The-parts-of-the-sample-container)
# 1. [The Dockerfile](#The-Dockerfile)
# 1. [Building and registering the container](#Building-and-registering-the-container)
# 1. [Testing your algorithm on your local machine](#Testing-your-algorithm-on-your-local-machine)
# 1. [Part 2: Training and Hosting your Algorithm in Amazon SageMaker](#Part-2:-Training-and-Hosting-your-Algorithm-in-Amazon-SageMaker)
# 1. [Set up the environment](#Set-up-the-environment)
# 1. [Create the session](#Create-the-session)
# 1. [Upload the data for training](#Upload-the-data-for-training)
# 1. [Training On SageMaker](#Training-on-SageMaker)
# 1. [Optional cleanup](#Optional-cleanup)
# 1. [Reference](#Reference)
#
# _or_ I'm impatient, just [let me see the code](#The-Dockerfile)!
#
# ## When should I build my own algorithm container?
#
# You may not need to create a container to bring your own code to Amazon SageMaker. When you are using a framework such as Apache MXNet or TensorFlow that has direct support in SageMaker, you can simply supply the Python code that implements your algorithm using the SDK entry points for that framework. This set of supported frameworks is regularly added to, so you should check the current list to determine whether your algorithm is written in one of these common machine learning environments.
#
# Even if there is direct SDK support for your environment or framework, you may find it more effective to build your own container. If the code that implements your algorithm is quite complex or you need special additions to the framework, building your own container may be the right choice.
#
# Some of the reasons to build an already supported framework container are:
# 1. A specific version isn't supported.
# 2. Configure and install your dependencies and environment.
# 3. Use a different training/hosting solution than provided.
#
# This walkthrough shows that it is quite straightforward to build your own container. So you can still use SageMaker even if your use case is not covered by the deep learning containers that we've built for you.
#
# ## Permissions
#
# Running this notebook requires permissions in addition to the normal `SageMakerFullAccess` permissions. This is because it creates new repositories in Amazon ECR. The easiest way to add these permissions is simply to add the managed policy `AmazonEC2ContainerRegistryFullAccess` to the role that you used to start your notebook instance. There's no need to restart your notebook instance when you do this, the new permissions will be available immediately.
#
# ## The example
#
# In this example we show how to package a custom TensorFlow container with a Python example which works with the CIFAR-10 dataset and uses TensorFlow Serving for inference. However, different inference solutions other than TensorFlow Serving can be used by modifying the docker container.
#
# In this example, we use a single image to support training and hosting. This simplifies the procedure because we only need to manage one image for both tasks. Sometimes you may want separate images for training and hosting because they have different requirements. In this case, separate the parts discussed below into separate Dockerfiles and build two images. Choosing whether to use a single image or two images is a matter of what is most convenient for you to develop and manage.
#
# If you're only using Amazon SageMaker for training or hosting, but not both, only the functionality used needs to be built into your container.
#
# [CIFAR-10]: http://www.cs.toronto.edu/~kriz/cifar.html
#
# ## The presentation
#
# This presentation is divided into two parts: _building_ the container and _using_ the container.
# # Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker
#
# ### An overview of Docker
#
# If you're familiar with Docker already, you can skip ahead to the next section.
#
# For many data scientists, Docker containers are a new technology. But they are not difficult and can significantly simply the deployment of your software packages.
#
# Docker provides a simple way to package arbitrary code into an _image_ that is totally self-contained. Once you have an image, you can use Docker to run a _container_ based on that image. Running a container is just like running a program on the machine except that the container creates a fully self-contained environment for the program to run. Containers are isolated from each other and from the host environment, so the way your program is set up is the way it runs, no matter where you run it.
#
# Docker is more powerful than environment managers like conda or virtualenv because (a) it is completely language independent and (b) it comprises your whole operating environment, including startup commands, and environment variable.
#
# A Docker container is like a virtual machine, but it is much lighter weight. For example, a program running in a container can start in less than a second and many containers can run simultaneously on the same physical or virtual machine instance.
#
# Docker uses a simple file called a `Dockerfile` to specify how the image is assembled. An example is provided below. You can build your Docker images based on Docker images built by yourself or by others, which can simplify things quite a bit.
#
# Docker has become very popular in programming and devops communities due to its flexibility and its well-defined specification of how code can be run in its containers. It is the underpinning of many services built in the past few years, such as [Amazon ECS].
#
# Amazon SageMaker uses Docker to allow users to train and deploy arbitrary algorithms.
#
# In Amazon SageMaker, Docker containers are invoked in a one way for training and another, slightly different, way for hosting. The following sections outline how to build containers for the SageMaker environment.
#
# Some helpful links:
#
# * [Docker home page](http://www.docker.com)
# * [Getting started with Docker](https://docs.docker.com/get-started/)
# * [Dockerfile reference](https://docs.docker.com/engine/reference/builder/)
# * [`docker run` reference](https://docs.docker.com/engine/reference/run/)
#
# [Amazon ECS]: https://aws.amazon.com/ecs/
#
# ### How Amazon SageMaker runs your Docker container
#
# Because you can run the same image in training or hosting, Amazon SageMaker runs your container with the argument `train` or `serve`. How your container processes this argument depends on the container.
#
# * In this example, we don't define an `ENTRYPOINT` in the Dockerfile so Docker runs the command [`train` at training time](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo.html) and [`serve` at serving time](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-inference-code.html). In this example, we define these as executable Python scripts, but they could be any program that we want to start in that environment.
# * If you specify a program as an `ENTRYPOINT` in the Dockerfile, that program will be run at startup and its first argument will be `train` or `serve`. The program can then look at that argument and decide what to do.
# * If you are building separate containers for training and hosting (or building only for one or the other), you can define a program as an `ENTRYPOINT` in the Dockerfile and ignore (or verify) the first argument passed in.
#
# #### Running your container during training
#
# When Amazon SageMaker runs training, your `train` script is run, as in a regular Python program. A number of files are laid out for your use, under the `/opt/ml` directory:
#
# /opt/ml
# ├── input
# │ ├── config
# │ │ ├── hyperparameters.json
# │ │ └── resourceConfig.json
# │ └── data
# │ └── <channel_name>
# │ └── <input data>
# ├── model
# │ └── <model files>
# └── output
# └── failure
#
# ##### The input
#
# * `/opt/ml/input/config` contains information to control how your program runs. `hyperparameters.json` is a JSON-formatted dictionary of hyperparameter names to values. These values are always strings, so you may need to convert them. `resourceConfig.json` is a JSON-formatted file that describes the network layout used for distributed training.
# * `/opt/ml/input/data/<channel_name>/` (for File mode) contains the input data for that channel. The channels are created based on the call to CreateTrainingJob but it's generally important that channels match algorithm expectations. The files for each channel are copied from S3 to this directory, preserving the tree structure indicated by the S3 key structure.
# * `/opt/ml/input/data/<channel_name>_<epoch_number>` (for Pipe mode) is the pipe for a given epoch. Epochs start at zero and go up by one each time you read them. There is no limit to the number of epochs that you can run, but you must close each pipe before reading the next epoch.
#
# ##### The output
#
# * `/opt/ml/model/` is the directory where you write the model that your algorithm generates. Your model can be in any format that you want. It can be a single file or a whole directory tree. SageMaker packages any files in this directory into a compressed tar archive file. This file is made available at the S3 location returned in the `DescribeTrainingJob` result.
# * `/opt/ml/output` is a directory where the algorithm can write a file `failure` that describes why the job failed. The contents of this file are returned in the `FailureReason` field of the `DescribeTrainingJob` result. For jobs that succeed, there is no reason to write this file as it is ignored.
#
# #### Running your container during hosting
#
# Hosting has a very different model than training because hosting is reponding to inference requests that come in via HTTP. In this example, we use [TensorFlow Serving](https://www.tensorflow.org/serving/), however the hosting solution can be customized. One example is the [Python serving stack within the scikit learn example](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/advanced_functionality/scikit_bring_your_own/scikit_bring_your_own.ipynb).
#
# Amazon SageMaker uses two URLs in the container:
#
# * `/ping` receives `GET` requests from the infrastructure. Your program returns 200 if the container is up and accepting requests.
# * `/invocations` is the endpoint that receives client inference `POST` requests. The format of the request and the response is up to the algorithm. If the client supplied `ContentType` and `Accept` headers, these are passed in as well.
#
# The container has the model files in the same place that they were written to during training:
#
# /opt/ml
# └── model
# └── <model files>
#
#
# ### The parts of the sample container
#
# The `container` directory has all the components you need to package the sample algorithm for Amazon SageMager:
#
# .
# ├── Dockerfile
# ├── build_and_push.sh
# └── cifar10
# ├── cifar10.py
# ├── resnet_model.py
# ├── nginx.conf
# ├── serve
# ├── train
#
# Let's discuss each of these in turn:
#
# * __`Dockerfile`__ describes how to build your Docker container image. More details are provided below.
# * __`build_and_push.sh`__ is a script that uses the Dockerfile to build your container images and then pushes it to ECR. We invoke the commands directly later in this notebook, but you can just copy and run the script for your own algorithms.
# * __`cifar10`__ is the directory which contains the files that are installed in the container.
#
# In this simple application, we install only five files in the container. You may only need that many, but if you have many supporting routines, you may wish to install more. These five files show the standard structure of our Python containers, although you are free to choose a different toolset and therefore could have a different layout. If you're writing in a different programming language, you will have a different layout depending on the frameworks and tools you choose.
#
# The files that we put in the container are:
#
# * __`cifar10.py`__ is the program that implements our training algorithm.
# * __`resnet_model.py`__ is the program that contains our Resnet model.
# * __`nginx.conf`__ is the configuration file for the nginx front-end. Generally, you should be able to take this file as-is.
# * __`serve`__ is the program started when the container is started for hosting. It simply launches nginx and loads your exported model with TensorFlow Serving.
# * __`train`__ is the program that is invoked when the container is run for training. Our implementation of this script invokes cifar10.py with our our hyperparameter values retrieved from /opt/ml/input/config/hyperparameters.json. The goal for doing this is to avoid having to modify our training algorithm program.
#
# In summary, the two files you probably want to change for your application are `train` and `serve`.
# ### The Dockerfile
#
# The Dockerfile describes the image that we want to build. You can think of it as describing the complete operating system installation of the system that you want to run. A Docker container running is quite a bit lighter than a full operating system, however, because it takes advantage of Linux on the host machine for the basic operations.
#
# For the Python science stack, we start from an official TensorFlow docker image and run the normal tools to install TensorFlow Serving. Then we add the code that implements our specific algorithm to the container and set up the right environment for it to run under.
#
# Let's look at the Dockerfile for this example.
# !cat container/Dockerfile
# ### Building and registering the container
#
# The following shell code shows how to build the container image using `docker build` and push the container image to ECR using `docker push`. This code is also available as the shell script `container/build-and-push.sh`, which you can run as `build-and-push.sh sagemaker-tf-cifar10-example` to build the image `sagemaker-tf-cifar10-example`.
#
# This code looks for an ECR repository in the account you're using and the current default region (if you're using a SageMaker notebook instance, this is the region where the notebook instance was created). If the repository doesn't exist, the script will create it.
# + language="sh"
#
# # The name of our algorithm
# algorithm_name=sagemaker-tf-cifar10-example
#
# cd container
#
# chmod +x cifar10/train
# chmod +x cifar10/serve
#
# account=$(aws sts get-caller-identity --query Account --output text)
#
# # Get the region defined in the current configuration (default to us-west-2 if none defined)
# region=$(aws configure get region)
# region=${region:-us-west-2}
#
# fullname="${account}.dkr.ecr.${region}.amazonaws.com/${algorithm_name}:latest"
#
# # If the repository doesn't exist in ECR, create it.
#
# aws ecr describe-repositories --repository-names "${algorithm_name}" > /dev/null 2>&1
#
# if [ $? -ne 0 ]
# then
# aws ecr create-repository --repository-name "${algorithm_name}" > /dev/null
# fi
#
# # Get the login command from ECR and execute it directly
# $(aws ecr get-login --region ${region} --no-include-email)
#
# # Build the docker image locally with the image name and then push it to ECR
# # with the full name.
#
# docker build -t ${algorithm_name} .
# docker tag ${algorithm_name} ${fullname}
#
# docker push ${fullname}
# -
# ## Testing your algorithm on your local machine
#
# When you're packaging you first algorithm to use with Amazon SageMaker, you probably want to test it yourself to make sure it's working correctly. We use the [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk) to test both locally and on SageMaker. For more examples with the SageMaker Python SDK, see [Amazon SageMaker Examples](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/sagemaker-python-sdk). In order to test our algorithm, we need our dataset.
# ## Download the CIFAR-10 dataset
# Our training algorithm is expecting our training data to be in the file format of [TFRecords](https://www.tensorflow.org/guide/datasets), which is a simple record-oriented binary format that many TensorFlow applications use for training data.
# Below is a Python script adapted from the [official TensorFlow CIFAR-10 example](https://github.com/tensorflow/models/tree/master/tutorials/image/cifar10_estimator), which downloads the CIFAR-10 dataset and converts them into TFRecords.
# ! python utils/generate_cifar10_tfrecords.py --data-dir=/tmp/cifar-10-data
# There should be three tfrecords. (eval, train, validation)
# ! ls /tmp/cifar-10-data
# ## SageMaker Python SDK Local Training
# To represent our training, we use the Estimator class, which needs to be configured in five steps.
# 1. IAM role - our AWS execution role
# 2. train_instance_count - number of instances to use for training.
# 3. train_instance_type - type of instance to use for training. For training locally, we specify `local`.
# 4. image_name - our custom TensorFlow Docker image we created.
# 5. hyperparameters - hyperparameters we want to pass.
#
# Let's start with setting up our IAM role. We make use of a helper function within the Python SDK. This function throw an exception if run outside of a SageMaker notebook instance, as it gets metadata from the notebook instance. If running outside, you must provide an IAM role with proper access stated above in [Permissions](#Permissions).
# +
from sagemaker import get_execution_role
role = get_execution_role()
# -
# ## Fit, Deploy, Predict
#
# Now that the rest of our estimator is configured, we can call `fit()` with the path to our local CIFAR10 dataset prefixed with `file://`. This invokes our TensorFlow container with 'train' and passes in our hyperparameters and other metadata as json files in /opt/ml/input/config within the container.
#
# After our training has succeeded, our training algorithm outputs our trained model within the /opt/ml/model directory, which is used to handle predictions.
#
# We can then call `deploy()` with an instance_count and instance_type, which is 1 and `local`. This invokes our Tensorflow container with 'serve', which setups our container to handle prediction requests through TensorFlow Serving. What is returned is a predictor, which is used to make inferences against our trained model.
#
# After our prediction, we can delete our endpoint.
#
# We recommend testing and training your training algorithm locally first, as it provides quicker iterations and better debuggability.
# Lets set up our SageMaker notebook instance for local mode.
# !/bin/bash ./utils/setup.sh
# +
from sagemaker.estimator import Estimator
hyperparameters = {'train-steps': 100}
instance_type = 'local'
estimator = Estimator(role=role,
train_instance_count=1,
train_instance_type=instance_type,
image_name='sagemaker-tf-cifar10-example:latest',
hyperparameters=hyperparameters)
estimator.fit('file:///tmp/cifar-10-data')
predictor = estimator.deploy(1, instance_type)
# -
# ## Making predictions using Python SDK
#
# To make predictions, we use an image that is converted using OpenCV into a json format to send as an inference request. We need to install OpenCV to deserialize the image that is used to make predictions.
#
# The JSON reponse will be the probabilities of the image belonging to one of the 10 classes along with the most likely class the picture belongs to. The classes can be referenced from the [CIFAR-10 website](https://www.cs.toronto.edu/~kriz/cifar.html). Since we didn't train the model for that long, we aren't expecting very accurate results.
# ! pip install opencv-python
# +
import cv2
import numpy
from sagemaker.predictor import json_serializer, json_deserializer
image = cv2.imread("data/cat.png", 1)
# resize, as our model is expecting images in 32x32.
image = cv2.resize(image, (32, 32))
data = {'instances': numpy.asarray(image).astype(float).tolist()}
# The request and response format is JSON for TensorFlow Serving.
# For more information: https://www.tensorflow.org/serving/api_rest#predict_api
predictor.accept = 'application/json'
predictor.content_type = 'application/json'
predictor.serializer = json_serializer
predictor.deserializer = json_deserializer
# For more information on the predictor class.
# https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/predictor.py
predictor.predict(data)
# -
predictor.delete_endpoint()
# # Part 2: Training and Hosting your Algorithm in Amazon SageMaker
# Once you have your container packaged, you can use it to train and serve models. Let's do that with the algorithm we made above.
#
# ## Set up the environment
# Here we specify the bucket to use and the role that is used for working with SageMaker.
# S3 prefix
prefix = 'DEMO-tensorflow-cifar10'
# ## Create the session
#
# The session remembers our connection parameters to SageMaker. We use it to perform all of our SageMaker operations.
# +
import sagemaker as sage
sess = sage.Session()
# -
# ## Upload the data for training
#
# We will use the tools provided by the SageMaker Python SDK to upload the data to a default bucket.
# +
WORK_DIRECTORY = '/tmp/cifar-10-data'
data_location = sess.upload_data(WORK_DIRECTORY, key_prefix=prefix)
# -
# ## Training on SageMaker
# Training a model on SageMaker with the Python SDK is done in a way that is similar to the way we trained it locally. This is done by changing our train_instance_type from `local` to one of our [supported EC2 instance types](https://aws.amazon.com/sagemaker/pricing/instance-types/).
#
# In addition, we must now specify the ECR image URL, which we just pushed above.
#
# Finally, our local training dataset has to be in Amazon S3 and the S3 URL to our dataset is passed into the `fit()` call.
#
# Let's first fetch our ECR image url that corresponds to the image we just built and pushed.
# +
import boto3
client = boto3.client('sts')
account = client.get_caller_identity()['Account']
my_session = boto3.session.Session()
region = my_session.region_name
algorithm_name = 'sagemaker-tf-cifar10-example'
ecr_image = '{}.dkr.ecr.{}.amazonaws.com/{}:latest'.format(account, region, algorithm_name)
print(ecr_image)
# +
from sagemaker.estimator import Estimator
hyperparameters = {'train-steps': 100}
instance_type = 'ml.m4.xlarge'
estimator = Estimator(role=role,
train_instance_count=1,
train_instance_type=instance_type,
image_name=ecr_image,
hyperparameters=hyperparameters)
estimator.fit(data_location)
predictor = estimator.deploy(1, instance_type)
# +
image = cv2.imread("data/cat.png", 1)
# resize, as our model is expecting images in 32x32.
image = cv2.resize(image, (32, 32))
data = {'instances': numpy.asarray(image).astype(float).tolist()}
predictor.accept = 'application/json'
predictor.content_type = 'application/json'
predictor.serializer = json_serializer
predictor.deserializer = json_deserializer
predictor.predict(data)
# -
# ## Optional cleanup
# When you're done with the endpoint, you should clean it up.
#
# All of the training jobs, models and endpoints we created can be viewed through the SageMaker console of your AWS account.
predictor.delete_endpoint()
# # Reference
# - [How Amazon SageMaker interacts with your Docker container for training](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo.html)
# - [How Amazon SageMaker interacts with your Docker container for inference](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-inference-code.html)
# - [CIFAR-10 Dataset](https://www.cs.toronto.edu/~kriz/cifar.html)
# - [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk)
# - [Dockerfile](https://docs.docker.com/engine/reference/builder/)
# - [scikit-bring-your-own](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/advanced_functionality/scikit_bring_your_own/scikit_bring_your_own.ipynb)
| advanced_functionality/tensorflow_bring_your_own/tensorflow_bring_your_own.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="s6q3p6nu8M1p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="fc756ea8-cc3f-442c-99f7-6dfc357c9e94" executionInfo={"status": "ok", "timestamp": 1591558015427, "user_tz": -330, "elapsed": 2105, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
from google.colab import drive
drive.mount('/gdrive')
# %cd /gdrive
# + id="-dP0zDo08Rs9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="e05c67da-aa8d-4f43-d241-d6d4472aa730" executionInfo={"status": "ok", "timestamp": 1591558016133, "user_tz": -330, "elapsed": 2766, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
import os
from gensim.models.keyedvectors import KeyedVectors
import numpy as np
os.chdir('/gdrive/My Drive/NLP_Project_Programs/Paper_Implementation')
os.listdir()
# + id="zbY_E8l4Cnov" colab_type="code" colab={}
wt = np.array([1,2,4])
# + id="BgowkFPkBXsW" colab_type="code" colab={}
words_list = ['book', 'paper', 'newspaper', 'report' , 'notebook' ]
# + id="Xytqo3Or8SmD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="16936909-1dc6-4823-c198-fee0e8cb9f58" executionInfo={"status": "ok", "timestamp": 1591558031752, "user_tz": -330, "elapsed": 18318, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
import argparse
import gzip
import math
import numpy
import re
import sys
from copy import deepcopy
isNumber = re.compile(r'\d+.*')
def norm_word(word):
if isNumber.search(word.lower()):
return '---num---'
elif re.sub(r'\W+', '', word) == '':
return '---punc---'
else:
return word.lower()
''' Read all the word vectors and normalize them '''
def read_word_vecs(filename):
wordVectors = {}
if filename.endswith('.gz'): fileObject = gzip.open(filename, 'r')
else: fileObject = open(filename, 'r')
for line in fileObject:
line = line.strip().lower()
word = line.split()[0]
wordVectors[word] = numpy.zeros(len(line.split())-1, dtype=float)
for index, vecVal in enumerate(line.split()[1:]):
wordVectors[word][index] = float(vecVal)
''' normalize weight vector '''
wordVectors[word] /= math.sqrt((wordVectors[word]**2).sum() + 1e-6)
sys.stderr.write("Vectors read from: "+filename+" \n")
return wordVectors
''' Read the PPDB word relations as a dictionary '''
def read_lexicon(filename):
lexicon = {}
for line in open(filename, 'r', encoding="utf8"):
words = line.lower().strip().split()
lexicon[norm_word(words[0])] = [norm_word(word) for word in words[1:]]
return lexicon
''' Write word vectors to file '''
'''
def print_word_vecs(wordVectors, outFileName):
sys.stderr.write('\nWriting down the vectors in '+outFileName+'\n')
outFile = open(outFileName, 'w')
for word, values in wordVectors.items():
outFile.write(word+' ')
for val in wordVectors[word]:
outFile.write('%.4f' %(val)+' ')
outFile.write('\n')
outFile.close()
'''
''' Retrofit word vectors to a lexicon '''
def retrofit(wordVecs, lexicon, numIters):
newWordVecs = deepcopy(wordVecs)
wvVocab = set(newWordVecs.keys())
loopVocab = wvVocab.intersection(set(lexicon.keys()))
count = 0
vec_list = []
for word in loopVocab:
if word in words_list :
count+=1
vec_list.append(wordVecs[word][:10])
word_set_matrix = np.array([vec for vec in vec_list])
matrices = [word_set_matrix]
#print(word_set_matrix)
#return
#if count == 5:
#print ("got all!!!!")
#return
for it in range(numIters):
# loop through every node also in ontology (else just use data estimate)
for word in loopVocab:
wordNeighbours = set(lexicon[word]).intersection(wvVocab)
numNeighbours = len(wordNeighbours)
#no neighbours, pass - use data estimate
if numNeighbours == 0:
continue
# the weight of the data estimate if the number of neighbours
newVec = numNeighbours * wordVecs[word]
# loop over neighbours and add to new vector (currently with weight 1)
for ppWord in wordNeighbours:
newVec += newWordVecs[ppWord]
newWordVecs[word] = newVec/(2*numNeighbours)
temp_list = []
for w in words_list:
temp_list.append(newWordVecs[w][:10])
temp_matrix = np.array([vec for vec in temp_list])
matrices.append(temp_matrix)
#return newWordVecs
return matrices
# if __name__=='__main__':
# parser = argparse.ArgumentParser()
# parser.add_argument("-i", "--input", type=str, default=None, help="Input word vecs")
# parser.add_argument("-l", "--lexicon", type=str, default=None, help="Lexicon file name")
# parser.add_argument("-o", "--output", type=str, help="Output word vecs")
# parser.add_argument("-n", "--numiter", type=int, default=10, help="Num iterations")
# args = parser.parse_args()
# wordVecs = read_word_vecs(args.input)
# lexicon = read_lexicon(args.lexicon)
# numIter = int(args.numiter)
# outFileName = args.output
# ''' Enrich the word vectors using ppdb and print the enriched vectors '''
# print_word_vecs(retrofit(wordVecs, lexicon, numIter), outFileName)
input_vector = 'Vector Representation of Words/3/3_word_vectors.txt'
lexicon = 'lexicons/wordnet-synonyms+.txt'
numiter = 10
#output = 'out_vec_.txt'
wordVecs = read_word_vecs(input_vector)
lexicon = read_lexicon(lexicon)
numIter = int(numiter)
#outFileName = output
''' Enrich the word vectors using ppdb and print the enriched vectors '''
Matrices = retrofit(wordVecs, lexicon, numIter)
# + id="pYxRiuyTJOpG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9adac951-109d-4db8-81e0-02e714f6715f" executionInfo={"status": "ok", "timestamp": 1591558031754, "user_tz": -330, "elapsed": 18299, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
print(len(Matrices))
# + id="tALnJyFvKimx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="59064431-08bc-4acf-8b06-3648fa4f6b5b" executionInfo={"status": "ok", "timestamp": 1591558031756, "user_tz": -330, "elapsed": 18255, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
min_arr = []
max_arr = []
for i in range(5):
min_arr.append(np.amin(Matrices[i]))
max_arr.append(np.amax(Matrices[i]))
print(min(min_arr))
print(max(max_arr))
# + id="_At5ACJiMq3q" colab_type="code" colab={}
# + id="vnbq9Fj7Ls49" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="0a9d5220-e27b-4e23-83fa-745277010136" executionInfo={"status": "ok", "timestamp": 1591558031762, "user_tz": -330, "elapsed": 18212, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
import matplotlib.pyplot as plt
import matplotlib.colors
import seaborn as sns
import imageio
from IPython.display import HTML
# + id="shCEc3QDLalp" colab_type="code" colab={}
my_cmap = matplotlib.colors.LinearSegmentedColormap.from_list("", ["red","yellow","green"])
# + id="EenGpfPOI32Y" colab_type="code" colab={}
def plot_heat_map(iter):
fig = plt.figure(figsize = (10,5))
sns.heatmap(Matrices[iter],annot = True,cmap = my_cmap, vmin = -0.5, vmax = 0.5)
plt.title("ITERATION-"+str(iter))
fig.canvas.draw()
image = np.frombuffer(fig.canvas.tostring_rgb(),dtype = 'uint8')
image = image.reshape(fig.canvas.get_width_height()[::-1] + (3,))
return image
# + id="WD200SVgJAqH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="78447361-1bee-4336-b6a3-12b8d1302903" executionInfo={"status": "ok", "timestamp": 1591558379894, "user_tz": -330, "elapsed": 7803, "user": {"displayName": "<NAME>16b179", "photoUrl": "", "userId": "14049130637097040281"}}
imageio.mimsave('./wordvecs_viz.gif', [plot_heat_map(i) for i in range(0,len(Matrices))],fps =1)
# + id="kbOYwcmqOtjw" colab_type="code" colab={"resources": {"http://localhost:8080/wordvecs_viz.gif": {"data": "<KEY>", "ok": false, "headers": [["content-length", "1449"], ["content-type", "text/html; charset=utf-8"]], "status": 404, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 37} outputId="c48e0539-3d0f-406c-caf2-73af14969aa9" executionInfo={"status": "ok", "timestamp": 1591558384606, "user_tz": -330, "elapsed": 4453, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
from IPython import display
HTML('<img src="wordvecs_viz.gif">')
# + id="iwiLKQhm8Sjg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="85b4db86-1e29-4aed-b023-948d6d7065d8" executionInfo={"status": "ok", "timestamp": 1591558039553, "user_tz": -330, "elapsed": 25857, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
a = np.array([[1,2,3],[11,2,23],[33,444,55]])
print(a)
# + id="wfLUkSNn8Sg0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="738de29d-8ba1-41fc-b13e-95dc8fa4d2a4" executionInfo={"status": "ok", "timestamp": 1591558039554, "user_tz": -330, "elapsed": 25833, "user": {"displayName": "<NAME> me16b179", "photoUrl": "", "userId": "14049130637097040281"}}
a = [3]
b = []
b.append(3)
print(a)
print(b)
# + id="yse_G4PJ8SeT" colab_type="code" colab={}
# + id="svj40E808SOj" colab_type="code" colab={}
| Paper_Implementation/GIF_heatmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/carloslme/Practical-Deep-Learning-Book/blob/master/Chapter_15_Keeping_Time%2C_Scheduling_Tasks%2C_and_Launching_Programs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="0n3nWfAZSi67"
# # The time Module
# Your computer’s system clock is set to a specific date, time, and time zone. The built-in time module allows your Python programs to read the system clock for the current time. The `time.time()` and `time.sleep()` functions are the most useful in the time module.
# + [markdown] id="MbU_ZYGYSyNz"
# ## The time.time() Function
# The `time.time()` function returns the number of seconds since that moment as a float value. (Recall that a float is just a number with a decimal point.) This number is called an epoch timestamp
# + colab={"base_uri": "https://localhost:8080/"} id="J59yEpLeSQVe" outputId="76d11ebb-3c3f-440c-bd26-ae53a195c72f"
import time
time.time()
# + [markdown] id="DVktEJsbTHau"
# The return value is how many seconds have passed between the Unix epoch and the moment `time.time()` was called.
#
# Epoch timestamps can be used to profile code, that is, measure how long a piece of code takes to run. If you call `time.time()` at the beginning of the code block you want to measure and again at the end, you can subtract the first timestamp from the second to find the elapsed time between those two calls.
# + colab={"base_uri": "https://localhost:8080/"} id="GIytqfwbTBT8" outputId="55525d16-afbc-4a33-c6dd-09fa816b78d0"
import time
def calcProd():
# Calculate the product of the first 100,000 numbers.
product = 1
for i in range(1, 100000):
product = product * i
return product
startTime = time.time()
prod = calcProd()
endTime = time.time()
print('The result is %s digits long.' % (len(str(prod))))
print('Took %s seconds to calculate.' % (endTime - startTime))
# + [markdown] id="9kNldaFqWqj8"
# ##The time.sleep() Function
# If you need to pause your program for a while, call the `time.sleep()` function and pass it the number of seconds you want your program to stay paused.
# + colab={"base_uri": "https://localhost:8080/"} id="jWmhBpg2U6y4" outputId="3d266f6f-a875-46bc-e929-9aa062030b89"
import time
for i in range(3):
print('Tick')
time.sleep(1)
print('Tock')
time.sleep(1)
# + id="84okim3gW43_"
time.sleep(5)
# + [markdown] id="IgHhg9CgXBBy"
# The `time.sleep()` function will block —that is, it will not return and release your program to execute other code—until after the number of seconds you passed to `time.sleep()` has elapsed.
#
# Be aware that pressing CTRL -C will not interrupt `time.sleep()` calls in IDLE. IDLE waits until the entire pause is over before raising the KeyboardInterrupt exception. To work around this problem, instead of having a single `time.sleep(30) `call to pause for 30 seconds, use a for loop to make 30 calls to `time.sleep(1)`
# + colab={"base_uri": "https://localhost:8080/", "height": 180} id="u8OvW16jXAB-" outputId="efe35e8f-22c1-4c4b-c024-5c4960175461"
for i in range(30):
time.sleep(1)
# + [markdown] id="FymooTSAXXuv"
# If you press CTRL -C sometime during these 30 seconds, you should see the KeyboardInterrupt exception thrown right away.
# + id="OlhgFFzyXUYt"
| Chapter_15_Keeping_Time,_Scheduling_Tasks,_and_Launching_Programs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
# Ler um arquivo de configuração e aplicar como filtro
config_file = '/mnt/sadatalakevcbr/config/Engemix/dualmining_taglist.csv'
# lê arquivo de configuração com a lista de Tags
config_tag_list = [ row[0] for row in spark.read.format("csv").option("header",True).load(config_file).collect()]
# Lê dados e aplica filtro
df_cleansed_tags = spark.read.format("delta").load(source_path)\
.filter(col("tag").isin(config_tag_list))
| pyspark/pyspark_read_config_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Project Euler: Problem 1
# + [markdown] nbgrader={}
# If we list all the natural numbers below 10 that are multiples of 3 or 5, we get 3, 5, 6 and 9. The sum of these multiples is 23.
#
# Find the sum of all the multiples of 3 or 5 below 1000.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
sum_of_multiples=0
for i in range (0,1000):
if i%3==0 or i%5==0:
sum_of_multiples = sum_of_multiples + i
print (sum_of_multiples)
# -
# I set sum_of_multiples equal to zero in order to start my sum at zero. I then used a for loop to loop through all the numbers in the range 0-1000. Then used an if statement, if a number in this range is evenly divisible by three or evenly divisble by 5 add that number to the sum_of_multiples. Then I printed the final sum_of_multiples.
# + deletable=false nbgrader={"checksum": "6e498cbe102f8b3c1bc4ebd777bcc952", "grade": true, "grade_id": "projecteuler1", "points": 10}
# This cell will be used for grading, leave it at the end of the notebook.
| assignments/assignment01/ProjectEuler1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from cortex import Cortex
import pydash
from cortex_common.types import EntityEvent, ListAttributeValue, DimensionalAttributeValue, StringAttributeValue
cortex= Cortex.client()
builder = cortex.builder()
# -
df = pd.read_csv("../../data/sample-member-profile.csv", dtype={'member_adress_zip':'str', 'profile_id' : 'str'})
df = df.astype('object')
list_members = df['member_id'][0:5]
df2 = df[df["member_id"] .isin(list_members) ]
df.head()
list_columns = df.columns.tolist()
cortex_schema_version = 'accelerators/Member'
cortex_schema_version
event_list = [
[EntityEvent(event = i,entityId = df['member_id'][idx],entityType = cortex_schema_version,properties = {'value': df[i][idx]}) for i in list_columns]
for idx in df.index]
event_list[0:10]
for event in event_list:
try:
profile = builder.profiles(cortex_schema_version).with_events(event).build()
print(profile)
except Exception as e:
print('Error:', e)
df1 = pd.read_csv('../../data/sample-alert-data.csv', dtype={ 'member_id' : 'str'})
df2 = pd.read_csv('../../data/sample-alert-acted.csv', dtype={ 'member_id' : 'str'})
df1.head()
df1 = df1.astype('object')
# +
member_id_list1 = []
event_list1 = []
for idx in df1.index:
if df1.iloc[idx]['member_id'] in member_id_list1:
break
a = df1[df1['member_id'] == df1.iloc[idx]['member_id']]
list_alerts = a.to_dict('records')
val_strings = []
for i in list_alerts:
i.pop('member_id')
i.pop('alert')
val = StringAttributeValue(value=i['alert_id'])
val_strings.append(val)
list_alerts = ListAttributeValue(value=val_strings)
event = EntityEvent(
event = 'alerts_generated',
entityId = df1['member_id'][idx],
entityType = cortex_schema_version,
properties = dict(list_alerts))
event_list1.append(event)
member_id_list1.append(df1.iloc[idx]['member_id'])
print(event_list1)
try:
profile = builder.profiles(cortex_schema_version).with_events(event_list1).build()
print(profile)
except Exception as e:
print('Error:', e)
# -
df2 = df2.astype('object')
df2.head()
# +
from cortex_common.types.attribute_values import Dimension, StringAttributeValue, CounterAttributeValue
member_id_list2 = []
event_list2 = []
for idx in df2.index:
if df2.iloc[idx]['member_id'] in member_id_list2:
break
a = df2[df2['member_id'] == df2.iloc[idx]['member_id']]
list_alerts = a.to_dict('records')
list_alerts_viewed = [pydash.merge(pydash.pick(alert, 'alert_id'), pydash.pick(alert,'alert_viewed')) for alert in list_alerts]
list_alerts_acted = [pydash.merge(pydash.pick(alert, 'alert_id'), pydash.pick(alert,'alert_acted')) for alert in list_alerts]
viewed = [ a["alert_id"] for a in list_alerts_viewed if a["alert_viewed"] == 1]
acted = [ a["alert_id"] for a in list_alerts_acted if a["alert_acted"] == 1]
#
attribute_for_alerts_viewed = DimensionalAttributeValue(value=[
Dimension(
dimensionId=alert["alert_id"],
dimensionValue=CounterAttributeValue(value=1 if alert["alert_id"] in viewed else 0),
)
for alert in list_alerts
], contextOfDimension=StringAttributeValue.detailed_schema_type().outerType,contextOfDimensionValue=CounterAttributeValue.detailed_schema_type())
attribute_for_alerts_acted = DimensionalAttributeValue(value=[
Dimension(
dimensionId=alert["alert_id"],
dimensionValue=CounterAttributeValue(value=1 if alert["alert_id"] in acted else 0),
)
for alert in list_alerts
], contextOfDimension=StringAttributeValue.detailed_schema_type().outerType,contextOfDimensionValue=CounterAttributeValue.detailed_schema_type().outerType)
# print(attribute_for_alerts_veiwed)
# print(list_alerts)
# list_alerts_viewed = ListAttributeValue(value=list_alerts_viewed)
# list_alerts_acted = ListAttributeValue(value=list_alerts_acted)
event_alert_viewed = EntityEvent(
event = 'alerts_viewed',
entityId = df2['member_id'][idx],
entityType = cortex_schema_version,
properties = dict(attribute_for_alerts_viewed))
event_alert_acted = EntityEvent(
event = 'alerts_acted',
entityId = df2['member_id'][idx],
entityType = cortex_schema_version,
properties = dict(attribute_for_alerts_acted))
event_list2.append(event_alert_viewed)
event_list2.append(event_alert_acted)
member_id_list2.append(df2.iloc[idx]['member_id'])
# print(event_list2)
print(event_list2)
try:
profile = builder.profile(cortex_schema_version).with_events(event_list2).build()
except Exception as e:
print('Error:', e)
# +
if ':' not in cortex_schema_version:
schema_version = cortex.profile_schema(cortex_schema_version).latest()._version
cortex_schema_version = "{}:{}".format(cortex_schema_version,schema_version)
profile_attrs = cortex.profile('1234567').latest("accelerators/Member")
# -
profile_attrs
| notebooks/name_parse/build_member_profile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
# -
ds = xr.open_mfdataset("/project/amp/jcaron/CPC_Tminmax/tmax.*.nc")
#ds = xr.open_mfdataset("/project/amp/akwilson/1979-2005/tmax.*.nc")
de = xr.open_mfdataset("/project/amp/brianpm/TemperatureExtremes/Regridded/f.e13.FAMIPC5CN.ne30_ne30.beta17.t3.cam.h1.TREFMXAV.19650101-20051231.regrid.nc")
tmax1 = de['TREFMXAV'].compute()
# +
#checker = (de['time.month'] == 12) & (de['time.day'] == 31)
#last_dec31 = np.asscalar(np.argwhere(checker.values == 1)[-1])
#print(last_dec31)
#trim_time = de.isel(time=slice(0,last_dec31+1))
#trim_time['time']
# -
tmax1_avg_by_month = tmax1.groupby('time.month').mean(dim='time')
month_names = ["JAN", "FEB", "MAR", "APR", "MAY", "JUN", "JUL", "AUG", "SEP", "OCT", "NOV", "DEC"]
lons, lats = np.meshgrid(de['lon'], de['lat']) # this creates arrays of longitude and latitude
# +
fig, ax = plt.subplots(figsize=(12,12), nrows=3, ncols=4, subplot_kw={"projection":ccrs.Robinson()}, constrained_layout=True)
for i, a in enumerate(ax.ravel()):
im = a.contourf(lons, lats, tmax1_avg_by_month.isel(month=i), transform=ccrs.PlateCarree(), cmap='inferno_r')
a.coastlines()
cbar = fig.colorbar(im, ax=a, shrink=0.5, orientation='horizontal')
# reduce ticks:
clr_ticks = cbar.get_ticks()
cbar.set_ticks(clr_ticks[::2])
# make the map global rather than have it zoom in to
# the extents of any plotted data
a.set_global()
a.set_title(month_names[i], loc='left', fontsize=10)
# +
fig0, ax0 = plt.subplots(subplot_kw={"projection":ccrs.Robinson()})
im0 = ax0.contourf(lons, lats, tmax1_avg_by_month.isel(month=8), transform=ccrs.PlateCarree(), cmap='inferno_r')
ax0.coastlines()
cbar0 = fig0.colorbar(im0, ax=ax0, shrink=0.5, orientation='horizontal')
print(dir(cbar0))
# reduce ticks:
clr_ticks = cbar0.get_ticks()
print(clr_ticks)
cbar0.set_ticks(clr_ticks[::2])
# # make the map global rather than have it zoom in to
# # the extents of any plotted data
ax0.set_global()
ax0.set_title(month_names[0], loc='left', fontsize=10)
# -
cmich = (43. + 35/60, 360. - (84. + 46/60))
atlanta = (33. + 45/60, 360. - (84. + 23/60))
atlanta_tmax = tmax.sel(lat=atlanta[0], lon=atlanta[1], method='nearest').isel(time=slice(0,-1))
atlanta_tmax.std()
atlanta_tmax['time']#[-1]
time = ds['time']
atlantaug = atlanta_tmax[(time.dt.month == 8)]
# +
# print (de['time'])
fig100, ax103 = plt.subplots(figsize=(10,3))
xax = np.arange(len(augatlanta_f['time']))
ax103.plot(xax, augatlanta_f)
fig104, ax104 = plt.subplots(figsize=(10,3))
gb = list(augatlanta_f.groupby('time.year'))
# print(gb)
xax = np.arange(len(gb[0][1]['time']))
#[ax104.plot(xax, gb[i][1], ('0.4')) for i,_ in enumerate(gb)]
plt.xlabel('Day')
plt.ylabel('Temperature[F]')
plt.title('Atlanta August Tmax Data from 1965 to 2005 ')
plt.axhline(augatlanta_f.mean(), color='k', linestyle='dashed', linewidth=1)
# -
blah = [gb[i][1] for i,_ in enumerate(gb)]
[i.shape for i in blah]
gb[0][1]['time']
# gb --> list of (year, data) pairs (tuples)
# gb[0] is the first (year, data)
# gb[0][1] is the 2nd entry => data in that pair That's a DataArray
# +
atlantaF= atlantaug1 -273.15
def degC_to_degF(C):
return atlantaF * (9./5.) + 32.
# -
augatlanta_f = degC_to_degF(atlantaF)
cmichjan_f.plot()
# +
result = plt.hist(cmichjan_f, bins=20, color='c', edgecolor='k', alpha=0.65)
plt.axvline(x=40.16, color='k', linestyle='dashed', linewidth=1)
plt.title('CMU January Tmax Data')
plt.xlabel('Temperature [F]')
plt.ylabel('Count')
plt.show()
#plt.text(1, 2, x=65.46)
#fig.savefig("/project/amp/akwilson/testdata/atlantajan_figure_001.png")
# -
np.nanquantile(augatlanta_f,.50)
np.nanquantile(augcmich_f,.50)
| copy_akwilson/19792005Hist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Goals
#
# ### 1. Estimating training time before running starting training is helpful when
# - You are using 3rd party GPU providors suchs as kaggle, colab which have limited resource
# - You planning to trin on a large dataset
# # Table of Contents
#
#
# ## [0. Install](#0)
#
#
# ## [1. Setup Default Params](#1)
#
#
# ## [2. Estimate Training Time](#2)
# <a id='0'></a>
# # Install Monk
#
# - git clone https://github.com/Tessellate-Imaging/monk_v1.git
#
# - cd monk_v1/installation && pip install -r requirements_cu9.txt
# - (Select the requirements file as per OS and CUDA version)
# !git clone https://github.com/Tessellate-Imaging/monk_v1.git
# Select the requirements file as per OS and CUDA version
# !cd monk_v1/installation && pip install -r requirements_cu9.txt
# ## Dataset - Pokemon Classification
# - https://www.kaggle.com/lantian773030/pokemonclassification
# ! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1rtUxa0o6e5YiUn8LHf6W7Rf3vuA5LEzU' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1rtUxa0o6e5YiUn8LHf6W7Rf3vuA5LEzU" -O pokemonclassification.zip && rm -rf /tmp/cookies.txt
# ! unzip -qq pokemonclassification.zip
# # Imports
# Monk
import os
import sys
sys.path.append("monk_v1/monk/");
#Using mxnet-gluon backend
from gluon_prototype import prototype
# ### Creating and managing experiments
# - Provide project name
# - Provide experiment name
gtf = prototype(verbose=1);
gtf.Prototype("sample-experiment-1", "sample-project-1");
# <a id='1'></a>
# # Setup Default Params
gtf.Default(dataset_path="pokemonclassification/train",
model_name="resnet18_v1",
num_epochs=5);
# <a id='2'></a>
# # Estimate Training Time
gtf.Estimate_Train_Time(num_epochs=50);
gtf.Estimate_Train_Time(num_epochs=500);
| study_roadmaps/1_getting_started_roadmap/7_extra_features/2) Estimate training time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def joker (x):
return x ** 2
print (joker(2))
# -
class student(object):
print('hello')
student()
# object 不是一个参数
class jxh(object):
def __init__(self,a,b):
print(a+b,a-b,a*b,a/b,a//b,a%b)
jxh(6,4)
class year(object):
def __init__(self,y):
if y>18:
print('岛国爱情片')
else:
print('动画片')
year(14)
# +
class year(object):
def __init__(self,y):
if y>18 and y<20:
print('4级片')
if y>20 and y<50:
print('岛国')
if y>50:
print('别看了')
year(19)
# +
class year(object):
def __init__(self,y):
if 18<=y<=20:
print('4级片')
elif 20<y<=50:
print('岛国')
else:
print('别看了')
year(19)
# -
class Jxh(object):
def __init__(self,a,b):
self.a = a
self.b = b
def func1(self):
if self.a >40:
print('老年')
elif self.a >18:
print('中年')
else:
print('青年')
def func2(self):
if self.b == "男":
print("男人")
elif self.b == "女":
print("女人")
else:
print("未知")
jxh = Jxh(12,"女人")
jxh.func1()
jxh.func2()
a=[1,5,3]
a.sort()
print(a)
# +
class Jxh(object):
def __init__(self,a):
self.a = a
def nly(self):
for i in range(2,self.a):
if self.a % i == 0:
print('非素数')
break
else:
print('素数')
name = Jxh(12)
name.nly()
# -
import numpy as np
res = np.random.choice(['典韦','赵云','鲁班'])
print(res)
class jy(object):
def __init__(self,a):
self.a = a
gh =input('》》')
if gh == ('人机'):
print('人机')
elif gh == ('多人对战'):
print('多人对战')
else:
print('开战')
def choose(self):
print()
name=jy('人机')
name.choose()
# +
class Jxh(object):
def __init__(self):
print('输入密码')
self.res = input()
def b (self) :
print('请再次输入密码')
red = input()
if self.res == red:
print('密码设置成功')
else:
print('错误')
J=Jxh()
J.b()
# +
class Num(object):
def __init__(self,a,b,c):
self.__a = a
self.__b = b
self.__c = c
@property
def a(self):
print(self.__a)
@a.setter
def a(self,a1):
self.__a = a1
def a1(self):
print(self.__a + self.__b + self.__c)
num=Num(1,5,5)
num.a = 1000
num.a1()
# +
class Num(object):
def __init__(self):
self._mimi = '哈哈哈'
self._mima = '123456'
@property
def mimi(self):
return self._mimi
@mimi.setter
def mimi(self,str_):
self._mimi = str_
num = Num()
print(num.mimi)
num.mimi = '啦啦'
print(num.mimi)
# -
class Num(object):
def __init__(self):
self.__a = 1000
def A (self):
print(self.__a)
num=Num()
print(num.A())
class Num(object):
__slots__ = ('a','b')
def __init__(self):
self.a = 1000
num=Num()
num.b = 1000
print(num.b)
# 例题:输入四条边判断它是否是一个正方形,如果是计算面积
class Zfx (object):
def __init__(self,a,b,c,d):
self.a = a
self.b = b
self.c = c
self.d = d
@staticmethod
def is_valid(a,b,c,d):
for i in [b,c,d]:
if i != a:
return False
else:
return True
def area(self):
if res == True:
area_=self.a*self.b
return area_
zfx=Zfx(8,8,8,8)
res = zfx.is_valid(8,8,8,8)
if res==True:
print(zfx.area())
class A(object):
def __init__(self):
self.a=10
self.a1=20
def sum_(self):
SUM_=self.a+self.a1
return SUM_
class B(A):
def __init__(self):
A. __init__(self)
def B1(self):
res = self.sum_()
print(res)
b=B()
b.B1()
class A(object):
def __init__(self):
self.a=111
def A1(self):
print('A1')
#在类名中写上继承的类
class B(A):
def __init__(self):
A. __init__(self)
self.a=200
def B1(self):
print('B1')
c=B()
print(c.a)
c.A1()
def deco(func):
def warp(a1,a2):
print('a1+a2',a1+a2)
return func(a1,a2)
return warp
@deco
def SUM(a1,a2):
print(a1,a2)
SUM(100,100)
from time import time,localtime,sleep
print(time())
class A(object):
def __init__(self):
self.jxh = 100
@classmethod
def a(cls):
return cls()
c = A.a()
print (c.jxh)
a = (x for x in range(100000000000) if x % 2== 0)
for i in range(100):
print(next(a))
# +
def jxh(func):
def warp(n1,n2,n3):
num = n1 + n2
return func(0,num,n3)
return warp
#装饰器将前两个数字求和,函数本身第三个参数乘上这个和
@jxh
def SUM(num1,num2,num3):
print(num1,num2,num3)
print(num2 * num3)
SUM(10,2,3)
# -
def foo( ):
def bar( ):
return 'hello'
return bar( )
a = foo( )
print (a)
| python01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wesleybeckner/technology_fundamentals/blob/main/C2%20Statistics%20and%20Model%20Creation/SOLUTIONS/SOLUTION_Tech_Fun_C2_S3_Inferential_Statistics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ZhMpsTQGPQIm"
# # Technology Fundamentals Course 2, Session 3: Inferential Statistics
#
# **Instructor**: <NAME>
#
# **Contact**: <EMAIL>
#
# **Teaching Assitants**: <NAME>, <NAME>
#
# **Contact**: <EMAIL>, <EMAIL>
#
# <br>
#
# ---
#
# <br>
#
# In this session we will look at the utility of EDA combined with inferential statistics.
#
# <br>
#
# ---
#
# + [markdown] id="jc0SbuFeQBwW"
# <a name='x.0'></a>
#
# ## 6.0 Preparing Environment and Importing Data
#
# [back to top](#top)
# + [markdown] id="biwnU2732lYG"
# <a name='x.0.1'></a>
#
# ### 6.0.1 Import Packages
#
# [back to top](#top)
# + id="XA8E1GTQQBwW" colab={"base_uri": "https://localhost:8080/"} outputId="f0bfd37a-6722-4c23-c78e-b81378e4a90e"
# The modules we've seen before
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import plotly.express as px
import seaborn as sns
# our stats modules
import random
import scipy.stats as stats
import statsmodels.api as sm
from statsmodels.formula.api import ols
import scipy
# + [markdown] id="2oNmcaxw2og4"
# <a name='x.0.2'></a>
#
# ### 6.0.2 Load Dataset
#
# [back to top](#top)
#
# For this session, we will use dummy datasets from sklearn.
# + id="aRWow_s8eSm7"
df = pd.read_csv('https://raw.githubusercontent.com/wesleybeckner/'\
'ds_for_engineers/main/data/truffle_margin/truffle_margin_customer.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 421} id="7P7VmDLOa5xa" outputId="b777e7d6-f3c8-4eb8-d34d-c25c2bf757c4"
df
# + id="I734bGQy10_r"
descriptors = df.columns[:-2]
# + colab={"base_uri": "https://localhost:8080/"} id="I3b_cORVpFAs" outputId="44fc2914-217e-48d1-a40b-26491a39fde0"
for col in descriptors:
print(col)
print(df[col].unique())
print()
# + [markdown] id="KKkK88MuiQby"
# ## 6.1 Many Flavors of Statistical Tests
#
# <p align="center">
# <img src="https://luminousmen.com/media/descriptive-and-inferential-statistics.jpeg" width=400px></img>
# <br>
# <small> https://luminousmen.com/post/descriptive-and-inferential-statistics </small>
# </p>
#
# >Descriptive statistics describes data (for example, a chart or graph) and inferential statistics allows you to make predictions (“inferences”) from that data. With inferential statistics, you take data from samples and make generalizations about a population - [statshowto](https://www.statisticshowto.com/probability-and-statistics/statistics-definitions/inferential-statistics/#:~:text=Descriptive%20statistics%20describes%20data%20(for,make%20generalizations%20about%20a%20population.)
#
# * **Moods Median Test**
# * [Kruskal-Wallis Test](https://sixsigmastudyguide.com/kruskal-wallis-non-parametric-hypothesis-test/) (Another comparison of Medians test)
# * T-Test
# * Analysis of Variance (ANOVA)
# * One Way ANOVA
# * Two Way ANOVA
# * MANOVA
# * Factorial ANOVA
#
# When do I use each of these? We will talk about this as we proceed through the examples. [This page](https://support.minitab.com/en-us/minitab/20/help-and-how-to/statistics/nonparametrics/supporting-topics/which-test-should-i-use/) from minitab has good rules of thumb on the subject.
#
#
# + [markdown] id="D5evicZ44rAA"
# ### 6.1.1 What is Mood's Median?
#
# > You can use Chi-Square to test for a goodness of fit (whether a sample of data represents a distribution) or whether two variables are related (using a contingency table, which we will create below!)
#
# **A special case of Pearon's Chi-Squared Test:** We create a table that counts the observations above and below the global median for two different groups. We then perform a *chi-squared test of significance* on this *contingency table*
#
# Null hypothesis: the Medians are all equal
#
# The chi-square test statistic:
#
# $x^2 = \sum{\frac{(O-E)^2}{E}}$
#
# Where $O$ is the observed frequency and $E$ is the expected frequency.
#
# **Let's take an example**, say we have three shifts with the following production rates:
# + id="_zJS1ehrKi32"
np.random.seed(42)
shift_one = [round(i) for i in np.random.normal(16, 3, 10)]
shift_two = [round(i) for i in np.random.normal(24, 3, 10)]
# + colab={"base_uri": "https://localhost:8080/"} id="49AvW9Sq7MKQ" outputId="7ff969ba-ee34-46e9-a9e6-79857e2bca3d"
print(shift_one)
print(shift_two)
# + id="1zS4K5hL7gON"
stat, p, m, table = scipy.stats.median_test(shift_one, shift_two, correction=False)
# + [markdown] id="KVzJKeBg9NNu"
# what is `median_test` returning?
# + colab={"base_uri": "https://localhost:8080/"} id="pLUCH4Rv73Qp" outputId="9f203559-103f-42cb-9d44-e4329bbc7443"
print("The perasons chi-square test statistic: {:.2f}".format(stat))
print("p-value of the test: {:.3f}".format(p))
print("the grand median: {}".format(m))
# + [markdown] id="_DJLQx0x-hGU"
# Let's evaluate that test statistic ourselves by taking a look at the contingency table:
# + colab={"base_uri": "https://localhost:8080/"} id="dH_mlqu0762C" outputId="77448817-4ed0-4d25-8025-a5c355feb73d"
table
# + [markdown] id="0Ox8X-A4-mnk"
# This is easier to make sense of if we order the shift times
# + colab={"base_uri": "https://localhost:8080/"} id="wT4_CjEL-pkg" outputId="b500833e-c4d5-49bc-9370-0ea92020139f"
shift_one.sort()
shift_one
# + [markdown] id="8QAfysw6-yJR"
# When we look at shift one, we see that 8 values are at or below the grand median.
# + colab={"base_uri": "https://localhost:8080/"} id="9udymrL9-xxF" outputId="a78904d4-4b03-4284-ccbb-32893aaae676"
shift_two.sort()
shift_two
# + [markdown] id="jDgDGYlQ-miZ"
# For shift two, only two are at or below the grand median.
#
# Since the sample sizes are the same, the expected value for both groups is the same, 5 above and 5 below the grand median. The chi-square is then:
#
# $X^2 = \frac{(2-5)^2}{5} + \frac{(8-5)^2}{5} + \frac{(8-5)^2}{5} + \frac{(2-5)^2}{5}$
#
# + colab={"base_uri": "https://localhost:8080/"} id="F9BEYdaW_m43" outputId="109d40a5-2b69-4b2b-eead-9fd3be5ef0bf"
(2-5)**2/5 + (8-5)**2/5 + (8-5)**2/5 + (2-5)**2/5
# + [markdown] id="4X2xYQkuExJ-"
# Our p-value, or the probability of observing the null-hypothsis, is under 0.05. We can conclude that these shift performances were drawn under seperate distributions.
#
# For comparison, let's do this analysis again with shifts of equal performances
# + colab={"base_uri": "https://localhost:8080/"} id="_bcZqXDmFHBV" outputId="18d224ec-3eb3-44a5-a897-8072b0052803"
np.random.seed(3)
shift_three = [round(i) for i in np.random.normal(16, 3, 10)]
shift_four = [round(i) for i in np.random.normal(16, 3, 10)]
stat, p, m, table = scipy.stats.median_test(shift_three, shift_four,
correction=False)
print("The pearsons chi-square test statistic: {:.2f}".format(stat))
print("p-value of the test: {:.3f}".format(p))
print("the grand median: {}".format(m))
# + [markdown] id="w3CJcRMUFVE3"
# and the shift raw values:
# + colab={"base_uri": "https://localhost:8080/"} id="crPZoQBMFXYQ" outputId="458fb6f4-87f1-483b-e6ad-4c79130cf49a"
shift_three.sort()
shift_four.sort()
print(shift_three)
print(shift_four)
# + colab={"base_uri": "https://localhost:8080/"} id="etY5ypiYFdxs" outputId="fe3c229a-4538-4ae1-beae-a27b3028b541"
table
# + [markdown] id="otPGkuW40I_f"
# ### 6.1.2 When to Use Mood's?
#
# **Mood's Median Test is highly flexible** but has the following assumptions:
#
# * Considers only one categorical factor
# * Response variable is continuous (our shift rates)
# * Data does not need to be normally distributed
# * But the distributions are similarly shaped
# * Sample sizes can be unequal and small (less than 20 observations)
#
# Other considerations:
#
# * Not as powerful as Kruskal-Wallis Test but still useful for small sample sizes or when there are outliers
# + [markdown] id="NRpmP6jWPHNj"
# #### 6.1.2.1 Exercise: Use Mood's Median Test
#
# + [markdown] id="BNHxgf12nGJs"
# ##### **Part A** Perform moods median test on Base Cake in Truffle data
#
# We're also going to get some practice with pandas groupby.
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="rHm8-bQnfDUS" outputId="a72c25bd-fd61-479f-d437-b23a2cef2632"
df[['Base Cake', 'EBITDA/KG']].head()
# + id="SaEEogoHjvHj"
# what is returned by this groupby?
gp = df.groupby('Base Cake')
# + [markdown] id="0K7e7MGEj5Ht"
# How do we find out? We could iterate through it:
# + colab={"base_uri": "https://localhost:8080/"} id="2LGkJEinj9SK" outputId="4aea071b-4448-481a-f505-bc3b2ae9fccb"
# seems to be a tuple of some sort
for i in gp:
print(i)
break
# + colab={"base_uri": "https://localhost:8080/"} id="lwMASrDRkTQj" outputId="d5a87e23-9ccc-4d2c-b518-5a6fac0f9546"
# the first object appears to be the group
print(i[0])
# the second object appears to be the df belonging to that group
print(i[1])
# + [markdown] id="gFOrTaPoknGx"
# going back to our diagram from our earlier pandas session. It looks like whenever we split in the groupby method, we create separate dataframes as well as their group label:
#
# <img src="https://swcarpentry.github.io/r-novice-gapminder/fig/12-plyr-fig1.png" width=500></img>
#
# Ok, so we know `gp` is separate dataframes. How do we turn them into arrays to then pass to `median_test`?
# + id="nVUYH5gFlWFj" colab={"base_uri": "https://localhost:8080/"} outputId="0fe0e412-48ed-440d-a1bd-35cad55f6396"
# complete this for loop
for i, j in gp:
# turn j into an array using the .values attribute
print(i, j['EBITDA/KG'].values)
# turn j into an array of the EBITDA/KG column and grab the values using .values attribute
# j --> grab EBITDA/KG --> turn into an array with .values
# print this to the screen
# + [markdown] id="QlP1VYjSlrOJ"
# After you've completed the previous step, turn this into a list comprehension and pass the result to a variable called `margins`
# + id="pCr2wjfpP31r"
# complete the code below
margins = [j['EBITDA/KG'].values for i,j in gp]
# + [markdown] id="TjN_QppNl4v9"
# Remember the list unpacking we did for the tic tac toe project? We're going to do the same thing here. Unpack the margins list for `median_test` and run the cell below!
# + colab={"base_uri": "https://localhost:8080/"} id="gJwZFD84PUBP" outputId="d62c9a84-8f32-449c-d62d-c5ec9a2a7929"
# complete the following line
stat, p, m, table = scipy.stats.median_test(*margins, correction=False)
print("The pearsons chi-square test statistic: {:.2f}".format(stat))
print("p-value of the test: {:.2e}".format(p))
print("the grand median: {:.2e}".format(m))
# + [markdown] id="-UjWU3c6SV1d"
# ##### **Part B** View the distributions of the data using matplotlib and seaborn
#
# What a fantastic statistical result we found! Can we affirm our result with some visualizations? I hope so! Create a boxplot below using pandas. In your call to `df.boxplot()` the `by` parameter should be set to `Base Cake` and the `column` parameter should be set to `EBITDA/KG`
# + colab={"base_uri": "https://localhost:8080/", "height": 409} id="BwdnbQWYQ4WL" outputId="bad403d7-c2c7-428a-a3ed-be9bb711c7f2"
# YOUR BOXPLOT HERE
df.boxplot(by='Base Cake', column='EBITDA/KG')
# + [markdown] id="AXPBuKkPmZpW"
# For comparison, I've shown the boxplot below using seaborn!
# + colab={"base_uri": "https://localhost:8080/", "height": 442} id="jetbzeP8R6UK" outputId="0db76e3e-e01a-47fa-ff69-bb8a89598263"
fig, ax = plt.subplots(figsize=(10,7))
ax = sns.boxplot(x='Base Cake', y='EBITDA/KG', data=df, color='#A0cbe8')
# + [markdown] id="J2cnPwn8SNq5"
# ##### **Part C** Perform Moods Median on all the other groups
# + colab={"base_uri": "https://localhost:8080/"} id="awIi7LRoppUd" outputId="498cafd8-f7d3-4803-caed-a50b0519f2c1"
ls = []
for i in range(10): # for loop initiation line
if i % 2 == 0:
ls.append(i**2) # actual task upon each loop
# ls
# + colab={"base_uri": "https://localhost:8080/"} id="EluB83eYp2Mg" outputId="8f464da8-1256-4547-fdc8-a6394c4c96e0"
ls = [i**2 for i in range(10) if i % 2 == 0]
# ls
# + colab={"base_uri": "https://localhost:8080/"} id="EVm8E8XcSmwI" outputId="f3da4309-e499-4075-c5df-47c5ab3f6097"
# Recall the other descriptors we have
descriptors
# + colab={"base_uri": "https://localhost:8080/"} id="H2ZozFp_SeWx" outputId="68a5b049-eca3-4156-e10d-35666b0d546e"
for desc in descriptors:
# YOUR CODE FORM MARGINS BELOW
margins = [j['EBITDA/KG'].values for i,j in df.groupby(desc)]
# UNPACK MARGINS INTO MEDIAN_TEST
stat, p, m, table = scipy.stats.median_test(*margins, correction=False)
print(desc)
print("The pearsons chi-square test statistic: {:.2f}".format(stat))
print("p-value of the test: {:e}".format(p))
print("the grand median: {}".format(m), end='\n\n')
# + [markdown] id="sYwTWkVCm7jM"
# ##### **Part D** Many boxplots
#
# And finally, we will confirm these visually. Complete the Boxplot for each group:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="eEFvkzyQZtP2" outputId="4629e0d9-b75d-4cd0-8d94-821882bba649"
for desc in descriptors:
fig, ax = plt.subplots(figsize=(10,5))
sns.boxplot(x=desc, y='EBITDA/KG', data=df, color='#A0cbe8', ax=ax)
# + [markdown] id="a2hegS_Q2g3K"
# ### 6.1.3 **Enrichment**: What is a T-test?
#
# There are 1-sample and 2-sample T-tests
#
# _(note: we would use a 1-sample T-test just to determine if the sample mean is equal to a hypothesized population mean)_
#
# Within 2-sample T-tests we have **_independent_** and **_dependent_** T-tests (uncorrelated or correlated samples)
#
# For independent, two-sample T-tests:
#
# * **_Equal variance_** (or pooled) T-test
# * `scipy.stats.ttest_ind(equal_var=True)`
# * **_Unequal variance_** T-test
# * `scipy.stats.ttest_ind(equal_var=False)`
# * also called ***Welch's T-test***
#
# <br>
#
# For dependent T-tests:
# * Paired (or correlated) T-test
# * `scipy.stats.ttest_rel`
#
# A full discussion on T-tests is outside the scope of this session, but we can refer to wikipedia for more information, including formulas on how each statistic is computed:
# * [student's T-test](https://en.wikipedia.org/wiki/Student%27s_t-test#Dependent_t-test_for_paired_samples)
# + [markdown] id="RUTirNIp0VeC"
# ### 6.1.4 **Enrichment**: Demonstration of T-tests
#
# [back to top](#top)
# + [markdown] id="fWgpdHu3Mp19"
# We'll assume our shifts are of **_equal variance_** and proceed with the appropriate **_independent two-sample_** T-test...
# + colab={"base_uri": "https://localhost:8080/"} id="5mxktlT2NB8v" outputId="71d62d31-5d7a-4ee6-d9b3-8a4112646ebb"
print(shift_one)
print(shift_two)
# + [markdown] id="NymtMrgQ0FaD"
# To calculate the T-test, we follow a slightly different statistical formula:
#
# $T=\frac{\mu_1 - \mu_2}{s\sqrt{\frac{1}{n_1} + \frac{1}{n_2}}}$
#
# where $\mu$ are the means of the two groups, $n$ are the sample sizes and $s$ is the pooled standard deviation, also known as the cummulative variance (depending on if you square it or not):
#
# $s= \sqrt{\frac{(n_1-1)\sigma_1^2 + (n_2-1)\sigma_2^2}{n_1 + n_2 - 2}}$
#
# where $\sigma$ are the standard deviations. What you'll notice here is we are combining the two variances, we can only do this if we assume the variances are somewhat equal, this is known as the *equal variances* t-test.
# + colab={"base_uri": "https://localhost:8080/"} id="vsol_nFrNXBZ" outputId="941d1312-3a2c-48a8-ab8e-a21d7b1675e4"
mean_shift_one = np.mean(shift_one)
mean_shift_two = np.mean(shift_two)
print(mean_shift_one, mean_shift_two)
# + colab={"base_uri": "https://localhost:8080/"} id="WQpbuu9sQ_Ky" outputId="e46c5ab0-c313-4640-c423-53b81c21035d"
com_var = ((np.sum([(i - mean_shift_one)**2 for i in shift_one]) +
np.sum([(i - mean_shift_two)**2 for i in shift_two])) /
(len(shift_one) + len(shift_two)-2))
print(com_var)
# + id="wOqV1VffNhFr"
T = (np.abs(mean_shift_one - mean_shift_two) / (
np.sqrt(com_var/len(shift_one) +
com_var/len(shift_two))))
# + colab={"base_uri": "https://localhost:8080/"} id="iGK1cLxvQl6e" outputId="2cadce9f-c373-45c9-f05e-cde49212be1a"
T
# + [markdown] id="eX2l-cJbB7fm"
# We see that this hand-computed result matches that of the `scipy` module:
# + colab={"base_uri": "https://localhost:8080/"} id="rgXWXjqIOPBm" outputId="4430f467-edd1-47de-a6fa-91e45050a64c"
scipy.stats.ttest_ind(shift_two, shift_one, equal_var=True)
# + [markdown] id="Z5R6Gl9-Kmt_"
# ### **Enrichment**: 6.1.5 What are F-statistics and the F-test?
#
# The F-statistic is simply a ratio of two variances, or the ratio of _mean squares_
#
# _mean squares_ is the estimate of population variance that accounts for the degrees of freedom to compute that estimate.
#
# We will explore this in the context of ANOVA
# + [markdown] id="QleTrLDgjYYy"
# ### 6.1.6 **Enrichment**: What is Analysis of Variance?
#
# ANOVA uses the F-test to determine whether the variability between group means is larger than the variability within the groups. If that statistic is large enough, you can conclude that the means of the groups are not equal.
#
# **The caveat is that ANOVA tells us whether there is a difference in means but it does not tell us where the difference is.** To find where the difference is between the groups, we have to conduct post-hoc tests.
#
# There are two main types:
# * One-way (one factor) and
# * Two-way (two factor) where factor is an indipendent variable
#
# <br>
#
# | Ind A | Ind B | Dep |
# |-------|-------|-----|
# | X | H | 10 |
# | X | I | 12 |
# | Y | I | 11 |
# | Y | H | 20 |
#
# <br>
#
# #### ANOVA Hypotheses
#
# * _Null hypothesis_: group means are equal
# * _Alternative hypothesis_: at least one group mean is different form the other groups
#
# ### ANOVA Assumptions
#
# * Residuals (experimental error) are normally distributed (test with Shapiro-Wilk)
# * Homogeneity of variances (variances are equal between groups) (test with Bartlett's)
# * Observations are sampled independently from each other
# * _Note: ANOVA assumptions can be checked using test statistics (e.g. Shapiro-Wilk, Bartlett’s, Levene’s test) and the visual approaches such as residual plots (e.g. QQ-plots) and histograms._
#
# ### Steps for ANOVA
#
# * Check sample sizes: equal observations must be in each group
# * Calculate Sum of Square between groups and within groups ($SS_B, SS_E$)
# * Calculate Mean Square between groups and within groups ($MS_B, MS_E$)
# * Calculate F value ($MS_B/MS_E$)
#
# <br>
#
# This might be easier to see in a table:
#
# <br>
#
# | Source of Variation | degree of freedom (Df) | Sum of squares (SS) | Mean square (MS) | F value |
# |-----------------------------|------------------------|---------------------|--------------------|-------------|
# | Between Groups | Df_b = P-1 | SS_B | MS_B = SS_B / Df_B | MS_B / MS_E |
# | Within Groups | Df_E = P(N-1) | SS_E | MS_E = SS_E / Df_E | |
# | total | Df_T = PN-1 | SS_T | | |
#
# Where:
# $$ SS_B = \sum_{i}^{P}{(\bar{y}_i-\bar{y})^2} $$
# <br>
# $$ SS_E = \sum_{ik}^{PN}{(\bar{y}_{ik}-\bar{y}_i)^2} $$
# <br>
# $$ SS_T = SS_B + SS_E $$
#
# Let's go back to our shift data to take an example:
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="NZEL5iUfTeNv" outputId="52d540dd-ef7f-4fad-f898-bc7c15394290"
shifts = pd.DataFrame([shift_one, shift_two, shift_three, shift_four]).T
shifts.columns = ['A', 'B', 'C', 'D']
shifts.boxplot()
# + [markdown] id="NItJ8vbPWFKo"
# #### 6.1.6.0 **Enrichment**: SNS Boxplot
#
# this is another great way to view boxplot data. Notice how sns also shows us the raw data alongside the box and whiskers using a _swarmplot_.
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="gBVbCptwUrdN" outputId="3e750968-620b-4259-92d1-e095f0386db9"
shift_melt = pd.melt(shifts.reset_index(), id_vars=['index'],
value_vars=['A', 'B', 'C', 'D'])
shift_melt.columns = ['index', 'shift', 'rate']
ax = sns.boxplot(x='shift', y='rate', data=shift_melt, color='#A0cbe8')
ax = sns.swarmplot(x="shift", y="rate", data=shift_melt, color='#79706e')
# + [markdown] id="TlgRvZlEGt2l"
# Anyway back to ANOVA...
# + colab={"base_uri": "https://localhost:8080/"} id="SX5wvWAxV_XC" outputId="fca50497-87c5-4092-8d9b-7edcc6c354ba"
fvalue, pvalue = stats.f_oneway(shifts['A'],
shifts['B'],
shifts['C'],
shifts['D'])
print(fvalue, pvalue)
# + [markdown] id="ywXmwS-aY4YM"
# We can get this in the format of the table we saw above:
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="sC_-cfKqYnQp" outputId="ee8daac7-5baa-4dc9-83b0-86d7edf54ace"
# get ANOVA table
import statsmodels.api as sm
from statsmodels.formula.api import ols
# Ordinary Least Squares (OLS) model
model = ols('rate ~ C(shift)', data=shift_melt).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
anova_table
# output (ANOVA F and p value)
# + [markdown] id="pW5Vx6VWdNxw"
# The **_Shapiro-Wilk_** test can be used to check the _normal distribution of residuals_. Null hypothesis: data is drawn from normal distribution.
# + colab={"base_uri": "https://localhost:8080/"} id="phB9lnHddQv1" outputId="986bfcb3-a80d-48cc-b142-9d19f5b50468"
w, pvalue = stats.shapiro(model.resid)
print(w, pvalue)
# + [markdown] id="0xRqtKixd1MT"
# We can use **_Bartlett’s_** test to check the _Homogeneity of variances_. Null hypothesis: samples from populations have equal variances.
# + colab={"base_uri": "https://localhost:8080/"} id="z2sya2JMd2IJ" outputId="d70f70ca-6105-482a-cdef-2585aeeb9908"
w, pvalue = stats.bartlett(shifts['A'],
shifts['B'],
shifts['C'],
shifts['D'])
print(w, pvalue)
# + [markdown] id="mJlzhEk7ar6t"
# #### 6.1.6.1 ANOVA Interpretation
#
# The _p_ value form ANOVA analysis is significant (_p_ < 0.05) and we can conclude there are significant difference between the shifts. But we do not know which shift(s) are different. For this we need to perform a post hoc test. There are a multitude of these that are beyond the scope of this discussion ([Tukey-kramer](https://www.real-statistics.com/one-way-analysis-of-variance-anova/unplanned-comparisons/tukey-kramer-test/) is one such test)
#
# <p align=center>
# <img src="https://media.tenor.com/images/4da4d46c8df02570a9a1219cac42bf27/tenor.gif"></img>
# </p>
# + [markdown] id="_vh_d3ENIImT"
# ### 6.1.7 Putting it all together
#
# In summary, there are many statistical tests at our disposal when performing inferential statistical analysis. In times like these, a simple decision tree can be extraordinarily useful!
#
# <img src="https://cdn.scribbr.com/wp-content/uploads//2020/01/flowchart-for-choosing-a-statistical-test.png" width=800px></img>
#
# <small>source: [scribbr](https://www.scribbr.com/statistics/statistical-tests/)</small>
# + [markdown] id="uNnqPwXqaAEA"
# ## 6.2 Evaluate statistical significance of product margin: a snake in the garden
# + [markdown] id="X1i4SQ0y2p4Q"
# ### 6.2.1 Mood's Median on product descriptors
#
# The first issue we run into with moods is... what?
#
# We can only perform moods on two groups at a time. How can we get around this?
#
# Let's take a look at the category with the fewest descriptors. If we remember, this was the Truffle Types.
# + colab={"base_uri": "https://localhost:8080/"} id="TqfEEjXbVCTo" outputId="5dd791fd-0bdd-4111-8d01-c592d6bf5d25"
df.columns
# + colab={"base_uri": "https://localhost:8080/"} id="iKAlINvT2n2E" outputId="8301b41c-1112-42f6-eed9-b200cdfa9bdd"
df['Truffle Type'].unique()
# + colab={"base_uri": "https://localhost:8080/"} id="Gsm0QCeywlou" outputId="5f4834dc-853e-4f54-d105-7d54f6eb74d4"
col = 'Truffle Type'
moodsdf = pd.DataFrame()
for truff in df[col].unique():
# for each
group = df.loc[df[col] == truff]['EBITDA/KG']
pop = df.loc[~(df[col] == truff)]['EBITDA/KG']
stat, p, m, table = scipy.stats.median_test(group, pop)
median = np.median(group)
mean = np.mean(group)
size = len(group)
print("{}: N={}".format(truff, size))
print("Welch's T-Test for Unequal Variances")
print(scipy.stats.ttest_ind(group, pop, equal_var=False))
welchp = scipy.stats.ttest_ind(group, pop, equal_var=False).pvalue
print()
moodsdf = pd.concat([moodsdf,
pd.DataFrame([truff,
stat, p, m, mean, median, size,
welchp, table]).T])
moodsdf.columns = [col, 'pearsons_chi_square', 'p_value',
'grand_median', 'group_mean', 'group_median', 'size', 'welch p',
'table']
# + [markdown] id="N747uVxhEGEP"
# ### Question 1: Moods Results on Truffle Type
#
# > What do we notice about the resultant table?
#
# * **_p-values_** Most are quite small (really low probability of achieving these table results under a single distribution)
# * group sizes: our Jelly Filled group is relatively small
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="6TQlsrBrw2zQ" outputId="8d6db584-dc34-42e4-a3f9-ab05a096c867"
sns.boxplot(x='Base Cake', y='EBITDA/KG', data=df)
# + colab={"base_uri": "https://localhost:8080/", "height": 143} id="DGaZNDDSxHeP" outputId="27b51784-fae7-47b7-84de-7432f86e7bcc"
moodsdf.sort_values('p_value')
# + [markdown] id="mawPQ2p5xW1O"
# We can go ahead and repeat this analysis for all of our product categories:
# + colab={"base_uri": "https://localhost:8080/"} id="X7PmQ04oVZRi" outputId="d066c7f2-cb80-4c58-d38f-56c59559fa95"
df.columns[:5]
# + id="HjTW8SGsvKdF" colab={"base_uri": "https://localhost:8080/"} outputId="e4dcd64b-235e-4133-a407-b5e53eeee1ce"
moodsdf = pd.DataFrame()
for col in df.columns[:5]:
for truff in df[col].unique():
group = df.loc[df[col] == truff]['EBITDA/KG']
pop = df.loc[~(df[col] == truff)]['EBITDA/KG']
stat, p, m, table = scipy.stats.median_test(group, pop)
median = np.median(group)
mean = np.mean(group)
size = len(group)
welchp = scipy.stats.ttest_ind(group, pop, equal_var=False).pvalue
moodsdf = pd.concat([moodsdf,
pd.DataFrame([col, truff,
stat, p, m, mean, median, size,
welchp, table]).T])
moodsdf.columns = ['descriptor', 'group', 'pearsons_chi_square', 'p_value',
'grand_median', 'group_mean', 'group_median', 'size', 'welch p',
'table']
print(moodsdf.shape)
# + id="h-4GZ5bSvn6f" colab={"base_uri": "https://localhost:8080/"} outputId="43feb4b5-5be6-4281-c605-67797c580b26"
moodsdf = moodsdf.loc[(moodsdf['welch p'] < 0.05) &
(moodsdf['p_value'] < 0.05)].sort_values('group_median')
moodsdf = moodsdf.sort_values('group_median').reset_index(drop=True)
print(moodsdf.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 361} id="V6Rm8Ju82TKd" outputId="57d37bb7-b7e3-4bdc-8f20-3a576154daa7"
moodsdf[-10:]
# + [markdown] id="PIiMWCRVGBsB"
# ### 6.2.2 **Enrichment**: Broad Analysis of Categories: ANOVA
#
#
# + [markdown] id="ytKRGYYWiSFX"
# Recall our "melted" shift data. It will be useful to think of getting our Truffle data in this format:
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="URjZgCMViMiw" outputId="1ca51502-e172-4566-9bbe-8aae64e690c4"
shift_melt.head()
# + id="WYvWTt4YixSw"
df.columns = df.columns.str.replace(' ', '_')
df.columns = df.columns.str.replace('/', '_')
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="6xBHomXrhqAm" outputId="55d31c27-dcba-41df-e694-186430f7dab6"
# get ANOVA table
# Ordinary Least Squares (OLS) model
model = ols('EBITDA_KG ~ C(Truffle_Type)', data=df).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
anova_table
# output (ANOVA F and p value)
# + [markdown] id="7A9JQbARjF5B"
# Recall the **_Shapiro-Wilk_** test can be used to check the _normal distribution of residuals_. Null hypothesis: data is drawn from normal distribution.
# + colab={"base_uri": "https://localhost:8080/"} id="OPiFlR0Ii7RB" outputId="dc554747-08df-48fe-8c31-1529307f9776"
w, pvalue = stats.shapiro(model.resid)
print(w, pvalue)
# + [markdown] id="f739OA5gjYGZ"
# And the **_Bartlett’s_** test to check the _Homogeneity of variances_. Null hypothesis: samples from populations have equal variances.
# + id="ozV8i6sLmBst" colab={"base_uri": "https://localhost:8080/"} outputId="82110757-583d-47ae-9c77-bdeea2596e6e"
gb = df.groupby('Truffle_Type')['EBITDA_KG']
gb
# + colab={"base_uri": "https://localhost:8080/"} id="ixHKhdRQjEkh" outputId="9c484ee1-0b37-43f2-84e2-cc1720df8f65"
w, pvalue = stats.bartlett(*[gb.get_group(x) for x in gb.groups])
print(w, pvalue)
# + [markdown] id="9XIDDKOBmp3L"
# Wow it looks like our data is not drawn from a normal distribution! Let's check this for other categories...
#
# We can wrap these in a for loop:
# + colab={"base_uri": "https://localhost:8080/", "height": 842} id="36ctJA_MmrzD" outputId="167595a4-50b3-4c8f-887b-277a93d7f26b"
for col in df.columns[:5]:
print(col)
model = ols('EBITDA_KG ~ C({})'.format(col), data=df).fit()
anova_table = sm.stats.anova_lm(model, typ=2)
display(anova_table)
w, pvalue = stats.shapiro(model.resid)
print("Shapiro: ", w, pvalue)
gb = df.groupby(col)['EBITDA_KG']
w, pvalue = stats.bartlett(*[gb.get_group(x) for x in gb.groups])
print("Bartlett: ", w, pvalue)
print()
# + [markdown] id="2bnum6yeqmy4"
# ### 6.2.3 **Enrichment**: Visual Analysis of Residuals: QQ-Plots
#
# This can be distressing and is often why we want visual methods to see what is going on with our data!
# + colab={"base_uri": "https://localhost:8080/", "height": 544} id="M3BHEncwo01Z" outputId="1e58f2fd-7e34-4041-acab-68986a35c429"
model = ols('EBITDA_KG ~ C(Truffle_Type)', data=df).fit()
#create instance of influence
influence = model.get_influence()
#obtain standardized residuals
standardized_residuals = influence.resid_studentized_internal
# res.anova_std_residuals are standardized residuals obtained from ANOVA (check above)
sm.qqplot(standardized_residuals, line='45')
plt.xlabel("Theoretical Quantiles")
plt.ylabel("Standardized Residuals")
plt.show()
# histogram
plt.hist(model.resid, bins='auto', histtype='bar', ec='k')
plt.xlabel("Residuals")
plt.ylabel('Frequency')
plt.show()
# + [markdown] id="b5QthjAmqy0k"
# We see that a lot of our data is swayed by extremely high and low values, so what can we conclude?
#
# > You need the right test statistic for the right job, in this case, we are littered with unequal variance in our groupings so we use the moods median and welch (unequal variance t-test) to make conclusions about our data
#
# + [markdown] id="1kO7TDSohI_b"
# # References
#
# * [<NAME> ANOVA](https://www.reneshbedre.com/blog/anova.html)
# * [Minitab ANOVA](https://blog.minitab.com/en/adventures-in-statistics-2/understanding-analysis-of-variance-anova-and-the-f-test)
# * [Analytics Vidhya ANOVA](https://www.analyticsvidhya.com/blog/2020/06/introduction-anova-statistics-data-science-covid-python/)
# * [Renesh Bedre Hypothesis Testing](https://www.reneshbedre.com/blog/hypothesis-testing.html)
# * [Real Statistics Turkey-kramer](https://www.real-statistics.com/one-way-analysis-of-variance-anova/unplanned-comparisons/tukey-kramer-test/)
# * [Mutual Information](https://www.kaggle.com/ryanholbrook/mutual-information)
| C2 Statistics and Model Creation/SOLUTIONS/SOLUTION_Tech_Fun_C2_S3_Inferential_Statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("pokemon-from-kaggle.csv")
df.head(2)
df.columns
df['type1'].unique()
df['type1'].value_counts()
df['type1'].value_counts().plot(kind="bar")
| Pandas - Frequency Count.ipynb |
# ---
# jupytext:
# text_representation:
# extension: .md
# format_name: myst
# format_version: '0.10'
# jupytext_version: 1.5.2
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Biomedical flows
#
# +++
#
# ```{margin}
# 
# ```
#
# +++
#
# I have been working from 2012-2016 as an [adjunct research scientist](http://www.simula.no/people/mikaelmo) at [Simula Research Laboratory](http://www.simula.no). Here I had the great honor to be working with the late
# [Prof. <NAME>](https://hplgit.com/homepage)
# {cite}`Valen-Sendstad2011,Mortensen2016,Mortensen2012_transitional,mortensen_2011_awr`,
# at the [Center for Biomedical Computing](http://cbc.simula.no/pub/).
#
# +++
#
# I have contributed quite a bit to the [FEniCS](https://www.fenicsproject.org) project, and the incompressible Navier-Stokes solver [Oasis](https://github.com/mikaem/Oasis) has been developed within the FEniCS framework. The solver has been developed for
# efficiency, with MPI, and it is written entirely in Python. The Oasis solver is documented in the
# *Computer Physics Communications paper*
# {cite}`mortensen2015oasis`, where we
# show that it may run as fast and accurate as the low-level finite volume
# C++ solvers OpenFOAM and CDP (Stanford).
#
# +++
#
# ```{sidebar} CSF flow
#
# 
#
# **Injected drug** inside a CSF channel, computed using Oasis.
# ```
#
# +++
#
# The Oasis solver has been used in a range of master theses (see [Teaching](teaching.html#sec:master)).
# For one of my master students, <NAME>, the thesis led to a journal paper on injecting
# drugs in the Cerebrospinal fluid (CSF) {cite}`haga_2017`. The animation on the right
# shows how an injected drug moves up and down inside the CSF channel. It can also
# be seen that, due to the very low diffusivity of the scalar drug, we modelled the
# scalar transport using Lagrangian particle tracking.
#
# I have been working quite a bit together with [<NAME>](https://www.simula.no/people/kvs) at Simula, on different aspects of biomedical
# flows. Our simulations on intracranial aneurysms
# {cite}`Valen-Sendstad2011`, actually reached the headlines of Norway's larges
# newspaper [VG](#vg), when we found a correlation between transition
# to turbulence and the risk of aneurysm rupture (more famously known as stroke).
# For this work I performed most of the simulations when still at [FFI](https://www.ffi.no).
#
# <div id="vg"></div>
#
# ```{sidebar} Headlines
#
# 
# **Headlines** in norwegian newspaper VG, 5/11-2011
# ```
#
# More recently we have been studying transition and
# mesh sensitivity in the FDA nozzle benchmark. In {cite}`Bergersen2018`
# we use both regular CFD and linear stability analysis to show that care must be
# taken when designing a CFD benchmark. Transition to turbulence can only
# come from a seed, or perturbation, and an ideal case like the FDA
# benchmark should not transtition at all unless some noise is added to the
# system. [Figure](#lsa) is showing an unstable eigenmode in the FDA
# bechmark, showing that transition should in deed occur at the Reynolds
# number=3500. Here I have conducted the stability simulations using the
# [dog](http://users.monash.edu.au/~bburn/semtex.html) linear stability
# analysis software package.
#
# <p>Linear stability analysis of the FDA benchmark. Showing the most unstable eigenmode.</p>
#
# 
# ## References
#
# ```{bibliography} ../../references.bib
# :filter: docname in docnames
# ```
| _build/jupyter_execute/content/research/biomedical.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// <a name="top"></a><img src="images/chisel_1024.png" alt="Chisel logo" style="width:480px;" />
// # Module 3 Interlude: Chisel Standard Library
// **Prev: [Generators: Collections](3.2_collections.ipynb)**<br>
// **Next: [Higher-Order Functions](3.3_higher-order_functions.ipynb)**
//
// ## Motivation
// Chisel is all about re-use, so it only makes sense to provide a standard library of interfaces (encouraging interoperability of RTL) and generators for commonly-used hardware blocks.
//
// ## Setup
val path = System.getProperty("user.dir") + "/source/load-ivy.sc"
interp.load.module(ammonite.ops.Path(java.nio.file.FileSystems.getDefault().getPath(path)))
import chisel3._
import chisel3.util._
import chisel3.iotesters.{ChiselFlatSpec, Driver, PeekPokeTester}
// ---
// # The Cheatsheet
// The [Chisel3 cheatsheet](https://github.com/freechipsproject/chisel-cheatsheet/releases/latest/download/chisel_cheatsheet.pdf) contains a summary of all the major hardware construction APIs, including some of the standard library utilities that we'll introduce below.
//
// # Decoupled: A Standard Ready-Valid Interface
// One of the commonly used interfaces provided by Chisel is `DecoupledIO`, providing a ready-valid interface for transferring data. The idea is that the source drives the `bits` signal with the data to be transferred and the `valid` signal when there is data to be transferred. The sink drives the `ready` signal when it is ready to accept data, and data is considered transferred when both `ready` and `valid` are asserted on a cycle.
//
// This provides a flow control mechanism in both directions for data transfer, including a backpressure mechanism.
//
// Note: `ready` and `valid` should not be combinationally coupled, otherwise this may result in unsynthesizable combinational loops. `ready` should only be dependent on whether the sink is able to receive data, and `valid` should only be dependent on whether the source has data. Only after the transaction (on the next clock cycle) should the values update.
//
// Any Chisel data can be wrapped in a `DecoupledIO` (used as the `bits` field) as follows:
//
// ```scala
// val myChiselData = UInt(8.W)
// // or any Chisel data type, such as Bool(), SInt(...), or even custom Bundles
// val myDecoupled = Decoupled(myChiselData)
// ```
//
// The above creates a new `DecoupledIO` Bundle with fields
// - `valid`: Output(Bool)
// - `ready`: Input(Bool)
// - `bits`: Output(UInt(8.W))
// ___
//
// The rest of the section will be structured somewhat differently from the ones before: instead of giving you coding exercises, we're going to give some code examples and testcases that print the circuit state. Try to predict what will be printed before just running the tests.
//
// ## Queues
//
// `Queue` creates a FIFO (first-in, first-out) queue with Decoupled interfaces on both sides, allowing backpressure. Both the data type and number of elements are configurable.
Driver(() => new Module {
// Example circuit using a Queue
val io = IO(new Bundle {
val in = Flipped(Decoupled(UInt(8.W)))
val out = Decoupled(UInt(8.W))
})
val queue = Queue(io.in, 2) // 2-element queue
io.out <> queue
}) { c => new PeekPokeTester(c) {
// Example testsequence showing the use and behavior of Queue
poke(c.io.out.ready, 0)
poke(c.io.in.valid, 1) // Enqueue an element
poke(c.io.in.bits, 42)
println(s"Starting:")
println(s"\tio.in: ready=${peek(c.io.in.ready)}")
println(s"\tio.out: valid=${peek(c.io.out.valid)}, bits=${peek(c.io.out.bits)}")
step(1)
poke(c.io.in.valid, 1) // Enqueue another element
poke(c.io.in.bits, 43)
// What do you think io.out.valid and io.out.bits will be?
println(s"After first enqueue:")
println(s"\tio.in: ready=${peek(c.io.in.ready)}")
println(s"\tio.out: valid=${peek(c.io.out.valid)}, bits=${peek(c.io.out.bits)}")
step(1)
poke(c.io.in.valid, 1) // Read a element, attempt to enqueue
poke(c.io.in.bits, 44)
poke(c.io.out.ready, 1)
// What do you think io.in.ready will be, and will this enqueue succeed, and what will be read?
println(s"On first read:")
println(s"\tio.in: ready=${peek(c.io.in.ready)}")
println(s"\tio.out: valid=${peek(c.io.out.valid)}, bits=${peek(c.io.out.bits)}")
step(1)
poke(c.io.in.valid, 0) // Read elements out
poke(c.io.out.ready, 1)
// What do you think will be read here?
println(s"On second read:")
println(s"\tio.in: ready=${peek(c.io.in.ready)}")
println(s"\tio.out: valid=${peek(c.io.out.valid)}, bits=${peek(c.io.out.bits)}")
step(1)
// Will a third read produce anything?
println(s"On third read:")
println(s"\tio.in: ready=${peek(c.io.in.ready)}")
println(s"\tio.out: valid=${peek(c.io.out.valid)}, bits=${peek(c.io.out.bits)}")
step(1)
} }
// ## Arbiters
// Arbiters routes data from _n_ `DecoupledIO` sources to one `DecoupledIO` sink, given a prioritization.
// There are two types included in Chisel:
// - `Arbiter`: prioritizes lower-index producers
// - `RRArbiter`: runs in round-robin order
//
// Note that Arbiter routing is implemented in combinational logic.
//
// The below example will demonstrate the use of the priority arbiter (which you will also implement in the next section):
Driver(() => new Module {
// Example circuit using a priority arbiter
val io = IO(new Bundle {
val in = Flipped(Vec(2, Decoupled(UInt(8.W))))
val out = Decoupled(UInt(8.W))
})
// Arbiter doesn't have a convenience constructor, so it's built like any Module
val arbiter = Module(new Arbiter(UInt(8.W), 2)) // 2 to 1 Priority Arbiter
arbiter.io.in <> io.in
io.out <> arbiter.io.out
}) { c => new PeekPokeTester(c) {
poke(c.io.in(0).valid, 0)
poke(c.io.in(1).valid, 0)
println(s"Start:")
println(s"\tin(0).ready=${peek(c.io.in(0).ready)}, in(1).ready=${peek(c.io.in(1).ready)}")
println(s"\tout.valid=${peek(c.io.out.valid)}, out.bits=${peek(c.io.out.bits)}")
poke(c.io.in(1).valid, 1) // Valid input 1
poke(c.io.in(1).bits, 42)
// What do you think the output will be?
println(s"valid input 1:")
println(s"\tin(0).ready=${peek(c.io.in(0).ready)}, in(1).ready=${peek(c.io.in(1).ready)}")
println(s"\tout.valid=${peek(c.io.out.valid)}, out.bits=${peek(c.io.out.bits)}")
poke(c.io.in(0).valid, 1) // Valid inputs 0 and 1
poke(c.io.in(0).bits, 43)
// What do you think the output will be? Which inputs will be ready?
println(s"valid inputs 0 and 1:")
println(s"\tin(0).ready=${peek(c.io.in(0).ready)}, in(1).ready=${peek(c.io.in(1).ready)}")
println(s"\tout.valid=${peek(c.io.out.valid)}, out.bits=${peek(c.io.out.bits)}")
poke(c.io.in(1).valid, 0) // Valid input 0
// What do you think the output will be?
println(s"valid input 0:")
println(s"\tin(0).ready=${peek(c.io.in(0).ready)}, in(1).ready=${peek(c.io.in(1).ready)}")
println(s"\tout.valid=${peek(c.io.out.valid)}, out.bits=${peek(c.io.out.bits)}")
} }
// # Misc Function Blocks
// Chisel Utils has some helpers that perform stateless functions.
//
// ## Bitwise Utilities
// ### PopCount
// PopCount returns the number of high (1) bits in the input as a `UInt`.
//
// ### Reverse
// Reverse returns the bit-reversed input.
Driver(() => new Module {
// Example circuit using Reverse
val io = IO(new Bundle {
val in = Input(UInt(8.W))
val out = Output(UInt(8.W))
})
io.out := PopCount(io.in)
}) { c => new PeekPokeTester(c) {
// Integer.parseInt is used create an Integer from a binary specification
poke(c.io.in, Integer.parseInt("00000000", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
poke(c.io.in, Integer.parseInt("00001111", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
poke(c.io.in, Integer.parseInt("11001010", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
poke(c.io.in, Integer.parseInt("11111111", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
} }
Driver(() => new Module {
// Example circuit using Reverse
val io = IO(new Bundle {
val in = Input(UInt(8.W))
val out = Output(UInt(8.W))
})
io.out := Reverse(io.in)
}) { c => new PeekPokeTester(c) {
// Integer.parseInt is used create an Integer from a binary specification
poke(c.io.in, Integer.parseInt("01010101", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=0b${peek(c.io.out).toInt.toBinaryString}")
poke(c.io.in, Integer.parseInt("00001111", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=0b${peek(c.io.out).toInt.toBinaryString}")
poke(c.io.in, Integer.parseInt("11110000", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=0b${peek(c.io.out).toInt.toBinaryString}")
poke(c.io.in, Integer.parseInt("11001010", 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=0b${peek(c.io.out).toInt.toBinaryString}")
} }
// ## OneHot encoding utilities
// OneHot is an encoding of integers where there is one wire for each value, and exactly one wire is high. This allows the efficient creation of some functions, for example muxes. However, behavior may be undefined if the one-wire-high condition is not held.
//
// The below two functions provide conversion between binary (`UInt`) and OneHot encodings, and are inverses of each other:
// - UInt to OneHot: `UIntToOH`
// - OneHot to UInt: `OHToUInt`
Driver(() => new Module {
// Example circuit using UIntToOH
val io = IO(new Bundle {
val in = Input(UInt(4.W))
val out = Output(UInt(16.W))
})
io.out := UIntToOH(io.in)
}) { c => new PeekPokeTester(c) {
poke(c.io.in, 0)
println(s"in=${peek(c.io.in)}, out=0b${peek(c.io.out).toInt.toBinaryString}")
poke(c.io.in, 1)
println(s"in=${peek(c.io.in)}, out=0b${peek(c.io.out).toInt.toBinaryString}")
poke(c.io.in, 8)
println(s"in=${peek(c.io.in)}, out=0b${peek(c.io.out).toInt.toBinaryString}")
poke(c.io.in, 15)
println(s"in=${peek(c.io.in)}, out=0b${peek(c.io.out).toInt.toBinaryString}")
} }
Driver(() => new Module {
// Example circuit using OHToUInt
val io = IO(new Bundle {
val in = Input(UInt(16.W))
val out = Output(UInt(4.W))
})
io.out := OHToUInt(io.in)
}) { c => new PeekPokeTester(c) {
poke(c.io.in, Integer.parseInt("0000 0000 0000 0001".replace(" ", ""), 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
poke(c.io.in, Integer.parseInt("0000 0000 1000 0000".replace(" ", ""), 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
poke(c.io.in, Integer.parseInt("1000 0000 0000 0001".replace(" ", ""), 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
// Some invalid inputs:
// None high
poke(c.io.in, Integer.parseInt("0000 0000 0000 0000".replace(" ", ""), 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
// Multiple high
poke(c.io.in, Integer.parseInt("0001 0100 0010 0000".replace(" ", ""), 2))
println(s"in=0b${peek(c.io.in).toInt.toBinaryString}, out=${peek(c.io.out)}")
} }
// ## Muxes
// These muxes take in a list of values with select signals, and output the value associated with the lowest-index select signal.
//
// These can either take a list of (select: Bool, value: Data) tuples, or corresponding lists of selects and values as arguments. For simplicity, the examples below only demonstrate the second form.
//
// ### Priority Mux
// A `PriorityMux` outputs the value associated with the lowest-index asserted select signal.
//
// ### OneHot Mux
// An `Mux1H` provides an efficient implementation when it is guaranteed that exactly one of the select signals will be high. Behavior is undefined if the assumption is not true.
Driver(() => new Module {
// Example circuit using PriorityMux
val io = IO(new Bundle {
val in_sels = Input(Vec(2, Bool()))
val in_bits = Input(Vec(2, UInt(8.W)))
val out = Output(UInt(8.W))
})
io.out := PriorityMux(io.in_sels, io.in_bits)
}) { c => new PeekPokeTester(c) {
poke(c.io.in_bits(0), 10)
poke(c.io.in_bits(1), 20)
// Select higher index only
poke(c.io.in_sels(0), 0)
poke(c.io.in_sels(1), 1)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
// Select both - arbitration needed
poke(c.io.in_sels(0), 1)
poke(c.io.in_sels(1), 1)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
// Select lower index only
poke(c.io.in_sels(0), 1)
poke(c.io.in_sels(1), 0)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
} }
Driver(() => new Module {
// Example circuit using Mux1H
val io = IO(new Bundle {
val in_sels = Input(Vec(2, Bool()))
val in_bits = Input(Vec(2, UInt(8.W)))
val out = Output(UInt(8.W))
})
io.out := Mux1H(io.in_sels, io.in_bits)
}) { c => new PeekPokeTester(c) {
poke(c.io.in_bits(0), 10)
poke(c.io.in_bits(1), 20)
// Select index 1
poke(c.io.in_sels(0), 0)
poke(c.io.in_sels(1), 1)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
// Select index 0
poke(c.io.in_sels(0), 1)
poke(c.io.in_sels(1), 0)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
// Select none (invalid)
poke(c.io.in_sels(0), 0)
poke(c.io.in_sels(1), 0)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
// Select both (invalid)
poke(c.io.in_sels(0), 1)
poke(c.io.in_sels(1), 1)
println(s"in_sels=${peek(c.io.in_sels)}, out=${peek(c.io.out)}")
} }
// ## Counter
// `Counter` is a counter that can be incremented once every cycle, up to some specified limit, at which point it overflows. Note that it is **not** a Module, and its value is accessible.
Driver(() => new Module {
// Example circuit using Mux1H
val io = IO(new Bundle {
val count = Input(Bool())
val out = Output(UInt(2.W))
})
val counter = Counter(3) // 3-count Counter (outputs range [0...2])
when(io.count) {
counter.inc()
}
io.out := counter.value
}) { c => new PeekPokeTester(c) {
poke(c.io.count, 1)
println(s"start: counter value=${peek(c.io.out)}")
step(1)
println(s"step 1: counter value=${peek(c.io.out)}")
step(1)
println(s"step 2: counter value=${peek(c.io.out)}")
poke(c.io.count, 0)
step(1)
println(s"step without increment: counter value=${peek(c.io.out)}")
poke(c.io.count, 1)
step(1)
println(s"step again: counter value=${peek(c.io.out)}")
} }
// ---
// # You're done!
//
// [Return to the top.](#top)
| 3.2_interlude.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sns
import os
from pandas_profiling import ProfileReport
from sqlalchemy import create_engine
from IPython.display import display
from typing import List
from pd_help_function import get_info_for_one_column,check_missing_value_columns, df_to_excel
from slidingGrind import slidingGrind
from insertForce import insertForce
from random_select_files import get_a_random_filepath_from_folder, get_files_from_folder, get_a_random_subfolder
# %matplotlib inline
# -
working_dict = r'D:\Github\at36_sliding_effort'
os.chdir(working_dict)
df = pd.read_excel(r'./cleaned_df/sliding_grind_meta_all_2021-06-04.xlsx')
df = df[df['max'] > 1] ## if df.max small than 1, means null data, we remove that
df.head()
get_info_for_one_column(df,'cycles')
get_info_for_one_column(df,'sn')
# +
df_heatmap = df.drop(columns=['st', 'fn', 'file_path',])
plt.subplots(figsize = (10,8))
sns.heatmap(df_heatmap.corr(method='pearson'), linewidth = 0.3 ,annot =False )
## https://en.wikipedia.org/wiki/Pearson_correlation_coefficient
# -
fig, ax = plt.subplots(figsize=(10,6))
sns.barplot(data = df, x = 'sn_group', y = 'mean', hue = 'cycles',palette="plasma", ax=ax)
df_groups = df[(df['sn_group'] == 1) | (df['sn_group'] == 2) |
(df['sn_group'] == 3) | (df['sn_group'] == 9)
]
fig, ax = plt.subplots(figsize=(10,6))
sns.lineplot(data = df_groups, x = 'cycles', y = 'mean', hue = 'sn_group',palette="plasma", ax=ax, estimator= 'mean', ci=1)
df_groups = df[(df['sn_group'] == 7) | (df['sn_group'] == 8) |
(df['sn_group'] == 9)
]
fig, ax = plt.subplots(figsize=(10,6))
sns.lineplot(data = df_groups, x = 'cycles', y = 'mean', hue = 'sn_group',palette="plasma", ax=ax, estimator= 'mean', ci=1)
ax = sns.relplot(data = df,x='cycles',y='mean',
kind='line',alpha=0.75,col="sn_group", col_wrap=3, palette="plasma",)
ax = sns.relplot(data = df,x='cycles',y='max',
kind='line',alpha=0.75,col="sn_group", col_wrap=3, palette="plasma",)
ax = sns.relplot(data = df,x='cycles',y='min',
kind='line',alpha=0.75,col="sn_group", col_wrap=3, palette="plasma",)
ax = sns.relplot(data = df,x='cycles',y='delta',
kind='line',alpha=0.75,col="sn_group", col_wrap=3, palette="plasma",)
ax = sns.relplot(data = df,x='cycles',y='std',
kind='line',alpha=0.75,col="sn_group", col_wrap=3, palette="plasma",)
df_ = df[df['sn_group'] ==9]
df__ = df_[df_['cycles'] ==0]
for file in df__['file_path']:
obj = slidingGrind(file)
obj.set_signal_position(-60,-10)
obj.df_signal = obj.calculate_moving_average(obj.df_signal, window_size = 50, step = 1) #! average 50 point to smooth the curve
obj.plot_signal()
# obj.print_stats()
# ## study on painting removal
# +
# df_paint = pd.read_excel(r'./data_raw/grinding_cycles_study/Paint_removal_cleaned.xlsx')
# +
# df_paint['sn_group'] = df_paint['partid'].apply(lambda x : str(x)[0])
# +
# df_plot = df_paint.groupby(by='sn_group').mean()
# df_plot.drop(columns = 'partid',inplace= True)
# sns.lineplot(data = df_plot.T,dashes = False)
# +
# df_plot = df_paint.groupby(by='sn_group').max()
# df_plot.drop(columns = 'partid',inplace= True)
# sns.lineplot(data = df_plot.T,dashes = False)
# -
| 02_griding_cycles_ds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## ${\textbf{Libraries}}$
# +
import numpy as np
import pandas as pd
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
import pickle
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# -
# ## ${\textbf{Data Import}}$
df_purchase = pd.read_csv('purchase data.csv')
# ## ${\textbf{Data Exploration}}$
df_purchase.head()
df_purchase.isnull().sum()
# ## ${\textbf{Data Segmentation}}$
# ### ${\textbf{Import Segmentation Model}}$
scaler = pickle.load(open('scaler.pickle', 'rb'))
pca = pickle.load(open('pca.pickle', 'rb'))
kmeans_pca = pickle.load(open('kmeans_pca.pickle', 'rb'))
# ### ${\textbf{Standardization}}$
features = df_purchase[['Sex', 'Marital status', 'Age', 'Education', 'Income', 'Occupation', 'Settlement size']]
df_purchase_segm_std = scaler.transform(features)
# ### ${\textbf{PCA}}$
df_purchase_segm_pca = pca.transform(df_purchase_segm_std)
# ### ${\textbf{K-means PCA}}$
purchase_segm_kmeans_pca = kmeans_pca.predict(df_purchase_segm_pca)
df_purchase_predictors = df_purchase.copy()
df_purchase_predictors['Segment'] = purchase_segm_kmeans_pca
# ## ${\textbf{Descriptive Analysis by Segments}}$
# ### ${\textbf{Data Analysis by Customer}}$
df_purchase_predictors.head()
temp1 = df_purchase_predictors[['ID', 'Incidence']].groupby(['ID'], as_index = False).count()
temp1 = temp1.set_index('ID')
temp1 = temp1.rename(columns = {'Incidence': 'N_Visits'})
temp1.head()
temp2 = df_purchase_predictors[['ID', 'Incidence']].groupby(['ID'], as_index = False).sum()
temp2 = temp2.set_index('ID')
temp2 = temp2.rename(columns = {'Incidence': 'N_Purchases'})
temp3 = temp1.join(temp2)
temp3.head()
temp3['Average_N_Purchases'] = temp3['N_Purchases'] / temp3['N_Visits']
temp3.head()
temp4 = df_purchase_predictors[['ID', 'Segment']].groupby(['ID'], as_index = False).mean()
temp4 = temp4.set_index('ID')
df_purchase_descr = temp3.join(temp4)
df_purchase_descr.head()
# ### ${\textbf{Segment Proportions}}$
segm_prop = df_purchase_descr[['N_Purchases', 'Segment']].groupby(['Segment']).count() / df_purchase_descr.shape[0]
segm_prop = segm_prop.rename(columns = {'N_Purchases': 'Segment Proportions'})
segm_prop.head()
plt.figure(figsize = (9, 6))
plt.pie(segm_prop['Segment Proportions'],
labels = ['Standard', 'Career-Focused', 'Fewer-Opportunities', 'Well-Off'],
autopct = '%1.1f%%',
colors = ('b', 'g', 'r', 'orange'))
plt.title('Segment Proportions')
# ### ${\textbf{Purchase Occasion and Purchase Incidence}}$
segments_mean = df_purchase_descr.groupby(['Segment']).mean()
segments_mean
segments_std = df_purchase_descr.groupby(['Segment']).std()
plt.figure(figsize = (9, 6))
plt.bar(x = (0, 1, 2, 3),
tick_label = ('Standard', 'Career-Focused', 'Fewer-Opportunities', 'Well-Off'),
height = segments_mean['N_Visits'],
yerr = segments_std['N_Visits'],
color = ('b', 'g', 'r', 'orange'))
plt.xlabel('Segment')
plt.ylabel('Number of Store Visits')
plt.title('Average Number of Store Visits by Segment')
plt.figure(figsize = (9, 6))
plt.bar(x = (0, 1, 2, 3),
tick_label = ('Standard', 'Career-Focused', 'Fewer-Opportunities', 'Well-Off'),
height = segments_mean['N_Purchases'],
yerr = segments_std['N_Purchases'],
color = ('b', 'g', 'r', 'orange'))
plt.xlabel('Segment')
plt.ylabel('Purchase Incidences')
plt.title('Number of Purchases by Segment')
# ## ${\textbf{Homework}}$
# ### ${\textbf{Average number of store visits by segments}}$
plt.figure(figsize = (9, 6))
plt.bar(x = (0, 1, 2, 3),
tick_label = ('Standard','Career Focused','Fewer Opportunities','Well-off'),
height = segments_mean['Average_N_Purchases'],
yerr = segments_std['Average_N_Purchases'],
color = ('b', 'g', 'r', 'orange'))
plt.xlabel('Segment')
plt.ylabel('Purchase Incidences')
plt.title('Average Number of Purchases by Segment')
| 21 - Customer Analytics in Python/7_Descriptive Analyses by Segments/3_Purchase Analytics Descriptive Statistics: Homework/Purchase Analytics Descriptive Analysis 8.2 Homework Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.6 64-bit (''pt1.9'': conda)'
# name: python3
# ---
# + [markdown] id="F77yOqgkX8p4"
# <a href="https://colab.research.google.com/github/open-mmlab/mmpose/blob/main/demo/MMPose_Tutorial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9_h0e90xzw0w"
# # MMPose Tutorial
#
# Welcome to MMPose colab tutorial! In this tutorial, we will show you how to
# - perform inference with an MMPose model
# - train a new mmpose model with your own datasets
#
# Let's start!
# + [markdown] id="bMVTUneIzw0x"
# ## Install MMPose
#
# We recommend to use a conda environment to install mmpose and its dependencies. And compilers `nvcc` and `gcc` are required.
# + colab={"base_uri": "https://localhost:8080/"} id="9dvKWH89zw0x" outputId="c3e29ad4-6a1b-4ef8-ec45-93196de7ffae"
# check NVCC version
# !nvcc -V
# check GCC version
# !gcc --version
# check python in conda environment
# !which python
# + colab={"base_uri": "https://localhost:8080/"} id="26-3yY31zw0y" outputId="7e6f3bae-7cf0-47b1-e6fd-9a8fd1cec453"
# install dependencies: (use cu111 because colab has CUDA 11.1)
# %pip install torch==1.10.0+cu111 torchvision==0.11.0+cu111 -f https://download.pytorch.org/whl/torch_stable.html
# install mmcv-full thus we could use CUDA operators
# %pip install mmcv-full -f https://download.openmmlab.com/mmcv/dist/cu111/torch1.10.0/index.html
# install mmdet for inference demo
# %pip install mmdet
# clone mmpose repo
# %rm -rf mmpose
# !git clone https://github.com/open-mmlab/mmpose.git
# %cd mmpose
# install mmpose dependencies
# %pip install -r requirements.txt
# install mmpose in develop mode
# %pip install -e .
# + colab={"base_uri": "https://localhost:8080/"} id="aIEhiA44zw0y" outputId="31e36b6e-29a7-4f21-dc47-22905c6a48ca"
# Check Pytorch installation
import torch, torchvision
print('torch version:', torch.__version__, torch.cuda.is_available())
print('torchvision version:', torchvision.__version__)
# Check MMPose installation
import mmpose
print('mmpose version:', mmpose.__version__)
# Check mmcv installation
from mmcv.ops import get_compiling_cuda_version, get_compiler_version
print('cuda version:', get_compiling_cuda_version())
print('compiler information:', get_compiler_version())
# + [markdown] id="KyrovOnDzw0z"
# ## Inference with an MMPose model
#
# MMPose provides high level APIs for model inference and training.
# + colab={"base_uri": "https://localhost:8080/", "height": 421} id="AaUNCi28zw0z" outputId="441a8335-7795-42f8-c48c-d37149ca85a8"
import cv2
from mmpose.apis import (inference_top_down_pose_model, init_pose_model,
vis_pose_result, process_mmdet_results)
from mmdet.apis import inference_detector, init_detector
local_runtime = False
try:
from google.colab.patches import cv2_imshow # for image visualization in colab
except:
local_runtime = True
pose_config = 'configs/body/2d_kpt_sview_rgb_img/topdown_heatmap/coco/hrnet_w48_coco_256x192.py'
pose_checkpoint = 'https://download.openmmlab.com/mmpose/top_down/hrnet/hrnet_w48_coco_256x192-b9e0b3ab_20200708.pth'
det_config = 'demo/mmdetection_cfg/faster_rcnn_r50_fpn_coco.py'
det_checkpoint = 'https://download.openmmlab.com/mmdetection/v2.0/faster_rcnn/faster_rcnn_r50_fpn_1x_coco/faster_rcnn_r50_fpn_1x_coco_20200130-047c8118.pth'
# initialize pose model
pose_model = init_pose_model(pose_config, pose_checkpoint)
# initialize detector
det_model = init_detector(det_config, det_checkpoint)
img = 'tests/data/coco/000000196141.jpg'
# inference detection
mmdet_results = inference_detector(det_model, img)
# extract person (COCO_ID=1) bounding boxes from the detection results
person_results = process_mmdet_results(mmdet_results, cat_id=1)
# inference pose
pose_results, returned_outputs = inference_top_down_pose_model(pose_model,
img,
person_results,
bbox_thr=0.3,
format='xyxy',
dataset=pose_model.cfg.data.test.type)
# show pose estimation results
vis_result = vis_pose_result(pose_model,
img,
pose_results,
dataset=pose_model.cfg.data.test.type,
show=False)
# reduce image size
vis_result = cv2.resize(vis_result, dsize=None, fx=0.5, fy=0.5)
if local_runtime:
from IPython.display import Image, display
import tempfile
import os.path as osp
with tempfile.TemporaryDirectory() as tmpdir:
file_name = osp.join(tmpdir, 'pose_results.png')
cv2.imwrite(file_name, vis_result)
display(Image(file_name))
else:
cv2_imshow(vis_result)
# + [markdown] id="mOulhU_Wsr_S"
# ## Train a pose estimation model on a customized dataset
#
# To train a model on a customized dataset with MMPose, there are usually three steps:
# 1. Support the dataset in MMPose
# 1. Create a config
# 1. Perform training and evaluation
#
# ### Add a new dataset
#
# There are two methods to support a customized dataset in MMPose. The first one is to convert the data to a supported format (e.g. COCO) and use the corresponding dataset class (e.g. TopdownCOCODataset), as described in the [document](https://mmpose.readthedocs.io/en/latest/tutorials/2_new_dataset.html#reorganize-dataset-to-existing-format). The second one is to add a new dataset class. In this tutorial, we give an example of the second method.
#
# We first download the demo dataset, which contains 100 samples (75 for training and 25 for validation) selected from COCO train2017 dataset. The annotations are stored in a different format from the original COCO format.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="tlSP8JNr9pEr" outputId="aee224ab-4469-40c6-8b41-8591d92aafb3"
# download dataset
# %mkdir data
# %cd data
# !wget https://openmmlab.oss-cn-hangzhou.aliyuncs.com/mmpose/datasets/coco_tiny.tar
# !tar -xf coco_tiny.tar
# %cd ..
# + colab={"base_uri": "https://localhost:8080/"} id="UDzqo6pwB-Zz" outputId="96bb444c-94c5-4b8a-cc63-0a94f16ebf95"
# check the directory structure
# !apt-get -q install tree
# !tree data/coco_tiny
# + colab={"base_uri": "https://localhost:8080/"} id="ef-045CUCdb3" outputId="5a39b30a-8e6c-4754-8908-9ea13b91c22b"
# check the annotation format
import json
import pprint
anns = json.load(open('data/coco_tiny/train.json'))
print(type(anns), len(anns))
pprint.pprint(anns[0], compact=True)
# + [markdown] id="r4Dt1io8D7m8"
# After downloading the data, we implement a new dataset class to load data samples for model training and validation. Assume that we are going to train a top-down pose estimation model (refer to [Top-down Pose Estimation](https://github.com/open-mmlab/mmpose/tree/master/configs/body/2d_kpt_sview_rgb_img/topdown_heatmap#readme) for a brief introduction), the new dataset class inherits `TopDownBaseDataset`.
# + id="WR9ZVXuPFy4v"
import json
import os
import os.path as osp
from collections import OrderedDict
import tempfile
import numpy as np
from mmpose.core.evaluation.top_down_eval import (keypoint_nme,
keypoint_pck_accuracy)
from mmpose.datasets.builder import DATASETS
from mmpose.datasets.datasets.base import Kpt2dSviewRgbImgTopDownDataset
@DATASETS.register_module()
class TopDownCOCOTinyDataset(Kpt2dSviewRgbImgTopDownDataset):
def __init__(self,
ann_file,
img_prefix,
data_cfg,
pipeline,
dataset_info=None,
test_mode=False):
super().__init__(
ann_file, img_prefix, data_cfg, pipeline, dataset_info, coco_style=False, test_mode=test_mode)
# flip_pairs, upper_body_ids and lower_body_ids will be used
# in some data augmentations like random flip
self.ann_info['flip_pairs'] = [[1, 2], [3, 4], [5, 6], [7, 8], [9, 10],
[11, 12], [13, 14], [15, 16]]
self.ann_info['upper_body_ids'] = (0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10)
self.ann_info['lower_body_ids'] = (11, 12, 13, 14, 15, 16)
self.ann_info['joint_weights'] = None
self.ann_info['use_different_joint_weights'] = False
self.dataset_name = 'coco_tiny'
self.db = self._get_db()
def _get_db(self):
with open(self.ann_file) as f:
anns = json.load(f)
db = []
for idx, ann in enumerate(anns):
# get image path
image_file = osp.join(self.img_prefix, ann['image_file'])
# get bbox
bbox = ann['bbox']
center, scale = self._xywh2cs(*bbox)
# get keypoints
keypoints = np.array(
ann['keypoints'], dtype=np.float32).reshape(-1, 3)
num_joints = keypoints.shape[0]
joints_3d = np.zeros((num_joints, 3), dtype=np.float32)
joints_3d[:, :2] = keypoints[:, :2]
joints_3d_visible = np.zeros((num_joints, 3), dtype=np.float32)
joints_3d_visible[:, :2] = np.minimum(1, keypoints[:, 2:3])
sample = {
'image_file': image_file,
'center': center,
'scale': scale,
'bbox': bbox,
'rotation': 0,
'joints_3d': joints_3d,
'joints_3d_visible': joints_3d_visible,
'bbox_score': 1,
'bbox_id': idx,
}
db.append(sample)
return db
def _xywh2cs(self, x, y, w, h):
"""This encodes bbox(x, y, w, h) into (center, scale)
Args:
x, y, w, h
Returns:
tuple: A tuple containing center and scale.
- center (np.ndarray[float32](2,)): center of the bbox (x, y).
- scale (np.ndarray[float32](2,)): scale of the bbox w & h.
"""
aspect_ratio = self.ann_info['image_size'][0] / self.ann_info[
'image_size'][1]
center = np.array([x + w * 0.5, y + h * 0.5], dtype=np.float32)
if w > aspect_ratio * h:
h = w * 1.0 / aspect_ratio
elif w < aspect_ratio * h:
w = h * aspect_ratio
# pixel std is 200.0
scale = np.array([w / 200.0, h / 200.0], dtype=np.float32)
# padding to include proper amount of context
scale = scale * 1.25
return center, scale
def evaluate(self, results, res_folder=None, metric='PCK', **kwargs):
"""Evaluate keypoint detection results. The pose prediction results will
be saved in `${res_folder}/result_keypoints.json`.
Note:
batch_size: N
num_keypoints: K
heatmap height: H
heatmap width: W
Args:
results (list(preds, boxes, image_path, output_heatmap))
:preds (np.ndarray[N,K,3]): The first two dimensions are
coordinates, score is the third dimension of the array.
:boxes (np.ndarray[N,6]): [center[0], center[1], scale[0]
, scale[1],area, score]
:image_paths (list[str]): For example, ['Test/source/0.jpg']
:output_heatmap (np.ndarray[N, K, H, W]): model outputs.
res_folder (str, optional): The folder to save the testing
results. If not specified, a temp folder will be created.
Default: None.
metric (str | list[str]): Metric to be performed.
Options: 'PCK', 'NME'.
Returns:
dict: Evaluation results for evaluation metric.
"""
metrics = metric if isinstance(metric, list) else [metric]
allowed_metrics = ['PCK', 'NME']
for metric in metrics:
if metric not in allowed_metrics:
raise KeyError(f'metric {metric} is not supported')
if res_folder is not None:
tmp_folder = None
res_file = osp.join(res_folder, 'result_keypoints.json')
else:
tmp_folder = tempfile.TemporaryDirectory()
res_file = osp.join(tmp_folder.name, 'result_keypoints.json')
kpts = []
for result in results:
preds = result['preds']
boxes = result['boxes']
image_paths = result['image_paths']
bbox_ids = result['bbox_ids']
batch_size = len(image_paths)
for i in range(batch_size):
kpts.append({
'keypoints': preds[i].tolist(),
'center': boxes[i][0:2].tolist(),
'scale': boxes[i][2:4].tolist(),
'area': float(boxes[i][4]),
'score': float(boxes[i][5]),
'bbox_id': bbox_ids[i]
})
kpts = self._sort_and_unique_bboxes(kpts)
self._write_keypoint_results(kpts, res_file)
info_str = self._report_metric(res_file, metrics)
name_value = OrderedDict(info_str)
if tmp_folder is not None:
tmp_folder.cleanup()
return name_value
def _report_metric(self, res_file, metrics, pck_thr=0.3):
"""Keypoint evaluation.
Args:
res_file (str): Json file stored prediction results.
metrics (str | list[str]): Metric to be performed.
Options: 'PCK', 'NME'.
pck_thr (float): PCK threshold, default: 0.3.
Returns:
dict: Evaluation results for evaluation metric.
"""
info_str = []
with open(res_file, 'r') as fin:
preds = json.load(fin)
assert len(preds) == len(self.db)
outputs = []
gts = []
masks = []
for pred, item in zip(preds, self.db):
outputs.append(np.array(pred['keypoints'])[:, :-1])
gts.append(np.array(item['joints_3d'])[:, :-1])
masks.append((np.array(item['joints_3d_visible'])[:, 0]) > 0)
outputs = np.array(outputs)
gts = np.array(gts)
masks = np.array(masks)
normalize_factor = self._get_normalize_factor(gts)
if 'PCK' in metrics:
_, pck, _ = keypoint_pck_accuracy(outputs, gts, masks, pck_thr,
normalize_factor)
info_str.append(('PCK', pck))
if 'NME' in metrics:
info_str.append(
('NME', keypoint_nme(outputs, gts, masks, normalize_factor)))
return info_str
@staticmethod
def _write_keypoint_results(keypoints, res_file):
"""Write results into a json file."""
with open(res_file, 'w') as f:
json.dump(keypoints, f, sort_keys=True, indent=4)
@staticmethod
def _sort_and_unique_bboxes(kpts, key='bbox_id'):
"""sort kpts and remove the repeated ones."""
kpts = sorted(kpts, key=lambda x: x[key])
num = len(kpts)
for i in range(num - 1, 0, -1):
if kpts[i][key] == kpts[i - 1][key]:
del kpts[i]
return kpts
@staticmethod
def _get_normalize_factor(gts):
"""Get inter-ocular distance as the normalize factor, measured as the
Euclidean distance between the outer corners of the eyes.
Args:
gts (np.ndarray[N, K, 2]): Groundtruth keypoint location.
Return:
np.ndarray[N, 2]: normalized factor
"""
interocular = np.linalg.norm(
gts[:, 0, :] - gts[:, 1, :], axis=1, keepdims=True)
return np.tile(interocular, [1, 2])
# + [markdown] id="gh05C4mBl_u-"
# ### Create a config file
#
# In the next step, we create a config file which configures the model, dataset and runtime settings. More information can be found at [Learn about Configs](https://mmpose.readthedocs.io/en/latest/tutorials/0_config.html). A common practice to create a config file is deriving from a existing one. In this tutorial, we load a config file that trains a HRNet on COCO dataset, and modify it to adapt to the COCOTiny dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="n-z89qCJoWwL" outputId="a3f6817e-b448-463d-d3df-2c5519efa99c"
from mmcv import Config
cfg = Config.fromfile(
'./configs/body/2d_kpt_sview_rgb_img/topdown_heatmap/coco/hrnet_w32_coco_256x192.py'
)
# set basic configs
cfg.data_root = 'data/coco_tiny'
cfg.work_dir = 'work_dirs/hrnet_w32_coco_tiny_256x192'
cfg.gpu_ids = range(1)
cfg.seed = 0
# set log interval
cfg.log_config.interval = 1
# set evaluation configs
cfg.evaluation.interval = 10
cfg.evaluation.metric = 'PCK'
cfg.evaluation.save_best = 'PCK'
# set learning rate policy
lr_config = dict(
policy='step',
warmup='linear',
warmup_iters=10,
warmup_ratio=0.001,
step=[17, 35])
cfg.total_epochs = 40
# set batch size
cfg.data.samples_per_gpu = 16
cfg.data.val_dataloader = dict(samples_per_gpu=16)
cfg.data.test_dataloader = dict(samples_per_gpu=16)
# set dataset configs
cfg.data.train.type = 'TopDownCOCOTinyDataset'
cfg.data.train.ann_file = f'{cfg.data_root}/train.json'
cfg.data.train.img_prefix = f'{cfg.data_root}/images/'
cfg.data.val.type = 'TopDownCOCOTinyDataset'
cfg.data.val.ann_file = f'{cfg.data_root}/val.json'
cfg.data.val.img_prefix = f'{cfg.data_root}/images/'
cfg.data.test.type = 'TopDownCOCOTinyDataset'
cfg.data.test.ann_file = f'{cfg.data_root}/val.json'
cfg.data.test.img_prefix = f'{cfg.data_root}/images/'
print(cfg.pretty_text)
# + [markdown] id="WQVa6wBDxVSW"
# ### Train and Evaluation
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["c50b2c7b3d58486d9941509548a877e4", "ae33a61272f84a7981bc1f3008458688", "a0bf65a0401e465393ef8720ef3328ac", "<KEY>", "210e7151c2ad44a3ba79d477f91d8b26", "<KEY>", "<KEY>", "9035c6e9fddd41d8b7dae395c93410a2", "1d31e1f7256d42669d76f54a8a844b79", "<KEY>", "<KEY>"]} id="XJ5uVkwcxiyx" outputId="0693f2e3-f41d-46a8-d3ed-1add83735f91"
from mmpose.datasets import build_dataset
from mmpose.models import build_posenet
from mmpose.apis import train_model
import mmcv
# build dataset
datasets = [build_dataset(cfg.data.train)]
# build model
model = build_posenet(cfg.model)
# create work_dir
mmcv.mkdir_or_exist(cfg.work_dir)
# train model
train_model(
model, datasets, cfg, distributed=False, validate=True, meta=dict())
# + [markdown] id="iY2EWSp1zKoz"
# Test the trained model. Since the model is trained on a toy dataset coco-tiny, its performance would be as good as the ones in our model zoo. Here we mainly show how to inference and visualize a local model checkpoint.
# + colab={"base_uri": "https://localhost:8080/", "height": 387} id="i0rk9eCVzT_D" outputId="722542be-ab38-4ca4-86c4-dce2cfb95c4b"
from mmpose.apis import (inference_top_down_pose_model, init_pose_model,
vis_pose_result, process_mmdet_results)
from mmdet.apis import inference_detector, init_detector
local_runtime = False
try:
from google.colab.patches import cv2_imshow # for image visualization in colab
except:
local_runtime = True
pose_checkpoint = 'work_dirs/hrnet_w32_coco_tiny_256x192/latest.pth'
det_config = 'demo/mmdetection_cfg/faster_rcnn_r50_fpn_coco.py'
det_checkpoint = 'https://download.openmmlab.com/mmdetection/v2.0/faster_rcnn/faster_rcnn_r50_fpn_1x_coco/faster_rcnn_r50_fpn_1x_coco_20200130-047c8118.pth'
# initialize pose model
pose_model = init_pose_model(cfg, pose_checkpoint)
# initialize detector
det_model = init_detector(det_config, det_checkpoint)
img = 'tests/data/coco/000000196141.jpg'
# inference detection
mmdet_results = inference_detector(det_model, img)
# extract person (COCO_ID=1) bounding boxes from the detection results
person_results = process_mmdet_results(mmdet_results, cat_id=1)
# inference pose
pose_results, returned_outputs = inference_top_down_pose_model(pose_model,
img,
person_results,
bbox_thr=0.3,
format='xyxy',
dataset='TopDownCocoDataset')
# show pose estimation results
vis_result = vis_pose_result(pose_model,
img,
pose_results,
kpt_score_thr=0.,
dataset='TopDownCocoDataset',
show=False)
# reduce image size
vis_result = cv2.resize(vis_result, dsize=None, fx=0.5, fy=0.5)
if local_runtime:
from IPython.display import Image, display
import tempfile
import os.path as osp
import cv2
with tempfile.TemporaryDirectory() as tmpdir:
file_name = osp.join(tmpdir, 'pose_results.png')
cv2.imwrite(file_name, vis_result)
display(Image(file_name))
else:
cv2_imshow(vis_result)
| demo/MMPose_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 7.17 Sizing of Pump for Oil Pipelines
# Crude oil (30 degree API) at 15.6 C with a viscosity of 75 Universal Saybolt seconds flows down a 12" Schedule 30 steel pipe with a flow rate of 1900 barrels/hour.
#
# The pipeline is 50 miles long, and the net elevation change is an increase of 2000 feet above the initial pump. The pump has an efficiency of 67%.
#
# Calculate the brake horsepower of the pump.
# +
from fluids.units import *
from math import pi
L = 50*u.miles
dH = 2000*u.foot
efficiency = 0.67
# Note in pint the default barrel is for US dry barrel
Q = 1900*u.oil_barrel/u.hour
mu = 12.5*u.cP
rho = 54.64*u.lb/u.ft**3
NPS, Di, Do, t = nearest_pipe(NPS=12, schedule='30')
A = 0.25*pi*Di**2
v = Q/A
Re = rho*v*Di/mu
print(Re.to_base_units())
fd = friction_factor(Re=Re, eD=0.0022*u.inch/Di)
print(fd)
K_tot = K_from_f(fd=fd, L=L, D=Di)
dP = dP_from_K(K=K_tot, rho=rho, V=v) + rho*dH*1*u.gravity
dP.to(u.psi), v.to(u.foot/u.s)
head = head_from_P(dP, rho).to(u.foot)
print('head = %s' %head)
power = Q*dP/efficiency
print('power = %s' %(power.to(u.hp)))
# -
# The listed values are 3406.5 feet and 1496 hp, however a shortcut formula is used there.
| docs/Examples/Crane TP 410 Solved Problems/7.17 Sizing of Pump for Oil Pipelines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pook
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="hUkBRdY8ndhZ"
from matplotlib.pyplot import imshow
import matplotlib.cm as cm
import matplotlib.pylab as plt
from tensorflow.keras.preprocessing.image import ImageDataGenerator
import tensorflow as tf
import numpy as np
import PIL
from PIL import ImageFilter
import cv2
import itertools
import random
from tensorflow import keras
import imutils
from imutils import paths
import os
from tensorflow.keras import optimizers
from tensorflow.keras.preprocessing.image import img_to_array
from sklearn.model_selection import train_test_split
from tensorflow.keras.utils import to_categorical
from tensorflow.keras import callbacks
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.layers import Conv2D, MaxPooling2D , UpSampling2D ,Conv2DTranspose
from tensorflow.keras import backend as K
# %matplotlib inline
# + colab={} colab_type="code" id="JHEynQv2ndhn"
def pil_image(img_path):
pil_im =PIL.Image.open(img_path).convert('L')
pil_im=pil_im.resize((105,105))
#imshow(np.asarray(pil_im))
return pil_im
# + [markdown] colab_type="text" id="1hbTCU2qndht"
# # Augumentation Steps
# 1) Noise
# 2) Blur
# 3) Perpective Rotation
# 4) Shading
# 5) Variable Character Spacing
# 6) Variable Aspect Ratio
# + colab={} colab_type="code" id="MLCHbBKsndhv"
def noise_image(pil_im):
# Adding Noise to image
img_array = np.asarray(pil_im)
mean = 0.0 # some constant
std = 5 # some constant (standard deviation)
noisy_img = img_array + np.random.normal(mean, std, img_array.shape)
noisy_img_clipped = np.clip(noisy_img, 0, 255)
noise_img = PIL.Image.fromarray(np.uint8(noisy_img_clipped)) # output
#imshow((noisy_img_clipped ).astype(np.uint8))
noise_img=noise_img.resize((105,105))
return noise_img
# + colab={} colab_type="code" id="5TPvDBV5ndh2"
def blur_image(pil_im):
#Adding Blur to image
blur_img = pil_im.filter(ImageFilter.GaussianBlur(radius=3)) # ouput
#imshow(blur_img)
blur_img=blur_img.resize((105,105))
return blur_img
# + colab={} colab_type="code" id="CIDSvv7Qndh6"
def affine_rotation(img):
#img=cv2.imread(img_path,0)
rows, columns = img.shape
point1 = np.float32([[10, 10], [30, 10], [10, 30]])
point2 = np.float32([[20, 15], [40, 10], [20, 40]])
A = cv2.getAffineTransform(point1, point2)
output = cv2.warpAffine(img, A, (columns, rows))
affine_img = PIL.Image.fromarray(np.uint8(output)) # affine rotated output
#imshow(output)
affine_img=affine_img.resize((105,105))
return affine_img
# + colab={} colab_type="code" id="VCy6ReUNndh_"
def gradient_fill(image):
#image=cv2.imread(img_path,0)
laplacian = cv2.Laplacian(image,cv2.CV_64F)
laplacian = cv2.resize(laplacian, (105, 105))
return laplacian
# + [markdown] colab_type="text" id="OBLjVHT9ndiF"
# ## Preparing Dataset
# + colab={} colab_type="code" id="6hc1RAaVndiI"
data_path = "TextRecognitionDataGenerator/font_patch2/"
data=[]
labels=[]
imagePaths = sorted(list(paths.list_images(data_path)))
random.seed(42)
random.shuffle(imagePaths)
# + colab={} colab_type="code" id="HYYzr_c1ndiN"
"""def conv_label(label):
if label == 'Lato':
return 0
elif label == 'Raleway':
return 1
elif label == 'Roboto':
return 2
elif label == 'Sansation':
return 3
elif label == 'Walkway':
return 4"""
num_labelss=7
def conv_label(label):
if label == 'kakugo_0':
return 0
elif label == 'kakugo_2':
return 1
elif label == 'kakugo_4':
return 2
elif label == 'kakugo_6':
return 3
elif label == 'kakugo_8':
return 4
elif label == 'marugo_4':
return 5
elif label == 'mincho_4':
return 6
# + colab={} colab_type="code" id="L5emmKqjd-3Q"
augument=["blur","noise","affine","gradient"]
a=itertools.combinations(augument, 4)
for i in list(a):
print(list(i))
# + colab={} colab_type="code" id="PIc22kLf4SAP"
counter=0
for imagePath in imagePaths:
label = imagePath.split(os.path.sep)[-2]
label = conv_label(label)
pil_img = pil_image(imagePath)
#imshow(pil_img)
# Adding original image
org_img = img_to_array(pil_img)
#print(org_img.shape)
data.append(org_img)
labels.append(label)
augument=["noise","blur","affine","gradient"]
for l in range(0,len(augument)):
a=itertools.combinations(augument, l+1)
for i in list(a):
combinations=list(i)
print(len(combinations))
temp_img = pil_img
for j in combinations:
if j == 'noise':
# Adding Noise image
temp_img = noise_image(temp_img)
elif j == 'blur':
# Adding Blur image
temp_img = blur_image(temp_img)
#imshow(blur_img)
elif j == 'affine':
open_cv_affine = np.array(pil_img)
# Adding affine rotation image
temp_img = affine_rotation(open_cv_affine)
elif j == 'gradient':
open_cv_gradient = np.array(pil_img)
# Adding gradient image
temp_img = gradient_fill(open_cv_gradient)
temp_img = img_to_array(temp_img)
data.append(temp_img)
labels.append(label)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="cFpIsgdHndit" outputId="084a49bf-ee2d-4067-cbde-90d9b42bd8c6"
data = np.asarray(data, dtype="float") / 255.0
labels = np.array(labels)
print("Success")
# partition the data into training and testing splits using 75% of
# the data for training and the remaining 25% for testing
(trainX, testX, trainY, testY) = train_test_split(data,
labels, test_size=0.25, random_state=42,stratify=labels)
# + colab={} colab_type="code" id="1NQr6OCQ_3qO"
# convert the labels from integers to vectors
trainY = to_categorical(trainY, num_classes=num_labelss)
testY = to_categorical(testY, num_classes=num_labelss)
# + colab={} colab_type="code" id="9omeq7fqryGW"
aug = ImageDataGenerator(rotation_range=30, width_shift_range=0.1,height_shift_range=0.1, shear_range=0.2, zoom_range=0.2,horizontal_flip=True)
# + colab={} colab_type="code" id="-vWihISP8kHV"
#K.set_image_dim_ordering('tf')
# + colab={} colab_type="code" id="DpDdwzQguqWR"
def create_model():
model=Sequential()
# Cu Layers
model.add(Conv2D(64, kernel_size=(48, 48), activation='relu', input_shape=(105,105,1)))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(128, kernel_size=(24, 24), activation='relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2DTranspose(128, (24,24), strides = (2,2), activation = 'relu', padding='same', kernel_initializer='uniform'))
model.add(UpSampling2D(size=(2, 2)))
model.add(Conv2DTranspose(64, (12,12), strides = (2,2), activation = 'relu', padding='same', kernel_initializer='uniform'))
model.add(UpSampling2D(size=(2, 2)))
#Cs Layers
model.add(Conv2D(256, kernel_size=(12, 12), activation='relu'))
model.add(Conv2D(256, kernel_size=(12, 12), activation='relu'))
model.add(Conv2D(256, kernel_size=(12, 12), activation='relu'))
model.add(Flatten())
model.add(Dense(4096, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(4096,activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(2383,activation='relu'))
model.add(Dense(num_labelss, activation='softmax'))
return model
# -
# ##START TRAINING
# + colab={} colab_type="code" id="LSUkpdoI2J-M"
batch_size = 128
epochs = 50
model= create_model()
sgd = optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
model.compile(loss='mean_squared_error', optimizer=sgd, metrics=['accuracy'])
# + colab={} colab_type="code" id="IH8DclwlLkOw"
early_stopping=callbacks.EarlyStopping(monitor='val_loss', min_delta=0, patience=10, verbose=0, mode='min')
filepath="top_model.h5"
checkpoint = callbacks.ModelCheckpoint(filepath, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
callbacks_list = [early_stopping,checkpoint]
# + colab={} colab_type="code" id="ZfjlSwNt73XO"
model.fit(trainX, trainY,shuffle=True,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(testX, testY),callbacks=callbacks_list)
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="QLtRqPzhLOUF" outputId="7de4cd06-136d-424b-dd2d-6d1d85487384"
score = model.evaluate(testX, testY, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# -
# ##START TEST
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="9oDaZS8LuWem" outputId="c308968b-442f-49da-a69e-1a87bdb1cf27"
from tensorflow.keras.models import load_model
model = load_model('hiragana_128_100_20220110-055412.h5')#hiragana_128_100_adam_20220110-055359.h5')
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="ltfB09zptlNN" outputId="e023e99b-eb5a-449a-e08f-b124b3f7f284"
score = model.evaluate(testX, testY, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# + colab={} colab_type="code" id="ov6w2Kmdv4dV"
img_path="TextRecognitionDataGenerator/sample2/こんにちは_3.jpg"
pil_im =PIL.Image.open(img_path).convert('L')
#pil_im=blur_image(pil_im)
pil_im=pil_im.resize((105,105))
org_img = img_to_array(pil_im)/255.0
# + colab={} colab_type="code" id="jN4su5FX3MzC"
"""def rev_conv_label(label):
if label == 0 :
return 'Lato'
elif label == 1:
return 'Raleway'
elif label == 2 :
return 'Roboto'
elif label == 3 :
return 'Sansation'
elif label == 4:
return 'Walkway'"""
def rev_conv_label(label):
if label == 0:
return 'kakugo_0'
elif label == 1:
return 'kakugo_2'
elif label == 2:
return 'kakugo_4'
elif label == 3:
return 'kakugo_6'
elif label == 4:
return 'kakugo_8'
elif label == 5:
return 'marugo_4'
elif label == 6:
return 'mincho_4'
# -
org_img.shape
# + colab={} colab_type="code" id="q1yBSPTh0ooD"
###3##33f3gaa##sasdfasaadata=[]
data=data.tolist()
data.append(org_img)
data = np.asarray(data, dtype="float") #/ 255.0
# + colab={} colab_type="code" id="JR2YCKaaznhT"
y = model.predict_classes(data)
# + colab={"base_uri": "https://localhost:8080/", "height": 268} colab_type="code" id="SQjS-Iv80iLc" outputId="75bc8aff-55d9-4675-bd36-96070ecfcf49"
label = rev_conv_label(int(y[0]))
fig, ax = plt.subplots(1)
ax.imshow(pil_im, interpolation='nearest', cmap=cm.gray)
ax.text(5, 5, label , bbox={'facecolor': 'white', 'pad': 10})
plt.show()
# -
print(trainX.shape,testX.shape)
1680+560
data.shape
2240/7/20
# +
label = rev_conv_label(int(y[0]))
fig1, ax1 = plt.subplots(1,5,figsize=(12,12))
ax1[0].imshow(data[0], cmap=cm.gray)
ax1[1].imshow(data[1], cmap=cm.gray)
ax1[2].imshow(data[2], cmap=cm.gray)
ax1[3].imshow(data[3], cmap=cm.gray)
ax1[4].imshow(data[4], cmap=cm.gray)
#ax.text(5, 5, label , bbox={'facecolor': 'white', 'pad': 10})
plt.show()
# -
label = rev_conv_label(int(y[0]))
fig1, ax1 = plt.subplots(1)
ax1.imshow(data[], interpolation='nearest', cmap=cm.gray)
#ax.text(5, 5, label , bbox={'facecolor': 'white', 'pad': 10})
plt.show()
| Font_Rec(DeepFont).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import all the necessary files!
import os
import zipfile
import random
import shutil
from os import getcwd
from shutil import copyfile
import tensorflow as tf
from tensorflow.keras import layers, Model
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.layers import Conv2D, Dense, MaxPooling2D, Flatten
# + jupyter={"outputs_hidden": true} tags=[]
# Download the inception v3 weights
# !wget --no-check-certificate \
# https://storage.googleapis.com/mledu-datasets/inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5 \
# -O inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5
path_inception = "inception_v3_weights_tf_dim_ordering_tf_kernels_notop.h5"
# Import the inception model
from tensorflow.keras.applications.inception_v3 import InceptionV3
# Create an instance of the inception model from the local pre-trained weights
local_weights_file = path_inception
pre_trained_model = InceptionV3(input_shape = (150, 150, 3),
include_top = False,
weights = None)
pre_trained_model.load_weights(local_weights_file)
# Make all the layers in the pre-trained model non-trainable
for layer in pre_trained_model.layers:
layer.trainable = False
# Print the model summary
pre_trained_model.summary()
# -
last_layer = pre_trained_model.get_layer("mixed7")
print('last layer output shape: ', last_layer.output_shape)
last_output = last_layer.output
# Define a Callback class that stops training once accuracy reaches 97.0%
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs={}):
if logs.get('accuracy') and (logs.get('accuracy')>0.97):
print("\nReached 99% accuracy so cancelling training!")
self.model.stop_training = True
# + jupyter={"outputs_hidden": true} tags=[]
from tensorflow.keras.optimizers import RMSprop
x = layers.Flatten()(last_output)
x = layers.Dense(1024, activation='relu')(x)
x = layers.Dropout(0.2)(x)
x = layers.Dense (1, activation='sigmoid')(x)
model = Model( pre_trained_model.input, x)
model.compile(optimizer = RMSprop(lr=0.0001),
loss = 'binary_crossentropy',
metrics = ['accuracy'])
model.summary()
# +
# Run this and see how many epochs it should take before the callback
# fires, and stops training at 97% accuracy
callbacks = myCallback()
history = model.fit_generator(
train_generator,
validation_data = validation_generator,
epochs = 3,
# validation_steps = 50,
verbose = 2,
callbacks = [callbacks])
# +
# %matplotlib inline
import matplotlib.pyplot as plt
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'r', label='Training accuracy')
plt.plot(epochs, val_acc, 'b', label='Validation accuracy')
plt.title('Training and validation accuracy')
plt.legend(loc=0)
plt.figure()
plt.show()
| Course2/Course_2_Week_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.preprocessing import Normalizer
from sklearn.neighbors import KNeighborsClassifier
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
import numpy as np
# +
features = ['duration','protocol_type','service','flag','src_bytes','dst_bytes','land','wrong_fragment','urgent','hot','num_failed_logins','logged_in','num_compromised','root_shell','su_attempted',
'num_root','num_file_creations','num_shells','num_access_files','num_outbound_cmds',
'is_host_login',
'is_guest_login',
'count',
'srv_count',
'serror_rate',
'srv_serror_rate',
'rerror_rate',
'srv_rerror_rate',
'same_srv_rate',
'diff_srv_rate',
'srv_diff_host_rate',
'dst_host_count',
'dst_host_srv_count',
'dst_host_same_srv_rate',
'dst_host_diff_srv_rate',
'dst_host_same_src_port_rate',
'dst_host_srv_diff_host_rate',
'dst_host_serror_rate',
'dst_host_srv_serror_rate',
'dst_host_rerror_rate',
'dst_host_srv_rerror_rate',
'intrusion_type']
print(len(features))
# -
data = pd.read_csv('dataset/kddcup.data_10_percent', names=features, header=None)
data.head(10)
print('The no of data points are:',data.shape[0])
print('The no of features are:',data.shape[1])
print('Some of the features are:',features[:10])
output = data['intrusion_type'].values
labels = set(output)
print('The different type of output labels are:',labels)
# # Data Cleaning
print('Null values in the dataset are: ',len(data[data.isnull().any(1)]))
data.drop_duplicates(subset=features, keep='first', inplace=True)
data.shape
plt.figure(figsize=(20,15))
class_distribution = data['intrusion_type'].value_counts()
class_distribution.plot(kind='bar')
plt.xlabel('Class')
plt.ylabel('Data points per Class')
plt.title('Distribution of yi in train data')
plt.grid()
plt.show()
# Most of the data points are from "normal" category which is around 60.33 %.
#
# In the categories that belong to bad connections, "neptune." 35.594 % and "back." 0.665 % have the highest no of data points.
#
# Classes "rootkit.", "loadmodule.", "ftp_write.", "multihop.", "phf.", "perl.", "spy." have the least no of data points with less than 10 data points per class.
# # Feature Extraction
data['num_outbound_cmds'].value_counts()
data.drop('num_outbound_cmds', axis=1, inplace=True)
data['is_host_login'].value_counts()
data.drop('is_host_login', axis=1, inplace=True)
# # Transformation of categorical values
# +
data['protocol_type'] = data['protocol_type'].astype('category')
data['service'] = data['service'].astype('category')
data['flag'] = data['flag'].astype('category')
cat_columns = data.select_dtypes(['category']).columns
data[cat_columns] = data[cat_columns].apply(lambda x: x.cat.codes)
# -
X = data.drop('intrusion_type', axis=1)
Y = data['intrusion_type']
data.replace(to_replace = ['ipsweep.', 'portsweep.', 'nmap.', 'satan.'], value = 'probe', inplace = True)
data.replace(to_replace = ['ftp_write.', 'guess_passwd.', 'imap.', 'multihop.', 'phf.', 'spy.', 'warezclient.', 'warezmaster.'], value = 'r2l', inplace = True)
data.replace(to_replace = ['buffer_overflow.', 'loadmodule.', 'perl.', 'rootkit.'], value = 'u2r', inplace = True)
data.replace(to_replace = ['back.', 'land.' , 'neptune.', 'pod.', 'smurf.', 'teardrop.'], value = 'dos', inplace = True)
# # Standardization
# +
from sklearn.preprocessing import StandardScaler
sScaler = StandardScaler()
rescaleX = sScaler.fit_transform(X)
names_inputed =features[0:39]
data = pd.DataFrame(data=rescaleX, columns=names_inputed)
# -
# # Normalization
#
from sklearn.preprocessing import Normalizer
norm = Normalizer()
X = norm.fit_transform(X)
# # Applying Machine Learning Algorithm
import datetime as dt
from sklearn.metrics import accuracy_score, confusion_matrix, roc_auc_score, precision_score, recall_score, f1_score, classification_report
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
from sklearn.externals import joblib
from sklearn.metrics import confusion_matrix
import seaborn as sn
import seaborn as sns
import matplotlib.pyplot as plt
# +
def confusion_matrix_func(Y_test, y_test_pred):
C = confusion_matrix(Y_test, y_test_pred)
cm_df = pd.DataFrame(C)
labels = ['dos','normal','probe', 'r2l', 'u2r']
sns.set(font_scale=1.1)
plt.figure(figsize=(8,6))
ax = sns.heatmap(cm_df, annot=True,
fmt='g', cmap='Blues', xticklabels=labels, yticklabels=labels)
plt.ylabel('Actual Class')
plt.xlabel('Predicted Class')
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
# +
results_test = dict()
results_test['accuracy'] = 0
results_test['precision'] = 0
results_test['recall'] = 0
results_test['f1_score'] = 0
def reset_result():
results_test['accuracy'] = 0
results_test['precision'] = 0
results_test['recall'] = 0
results_test['f1_score'] = 0
print('Prediction on test data:')
def print_result():
print('Accuracy score is:')
print(results_test['accuracy'] / 5)
print('='*50)
print('Precision score is:')
print(results_test['precision'] / 5)
print('='*50)
print('Recall score is:')
print(results_test['recall'] / 5)
print('='*50)
print('F1-score is:')
print(results_test['f1_score'] / 5)
return results_test
# -
def model(model_name, X_train, Y_train, X_test, Y_test):
model_name.fit(X_train, Y_train)
y_test_pred = model_name.predict(X_test)
print('='*50)
print('Classification Report: ')
result_classification_report = classification_report(Y_test, y_test_pred)
print(result_classification_report)
results_test['accuracy'] += accuracy_score(Y_test, y_test_pred)
results_test['precision'] += precision_score(Y_test, y_test_pred, average='weighted')
results_test['recall'] += recall_score(Y_test, y_test_pred, average='weighted')
results_test['f1_score'] += f1_score(Y_test, y_test_pred, average='weighted')
print('Confusion Matrix is:')
confusion_matrix_func(Y_test, y_test_pred)
X = np.array(X)
Y = np.array(Y)
def kfold_validation(classifer):
kf = KFold(n_splits=5)
start = dt.datetime.now()
reset_result()
for train_index, test_index in kf.split(X, Y):
X_train, X_test = X[train_index], X[test_index]
Y_train, Y_test = Y[train_index], Y[test_index]
model(classifer, X_train, Y_train, X_test, Y_test)
print_result()
print('Completed')
print('Total time:',dt.datetime.now()-start)
# %matplotlib inline
# # Model 1: Decision Tree
# +
from sklearn import tree
from sklearn.tree import DecisionTreeClassifier
decision_tree = DecisionTreeClassifier(criterion='gini',class_weight='balanced')
kfold_validation(decision_tree)
# +
from sklearn.ensemble import RandomForestClassifier
randomForest = RandomForestClassifier(n_estimators=100)
kfold_validation(randomForest)
# -
# # Model 3: Naive Bayes
from sklearn.naive_bayes import GaussianNB
gaussian_nb = GaussianNB()
kfold_validation(gaussian_nb)
# # Model 4: KNN
# +
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(3)
kfold_validation(knn)
| ML_Intrusion_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # I. Introduction
#
# 
# As illustrated in the lectures, simulated annealing is a probablistic technique used for finding an approximate solution to an optimization problem--one of the simplest "gradient-free" optimization techniques. In this exercise you will check your understanding by implementing [simulated annealing](https://en.wikipedia.org/wiki/Simulated_annealing) to solve the [Traveling Salesman Problem](https://en.wikipedia.org/wiki/Travelling_salesman_problem) (TSP) between US state capitals. Briefly, the TSP is an optimization problem that seeks to find the shortest path passing through every city exactly once. In our example the TSP path is defined to start and end in the same city (so the path is a closed loop).
#
# Image Source: [Simulated Annealing - By Kingpin13 (Own work) [CC0], via Wikimedia Commons (Attribution not required)](https://commons.wikimedia.org/wiki/File:Hill_Climbing_with_Simulated_Annealing.gif)
# ## Overview
#
# Students should read through the code, then:
#
# 0. Implement the `simulated_annealing()` main loop function in Section II
# 0. Complete the `TravelingSalesmanProblem` class by implementing the `successors()` and `get_value()` methods in section III
# 0. Complete the `schedule()` function to define the temperature schedule in Section IV
# 0. Use the completed algorithm and problem description to experiment with simulated annealing to solve larger TSP instances on the map of US capitals
# +
import json
import copy
import numpy as np # contains helpful math functions like numpy.exp()
import numpy.random as random # see numpy.random module
# import random # alternative to numpy.random module
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# %matplotlib inline
# +
"""Read input data and define helper functions for visualization."""
# Map services and data available from U.S. Geological Survey, National Geospatial Program.
# Please go to http://www.usgs.gov/visual-id/credit_usgs.html for further information
map = mpimg.imread("map.png") # US States & Capitals map
# List of 30 US state capitals and corresponding coordinates on the map
with open('capitals.json', 'r') as capitals_file:
capitals = json.load(capitals_file)
capitals_list = list(capitals.items())
def show_path(path, starting_city, w=12, h=8):
"""Plot a TSP path overlaid on a map of the US States & their capitals."""
x, y = list(zip(*path))
_, (x0, y0) = starting_city
plt.imshow(map)
plt.plot(x0, y0, 'y*', markersize=15) # y* = yellow star for starting point
plt.plot(x + x[:1], y + y[:1]) # include the starting point at the end of path
plt.axis("off")
fig = plt.gcf()
fig.set_size_inches([w, h])
# -
# ## II. Simulated Annealing -- Main Loop
#
# The main loop of simulated annealing repeatedly generates successors in the neighborhood of the current state and considers moving there according to an acceptance probability distribution parameterized by a cooling schedule. See the [simulated-annealing function](https://github.com/aimacode/aima-pseudocode/blob/master/md/Simulated-Annealing.md) pseudocode from the AIMA textbook online at github. Note that our Problem class is already a "node", so the MAKE-NODE line is not required.
def simulated_annealing(problem, schedule):
"""The simulated annealing algorithm, a version of stochastic hill climbing
where some downhill moves are allowed. Downhill moves are accepted readily
early in the annealing schedule and then less often as time goes on. The
schedule input determines the value of the temperature T as a function of
time. [Norvig, AIMA Chapter 3]
Parameters
----------
problem : Problem
An optimization problem, already initialized to a random starting state.
The Problem class interface must implement a callable method
"successors()" which returns states in the neighborhood of the current
state, and a callable function "get_value()" which returns a fitness
score for the state. (See the `TravelingSalesmanProblem` class below
for details.)
schedule : callable
A function mapping time to "temperature". "Time" is equivalent in this
case to the number of loop iterations.
Returns
-------
Problem
An approximate solution state of the optimization problem
Notes
-----
(1) DO NOT include the MAKE-NODE line from the AIMA pseudocode
(2) Modify the termination condition to return when the temperature
falls below some reasonable minimum value (e.g., 1e-10) rather than
testing for exact equality to zero
See Also
--------
AIMA simulated_annealing() pseudocode
https://github.com/aimacode/aima-pseudocode/blob/master/md/Simulated-Annealing.md
"""
Current = problem
for t in range(1,1000000):
T=schedule(t)
if T < 1e-10: return Current
successors = Current.successors()
Next = random.choice(successors)
deltaE = Next.get_value() - Current.get_value()
if deltaE > 0:
Current = Next
else:
prob = np.exp(deltaE/T)
if random.random() < prob:
Current = Next
# ## III. Representing the Problem
#
# In order to use simulated annealing we need to build a representation of the problem domain. The choice of representation can have a significant impact on the performance of simulated annealing and other optimization techniques. Since the TSP deals with a closed loop that visits each city in a list once, we will represent each city by a tuple containing the city name and its position specified by an (x,y) location on a grid. The _state_ will then consist of an ordered sequence (a list) of the cities; the path is defined as the sequence generated by traveling from each city in the list to the next in order. By default you should use the Euclidean distance metric to measure the path length.
class TravelingSalesmanProblem:
"""Representation of a traveling salesman optimization problem. The goal
is to find the shortest path that visits every city in a closed loop path.
Students should only need to implement or modify the successors() and
get_values() methods.
Parameters
----------
cities : list
A list of cities specified by a tuple containing the name and the x, y
location of the city on a grid. e.g., ("Atlanta", (585.6, 376.8))
Attributes
----------
names
coords
path : list
The current path between cities as specified by the order of the city
tuples in the list.
"""
def __init__(self, cities):
self.path = copy.deepcopy(cities)
def copy(self):
"""Return a copy of the current board state."""
new_tsp = TravelingSalesmanProblem(self.path)
return new_tsp
@property
def names(self):
"""Strip and return only the city name from each element of the
path list. For example,
[("Atlanta", (585.6, 376.8)), ...] -> ["Atlanta", ...]
"""
names, _ = zip(*self.path)
return names
@property
def coords(self):
"""Strip the city name from each element of the path list and return
a list of tuples containing only pairs of xy coordinates for the
cities. For example,
[("Atlanta", (585.6, 376.8)), ...] -> [(585.6, 376.8), ...]
"""
_, coords = zip(*self.path)
return coords
def successors(self):
"""Return a list of states in the neighborhood of the current state by
switching the order in which any adjacent pair of cities is visited.
For example, if the current list of cities (i.e., the path) is [A, B, C, D]
then the neighbors will include [A, B, D, C], [A, C, B, D], [B, A, C, D],
and [D, B, C, A]. (The order of successors does not matter.)
In general, a path of N cities will have N neighbors (note that path wraps
around the end of the list between the first and last cities).
Returns
-------
list<Problem>
A list of TravelingSalesmanProblem instances initialized with their list
of cities set to one of the neighboring permutations of cities in the
present state
"""
current_path = copy.deepcopy(self.path)
neigh = []
for i in range(len(self.path)):
self.path = copy.deepcopy(current_path)
self.path[i-1], self.path[i] = self.path[i], self.path[i-1]
neigh.append(self.copy())
self.path = current_path
return neigh
def get_value(self):
"""Calculate the total length of the closed-circuit path of the current
state by summing the distance between every pair of adjacent cities. Since
the default simulated annealing algorithm seeks to maximize the objective
function, return -1x the path length. (Multiplying by -1 makes the smallest
path the smallest negative number, which is the maximum value.)
Returns
-------
float
A floating point value with the total cost of the path given by visiting
the cities in the order according to the self.cities list
Notes
-----
(1) Remember to include the edge from the last city back to the
first city
(2) Remember to multiply the path length by -1 so that simulated
annealing finds the shortest path
"""
Acc_Dist = 0.0
Coord = self.coords
for i in range(len(self.path)):
dist = np.sqrt((Coord[i][0]-Coord[i-1][0])**2 + (Coord[i][1]-Coord[i-1][1])**2)
Acc_Dist += dist
Acc_Dist *= (-1.0)
return Acc_Dist
# ### Testing TravelingSalesmanProblem
# The following tests should validate the class constructor and functionality of the `successors()` and `get_value()` methods.
# Construct an instance of the TravelingSalesmanProblem
test_cities = [('DC', (11, 1)), ('SF', (0, 0)), ('PHX', (2, -3)), ('LA', (0, -4))]
tsp = TravelingSalesmanProblem(test_cities)
assert(tsp.path == test_cities)
# Test the successors() method -- no output means the test passed
successor_paths = [x.path for x in tsp.successors()]
assert(all(x in [[('LA', (0, -4)), ('SF', (0, 0)), ('PHX', (2, -3)), ('DC', (11, 1))],
[('SF', (0, 0)), ('DC', (11, 1)), ('PHX', (2, -3)), ('LA', (0, -4))],
[('DC', (11, 1)), ('PHX', (2, -3)), ('SF', (0, 0)), ('LA', (0, -4))],
[('DC', (11, 1)), ('SF', (0, 0)), ('LA', (0, -4)), ('PHX', (2, -3))]]
for x in successor_paths))
# Test the get_value() method -- no output means the test passed
assert(np.allclose(tsp.get_value(), -28.97, atol=1e-3))
# ## IV. Define the Temperature Schedule
#
# The most common temperature schedule is simple exponential decay:
# $T(t) = \alpha^t T_0$
#
# (Note that this is equivalent to the incremental form $T_{i+1} = \alpha T_i$, but implementing that form is slightly more complicated because you need to preserve state between calls.)
#
# In most cases, the valid range for temperature $T_0$ can be very high (e.g., 1e8 or higher), and the _decay parameter_ $\alpha$ should be close to, but less than 1.0 (e.g., 0.95 or 0.99). Think about the ways these parameters effect the simulated annealing function. Try experimenting with both parameters to see how it changes runtime and the quality of solutions.
#
# You can also experiment with other schedule functions -- linear, quadratic, etc. Think about the ways that changing the form of the temperature schedule changes the behavior and results of the simulated annealing function.
# +
# These are presented as globals so that the signature of schedule()
# matches what is shown in the AIMA textbook; you could alternatively
# define them within the schedule function, use a closure to limit
# their scope, or define an object if you would prefer not to use
# global variables
alpha = 0.95
temperature=1e4
def schedule(time):
return (alpha**time)*temperature
# -
# ### Testing the Temperature Schedule
# The following tests should validate the temperature schedule function and perform a simple test of the simulated annealing function to solve a small TSP test case
# test the schedule() function -- no output means that the tests passed
assert(np.allclose(alpha, 0.95, atol=1e-3))
assert(np.allclose(schedule(0), temperature, atol=1e-3))
assert(np.allclose(schedule(10), 5987.3694, atol=1e-3))
# Failure implies that the initial path of the test case has been changed
assert(tsp.path == [('DC', (11, 1)), ('SF', (0, 0)), ('PHX', (2, -3)), ('LA', (0, -4))])
result = simulated_annealing(tsp, schedule)
print("Initial score: {}\nStarting Path: {!s}".format(tsp.get_value(), tsp.path))
print("Final score: {}\nFinal Path: {!s}".format(result.get_value(), result.path))
assert(tsp.path != result.path)
assert(result.get_value() > tsp.get_value())
# ## V. Run Simulated Annealing on a Larger TSP
# Now we are ready to solve a TSP on a bigger problem instance by finding a shortest-path circuit through several of the US state capitals.
#
# You can increase the `num_cities` parameter up to 30 to experiment with increasingly larger domains. Try running the solver repeatedly -- how stable are the results?
# Create the problem instance and plot the initial state
num_cities = 30
capitals_tsp = TravelingSalesmanProblem(capitals_list[:num_cities])
starting_city = capitals_list[0]
print("Initial path value: {:.2f}".format(-capitals_tsp.get_value()))
print(capitals_list[:num_cities]) # The start/end point is indicated with a yellow star
show_path(capitals_tsp.coords, starting_city)
# set the decay rate and initial temperature parameters, then run simulated annealing to solve the TSP
alpha = 0.95
temperature=1e6
result = simulated_annealing(capitals_tsp, schedule)
print("Final path length: {:.2f}".format(-result.get_value()))
print(result.path)
show_path(result.coords, starting_city)
# ### Experiments (Optional)
# Here are some ideas for additional experiments with various settings and parameters once you've completed the lab.
#
# - Change the number of cities in the final map (between 10 and 30). How are your results affected? Why?
# - Change the alpha and temperature parameters. How do they affect the results?
# - Use a different schedule function (something other than exponential decay). Is the algorithm still effective?
# - Use a different successors function; e.g., generate successors of a state by swapping _any_ pair of cities in the path, rather than only adjacent cities, or reversing part of the path (e.g., reverse the BCD sequence in [ABCDE] to get [ADCBE] as the successor). Try defining your own successor function. What effect does the change have?
# - Use a different distance metric for get_value (e.g., we used the L2-norm (Euclidean distance), try the L1-norm (manhattan distance) or L$\infty$-norm (uniform norm)
#
# Share and discuss your results with others in the forums!
| Simulated_Annealing Algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # QUESTION 1
#
# Using a lambda expression, complete the mul_by_num function. This function should take an argument and return a one argument function that multiplies any value passed to it by the original number. Its body must be one line long:
#
# def mul_by_num(num):
#
# > Returns a function that takes one argument and returns num times that argument.
#
# x = mul_by_num(5)
# y = mul_by_num(2)
# x(3)
# 15
# y(-4)
# -8
# *** YOUR CODE HERE ***
# return ______
# +
def mul_by_num(num):
return lambda a : a * num
x = mul_by_num(5)
y = mul_by_num(2)
print(x(3))
print(y(-4))
# -
# # QUESTION 2
# The Fibonacci numbers are the numbers in the following integer sequence.
# >0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89, 144, ……..
#
# In mathematical terms, the sequence Fn of Fibonacci numbers is defined by the recurrence relation:
#
# > Fn = Fn-1 + Fn-2 with seed values F0 = 0 and F1 = 1.
#
# Find the series of Fibonacci numbers using lambda function.
# +
from functools import reduce
fib = lambda n: reduce(lambda x, _: x+[x[-1]+x[-2]],
range(n-2), [0, 1])
print(fib(5))
# -
fib = lambda n: n if n < 2 else fib(n-1) + fib(n-2)
fib(5)
import math
def fib(nr):
ratio = (1 + math.sqrt(5)) / 2
return int(ratio ** nr / math.sqrt(5) + 0.5)
fib(5)
fib = lambda x,y=[1,1]:([(y.append(y[-1]+y[-2]),y[-1])[1] for i in range(1+x-len(y))],y[x])[1]
fib(5)
fib = lambda n, x=0, y=1 : x if not n else fib(n-1, y, x+y)
fib(5)
fib = (lambda n, fib=[0,1]: fib[:n]+[fib.append(fib[-1] + fib[-2]) or fib[-1] for i in range(n-len(fib))])
fib(10)
fib = lambda x,y=[1,1]:[1]*x if (x<2) else ([y.append(y[q-1] + y[q-2]) for q in range(2,x)],y)[1]
fib(5)
# # QUESTION 3
# Create a script that check if a page is present on the server or return an error. Use the urllib seen during the lecture.
# +
import urllib.request
with urllib.request.urlopen ( 'http://daaru.com/' ) as res:
print("Headers:{}".format(res.info()))
# -
# # QUESTION 4
# Write a program to get the current weather of a city given in input. You can use the following API documentation:
#
# [https://openweathermap.org/current]
#
# > Hint: The GET request should have the following string appended at the end of the query for auth: APPID=<KEY>
#
# You should retrieve for the city:
#
# Temperature: 12.32°C
# Wind speed: 8.7 m/s
# Description: moderate rain
# Weather: Rain
import requests
response = requests.get("https://pro.openweathermap.org/data/2.5/climate/month?q=London&appid={b35975e18dc93725acb092f7272cc6b8}")
print(response.json())
# api.openweathermap.org/data/2.5/weather?q={city name}&appid={your api key}
# # QUESTION 5
# Write a program to read the xml file people.xml and output a csv file and json file with the same information.
#
# Validate the Json using [https://jsonlint.com/]
#
# Import the CSV using Excel
# +
import xml.etree.ElementTree as ET
tree = ET.parse('people.xml')
root = tree.getroot()
print('\nAll item data:')
for elem in root:
for subelem in elem:
print(subelem.text)
# +
import json
import xmltodict
with open("people.xml",'r') as f:
xmlString = f.read()
jsonString = json.dumps(xmltodict.parse(xmlString), indent=4)
with open('people_json.json', 'w', encoding='utf-8') as f:
json.dump(jsonString, f, ensure_ascii=False, indent=4)
# print(jsonString)
# +
from xml.etree import ElementTree
import os
import csv
tree = ElementTree.parse('people.xml')
root = tree.getroot()
people_data = open('people.csv', 'w', newline='', encoding='utf-8')
csvwriter = csv.writer(people_data)
people_data_header = []
count==0
for member in root.findall('Resident'):
people = []
address_list = []
if count==0:
name = member.find('Name').tag
people_data_header.append(name)
PhoneNumber = member.find('PhoneNumber').tag
people_data_header.append(PhoneNumber)
EmailAddress = member.find('EmailAddress').tag
people_data_header.append(EmailAddress)
Address = member[3].tag
people_data_header.append(Address)
csvwriter.writerow(people_data_header)
count += 1
name = member.find('Name').text
people.append(name)
PhoneNumber = member.find('PhoneNumber').text
people.append(PhoneNumber)
EmailAddress = member.find('EmailAddress').text
people.append(EmailAddress)
StreetLine1 = member[3][0].text
address_list.append(StreetLine1)
City = member[3][1].text
address_list.append(City)
StateCode = member[3][2].text
address_list.append(StateCode)
PostalCode = member[3][3].text
address_list.append(PostalCode)
people.append(address_list)
csvwriter.writerow(people)
people_data.close()
# -
| .ipynb_checkpoints/Lab3-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# <img src='./img/LogoWekeo_Copernicus_RGB_0.png' alt='Logo EU Copernicus EUMETSAT' align='right' width='20%'></img>
# + [markdown] Collapsed="false"
# <a href="./00_index.ipynb"><< Index</a><br>
# <a href="./20_Sentinel5P_TROPOMI_NO2_L2_retrieve.ipynb"><< 20 - Sentinel-5P NO<sub>2</sub> - Nitrogen Dioxide - Retrieve</a>
# -
# <div class="alert alert-block alert-warning">
# <b>LOAD, BROWSE AND VISUALIZE</b></div>
# + [markdown] Collapsed="false"
# # Copernicus Sentinel-5 Precursor (Sentinel-5P) - NO<sub>2</sub>
# + [markdown] Collapsed="false"
# The subsequent example introduces you to Sentinel-5P data in general and the total column of NO<sub>2</sub> sensed by Sentinel-5P in specific. NO<sub>2</sub> is useful for monitoring air pollution. The example is based on elevated nitrogen dioxide levels in Europe which occurred in February 2021.
#
# + [markdown] Collapsed="false"
# #### Module outline:
# * [1 - Load and browse Sentinel-5P TROPOMI data](#load_s5P)
# * [2 - Create a geographical subset](#geographical_subset)
# * [3 - Visualize Sentinel-5P NO<sub>2</sub> data](#visualize_s5P)
# + [markdown] Collapsed="false"
# #### Load required libraries
# + Collapsed="false"
# %matplotlib inline
import os
import xarray as xr
import numpy as np
import netCDF4 as nc
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
import cartopy.crs as ccrs
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import cartopy.feature as cfeature
from matplotlib.axes import Axes
from cartopy.mpl.geoaxes import GeoAxes
GeoAxes._pcolormesh_patched = Axes.pcolormesh
# -
# #### Load helper functions
# %run ./functions.ipynb
# + [markdown] Collapsed="false"
# <hr>
# + [markdown] Collapsed="false"
# ## <a id="load_s5P"></a>Load and browse Sentinel-5P data
# -
# A Sentinel-5P file is organised in two groups: `PRODUCT` and `METADATA`. The `PRODUCT` group stores the main data fields of the product, including `latitude`, `longitude` and the variable itself. The `METADATA` group provides additional metadata items.
#
# Sentinel-5P variables have the following dimensions:
# * `scanline`: the number of measurements in the granule / along-track dimension index
# * `ground_pixel`: the number of spectra in a measurement / across-track dimension index
# * `time`: time reference for the data
# * `corner`: pixel corner index
# * `layer`: this dimension indicates the vertical grid of profile variables
#
# Sentinel-5P TROPOMI data is disseminated in `netCDF`. You can load a `netCDF` file with the `open_dataset()` function of the xarray library. In order to load the variable as part of a Sentinel-5P data files, you have to specify the following keyword arguments:
# - `group='PRODUCT'`: to load the `PRODUCT` group
#
# Let us load a Sentinel-5P TROPOMI data file as `xarray.Dataset` from 5 February 2021 and inspect the data structure:
# + Collapsed="false"
s5P = xr.open_dataset('./data/S5P_OFFL_L2__NO2____20210205T104439_20210205T122609_17182_01_010400_20210207T042548.nc', group='PRODUCT')
s5P
# + [markdown] Collapsed="false"
# You see that a Sentinel-5P data file consists of eight dimensions and twelve data variables:
#
# * **Dimensions**:
# * `scanline`
# * `ground_pixel`
# * `time`
# * `corner`
# * `polynomial_exponents`
# * `intensity_offset_polynomial_exponents`
# * `layer`
# * `vertices`
# -
# * **Data variables**:
# * `delta_time`: the offset of individual measurements within the granule, given in milliseconds
# * `time_utc`: valid time stamp of the data
# * `qa_value`: quality descriptor, varying between 0 (nodata) and 1 (full quality data).
# * `nitrogendioxide_tropospheric_column`: Vertically integrated NO<sub>2</sub> column density
# * `nitrogendioxide_tropospheric_column_precision`: Standard error of the vertically integrated NO<sub>2</sub> column
#
# As well as a few other variables:
# * `nitrogendioxide_tropospheric_column_precision_kernel`
# * `averaging_kernel`
# * `air_mass_factor_troposphere`
# * `air_mass_factor_total`
# * `tm5_tropopause_layer_index`
# * `tm5_constant_a`
# * `tm5_constant_b`
# + [markdown] Collapsed="false"
# You can specify one variable of interest and get more detailed information about the variable. E.g. `nitrogendioxide_total_column` is the atmosphere mole content of NO<sub>2</sub>, has the unit `mol m-2` (which means `mol per m2`), and has three dimensions, `time`, `scanline` and `groundpixel` respectively.
# -
s5P_no2 = s5P['nitrogendioxide_tropospheric_column']
s5P_no2
# You can do this for the available variables, but also for the dimensions latitude and longitude.
latitude = s5P_no2.latitude
latitude
# + [markdown] Collapsed="false"
# <br>
# -
longitude = s5P_no2.longitude
longitude
# + [markdown] Collapsed="false"
# You can retrieve the array values of the variable with squared bracket: `[:,:,:]`. One single time step can be selected by specifying one value of the time dimension, e.g. `[0,:,:]`.
# -
s5P_no2_0502 = s5P_no2[0,:,:]
s5P_no2_0502
# The attributes of the data array hold the entry `multiplication_factor_to_convert_to_molecules_percm2`, which is a conversion factor that has to be applied to convert the data from `mol per m2` to `molecules per cm2`.
#
conversion_factor = s5P_no2_0502.multiplication_factor_to_convert_to_molecules_percm2
conversion_factor
# Additionally, you can save the attribute `longname`, which you can make use of when visualizing the data.
# + Collapsed="false"
longname = s5P_no2_0502.long_name
longname
# + [markdown] Collapsed="false"
# ## <a id='geographical_subset'></a>Create a geographical subset
# + [markdown] Collapsed="false"
# You can zoom into a region by specifying a `bounding box` of interest. Let's set the extent to Europe with the following bounding box information:
# -
latmin = 28.
latmax = 71.
lonmin = -22.
lonmax = 43
# You can use the function [generate_geographical_subset()](./functions.ipynb#generate_geographical_subset) to subset an xarray DataArray based on a given bounding box.
s5P_no2_subset = generate_geographical_subset(s5P_no2_0502, latmin, latmax, lonmin, lonmax)
s5P_no2_subset
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# ## <a id="plotting_s5P"></a>Plotting example - Sentinel-5P TROPOMI data
# + [markdown] Collapsed="false"
# You can plot data arrays of type `numpy` with matplotlib's `pcolormesh` function. In combination with the library [cartopy](https://scitools.org.uk/cartopy/docs/latest/), you can produce high-quality maps.
# + [markdown] Collapsed="false"
# In order to make it easier to visualize the NO<sub>2</sub> values, we apply the conversion factor to the DataArray. This converts the NO<sub>2</sub> values from mol per m<sup>2</sup> to molecules per cm<sup>2</sup>.
# -
s5P_no2_converted = s5P_no2_subset*conversion_factor
s5P_no2_converted
# The next step is to visualize the dataset. You can use the function [visualize_pcolormesh](../functions.ipynb#visualize_pcolormesh), which makes use of matploblib's function `pcolormesh` and the [Cartopy](https://scitools.org.uk/cartopy/docs/latest/) library.
#
# With `?visualize_pcolormesh` you can open the function's docstring to see what keyword arguments are needed to prepare your plot.
# ?visualize_pcolormesh
# Now, let us apply the [visualize_pcolormesh](./functions#visualize_pcolormesh) function and visualize the vertically integrated NO<sub>2</sub> column sensored from the Sentinel-5P satellite on 5 February 2021.
#
# Note: Multiplying the `DataArray` values with `1e-15` improves the readibility of the map legend.
visualize_pcolormesh(data_array=s5P_no2_converted*1e-15,
longitude=s5P_no2_converted.longitude,
latitude=s5P_no2_converted.latitude,
projection=ccrs.PlateCarree(),
color_scale='viridis',
unit='*1e-15 molecules per cm2',
long_name=longname + ' ' + str(s5P_no2_converted.time.data),
vmin=0,
vmax=35,
lonmin=lonmin,
lonmax=lonmax,
latmin=latmin,
latmax=latmax,
set_global=False)
# + [markdown] Collapsed="false"
# <br>
# + [markdown] Collapsed="false"
# <a href="./00_index.ipynb"><< Index</a><br>
# <a href="./20_Sentinel5P_TROPOMI_NO2_L2_retrieve.ipynb"> << 20 - Sentinel-5P NO<sub>2</sub> - Nitrogen Dioxide - Retrieve</a>
# + [markdown] Collapsed="false"
# <hr>
# + [markdown] Collapsed="false"
# <p><img src='./img/all_partners_wekeo.png' align='left' alt='Logo EU Copernicus' width='100%'></img><p>
| atmosphere/20210506_wekeo_webinar/21_Sentinel5P_TROPOMI_NO2_L2_load_browse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exam 2
# ## <NAME>
# ### Problem 1: Controls
# I have created a function that can check if the number of lines of both files are the same and, in that case, if the order of the entries regarding the AGI Codes is the same.
def lines_control(file_1_lines, file_2_lines):
"""" This function compares the lines
number and order of the given files."""
same = False
if len(file_1_lines) == len (file_2_lines):
print ("Both files have the same number of lines.")
same = True
else:
print("The given files have not the same number of lines.")
if same == True:
dif = 0
for line in range(1, len(file_1_lines)):
line_G = file_1_lines[line].split("\t")[0]
line_L = file_2_lines[line].split("\t")[0]
if line_G != line_L:
dif = 1
break
if dif == 0:
print("All lines of both files are in the same sequence (order)!")
else:
print("The lines of both files are not in the same sequence (order).")
# +
germplasm_file = open("Germplasm.tsv", "r")
locusgene_file = open("LocusGene.tsv", "r")
germplasm_lines = germplasm_file.readlines()
locusgene_lines = locusgene_file.readlines()
lines_control(germplasm_lines, locusgene_lines)
# -
# ### Problem 2: Design and create the database
# %load_ext sql
# %sql mysql+pymysql://root:root@127.0.0.1:3306/mysql
# #%sql drop database exam2_DB
# %sql create database exam2_DB;
# +
import pymysql.cursors
#Connection is established
connection = pymysql.connect(host='localhost',
user='root',
password='<PASSWORD>',
db='exam2_DB',
charset='utf8mb4',
cursorclass=pymysql.cursors.DictCursor)
# +
def create_function(query_list):
"""This function creates the tables following the
queries in the given list"""
for query in query_list:
try:
with connection.cursor() as cursor:
sql = query
cursor.execute(sql)
except:
print("Oops! An error ocurred. Try again.")
query_1 = "create table Germplasm(AGI_code VARCHAR(20) NOT NULL PRIMARY KEY, Germplasm VARCHAR(50) NOT NULL, Phenotype VARCHAR(1000) NOT NULL, PUBMED INT NOT NULL)"
query_2 = "create table LocusGene(AGI_code VARCHAR(20) NOT NULL PRIMARY KEY REFERENCES Germplasm(AGI_code), Gene VARCHAR(20) NOT NULL, Protein_Length INT NOT NULL)"
# Due to the 1:1 relationship between tables, AGI_code of LocusGene table references to the Germplasm one
query_list = [query_1, query_2]
create_function(query_list)
# -
# ### Problem 3: Fill the database
# +
def insert_function(file_1_lines, file_2_lines, query1, query2):
"""This function fills the tables folowing the specified queries.
It obtains the information from the given files."""
for line in range(1, len(file_1_lines)):
AGI_code, germplasm, phenotype, pubmed = file_1_lines[line].split("\t")
AGI_code, gene, ProtLen = file_2_lines[line].split("\t")
#Note that AGI_code variable contains exactly the same information in both cases
try:
with connection.cursor() as cursor:
sql = query1.format(AGI_code, germplasm, phenotype, pubmed)
cursor.execute(sql)
sql = query2.format(AGI_code, gene, ProtLen)
cursor.execute(sql)
except:
print("Oops! An error ocurred. Try again.")
query1 = """INSERT INTO Germplasm(AGI_code, Germplasm, Phenotype, PUBMED)
VALUES ("{}","{}","{}", {})"""
query2 = """INSERT INTO LocusGene(AGI_code, Gene, Protein_Length) VALUES ("{}","{}", {})"""
insert_function(germplasm_lines, locusgene_lines, query1, query2)
# -
germplasm_file.close()
locusgene_file.close()
# ### Problem 4: Create reports, written to a file
def reports_function(reports_file, write_append, report_number, query):
""" This function executes the specified query in order to get some information from the data base,
writting it to a indicated file."""
reports_file = open(reports_file, write_append)
reports_file.write("Report {}\n\n".format(report_number))
try:
with connection.cursor() as cursor:
sql = query
cursor.execute(sql)
results = cursor.fetchall()
header = results[0].keys()
reports_file.write('\t'.join([str(x) for x in header]))
reports_file.write("\n")
for row in results:
reports_file.write('\t'.join([str(x) for x in row.values()]))
reports_file.write("\n")
reports_file.write("\n\n")
except:
print("Oops! An error ocurred. Try again.")
reports_file.close()
# +
query1 = """SELECT * FROM Germplasm, LocusGene
WHERE Germplasm.AGI_code = LocusGene.AGI_code;"""
reports_function("Reports_file.tsv", "w", 1, query1)
# +
query2 = """SELECT * FROM Germplasm, LocusGene
WHERE (LocusGene.gene = "SKOR" OR LocusGene.gene = "MAA3")
AND Germplasm.AGI_code = LocusGene.AGI_code;"""
reports_function("Reports_file.tsv", "a", 2, query2)
# +
query3 = """SELECT SUBSTRING(Germplasm.AGI_code, 3,1) AS 'Chromosomes', count(*) AS 'Number of entries' FROM Germplasm
GROUP BY SUBSTRING(Germplasm.AGI_code, 3,1);"""
reports_function("Reports_file.tsv", "a", 3, query3)
# +
query4 = """SELECT SUBSTRING(LocusGene.AGI_code, 3,1) AS 'Chromosomes', AVG(Protein_Length) AS 'Protein_length_mean' FROM LocusGene
GROUP BY SUBSTRING(LocusGene.AGI_code, 3,1);"""
reports_function("Reports_file.tsv", "a", 4, query4)
# -
connection.close()
| Exam_2_answers_Andrea_Escolar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="Python 3 (ipykernel)"
# ## 1D Convolution
# + kernel="Python 3 (ipykernel)" tags=[]
import numpy as np
def MatrixConvolution1D():
a=np.array([1,3,5,7,6])
b=np.array([1,2,3])
m = len(a)
n = len(b)
convolution= np.zeros(m+n-1)
for i in range(1,(m+n-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for j in range(max(1,i+1-n),min(i,m)+1): #S={max(1,i+1-n),....,min(i,n)}
convolution[i-1]=convolution[i-1]+a[j-1]*b[(i-1)-j+1]
print("Convolution 1D using step by step")
print(convolution)
conv = np.convolve(a,b)
print("Convolution 1D using numpy method")
print(conv)
MatrixConvolution1D()
# + [markdown] kernel="Python 3 (ipykernel)"
# ## 2D Convolution
# + kernel="Python 3 (ipykernel)" tags=[]
import numpy as np
import scipy as scp
from scipy import signal
def MatrixConvolution2D():
a=[[1,0,1],[4,3,1],[-1,0,2],[3,0,-7]]
b=[[1,-1,2,3],[-4,0,1,5],[3,2,-1,0]]
m1,n1=np.shape(a)
m2,n2=np.shape(b)
convolution= np.zeros((m1+m2-1,n1+n2-1))
for j in range(1,(m1+m2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,(n1+n2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
print("Convolution 2D using step by step")
print(convolution)
conv= signal.convolve2d(a,b,mode='full',boundary='fill',fillvalue=0)
print("Convolution 2D using scipy method")
print(conv)
MatrixConvolution2D()
# + [markdown] kernel="Python 3 (ipykernel)"
# ## Gaussian Filter
# + kernel="Python 3 (ipykernel)" tags=[]
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
def Convolution(a,b,type_convolution):
m1,n1=np.shape(a)
m2,n2=np.shape(b)
convolution= np.zeros((m1+m2-1,n1+n2-1))
if type_convolution=="full":
for j in range(1,(m1+m2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,(n1+n2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
else:
for j in range(1,m1+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,n1+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
return convolution
def GaussianFilter():
image=plt.imread('child2.jpg')
image=image.astype(float)
image=np.asarray(image)
image=np.clip(image, 0, 255)
filterG=(1/16)*np.array([[1, 2, 1],[2, 4, 2],[1, 2, 1]])
fig = plt.figure()
image=image.astype(np.uint8)#image at the moment is
#an array of type flot and here it becomes type uint8
#convolution= signal.convolve2d(image,filterG,mode='full',boundary='fill',fillvalue=0)
convolution=Convolution(image,filterG,"same")
convolution=np.clip(convolution, 0, 255)
convolution=convolution.astype(np.uint8)#image at the moment is
#an array of type flot and here it becomes type uint8
convolution=np.asarray(convolution)
ax1 = fig.add_subplot(1,2,1)
ax2 = fig.add_subplot(1,2,2)
ax1.set_title("Original Image")
ax2.set_title("Image with Gaussian Filter")
ax1.imshow(image,cmap='gray')
ax2.imshow(convolution,cmap='gray',vmin=0,vmax=255)
GaussianFilter()
# + [markdown] kernel="Python 3 (ipykernel)"
# ## Sobel Filter
# + kernel="Python 3 (ipykernel)"
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
def Convolution(a,b,type_convolution):
m1,n1=np.shape(a)
m2,n2=np.shape(b)
convolution= np.zeros((m1+m2-1,n1+n2-1))
if type_convolution=="full":
for j in range(1,(m1+m2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,(n1+n2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
else:
for j in range(1,m1+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,n1+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
return convolution
def SobelFilter():
image=plt.imread('baby_yoda.jpg')
image=image.astype(float)
image=np.asarray(image)
image=np.clip(image, 0, 255)
#filterS=np.array([[-1, -2, -1],[0, 0, 0],[1, 2, 1]])
filterBx=np.array([[-1, -2, -1],[0, 0, 0],[1, 2, 1]])
filterBy=np.array([[-1, 0, 1],[-2, 0, 2],[-1, 0, 1]])
fig = plt.figure()
image=image.astype(np.uint8)#image at the moment is
#an array of type flot and here it becomes type uint8
#Cx= signal.convolve2d(image,filterBx,mode='full',boundary='fill',fillvalue=0)
#Cy= signal.convolve2d(image,filterBy,mode='full',boundary='fill',fillvalue=0)
Cx=Convolution(image,filterBx,"same")
Cy=Convolution(image,filterBy,"same")
Cx=np.clip(Cx, 0, 255)
#an array of type flot and here it becomes type uint8
Cy=np.clip(Cy, 0, 255)
#an array of type flot and here it becomes type uint8
C=np.sqrt(Cx**2+Cy**2)
C=np.asarray(C)
C=C.astype(np.uint8)
#an array of type flot and here it becomes type uint8
ax1 = fig.add_subplot(1,2,1)
ax2 = fig.add_subplot(1,2,2)
ax1.set_title("Original Image")
ax2.set_title("Image with Sobel Filter")
ax1.imshow(image,cmap='gray')
ax2.imshow(C,cmap='gray',vmin=0,vmax=255)
SobelFilter()
# + [markdown] kernel="Python 3 (ipykernel)"
# ## Emphasize edges
# + kernel="Python 3 (ipykernel)"
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
def Convolution(a,b,type_convolution):
m1,n1=np.shape(a)
m2,n2=np.shape(b)
if type_convolution=="full":
convolution= np.zeros((m1+m2-1,n1+n2-1))
for j in range(1,(m1+m2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,(n1+n2-1)+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
else:
convolution= np.zeros((m1,n1))
for j in range(1,m1+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for k in range(1,n1+1):#Starts in 1 because that is the way the method works, so, is necesary to adjust
#the indexes
for p in range(max(1,j-m2+1),min(j,m1)+1):#S={max(1,j-m2+1),....,min(j,m1)
for q in range(max(1,k-n2+1),min(k,n1)+1):#S={max(1,k-n2+1),....,min(k,n1)
convolution[j-1][k-1]=convolution[j-1][k-1]+(a[p-1][q-1])*(b[(j-1)-(p)+1][(k-1)-(q)+1])
return convolution
def EmphasizeEdges():
image=plt.imread('baby_yoda.jpg')
image=image.astype(float)
image=np.asarray(image)
image=np.clip(image, 0, 255)
filterEE=np.array([[1, 1, 1],[1, -8, 1],[1, 1, 1]])
fig = plt.figure()
image=image.astype(np.uint8)#image at the moment is
#an array of type flot and here it becomes type uint8
#convolution= signal.convolve2d(image,filterG,mode='full',boundary='fill',fillvalue=0)
convolution=Convolution(image,filterEE,"same")
c=1
D=image+c*convolution;
D=np.asarray(D)
D=D.astype(np.uint8)#image at the moment is
#an array of type flot and here it becomes type uint8
ax1 = fig.add_subplot(1,2,1)
ax2 = fig.add_subplot(1,2,2)
ax1.set_title("Original Image")
ax2.set_title("Emphasize Edges")
ax1.imshow(image,cmap='gray')
ax2.imshow(D,cmap='gray')
EmphasizeEdges()
# + [markdown] kernel="Python 3 (ipykernel)"
# ## DFT-2D
# + kernel="Python 3 (ipykernel)"
import numpy as np
import matplotlib.pyplot as plt
def im2double(image):
number_decimals=7
info=np.iinfo(image.dtype)
out=image.astype(float) / info.max
out=out.round(number_decimals)
return out
def DFT2D():
image=plt.imread('chest.jpg')
image_copy=image
image=np.asarray(image)
image=im2double(image)
DFT2D=np.fft.fft2(image)
DFT2D_Shift=np.fft.fftshift(DFT2D)
result_DFT2D=np.log(1+np.abs(DFT2D))
result_DFT2DShift=np.log(1+np.abs(DFT2D_Shift))
fig = plt.figure()
ax1 = fig.add_subplot(1,5,1)
ax2 = fig.add_subplot(1,5,3)
ax3 = fig.add_subplot(1,5,5)
ax1.set_title("Original Image")
ax2.set_title("DFT-2D")
ax3.set_title("DFT-2D Shift")
ax1.imshow(image_copy,cmap='gray')
ax2.imshow(result_DFT2D,cmap='gray',vmin=0,vmax=10)
ax3.imshow(result_DFT2DShift,cmap='gray',vmin=0,vmax=10)
DFT2D()
# + [markdown] kernel="Python 3 (ipykernel)"
# ## Gaussian Filter DFT-2D
# + kernel="Python 3 (ipykernel)"
import numpy as np
import matplotlib.pyplot as plt
def im2double(image):
number_decimals=7
info=np.iinfo(image.dtype)
out=image.astype(float) / info.max
out=out.round(number_decimals)
return out
def GaussianFilterDFT2D():
image=plt.imread('edificio_china.jpg')
image_copy=image
m,n=np.shape(image)
image = im2double(image)
#Calculation of DFT-2D
F = np.fft.fft2(image)
F_shift = np.fft.fftshift(F)
#Calculate the Gaussian Filter
D = np.zeros((m,n))
for i in range(1,m+1):
for j in range(1,n+1):
D[i-1,j-1] = np.sqrt(i**2+j**2)
H = np.zeros((m,n))
sigma = 50
for i in range(1,m+1):
for j in range(1,n+1):
H[i-1,j-1] = np.exp(-(D[i-1][j-1])**2 / (2 * sigma**2))
index1=int(np.floor(m/2))
index2=int(np.floor(n/2))
HSI = H[:index1, :index2]
HSD = np.transpose(np.rot90(HSI,1))
HID = np.rot90(HSI,2)
HII = np.transpose(np.rot90(HSI,3))
m1, n1 = np.shape(HSI)
H[0:m1, n-n1:n] = HSD
H[m-m1:m+1, n-n1:n] = HID
H[m-m1:m+1, 0:n1] = HII
H = np.asarray(H)
F = np.asarray(F)
#Apply the filter
H_shift = np.fft.fftshift(H)
DFT2_filt = F*H
FM_shift = np.fft.fftshift(DFT2_filt)
#Filtered Image
I_new = np.abs(np.fft.ifft2(DFT2_filt));
result_F=np.log(1+np.abs(F_shift))
result_FM=np.log(1+np.abs(FM_shift))
fig = plt.figure()
ax1 = fig.add_subplot(3,3,1)
ax2 = fig.add_subplot(3,3,3)
ax3 = fig.add_subplot(3,3,7)
ax4 = fig.add_subplot(3,3,9)
ax1.set_title("Original Image")
ax2.set_title("Image DFT-2D (Shift)")
ax3.set_title("Image DFT-2D with Gaussian Filter")
ax4.set_title("Image with Gaussian Filter")
ax1.imshow(image_copy,cmap='gray')
ax2.imshow(result_F,cmap='gray',vmin=0,vmax=10)
ax3.imshow(result_FM,cmap='gray',vmin=0,vmax=10)
ax4.imshow(I_new,cmap='gray')
GaussianFilterDFT2D()
# + kernel="Octave"
| Catalogo2_Hernandez_Brenes_Porras/Convolution, spatial domain filters and low pass frequency domain filters/MetodosPythonCatalogo2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="logo_img/logo.png" height=50 width=50 align="right"/><br/><br/>
#
#
#
# # IST 5520: Data Science and ML in Python
# ## Project Report: Group 4
#
# <h3>Group Members: <font size=3><i><NAME>, <NAME>, <NAME>, <NAME>, <NAME> </i></font></h3>
#
#
#
# # 1. Introduction
#
# ## 1.1 Background
# Greenhouse gases (GHG) are responsible for trapping heat, thereby making the planet warmer. For the past 150 years, the sharp global increase in GHG in the atmosphere can be primarily attributed to anthropogenic or human-induced activities. In the United States, the largest source of GHG emissions from human activities includes fossil fuel burning, which acts as the predominant energy source in the transportation sector and is also used to generate electricity and heat. The transportation sector generates the largest share of GHG emissions (29% of 2019 GHG emissions) from burning fossil fuels (mostly petroleum by-products such as gasoline and diesel) for our cars, trucks, ships, trains, and planes.
#
#
# The Environmental Protection Agency (EPA) emissions score (smog rating) reflects vehicle tailpipe emissions (CO$_2$) that contribute to local and regional air pollution, creating problems such as smog, haze, and health issues. CO$_2$ emissions typically constitute 99% of the tailpipe emissions of greenhouse gases. Essentially, the higher the rating, the cleaner the car is, and this rating is a direct proxy for the CO$_2$ emissions.
#
# ## 1.2 Problem Description and Research Questions
# This project aims to build a machine learning model to predict the EPA emission score from different vehicles operating in the United States. Essentially, this is a classification problem wherein we will utilize the EPA vehicle and emission data sets to predict the EPA emission score (a discrete quantity). We will also be analyzing the factors having the most impact on this score which varies from 1 (worst) to 10 (best). In addition, we will compare different machine learning models to assess their performance on the data sets. Such kind of modeling and analysis, if pushed to a production environment, could enable policymakers to address critical issues in the sustainable transportation industry. In this regard, we have to answer the following research question:
#
# <i> Which predictors are most suited for predicting the EPA emission score? </i>
#
#
# Some specific research questions include:
#
# <i>a. Which machine learning model works best for this data set?</i>
#
# <i>b. Which vehicle class has poor emission ratings? </i>
#
#
# ## 1.3 Classification Analysis
# The vehicle and car emissions data sets can model a classification problem where we need to predict the EPA emission score (a unitless quantity having integral values in the [1,10] interval) based on the explanatory variables. In this project, we deal with a high-dimensional classification problem concerned with prediction (discrete EPA emission score) and inference.
#
# ## 1.4 Potential Problems and Challenges
# The major challenge in this project is to perform feature engineering. Appropriate data-preprocessing and feature engineering must be performed to reduce the data dimensionality required to address the above research questions suitably.
#
# ## 1.5 Tentative Timeline
#
# |Phase|Activities|Completion|
# |----|----------|--------|
# |Data Collection| Collecting the required data set for the business analytics project|September 24, 2021|
# |Kickoff|Understanding the project requirements and elicitation|October 1, 2021|
# |Data Management|Data cleaning, pre-processing|October 29, 2021|
# |Full Data Analysis|Evaluating different ML models|November 20, 2021|
# |Project Submission|Report writing, preparing presentation, and proof-reading|December 5, 2021|
#
# # 2. Data Source and Collection
#
# ## 2.1 Data Source
#
# This dataset can be found at https://www.fueleconomy.gov/feg/ws/index.shtml. The vehicle and emission data sets need to be linked based on the vehicle ID. The vehicle data set contains 83 explanatory variables (columns) that provide detailed car specifications and has 44075 rows. As for the emissions data set, it has 8 features and 42442 rows. Some of the important features are listed below:
#
# |Data Set|Feature Name|Details|
# |----|----------|--------|
# |emissions|score|EPA 1-10 smog rating for fuelType1 (target variable)|
# |vehicle|fuelType1|For single fuel vehicles, this will be the only fuel. For dual fuel vehicles, this will be the conventional fuel.|
# |vehicle|highway08|Highway MPG for fuelType1|
# |vehicle|barrels08|Annual petroleum consumption in barrels for fuelType1|
# |vehicle|year|Model year|
# |vehicle|VClass|EPA vehicle size class|
# |vehicle|phevBlended|If True, this vehicle operates on a blend of gasoline and electricity in charge depleting mode|
#
#
#
# ## 2.2 Collection
#
# We found this data set on [EPA fuel economy portal](https://www.fueleconomy.gov/). The EPA has generated [annual reports](https://www.fueleconomy.gov/feg/pdfs/guides/FEG2021.pdf), but there are no existing publicly available notebooks having detailed machine learning workflows. Moreover, in conjunction, these two data sets satisfy all the other project requirements ($\ge$20 columns and $\ge$1000 rows) listed as part of this group assignment.
#
# # 3. Data Management
# import required libraries for data management
import pandas as pd
import numpy as np
import seaborn as sns
import plotly.express as px
import matplotlib.pyplot as plt
from sklearn.preprocessing import RobustScaler
from sklearn.decomposition import PCA
# ## 3.1 Data Cleaning
# Read data sets
vehicle_df = pd.read_csv('data/vehicles.csv', low_memory=False)
emissions_df = pd.read_csv('data/emissions.csv')
# Get vehicle data info and description
vehicle_df.info()
vehicle_df.describe().T
# We observe that there are several missing values (NaNs) in the vehicle data. However, there are even more NaN values because some features have -1 or 0 as values which represent no data. So, we have to replace these -1s and 0s with NaN.
# Replace -1 with NaNs
vehicle_df = vehicle_df.replace([-1, 0], np.nan)
vehicle_df.info()
vehicle_df.describe().T
# Check NaN values for vehicle
percent_nan = vehicle_df.isnull().sum() * 100 / vehicle_df.shape[0]
percent_nan
# Since want to include details about all types of cars, we can use the above colums (some of which we identified to be of most use). We can simply drop some columns such as modifiedOn and createdOn as these will not be useful predictors.
# Columns with no missing values
non_nan_cols = percent_nan[percent_nan == 0]
non_nan_cols
selected_columns = non_nan_cols.index[:-2]
print(selected_columns)
vehicle_df = vehicle_df[selected_columns]
vehicle_df.head()
# Get emissions data info and description
emissions_df.info()
emissions_df.describe().T
# Here, we only need the id and score columns. So, we can drop the rest.
emissions_df = emissions_df[['id', 'score']]
emissions_df.describe().T
# Although, we have no missing values, the score column has -12.0 as the minimum value implying there might be incorrect values lying outisde the [1, 10] interval.
incorrect_scores = emissions_df.score[emissions_df.score < 1].size
print('There are {} incorrect emission scores'.format(incorrect_scores))
# Remove incorrect scores
emissions_df = emissions_df[~(emissions_df.score < 1)]
emissions_df.describe().T
# Now, we have to join the two data frames based on id.
# We perform inner join because we only need matching car ids.
vehicle_emissions_df = vehicle_df.join(emissions_df.set_index('id'), on='id', how='inner').reset_index(drop=True)
vehicle_emissions_df.head()
# Remove ID column and drop duplicate rows
vehicle_emissions_df = vehicle_emissions_df.drop(columns=['id'])
vehicle_emissions_df = vehicle_emissions_df.drop_duplicates().reset_index(drop=True)
vehicle_emissions_df.head()
vehicle_emissions_df.describe()
vehicle_emissions_df.to_csv('data/Vehicle_Emissions.csv', index=False)
# Now, we have a cleaned data set having 28556 non-redundant observations that combines both vehicles and emissions data sets.
# ## 3.2 Data Transformation
# Read the cleaned data to avoid recomputation
ve_df = pd.read_csv('data/Vehicle_Emissions.csv')
ve_df.head()
# First we check VClass categorical variable. This is an imporant variable as vehicle size is directly responsible for mileage and hence, emission score. We have to aggregate this column in to a higher level because there are two many categories.
ve_df.VClass.value_counts()
# +
# vclass_1 = [
# 'Compact Cars',
# 'Subcompact Cars',
# 'Two Seaters',
# 'Minicompact Cars'
# ]
# vclass_2 = [
# 'Midsize Cars',
# 'Utility Vehicle',
# ]
# for vclass in ve_df.VClass.unique():
# selection = ve_df['VClass'] == vclass
# if vclass in vclass_1:
# ve_df.loc[selection, 'VClass'] = 1
# elif any(v in vclass for v in vclass_2):
# ve_df.loc[selection, 'VClass'] = 2
# else:
# ve_df.loc[selection, 'VClass'] = 3
# ve_df.VClass.value_counts()
# +
vclass_1 = [
'Standard Pickup Trucks 2WD',
'Standard Pickup Trucks 4WD'
]
vclass_2 = [
'Vans, Cargo Type',
'Vans, Passenger Type'
]
for vclass in ve_df.VClass.unique():
selection = ve_df['VClass'] == vclass
if vclass in vclass_1:
ve_df.loc[selection, 'VClass'] = 'Standard Pickup Trucks'
elif vclass in vclass_2:
ve_df.loc[selection, 'VClass'] = 'Vans'
else:
ve_df.loc[selection, 'VClass'] = 'Other Vehicles'
ve_df.VClass.value_counts()
# -
# So, we have made 3 categories out of the initial 24! Now, we check the fuelType1 column
ve_df.fuelType1.value_counts()
# We can combine midgrade and regular gasolines as 'Regular Gasoline'. We can also combine 'Electricity' and 'Natural Gas' as 'Green Fuel'.
for ft in ve_df.fuelType1.unique():
selection = ve_df['fuelType1'] == ft
if ft == 'Midgrade Gasoline':
ve_df.loc[selection, 'fuelType1'] = 'Regular Gasoline'
elif ft in ['Electricity', 'Natural Gas']:
ve_df.loc[selection, 'fuelType1'] = 'Green Fuel'
ve_df.fuelType1.value_counts()
ve_df.phevBlended.value_counts()
# +
# We drop columns that won't be useful or are redundant
ve_df = ve_df.drop(columns=['fuelType', 'make', 'model'])
# Set int types to year and score
ve_df.year = ve_df.year.astype(int)
ve_df.score = ve_df.score.astype(int)
ve_df.to_csv('data/VE_Transformed.csv', index=False)
ve_df.head()
# -
# Note that, so far, we have handled missing values and categorical variables. We also handled incorrect values. Next, as a part of data visualization, we will handle outliers.
# ## 3.3 Dimensionality Reduction
ve_df = pd.read_csv('data/VE_Transformed.csv')
ve_df.VClass = ve_df.VClass.astype(str)
ve_df.phevBlended = ve_df.phevBlended.astype(bool)
ve_df.head()
# ### 3.3.1 Data Tabularization
# +
# Relative frequency table
pd.crosstab(index=ve_df.score, columns="Percent") * 100 / pd.crosstab(index=ve_df.score, columns="Percent").sum()
# -
# From the relative frequency table, we see that emission scores 5 and 6 are the most frequent.
# +
# Contingency Table
pd.crosstab(ve_df.fuelType1, ve_df.score, margins=False)
# -
# The contingency table shows that Green Fuel vehicles mostly receive a score of 10 (cleanest). Premium Gasoline and Regular Gasoline vehicles have a wide range of scores. There are no diesel vehicles having an emission score of more than 6.
# ### 3.3.2 Correlation Analysis
# Correlation table
ve_df.corr(method='pearson')
# +
# Correlation heatmap
# %matplotlib inline
ve_corr = ve_df.corr()
mask = np.zeros_like(ve_corr)
mask[np.triu_indices_from(mask)] = True
sns.heatmap(ve_corr, cmap='Spectral', vmax=1.0, vmin=-1.0 , mask = mask, linewidths=2)
plt.yticks(rotation=0)
plt.xticks(rotation=45)
plt.savefig('correlation_heatmap.png', dpi=600, bbox_inches='tight')
plt.show()
# -
# From the correlation analysis, we see that barrels08 (annual petroleum consumption in barrels) shows the strongest correlation with emisisons score, whereas, phevBlended (hybrid car or not) shows the least correlation.Intriguingly, we observe that most of our selected variables exhibit a relatively high correlation (either positive or negative) with the emissions score.
# ### 3.3.3 Distribution and Cross-Comparison Analysis
# Kernel density estimate
ax = sns.kdeplot(ve_df.score)
# We see that score exhibits a multimodal distribution.
# violin plot
ax = sns.violinplot(x=ve_df.score)
# The violinplot shows that the majority of the scores are either 5 or 6.
sns.boxplot(x='score', y='fuelType1', data=ve_df)
# From the above boxplot, we see that Diesel cars have lower scores without any outliers. For Gasoline cars, the median score is 5.
sns.boxplot(x='score', y='year', data=ve_df)
# From this boxplot, we notice that as we move ahead with time, the emission scores increase. This is a realistic observation as newer car makes have better fuel efficiency and emissions standards. Therefore, we should treat year as a numeric variable and not categorical.
plt.rcParams["figure.figsize"] = (10,3)
sns.boxplot(x='VClass', y='score', data=ve_df)
# We see that there is very little score variation between the vehicle classes. Thus, this variable can also be dropped.
ax = sns.barplot(x="score", y="barrels08", ci='sd', data=ve_df, capsize=.2)
# The score vs barrels08 barplot shows that the higher the score is, the annual fuel consumption (in barrels) is lower. Noticeably, green vehicles, have the least consumption.
#
# The correlation and distribution analysis suggests removing phevBlended and VClass.So, we remove them.
ve_df = ve_df.drop(columns=['phevBlended', 'VClass'])
# ### 3.3.4 Outlier detection using Boxplot rule
# +
q1 = ve_df.score.quantile(0.25)
q3 = ve_df.score.quantile(0.75)
# Calculate the interquantile range (IQR)
IQR = q3 - q1
ll = q1 - 1.5 * IQR
ul = q3 + 1.5 * IQR
# -
ll_outliers = ve_df.score < ll
ul_outliers = ve_df.score > ul
print('Lower limit outliers:', ll_outliers.sum())
print('Upper limit outliers:', ul_outliers.sum())
print('Total outliers:', ll_outliers.sum() + ul_outliers.sum())
# Although we detect 7046 outliers, we are not removing these at this moment. Since there are a sizeable number of outliers, removing them will reduce the data set size. We can use ensemble machine learning algorithms such as Random Forests which are somewhat robust to outliers. Later on, if we do no get satisfactory model performance, we can remove these outliers.
ve_df.to_csv('data/VE_Cleaned.csv', index=False)
# ### 3.3.5 Principal Component Analysis
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from sklearn.preprocessing import RobustScaler
ve_df = pd.read_csv('data/VE_Cleaned.csv')
ve_df.head()
# #### Data Normalization
# Data normalization
robust_scaler = RobustScaler()
ve_norm_arr = robust_scaler.fit_transform(ve_df.loc[:, ve_df.columns != 'fuelType1'])
ve_norm_df = pd.DataFrame(ve_norm_arr)
ve_norm_df.describe().T
# #### Full-component PCA solution
# Fit linear Kernel
pca_linear = PCA(n_components=7)
pca_linear.fit(ve_norm_arr)
#Cumulative Variance
var_pca = np.cumsum(np.round(pca_linear.explained_variance_ratio_, decimals=4) * 100)
print(var_pca)
# Scree plot
var_pca = pd.DataFrame(var_pca, index=np.arange(1, 8))
plt.plot(var_pca, color='blue')
plt.title('Scree Plot')
plt.xlabel('Number of Principal Components')
plt.ylabel('Cumulative Variance Explained (%)')
plt.savefig('scree_plot.png', dpi=600, bbox_inches='tight')
# From the cummulative variance data and the screen plot obtained using linear PCA, we find that 3 components are sufficient to explain 96% of the variance. Therefore, 3 orthogonal (uncorrelated) components can keep more than 95% variance of the 7 variables in the original dataset. This implies we would be obtaining a good solution.
pca_linear = PCA(n_components=3)
ve_pca_transform_arr = pca_linear.fit_transform(ve_norm_arr)
ve_pca_transform_df = pd.DataFrame(ve_pca_transform_arr)
ve_pca_transform_df.columns = ['PC1', 'PC2', 'PC3']
ve_pca_transform_df.head(10)
ve_norm_df.corr()
ve_pca_transform_df.corr()
# When we compare the normalized data correlation and the PCA transformed correlation, we see that the principal components are almost uncorrelated with each other. Earlier, e.g, city08 (city MPG for fuelType1) had a high correlation with comb08 (combined MPG for fuelType1).
# PCA loadings
pd.DataFrame(pca_linear.components_.transpose(),
index=ve_df.loc[:, ve_df.columns != 'fuelType1'].columns,
columns=ve_pca_transform_df.columns)
# #### Visualizing PCA Components
# +
# %matplotlib widget
# %matplotlib inline
ve_df.score = pd.Categorical(ve_df.score)
score_color = ve_df.score.cat.codes
# Plot initialisation
fig = plt.figure(figsize=(480/96, 480/96), dpi=96)
ax = fig.add_subplot(111, projection='3d')
ax.scatter(ve_pca_transform_df['PC1'], ve_pca_transform_df['PC2'], ve_pca_transform_df['PC3'],
c=score_color, cmap="tab20c", s=60)
# make simple, bare axis lines through space:
xAxisLine = ((min(ve_pca_transform_df['PC1']), max(ve_pca_transform_df['PC1'])), (0, 0), (0,0))
ax.plot(xAxisLine[0], xAxisLine[1], xAxisLine[2], 'r')
yAxisLine = ((0, 0), (min(ve_pca_transform_df['PC2']), max(ve_pca_transform_df['PC2'])), (0,0))
ax.plot(yAxisLine[0], yAxisLine[1], yAxisLine[2], 'r')
zAxisLine = ((0, 0), (0,0), (min(ve_pca_transform_df['PC3']), max(ve_pca_transform_df['PC3'])))
ax.plot(zAxisLine[0], zAxisLine[1], zAxisLine[2], 'r')
# label the axes
ax.set_xlabel("PC1")
ax.set_ylabel("PC2")
ax.set_zlabel("PC3")
plt.title("Total Explained Variance: {:.2f}%".format(pca_linear.explained_variance_ratio_.sum() * 100))
plt.show()
# -
# Note that for PCA analysis, we leave out fuelType1 even though it is an important variable.
# # 4. Predictive Modeling
# ## 4.1 Loading cleaned data
import pandas as pd
# Note that we are not using dimension reduced data sets from PCA or correlation analysis.
ve_df_all = pd.read_csv('data/Vehicle_Emissions.csv')
ve_df = pd.read_csv('data/VE_Cleaned.csv')
ve_df['VClass'] = ve_df_all['VClass'] # Getting original VClass
# +
# Creating dummies
ve_df = pd.get_dummies(ve_df, columns=['fuelType1', 'VClass'])
# -
ve_df.head()
# ## 4.2 Data splitting
from sklearn.model_selection import train_test_split
y = ve_df['score'].copy()
X = ve_df.drop(columns='score')
random_state = 0
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
random_state=random_state,
stratify=y
)
# ## 4.3 Training and Test Data Normalization
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train_norm = scaler.fit_transform(X_train.to_numpy())
X_test_norm = scaler.transform(X_test.to_numpy())
# ## 4.4 Model Tuning on Local System
# ### 4.4.1 Random Forests
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV, RepeatedKFold
# +
from sklearn.metrics import make_scorer, cohen_kappa_score
score_avg = 'macro'
kappa_scorer = make_scorer(cohen_kappa_score)
scoring_metrics = {
'f1_' + score_avg: 'f1_' + score_avg,
'precision_' + score_avg: 'precision_' + score_avg,
'recall_' + score_avg: 'recall_' + score_avg,
'balanced_accuracy': 'balanced_accuracy',
'kappa': kappa_scorer
}
scoring_metrics
# -
param_dict = {
'n_estimators': [500],
'max_features': [5, 8],
'max_depth': [None],
'max_samples': [0.8],
'min_samples_leaf': [1],
'criterion': ['gini', 'entropy'],
'class_weight': ['balanced'] #, 'balanced_subsample']
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = RandomForestClassifier(n_jobs=-1, random_state=random_state)
model_grid = GridSearchCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_rf = model_grid.fit(X_train_norm, y_train)
# +
# User defined function for computing model metrics
import numpy as np
from sklearn.metrics import roc_auc_score, balanced_accuracy_score, f1_score, precision_score, recall_score, cohen_kappa_score
from sklearn.metrics import classification_report
def calc_error_metrics(gs_obj, X_test, y_test, score_avg='macro'):
"""
Calculate train, validation, test error metrics, and classification report (for the test data)
:param gs_obj: Fitted GridSearchCV object
:param X_test: X_test
:param y_test: y_test
:param score_avg: Scikit-learn scorer avergaging for multi-class classification
:return Pandas dataframe containing the error metrics and classification report
"""
# auc_list = []
f1_list = []
precision_list = []
recall_list = []
ba_list = []
kappa_list = []
search_data = ['mean_train', 'mean_test']
scores = gs_obj.cv_results_
for sd in search_data:
# auc = scores['{}_roc_auc_{}'.format(sd, score_avg)].mean()
f1 = scores['{}_f1_{}'.format(sd, score_avg)].mean()
precision = scores['{}_precision_{}'.format(sd, score_avg)].mean()
recall = scores['{}_recall_{}'.format(sd, score_avg)].mean()
ba = scores['{}_balanced_accuracy'.format(sd)].mean()
kappa = scores['{}_kappa'.format(sd)].mean()
# auc_list.append(auc)
f1_list.append(f1)
precision_list.append(precision)
recall_list.append(recall)
ba_list.append(ba)
kappa_list.append(kappa)
best_model = gs_obj.best_estimator_
y_pred = best_model.predict(X_test)
# y_pred_proba = best_model.predict_proba(X_test)
labels = np.unique(y_pred)
# test_auc = roc_auc_score(y_test, y_pred, labels=labels, average=score_avg)
test_f1 = f1_score(y_test, y_pred, labels=labels, average=score_avg)
test_precision = precision_score(y_test, y_pred, labels=labels, average=score_avg)
test_recall = recall_score(y_test, y_pred, labels=labels, average=score_avg)
test_ba = balanced_accuracy_score(y_test, y_pred)
test_kappa = cohen_kappa_score(y_test, y_pred, labels=labels)
# auc_list.append(test_auc)
f1_list.append(test_f1)
precision_list.append(test_precision)
recall_list.append(test_recall)
ba_list.append(test_ba)
kappa_list.append(test_kappa)
results_dict = {
'Data': ['Train', 'Validation', 'Test'],
# 'ROC_AUC': auc_list,
'F1 Score': f1_list,
'Precision': precision_list,
'Recall': recall_list,
'Balanced Accuracy': ba_list,
'Kappa Score': kappa_list
}
results_df = pd.DataFrame(data=results_dict)
report = classification_report(y_test, y_pred, target_names=labels.astype(str), labels=labels)
return results_df, report
# -
metrics_df, report = calc_error_metrics(search_rf, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_rf.best_params_)
# ## 4.5 Model Tuning on the Missouri S&T Foundry HPC
# We use Dask to perform model tuning on a distributed computing environment
# ### 4.5.1 LightGBM
from dask_ml.model_selection import GridSearchCV as DaskGCV
from dask.distributed import Client
from dask_jobqueue import SLURMCluster
from lightgbm import LGBMClassifier
param_dict = {
'boosting_type': ['gbdt'],
'n_estimators': [300, 500],
'max_depth': [10, 20, 30, -1],
'max_bin': [127, 255],
'min_data_in_bin': [3, 10, 20],
'learning_rate': [0.05, 0.1],
'subsample': [1, 0.8],
'colsample_bytree': [1, 0.5],
'colsample_bynode': [1],
'path_smooth': [0, 0.1, 0.2],
'num_leaves': [31, 63],
'min_child_samples': [20],
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = LGBMClassifier(
tree_learner='feature',
class_weight='balanced',
random_state=random_state,
# deterministic=True, force_row_wise=True
)
cluster = SLURMCluster(
cores=40,
processes=1,
memory="50G",
walltime="02:00:00",
interface='ib0',
env_extra=['#SBATCH --out=IST-5520-Foundry-Dask-%j.out']
)
cluster.adapt(
minimum=10, maximum=50,
minimum_jobs=10, maximum_jobs=50,
minimum_memory='40G', maximum_memory='50G'
)
dask_client = Client(cluster)
print('Waiting for dask workers...')
dask_client.wait_for_workers(10)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv,
refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_dask_lgbm = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_dask_lgbm, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_dask_lgbm.best_params_)
dask_client.shutdown()
# ### 4.5.2 Random Forests
param_dict = {
'n_estimators': [300, 500],
'max_features': [3, 5, 10, 20],
'max_depth': [6, 10, None],
'max_samples': [0.8, 0.5, 0.9, None],
'min_samples_leaf': [1],
'criterion': ['gini', 'entropy'],
'class_weight': ['balanced', 'balanced_subsample']
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = RandomForestClassifier(
n_jobs=-1,
random_state=random_state)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv,
refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_rf = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_rf, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_rf.best_params_)
# ### 4.5.3 Support Vector Machine
from sklearn.svm import SVC
param_dict = {
'C': [0.5, 0.8, 1],
'gamma': ['scale', 'auto', 0.8],
'tol': [1e-3, 1e-2],
'break_ties': [True, False]
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = SVC(class_weight='balanced', random_state=random_state)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_svm = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_svm, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_svm.best_params_)
# ### 4.5.4 Extra Trees Classifier
from sklearn.ensemble import ExtraTreesClassifier
param_dict = {
'n_estimators': [300, 500],
'max_features': [3, 5, 7, 10],
'max_depth': [6, 10, None],
'max_samples': [0.8, 0.5, 0.9, None],
'min_samples_leaf': [1],
'criterion': ['gini', 'entropy'],
'class_weight': ['balanced', 'balanced_subsample']
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = ExtraTreesClassifier(
class_weight='balanced',
random_state=random_state,
n_jobs=-1
)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_et = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_et, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_et.best_params_)
# ### 4.5.5 MLP Classifier
from sklearn.neural_network import MLPClassifier
param_dict = {
'hidden_layer_sizes': [(512, 256), (256, 128, 64), (64, 32, 16, 8)],
'learning_rate_init': [1e-3, 1e-2],
'max_iter': [100, 200],
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = MLPClassifier(random_state=random_state)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_mlp = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_mlp, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_mlp.best_params_)
import matplotlib.pyplot as plt
plt.plot(search_mlp.best_estimator_.loss_curve_)
plt.show()
# #### 4.5.6 Boosted Random Forests
param_dict = {
'n_estimators': [300, 500],
'max_depth': [10, 20, -1],
'max_bin': [127, 255],
'min_data_in_bin': [3, 10, 20],
'learning_rate': [0.05, 0.1],
'subsample': [0.9, 0.5],
'colsample_bytree': [1, 0.5],
'colsample_bynode': [1],
'path_smooth': [0, 0.1, 0.2],
'num_leaves': [31, 63],
'min_child_samples': [20],
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = LGBMClassifier(
boosting_type='rf',
bagging_freq=1,
class_weight='balanced',
random_state=random_state
)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_brf = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_brf, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_brf.best_params_)
# ### 4.5.7 Multinomial Logistic Regression
from sklearn.linear_model import LogisticRegression
y = ve_df['score'].copy()
X = ve_df.drop(columns='score')
X['const'] = 1
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
random_state=random_state,
stratify=y
)
scaler = MinMaxScaler()
X_train_norm = scaler.fit_transform(X_train.to_numpy())
X_test_norm = scaler.transform(X_test.to_numpy())
from sklearn.model_selection import RepeatedKFold
param_dict = {
'tol': [1e-4, 1e-3, 1e-2],
'C': [0.5, 0.8, 1],
'solver': ['saga', 'lbfgs'],
'max_iter': [100, 200, 300, 500]
}
cv = RepeatedKFold(n_splits=5, n_repeats=1, random_state=random_state)
model = LogisticRegression(
class_weight='balanced',
random_state=random_state,
multi_class='multinomial',
n_jobs=-1
)
model_grid = DaskGCV(
estimator=model,
param_grid=param_dict,
scoring=scoring_metrics,
n_jobs=-1, cv=cv, refit=scoring_metrics['f1_' + score_avg],
return_train_score=True
)
search_mlr = model_grid.fit(X_train_norm, y_train)
metrics_df, report = calc_error_metrics(search_mlr, X_test_norm, y_test, score_avg)
metrics_df
print(report)
print(search_mlr.best_params_)
# # 5. Final Model
# ## 5.1 Get Training and Test Data
import pandas as pd
# Note that we are not using dimension reduced data sets from PCA or correlation analysis.
ve_df_all = pd.read_csv('data/Vehicle_Emissions.csv')
ve_df = pd.read_csv('data/VE_Cleaned.csv')
ve_df['VClass'] = ve_df_all['VClass'] # Getting original VClass
# +
# Creating dummies
ve_df = pd.get_dummies(ve_df, columns=['fuelType1', 'VClass'])
# -
from sklearn.model_selection import train_test_split
y = ve_df['score'].copy()
X = ve_df.drop(columns='score')
random_state = 0
X_train, X_test, y_train, y_test = train_test_split(
X, y,
test_size=0.3,
random_state=random_state,
stratify=y
)
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
X_train_norm = scaler.fit_transform(X_train.to_numpy())
X_test_norm = scaler.transform(X_test.to_numpy())
# ## 5.1 Fitting with the best parameters
param_dict = {
'boosting_type': 'gbdt',
'colsample_bynode': 1,
'colsample_bytree': 1,
'learning_rate': 0.05,
'max_bin': 127,
'max_depth': 10,
'min_child_samples': 20,
'min_data_in_bin': 3,
'n_estimators': 300,
'num_leaves': 31,
'path_smooth': 0.2,
'subsample': 1
}
random_state = 0
score_avg = 'macro'
# +
from lightgbm import LGBMClassifier
model = LGBMClassifier(
tree_learner='feature',
class_weight='balanced',
random_state=random_state,
)
model.set_params(**param_dict)
# -
model.fit(X_train_norm, y_train)
# +
import matplotlib.pyplot as plt
# %matplotlib inline
f_dict = {
'Feature': X_train.columns,
'Importance': model.feature_importances_
}
feature_df = pd.DataFrame(data=f_dict).sort_values(by='Importance', ascending=False)
feature_df = feature_df[feature_df.Importance >= 1000]
plt.rcParams["figure.figsize"] = (10, 5)
feature_df.set_index('Feature').plot.bar(rot=45)
# -
# ## 5.2 Test Data Predictions
# +
import numpy as np
from sklearn.metrics import roc_auc_score, balanced_accuracy_score, f1_score, precision_score, recall_score, cohen_kappa_score
from sklearn.metrics import classification_report
y_pred = model.predict(X_test_norm)
y_pred_proba = model.predict_proba(X_test)
labels = np.unique(y_pred)
test_auc = roc_auc_score(y_test, y_pred_proba, labels=labels, average=score_avg, multi_class='ovr')
test_f1 = f1_score(y_test, y_pred, labels=labels, average=score_avg)
test_precision = precision_score(y_test, y_pred, labels=labels, average=score_avg)
test_recall = recall_score(y_test, y_pred, labels=labels, average=score_avg)
test_ba = balanced_accuracy_score(y_test, y_pred)
test_kappa = cohen_kappa_score(y_test, y_pred, labels=labels)
results_dict = {
'ROC_AUC': [test_auc],
'F1 Score': [test_f1],
'Precision': [test_precision],
'Recall': [test_recall],
'Balanced Accuracy': [test_ba],
'Kappa Score': [test_kappa]
}
results_df = pd.DataFrame(data=results_dict)
report = classification_report(y_test, y_pred, target_names=labels.astype(str), labels=labels)
# -
results_df.T
print(report)
# +
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
import matplotlib.pyplot as plt
# %matplotlib inline
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
import itertools
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.tight_layout()
# +
cnf_matrix = confusion_matrix(y_test, y_pred,labels=model.classes_)
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plt.rcParams["figure.figsize"] = (20, 20)
plt.rcParams["font.size"] = 18
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=model.classes_)
# -
# # 6. Summary
# We developed a machine learning-based predictive model to predict the vehicle emission score or smog rating (vehicle tailpipe CO2 emission) as defined by the Environmental Protection Agency (EPA). Essentially, the task is to perform a multi-class classification of the emission score, a discrete unitless integer quantity, varying from 1 (worst) to 10 (best). For this purpose, we collected the ‘vehicle’ and ‘emissions’ data sets from the EPA portal [1], [2].
#
# As of now, the ‘vehicle’ and ‘emissions’ data contain around 44000 and 42000 records each with 83 and 8 features or columns, respectively. However, these data sets must be joined based on the vehicle id column which reduces the number of available observations because of missing ids. Moreover, the emission score had incorrect entries (negative scores) which were removed. Hence, after the data merging and cleaning steps, we had around 28000 observations.
#
# Thereafter, for categorical data transformation, we aggregated the fuelType1 column by combining midgrade and regular gasolines as 'Regular Gasoline' and 'Electricity' and 'Natural Gas' as 'Green Fuel'. Next, we performed correlation and distribution analysis, using which we were able to answer one of the sub research questions. We found that Standard Pickup Trucks and Vans (both cargo and passenger types) had the lowest emission scores in general. In addition, it was confirmed from boxplot analysis that, diesel vehicles received the lowest scores.
#
# An interesting observation from our boxplot outlier analysis was that all scores except those in the interval [4, 7] were marked as outliers. Hence, our initial choice of not performing outlier removal as part of the second milestone was justifiable. Similarly, we found principal component analysis (PCA) to be of not much use possibly because of strong non-linearity in the data set. We may need to apply kernel PCA-based approaches for appropriately investigating the benefits of PCA.
#
# The final component of this project was the predictive modeling where we compared seven classification models. These are Multinomial Logistic Regression (MLR), Support Vector Machine (SVM), Gradient Boosting Trees (GBT), Random Forests (RF), Boosted Random Forests (BRF), Extremely Randomized Trees (ETR), and Multi-Layer Perceptron (MLP). For the GBT and BRF algorithms, we used the LightGBM [3] package and used scikit-learn for the rest.
#
# One issue with the given data set is extremely high class imbalance. For example, the EPA score 4 only has 6 samples (0.02%) in total while 40% of the total emission scores are marked as 5. Another potential problem was the data dimensionality (35 predictors in total including dummies from the VClass column).
#
# Therefore, we preferred using ensemble machine learning algorithms that support class weights as parameters and provide feature selection on the fly. Moreover, we only have 28000 observations, so going for deep learning models is overkill.
#
# In order to appropriately compare all the models, we scaled the features in the [0, 1] interval using MinMaxScaler. Next, to train the models, we initially used the scikit-learn GridSearchCV with RepeatedKFold having five-folds and a single repeat (we tried multiple repeats too, but the results were similar). Here, we could not use the stratified version because class 4 only had four observations in the training data. This was also a reason why we used 70-30 split of the training and test data. If we used 80% as training, then class 4 only had a single observation in the test data.
#
# The model evaluation was based on the similarity of the training and validation F1 score to reduce overfitting. We then checked the F1 for the test data. For additional interpretability, we also showed balanced accuracy, precision, recall, and the Cohen’s kappa score. Note that F1 was the metric that was optimized. For, F1, precision, and recall, we used the macro averaging as part of the GridSearch scoring metrics. In this context, we had NaN issues with the ROC AUC macro OVR (one-versus-rest) scoring metric through GridSearchCV. That is why we only show the ROC AUC for the test data of the final model.
#
# While running the scikit-learn GridSearchCV with different hyperparameters of the ensemble learning algorithms and MLP on an Alienware M17 R1 (personal PC), we noticed that for many parameter values, even 8 hrs of computation was not sufficient. As a result, we switched to the Missouri S&T Foundry High-Performance Computing cluster and used Dask [4] to train our models in a distributed computing environment. This resulted in a significant reduction of computing time wherein we could tune several parameters in a very short time. For example, the GBM model training took around 6 hrs on the Alienware and took only 15 minutes using Dask.
#
#
# Finally, we found that the GBM shown in section 5 has the best performance in terms of all the error metrics for the test data.
# # References
# [1] EPA, “EPA Fuel Economy,” 2021. https://www.fueleconomy.gov/ (accessed Dec. 05, 2021).
#
# [2] EPA, “Fuel Economy Guide,” 2021. [Online]. Available: https://www.fueleconomy.gov/feg/pdfs/guides/FEG2021.pdf.
#
# [3] <NAME> et al., “LightGBM: A Highly Efficient Gradient Boosting Decision Tree,” in Advances in Neural Information Processing Systems, 2017, vol. 30, [Online]. Available: https://proceedings.neurips.cc/paper/2017/file/6449f44a102fde848669bdd9eb6b76fa-Paper.pdf.
#
# [4] <NAME>, “Dask: Parallel Computation with Blocked algorithms and Task Scheduling,” Proc. 14th Python Sci. Conf, 2015, Accessed: 29-Nov-2021. [Online]. Available: http://conference.scipy.org/proceedings/scipy2015/pdfs/matthew_rocklin.pdf.
#
| Analysis&Predictive Modelling of Proj.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Patch
from minisom import MiniSom
from sklearn.preprocessing import minmax_scale, scale
from sklearn import preprocessing
import settings
# -
png_name = 'icd_food_control'
regression_targets = f'{settings.raw_data_folder}control/regression_targets.csv'
data = f'{settings.processed_data_folder}{png_name}_selected_features.csv'
df = pd.read_csv(data, index_col=0)
targets = pd.read_csv(regression_targets)
merged = pd.merge(df, targets, on='Country')
merged
def scale_df(df):
x = df.values # returns a numpy array
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df_scaled = pd.DataFrame(x_scaled)
return df_scaled
columns_for_scale = merged.columns
columns_for_scale = columns_for_scale.tolist()
columns_for_scale.remove('Country')
countries = merged['Country']
scaled = scale_df(merged[columns_for_scale])
scaled.columns = columns_for_scale
features_for_som = columns_for_scale
features_for_som.remove('Deaths')
scaled['Country'] = countries
scaled['binned'] = pd.cut(scaled['Deaths'], 3, labels = ['small', 'medium', 'large'])
df_features_scaled_not_nan = scaled[scaled['binned'].notna()]
all_countries = df_features_scaled_not_nan['Country']
category_color = {'small': 'darkgreen',
'medium': '#ffa500',
'large': 'red'
}
colors_dict = {}
for c, dm in zip(all_countries, df_features_scaled_not_nan['binned']):
colors_dict[c] = category_color[dm]
# + jupyter={"source_hidden": true}
country_codes = {'Afghanistan': 'AF',
'Albania': 'AL',
'Algeria': 'DZ',
'Angola': 'AO',
'Argentina': 'AR',
'Armenia': 'AM',
'Australia': 'AU',
'Austria': 'AT',
'Azerbaijan': 'AZ',
'Bahrain': 'BH',
'Bangladesh': 'BD',
'Belarus': 'BY',
'Belgium': 'BE',
'Benin': 'BJ',
'Bhutan': 'BT',
'Bolivia': 'BO',
'Bosnia and Herzegovina': 'BA',
'Botswana': 'BW',
'Brazil': 'BR',
'Bulgaria': 'BG',
'Burkina Faso': 'BF',
'Burundi': 'BI',
'Cambodia': 'KH',
'Cameroon': 'CM',
'Canada': 'CA',
'Cape Verde': 'CV',
'Central African Republic': 'CF',
'Chad': 'TD',
'Chile': 'CL',
'China': 'CN',
'Colombia': 'CO',
'Comoros': 'KM',
'Costa Rica': 'CR',
'Croatia': 'HR',
'Cuba': 'CU',
'Cyprus': 'CY',
'Czechia': 'CZ',
'Democratic Republic of the Congo': 'CD',
'Denmark': 'DK',
'Djibouti': 'DJ',
'Dominican Republic': 'DO',
'Ecuador': 'EC',
'Egypt': 'EG',
'El Salvador': 'SV',
'Equatorial Guinea': 'GQ',
'Eritrea': 'ER',
'Estonia': 'EE',
'Ethiopia': 'ET',
'Fiji': 'FJ',
'Finland': 'FI',
'France': 'FR',
'Gabon': 'GA',
'Gambia': 'GM',
'Georgia': 'GE',
'Germany': 'DE',
'Ghana': 'GH',
'Greece': 'GR',
'Guatemala': 'GT',
'Guinea': 'GN',
'Guinea-Bissau': 'GW',
'Guyana': 'GY',
'Haiti': 'HT',
'Honduras': 'HN',
'Hong Kong': 'HK',
'Hungary': 'HU',
'Iceland': 'IS',
'India': 'IN',
'Indonesia': 'ID',
'Iran (Islamic Republic of)': 'IR',
'Iraq': 'IQ',
'Ireland': 'IE',
'Israel': 'IL',
'Italy': 'IT',
'Ivory Coast': 'IC',
'Jamaica': 'JM',
'Japan': 'JP',
'Jordan': 'JO',
'Kazakhstan': 'KZ',
'Kenya': 'KE',
'Kuwait': 'KW',
'Kyrgyzstan': 'KG',
'Laos': 'LA',
'Latvia': 'LV',
'Lebanon': 'LB',
'Lesotho': 'LS',
'Liberia': 'LR',
'Libya': 'LY',
'Lithuania': 'LT',
'Luxembourg': 'LU',
'Macedonia': 'MK',
'Madagascar': 'MG',
'Malawi': 'MW',
'Malaysia': 'MY',
'Mali': 'ML',
'Malta': 'MT',
'Mauritania': 'MR',
'Mauritius': 'MU',
'Mexico': 'MX',
'Republic of Moldova': 'MD',
'Mongolia': 'MN',
'Montenegro': 'ME',
'Morocco': 'MA',
'Mozambique': 'MZ',
'Myanmar': 'MM',
'Namibia': 'NA',
'Nepal': 'NP',
'Netherlands': 'NL',
'New Zealand': 'NZ',
'North Macedonia': 'NM',
'Nicaragua': 'NI',
'Niger': 'NE',
'Nigeria': 'NG',
'North Korea': 'KP',
'Norway': 'NO',
'Oman': 'OM',
'Pakistan': 'PK',
'Palestine': 'PS',
'Panama': 'PA',
'Papua New Guinea': 'PG',
'Paraguay': 'PY',
'Peru': 'PE',
'Philippines': 'PH',
'Poland': 'PL',
'Portugal': 'PT',
'Qatar': 'QA',
'Republic of China (Taiwan)': 'TW',
'Republic of the Congo': 'CG',
'Romania': 'RO',
'Russia': 'RU',
'Rwanda': 'RW',
'Saudi Arabia': 'SA',
'Senegal': 'SN',
'Serbia': 'RS',
'Sierra Leone': 'SL',
'Singapore': 'SG',
'Slovakia': 'SK',
'Slovenia': 'SI',
'South Africa': 'ZA',
'South Korea': 'KR',
'Spain': 'ES',
'Sri Lanka': 'LK',
'Sudan': 'SD',
'Suriname': 'SR',
'Swaziland': 'SZ',
'Sweden': 'SE',
'Switzerland': 'CH',
'Syria': 'SY',
'Tajikistan': 'TJ',
'Tanzania': 'TZ',
'Thailand': 'TH',
'Timor-Leste': 'TL',
'Togo': 'TG',
'Trinidad and Tobago': 'TT',
'Tunisia': 'TN',
'Turkey': 'TR',
'Turkmenistan': 'TM',
'Uganda': 'UG',
'Ukraine': 'UA',
'United Arab Emirates': 'AE',
'United Kingdom': 'GB',
'United States of America': 'US',
'Uruguay': 'UY',
'Uzbekistan': 'UZ',
'Venezuela (Bolivarian Republic of)': 'VE',
'Vietnam': 'VN',
'Yemen': 'YE',
'Zambia': 'ZM',
'Zimbabwe': 'ZW',
'Bahamas': 'BHMS',
'Saint Vincent and the Grenadines': 'SVG',
'Barbados': 'BRBDS',
'Antigua and Barbuda' : 'AAB',
'Maldives' : 'MDV',
'Belize': 'BZE',
'Saint Lucia': 'SL',
'Grenada': 'GDA',
'Lao People\'s Democratic Republic':'LPDP',
'United Republic of Tanzania': 'TNZ',
'Russian Federation': 'Russia',
'Saint Kitts and Nevis' : 'SK',
'Uganda': 'UG',
'Sao Tome and Principe': 'STP',
'Solomon Islands': 'SI',
'Cote d\'Ivoire': 'CI'}
# +
feature_names = features_for_som
X = df_features_scaled_not_nan[feature_names].values
size = 8
som = MiniSom(size, size, len(X[0]),
neighborhood_function='gaussian', sigma=1.5,
random_seed=1)
som.pca_weights_init(X)
som.train_random(X, 2500, verbose=True)
# +
def shorten_country(c):
if len(c) > 8 and c in country_codes:
return country_codes[c]
else:
return c
country_map = som.labels_map(X, all_countries)
plt.figure(figsize=(14, 14))
for p, countries in country_map.items():
countries = list(countries)
x = p[0] + .1
y = p[1] - .3
for i, c in enumerate(countries):
off_set = (i+1)/len(countries) + 0.15
plt.text(x, y+off_set, shorten_country(c), color=colors_dict[c], fontsize=10)
plt.pcolor(som.distance_map().T, cmap='gray_r', alpha=.2)
plt.xticks(np.arange(size+1))
plt.yticks(np.arange(size+1))
plt.grid()
legend_elements = [Patch(facecolor=clr,
edgecolor='w',
label=l) for l, clr in category_color.items()]
plt.legend(handles=legend_elements, loc='center left', bbox_to_anchor=(1, .95))
plt.savefig(f'./{png_name}_{size}x{size}.png', dpi=300, bbox_inches = "tight")
plt.show()
# -
import math
W = som.get_weights()
size_plot = math.sqrt(len(feature_names))
size_plot = math.floor(size_plot)
plt.figure(figsize=(20, 20))
for i, f in enumerate(feature_names):
plt.subplot(size_plot+1, size_plot+1, i+1)
plt.title(f)
plt.pcolor(W[:,:,i].T, cmap='coolwarm')
plt.xticks(np.arange(size_plot+1))
plt.yticks(np.arange(size_plot+1))
plt.tight_layout()
plt.savefig(f'./{png_name}_{size}x{size}_weights.png', dpi=300, bbox_inches = "tight")
plt.show()
# +
Z = np.zeros((size, size))
plt.figure(figsize=(8, 8))
for i in np.arange(som._weights.shape[0]):
for j in np.arange(som._weights.shape[1]):
feature = np.argmax(W[i, j , :])
plt.plot([j+.5], [i+.5], 'o', color='C'+str(feature),
marker='s', markersize=24)
legend_elements = [Patch(facecolor='C'+str(i),
edgecolor='w',
label=f) for i, f in enumerate(feature_names)]
plt.legend(handles=legend_elements,
loc='center left',
bbox_to_anchor=(1, .95))
plt.xlim([0, size])
plt.ylim([0, size])
plt.savefig(f'./{png_name}_{size}x{size}_decisions.png', dpi=300, bbox_inches = "tight")
plt.show()
# -
| self_organized_maps/som_icd_food_control_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
# +
import sys
SOURCE_DIR = os.path.dirname(os.path.dirname(os.path.abspath(__name__)))
sys.path.insert(0, SOURCE_DIR)
# -
import tensorflow as tf
import malaya_speech
import malaya_speech.train
from malaya_speech.train.model import melgan
import malaya_speech.config
import numpy as np
melgan_config = malaya_speech.config.melgan_config
generator = melgan.Generator(
melgan.GeneratorConfig(**melgan_config['melgan_generator_params']),
name='melgan-generator',
)
discriminator = melgan.MultiScaleDiscriminator(
melgan.DiscriminatorConfig(**melgan_config['melgan_discriminator_params']),
name='melgan-discriminator',
)
mels_loss = melgan.loss.TFMelSpectrogram()
y = tf.placeholder(tf.float32, (None, None))
x = tf.placeholder(tf.float32, (None, None, 80))
y_hat = generator(x)
p_hat = discriminator(y_hat)
p = discriminator(tf.expand_dims(y, -1))
from malaya_speech.train.loss import calculate_2d_loss, calculate_3d_loss
mse_loss = tf.keras.losses.MeanSquaredError(
reduction='none'
)
mae_loss = tf.keras.losses.MeanAbsoluteError(
reduction='none'
)
# +
adv_loss = 0.0
for i in range(len(p_hat)):
adv_loss += calculate_3d_loss(
tf.ones_like(p_hat[i][-1]), p_hat[i][-1], loss_fn=mse_loss
)
adv_loss /= i + 1
fm_loss = 0.0
for i in range(len(p_hat)):
for j in range(len(p_hat[i]) - 1):
fm_loss += calculate_3d_loss(
p[i][j], p_hat[i][j], loss_fn=mae_loss
)
fm_loss /= (i + 1) * (j + 1)
adv_loss += 10 * fm_loss
spect_loss = calculate_2d_loss(
y, tf.squeeze(y_hat, -1), loss_fn = mels_loss
)
# -
real_loss = 0.0
fake_loss = 0.0
for i in range(len(p)):
real_loss += calculate_3d_loss(
tf.ones_like(p[i][-1]), p[i][-1], loss_fn=mse_loss
)
fake_loss += calculate_3d_loss(
tf.zeros_like(p_hat[i][-1]), p_hat[i][-1], loss_fn=mse_loss
)
real_loss /= i + 1
fake_loss /= i + 1
dis_loss = real_loss + fake_loss
adv_loss, dis_loss, spect_loss
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# +
# adv_loss_, dis_loss_ = sess.run([adv_loss, dis_loss],
# feed_dict = {x: np.random.uniform(size=(1,200,80)),
# y: np.random.uniform(size=(1,51200))})
# +
# y_hat_, loss, p_, p_hat_ = sess.run([y_hat, p_hat, p],
# feed_dict = {x: np.random.uniform(size=(1,200,80)),
# y: np.random.uniform(size=(1,51200))})
# -
sess.run(y_hat, feed_dict = {x: np.random.uniform(size=(1,200,80))}).shape
| test/test-melgan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#export
import torch
# +
# default_exp functions
# -
# # Functions
#
# > This module contains implementations of both traditional survival analysis functions, as well as the loss functions associated with uncensored data, as defined in the original DRSA paper.
#hide
from nbdev.showdoc import *
import pytest
# ## Survival Analysis Functions
#
# Following the notation used in the the [DRSA paper](https://arxiv.org/pdf/1809.02403.pdf), we define the following:
#
# * Let $z$ be the true occurrence time for the event of interest.
#
# * Let $t$ be the time that a given data point was observed.
#
# * For each observation, there exist $L$ time slices, ie $0 < t_1 < t_2 < \dots < t_L$, at which we either observe the event (uncensored) or do not (censored).
#
# * Let $V_l = (t_{l-1}, t_l]$ be the set of all disjoint intervals with $l = 1, 2, \dots, L$.
# +
#exporti
def assert_correct_input_shape(h):
if len(h.shape) != 3:
raise ValueError(f"h is of shape {h.shape}. It is expected that h is of shape (batch size, sequence_length, 1), as this is most amenable to use in training neural nets with pytorch.")
def assert_correct_output_shape(q, batch_size):
if q.shape != torch.Size([batch_size, 1]):
raise ValueError(f"q is of shape {q.shape}. It is expected that q is of shape (batch_size, 1)")
# -
# ### Discrete Survival function
#
# Though it's given its own name is survival analysis, the survival function is simply calculated as $1 - \text{CDF}(z)$. In the discrete, empirical case, the survival function is estimated as follows (this is equation (5) in the paper).
#
# $$ S(t_l) = Pr(z > t_l) = \sum_{j > l}Pr(z\in V_j) $$
# +
#export
def survival_rate(h):
"""
Given the predicted conditional hazard rate, this function estimates
the survival rate.
*input*:
* `h`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`, as this is most amenable to use in training neural nets with pytorch.
_output_:
* `s`:
- type: `torch.tensor`
- estimated survival rate at time t.
- note: `s.shape == (batch_size, 1)`
"""
assert_correct_input_shape(h)
s = (1-h).prod(dim=1)
return s
# -
# example
h1 = torch.tensor([[0.001],
[0.5],
[0.55],
[0.15],
[0.15],
[0.15],
[0.15],
[0.9]], requires_grad=True)
h2 = torch.tensor([[0.001],
[0.005],
[0.1],
[0.11],
[0.12],
[0.15],
[0.15],
[0.9]], requires_grad=True)
h = torch.stack([h1, h2], dim=0)
survival_rate(h)
# +
#hide
# survival rate tests
def test_survival_rate(h):
# shape should be 3-d
with pytest.raises(ValueError):
s = survival_rate(h[0, :, :])
batch_size, length, _ = h.shape
s = survival_rate(h)
# output should have shape (batch_size, 1)
assert_correct_output_shape(s, batch_size)
# testing correct output
torch.testing.assert_allclose(s, torch.tensor([[0.0117], [0.0506]]), rtol=1e-3, atol=1e-3)
test_survival_rate(h)
# -
# ### Discrete Event Rate function
#
# The event rate function is calculated as $\text{CDF}(z)$. In the discrete, empirical case, it is estimated as follows (this is equation (5) in the paper).
#
# $$ W(t_l) = Pr(z \leq t_l) = \sum_{j\leq l}Pr(z\in V_j) $$
# +
#export
def event_rate(h):
"""
Given the predicted conditional hazard rate, this function estimates
the event rate.
*input*:
* `h`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`, as this is most amenable to use in training neural nets with pytorch.
_output_:
* `w`:
- type: `torch.tensor`
- estimated survival rate at time t.
- note: `w.shape == (batch_size, 1)`
"""
assert_correct_input_shape(h)
w = 1-survival_rate(h)
return w
# -
# example
event_rate(h)
# +
#hide
# event rate tests
def test_event_rate(h):
# shape should be 3-d
with pytest.raises(ValueError):
w = event_rate(h[0, :, :])
batch_size, length, _ = h.shape
w = event_rate(h)
# output should have shape (batch_size, 1)
assert_correct_output_shape(w, batch_size)
# testing correct output
torch.testing.assert_allclose(w, torch.tensor([[0.9883], [0.9494]]), rtol=1e-3, atol=1e-3)
test_event_rate(h)
# -
# ### Discrete Event Time Probability function
#
# The event time probability function is calculated as $\text{PDF}(z)$. In the discrete, empirical case, it is estimated as follows (this is equation (6) in the paper).
#
# $$p_l = Pr(z\in V_t) = W(t_l) - W(t_{l-1}) = S(t_{l-1}) - S(t_{l})$$
# +
#export
def event_time(h):
"""
Given the predicted conditional hazard rate, this function estimates
the probability that the event occurs at time t.
*input*:
* `h`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`, as this is most amenable to use in training neural nets with pytorch.
_output_:
* `p`:
- type: `torch.tensor`
- estimated probability of event at time t.
- note: `p.shape == (batch_size, 1)`
"""
assert_correct_input_shape(h)
p = h[:, -1, :] * survival_rate(h[:, :-1, :])
return p
# -
# example
event_time(h)
# +
#hide
# event time tests
def test_event_time(h):
# shape should be 3-d
with pytest.raises(ValueError):
p = event_time(h[0, :, :])
batch_size, length, _ = h.shape
p = event_time(h)
# output should have shape (batch_size, 1)
assert_correct_output_shape(p, batch_size)
# testing correct output
torch.testing.assert_allclose(p, torch.tensor([[0.1056], [0.4556]]), rtol=1e-3, atol=1e-3)
test_event_time(h)
# -
# ### Discrete Conditional Hazard Rate
#
# The conditional hazard rate is the quantity which will be predicted at each time step by a recurrent survival analysis model. In the discrete, empirical case, it is estimated as follows (this is equation (7) in the paper).
#
# $$h_l = Pr(z\in V_l | z > t_{l-1}) = \frac{Pr(z\in V_l)}{Pr(z>t_{l-1})} = \frac{p_l}{S(t_{l-1})}$$
# ## Log Survival Analysis Functions
#
# We additionally define the log of each of the traditional survival analysis functions, which prove useful for computational stability, being that we need to multiply many float point decimal values together.
# ### Log Survival Function
# +
#export
def log_survival_rate(h):
"""
Given the predicted conditional hazard rate, this function estimates
the log survival rate.
*input*:
* `h`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`, as this is most amenable to use in training neural nets with pytorch.
_output_:
* `s`:
- type: `torch.tensor`
- estimated log survival rate at time t.
- note: `s.shape == (batch_size, 1)`
"""
assert_correct_input_shape(h)
s = (1-h).log().sum(dim=1)
return s
# -
#example
log_survival_rate(h)
# +
#hide
# log survival rate tests
def test_log_survival_rate(h):
# shape should be 3-d
with pytest.raises(ValueError):
s = log_survival_rate(h[0, :, :])
batch_size, length, _ = h.shape
s = log_survival_rate(h)
# output should have shape (batch_size, 1)
assert_correct_output_shape(s, batch_size)
# testing correct output
torch.testing.assert_allclose(s, survival_rate(h).log(), rtol=1e-3, atol=1e-3)
test_log_survival_rate(h)
# -
# ### Log Event Rate Function
# +
#export
def log_event_rate(h):
"""
Given the predicted conditional hazard rate, this function estimates
the log event rate.
*input*:
* `h`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`, as this is most amenable to use in training neural nets with pytorch.
_output_:
* `w`:
- type: `torch.tensor`
- estimated log survival rate at time t.
- note: `w.shape == (batch_size, 1)`
"""
assert_correct_input_shape(h)
# w = event_rate(h).log() # numerically unstable, darn probabilities
w = (1 - log_survival_rate(h).exp()).log() # numerically stable
return w
# -
# example
log_event_rate(h)
# +
#hide
# log event rate tests
def test_log_event_rate(h):
# shape should be 3-d
with pytest.raises(ValueError):
w = log_event_rate(h[0, :, :])
batch_size, length, _ = h.shape
w = log_event_rate(h)
# output should have shape (batch_size, 1)
assert_correct_output_shape(w, batch_size)
# testing correct output
torch.testing.assert_allclose(w, event_rate(h).log(), rtol=1e-3, atol=1e-3)
test_log_event_rate(h)
# -
# ### Log Event Time Function
# +
#export
def log_event_time(h):
"""
Given the predicted conditional hazard rate, this function estimates
the log probability that the event occurs at time t.
*input*:
* `h`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`, as this is most amenable to use in training neural nets with pytorch.
_output_:
* `p`:
- type: `torch.tensor`
- estimated log probability of event at time t.
- note: `p.shape == (batch_size, 1)`
"""
assert_correct_input_shape(h)
p = torch.log(h[:, -1, :]) + log_survival_rate(h[:, :-1, :])
return p
# -
# example
log_event_time(h)
# +
#hide
# log event time tests
def test_log_event_time(h):
# shape should be 3-d
with pytest.raises(ValueError):
p = log_event_time(h[0, :, :])
batch_size, length, _ = h.shape
p = log_event_time(h)
# output should have shape (batch_size, 1)
assert_correct_output_shape(p, batch_size)
# testing correct output
torch.testing.assert_allclose(p, event_time(h).log(), rtol=1e-3, atol=1e-3)
test_log_event_time(h)
# -
# ## Loss Functions
#
# Now, we define the transform these generic survival analysis functions into loss functions that can be automatically differentiated by PyTorch, in order to train a Deep Recurrent Survival Analysis model.
#
#
# We make a few notes below:
#
# 1. The functions below adhere to the common pattern used across all of [`PyTorch`'s loss functions](https://pytorch.org/docs/stable/nn.functional.html#loss-functions), which is to take two arguments named `input` and `target`. We note, however, that due to the nature of this survival data, the target is inherent to the data structure and thus unnecessary.
#
# 2. The original DRSA paper defines 3 loss functions, 2 of which are directed towards uncensored data, and 1 of which applies to censored data. This library's focus is on DRSA models using only uncensored data, so those are the only lossed we'll be defining.
# ### Event Time Loss
# +
#export
def event_time_loss(input, target=None):
"""
Loss function applied to uncensored data in order
to optimize the PDF of the true event time, z
input:
* `input`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`
* `target`:
- unused, only present to mimic pytorch loss functions
output:
* `evt_loss`:
- type: `torch.tensor`
- Loss associated with how wrong each predicted probability was at each time step
"""
assert_correct_input_shape(input)
evt_loss = -log_event_time(input).mean(dim=0).squeeze()
return evt_loss
# -
# example
event_time_loss(h)
# +
#hide
# event time loss tests
def test_event_time_loss(input, target=None):
evt_loss = event_time_loss(input)
# testing correct output
torch.testing.assert_allclose(evt_loss, torch.tensor(1.5171), rtol=1e-3, atol=1e-3)
test_event_time_loss(h)
# -
# ### Event Rate Loss
# +
#export
def event_rate_loss(input, target=None):
"""
Loss function applied to uncensored data in order
to optimize the CDF of the true event time, z
input:
* `input`:
- type: `torch.tensor`,
- predicted conditional hazard rate, at each observed time step.
- note: `h.shape == (batch size, 1, 1)`
* `target`:
- unused, only present to mimic pytorch loss functions
output:
* `evr_loss`:
- type: `torch.tensor`
- Loss associated with how cumulative predicted probabilities differ from the ground truth labels.
"""
assert_correct_input_shape(input)
evr_loss = -log_event_rate(input).mean(dim=0).squeeze()
return evr_loss
# -
# example
event_rate_loss(h)
# +
#hide
# event rate loss tests
def test_event_rate_loss(input, target=None):
evr_loss = event_rate_loss(input)
# testing correct output
torch.testing.assert_allclose(evr_loss, torch.tensor(0.0319), rtol=1e-3, atol=1e-3)
test_event_rate_loss(h)
| notebooks/00_functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
df_morning = pd.read_csv('data/gang_morning.csv')
df_morning
# -
# %run -i "runTCDF.py" --help
"""Run TCDF"""
# %matplotlib inline
# %run -i "runTCDF.py" --data data/gang_morning.csv --cuda --significance 0.99 --hidden_layers 2 --kernel_size 2 --log_interval 50 --epochs 100 --plot --dilation_coefficient 2
"""Run TCDF"""
# %matplotlib inline
# %run -i "runTCDF.py" --data data/gang_morning.csv --cuda --significance 0.8 --hidden_layers 2 --kernel_size 2 --log_interval 50 --epochs 100 --plot --dilation_coefficient 2
"""Run TCDF"""
# %matplotlib inline
# %run -i "runTCDF.py" --data data/gang_morning.csv --cuda --significance 0.8 --hidden_layers 2 --kernel_size 3 --log_interval 500 --epochs 1000 --plot --dilation_coefficient 3
"""Run TCDF"""
# %matplotlib inline
# %run -i "runTCDF.py" --data data/gang_morning.csv --cuda --significance 0.8 --hidden_layers 2 --kernel_size 2 --log_interval 500 --epochs 1000 --plot --dilation_coefficient 2
| cs224w/TCDF-test1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# importacion general de librerias y de visualizacion (matplotlib y seaborn)
import pandas as pd
import numpy as np
import random
import re
pd.options.display.float_format = '{:20,.2f}'.format # suprimimos la notacion cientifica en los outputs
import warnings
warnings.filterwarnings('ignore')
# -
train_data = pd.read_csv('~/Documents/Datos/DataSets/TP2/train_featured.csv')
test_data = pd.read_csv('~/Documents/Datos/DataSets/TP2/test_featured.csv')
test_data['clean_text'].fillna("", inplace=True)
hashtag_dic = {}
for ls in train_data.hashtags.str.split():
try:
for hashtag in ls:
try:
hashtag_dic[hashtag] += 1
except KeyError:
hashtag_dic[hashtag] = 1
except TypeError:
pass
hashtag_dic = {k: v for k, v in sorted(hashtag_dic.items(), key=lambda item: item[1], reverse=True)}
hashtag_dic = {k: v for k, v in hashtag_dic.items() if v >= 10}
hashtag_dic
for x in hashtag_dic.keys():
train_data[x] = 0
test_data[x] = 0
def one_hot_hashtags(row):
try:
for x in row.hashtags.split():
if x in row:
row[x] = 1
except AttributeError:
pass
return row
train_data = train_data.apply(one_hot_hashtags, axis=1)
train_data.head()
test_data = test_data.apply(one_hot_hashtags, axis=1)
test_data.head()
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics import accuracy_score
import xgboost as xgb
from sklearn.preprocessing import OneHotEncoder
train, train_test = train_test_split(train_data, test_size=0.5)
text_clf = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
text_clf = text_clf.fit(train.clean_text, train.target_label)
train_predicted = text_clf.predict(train_test.clean_text)
train_predicted_proba = text_clf.predict_proba(train_test.clean_text)
np.mean(train_predicted == train_test.target_label)
test_predicted = text_clf.predict(test_data.clean_text)
test_predicted_proba = text_clf.predict_proba(test_data.clean_text)
train_test['NB_target_proba_not'] = train_predicted_proba.T[0]
test_data['NB_target_proba_not'] = test_predicted_proba.T[0]
text_clf_svm = Pipeline([('vect', CountVectorizer()),
('tfidf', TfidfTransformer()),
('clf-svm', SGDClassifier(loss='log', penalty='l2',
alpha=1e-3, random_state=42)),
])
text_clf_svm = text_clf_svm.fit(train.clean_text, train.target_label)
train_predicted = text_clf_svm.predict(train_test.clean_text)
train_predicted_proba = text_clf_svm.predict_proba(train_test.clean_text)
np.mean(train_predicted == train_test.target_label)
test_predicted = text_clf_svm.predict(test_data.clean_text)
test_predicted_proba = text_clf_svm.predict_proba(test_data.clean_text)
train_test['SVM_target_proba_not'] = train_predicted_proba.T[0]
test_data['SVM_target_proba_not'] = test_predicted_proba.T[0]
train_test.drop(["id_original","keyword_original","location_original","text_original","hashtags","labels","clean_text"], axis=1, inplace=True)
real_test_data = test_data.drop(["id_original","keyword_original","location_original","text_original","hashtags","labels","clean_text"], axis=1)
train, test = train_test_split(train_test, test_size=0.2)
model_xgb = xgb.XGBClassifier(n_estimators=300, colsample_bytree=0.5, learning_rate=0.1, max_depth=11)
model_xgb.fit(train.drop(['target_label'], axis=1), train.target_label)
test_prediction = model_xgb.predict(test.drop(['target_label'], axis=1))
print("Accuracy score: %f" % (accuracy_score(test.target_label, test_prediction)))
pd.DataFrame(model_xgb.feature_importances_, index=train.drop(['target_label'], axis=1).columns, columns=["importancia"]).\
sort_values(by="importancia",ascending=False)
model_xgb = xgb.XGBClassifier(n_estimators=300, colsample_bytree=0.5, learning_rate=0.1, max_depth=11)
model_xgb.fit(train_test.drop(['target_label'], axis=1), train_test.target_label)
real_test_prediction = model_xgb.predict(real_test_data)
test_data['target'] = real_test_prediction
test_data[['id_original', 'target']].rename(columns={'id_original': 'id'}).to_csv('~/Documents/Datos/DataSets/TP2/res_XGB_1.csv', index=False)
| TP2/Alejo/3_XGBOOST_hashtag_encoding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="9MM5cpotg1r-"
# This is a companion notebook for the book [Deep Learning with Python, Second Edition](https://www.manning.com/books/deep-learning-with-python-second-edition?a_aid=keras&a_bid=76564dff). For readability, it only contains runnable code blocks and section titles, and omits everything else in the book: text paragraphs, figures, and pseudocode.
#
# **If you want to be able to follow what's going on, I recommend reading the notebook side by side with your copy of the book.**
#
# This notebook was generated for TensorFlow 2.6.
# + [markdown] id="xHRSsgpVg1sA"
# # The mathematical building blocks of neural networks
# + [markdown] id="JoRZ6c6Rg1sA"
# ## A first look at a neural network
# + [markdown] id="0W1ivKy5g1sA"
# **Loading the MNIST dataset in Keras**
# + id="FL6r2x8bg1sA" outputId="be74a974-df32-4684-bb73-960e18dbb9a6" colab={"base_uri": "https://localhost:8080/"}
from tensorflow.keras.datasets import mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# + id="JEq08joxg1sB" outputId="34aa852d-2702-43a0-8a64-4bbdae07b7f7" colab={"base_uri": "https://localhost:8080/"}
train_images.shape
# + id="SECoEKbGg1sB" outputId="a9c38f57-6db9-4419-cbc1-50dc176c4680" colab={"base_uri": "https://localhost:8080/"}
len(train_labels)
# + id="Q2tXQUB0g1sB" outputId="7f1ca2b2-e070-460a-c3e7-a8d8838c676c" colab={"base_uri": "https://localhost:8080/"}
train_labels
# + id="Xdl3Cskrg1sB" outputId="b44749f9-a1be-410f-c17f-0ad7ccbc204c" colab={"base_uri": "https://localhost:8080/"}
test_images.shape
# + id="fwrznngsg1sB" outputId="c8078183-85da-4381-9d9a-c34b973de268" colab={"base_uri": "https://localhost:8080/"}
len(test_labels)
# + id="FDCdaXOsg1sC" outputId="e15a6949-9873-48d5-c683-cf18470d337f" colab={"base_uri": "https://localhost:8080/"}
test_labels
# + [markdown] id="xe_6LWGMg1sC"
# **The network architecture**
# + id="UOBHD0uAg1sC"
from tensorflow import keras
from tensorflow.keras import layers
model = keras.Sequential([
layers.Dense(512, activation="relu"),
layers.Dense(10, activation="softmax")
])
# + [markdown] id="J5IgD_n4g1sC"
# **The compilation step**
# + id="giglsU_qg1sD"
model.compile(optimizer="rmsprop",
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# + [markdown] id="4IsUqOSOg1sD"
# **Preparing the image data**
# + id="Oo7z79kgg1sD"
train_images = train_images.reshape((60000, 28 * 28))
train_images = train_images.astype("float32") / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype("float32") / 255
# + [markdown] id="oP60Brytg1sE"
# **"Fitting" the model**
# + id="8dQDQbv9g1sE" outputId="e51bb315-bb2f-4459-95da-80c46c170674" colab={"base_uri": "https://localhost:8080/"}
model.fit(train_images, train_labels, epochs=5, batch_size=128)
# + [markdown] id="-_QOe66Kg1sE"
# **Using the model to make predictions**
# + id="eXb0DCq8g1sE" outputId="95ebb609-6341-44b8-c615-83eaa29f9f87" colab={"base_uri": "https://localhost:8080/"}
test_digits = test_images[0:10]
predictions = model.predict(test_digits)
predictions[0]
# + id="cZMXjCfpg1sE" outputId="1b6aa813-cc4d-4c50-b9c9-25ffe71c6df5" colab={"base_uri": "https://localhost:8080/"}
predictions[0].argmax()
# + id="eMG-mOMLg1sF" outputId="1fec9ff3-0273-4592-947f-d7445ac5f114" colab={"base_uri": "https://localhost:8080/"}
predictions[0][7]
# + id="bKaFCXjrg1sF" outputId="22c7c397-4335-4670-adc7-b7d80ba64cb1" colab={"base_uri": "https://localhost:8080/"}
test_labels[0]
# + [markdown] id="xP30N8jtg1sF"
# **Evaluating the model on new data**
# + id="nGd1CAv_g1sF" outputId="d91559e4-3479-4158-a314-0c985e9ea554" colab={"base_uri": "https://localhost:8080/"}
test_loss, test_acc = model.evaluate(test_images, test_labels)
print(f"test_acc: {test_acc}")
# + [markdown] id="5EbftRpQg1sF"
# ## Data representations for neural networks
# + [markdown] id="GO06fP6kg1sF"
# ### Scalars (rank-0 tensors)
# + id="rs4rqL8tg1sF" outputId="700e287c-0927-48d7-dd44-300ec0a93e6b" colab={"base_uri": "https://localhost:8080/"}
import numpy as np
x = np.array(12)
x
# + id="iTCEVo_Mg1sG" outputId="a3fc79bc-960e-436d-8b23-4abf5decd108" colab={"base_uri": "https://localhost:8080/"}
x.ndim
# + [markdown] id="j1tuWK_Hg1sG"
# ### Vectors (rank-1 tensors)
# + id="JtGbpcWug1sH" outputId="311f154e-08cb-4512-b9d8-868694e29622" colab={"base_uri": "https://localhost:8080/"}
x = np.array([12, 3, 6, 14, 7])
x
# + id="Hb-kh3Eyg1sI" outputId="f91d1a36-d6bf-4f68-860e-ce3b5c5ee36e" colab={"base_uri": "https://localhost:8080/"}
x.ndim
# + [markdown] id="ikbiXQzGg1sJ"
# ### Matrices (rank-2 tensors)
# + id="CbtqYd5Hg1sK" outputId="6b2350ea-87cf-4870-d1fd-9be2dc83dabb" colab={"base_uri": "https://localhost:8080/"}
x = np.array([[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]])
x.ndim
# + [markdown] id="okjVS886g1sK"
# ### Rank-3 and higher-rank tensors
# + id="RFyQ1oXEg1sL"
x = np.array([[[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]],
[[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]],
[[5, 78, 2, 34, 0],
[6, 79, 3, 35, 1],
[7, 80, 4, 36, 2]]])
x.ndim
# + [markdown] id="21P4wyIwg1sL"
# ### Key attributes
# + id="m3xuh4Wsg1sL"
from tensorflow.keras.datasets import mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
# + id="7iONXsH4g1sL" outputId="8dcd24e1-b8e2-42e1-a8f8-6be7aaa3ddf8" colab={"base_uri": "https://localhost:8080/"}
train_images.ndim
# + id="rWiEkEQsg1sL" outputId="59d885df-3cbd-44fa-b8ee-18f51af101e1" colab={"base_uri": "https://localhost:8080/"}
train_images.shape
# + id="sNjmymS6g1sM" outputId="702ee372-b683-4af7-d55a-6999be949f67" colab={"base_uri": "https://localhost:8080/"}
train_images.dtype
# + [markdown] id="UoLRnasBg1sM"
# **Displaying the fourth digit**
# + id="aYBe5JyYg1sM" outputId="841838d7-5710-403c-e2c8-a738f65d798b" colab={"base_uri": "https://localhost:8080/", "height": 265}
import matplotlib.pyplot as plt
digit = train_images[4]
plt.imshow(digit, cmap=plt.cm.binary)
plt.show()
# + id="kJt34pzNg1sN" outputId="6e292759-c187-4cb1-e188-2a36d2ddf55d" colab={"base_uri": "https://localhost:8080/"}
train_labels[4]
# + [markdown] id="DTp-qzrBg1sN"
# ### Manipulating tensors in NumPy
# + id="yBU9lQ7Lg1sN" outputId="b4abb10c-5ec1-4c1f-cfc0-356fcb7eeb42" colab={"base_uri": "https://localhost:8080/"}
my_slice = train_images[10:100]
my_slice.shape
# + id="kavfm75tg1sN" outputId="7f6c6c1c-f07a-4445-9205-11b617a7c6c3" colab={"base_uri": "https://localhost:8080/"}
my_slice = train_images[10:100, :, :]
my_slice.shape
# + id="b6Xo7_-bg1sN" outputId="676e0c04-6a08-43f3-99e7-05fadd083d96" colab={"base_uri": "https://localhost:8080/"}
my_slice = train_images[10:100, 0:28, 0:28]
my_slice.shape
# + id="LTaORavAg1sN"
my_slice = train_images[:, 14:, 14:]
# + id="i_cGGmErg1sN"
my_slice = train_images[:, 7:-7, 7:-7]
# + [markdown] id="HV4Qhp3_g1sN"
# ### The notion of data batches
# + id="fD1128MUg1sN"
batch = train_images[:128]
# + id="zl7H5uSog1sO"
batch = train_images[128:256]
# + id="v_MPAAWeg1sO"
n = 3
batch = train_images[128 * n:128 * (n + 1)]
# + [markdown] id="8DPy9negg1sO"
# ### Real-world examples of data tensors
# + [markdown] id="ZOxUABclg1sO"
# ### Vector data
# + [markdown] id="aWgw5kVtg1sO"
# ### Timeseries data or sequence data
# + [markdown] id="ykFV1IqTg1sO"
# ### Image data
# + [markdown] id="3ITS9y2Gg1sO"
# ### Video data
# + [markdown] id="tjIrKTDag1sO"
# ## The gears of neural networks: tensor operations
# + [markdown] id="1xVBrrhPg1sO"
# ### Element-wise operations
# + id="Z9zlNSYfg1sO"
def naive_relu(x):
assert len(x.shape) == 2
x = x.copy()
for i in range(x.shape[0]):
for j in range(x.shape[1]):
x[i, j] = max(x[i, j], 0)
return x
# + id="JGgsvXU9g1sO"
def naive_add(x, y):
assert len(x.shape) == 2
assert x.shape == y.shape
x = x.copy()
for i in range(x.shape[0]):
for j in range(x.shape[1]):
x[i, j] += y[i, j]
return x
# + id="vI81x3Qfg1sP" outputId="7bc507ad-9125-4613-b55d-f7b67f9eaf9b" colab={"base_uri": "https://localhost:8080/"}
import time
x = np.random.random((20, 100))
y = np.random.random((20, 100))
t0 = time.time()
for _ in range(1000):
z = x + y
z = np.maximum(z, 0.)
print("Took: {0:.2f} s".format(time.time() - t0))
# + id="CN9alajNg1sP" outputId="1b0b06e2-5927-4efd-84aa-39a73c011e9c" colab={"base_uri": "https://localhost:8080/"}
t0 = time.time()
for _ in range(1000):
z = naive_add(x, y)
z = naive_relu(z)
print("Took: {0:.2f} s".format(time.time() - t0))
# + [markdown] id="oMY_ah5Jg1sP"
# ### Broadcasting
# + id="PHqE9hdXg1sP"
import numpy as np
X = np.random.random((32, 10))
y = np.random.random((10,))
# + id="jaTTuO6wg1sP"
y = np.expand_dims(y, axis=0)
# + id="eFJROHrPg1sP"
Y = np.concatenate([y] * 32, axis=0)
# + id="-KyT8wvKg1sP"
def naive_add_matrix_and_vector(x, y):
assert len(x.shape) == 2
assert len(y.shape) == 1
assert x.shape[1] == y.shape[0]
x = x.copy()
for i in range(x.shape[0]):
for j in range(x.shape[1]):
x[i, j] += y[j]
return x
# + id="Y9N0MKTyg1sP"
import numpy as np
x = np.random.random((64, 3, 32, 10))
y = np.random.random((32, 10))
z = np.maximum(x, y)
# + [markdown] id="2xgKf01hg1sP"
# ### Tensor product
# + id="FpdBnpdIg1sP"
x = np.random.random((32,))
y = np.random.random((32,))
z = np.dot(x, y)
# + id="0mVtPwx2g1sP"
def naive_vector_dot(x, y):
assert len(x.shape) == 1
assert len(y.shape) == 1
assert x.shape[0] == y.shape[0]
z = 0.
for i in range(x.shape[0]):
z += x[i] * y[i]
return z
# + id="6UCIw5mDg1sQ"
def naive_matrix_vector_dot(x, y):
assert len(x.shape) == 2
assert len(y.shape) == 1
assert x.shape[1] == y.shape[0]
z = np.zeros(x.shape[0])
for i in range(x.shape[0]):
for j in range(x.shape[1]):
z[i] += x[i, j] * y[j]
return z
# + id="DiL4hp1ug1sQ"
def naive_matrix_vector_dot(x, y):
z = np.zeros(x.shape[0])
for i in range(x.shape[0]):
z[i] = naive_vector_dot(x[i, :], y)
return z
# + id="voGFvayCg1sQ"
def naive_matrix_dot(x, y):
assert len(x.shape) == 2
assert len(y.shape) == 2
assert x.shape[1] == y.shape[0]
z = np.zeros((x.shape[0], y.shape[1]))
for i in range(x.shape[0]):
for j in range(y.shape[1]):
row_x = x[i, :]
column_y = y[:, j]
z[i, j] = naive_vector_dot(row_x, column_y)
return z
# + [markdown] id="87s1tOlcg1sQ"
# ### Tensor reshaping
# + id="vJzQ92sTg1sQ"
train_images = train_images.reshape((60000, 28 * 28))
# + id="VsKlMohCg1sQ" outputId="bb2d2551-6fdb-4ffb-d95d-dd1127bae03b" colab={"base_uri": "https://localhost:8080/"}
x = np.array([[0., 1.],
[2., 3.],
[4., 5.]])
x.shape
# + id="lDntXx80g1sQ" outputId="b1f257d7-1aa7-4096-d23e-119943c551c0" colab={"base_uri": "https://localhost:8080/"}
x = x.reshape((6, 1))
x
# + id="DbRnGHY1g1sQ" outputId="4fce4e8d-98f6-4d72-faea-af302a28c528" colab={"base_uri": "https://localhost:8080/"}
x = np.zeros((300, 20))
x = np.transpose(x)
x.shape
# + [markdown] id="4_yeezrwg1sQ"
# ### Geometric interpretation of tensor operations
# + [markdown] id="j8JZSdf7g1sQ"
# ### A geometric interpretation of deep learning
# + [markdown] id="-KqYgF79g1sQ"
# ## The engine of neural networks: gradient-based optimization
# + [markdown] id="x9wCEdRNg1sQ"
# ### What's a derivative?
# + [markdown] id="dhTiQKdsg1sQ"
# ### Derivative of a tensor operation: the gradient
# + [markdown] id="6tOxhhihg1sQ"
# ### Stochastic gradient descent
# + [markdown] id="v-aqPU77g1sQ"
# ### Chaining derivatives: The Backpropagation algorithm
# + [markdown] id="dB2y9Mubg1sQ"
# #### The chain rule
# + [markdown] id="ccabX_UNg1sR"
# #### Automatic differentiation with computation graphs
# + [markdown] id="23RssObFg1sR"
# #### The gradient tape in TensorFlow
# + id="iRg1Lty4g1sR"
import tensorflow as tf
x = tf.Variable(0.)
with tf.GradientTape() as tape:
y = 2 * x + 3
grad_of_y_wrt_x = tape.gradient(y, x)
# + id="svzSWzNhg1sR"
x = tf.Variable(tf.random.uniform((2, 2)))
with tf.GradientTape() as tape:
y = 2 * x + 3
grad_of_y_wrt_x = tape.gradient(y, x)
# + id="TBANtOG8g1sR"
W = tf.Variable(tf.random.uniform((2, 2)))
b = tf.Variable(tf.zeros((2,)))
x = tf.random.uniform((2, 2))
with tf.GradientTape() as tape:
y = tf.matmul(x, W) + b
grad_of_y_wrt_W_and_b = tape.gradient(y, [W, b])
# + [markdown] id="eIY5M7lSg1sR"
# ## Looking back at our first example
# + id="s4CJCYpSg1sR"
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images.reshape((60000, 28 * 28))
train_images = train_images.astype("float32") / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype("float32") / 255
# + id="z3ZPT1d9g1sR"
model = keras.Sequential([
layers.Dense(512, activation="relu"),
layers.Dense(10, activation="softmax")
])
# + id="8zhTAwsug1sR"
model.compile(optimizer="rmsprop",
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# + id="zZuE8EPZg1sR" outputId="f88392da-1e25-4b80-f348-bb4ad075d87a" colab={"base_uri": "https://localhost:8080/"}
model.fit(train_images, train_labels, epochs=5, batch_size=128)
# + [markdown] id="vX9_wNVlg1sR"
# ### Reimplementing our first example from scratch in TensorFlow
# + [markdown] id="rFXnMgl8g1sR"
# #### A simple Dense class
# + id="A6W1NxBHg1sR"
import tensorflow as tf
class NaiveDense:
def __init__(self, input_size, output_size, activation):
self.activation = activation
w_shape = (input_size, output_size)
w_initial_value = tf.random.uniform(w_shape, minval=0, maxval=1e-1)
self.W = tf.Variable(w_initial_value)
b_shape = (output_size,)
b_initial_value = tf.zeros(b_shape)
self.b = tf.Variable(b_initial_value)
def __call__(self, inputs):
return self.activation(tf.matmul(inputs, self.W) + self.b)
@property
def weights(self):
return [self.W, self.b]
# + [markdown] id="f4HRbJaig1sS"
# #### A simple Sequential class
# + id="pCH86zn1g1sS"
class NaiveSequential:
def __init__(self, layers):
self.layers = layers
def __call__(self, inputs):
x = inputs
for layer in self.layers:
x = layer(x)
return x
@property
def weights(self):
weights = []
for layer in self.layers:
weights += layer.weights
return weights
# + id="l53RG-bWg1sS"
model = NaiveSequential([
NaiveDense(input_size=28 * 28, output_size=512, activation=tf.nn.relu),
NaiveDense(input_size=512, output_size=10, activation=tf.nn.softmax)
])
assert len(model.weights) == 4
# + [markdown] id="A1PupGA1g1sS"
# #### A batch generator
# + id="MC21aCDeg1sS"
import math
class BatchGenerator:
def __init__(self, images, labels, batch_size=128):
assert len(images) == len(labels)
self.index = 0
self.images = images
self.labels = labels
self.batch_size = batch_size
self.num_batches = math.ceil(len(images) / batch_size)
def next(self):
images = self.images[self.index : self.index + self.batch_size]
labels = self.labels[self.index : self.index + self.batch_size]
self.index += self.batch_size
return images, labels
# + [markdown] id="S5EbKuHbg1sS"
# ### Running one training step
# + id="vv_e9JOAg1sS"
def one_training_step(model, images_batch, labels_batch):
with tf.GradientTape() as tape:
predictions = model(images_batch)
per_sample_losses = tf.keras.losses.sparse_categorical_crossentropy(
labels_batch, predictions)
average_loss = tf.reduce_mean(per_sample_losses)
gradients = tape.gradient(average_loss, model.weights)
update_weights(gradients, model.weights)
return average_loss
# + id="hFh4Mu1ug1sS"
learning_rate = 1e-3
def update_weights(gradients, weights):
for g, w in zip(gradients, weights):
w.assign_sub(g * learning_rate)
# + id="honLPZDng1sS"
from tensorflow.keras import optimizers
optimizer = optimizers.SGD(learning_rate=1e-3)
def update_weights(gradients, weights):
optimizer.apply_gradients(zip(gradients, weights))
# + [markdown] id="aov4CNQ4g1sS"
# ### The full training loop
# + id="ckJHr3_vg1sS"
def fit(model, images, labels, epochs, batch_size=128):
for epoch_counter in range(epochs):
print(f"Epoch {epoch_counter}")
batch_generator = BatchGenerator(images, labels)
for batch_counter in range(batch_generator.num_batches):
images_batch, labels_batch = batch_generator.next()
loss = one_training_step(model, images_batch, labels_batch)
if batch_counter % 100 == 0:
print(f"loss at batch {batch_counter}: {loss:.2f}")
# + id="j_VHCs1hg1sS" outputId="59b20f5c-e5bf-4578-d371-494b2834c05f" colab={"base_uri": "https://localhost:8080/"}
from tensorflow.keras.datasets import mnist
(train_images, train_labels), (test_images, test_labels) = mnist.load_data()
train_images = train_images.reshape((60000, 28 * 28))
train_images = train_images.astype("float32") / 255
test_images = test_images.reshape((10000, 28 * 28))
test_images = test_images.astype("float32") / 255
fit(model, train_images, train_labels, epochs=10, batch_size=128)
# + [markdown] id="xq6yXBw_g1sS"
# ### Evaluating the model
# + id="xoQorv2_g1sS" outputId="4a77840f-1a84-4e4c-bab5-8035e2b0f394" colab={"base_uri": "https://localhost:8080/"}
predictions = model(test_images)
predictions = predictions.numpy()
predicted_labels = np.argmax(predictions, axis=1)
matches = predicted_labels == test_labels
print(f"accuracy: {matches.mean():.2f}")
# + [markdown] id="2TMivNIyg1sT"
# ## Summary
| chapter02_mathematical-building-blocks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="oW0JBwpOCSGs"
"""
Reconstructiong MNIST images using a Deep Convolution GAN
This is the most challenging thing i've done till the date 22/09/2018
"""
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="HCaMSn7yCk8v"
# creating minibatches
def mini_batch(X, size):
# generate random integers of shape size*1 and in range 0 to len(X)
# treat x_bat as indices and return values from X at those indices
x_bat = X[np.random.randint(len(X), size = (size,1))]
x_bat = x_bat.reshape(size, 28, 28, 1)
return x_bat
# + colab={"base_uri": "https://localhost:8080/", "height": 522} colab_type="code" id="SK30bCdIClYW" outputId="2ec322b7-a016-46d0-f7b1-22120ea2af20"
# importing dataset
mnist = tf.keras.datasets.mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
del(y_train, y_test)
fig = plt.figure(figsize=(8,8))
col = 4
row = 4
j = 0
for i in range(1, row*col+1):
fig.add_subplot(row, col, i)
plt.imshow(x_train[j+100])
j+=1
plt.show()
# + colab={} colab_type="code" id="J-EDTCfjCmza"
tf.reset_default_graph()
# input and labels placeholders
fake_x = tf.placeholder(tf.float32, [None, 4,4,1])
real_x = tf.placeholder(tf.float32, [None, 28,28,1])
# generator network weights
gen_wt = {'w_c1': tf.get_variable('w_c1', [3,3,1,64], initializer=tf.contrib.layers.xavier_initializer()),
'w_c2': tf.get_variable('w_c2', [3,3,64,128], initializer=tf.contrib.layers.xavier_initializer()),
'w_c3': tf.get_variable('w_c3', [3,3,128,128], initializer=tf.contrib.layers.xavier_initializer()),
'w_c4': tf.get_variable('w_c4', [3,3,128,1], initializer=tf.contrib.layers.xavier_initializer())
}
# discriminator network weights
dis_wt = {'d_c1': tf.get_variable('d_c1', [3,3,1,32], initializer=tf.contrib.layers.xavier_initializer()),
'd_c2': tf.get_variable('d_c2', [3,3,32,64], initializer=tf.contrib.layers.xavier_initializer()),
'd_c3': tf.get_variable('d_c3', [3,3,64,64], initializer=tf.contrib.layers.xavier_initializer()),
'd_c4': tf.get_variable('d_c4', [3,3,64,1], initializer=tf.contrib.layers.xavier_initializer())
}
# + colab={} colab_type="code" id="l7sVLZzeCooX"
# generator network
with tf.device('/gpu:0'):
def generator(noise):
# changed relu to tanh
conv_1 = tf.nn.tanh(tf.nn.conv2d(noise, gen_wt['w_c1'], strides=[1,1,1,1], padding='SAME'))
up_scale_1 = tf.image.resize_images(conv_1, [14,14], method = tf.image.ResizeMethod.BILINEAR)
conv_2 = tf.nn.tanh(tf.nn.conv2d(up_scale_1, gen_wt['w_c2'], strides=[1,1,1,1], padding='SAME'))
drop_2 = tf.nn.dropout(conv_2, 0.8)
up_scale_2 = tf.image.resize_images(drop_2, [28,28], method = tf.image.ResizeMethod.BILINEAR)
conv_3 = tf.nn.tanh(tf.nn.conv2d(up_scale_2, gen_wt['w_c3'], strides=[1,1,1,1], padding='SAME'))
generator = tf.nn.tanh(tf.nn.conv2d(conv_3, gen_wt['w_c4'], strides=[1,1,1,1], padding='SAME'))
return generator
# discriminator network
def discriminator(imgs):
## changed activation from leaky relu to tanh
## changed avg_pool to strided convolution
c1 = tf.nn.tanh(tf.nn.conv2d(imgs, dis_wt['d_c1'], strides=[1,1,1,1], padding='SAME'))
# m1 = tf.nn.avg_pool(c1, ksize=[1,2,2,1], strides=[1,2,2,1], padding='SAME')
c2 = tf.nn.tanh(tf.nn.conv2d(c1, dis_wt['d_c2'], strides=[1,2,2,1], padding='SAME'))
# m2 = tf.nn.avg_pool(c2, ksize=[1,2,2,1], strides=[1,2,2,1], padding='SAME')
d2 = tf.nn.dropout(c2, 0.8)
c3 = tf.nn.tanh(tf.nn.conv2d(d2, dis_wt['d_c3'], strides=[1,2,2,1], padding='SAME'))
#m3 = tf.nn.avg_pool(c3, ksize=[1,2,2,1], strides=[1,2,2,1], padding='SAME')
d3 = tf.nn.dropout(c3, 0.8)
f4 = tf.layers.flatten(d3)
d4 = tf.layers.dense(f4, 16, activation='sigmoid', trainable=True)
discriminator = tf.layers.dense(d4, 1, activation='sigmoid', trainable=True)
return discriminator
# loss and optimization
# generate fake images from noise
g_output = generator(fake_x)
# use the discriminator to differentiate between real and fake images
real_prob = discriminator(real_x)
fake_prob = discriminator(g_output)
# loss function for discriminator
d_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=real_prob, labels=tf.zeros_like(real_prob)) +
tf.nn.sigmoid_cross_entropy_with_logits(logits=fake_prob, labels=tf.ones_like(fake_prob)))
# loss function for generator
g_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits=fake_prob, labels=tf.ones_like(fake_prob)))
# optimizer for discriminator
d_ops = tf.train.AdamOptimizer(0.0001).minimize(d_loss)
# optimizer for generator
g_ops = tf.train.AdamOptimizer(0.0001).minimize(g_loss)
# + colab={} colab_type="code" id="sxmhf8W9CqTz"
# hyperparameters
sample_size = 512
epochs = 1000
s = tf.Session(config=tf.ConfigProto(log_device_placement=True))
s.run(tf.global_variables_initializer())
g_l = []
d_l = []
# here goes the training
for i in range(epochs):
# create noise sample
noise = np.random.normal(0, 0.1,[sample_size, fake_x.shape[1], fake_x.shape[2], fake_x.shape[3]])
# create sample of real images
real_imgs = mini_batch(x_train, sample_size)/255.0
for j in range(1):
# training the discriminator
s.run(d_ops, {real_x: real_imgs, fake_x: noise})
d_l.append(s.run(d_loss, {real_x: real_imgs, fake_x: noise}))
# training the generator
s.run(g_ops, {fake_x: noise})
g_l.append(s.run(g_loss, {fake_x: noise}))
# + colab={"base_uri": "https://localhost:8080/", "height": 361} colab_type="code" id="vMsCvA6BeMfA" outputId="a74507b1-56ca-4e26-b068-0379e17a90ff"
# plotting loss
plt.plot(range(0,len(d_l)), d_l, 'g', label='discriminator loss')
plt.plot(range(0,len(g_l)), g_l, 'b', label='generator loss')
plt.legend(loc='upper right')
plt.xlabel('Iteration')
plt.ylabel('Loss')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 485} colab_type="code" id="gprTpLhWRWLU" outputId="6dfd8fc0-1990-4b72-c0a7-6600a85e3684"
pred = s.run(g_output, {fake_x: np.random.normal(0, 0.1,[sample_size,4,4,1])}).reshape(sample_size, 28,28)
fig2 = plt.figure(figsize=(8,8))
col = 4
row = 4
j = 0
for i in range(1, row*col+1):
fig2.add_subplot(row, col, i)
plt.imshow(pred[j])
j+=1
plt.show()
# + [markdown] colab_type="text" id="TaxVilW8w5Tq"
# ### It's quite a challenging thing to train a GAN...
| DCGAN/MNIST_GAN_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dscohen75/DS-Unit-2-Linear-Models/blob/master/module3-ridge-regression/Copy_of_LS_DS_213_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vMEHeozLT1Pe" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 1, Module 3*
#
# ---
# + [markdown] colab_type="text" id="7IXUfiQ2UKj6"
# # Ridge Regression
#
# ## Assignment
#
# We're going back to our other **New York City** real estate dataset. Instead of predicting apartment rents, you'll predict property sales prices.
#
# But not just for condos in Tribeca...
#
# - [ ] Use a subset of the data where `BUILDING_CLASS_CATEGORY` == `'01 ONE FAMILY DWELLINGS'` and the sale price was more than 100 thousand and less than 2 million.
# - [ ] Do train/test split. Use data from January — March 2019 to train. Use data from April 2019 to test.
# - [ ] Do one-hot encoding of categorical features.
# - [ ] Do feature selection with `SelectKBest`.
# - [ ] Fit a ridge regression model with multiple features. Use the `normalize=True` parameter (or do [feature scaling](https://scikit-learn.org/stable/modules/preprocessing.html) beforehand — use the scaler's `fit_transform` method with the train set, and the scaler's `transform` method with the test set)
# - [ ] Get mean absolute error for the test set.
# - [ ] As always, commit your notebook to your fork of the GitHub repo.
#
# The [NYC Department of Finance](https://www1.nyc.gov/site/finance/taxes/property-rolling-sales-data.page) has a glossary of property sales terms and NYC Building Class Code Descriptions. The data comes from the [NYC OpenData](https://data.cityofnewyork.us/browse?q=NYC%20calendar%20sales) portal.
#
#
# ## Stretch Goals
#
# Don't worry, you aren't expected to do all these stretch goals! These are just ideas to consider and choose from.
#
# - [ ] Add your own stretch goal(s) !
# - [ ] Instead of `Ridge`, try `LinearRegression`. Depending on how many features you select, your errors will probably blow up! 💥
# - [ ] Instead of `Ridge`, try [`RidgeCV`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.RidgeCV.html).
# - [ ] Learn more about feature selection:
# - ["Permutation importance"](https://www.kaggle.com/dansbecker/permutation-importance)
# - [scikit-learn's User Guide for Feature Selection](https://scikit-learn.org/stable/modules/feature_selection.html)
# - [mlxtend](http://rasbt.github.io/mlxtend/) library
# - scikit-learn-contrib libraries: [boruta_py](https://github.com/scikit-learn-contrib/boruta_py) & [stability-selection](https://github.com/scikit-learn-contrib/stability-selection)
# - [_Feature Engineering and Selection_](http://www.feat.engineering/) by Kuhn & Johnson.
# - [ ] Try [statsmodels](https://www.statsmodels.org/stable/index.html) if you’re interested in more inferential statistical approach to linear regression and feature selection, looking at p values and 95% confidence intervals for the coefficients.
# - [ ] Read [_An Introduction to Statistical Learning_](http://faculty.marshall.usc.edu/gareth-james/ISL/ISLR%20Seventh%20Printing.pdf), Chapters 1-3, for more math & theory, but in an accessible, readable way.
# - [ ] Try [scikit-learn pipelines](https://scikit-learn.org/stable/modules/compose.html).
# + colab_type="code" id="o9eSnDYhUGD7" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# Ignore this Numpy warning when using Plotly Express:
# FutureWarning: Method .ptp is deprecated and will be removed in a future version. Use numpy.ptp instead.
import warnings
warnings.filterwarnings(action='ignore', category=FutureWarning, module='numpy')
# + colab_type="code" id="QJBD4ruICm1m" colab={}
import pandas as pd
import pandas_profiling
# Read New York City property sales data
df = pd.read_csv(DATA_PATH+'condos/NYC_Citywide_Rolling_Calendar_Sales.csv',
parse_dates=['SALE DATE'],
index_col='SALE DATE'
)
# Change column names: replace spaces with underscores
df.columns = [col.replace(' ', '_') for col in df]
# SALE_PRICE was read as strings.
# Remove symbols, convert to integer
df['SALE_PRICE'] = (
df['SALE_PRICE']
.str.replace('$','')
.str.replace('-','')
.str.replace(',','')
.astype(int)
)
# + id="PcCX0wP3T1Pm" colab_type="code" colab={}
# BOROUGH is a numeric column, but arguably should be a categorical feature,
# so convert it from a number to a string
df['BOROUGH'] = df['BOROUGH'].astype(str)
# + id="ghXKkA-pT1Pp" colab_type="code" colab={}
# Reduce cardinality for NEIGHBORHOOD feature
# Get a list of the top 10 neighborhoods
top10 = df['NEIGHBORHOOD'].value_counts()[:10].index
# At locations where the neighborhood is NOT in the top 10,
# replace the neighborhood with 'OTHER'
df.loc[~df['NEIGHBORHOOD'].isin(top10), 'NEIGHBORHOOD'] = 'OTHER'
# + id="eDu0jIHUT1Pr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="be8a8f70-12f9-43ff-ff03-9821065ce4b4"
top10
# + [markdown] id="ZTDmxAxlWyhM" colab_type="text"
# - Use a subset of the data where BUILDING_CLASS_CATEGORY == '01 ONE FAMILY DWELLINGS' and the sale price was more than 100 thousand and less than 2 million.
# - Do train/test split. Use data from January — March 2019 to train. Use data from April 2019 to test.
# - Do one-hot encoding of categorical features.
# - Do feature selection with SelectKBest.
# - Fit a ridge regression model with multiple features. Use the normalize=True parameter (or do feature scaling beforehand — use the scaler's fit_transform method with the train set, and the scaler's transform method with the test set)
# - Get mean absolute error for the test set.
# + id="DknArVX9XFpF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="4b3094f8-4f06-4304-c6a1-19effb29616b"
df.columns
# + id="TJxtHmXFYMSb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="163650e5-9690-48ee-fccf-025caa6c6d9f"
type(df['SALE_PRICE'])
# + id="YX448ix-YS_1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 476} outputId="077452ed-33f4-4322-d9db-d2b2befcb13a"
df.info()
# + id="Yr0OUWuPXIwp" colab_type="code" colab={}
onefam = df[df['BUILDING_CLASS_CATEGORY']=='01 ONE FAMILY DWELLINGS']
# + id="AW7gEJUtZ6iV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b94f8ed5-0fbd-4546-8380-4c5d3021ccd0"
onefam.shape
# + id="BuAyfyNMZpTi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="70c7b3aa-1279-4a56-db8a-52cfe101c0cf"
mask = (onefam['SALE_PRICE'] > 100000) & (onefam['SALE_PRICE'] < 2000000)
onefam = onefam.loc[mask]
onefam.shape
# + id="O3vJJ8Kam4lv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="64955b16-cd60-4f52-ad18-b864535585a8"
onefam.head()
# + id="Nkw9-mqfoFEF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="9fcd2ef6-296b-4388-a79b-d05b732ff9b6"
onefam['BOROUGH'].value_counts()
# + id="v12UU4gEoKjA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="75696c56-c8fc-4400-d0e2-8e134c6d64e5"
onefam['TAX_CLASS_AT_PRESENT'].value_counts()
# + id="RTY6qJiwod9O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 255} outputId="58d3ee63-4bfb-42bf-a9c7-f1829d6aa557"
onefam['BUILDING_CLASS_AT_PRESENT'].value_counts()
# + id="XeP8vAbZoqRw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="4eb247c5-28a3-4233-abe4-2032573f8438"
onefam['ADDRESS'].value_counts()
# + id="Y93k0KG1owT1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="eec4558c-7aa3-4ffc-88aa-3e21f4be68f0"
onefam['LAND_SQUARE_FEET'].value_counts()
# + id="16hMF-8Go8OB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="97b5de46-fff9-40a9-ee69-c068b4eea041"
onefam['BUILDING_CLASS_AT_TIME_OF_SALE'].value_counts()
# + id="R--w1t33pKST" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="a44468d6-5381-4457-c1c9-63a39e3d4f7c"
onefam['TAX_CLASS_AT_TIME_OF_SALE'].value_counts()
# + id="vRpiEW49rznJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="19922550-942b-44dd-de54-469830d7e961"
onefam['ZIP_CODE'].value_counts()
# + id="1yF2zpAIqA59" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="a6636651-068f-4e51-8ef3-81cec3e9248c"
onefam['LAND_SQUARE_FEET'].value_counts()
# + id="Y7imLCKyqWcV" colab_type="code" colab={}
# LAND_SQUARE_FEET was read as strings.
# Remove symbols, convert to integer
onefam['LAND_SQUARE_FEET'] = (
onefam['LAND_SQUARE_FEET'].str.replace(',','').astype(int))
# + id="iU35HxMtqrAb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="3eb1df54-220b-4a28-ded3-779f0ba1866e"
onefam['LAND_SQUARE_FEET'].describe()
# + id="zWxOJ74omoM9" colab_type="code" colab={}
# Remove certain columns because they are too high in cardinality,
# Or all the same
target = 'SALE_PRICE'
y = onefam[target]
X = onefam.drop([target] + ['EASE-MENT','BUILDING_CLASS_CATEGORY', 'ADDRESS', 'APARTMENT_NUMBER', 'TAX_CLASS_AT_TIME_OF_SALE'], axis =1)
# + id="r1G9zNLgsjMR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="4b66fa64-ed56-4d31-d11a-2b409d2bfa2f"
print(y.shape)
print(X.shape)
# + [markdown] id="yYM0q-u-stR5" colab_type="text"
# Split into training and validation sets:
# + id="D_ELgnCZZVWE" colab_type="code" colab={}
# Use data from January — March 2019 to train.
# Use data from April 2019 to test.
cutoff = '2019-04-01'
mask = onefam.index < cutoff
X_train, y_train = X.loc[mask], y.loc[mask]
X_val, y_val = X.loc[~mask], y.loc[~mask]
# + id="JlEPTNOmtuoG" colab_type="code" colab={}
assert X_train.shape[0] + X_val.shape[0] == X.shape[0]
# + [markdown] id="s71im-7vufUJ" colab_type="text"
# Establish a baseline
# + id="XwiX-Wc7utN2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="46be8a7d-4858-4daf-b3fb-277bd6b88652"
y_train.hist()
# why wouldn't this plot show?
# + id="DY1iwFKft7IR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="826aa270-4a18-435f-d7ef-42977cc0367b"
y_train.mean()
# + id="Dx0mR4jIuk6J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e0f05445-5792-452e-e6e2-7a402e91eb41"
from sklearn.metrics import mean_absolute_error
print('Baseline MAE:', mean_absolute_error(y_train, [y_train.mean()]*len(y_train)))
# + [markdown] id="uq7MDojwx3br" colab_type="text"
# Build Model
# + id="tau_IdIbv8V6" colab_type="code" colab={}
from sklearn.linear_model import LinearRegression
from category_encoders import OneHotEncoder
# + [markdown] id="kk9NbpRxyF82" colab_type="text"
# Tranform the categorical variables:
# + id="pQj3YFjLyCXX" colab_type="code" colab={}
catvars = ['BOROUGH', 'NEIGHBORHOOD', 'TAX_CLASS_AT_PRESENT', 'BUILDING_CLASS_AT_PRESENT','BUILDING_CLASS_AT_TIME_OF_SALE']
# + id="kmeg61zYynun" colab_type="code" colab={}
# Instantiate transformer
transformer_1 = OneHotEncoder(use_cat_names=True, cols=catvars)
# + id="K716jRzJy4VE" colab_type="code" colab={}
# Fit transformer to the data
transformer_1.fit(X_train)
# Transform the training data
XT_train = transformer_1.transform(X_train)
# + id="gu2Zbc700CPL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="743405aa-77da-4303-b7e1-bbd8b443ce36"
XT_train.shape
# + id="Hy3WZsavzvs1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 442} outputId="4d044f63-1387-431f-8288-f44229820946"
XT_train.columns
# + [markdown] id="-w7xSFBf0N-T" colab_type="text"
# ## That's a lot of columns!
# + [markdown] id="p_cdk9xH0Scx" colab_type="text"
# ## Select K Best:
# + id="HsBQIn8mz1ZJ" colab_type="code" colab={}
from sklearn.feature_selection import SelectKBest
# + id="aO613b7504Ma" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="37bfa77a-57b4-44d3-cfda-0a4cfd4331fa"
# Instantiate the transformer
transformer_2 = SelectKBest(k=15)
# Fit transformer to the training data
transformer_2.fit(XT_train, y_train)
# Transform the training data
XTT_train = transformer_2.transform(XT_train)
# + [markdown] id="6aDtTSfI25oV" colab_type="text"
# ## Predictor - Ridge Regression
# + id="XiDv1pWA4YKr" colab_type="code" colab={}
from sklearn.linear_model import Ridge
# + id="x8UXxxn925Sw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="2dbdfb7f-5b4d-46ba-a9d5-f18d85583fd4"
# Instantiate model
predictor = Ridge(normalize=True)
# Fit model to training data
predictor.fit(XTT_train, y_train)
# + id="n1rDEk2F4xWZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4f3ac0ce-16d9-4ea7-ebc4-fc47cc3de71b"
# Make predictions with the model
# from the twice-transformed training data
y_pred = predictor.predict(XTT_train)
# Calculate MAE
print('Training MAE:', mean_absolute_error(y_train, y_pred))
# + id="MfqosIQ36Ydx" colab_type="code" colab={}
# Transform the validation data with the same 2 transformers
XT_val = transformer_1.transform(X_val)
XTT_val = transformer_2.transform(XT_val)
# + id="4kDNNRJr6x-M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="766f5aef-6913-4bca-a7b8-544a55a543e4"
# Make predictions from the transformed validation data
#
y_pred_val = predictor.predict(XTT_val)
# Calculate MAE
print('Training MAE:', mean_absolute_error(y_val, y_pred_val))
| module3-ridge-regression/Copy_of_LS_DS_213_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/DgBack/House-Price-Prediction/blob/main/House_Price_Prediction.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Fzn2tpjGVA2b"
# Importing the Dependencies
# + id="uf4OG7G5UwKD"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn.datasets
from sklearn.model_selection import train_test_split
from xgboost import XGBRegressor
from sklearn import metrics
# + [markdown] id="4SotPY6-XOF0"
# Importing the Boston House Price Dataset
# + id="Y9Mr4bL4XYm0"
house_price_dataset = sklearn.datasets.load_boston()
# + id="fv2DsrYRXkPZ"
print(house_price_dataset)
# + id="n4cOPvZpXtFZ"
# Loading the dataset to a Pandas DataFrame
house_price_dataframe = pd.DataFrame(house_price_dataset.data, columns = house_price_dataset.feature_names)
# + id="ptXMBg58YySI"
#Print firt 5 rows of our DataFrame
house_price_dataframe.head()
# + id="dK4_we69dhia"
# add the target (price) column to the DataFrame
house_price_dataframe['price'] = house_price_dataset.target
# + id="04JIHSCeecfe"
#checking the number of rows and columns in the data frame
house_price_dataframe.shape
# + id="kh4KbyIDeytx"
# check for missing values
house_price_dataframe.isnull().sum()
# + id="hhZPJEoUfG0l"
# statistical measures of the dataset
house_price_dataframe.describe()
# + id="ngQXrHa8jN4J"
correlation = house_price_dataframe.corr()
print(correlation)
# + [markdown] id="irobNKAajd3Y"
# Splitting the data and target
# + id="0ifByK87jdZy"
X = house_price_dataframe.drop(['price'], axis=1)
Y = house_price_dataframe['price']
# + id="6A1EZcA8jkdD"
print(X)
print(Y)
# + [markdown] id="fJ6Ps3WFj2nD"
# Splitting the data into Training data and Test data
#
# + id="AE2hPpX2jqws"
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=0.2, random_state=2)
# + id="Cy-NCI7lkRPy"
print(X.shape, X_train.shape, X_test.shape)
# + [markdown] id="QHV7mRfkkdOq"
# Model Trainning
# + [markdown] id="PrBYkIalkgP3"
# XGBoost Regressor
# + id="33h8wMStkcuS"
# loading the model
model = XGBRegressor()
# + id="QmVaOhFIkz0_"
# Trainning model with X_train
model.fit(X_train, Y_train)
# + [markdown] id="kPjAxDytlEn0"
# Prediction on training data
# + id="3rvxTnqSk_08"
# accuracy fo prediction on training data
training_data_prediction = model.predict(X_train)
print(training_data_prediction)
# + id="fewOB05xmZrq"
# R squared error
score_1 = metrics.r2_score(Y_train, training_data_prediction)
# Mean Absolute Error
score_2 = metrics.mean_absolute_error(Y_train, training_data_prediction)
print("R squared error : ", score_1)
print('Mean Absolute Error : ', score_2)
# + id="a9BlxxOqmpb2"
# accuracy for prediction on test data
test_data_prediction = model.predict(X_test)
# + id="1hU4c0MTmrsN"
# R squared error
score_1 = metrics.r2_score(Y_test, test_data_prediction)
# Mean Absolute Error
score_2 = metrics.mean_absolute_error(Y_test, test_data_prediction)
print("R squared error : ", score_1)
print('Mean Absolute Error : ', score_2)
| House_Price_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env-tf2
# language: python
# name: env-tf2
# ---
# # Unsupervised Learning
#
# In unsupervised learning, labels are either not available or are not used for learning. Clustering is a form of unsupervised learning whereby, without labels, the data is characterized by constraints alone.
# LTN can formulate such constraints, such as for example:
# - clusters should be disjoint,
# - every example should be assigned to a cluster,
# - a cluster should not be empty,
# - if the points are near, they should belong to the same cluster,
# - if the points are far, they should belong to different clusters.
import logging; logging.basicConfig(level=logging.INFO)
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import logictensornetworks as ltn
# ## Data
# +
# loading data
nr_of_clusters = 4
nr_of_points_x_cluster = 50
clst_ids = range(nr_of_clusters)
close_threshold = 0.2
distant_threshold = 1.0
margin = .2
mean = [np.random.uniform([-1+margin,-1+margin],[0-margin,0-margin],2),
np.random.uniform([0+margin,-1+margin],[1-margin,0-margin],2),
np.random.uniform([-1+margin,0+margin],[0-margin,1-margin],2),
np.random.uniform([0+margin,0+margin],[1-margin,1-margin],2)]
cov = np.array([[[.01,0],[0,.01]]]*nr_of_clusters)
cluster_data = {}
for i in clst_ids:
cluster_data[i] = np.random.multivariate_normal(mean=mean[i],cov=cov[i],size=nr_of_points_x_cluster)
data = np.concatenate([cluster_data[i] for i in clst_ids])
for i in clst_ids:
plt.scatter(cluster_data[i][:, 0], cluster_data[i][:, 1])
# -
# ## Language
Not = ltn.Wrapper_Connective(ltn.fuzzy_ops.Not_Std())
And = ltn.Wrapper_Connective(ltn.fuzzy_ops.And_Prod())
Or = ltn.Wrapper_Connective(ltn.fuzzy_ops.Or_ProbSum())
Implies = ltn.Wrapper_Connective(ltn.fuzzy_ops.Implies_Reichenbach())
Equiv = ltn.Wrapper_Connective(ltn.fuzzy_ops.Equiv(ltn.fuzzy_ops.And_Prod(),ltn.fuzzy_ops.Implies_Reichenbach()))
Forall = ltn.Wrapper_Quantifier(ltn.fuzzy_ops.Aggreg_pMeanError(p=4),semantics="forall")
Exists = ltn.Wrapper_Quantifier(ltn.fuzzy_ops.Aggreg_pMean(p=6),semantics="exists")
# +
from tensorflow.keras import layers
class MLP_classifier(tf.keras.Model):
""" Model to call as P(x,class) """
def __init__(self, n_classes, single_label, hidden_layer_sizes=(16,16,16)):
super(MLP_classifier, self).__init__()
self.denses = [layers.Dense(s, activation="elu") for s in hidden_layer_sizes]
self.dense_class = layers.Dense(n_classes)
self.to_probs = tf.nn.softmax if single_label else tf.math.sigmoid
def call(self, inputs):
x, c = inputs[0], inputs[1]
for dense in self.denses:
x = dense(x)
logits = self.dense_class(x)
probs = self.to_probs(logits)
indices = tf.cast(c, tf.int32)
return tf.gather(probs, indices, batch_dims=1)
C = ltn.Predicate(MLP_classifier(nr_of_clusters, single_label=True))
cluster = ltn.variable("cluster",clst_ids)
x = ltn.variable("x",data)
y = ltn.variable("y",data)
# -
# Notice that we use a clustering predicate that ends in a `softmax` layer (`single_label=True`), which returns mutually-exclusive probabilities for each cluster.
# Therefore, there is no explicit constraint in the knowledgebase about clusters being disjoint.
#
# Notice also the use of guarded quantifiers in the two last axioms: all the pairs of points with Euclidean distance lower (resp. higher) than some threshold should belong in the same cluster (resp.should not). The thresholds (`0.2` and `1.0` in this case) do not have to explicitly define all existing pairs of points as "close" or "distant"; they only define *some of* the closest and most distant pairs of points.
# +
formula_aggregator = ltn.fuzzy_ops.Aggreg_pMeanError(p=2)
eucl_dist = lambda x,y: tf.expand_dims(tf.norm(x-y,axis=1),axis=1) # function measuring euclidian distance
# defining the theory
@tf.function
def axioms(p_exists):
axioms = [
Forall(x, Exists(cluster, C([x,cluster]),p=p_exists)),
Forall(cluster, Exists(x, C([x,cluster]),p=p_exists)),
Forall([cluster,x,y], Equiv(C([x,cluster]),C([y,cluster])),
mask_vars=[x,y],
mask_fn=lambda mask_vars: eucl_dist(mask_vars[0],mask_vars[1]) < close_threshold),
Forall([cluster,x,y], Not(And(C([x,cluster]),C([y,cluster]))),
mask_vars=[x,y],
mask_fn=lambda mask_vars: eucl_dist(mask_vars[0],mask_vars[1]) > distant_threshold)
]
axioms = tf.stack(axioms)
sat_level = formula_aggregator(axioms)
return sat_level, axioms
# -
# first call to build the graph
axioms(p_exists=6)
# ## Training
# +
trainable_variables = C.trainable_variables
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
for epoch in range(1000):
if epoch <= 100:
p_exists = 1
else:
p_exists = 6
with tf.GradientTape() as tape:
loss_value = 1. - axioms(p_exists=p_exists)[0]
grads = tape.gradient(loss_value, trainable_variables)
optimizer.apply_gradients(zip(grads, trainable_variables))
if epoch%100 == 0:
print("Epoch %d: Sat Level %.3f"%(epoch, axioms(p_exists=p_exists)[0]))
print("Training finished at Epoch %d with Sat Level %.3f"%(epoch, axioms(p_exists=p_exists)[0]))
# -
# ## Print result
plt.rcParams['font.size'] = 12
# +
x0 = data[:, 0]
x1 = data[:, 1]
prC = [C.model([data, tf.constant([[i]]*len(data))]) for i in clst_ids]
n = 2
m = (nr_of_clusters + 1) // n + 1
fig = plt.figure(figsize=(10, m * 3))
plt.subplots_adjust(wspace=0.2,hspace=0.3)
ax = plt.subplot2grid((m,8),(0,2),colspan=4)
ax.set_title("groundtruth")
for i in clst_ids:
ax.scatter(cluster_data[i][:, 0], cluster_data[i][:, 1])
for i in clst_ids:
fig.add_subplot(m, n, i + 3)
plt.title("C" + str(i) + "(x)")
plt.scatter(x0, x1, c=prC[i],vmin=0,vmax=1)
plt.colorbar()
# plt.savefig("ex_clustering_test.pdf")
plt.show()
| examples/clustering/clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dpk-a7/Deep-learning/blob/main/Transformers_Sentiment_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="DjSFbpMeqvFb" outputId="71805ea1-de36-4efd-edb4-d73f64fe168a"
# !pip install -q transformers
# + id="qrA_bOSmq5q5"
from transformers import pipeline
# + colab={"base_uri": "https://localhost:8080/", "height": 218, "referenced_widgets": ["e8a109b152d1433fa53999a48a55063c", "2a84439b5a9f465ea8b92f3a221ed0a6", "8fbcea5c129e41e785e588a65352ee54", "f588918edd3a4a1fba495a9556ab2b21", "2<PASSWORD>", "1481ee0fb5c5428ab913b6b9f7c45792", "17881673e1994651b27f39a87c364b29", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "7b64490725ec47be897736f5a2597d5b", "<KEY>", "<KEY>", "<KEY>", "cca72052441042b49cee61023f4d5a66", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "395ad24bee054e468c235d3711da97e7", "e95736cefb1049fead2e8d9721e5880b", "<KEY>", "<KEY>", "<KEY>", "a6de0d64ec874d9f8efbc2ab60d13e42", "2cef320980044cff8b65ea624370c1bf", "500a158dfac341b0ae7e273ab9956936", "7603119ce1874d4cac5350919c899181", "<KEY>", "a9c4aa7078b446a58387f2f8c592df91"]} id="S-WWXV6gq_Nm" outputId="f1c333c3-1a22-4f14-ed81-52b460d724ea"
sent_pipeline = pipeline("sentiment-analysis")
# + colab={"base_uri": "https://localhost:8080/"} id="4Bd6iu7BrHTN" outputId="ea3203a1-619a-4fb1-8f36-051607912d89"
sent_pipeline("There is a joy when using transfomers - Deepak")
# + colab={"base_uri": "https://localhost:8080/"} id="29Kp0jqsrZ7n" outputId="a80750a7-7410-42e8-e106-fb53abf73b36"
sent_pipeline("I dont like pytorch because there is soo much in it, but its the best")
# + colab={"base_uri": "https://localhost:8080/"} id="qy_B5-rrrq3P" outputId="775510e1-4643-47c1-c98f-289bf8f5eac8"
sent_pipeline("I dont like pytorch because there is soo much in it")
# + id="pN1JZEyKryIv"
| Transformers_Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import paper
import database
import importlib
importlib.reload(database)
db = database.Database()
citations = [
"<NAME>, <NAME>, Towards compressing web graphs, in: In \
Proceedings of the IEEE Data Compression Conference (DCC), 2001, pp. 203–212.",
"<NAME>, A graph distance metric based on the maximum common subgraph, \
Pattern Recognit. Lett. 19 (3) (1998) 255–259.",
"<NAME>, <NAME>, <NAME>, <NAME>, \
NetSimile: A Scalable Approach to Size-Independent Network Similarity"
]
for citation in citations:
db.add_paper(paper.Paper(), citation)
# -
print(db.all_papers.keys())
for p_hash in db.all_papers:
print
with open("testfile.txt", 'w', errors='backslashreplace') as output:
mystr = "<NAME> and <NAME>\u0107"
output.write(mystr)
mystr = mystr.replace('\\', '\\\\')
output.write(mystr)
| .ipynb_checkpoints/Networkx fiddling-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
# Activities are the class labels
# It is a 6 class classification
ACTIVITIES = {
0: 'WALKING',
1: 'WALKING_UPSTAIRS',
2: 'WALKING_DOWNSTAIRS',
3: 'SITTING',
4: 'STANDING',
5: 'LAYING',
}
# Utility function to print the confusion matrix
def confusion_matrix(Y_true, Y_pred):
Y_true = pd.Series([ACTIVITIES[y] for y in np.argmax(Y_true, axis=1)])
Y_pred = pd.Series([ACTIVITIES[y] for y in np.argmax(Y_pred, axis=1)])
return pd.crosstab(Y_true, Y_pred, rownames=['True'], colnames=['Pred'])
# -
# Data directory
DATADIR = 'UCI_HAR_Dataset'
# Raw data signals
# Signals are from Accelerometer and Gyroscope
# The signals are in x,y,z directions
# Sensor signals are filtered to have only body acceleration
# excluding the acceleration due to gravity
# Triaxial acceleration from the accelerometer is total acceleration
SIGNALS = [
"body_acc_x",
"body_acc_y",
"body_acc_z",
"body_gyro_x",
"body_gyro_y",
"body_gyro_z",
"total_acc_x",
"total_acc_y",
"total_acc_z"
]
# +
# Utility function to read the data from csv file
def _read_csv(filename):
return pd.read_csv(filename, delim_whitespace=True, header=None)
# Utility function to load the load
def load_signals(subset):
signals_data = []
for signal in SIGNALS:
filename = f'UCI_HAR_Dataset/{subset}/Inertial Signals/{signal}_{subset}.txt'
signals_data.append(
_read_csv(filename).as_matrix()
)
# Transpose is used to change the dimensionality of the output,
# aggregating the signals by combination of sample/timestep.
# Resultant shape is (7352 train/2947 test samples, 128 timesteps, 9 signals)
return np.transpose(signals_data, (1, 2, 0))
# +
def load_y(subset):
"""
The objective that we are trying to predict is a integer, from 1 to 6,
that represents a human activity. We return a binary representation of
every sample objective as a 6 bits vector using One Hot Encoding
(https://pandas.pydata.org/pandas-docs/stable/generated/pandas.get_dummies.html)
"""
filename = f'UCI_HAR_Dataset/{subset}/y_{subset}.txt'
y = _read_csv(filename)[0]
return pd.get_dummies(y).as_matrix()
# -
def load_data():
"""
Obtain the dataset from multiple files.
Returns: X_train, X_test, y_train, y_test
"""
X_train, X_test = load_signals('train'), load_signals('test')
y_train, y_test = load_y('train'), load_y('test')
return X_train, X_test, y_train, y_test
import tensorflow as tf
tf.__version__
# Importing tensorflow
np.random.seed(42)
#import tensorflow as tf
tf.set_random_seed(42)
# Configuring a session
session_conf = tf.ConfigProto(
intra_op_parallelism_threads=1,
inter_op_parallelism_threads=1
)
# Import Keras
from keras import backend as K
sess = tf.Session(graph=tf.get_default_graph(), config=session_conf)
K.set_session(sess)
# +
# Importing libraries
from keras.models import Sequential
from keras.layers import LSTM
from keras.layers.core import Dense, Dropout
from keras.layers import LSTM , BatchNormalization
from keras.regularizers import L1L2
import matplotlib.pyplot as plt
plt.rcParams["font.family"] = 'DejaVu Sans'
import seaborn as sns
# To be able to save images on server
import matplotlib
matplotlib.use('Agg')
from matplotlib import pyplot
import warnings
warnings.filterwarnings("ignore")
warnings.simplefilter(action='ignore', category=FutureWarning)
warnings.simplefilter(action='ignore', category=UserWarning)
import itertools
from datetime import datetime
# -
# Utility function to count the number of classes
def _count_classes(y):
return len(set([tuple(category) for category in y]))
# Loading the train and test data
X_train, X_test, Y_train, Y_test = load_data()
# +
timesteps = len(X_train[0])
input_dim = len(X_train[0][0])
n_classes = _count_classes(Y_train)
print(timesteps)
print(input_dim)
print(len(X_train))
# -
def plot_train_cv_loss(trained_model, epochs, colors=['g']):
fig, ax = plt.subplots(1,1)
ax.set_xlabel('epoch')
ax.set_ylabel('Categorical Crossentropy Loss')
x_axis_values = list(range(1,epochs+1))
validation_loss = trained_model.history['val_loss']
train_loss = trained_model.history['loss']
ax.plot(x_axis_values, validation_loss, 'g', label="Validation Loss")
ax.plot(x_axis_values, train_loss, 'r', label="Train Loss")
plt.legend()
plt.grid()
fig.canvas.draw()
# ## Trail 4
# +
n_epochs = 30
n_batch = 16
n_classes = _count_classes(Y_train)
# Bias regularizer value - we will use elasticnet
reg = L1L2(0.01, 0.01)
# -
model = Sequential()
model.add(LSTM(32, input_shape=(timesteps, input_dim), return_sequences=True, bias_regularizer=reg))
model.add(BatchNormalization())
model.add(Dropout(0.50))
model.add(LSTM(32))
model.add(Dropout(0.50))
model.add(Dense(n_classes, activation='sigmoid'))
print("Model Summary: ")
model.summary()
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# +
start = datetime.now()
# Training the model
trained_model = model.fit(X_train,
Y_train,
batch_size=n_batch,
validation_data=(X_test, Y_test),
epochs=n_epochs)
print("\n Time Taken: ",datetime.now() - start)
# -
# Confusion Matrix
print(confusion_matrix(Y_test, model.predict(X_test)))
plot_train_cv_loss(trained_model, n_epochs)
score = model.evaluate(X_test, Y_test)
score
# ## Observation
## this model is works well and we still need to improve the score
# ## Trail 5
# +
n_epochs = 30
n_batch = 16
n_classes = _count_classes(Y_train)
# Bias regularizer value - we will use elasticnet
reg = L1L2(0.01, 0.01)
# -
model1 = Sequential()
model1.add(LSTM(64, input_shape=(timesteps, input_dim), return_sequences=True, bias_regularizer=reg))
model1.add(BatchNormalization())
model1.add(Dropout(0.50))
model1.add(LSTM(32))
model1.add(Dropout(0.50))
model1.add(Dense(n_classes, activation='relu'))
print("Model Summary: ")
model1.summary()
model1.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# +
start = datetime.now()
# Training the model
trained_model = model1.fit(X_train,
Y_train,
batch_size=n_batch,
validation_data=(X_test, Y_test),
epochs=n_epochs)
print("\n Time Taken: ",datetime.now() - start)
# -
plot_train_cv_loss(trained_model, n_epochs)
# Confusion Matrix
print(confusion_matrix(Y_test, model1.predict(X_test)))
score1 = model1.evaluate(X_test, Y_test)
score1
# ## Observation
# +
# with the relu activation layer the reults are gone worse
#the test accuracy is 16.83 %
# the model not even calculated the val_loss
# here i can conclude that the model with relu activtion is not best for the this case
# -
# ## Trail 6
# Initializing parameters
epochs = 30
batch_size = 16
n_hidden = 32
model2 = Sequential()
# Configuring the parameters
model2.add(LSTM(n_hidden, input_shape=(timesteps, input_dim)))
# Adding a dropout layer
model2.add(Dropout(0.5))
# Adding a dense output layer with sigmoid activation
model2.add(Dense(n_classes, activation='sigmoid'))
model2.summary()
# Compiling the model
model2.compile(loss='categorical_crossentropy',
optimizer='adagrad',
metrics=['accuracy'])
model2.fit(X_train,
Y_train,
batch_size=batch_size,
validation_data=(X_test, Y_test),
epochs=epochs)
print(confusion_matrix(Y_test, model2.predict(X_test)))
score = model2.evaluate(X_test, Y_test)
score
# ## Observation
# +
# the optimiser adgrad also failed to calculate the better result
# when we obser the confusion matrix
# my model is unable to decet the walking upstaris
# this model has given the test accurac of 60.909%only
# -
| HRA_LSTM2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Deutsch-Jozsa algorithm
#
# The [Deutsch-Jozsa algorithm](https://en.wikipedia.org/wiki/Deutsch%E2%80%93Jozsa_algorithm) is the classical demonstration of the quantum parallelism: the application of a function to all the quantum states in a single step.
#
# The main idea is to find if a binary function:
# $$f:Z_{2^n} \to Z_2$$
# that is known to be *constant* (for every x the result is equal, or 0 or 1) or *balanced* (produces the same number of 0s and 1s), is constant or balanced.
#
# To compute it, the value of the function is encoded in the phase of one state as:
# $$U_f(|x>) = (-1)^{f(x)} |x>$$
#
# Where $$U_f$$ is named as *oracle*. Here, you will use a simple case:
# $$f(x) = x\%2 $$
#
# so, using the integer or binary representation of one n-qubit state
#
# $$U_f(|00...00\rangle)=(-1)^{f(0)}|0\rangle= +|0\rangle$$
# $$U_f(|00...01\rangle)=(-1)^{f(1)}|1\rangle=-|1\rangle$$
# $$U_f(|00...10\rangle)=(-1)^{f(2)}|2\rangle= +|2\rangle$$
# $$U_f(|00...11\rangle)=(-1)^{f(3)}|3\rangle=-|3\rangle$$
# $$etc.$$
#
# The algorithm has four steps:
#
# 1. Starting from a $|0\rangle$ state of *n* qubits, apply a Walsh-Hadamard transformation to create the state $|\Phi\rangle=\frac{1}{\sqrt(2^n)}\sum_{i=0}^{2^n-1}|i\rangle$
# 2. Apply the desired oracle.
# 3. Uncompute operation 1
# 4. Measure all qubits and compare with state |0>
#
import projectq
from projectq.cengines import MainEngine
from projectq.ops import H, Z, X, Measure, All
# Start the Engine and allocate an even number of qubits (by default, 4)
eng=MainEngine()
qreg=eng.allocate_qureg(4)
# Create a superposition of all the posible states with the number of allocated qubits, applying a Walsh-Hadamard operator. The cheat() method will print the amplitudes for all the states: |0000>,|0001>,etc.
All(H)|qreg
eng.flush()
eng.backend.cheat()
# To create the Oracle for the function, it is enought to compute Z applied to first qubit (qubit 0 in your quantum register), this means, for *n* qubits:
#
# $$ I^{\otimes n-1}\otimes Z $$
#
# For 2 qubits this operator, in matrix form, looks as:
#
# $$ I \otimes Z = \begin{bmatrix}1&0\\0&1\end{bmatrix}\otimes\begin{bmatrix}1&0\\0&-1\end{bmatrix}=
# \begin{bmatrix}1&0&0&0\\0&-1&0&0\\0&0&1&0\\0&0&0&-1\end{bmatrix}$$
#
# Applying it to the generated state:
# $$|\Phi\rangle = \frac{1}{2}\begin{bmatrix}1\\1\\1\\1\\\end{bmatrix}=\frac{1}{2}(|0\rangle+|1\rangle+|2\rangle+|3\rangle)$$
#
# yields:
#
# $$|\phi\rangle = \frac{1}{2}\begin{bmatrix}1\\-1\\1\\-1\\\end{bmatrix}$$
#
# or, using the integer representation:
#
# $$|\phi\rangle = \frac{1}{2}(|0\rangle-|1\rangle+|2\rangle-|3\rangle)$$
#
#
Z|qreg[0]
#
#Other oracles you can check
#
#X|qreg[0] #Constant
#All(X)|qreg #Constant
#All(Z)|qreg #balanced
eng.flush()
eng.backend.cheat()
# Uncompute the superposition
All(H)|qreg
eng.flush()
eng.backend.cheat()
# Measure the results. If it is **constant**, the result must be |0000>=|0>. If it is **balanced**, the result is different of |0>
All(Measure) | qreg
eng.flush()
output=0
for i in qreg:
output+=int(i)
print("The result is %s"%("balanced" if output else "constant"))
# If you want to check that the result is 0 if is constant, substitute the gate Z in the oracle by X.
# If you want to test with another balanced function, substitute Z by All(Z), but substitute also qreg[0] by qreg
| Notebooks/Deutsch-Jozsa_algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import multiprocessing as m
import random as r
import time
import numpy as np
def test():
r.seed()
time.sleep(1)
k = r.randint(10,100) + r.randint(10,100)
print(k)
test()
p1 = m.Process(target = test)
p2 = m.Process(target = test)
p1.start()
p2.start()
# +
t1 = time.time()
for i in range(10):
test()
t2 = time.time()
print('Done in {:.4f} seconds'.format(t2-t1))
# +
ranProcesses = []
tic = time.time()
for i in range(10):
p = m.Process(target = test)
p.start()
ranProcesses.append(p)
for process in ranProcesses:
process.join()
toc = time.time()
print('Done in {:.4f} seconds'.format(toc-tic))
# -
def testDummy(i):
return test()
# +
t3 = time.time()
pool = m.Pool(10)
pool.map(testDummy, range(1,11))
pool.close()
t4 = time.time()
print('Done in {:.4f} seconds'.format(t4-t3))
# -
def generate_data(num_data):
# AWGN
v = 0.1*np.random.normal(0,1,num_data+2)
# Initial conditions
d_true =[0.1 , 0.1]
d = [d_true[0] + v[0], d_true[1] + v[1]]
# Grab new data
new_d_true = lambda d: d.append((0.8 - 0.5 * np.exp(-(d[-1]**2)))*d[-1] - (0.3 + 0.9*np.exp(-(d[-1]**2)))*d[-2] + 0.1*np.sin(np.pi*d[-1]))
for i in range(2,num_data+2):
new_d_true(d_true)
d.append(d_true[-1] + v[i])
u = np.hstack((np.array(d[0:num_data]).reshape(num_data,1),np.array(d[1:num_data+1]).reshape(num_data,1)))
d_true = d_true[2::]
d = d[2::]
return np.array(u), np.array(d),np.array(d_true)
u, d, dTrue = generate_data(10)
| FederatedLearing/Multi-Process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="6q7dDydT7n2d" outputId="be47d969-c754-4cdd-f6d5-db210bcb30b6" colab={"base_uri": "https://localhost:8080/", "height": 255}
# !wget http://www.manythings.org/anki/cmn-eng.zip
# !unzip -d ./cmn-eng cmn-eng.zip
# + colab_type="code" id="Zo7QCacp7n2s" colab={}
seed = 2020
# + colab_type="code" id="05LtExs57n20" colab={}
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
import time
import math
import random
# + id="B7h691pQdmyW" colab_type="code" colab={}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + [markdown] colab_type="text" id="QW9iTQeF7n2-"
# # 读取数据
# + colab_type="code" id="9TBhsBe27n3A" outputId="3cbf44b8-3dc7-490d-810a-3195ba8f38ad" colab={"base_uri": "https://localhost:8080/", "height": 102}
# 每一行数据如下
# 'Hi.\t嗨。\tCC-BY 2.0 (France) Attribution: tatoeba.org #538123 (CM) & #891077 (Martha)'
with open('./cmn-eng/cmn.txt', 'r', encoding='utf-8') as f:
data = f.read()
data = data.strip()
data = data.split('\n')
print('样本数:\n', len(data))
print('\n样本示例:')
data[0]
# + colab_type="code" id="vXXhro447n3F" outputId="<PASSWORD>-ee<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 102}
# 分割英文数据和中文数据
en_data = [line.split('\t')[0] for line in data]
ch_data = [line.split('\t')[1] for line in data]
print('英文数据:\n', en_data[:10])
print('\n中文数据:\n', ch_data[:10])
# + colab_type="code" id="k1ioMRh17n3O" outputId="<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 102}
# 按字符级切割,并添加<eos>
en_token_list = [[char for char in line]+["<eos>"] for line in en_data]
ch_token_list = [[char for char in line]+["<eos>"] for line in ch_data]
print('英文数据:\n', en_token_list[:2])
print('\n中文数据:\n', ch_token_list[:2])
# + colab_type="code" id="wPAejxm17n3T" colab={}
# 基本字典
basic_dict = {'<pad>':0, '<unk>':1, '<bos>':2, '<eos>':3}
# 分别生成中英文字典
en_vocab = set(''.join(en_data))
en2id = {char:i+len(basic_dict) for i, char in enumerate(en_vocab)}
en2id.update(basic_dict)
id2en = {v:k for k,v in en2id.items()}
# 分别生成中英文字典
ch_vocab = set(''.join(ch_data))
ch2id = {char:i+len(basic_dict) for i, char in enumerate(ch_vocab)}
ch2id.update(basic_dict)
id2ch = {v:k for k,v in ch2id.items()}
# + colab_type="code" id="jLIjhvZt7n3e" outputId="96e35b74-b2d5-424e-ac0d-425090c11275" colab={"base_uri": "https://localhost:8080/", "height": 51}
# 利用字典,映射数据
en_num_data = [[en2id[en] for en in line ] for line in en_token_list]
ch_num_data = [[ch2id[ch] for ch in line] for line in ch_token_list]
print('char:', en_data[1])
print('index:', en_num_data[1])
# + [markdown] colab_type="text" id="IZ5ZtQfN7n3j"
# # 表示为Dataset
# + colab_type="code" id="PPTYSlOw7n3k" colab={}
class TranslationDataset(Dataset):
def __init__(self, src_data, trg_data):
self.src_data = src_data
self.trg_data = trg_data
assert len(src_data) == len(trg_data), \
"numbers of src_data and trg_data must be equal!"
def __len__(self):
return len(self.src_data)
def __getitem__(self, idx):
src_sample =self.src_data[idx]
src_len = len(self.src_data[idx])
trg_sample = self.trg_data[idx]
trg_len = len(self.trg_data[idx])
return {"src": src_sample, "src_len": src_len, "trg": trg_sample, "trg_len": trg_len}
# + colab_type="code" id="yUKfKfpc7n3p" colab={}
def padding_batch(batch):
"""
input: -> list of dict
[{'src': [1, 2, 3], 'trg': [1, 2, 3]}, {'src': [1, 2, 2, 3], 'trg': [1, 2, 2, 3]}]
output: -> dict of tensor
{
"src": [[1, 2, 3, 0], [1, 2, 2, 3]].T
"trg": [[1, 2, 3, 0], [1, 2, 2, 3]].T
}
"""
src_lens = [d["src_len"] for d in batch]
trg_lens = [d["trg_len"] for d in batch]
src_max = max([d["src_len"] for d in batch])
trg_max = max([d["trg_len"] for d in batch])
for d in batch:
d["src"].extend([en2id["<pad>"]]*(src_max-d["src_len"]))
d["trg"].extend([ch2id["<pad>"]]*(trg_max-d["trg_len"]))
srcs = torch.tensor([pair["src"] for pair in batch], dtype=torch.long, device=device)
trgs = torch.tensor([pair["trg"] for pair in batch], dtype=torch.long, device=device)
batch = {"src":srcs.T, "src_len":src_lens, "trg":trgs.T, "trg_len":trg_lens}
return batch
# + [markdown] colab_type="text" id="aw8EeZBB9mrK"
# # 不用Attention机制
# + [markdown] colab_type="text" id="F19j3Svn7n3v"
# ## 模型
# + colab_type="code" id="8qsKXeDY7n3w" colab={}
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, hid_dim, n_layers, dropout=0.5, bidirectional=True):
super(Encoder, self).__init__()
self.hid_dim = hid_dim
self.n_layers = n_layers
self.embedding = nn.Embedding(input_dim, emb_dim)
self.gru = nn.GRU(emb_dim, hid_dim, n_layers, dropout=dropout, bidirectional=bidirectional)
def forward(self, input_seqs, input_lengths, hidden):
# input_seqs = [seq_len, batch]
embedded = self.embedding(input_seqs)
# embedded = [seq_len, batch, embed_dim]
packed = torch.nn.utils.rnn.pack_padded_sequence(embedded, input_lengths, enforce_sorted=False)
outputs, hidden = self.gru(packed, hidden)
outputs, output_lengths = torch.nn.utils.rnn.pad_packed_sequence(outputs)
# outputs = [seq_len, batch, hid_dim * n directions]
# output_lengths = [batch]
return outputs, hidden
# + colab_type="code" id="3WdZchKu7n33" colab={}
class Decoder(nn.Module):
def __init__(self, output_dim, emb_dim, hid_dim, n_layers, dropout=0.5, bidirectional=True):
super(Decoder, self).__init__()
self.output_dim = output_dim
self.hid_dim = hid_dim
self.n_layers = n_layers
self.embedding = nn.Embedding(output_dim, emb_dim)
self.gru = nn.GRU(emb_dim, hid_dim, n_layers, dropout=dropout, bidirectional=bidirectional)
if bidirectional:
self.fc_out = nn.Linear(hid_dim*2, output_dim)
else:
self.fc_out = nn.Linear(hid_dim, output_dim)
self.dropout = nn.Dropout(dropout)
self.softmax = nn.LogSoftmax(dim=1)
def forward(self, token_inputs, hidden):
# token_inputs = [batch]
batch_size = token_inputs.size(0)
embedded = self.dropout(self.embedding(token_inputs).view(1, batch_size, -1))
# embedded = [1, batch, emb_dim]
output, hidden = self.gru(embedded, hidden)
# output = [1, batch, n_directions * hid_dim]
# hidden = [n_layers * n_directions, batch, hid_dim]
output = self.fc_out(output.squeeze(0))
output = self.softmax(output)
# output = [batch, output_dim]
return output, hidden
# + colab_type="code" id="tkDLZQVR7n4A" colab={}
class Seq2Seq(nn.Module):
def __init__(self,
encoder,
decoder,
device,
predict=False,
basic_dict=None,
max_len=100
):
super(Seq2Seq, self).__init__()
self.device = device
self.encoder = encoder
self.decoder = decoder
self.predict = predict # 训练阶段还是预测阶段
self.basic_dict = basic_dict # decoder的字典,存放特殊token对应的id
self.max_len = max_len # 翻译时最大输出长度
self.enc_n_layers = self.encoder.gru.num_layers
self.enc_n_directions = 2 if self.encoder.gru.bidirectional else 1
self.dec_n_directions = 2 if self.decoder.gru.bidirectional else 1
assert encoder.hid_dim == decoder.hid_dim, \
"Hidden dimensions of encoder and decoder must be equal!"
assert encoder.n_layers == decoder.n_layers, \
"Encoder and decoder must have equal number of layers!"
assert self.enc_n_directions >= self.dec_n_directions, \
"If decoder is bidirectional, encoder must be bidirectional either!"
def forward(self, input_batches, input_lengths, target_batches=None, target_lengths=None, teacher_forcing_ratio=0.5):
# input_batches = target_batches = [seq_len, batch]
batch_size = input_batches.size(1)
BOS_token = self.basic_dict["<bos>"]
EOS_token = self.basic_dict["<eos>"]
PAD_token = self.basic_dict["<pad>"]
# 初始化
encoder_hidden = torch.zeros(self.enc_n_layers*self.enc_n_directions, batch_size, self.encoder.hid_dim, device=self.device)
# encoder_output = [seq_len, batch, hid_dim * n directions]
# encoder_hidden = [n_layers*n_directions, batch, hid_dim]
encoder_output, encoder_hidden = self.encoder(
input_batches, input_lengths, encoder_hidden)
# 初始化
decoder_input = torch.tensor([BOS_token] * batch_size, dtype=torch.long, device=self.device)
if self.enc_n_directions == self.dec_n_directions:
decoder_hidden = encoder_hidden
else:
L = encoder_hidden.size(0)
decoder_hidden = encoder_hidden[range(0, L, 2)] + encoder_hidden[range(1, L, 2)]
if self.predict:
# 预测阶段使用
# 一次只输入一句话
assert batch_size == 1, "batch_size of predict phase must be 1!"
output_tokens = []
while True:
decoder_output, decoder_hidden = self.decoder(
decoder_input, decoder_hidden
)
# [1, 1]
topv, topi = decoder_output.topk(1)
decoder_input = topi.squeeze(1) # 上一个预测作为下一个输入
output_token = topi.squeeze().detach().item()
if output_token == EOS_token or len(output_tokens) == self.max_len:
break
output_tokens.append(output_token)
return output_tokens
else:
# 训练阶段
max_target_length = max(target_lengths)
all_decoder_outputs = torch.zeros((max_target_length, batch_size, self.decoder.output_dim), device=self.device)
for t in range(max_target_length):
use_teacher_forcing = True if random.random() < teacher_forcing_ratio else False
if use_teacher_forcing:
# decoder_output = [batch, output_dim]
# decoder_hidden = [n_layers*n_directions, batch, hid_dim]
decoder_output, decoder_hidden = self.decoder(
decoder_input, decoder_hidden
)
all_decoder_outputs[t] = decoder_output
decoder_input = target_batches[t] # 下一个输入来自训练数据
else:
decoder_output, decoder_hidden = self.decoder(
decoder_input, decoder_hidden
)
# [batch, 1]
topv, topi = decoder_output.topk(1)
all_decoder_outputs[t] = decoder_output
decoder_input = topi.squeeze(1) # 下一个输入来自模型预测
loss_fn = nn.NLLLoss(ignore_index=PAD_token)
loss = loss_fn(
all_decoder_outputs.reshape(-1,self.decoder.output_dim ), # [batch*seq_len, output_dim]
target_batches.reshape(-1) # [batch*seq_len]
)
return loss
# + [markdown] colab_type="text" id="DGIK_4IO7n4G"
# ## 训练和预测代码
# + colab_type="code" id="TvUaEp0Y7n4H" colab={}
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + colab_type="code" id="FcB6O0cJ7n4P" colab={}
def train(
model,
data_loader,
optimizer,
clip=1,
teacher_forcing_ratio=0.5,
print_every=None # None不打印
):
model.predict = False
model.train()
if print_every == 0:
print_every = 1
print_loss_total = 0 # 每次打印都重置
start = time.time()
epoch_loss = 0
for i, batch in enumerate(data_loader):
# shape = [seq_len, batch]
input_batchs = batch["src"]
target_batchs = batch["trg"]
# list
input_lens = batch["src_len"]
target_lens = batch["trg_len"]
optimizer.zero_grad()
loss = model(input_batchs, input_lens, target_batchs, target_lens, teacher_forcing_ratio)
print_loss_total += loss.item()
epoch_loss += loss.item()
loss.backward()
# 梯度裁剪
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
optimizer.step()
if print_every and (i+1) % print_every == 0:
print_loss_avg = print_loss_total / print_every
print_loss_total = 0
print('\tCurrent Loss: %.4f' % print_loss_avg)
return epoch_loss / len(data_loader)
# + colab_type="code" id="DgYv7rsp7n4V" colab={}
def evaluate(
model,
data_loader,
print_every=None
):
model.predict = False
model.eval()
if print_every == 0:
print_every = 1
print_loss_total = 0 # 每次打印都重置
start = time.time()
epoch_loss = 0
with torch.no_grad():
for i, batch in enumerate(data_loader):
# shape = [seq_len, batch]
input_batchs = batch["src"]
target_batchs = batch["trg"]
# list
input_lens = batch["src_len"]
target_lens = batch["trg_len"]
loss = model(input_batchs, input_lens, target_batchs, target_lens, teacher_forcing_ratio=0)
print_loss_total += loss.item()
epoch_loss += loss.item()
if print_every and (i+1) % print_every == 0:
print_loss_avg = print_loss_total / print_every
print_loss_total = 0
print('\tCurrent Loss: %.4f' % print_loss_avg)
return epoch_loss / len(data_loader)
# + colab_type="code" id="ldcZM35r7n4h" colab={}
def translate(
model,
sample,
idx2token=None
):
model.predict = True
model.eval()
# shape = [seq_len, 1]
input_batch = sample["src"]
# list
input_len = sample["src_len"]
output_tokens = model(input_batch, input_len)
output_tokens = [idx2token[t] for t in output_tokens]
return "".join(output_tokens)
# + [markdown] colab_type="text" id="r9luHxDc7n4q"
# ## 开始训练
# + colab_type="code" id="QSLPWwzT7n4s" colab={}
INPUT_DIM = len(en2id)
OUTPUT_DIM = len(ch2id)
# 超参数
BATCH_SIZE = 32
ENC_EMB_DIM = 256
DEC_EMB_DIM = 256
HID_DIM = 512
N_LAYERS = 2
ENC_DROPOUT = 0.5
DEC_DROPOUT = 0.5
LEARNING_RATE = 1e-4
N_EPOCHS = 200
CLIP = 1
bidirectional = True
enc = Encoder(INPUT_DIM, ENC_EMB_DIM, HID_DIM, N_LAYERS, ENC_DROPOUT, bidirectional)
dec = Decoder(OUTPUT_DIM, DEC_EMB_DIM, HID_DIM, N_LAYERS, DEC_DROPOUT, bidirectional)
model = Seq2Seq(enc, dec, device, basic_dict=basic_dict).to(device)
## encoder和encoder设置相同的学习策略
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
# ## encoder和encoder设置不同的学习策略
# optimizer_grouped_parameters = [
# {'params': [p for n, p in model.named_parameters() if 'encoder' in n], 'lr': LEARNING_RATE},
# {'params': [p for n, p in model.named_parameters() if 'decoder' in n], 'lr': LEARNING_RATE*2}
# ]
# optimizer = optim.Adam(optimizer_grouped_parameters)
# + colab_type="code" id="xHDdqB2t7n43" colab={}
# 数据集
train_set = TranslationDataset(en_num_data, ch_num_data)
train_loader = DataLoader(train_set, batch_size=BATCH_SIZE, collate_fn=padding_batch)
# + id="39nVqNwVdm2R" colab_type="code" colab={}
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_loader, optimizer, CLIP)
valid_loss = evaluate(model, train_loader)
end_time = time.time()
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), 'en2ch-model.pt')
if epoch %2 == 0:
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Val. Loss: {valid_loss:.3f}')
# + [markdown] id="f7AKhdoGdm2g" colab_type="text"
# ## 模型使用
# + id="oqXQO8ezdm2h" colab_type="code" colab={} outputId="31464c6f-82ed-46a5-a710-7e01e6f487e0"
print("best valid loss:", best_valid_loss)
# 加载最优权重
model.load_state_dict(torch.load('en2ch-model.pt'))
# + id="twr9NGHzdm2w" colab_type="code" colab={} outputId="fa4ed3e9-37bf-4f72-c133-a9d93a4c926e"
random.seed(seed)
for i in random.sample(range(len(en_num_data)), 10): # 随机看10个
en_tokens = list(filter(lambda x: x!=0, en_num_data[i])) # 过滤零
ch_tokens = list(filter(lambda x: x!=3 and x!=0, ch_num_data[i])) # 和机器翻译作对照
sentence = [id2en[t] for t in en_tokens]
print("【原文】")
print("".join(sentence))
translation = [id2ch[t] for t in ch_tokens]
print("【原文】")
print("".join(translation))
test_sample = {}
test_sample["src"] = torch.tensor(en_tokens, dtype=torch.long, device=device).reshape(-1, 1)
test_sample["src_len"] = [len(en_tokens)]
print("【机器翻译】")
print(translate(model, test_sample, id2ch), end="\n\n")
# + colab_type="code" id="ANbudxH17n5Z" outputId="131e7f2c-2d6b-4deb-9406-6f1368ec0554" colab={}
random.seed(seed*2)
for i in random.sample(range(len(en_num_data)), 10): # 随机看10个
en_tokens = list(filter(lambda x: x!=0, en_num_data[i])) # 过滤零
ch_tokens = list(filter(lambda x: x!=3 and x!=0, ch_num_data[i])) # 和机器翻译作对照
sentence = [id2en[t] for t in en_tokens]
print("【原文】")
print("".join(sentence))
translation = [id2ch[t] for t in ch_tokens]
print("【原文】")
print("".join(translation))
test_sample = {}
test_sample["src"] = torch.tensor(en_tokens, dtype=torch.long, device=device).reshape(-1, 1)
test_sample["src_len"] = [len(en_tokens)]
print("【机器翻译】")
print(translate(model, test_sample, id2ch), end="\n\n")
# + [markdown] colab_type="text" id="Es1g11r09g9v"
# # Attention机制
# + [markdown] colab_type="text" id="0h3IhpdS7n5j"
# ## 模型
# + colab_type="code" id="SBpm6azx_FLe" colab={}
class Encoder(nn.Module):
def __init__(self, input_dim, emb_dim, hid_dim, n_layers, dropout=0.5, bidirectional=True):
super(Encoder, self).__init__()
self.hid_dim = hid_dim
self.n_layers = n_layers
self.embedding = nn.Embedding(input_dim, emb_dim)
self.gru = nn.GRU(emb_dim, hid_dim, n_layers, dropout=dropout, bidirectional=bidirectional)
def forward(self, input_seqs, input_lengths, hidden):
# input_seqs = [seq_len, batch]
embedded = self.embedding(input_seqs)
# embedded = [seq_len, batch, embed_dim]
packed = torch.nn.utils.rnn.pack_padded_sequence(embedded, input_lengths, enforce_sorted=False)
outputs, hidden = self.gru(packed, hidden)
outputs, output_lengths = torch.nn.utils.rnn.pad_packed_sequence(outputs)
# outputs = [seq_len, batch, hid_dim * n directions]
# output_lengths = [batch]
return outputs, hidden
# + colab_type="code" id="rMbsHQa38Kv1" colab={}
class Attn(nn.Module):
def __init__(self, method, hidden_size):
super(Attn, self).__init__()
self.method = method
if self.method not in ['dot', 'general', 'concat']:
raise ValueError(self.method, "is not an appropriate attention method.")
self.hidden_size = hidden_size
if self.method == 'general':
self.attn = nn.Linear(self.hidden_size, hidden_size)
elif self.method == 'concat':
self.attn = nn.Linear(self.hidden_size * 2, hidden_size)
self.v = nn.Parameter(torch.FloatTensor(hidden_size))
def dot_score(self, hidden, encoder_output):
return torch.sum(hidden * encoder_output, dim=2) # [seq_len, batch]
def general_score(self, hidden, encoder_output):
energy = self.attn(encoder_output) # [seq_len, batch, hid_dim]
return torch.sum(hidden * energy, dim=2) # [seq_len, batch]
def concat_score(self, hidden, encoder_output):
# hidden.expand(encoder_output.size(0), -1, -1) -> [seq_len, batch, N]
energy = self.attn(torch.cat((hidden.expand(encoder_output.size(0), -1, -1), encoder_output), 2)).tanh()
# energy = [sql_len, batch, hidden_size]
return torch.sum(self.v * energy, dim=2) # [seq_len, batch]
def forward(self, hidden, encoder_outputs):
# hidden = [1, batch, n_directions * hid_dim]
# encoder_outputs = [seq_len, batch, hid dim * n directions]
if self.method == 'general':
attn_energies = self.general_score(hidden, encoder_outputs)
elif self.method == 'concat':
attn_energies = self.concat_score(hidden, encoder_outputs)
elif self.method == 'dot':
attn_energies = self.dot_score(hidden, encoder_outputs)
attn_energies = attn_energies.t() # [batch, seq_len]
return F.softmax(attn_energies, dim=1).unsqueeze(1) # softmax归一化# [batch, 1, seq_len]
# + colab_type="code" id="lLbqgxNg8XT4" colab={}
class AttnDecoder(nn.Module):
def __init__(self, output_dim, emb_dim, hid_dim, n_layers=1, dropout=0.5, bidirectional=True, attn_method="general"):
super(AttnDecoder, self).__init__()
self.output_dim = output_dim
self.emb_dim = emb_dim
self.hid_dim = hid_dim
self.n_layers = n_layers
self.dropout = dropout
self.embedding = nn.Embedding(output_dim, emb_dim)
self.embedding_dropout = nn.Dropout(dropout)
self.gru = nn.GRU(emb_dim, hid_dim, n_layers, dropout=dropout, bidirectional=bidirectional)
if bidirectional:
self.concat = nn.Linear(hid_dim * 2 * 2, hid_dim*2)
self.out = nn.Linear(hid_dim*2, output_dim)
self.attn = Attn(attn_method, hid_dim*2)
else:
self.concat = nn.Linear(hid_dim * 2, hid_dim)
self.out = nn.Linear(hid_dim, output_dim)
self.attn = Attn(attn_method, hid_dim)
self.softmax = nn.LogSoftmax(dim=1)
def forward(self, token_inputs, last_hidden, encoder_outputs):
batch_size = token_inputs.size(0)
embedded = self.embedding(token_inputs)
embedded = self.embedding_dropout(embedded)
embedded = embedded.view(1, batch_size, -1) # [1, B, hid_dim]
gru_output, hidden = self.gru(embedded, last_hidden)
# gru_output = [1, batch, n_directions * hid_dim]
# hidden = [n_layers * n_directions, batch, hid_dim]
# encoder_outputs = [sql_len, batch, hid dim * n directions]
attn_weights = self.attn(gru_output, encoder_outputs)
# attn_weights = [batch, 1, sql_len]
context = attn_weights.bmm(encoder_outputs.transpose(0, 1))
# [batch, 1, hid_dim * n directions]
# LuongAttention
gru_output = gru_output.squeeze(0) # [batch, n_directions * hid_dim]
context = context.squeeze(1) # [batch, n_directions * hid_dim]
concat_input = torch.cat((gru_output, context), 1) # [batch, n_directions * hid_dim * 2]
concat_output = torch.tanh(self.concat(concat_input)) # [batch, n_directions*hid_dim]
output = self.out(concat_output) # [batch, output_dim]
output = self.softmax(output)
return output, hidden, attn_weights
# + colab_type="code" id="y94lk1qX_Cxn" colab={}
class Seq2Seq(nn.Module):
def __init__(self,
encoder,
decoder,
device,
predict=False,
basic_dict=None,
max_len=100
):
super(Seq2Seq, self).__init__()
self.device = device
self.encoder = encoder
self.decoder = decoder
self.predict = predict # 训练阶段还是预测阶段
self.basic_dict = basic_dict # decoder的字典,存放特殊token对应的id
self.max_len = max_len # 翻译时最大输出长度
assert encoder.hid_dim == decoder.hid_dim, \
"Hidden dimensions of encoder and decoder must be equal!"
assert encoder.n_layers == decoder.n_layers, \
"Encoder and decoder must have equal number of layers!"
assert encoder.gru.bidirectional == decoder.gru.bidirectional, \
"Decoder and encoder must had same value of bidirectional attribute!"
def forward(self, input_batches, input_lengths, target_batches=None, target_lengths=None, teacher_forcing_ratio=0.5):
# input_batches = [seq_len, batch]
# target_batches = [seq_len, batch]
batch_size = input_batches.size(1)
BOS_token = self.basic_dict["<bos>"]
EOS_token = self.basic_dict["<eos>"]
PAD_token = self.basic_dict["<pad>"]
# 初始化
enc_n_layers = self.encoder.gru.num_layers
enc_n_directions = 2 if self.encoder.gru.bidirectional else 1
encoder_hidden = torch.zeros(enc_n_layers*enc_n_directions, batch_size, self.encoder.hid_dim, device=self.device)
# encoder_outputs = [input_lengths, batch, hid_dim * n directions]
# encoder_hidden = [n_layers*n_directions, batch, hid_dim]
encoder_outputs, encoder_hidden = self.encoder(
input_batches, input_lengths, encoder_hidden)
# 初始化
decoder_input = torch.tensor([BOS_token] * batch_size, dtype=torch.long, device=self.device)
decoder_hidden = encoder_hidden
if self.predict:
# 一次只输入一句话
assert batch_size == 1, "batch_size of predict phase must be 1!"
output_tokens = []
while True:
decoder_output, decoder_hidden, decoder_attn = self.decoder(
decoder_input, decoder_hidden, encoder_outputs
)
# [1, 1]
topv, topi = decoder_output.topk(1)
decoder_input = topi.squeeze(1).detach()
output_token = topi.squeeze().detach().item()
if output_token == EOS_token or len(output_tokens) == self.max_len:
break
output_tokens.append(output_token)
return output_tokens
else:
max_target_length = max(target_lengths)
all_decoder_outputs = torch.zeros((max_target_length, batch_size, self.decoder.output_dim), device=self.device)
for t in range(max_target_length):
use_teacher_forcing = True if random.random() < teacher_forcing_ratio else False
if use_teacher_forcing:
# decoder_output = [batch, output_dim]
# decoder_hidden = [n_layers*n_directions, batch, hid_dim]
decoder_output, decoder_hidden, decoder_attn = self.decoder(
decoder_input, decoder_hidden, encoder_outputs
)
all_decoder_outputs[t] = decoder_output
decoder_input = target_batches[t] # 下一个输入来自训练数据
else:
decoder_output, decoder_hidden, decoder_attn = self.decoder(
decoder_input, decoder_hidden, encoder_outputs
)
# [batch, 1]
topv, topi = decoder_output.topk(1)
all_decoder_outputs[t] = decoder_output
decoder_input = topi.squeeze(1).detach() # 下一个输入来自模型预测
loss_fn = nn.NLLLoss(ignore_index=PAD_token)
loss = loss_fn(
all_decoder_outputs.reshape(-1, self.decoder.output_dim), # [batch*seq_len, output_dim]
target_batches.reshape(-1) # [batch*seq_len]
)
return loss
# + [markdown] colab_type="text" id="kLS0uKiu9WtW"
# ## 开始训练
# + colab_type="code" id="3ALxD62K8a-s" colab={}
INPUT_DIM = len(en2id)
OUTPUT_DIM = len(ch2id)
# 超参数
BATCH_SIZE = 32
ENC_EMB_DIM = 256
DEC_EMB_DIM = 256
HID_DIM = 512
N_LAYERS = 2
ENC_DROPOUT = 0.5
DEC_DROPOUT = 0.5
LEARNING_RATE = 1e-4
N_EPOCHS = 200
CLIP = 1
bidirectional = True
attn_method = "general"
enc = Encoder(INPUT_DIM, ENC_EMB_DIM, HID_DIM, N_LAYERS, ENC_DROPOUT, bidirectional)
dec = AttnDecoder(OUTPUT_DIM, DEC_EMB_DIM, HID_DIM, N_LAYERS, DEC_DROPOUT, bidirectional, attn_method)
model = Seq2Seq(enc, dec, device, basic_dict=basic_dict).to(device)
optimizer = optim.Adam(model.parameters(), lr=LEARNING_RATE)
# + colab_type="code" id="A1HXlUlz9_oV" colab={}
# 数据集
train_set = TranslationDataset(en_num_data, ch_num_data)
train_loader = DataLoader(train_set, batch_size=BATCH_SIZE, collate_fn=padding_batch)
# + colab_type="code" id="GbnYsq9p-GbI" colab={}
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_loader, optimizer, CLIP)
valid_loss = evaluate(model, train_loader)
end_time = time.time()
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), "en2ch-attn-model.pt")
if epoch %2 == 0:
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Val. Loss: {valid_loss:.3f}')
# + id="2aw5nHpOdm5J" colab_type="code" colab={} outputId="f41efc81-8de2-4c84-a4f4-437fda6b5dd6"
print("best valid loss:", best_valid_loss)
# 加载最优权重
model.load_state_dict(torch.load("en2ch-attn-model.pt"))
# + id="8hGVhGBydm5R" colab_type="code" colab={} outputId="0ebf3daa-4717-42aa-f0ed-c535bb5d2248"
random.seed(seed)
for i in random.sample(range(len(en_num_data)), 10): # 随机看10个
en_tokens = list(filter(lambda x: x!=0, en_num_data[i])) # 过滤零
ch_tokens = list(filter(lambda x: x!=3 and x!=0, ch_num_data[i])) # 和机器翻译作对照
sentence = [id2en[t] for t in en_tokens]
print("【原文】")
print("".join(sentence))
translation = [id2ch[t] for t in ch_tokens]
print("【原文】")
print("".join(translation))
test_sample = {}
test_sample["src"] = torch.tensor(en_tokens, dtype=torch.long, device=device).reshape(-1, 1)
test_sample["src_len"] = [len(en_tokens)]
print("【机器翻译】")
print(translate(model, test_sample, id2ch), end="\n\n")
# + id="k0_5yNr-dm5a" colab_type="code" colab={}
| pytorch/Seq2SeqForTranslation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os.path as osp
import re
import glob
# %load_ext dotenv
# %dotenv env.sh
# %run -m cytokit_nb.keyence
df = pd.read_csv('experiments-base.csv')
df
# +
def get_grids(r):
grids = glob.glob(osp.join(os.environ['EXP_GROUP_RAW_DIR'], r['dir'], 'XY*'))
return [g.split(os.sep)[-1] for g in grids]
dfg = df.copy()
dfg['grid'] = df.apply(get_grids, axis=1)
# Stack grid list into rows
dfg = dfg.set_index(['name', 'dir'])['grid'].apply(pd.Series).stack().rename('grid').reset_index(level=-1, drop=True).reset_index()
# Assign replicate number
dfg = dfg.groupby(['name', 'dir'], group_keys=False)\
.apply(lambda g: g.assign(rep=pd.Categorical(g['grid'], ordered=True, categories=g['grid'].sort_values()).codes + 1))
dfg.head()
# -
dfg.groupby(['name', 'dir'])['grid'].unique()
dfg.groupby(['name', 'dir'])['rep'].unique()
# +
def get_row(r, info, base_conf):
return r.append(pd.Series({
'z_pitch': info['z_pitch'].iloc[0],
'n_tiles': info['tile'].nunique(),
'n_z': info['z'].nunique(),
'n_ch': info['ch'].nunique(),
'chs': tuple(sorted(info['ch'].unique())),
'conf': 'experiment.yaml'
})).sort_index()
def add_info(r):
r = r.copy()
path = osp.join(os.environ['EXP_GROUP_RAW_DIR'], r['dir'], r['grid'])
info = analyze_keyence_dataset(path)
base_conf = 'experiment.yaml'
return get_row(r, info, base_conf)
dfi = pd.DataFrame([
add_info(r)
for _, r in dfg.iterrows()
])
dfi.head()
# -
dfi['n_tiles'].unique()
dfi['z_pitch'].unique()
dfi['chs'].unique()
assert dfi['chs'].nunique() == 1
assert dfi['n_ch'].nunique() == 1
assert dfi['z_pitch'].nunique() == 1
assert dfi['n_tiles'].nunique() == 1
dfe = dfi.copy()
dfe = dfe.rename(columns={'name': 'cond'})
dfe = dfe.drop('chs', axis=1)
dfe.insert(0, 'name', dfe.apply(lambda r: '{}-{}-{}'.format(r['cond'], r['grid'], r['rep']), axis=1))
dfe.head()
dfe
dfe.to_csv('experiments.csv', index=False)
| analysis/spheroid/20190822-co-culture/exp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # The mplot3d Toolkit
#
#
# Generating 3D plots using the mplot3d toolkit.
#
# .. currentmodule:: mpl_toolkits.mplot3d
# :backlinks: none
#
#
# Getting started
# ---------------
# An Axes3D object is created just like any other axes using
# the projection='3d' keyword.
# Create a new :class:`matplotlib.figure.Figure` and
# add a new axes to it of type :class:`~mpl_toolkits.mplot3d.Axes3D`::
#
# import matplotlib.pyplot as plt
# from mpl_toolkits.mplot3d import Axes3D
# fig = plt.figure()
# ax = fig.add_subplot(111, projection='3d')
#
# .. versionadded:: 1.0.0
# This approach is the preferred method of creating a 3D axes.
#
# <div class="alert alert-info"><h4>Note</h4><p>Prior to version 1.0.0, the method of creating a 3D axes was
# different. For those using older versions of matplotlib, change
# ``ax = fig.add_subplot(111, projection='3d')``
# to ``ax = Axes3D(fig)``.</p></div>
#
# See the `toolkit_mplot3d-faq` for more information about the mplot3d
# toolkit.
#
#
# Line plots
# ====================
# .. automethod:: Axes3D.plot
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_lines3d_001.png
# :target: ../../gallery/mplot3d/lines3d.html
# :align: center
# :scale: 50
#
# Lines3d
#
#
# Scatter plots
# =============
# .. automethod:: Axes3D.scatter
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_scatter3d_001.png
# :target: ../../gallery/mplot3d/scatter3d.html
# :align: center
# :scale: 50
#
# Scatter3d
#
#
# Wireframe plots
# ===============
# .. automethod:: Axes3D.plot_wireframe
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_wire3d_001.png
# :target: ../../gallery/mplot3d/wire3d.html
# :align: center
# :scale: 50
#
# Wire3d
#
#
# Surface plots
# =============
# .. automethod:: Axes3D.plot_surface
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_surface3d_001.png
# :target: ../../gallery/mplot3d/surface3d.html
# :align: center
# :scale: 50
#
# Surface3d
#
# Surface3d 2
#
# Surface3d 3
#
#
# Tri-Surface plots
# =================
# .. automethod:: Axes3D.plot_trisurf
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_trisurf3d_001.png
# :target: ../../gallery/mplot3d/trisurf3d.html
# :align: center
# :scale: 50
#
# Trisurf3d
#
#
#
# Contour plots
# =============
# .. automethod:: Axes3D.contour
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_contour3d_001.png
# :target: ../../gallery/mplot3d/contour3d.html
# :align: center
# :scale: 50
#
# Contour3d
#
# Contour3d 2
#
# Contour3d 3
#
#
# Filled contour plots
# ====================
# .. automethod:: Axes3D.contourf
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_contourf3d_001.png
# :target: ../../gallery/mplot3d/contourf3d.html
# :align: center
# :scale: 50
#
# Contourf3d
#
# Contourf3d 2
#
# .. versionadded:: 1.1.0
# The feature demoed in the second contourf3d example was enabled as a
# result of a bugfix for version 1.1.0.
#
#
# Polygon plots
# ====================
# .. automethod:: Axes3D.add_collection3d
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_polys3d_001.png
# :target: ../../gallery/mplot3d/polys3d.html
# :align: center
# :scale: 50
#
# Polys3d
#
#
# Bar plots
# ====================
# .. automethod:: Axes3D.bar
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_bars3d_001.png
# :target: ../../gallery/mplot3d/bars3d.html
# :align: center
# :scale: 50
#
# Bars3d
#
#
# Quiver
# ====================
# .. automethod:: Axes3D.quiver
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_quiver3d_001.png
# :target: ../../gallery/mplot3d/quiver3d.html
# :align: center
# :scale: 50
#
# Quiver3d
#
#
# 2D plots in 3D
# ====================
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_2dcollections3d_001.png
# :target: ../../gallery/mplot3d/2dcollections3d.html
# :align: center
# :scale: 50
#
# 2dcollections3d
#
#
# Text
# ====================
# .. automethod:: Axes3D.text
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_text3d_001.png
# :target: ../../gallery/mplot3d/text3d.html
# :align: center
# :scale: 50
#
# Text3d
#
#
# Subplotting
# ====================
# Having multiple 3D plots in a single figure is the same
# as it is for 2D plots. Also, you can have both 2D and 3D plots
# in the same figure.
#
# .. versionadded:: 1.0.0
# Subplotting 3D plots was added in v1.0.0. Earlier version can not
# do this.
#
# .. figure:: ../../gallery/mplot3d/images/sphx_glr_subplot3d_001.png
# :target: ../../gallery/mplot3d/subplot3d.html
# :align: center
# :scale: 50
#
# Subplot3d
#
# Mixed Subplots
#
#
| python/learn/matplotlib/tutorials_jupyter/toolkits/mplot3d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.keras.models import load_model
from tkinter import *
import tkinter as tk
import win32gui
from PIL import ImageGrab, Image, ImageTk
# +
model = load_model('mnist-4-5.h5') #5 is the best till now
def predict_digit(img):
#resize image to 75x75 pixels
img = img.resize((100, 100))
#convert rgb to grayscale
img = img.convert('L')
img = np.array(img)
# print('Image to be predicted...')
# plt.imshow(img, cmap = 'gray')
# plt.show()
#reshaping to support our model input and normalizing
img = img.reshape(1, 100, 100, 1).astype('float32')
img = img/255.0
#predicting the class
res = model.predict([img])
# print(res)
digits_res = res[0].flatten()
digits = digits_res.argsort()[-2:][::-1]
# print(digits)
digits_acc = [digits_res[digits[0]], digits_res[digits[1]]]
# print(digits_acc)
boxes = res[1].flatten()
div = 100
box = np.array([boxes[1], (boxes[0]*100 - 28)/div, (boxes[1]*100 + 28)/div, boxes[0],
boxes[3], (boxes[2]*100 - 28)/div, (boxes[3]*100 + 28)/div, boxes[2]])
boxes = box.reshape((2, 4))
# print(boxes)
return digits, digits_acc, boxes
class App(tk.Tk):
def __init__(self):
tk.Tk.__init__(self)
self.x = self.y = 0
# Creating elements
self.title('Handwritten Digit Recognition and Localization')
self.canvas1 = tk.Canvas(self, width=500, height=500, bg = 'black', cursor="cross")
self.label1 = tk.Label(self, text="Draw..", font=("Helvetica", 48))
self.label2 = tk.Label(self, text="Draw..", font=("Helvetica", 48))
self.classify_btn = tk.Button(self, text = "Recognise", command = self.classify_handwriting)
self.button_clear = tk.Button(self, text = "Clear", command = self.clear_all)
# Grid structure
self.canvas1.grid(row=0, column=0, pady=0, sticky=W, rowspan = 2, columnspan = 1)
self.label1.grid(row=0, column=1,pady=0, padx=0)
self.label2.grid(row = 1, column = 1, pady = 0, padx = 0)
self.classify_btn.grid(row=2, column=1, pady=2, padx = 2)
self.button_clear.grid(row=2, column=0, pady=2)
self.canvas1.bind("<B1-Motion>", self.draw_lines) # event handler for mouse events
def clear_all(self):
self.canvas1.delete("all")
def classify_handwriting(self):
# code to convert drawing on canvas to an image
HWND = self.canvas1.winfo_id() # get the handle of the canvas
rect = win32gui.GetWindowRect(HWND) # get the coordinate of the canvas
self.im = ImageGrab.grab(rect)
# predict what the image is...
digits, acc, boxes = predict_digit(self.im)
w, h = self.im.size
box01 = boxes[0][0] * h # height (y_min)
box02 = boxes[0][1] * w # width (x_min)
box03 = boxes[0][2] * h # height (y_max)
box04 = boxes[0][3] * w # width (x_max)
box11 = boxes[1][0] * h # height (y_min)
box12 = boxes[1][1] * w # width (x_min)
box13 = boxes[1][2] * h # height (y_max)
box14 = boxes[1][3] * w # width (x_max)
self.img = ImageTk.PhotoImage(self.im)
self.canvas1.create_image((w/2, h/2), image=self.img, anchor = "center")
self.canvas1.create_rectangle(box02, box01, box04, box03, outline = 'red')
self.canvas1.create_rectangle(box12, box11, box14, box13, outline = 'red')
string1 = str(digits[0])+ ',' + str(int(acc[0] * 100)) + '%'
string2 = str(digits[1])+ ',' + str(int(acc[1] * 100)) + '%'
self.label1.configure(text = string1)
if acc[1] > 0.1:
self.label2.configure(text = string2)
else:
self.label2.configure(text = '!')
# if box01<=20:
# self.canvas1.create_text(box02+35, box03 +15, fill = 'red', text = string1,
# font="Times 20 italic bold")
# elif box02<=20:
# self.canvas1.create_text(box04-15, box01-15, fill = 'red', text = string1,
# font="Times 20 italic bold")
# else:
# self.canvas1.create_text(box02+35, box01-15, fill = 'red', text = string1,
# font="Times 20 italic bold")
# if box11<=20:
# self.canvas1.create_text(box12+35, box13 +15, fill = 'red', text = string2,
# font="Times 20 italic bold")
# elif box12<=20:
# self.canvas1.create_text(box14-15, box11-15, fill = 'red', text = string2,
# font="Times 20 italic bold")
# else:
# self.canvas1.create_text(box12+35, box11-15, fill = 'red', text = string2,
# font="Times 20 italic bold")
def draw_lines(self, event):
# draw on the canvas
self.x = event.x
self.y = event.y
r= 6 # control the width of strokes
self.canvas1.create_oval(self.x+r, self.y+r, self.x - r, self.y - r, fill='white',outline = 'white')
app = App()
mainloop()
# -
| MNIST-4/App.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3xEPIoQ2VdUI"
# ## Heart Disease Prediction
# + id="2DaqkLrTVlrF"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib import rcParams
from matplotlib.cm import rainbow
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
# + id="XZ2rxf_TWFEV"
from sklearn.neighbors import KNeighborsClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
# + id="aFcM5RpAXMRl"
url='https://raw.githubusercontent.com/krishnaik06/Predicting-Heart-Disease/master/dataset.csv'
df=pd.read_csv(url)
# + id="7Wd9_8O6XYdJ" outputId="6b64925d-8425-4a8f-e03c-21190a588a65" colab={"base_uri": "https://localhost:8080/"}
df.info()
# + id="ASDtMJmsXgV7" outputId="5156f56d-ff97-4ba3-ad11-a26ca7717d00" colab={"base_uri": "https://localhost:8080/", "height": 317}
df.describe()
# + [markdown] id="EQR_F9m-X0WF"
# ## Feature Selection
# + id="_2D0he5TXvb-" outputId="fb9081c4-e8ac-4bdf-ed38-cb0744ad7cc1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import seaborn as sns
#get correlations of each features in dataset
corrmat = df.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(20,20))
#plot heat map
g=sns.heatmap(df[top_corr_features].corr(),annot=True,cmap="RdYlGn")
# + id="--ZzR2WVY1Fv" outputId="cb5e2208-2bb9-45f5-cc48-df3ac5b7e2d7" colab={"base_uri": "https://localhost:8080/", "height": 570}
df.hist()
# + id="XOualfjTarKs" outputId="ffc2bf05-6d96-48dd-9b92-0ed93a2f21fa" colab={"base_uri": "https://localhost:8080/", "height": 296}
sns.set_style('whitegrid')
sns.countplot(x='target',data=df,palette='RdBu_r')
# + id="OacqB8Coa4gK"
dataset = pd.get_dummies(df, columns = ['sex', 'cp', 'fbs', 'restecg', 'exang', 'slope', 'ca', 'thal'])
# + id="FvaC1DcNbBqY"
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
standardScaler = StandardScaler()
columns_to_scale = ['age', 'trestbps', 'chol', 'thalach', 'oldpeak']
dataset[columns_to_scale] = standardScaler.fit_transform(dataset[columns_to_scale])
# + id="GB6KylUKbPPb" outputId="b8d6a598-b956-4def-98ec-a9a119de51b9" colab={"base_uri": "https://localhost:8080/", "height": 224}
dataset.head()
# + id="gDjZ7mhmbTnJ"
y= dataset['target']
X= dataset.drop(['target'],axis=1)
# + id="sIhPt9ASbzh8"
from sklearn.model_selection import cross_val_score
knn_scores = []
for k in range(1,21):
knn_classifier = KNeighborsClassifier(n_neighbors = k)
score=cross_val_score(knn_classifier,X,y,cv=10)
knn_scores.append(score.mean())
# + id="sulLs57vb5nV" outputId="c4453b91-f655-4444-f75c-08c4904f5311" colab={"base_uri": "https://localhost:8080/", "height": 312}
plt.plot([k for k in range(1, 21)], knn_scores, color = 'red')
for i in range(1,21):
plt.text(i, knn_scores[i-1], (i, knn_scores[i-1]))
plt.xticks([i for i in range(1, 21)])
plt.xlabel('Number of Neighbors (K)')
plt.ylabel('Scores')
plt.title('K Neighbors Classifier scores for different K values')
# + id="agL8pgIbcTxh"
knn_classifier = KNeighborsClassifier(n_neighbors = 12)
score=cross_val_score(knn_classifier,X,y,cv=10)
# + id="Ver776AccXRp" outputId="c9abdfee-f712-47e1-ba41-ffae4eeac0f4" colab={"base_uri": "https://localhost:8080/"}
score.mean()
# + [markdown] id="Bj8KwAVac1sl"
# # Random Forest Classifier
# + id="9xtk8dnmcapU"
from sklearn.ensemble import RandomForestClassifier
# + id="xHwmLHxDdC7K"
randomforest_classifier= RandomForestClassifier(n_estimators=10)
score=cross_val_score(randomforest_classifier,X,y,cv=10)
# + id="TKC76vaSdIyY" outputId="951f8b6c-617e-4c01-b966-48d9c03ce3c7" colab={"base_uri": "https://localhost:8080/"}
score.mean()
# + [markdown] id="mzXCYkcUdQlk"
# # Decision Tree Classifier
# + id="kIVvmU9ndWrk"
from sklearn import tree
# + id="d06kFvyteLlw"
decisiontree_classifier = tree.DecisionTreeClassifier(random_state=200,max_depth=3,min_samples_leaf=5)
score=cross_val_score(decisiontree_classifier,X,y,cv=10)
# + id="EmjESlTDe08J" outputId="9c9a65eb-09b2-4f35-deb2-7805a8b9b18a" colab={"base_uri": "https://localhost:8080/"}
score.mean()
# + id="ePmVpYUee3NK"
| Heart_Disease_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PyFME Validation : comparing response with eigenvalue analysis
from pyfme.aircrafts import LinearB747, Cessna172, SimplifiedCessna172
from pyfme.models import EulerFlatEarth
import numpy as np
nl = np.linalg
import matplotlib.pyplot as plt
from pyfme.environment.atmosphere import ISA1976, SeaLevel
from pyfme.environment.wind import NoWind
from pyfme.environment.gravity import VerticalConstant
from pyfme.environment import Environment
from pyfme.utils.trimmer import steady_state_trim
from pyfme.models.state.position import EarthPosition
from pyfme.simulator import Simulation
from pyfme.utils.coordinates import wind2body, body2wind
from pyfme.utils.input_generator import Constant
# %load_ext autoreload
# %autoreload 2
# We start by defining the airplane, the environment and the trim position.
aircraft = SimplifiedCessna172()
atmosphere = SeaLevel()
gravity = VerticalConstant()
wind = NoWind()
environment = Environment(atmosphere, gravity, wind)
pos = EarthPosition(x=0, y=0, height=1000)
psi = 0.5 # rad
TAS = 45 # m/s
controls0 = {'delta_elevator': 0, 'delta_aileron': 0, 'delta_rudder': 0, 'delta_t': 0.5}
trimmed_state, trimmed_controls = steady_state_trim(
aircraft,
environment,
pos,
psi,
TAS,
controls0
)
environment.update(trimmed_state)
# Then we linearize the model around the trim condition. It gives us two matrices for lateral and longitudinal small perturbations.
# Under the hood, the code computes dimensional stability derivatives using numerical differentiation, and then uses the analytical formulas for the linearized system given in [1].
system = EulerFlatEarth(t0=0, full_state=trimmed_state)
A_long, A_lat = system.linearized_model(trimmed_state, aircraft, environment, trimmed_controls)
long_val, long_vec=nl.eig(A_long)
long_val = np.expand_dims(long_val, axis = 0)
print(f"Longitudinal eigenvalues : ")
for l in long_val[0]:
print(l)
# As expected we find two damped oscillatory modes in the longitudinal dynamics: phugoid and short period.
lat_val, lat_vec=nl.eig(A_lat)
lat_val = np.expand_dims(lat_val, axis = 0)
print(f"Lateral eigenvalues : ")
for l in lat_val[0]:
print(l)
# In the lateral case, we get one oscillatory mode (dutch roll), a stable rolling convergence and an unstable - but very slow - spiral mode.
# # Eigenvalue trajectories
# We can compute the predicted trajectories for small perturbations.
# helper function to go from stability to body axis
def linear_stab_2_body(long_state=np.zeros(4), lat_state=np.zeros(4), u0=0, theta0=0,alpha0=0, beta0=0):
# velocities
v = wind2body(np.array([long_state[0] + u0, lat_state[0], long_state[1]]), alpha=alpha0, beta=beta0)
# Roll rates
r = wind2body(np.array([lat_state[1], long_state[2], lat_state[2]]), alpha=alpha0, beta=beta0)
long_stateB = np.copy(long_state)
lat_stateB = np.copy(lat_state)
long_stateB[0] = v[0]
long_stateB[1] = v[2]
long_stateB[2] = r[1]
long_stateB[3] += theta0
lat_stateB[0] = v[1]
lat_stateB[1] = r[0]
lat_stateB[2] = r[2]
return long_stateB.real, lat_stateB.real
# Reference conditions
alpha = np.arctan2(trimmed_state.velocity.w, trimmed_state.velocity.u)
beta = np.arcsin(trimmed_state.velocity.v/nl.norm(trimmed_state.velocity.vel_body))
u, v, w = body2wind(trimmed_state.velocity.vel_body, alpha, 0)
theta0 = trimmed_state.attitude.theta*1.0
# ## Longitudinal case
# We can pick any perturbation from the equilibrium. Here I chose something along the eigenvectors of A_long.
# The result will be a weighted sum of eigenvector*exp(eigenvalue*t). The weights are determined by the initial condition.
long_perturbation = (long_vec.T[2] + long_vec.T[3])/1000
C = nl.lstsq(a=long_vec,b=long_perturbation.real)[0].real
T = 100;
t_long = np.linspace(0,T,1000)
N = len(t_long)
X_long = np.zeros((N,4))
for i in range(N):
x_stab = (long_vec*np.exp(long_val*t_long[i])).dot(C)
X_long[i,:] = linear_stab_2_body(long_state=x_stab.real, alpha0=alpha, u0=u, theta0 = theta0)[0]
# ## Lateral case
# Again we can pick any perturbation
lat_perturbation = (lat_vec.T[0])
C = nl.lstsq(a=lat_vec,b=lat_perturbation.real)[0].real
t_lat = np.linspace(0,10,100)
N = len(t_lat)
X_lat = np.zeros((N,4))
for i in range(N):
x_stab = (lat_vec*np.exp(lat_val*t_lat[i])).dot(C)
X_lat[i,:] = linear_stab_2_body(lat_state=x_stab.real, beta0=beta, alpha0=alpha, u0=u, theta0 = theta0)[1]
# # Simulation
controls = {
'delta_elevator': Constant(trimmed_controls['delta_elevator']),
'delta_aileron': Constant(trimmed_controls['delta_aileron']),
'delta_rudder': Constant(trimmed_controls['delta_rudder']),
'delta_t': Constant(trimmed_controls['delta_t'])
}
# ## Longitudinal case
# We perturbate the trimmed state and run the simulation
# Perturbate the trimmed state
trimmed_state.cancel_perturbation()
p = linear_stab_2_body(long_state=long_perturbation.real, alpha0=alpha)[0]
trimmed_state.perturbate(np.array([p[0],0,p[1]]), 'velocity')
trimmed_state.perturbate(np.array([0,p[2],0]), 'angular_vel')
trimmed_state.perturbate(np.array([p[3],0,0]), 'attitude') # /!\ Convention theta, phi, psi
trimmed_state.save_to_json('long_perturb.json')
environment.update(trimmed_state)
system = EulerFlatEarth(t0=0, full_state=trimmed_state)
sim = Simulation(aircraft, system, environment, controls)
r_long = sim.propagate(100)
trimmed_state.cancel_perturbation();
# ## Lateral case
# Perturbate the trimmed state
trimmed_state.cancel_perturbation()
p = linear_stab_2_body(lat_state=lat_perturbation.real, alpha0=alpha, beta0=beta)[1]
trimmed_state.perturbate(np.array([0,p[0],0]), 'velocity')
trimmed_state.perturbate(np.array([p[1],0,p[2]]), 'angular_vel')
trimmed_state.perturbate(np.array([0,p[3],0]), 'attitude') # /!\ Convention theta, phi, psi
environment.update(trimmed_state)
system = EulerFlatEarth(t0=0, full_state=trimmed_state)
sim = Simulation(aircraft, system, environment, controls)
r_lat = sim.propagate(10)
trimmed_state.cancel_perturbation();
# # Compare outputs
# +
# Longitudinal case
plt.plot(t_long, X_long[:,0], '.', label='eigenvalue analysis')
plt.plot(r_long.u, label='PyFME simulation')
plt.legend()
plt.title("Horizontal velocity")
plt.show()
plt.plot(t_long, X_long[:,1], '.', label='eigenvalue analysis')
plt.plot(r_long.w, label='PyFME simulation')
plt.legend()
plt.title("Vertical velocity")
plt.show()
plt.plot(t_long, X_long[:,2], '.', label='eigenvalue analysis')
plt.plot(r_long.q, label='PyFME simulation')
plt.legend()
plt.title("Pitch rate")
plt.show()
plt.plot(t_long, X_long[:,3], '.', label='eigenvalue analysis')
plt.plot(r_long.theta, label='PyFME simulation')
plt.legend()
plt.title("Pitch angle")
plt.show()
# +
# Lateral case
plt.plot(t_lat, X_lat[:,0], '.', label='eigenvalue analysis')
plt.plot(r_lat.v, label='PyFME simulation')
plt.legend()
plt.title("Lateral velocity")
plt.show()
plt.plot(t_lat, X_lat[:,1], '.', label='eigenvalue analysis')
plt.plot(r_lat.p, label='PyFME simulation')
plt.legend()
plt.title("Roll rate")
plt.show()
plt.plot(t_lat, X_lat[:,2], '.', label='eigenvalue analysis')
plt.plot(r_lat.r, label='PyFME simulation')
plt.legend()
plt.title("Yaw rate")
plt.show()
plt.plot(t_lat, X_lat[:,3], '.', label='eigenvalue analysis')
plt.plot(r_lat.phi, label='PyFME simulation')
plt.legend()
plt.title("Heading angle")
plt.show()
# -
aircraft.inertia
# # References
# Dynamics of Flight, Stability and Control, Etkin and Reid
| .ipynb_checkpoints/PyFME vs Eigenvalue analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="figures/DLlogosmall.png">
#
# *This notebook contains an excerpt from the [Deep Learning with Tensorflow 2.0](https://www.adhiraiyan.org/DeepLearningWithTensorflow.html) by <NAME>. The code is released under the [MIT license](https://opensource.org/licenses/MIT) and is available for FREE [on GitHub](https://github.com/adhiraiyan/DeepLearningWithTF2.0).*
#
# *Open Source runs on love, laughter and a whole lot of coffee. Consider buying me [one](https://www.buymeacoffee.com/mmukesh) if you find this content useful!*
#
#
# <!--NAVIGATION-->
# < [03.00 - Probability and Information Theory](03.00-Probability-and-Information-Theory.ipynb) | [Contents](Index.ipynb) |
#
# <a href="https://colab.research.google.com/github/adhiraiyan/DeepLearningWithTF2.0/blob/master/notebooks/04.00-Numerical-Computation.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# + [markdown] colab_type="text" id="DF-azaVjrT3v"
# # 04.00 - Numerical Computation
# + [markdown] colab_type="text" id="WCLFmnilZJC6"
# We now use computers for a wide range of reasons, from watching movies to reading books to playing games, but originally computers were designed and used to solve computational problems.
#
# Numerical analysis or scientific computing is defines as the study of approximation techniques for numerically solving mathematical problems.
#
# __Numerical Computation__ is necessary for problem solving in that very few mathematical problems have a closed form solution. If an equation solves a given problem in terms of functions and mathematical operations from a given generally-accepted set in a finite number of standard operations, it is said to be closed form. But since most of the problems we deal in real life are non closed form, we use numerical methods to solve it.
#
# Linear equations, linear programming, optimization and numerical partial differential equations are main branches of numerical computation. These may sound far off from what you deal with in daily life so let me give you few examples, you know how plane ticket prices seem to go up whenever they want, that is an optimization problem, Google's page rank that ranks web pages is an eigenvector of a matrix of order about 3 billion, all these problems are solved using numerical computation. We will look at some of those methods in this chapter.
#
# Optimization and solving systems of linear equations is at the heart of almost all machine learning and statistical techniques. These algorithms usually require a high amount of numerical computation. These evaluations can be difficult when the function involves real numbers, which can't be represented precisely using a finite amount of memory, which brings us to our first section.
# + colab={} colab_type="code" id="0x5dgEesF1XN"
"""
At the moment of writing (06.06.2019) the only build that supports tensorflow probability is the tensorflow
nightly build so we will use that to install tensorflow 2.0 and tensorflow probability.
"""
# Install tensorflow 2.0 and tensorflow probability from the nightly build
# !pip install --upgrade tf-nightly-2.0-preview tfp-nightly
# + colab={} colab_type="code" id="05SSSlwwSW2f"
# Imports
import tensorflow as tf
import tensorflow_probability as tfp
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
# plt axis colors setup
plt.rc_context({'axes.edgecolor':'orange', 'xtick.color':'red', 'ytick.color':'red', 'text.color':'orange'})
# + [markdown] colab_type="text" id="hQYa3NWorU1v"
# # 04.01 - Overflow and Underflow
# + [markdown] colab_type="text" id="TZIi9BksaDMc"
# Representing infinitely many real numbers with a finite number of bit patterns represents a fundamental difficulty in performing continuous math on a digital computer. This means that for almost all real numbers we incur some approximation error in the form of rounding error. Rounding error is problematic when it compounds across many operations, and can cause algorithms that work in theory to fail in practice if they are not designed to minimize the accumulation of rounding error.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="vG3Nnyq_rVoC" outputId="6358e9d6-cd32-41e3-e2c7-aef8e7038a89"
"""
In the same way, no matter how many base 2 digits you’re willing to use, the decimal value 0.1 cannot be represented
exactly as a base 2 fraction. In base 2, 1/10 is the infinitely repeating fraction
0.0001100110011001100110011001100110011001100110011...
One illusion may beget another. For example, since 0.1 is not exactly 1/10, summing three values of 0.1 may not
yield exactly 0.3, either:
Also, since the 0.1 cannot get any closer to the exact value of 1/10 and 0.3 cannot get any closer to the exact
value of 3/10, then pre-rounding with round() function cannot help:
Though the numbers cannot be made closer to their intended exact values, the round() function can be useful for
post-rounding so that results with inexact values become comparable to one another:
Binary floating-point arithmetic holds many surprises like this.
"""
x = 0.1
y = 0.2
print("x + y = {}".format(x + y))
print("Rounded x + y: {}".format(round(x + y, 1)))
print("Check if .1 + .1 +.1 == .3: {}".format(1 + .1 + .1 == .3))
print("What if we pre round before adding: {}".format(round(.1, 1) + round(.1, 1) + round(.1, 1) == round(.3, 1)))
print("What if we post round after adding: {}".format(round(.1 + .1 + .1, 10) == round(.3, 10)))
# + [markdown] colab_type="text" id="0eH-1wLub-VN"
# __Underflow__: occurs when numbers near zero are rounded to zero. This can be particularly devastating, think of division by zero, some software environments will raise an exception but other will result with a placeholder not a number value.
#
# __Overflow__: occurs when numbers with large magnitude are approximated as $\infty$ or $- \infty$.
#
# One function that must be stabilized against underflow and overflow is the __softmax function__:
#
# $$\color{orange}{\text{softmax}(x)_i = \frac{exp(x_j)}{\sum_{j=1}^n exp(x_j)} \tag{1}}$$
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="hoFPmxjob_AS" outputId="69e6d670-f8da-4bb0-abbc-66bd95f1b704"
def softmax(x, solve=False):
"""Softmax implementation"""
if solve:
z = x-max(x)
else:
z = x
numerator = tf.math.exp(z)
denominator = tf.math.reduce_sum(numerator)
return tf.divide(numerator, denominator)
# Underflow example
"""
If c is very negative, exp(c) will underflow, meaning the denominator will become 0
"""
underflow = tf.constant([-12345, -67890, -99999999], dtype=tf.float32)
print("Softmax Underflow {}".format(softmax(underflow, solve=False)))
# Overflow example
"""
When c is very large and positive, exp(c) will overflow and the expression ends up being undefined
"""
overflow = tf.constant([12345, 67890, 99999999], dtype=tf.float32)
print("Softmax Overflow {}".format(softmax(overflow, solve=False)))
# Solution
"""
Both of these can be solved by evaluating softmax(z) where z = x - max_i x_i.
This works because subtracting max results in the largest argument to exp being 0, getting rid of overflow
and atleast one term in the denominator has a value of 1, which gets rid of underflow
Compare the overflow and underflow examples
"""
underflow = tf.constant([-12345, -67890, -99999999], dtype=tf.float32)
print("Underflow Solved: {}".format(softmax(underflow, solve=True)))
overflow = tf.constant([12345, 67890, 99999999], dtype=tf.float32)
print("Overflow Solved: {}".format(softmax(overflow, solve=True)))
# compare the solution with the tensorflow softmax implementation
underflow_softmax_tf = tf.nn.softmax(underflow, axis=None)
overflow_softmax_tf = tf.nn.softmax(overflow, axis=None)
print("Tensorflow Softmax Underflow: {} \nTensorflow Softmax Overflow: {}".format(underflow_softmax_tf, overflow_softmax_tf))
# + [markdown] colab_type="text" id="d3uIn1lPrZ6-"
# # 04.02 - Poor Conditioning
# + [markdown] colab_type="text" id="TSwxWe30iCyb"
# Conditioning refers to how rapidly a function changes with respect to small changes in its inputs. Functions that change rapidly when their inputs are perturbed slightly can be problematic for scientific computation because rounding errors in the inputs can result in large changes in the output.
#
# For example, the function $f(x) = A^{-1}x$. When $A \in \mathbb{R}^{n \times n}$ has an eigenvalue decomposition, its __condition number__ is:
#
# $$\color{orange}{max_{i, j} \Biggr | \frac{\lambda_i}{\lambda_j} \Biggr | \tag{2}}$$
#
# This is the ratio of the magnitude of the largest and smallest eigenvalue. When this number is large, matrix inversion is particularly sensitive to error in the input.
#
# This sensitivity is an intrinsic property of the matrix itself, not the result of rounding error during matrix inversion. Poorly conditioned matrices amplify pre-existing errors when we multiply by the true matrix inverse. In practice, the error will be compounded further by numerical errors in the inversion process itself.
# + colab={"base_uri": "https://localhost:8080/", "height": 255} colab_type="code" id="1UEpG4o-rapB" outputId="de8672b3-7b40-44d1-cf7d-a86e1841209b"
A = tf.constant([[4.1, 2.8], [9.7, 6.6]], dtype=tf.float32)
b = tf.constant([[4.1], [9.7]], dtype=tf.float32)
print("Matrix A: \n{}\n".format(A))
# solve for x, from Ax=b, x = A^(-1) b
x = tf.linalg.matmul(tf.linalg.inv(A), b)
print("Value of x: \n{}\n".format(x))
# Now lets see what happens if we add 0.01 to the first component of b
b2 = tf.constant([[4.11], [9.7]], dtype=tf.float32)
# We can also use tf.linalg.solve to solve a systems of linear equations
x_solve = tf.linalg.solve(A, b2)
print("Solving for new x: \n{}".format(x_solve))
print("We can see how the solution changes dramatically for a small increase in value\n")
# let's now calculate the condition number for matrix A using ||A|| * ||A^-1||
condition_A = tf.math.multiply(tf.norm(A), tf.norm(tf.linalg.inv(A)))
print("Condition Number of A: {}".format(condition_A))
# + [markdown] colab_type="text" id="o7BG2Q6_rbI2"
# # 04.03 - Gradient-Based Optimization
# + [markdown] colab_type="text" id="BFlhfprht5BQ"
# Most deep learning algorithms involve optimization of some sort. Optimization refers to the task of either minimizing or maximizing some function $f(x)$ by altering $x$. We usually phrase most optimization problems in terms of minimizing $f(x)$. Maximization may be accomplished via a minimization algorithm by minimizing $-f(x)$.
#
# The function we want to minimize or maximize is called the __objective function__ or __criterion__. When we are minimizing it, we may also call it the __cost function, loss function__, or __error function__.
#
# We often denote the value that minimizes or maximizes a function with a superscript *. For example we might say $x^* = arg \ min \ f(x)$.
#
#
# + [markdown] colab_type="text" id="3DO2wa2-vJ1T"
# Suppose we have a function $y = f(x)$, where both _x_ and _y_ are real numbers. The __derivative__ of this function is denoted as $f^{'}(x)$ or as $\frac{dy}{dx}$. The derivative $f^{'}(x)$ gives the slope of $f(x)$ at the point _x_. In other words, it specifies how to scale a small change in the input to obtain the corresponding change in the output:
#
# $$\color{orange}{f(x + \epsilon) \approx f(x) + \epsilon f^{'}(x) \tag{3}}$$
# + [markdown] colab_type="text" id="QyWiLUPEyPeY"
# __Gradient descent__ is the technique of moving in small steps with the opposite sign of the derivative to reduce $f(x)$.
#
#
# 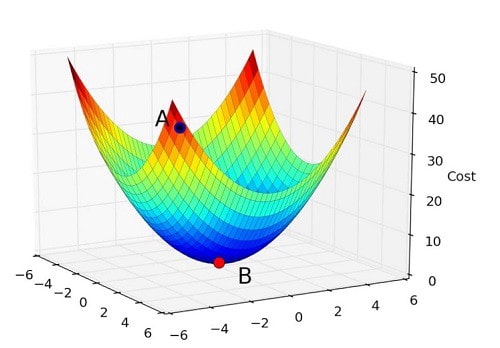
#
# + [markdown] colab_type="text" id="lKdh5_kZ3tMA"
# - When $f^{'}(x) = 0$, the derivative provides no information about which direction to move. These points are known as __critical points__, or __stationary points__.
#
# - A __local minimum__ is a points where $f(x)$ is lower than all neighboring points, so it is no longer possible to decrease $f(x)$ by making infinitesimal steps.
#
# - A __local maximum__ is a point where $f(x)$ is higher than all neighboring points, so it is not possible to increase $f(x)$ by making infinitesimal steps.
#
# - Some critical points are neither maxima or minima, these are knows as __saddle points__.
#
# - A point that obtains the absolute lowest value of $f(x)$ is a __global minimum__.
# + colab={"base_uri": "https://localhost:8080/", "height": 500} colab_type="code" id="8rP-VIP3r7OY" outputId="41d5568d-c644-4be6-8f05-deea5f5d97c3"
"""
Let's see how these look like by plotting these. Note that I am using numpy for these plots but we will start
using tensorflow in the upcoming sections. Also, taking the derivatives and most optimizations are
easily done with packages like numpy, scipy or sympy since these are dedicated scientific libraries
whereas tensorflow is a machine learning framework but I am working on bringing a scientific extension to the
tensorflow, please see (https://github.com/mukeshmithrakumar/scientific).
This is work in progress and contributions are welcome.
"""
polynomial = np.poly1d([2,-4,-28,62,122,-256,-196,140,392,240,72]) # Create a one-dimensional polynomial class
polynomial_root = polynomial.deriv().r # Return a derivative of this polynomial with the roots
r_crit = polynomial_root[polynomial_root.imag==0].real # Return the real part of the complex argument
test = polynomial.deriv(2)(r_crit)
# local minima
x_min = r_crit[test>0]
y_min = polynomial(x_min)
plt.plot(x_min, y_min, 'o')
plt.text(1.5, 200, 'local minima')
# local maxima
x_max = r_crit[test<0]
y_max = polynomial(x_max)
plt.plot(x_max, y_max, 'o', color='r')
plt.text(0.5, 800, 'local maxima')
#global maxima
plt.text(-1.5, 1600, 'global maxima')
# global minima
xc = np.arange(-2.5, 2.6, 0.02)
yc = polynomial(xc)
plt.plot( xc, yc)
plt.xlim([-2.5,2.5])
plt.text(-1, 400, 'global minima')
# Saddle Point
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
plot_args = {'rstride': 1, 'cstride': 1, 'cmap':"Blues_r",
'linewidth': 0.4, 'antialiased': True,
'vmin': -1, 'vmax': 1}
x, y = np.mgrid[-1:1:31j, -1:1:31j]
z = x**2 - y**2
ax.plot_surface(x, y, z, **plot_args)
ax.view_init(azim=-60, elev=30)
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
ax.set_zlim(-1, 1)
plt.xticks([-1, -0.5, 0, 0.5, 1], [r"$-1$", r"$-1/2$", r"$0$", r"$1/2$", r"$1$"])
plt.yticks([-1, -0.5, 0, 0.5, 1],[r"$-1$", r"$-1/2$", r"$0$", r"$1/2$", r"$1$"])
ax.set_zticks([-1, -0.5, 0, 0.5, 1])
ax.set_zticklabels([r"$-1$", r"$-1/2$", r"$0$", r"$1/2$", r"$1$"])
ax.scatter([0], [0], [.3], s=100, color = 'r', marker="o")
plt.title("Saddle Point")
plt.show()
# + [markdown] colab_type="text" id="QxZLNQfM45Az"
# In deep learning we often deal with multidimensional inputs with functions that may have many local minima that are not optimal and many saddle points surrounded by very flat regions. This makes optimization difficult, we therefore usually settle for finding a value of $f$ that is very low but not necessarily minimal in any formal sense.
#
# For functions with multiple inputs, we must make use of the concept of __partial derivatives__. The partial derivative $\frac{\partial}{\partial x_i} f(x)$ measures how $f$ changes as only the variable $x_i$ increases at point __x__. The __gradient__ generalizes the notion of derivative to the case where the derivative is with respect to a vector: the gradient of $f$ is the vector containing all the partial derivatives, denoted $\nabla_x f(x)$. Element $i$ of the gradient is the partial derivative of $f$ with respect to $x_i$.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="sSCjTBlO45am" outputId="e9c30869-e50b-4227-b485-2081b75a41b9"
"""
Lets say we have a function z = f(x, y) = 2x - y: The partial derivatives of this equation is:
dz_dx = 2, dz_dy = -1. Let's see how we can calculate this in tensorflow using GradientTape.
GradientTape records operations for automatic differentiation and we use it with persistent=True
to allow multiple calls to the gradient(). Note that since we are using constants, we need to
ensure that these are being traced when we do the GradientTape, hence we use a watch method.
But when we start working with variables, we don't need to have a watch.
"""
x = tf.constant(1.0)
y = tf.constant(2.0)
with tf.GradientTape(persistent=True) as g:
g.watch(y)
g.watch(x)
out = tf.subtract(2*x, y)
dz_dx = g.gradient(out, x)
dz_dy = g.gradient(out, y)
del g # Drop the reference to the tape
print("Partial Derivative of dz_dx: {} \nPartial Derivative of dz_dy: {} ".format(dz_dx, dz_dy))
# + [markdown] colab_type="text" id="LHTc_dDB45u7"
# The __directional derivative__ in direction $u$ (a unit vector) is the slope of the function $f$ in direction $u$. In other words, the directional derivative is the derivative of the function $f(x + \alpha u)$ with respect to $\alpha$, evaluated at $\alpha = 0$. Using the chain rule, we can see that $\frac{\partial}{\partial \alpha} f(x + \alpha u)$ evaluates to $u^{\top} \nabla_x f(x)$ when $\alpha = 0$.
# + [markdown] colab_type="text" id="YwOYvtojQ5DE"
# To minimize $f$, we would like to find the direction in which $f$ decreases the fastest. We can do this using the directional derivative:
#
# $$\color{orange}{min_{u, u^{\top} \ u=1} u^{\top} \nabla_x f(x) \tag{4}}$$
#
# $$\color{orange}{=min_{u, u^{\top} \ u=1} \|u\|_2 \ \| \nabla_x f(x) \|_2 \ cos \theta \tag{5}}$$
#
# where $\theta$ is the angle between $u$ and the gradient. Substituting in $\|u\|_2 = 1$ and ignoring factors that do not depend on $u$, this simplifies to $min_u cos \theta$. This is minimized when $u$ points in the opposite direction as the gradient. in other words, the gradient points directly uphill, and the negative gradient points directly downhill. We can decrease $f$ by moving in the direction of the negative gradient. This is known as the __method of steepest descent__ or __gradient descent__.
#
#
# Well, if you take away all the math, gradient descent simply states that if you move in the opposite direction of a functions gradient, you will be able to decrease it. Think about it this way, if you have climbed a mountain and you want to hike down, you know the fastest way down is from the way the mountain is steepest right. If you have skied, one of the fastest ways to reach the bottom is by skiing from the steepest parts. This is exactly what we do, we find the steepest part and then move in the opposite direction.
#
#
# Steepest descent proposes a new point:
#
# $$\color{orange}{x^{'} = x - \epsilon \nabla_x f(x) \tag{6}}$$
#
# where $\epsilon$ is the __learning rate__, a positive scalar determining the size of the step.
# + colab={"base_uri": "https://localhost:8080/", "height": 439} colab_type="code" id="ANTmu6Ric-C3" outputId="c8906429-3a74-418f-be69-c5ab4260297d"
"""
f(x,y) = x^2 + y^2 + xsiny + ysinx
f'(x, y) = (2 x + y cos(x) + sin(y)) dx + (2 y + x cos(y) + sin(x)) dy
If you know chain rule, this can be easily differentiated but if you don't, you don't have to worry, tensorflow
will take care of any kind of function for you using Gradient Tape
"""
def f(x, y):
return tf.pow(x, 2) + tf.pow(y, 2) + tf.multiply(x, tf.math.sin(y)) + tf.multiply(y, tf.math.sin(x))
def grad(x, y):
with tf.GradientTape() as t:
t.watch(x)
out = f(x, y)
return t.gradient(out, x)
a = 0.1 # learning rate
x0 = tf.constant(1.0)
y0 = tf.constant(1.0)
print("Starting x value: {}".format(x0))
update = []
for i in range(41):
x0 -= a * grad(x0, y0)
update.append(x0)
plt.plot(i, update[-1], color='r', marker='.')
if i%5 == 0:
print("Iteration: {} x: {}".format(i, x0))
plt.grid()
# + [markdown] colab_type="text" id="-dJzWgUHl0dn"
# Below is an animation of how our gradient takes small steps and finds the global minimum.
#
# 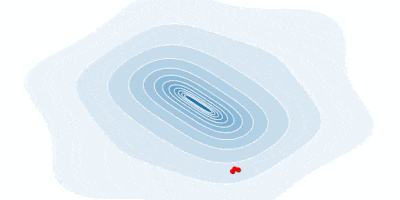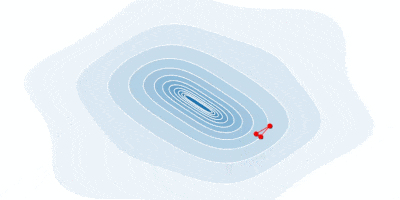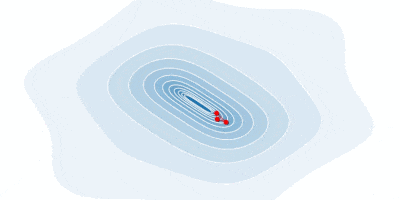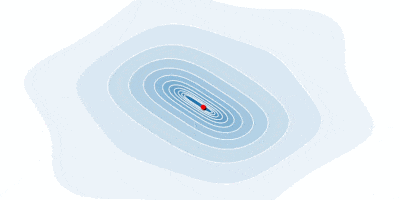
#
# In some cases, we may be able to avoid running this iterative algorithm and just jump directly to the critical point by solving the equation $\nabla_x f(x) = 0$ for $x$.
#
# Although gradient descent is limited to optimization in continuous spaces, the general concept of repeatedly making a small move (that is approximately the best small move) toward better configurations can be generalized to discrete spaces. Ascending an objective function of discrete parameters is called __hill climbing__.
# + [markdown] colab_type="text" id="eUj0maERUrX0"
# Haha, I had to put this, the perfect analogy for gradient descent would be have you seen these videos where people roll down the hill,
#
# 
#
# Gradient descent is exactly that, not everyone reaches the bottom (global minimum), they get stuck in pits (local minimum), where you start from the top (initial values) plays a huge part in the path you take and how fast you roll down (learning rate) determines how fast you reach the bottom and if you could potentially jump out of the pits. If you haven't watched these videos, take a look at [this](https://www.youtube.com/watch?v=fiaWyG7iKes) btw, that is just crazy.
# + [markdown] colab_type="text" id="cPF1brGEmjSQ"
# ## 4.3.1 Beyond the Gradient: Jacobian and Hessian Matrices
# + [markdown] colab_type="text" id="wggGPsxTU681"
# Suppose we need to find all the partial derivatives of a function whose input and output are both vectors. The matrix containing all such partial derivatives is known as a __Jacobian matrix__. Specifically, if we have a function $f : \mathbb{R}^m \rightarrow \mathbb{R}^n$, then the Jacobian matrix $J \in \mathbb{R}^{n \times m}$ of $f$ is defined such that $J_{i, j} = \frac{\partial}{\partial x_j} f(x)$.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="KmaMBMQEbI8o" outputId="12313714-7eec-494f-97b4-e995cce971eb"
"""
Using Tensorflow GradientTape, we can easily find the Jacobian of any vector using "jacobian(target, sources)".
In this example for f(x) = x^2 we will look at the Jacobian when x = [1.0, 2.0]
"""
with tf.GradientTape() as g:
x = tf.constant([1.0, 2.0])
g.watch(x)
y = x * x
jacobian = g.jacobian(y, x)
print("Jacobian \n{}".format(jacobian))
del g # Drop the reference to the tape
# + [markdown] colab_type="text" id="ux1iEgS4VAOY"
# We are also sometimes interested in a derivative of a derivative. This is known
# as a second derivative. For example, for a function $f : \mathbb{R}^n \rightarrow \mathbb{R}$, the derivative with respect to $x_i$ of the derivative of $f$ with respect to $x_j$ is denoted as $\frac{\partial^2}{\partial x_i \partial x_j} f$. In a single dimension we denote $\frac{d^2}{dx^2} f$ by $f^{''}(x)$. The second derivative tells
# us how the first derivative will change as we vary the input. This is important because it tells us whether a gradient step will cause as much of an improvement as we would expect based on the gradient alone. We can think of the second derivative as measuring __curvature__.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="6zwVXOwAGfm-" outputId="07e8cf3e-f4b7-4836-cace-311cba7ea817"
"""
We will use tf.GradientTape() to calculate the second derivative of the function
f(x) = x^3
f'(x) = 3x^2
f''(x) = 6x
"""
x = tf.Variable(1.0)
with tf.GradientTape() as t:
with tf.GradientTape() as t2:
y = x * x * x
dy_dx = t2.gradient(y, x)
d2y_dx2 = t.gradient(dy_dx, x)
print("First Derivative of f(x): {} \nSecond Derivative of f(x): {}".format(dy_dx, d2y_dx2))
del t, t2 # Drop the reference to the tape
# + [markdown] colab_type="text" id="yZ_kHUupLPTZ"
# Below is a plot of our function f(x) = x^3, f'(x) = 3x^2 and f''(x)=6x:
#
# 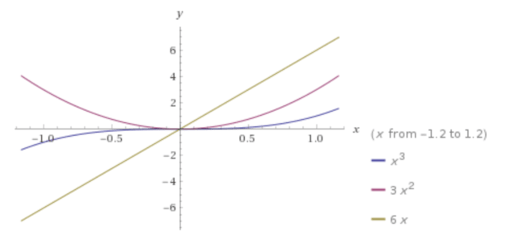
#
# + [markdown] colab_type="text" id="GQf7lXob0s5N"
# The second derivative can be used to determine whether a critical point is
# a local maximum, a local minimum, or a saddle point.
#
# - When $f^{'}(x) = 0$ and $f^{''}(x) > 0$, we can conclude that $x$ is a local minimum.
#
# - When $f^{'}(x) = 0$ and $f^{''}(x) < 0$, we can conclude that $x$ is a local maximum.
#
# - If $f^{''}(x) = 0$ then $x$ may be a saddle point or a part of a flat region.
#
# + [markdown] colab_type="text" id="UQhqZG7BVA4k"
# - If the second derivative is zero, then there is no curvature. It is a perfectly flat line, and its value can be predicted using only the gradient. If the gradient is 1, then we can make a step of size $\epsilon$ along the negative gradient, and the cost function will decrease by $\epsilon$.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="lY56ugKxQF7b" outputId="43f66c6d-0f3a-4c8a-df0f-b6c0ab46e4be"
# Plot where second derivative is zero
x0 = tf.range(-15., 15., 1.)
x = tf.Variable(x0)
def f(x):
return tf.pow(x, 2)
def second_derivative(x):
"""
Note here since x is a variable we are not calling the watch method.
Since we are taking the second derivative, we need to call the GradientTape twice and use the
gradient() method to obtain the gradients.
"""
with tf.GradientTape() as t:
with tf.GradientTape() as t2:
out = f(x)
dy_dx = t2.gradient(out, x)
d2y_dx2 = t.gradient(dy_dx, x)
return d2y_dx2
# Original Function f(x) = x^2
plt.plot(x0, f(x0), color='dodgerblue')
# Second Derivative of f(x)
d2 = second_derivative(x)
plt.plot(x0, d2, color='#FF9A13')
plt.grid()
# + [markdown] colab_type="text" id="f-7imtqRYtJ1"
#
# - If the second derivative is negative, the function curves downward, so the cost function will actually decrease by more than $\epsilon$.
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="8XmRGGIqYseM" outputId="aaf14c75-8428-409e-eb4b-2747cfaf56f5"
# Plot where second derivative is negative
x0 = tf.range(-10., 10., .5)
x = tf.Variable(x0)
def f(x):
return -(tf.pow(x, 4))
def second_derivative(x):
with tf.GradientTape() as t:
with tf.GradientTape() as t2:
out = f(x)
dy_dx = t2.gradient(out, x)
d2y_dx2 = t.gradient(dy_dx, x)
return d2y_dx2
# Original Function f(x): -x^4
plt.plot(x0, f(x0), color='dodgerblue')
# Second Derivative of f(x)
d3 = second_derivative(x)
plt.plot(x0, d3, color='#FF9A13')
plt.grid()
# + [markdown] colab_type="text" id="eUIA6TSYa6ok"
# - If the second derivative is positive, the function curves upward, so the cost function can decrease by less than $\epsilon$.
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="DkBr2AC3VBSn" outputId="c07105d7-b7ea-4d34-e8aa-f43d1861f455"
# Plot where second derivative is positive
x0 = tf.range(-10., 10., .5)
x = tf.Variable(x0)
def f(x):
return (tf.pow(x, 4))
def second_derivative(x):
with tf.GradientTape() as t:
with tf.GradientTape() as t2:
out = f(x)
dy_dx = t2.gradient(out, x)
d2y_dx2 = t.gradient(dy_dx, x)
return d2y_dx2
# Original Function f(x): x^4
plt.plot(x0, f(x0), color='dodgerblue')
# Second Derivative of f(x)
d3 = second_derivative(x)
plt.plot(x0, d3, color='#FF9A13')
plt.grid()
# + [markdown] colab_type="text" id="WnvCkcytVBkC"
# When our function has multiple input dimensions, there are many second derivatives. These derivatives can be collected together into a matrix called the __Hessian matrix__. The Hessian matrix $H (f)(x)$ is defined such that:
#
# $$\color{orange}{H(f)(x)_{i, j} = \frac{\partial^2}{\partial x_i \partial x_j} f(x) \tag{7}}$$
#
# Equivalently, the Hessian is the Jacobian of the gradient.
#
# If you have been following everything upto now, this should make sense and if it does feel free to skip to the code, but sometimes it's better to recap, so let me explain what Hessian and Jacobian mean and how they are related, starting with the gradients in one dimension, a gradient is simply the slope of a function in a direction, if you have multiple direction (inputs), for each direction there is a different rate of change for each direction and gradient is simply a collection of all of this. Think of throwing a ball, the ball goes further (x direction) and higher (z direction) and if you take the derivative of the directions with respect to time, you get the individual velocities in each direction and the gradient is a matrix of such values. This matrix of gradients is called the Jacobian. Now, let's say you take the derivative of the velocities again, you end up with the acceleration of the ball in each direction (x and z) and this is the Hessian matrix, simple as that.
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="DLELEdV0cYV2" outputId="dea4efa0-5d95-4b49-fdd1-26bf76d4c50f"
"""
NOTE: As of 06.14.2019 tf.hessians doesn't work with tf eager mode and the tf.GradientTape doesn't
have a method hessians. There is a bug report open in GitHub
(https://github.com/tensorflow/tensorflow/issues/29781) so as soon as that is resolved the following
code will work and I will fix this cell.
>>> f = 100*(y - x**2)**2 + (1 - x)**2
>>> x = tf.Variable([1., 1.])
>>> hessian_matrix = tf.hessians(f, x)
>>> print(hessian_matrix)
>>> with tf.GradientTape() as t:
... out = f
>>> hessian = t.hessians(out, x)
But for now, we will look into another way we can get the Hessian using the
equivalency of the Hessian being the gradient of the Jacobian for f(x,y)= x^3 - 2xy + y^6 and for
points x = 1, y = 2, but note that this will return the sum of each column, the final
Hessian Matrix will be:
[[ 6., -2. ],
[-2., -480.]]
I specifically took this example from Khan academy so if you want to see how the derivation is calculated,
take a look at:
(https://www.khanacademy.org/math/multivariable-calculus/applications-of-multivariable-derivatives/quadratic-approximations/a/the-hessian)
"""
def f(x): return tf.pow(x[0], 3) - 2*x[0]*x[1] - tf.pow(x[1], 6)
x = tf.Variable([1., 2.], dtype=tf.float32)
with tf.GradientTape() as h:
with tf.GradientTape() as g:
out = f(x)
jacobian = g.jacobian(out, x) # Calls the Jacobian method
hessian = h.gradient(jacobian, x) # We take the gradient of the Jacobian to get the Hessian Matrix
print("Jacobian: \n{} \nHessian: \n{}".format(jacobian, hessian))
# + [markdown] colab_type="text" id="ZESHQxY8VCf4"
# Anywhere that the second partial derivatives are continuous, the differential
# operators are commutative, this implies that:
#
# $$\color{orange}{H_{i, j} = H_{j, i} \tag{8}}$$
#
# so the Hessian matrix is symmetric at such points.
#
# Most of the functions we encounter in the context of deep learning have a symmetric Hessian almost everywhere. Because the Hessian matrix is real and symmetric, we can decompose it into a set of real eigenvalues and an orthogonal basis of eigenvectors. If you don't recall Eigenvalues and Eigenvectors, I strongly suggest you review [Eigendecomposition](https://www.adhiraiyan.org/deeplearning/02.00-Linear-Algebra#7). But for the sake of completion, the eigenvectors of matrix M are vectors that do not change direction when multiplied with M , and the eigenvalues represent the change in length of the eigenvector when multiplied with M .
#
# The second derivative in a specific direction represented by a unit vector $d$ is given by $d^{\top} H d$. When $d$ is the eigenvector of $H$, the second derivative in that direction is given by the corresponding eigenvalue. For other directions of $d$, the directional second derivative is a weighted average of all the eigenvalues, with weights between 0 and 1, and eigenvectors that have a smaller angle with $d$ receiving more weight. The maximum eigenvalue determines the maximum second derivative, and the minimum eigenvalue determines the minimum second derivative.
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="_Mqr5Uz7VC2B" outputId="931bdfaf-4604-488e-d1d1-6a613011c76a"
# Eigenvalue and Eigenvectors for the Hessian matrix
hessian = tf.constant([[ 6., -2.], [-2., -480.]])
eigenvalues, eigenvectors = tf.linalg.eigh(hessian)
print("Eigenvalue for the Hessian: \n{} \n\nEigenvectors for the Hessian: \n{}".format(eigenvalues, eigenvectors))
# + [markdown] colab_type="text" id="FdS983650tV2"
# The (directional) second derivative tells us how well we can expect a gradient
# descent step to perform. We can make a second-order Taylor series approximation to the function $f(x)$ around the current point $x^{(0)}$ and if we use a learning rate of $\epsilon$, then the new point $x$ will be given by $x^{(0)} - \epsilon g$ which gives:
#
# $$\color{orange}{f(x^{(0)} - \epsilon g) \approx f(x^{(0)}) - \epsilon g^{\top}g + \frac{1}{2} \epsilon^2 g^{\top} Hg \tag{9}}$$
#
# I know, that looks complicated, let me explain. There are three terms here: the original value of the function, the expected improvement due to the slope of the function, and the correction we must apply to account for the curvature of the function. What you need to know is:
#
# - When this last term is too large, the gradient descent step can actually move uphill.
#
# - When it is zero or negative, the Taylor series approximation predicts that increasing $\epsilon$ forever will decrease $f$ forever.
#
# Take a look at the image below. The graph on the left, which is the first derivative of a one dimensional function and the graph on the right is the second derivative of the same function.
#
# 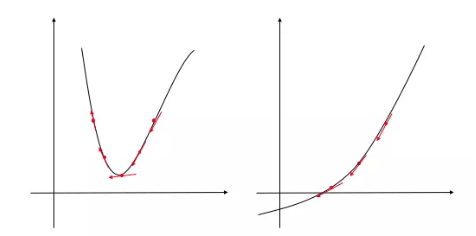
#
# In the left diagram, the slope changes a lot, while it is more stable in the right diagram. As you can see, the rate of change of the slope corresponds to how “curved” each loss function is. The sharper the curve, the more rapidly the slope changes.
#
# + [markdown] colab_type="text" id="Nnzwd5x_0sNt"
# In multiple dimensions, we need to examine all the second derivatives of the function. Using the eigendecomposition of the Hessian matrix, we can generalize the second derivative test to multiple dimensions. At a critical point, where $\nabla_x f(x) = 0$, we can examine the eigenvalues of the Hessian to determine whether the critical point is a local maximum, local minimum, or saddle point.
#
# - When the Hessian is positive definite (all its eigenvalues are positive), the point is a local minima.
#
# - When the Hessian is negative definite (all its eigenvalues are negative), the point is a local maximum.
#
# - When all the nonzero eigenvalues have the same sign but atleast one eigenvalue is zero, the derivative test can be inconclusive, like the univariate second derivative test.
#
# + [markdown] colab_type="text" id="eo0wvHXH0rf8"
# In multiple dimensions, there is a different second derivative for each direction at a single point. The condition number of the Hessian at this point measures how much the second derivatives differ from each other. When the Hessian has a poor condition number, gradient descent performs poorly. This is because in one direction, the derivative increases rapidly, while in another direction, it increases slowly. Gradient descent is unaware of this change in the derivative, so it does not know that it needs to explore preferentially in the direction where the derivative remains negative for longer. Poor condition number also makes choosing a good step size difficult. The step size must be small enough to avoid overshooting the minimum and going uphill in directions with strong positive curvature. This usually means that the step size is too small to make significant progress in other directions with less curvature.
# + [markdown] colab_type="text" id="2VeHZB7ZBMYy"
# This issue can be resolved by using information from the Hessian matrix to guide the search. The simplest method for doing so is known as __Newton’s method__. Newton’s method is based on using a second-order Taylor series expansion to approximate $f(x)$ near some point $x^{(0)}$ and by solving for the critical point, we obtain:
#
# $$\color{orange}{x^* = x^{(0)} - H(f)(x^{(0)})^{-1} \ \nabla_x f(x^{(0)}) \tag{10}}$$
#
# - When $f$ is positive definite quadratic function, Newton's method consists of applying the above equation once to jump to the minimum of the function directly.
#
# - When $f$ is not truly quadratic but can be locally approximated as a positive definite quadratic, Newton’s method consists of applying the above equation multiple times.
#
# Iteratively updating the approximation and jumping to the minimum of the approximation can reach the critical point much faster than gradient descent would. This is a useful property near a local minimum, but it can be a harmful property near a saddle point.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="i8GvcXFY9kEM" outputId="50150f13-0234-49db-fdcc-0a98d56a8cfe"
"""
Let's see how Newton's method can be used to solve a quadratic function by applying it once.
We start with a quadratic function x^2 + 5y^2 and when you differentiate it, you end up with 2x + 10y
which would result in a Hessian of [[2 0], [0, 10]] because the derivative of 2x + 10y is 2 and 10.
"""
def quad(x):
return ((x[1:])**2.0 + 5*(x[:-1])**2.0)
def quad_grad(x,y):
return (tf.Variable([2.0*x, 10.0*y]))
# Create x and y values
x = tf.linspace(-20.0, 20.0, 100)
y = tf.linspace(-20.0, 20.0, 100)
# Broadcasts parameters for evaluation on an N-D grid
X, Y = tf.meshgrid(x, y)
# Reshape X and Y to pack along first dim (row wise) and apply the quadratic function and reshape it to get the original dimension
Z = tf.reshape(quad(tf.stack([tf.reshape(X, [10000,]), tf.reshape(Y, [10000,])], axis=0)), [100, 100])
# Take the inverse of the Hessian: (1/2, 1/10)
H_inv = - tf.constant([[0.5, 0], [0, 0.1]])
plt.figure(figsize=(12,4))
plt.subplot(121)
plt.contour(X,Y,Z);
plt.title("Steepest Descent");
step = -0.25
X0 = 10.0
Y0 = 1.0
# Here we calculate the Gradient of our function and take the dot product between the gradient and the Hessian inverse
N_grad = tf.tensordot(H_inv, quad_grad(X0,Y0), axes=1)
sgrad = step*quad_grad(X0,Y0)
plt.quiver(X0, Y0, sgrad[0], sgrad[1], color='red',angles='xy',scale_units='xy',scale=1);
X1 = X0 + sgrad[0]
Y1 = Y0 + sgrad[1]
sgrad = step*quad_grad(X1,Y1)
plt.quiver(X1.numpy(), Y1.numpy(), sgrad[0].numpy(), sgrad[1].numpy(), color='green', angles='xy', scale_units='xy', scale=1);
X2 = X1 + sgrad[0]
Y2 = Y1 + sgrad[1]
sgrad = step*quad_grad(X2,Y2)
plt.quiver(X2.numpy(), Y2.numpy(), sgrad[0].numpy(), sgrad[1].numpy(), color='purple',angles='xy',scale_units='xy',scale=1);
plt.subplot(122)
plt.contour(X,Y,Z);
plt.title("Newton's Method")
plt.quiver(X0, Y0, N_grad[0], N_grad[1], color='purple',angles='xy',scale_units='xy',scale=1);
# + [markdown] colab_type="text" id="KBVK0lNS7z7f"
#
# Optimization algorithms that use only the gradient, such as gradient descent, are called __first-order optimization algorithms__. Optimization algorithms that also use the Hessian matrix, such as Newton’s method, are called __second-order optimization algorithms__.
# + [markdown] colab_type="text" id="uxbCCGRhP202"
# Deep learning algorithms tend to lack guarantees because the family of functions used in deep learning is quite complicated. But sometimes we gain some guarantees by restricting ourselves to functions that are either __Lipschitz continuous__ or have Lipschitz continuous derivatives. A Lipschitz function is a function $f$ whose rate of change is bounded by a __Lipschitz constant__ $\mathcal{L}$:
#
# $$\color{orange}{\forall x, \forall y, |f(x) - f(y)| \leq \mathcal{L} \| x - y \|_2 \tag{11}}$$
#
# This property is useful because it enables us to quantify our assumption that a small change in the input made by an algorithm such as gradient descent will have a small change in the output.
# + [markdown] colab_type="text" id="ci5LjG2Trb90"
# # 04.04 - Constrained Optimization
# + [markdown] colab_type="text" id="enaamo6eVGrr"
# Sometimes we may wish to find the maximal or minimal value of $f(x)$ for values of $x$ in some set $\mathbb{S}$. This is knows as __constrained optimization__. Points that lie within the set $\mathbb{S}$ are called __feasible__ points in constrained optimization technology.
#
# We often wish to find a solution that is small in some sense. A common approach in such situations is to impose a norm constraint, such as $\| x \| \leq 1$. One simple approach to constrained optimization is simply to modify gradient descent taking the constraint into account.
#
# + [markdown] colab_type="text" id="PkBfy4NuVHH4"
# A more sophisticated approach is to design a different, unconstrained optimization problem whose solution can be converted into a solution to the original, constrained optimization problem. This approach requires creativity; the transformation between optimization problems must be designed specifically for each case we encounter.
#
# The __Karush–Kuhn–Tucker (KKT)__ approach provides a very general
# solution to constrained optimization. With the KKT approach, we introduce
# a new function called the __generalized Lagrangian__ or __generalized Lagrange function__.
#
# To define the Lagrangian, we first need to describe $\mathbb{S}$ in terms of equations and inequalities. We want a description of $\mathbb{S}$ in terms of m functions $g^{(i)}$ and $n$ functions $h^{(j)}$ so that $\mathbb{S} = \{ x \ \mid \ \forall i, g^{(i)} (x) = 0 \ \text{and} \ \forall j, h^{(j)} (x) \leq o) \}$. The equations involving $g^{(i)}$ are called __equality constraints__, and the inequalities involving $h^{(j)}$ are called __inequality constraints__.
#
# We introduce new variables $\lambda_i$ and $\alpha_j$ for each constraint, these are called the KKT multipliers. The generalized Lagrangian is then defined as:
#
# $$\color{orange}{L(x, \lambda, \alpha) = f(x) + \displaystyle\sum_i \lambda_i g^{(i)} (x) + \displaystyle\sum_j \alpha_j h^{(j)} (x) \tag{12}}$$
#
# We can now solve a constrained minimization problem using unconstrained optimization of the generalized Lagrangian. As long as at least one feasible point exists and $f(x)$ is not permitted to have $\infty$.
# + colab={"base_uri": "https://localhost:8080/", "height": 286} colab_type="code" id="huEEYquLO2Kb" outputId="3390b120-67cb-4ed4-a62f-84aedaf0a133"
"""
Let's see how this looks by plotting a function f(x) = −(2xy+2x−x^2−2y^2) with constraints
x^3−y = 0 and y−(x−1)^4−2≥ 0 with bounds 0.5, 1.5, 1.5, 2.5. The solution will be where the
two constraints meet inside the bound.
"""
def f(x):
return -(2*x[0]*x[1] + 2*x[0] - x[0]**2 - 2*x[1]**2)
x = np.linspace(0, 3, 100)
y = np.linspace(0, 3, 100)
X, Y = np.meshgrid(x, y)
Z = f(np.vstack([X.ravel(), Y.ravel()])).reshape((100,100))
plt.contour(X, Y, Z, np.arange(-1.99,10, 1), cmap='jet');
plt.plot(x, x**3, 'k:', linewidth=2, color='r') # constraint 1: x^3−y = 0
plt.plot(x, (x-1)**4+2, 'k:', linewidth=2, color='b') # constraint 2: y−(x−1)^4−2≥ 0
plt.fill([0.5,0.5,1.5,1.5], [2.5,1.5,1.5,2.5], alpha=0.3) # bounds 0.5, 1.5, 1.5, 2.5
plt.axis([0,3,0,3])
# + [markdown] colab_type="text" id="XYXBAbc0VHpM"
# To perform constrained maximization, we can construct the generalized
# Lagrange function of $- f(x)$, which leads to this optimization problem:
#
# $$\color{orange}{min_x max_{\lambda} max_{\alpha, \alpha \geq 0} = - f(x) + \displaystyle\sum_i \lambda_i g^{(i)} (x) + \displaystyle\sum_j \alpha_j h^{(j)} (x) \tag{13}}$$
#
# The sign of the term for the equality constraints does not matter; we may define it with addition or subtraction as we wish.
# + [markdown] colab_type="text" id="nXOGjZgOzL55"
# A simple set of properties describe the optimal points of constrained optimization problems. These properties are called the Karush-Kuhn-Tucker (KKT) conditions. They are necessary conditions, but not always sufficient conditions, for a point to be optimal. The conditions are:
#
# - The gradient of the generalized Lagrangian is zero.
#
# - All constraints on both $x$ and the KKT multipliers are satisfied
#
# - The inequality constraints exhibit "complementary slackness": $\alpha \odot h(x) = 0$
# + [markdown] colab_type="text" id="YruCMnZDruwB"
# # 04.05 - Example: Linear Least Squares
# + [markdown] colab_type="text" id="97g_Kjn6VLRB"
# Let's see how we can find the value of $x$ that minimizes:
#
# $$\color{orange}{f(x) = \frac{1}{2} \| Ax - b \|_2^2 \tag{14}}$$
#
# using gradient based optimization.
#
# First we need to obtain the gradient:
#
# $$\color{orange}{\nabla_x f(x) = A^{\top} (Ax - b) = A^{\top} Ax - A^{\top} b \tag{15}}$$
#
#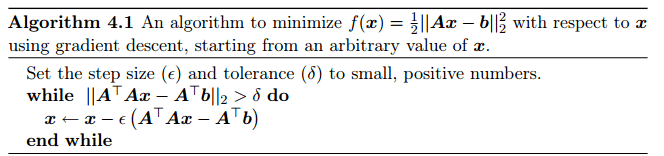 
#
# One can also solve this problem using Newton’s method. In this case, because
# the true function is quadratic, the quadratic approximation employed by Newton’s method is exact, and the algorithm converges to the global minimum in a single step.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 303} colab_type="code" id="dgrix6ojcvXQ" outputId="58842efa-7f97-4e4e-d013-843b6c33eafb"
"""
There are two ways to solve the above system. Direct method and iterative method.
We start with the iterative method. The optimal solution is where the gradient becomes zero
therefore it is at x = (A^T b)*(A^T A)^{-1} and this is what we will be calculating
"""
# Generate random x and y data
x_vals = np.linspace(0., 10., 100)
y_vals = x_vals + np.random.normal(loc=0, scale=1, size=100)
x_vals_column = np.transpose(np.matrix(x_vals))
ones_column = tf.ones([100, 1], dtype=tf.float32)
# tensorflow needs its data types in float so we cast the dtypes to float
A_tensor = tf.dtypes.cast(tf.concat([x_vals_column, ones_column], axis=1), tf.float32)
Y_tensor = tf.dtypes.cast(tf.reshape(tf.transpose(y_vals), [100, 1]), tf.float32)
# Iterative method
tA_A = tf.matmul(A_tensor, A_tensor, transpose_a=True) # We calculate A^T A
tA_A_inv = tf.linalg.inv(tA_A) # And take the inverse of it (A^TA)^{-1}
product = tf.matmul(tA_A_inv, A_tensor, transpose_b=True) # Then multiply it with A to yield (A^TA)^{-1} A
A_eval = tf.matmul(product, Y_tensor) # Finally we find (A^TA)^{-1}*A*b
m_slope = A_eval[0][0]
b_intercept = A_eval[1][0]
print('slope (m): ' + str(m_slope))
print('intercept (b): ' + str(b_intercept))
# Now for each x_val we find the best fit line
best_fit = []
for i in x_vals:
best_fit.append(m_slope * i + b_intercept)
plt.plot(x_vals, y_vals, 'o', label='Data')
plt.plot(x_vals, best_fit, 'r-', label='Least square fit', linewidth=3)
plt.legend(loc='upper left')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 291} colab_type="code" id="jzknk-aNO20g" outputId="15509a75-2405-499b-b11b-eaeb0b79ff65"
"""
Next, we try the direct method for another dataset using tensorflows least squares class for function
f(x) = 1 + 2x + 3x^2
"""
# define true model parameters
x = np.linspace(-1, 1, 100)
a, b, c = 1, 2, 3
y_exact = a + b * x + c * x**2
# simulate noisy data points
m = 100
X = 1 - 2 * np.random.rand(m)
Y = tf.reshape(tf.convert_to_tensor(a + b * X + c * X**2 + np.random.randn(m)), [100, 1])
A = tf.stack([X**0, X**1, X**2], axis=1)
# Solving directly using tensorflow's least sqaures.
sol = tf.linalg.lstsq(A, Y)
y_fit = sol[0] + sol[1] * x + sol[2] * x**2
fig, ax = plt.subplots(figsize=(12, 4))
ax.plot(X, Y, 'go', alpha=0.5, label='Simulated data')
ax.plot(x, y_exact, 'k', lw=2, label='True value $y = 1 + 2x + 3x^2$')
ax.plot(x, y_fit, 'b', lw=2, label='Least square fit')
ax.set_xlabel(r"$x$", fontsize=18)
ax.set_ylabel(r"$y$", fontsize=18)
ax.legend(loc=2);
# + [markdown] colab_type="text" id="PZH1Y9orgZWI"
# Now, suppose we wish to minimize the same function, but subject to the constraint $x^{\top} x \leq 1$. To do so, we introduce the Lagrangian:
#
# $$\color{orange}{L(x, \lambda) = f(x) + \lambda (x^{\top} x - 1) \tag{16}}$$
#
# We can now solve the problem:
#
# $$\color{orange}{min_x max_{\lambda, \lambda \geq 0} L(x, \lambda) \tag{17}}$$
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="xQvOm7Q6c8V8" outputId="51b88236-a259-4d24-ff22-c0df7156e352"
"""
Let' take an example: Maximize f(x,y,z)=xy+yz subject to the constraints x+2y=6 and x−3z=0.
We start by setting up the equation:
F(x,y,z,λ,α) = xy + yz − λ(x+2y−6) + α(x-3z)
Now set the partial derivatives to zero and solve the following set of equations:
y − λ − α = 0
x + z − 2λ = 0
y + 3α = 0
x + 2y −6 = 0
x − 3z = 0
which is a linear equation in x,y,z,λ,μ, Now this can be put into the matrix equation:
[0, 1, 0, -1, -1] [x] = [0]
[1, 0, 1, -2, 0] [y] = [0]
[0, 1, 0, 0, 3] [z] = [0]
[1, 2, 0, 0, 0] [λ] = [6]
[1, 0,-3, 0, 0] [μ] = [0]
"""
matrix = tf.constant([
[0, 1, 0, -1, -1],
[1, 0, 1, -2, 0],
[0, 1, 0, 0, 3],
[1, 2, 0, 0, 0],
[1, 0,-3, 0, 0]], dtype=tf.float32)
rhs = tf.constant([[0],[0],[0],[6],[0]], dtype=tf.float32)
solve = tf.linalg.solve(matrix, rhs)
print("Solving the constrained optimization using Lagrange multipliers yield x: \n{}".format(solve))
# + [markdown] colab_type="text" id="_JaAxGaic-aa"
# The smallest norm solution to the unconstrained least squares problem may be found using the Moore-Penrose pseudoinverse: $x = A^+ b$. If this point is feasible, then it is the solution to the constrained problem.
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="ZYo5HSoQgZ2G" outputId="a8a044f6-4578-4cf6-8331-a12a301338ea"
# Moore-penrose pseudoinverse solution to the above problem
mpp_solution = tfp.math.pinv(matrix)
print(mpp_solution)
# + [markdown] colab_type="text" id="hnKwfKJ2jLbc"
# If the Moore-Penrose pseudoinverse solution is not feasible, we find the solution where the constraint is active by differetiating the Lagrangian with respect to $x$ and solving for $x$, we obtain:
#
# $$\color{orange}{x = (A^{\top} A + 2 \lambda I)^{-1} A^{\top} b \tag{18}}$$
#
# We can then find the magnitude of $\lambda$ by performing gradient ascent using:
#
# $$\color{orange}{\frac{\partial}{\partial \lambda} L(x, \lambda) = x^{\top} x - 1 \tag{19}}$$
#
# When the norm of $x$ exceeds 1, this derivative is positive, so to follow the derivative uphill and increase the Lagrangian with respect to $\lambda$, we increase $\lambda$. Because the coefficient on the $x^{\top} x$ penatly has increased, solving the linear equation for $x$ will now yield a solution with a smaller norm. The process of solving the linear equation and adjusting $\lambda$ continues until $x$ has the correct norm and the derivative on $\lambda$ is 0.
#
#
# + [markdown] colab_type="text" id="LU--mNoUad4S"
# # 💫 Congratulations
#
# You have successfully completed Chapter 4 Numerical Computation of [Deep Learning with Tensorflow 2.0](https://www.adhiraiyan.org/DeepLearningWithTensorflow.html). To recap, we went through the following concepts:
#
# - Numerical Computation
# - Overflow and Underflow
# - Poor Conditioning
# - Gradient Based Optimization
# - Constrained Optimization
# - Example: Linear Least Squares
#
#
# If you like to read more about Numerical Computation take a look at [Numerical Methods
# for Engineers by <NAME>](http://mechfamilyhu.net/download/uploads/mech144232415981.pdf) or [Numerical Methods for
# Computational Science and Engineering by Prof. <NAME>](http://www.sam.math.ethz.ch/~hiptmair/tmp/NumCSE/NumCSE15.pdf).
#
# This concludes the mathematical preliminaries that we use to develop machine learning algorithms. We are now ready to build and analyze some full-fledged machine learning systems. I am excited, are you 😍.
# -
# <!--NAVIGATION-->
# < [03.00 - Probability and Information Theory](03.00-Probability-and-Information-Theory.ipynb) | [Contents](Index.ipynb) |
#
# <a href="https://colab.research.google.com/github/adhiraiyan/DeepLearningWithTF2.0/blob/master/notebooks/04.00-Numerical-Computation.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
| notebooks/04.00-Numerical-Computation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Inferential Statistics Report
#
# In this brief document, we summarize the inferential statistics analysis and finding from the Jupyter notebook `Inferential-Statistics.ipynb`.
#
#
# ## Significance level ($\alpha$)
#
# In all of the hypothesis tests in our analysis, we used a significance level of $\alpha = 0.05$.
#
# ## Our Questions
# The questions that we tried to answer were as follows:
# 1. Is the proportion of defaults the same for men and women?
# 2. Is age a significant predictor of default?
# 3. Is credit limit a significant predictor of default?
# 4. Is the ratio of $\left(\frac{\text{bill amount}}{\text{credit limit}}\right)$ a significant predictor of default? Here, "$\text{bill amount}$" stands for past credit card bill amounts.
# 5. Is the ratio of $\left(\frac{\text{bill amount} - \text{pay amount}}{\text{credit limit}}\right)$ a significant predictor of default?
#
#
# ### 1. Is the proportion of defaults the same for men and women?
#
# We wanted to test whether the proportion of defaults the same for men and women.
#
# Let $p_m$ represent the proportion of defaults for men.
#
# Let $p_w$ represent the proportion of defaults for women.
#
# Our null and alternative hypotheses are as follows:
#
# + $H_0$: $p_m = p_w$
#
# + $H_1$: $p_m \neq p_w$
#
# We used bootstrapping to test our null hypothesis. We reject the null hypothesis that $p_m = p_w$.
#
# ### 2. Is age a significant predictor of default?
#
# We conducted a logistic regression where age was the predictor variable and default status was the target variable.
#
# We used the implementation of logistic regression in the `glm` package in the R language. We chose to use `glm`'s implementation because it calculates the p-values associated with each regression coefficient. The logistic regression implemented in `scikit-learn` does not calculate these p-values.
#
# We used the `rpy2` Python library to call R from within Python.
#
# From our regression results, we found that the p-value for the regression coefficient was 0.0162, which is less than $\alpha = 0.05$. Therefore, we conclude that age was a statistically significant predictor of default.
#
# The regression coefficient was positive, implying that the log-odds of default increase as age increases.
#
#
# ### 3. Is credit limit a significant predictor of default?
#
# We conducted a logistic regression where credit limit is the predictor variable and default status is the target variable.
#
# From our regression results, we found that the p-value for the regression coefficient was less than $2 \times 10^{-16}$, which is less than $\alpha = 0.05$. Therefore, we concluded that credit limit was a statistically significant predictor of default.
#
# The regression coefficient was negative, implying that the log-odds of default decrease as credit limit increases.
#
# ### 4. Is the ratio of $\left(\frac{\text{bill amount}}{\text{credit limit}}\right)$ a significant predictor of default?
#
# Here, "$\text{bill amount}$" stands for past credit card bill amounts.
#
# We conducted a logistic regression where the ratio of $\left(\frac{\text{bill amount}}{\text{credit limit}}\right)$ was the predictor variable and default status was the target variable.
#
# There are six bill amount features: 'BILL_AMT1', 'BILL_AMT2', ..., & 'BILL_AMT6'.
#
# From our regression results, we found that the p-value for the regression coefficient of each bill amount ratio was less than $2 \times 10^{-16}$, which is less than $\alpha = 0.05$. Therefore, we concluded that, for each of the 6 bill amounts, the ratio of $\left(\frac{\text{bill amount}}{\text{credit limit}}\right)$ was a statistically significant predictor of default.
#
# The regression coefficients were positive, implying that the log-odds of default increase as the ratio of $\left(\frac{\text{bill amount}}{\text{credit limit}}\right)$ increases.
#
#
# ### 5. Is the ratio of $\left(\frac{\text{bill amount} - \text{pay amount}}{\text{credit limit}}\right)$ a significant predictor of default?
#
# We conducted a logistic regression where the ratio of $\left(\frac{\text{bill amount} - \text{pay amount}}{\text{credit limit}}\right)$ was the predictor variable and default status was the target variable.
#
# From our regression results, we found that the p-value for the regression coefficient of each ratio is less than $2 \times 10^{-16}$, which is less than $\alpha = 0.05$. Therefore, we concluded that, for each of the 6 (bill amount, pay amount) pairs, the ratio of $\left(\frac{\text{bill amount} - \text{pay amount}}{\text{credit limit}}\right)$ was a statistically significant predictor of default.
#
# The regression coefficients were positive, implying that the log-odds of default increase as the ratio of
# $\left(\frac{\text{bill amount} - \text{pay amount}}{\text{credit limit}}\right)$ increases.
#
| reports/Inferential-Statistics-Report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # FashionMNIST
# Load images from the [Fashion-MNIST data](https://github.com/zalandoresearch/fashion-mnist)
#
#
# The dataset comprised of 60,000 small square 28x28 pixel grayscale images of items of 10 types of clothing with 0-9 class labels.
# class labels:
# * 0: T-shirt/top
# * 1: Trouser
# * 2: Pullover
# * 3: Dress
# * 4: Coat
# * 5: Sandal
# * 6: Shirt
# * 7: Sneaker
# * 8: Bag
# * 9: Ankle boot
#
# ### Load the Fashion-MNIST data
# * Use ``torch.utils.data.dataset``
# * Data path: data
# * Apply transformations to the data (turning all images into Tensor's for training a NN
#
# ## Import the Necessary Packages
# +
# basic torch libraries
import torch
import torchvision
# data loading and transforming
from torchvision.datasets import FashionMNIST
from torch.utils.data import DataLoader
from torchvision import transforms
# basic libraries
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### The output of ``torchvision`` are PILImage images of range [0, 1]
# * Transform them to Tensor for input into a CNN
# +
# Defin a transform to read the data in as a Tensor
data_transform = transforms.ToTensor()
# Choose the training and test datasets
path = './data'
train_data = FashionMNIST(root=path,
train=True,
download=False,
transform=data_transform)
# Print out some stats about the training data
print('Train data, number of images', len(train_data))
# -
# ## Data iteration and batching
# ``torch.utils.data.DataLoader`` is an iterator that allows to batch and shuffle the data
#
# +
# shuffle the data and load in image/label data in batches of size 20
# Depends on large or small size of batch size will affect the loss
batch_size = 20
train_loader = DataLoader(train_data, batch_size=batch_size, shuffle=True)
# specify the image classes
classes = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat', 'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# -
# Using ``dataiter.next()`` for cell iterates over the training dataset of loaded a random batch image/label data.
#
# Plots the batch of images and labels in a ``2*batch_size/2`` grid.
#
# +
# obtain one batch of training images
# iter
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert to numpy
# plot the images in the batch with labels
fig = plt.figure(figsize=(25, 4)) # fig size
for idx in np.arange(batch_size):
ax = fig.add_subplot(2, batch_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title(classes[labels[idx]])
# -
# ## View an image
# * Normalize
# * grayscale image
#
# ### Normalization
# Normalization ensures that, as we go through a feedforward and then backpropagation step in training our CNN, that each image feature will fall within a similar range of values and not overly activate any particular layer in our network. During the feedfoward step, a network takes in an input image and multiplies each input pixel by some convolutional filter weights (and adds biases!), then it applies some activation and pooling functions. Without normalization, it's much more likely that the calculated gradients in the backpropagaton step will be quite large and cause our loss to increase instead of converge
#
#
# +
# select an image by index
idx = 2
img = np.squeeze(images[idx])
# display the pixel values in the image
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
# -
| 03_Fashion_MNIST/FashionMNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Betelgeuse apparent magnitude over the last 50 years
# Author: <NAME>, Ph.D.
# Date: 2020-01-08
# Source: AAVSO Database https://www.aavso.org/data-download
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import datetime as dt
# %matplotlib inline
# +
# Load data
fname = "AAVSO_Betelgeuse_50yrs.csv"
df = pd.read_csv(fname)
# Include only broadband visible observations
df = df[df["Band"] == "Vis."]
# Cast magnitude to numeric where possible
df["Magnitude"] = pd.to_numeric(df["Magnitude"], errors="coerce")
# Convert Julian dates to YYYY-MM-DD
df["Date"] = dt. df["JD"]
# -
df.head(10)
df.tail(10)
df.plot(x="JD", y="Magnitude", kind="scatter", figsize=(20,10), ylim=(2.5,-0.5))
df.plot(x="JD", y="Magnitude", kind="scatter", figsize=(20,10), xlim=(2458000.0, np.max(df["JD"])+10), ylim=(2.5,-0.5))
| Betelgeuse_Variability_50yrs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Inspecting training data
#
# ## Background
#
# Prior to training a machine learning classifier, it can be useful to understand which of our feature layers are most useful for distinguishing between classes. The feature layers the model is trained on form the **knowledge base** of the algorithm. We can explore this knowledge base using class-specific [violin plots](https://en.wikipedia.org/wiki/Violin_plot#:~:text=A%20violin%20plot%20is%20a,by%20a%20kernel%20density%20estimator.), and through a dimensionality reduction approach called [principal-components analysis](https://builtin.com/data-science/step-step-explanation-principal-component-analysis). The latter transforms our large dataset with lots of variables into a smaller dataset with fewer variables (while still preserving much of the variance), this allows us to visualise a very complex dataset in a relatively intuitive and straightforward manner.
#
# ## Description
#
# Using the training data written to file in the previous notebook, [1_Extract_training_data](1_Extract_training_data.ipynb), this notebook will:
#
# 1. Plot class-specific violin plots for each of the feature layers in the training data.
# 2. Plot the importance of each feature after applying a model to the data.
# 3. Calculate the first two and three prinicpal components of the dataset and plot them as 2D and 3D scatter plots.
#
# ***
# ## Getting started
#
# To run this analysis, run all the cells in the notebook, starting with the "Load packages" cell.
# ## Load packages
# %matplotlib inline
import random
import numpy as np
import pandas as pd
import seaborn as sns
from pprint import pprint
import matplotlib.pyplot as plt
from matplotlib.patches import Patch
from sklearn.decomposition import PCA
from mpl_toolkits.mplot3d import Axes3D
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
# ## Analysis Parameters
#
# * `training_data`: Name and location of the training data `.txt` file output from runnning `1_Extract_training_data.ipynb`
# * `class_dict`: A dictionary mapping the 'string' name of the classes to the integer values that represent our classes in the training data (e.g. `{'crop': 1., 'noncrop': 0.}`)
# * `field`: This is the name of column in the original training data shapefile that contains the class labels. This is provided simply so we can remove this attribute before we plot the data
# +
training_data = "results/training_data/southern_training_data_20211108.txt"
class_dict = {'crop':1, 'noncrop':0}
field = 'Class'
# -
# ## Import training data
#
# +
# load the data
model_input = np.loadtxt(training_data)
# load the column_names
with open(training_data, 'r') as file:
header = file.readline()
column_names = header.split()[1:]
# Extract relevant indices from training data
model_col_indices = [column_names.index(var_name) for var_name in column_names[1:]]
# -
# ## Data Wrangling
#
# This cell extracts each class in the training data array and assigns it to a dictionary, this step will allow for cleanly plotting our data
dfs = {}
for key, value in class_dict.items():
print(key, value)
# extract values for class from training data
arr = model_input[model_input[:,0]==value]
# create a pandas df for ease of use later
df = pd.DataFrame(arr).rename(columns={i:column_names[i] for i in range(0,len(column_names))}).drop(field, axis=1)
# Scale the dataframe
scaled_df = StandardScaler(with_mean=False).fit_transform(df)
scaled_df = pd.DataFrame(scaled_df, columns=df.columns)
dfs.update({key:scaled_df})
# ## Feature layer violin plots
#
# The code here will generate class-specific violin plots for each feature side-by-side so we can see how seperable the features are between the classes. Features that distinguish between crop and non-crop will have medians and distributions that do not overlap too much
# +
#generate a random list of colors same length as num of classes
get_colors = lambda n: list(map(lambda i: "#" + "%06x" % random.randint(0, 0xFFFFFF),range(n)))
colors = get_colors(len(dfs))
#generate list of offsets & widths for plotting
start=-0.2
end=0.2
offsets = list(np.linspace(start,end,len(dfs)))
if len(dfs) == 2:
width=0.4
else:
width=np.abs(offsets[0] - offsets[1])
#create figure and axes
fig, ax = plt.subplots(figsize=(40,8))
for key, color, offset in zip(dfs,colors, offsets):
#create violin plots
pp = ax.violinplot(dfs[key].values,
showmedians=True,
positions=np.arange(dfs[key].values.shape[1])+offset, widths=width
)
# change the colour of the plots
for pc in pp['bodies']:
pc.set_facecolor(color)
pc.set_edgecolor(color)
pc.set_alpha(1)
#change the line style in the plots
for partname in ('cbars','cmins','cmaxes','cmedians'):
vp = pp[partname]
vp.set_edgecolor('black')
vp.set_linewidth(1)
#tidy the plot, add a title and legend
ax.set_xticks(np.arange(len(column_names[1:])))
ax.set_xticklabels(column_names[1:])
ax.set_ylim(-5.0,12)
ax.set_xlim(-0.5,len(column_names[1:])-.5)
ax.set_ylabel("Scaled Values", fontsize=14)
ax.set_xlabel("Feature Layers", fontsize=14)
ax.set_title("Training Data Knowledge-Base", fontsize=14)
ax.legend([Patch(facecolor=c) for c in colors], [key for key in dfs], loc='upper right');
# -
# # Feature Importance
#
# Here we extract classifier estimates of the relative importance of each feature for training the classifier. This is useful for potentially selecting a subset of input bands/variables for model training/classification (i.e. optimising feature space). However, in this case, we are not selecting a subset of features, but rather just trying to understand the importance of each feature.
#
# Results will be presented in asscending order with the most important features listed last. Importance is reported as a relative fraction between 0 and 1.
model=RandomForestClassifier()
model.fit(model_input[:, model_col_indices], model_input[:, 0])
# +
order = np.argsort(model.feature_importances_)
plt.figure(figsize=(35,10))
plt.bar(x=np.array(column_names[1:])[order],
height=model.feature_importances_[order])
# plt.bar(x=column_names[1:], height=model.feature_importances_)
plt.gca().set_ylabel('Importance', labelpad=10)
plt.gca().set_xlabel('Feature', labelpad=10)
plt.tight_layout()
# -
# ## Principal Component Analysis
#
# The code below will calculate and plot the first two and three principal components of our training dataset.
#
# The first step is to standardise our data to express each feature layer in terms of mean and standard deviation, this is necessary because principal component analysis is quite sensitive to the variances of the initial variables. If there are large differences between the ranges of initial variables, those variables with larger ranges will dominate over those with small ranges (For example, a variable that ranges between 0 and 100 will dominate over a variable that ranges between 0 and 1), which will lead to biased results. So, transforming the data to comparable scales can prevent this problem. We do this using sklearn's `StandardScalar` function which will normalise the values in an array to the array's mean and standard deviation via the formuala: `z = (x-u/s)`, where `u` is the mean of and `s` is the standard deviation.
# Compute the mean and variance for each feature
x = StandardScaler().fit_transform(model_input[:,1:])
# ### Conduct the PCAs
# +
#two components
pca2 = PCA(n_components=2)
pca2_fit = pca2.fit_transform(x)
#three PCA components
pca3 = PCA(n_components=3)
pca3_fit = pca3.fit_transform(x)
#add back to df
pca2_df = pd.DataFrame(data = pca2_fit,
columns = ['PC1', 'PC2'])
pca3_df = pd.DataFrame(data = pca3_fit,
columns = ['PC1', 'PC2', 'PC3'])
# concat with classes
result2 = pd.concat([pca2_df, pd.DataFrame({'class':model_input[:,0]})], axis=1)
result3 = pd.concat([pca3_df, pd.DataFrame({'class':model_input[:,0]})], axis=1)
# -
a2,b2 = pca2.explained_variance_ratio_
a3,b3,c3 = pca3.explained_variance_ratio_
print("Variance explained by two principal components = " + str(round((a2+b2)*100, 2))+" %")
print("Variance explained by three principal components = " + str(round((a3+b3+c3)*100, 2))+" %")
# #### Plot both 2 & 3 principal components
#
# +
if len(result2) > 5000:
result2=result2.sample(n=5000)
if len(result3) > 5000:
result3=result3.sample(n=5000)
fig = plt.figure(figsize=(18,8))
fig.suptitle('Training data: Principal Component Analysis', fontsize=14)
# First subplot
ax = fig.add_subplot(1, 2, 1)
scatter1=sns.scatterplot(x="PC1", y="PC2",
data=result2,
hue='class',
hue_norm=tuple(np.unique(result2['class'])),
palette='viridis',
legend=False,
alpha=0.7,
ax=ax
)
ax.set_title('Two Principal Components', fontsize=14)
ax.grid(True)
# Second subplot
ax = fig.add_subplot(1, 2, 2, projection='3d')
scatter2 = ax.scatter(result3['PC1'], result3['PC2'], result3['PC3'],
c=result3['class'], s=60, alpha=0.5)
# make simple, bare axis lines through space:
xAxisLine = ((min(result3['PC1']), max(result3['PC1'])), (0, 0), (0,0))
ax.plot(xAxisLine[0], xAxisLine[1], xAxisLine[2], 'r')
yAxisLine = ((0, 0), (min(result3['PC2']), max(result3['PC2'])), (0,0))
ax.plot(yAxisLine[0], yAxisLine[1], yAxisLine[2], 'r')
zAxisLine = ((0, 0), (0,0), (min(result3['PC3']), max(result3['PC3'])))
ax.plot(zAxisLine[0], zAxisLine[1], zAxisLine[2], 'r')
ax.set_title('Three Principal Components', fontsize=14)
ax.set_xlabel("PC1")
ax.set_ylabel("PC2")
ax.set_zlabel("PC3");
# -
# ## Next steps
#
# To continue working through the notebooks in this `Southern Africa Cropland Mask` workflow, go to the next notebook `3_Train_fit_evaluate_classifier.ipynb`.
#
# 1. [Extract_training_data](1_Extract_training_data.ipynb)
# 2. **Inspect_training_data (this notebook)**
# 3. [Train_fit_evaluate_classifier](3_Train_fit_evaluate_classifier.ipynb)
#
# ***
#
# ## Additional information
#
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
# Digital Earth Africa data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
#
# **Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
# If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/digitalearthafrica/deafrica-sandbox-notebooks).
#
# **Last modified:** Dec 2020
#
| testing/southern_cropmask/2_Inspect_training_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# !pip install pandas
import sympy as sym
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
sym.init_printing()
# -
# ## Correlación
#
#
# La correlación entre las señales $f(t)$ y $g(t)$ es una operación que indica cuán parecidas son las dos señales entre sí.
#
# \begin{equation}
# (f \; \circ \; g)(\tau) = h(\tau) = \int_{-\infty}^{\infty} f(t) \cdot g(t + \tau) \; dt
# \end{equation}
#
# Observe que la correlación y la convolución tienen estructura similares.
#
# \begin{equation}
# f(t) * g(t) = \int_{-\infty}^{\infty} f(\tau) \cdot g(t - \tau) \; d\tau
# \end{equation}
# ## Señales periódicas
#
# La señal $y(t)$ es periódica si cumple con la condición $y(t+nT)=y(t)$ para todo $n$ entero. En este caso, $T$ es el periodo de la señal.
#
# 
#
# La señal seno es la oscilación más pura que se puede expresar matemáticamente. Esta señal surge al considerar la proyección de un movimiento circular uniforme.
#
#
# ## Serie de Fourier
#
#
# Si se combinan apropiadamente un conjunto de oscilaciones puras, como combinaciones lineales de señales desplazadas y escaladas en tiempo y amplitud, podría recrearse cualquiér señal periódica. Esta idea da lugar a las series de Fourier.
#
# \begin{equation}
# y(t) = \sum_{n=0}^{\infty} C_n \cdot cos(n \omega_0 t - \phi_n)
# \end{equation}
#
# La señal $y(t)$ es igual a una combinación de infinitas señales coseno, cada una con una amplitud $C_n$, una frecuencia $n \omega_0$ y un desfase $\phi_n$.
#
# También puede expresarse como:
#
# \begin{equation}
# y(t) = \sum_{n=0}^{\infty} A_n \cdot cos(n \omega_0 t) + B_n \cdot sin(n \omega_0 t)
# \end{equation}
#
# La serie queda definida si se encuentran los valores apropiados de $A_n$ y $B_n$ para todos los valores de $n$.
#
# Observe que:
# - $A_n$ debe ser más grande si $y(t)$ se "parece" más a un cos.
# - $B_n$ debe ser más grande si $y(t)$ se "parece" más a un sin.
# \begin{equation}
# y(t) = \sum_{n=0}^{\infty} A_n \cdot cos(n \omega_0 t) + B_n \cdot sin(n \omega_0 t)
# \end{equation}
#
# \begin{equation}
# (f \; \circ \; g)(\tau) = \int_{-\infty}^{\infty} f(t) \cdot g(t + \tau) \; dt
# \end{equation}
#
# \begin{equation}
# (y \; \circ \; sin_n)(\tau) = \int_{-\infty}^{\infty} y(t) \cdot sin(n \omega_0(t + \tau)) \; dt
# \end{equation}
#
#
# Considerando:
# - $\tau=0$ para no incluir desfases.
# - la señal $y(t)$ es periódica con periodo $T$.
#
# \begin{equation}
# (y \; \circ \; sin_n)(0) = \frac{1}{T} \int_{0}^{T} y(t) \cdot sin(n \omega_0 t) \; dt
# \end{equation}
#
# Esta expresión puede interpretarse como el parecido de una señal $y(t)$ a la señal $sin$ con crecuencia $n \omega_0$ promediado a lo largo de un periodo sin desfase del seno.
# Retomando la idea inicial
#
# \begin{equation}
# y(t) = \sum_{n=0}^{\infty} A_n \cdot cos(n \omega_0 t) + B_n \cdot sin(n \omega_0 t)
# \end{equation}
#
# donde
# \begin{equation}
# A_n = \frac{1}{T} \int_{0}^{T} y(t) \cdot cos(n \omega_0 t) \; dt
# \end{equation}
#
# \begin{equation}
# B_n = \frac{1}{T} \int_{0}^{T} y(t) \cdot sin(n \omega_0 t) \; dt
# \end{equation}
# Se recomienda al estudiante que encuentre la relación entre las Series anteriores y la siguiente alternativa para representar la Series de Fourier.
#
# \begin{equation}
# y(t) = \sum_{n=-\infty}^{\infty} C_n \cdot e^{j n \omega_0 t}
# \end{equation}
#
# donde
#
# \begin{equation}
# C_n = \frac{1}{T} \int_{0}^{T} y(t) \cdot e^{j n \omega_0 t} \; dt
# \end{equation}
# Los valores $C_n$ son el espectro de la señal periódica $y(t)$ y son una representación en el dominio de la frecuencia.
# **Ejemplo # 1**
#
# La señal $y(t) = sin(2 \pi t)$ es en sí misma una oscilación pura de periodo $T=1$.
# +
# Se define y como el seno de t
t = sym.symbols('t', real=True)
#T = sym.symbols('T', real=True)
T = 1
nw = sym.symbols('n', real=True)
delta = sym.DiracDelta(nw)
w0 = 2 * sym.pi / T
y = t
# y = 4*sym.sin(w0*t + 0.5) - 10
# y = sym.sin(w0*t)
# y = (t-0.5)*(t-0.5)
y
# -
# Aunque la sumatoria de las series de Fourier incluye infinitos términos, solamente se tomaran las primeras 3 componentes.
# +
n_max = 3
y_ser = 0
C = 0
ns = range(-n_max,n_max+1)
espectro = pd.DataFrame(index = ns,
columns= ['C','C_np','C_real','C_imag','C_mag','C_ang'])
for n in espectro.index:
C_n = (1/T)*sym.integrate(y*sym.exp(-1j*n*w0*t), (t,0,T)).evalf()
C = C + C_n*delta.subs(nw,nw-n)
y_ser = y_ser + C_n*sym.exp(1j*n*w0*t)
espectro['C'][n]=C_n
C_r = float(sym.re(C_n))
C_i = float(sym.im(C_n))
espectro['C_real'][n] = C_r
espectro['C_imag'][n] = C_i
espectro['C_np'][n] = complex(C_r + 1j*C_i)
espectro['C_mag'][n] = np.absolute(espectro['C_np'][n])
espectro['C_ang'][n] = np.angle(espectro['C_np'][n])
espectro
# -
# La señal reconstruida con un **n_max** componentes
y_ser
# +
plt.rcParams['figure.figsize'] = 7, 2
#g1 = sym.plot(y, (t,0,1), ylabel=r'Amp',show=False,line_color='blue',legend=True, label = 'y(t) original')
#g2 = sym.plot(sym.re(y_ser), (t,-1,2), ylabel=r'Amp',show=False,line_color='red',legend=True, label = 'y(t) reconstruida')
g1 = sym.plot(y, (t,0,1), ylabel=r'Amp',show=False,line_color='blue')
g2 = sym.plot(sym.re(y_ser), (t,-1,2), ylabel=r'Amp',show=False,line_color='red')
g1.extend(g2)
g1.show()
# -
C
plt.rcParams['figure.figsize'] = 7, 4
plt.stem(espectro.index,espectro['C_mag'])
# **Ejercicio**
#
# Use las siguientes funciones para definir un periodo de una señal periódica con periodo $T=1$:
#
# \begin{equation}
# y_1(t) = \begin{cases}
# -1 & 0 \leq t < 0.5 \\
# 1 & 0.5 \leq t < 1
# \end{cases}
# \end{equation}
#
#
# \begin{equation}
# y_2(t) = t
# \end{equation}
#
# \begin{equation}
# y_3(t) = 3 sin(2 \pi t)
# \end{equation}
#
# Varíe la cantidad de componentes que reconstruyen cada función y analice la reconstrucción obtenida y los valores de $C_n$
| .ipynb_checkpoints/04_Series_de_Fourier-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (SALib-impact)
# language: python
# name: salib-impact
# ---
# +
from SALib.test_functions import Sobol_G
from SALib import ProblemSpec
# Problem specification for the Sobol_G test function
sp = ProblemSpec({
'names': ['x1', 'x2', 'x3', 'x4', 'x5', 'x6', 'x7', 'x8'],
'groups': None,
'dists': None,
'bounds': [[0.0, 1.0],] * 8,
'outputs': ['Y'] # Optional output name
})
(sp.sample_morris(1000, num_levels=4)
.evaluate(Sobol_G.evaluate)
.analyze_morris(num_levels=4,
num_resamples=100))
# Show table of results
print(sp)
# Display plot of results
sp.plot();
# +
X = sp.samples
Y = sp.results
S = sp.analysis
# sp.set_samples(X)
# sp.samples = X
# -
sp.results
# +
# X = sp.samples
# Y = sp.results
# S = sp.analysis
sp.samples = X
sp.results = Y
sp.set_
# -
# Same as above without comments for publication
# +
from SALib.test_functions import Sobol_G
from SALib import ProblemSpec
sp = ProblemSpec({
'names': ['x1', 'x2', 'x3', 'x4', 'x5', 'x6', 'x7', 'x8'],
'groups': None,
'bounds': [[0.0, 1.0],] * 8,
'outputs': ['Y']
})
(sp.sample_morris(1000, num_levels=4)
.evaluate(Sobol_G.evaluate)
.analyze_morris(num_levels=4,
num_resamples=100))
print(sp)
ax = sp.plot()
# -
# `sp` is a custom dictionary object to maintain backwards-compatibility with the earlier procedural approach
sp
# Procedural style
# +
sp = ProblemSpec({
'names': ['x1', 'x2', 'x3', 'x4', 'x5', 'x6', 'x7', 'x8'],
'groups': None,
'bounds': [[0.0, 1.0],] * 8,
'outputs': ['Y']
})
sp.sample_morris(1000, num_levels=4)
sp.evaluate(Sobol_G.evaluate)
sp.analyze_morris(num_levels=4, num_resamples=100)
print(sp)
sp.plot()
# -
# ?sp.sample_
| notebooks/Box2b.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#all_slow
# -
from fastai2.vision.all import *
# # Tutorial - Migrating from Catalyst
#
# > Incrementally adding fastai goodness to your Catalyst training
# ## Catalyst code
# We're going to use the MNIST training code from Catalyst's README (as at August 2020), converted to a module.
from migrating_catalyst import *
# To use it in fastai, we first convert the Catalyst dict into a `DataLoaders` object:
data = DataLoaders(loaders['train'], loaders['valid']).cuda()
# ### Using callbacks
# In the Catalyst code, a training loop is defined manually, which is where the input tensor is flattened. In fastai, there's no need to define your own training loop - you can insert your own code into any part of the training process by using a callback, which can even modify data, gradients, the loss function, or anything else in the training loop:
@before_batch_cb
def cb(self, xb, yb): return (xb[0].view(xb[0].size(0), -1),),yb
# The Catalyst example also modifies the training loop to add metrics, but you can pass these directly to your `Learner` in fastai:
metrics=[accuracy,top_k_accuracy]
learn = Learner(data, model, loss_func=F.cross_entropy, opt_func=Adam,
metrics=metrics, cbs=cb)
# You can now fit your model. fastai supports many schedulers. We recommend using 1cycle:
learn.fit_one_cycle(1, 0.02)
# As you can see, migrating from Catalyst allowed us to replace 17 lines of code (in `CustomRunner`) with just 3 lines, and doesn't require you to change any of your existing data pipelines, optimizers, loss functions, models, etc. Once you've made this change, you can then benefit from fastai's rich set of callbacks, transforms, visualizations, and so forth.
#
# Note that fastai is very different from Catalyst, in that it is much more than just a training loop (although we're only using the training loop in this example) - it is a complete framework including GPU-accelerated transformations, end-to-end inference, integrated applications for vision, text, tabular, and collaborative filtering, and so forth. You can use any part of the framework on its own, or combine them together, as described in the [fastai paper](https://arxiv.org/abs/2002.04688).
# ### Changing the model
# Instead of using callbacks, in this case you can also simply change the model. Here we pull the `view()` out of the training loop, and into the model, using fastai's `Flatten` layer:
model = nn.Sequential(
Flatten(),
torch.nn.Linear(28 * 28, 10))
# We can now create a `Learner` and train without using any callbacks:
learn = Learner(data, model, loss_func=F.cross_entropy, opt_func=Adam, metrics=metrics)
learn.fit_one_cycle(1, 0.02)
| nbs/examples/migrating_catalyst.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 1. Adapt the Gradient_univ for polynomial 2x**2 +2x +1
# 2. Adapt the program for x**4
# +
from sympy import *
from sympy.matrices import Matrix
import numpy as np
from matplotlib import pyplot as plt
x = Symbol('x')
# Function
y = 2*x**2 +2*x + 1
# First derivative with respect to x
yprime = y.diff(x)
# Initial sequence
theta = 2
theta2 = 0
# Initiale the step size
alpha = .001
iterations = 0
check = 0
precision = 1/100000
iterationsMax = 100000
while True:
theta2 = theta - alpha*yprime.evalf(subs={x:theta})
iterations += 1
# If we make too much iterations our program
# stops and we need to check it to be sure the
# parameters are correct and it is working properly
if iterations > iterationsMax:
print("Too many iterations")
break
if abs(theta - theta2) < precision:
break
theta = theta2
print("Number of iterations:",iterations,"value of theta:",theta2,sep=" ")
plt.plot(theta,y.evalf(subs={x:theta}),marker='o',color='r')
space = np.linspace(-5,5,100)
data = np.array([y.evalf(subs={x: value}) for value in space])
plt.plot(space, data)
plt.show()
# +
# Function
y = x**4
# First derivative with respect to x
yprime = y.diff(x)
# Initial sequence
theta = 2
theta2 = 0
# Initiale the step size
alpha = .0001
iterations = 0
check = 0
precision = 1/100000
iterationsMax = 100000
while True:
theta2 = theta - alpha*yprime.evalf(subs={x:theta})
iterations += 1
# If we make too much iterations our program
# stops and we need to check it to be sure the
# parameters are correct and it is working properly
if iterations > iterationsMax:
print("Too many iterations")
break
if abs(theta - theta2) < precision:
break
theta = theta2
print("Number of iterations:",iterations,"value of theta:",theta2,sep=" ")
plt.plot(theta,y.evalf(subs={x:theta}),marker='o',color='r')
space = np.linspace(-5,5,100)
data = np.array([y.evalf(subs={x: value}) for value in space])
plt.plot(space, data)
plt.show()
# -
# 1. Make a 2 by 2 matrix of second partial derivatives (Hessian)
# 2. Adapt the program for a function of 2 variables. Precision?
# +
x,y = symbols('x y')
#function
z = x**2 + y**2
# Hessian
f11 = z.diff(x).diff(x)
f12 = z.diff(x).diff(y)
f21 = z.diff(y).diff(x)
f22 = z.diff(y).diff(y)
hessian = np.array([[f11, f12], [f21, f22]])
H = sympy.Matrix(hessian)
det = H.det()
print(hessian)
print('The determinant is', det)
if det >= 0:
print('The matrix positive semidefinite!')
else:
print('The matrix is not positive semidefinite!')
#det = f11 * f22 - f12 * f21
# +
# Function
z = x**2 + y**2 + 1
zprimex = z.diff(x)
zprimey = z.diff(y)
# Initial sequence
thetaX = 2
thetaY = 2
theta2X = 0
theta2Y = 0
# Initiale the step size
alpha = .0001
iterations = 0
check = 0
precision = 1/100000
iterationsMax = 100000
while True:
theta2X = thetaX - alpha*zprimex.evalf(subs={x:thetaX, y:thetaY})
theta2Y = thetaY - alpha*zprimey.evalf(subs={x:thetaX, y:thetaY})
iterations += 1
# If we make too much iterations our program
# stops and we need to check it to be sure the
# parameters are correct and it is working properly
if iterations > iterationsMax:
print("Too many iterations")
break
if ((thetaX - theta2X)**2 + (thetaY - theta2Y)**2) < precision**2:
break
thetaX = theta2X
thetaY = theta2Y
print("Number of iterations:",iterations,"value of theta:", theta2X, theta2Y,sep=" ")
#plt.plot(theta,y.evalf(subs={x:theta}),marker='o',color='r')
#space = np.linspace(-5,5,100)
#data = np.array([y.evalf(subs={x: value}) for value in space])
#plt.plot(space, data)
#plt.show()
# +
# Crazy Function
z = log(1-y**2 + x**4)
zprimex = z.diff(x)
zprimey = z.diff(y)
# Initial sequence
thetaX = 2
thetaY = 2
theta2X = 0
theta2Y = 0
# Initiale the step size
alpha = .0001
iterations = 0
check = 0
precision = 1/100000
iterationsMax = 100000
while True:
theta2X = thetaX - alpha*zprimex.evalf(subs={x:thetaX, y:thetaY})
theta2Y = thetaY - alpha*zprimey.evalf(subs={x:thetaX, y:thetaY})
iterations += 1
# If we make too much iterations our program
# stops and we need to check it to be sure the
# parameters are correct and it is working properly
if iterations > iterationsMax:
print("Too many iterations")
break
if ((thetaX - theta2X)**2 + (thetaY - theta2Y)**2) < precision**2:
break
thetaX = theta2X
thetaY = theta2Y
print("Number of iterations:",iterations,"value of theta:", theta2X, theta2Y,sep=" ")
#plt.plot(theta,y.evalf(subs={x:theta}),marker='o',color='r')
#space = np.linspace(-5,5,100)
#data = np.array([y.evalf(subs={x: value}) for value in space])
#plt.plot(space, data)
#plt.show()
# -
| Exercises/DaigleGradientDescentExercise.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Java
# language: java
# name: java
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Exemple de notebook avec Java
# -
# ## Explications
#
# Le kernel Java n'est pas installé par défaut avec Jupyter.
#
# Il faut installer [IJava](https://github.com/SpencerPark/IJava).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exemples
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Un exemple écrit par un de mes anciens élèves
#
# Cf. [ce petit challenge arithmético-algorithmique](https://perso.crans.org/besson/notebooks/F%C3%A9vrier%202021%20un%20mini%20challenge%20arithm%C3%A9tico-algorithmique.html).
# +
// ceci est du code Java 9 et pas Python !
// On a besoin des dépendances suivantes :
import java.util.Calendar; // pour Calendar.FEBRUARY, .YEAR, .MONDAY
import java.util.GregorianCalendar; // pour
import java.util.stream.IntStream; // pour cet IntStream
IntStream.rangeClosed(1994, 2077)
//.parallel() // ce .parallel() est inutile, il y a trop peu de valeurs
.mapToObj(i -> new GregorianCalendar(i, Calendar.FEBRUARY, 1))
.filter(calendar -> !calendar.isLeapYear(calendar.get(Calendar.YEAR)))
.filter(calendar -> calendar.get(Calendar.DAY_OF_WEEK) == Calendar.MONDAY)
.count();
# + [markdown] slideshow={"slide_type": "subslide"}
# ### D'autres exemples ?
#
# TODO: plus tard !
#
# TODO prendre quelques cellules de notebooks dans <https://perso.crans.org/besson/teach/INF1_L1_Rennes1_2020-21/>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Pour en apprendre plus
#
# - Ce WikiBooks : <https://fr.wikibooks.org/wiki/Programmation_Java> (sous licence libre) ;
| notebooks/Exemple de notebook avec Java.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
from IPython.display import Image
Image("C:\\Users\\User\\Desktop\\XGB.png")
# -
# Importing needed packages
from numpy import loadtxt
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
#Importing various models to compare
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
# +
# loading data from csv
mydata = loadtxt('C:\\Users\\User\\Desktop\\pima-indians-diabetes.csv', delimiter=",")
# spliting data into independent and dependent features
Independent_features = mydata[:,0:8]
Dependent_features = mydata[:,8]
# -
import pandas as pd
pd.DataFrame(mydata).shape
# split data into train and test sets
seed = 1
X_train, X_test, y_train, y_test = train_test_split(Independent_features, Dependent_features, test_size=0.20, random_state=seed)
# +
#Running various models
models = []
models.append(('LogisticRegression', LogisticRegression()))
models.append(('KNN', KNeighborsClassifier()))
models.append(('SVM', SVC()))
models.append(('XGB',XGBClassifier(eta=0.01,gamma=10))) #eta = 0.01,gamma = 10
import time
# evaluate each model in turn
results = []
names = []
scoring = 'accuracy'
for name, model in models:
start_time = time.time()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
# evaluate predictions
accuracy = accuracy_score(y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0),name)
print("--- %s seconds ---" % (time.time() - start_time))
# -
| Examples/XGBoost/XGBoost Demonstration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: encn423_env
# language: python
# name: python3
# ---
# # Wiki whā (4) tutorial Python notebook
#
# Use this notebook to help you code and calculate answers to the Week 4 tutorial questions. We'll assume you're familar with Python operations now, so get straight to it.
#
# Remeber, useful equations for the tutorial should be in your class notes or the lecture summary notes on Learn.
#
# Ready, set, code!
#
# Load packages - add any others you might need
import pandas as pd
import numpy as np
import math
# +
# Begin code!
# Question 1
cap = 30 #MW
T_h = 171+273
T_c = 15+273
#calculate carnot and PP efficiency
eta_c = 1-(T_c/T_h)
eta_PP = 0.5*eta_c
print('The carnot efficiency is {eff_c:.0%} and the power plant efficiency is {eff:.0%}'.format(eff_c=eta_c,eff=eta_PP))
#power output at 35C and decrease in efficiency
T_c2 = 35+273
eta_PP2 = 0.5*(1-(T_c2/T_h))
eff_decrease = 1-(eta_PP2/eta_PP)
print('At 35 degrees C, the decrease in efficiency is {eff_d:.0%}'.format(eff_d=eff_decrease))
#percent increase in efficiency with spray cooling
T_wb = 13+273
T_c3 = T_c2 - (T_c2-T_wb)*0.6
eta_PP3 = 0.5*(1-(T_c3/T_h))
eff_increase = (eta_PP3/eta_PP2)-1
print('With spray cooling, the efficiency increases {eff_i:.0%}'.format(eff_i=eff_increase))
#turbine efficiency
cap_t = cap/5 * 1000 #kW = kJ/s
H_steam = 2804 #kJ/kg
x_iso = 0.74
x_act = 0.82
H_v = 2591 #kJ/kg
H_l = 209 #kJ/kg
H_iso = x_iso*H_v + (1-x_iso)*H_l
H_act = x_act*H_v + (1-x_act)*H_l
eta_turbine = (H_steam - H_act)/(H_steam - H_iso)
print('The efficiency of the turbine is {eff_t:.0%}'.format(eff_t=eta_turbine))
#calculate mass flow rate
m_dot = cap_t/(H_steam-H_act) #kg/s
print('The mass flow rate through each turbine is {m:.2f} kg/s'.format(m=m_dot))
# +
# Question 2
T_g = 12+273
T_house = 21+273
T_h = 34.8+273
T_c = -34.4+273
#In summer
COP_g = T_g/(T_house-T_g)
COP_a = T_house/(T_h-T_house)
print('In summer, the GSHP has a COP of {cop_g:.2f} and the air HP has a COP of {cop_a:.2f}'.format(cop_g=COP_g, cop_a=COP_a))
#In winter
COP_g = T_house/(T_house-T_g)
COP_a = T_house/(T_house-T_c)
print('In winter, the GSHP has a COP of {cop_g:.2f} and the air HP has a COP of {cop_a:.2f}'.format(cop_g=COP_g, cop_a=COP_a))
# +
# Question 2 continued
heatloss = 105 #MJ/h
# load data
temp_df = pd.read_csv('calgary_hourly.csv')
#calculate COPs at each hour
temp_df['COP_a'] = [T_house/(T_house-(t+273)) * 0.7 if t < 21 else (t/((t+273)-T_house) * 0.7 if t > 21 else 0) for t in temp_df['temperature_degC']]
temp_df['COP_g'] = [T_house/(T_house-T_g) * 0.7 if t < 21 else T_g/(T_house-T_g)*0.7 for t in temp_df['temperature_degC']]
#calculate electricity
temp_df['e_air'] = [heatloss/60/60/cop*1000 if cop>0 else 0 for cop in temp_df['COP_a']]
temp_df['e_ground'] = [heatloss/60/60/cop*1000 for cop in temp_df['COP_g']]
#total costs
cost_air = temp_df['e_air'].sum()*0.14 #$
cost_ground = temp_df['e_ground'].sum()*0.14 #$
print('The air HP would cost ${c_a:.2f} per year and the GSHP would cost ${c_g:.2f} per year'.format(c_a=cost_air,c_g=cost_ground))
# +
#CHALLENGE
temp_df['date'] = pd.to_datetime(temp_df['date'])
temp_df['time_local'] = pd.to_datetime(temp_df['time_local'])
temp_df['month'] = [m.month for m in temp_df['date']]
temp_df['hour'] = [h.hour for h in temp_df['time_local']]
#set up cost structure
cost_w = [0.06,0.06,0.06,0.06,0.06,0.06,0.16,0.16,0.16,0.16,0.08,0.08,0.08,0.08,0.08,0.08,0.08,0.2,0.2,0.2,0.2,0.2,0.2,0.06]
cost_s = [0.04,0.04,0.04,0.04,0.04,0.04,0.14,0.14,0.14,0.14,0.07,0.07,0.07,0.07,0.07,0.07,0.07,0.18,0.18,0.18,0.18,0.18,0.18,0.04]
#summarise our electricity data for air and ground heating, in kWh
elec_air = pd.pivot_table(temp_df,index=['month'],columns=['hour'],values=['e_air'],aggfunc='sum')
elec_ground = pd.pivot_table(temp_df,index=['month'],columns=['hour'],values=['e_ground'],aggfunc='sum')
cost_a = 0
cost_g = 0
for c in range(0,len(elec_air.index)):
if c in [0,1,2,9,10,11]:
cost_a += np.sum(elec_air.iloc[c,:]*cost_w)
cost_g += np.sum(elec_ground.iloc[c,:]*cost_w)
else:
cost_a += np.sum(elec_air.iloc[c,:]*cost_s)
cost_g += np.sum(elec_ground.iloc[c,:]*cost_s)
#some rounding differences between excel and python pivoting -- acceptable
print('The air HP would cost ${c_a:.2f} per year and the GSHP would cost ${c_g:.2f} per year'.format(c_a=cost_a,c_g=cost_g))
# -
| Wiki_wha_sln.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import glob
import uuid
import datetime
import zipfile
import tqdm
import netCDF4
import pandas as pd
import xarray as xr
import dask
import dask.bag as db
from dask.diagnostics import ProgressBar
# -
def good_keys():
keep_keys = ['time',
'range',
'azimuth',
'elevation',
'radar_echo_classification',
'radar_estimated_rain_rate',
'velocity',
'total_power',
'reflectivity',
'cross_correlation_ratio',
'differential_reflectivity',
'corrected_differential_reflectivity',
'differential_phase',
'corrected_differential_phase',
'corrected_specific_differential_phase',
'spectrum_width',
'signal_to_noise_ratio',
'sweep_number',
'fixed_angle',
'sweep_start_ray_index',
'sweep_end_ray_index',
'sweep_mode',
'prt_mode',
'prt',
'nyquist_velocity',
'unambiguous_range',
'radar_beam_width_h',
'radar_beam_width_v',
'latitude',
'longitude',
'altitude',
'time_coverage_start',
'time_coverage_end',
'time_reference',
'volume_number',
'platform_type',
'instrument_type',
'primary_axis']
return keep_keys
def get_metadata(radar_start_date, radar_end_date):
maxlon = 132.385
minlon = 129.703
maxlat = -10.941
minlat = -13.552
origin_altitude = '50'
origin_latitude = '-12.2491'
origin_longitude = '131.0444'
unique_id = str(uuid.uuid4())
fieldnames = ['radar_echo_classification',
'radar_estimated_rain_rate',
'velocity',
'total_power',
'reflectivity',
'cross_correlation_ratio',
'differential_reflectivity',
'corrected_differential_reflectivity',
'differential_phase',
'corrected_differential_phase',
'corrected_specific_differential_phase',
'spectrum_width',
'signal_to_noise_ratio']
metadata = {'Conventions': "CF-1.6, ACDD-1.3",
'acknowledgement': 'This work has been supported by the U.S. Department of Energy Atmospheric Systems Research Program through the grant DE-SC0014063. Data may be freely distributed.',
'country': 'Australia',
'creator_email': '<EMAIL>',
'creator_name': '<NAME>',
'creator_url': 'github.com/vlouf',
'date_modified': datetime.datetime.now().isoformat(),
'field_names': ", ".join(fieldnames),
"geospatial_bounds": f"POLYGON(({minlon:0.6} {minlat:0.6},{minlon:0.6} {maxlat:0.6},{maxlon:0.6} {maxlat:0.6},{maxlon:0.6} {minlat:0.6},{minlon:0.6} {minlat:0.6}))",
'geospatial_lat_max': f'{maxlat:0.6}',
'geospatial_lat_min': f'{minlat:0.6}',
'geospatial_lat_units': "degrees_north",
'geospatial_lon_max': f'{maxlon:0.6}',
'geospatial_lon_min': f'{minlon:0.6}',
'geospatial_lon_units': "degrees_east",
'id': unique_id,
'institution': 'Bureau of Meteorology',
'instrument': 'radar',
'instrument_name': 'CPOL',
'instrument_type': 'radar',
'keywords': 'radar, tropics, Doppler, dual-polarization',
'licence': "Freely Distributed",
'naming_authority': 'au.org.nci',
'origin_altitude': origin_altitude,
'origin_latitude': origin_latitude,
'origin_longitude': origin_longitude,
'platform_is_mobile': 'false',
'processing_level': 'b1',
'project': "CPOL",
'publisher_name': "NCI",
'publisher_url': "nci.gov.au",
'references': 'doi:10.1175/JTECH-D-18-0007.1',
'site_name': 'Gunn Pt',
'source': 'radar',
'state': "NT",
'standard_name_vocabulary': 'CF Standard Name Table v71',
'summary': "Volumetric scan from CPOL dual-polarization Doppler radar (Darwin, Australia)",
'time_coverage_start': str(radar_start_date),
'time_coverage_end': str(radar_end_date),
'time_coverage_duration': "P10M",
'time_coverage_resolution': "PT10M",
'title': "radar PPI volume from CPOL",
'uuid': unique_id}
return metadata
def mkdir(path):
try:
os.mkdir(path)
except FileExistsError:
pass
return None
def extract_zip(inzip, path):
dates = os.path.basename(inzip).replace('.zip', '')
with zipfile.ZipFile(inzip) as zid:
zid.extractall(path=path)
namelist = [os.path.join(path, f) for f in zid.namelist()]
return dates, namelist
def update_dataset(radar_file, path):
dset = xr.open_dataset(radar_file)
radar_start_date = dset.time[0].values
radar_end_date = dset.time[-1].values
fname = "twp10cpolppi.b1.{}00.nc".format(pd.Timestamp(radar_start_date).strftime("%Y%m%d.%H%M"))
outfilename = os.path.join(path, fname)
if os.path.exists(outfilename):
print(f'File already exists.')
return None
keep_keys = good_keys()
keylist = [k for k in dset.variables.keys()]
for k in keylist:
if k not in keep_keys:
dset = dset.drop(k)
metadata = get_metadata(radar_start_date, radar_end_date)
metadata['product_version'] = "v" + dset.attrs['product_version']
metadata['version'] = "v" + dset.attrs['product_version']
metadata['date_created'] = dset.attrs['created'],
metadata['history'] = "created by <NAME> on raijin.nci.org.au at " + dset.attrs['created'] + " using Py-ART",
dset.attrs = metadata
dset.to_netcdf(outfilename, encoding={k:{'zlib': True} for k in dset.variables.keys()})
if not os.path.exists(outfilename):
print(f'Output file does not exist !!!.')
return None
del dset
return radar_file
def remove(flist):
for f in flist:
if f is None:
continue
try:
os.remove(f)
except FileNotFoundError:
pass
return None
zipdir = '/scratch/kl02/vhl548'
ziplist = sorted(glob.glob('/g/data/hj10/admin/cpol_level_1b/v2018/ppi/1998/*.zip'))
# +
# dates, namelist = extract_zip(ziplist[0], zipdir)
# outpath = f'/scratch/kl02/vhl548/tmpcpol/{dates}'
# mkdir(outpath)
# -
for zfile in tqdm.tqdm_notebook(ziplist[1:]):
dates, namelist = extract_zip(zfile, zipdir)
outpath = f'/scratch/kl02/vhl548/tmpcpol/{dates}'
mkdir(outpath)
bag = db.from_sequence([(n, outpath) for n in namelist]).starmap(update_dataset)
with ProgressBar():
rslt = bag.compute()
remove(rslt)
| notebook/Unzip - Zip folders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RL Environment
# language: python
# name: rl_env
# ---
# +
import numpy as np
from scipy import stats
import torch
import torch.optim as optim
import multiprocessing as mp
import logging
from datetime import datetime
from uniswapv3_simulator.pool import Uniswapv3Pool
from uniswapv3_simulator.optimization.environments import OneStepEnvironment, ScaleWrapper
from uniswapv3_simulator.optimization.ddpg.ddpg import (
DDPG,
DDPGTrainer,
DeepActorModel,
TrainArgs
)
from uniswapv3_simulator.optimization.ddpg.exploration_noise import GaussianProcess
from uniswapv3_simulator.optimization.ddpg.schedulers import ExponentialScheduler
timestamp = datetime.now().strftime('%y%m%d%H%M%S')
logging.basicConfig(
level=logging.INFO,
filename=f'./logs/rl_test_{timestamp}.log'
)
logging.getLogger('optimization').setLevel(logging.DEBUG)
# +
SEED = 1234
seed_seq = np.random.SeedSequence(entropy=SEED)
seeds = seed_seq.generate_state(8)
init_price = 100
liquidity_bins = [70, 80, 90, 100, 110, 120, 130]
# fees = stats.uniform(1e-4, 0.01 - 1e-4)
# mu = stats.uniform(-0.05, 0.1)
# sigma = stats.uniform(1e-4, 0.1 - 1e-4)
# alpha = stats.randint(1, 100 + 1)
# beta = stats.randint(100, 1000 + 1)
fees = stats.uniform(0.01, 0.0)
mu = stats.uniform(0.0, 0.0)
sigma = stats.uniform(0.05, 0.0)
alpha = stats.randint(50, 50 + 1)
beta = stats.randint(500, 500 + 1)
fees.random_state = seeds[0]
mu.random_state = seeds[1]
sigma.random_state = seeds[2]
alpha.random_state = seeds[3]
beta.random_state = seeds[4]
n_sims_per_step = 500
n_jobs = mp.cpu_count() - 1
env = OneStepEnvironment(
init_price, liquidity_bins,
fees, mu, sigma, alpha, beta,
n_sims_per_step=n_sims_per_step,
n_jobs=n_jobs, seed=seeds[5]
)
# -
print('Random Variables')
print(f'fees: mean={fees.mean():,.4f}, std={fees.std():,.4f}')
print(f'mu: mean={mu.mean():,.4f}, std={mu.std():,.4f}')
print(f'sigma: mean={sigma.mean():,.4f}, std={sigma.std():,.4f}')
print(f'alpha: mean={alpha.mean():,.2f}, std={alpha.std():,.2f}')
print(f'beta: mean={beta.mean():,.2f}, std={beta.std():,.2f}')
# +
def obs_scale_fn(obs):
# mu = np.array([0.0051, 0.0000, 0.0501, 50.50, 550.00])
# sigma = np.array([0.0029, 0.0289, 0.0288, 28.87, 260.10])
mu = np.zeros(5)
sigma = np.array([0.01, 1.0, 0.05, 50, 500])
return (obs - mu) / sigma
def action_scale_fn(action):
return action * 5e+4
def reward_scale_fn(reward):
return reward
env = ScaleWrapper(env, obs_scale_fn, action_scale_fn, reward_scale_fn)
# -
torch.manual_seed(seeds[6])
action_size = len(liquidity_bins) - 1
model = DeepActorModel(5, action_size, (128, 64), (128, 64))
agent = DDPG(
model=model,
gamma=0.99,
tau=1e-3,
optimizer=optim.Adam,
actor_optimizer_kwargs={
'lr': 1e-4,
'weight_decay': 1e-5
},
critic_optimizer_kwargs={
'lr': 1e-3,
'weight_decay': 1e-5
},
clip_gradients=5.0
)
train_args = TrainArgs(
train_steps=1000,
batch_size=64,
memory_size=100000,
exploration_noise=GaussianProcess,
noise_kwargs={
'size': (action_size, ),
'std': ExponentialScheduler(0.2, 0.01, 0.997)
},
update_start=50,
update_freq=4,
clip_actions=(1e-6, np.inf),
seed=seeds[7]
)
trainer = DDPGTrainer(agent, env, train_args)
model
# %%time
rewards = trainer.train()
| archive/RL Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# ## Coach Instructions
#
# This Python notebook can be used before the Hack to prepare data files for the participants. The intent is for the coach to get the latest Open Powerlitfing data from their website (linked in the challenge files). In order to suport the challenge structure of implementing an inital load followed by incremental loads, it is necessary to split up the full data set.
#
# Starting with the full data set, this notebook will extract the most recent meet activity (by date) and save it off to separate files. These files will simulate new, incremental data for the Hack team's incremental loads. All remaining historical data will be saved as the initial data load file.
#
# The coach can decide how many daily incremental files to create by changing the value of the <b>numDaysToSeparate</b> variable.
#
# Coach Data Preparation Steps:
#
# 1. Create a Synapse Workspace. Create a Spark pool in the workspace to run this notebook.
# 2. Save the latest Open Powerlifting data file to the root of the workspace storage account. Put the path to that file in the <b>pathToOpenPowerliftingCSV</b> variable in the first code cell.
# 3. Set the number of daily meet result files to create in the <b>numDaysToSeparate</b> variable in the second code cell.
# 4. Run the notebook. Daily meet result files (incremental loads) will be created in a subfolder called <b>dailyReports</b> in the workspace storage account. All remaining historical data will be saved in a single file in a subfolder called <b>initialData</b>.
#
# At the coach's discretion, this work can either be done by the coach before the Hack or by the Hack team as part of Challenge 1.
#
# ### File Format
# It is possible that the format of the Open Powerlifting data file will change over time. This notebook is only dependent on the Date and MeetName fields in that file. Provided those fields are present, with those names, the notebook should still produce the desired results with whatever the current format is.
#
#
#
# +
## Replace this variable with full path to the OPL data file in the root container of the Synapse Workspace storage account
pathToOpenPowerliftingCSV = "abfss://<rootContainerName>@<synpaseWorkspaceName>.dfs.core.windows.net/openpowerlifting-2021-01-05.csv"
df = spark.read.csv(pathToOpenPowerliftingCSV, header=True)
##display(df.limit(10))
df.createOrReplaceTempView("LiftingResults")
# + diagram={"activateDiagramType": 1, "aggData": "{\"Meets\":{\"2020-12-17\":2,\"2020-12-18\":3,\"2020-12-19\":16,\"2020-12-20\":3,\"2020-12-26\":1}}", "chartConfig": {"aggByBackend": false, "aggregation": "SUM", "category": "bar", "keys": ["Date"], "values": ["Meets"], "xLabel": "Date", "yLabel": "Meets"}, "isSql": false, "isSummary": false, "previewData": {"filter": null}}
numDaysToSeparate = 5
dfMeetsByDate = spark.sql("SELECT Date, COUNT(DISTINCT MeetName) AS Meets, COUNT(*) AS Participants FROM LiftingResults GROUP BY Date ORDER BY Date DESC")
listMostRecentDates = dfMeetsByDate.take(numDaysToSeparate)
##dfMeetsByDate.show()
display(listMostRecentDates)
# +
from notebookutils import mssparkutils
from pyspark.sql.functions import col, asc,desc
##Prep target folder path for daily reports
if list(filter(lambda x : x.name == "dailyReports", mssparkutils.fs.ls("/"))):
mssparkutils.fs.rm("/dailyReports", True)
mssparkutils.fs.mkdirs("/dailyReports")
##Iterate through the N most recent dates and create a CSV file per date, in the target folder path
for row in listMostRecentDates:
activityDate = row[0]
outputFilename = "daily-results-" + activityDate + ".csv"
outputFullPath = "/" + outputFilename
df.where(df.Date == activityDate).coalesce(1).write.mode("overwrite").option("header", "true").option("emptyValue", "").csv(outputFullPath)
files = mssparkutils.fs.ls(outputFullPath)
partFilename = list(filter(lambda x : x.name.endswith("csv"), files))
for filename in partFilename:
mssparkutils.fs.mv(filename.path, "/dailyReports/" + outputFilename)
mssparkutils.fs.rm(outputFullPath, True)
##Done with daily reports
##Prep target folder path for initial data. This is simply the OpenPowerLifting dataset with the N most recent dates removed
if list(filter(lambda x : x.name == "initialData", mssparkutils.fs.ls("/"))):
mssparkutils.fs.rm("/initialData", True)
mssparkutils.fs.mkdirs("/initialData")
outputFilename = "openpowerlifting-initial-data.csv"
outputFullPath = "/" + outputFilename
##Filter the dataframe to exclude the N most recent dates and write it out to a single CSV in the \initalData folder
initialDataBeforeThisDate = listMostRecentDates[numDaysToSeparate - 1][0]
dfInitialData = df.where(df.Date < initialDataBeforeThisDate).orderBy(col("Date").desc())
dfInitialData.coalesce(1).write.mode("overwrite").option("header", "true").option("emptyValue", "").csv(outputFullPath)
files = mssparkutils.fs.ls(outputFullPath)
partFilename = list(filter(lambda x : x.name.endswith("csv"), files))
for filename in partFilename:
mssparkutils.fs.mv(filename.path, "/initialData/" + outputFilename)
mssparkutils.fs.rm(outputFullPath, True)
##Done with initial data
| 024-DoYouEvenSynapse/Coach/Resources/Lab Data Prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: ipykernel_py2
# ---
Participants = ['John', 'Leila', 'Gregory', 'Cate']
Participants
print Participants[1]
Participants[-1]
Participants[-2]
Participants[3]='Maria'
Participants
del Participants[2]
Participants
Participants[2]
| 11 - Introduction to Python/7_Sequences/1_Lists (8:18)/Lists - Lecture_Py2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyzing the UncertaintyForest Class by Reproducing Figure 1
#
# This set of three tutorials (`uncertaintyforest_running_example.ipynb`,`uncertaintyforest_fig1.ipynb`, and `uncertaintyforest_fig2.ipynb`) will explain the UncertaintyForest class. After following these tutorials, you should have the ability to run UncertaintyForest on your own machine and generate Figures 1 and 2 from [this paper](https://arxiv.org/pdf/1907.00325.pdf).
#
# If you haven't seen it already, take a look at other tutorials to setup and install the ProgLearn package: `installation_guide.ipynb`.
#
# *Goal: Run the UncertaintyForest class to produce the results from Figure 1*
#
# *Note: Figure 1 refers to Figure 1 from [this paper](https://arxiv.org/pdf/1907.00325.pdf)*
# ## Import Required Packages
# +
import numpy as np
from sklearn.ensemble import RandomForestClassifier
from sklearn.calibration import CalibratedClassifierCV
from proglearn.forest import UncertaintyForest
from functions.unc_forest_tutorials_functions import generate_data, estimate_posterior, plot_posterior, plot_variance, plot_fig1
# -
# ## Specify Parameters
# +
# The following are two sets of parameters.
# The first are those that were actually used to produce Figure 1.
# These take a long time to actually run since there are 6000 data points.
# Below those, you'll find some testing parameters so that you can see the results more quickly.
# Here are the "Real Parameters"
#n = 6000
#mean = 1
#var = 1
#num_trials = 100
#X_eval = np.linspace(-2, 2, num = 30).reshape(-1, 1)
#n_estimators = 300
#num_plotted_trials = 10
# Here are the "Test Parameters"
n = 300 # number of data points
mean = 1 # mean of the data
var = 1 # variance of the data
num_trials = 3 # number of trials to run
X_eval = np.linspace(-2, 2, num = 10).reshape(-1, 1) # the evaluation span (over X) for the plot
n_estimators = 200 # the number of estimators
num_plotted_trials = 2 # the number of "fainter" lines to be displayed on the figure
# -
# ## Specify Learners
#
# Now, we'll specify which learners we'll compare. Figure 1 uses three different learners.
# +
# Algorithms used to produce Figure 1
algos = [
{
'instance': RandomForestClassifier(n_estimators = n_estimators),
'label': 'CART',
'title': 'CART Forest',
'color': "#1b9e77",
},
{
'instance': CalibratedClassifierCV(base_estimator=RandomForestClassifier(n_estimators = n_estimators // 5),
method='isotonic',
cv = 5),
'label': 'IRF',
'title': 'Isotonic Reg. Forest',
'color': "#fdae61",
},
{
'instance': UncertaintyForest(n_estimators = n_estimators, tree_construction_proportion = 0.4, kappa = 3.0),
'label': 'UF',
'title': 'Uncertainty Forest',
'color': "#F41711",
},
]
# Plotting parameters
parallel = True
# -
# ## Generate predicted posteriors
#
# Now, we'll run the code to obtain the results that will be displayed in Figure 1.
# This is the code that actually generates data and predictions.
for algo in algos:
algo['predicted_posterior'] = estimate_posterior(algo, n, mean, var, num_trials, X_eval, parallel = parallel)
# ## Create Figure 1
plot_fig1(algos, num_plotted_trials, X_eval)
| docs/tutorials/uncertaintyforest_fig1.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// <a name="top"></a><img src="images/chisel_1024.png" alt="Chisel logo" style="width:480px;" />
// ##### Module 2.6: More on ChiselTest
// **Prev: [Putting it all Together: An FIR Filter](2.5_exercise.ipynb)**<br>
// **Next: [Generators: Parameters](3.1_parameters.ipynb)**
//
// ## Motivation
// The Chisel team has been working on an improved testing framework. "ChiselTest", it provides the following improvements .
//
// - suitable for both unit tests and system integration tests
// - designed for composable abstractions and layering
// - highly usable, encouraging unit tests by making it as easy, painless (avoiding boilerplate and other nonsense), and useful as possible to write them
//
// ### Planned
// - ablity to target multiple backends and simulators (possibly requiring a link to Scala, if the testvector is not static, or using a limited test constructing API subset, when synthesizing to FPGA)
// - will be included in base chisel3, to avoid packaging and dependency nightmares
//
//
// ## Setup
val path = System.getProperty("user.dir") + "/source/load-ivy.sc"
interp.load.module(ammonite.ops.Path(java.nio.file.FileSystems.getDefault().getPath(path)))
import chisel3._
import chisel3.util._
import chisel3.experimental._
import chisel3.experimental.BundleLiterals._
import chisel3.tester._
import chisel3.tester.RawTester.test
// >This bootcamp requires some slight differences from the imports you might see
// elsewhere for chisel. The `import chisel3.tester.RawTester.test` brings in
// version of `test(...)` below that is designed specifically for the bootcamp
// ---
// # Basic Tester implementation
//
// ChiselTest starts with the same basic operations as iotesters. Here's a brief summary of the basic
// functionality mapping between the older iotesters and the new ChiselTest
//
// | | iotesters | ChiselTest |
// | :---- | :--- | :--- |
// | poke | poke(c.io.in1, 6) | c.io.in1.poke(6.U) |
// | peek | peek(c.io.out1) | c.io.out1.peek() |
// | expect | expect(c.io.out1, 6) | c.io.out1.expect(6.U) |
// | step | step(1) | c.io.clock.step(1) |
// | initiate | Driver.execute(...) { c => | test(...) { c => |
//
// Let's start by looking at the simple pass through module from 2.1
// Chisel Code, but pass in a parameter to set widths of ports
class PassthroughGenerator(width: Int) extends Module {
val io = IO(new Bundle {
val in = Input(UInt(width.W))
val out = Output(UInt(width.W))
})
io.out := io.in
}
// Using the old style a simple test would look like this
//
// ```scala
// val testResult = Driver(() => new Passthrough()) {
// c => new PeekPokeTester(c) {
// poke(c.io.in, 0) // Set our input to value 0
// expect(c.io.out, 0) // Assert that the output correctly has 0
// poke(c.io.in, 1) // Set our input to value 1
// expect(c.io.out, 1) // Assert that the output correctly has 1
// poke(c.io.in, 2) // Set our input to value 2
// expect(c.io.out, 2) // Assert that the output correctly has 2
// }
// }
// assert(testResult) // Scala Code: if testResult == false, will throw an error
// println("SUCCESS!!") // Scala Code: if we get here, our tests passed!
// ```
//
//
test(new PassthroughGenerator(16)) { c =>
c.io.in.poke(0.U) // Set our input to value 0
c.io.out.expect(0.U) // Assert that the output correctly has 0
c.io.in.poke(1.U) // Set our input to value 1
c.io.out.expect(1.U) // Assert that the output correctly has 1
c.io.in.poke(2.U) // Set our input to value 2
c.io.out.expect(2.U) // Assert that the output correctly has 2
}
// >Just to illustrate the way ChiselTest advances the clock we can
// add some `step` operations to the previous examples.
test(new PassthroughGenerator(16)) { c =>
c.io.in.poke(0.U) // Set our input to value 0
c.clock.step(1) // advance the clock
c.io.out.expect(0.U) // Assert that the output correctly has 0
c.io.in.poke(1.U) // Set our input to value 1
c.clock.step(1) // advance the clock
c.io.out.expect(1.U) // Assert that the output correctly has 1
c.io.in.poke(2.U) // Set our input to value 2
c.clock.step(1) // advance the clock
c.io.out.expect(2.U) // Assert that the output correctly has 2
}
// ---
// ## What to notice in the above example
//
// ChiselTest's `test` method requires a bit less boiler plate. What was the `PeekPokeTester` is now
// built into the process.
//
// The `poke` and `expect` methods are now part of each individual `io` element.
// This gives important hints the the tester to make better checking of types.
// The `peek` and `step` operations are also now methods on `io` elements.
//
// Another difference is that values poked and expected are Chisel literals.
// Although pretty simple here, it also provides stronger checking in more advanced and interesting examples.
// This will be further enhanced with coming improvements in the ability to specify `Bundle` literals
//
//
// # Modules with Decoupled Interfaces
// In this section we will look at some of the tester2's tools for working with `Decoupled` interfaces.
// `Decoupled` takes a chisel data type and provides it with `ready` and `valid` signals.
// ChiselTest provides some nice tools for automating and reliably testing these interfaces.
//
// ## A queue example
// The `QueueModule` passes through data whose type is determined by `ioType`. There are `entries` state elements inside the `QueueModule` meaning it can hold that many elements before it exerts backpressure.
case class QueueModule[T <: Data](ioType: T, entries: Int) extends MultiIOModule {
val in = IO(Flipped(Decoupled(ioType)))
val out = IO(Decoupled(ioType))
out <> Queue(in, entries)
}
// > Note the `case` class modifer is not generally required but seems to be in order for
// this example to be re-used in multiple cells in Jupyter
// ## EnqueueNow and expectDequeueNow
// *ChiselTest* has some built in methods for dealing with circuits with Decoupled interfaces in the IOs. In this example we will see how to insert and extract values from the `queue`.
//
// | method | description |
// | :--- | :--- |
// | enqueueNow | Add (enqueue) one element to a `Decoupled` input interface |
// | expectDequeueNow | Removes (dequeues) one element from a `Decoupled` output interface |
// ---
//
//
// >Note: There is some required boiler plate `initSource`, `setSourceClock`, etc that is necessary to ensure that the `ready` and `valid` fields are
// all initialized correctly at the beginning of the test.
//
test(QueueModule(UInt(9.W), entries = 200)) { c =>
// Example testsequence showing the use and behavior of Queue
c.in.initSource()
c.in.setSourceClock(c.clock)
c.out.initSink()
c.out.setSinkClock(c.clock)
val testVector = Seq.tabulate(200){ i => i.U }
testVector.zip(testVector).foreach { case (in, out) =>
c.in.enqueueNow(in)
c.out.expectDequeueNow(out)
}
}
// ## EnqueueSeq and DequeueSeq
// Now we are going to introduce two new methods that deal with enqueuing and dequeuing operations in single operations.
//
// | method | description |
// | :--- | :--- |
// | enqueueSeq | Continues to add (enqueue) elements from the `Seq` to a `Decoupled` input interface, one at a time, until the sequence is exhausted |
// | expectDequeueSeq | Removes (dequeues) elements from a `Decoupled` output interface, one at a time, and compares each one to the next element of the `Seq` |
// ---
// > Note: The example below works fine but, as written, the `enqueueSeq` must finish before the `expectDequeueSeq` can begin. This example would fail if the `testVector`'s size is made larger than the queue depth, because the queue would fill up and not be able to complete the `enqueueSeq`. Try it yourself to see what the failure looks like. In the next section we will show to construct this type of test properly.
//
test(QueueModule(UInt(9.W), entries = 200)) { c =>
// Example testsequence showing the use and behavior of Queue
c.in.initSource()
c.in.setSourceClock(c.clock)
c.out.initSink()
c.out.setSinkClock(c.clock)
val testVector = Seq.tabulate(100){ i => i.U }
c.in.enqueueSeq(testVector)
c.out.expectDequeueSeq(testVector)
}
// > One more important take away from the last section is that the functions we just saw, `enqueueNow`,
// `enqueueSeq`, `expectDequeueNow`, and `expectDequeueSeq` are not complicated special case logic in ChiselTest.
// Rather they are examples of the kinds of harness building that ChiselTest encourages you to build from the ChiselTest primitives. To see how these methods are implemented check out [TestAdapters.scala](https://github.com/ucb-bar/chisel-testers2/blob/d199c5908828d0be5245f55fce8a872b2afb314e/src/main/scala/chisel3/tester/TestAdapters.scala)
// # Fork and Join in ChiselTest
// In this section we will look at running sections of a unit test concurrently. In order to do this we will introduce two new features of testers2.
//
// | method | description |
// | :--- | :--- |
// | fork | launches a concurrent code block, additional forks can be run concurrently to this one via the .fork appended to end of the code block of the preceeding fork |
// | join | re-unites multiple related forks back into the calling thread |
// ---
//
// In the example below two `fork`s are chained together, and then `join`ed. In the first `fork` block the `enqueueSeq` will continue to add elements until exhausted. The second `fork` block will `expectDequeueSeq` on each cycle when data is available.
//
// >The threads created by fork are run in a deterministic order, largely according to their order as specified in code, and certain bug-prone operations that depend on other threads are forbidden with runtime checks.
//
// +
test(QueueModule(UInt(9.W), entries = 200)) { c =>
// Example testsequence showing the use and behavior of Queue
c.in.initSource()
c.in.setSourceClock(c.clock)
c.out.initSink()
c.out.setSinkClock(c.clock)
val testVector = Seq.tabulate(300){ i => i.U }
fork {
c.in.enqueueSeq(testVector)
}.fork {
c.out.expectDequeueSeq(testVector)
}.join()
}
// -
// ## Using Fork and Join with GCD
// In this section we will use the fork join methods to implement tests of *Greatest Common Denominator* **GCD**.
// Let's start by defining our IO bundles. We are going to add a bit of boiler plate here to allow us to use `Bundle` *literals*. As the comments say, it is hoped that we will soon have support for autogeneration of the literal support code.
class GcdInputBundle(val w: Int) extends Bundle {
val value1 = UInt(w.W)
val value2 = UInt(w.W)
}
class GcdOutputBundle(val w: Int) extends Bundle {
val value1 = UInt(w.W)
val value2 = UInt(w.W)
val gcd = UInt(w.W)
}
// Now let's look at a *Decoupled* version of **GCD**. We've modified it a bit here to use the `Decoupled` wrapper that adds a `ready` and a `valid` signal to the input and output Bundle. The `Flipped` wrapper takes the `Decoupled` `GcdInputBundle` which by default is created as an output and converts each field to the opposite direction (recursively). The data elements of the bundled arguments to `Decoupled` are placed in the top level field `bits`.
/**
* Compute GCD using subtraction method.
* Subtracts the smaller of registers x and y from the larger until register y is zero.
* value input register x is then the Gcd
* returns a packet of information with the two input values and their GCD
*/
class DecoupledGcd(width: Int) extends MultiIOModule {
val input = IO(Flipped(Decoupled(new GcdInputBundle(width))))
val output = IO(Decoupled(new GcdOutputBundle(width)))
val xInitial = Reg(UInt())
val yInitial = Reg(UInt())
val x = Reg(UInt())
val y = Reg(UInt())
val busy = RegInit(false.B)
val resultValid = RegInit(false.B)
input.ready := ! busy
output.valid := resultValid
output.bits := DontCare
when(busy) {
// during computation keep subtracting the smaller from the larger
when(x > y) {
x := x - y
}.otherwise {
y := y - x
}
when(y === 0.U) {
// when y becomes zero computation is over, signal valid data to output
output.bits.value1 := xInitial
output.bits.value2 := yInitial
output.bits.gcd := x
output.valid := true.B
busy := false.B
}
}.otherwise {
when(input.valid) {
// valid data available and no computation in progress, grab new values and start
val bundle = input.deq()
x := bundle.value1
y := bundle.value2
xInitial := bundle.value1
yInitial := bundle.value2
busy := true.B
resultValid := false.B
}
}
}
// Our test looks pretty much the same as the earlier Queue tests.
// But there's more going on because the computation take multiple cycles so the input enqueue process is blocked as each GCD is computed.
// The good news is that test side of this is simple and consistent across different Decoupled circuits.
//
// Also introduced here is the new Chisel3 `Bundle` literal notation. Consider the line
// ```scala
// new GcdInputBundle(16)).Lit(_.value1 -> x.U, _.value2 -> y.U)
// ```
// `GcdInputBundle` defined above has two fields `value1` and `value2`.
// We create a bundle literal by first creating a bundle then calling its `.Lit` method.
// That method takes a variable argument list of key/value pairs, where the key (e.g. `_.value`) is the field name and the value (e.g. x.U) is a chisel hardware literal, the Scala `Int` x is converted into a Chisel `UInt` literal.
// The `_.` in front of the field name is necessary to bind the name value to the bundle internals.
//
// >This may not be the perfect notation but in extensive development discussions it was viewed as
// the best balance between minimizing boilerplate and the notational limitations available in Scala.
//
test(new DecoupledGcd(16)) { dut =>
dut.input.initSource().setSourceClock(dut.clock)
dut.output.initSink().setSinkClock(dut.clock)
val testValues = for { x <- 1 to 10; y <- 1 to 10} yield (x, y)
val inputSeq = testValues.map { case (x, y) =>
(new GcdInputBundle(16)).Lit(_.value1 -> x.U, _.value2 -> y.U)
}
val resultSeq = testValues.map { case (x, y) =>
new GcdOutputBundle(16).Lit(_.value1 -> x.U, _.value2 -> y.U, _.gcd -> BigInt(x).gcd(BigInt(y)).U)
}
fork {
dut.input.enqueueSeq(inputSeq)
}.fork {
dut.output.expectDequeueSeq(resultSeq)
}.join()
}
// ---
// # You're done!
//
// [Return to the top.](#top)
| 2.6_chiseltest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# +
import os
import math
import datetime
import numpy as np
import pandas as pd
import time
import os.path
from shapely.geometry import MultiLineString
# -
# ## step 1 obtain OSM network
# +
"Get the OSM Network"
network_type = 'auto'
import osm2gmns as og
net = og.getNetFromOSMFile('tmc.osm',network_type=(network_type), default_lanes=True, default_speed=True)
og.consolidateComplexIntersections(net)
og.outputNetToCSV(net, output_folder=os.getcwd())
# -
# ## step 2 Convert tmc Data into node and link
"Convert TMC Data into GMNS Format"
tmc = pd.read_csv('TMC_Identification.csv')
tmc = tmc.drop_duplicates(subset=['direction','road_order']).sort_values(by=['direction','road_order'])
tmc = tmc.reset_index()
tmc = tmc.drop(['index'], 1)
# +
'''build node.csv'''
print('converting tmc data into gmns format...')
p=1
node_tmc = pd.DataFrame()
node_tmc['name'] = None
node_tmc['x_coord'] = None
node_tmc['y_coord'] = None
node_tmc['z_coord'] = None
node_tmc['node_type'] = None
node_tmc['ctrl_type'] = None
node_tmc['zone_id'] = None
node_tmc['geometry'] = None
for i in range(0,len(tmc)-1):
if tmc.loc[i+1,'road_order'] > tmc.loc[i,'road_order']:
node_tmc = node_tmc.append({'name': tmc.loc[i,'tmc'],\
'x_coord': tmc.loc[i,'start_longitude'], \
'y_coord': tmc.loc[i,'start_latitude'],\
'z_coord': None,\
'node_type': 'tmc_start',\
'ctrl_type': None,\
'zone_id': None,\
'geometry': "POINT (" + tmc.loc[i,'start_longitude'].astype(str) + " " + tmc.loc[i,'start_latitude'].astype(str) +")"}, ignore_index=True)
else:
node_tmc = node_tmc.append({'name': tmc.loc[i,'tmc'],\
'x_coord': tmc.loc[i,'start_longitude'], \
'y_coord': tmc.loc[i,'start_latitude'],\
'z_coord': None,\
'node_type': 'tmc_start',\
'ctrl_type': None,\
'zone_id': None,\
'geometry': "POINT (" + tmc.loc[i,'start_longitude'].astype(str) + " " + tmc.loc[i,'start_latitude'].astype(str) +")"}, ignore_index=True)
node_tmc = node_tmc.append({'name': tmc.loc[i,'tmc']+'END',\
'x_coord': tmc.loc[i,'end_longitude'], \
'y_coord': tmc.loc[i,'end_latitude'],\
'z_coord': None,\
'node_type': 'tmc_end',\
'ctrl_type': None,\
'zone_id': None,\
'geometry': "POINT (" + tmc.loc[i,'end_longitude'].astype(str) + " " + tmc.loc[i,'end_latitude'].astype(str) +")"}, ignore_index=True)
if i > p/10 * len(tmc):
print(str(p*10)+"%"+' nodes completed!')
p = p + 1
node_tmc = node_tmc.append({'name': tmc.loc[i+1,'tmc'],\
'x_coord': tmc.loc[i+1,'start_longitude'], \
'y_coord': tmc.loc[i+1,'start_latitude'],\
'z_coord': None,\
'node_type': 'tmc_start',\
'ctrl_type': None,\
'zone_id': None,\
'geometry': "POINT (" + tmc.loc[i+1,'start_longitude'].astype(str) + " " + tmc.loc[i+1,'start_latitude'].astype(str) +")"}, ignore_index=True)
node_tmc = node_tmc.append({'name': tmc.loc[i+1,'tmc']+'END',\
'x_coord': tmc.loc[i+1,'end_longitude'], \
'y_coord': tmc.loc[i+1,'end_latitude'],\
'z_coord': None,\
'node_type': 'tmc_end',\
'ctrl_type': None,\
'zone_id': None,\
'geometry': "POINT (" + tmc.loc[i+1,'end_longitude'].astype(str) + " " + tmc.loc[i+1,'end_latitude'].astype(str) +")"}, ignore_index=True)
node_tmc.index.name = 'node_id'
node_tmc.index += 100000001 #index from 0
node_tmc.to_csv('node_tmc.csv')
print('node_tmc.csv (' + str(len(node_tmc)) + ' nodes' + ') generated!')
# -
node_tmc
# +
'''build link_tmc.csv'''
p = 1
link_tmc = pd.DataFrame()
link_tmc['name'] = None
link_tmc['corridor_id'] = None
link_tmc['corridor_link_order'] = None
link_tmc['from_node_id'] = None
link_tmc['to_node_id'] = None
link_tmc['directed'] = None
link_tmc['geometry_id'] = None
link_tmc['geometry'] = None
link_tmc['dir_flag'] = None
link_tmc['length'] = None
link_tmc['grade'] = None
link_tmc['facility_type'] = None
link_tmc['capacity'] = None
link_tmc['free_speed'] = None
link_tmc['lanes'] = None
for i in range(0,len(tmc)):
link_tmc = link_tmc.append({'name': tmc.loc[i,'tmc'],\
'corridor_id': tmc.loc[i,'road']+'_'+tmc.loc[i,'direction'],\
'corridor_link_order' : tmc.loc[i,'road_order'],\
'from_node_id': node_tmc[(node_tmc['x_coord']==tmc.loc[i,'start_longitude']) & (node_tmc['y_coord']==tmc.loc[i,'start_latitude'])].index.values[0], \
'to_node_id': node_tmc[(node_tmc['x_coord']==tmc.loc[i,'end_longitude']) & (node_tmc['y_coord']==tmc.loc[i,'end_latitude'])].index.values[0],\
'directed': 1,\
'geometry_id': None,\
'geometry': "LINESTRING (" + tmc.loc[i,'start_longitude'].astype(str) + " " + tmc.loc[i,'start_latitude'].astype(str) + "," +\
tmc.loc[i,'end_longitude'].astype(str) +" "+ tmc.loc[i,'end_latitude'].astype(str) + ")",\
'dir_flag': 1,\
'length': tmc.loc[i,'miles'],\
'grade': None,\
'facility_type': 'interstate' if tmc.loc[i,'road'][0] == 'I'else None ,\
'capacity':None,\
'free_speed':None,\
'lanes': None}, ignore_index=True)
if i > p/10 * len(tmc):
print(str(p*10)+"%"+' links completed!')
p = p + 1
link_tmc.index.name = 'link_id'
link_tmc.index += 100000001
link_tmc.to_csv('link_tmc.csv')
print('link_tmc.csv (' + str(len(link_tmc)) + ' links' + ') generated!')
# -
link_tmc
# +
## reading by detid
reading = pd.read_csv('Reading.csv')
reading = reading.loc[0:5000]
reading_dict = {}
gp = reading.groupby('tmc_code')
for key, form in gp:
reading_dict[key] = {
'measurement_tstamp':form['measurement_tstamp'].tolist(),
'speed':form['speed'].tolist()
}
# +
'''build measurement_tmc.csv'''
measurement_tmc = pd.DataFrame()
measurement_tmc['link_id_tmc'] = None
measurement_tmc['corridor_id'] = None
measurement_tmc['corridor_link_order'] = None
measurement_tmc['from_node_id'] = None
measurement_tmc['to_node_id'] = None
measurement_tmc['time_period'] = None
measurement_tmc['date'] = None
measurement_tmc['geometry'] = None
measurement_tmc['volume'] = None
measurement_tmc['travel_time'] = None
measurement_tmc['speed'] = None
measurement_tmc['reference_speed'] = None
measurement_tmc['density'] = None
measurement_tmc['queue'] = None
measurement_tmc['notes'] = None
k=0
p=1
measurement_tmc_dict = {}
for i in link_tmc.index:
try:
reading_dict_selected = reading_dict[link_tmc['name'][i]]
for j in range(0,len(reading_dict_selected['measurement_tstamp'])):
measurement_tmc_dict[k] = {'link_id_tmc': i,\
'corridor_id': link_tmc['corridor_id'][i],\
'corridor_link_order' : link_tmc['corridor_link_order'][i],\
'from_node_id': link_tmc.loc[i,'from_node_id'], \
'to_node_id': link_tmc.loc[i,'to_node_id'], \
'time_period': reading_dict_selected['measurement_tstamp'][j][11:13]+\
reading_dict_selected['measurement_tstamp'][j][14:16]+'_'+\
reading_dict_selected['measurement_tstamp'][j+1][11:13]+\
reading_dict_selected['measurement_tstamp'][j+1][14:16],\
'date': reading_dict_selected['measurement_tstamp'][j][:10],\
'geometry': link_tmc['geometry'][i],\
'volume': None,\
'travel_time': None,\
'speed':round(np.mean(reading_dict_selected['speed'][j:j+1])),\
'reference_speed': None,\
'density': None,\
'queue': None,\
'notes': None }
k += 1
except:
measurement_tmc_dict[k] = {'link_id_tmc': i,\
'corridor_id': link_tmc['corridor_id'][i],\
'corridor_link_order' : link_tmc['corridor_link_order'][i],\
'from_node_id': link_tmc.loc[i,'from_node_id'], \
'to_node_id': link_tmc.loc[i,'to_node_id'], \
'time_period': None,\
'date': None,\
'geometry': link_tmc['geometry'][i],\
'volume': None,\
'travel_time': None,\
'speed': None,\
'reference_speed': None,\
'density': None,\
'queue': None,\
'notes': None }
k += 1
if link_tmc.index.get_loc(i) > p/10 * len(link_tmc):
print(str(p*10)+"%"+' measurement_tmc completed!')
p = p + 1
measurement_tmc = pd.DataFrame(measurement_tmc_dict).transpose()
measurement_tmc = measurement_tmc.dropna(subset=['time_period']) #remove na at the end of day of unrecorded ones
measurement_tmc = measurement_tmc.reset_index()
measurement_tmc = measurement_tmc.drop(['index'], 1)
measurement_tmc.to_csv('measurement_tmc.csv',index = False)
print('measurement_tmc.csv generated!')
# -
measurement_tmc
# ## step 3 mapping osm network to tmc network
link_base = pd.read_csv('link.csv', low_memory=False)
link_base = link_base[link_base['link_type_name'].isin(['motorway','trunk'])]
# link_base = link_base[link_base['link_type_name'].isin(['motorway','trunk','primary','secondary'])]
link_base = link_base.reset_index()
link_base = link_base.drop(['index'], 1)
link_base
def LLs2Dist(lon1, lat1, lon2, lat2): #WGS84 transfer coordinate system to distance(meter) #xy #credit to xtHuang0927
R = 6371
dLat = (lat2 - lat1) * math.pi / 180.0
dLon = (lon2 - lon1) * math.pi / 180.0
a = math.sin(dLat / 2) * math.sin(dLat/2) + math.cos(lat1 * math.pi / 180.0) * math.cos(lat2 * math.pi / 180.0) * math.sin(dLon/2) * math.sin(dLon/2)
c = 2 * math.atan2(math.sqrt(a), math.sqrt(1-a))
distance = R * c * 1000
return distance
def getDegree(latA, lonA, latB, lonB):
"""
Args:
point p1(latA, lonA)
point p2(latB, lonB)
Returns:
bearing between the two GPS points,
default: the basis of heading direction is north
"""
radLatA = math.radians(latA)
radLonA = math.radians(lonA)
radLatB = math.radians(latB)
radLonB = math.radians(lonB)
dLon = radLonB - radLonA
y = math.sin(dLon) * math.cos(radLatB)
x = math.cos(radLatA) * math.sin(radLatB) - math.sin(radLatA) * math.cos(radLatB) * math.cos(dLon)
brng = np.degrees(math.atan2(y, x))
brng = (brng + 360) % 360
# brng = 360 - brng
return brng
link_tmc
link_base
# +
multiline_string_tmc_list = []
multiline_string_tmc_list_sub = []
for j in link_tmc.index:
link_tmc_geometry_list = link_tmc.loc[j,'geometry'][12:-1].split(",")
for link_tmc_geometry in link_tmc_geometry_list:
multiline_string_tmc_list_sub.append((float(link_tmc_geometry.split(" ")[0]),float(link_tmc_geometry.split(" ")[1])))
multiline_string_tmc_list_sub = tuple(multiline_string_tmc_list_sub)
multiline_string_tmc_list.append(multiline_string_tmc_list_sub)
multiline_string_tmc_list_sub = []
multiline_string_base_list = []
multiline_string_base_list_sub = []
for j in link_base.index:
link_base_geometry_list = link_base.loc[j,'geometry'][12:-1].split(", ")
for link_base_geometry in link_base_geometry_list:
multiline_string_base_list_sub.append((float(link_base_geometry.split(" ")[0]),float(link_base_geometry.split(" ")[1])))
multiline_string_base_list_sub = tuple(multiline_string_base_list_sub)
multiline_string_base_list.append(multiline_string_base_list_sub)
multiline_string_base_list_sub = []
from shapely.geometry import MultiLineString
line_tmc = MultiLineString(multiline_string_tmc_list)
line_base = MultiLineString(multiline_string_base_list)
#bbox = left,bottom,right,top = min Longitude , min Latitude , max Longitude , max Latitude
if (line_tmc.bounds[0] > line_base.bounds[0]) & (line_tmc.bounds[1] > line_base.bounds[1]) \
& (line_tmc.bounds[2] < line_base.bounds[2]) & (line_tmc.bounds[3] < line_base.bounds[3]):
matching_tmc2gmns_dict = {}
k = 0
p = 1
for j in link_tmc.index:
lon_tmc_list = []
lat_tmc_list = []
link_tmc_geometry_list = link_tmc.loc[j,'geometry'][12:-1].split(",")
for link_tmc_geometry in link_tmc_geometry_list:
lon_tmc_list.append(float(link_tmc_geometry.split(" ")[0]))
lat_tmc_list.append(float(link_tmc_geometry.split(" ")[1]))
center_tmc_lon = np.mean(lon_tmc_list)
center_tmc_lat = np.mean(lat_tmc_list)
tmc_lon_1 = lon_tmc_list[0]
tmc_lon_2 = lon_tmc_list[-1]
tmc_lat_1 = lat_tmc_list[0]
tmc_lat_2 = lat_tmc_list[-1]
if getDegree(tmc_lat_1,tmc_lon_1,tmc_lat_2,tmc_lon_2)>180:
angle_tmc = getDegree(tmc_lat_2,tmc_lon_2,tmc_lat_1,tmc_lon_1)
else:
angle_tmc = getDegree(tmc_lat_1,tmc_lon_1,tmc_lat_2,tmc_lon_2)
distance_list = []
angle_list = []
for i in range(len(link_base)):
lon_list = []
lat_list = []
link_geometry_list = link_base.loc[i,'geometry'][12:-1].split(", ")
for link_geometry in link_geometry_list:
lon_list.append(float(link_geometry.split(" ")[0]))
lat_list.append(float(link_geometry.split(" ")[1]))
'''distance'''
center_lon = np.mean(lon_list)
center_lat = np.mean(lat_list)
distance_list.append(LLs2Dist(center_lon, center_lat, center_tmc_lon, center_tmc_lat))
'''angle '''
base_lon_1 = lon_list[0]
base_lon_2 = lon_list[-1]
base_lat_1 = lat_list[0]
base_lat_2 = lat_list[-1]
if getDegree(base_lat_1,base_lon_1,base_lat_2,base_lon_2)>180:
angle_base = getDegree(base_lat_2,base_lon_2,base_lat_1,base_lon_1)
else:
angle_base = getDegree(base_lat_1,base_lon_1,base_lat_2,base_lon_2)
if abs(angle_tmc - angle_base) >= 90:
relative_angle = 180 - abs(angle_tmc - angle_base)
else:
relative_angle = abs(angle_tmc - angle_base)
angle_list.append(relative_angle)
small_angle_list = [i for i, value in enumerate(angle_list) if value < 45]
df_distance = pd.DataFrame({'distance':distance_list})
nearest_index = df_distance.loc[small_angle_list].idxmin().values[0]
matching_tmc2gmns_dict[k] = {'name_tmc':link_tmc.loc[j]['name'],\
'corridor_id_tmc':link_tmc.loc[j]['corridor_id'],\
'link_id_tmc':link_tmc.loc[[j]].index.values[0],\
'from_node_id_tmc':link_tmc.loc[j]['from_node_id'],\
'to_node_id_tmc':link_tmc.loc[j]['to_node_id'],\
'category_id_tmc':link_tmc.index.get_loc(j)+1,\
'geometry_tmc':link_tmc.loc[j]['geometry'],\
'name_base':link_base['name'][nearest_index],\
'link_id_base':link_base['link_id'][nearest_index],\
'from_node_id_base':link_base['from_node_id'][nearest_index],\
'to_node_id_base':link_base['to_node_id'][nearest_index],\
'category_id_base':link_tmc.index.get_loc(j)+1,\
'geometry_base':link_base['geometry'][nearest_index],\
'distance':min(distance_list),\
'geometry_tmc_base':'MULTILINESTRING ('+ link_tmc.loc[j]['geometry'][11:] + \
', ' + link_base['geometry'][nearest_index][11:]+')'}
k += 1
if link_tmc.index.get_loc(j) > p/10 * len(link_tmc):
print(str(p*10)+"%"+' matching completed!')
p = p + 1
matching_tmc2gmns = pd.DataFrame(matching_tmc2gmns_dict).transpose()
matching_tmc2gmns.to_csv('matching_tmc2gmns.csv',index = False)
print('matching_tmc2gmns.csv generated!')
else:
print('base map cannot cover all TMC links, please use larger base map')
# -
matching_tmc2gmns
# ## step 4 generating measurement_osm
# +
link_base = pd.read_csv('link.csv', low_memory=False)
link_base = link_base[link_base['link_type_name'].isin(['motorway','trunk','primary','secondary'])]
link_base = link_base.reset_index()
link_base = link_base.drop(['index'], 1)
'''build measurement_base.csv'''
measurement_base = pd.DataFrame()
measurement_base['link_id'] = None
measurement_base['osm_way_id'] = None
measurement_base['from_node_id'] = None
measurement_base['to_node_id'] = None
measurement_base['lanes'] = None
measurement_base['length'] = None
measurement_base['time_period'] = None
measurement_base['date'] = None
measurement_base['geometry'] = None
measurement_base['volume'] = None
measurement_base['speed'] = None
measurement_base['ip_address'] = None
k=0
p=1
measurement_base_dict = {}
for i in matching_tmc2gmns.index:
try:
measurement_tmc_selected = measurement_tmc[measurement_tmc['link_id_tmc'] == matching_tmc2gmns['link_id_tmc'][i]]
link_base_selected = link_base[link_base['link_id'] == matching_tmc2gmns['link_id_base'][i]]
for j in measurement_tmc_selected.index:
measurement_base_dict[k] = {'link_id': link_base_selected['link_id'].values[0],\
'osm_way_id':link_base_selected['osm_way_id'].values[0],\
'from_node_id': link_base_selected['from_node_id'].values[0],\
'to_node_id': link_base_selected['to_node_id'].values[0],\
'lanes': link_base_selected['lanes'].values[0], \
'length': link_base_selected['length'].values[0], \
'link_type_name': link_base_selected['link_type_name'].values[0], \
'time_period': measurement_tmc_selected['time_period'][j],\
'date': measurement_tmc_selected['date'][j],\
'geometry': link_base_selected['geometry'].values[0],\
'volume': None,\
'speed': measurement_tmc_selected['speed'][j],\
'ip_address': 'www.openstreetmap.org/?way=' + str(link_base_selected['osm_way_id'].values[0])}
k += 1
except:
measurement_base_dict[k] = {'link_id': link_base_selected['link_id'].values[0],\
'osm_way_id':link_base_selected['osm_way_id'].values[0],\
'from_node_id': link_base_selected['from_node_id'].values[0],\
'to_node_id': link_base_selected['to_node_id'].values[0],\
'lanes': link_base_selected['lanes'].values[0], \
'length': link_base_selected['length'].values[0], \
'link_type_name': link_base_selected['link_type_name'].values[0], \
'time_period':None,\
'date': None,\
'geometry': link_base_selected['geometry'].values[0],\
'volume': None,\
'speed': None,\
'ip_address': 'www.openstreetmap.org/?way=' + str(link_base_selected['osm_way_id'].values[0])}
k += 1
if i+1 > p/10 * len(matching_tmc2gmns.index):
print(str(p*10)+"%"+' measurement_base completed!')
p = p + 1
measurement_base = pd.DataFrame(measurement_base_dict).transpose()
measurement_base.to_csv('measurement_base.csv',index = False)
print('measurement_base.csv generated!')
# -
measurement_base
print( __name__)
print(os.path.dirname(os.path.realpath('__file__')))
print(os.path.realpath('__file__'))
print(os.getcwd())
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import shapely.geometry as geom
import numpy as np
import pandas as pd
import geopandas as gpd
lines = gpd.GeoSeries(
[geom.LineString(((1.4, 3), (0, 0))),
geom.LineString(((1.1, 2.), (0.1, 0.4))),
geom.LineString(((-0.1, 3.), (1, 2.)))])
# 10 points
n = 10
points = gpd.GeoSeries([geom.Point(x, y) for x, y in np.random.uniform(0, 3, (n, 2))])
# Put the points in a dataframe, with some other random column
df_points = gpd.GeoDataFrame(np.array([points, np.random.randn(n)]).T)
df_points.columns = ['Geometry', 'Property1']
points.plot()
lines.plot()
# -
min_dist = np.empty(n)
for i, point in enumerate(points):
min_dist[i] = np.min([point.distance(line) for line in lines])
df_points['min_dist_to_lines'] = min_dist
df_points.head(3)
# +
from shapely.geometry import Point, LineString
import geopandas
line1 = LineString([
Point(0, 0),
Point(0, 1),
Point(1, 1),
Point(1, 2),
Point(3, 3),
Point(5, 6),
])
line2 = LineString([
Point(5, 3),
Point(5, 5),
Point(9, 5),
Point(10, 7),
Point(11, 8),
Point(12, 12),
])
line3 = LineString([
Point(9, 10),
Point(10, 14),
Point(11, 12),
Point(12, 15),
])
print(line1.distance(line2))
print(LineString([Point(0, 0),Point(20, 0),]).distance(Point(9, 10)))
# -
gs = geopandas.GeoSeries([line1, line2, line3])
gs.distance(gs.shift()) #the distances from line1 to line2, and line2 to line3:
# +
#for a line-segment AB, and a point C (all points are tuples/lists)
pdist=lambda A,B:((A[0]-B[0])**2+(A[1]-B[1])**2)**(1/2)
def dist(A,B,C):
c=pdist(A,B)
rat=((C[0]-A[0])*(B[0]-A[0])+(C[1]-A[1])*(B[1]-A[1]))/c/c
if rat<0 or rat>1:
return None,None
a=pdist(B,C)
b=pdist(A,C)
s=(a+b+c)/2
alt=2*(s*(s-a)*(s-b)*(s-c))**(1/2)/c
return alt,rat
dist((0, 0),(20, 0),(10, 10))
# -
| tmc2gmns_final_restruct_v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/kundajelab/dragonn/blob/master/paper_supplement/PrimerTutorial%205%20-%20Functional%20variant%20characterization%20for%20non-coding%20SNPs%20within%20the%20SPI1%20motif.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="XIu-8nkCV4zm"
# # How to train your DragoNN tutorial 5:
# ## Functional variant characterization for non-coding SNPs within the SPI1 motif
#
# This tutorial is a supplement to the DragoNN manuscript.
#
# This tutorial will take 2 - 3 hours if executed on a GPU.
#
# ## Outline<a name='outline'>
# <ol>
# <li><a href=#1>Input data: SPI1 ChiP-seq and experimental bQTL data</a></li>
# <li><a href=#2>Genomewide classification and regression labels for SPI1 TF ChiPseq</a></li>
# <li><a href=#3>Optional: Download pre-generated models and test-set predictions</a></li>
# <li><a href=#4>Genome-wide classification for SPI1</a></li>
# <li><a href=#5>Genome-wide regression for SPI1</a></li>
# <li><a href=#6>Genome-wide interpretation of true positive predictions in SPI1, with DeepLIFT</a></li>
# <li><a href=#7>Recovering bQTL effect sizes: Classification vs Regression</a></li>
# <li><a href=#8>Model-predicted SNP effect sizes vs bQTL effect sizes</a></li>
# <li><a href=#a>Kat's architecture: Classification Model</a></li>
# <li><a href=#b>Kat's architecture: Regression Model</a></li>
# <li><a href=#9>Conclusions</a></li>
# <li><a href=#10>Save tutorial outputs</a></li>
# </ol>
# Github issues on the [dragonn repository](https://github.com/kundajelab/dragonn) with feedback, questions, and discussion are always welcome.
#
# + colab={} colab_type="code" id="72XlYRZBluGr"
# If you don't have bedtools installed in your environment (i.e. Google Colab), uncomment and run the command below
# #!apt-get install bedtools
# #!pip install pybedtools
# + colab={} colab_type="code" id="FmftiCCDV4zo"
#uncomment the lines below if you are running this tutorial from Google Colab
# #!pip install dragonn>=0.2.2
# + colab={} colab_type="code" id="fyLzeiF5V4zq"
# Making sure our results are reproducible
from numpy.random import seed
seed(1234)
from tensorflow import set_random_seed
set_random_seed(1234)
# + colab={} colab_type="code" id="8M6gdfuJV4zu"
#load dragonn tutorial utilities
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
from dragonn.tutorial_utils import *
# + [markdown] colab_type="text" id="djDLAi21V4zy"
# ## Input data <a name='1'>
# <a href=#outline>Home</a>
#
# This tutorial uses the same in vivo SPI1 transcription factor CHiP-seq dataset that was used in [Tutorial 4](https://colab.research.google.com/github/kundajelab/dragonn/blob/keras_2.2_tensorflow_1.6_purekeras/paper_supplement/PrimerTutorial%204%20-%20Interpreting%20predictive%20sequence%20features%20in%20in-vivo%20TF%20binding%20events.ipynb). Our goal is to compare predicted variant effect sizes from classification and regression models against experimental bQTL data. The bQTL data in this way serves as a "gold-standard" validation that in silico mutagenesis on the deep learning inputs leads to correct variant effect size prediction. We will use bQTL data that has been intersected with SPI1 CISBP genome motif annotations.
# + colab={"base_uri": "https://localhost:8080/", "height": 413} colab_type="code" id="O707uf21V4zy" outputId="f5ca2dbc-9594-4a62-aa67-97190945d622"
# SPI1, optimal IDR thresholded peaks, Myers lab, hg19
# https://www.encodeproject.org/experiments/ENCSR000BGQ/
# !wget -O SPI1.narrowPeak.gz http://mitra.stanford.edu/kundaje/projects/dragonn/dragonn_gm12878_pipeline/spi1_ENCSR000BGQ/cromwell-executions/chip/bb0c3c5a-3889-43fe-a218-05851cecc74a/call-reproducibility_idr/execution/optimal_peak.regionPeak.gz
#Fold change bigWig track for the SPI1 dataset:
# !wget -O SPI1.pooled.fc.bigWig http://mitra.stanford.edu/kundaje/projects/dragonn/dragonn_gm12878_pipeline/spi1_ENCSR000BGQ/cromwell-executions/chip/bb0c3c5a-3889-43fe-a218-05851cecc74a/call-macs2_pooled/execution/ENCFF000OBU.Rep1.merged.nodup.pooled_x_ENCFF000OCW.Control.Rep1.merged.nodup.fc.signal.bigwig
## Download the hg19 chromsizes file (We only use chroms 1 -22, X, Y for training)
# !wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.chrom.sizes
## Download the hg19 fasta reference genome (and corresponding .fai index)
# !wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.genome.fa.gz
# !wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.genome.fa.gz.fai
# !wget http://mitra.stanford.edu/kundaje/projects/dragonn/hg19.genome.fa.gz.gzi
# + colab={"base_uri": "https://localhost:8080/", "height": 215} colab_type="code" id="-YwnqCV-V4z2" outputId="85791b16-8647-45ff-cd8b-cf281d618350"
# Download bQTL experimental data for SPI1 loci
# !wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.bQTLs.txt.gz
# + [markdown] colab_type="text" id="sp9mi-6_V4z4"
# ## Generating genome-wide classification and regression labels <a name='2'>
# <a href=#outline>Home</a>
# + [markdown] colab_type="text" id="Zmt5OJP_V4z5"
# We will use the *genomewide_labels* function from the [seqdataloader](https://github.com/kundajelab/seqdataloader) package to generate positive and negative labels for the TF-ChIPseq peaks across the genome. We will treat each sample as a task for the model and compare the performance of the model on SPI1 task in the single-tasked and multi-tasked setting.
# + colab={} colab_type="code" id="SLGpH2rOV4z6"
from seqdataloader import *
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="u2wpRugxV4z7" outputId="c695d444-b6d0-408b-f294-6afcdc6b1033"
## seqdataloader accepts an input file, which we call SPI1.tasks.tsv, with task names in column 1, corresponding
## peak files in column 2, and the signal track in column 3. In this tutorial, the task file will have a single task entry for the SPI1 TF CHiP-seq
with open("SPI1.task.tsv",'w') as f:
f.write("SPI1\tSPI1.narrowPeak.gz\tSPI1.pooled.fc.bigWig\n")
f.close()
# !cat SPI1.task.tsv
# + [markdown] colab_type="text" id="-pqz2oVGV4z_"
# With the parameter configuration below, seqdataloader splits the genome into 1kb regions, with a stride of 50. Each 1kb region is centered at a 200 bp bin, with a left flank of 400 bases and a right flank of 400 bases.
#
# * In the classification case, each 200 bp bin is labeled as positive if a narrowPeak summit overlaps with it. The bin is labeled negative if there is no overlap with the narrowPeak.
# * In the regression case, the asinh(mean coverage) in the 200 bp bin is computed.
#
# + [markdown] colab_type="text" id="e--f8QWuV4z_"
# **Note**: The label generation may take 10 - 15 minutes to complete. If you prefer not to wait, you can download the
# pre-generated classification and regression labels for the training, validation, and test sets by uncommenting the code below:
# + colab={"base_uri": "https://localhost:8080/", "height": 1205} colab_type="code" id="LMG9IzPnV40A" outputId="aca1e05d-dc62-416e-b22f-b68a53aaf3f7"
## Classification labels
# ! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.train.classification.hdf5
# ! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.valid.classification.hdf5
# ! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.test.classification.hdf5
## Regression labels
# ! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.train.regression.hdf5
# ! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.valid.regression.hdf5
# ! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.test.regression.hdf5
# + [markdown] colab_type="text" id="_lleRdAaV40B"
# If you prefer to generate the labels from scratch, execute the two code cell below:
# + colab={} colab_type="code" id="erh3B4hIV40D"
# Generate genome-wide classification labels
#1) Training set: all chromosomes with the exception of 1,2, and 19 in our training set. Also, the dataset does not
# include chromosome Y, so we exclude it as well.
train_set_params={
'task_list':"SPI1.task.tsv",
'outf':"SPI1.train.classification.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_exclude':['chr1','chr2','chr19','chrY'],
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':4,
'subthreads':4,
'allow_ambiguous':False,
'labeling_approach':'peak_summit_in_bin_classification'
}
genomewide_labels(train_set_params)
#2) Validation set: Chromosome 1
valid_set_params={'task_list':"SPI1.task.tsv",
'outf':"SPI1.valid.classification.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_keep':'chr1',
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':1,
'subthreads':4,
'allow_ambiguous':False,
'labeling_approach':'peak_summit_in_bin_classification'
}
genomewide_labels(valid_set_params)
#3) Test set: Chromosomes 2, 19
test_set_params={
'task_list':"SPI1.task.tsv",
'outf':"SPI1.test.classification.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_keep':['chr2','chr19'],
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':2,
'subthreads':4,
'allow_ambiguous':False,
'labeling_approach':'peak_summit_in_bin_classification'
}
genomewide_labels(test_set_params)
# + colab={} colab_type="code" id="PBqbysuAV40G"
# Generate regression labels genome-wide
#1) Training set: all chromosomes with the exception of 1,2, and 19 in our training set
train_set_params={
'task_list':"SPI1.task.tsv",
'outf':"SPI1.train.regression.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_exclude':['chr1','chr2','chr19','chrY'],
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':4,
'subthreads':4,
'allow_ambiguous':False,
'labeling_approach':'all_genome_bins_regression'
}
genomewide_labels(train_set_params)
#2) Validation set: Chromosome 1
valid_set_params={'task_list':"SPI1.task.tsv",
'outf':"SPI1.valid.regression.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_keep':'chr1',
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':1,
'subthreads':4,
'allow_ambiguous':False,
'labeling_approach':'all_genome_bins_regression'
}
genomewide_labels(valid_set_params)
#3) Test set: Chromosomes 2, 19
test_set_params={
'task_list':"SPI1.task.tsv",
'outf':"SPI1.test.regression.hdf5",
'output_type':'hdf5',
'chrom_sizes':'hg19.chrom.sizes',
'chroms_to_keep':['chr2','chr19'],
'bin_stride':50,
'left_flank':400,
'right_flank':400,
'bin_size':200,
'threads':2,
'subthreads':4,
'allow_ambiguous':False,
'labeling_approach':'all_genome_bins_regression'
}
genomewide_labels(test_set_params)
# + [markdown] colab_type="text" id="x1m9HsgXV40J"
# Let's examine the files that were generated:
# + colab={"base_uri": "https://localhost:8080/", "height": 390} colab_type="code" id="Q0SgKnZtV40J" outputId="5e5501e6-61e3-44e1-f6ca-94e70f3ff63f"
#The code generates bed file outputs with a label of 1 or 0 for each 1kb
# genome bin for each task. Note that the bins are shifted with a stride of 50.
pd.read_hdf("SPI1.train.classification.hdf5",start=1000000,stop=1000010)
# + colab={"base_uri": "https://localhost:8080/", "height": 390} colab_type="code" id="weH35tmhV40N" outputId="a8aea293-f560-400f-cb2f-2d0085625934"
pd.read_hdf("SPI1.train.regression.hdf5",start=1000000,stop=1000010)
# + [markdown] colab_type="text" id="FKHBBFpRV40Q"
# ## Optional: Download pre-generated models and test-set predictions <a name='3'>
# <a href=#outline>Home</a>
#
# Next, we will train classification and regression models to predict TF CHiP-seq peaks for SPI1. If you want to skip straight to model interpretation and bQTL analysis, you can download the pre-trained models by uncommenting the
# block of code below.
# + colab={} colab_type="code" id="CqyIROINV40R"
from keras.models import load_model
## Download classification model
# #! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.classification.model.hdf5
spi1_classification_model=load_model("SPI1.kat.classification.model.hdf5")
## Download regression model
# #! wget http://mitra.stanford.edu/kundaje/projects/dragonn/SPI1.regression.model.hdf5
spi1_regression_model=load_model("SPI1.kat.regression.model.hdf5")
## Get test set classification model and regression model predictions
#import h5py
#test_set_predictions=h5py.File("SPI1.test.predictions.hdf5")
#spi1_test_classification_predictions=test_set_predictions['classification'].value
#spi1_test_regression_predictions=test_set_predictions['regression'].value
# + [markdown] colab_type="text" id="FZcwz5AmV40U"
# ## Genome-wide classification model <a name='4'>
# <a href=#outline>Home</a>
#
# + colab={} colab_type="code" id="ZeBBukYGV40V"
#To prepare for model training, we import the necessary functions and submodules from keras
from keras.models import Sequential
from keras.layers.core import Dropout, Reshape, Dense, Activation, Flatten
from keras.layers.convolutional import Conv2D, MaxPooling2D
from keras.optimizers import Adadelta, SGD, RMSprop;
import keras.losses;
from keras.constraints import maxnorm;
from keras.layers.normalization import BatchNormalization
from keras.regularizers import l1, l2
from keras.callbacks import EarlyStopping, History
from keras import backend as K
K.set_image_data_format('channels_last')
# + colab={} colab_type="code" id="CXtJhYf2V40Z"
from concise.metrics import tpr, tnr, fpr, fnr, precision, f1
def initialize_classification_model(ntasks=1):
#Define the model architecture in keras (regularized, 3-layer convolution model followed by 1 dense layer)
model=Sequential()
model.add(Conv2D(filters=15,kernel_size=(1,10),input_shape=(1,1000,4)))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(MaxPooling2D(pool_size=(1,35)))
model.add(Conv2D(filters=15,kernel_size=(1,10)))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Conv2D(filters=15,kernel_size=(1,10)))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(ntasks))
model.add(Activation("sigmoid"))
##compile the model, specifying the Adam optimizer, and binary cross-entropy loss.
model.compile(optimizer='adam',loss='binary_crossentropy',
metrics=[tpr,
tnr,
fpr,
fnr,
precision,
f1])
return model
# + [markdown] colab_type="text" id="XY-g6ik9V40c"
# We create generators for the training and validation data:
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="5PrSMLLiV40d" outputId="d7ae2438-6e8e-4fc0-c1b3-6bde39667604"
#create the generators, upsample positives to ensure they constitute 30% of each batch
from dragonn.generators import *
spi1_train_classification_gen=DataGenerator("SPI1.train.classification.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3, batch_size=256)
spi1_valid_classification_gen=DataGenerator("SPI1.valid.classification.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3, batch_size=256)
# + colab={} colab_type="code" id="NLRfBhebV40e"
#Train the SPI1 classification model
spi1_classification_model=initialize_classification_model()
## use the keras fit_generator function to train the model with early stopping after 3 epochs
history_classification=spi1_classification_model.fit_generator(spi1_train_classification_gen,
validation_data=spi1_valid_classification_gen,
steps_per_epoch=10000,
validation_steps=5000,
epochs=150,
verbose=1,
use_multiprocessing=True,
workers=40,
max_queue_size=100,
callbacks=[EarlyStopping(patience=3,restore_best_weights=True),History()])
# + colab={} colab_type="code" id="DN2bFz_aV40h" outputId="323bc266-fc71-4e30-e277-b2db3c3e9a0a"
## Plot the learning curves for SPI1
from dragonn.tutorial_utils import plot_learning_curve
plot_learning_curve(history_classification)
# + [markdown] colab_type="text" id="YoP336y_V40k"
# We now measure how well the model performed by calculating performance metrics on the test splits across the whole genome.
# + colab={} colab_type="code" id="ZF6VNZH3V40k" outputId="101ca0a8-db49-44cc-c133-e799b070e1e0"
from dragonn.generators import *
spi1_test_classification_gen=DataGenerator("SPI1.test.classification.hdf5",
"hg19.genome.fa.gz",
upsample=False,
add_revcomp=False,
batch_size=1000,
tasks=['SPI1'])
spi1_test_classification_predictions=spi1_classification_model.predict_generator(spi1_test_classification_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
spi1_test_classification_truth=spi1_test_classification_gen.data
# + colab={} colab_type="code" id="1QGwsaLsV40n" outputId="829e46ff-12f4-42c9-c18c-2d9c66a22e97"
spi1_test_classification_predictions.shape
# + colab={} colab_type="code" id="9LAq8j_MV40r" outputId="d1b31200-f1a6-493b-ad99-6c0c62b65645"
spi1_test_classification_truth.shape
# + colab={} colab_type="code" id="X83Mt6VXV40t" outputId="b7b4fe79-812d-463a-f2f1-611122a6fa0e"
## Generate a ClassificationResult object to print performance metrics on held-out test set
from dragonn.metrics import ClassificationResult
print(ClassificationResult(spi1_test_classification_truth.values.astype(bool),spi1_test_classification_predictions))
# + colab={} colab_type="code" id="jBvkHg7zV40y"
#save the models
spi1_classification_model.save("SPI1.classification.model.hdf5")
# -
# + [markdown] colab_type="text" id="b0sibNElV403"
# ## Genome-wide regression model <a name='5'>
# <a href=#outline>Home</a>
# + colab={} colab_type="code" id="nM0-1aa9V404"
def initialize_regression_model(ntasks=1):
#Define the model architecture in keras (regularized, 3-layer convolution model followed by 1 dense layer)
model=Sequential()
model.add(Conv2D(filters=15,kernel_size=(1,10),input_shape=(1,1000,4)))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(MaxPooling2D(pool_size=(1,35)))
model.add(Conv2D(filters=10,kernel_size=(1,10)))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Conv2D(filters=5,kernel_size=(1,10)))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(ntasks))
##compile the model, specifying the Adam optimizer, and binary cross-entropy loss.
model.compile(optimizer='adam',loss='mse')
return model
# + colab={} colab_type="code" id="zO-uKK8GV407" outputId="04c1a178-b8ee-4b14-c944-328c17ecec9c"
#we want to determine a threshold for upsampling the non-zero bins in a given batch
# extract 5 million datapoints from the training data and observe the distribution of non-zero signal values
sample=pd.read_hdf("SPI1.train.regression.hdf5",start=0,stop=5000000)
nonzero_sample=sample[sample.max(axis=1)>0]
print(nonzero_sample.shape)
nonzero_sample.hist(bins=100)
# + [markdown] colab_type="text" id="F5Raqyz3V40_"
# This suggests that 0.1 is a reasonable threshold for upsampling signal bins in regression
# + colab={} colab_type="code" id="JSjGIlYTV41B"
#create the generators, no upsampling of positives is used for regression.
from dragonn.generators import *
spi1_train_regression_gen=DataGenerator("SPI1.train.regression.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3,upsample_thresh=0.01)
spi1_valid_regression_gen=DataGenerator("SPI1.valid.regression.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3,upsample_thresh=0.01)
# + colab={} colab_type="code" id="P3pSs4TYV41E" outputId="bb50b654-2336-4b6d-d16c-a8a15672f379"
#Train the SPI1 regression model
spi1_regression_model=initialize_regression_model()
## use the keras fit_generator function to train the model with early stopping after 3 epochs
history_regression=spi1_regression_model.fit_generator(spi1_train_regression_gen,
validation_data=spi1_valid_regression_gen,
steps_per_epoch=10000,
validation_steps=5000,
epochs=150,
verbose=1,
use_multiprocessing=True,
workers=40,
max_queue_size=100,
callbacks=[EarlyStopping(patience=3,restore_best_weights=True),History()])
# + colab={} colab_type="code" id="EFxneNJqV41H" outputId="472b64dc-f85d-49e6-8bd0-4d023374e04a"
plot_learning_curve(history_regression)
# + colab={} colab_type="code" id="_o3I4gN7V41K" outputId="da95db30-a6ad-4043-86c6-ed587a44277b"
from dragonn.generators import *
spi1_test_regression_gen=DataGenerator("SPI1.test.regression.hdf5",
"hg19.genome.fa.gz",
upsample=False,
add_revcomp=False,
batch_size=1000,
tasks=['SPI1'])
spi1_test_regression_predictions=spi1_regression_model.predict_generator(spi1_test_regression_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
spi1_test_regression_truth=spi1_test_regression_gen.data
# + colab={} colab_type="code" id="QMbEZo8XGd2m"
## find the indices of the non-zero coverage bins
nonzero_bins=spi1_test_regression_truth.max(axis=1)>0
# + colab={} colab_type="code" id="-byNK4qMV41N" outputId="aa18b7e6-6ee2-439d-ca0b-d68d2a7aa8a1"
#Calculate spearman and pearson correlation between truth labels and predictions
from scipy.stats import pearsonr, spearmanr
corr_pearson=pearsonr(spi1_test_regression_truth,spi1_test_regression_predictions)
corr_spearman=spearmanr(spi1_test_regression_truth,spi1_test_regression_predictions)
print("Pearson correlation on test set:"+str(corr_pearson))
print("Spearman correlation on test set:"+str(corr_spearman))
# + colab={} colab_type="code" id="YipLiXRFGd2t" outputId="f7e2d334-30b4-4661-d8bb-23832c8c5763"
# Calculate the spearman and pearson correlation, restricted to non-zero bins
corr_pearson_nonzero_bins=pearsonr(spi1_test_regression_truth[nonzero_bins],spi1_test_regression_predictions[nonzero_bins])
corr_spearman_nonzero_bins=spearmanr(spi1_test_regression_truth[nonzero_bins],spi1_test_regression_predictions[nonzero_bins])
print("Pearson correlation on test set:"+str(corr_pearson_nonzero_bins))
print("Spearman correlation on test set:"+str(corr_spearman_nonzero_bins))
# + colab={} colab_type="code" id="kVEfZ1d0V41O"
#There is some overfitting, let's save this model and see if we can do better
spi1_regression_model.save("SPI1.regression.model.hdf5")
# + colab={} colab_type="code" id="UGW2FQrgGd2y" outputId="e3069d0a-65d3-4c8c-be0d-eef77ed83581"
spi1_test_regression_truth.values[0:10].squeeze()
# + colab={} colab_type="code" id="SKa34qjbGd21"
test_df=pd.DataFrame({"Observed":list(spi1_test_regression_truth.values.squeeze()),
"Predicted":list(spi1_test_regression_predictions.squeeze())})
# + colab={} colab_type="code" id="L1tUvhU5Gd27"
test_df_nonzero=pd.DataFrame({"Observed":list(spi1_test_regression_truth[nonzero_bins].values.squeeze()),
"Predicted":list(spi1_test_regression_predictions[nonzero_bins].squeeze())})
# + colab={} colab_type="code" id="Xqvik10lGd29" outputId="4d35b751-13ce-4555-f798-916765d4e981"
import plotnine
from plotnine import *
print((ggplot(test_df,aes(x="Observed",y="Predicted"))
+geom_bin2d(bins=100)
+theme_bw()
+xlab("Observed asinh(mean coverage in FC bigWig")
+ylab("Model prediction")
+ggtitle("SPI1 regression model test set prediction")))
print((ggplot(test_df_nonzero,aes(x="Observed",y="Predicted"))
+geom_bin2d(bins=100)
+theme_bw()
+xlab("Observed asinh(mean coverage in FC bigWig")
+ylab("Model prediction")
+ggtitle("SPI1 regression model test set prediction: bins with nonzero coverage")))
# + colab={} colab_type="code" id="TOBvin88Gd3A" outputId="5fc16ecd-16d2-47f0-a39f-e91244e4d9cb"
# Plot observed vs predicted regression values
plt.scatter(spi1_test_regression_truth, spi1_test_regression_predictions, alpha=0.01)
plt.xlabel("Observed asinh(mean coverage in FC bigWig)")
plt.ylabel("Model prediction")
plt.title("SPI1 regression model test set prediction")
plt.show()
# + colab={} colab_type="code" id="WCgR4DmGGd3C" outputId="82246537-1073-48b1-81f7-1dc8f2a4b178"
# Plot observed vs predicted regression values for the nonzero bins
plt.scatter(spi1_test_regression_truth, spi1_test_regression_predictions, alpha=0.01)
plt.xlabel("Observed asinh(mean coverage in FC bigWig) for bins ")
plt.ylabel("Model prediction")
plt.title("SPI1 regression model test set prediction: bins with nonzero coverage")
plt.show()
# + [markdown] colab_type="text" id="48jGNSBtV41R"
# ## Genome-wide interpretation of true positive predictions in SPI1, with DeepLIFT <a name='6'>
# <a href=#outline>Home</a>
#
# ### Classification Model
# + colab={} colab_type="code" id="RWumi0mQV41S"
#get the true positive predictions with a threshold of 0.9 (i.e. high confidence true positive predictions)
spi1_test_classification_truth_bool=spi1_test_classification_truth.values.astype(bool)
true_pos_spi1=spi1_test_classification_truth[spi1_test_classification_truth_bool*spi1_test_classification_predictions >0.9]
true_pos_spi1.head
# + colab={} colab_type="code" id="ecJa0a2HV41U" outputId="bb860e8e-f083-41e3-ed79-e66bb8aec2ac"
true_pos_spi1.shape
# + colab={} colab_type="code" id="UH5WvDmXV41W" outputId="c549e164-1087-4f2b-93bd-7f0677075090"
from dragonn.utils import one_hot_from_bed
deep_lift_input_spi1=one_hot_from_bed([i for i in true_pos_spi1.index],"hg19.genome.fa.gz")
deep_lift_input_spi1.shape
# + colab={} colab_type="code" id="41dInIg-V41Y"
from dragonn.tutorial_utils import deeplift
# + colab={} colab_type="code" id="hViDtFpGV41a"
deep_lift_scores_spi1=deeplift(spi1_classification_model,deep_lift_input_spi1)
# + colab={} colab_type="code" id="mFkS3UwpV41c" outputId="953cdbc2-d7c0-4a17-d6d1-227acb47c325"
deep_lift_scores_spi1.shape
# + [markdown] colab_type="text" id="skCZQ8j0V41e"
# Let's plot a few of the DeepLIFT tracks and see if the model successfully learned SPI1:
# + colab={} colab_type="code" id="Srt-dPa_V41e"
from dragonn.tutorial_utils import plot_seq_importance
# + colab={} colab_type="code" id="nhcSoq48V41h" outputId="fd5de43b-d65f-4abb-d998-7115c5ad1c1e"
plot_seq_importance(deep_lift_scores_spi1[0],deep_lift_input_spi1[0])
# + colab={} colab_type="code" id="eyejvCeWV41j" outputId="9985d83c-d05f-4979-f472-bb352cce2f80"
plot_seq_importance(deep_lift_scores_spi1[1],deep_lift_input_spi1[1])
# + colab={} colab_type="code" id="CyuioSW_V41o" outputId="75f83e18-bf46-475d-c43b-22ab8cdc56d9"
plot_seq_importance(deep_lift_scores_spi1[2],deep_lift_input_spi1[2])
# + [markdown] colab_type="text" id="AfkuGJTGV41r"
# Let's zoom in to the center of one sequence so that it is easier to distinguish the motif:
# + colab={} colab_type="code" id="YCr3vtPeV41s" outputId="a04a10b2-ff18-44cb-f918-005872cb3b59"
plot_seq_importance(deep_lift_scores_spi1[2].squeeze()[550:650],deep_lift_input_spi1[2].squeeze()[550:650])
# + [markdown] colab_type="text" id="tiw5YPInV41v"
# If we query the sequence "CACTTCCCCT" in the [TomTom](http://meme-suite.org/tools/tomtom) software from the MEME suite, we find that the motif is a good match for SPIB:
# <img src="https://github.com/kundajelab/dragonn/blob/master/paper_supplement/tutorial_images/SPI1.Tut4.png?raw=1" alt="SPI12TomTom" width="400"/>
#
# + [markdown] colab_type="text" id="kEUxg-r1V41x"
# ### Regression model
# + colab={} colab_type="code" id="bEJudwWXV41y" outputId="f2395f38-be1f-4b76-95a3-91cc85efa471"
#Sanity-check that the model is learning the SPI1 motif by running DeepLIFT on True Positives with high confidence (>0.9)
#get the true positive predictions
true_pos=spi1_test_regression_truth[(spi1_test_regression_truth.values*spi1_test_regression_predictions)>2]
true_pos.shape
# + colab={} colab_type="code" id="z5dpuvzLV413" outputId="c0db98aa-22bf-4e44-f15f-0a75df7c9239"
deep_lift_input=one_hot_from_bed([i for i in true_pos.index],"hg19.genome.fa.gz")
deep_lift_input.shape
# + colab={} colab_type="code" id="D-3kc90ZV416" outputId="514a9b2b-0f73-45ac-fe0c-b695bb182dbb"
help(deeplift)
# + colab={} colab_type="code" id="YeuZFMZqV41-"
deep_lift_scores_spi1=deeplift(spi1_regression_model,deep_lift_input_spi1,target_layer_idx=-1)
# + colab={} colab_type="code" id="8qigmzDOV41-" outputId="1ef6a324-0b04-4000-a658-87c75cb0c50d"
plot_seq_importance(deep_lift_scores_spi1[0],deep_lift_input_spi1[0])
# + colab={} colab_type="code" id="EPHe9I8VV42A" outputId="b7ee3839-2d4e-466b-e2f7-ede00db8a16b"
plot_seq_importance(deep_lift_scores_spi1[1],deep_lift_input_spi1[1])
# + colab={} colab_type="code" id="bogEKZN2V42C" outputId="6465c01d-b842-4217-9766-aca7a14797b2"
plot_seq_importance(deep_lift_scores_spi1[2],deep_lift_input_spi1[2])
# + colab={} colab_type="code" id="Ck63kAsAV42F" outputId="5353b1fb-b881-4515-f931-96f49f196045"
plot_seq_importance(deep_lift_scores_spi1[2].squeeze()[550:650],deep_lift_input_spi1[2].squeeze()[550:650])
# + [markdown] colab_type="text" id="PWqxtR6NV42I"
# The motif learned by the regression model matches the canonical SPI1 motif, though the deepLIFT tracks are noisier compared to those for the classification model.
#
# + [markdown] colab_type="text" id="-PGA_k3RV42J"
# ## Recovering bQTL effect sizes: Classification vs Regression <a name='7'>
# <a href=#outline>Home</a>
# + colab={} colab_type="code" id="GROAoPZDV42J"
from dragonn.generators import *
bqtl_ref_gen=BQTLGenerator("SPI1.bQTLs.txt.gz","hg19.genome.fa.gz","POSTallele")
bqtl_alt_gen=BQTLGenerator("SPI1.bQTLs.txt.gz","hg19.genome.fa.gz","ALTallele")
# + colab={} colab_type="code" id="GJq0Ic_8V42L" outputId="f27e9d3e-87a2-479f-90ca-320bf3066fc0"
bqtl_ref_classification_predictions=spi1_classification_model.predict_generator(bqtl_ref_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
# + colab={} colab_type="code" id="xlPJpmmLV42L" outputId="bf8b8ff1-0268-49af-ce7d-9d8875e2ae3f"
bqtl_alt_classification_predictions=spi1_classification_model.predict_generator(bqtl_alt_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
bqtl_ref_classification_truth=bqtl_ref_gen.data['pvalue']
# + colab={} colab_type="code" id="MGXJaSvPV42N" outputId="32d58541-15ab-4c82-90d5-b3118ed18520"
print(bqtl_ref_classification_predictions.shape)
print(bqtl_alt_classification_predictions.shape)
print(bqtl_ref_classification_truth.shape)
# + colab={} colab_type="code" id="1NpOM03tV42P" outputId="9e58fdff-c60a-463a-fa43-2a6979d96064"
bqtl_ref_regression_predictions=spi1_regression_model.predict_generator(bqtl_ref_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
bqtl_alt_regression_predictions=spi1_regression_model.predict_generator(bqtl_alt_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
# + colab={} colab_type="code" id="hi_Pr6jcV42Q" outputId="62710f23-734d-4ea4-b9a9-e1c28948d597"
plt.scatter(bqtl_ref_classification_predictions, bqtl_alt_classification_predictions, alpha=0.01)
plt.xlabel("Ref")
plt.ylabel("Alt")
plt.title("BQTL Classification Model Predictions")
plt.show()
# + colab={} colab_type="code" id="tg-ZRo1tV42R" outputId="9cb78eaf-ed5f-4fe4-ca02-6f927b07bcca"
plt.scatter(bqtl_ref_regression_predictions, bqtl_alt_regression_predictions, alpha=0.01)
plt.xlabel("Ref")
plt.ylabel("Alt")
plt.title("BQTL Regression Model Predictions")
plt.show()
# + [markdown] colab_type="text" id="yxG07_SzV42T"
# ## Model-predicted SNP effect sizes vs bQTL effect sizes <a name='8'>
# <a href=#outline>Home</a>
# + colab={} colab_type="code" id="YdFUgn60V42T"
logpval=np.log10(bqtl_ref_classification_truth.values)
delta=bqtl_alt_classification_predictions-bqtl_ref_classification_predictions
# + [markdown] colab_type="text" id="WdQsnAM7Gd4B"
# ## Kat's Model Architecture (Classification)<a name='a'>
# <a href=#outline>Home</a>
# + colab={} colab_type="code" id="7yIyOg-AGd4B"
from concise.metrics import tpr, tnr, fpr, fnr, precision, f1
from keras.constraints import max_norm
def initialize_kat_classification_model(ntasks=1):
#Define the model architecture in keras (regularized, 3-layer convolution model followed by 1 dense layer)
model=Sequential()
model.add(Conv2D(filters=50,kernel_size=(1,15),padding="same", kernel_constraint=max_norm(7.0,axis=-1),input_shape=(1,1000,4)))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Conv2D(filters=50,kernel_size=(1,15),padding="same"))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Conv2D(filters=50,kernel_size=(1,13),padding="same"))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(1,40)))
model.add(Flatten())
model.add(Dense(50))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Dense(ntasks))
model.add(Activation("sigmoid"))
##compile the model, specifying the Adam optimizer, and binary cross-entropy loss.
model.compile(optimizer='adam', loss='binary_crossentropy',
metrics=[tpr,
tnr,
fpr,
fnr,
precision,
f1])
return model
# + colab={} colab_type="code" id="EG64R1eNGd4C"
#create the generators, upsample positives to ensure they constitute 30% of each batch
from dragonn.generators import *
spi1_train_classification_gen=DataGenerator("SPI1.train.classification.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3, batch_size=256)
spi1_valid_classification_gen=DataGenerator("SPI1.valid.classification.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3, batch_size=256)
# + colab={} colab_type="code" id="3SeWRHimGd4D" outputId="7516c239-52e8-4c19-dafc-ee32c264808c"
#Train the SPI1 classification model
spi1_kat_classification_model=initialize_kat_classification_model()
## use the keras fit_generator function to train the model with early stopping after 3 epochs
history_kat_classification=spi1_kat_classification_model.fit_generator(spi1_train_classification_gen,
validation_data=spi1_valid_classification_gen,
steps_per_epoch=10000,
validation_steps=5000,
epochs=150,
verbose=1,
use_multiprocessing=True,
workers=40,
max_queue_size=100,
callbacks=[EarlyStopping(patience=3,restore_best_weights=True),History()])
# + colab={} colab_type="code" id="a3D2uhXwGd4E" outputId="b5450d6e-447b-4a4a-e5c0-19890b4949bc"
## Plot the learning curves for SPI1
from dragonn.tutorial_utils import plot_learning_curve
plot_learning_curve(history_kat_classification)
# + colab={} colab_type="code" id="Jq6tLmcTGd4G" outputId="9757df93-a9e1-45c9-85f9-8cc5c51fd926"
from dragonn.generators import *
spi1_test_classification_gen=DataGenerator("SPI1.test.classification.hdf5",
"hg19.genome.fa.gz",
upsample=False,
add_revcomp=False,
batch_size=1000,
tasks=['SPI1'])
spi1_test_classification_predictions=spi1_kat_classification_model.predict_generator(spi1_test_classification_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
spi1_test_classification_truth=spi1_test_classification_gen.data
# + colab={} colab_type="code" id="-fWPY4DoGd4I" outputId="ab351e3b-7f55-465f-8f74-a5e8b7981d18"
## Generate a ClassificationResult object to print performance metrics on held-out test set
from dragonn.metrics import ClassificationResult
print(ClassificationResult(spi1_test_classification_truth.values.astype(bool),spi1_test_classification_predictions))
# + [markdown] colab_type="text" id="dPWznGgeGd4K"
# ## Kat's Model Architecture (Regression)<a name='b'>
# <a href=#outline>Home</a>
# + colab={} colab_type="code" id="2Nt_b4_BGd4K"
def initialize_kat_regression_model(ntasks=1):
#Define the model architecture in keras (regularized, 3-layer convolution model followed by 1 dense layer)
model=Sequential()
model.add(Conv2D(filters=50,kernel_size=(1,15),padding="same", kernel_constraint=max_norm(7.0,axis=-1),input_shape=(1,1000,4)))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Conv2D(filters=50,kernel_size=(1,15),padding="same"))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Conv2D(filters=50,kernel_size=(1,13),padding="same"))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(1,40)))
model.add(Flatten())
model.add(Dense(50))
model.add(BatchNormalization(axis=-1))
model.add(Activation('relu'))
model.add(Dropout(0.2))
model.add(Dense(ntasks))
##compile the model, specifying the Adam optimizer, and binary cross-entropy loss.
model.compile(optimizer='adam',loss='mse')
return model
# + colab={} colab_type="code" id="Wm8P3ABcGd4L"
#create the generators, no upsampling of positives is used for regression.
from dragonn.generators import *
spi1_train_regression_gen=DataGenerator("SPI1.train.regression.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3,upsample_thresh=0.01)
spi1_valid_regression_gen=DataGenerator("SPI1.valid.regression.hdf5","hg19.genome.fa.gz",upsample_ratio=0.3,upsample_thresh=0.01)
# + colab={} colab_type="code" id="O2xnIWHXGd4M" outputId="716c5c00-0b5b-417d-aa33-72cd483c7546"
#Train the SPI1 regression model
spi1_kat_regression_model=initialize_kat_regression_model()
## use the keras fit_generator function to train the model with early stopping after 3 epochs
history_kat_regression=spi1_kat_regression_model.fit_generator(spi1_train_regression_gen,
validation_data=spi1_valid_regression_gen,
steps_per_epoch=10000,
validation_steps=5000,
epochs=150,
verbose=1,
use_multiprocessing=True,
workers=40,
max_queue_size=100,
callbacks=[EarlyStopping(patience=3,restore_best_weights=True),History()])
# + colab={} colab_type="code" id="S2sQWdgtGd4O" outputId="3db4acbc-8de5-4d24-96da-3c4c41eac26a"
plot_learning_curve(history_kat_regression)
# + colab={} colab_type="code" id="JvSYpEFZGd4P" outputId="7fcf68b2-7f0e-4191-ae45-71948da1cd3f"
from dragonn.generators import *
spi1_test_regression_gen=DataGenerator("SPI1.test.regression.hdf5",
"hg19.genome.fa.gz",
upsample=False,
add_revcomp=False,
batch_size=1000,
tasks=['SPI1'])
spi1_test_regression_predictions=spi1_kat_regression_model.predict_generator(spi1_test_regression_gen,
max_queue_size=5000,
workers=40,
use_multiprocessing=True,
verbose=1)
spi1_test_regression_truth=spi1_test_regression_gen.data
# + colab={} colab_type="code" id="2vIyYceoGd4R"
## find the indices of the non-zero coverage bins
nonzero_bins=spi1_test_regression_truth.max(axis=1)>0
# + colab={} colab_type="code" id="pJ8NbtgWGd4T" outputId="8a0c8f6c-512f-419e-dd49-8002895cdeb7"
#Calculate spearman and pearson correlation between truth labels and predictions
from scipy.stats import pearsonr, spearmanr
corr_pearson=pearsonr(spi1_test_regression_truth,spi1_test_regression_predictions)
corr_spearman=spearmanr(spi1_test_regression_truth,spi1_test_regression_predictions)
print("Pearson correlation on test set:"+str(corr_pearson))
print("Spearman correlation on test set:"+str(corr_spearman))
# + colab={} colab_type="code" id="QpnVA7VsGd4U" outputId="94a59775-862e-48e4-f036-d7d3fb4d7654"
# Calculate the spearman and pearson correlation, restricted to non-zero bins
corr_pearson_nonzero_bins=pearsonr(spi1_test_regression_truth[nonzero_bins],spi1_test_regression_predictions[nonzero_bins])
corr_spearman_nonzero_bins=spearmanr(spi1_test_regression_truth[nonzero_bins],spi1_test_regression_predictions[nonzero_bins])
print("Pearson correlation on test set:"+str(corr_pearson_nonzero_bins))
print("Spearman correlation on test set:"+str(corr_spearman_nonzero_bins))
# + colab={} colab_type="code" id="KCaEkJICGd4V"
test_df=pd.DataFrame({"Observed":list(spi1_test_regression_truth.values.squeeze()),
"Predicted":list(spi1_test_regression_predictions.squeeze())})
# + colab={} colab_type="code" id="9kktZbRCGd4W"
test_df_nonzero=pd.DataFrame({"Observed":list(spi1_test_regression_truth[nonzero_bins].values.squeeze()),
"Predicted":list(spi1_test_regression_predictions[nonzero_bins].squeeze())})
# + colab={} colab_type="code" id="af-4fxu0Gd4X" outputId="bc675856-7680-4a00-d016-a4e0fe3e6d55"
import plotnine
from plotnine import *
print((ggplot(test_df,aes(x="Observed",y="Predicted"))
+geom_bin2d(bins=100)
+theme_bw()
+xlab("Observed asinh(mean coverage in FC bigWig")
+ylab("Model prediction")
+ggtitle("SPI1 regression model test set prediction")))
print((ggplot(test_df_nonzero,aes(x="Observed",y="Predicted"))
+geom_bin2d(bins=100)
+theme_bw()
+xlab("Observed asinh(mean coverage in FC bigWig")
+ylab("Model prediction")
+ggtitle("SPI1 regression model test set prediction: bins with nonzero coverage")))
# + [markdown] colab_type="text" id="cZnRq3SzGd4Z"
# ## Kat's Model DeepLIFT profiles (Classification)
# + colab={} colab_type="code" id="UdWiImsYGd4Z"
spi1_test_classification_truth_bool=spi1_test_classification_truth.values.astype(bool)
true_pos_spi1=spi1_test_classification_truth[spi1_test_classification_truth_bool*spi1_test_classification_predictions >0.9]
# + colab={} colab_type="code" id="6iSq-J-iGd4a" outputId="578fdbcd-8cd1-4d5d-c6e2-4d28234b6610"
from dragonn.utils import one_hot_from_bed
deep_lift_input_spi1=one_hot_from_bed([i for i in true_pos_spi1.index],"hg19.genome.fa.gz")
deep_lift_input_spi1.shape
# + colab={} colab_type="code" id="XRIkb8oFGd4c"
from dragonn.tutorial_utils import deeplift, plot_seq_importance
deep_lift_scores_spi1=deeplift(spi1_kat_classification_model,deep_lift_input_spi1)
# + colab={} colab_type="code" id="sCAY5G5JGd4g" outputId="580aca0a-9a35-4cc8-84bc-15e9bf399d56"
plot_seq_importance(deep_lift_scores_spi1[0],deep_lift_input_spi1[0])
plot_seq_importance(deep_lift_scores_spi1[1],deep_lift_input_spi1[1])
plot_seq_importance(deep_lift_scores_spi1[2],deep_lift_input_spi1[2])
# + colab={} colab_type="code" id="fefAyEp4Gd4h" outputId="dca31e18-7f19-48bd-e76d-86a98e6cb221"
plot_seq_importance(deep_lift_scores_spi1[2].squeeze()[400:500],deep_lift_input_spi1[2].squeeze()[400:500])
# + [markdown] colab_type="text" id="NaizmyYbGd4i"
# If we query the sequence "GTTTCACTTCTGCAAA" in the [TomTom](http://meme-suite.org/tools/tomtom) software from the MEME suite, we find that the motif is a good match (p=3.55e-03) for SPIB:
# <img src="https://github.com/kundajelab/dragonn/blob/master/paper_supplement/tutorial_images/SPIB.Kat.png?raw=1" alt="SPI12TomTom" width="400"/>
# + [markdown] colab_type="text" id="RFlubWfKGd4i"
# ## Kat's Model DeepLIFT profiles (Regression)
# + colab={} colab_type="code" id="5DdIebdVGd4j" outputId="450f7622-0471-49af-fcb4-cd442ec04c57"
#Sanity-check that the model is learning the SPI1 motif by running DeepLIFT on True Positives with high confidence (>0.9)
#get the true positive predictions
true_pos=spi1_test_regression_truth[(spi1_test_regression_truth.values*spi1_test_regression_predictions)>4]
true_pos.shape
# + colab={} colab_type="code" id="My8t2AvgGd4k" outputId="65d55c2a-48ca-4878-f554-5dcf00e33c9f"
deep_lift_input=one_hot_from_bed([i for i in true_pos.index],"hg19.genome.fa.gz")
deep_lift_input.shape
# + colab={} colab_type="code" id="PzURc6r9Gd4l"
deep_lift_scores_spi1=deeplift(spi1_regression_model,deep_lift_input_spi1,target_layer_idx=-1)
# + colab={} colab_type="code" id="tqrN9urUGd4n" outputId="709f3bef-fe1b-40d3-b775-d20740a84deb"
plot_seq_importance(deep_lift_scores_spi1[0],deep_lift_input_spi1[0])
plot_seq_importance(deep_lift_scores_spi1[1],deep_lift_input_spi1[1])
plot_seq_importance(deep_lift_scores_spi1[2],deep_lift_input_spi1[2])
# + colab={} colab_type="code" id="leSwvZWFGd4n" outputId="848dd89f-eaa3-4872-ae04-3301ac0fff7d"
plot_seq_importance(deep_lift_scores_spi1[2].squeeze()[400:500],deep_lift_input_spi1[2].squeeze()[400:500])
# + [markdown] colab_type="text" id="QMBWHvnZV42V"
# ## Conclusions <a name='9'>
# <a href=#outline>Home</a>
# + [markdown] colab_type="text" id="PFyKUGQnV42X"
# ## Save tutorial outputs <a name='10'>
# <a href=#outline>Home</a>
#
# We save the models and test set predictions generated in this tutorial to an hdf5 file so that they can be loaded more readily in the future.
# + colab={} colab_type="code" id="WBNQNhg3V42X"
#save the models
#spi1_kat_classification_model.save("SPI1.kat.classification.model.hdf5")
#spi1_kat_regression_model.save("SPI1.kat.regression.model.hdf5")
#spi1_classification_model.save("SPI1.classification.model.hdf5")
#spi1_regression_model.save("SPI1.regression.model.hdf5")
#save the test predictions
import h5py
test_set_predictions=h5py.File("SPI1.test.kat.predictions.hdf5",'w')
test_set_predictions.create_dataset("classification",data=spi1_test_classification_predictions)
test_set_predictions.create_dataset("regression",data=spi1_test_regression_predictions)
test_set_predictions.close()
# + colab={} colab_type="code" id="LOPcHHY5V42a"
| tutorials/Supplementary Primer Tutorial 1 - Effects of Suboptimal Architecture on Functional variant characterization for non-coding SNPs within the SPI1 motif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.005862, "end_time": "2020-11-13T12:23:29.149526", "exception": false, "start_time": "2020-11-13T12:23:29.143664", "status": "completed"} tags=[]
# SHARED NOTEBOOK : Espaço para postarmos as melhorias incrementais do projeto (a partir de desenvolvimento nos nossos respectivos notebooks privados).
#
# Não se esqueçam de salvar as versões após as mudanças!
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 1.13421, "end_time": "2020-11-13T12:23:30.288429", "exception": false, "start_time": "2020-11-13T12:23:29.154219", "status": "completed"} tags=[]
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_score
from sklearn.preprocessing import StandardScaler
from datetime import datetime
import calendar
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" papermill={"duration": 11.042265, "end_time": "2020-11-13T12:23:41.337047", "exception": false, "start_time": "2020-11-13T12:23:30.294782", "status": "completed"} tags=[]
data = pd.read_csv('donors.csv', low_memory=False)
# -
#
# PROMOTION CODES
# 1st 2 bytes of the code refers to the year of the
# mailing while 3rd and 4th bytes refer to the
# following promotion codes/types:
#
# LL mailings had labels only
# WL mailings had labels only
# CC mailings are calendars with stickers but do
# not have labels
# FS mailings are blank cards that fold into
# thirds with labels
# NK mailings are blank cards with labels
# SK mailings are blank cards with labels
# TK mailings have thank you printed on the
# outside with labels
# GK mailings are general greeting cards (an
# assortment of birthday, sympathy, blank, & get
# well) with labels
# XK mailings are Christmas cards with labels
# X1 mailings have labels and a notepad
# G1 mailings have labels and a notepad
#
# This information could certainly be used to calculate
# several summary variables that count the number of
# occurrences of various types of promotions received
# in the most recent 12-36 months, etc.
#
#
# + papermill={"duration": 0.193601, "end_time": "2020-11-13T12:23:41.535965", "exception": false, "start_time": "2020-11-13T12:23:41.342364", "status": "completed"} tags=[]
adate = data[['Unnamed: 0','ADATE_2','ADATE_3','ADATE_4','ADATE_5','ADATE_6','ADATE_7','ADATE_8','ADATE_9','ADATE_10','ADATE_11','ADATE_12','ADATE_13','ADATE_14','ADATE_15','ADATE_16','ADATE_17','ADATE_18','ADATE_19','ADATE_20','ADATE_21','ADATE_22','ADATE_23','ADATE_24']]
adate.columns = ['ID','17NK','16NK','16TK','16SK','16LL','16G1','16GK','16CC','16WL','16X1','16XK','15FS','15NK','15TK','15LL','15G1','15GK','15CC','15WL','15X1','15XK','14FS','14NK']
adate # Date the 17NK promotion was mailed
# -
adate.iloc[36]
rfa = data[['Unnamed: 0','RFA_2','RFA_3','RFA_4','RFA_5','RFA_6','RFA_7','RFA_8','RFA_9','RFA_10','RFA_11','RFA_12','RFA_13','RFA_14','RFA_15','RFA_16','RFA_17','RFA_18','RFA_19','RFA_20','RFA_21','RFA_22','RFA_23','RFA_24']]
rfa.columns = ['ID','17NK','16NK','16TK','16SK','16LL','16G1','16GK','16CC','16WL','16X1','16XK','15FS','15NK','15TK','15LL','15G1','15GK','15CC','15WL','15X1','15XK','14FS','14NK']
rfa.drop(columns = 'ID') # Donor's RFM status as of 17NK promotion date
rdate = data[['Unnamed: 0','RDATE_3','RDATE_4','RDATE_5','RDATE_6','RDATE_7','RDATE_8','RDATE_9','RDATE_10','RDATE_11','RDATE_12','RDATE_13','RDATE_14','RDATE_15','RDATE_16','RDATE_17','RDATE_18','RDATE_19','RDATE_20','RDATE_21','RDATE_22','RDATE_23','RDATE_24']]
rdate.columns = ['ID','16NK','16TK','16SK','16LL','16G1','16GK','16CC','16WL','16X1','16XK','15FS','15NK','15TK','15LL','15G1','15GK','15CC','15WL','15X1','15XK','14FS','14NK']
rdate = rdate.drop(columns = 'ID') # Date the gift was received for 16NK
lastgiftdate = []
for i in range(0,len(rdate)):
if rdate.iloc[i].isnull().sum() == 22:
lastgiftdate.append(0)
else:
lastgiftdate.append(rdate.iloc[i][rdate.iloc[i].notnull()][0])
rdate['lastgiftdate'] = lastgiftdate
rdate.loc[:,'lastgiftdate'] = rdate.loc[:,'lastgiftdate'].map(lambda x: datetime.strptime(x, '%Y-%m-%d')
if not isinstance(x, int) else 0)
#Last promotion date
lastpromotiondate = datetime.strptime('2017-06-01', '%Y-%m-%d')
Recency = []
for i in rdate['lastgiftdate']:
if i == 0:
Recency.append(None)
else:
Recency.append((lastpromotiondate.year - i.year)*12 + (lastpromotiondate.month - i.month))
ramnt = data[['Unnamed: 0','RAMNT_3','RAMNT_4','RAMNT_5','RAMNT_6','RAMNT_7','RAMNT_8','RAMNT_9','RAMNT_10','RAMNT_11','RAMNT_12','RAMNT_13','RAMNT_14','RAMNT_15','RAMNT_16','RAMNT_17','RAMNT_18','RAMNT_19','RAMNT_20','RAMNT_21','RAMNT_22','RAMNT_23','RAMNT_24']]
ramnt.columns = ['ID','16NK','16TK','16SK','16LL','16G1','16GK','16CC','16WL','16X1','16XK','15FS','15NK','15TK','15LL','15G1','15GK','15CC','15WL','15X1','15XK','14FS','14NK']
ramnt = ramnt.drop(columns = 'ID') #Dollar amount of the gift for 16NK
Monetary = ramnt.sum(axis=1)
Frequency = ramnt.count(axis=1)
data['Recency'] = Recency
data['Frequency'] = Frequency
data['Monetary'] = Monetary
RFM_Promotion = data.loc[:,['Recency','Frequency','Monetary']]
RFM_Promotion.dropna(inplace = True)
RFM_Promotion['Recency_star'] = pd.qcut(RFM_Promotion['Recency'].rank(method='first'), q=5 , labels= (5,4,3,2,1))
RFM_Promotion['Recency_star'] = RFM_Promotion['Recency_star'].astype(int)
RFM_Promotion['Frequency_star'] = pd.qcut(RFM_Promotion['Frequency'].rank(method='first'), q=5 , labels= range(1,6))
RFM_Promotion['Frequency_star'] = RFM_Promotion['Frequency_star'].astype(int)
RFM_Promotion['Monetary_star'] = pd.qcut(RFM_Promotion['Monetary'].rank(method='first'), q=5 , labels= range(1,6))
RFM_Promotion['Monetary_star'] = RFM_Promotion['Monetary_star'].astype(int)
RFM_Promotion['Stars'] = (RFM_Promotion['Recency_star'].astype(str) +
RFM_Promotion['Frequency_star'].astype(str) +
RFM_Promotion['Monetary_star'].astype(str))
RFM_Promotion
kmeans_promotion = KMeans(init='k-means++', n_clusters=8, n_init=10, max_iter=300)
kmeans_promotion.fit(RFM_Promotion.loc[:, ['Recency_star','Frequency_star', 'Monetary_star']])
kmeans_promotion.n_iter_
kmeans_promotion.cluster_centers_
RFM_Promotion['Cluster'] = kmeans_promotion.labels_
RFM_Promotion_table = RFM_Promotion.groupby(['Cluster']).agg(['mean'])
RFM_Promotion_table['Total'] = RFM_Promotion_table.loc[:,['Recency_star','Frequency_star', 'Monetary_star']].sum(axis = 1)
round(RFM_Promotion_table.sort_values(by = 'Total',ascending = False),2)
# # RFM LIFE TIME
#transform to datetime
data['LASTDATE'] = data['LASTDATE'].map(lambda x: datetime.strptime(x, '%Y-%m-%d'))
#Last promotion date
lastpromotiondate = datetime.strptime('2017-06-01', '%Y-%m-%d')
lastpromotiondate.year
#Date associated with the most recent gift
data['Recency'] = (lastpromotiondate.year - data['LASTDATE'].dt.year)*12 + (lastpromotiondate.month - data['LASTDATE'].dt.month)
#Number of lifetime gifts to date
data['Frequency'] = data['NGIFTALL']
#Average dollar amount of gifts to date
data['Monetary'] = data['AVGGIFT']
RFM = data.loc[:, ['Recency', 'Frequency', 'Monetary']]
RFM
RFM['Recency_star'] = pd.qcut(RFM['Recency'].rank(method='first'), q=5 , labels= (5,4,3,2,1))
RFM['Recency_star'] = RFM['Recency_star'].astype(int)
RFM['Frequency_star'] = pd.qcut(RFM['Frequency'].rank(method='first'), q=5 , labels= range(1,6))
RFM['Frequency_star'] = RFM['Frequency_star'].astype(int)
RFM['Monetary_star'] = pd.qcut(RFM['Monetary'].rank(method='first'), q=5 , labels= range(1,6))
RFM['Monetary_star'] = RFM['Monetary_star'].astype(int)
RFM['Stars'] = (RFM['Recency_star'].astype(str) + RFM['Frequency_star'].astype(str) + RFM['Monetary_star'].astype(str))
RFM
kmeans = KMeans(init='k-means++', n_clusters=8, n_init=10, max_iter=300)
kmeans.fit(RFM.loc[:, ['Recency_star','Frequency_star', 'Monetary_star']])
kmeans.n_iter_
kmeans.cluster_centers_
RFM['Cluster'] = kmeans.labels_
RFM.groupby(['Cluster']).agg(['min','max','mean'])
RFM_table = RFM.groupby(['Cluster']).agg(['mean'])
RFM_table['Total'] = RFM_table.loc[:,['Recency_star','Frequency_star', 'Monetary_star']].sum(axis = 1)
round(RFM_table.sort_values(by = 'Total',ascending = False),2)
round(RFM_Promotion_table.sort_values(by = 'Total',ascending = False),2)
data['DATASRCE']
kmeans.inertia_
kmeans_promotion.inertia_
# # Cluster Metric Tests
from yellowbrick.cluster import KElbowVisualizer
from yellowbrick.cluster.elbow import kelbow_visualizer
X = RFM.loc[:, ['Recency_star','Frequency_star', 'Monetary_star']]
kelbow_visualizer(KMeans(init='k-means++', n_init=10, max_iter=300), X, k=(4,12))
X = RFM_Promotion.loc[:, ['Recency_star','Frequency_star', 'Monetary_star']]
kelbow_visualizer(KMeans(init='k-means++', n_init=10, max_iter=300), X, k=(4,12))
# +
X = RFM.loc[:, ['Recency_star','Frequency_star', 'Monetary_star']]
model = KMeans(init='k-means++', n_init=10, max_iter=300)
visualizer = KElbowVisualizer(
model, k=(4,12), metric='silhouette', timings=False
)
visualizer.fit(X) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# +
X = RFM_Promotion.loc[:, ['Recency_star','Frequency_star', 'Monetary_star']]
model = KMeans(init='k-means++', n_init=10, max_iter=300)
visualizer = KElbowVisualizer(
model, k=(4,12), metric='silhouette', timings=False
)
visualizer.fit(X) # Fit the data to the visualizer
visualizer.show() # Finalize and render the figure
# -
| Cluster_RFM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy.fftpack as fftpack
from astropy.table import Table
import matplotlib.pyplot as plt
import matplotlib.font_manager as font_manager
from stingray.events import EventList
from stingray.lightcurve import Lightcurve
from stingray import Powerspectrum, AveragedPowerspectrum
# %matplotlib inline
font_prop = font_manager.FontProperties(size=16)
# # Problem 1: damped harmonic oscillator example
# Generating a light curve
dt = 0.0001 # time step, in seconds
duration = 200 # length of time, in seconds
omega = 2*np.pi # angular frequency, in radians
phi = 0.0 # offset angle, in radians
# ## 1a. Compute the time steps and a cosine harmonic with the above-defined properties.
# For plotting ease below, save them as `time` and `oscill`.
# ## 1b. Compute four exponentially damped versions of the harmonic oscillation.
# $$D(t_i) = e^{-\zeta t_i}H(t_i)$$
# where $H(t_i)$ is your harmonic oscillating time series.
#
# Pick your own four $\zeta$ values. I recommend values between 0.01 and 1.
# Save them as `damp1`, `damp2`, etc.
zeta1 = 0.01
damp1 = np.exp(-time * zeta1) * oscill
# Make 3 more damped harmonic oscillators with your own pick of zeta:
# ## 1c. Plot them all on top of each other.
fig, ax = plt.subplots(1, 1, figsize=(8, 4), dpi=300)
ax.plot(time, oscill, lw=2, linestyle='-', color='black')
ax.plot(time, damp1, lw=2, linestyle='-', color='orange')
ax.plot(time, damp2, lw=2, linestyle='-.', color='blue')
ax.plot(time, damp3, lw=2, linestyle='--', color='magenta')
ax.plot(time, damp4, lw=2, linestyle='-', color='green')
ax.set_xlim(0,8)
ax.set_xlabel("Time (seconds)", fontproperties=font_prop)
ax.set_ylabel("Amplitude", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# ## 1d. Take the power spectrum of the harmonic and 4 damped harmonic time series.
# The power $P$ at each frequency $\nu_i$, for the Fourier transform $F$, is $$P(\nu_i)=|F(\nu_i)|^2$$
pow_oscill = np.abs(fftpack.fft(oscill)) ** 2
# Now you take the power spectrum of the damped harmonic time series. Again, for plotting ease, save as `pow_damp1`, etc.
# Test out what happens if you don't use 'abs'. What data type do you get?
type(pow_damp1[2])
# ## 1e. Plot them!
# Notice the trend between the width of the peak in the power spectrum, and the strength of the damping factor.
# +
freq = fftpack.fftfreq(len(time), d=dt)
nyq_ind = int(len(time)/2.) # the index of the last positive Fourier frequency
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(freq[0:nyq_ind], pow_oscill[0:nyq_ind].real/3e9,
lw=2, drawstyle='steps-mid', color='black')
ax.plot(freq[0:nyq_ind], pow_damp1[0:nyq_ind].real/1e9,
lw=2, drawstyle='steps-mid', linestyle='-', color='orange')
ax.plot(freq[0:nyq_ind], pow_damp2[0:nyq_ind].real/1e9,
lw=2, drawstyle='steps-mid', linestyle='-.', color='blue')
ax.plot(freq[0:nyq_ind], pow_damp3[0:nyq_ind].real/1e9,
lw=2, drawstyle='steps-mid', linestyle='--', color='magenta')
ax.plot(freq[0:nyq_ind], pow_damp4[0:nyq_ind].real/1e9,
lw=2, drawstyle='steps-mid', color='green')
ax.set_xlim(0.5, 1.5)
ax.set_ylim(1e-3, 5e2)
ax.set_yscale('log')
ax.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax.set_ylabel("Amplitude (arbitrary)", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# -
# # 2. Problem 2: Same idea, using Stingray!
# Now that you see the math behind the power spectrum, let's start using the spectral-timing library Stingray. Pull up the documentation to help you along.
# ## 2a. Make a simple `Lightcurve` object from `oscill`.
lc = Lightcurve(time, oscill)
# *Look at the warnings!* You already know your timestep `dt` and you know that you lightcurve is sorted (time always increases from lower indices to higher indices), so you can skip those checks here. Look at the Stingray documentation to see how to set these parameters.
# ## 2b. Try making a power spectrum of that lightcurve using `Powerspectrum`.
ps = Powerspectrum(lc)
print(ps)
# Ok, you probably see a ValueError above. Let's rethink this.
#
# The difference between our previous rough-and-tumble power spectrum (squaring the absolute value of the Fourier transform) and Stingray's `Powerspectrum` is that Stingray expects its data to be photon counts. Our sample data goes negative (since we were doing a simple case of deviations from a mean value of 0), but Stingray knows that you can't detect negative photons!
# ## 2c. Getting the sample data in the right format for Stingray
# So, to make our data fit Stingray's expectation, multiply our light curve `oscill` by a scaling factor and add a mean photon count rate value to that scaled light curve (anywhere from 100 to 1000 is a reasonable X-ray photon counts/second). Since Stingray expects the count rate as counts per time bin (not counts per second - pay attention to units!), the counts must be an integers. Hint: `np.rint` can be a helpful method.
# Plot `.time` vs `.counts`
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(lc.time, lc.counts, lw=2, drawstyle='steps-mid', color='black')
ax.set_xlim(0,8)
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Amplitude (photon counts)", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# ## 2d. Now make a power spectrum of that light curve using Stingray
# (redoing 2b, this time without an error)
# ### 2d.i. Do the same 2a-2d for damp1 through damp4, making a scaled light curve and power spectrum.
# Call them `lc1`, `ps1`, `lc2`, `ps2`, etc.
lc1 = Lightcurve(time, np.rint(damp1*2)+2, dt=dt, skip_checks=True)
ps1 = Powerspectrum(lc1)
lc2 = Lightcurve(time, np.rint(damp2*2)+2, dt=dt, skip_checks=True)
ps2 = Powerspectrum(lc2)
lc3 = Lightcurve(time, np.rint(damp3*2)+2, dt=dt, skip_checks=True)
ps3 = Powerspectrum(lc3)
lc4 = Lightcurve(time, np.rint(damp4*2)+2, dt=dt, skip_checks=True)
ps4 = Powerspectrum(lc4)
# Plot the power spectra! No need to compute the Nyquist frequency like we did in problem 1, since Stingray's default is only to keep and plot the positive Fourier frequencies.
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(ps.freq, ps.power, lw=2, drawstyle='steps-mid', color='black')
ax.plot(ps1.freq, ps1.power, lw=2, drawstyle='steps-mid', linestyle='-', color='orange')
ax.plot(ps2.freq, ps2.power, lw=2, drawstyle='steps-mid', linestyle='-.', color='blue')
ax.plot(ps3.freq, ps3.power, lw=2, drawstyle='steps-mid', linestyle='--', color='magenta')
ax.plot(ps4.freq, ps4.power, lw=2, drawstyle='steps-mid', color='green')
ax.set_xlim(0.5, 1.5)
ax.set_ylim(1e-4, 5e2)
ax.set_yscale('log')
ax.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax.set_ylabel("Amplitude (arbitrary)", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# Remember, the reason we are plotting right around 1 Hz is because we defined the time series to have that frequency. With real data, you don't want to zoom in your plots like that initially.
# # Problem 3: Analyzing *NICER* data of the black hole X-ray binary MAXI J1535-571
# Import it with astropy tables from the fits file "J1535_evt.fits", and call it `j1535`.
j1535 = Table.read("./J1535_evt.fits", format='fits')
# The data have come to us as an 'event list', meaning that it's a list of the time at which a photon was detected (in seconds, in spacecraft clock time) and the energy of the photon (a detector channel integer; channel/100=photon energy in keV).
# ## 3a. Turn this list of photons into an evenly-spaced light curve
# ### 3a.i.
# First, clean it up a little by only keeping photons with energies greater than 1 keV and less than 12 keV, using array masking.
print(len(j1535))
energy_mask = (j1535['ENERGY'] >= 100) & (j1535['ENERGY'] <= 1200)
j1535 = j1535[energy_mask]
print(len(j1535))
# Printing the lengths to show how many events were removed with this filter.
#
# ### 3a.ii.
# Use Stingray's method `Lightcurve.make_lightcurve` to turn this event list into a light curve with evenly spaced time bins and photon counts per bin. Pick a light curve time resolution of `dt=1/8`seconds to start with.
# These things might take a second; you're using half a million time bins in your light curve! I sometimes check the min and max of a light curve, to be sure that there wasn't an error.
np.max(lc_j1535.countrate)
np.min(lc_j1535.countrate)
# ## 3b. Let's try taking the power spectrum of it.
# Plot it!
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(ps.freq, ps.power, lw=1, drawstyle='steps-mid', color='black')
ax.set_yscale('log')
ax.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax.set_ylabel("Power/Hz", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# It's ugly! But more importantly, you can't get useful science out of it.
# ## What's going on?
# 1. There are gaps in the light curve (see below) due to the orbit of the spacecraft (and occasionally stuff gets in the way). This has the effect of inserting top-hat windows into our function, which give the lumpy bumps at ~0.25 Hz. So, we need to break the light curve up into shorter segments that won't have weird drop-outs.
# 2. There is a giant DC component at $\nu=0$. This is not astrophysical in origin, but from the mean of the light curve.
# 3. Power spectra are often plotted on log-log scales, but the power gets really noisy and 'scattery' at higher frequencies.
# 4. The eagle-eyed observer will notice that we can only go up to a Nyquist frequency of 4 Hz. There are interesting astrophysical signals above 4 Hz, but if we did smaller `dt` with keeping the very long segment length, we'd have >1 million time bins, which can be asking a lot of a laptop processor.
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(lc.time, lc.countrate, lw=2, drawstyle='steps-mid', color='black')
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Amplitude (counts/s)", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# ## 3c. Segments!
# ## 3c.i. GTIs
# Sometimes, the detector is on and recording photons, but it's pointed too close to the Earth, or a structure on the spacecraft is occulting part of the view, or the instrument is moving through a zone of high particle background, or other things. The times when these things happen are recorded, and in data reduction you make a list of Good Time Intervals, or GTIs, which is when you can use good science data. I made a list of GTIs for this data file that are longer than 4 seconds long, which you can read in from "J1535_gti.fits", and call it `gti_tab`.
# Stingray needs the gtis as a list of start and stop time pairs.
gtis = [[i,j] for (i,j) in zip(gti_tab['START'], gti_tab['STOP'])]
# ### 3c.ii. Segments
# Not only do we want to only use data in the GTIs, but we want to split the light curve up into multiple equal-length segments, take the power spectrum of each segment, and average them together, using `AveragedPowerspectrum`. By using shorter time segments like `segment_size=32` seconds, we can use a finer `dt` like 1/64 sec on the light curves, without having so many bins for each computation that our computer grinds to a halt. There is the added bonus that the noise amplitudes will tend to cancel each other out, and the signal amplitudes will add, and we get better signal-to-noise! When calculating this averaged power spectrum here, use `norm=none`.
#
# Make a new `Lightcurve` object of the data and the averaged power spectrum of that lightcurve with these recommended properties.
#
# As you learned in lecture, setting the length of the segment determines the lowest frequency you can probe, but for stellar-mass compact objects where we're usually interested in variability above ~0.1 Hz, this is an acceptable trade-off.
# Plot the light curve and its corresponding power spectrum! Note that the Good Times Intervals are saved to the `Lightcurve` object, but won't appear to be applied to the plotted data.
# The counts per second should be the same, regardless of your time binning!
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(lc_new.time, lc_new.countrate, lw=2, drawstyle='steps-mid', color='black')
# ax.set_xlim(0,8)
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Amplitude (counts/s)", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.plot(ps_new.freq, ps_new.power, lw=1, drawstyle='steps-mid', color='black')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax.set_ylabel("Power/Hz", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# Now, we've also applied logarithmic scales to the x and y axes in addition to using the GTIs. You can see something just to the left 10 Hz much clearer!
# The sharp signal in the lowest frequency bin is called the 'DC component', which is not astrophysical and arises from the mean count rate of the light curve. For ease, we typically plot these starting at frequency bin index 1 instead of index 0. If you're calculating your own power spectra with Fourier transforms outside of Stingray, subtract the mean counts/s from the light curve (in counts/s) before taking the Fourier transform.
# ## 3d. Error on average power
# The average power at a particular frequency has a chi-squared distribution with two degrees of freedom about the true underlying power spectrum. So, error is the value divided by the root of the number of segments (`M` in Stingray). A big reason why we love power spectra(/periodograms) is because this is so straight forward!
#
# $\text{error} = \frac{\text{power}}{\sqrt{M}}$
#
# One way to intuitively check if your errors are way-overestimated or way-underestimated is whether the size of the error bar looks commeasurate with the amount of bin-to-bin scatter of power at neighbouring frequencies.
ps_new.power_err
# Plotting, this time with `ax.errorbar` instead of `ax.plot`.
fig, ax = plt.subplots(1, 1, figsize=(8, 5), dpi=300, tight_layout=True)
ax.errorbar(ps_new.freq[1:], ps_new.power[1:], yerr=ps_new.power_err[1:],
lw=1, drawstyle='steps-mid', color='black')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax.set_ylabel("Power/Hz", fontproperties=font_prop)
ax.tick_params(axis='both', which='major', labelsize=16,
top=True, right=True, bottom=True, left=True)
plt.show()
# The thing at ~8 Hz is a low-frequency QPO, and the hump at-and-below 1 Hz is broadband noise! Now that you've got the basic analysis step complete, we'll focus on plotting the data in a meaningful way so you can easily extract information about the QPO and noise.
# ## 3e. Re-binning
# We often plot power spectra on log-log scaled axes (so, log on both the X and Y), and you'll notice that there's a big noisy part above 10 Hz. It is common practice to bin up the power spectrum `logarithmically` (which is like making it equal-spaced in when log-plotted).
#
# For this written example, I'll use a re-binning factor of 0.03 (or 3%). If new bin 1 has the width of one old bin, new bin 2 will be some 3% of a bin wider. New bin 3 will be 3% wider than *that* (the width of new bin 2), etc. For the first couple bins, this will round to one old bin (since you can only have an integer number of bins), but eventually a new bin will be two old bins, then more and more as you move higher in frequency. If the idea isn't quite sticking, try drawing out a representation of old bins and how the new bins get progressively larger by the re-binning factor.
#
# For a given new bin `x` that spans indices `a` to `b` in the old bin array:
# $$\nu_{x} = \frac{1}{b-a}\sum_{i=a}^{b}\nu_{i}$$
# $$P_{x} = \frac{1}{b-a}\sum_{i=a}^{b}P_{i}$$
# $$\delta P_{x} = \frac{1}{b-a}\sqrt{\sum_{i=a}^{b}(\delta P_{i})^{2}}$$
#
# Thanks to Stingray, you don't need to code up these equations! Try using the `rebin` method for linear re-binning in frequency and `rebin_log` for logarithmically re-binning.
fig, ax2 = plt.subplots(1,1, figsize=(9,6))
ax2.errorbar(rb_ps.freq, rb_ps.power, yerr=rb_ps.power_err, lw=1,
drawstyle='steps-mid', color='black')
ax2.set_xscale('log')
ax2.set_yscale('log')
ax2.set_xlim(0.1, rb_ps.freq[-1])
ax2.set_xlabel(r'Frequency (Hz)', fontproperties=font_prop)
ax2.set_ylabel(r'Power/Hz', fontproperties=font_prop)
ax2.tick_params(axis='x', labelsize=16, bottom=True, top=True,
labelbottom=True, labeltop=False)
ax2.tick_params(axis='y', labelsize=16, left=True, right=True,
labelleft=True, labelright=False)
plt.show()
# Play around with a few different values of the re-bin factor `f` to see how it changes the plotted power spectrum. 1 should give back exactly what you put in, and 1.1 tends to bin things up quite a lot.
# Congratulations! You can make great-looking power spectra! Now, go back to part 3c. and try 4 or 5 different combinations of `dt` and `seg_length`. What happens when you pick too big of a `dt` to see the QPO frequency? What if your `seg_length` is really short?
#
# One of the most important things to notice is that for a real astrophysical signal, the QPO (and low-frequency noise) are present for a variety of different `dt` and `seg_length` parameters.
# ## 3g. Normalization
# The final thing standing between us and a publication-ready power spectrum plot is the normalization of the power along the y-axis. The normalization that's commonly used is fractional rms-squared normalization. For a power spectrum created from counts/second unit light curves, the equation is:
# $$P_{frac} = P \times \frac{2*dt}{N * mean^2}$$
# `P` is the power we already have,
# `dt` is the time step of the light curve,
# `N` is the number of bins in one segment, and
# `mean` is the mean count rate (in counts/s) of the light curve.
#
# Stingray already knows this equation! Look in its documentation for normalizations. After you remake your average power spectrum from 3c.ii. with `norm=frac`, don't forget to re-bin it!
fig, ax = plt.subplots(1,1, figsize=(9,6))
ax.errorbar(rb_ps.freq, rb_ps.power, yerr=rb_ps.power_err, lw=1,
drawstyle='steps-mid', color='black')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlim(0.1, rb_ps.freq[-1])
ax.set_xlabel(r'Frequency (Hz)', fontproperties=font_prop)
ax.set_ylabel(r'Power [(rms/mean$^{2}$)/Hz]', fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16, bottom=True, top=True,
labelbottom=True, labeltop=False)
ax.tick_params(axis='y', labelsize=16, left=True, right=True,
labelleft=True, labelright=False)
plt.show()
# ## 3h. Poisson noise level
# Notice that the Poisson noise is a power law with slope 0 at high frequencies. With this fractional rms-squared normalization, we can predict the power of the Poisson noise level from the mean counts/s rate of the light curve!
# $$P_{noise} = 2/meanrate$$
#
# Compute this noise level (call it `poissnoise`), and plot it with the power spectrum.
fig, ax = plt.subplots(1,1, figsize=(9,6))
ax.errorbar(rb_ps.freq, rb_ps.power, yerr=rb_ps.power_err, lw=1,
drawstyle='steps-mid', color='black')
ax.hlines(poissnoise, rb_ps.freq[0], rb_ps.freq[-1], color='red')
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlim(0.1, rb_ps.freq[-1])
ax.set_xlabel(r'Frequency (Hz)', fontproperties=font_prop)
ax.set_ylabel(r'Power [(rms/mean$^{2}$)/Hz]', fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16, bottom=True, top=True,
labelbottom=True, labeltop=False)
ax.tick_params(axis='y', labelsize=16, left=True, right=True,
labelleft=True, labelright=False)
plt.show()
# Your horizontal Poisson noise line should be really close to the power at and above ~10 Hz.
# ## 3i. For plotting purposes, we sometimes subtract the Poisson noise level from the power before plotting.
# Once we've done this and removed the noise, we can also plot the data in units of Power, instead of Power/Hz, by multiplying the power by the frequency. Recall that following the propagation of errors, you will need to multiply the error by the frequency as well, but not subtract the Poisson noise level there.
# Beautiful! This lets us see the components clearly above the noise and see their *relative* contributions to the power spectrum (and thus to the light curve).
# ## Recap of what you learned in problem 3:
# You are now able to take a light curve, break it into appropriate segments using the given Good Time Intervals, compute the average power spectrum (without weird aliasing artefacts), and plot it in such away that you can see the signals clearly.
# # Problem 4: It's pulsar time
# We are going to take these skills and now work on two different observations of the same source, the ultra-luminous X-ray pulsar Swift J0243.6+6124. The goal is for you to see how different harmonics in the pulse shape manifest in the power spectrum.
# ## 4a. Load the data and GTI
# Using the files J0243-122_evt.fits and J0243-134_evt.fits, and the corresponding *x*_gti.fits. Call them `j0243_1`, `gti_1`, `j0243_2`, and `gti_2`.
# Look back to problem 3 for help with syntax.
j0243_1 = Table.read("./J0243-122_evt.fits", format='fits')
gti_1 = Table.read("./J0243-122_gti.fits", format='fits')
j0243_2 = Table.read("./J0243-134_evt.fits", format='fits')
gti_2 = Table.read("./J0243-134_gti.fits", format='fits')
# ## 4b. Apply a mask to remove energies below 0.5 keV and above 12 keV.
# Again, look to problem 3 for help with syntax.
# ## 4c. Make the average power spectrum for each data file.
# Go through in the same way as 3c. The spin period is 10 seconds, so I don't recommend using a segment length shorter than that (try 64 seconds). Since the period is quite long (for a pulsar), you can use a longer `dt`, like 1/8 seconds, and use `frac` normalization. Use the same segment length and dt for both data sets. Re-bin your averaged power spectrum.
fig, ax = plt.subplots(1,1, figsize=(9,6))
ax.errorbar(ps_1.freq, ps_1.power, lw=1,
drawstyle='steps-mid', color='purple')
ax.errorbar(ps_2.freq, ps_2.power, lw=1,
drawstyle='steps-mid', color='green')
## Plotting without error bars for now
ax.set_xscale('log')
ax.set_yscale('log')
ax.set_xlim(0.01, 6)
ax.set_xlabel(r'Frequency (Hz)', fontproperties=font_prop)
ax.set_ylabel(r'Power (rms/mean$^{2}$)', fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16, bottom=True, top=True,
labelbottom=True, labeltop=False)
ax.tick_params(axis='y', labelsize=16, left=True, right=True,
labelleft=True, labelright=False)
plt.show()
# Side note: if you don't normalize them (`none`), notice how the countrate of the light curve correlates with the power.
# ## 4d. Make a phase-folded light curve the brute force way
# ### 4d.i. Spin period
# Determine the spin period from the frequency of the lowest (fundamental) tone in the power spectrum. Remember that period=1/f. Hint: `np.argmax` is a great function for quick, brute-force things.
spin_f = ps_2.freq[np.argmax(ps_2.power[0:10])]
period = 1./spin_f
# ### 4d.ii. Relative phases
# Use the modulo operator of the light curve (starting it at time zero, the first element in the time array) to determine the relative phase of each photon event, then divide by the period to have relative phase from 0 to 1.
rel_time1 = np.asarray(j0243_1['TIME']) - j0243_1['TIME'][0]
rel_phase1 = (rel_time1 % period) / period
rel_time2 = np.asarray(j0243_2['TIME']) - j0243_2['TIME'][0]
rel_phase2 = (rel_time2 % period) / period
# ### 4d.iii. Binning
# Make an array of 20 phase bins and put the relative phases in their phase bins with `np.histogram`. Call the results `phase1` and `bins1` for the first data set, and `phase2` and `bins2` for the second.
# ### 4d.iv. Plot the light curve next to its accompanying power spectrum
fig, ax = plt.subplots(1,1, figsize=(9,6))
ax.plot(bins1[0:-1], phase1, lw=1, color='purple')
ax.set_xlabel(r'Relative phase', fontproperties=font_prop)
ax.set_ylabel(r'Counts per phase bin', fontproperties=font_prop)
ax.set_xlim(0, 1)
ax.tick_params(axis='x', labelsize=16, bottom=True, top=True,
labelbottom=True, labeltop=False)
ax.tick_params(axis='y', labelsize=16, left=True, right=True,
labelleft=True, labelright=False)
plt.show()
fig, ax = plt.subplots(1,1, figsize=(9,6))
ax.plot(bins2[0:-1], phase2, lw=1, color='green')
ax.set_xlabel(r'Relative phase', fontproperties=font_prop)
ax.set_ylabel(r'Counts per phase bin', fontproperties=font_prop)
ax.set_xlim(0, 1)
ax.tick_params(axis='x', labelsize=16, bottom=True, top=True,
labelbottom=True, labeltop=False)
ax.tick_params(axis='y', labelsize=16, left=True, right=True,
labelleft=True, labelright=False)
plt.show()
# Though these are very quickly made phase-folded light curves, you can see how the light curve with stronger harmonic content shows more power at the harmonic frequency in the power spectrum, and the light curve that's more asymmetric in rise and fall times (number 1) shows power at higher harmonics!
#
# If you want to see what a real phase-folded pulse profile looks like for these data, check out the beautiful plots in Wilson-Hodge et al. 2018: https://ui.adsabs.harvard.edu/abs/2018ApJ...863....9W/abstract
# Data set 1 has an observation ID that ends in 122 and corresponds to MJD 58089.626, and data set 2 has an observation ID that ends in 134 and corresponds to MJD 58127.622.
# # Bonus challenges:
# ### 5. Dynamical power spectrum (/spectrogram):
# Instead of averaging the power spectra at each segment, save it into a dynamical power spectrum (also called a spectrogram) using `DynamicalPowerspectrum` in Stingray. Apply the normalization (see if you can re-bin it), then make a 3d plot with frequency along the y-axis, segment (which corresponds to elapsed time) along the x-axis, and power as the colormap. Don't subtract the Poisson noise before plotting here, since some segments will have noisy power below the Poisson noise level, and then you're trying to plot negative numbers on a log scale, which is a very bad idea.
#
# This approach is useful if you think the QPO turns on and off rapidly (high-frequency QPOs do this) or is changing its frequency on short timescales. If the frequency is changing, this can artificially broaden the Lorentzian-shaped peak we see in the average power spectrum. Or, sometimes it's intrinsically broad. A look at the dynamical power spectrum will tell you! This will be most interesting on the black hole J1535 data, but could be done for both objects.
# ### 6. Energy bands:
# Make and plot power spectra of the same object using light curves of different energy bands. For example, try 1-2 keV, 2-4 keV, and 4-12 keV. Try to only loop through the event list once as you do the analysis for all three bands. What do you notice about the energy dependence of the signal?
# ### 7. Modeling:
# Using astropy.modeling or stingray.modeling (or your own preferred modeling package), fit the power spectrum of the black hole J1535 with a Lorentzian for the QPO, a few Lorentzians for the low-frequency broadband noise, and a power law for the Poisson noise level. In papers we often report the centroid frequency and the full-width at half maximum (FWHM) of the QPO Lorentzian model. How would you rule out the presence of a QPO at, e.g., 12 Hz?
# ### 8. Data visualization:
# Add a legend to the power spectra plot in problem 1, so that the label for the color gives the corresponding $\zeta$.
# ### 9. Poisson noise:
# Go through problem 2 and use `np.random.poisson` to apply Poisson noise to the signals (`oscill` and the four `damp`), and take the power spectra again, and plot them. Then try using the information in problem 3 about the Poisson noise level as it relates to the average count rate of the light curve to calculate and plot them together.
# ### 10. Stingray phase-folded light curves:
# Looking through the Stingray documentation (and possibly HENDRICS), find a more elegant way to make phase-folded pulsar light curves.
| tutorials/abigail_timeseries/time_series_workbook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''env'': conda)'
# name: python3
# ---
# +
import numpy as np
import gym
import random
import matplotlib.pyplot as plt
env = gym.make("Taxi-v3").env
env.render()
# -
print(f'action space is :{env.action_space}')
print(f'state space is: {env.observation_space}')
env.reset()
env.s = env.encode(3,1,1,1)
print(f'state: {env.s}')
env.render()
env.reset()
env.s = env.encode(0,4,4,1)
print(f'state: {env.s}')
env.render()
print(f'state: {env.P[env.s]}')
# initialize q table
q = np.zeros((env.observation_space.n, env.action_space.n))
q_pre = np.zeros((env.observation_space.n, env.action_space.n)) # to check convergence when training
print(f'current q table shape is: {q.shape}')
# hyper parameters
alpha = 0.5
gamma = 0.95
epsilon = 0.1
def epsilon_greedy_policy(env, state, q, epsilon):
'''
epsilon greedy policy for q learning to generate actions
'''
if random.uniform(0,1) < epsilon:
return env.action_space.sample()
else:
return np.argmax(q[state])
def update_q_table(q, pre_state, action, reward, next_state, alpha, gamma):
'''
'''
next_max = np.max(q[next_state]) # max state-action value for next state
# print(f'current status is: {type(q[pre_state,action])},{type(alpha)},{type(reward)},{type(gamma)},{type(next_max)}')
q[pre_state,action] = q[pre_state,action] + alpha * (reward + gamma * next_max - q[pre_state,action])
# +
#q learning
reward_record = []
error_record = []
# loop for each episode:
for episode in range(5000):
r = 0
state = env.reset()
while True:# loop for each step of episode
# choose A from S using policy derived from Q(e.g, epsilon greedy policy)
action = epsilon_greedy_policy(env,state,q,epsilon)
# take action A, observe R, S'
next_state, reward, done, _ = env.step(action)
# update Q(S,A)
update_q_table(q,state,action,reward,next_state,alpha,gamma)
# S<--S'
state = next_state
r += reward
if done:
break
reward_record.append(r)
error = 0
for i in range(q.shape[0]):
error = error + np.sum(np.abs(q[i]-q_pre[i]))
# print(f'{np.abs(q[i]-q_pre[i])},{np.sum(np.abs(q[i]-q_pre[i]))}')
error_record.append(error)
q_pre = np.copy(q)
if episode%100 == 0:
print(f'{episode}th episode: {r}, {error}')
#close game env
env.close()
#plot diagram
plt.plot(list(range(5000)),reward_record)
plt.show()
plt.plot(list(range(5000)),error_record)
plt.show()
# -
def epsilon_greedy_policy_sarsa(env, state, sarsa, epsilon):
'''
epsilon greedy policy for q learning to generate actions
'''
if random.uniform(0,1) < epsilon:
return env.action_space.sample()
else:
return np.argmax(sarsa[state])
def update_sarsa_table(sarsa, state, action, reward, next_state, next_action, alpha, gamma):
'''
update sarsa state-action pair value, main difference from q learning is that it uses epsilon greedy policy
return action
'''
next_max = sarsa[next_state,next_action] # corresponding action-state value to current action
# print(f'current status is: {type(q[pre_state,action])},{type(alpha)},{type(reward)},{type(gamma)},{type(next_max)}')
sarsa[state,action] = sarsa[state,action] + alpha * (reward + gamma * next_max - sarsa[state,action])
# +
env = gym.make("Taxi-v3")
# sarsa learning
# initialize sarsa table
sarsa = np.zeros((env.observation_space.n, env.action_space.n))
sarsa_pre = np.zeros((env.observation_space.n, env.action_space.n)) # to check convergence when training
sarsa_reward_record = []
sarsa_error_record = []
# loop for each episode:
for episode in range(5000):
r = 0
state = env.reset()
# choose A from S using policy derived from Q(e.g, epsilon greedy policy)
action = epsilon_greedy_policy_sarsa(env, state, sarsa, epsilon)
while True:# loop for each step of episode
# take action A, observe R, S'
next_state, reward, done, _ = env.step(action)
# choose action A' from S' using policy derived from Q(e.g, epsilon greedy policy)
next_action = epsilon_greedy_policy_sarsa(env, state, sarsa, epsilon)
# update Q(S,A)
update_sarsa_table(sarsa, state, action, reward, next_state, next_action, alpha, gamma)
# S<--S'
state = next_state
action = next_action
r += reward
if done:
break
sarsa_reward_record.append(r)
error = 0
for i in range(sarsa.shape[0]):
error = error + np.sum(np.abs(sarsa[i]-sarsa_pre[i]))
# print(f'{np.abs(q[i]-q_pre[i])},{np.sum(np.abs(q[i]-q_pre[i]))}')
sarsa_error_record.append(error)
sarsa_pre = np.copy(sarsa)
if episode%100 == 0:
print(f'{episode}th episode: {r}, {error}')
#close game env
env.close()
#plot diagram
plt.plot(list(range(5000)),sarsa_reward_record)
plt.plot(list(range(5000)),reward_record)
plt.show()
plt.plot(list(range(5000)),sarsa_error_record)
plt.plot(list(range(5000)),error_record)
plt.show()
# +
import numpy as np
import gym
# FrozenLake-v0 gym environment
# env = gym.make('FrozenLake-v0')
env = gym.make("Taxi-v3")
# Parameters
epsilon = 0.1
total_episodes = 5000
# max_steps = 100
alpha = 0.1
gamma = 0.6
#Initializing the Q-vaue
Q = np.zeros((env.observation_space.n, env.action_space.n))
Q_pre = np.copy(Q)
test_reward = []
test_error = []
# Function to choose the next action with episolon greedy
def choose_action(state):
action=0
if np.random.uniform(0, 1) < epsilon:
action = env.action_space.sample()
else:
action = np.argmax(Q[state])
return action
#Initializing the reward
reward=0
# Starting the SARSA learning
for episode in range(total_episodes):
# t = 0
state1 = env.reset()
# action1 = choose_action(state1)
action1 = epsilon_greedy_policy_sarsa(env, state1, Q, epsilon)
r = 0
# while t < max_steps:
while True:
# Visualizing the training
# env.render()
# Getting the next state
state2, reward, done, info = env.step(action1)
#Choosing the next action
# action2 = choose_action(state2)
action2 = epsilon_greedy_policy_sarsa(env, state2, Q, epsilon)
#Learning the Q-value
# Q[state1, action1] = Q[state1, action1] + alpha * (reward + gamma * Q[state2, action2] - Q[state1, action1])
update_sarsa_table(Q, state1, action1, reward, state2, action2, alpha, gamma)
state1 = state2
action1 = action2
#Updating the respective vaLues
# t += 1
r += reward
#If at the end of learning process
if done:
break
test_reward.append(r)
error = 0
for i in range(Q.shape[0]):
error = error + np.sum(np.abs(Q[i]-Q_pre[i]))
# print(f'{np.abs(q[i]-q_pre[i])},{np.sum(np.abs(q[i]-q_pre[i]))}')
test_error.append(error)
Q_pre = np.copy(Q)
if episode%100 == 0:
print(f'current reward is: {r}, {error}')
#close game env
env.close()
#plot diagram
plt.plot(list(range(5000)),reward_record)
plt.plot(list(range(5000)),test_reward)
plt.show()
plt.plot(list(range(5000)),error_record)
plt.plot(list(range(5000)),test_error)
plt.show()
# +
#double q learning
env = gym.make("Taxi-v3")
# initialize q table
q1 = np.zeros((env.observation_space.n, env.action_space.n))
q2 = np.zeros((env.observation_space.n, env.action_space.n))
q1_pre = np.zeros((env.observation_space.n, env.action_space.n)) # to check convergence when training
q2_pre = np.zeros((env.observation_space.n, env.action_space.n)) # to check convergence when training
# reward and error record
d_reward_record = []
d_error_record = []
# loop for each episode:
for episode in range(5000):
r = 0
state = env.reset()
while True:# loop for each step of episode
# choose A from S using policy derived from Q1+Q2(e.g, epsilon greedy policy)
action = epsilon_greedy_policy(env,state,q1+q2,epsilon)
# take action A, observe R, S'
next_state, reward, done, _ = env.step(action)
# with 0.5 probability:
if random.uniform(0,1) < 0.5:
update_q_table(q1,state,action,reward,next_state,alpha,gamma)
else:
update_q_table(q2,state,action,reward,next_state,alpha,gamma)
# S<--S'
state = next_state
r += reward
if done:
break
d_reward_record.append(r)
error = 0
for i in range(q.shape[0]):
error = error + 0.5 * np.sum(np.abs(q1[i]-q1_pre[i])) + 0.5 * np.sum(np.abs(q2[i]-q2_pre[i]))
# print(f'{np.abs(q[i]-q_pre[i])},{np.sum(np.abs(q[i]-q_pre[i]))}')
d_error_record.append(error)
q1_pre = np.copy(q1)
q2_pre = np.copy(q2)
if episode%100 == 0:
print(f'{episode}th episode: {r}, {error}')
#close game env
env.close()
#plot diagram
plt.plot(list(range(5000)),reward_record)
plt.plot(list(range(5000)),d_reward_record)
plt.show()
plt.plot(list(range(5000)),error_record)
plt.plot(list(range(5000)),d_error_record)
plt.show()
# +
#monte carlo method
env = gym.make("Taxi-v3")
def get_probs(q, env, epsilon):
'''
get the probability of taking the best known action according to epsion
'''
actions = np.argmax(q,dims=1)
policy_s = np.ones((env.observation_space.n, env.action_space.n))*epsilon/env.action_space.n
policy_s[:,actions] = 1 - epsilon + (epsilon / env.action_space.n)
return policy_s
def update_Q(env, episodes, Q, q):
for (s,a,r) in episodes:
sum = 0
for x in Q[(s,a)]:
sum += x
avg = sum/len(Q[(s,a)])
q[s,a] = avg
def run(env, Q, q, epsilon, gamma,r):
episodes = []
state = env.reset()
G = 0
while True:# loop for each step of episode
probs = get_probs(q, env, epsilon) # get the current behavior policy
action = np.random.choice(np.arange(env.action_space.n),p=probs)\
if state in Q else env.action_space.sample()
next_state, reward, done, _ = env.step(action)
episodes.append((state, action, reward))
G = gamma * G + reward
Q[(state, action)].append(G)
r += reward
state = next_state
if done:
break
return episodes
# initialize q table
q = np.zeros((env.observation_space.n, env.action_space.n))
Q = {}
# reward and error record
q_reward_record = []
q_error_record = []
# loop for each episode:
for episode in range(5000):
r = 0
state = env.reset()
episodes = run(env,Q,epsilon,gamma,r)
update_Q(env,episodes,Q,q)
if episode%100 == 0:
print(f'{episode}th episode: {r}')
# d_reward_record.append(r)
# error = 0
# for i in range(q.shape[0]):
# error = error + 0.5 * np.sum(np.abs(q1[i]-q1_pre[i])) + 0.5 * np.sum(np.abs(q2[i]-q2_pre[i]))
# # print(f'{np.abs(q[i]-q_pre[i])},{np.sum(np.abs(q[i]-q_pre[i]))}')
# d_error_record.append(error)
# q1_pre = np.copy(q1)
# q2_pre = np.copy(q2)
# if episode%100 == 0:
# print(f'{episode}th episode: {r}, {error}')
# #close game env
# env.close()
# #plot diagram
# plt.plot(list(range(5000)),reward_record)
# plt.plot(list(range(5000)),d_reward_record)
# plt.show()
# plt.plot(list(range(5000)),error_record)
# plt.plot(list(range(5000)),d_error_record)
# plt.show()
| TD/td_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import requests
import io
plt.rcParams.update({'font.size': 14})
# # European Monetary Union #
# ## M2 Monetary Aggregate ##
# + tags=[]
# Fetch data from the ECB
# https://sdw.ecb.europa.eu/browse.do?node=bbn3501
resp = requests.get('https://sdw.ecb.europa.eu/quickviewexport.do?SERIES_KEY=117.BSI.M.U2.N.V.M20.X.1.U2.2300.Z01.E&type=csv')
fp = io.StringIO(resp.text)
df = pd.read_csv(fp, skiprows=5, header=None, names=['month', 'm2_million_euro'])
# -
df['date'] = pd.to_datetime(df['month'], format='%Y%b')
df = df[['date', 'm2_million_euro']]
df['m2'] = df['m2_million_euro'] / 1e3
m2 = df
def plot_m2(df, start_date, zone, currency):
sel = df[df['date'] > start_date].copy()
sel = sel.sort_values('date', ascending=False).reset_index(drop=True)
sel['value_ratio'] = sel['m2'].loc[len(sel)-1] / sel['m2']
fig, ax = plt.subplots(figsize=(12, 7))
sel.plot(x='date', y='m2', label=f'{zone} M2 Monetary Aggregate', ax=ax)
plt.ylabel(f'Billion {currency}')
plt.xlabel('')
plt.grid(linestyle=':')
ax2 = sel.plot(x='date', y='value_ratio', label='Inverted M2, norm. to 1', ax=ax, secondary_y=True, color = 'red', linestyle='--')
ax.grid(linestyle=':')
#ax2.grid(None)
#plt.legend(['Eurozone M2 Monetary Aggregate'])
plot_m2(m2, '2015-01-01', 'Eurozone', 'Euros')
plt.title('Euro Monetary Inflation since Jan. 2015')
plt.show()
plot_m2(m2, '2019-12-31', 'Eurozone', 'Euros')
plt.title('Monetary Inflation since Jan. 2020')
plt.show()
# ## M2 Relative to GDP ##
# Fetch data from the ECB
# https://sdw.ecb.europa.eu/browse.do?node=bbn3501
resp = requests.get('https://sdw.ecb.europa.eu/quickviewexport.do?SERIES_KEY=139.AME.A.EA19.1.0.0.0.OVGD&type=csv')
fp = io.StringIO(resp.text)
df = pd.read_csv(fp, skiprows=5, header=None, names=['year', 'gdp'])
gdp = df
gdp['date'] = pd.to_datetime(gdp['year'], format='%Y')
def plot_gdp(m2, gdp, start_date, end_date, currency):
sel = m2[m2['date'] > '2010-01-01'].copy()
sel = sel.merge(gdp, on=['date'], how='outer')
sel = sel[['date', 'm2', 'gdp']]
sel = sel.sort_values('date').reset_index(drop=True)
sel['gdp'] = sel['gdp'].fillna(method='ffill')
sel = sel[sel['date'] > start_date]
sel = sel[sel['date'] < end_date]
sel = sel.reset_index(drop=True)
sel['m2_adjusted'] = (sel['m2'] / sel['m2'].loc[0]) * (sel['gdp'].loc[0] / sel['gdp'])
fig, ax = plt.subplots(figsize=(12, 7))
sel.plot(x='date', y='m2', label='M2 Monetary Aggregate', ax=ax, color='#1f77b4')
sel.plot(x='date', y='gdp', label='GDP', ax=ax, color='#ff7f0e', linestyle=':')
plt.ylabel(f'Billion {currency}')
plt.grid(linestyle=':')
ax2 = sel.plot(x='date', y='m2_adjusted', label='M2 Monetary Aggregate, adjusted for GDP', ax=ax, secondary_y=True, linestyle='--', color='#1f77b4')
ax.grid(linestyle=':')
plt.xlabel('')
plot_gdp(m2, gdp, '2015-01-01', '2022-01-01', 'Euros')
plt.title('Eurozone M2 Monetary Aggregate')
plt.show()
# ## Market Valuation Relative to M2 ##
msci = pd.read_csv('./csv/LU1646360971-MFE.csv')
msci['date'] = pd.to_datetime(msci['date'])
def plot_market(mkt, m2, start_date, label):
sel = mkt[mkt.date > start_date][['date', 'close']]
sel = sel.merge(m2, on=['date'], how='outer')
sel['mfe'] = sel['close']
sel = sel[['date', 'mfe', 'm2']]
sel = sel.sort_values('date').reset_index(drop=True)
sel['m2'] = sel['m2'].fillna(method='ffill')
sel = sel[sel['date'] > start_date]
sel = sel.sort_values('date').reset_index(drop=True)
sel['mfe_adjusted'] = (sel['mfe'] / sel['mfe'].loc[0]) * (sel['m2'].loc[0] / sel['m2'])
fig, ax = plt.subplots(figsize=(12, 7))
sel.plot(x='date', y='mfe', label=label, ax=ax)
plt.ylabel('Euros')
plt.grid(linestyle=':')
ax2 = sel.plot(x='date', y='mfe_adjusted', label=f'{label}, adjusted for M2', ax=ax, secondary_y=True, color = 'red', linestyle='--')
ax.grid(linestyle=':')
plt.xlabel('')
plot_market(msci, m2, '2015-01-01', 'Lyxor MSCI EMU Dist. (MFE)')
plt.title('MSCI EMU Valuation')
plt.show()
# # United States of America #
# ## M2 Monetary Aggregate ##
# Fetch data from FRED
resp = requests.get(
'https://fred.stlouisfed.org/graph/fredgraph.csv?bgcolor=%23e1e9f0&chart_type=line&drp=0&fo=open%20sans&graph_bgcolor=%23ffffff&'
'height=450&mode=fred&recession_bars=on&txtcolor=%23444444&ts=12&tts=12&width=1168&nt=0&thu=0&trc=0&show_legend=yes&'
'show_axis_titles=yes&show_tooltip=yes&id=M2SL&scale=left&cosd=1959-01-01&coed=2021-02-01'
'&line_color=%234572a7&link_values=false&line_style=solid&mark_type=none&mw=3&lw=2&ost=-99999&oet=99999&mma=0&fml=a&fq=Monthly'
'&fam=avg&fgst=lin&fgsnd=2020-02-01&line_index=1&transformation=lin&vintage_date=2021-04-03&revision_date=2021-04-03&nd=1959-01-01'
)
fp = io.StringIO(resp.text)
df = pd.read_csv(fp)
usm2 = df
usm2.columns = [c.lower() for c in usm2.columns]
usm2['date'] = pd.to_datetime(usm2['date'])
usm2 = usm2.rename(
columns = {'m2sl': 'm2'}
)
plot_m2(usm2, '2015-01-01', 'US', 'Dollars')
plt.title('Dollar Monetary Inflation since Jan. 2015')
plt.show()
plot_m2(usm2, '2019-12-31', 'US', 'Dollars')
plt.title('Dollar Monetary Inflation since Jan. 2020')
plt.show()
# ## M2 Relative to GDP ##
# Fetch data from FRED
resp = requests.get(
'https://fred.stlouisfed.org/graph/fredgraph.csv?bgcolor=%23e1e9f0&chart_type=line&drp=0&fo=open%20sans&graph_bgcolor=%23ffffff&'
'height=450&mode=fred&recession_bars=on&txtcolor=%23444444&ts=12&tts=12&width=1168&nt=0&thu=0&trc=0&show_legend=yes&show_axis_titles=yes&show_tooltip=yes&'
'id=GDP&scale=left&cosd=1947-01-01&coed=2020-10-01&line_color=%234572a7&link_values=false&line_style=solid&mark_type=none&mw=3&lw=2&ost=-99999&oet=99999&mma=0&'
'fml=a&fq=Quarterly&fam=avg&fgst=lin&fgsnd=2020-02-01&line_index=1&transformation=lin&vintage_date=2021-04-03&revision_date=2021-04-03&nd=1947-01-01'
)
fp = io.StringIO(resp.text)
df = pd.read_csv(fp)
usgdp = df
usgdp.columns = [c.lower() for c in usgdp.columns]
usgdp['date'] = pd.to_datetime(usgdp['date'])
plot_gdp(usm2, usgdp, '2015-01-01', '2022-01-01', 'Dollars')
plt.title('US M2 Monetary Aggregate')
plt.show()
# ## Market Valuation Relative to M2 ##
spy = pd.read_csv('./csv/US78462F1030-SPY.csv')
spy['date'] = pd.to_datetime(spy['date'])
plot_market(spy, usm2, '2015-01-01', 'SPDR® S&P 500 (SPY)')
plt.title('S&P 500 Valuation')
plt.show()
| M2 Monetary Aggregate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from modelconductor.modelhandler import VariableLengthHistoricalRegressionModelHandler
target_keys = ["A_pred", "time_pred"]
input_keys = ["A"]
control_keys = ['time']
min_window_size = 1
step_size = 0.01
model=VariableLengthHistoricalRegressionModelHandler(
target_keys=target_keys,
input_keys=input_keys,
control_keys=control_keys,
min_window_size=min_window_size, step_size=step_size)
# +
import asyncio
import nest_asyncio
nest_asyncio.apply()
from modelconductor.experiment_mqtt import MqttExperiment
ex = MqttExperiment()
ex.model = model
headers = ex.model.input_keys + ex.model.target_keys + ex.model.control_keys
ex.logger = ex.initiate_logging(headers=headers)
# -
ex.run()
log_path = ex.log_path #"experiment_2020-12-10_02-42-11.log" #
import pandas as pd
results = pd.read_csv(log_path)
results.head()
inputs = pd.read_csv(log_path + "_inputs", names = ["time"] + input_keys + ["shit"])
inputs.head()
inputs.tail()
from matplotlib import pyplot as plt
plt.plot(inputs['time'], inputs['A'], label="actual value")
plt.plot(results['time_pred'], results['A_pred'], label="predicted value")
plt.legend()
plt.xlabel('time (s)')
plt.savefig('line_plot.pdf')
plt.show()
plt.savefig('line_plot.pdf')
| egs/skl_historical_poly_regression_variable_window_overmqtt/skl_historical_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# name: python3
# ---
import laspy
new_las = laspy.create(file_version="1.2", point_format=3)
print(new_las)
print(type(new_las.X))
# +
import json
# ALREADY WITTEN FUNCS HERE
def list_dump(input_list,outpath,mode='w+'):
with open(outpath,mode) as list_writer:
for item in input_list:
list_writer.write(str(item)+'\n')
def print_rem_time_info(total_it,curent_it,ref_time,chunk_size=1):
# "it" stands for 'iteration'
it_time = time.time()-ref_time
rem_its = total_it-curent_it
rem_time = it_time * rem_its
print("{} iter took {:.4f} seconds, estimated remaining time: {:.4f} minutes or {:.4f} hours, iteration {} of {}".format(chunk_size,it_time*chunk_size,rem_time/60.0,rem_time/3600.0,curent_it,total_it))
# +
import subprocess
# NEW FUNCS HERE
def parse_json(jsonpath):
with open(jsonpath) as reader:
return json.load(reader)
def gdal_vrt_from_filelist(filelistpath,outputpath,print_runstring=True):
runstring = f'gdalbuildvrt -input_file_list {filelistpath} {outputpath}'
if(print_runstring):
print(runstring)
subprocess.run(runstring,shell=True)
def gdal_translate_dem(imagepath,outpath,bbox_minxy_maxxy=None,outformat='XYZ',boundaries_epsg='4326',custom_cellsize=None,enlargement_percent=None,print_runstring=True):
'''
bbox must be given as a list in guessed order
'''
bbox_part = ''
if bbox_minxy_maxxy:
ulx = bbox_minxy_maxxy[0]
uly = bbox_minxy_maxxy[3]
lrx = bbox_minxy_maxxy[1]
lry = bbox_minxy_maxxy[2]
bbox_part = f'-projwin {ulx} {uly} {lrx} {lry} -projwin_srs "EPSG:{boundaries_epsg}"'
cellsize_part = ''
if custom_cellsize and (not enlargement_percent):
cellsize_part = f'-r bilinear -tr {custom_cellsize} {custom_cellsize}'
enlargement_part = ''
if not custom_cellsize and enlargement_percent:
cellsize_part = f'-r bilinear -outsize {enlargement_percent}% {enlargement_percent}%'
runstring = f'gdal_translate {bbox_part} {cellsize_part} {enlargement_part} -of {outformat} {imagepath} {outpath}'
if(print_runstring):
print(runstring)
subprocess.run(runstring,shell=True)
def get_raster_val_at_geoXY(x,y,rasterpath):
runstring = f'gdallocationinfo -valonly -geoloc {rasterpath} {x} {y}'
ret = subprocess.run(runstring,shell=True, stdout=subprocess.PIPE).stdout.decode('utf-8')
return float(ret.strip('\n'))
def string_to_floatlist(inputstring):
return list(map(float,line.strip('\n').split(' ')))
# +
jsonpath = '/home/kaue/data/sanit3d_out/temporary/dtm_raster_data_dump.json'
dtm_dict = parse_json(jsonpath)
dtm_filename_list = list(dtm_dict['downloaded_image_path'].values())
# +
jsonpath = '/home/kaue/data/sanit3d_out/temporary/dsm_raster_data_dump.json'
dsm_dict = parse_json(jsonpath)
dsm_filename_list = list(dsm_dict['downloaded_image_path'].values())
# -
dtm_filename_list
dsm_filename_list
# +
dtm_filelist_name = 'test_dtm_list.txt'
dtm_vrt_path = 'test_vrt_dtm.vrt'
list_dump(dtm_filename_list,dtm_filelist_name)
dsm_filelist_name = 'test_dsm_list.txt'
dsm_vrt_path = 'test_vrt_dsm.vrt'
list_dump(dsm_filename_list,dsm_filelist_name)
# -
gdal_vrt_from_filelist(dtm_filelist_name,dtm_vrt_path)
gdal_vrt_from_filelist(dsm_filelist_name,dsm_vrt_path)
# +
lgt_min = -40.3397827083
lgt_max = -40.3353356058
lat_min = -20.3214263926
lat_max = -20.3176282943
bbox_test = [lgt_min,lgt_max,lat_min,lat_max]
# +
# tests 232 * 211 = 48952 spected points
dtm_xyz_path = 'dtm_test.xyz'
gdal_translate_dem(dtm_vrt_path,dtm_xyz_path,bbox_test)
# +
dsm_xyz_path = 'dsm_test.xyz'
gdal_translate_dem(dsm_vrt_path,dsm_xyz_path,bbox_test,custom_cellsize=0.5)
# + tags=["outputPrepend"]
import time
Xlist = []
Ylist = []
Zlist = []
Clist = []
with open(dtm_xyz_path) as reader:
for i,line in enumerate(reader):
if i % 100 == 0:
t1 = time.time()
X,Y,Z = string_to_floatlist(line)
Xlist.append(X)
Ylist.append(Y)
Zlist.append(Z)
Clist.append(2)
dsm_z = get_raster_val_at_geoXY(X,Y,dsm_vrt_path)
if(dsm_z - Z) > 2:
Xlist.append(X)
Ylist.append(Y)
Zlist.append(dsm_z)
Clist.append(0)
if i % 100 == 0:
print_rem_time_info(48672,i,t1,100)
# -
# +
import laspy
import numpy as np
# my_data_xx, my_data_yy = np.meshgrid(np.linspace(-20, 20, 15), np.linspace(-20, 20, 15))
# my_data_zz = my_data_xx ** 2 + 0.25 * my_data_yy ** 2
# my_data = np.hstack((my_data_xx.reshape((-1, 1)), my_data_yy.reshape((-1, 1)), my_data_zz.reshape((-1, 1))))
las = laspy.create(file_version="1.2", point_format=3)
# las.header.offsets = np.min(my_data, axis=0)
# las.header.scales = [0.1, 0.1, 0.1]
las.x = np.array(Xlist)
las.y = np.array(Ylist)
las.z = np.array(Zlist)
las.classification = np.array(Clist,dtype=np.intc)
las.write("sample_las.las")
# -
np.array(Clist,dtype=np.intc)
# +
# generate terrain polygon
import geopandas as gpd
polygons_geojson_path = '/home/kaue/data/sanit3d_out/temporary/buildings_osm.geojson'
as_gdf = gpd.read_file(polygons_geojson_path)
# +
from shapely import wkt
small_bbox_wkt = 'POLYGON((-40.3397827083 -20.3176282943, -40.3353356058 -20.3176282943, -40.3353356058 -20.3214263926, -40.3397827083 -20.3214263926, -40.3397827083 -20.3176282943))'
bbox_polygon = wkt.loads(small_bbox_wkt)
# -
bbox_polygon
# +
crs_proj = 'EPSG:31984'
data2 = {'name':['bbox'],'geometry':[bbox_polygon]}
bbox_gdf = gpd.GeoDataFrame(data2,crs="EPSG:4326")
bbox_proj = bbox_gdf.to_crs(crs_proj)
# -
clipped = as_gdf.intersection(bbox_polygon)
clipped = clipped[~clipped.is_empty]
# +
# clipping
crs_proj = 'EPSG:31984'
clipped.to_file('clipped_wgs84.geojson',driver='GeoJSON')
clipped.to_crs(crs_proj).to_file('clipped_31984.geojson',driver='GeoJSON')
# +
# symettrical_difference
ground_pol = bbox_gdf.symmetric_difference(clipped.unary_union)
ground_pol.to_file('ground_pol_wgs84.geojson',driver='GeoJSON')
# -
ground_pol.to_crs(crs_proj).to_file('ground_pol_31984.geojson',driver='GeoJSON')
# + tags=[]
# clipping thematic layers:
import geopandas as gpd
import glob, os
for filepath in glob.glob('/home/kaue/data/sanit3d_out/temporary/geonode:asb*'):
print(filename)
filename = filepath.split('cesan_')[1].split('_1712')[0]
filename = 'sample_'+filename+'.geojson'
lyr_gdf = gpd.read_file(filepath)
# lyr_gdf.intersection(bbox_proj).to_file(filename,driver='GeoJSON')
res = gpd.overlay(lyr_gdf,bbox_proj,how='intersection')
if not res.empty:
res.to_file(filename,driver='GeoJSON')
# -
| tests/tests_laspy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VacationPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import os
import ipywidgets as widgets
import geopy
# Import API key
import mapkey
# Configure gmaps
import gmaps
gmaps.configure(api_key='mapkey')
# -
# ### Store Part I results into DataFrame
# * Load the csv exported in Part I to a DataFrame
# +
output_df = pd.read_csv('output_data.csv')
vacation_df = output_df.drop(columns=["Unnamed: 0"])
vacation_df
# -
# ### Humidity Heatmap
# * Configure gmaps.
# * Use the Lat and Lng as locations and Humidity as the weight.
# * Add Heatmap layer to map.
# ### Create new DataFrame fitting weather criteria
# * Narrow down the cities to fit weather conditions.
# * Drop any rows will null values.
# +
vacation_df = pd.DataFrame(vacation_df, columns = ["City","Lat","Lng","Max Temp", "Wind Speed", "Humidity","Cloudiness"])
temp = (vacation_df["Max Temp"] <= 90) & (new_types_df["Max Temp"] > 75)
wind = vacation_df["Wind Speed"] < 10
humidity = vacation_df["Humidity"] < 10
cloudiness = vacation_df["Cloudiness"] == 0
# new_types_df[max_temp & wind_speed & cloudiness]
idealvacation_df[temp & wind & humidity & cloudiness]
# -
# ### Hotel Map
# * Store into variable named `hotel_df`.
# * Add a "Hotel Name" column to the DataFrame.
# * Set parameters to search for hotels with 5000 meters.
# * Hit the Google Places API for each city's coordinates.
# * Store the first Hotel result into the DataFrame.
# * Plot markers on top of the heatmap.
| VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# NumPy uses vectorization via broadcasting (avoiding loops)
import numpy as np
# ## DataTypes & Attributes
# NumPy's main datatype is ndarray
a1 = np.array([1, 2, 3])
a1
type(a1)
a2 = np.array([[1, 2],
[3, 4]])
a2
a1.shape
a2.shape
a1.ndim
a2.ndim
a1.dtype
a2.dtype
a1.size
a2.size
a3 = np.array([[[1, 2, 3],
[4, 5, 6,],
[7, 8, 9]],
[[1, 2, 3],
[4, 5, 6],
[7, 8, 9]]])
a3
a3.shape
a3.ndim
a3.size
# ## Create a DataFrame from a NumPy array
a4 = np.array([[1, 2, 3],
[4, 5, 6],
[7, 8, 9]])
# +
import pandas as pd
df = pd.DataFrame(a4)
df
# -
# ## Creating Arrays
ones = np.ones((2, 3), int)
ones
zeros = np.zeros((2, 3), int)
zeros
range_array = np.arange(0, 12, 2)
range_array
random_array = np.random.randint(0, 11)
random_array
r_array = np.random.randint(0, 11, size=(3, 5))
r_array
r3 = np.random.random((5, 3))
r3
# ## Pseudo-random numbers
# NumPy generates pseudo-random numbers
# Imagine that we have a notebook where we generate random numbers for an experiment, if we share this notebook we want
# the other person can replicate the inpunts, that means we want the random numbers can be replicate, for this case
# we can use a seed:
np.random.seed(seed=0) # create random numbers trackable with the seed
r4 = np.random.randint(10, size=(4, 4))
r4
# ## Viewing arrays and matrices
a1 = np.random.randint(1, 11, size=(3, 5))
a1
a2 = np.random.randint(1, 10, size=(1, 10))
a2
np.unique(a2)
np.unique(a1)
a3 = np.array([[1, 0, 0],
[1, 0, 0],
[2, 1, 3]])
np.unique(a3)
# Return the unique rows of a 2D array
np.unique(a3, axis=0)
np.unique(a3, axis=1)
a2
a4 = np.random.randint(1, 11, size=(2, 3, 4))
a4
a4[0]
a4[:]
# I want the seven of the first matrix
a4[0][1][2]
a4[0, 1, 2]
a5 = np.array([[[1, 2, 3],
[4, 5, 6],
[7, 8, 9]],
[[10, 11, 12],
[13, 14, 15],
[16, 17, 18]]])
a5
a5[:2, :2, :2]
a5[:2, 0, :]
a6 = np.random.randint(1, 11, size=(2, 3, 4, 5))
a6
# Printing the first number 2
a6[0, 0, 1, 4]
# ## Manipulating & Comparing arrays
# ### Arithmetic
a1 = np.array([1, 2, 3])
a1
ones = np.ones(3, dtype=int)
ones
a1 + ones
a1 - ones
3 * a1
a1 = np.array([1, 2, 3])
a2 = np.array([4, 5, 6])
a2 * a1
a1 = np.array([[1, 2],
[3, 4]])
a2 = np.array([[5, 6],
[7, 8]])
a1 * a2
a1 = np.array([1, 3, 5, 6, 8])
a1 % 2
# ### Aggregation
# Aggregation = performing the same operation on a number of things.
big_array = np.random.random(1_000_000)
big_array[:10]
big_array.size
# %timeit sum(big_array)
# %timeit np.sum(big_array)
a1 = np.random.randint(1, 10, size=(3, 3))
a1
np.sum(a1)
np.mean(a1)
np.min(a1)
np.max(a1)
# **standart deviation = a measure of how spread out a group of numbers is from the mean** <br>
# **standart deviation = square_root(variance)**
np.sqrt(np.var(a1)), np.std(a1)
# **variance = measure of the average degree to which each number is different to the mean** <br>
# **Higher variance = wider range of numbers.** <br>
# **Lower variance = lower range of numbers.**
np.var(a1)
# Demo of std and var
high_var_array = np.array([1, 100, 200, 300, 4_000, 5_000])
low_var_array = np.array([2, 4, 6, 8, 10])
np.var(high_var_array), np.var(low_var_array)
import matplotlib.pyplot as plt
plt.hist(high_var_array)
plt.show()
plt.hist(low_var_array)
plt.show()
# ## Reshaping & transposing
a = np.array([[1, 2, 3],
[4, 5, 6],
[7, 8, 9]])
a
a.T
a
# ## Dot Product
a1 = np.array([[1, 2, 3],
[4, 5, 6]])
a2 = np.array([[10, 11],
[20, 21],
[30, 31]])
# Matrix Product
np.dot(a1, a2)
v1 = np.array([1, 2, 3])
v2 = np.array([4, 5, 6])
np.dot(v1, v2)
np.dot(v1.T, v2)
v1 = np.array([[1, 2, 3]]).T
v1
v2 = np.array([[4, 5, 6]])
v2
np.dot(v1, v2)
| notebooks/.ipynb_checkpoints/NumPy-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import StandardScaler
df = pd.read_csv('creditcard.csv')
df.head()
df.describe()
Scaled_Amount = df[['Amount']]
scaler = StandardScaler().fit(Scaled_Amount.values)
Scaled_Amount = scaler.transform(Scaled_Amount.values)
df['Amount'] = Scaled_Amount
X = df.drop(columns = 'Class')
Y = df['Class']
seed = 7
test_size = 0.2
X_train ,X_test, Y_train, Y_test = train_test_split(X,Y, test_size =test_size, random_state = seed)
model = XGBClassifier()
model.fit(X_train,Y_train)
Y_pred = model.predict(X_test)
predictions = [round(value) for value in Y_pred]
accuracy = accuracy_score(Y_test, predictions)
print('Accuracy: %.2f%%' % (accuracy*100.0))
| Projects/Fraud Detection Project/.ipynb_checkpoints/Fraud Detection - XGB -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/thiagosantos346/PNL_MODELS/blob/master/pnl_multlabel.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="iRa4Ulvmnt8h" colab_type="text"
# # Processamento de linguagem natural : Atributos multi-rotulado.
# + [markdown] id="71FMPfZFSany" colab_type="text"
# ## Importação dos dados:
# + id="x9zLGeCwbE4b" colab_type="code" colab={}
import pandas as pd
# + id="U1owgoRjbtG1" colab_type="code" colab={}
data_set = pd.read_csv('/content/drive/My Drive/Colab Notebooks/stackoverflow/stackoverflow_perguntas.csv')
# + id="ikgeVKgbb0Ag" colab_type="code" outputId="ec276e18-1a17-4c5e-e34f-1d64c6db24bf" colab={"base_uri": "https://localhost:8080/", "height": 204}
data_set.head()
# + [markdown] id="7FvFrKu5SSqm" colab_type="text"
# ## Limpando os dados:
# + [markdown] id="s3FZN3gmoLCK" colab_type="text"
# Aparentemente, temos uma classificação com multiplos rotulos, pois vendo a coluna tags, notamos que na linha 4, existe dois rotulos para a pergunta...
# + [markdown] id="Ro-ayIPzoSEE" colab_type="text"
# Agoras é interessante saber também o tamanho do nosso problema...
# - Vamos ver o tamanho total do conjunto de dados(dataset)
# - Depois a quantidade de tags únicas...
#
# + id="ygK_jrCyn5xK" colab_type="code" outputId="b9cb9f0f-d137-42a8-a50b-6bc0480ee34a" colab={"base_uri": "https://localhost:8080/", "height": 34}
dataset_size = len(data_set)
message_size = 'Número de linhas no dataset:{}.'.format(dataset_size)
print (message_size)
# + id="nzGUco92p1WG" colab_type="code" outputId="979bfc0e-31ad-4c56-d575-a0d42b638248" colab={"base_uri": "https://localhost:8080/", "height": 204}
tag_list = data_set.Tags
size = len(tag_list.unique())
tag_list = tag_list.unique()
message_size = 'Número de valores únicos:{size}.\n Valores :\n {values}'
message_size = message_size.format(size=size, values=tag_list)
print(message_size)
# + [markdown] id="KtAAIz-JqPCV" colab_type="text"
# Feito isso podemos notar alguns problemas que devemos tratar:
# - A ordem das palavras estão gerando novos valores únicos;
# - Erros e ruidos tamabém produzem novos valores únicos;
#
# + id="t09ftmwVrr-P" colab_type="code" outputId="793b12f9-ed4f-473c-db8d-0f9150cf5a55" colab={"base_uri": "https://localhost:8080/", "height": 0}
def get_clean_tag_list(tags_list):
cleaned_tag_list = list()
for tags in tags_list:
tags_splited = tags.split()
for tag in tags_splited:
if tag not in cleaned_tag_list:
cleaned_tag_list.append(tag)
return cleaned_tag_list
cleaned_tag_list = get_clean_tag_list(tag_list)
print(cleaned_tag_list)
# + id="cJtOzZuUKS-T" colab_type="code" outputId="4cf3da06-26f7-48f8-fa27-af8bac9acf84" colab={"base_uri": "https://localhost:8080/", "height": 0}
print(data_set['Tags'])
# + id="mjYuYluSDkUJ" colab_type="code" colab={}
def count_label_in_row(label_list, data_set, column_name='Tags'):
for tag in label_list:
count_list = list()
for column in data_set[column_name]:
if tag in column:
value = 1
else:
value = 0
count_list.append(value)
data_set[tag] = count_list
# + id="yn9jLBkzI_16" colab_type="code" outputId="8210f578-885c-4b57-df21-edce9ab4add7" colab={"base_uri": "https://localhost:8080/", "height": 0}
count_label_in_row(cleaned_tag_list, data_set)
data_set.head(100)
# + [markdown] id="w__PQxpCQuYs" colab_type="text"
# Agora que criamos uma coluna para cada tag unica, e marcamos o número de ocorrências dessa tag na linha podemos prosseguir para dividir as massa de dados para treino e teste.
# + [markdown] id="afdjakQQSGjp" colab_type="text"
# ## Separação do conjunto de dados para teste e treino:
# + [markdown] id="WOMioUNvTmT2" colab_type="text"
# ### Zipar as colunas
# 1. Temos uma pequena diferença da divisão feita nos modelos mais simples:
# - Temos 4 colunas para representar apenas uma entidade;
# - Nosso divisor precisa de um atributo que represente as 4 colunas em apenas uma.
#
# Sendo assim ao dividir os dados de teste e treino poderiamos ficar com dados
# não proporcionais a realidade, isso é uma divisão de dados que não contivesse uma das tags que queremos treinar o nosos modelo, a ideia é dividir os dados de maneira que não falte entidades na massa de dados para treino, evitando assim testar um valor ao qual o modelo não foi treinado.
#
# 2. Solução:
# - Criar apenas uma coluna contendo o crusamento dos valores da lita.
# - Para isso vamos usar uma função de zipagens dos dados a funcção `zip()`.
# - Assin contruindo uma coluna de tupulas das cominações de de cada rotulo que queremos prover.
#
# + id="43b8e_K2WDXs" colab_type="code" outputId="2b147428-dedf-4fa5-bda5-d37fecaff705" colab={"base_uri": "https://localhost:8080/", "height": 204}
l1 = data_set[cleaned_tag_list[0]]
l2 = data_set[cleaned_tag_list[1]]
l3 = data_set[cleaned_tag_list[2]]
l4 = data_set[cleaned_tag_list[3]]
ziped_tag_list = list(zip( l1, l2, l3, l4))
data_set['all_tagas_tupules'] = ziped_tag_list
data_set.head()
# + [markdown] id="NyaWSSKJWEjj" colab_type="text"
# ### Seprar os dados:
# + id="fwrVcJmhQron" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
import random
fit_input_dataset, test_input_dataset, fit_output_dataset, test_output_dataset = train_test_split(
data_set['Perguntas'],
data_set['all_tagas_tupules'],
test_size=0.2,
random_state=random.randint(1, 1000)
)
# + [markdown] id="ApWg2-68ah6y" colab_type="text"
# #### Vetorização do input de treino:
# + [markdown] id="vcFj38quawwV" colab_type="text"
# ##### TF-IDF
# + [markdown] id="jgDi4E0qbcLn" colab_type="text"
# ###### Configurar o vetorizador :
# - Vamos definir o número maximo de linhas no vetor com `max_features`.
# - Vamos retirar as palavras que se repetem muinto com `max_df`.
# + id="6p3Wtt64TDnC" colab_type="code" colab={}
from sklearn.feature_extraction.text import TfidfVectorizer
MAX_FEATURES = 5000
MAX_DISTANCE_FREQUENCIE = 0.85
vectorizer = TfidfVectorizer(max_features=MAX_FEATURES, max_df=MAX_DISTANCE_FREQUENCIE)
# + [markdown] id="vV_KcnhXc3lJ" colab_type="text"
# ###### Vamos criar o vetor de frequência de palavras no texto:
# + id="W6ln69tqdI9n" colab_type="code" colab={}
vectorizer.fit(data_set['Perguntas'])
fit_input_dataset_tfidf = vectorizer.transform(fit_input_dataset)
test_input_dataset_tfidf = vectorizer.transform(test_input_dataset)
# + id="E1tfAsDWdqkr" colab_type="code" outputId="d8040b0a-2582-43be-e89d-f813f69f29a1" colab={"base_uri": "https://localhost:8080/", "height": 0}
print(fit_input_dataset_tfidf.shape)
print(test_input_dataset_tfidf.shape)
# + [markdown] id="bdqTo80TePz8" colab_type="text"
# Como vemos foram geradas 5000 linhas, e combinadas com a frequência da proporção de pareto 80/20, para treino e teste.
# + [markdown] id="2dUCci40R4IR" colab_type="text"
# ## Treinando o modelo:
# + [markdown] id="L4DgTvO33SxF" colab_type="text"
# **Relevância Binária**
# + [markdown] id="q4TYvkqynyOj" colab_type="text"
# Essa estratégia se base em uma catégorização binaria simples, porem aplicada a cada uma das tupulas que queremos testar.
# + [markdown] id="DDArHY8voJpH" colab_type="text"
# ### Classificador:
# Usáremos o algoritmo **OneVsRestClassifier**, que tem como **Requesitos**:
# - Um **Estimador**, usaremos a **Regreção lógistica**.
# + [markdown] id="cMvqO0MTpPOP" colab_type="text"
# ### Regreção Lógistica :
# + id="K3Ib1jbIo_yK" colab_type="code" colab={}
from sklearn.linear_model import LogisticRegression
instance_LogisticRegression = LogisticRegression(solver='lbfgs')
# + [markdown] id="yh49bKzApUkt" colab_type="text"
# ### OneVsOneClassifier
# + id="Ce_kPSh7oBBm" colab_type="code" colab={}
from sklearn.multiclass import OneVsRestClassifier
instance_OneVsRestClassifier = OneVsRestClassifier(instance_LogisticRegression)
# + [markdown] id="VErqoqrCqLtb" colab_type="text"
# ### Tranformação pre-treino:
# + [markdown] id="Y9uZJrkLqXzW" colab_type="text"
# O nosso classificador recebe como entrada um arranjo binário, porem os nosso dados de treinos são do tipo serie, para resolver isso precisamos converter esses dados em um arranjo binário ou uma matriz esparsa.
# + id="BZS5XgLpqqLS" colab_type="code" outputId="66afc383-29b7-4998-edc5-6ee1d2cbae53" colab={"base_uri": "https://localhost:8080/", "height": 0}
type(fit_output_dataset)
# + [markdown] id="hnI4YLPcuW16" colab_type="text"
# Além dessa transformação, precisamos fazer as tupulas se tornarem listas, por isso o list em volta dos dados de treino e test de saída.
# + id="K06zC1oUrKN8" colab_type="code" colab={}
import numpy as np
array_fit_output_dataset = np.asarray(list(fit_output_dataset))
array_test_output_dataset = np.asarray(list(test_output_dataset))
# + id="Bf8DkUNzryZj" colab_type="code" outputId="654633a6-16e0-46b9-959b-f7c44626f614" colab={"base_uri": "https://localhost:8080/", "height": 0}
len(array_test_output_dataset)
# + [markdown] id="zYgWN69IqA0l" colab_type="text"
# ### Executando o Treinamento :
# + [markdown] id="eKZ3OKhjvIZZ" colab_type="text"
# Agora basta treinar o modelo com os TF-IDF e os Arrays de saída, criados anteriormente:
# + id="WB1makMAp_-R" colab_type="code" outputId="b447e10b-5929-449f-8184-a221cb915e54" colab={"base_uri": "https://localhost:8080/", "height": 0}
instance_OneVsRestClassifier.fit(fit_input_dataset_tfidf, array_fit_output_dataset)
# + [markdown] id="BKiNkhhQS_TI" colab_type="text"
# ## Avaliando o modelo
# + [markdown] id="rcfqPROW3qOR" colab_type="text"
# ### Acurácia :
# + [markdown] id="MJnSJmV10hlt" colab_type="text"
# A acurácia não é o melhor meio de avaliar o nosso modelo, pois é baseda em `Exact Match` que seria uma combinação exata das 4 colunas que queremos
# predizer.
# + id="DnDXdyUsxPFv" colab_type="code" outputId="6389590d-5fcf-48ee-8fa4-ee64b8a4a3d0" colab={"base_uri": "https://localhost:8080/", "height": 34}
score = instance_OneVsRestClassifier.score(test_input_dataset_tfidf, array_test_output_dataset)
mensage_model_acuracy = 'Acurácia atribuida : {0: .2f}%'.format(score*100)
print(mensage_model_acuracy)
# + [markdown] id="_2ceYTcx1Bxu" colab_type="text"
# Só para termos uma noção, vamos ver quantas combinações possiveis existem no nosso resultado.
# + id="mAJZlNrG1MSq" colab_type="code" outputId="2255f6ed-70b7-40d5-9776-aa4a1dd085f1" colab={"base_uri": "https://localhost:8080/", "height": 34}
choices = len(data_set.all_tagas_tupules.unique())
print(choices)
# + [markdown] id="HH4pAdkg1dqh" colab_type="text"
# O nosso total é:
# + id="a0L9zhhu1iYJ" colab_type="code" outputId="d5bbbb18-8ceb-44c0-96cd-c00a666f67d2" colab={"base_uri": "https://localhost:8080/", "height": 34}
options = len(data_set.all_tagas_tupules)
print(options)
# + [markdown] id="QNZQRBjr1lsq" colab_type="text"
# Com uma escolha aleátoria termismos a seguinte chances de acertar:
# + id="44w82oVG1k4Z" colab_type="code" outputId="15448ddf-7ca2-4fbd-ac39-6c7816f73fec" colab={"base_uri": "https://localhost:8080/", "height": 0}
mensage_random_choice = 'Acurácia atribuida : {0: .2f}%'.format((choices/options) * 100)
print(mensage_random_choice )
# + [markdown] id="D1SCeFKB3kVk" colab_type="text"
# ### Hamming Loss
# + [markdown] id="Ppzu7l7K3zPM" colab_type="text"
# Esse metodo de avaliação trabalha com distância de elementos por linha, em outras palavras se tenho um **total de 4 atributos** para predizer e foram **obitidos 3 elementos corretos** a distância entre o total e o números de acerto é de **`` 4 - 3 = 1 ``**.
# Quanto mais proxima de zero for essa distância melhor será os nossos resultados.
# + id="31YFPYWZETrV" colab_type="code" colab={}
from sklearn.metrics import hamming_loss
prediction_oneVsRest = instance_OneVsRestClassifier.predict(test_input_dataset_tfidf)
hamming_loss_oneVsRest = hamming_loss(array_test_output_dataset, prediction_oneVsRest)
# + id="pNnhkeTqGfnA" colab_type="code" outputId="b276636d-1cfa-494c-e071-42248d795d65" colab={"base_uri": "https://localhost:8080/", "height": 34}
mensage_hamming_loss_distance = 'Distância entre real e previsto(Hamming Loss): {0: .2f}.'.format(hamming_loss_oneVsRest)
print(mensage_hamming_loss_distance )
# + [markdown] id="GbaEBRHEHUmS" colab_type="text"
# Podemos ver também a correlação entre cada uma das variaveis, já que nesse modelo esstas foram tratadas de forma indepêndentes.
# + id="lmSSRq5pHeiW" colab_type="code" outputId="d4cb81cf-a0a7-4b49-83ce-1cca5399e9f6" colab={"base_uri": "https://localhost:8080/", "height": 173}
data_set.corr()
# + [markdown] id="TfvavIjFH-gn" colab_type="text"
# Notamos que nenhuma das variáveis da matriz veio com 0, existe pelo menos uma correlação inversa, o que levanta a duvida, seria possivel melhorar o desempenho desse modelo, levando enconta essas correlações?
# + [markdown] id="M9ITk0ATKoBo" colab_type="text"
# ## Refinando o modelo:
# + [markdown] id="HIBVoNpXJzoD" colab_type="text"
# ### Classificação em Cadeia
# + [markdown] id="n-L-TJ_jKuR1" colab_type="text"
# Vamos usar a estrátegia de classificação em cadadeia(Cadeias de Markov).
# - Antes :
# - Entrada : Uma pergunta.
# - Saída : A classifcação de todas as tags.
# - Agora :
# - Entrada : A pergunta, e classificação do elemento anterior, a parti do segundo elemento.
# - Saida : A classificação individual de cada elemento, baseda nas classificações anteriores.
#
# + [markdown] id="J66M3LiyWcKg" colab_type="text"
# Antes vamos instalar a biblioteca responsavel para trabalhar com muil-label em cadeia o ``scikit-multilearn``.
# + id="gHlU9oFEK3M-" colab_type="code" outputId="4e534244-a77f-4923-be40-a6a718aabbb3" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !pip install scikit-multilearn
# + [markdown] id="-1xLwupHWo7V" colab_type="text"
# Vamos importar e estanciar o nosso pacote, para isso vamos usar a instância de regreção logistica já crida anteriormente.
# + id="vQSXT3ECWny5" colab_type="code" colab={}
from skmultilearn.problem_transform import ClassifierChain
instance_of_classifier_chain = ClassifierChain(instance_LogisticRegression)
# + [markdown] id="xeQRcMILXvos" colab_type="text"
# #### Treinamento do modelo:
# + id="vp7PmbH2X4P-" colab_type="code" outputId="d9cfeeb0-350a-4f18-d2a1-0b278f59bb06" colab={"base_uri": "https://localhost:8080/", "height": 170}
instance_of_classifier_chain.fit(fit_input_dataset_tfidf, array_fit_output_dataset)
# + [markdown] id="HlqK61JNX9jd" colab_type="text"
# #### Avaliando o modelo:
# + [markdown] id="hWjU-ChvYVT5" colab_type="text"
# #### Acurácia
# + id="Wk5emrLxYIrU" colab_type="code" outputId="7a108a14-4d7c-4713-af5e-5c24b515b503" colab={"base_uri": "https://localhost:8080/", "height": 34}
score = instance_of_classifier_chain.score(test_input_dataset_tfidf, array_test_output_dataset)
mensage_model_acuracy = 'Acurácia atribuida : {0: .2f}%'.format(score*100)
print(mensage_model_acuracy)
# + [markdown] id="oiKQRUJHYYP_" colab_type="text"
# #### Hamming Loss
# + id="A5lnvT3hYa1Y" colab_type="code" outputId="09e4ad9a-e2ec-4ede-e8e5-2f680e6289f2" colab={"base_uri": "https://localhost:8080/", "height": 34}
prediction_classifier_chain = instance_of_classifier_chain.predict(test_input_dataset_tfidf)
hamming_loss_classifier_chain = hamming_loss(array_test_output_dataset, prediction_classifier_chain)
mensage_hamming_loss_distance = 'Distância entre real e previsto(Hamming Loss): {0: .2f}.'.format(hamming_loss_classifier_chain)
print(mensage_hamming_loss_distance )
# + [markdown] id="TesLcZdMZLiH" colab_type="text"
# - Ao comparar os dois modelos treiandos vemos que a ditância de Hamming Loss ficou maior, ou seja estamos nos ditâncido do valor real dos resultados na visão individual dos atributos.
# - Já à acurácia se tonou maior o que indica que linhas completas dos atributos completos estão preditas com maior eficiência.
#
# + [markdown] id="xz7yrfDzJ-Zo" colab_type="text"
# ### ML-KNN
# + [markdown] id="GolLuF3FKFym" colab_type="text"
# Essa é uma outra opção para executar as mesma predições, porem usando um classificador baseado em visionhos mais proximo, o código é bastante parecedio ao anterior, fica aqui só como forma de ver que da para usar muito outros algoritmos com os dados tratados e avaliar qual foi o que se saiu melhor.
# + id="JlJQRiRPKGLQ" colab_type="code" colab={}
from skmultilearn.adapt import MLkNN
# + id="vcfmXB5hK4-W" colab_type="code" colab={}
instance_of_MLkNN = MLkNN()
# + id="SwvUClBeLLC_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0e2ee894-c020-40bb-9c83-bc24dbfb54f5"
instance_of_MLkNN.fit(fit_input_dataset_tfidf, array_fit_output_dataset)
# + id="o0u5UJtkK_ze" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b232e901-048b-471a-e574-62254c797914"
score = instance_of_MLkNN.score(test_input_dataset_tfidf, array_test_output_dataset)
mensage_model_acuracy = 'Acurácia atribuida : {0: .2f}%'.format(score*100)
print(mensage_model_acuracy)
# + id="vpTitLz0LXzx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="260d687f-4cff-4443-e4f6-5e4cf83fdc21"
prediction_mlknn = instance_of_MLkNN.predict(test_input_dataset_tfidf)
hamming_loss_mlknn = hamming_loss(array_test_output_dataset, prediction_mlknn)
mensage_hamming_loss_distance = 'Distância entre real e previsto(Hamming Loss): {0: .2f}.'.format(hamming_loss_classifier_chain)
print(mensage_hamming_loss_distance )
| pnl_multlabel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Local Image Classification Training - Bars
#
# Classifies the bars to extract the bits they represent.
BARS_FOLDER = "../bars/"
TRAIN_FOLDER = "../bars/train/"
TEST_FOLDER = "../bars/test/"
AUGMENT_FOLDER = "../bars/augmented/"
PARTITION_SCRIPT = "../tensorflow/scripts/preprocessing/partition_dataset.py"
import pandas as pd
from os import listdir
from os.path import isfile, join
import os
train_paths = [TRAIN_FOLDER + f for f in listdir(TRAIN_FOLDER) if isfile(join(TRAIN_FOLDER, f))]
test_paths = [TEST_FOLDER + f for f in listdir(TEST_FOLDER) if isfile(join(TEST_FOLDER, f))]
# ## Create a DataFrame with Paths and Labels
# +
train_df = pd.DataFrame({'path': train_paths})
train_df["dec"] = train_df.apply(lambda x: int(x['path'].split("/")[-1].split("_")[0]), axis=1)
train_df["bin"] = train_df.apply(lambda x: bin(x['dec']), axis=1)
train_df["class"] = train_df.apply(lambda x: [i for i in reversed(range(20)) if (x['dec'] & 1 << i) != 0], axis=1)
print(train_df.shape)
train_df.head()
# +
test_df = pd.DataFrame({'path': test_paths})
test_df["dec"] = test_df.apply(lambda x: int(x['path'].split("/")[-1].split("_")[0]), axis=1)
test_df["bin"] = test_df.apply(lambda x: bin(x['dec']), axis=1)
test_df["class"] = test_df.apply(lambda x: [i for i in reversed(range(20)) if (x['dec'] & 1 << i) != 0], axis=1)
print(test_df.shape)
test_df.head()
# -
# ## Define the Image Generators
from tensorflow.keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(
rotation_range = 4,
zoom_range = 0.02,
brightness_range=[0.15,0.85],
channel_shift_range=0.2,
shear_range = 0.2,
horizontal_flip = False,
vertical_flip = False,
fill_mode = 'nearest')
target_size = (450, 100)
train_generator = train_datagen.flow_from_dataframe(
dataframe = train_df,
directory = ".",
target_size = target_size,
batch_size = 64,
x_col = 'path',
y_col = 'class',
class_mode = 'categorical')
# save_to_dir = '../augmented_bars/'
test_datagen = ImageDataGenerator()
test_generator = test_datagen.flow_from_dataframe(
dataframe = test_df,
directory = ".",
target_size = target_size,
shuffle = False,
x_col = 'path',
y_col = 'class',
class_mode = 'categorical')
# ## Define the Model
#
# ### Load model for transfer learning
# +
from tensorflow.keras.applications import VGG19
model = VGG19(include_top=False, input_shape=(450,100,3))
model.trainable = False
# -
model.summary()
# ## Add custom Dense Layers
# +
import tensorflow as tf
from tensorflow.keras import layers, Model
x = model.get_layer('block4_pool').output #remove last conv layers block
# Flatten layer
flatten = layers.Flatten()
# Create output layer with 20 nodes and sigmoid activation to handle multiple classes and labels
bits_pred = layers.Dense(20, activation = 'sigmoid', name='bits_pred')
output = bits_pred(flatten(x))
# Create an instance of a Model
model_final = Model(model.input, output)
# compile
model_final.compile(loss = 'binary_crossentropy',
optimizer='adam',
metrics = [tf.keras.metrics.BinaryAccuracy(threshold=0.5)])
# -
model_final.summary()
# ## Define early stopping
#
# I want the model to stop, only when the model started overfitting
# +
from tensorflow.keras.callbacks import EarlyStopping
early_stopping = EarlyStopping(monitor='val_loss',
min_delta = 0,
patience = 5,
verbose = 1,
mode ='min',
restore_best_weights = True)
# -
# ## Train the model
callbacks = [early_stopping]
history = model_final.fit(
train_generator,
steps_per_epoch = 28,
epochs = 9999, # We only want the training to stop after the early stopping condition is met
validation_data = test_generator,
verbose = 2,
callbacks = callbacks)
# ## Evaluate
# +
import matplotlib.pyplot as plt
def plot_history(history):
f = plt.figure(figsize = (15,6))
ax = f.add_subplot(121)
ax.plot(history.history['binary_accuracy'])
ax.plot(history.history['val_binary_accuracy'])
ax.legend(['training accuracy', 'validation accuracy'],
bbox_to_anchor = (0, 1.02, 1, 0.2),
loc = "lower left",
mode = "expand",
ncol = 2)
ax2 = f.add_subplot(122)
ax2.plot(history.history['loss'])
ax2.plot(history.history['val_loss'])
ax2.legend(['training loss', 'validation loss'],
bbox_to_anchor = (0, 1.02, 1, 0.2),
loc = "lower left",
mode = "expand",
ncol = 2)
# -
plot_history(history)
# +
import numpy as np
pred_y = model_final.predict(test_generator)
# -
pred_y = pred_y > 0.5
pred_y[0]
true_y = test_df['class']
true_y[0]
# +
bt = [0] * 20
bf = [0] * 20
fp = [0] * 20
fn = [0] * 20
for t_y, p_y in zip(true_y, pred_y):
p_y = np.where(p_y)[0]
for i in range(20):
if i in t_y and i in p_y:
bt[i] += 1
elif i not in t_y and i not in p_y:
bf[i] += 1
elif i in t_y and i not in p_y:
fn[i] += 1
else:
fp[i] += 1
print("true positive:", bt, "\ntrue negative:", bf, "\nfalse positive:", fp, "\nfalse negative:", fn)
# -
model_final.evaluate(test_generator)
# +
import seaborn as sns
f = plt.figure(figsize = (7,50))
sum_ax = f.add_subplot(6, 1, 1)
sns.heatmap([[sum(bt), sum(fp)],
[sum(fn), sum(bf)]],
annot = True,
cbar = False,
xticklabels = ["True", "False"],
yticklabels = ["True", "False"],
ax = sum_ax,
fmt='d')
sum_ax.set_title('All Bits')
sum_ax.set_ylabel("Predicted")
sum_ax.set_xlabel("Actual")
for i in range(20):
ax = f.add_subplot(12, 2, i + 5)
sns.heatmap([[bt[i], fp[i]],
[fn[i], bf[i]]],
annot = True,
cbar = False,
xticklabels = ["True", "False"],
yticklabels = ["True", "False"],
ax = ax,
fmt='d')
ax.set_title('Bit {}'.format(i))
ax.set_ylabel("Predicted")
ax.set_xlabel("Actual")
# -
# ## Save model
model_final.save('bar_cat_model_943')
| Chapter3/Bar Categorization/Bar Categorization-VGG19-nolast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Spark tutorial
#
# ## Setup
# Import the `.ipynb` file into Jupyter and open this as usual. MAKE SURE YOU ARE ON THE VPN IF YOU USE YOUR LAPTOP!
#
# First, we will install a package which will allow us to access Spark with the Python API: `findspark`. Execute the code in the cell below to install `findspark`.
#
# + language="bash"
# pip install findspark
# -
# With this new package, when we run `findspark.init()`, it creates a Pyspark environment we can run Spark applications in, as follows.
# +
import findspark
findspark.init('/home/comp6235/spark-2.4.0-bin-hadoop2.7')
import pyspark
from pyspark import SparkContext, SparkConf
# -
# We created a `SparkContext` above, which represents a connecting to a computing cluster. If you start the PySprak shell, it will be created automatically as `sc`, but here we need to create it manually.
#
# With the `SparkContext` object, we can create RDDs. It is also possible to configure the `SparkContext` with a `SparkConf` object passed as a paprameter:
# +
conf = SparkConf().setMaster('local').setAppName('My application')
sc = SparkContext(conf=conf)
sc
# -
# The output of `sc` gives you a link to Spark UI of your context. You can use this to visualise the jobs you're running.
#
# Note that you can only create one of these `SparkContext` objects at a time. If you wish to run it again, you will need to call `sc.stop()` to get rid of the old one and to start it running again.
#
# With our `SparkContext`, we can now start creating an RDD. There are two ways of creating an RDD: through an internal object or class, or an external dataset. For this, we are going to use a text file containing the text of the complete Sherlock Holmes stories. I have left a copy of the text at http://edshare.soton.ac.uk/19180/1/holmes_complete.txt. Download a copy of ths file into the same directory as your tutorial.
#
# We call the function `sc.textFile`, and this returns an RDD:
rdd = sc.textFile('holmes_complete.txt')
rdd
# A partition is a logical chunk of a large distributed data set. This represents the amount of parallel tasks you are going to be using. By default it goes to one per core, but you can set it by adding a parameter to the RDD object you created. We can see the amount of partitions we have by running the cell below:
rdd.getNumPartitions()
# ## Actions and transformations
#
# You can see the output refers to an RDD object, but nothing has happened yet, because the RDD uses lazy processing and does not do anything until it has to. By contrast, if we apply an action like `count()` or `reduce()` then it will return an object rather than an RDD which is illustrative that some processing has occurred. We will call an action `count()` here which returns the amount of lines in the RDD:
rdd.count()
# We can see how the transformation returns an RDD by executing the next cell. In this we are using the `filter()` transformation. The filter is using Python `lambda` expressions to identify any instances of the name "_Holmes_" inside the text file. A lambda expression is like an anonymous function, it would also be possible to define a function and pass that as a parameter instead.
holmes = rdd.filter(lambda line: "Holmes" in line)
print(holmes.count())
# If you need a bit more of an introduction to lambda functions, check out http://www.diveintopython.net/power_of_introspection/lambda_functions.html. These are very useful in Spark, so it's worth knowing how they work. In the next cell, try and get the equivalent outcome by defining a function and passing it as a parameter to the `filter` function. Call the output `holmes_2`.
# +
# YOUR CODE HERE
holmes_2 = rdd.filter(lambda line: "Holmes" in line)
print(holmes_2.count())
def filter2(rdd):
for
# -
# This returns a new RDD object, and does not mutate the initial RDD object either - despite referring to a filter operation on it. The following will return the first line of the document (which includes the name "Holmes". Note that it will not operate on `rdd`, but rather will operate on `holmes`, as can be seen from executing the following cell.
#
# Look at the output below, and notice how the amount of lines on the `rdd` object remains the same as above in contrast to the amount of lines in `holmes`.
print(rdd.count())
print(holmes.count())
# If we wish, we can perform set operations on the data. We will demonstrate this by creating another RDD object, containing only lines which have the name "_Watson_" in them, and then get the intersection and union of the two.
#
# Create the `watson` RDD object, and then call the `first()` action to return the first line, and then call the `count()` to see how many lines there are in the book which satisfy that filter.
# YOUR CODE HERE
# New RDD and show the first line here
watson_rdd= sc.textFile('holmes_complete.txt')
watson_rdd
# YOUR CODE HERE
# How many lines in the Watson RDD?
watson= watson_rdd.filter(lambda line: "Watson" in line)
watson.first()
print(watson.count())
# We can create a new RDD by calling the intersection or the union of the two:
holmes_and_watson = holmes.intersection(watson)
holmes_and_watson
holmes_and_watson.count()
# Try and find the union of lines with the name "Holmes" and the name "Watson":
# YOUR CODE HERE
holmes_and_watson_common = holmes_and_watson.filter(lambda line: "Holmes" and "Watson" in line)
# YOUR CODE HERE
# How many lines in the file?
holmes_and_watson_common.count()
holmes_or_watson = holmes.union(watson)
holmes_or_watson
holmes_or_watson.count()
# ## Lineage
#
# We discussed lineage: the process by which RDDs are connected. By creating these different RDDs, we are able to get an idea of the lineage of the We can get an idea of the lineage of an RDD object by calling the function `toDebugString()`. Call this on the union of the `holmes` and `watson` RDDS:
holmes_or_watson.toDebugString()
# holmes_and_watson.toDebugString()
# Alternatively, there is a UI listening on port `4040` as part of Spark (see above where we printed the output of `sc`). If you open that in a new tab, and click on the "jobs" you can click through to see the visualisation of the DAG which is how the lineage is represented.
# ## MapReduce
#
# Spark started off as a batch processing library, although one more efficient than Hadoop, and as such it is possible to run MapReduce tasks. This is done on the RDD object through the `map()` and `flatMap()` functions, (which are transformations) and the `reduce` function which is an action.
#
# Both `map()` and `flatMap()` transformation take a function as an argument, and applies it to each element in the RDD. `map()` returns a new RDD the same size as the original RDD, whereas `flatMap()` transforms the RDD of length _n_ into _n_ collections, and flattens these into a single RDD of results. See https://stackoverflow.com/questions/22350722/what-is-the-difference-between-map-and-flatmap-and-a-good-use-case-for-each for a good explanation.
#
# The typical "Hello, world" example, as last week, is that of word count. We will do a word count on the text we have in the `holmes_complete.txt` file, starting with the `rdd` variable we already have.
#
# It's just text this time, so we don't have to parse through a CSV file which should make it a bit easier. We will start with applying the `flatMap` to get the words tokenised. The `take(n)` function gives us a list of the first `n` items in the RDD which can be useful for debugging.
word_split = rdd.flatMap(lambda line: line.split(' '))
word_split.take(10)
# Using `map` on `word_split`, create a new RDD which outputs the words as a series of tuples of length two, each containing the word and the number 1. Then, output the first 30 pairs.
# YOUR CODE HERE
word_count_1=word_split.map(lambda line:(line,1))
word_count_1.take(30)
# Now we have the mapped part of the MapReduce function, the next step is to get the `reduce()` action. More specifically, we need to run the `reduceByKey()`. From the documentation:
# <blockquote>
# When called on a dataset of <code>(K, V)</code> pairs, returns a dataset of <code>(K, V)</code> pairs where the values for each key are aggregated using the given reduce function func, which must be of type <code>(K, V) => V</code>. Like in </code>groupByKey</code>, the number of reduce tasks is configurable through an optional second argument.
# </blockquote>
reduced_words = word_count_1.reduceByKey(lambda a,b: a + b)
reduced_words.take(10)
# Although we can see that there is a wordcount, it would be more useful for it to be done in order. Here, we use the `sortBy` function, which takes a function and operates on the second part of the tuple (the value) in order to sort in descending order by amount of occurrences.
reduced_words.sortBy(lambda a: -a[1]).take(10)
# ## Extra challenge
#
# * You will notice that there are many common words in the output which are not very useful for analysis. Try and figure out a way of getting rid of these words from the output. They are known as "stop words"
# * Clear the whitespace from the document
#
# ### Streaming
#
# If you are interested in seeing how the streaming works, check out the following link for streaming on Twitter data https://www.linkedin.com/pulse/apache-spark-streaming-twitter-python-laurent-weichberger
| Lab6_Spark/spark_ownpractice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <b>GOAL</b>: Visualize and Prepare features to feed into ML Models
# # Table of contents | Summary
# 1. [ <i>Import</i> Libraries ](#libs)
# 2. [<i>Read in</i> Training and Testing Data via (csv) & <i>Merge</i>](#Read-In-Data)
# 3. [<i>Load</i> Helper Functions](#helpers)
# 4. [<i>Understanding the Data</i>: Missing Values, Shape, Duplicates](#understanding)
# 1. [CHECK for Duplicates](#duplicates)
# 2. [DROP 'jobId' as the unique identifier](#dropid)
# 5. [<i>Split Train df</i> into Numerical and Categorical df's](#splitdf)
# 1. [UNDERSTAND Distribution of Numerical Data](#numdist)
# 6. [Understanding <i>Categorical</i> Features vs. Salary](#catvsalary)
# 1. [One Hot Encode (cat df) to feed into ML model](#ohe)
# 7. [Understanding <i>Numerical</i> Features vs. Salary](#numvsalary)
# 8. [Find outliers in the Target variable: LOW and HIGH](#outlierhead)
# 1. [UPPER outliers](#upperout)
# 2. [LOWER outliers](#lowerout)
# 3. [DROP outliers from X_full dataframe](#dropout)
# 11. [<i>Create</i> Train and Target tables w/ train_test_split](#train)
# 1. [<i>Create</i> Validation Set from Train data](#val)
# 12. [<i>Pickle</i> Train/ Validation/ and Test Data](#pickle)
# ## 1. <i>Import</i> Libraries <a name = "libs"></a>
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
import pickle
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
__author__ = "<NAME>"
__email__ = <EMAIL>"
# -
# ## 2. <i>Read in</i> Training and Testing Data via (csv) & <i>Merge</i><a name="Read-In-Data"></a>
# +
train_features = pd.read_csv('train_features.csv')
test_features = pd.read_csv('test_features.csv')
train_sal = pd.read_csv("train_salaries.csv") #contains only jobID and Salary columns -> merge with Train_Features
# -
#Merge Training Datasets on 'jobId'
train_full = train_features.merge(train_sal, on = 'jobId')
train_full.head()
# Understand that **"jobId" & "companyId"** are ***unique identifiers*** for each survey respondant and company
# ## 3. <i>Load</i> Helper Functions <a name = "helpers"></a>
# +
def feature_order(feature, ascending = True):
''' Return s a list of the feature from LOW -> HIGH'''
grouped_df = train_full.groupby(feature)['salary'].mean().sort_values().reset_index()
if ascending == False:
sorted_df = grouped_df.sort_values(by = 'salary', ascending = False).reset_index()
return sorted_df[feature].tolist()
else:
sorted_df = grouped_df.sort_values(by = 'salary', ascending = True).reset_index()
return sorted_df[feature].tolist()
def plot_viz(feature):
''' Plots Feature against Salary (Boxenplot)'''
plt.figure(figsize = (20,7))
plt.title(feature + ' vs. Salary')
plt.ylabel('Salary')
plt.xlabel(feature)
if feature == 'milesFromMetropolis':
return sns.boxenplot(x = feature, y = 'salary', data = train_full, order = feature_order(feature, False))
if feature == 'companyId':
plt.xticks(rotation = 90)
return sns.boxenplot(x = feature, y = 'salary', data = train_full, order = feature_order(feature))
def find_outliers(df, feature, lower = False, view = 10):
''' Returns the Top 10 lowest or highest outlier employees'''
describe_df = df[feature].describe().reset_index()
if lower == True:
q1 = describe_df.iloc[4][feature]
return df.loc[df[feature] < q1].sort_values(by= feature, ascending = True).head(view)
else:
q3 = describe_df.iloc[6][feature]
return df.loc[df[feature] > q3].sort_values(by= feature, ascending = False).head(view)
# -
# ## 4. <i>Understanding the Data</i>: Missing Values, Shape, Duplicates<a name ="understanding"></a>
train_full.info()
print("There are also NO missing values as shown in the Non-null column")
print('There are {} Train samples and {} Test samples'.format(train_full.shape[0], test_features.shape[0]))
# ### A. CHECK for Duplicates <a name = "duplicates"></a>
train_full[train_full.duplicated()]
# There are no Duplicate Values
# ### B. DROP 'jobId' as the unique identifier <a name = "dropid"></a>
train_full = train_full.drop('jobId', axis = 1)
train_full.head()
figure = plt.figure(figsize = (13,7))
num_corr = train_full.corr()
mask = np.triu(np.ones_like(num_corr, dtype=bool))
sns.heatmap(data= num_corr, annot = True, cmap = 'Blues', mask = mask)
plt.xticks(rotation = 45)
plt.show()
# ## 5. <i>Split Train df</i> into Numerical and Categorical df's<a name = "splitdf"></a>
cat = train_full.select_dtypes(include= 'object')
cat.head()
numeric = train_full.select_dtypes(exclude= 'object')
numeric.head()
# ### A. UNDERSTAND Distribution of Numerical Data<a name = "numdist"></a>
for col in numeric.columns.tolist():
numeric[col].hist(alpha = 0.3)
plt.legend([col for col in numeric.columns])
plt.title('# of Employees per Years of Experience')
plt.ylabel('Counts')
plt.xlabel('Years Experience')
numeric['yearsExperience'].hist(alpha = 0.3)
# **Summary**: Salary follows a relatively ***Normal Gaussian Distribution***.
# - yearsExperience and milesFromMetropolis are **uniformly distributed** and relatively **symmetrical**
# ## 6. Understanding <i>Categorical</i> Features vs. Salary <a name = "catvsalary"></a>
plot_viz('companyId')
# **Summary**: Salary tends to be relatively ***constant*** across different companies.
plot_viz('jobType')
# **Summary**: Salary ***increases*** as one gets a better jobType.
plot_viz('degree')
# **Summary**: Salary ***increases*** the more advanced a degree one has.
plot_viz('major')
# **Summary**: Salary is ***higher on average*** for Math, Business, and Engineering majors.
plot_viz('industry')
# **Summary**: Salary is ***higher on average*** for employees in Web, Finance, and Oil industries.
# ## 7. Understanding <i>Numerical</i> Features vs. Salary <a name = "numvsalary"></a>
# Find the correlation of the numerical features in the Training Data
train_full.corr()
plot_viz('yearsExperience')
fig, ax0 = plt.subplots(figsize= (20,7))
ax0 = sns.scatterplot(x = 'yearsExperience', y = 'salary', data= train_full, hue = 'jobType',
palette= 'Set2', alpha = 0.4, hue_order= feature_order('jobType',ascending = False))
ax1 = ax0.twiny()
ax1 = sns.lineplot(x = 'yearsExperience', y = 'salary', data= train_full,
hue = 'jobType', palette= 'Set2', alpha = 0.6, legend= False)
# **Summary**: YearsExperience & jobType have a ***positive*** correlations with salary: Salary **increases** with more experience and a better title in the organization.
plot_viz('milesFromMetropolis')
# **Summary**: milesFromMetropolis has a ***negative*** correlation with salary: Employees get paid less the more miles they travel to work. <t>This trend is also consistent with the *correlation table* above.
plt.figure(figsize = (15,7))
plt.title('Number of different jobTypes in the Dataset')
plt.ylabel('Counts')
train_full.groupby('jobType')['salary'].count().sort_values(ascending = True).plot(kind = 'bar')
# **Summary**: The Distribution of jobTypes is relatively *equal*
# ## 8. Find outliers in the Target variable: LOW and HIGH<a name = "outlierhead"></a>
# ### UPPER outliers<a name = "upperout"></a>
upper_outliers = find_outliers(train_full, feature = 'salary', lower = False)
upper_outliers
# **Summary**: Nothing totally unusual about the upper-bound outliers. There are some high paying salaries for individuals with only 'Bachelors' degrees but often they are CEOs and so this inflated salary is justified.
#
# With more time we would learn more information about these unique employees with lower education and a higher position in the company
# ### LOWER outliers<a name = "lowerout"></a>
#LOW
lower_outliers = find_outliers(train_full, 'salary', lower = True, view = 5)
lower_outliers
# **Summary**: Suspicious to find salaries = 0, higher levels positions such as "Vice Presidents" and "CTOs" should earn some money.
# Similarly we assume there are no unpaid internships for Junior jobTypes. But in reality we would check with the data source or data collection team to inquire more about these outliers.
# ### DROP outliers from X_full dataframet <a name = "dropout"></a>
outlier_indices = lower_outliers.reset_index().iloc[:5]['index'] #locate outliers
''' Drop outliers from Full Train and Test Data'''
X_full = train_full.drop(outlier_indices)
test_features = test_features.drop(outlier_indices)
# ## 9. <i>Create</i> Train and Target tables w/ train_test_split <a name = "train"></a>
y_full = X_full['salary']
X_Full = X_full.drop('salary', axis = 1)
X_train, X_test, y_train, y_test = train_test_split(X_Full, y_full, shuffle = True,
test_size = 0.2, random_state = 42)
# **Result:** Allocates 20% of the data for testing while using the remaining 80% for training the models
# ### Create Validation Set from Train data <a name = "val"></a>
# - Creates a 60k/20k/20k split for Train/Validation/Test data
X_train, X_valid, y_train, y_valid = train_test_split(X_train, y_train, test_size=0.25, random_state=42)
# ## 10. <i>Pickle</i> Train/ Validation/ and Test Data<a name = "pickle"></a>
'''Pickle Training Data'''
with open('../salary prediction data/Pickle Data/TRAIN_X.pkl', 'wb') as temp0:
pickle.dump(X_train, temp0)
with open('../salary prediction data/Pickle Data/TRAIN_Y.pkl', 'wb') as temp1:
pickle.dump(y_train, temp1)
'''Pickle Validation Data'''
with open('../salary prediction data/Pickle Data/VALID_X.pkl', 'wb') as temp2:
pickle.dump(X_valid, temp2)
with open('../salary prediction data/Pickle Data/VALID_y.pkl', 'wb') as temp3:
pickle.dump(y_valid, temp3)
'''Pickle Test Data'''
with open('../salary prediction data/Pickle Data/TEST_X.pkl', 'wb') as temp4:
pickle.dump(X_test, temp4)
with open('../salary prediction data/Pickle Data/TEST_Y.pkl', 'wb') as temp5:
pickle.dump(y_test, temp5)
# **Foreward:** Save the split data for training, validating, and testing the models in the 'Model Training' notebook
| Exploratory Data Analysis (EDA).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Essex County, MA Public Schools DOE + Census Bureau data
# Merging datasets from Massachusetts Department of Education (DOE) and U.S. Census Bureau data for school districts and towns in Essex County
# Imports
#Import libraries
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# import packages
from plotnine import *
# +
# # %%R
# # My commonly used R imports
# require('ggplot2')
# require('readr')
# require('extrafont')
# require('dplyr')
# require('reshape2')
# # require('theme538')
# -
pd.set_option("display.max_columns", None)
# ### Importing School and District Profiles from Mass. Department of Education (DOE)
# 2020-21 Class Size by Race/Ethnicity Report
df = pd.read_excel("ma_doe_files//ClassSizebyRaceEthnicity.xlsx")
df.columns = df.columns.str.lower().str.replace(" ", "_")
df.head(100)
# ###### 2021 MCAS Achievement Results
df_mcas = pd.read_excel("ma_doe_files//mcas.xlsx")
df_mcas.columns = df_mcas.columns.str.lower().str.replace(" ", "_")
df_mcas
# +
# note: MCAs scores double up districts because there are multi subjects tested per district, so it's not included in the full dataframe
# df = pd.merge(df, df_mcas, on='district_code', how='outer')
# df
# -
# ###### 2020-21 Advanced Placement Performance - All Subjects - All Students
df_ap_performance = pd.read_excel("ma_doe_files//ap_performance.xlsx")
df_ap_performance.columns = df_ap_performance.columns.str.lower().str.replace(" ", "_")
df_ap_performance
df = pd.merge(df, df_ap_performance, on='district_code', how='outer')
df
# ###### 2020-21 Advanced Placement Participation - All Subjects - All Students
df_ap_participation = pd.read_excel("ma_doe_files//ap_participation.xlsx")
df_ap_participation.columns = df_ap_participation.columns.str.lower().str.replace(" ", "_")
df_ap_participation
df = pd.merge(df, df_ap_participation, on='district_code', how='outer')
df
# ###### 2019-20 HS Graduates Attending Institutions of Higher Education (District) - All Students - All Colleges and Universities
df_grads_college = pd.read_excel("ma_doe_files//Gradsattendingcollege.xlsx")
df_grads_college.columns = df_grads_college.columns.str.lower().str.replace(" ", "_")
df_grads_college
df = pd.merge(df, df_grads_college, on='district_code', how='outer')
df
# ###### 2020-21 SAT Performance Report - All Students
df_sats = pd.read_excel("ma_doe_files//sat_performance.xlsx")
df_sats.columns = df_sats.columns.str.lower().str.replace(" ", "_")
df_sats
df = pd.merge(df, df_sats, on='district_code', how='outer')
df
# ###### 2020 Graduation Rate (District) All Students 4-Year Graduation Rate
df_grad_rates = pd.read_excel("ma_doe_files//gradrates.xlsx")
df_grad_rates.columns = df_grad_rates.columns.str.lower().str.replace(" ", "_")
df_grad_rates
df = pd.merge(df, df_grad_rates, on='district_code', how='outer')
df
# ###### Preliminary FY20 Per Pupil Expenditures by School (imported but not merged with larger dataset)
# Note: Data gathered by school, not merged with dataset but interesting to look at
df_ppx = pd.read_excel("ma_doe_files//preliminary-school-ppx-1tab_edited.xlsx")
df_ppx.columns = df_ppx.columns.str.lower().str.replace(" ", "_")
df_ppx.head(10)
# +
# saving a copy of the merged dataset
# df.to_csv('output_MA_DOE_profiles2.csv', index = False)
# -
df = pd.read_csv("processed//MA_DOE_profiles_edited.csv", encoding="utf-8")
df
df[df.district_name_w_charters.str.contains("Manchester")]
# ### Combining DOE data with town, school district, and high school name
# ##### Essex County MA Public High Schools List — compiled from Mass. Secretary of State's list
# MA: A Listing of Counties and the Cities and Towns Within
# source: https://www.sec.state.ma.us/cis/cisctlist/ctlistcoun.htm
df_essex_county = pd.read_excel("handmade//Essex_county_public_hs2.xlsx")
df_essex_county.columns = df_essex_county.columns.str.lower().str.replace(" ", "_")
df_essex_county.head(5)
df = pd.merge(df, df_essex_county, on='district_name_x', how='inner')
df.head(5)
df = df.rename(columns={'towns_essex_county_ma': 'towns'})
df.sort_values(by = 'towns', ascending=True).head(5)
# Checking + Cleaning dataset
df.head(3)
# +
# df.to_csv('output_MA_DOE_Essex_county_profiles.csv', index = False)
# -
# ##### Safety Check: Using the backup merged dataset as input in case something goes wrong
df = pd.read_csv("processed//input_MA_DOE_Essex_county_profiles.csv", encoding="utf-8")
df.head()
# ##### Merging U.S. Census bureau data for Essex County
# +
# Census.gov Massachusetts Basic Facts
# data source: https://www.census.gov/quickfacts/fact/table/swampscotttownessexcountymassachusetts/INC110219
# +
# Note: was not able to download data for the towns: Essex, Nahant, Wenham, West Newbury,
# however these towns are in regional school districts so I substituted census info for another town in the district
# Wenham = Hamilton, West Newbury = Groveland, Essex = Manchester-by-the-sea, Nahant = Swampscott
# -
df_ma_essex_county_census = pd.read_csv("ma_ec_census//MA_essex_county_census_info4.csv", encoding="utf-8")
df_ma_essex_county_census.head(5)
df_ma_essex_county_census.columns = df_ma_essex_county_census.columns.str.lower().str.replace(" ", "_")
df_ma_essex_county_census.head(3)
# ##### Cleaning up the data: renaming and adding new columns
df_ma_essex_county_census = df_ma_essex_county_census.rename(columns={'median_household_income_(in_2019_dollars),_2015-2019': 'median_household_income'})
df_ma_essex_county_census['median_household_income'] = df_ma_essex_county_census['median_household_income'].astype(int)
df = pd.merge(df, df_ma_essex_county_census, on='towns', how='outer')
df = df.rename(columns={'open_data_channel_type': 'channel'})
df.columns = df.columns.str.replace(",", "")
df.head(3)
df.columns = df.columns.str.replace("%", "pct")
df.columns = df.columns.str.replace("#", "num")
df = df.rename(columns={'african_american_%': 'scl_african_american_pct'})
df = df.rename(columns={'asian_%': 'scl_asian_pct'})
df = df.rename(columns={'hispanic_%': 'scl_hispanic_pct'})
df = df.rename(columns={'white_%': 'scl_white_pct'})
df = df.rename(columns={'native_american_%': 'scl_native_american_pct'})
df = df.rename(columns={'native_hawaiian,_pacific_islander_%%': 'scl_native_hawaiian_pacific_islander_pct'})
df = df.rename(columns={'multi-race,_non-hispanic_%': 'scl_multi_race_non_hispanic_pct'})
df = df.rename(columns={'native_hawaiian_pacific_islander_%': 'scl_native_hawaiian_pacific_islander_pct'})
df = df.rename(columns={'multi-race_non-hispanic_%': 'scl_multi_race_non_hispanic_pct'})
df = df.rename(columns={'%_score_1-2_x': 'ap_one_to_two_pct'})
df = df.rename(columns={'%_score_3-5_x': 'ap_three_to_five_pct'})
df = df.rename(columns={'attending_coll./univ._(%)': 'attending_coll/uni_pct'})
df.head()
df.shape
# +
# save merged full dataset
# df.to_csv('output_MA_DOE_Essex_county_profiles_w_census.csv', index = False)
# -
df_ma_essex_county_census.head(5)
df_ma_essex_county_census_subset = df_ma_essex_county_census[['towns', 'fact', 'median_household_income']]
df_ma_essex_county_census_subset.head(5)
df_ma_essex_county_census_subset.sort_values(by = 'median_household_income', ascending=False)
| ec-school-data/essex_county_public_schools_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ky6t2Ic6-VN-" colab_type="text"
# # Internal resistance of a battery
#
# We use the simple AAA battery from the remote for projector/presentations. The voltage without load (equal to e.m.f. $\varepsilon$) is $V = 1.498 \text{V}$. If we use a resistor with $R = 1007 \Omega$ parallel as load, the voltage drops to $V_{ext} = 1.462 \text{V}$.
#
# ## Calculate the internal resistance
#
# We use the formula from the booklet 5.3:
#
# - $\varepsilon = I(R + r)$
#
# Here $\varepsilon$ is the e.m.f. of the cell, $r$ the internal resistance and $R$ the external resistance or load resistor. First we have to find the current in this circuit and use the values for the load resistor in combination with Ohm's law:
#
# - $R = \frac{V}{I}$
#
# And with the given $V_{ext} = 1.462 \text{V}$ and $R = 1007 \Omega$ we get
# - $I = \frac{V_{ext}}{R} = \frac{1.462}{1007} = 1.452 \text{mA}$
#
# The internal voltage drop $V_{int}$ over the internal resistance is calculated, using Kirchoff's rule:
#
# - $\varepsilon = V_{int} + V_{ext} ~~\longrightarrow ~~ V_{int} = \varepsilon - V_{ext} = 1.498 - 1.462 = 0.036 \text{V}$
#
# Finally the internal resistance is calculated, using Ohm's law:
#
# - $r = \frac{V_{int}}{I} = \frac{0.036}{0.001452} = 24.8 \Omega$
#
# __Answer:__ The internal resistance of the AAA battery is $24.8 \Omega$.
# + id="9Ax_fQOB-Tre" colab_type="code" outputId="8715f85d-4d07-4df3-ac8f-b7fd0862ed44" colab={"base_uri": "https://localhost:8080/", "height": 34}
emf = 1.498 # emf of the battery, in Volt
V_ext = 1.462 # external voltage of battery under load of 1 kOhm resistor, in Volt
R = 1007 # resistance of load, in Ohm
I = V_ext / R
V_int = emf - V_ext
r = V_int / I
print("The internal resistance is {:.1f} Ohm.".format(r))
| internal-resistance-battery/internal_resistance_battery_updated.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import cx_Oracle
import os
os.environ["NLS_LANG"] = "SIMPLIFIED CHINESE_CHINA.UTF8"
# 连接数据库,使用详细信息连接
conn = cx_Oracle.connect("system/xxx@192.168.1.xxx/xxx")
# 获取游标
cursor = conn.cursor()
# 执行SQL
recs = cursor.execute("select sysdate from dual")
# 提取游标
recs.fetchone()
# 获取游标所有记录
recs.fetchall()
# 关闭游标
cursor.close()
# 关闭连接
conn.close()
# +
# 使用tnsname连接,登录为SYSDBA
# conn2 = cx_Oracle.connect("sys/oracle@orabiz217", mode=cx_Oracle.SYSDBA)
# conn2.close()
# -
# 使用tnsname连接
conn3 = cx_Oracle.connect("system/xxx@xxx")
cur3 = conn3.cursor()
# 调用函数,指明返回值类型
tradingday = cur3.callfunc("xxx", cx_Oracle.STRING)
# 具体调用过程请见参考文献[1]
print tradingday
# 调用函数,指明返回值类型及入参
last_tradingday = cur3.callfunc("xxx", cx_Oracle.STRING, [tradingday])
# 具体调用过程请见参考文献[1]
print last_tradingday
# 调用存储过程,指定输入输出参数
# 初始化输入输出参数
x = cur3.callproc("xxx",
[xxx])
# 具体调用过程请见参考文献[1]
print x
# 直接打印中文没有问题,最好在中文字符串之前声明是Unicode编码
print "中文"
print u"中文"
# list中的中文字符串会以字符编码的形式输出
x = [1, 2, 3, 4, 5, 6, 7, "成功"]
x
print x
# '\xb3\xc9\xb9\xa6'
x[7]
# 可以使用如下形式进行转码,实际输出乱码
print x[7].decode('gbk').encode("utf-8")
# 解码为GBK,实际输出乱码
print x[7].decode('gbk')
s = x[7].decode('gbk')
# 输出Unicode编码
s
# 输出乱码
print s
# 输出中文
print x[7]
# 解码正确,说明list默认使用utf-8编码
su = x[7].decode("utf-8")
# 输出中文
print su
# # 参考文献
# 1. http://blog.csdn.net/my2010sam/article/details/20724001 cx_Oracle操作Oracle数据库
| Python Oracle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def divide(x,y):
if y == 0:
raise ZeroDivisionError('division by zero')
quotient = 0
power = 32
yPower = y << power
remainder = x
while remainder >= y:
while yPower > remainder:
yPower >>= 1
power -= 1
quotient += 1 << power
remainder -= yPower
return quotient
# time complexity of O(N)
# shaves off the remainder, no decimals
# -
divide(6,3)
divide(10,3)
| Daily/division_without_div.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="_NliWKN6rBM2"
# ##### Copyright 2018 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="wMbda6-vqkeG"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="SwuyEwn-b0YH"
# # Guided ES Demo
#
# This is a fully self-contained notebook that reproduces the toy example in Fig.1 of the guided evolutionary strategies paper.
#
# The main code is in the 'Algorithms' section below.
#
# Contact: <EMAIL>
#
# Date: 6/22/18
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="w_OmAdfRPZoz" slideshow={"slide_type": "slide"}
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_probability as tfp
# -
print(f'tensorflow version: {tf.__version__}')
print(f'tensorflow_probability version: {tfp.__version__}')
# + [markdown] colab_type="text" id="7Su4I_YWlGAb"
# ## Helper functions
# + [markdown] colab_type="text" id="X-cRO-ncyB_u"
# ### Antithetic sampler
#
# Creates custom getters for perturbing variables.
#
# These are used to evaluate f(x + epsilon), where epsilon is some perturbation applied to the parameters, x.
#
# This also stores the sampled noise (epsilon) in a dictionary, since we need to reuse the noise for the negative sample, when we want to compute f(x - epsilon). (note: this is where the name `antithetic` comes from)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="iV_z02iDx5Xb"
class AntitheticSampler(object):
def __init__(self, distributions):
"""Antithetic perturbations.
Generates samples eta, and two custom getters that return
(x + eta) and (x - eta)
for a variable x.
This is used to evaluate a loss at perturbed parameter values, e.g.:
[f(x+eta), f(x-eta)]
"""
# stores the sampled noise
self.perturbations = {}
# store the distributions
self.distributions = distributions
def pos_getter(self, getter, name, *args, **kwargs):
"""Custom getter for positive perturbation"""
# get the variable
variable = getter(name, *args, **kwargs)
# check if we have pulled this variable before
if name not in self.perturbations:
# generate a noise sample and store it
self.perturbations[name] = self.distributions[name].sample()
# return the perturbed variable
return variable + tf.reshape(self.perturbations[name], variable.shape)
def neg_getter(self, getter, name, *args, **kwargs):
"""Custom getter for negative perturbation"""
# get the variable
variable = getter(name, *args, **kwargs)
# check if we have pulled this variable before
if name not in self.perturbations:
# generate a noise sample and store it
self.perturbations[name] = self.distributions[name].sample(shape=variable.shape)
# return the perturbed variable
return variable - tf.reshape(self.perturbations[name], variable.shape)
# + [markdown] colab_type="text" id="bYheIglf2clB"
# ### Noise distributions
#
# We draw perturbations of parameters from either a diagonal covariance (the standard evolutionary strategies algorithm), or from a diagonal plus low rank covariance (guided ES).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="pLYwc0ol2bL9"
mvn_diag = tfp.distributions.MultivariateNormalDiag
mvn_lowrank = tfp.distributions.MultivariateNormalDiagPlusLowRank
# + [markdown] colab_type="text" id="HKQinU4RT6md"
# ## Algorithms
# + [markdown] colab_type="text" id="2NRbt7DHT8S1"
# ### Gradient descent
#
# As a baseline, we will compare against running gradient descent directly on the biased gradients.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="CCqaG7bgT-Tc"
def gradient_descent(loss_fn, grads_and_vars):
return grads_and_vars
# + [markdown] colab_type="text" id="7oRZ6-lFlq3Q"
# ### Evolutionary strategies
#
# To compute descent directions using evolutionary strategies, we will use the antithetic sampler defined above.
#
# This will let us perturb model parameters centered on the current iterate.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="LeIL5S017KLv"
def evostrat_update(loss_fn, dists, grads_and_vars, beta, sigma):
"""Function to compute the evolutionary strategies.
See the guided ES paper for details on the method.
Args:
loss_fn: function that builds the graph that computes the loss. loss_fn,
when called, returns a scalar loss tensor.
dists: dict mapping from variable names to distributions for perturbing
those variables.
grads_and_vars: list of (gradient, variable) tuples. The gradient and
variable are tensors of the same shape. The gradient may be biased (it
is not necessarily the gradient of the loss_fn).
beta: float, scale hyperparameter of the guided ES algorithm.
sigma: float, controls the overall std. dev. of the perturbation
distribution.
Returns:
updates_and_vars: a list of (update, variable) tuples contaniing the
estimated descent direction (update) and variable for each variable to
optimize. (This list will be passed to a tf.train.Optimizer instance).
"""
# build the antithetic sampler
anti = AntitheticSampler(dists)
# evaluate the loss at different parameters
with tf.variable_scope('', custom_getter=anti.pos_getter):
y_pos = loss_fn()
with tf.variable_scope('', custom_getter=anti.neg_getter):
y_neg = loss_fn()
# use these losses to compute the evolutionary strategies update
c = beta / (2 * sigma ** 2)
updates_and_vars = [
(c * tf.reshape(anti.perturbations[v.op.name], v.shape) * (y_pos - y_neg), v)
for _, v in grads_and_vars]
return updates_and_vars
# + [markdown] colab_type="text" id="JHkc2vsAUFMZ"
# ### Vanilla ES
#
# Vanilla ES is the standard evolutionary strategies algorithm. It uses a diagonal covariance matrix for perturbing parameters.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="gTb30i9-UES6"
def vanilla_es(loss_fn, grads_and_vars, sigma=0.1, beta=1.0):
def vardist(v):
n = v.shape[0]
scale_diag = (sigma / tf.sqrt(tf.cast(n, tf.float32))) * tf.ones(n)
return mvn_diag(scale_diag=scale_diag)
# build distributions
dists = {v.op.name: vardist(v) for _, v in grads_and_vars}
updates_and_vars = evostrat_update(loss_fn, dists, grads_and_vars, beta, sigma)
return updates_and_vars
# + [markdown] colab_type="text" id="O0Hi-L1iY0jb"
# ### Guided ES
#
# Guided ES is our proposed method. It uses a diagonal plus low-rank covariance matrix for drawing perturbations, where the low-rank subspace is spanned by the available gradient information.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="saQTW0-BY2Hj"
def guided_es(loss_fn, grads_and_vars, sigma=0.1, alpha=0.5, beta=1.0):
def vardist(grad, variable):
"""Builds the sampling distribution for the given variable."""
n = tf.cast(variable.shape[0], tf.float32)
k = 1
a = sigma * tf.sqrt(alpha / n)
c = sigma * tf.sqrt((1-alpha) / k)
b = tf.sqrt(a ** 2 + c ** 2) - a
scale_diag = a * tf.ones(tf.cast(n, tf.int32))
perturb_diag = b * tf.ones(1,)
perturb_factor, _ = tf.qr(grad)
return mvn_lowrank(scale_diag=scale_diag,
scale_perturb_factor=perturb_factor,
scale_perturb_diag=perturb_diag)
dists = {v.op.name: vardist(g, v) for g, v in grads_and_vars}
# antithetic getter
updates_and_vars = evostrat_update(loss_fn, dists, grads_and_vars, beta, sigma)
return updates_and_vars
# + [markdown] colab_type="text" id="oTnAPygL6yg7"
# ## Tasks
# + [markdown] colab_type="text" id="Wu3NR6Tk6zlh"
# ### Perturbed quadratic
#
# This is a toy problem where we explicitly add bias and variance to the gradient
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="OyFvnj_O1HwT"
def generate_problem(n, m, seed=None):
rs = np.random.RandomState(seed=seed)
# sample a random problem
A = rs.randn(m, n)
b = rs.randn(m, 1)
grad_bias = rs.randn(n, 1)
return A, b, grad_bias
def perturbed_quadratic(n, m, problem_seed):
tf.reset_default_graph()
# generate problem
A_np, b_np, bias_np = generate_problem(n, m, seed=problem_seed)
A = tf.convert_to_tensor(A_np, dtype=tf.float32)
b = tf.convert_to_tensor(b_np, dtype=tf.float32)
# sample gradient bias and noise
grad_bias = 1.0 * tf.nn.l2_normalize(tf.convert_to_tensor(bias_np, dtype=tf.float32))
grad_noise = 1.5 * tf.nn.l2_normalize(tf.random_normal(shape=(n, 1)))
# compute loss
def loss_fn():
with tf.variable_scope('perturbed_quadratic', reuse=tf.AUTO_REUSE):
x = tf.get_variable('x', shape=(n, 1), initializer=tf.zeros_initializer)
resid = tf.matmul(A, x) - b
return 0.5*tf.norm(resid)**2 / float(m)
# compute perturbed gradient
with tf.variable_scope('perturbed_quadratic', reuse=tf.AUTO_REUSE):
x = tf.get_variable('x', shape=(n, 1), initializer=tf.zeros_initializer)
err = tf.matmul(tf.transpose(A), tf.matmul(A, x) - b) / float(m)
grad = err + (grad_bias + grad_noise) * tf.norm(err)
grads_and_vars = [(grad, x)]
return loss_fn, grads_and_vars
# + [markdown] colab_type="text" id="KnIJv_nhYI8p"
# ## Demo
# + [markdown] colab_type="text" id="JSAEHU44YKWx"
# ### Vanilla ES
#
# First, we run minimize the problem using vanilla evolutionary strategies.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="ZstfesU_1UQ0"
tf.reset_default_graph()
loss_fn, gav = perturbed_quadratic(1000, 2000, 2)
updates = vanilla_es(loss_fn, gav, sigma=0.1, beta=1.0)
opt = tf.train.GradientDescentOptimizer(0.2)
train_op = opt.apply_gradients(updates)
loss = loss_fn()
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} colab_type="code" executionInfo={"elapsed": 14006, "status": "ok", "timestamp": 1529723926412, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-BIjYFsOMLzc/AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="c-IQL75AVvzG" outputId="bd759007-423b-4cf7-e1b6-3b095a0b7d64"
# train
fobj = []
for k in range(10000):
f, _ = sess.run([loss, train_op])
fobj.append(f)
# store results for plotting
ves = np.array(fobj).copy()
sess.close()
# + [markdown] colab_type="text" id="foD-xrVrYNLw"
# ### Gradient descent
#
# Our next baseline is gradient descent, applied directly to the biased gradients.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 73} colab_type="code" executionInfo={"elapsed": 466, "status": "ok", "timestamp": 1529723992728, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-BIjYFsOMLzc/AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="HECrHTN5YS04" outputId="b593e567-5396-44d7-d84e-51f3b2afc2a0"
tf.reset_default_graph()
loss_fn, gav = perturbed_quadratic(1000, 2000, 2)
updates = gradient_descent(loss_fn, gav)
opt = tf.train.GradientDescentOptimizer(5e-3)
train_op = opt.apply_gradients(updates)
loss = loss_fn()
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} colab_type="code" executionInfo={"elapsed": 15736, "status": "ok", "timestamp": 1529724009245, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-BIjYFsOMLzc/AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="w3mJUgN7YS1P" outputId="ec5f0508-1499-4611-e5e7-90fc83d6d8e0"
# train
fobj = []
for k in range(10000):
f, _ = sess.run([loss, train_op])
fobj.append(f)
# store results for plotting
gd = np.array(fobj).copy()
sess.close()
# + [markdown] colab_type="text" id="qDhl6pjMZrCM"
# ### Guided ES
#
# Finally, we will run the same problem using the guided evolutionary strategies method.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 73} colab_type="code" executionInfo={"elapsed": 792, "status": "ok", "timestamp": 1529723943588, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-<KEY>AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="6gdZXM1vZuP_" outputId="c60112aa-f659-4929-ef33-d312eb3c93f9"
tf.reset_default_graph()
loss_fn, gav = perturbed_quadratic(1000, 2000, 2)
updates = guided_es(loss_fn, gav, sigma=0.1, alpha=0.5, beta=2.0)
opt = tf.train.GradientDescentOptimizer(0.2)
train_op = opt.apply_gradients(updates)
loss = loss_fn()
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 17} colab_type="code" executionInfo={"elapsed": 22555, "status": "ok", "timestamp": 1529723966176, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-BIjYFsOMLzc/AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="GglO7KDIZuQJ" outputId="156ffa99-bc23-4b42-e502-bc253afec563"
# train
fobj = []
for k in range(10000):
f, _ = sess.run([loss, train_op])
fobj.append(f)
# store results for plotting
ges = np.array(fobj).copy()
sess.close()
# + [markdown] colab_type="text" id="qEyLWBorYbe6"
# ### Plots
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 91} colab_type="code" executionInfo={"elapsed": 1400, "status": "ok", "timestamp": 1529723967604, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-BIjYFsOMLzc/AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="QrbAs1xbYPO2" outputId="b4790754-c476-4ce6-b7df-2dcbfcbbc2fc"
A, b, _ = generate_problem(1000, 2000, seed=2)
xstar = np.linalg.lstsq(A, b, rcond=None)[0]
f_opt = (0.5/2000) * np.linalg.norm(np.dot(A, xstar) - b) ** 2
# + [markdown] colab_type="text" id="DWJRegoBqHB_"
# As we see in the plot below, Guided ES combines the benefits of gradient descent (quick initial descent) and vanilla evolutionary strategies (converges on the true solution).
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 411} colab_type="code" executionInfo={"elapsed": 529, "status": "ok", "timestamp": 1529723977894, "user": {"displayName": "<NAME>", "photoUrl": "//lh6.googleusercontent.com/-BIjYFsOMLzc/AAAAAAAAAAI/AAAAAAAAADM/YvP5qhVshG8/s50-c-k-no/photo.jpg", "userId": "101494304217710169832"}, "user_tz": 420} id="9cax5P2RWIIn" outputId="e8a5b36e-a1b6-49dc-9f77-b7c7c3915fd8"
COLORS = {'ges': '#7570b3', 'ves': '#1b9e77', 'sgdm': '#d95f02'}
plt.figure(figsize=(8, 6))
plt.plot(ves - f_opt, color=COLORS['ves'], label='Vanilla ES')
plt.plot(gd - f_opt, color=COLORS['sgdm'], label='Grad. Descent')
plt.plot(ges - f_opt, color=COLORS['ges'], label='Guided ES')
plt.legend(fontsize=16, loc=0)
plt.xlabel('Iteration', fontsize=16)
plt.ylabel('Loss', fontsize=16)
plt.title('Demo of Guided Evolutionary Strategies', fontsize=16);
# -
| Guided_Evolutionary_Strategies_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:adventofcode]
# language: python
# name: conda-env-adventofcode-py
# ---
# # Evil solution
evil = lambda d,p=[0,0],o=0:sum(abs(a) for a in p) if len(d)==0 else evil(d[1:],p=[a+int(d[0][1:])*b for a,b in zip(p,[(1,0),(0,-1),(-1,0),(0,1)][[[1,3],[2,0],[3,1],[0,2]][o][0 if d[0][0]=='L' else 1]])],o=[[1,3],[2,0],[3,1],[0,2]][o][0 if d[0][0]=='L' else 1])
# # Test examples
evil(["R2", "L3"])
evil(["R2", "R2", "R2"])
evil(["R5", "L5", "R5", "R3"])
# # Process input
with open("inputs/day1a.txt") as fd:
print("Easter Bunny HQ is {} blocks away".format(evil([i.strip() for i in fd.read().split(',')])))
| 2016/jordi/Day 1 - Evil solution in one line.ipynb |