code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # A NumbaPro Mandelbrot Example
# This notebook was written by <NAME> based on code examples from Continuum Analytics that I modified somewhat. This is an example that demonstrates accelerating a Mandelbrot fractal computation using "CUDA Python" with NumbaPro.
#
# Let's start with a basic Python Mandelbrot set. We use a numpy array for the image and display it using pylab imshow.
import numpy as np
from pylab import imshow, show
from timeit import default_timer as timer
# The `mandel` function performs the Mandelbrot set calculation for a given (x,y) position on the imaginary plane. It returns the number of iterations before the computation "escapes".
def mandel(x, y, max_iters):
"""
Given the real and imaginary parts of a complex number,
determine if it is a candidate for membership in the Mandelbrot
set given a fixed number of iterations.
"""
c = complex(x, y)
z = 0.0j
for i in range(max_iters):
z = z*z + c
if (z.real*z.real + z.imag*z.imag) >= 4:
return i
return max_iters
# `create_fractal` iterates over all the pixels in the image, computing the complex coordinates from the pixel coordinates, and calls the `mandel` function at each pixel. The return value of `mandel` is used to color the pixel.
def create_fractal(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = mandel(real, imag, iters)
image[y, x] = color
# Next we create a 1024x1024 pixel image as a numpy array of bytes. We then call `create_fractal` with appropriate coordinates to fit the whole mandelbrot set.
# +
image = np.zeros((1024, 1536), dtype = np.uint8)
start = timer()
create_fractal(-2.0, 1.0, -1.0, 1.0, image, 20)
dt = timer() - start
print "Mandelbrot created in %f s" % dt
imshow(image)
show()
# -
# You can play with the coordinates to zoom in on different regions in the fractal.
create_fractal(-2.0, -1.7, -0.1, 0.1, image, 20)
imshow(image)
show()
# ## Faster Execution with Numba
# [Numba](https://github.com/numba/numba) is a Numpy-aware dynamic Python compiler based on the popular LLVM compiler infrastructure.
#
# Numba is an Open Source NumPy-aware optimizing compiler for Python sponsored by Continuum Analytics, Inc. It uses the remarkable compiler infrastructure to compile Python syntax to machine code. It is aware of NumPy arrays as typed memory regions and so can speed-up code using NumPy arrays, such as our Mandelbrot functions.
#
# The simplest way to use Numba is to decorate the functions you want to compile with @autojit. Numba will compile them for the CPU (if it can resolve the types used).
# +
from numba import autojit
@autojit
def mandel(x, y, max_iters):
"""
Given the real and imaginary parts of a complex number,
determine if it is a candidate for membership in the Mandelbrot
set given a fixed number of iterations.
"""
c = complex(x, y)
z = 0.0j
for i in range(max_iters):
z = z*z + c
if (z.real*z.real + z.imag*z.imag) >= 4:
return i
return max_iters
@autojit
def create_fractal(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = mandel(real, imag, iters)
image[y, x] = color
# -
# Let's run the `@autojit` code and see if it is faster.
# +
image = np.zeros((1024, 1536), dtype = np.uint8)
start = timer()
create_fractal(-2.0, 1.0, -1.0, 1.0, image, 20)
dt = timer() - start
print "Mandelbrot created in %f s" % dt
imshow(image)
show()
# -
# On my desktop computer, the time to compute the 1024x1024 mandelbrot set dropped from 6.92s down to 0.06s. That's a speedup of 115x! The reason this is so much faster is that Numba uses Numpy type information to convert the dynamic Python code into statically compiled machine code, which is many times faster to execute than dynamically typed, interpreted Python code.
# ## Even Bigger Speedups with CUDA Python
# [Anaconda](https://store.continuum.io/cshop/anaconda/), from Continuum Analytics, is a "completely free enterprise-ready Python distribution for large-scale data processing, predictive analytics, and scientific computing." [Anaconda Accelerate](https://store.continuum.io/cshop/accelerate/) is an add-on for Anaconda that includes the NumbaPro Python compiler.
#
# [NumbaPro](http://docs.continuum.io/numbapro/index.html) is an enhanced Numba that targets multi-core CPUs and GPUs directly from simple Python syntax, providing the performance of compiled parallel code with the productivity of the Python language.
#
# CUDA Python
# -----------
#
# In addition to various types of automatic vectorization and generalized Numpy Ufuncs, NumbaPro also enables developers to access the CUDA parallel programming model using Python syntax. With CUDA Python, you use parallelism explicitly just as in other CUDA languages such as CUDA C and CUDA Fortran.
#
# Let's write a CUDA version of our Python Mandelbrot set. We need to import `cuda` from the `numbapro` module. Then, we need to create a version of the mandel function compiled for the GPU. We can do this without any code duplication by calling `cuda.jit` on the function, providing it with the return type and the argument types, and specifying `device=True` to indicate that this is a function that will run on the GPU *device*.
#
# +
from numbapro import cuda
from numba import *
mandel_gpu = cuda.jit(restype=uint32, argtypes=[f8, f8, uint32], device=True)(mandel)
# -
# In CUDA, a *kernel* is a function that runs in parallel using many threads on the device. We can write a kernel version of our mandelbrot function by simply assuming that it will be run by a *grid* of threads. NumbaPro provides the familiar CUDA `threadIdx`, `blockIdx`, `blockDim` and `gridDim` intrinsics, as well as a `grid()` convenience function which evaluates to `blockDim * blockIdx + threadIdx`.
#
# Our example juse needs a minor modification to compute a grid-size stride for the x and y ranges, since we will have many threads running in parallel. We just add these three lines:
#
# startX, startY = cuda.grid(2)
# gridX = cuda.gridDim.x * cuda.blockDim.x;
# gridY = cuda.gridDim.y * cuda.blockDim.y;
#
# And we modify the range in the `x` loop to use `range(startX, width, gridX)` (and likewise for the `y` loop).
#
# We decorate the function with @cuda.jit, passing it the type signature of the function. Since kernels cannot have a return value, we do not need the `restype` argument.
@cuda.jit(argtypes=[f8, f8, f8, f8, uint8[:,:], uint32])
def mandel_kernel(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
startX, startY = cuda.grid(2)
gridX = cuda.gridDim.x * cuda.blockDim.x;
gridY = cuda.gridDim.y * cuda.blockDim.y;
for x in range(startX, width, gridX):
real = min_x + x * pixel_size_x
for y in range(startY, height, gridY):
imag = min_y + y * pixel_size_y
image[y, x] = mandel_gpu(real, imag, iters)
# #### Device Memory
# CUDA kernels must operate on data allocated on the device. NumbaPro provides the cuda.to_device() function to copy a Numpy array to the GPU.
#
# d_image = cuda.to_device(image)
#
# The return value (`d_image`) is of type DeviceNDArray, which is a subclass of numpy.ndarray, and provides the `to_host()` function to copy the array back from GPU to CPU memory
#
# d_image.to_host()
# #### Launching Kernels
# To launch a kernel on the GPU, we must configure it, specifying the size of the grid in blocks, and the size of each thread block. For a 2D image calculation like the Mandelbrot set, we use a 2D grid of 2D blocks. We'll use blocks of 32x8 threads, and launch 32x16 of them in a 2D grid so that we have plenty of blocks to occupy all of the multiprocessors on the GPU.
#
# Putting this all together, we launch the kernel like this.
# +
gimage = np.zeros((1024, 1536), dtype = np.uint8)
blockdim = (32, 8)
griddim = (32,16)
start = timer()
d_image = cuda.to_device(gimage)
mandel_kernel[griddim, blockdim](-2.0, 1.0, -1.0, 1.0, d_image, 20)
d_image.to_host()
dt = timer() - start
print "Mandelbrot created on GPU in %f s" % dt
imshow(gimage)
show()
# -
# You may notice that when you ran the above code, the image was generated almost instantly. On the NVIDIA Tesla K20c GPU installed in my desktop, it ran in 311 milliseconds, which is an additional 19.3x speedup over the `@autojit` (compiled CPU) code, or a total of over 2000x faster than interpreted Python code.
| notebooks/.ipynb_checkpoints/mandelbrot_numbapro-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/jonkrohn/DLTFpT/blob/master/notebooks/transfer_learning_in_tensorflow.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="0kbY0KYfXUvL"
# # Transfer Learning
#
# In this notebook, we load a pre-trained model (in this case, VGGNet19) and finetune it for a new task: detecting hot dogs.
# + [markdown] colab_type="text" id="gwkO_cNnXUvQ"
# #### Load dependencies
# + colab={} colab_type="code" id="QopB53apXUvV"
from tensorflow.keras.applications.vgg19 import VGG19 # new!
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Flatten
from tensorflow.keras.preprocessing.image import ImageDataGenerator # new!
# + [markdown] colab_type="text" id="yv-1x_UVXUvf"
# #### Load the pre-trained VGG19 model
# + colab={} colab_type="code" id="8EBF8Nq_XUvh"
vgg19 = VGG19(include_top=False,
weights='imagenet',
input_shape=(224,224,3),
pooling=None)
# + [markdown] colab_type="text" id="_Nas5PppXUvo"
# #### Freeze all the layers in the base VGGNet19 model
# + colab={} colab_type="code" id="OIfAuTcpXUvq"
for layer in vgg19.layers:
layer.trainable = False
# + [markdown] colab_type="text" id="MbKPfaJ4XUvw"
# #### Add custom classification layers
# + colab={} colab_type="code" id="1UwPBdAAXUvy"
# Instantiate the sequential model and add the VGG19 model:
model = Sequential()
model.add(vgg19)
# Add the custom layers atop the VGG19 model:
model.add(Flatten())
model.add(Dropout(0.5))
model.add(Dense(2, activation='softmax'))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="bcxO9igkahgS" outputId="0ac37247-f4ad-440b-ebdb-a9c06f56bc7c"
model.summary()
# + [markdown] colab_type="text" id="0hRykcyAXUv3"
# #### Compile the model for training
# + colab={} colab_type="code" id="8lMeMTHYXUv5"
model.compile(optimizer='nadam', loss='categorical_crossentropy', metrics=['accuracy'])
# + [markdown] colab_type="text" id="4qjso4EqXUwA"
# #### Prepare the data for training
# + colab={} colab_type="code" id="MmFfw5WyXUwC"
# You can comment out the two lines of code below if you executed them on
# a previous run of the notebook. The wget command downloads the data and the
# tar command extracts the data from a compressed file format.
# ! wget -c https://www.dropbox.com/s/86r9z1kb42422up/hot-dog-not-hot-dog.tar.gz
# ! tar -xzf hot-dog-not-hot-dog.tar.gz
# + colab={} colab_type="code" id="AXNPazweXUwH"
# Instantiate two image generator classes:
train_datagen = ImageDataGenerator(
rescale=1.0/255,
data_format='channels_last',
rotation_range=30,
horizontal_flip=True,
fill_mode='reflect')
valid_datagen = ImageDataGenerator(
rescale=1.0/255,
data_format='channels_last')
# + colab={} colab_type="code" id="GkC9jS5YXUwL"
# Define the batch size:
batch_size=32
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="gXyJj7ewXUwR" outputId="9b479e01-c188-4ad6-e90f-6ec454975abc"
# Define the train and validation generators:
train_generator = train_datagen.flow_from_directory(
directory='./hot-dog-not-hot-dog/train',
target_size=(224, 224),
classes=['hot_dog','not_hot_dog'],
class_mode='categorical',
batch_size=batch_size,
shuffle=True,
seed=42)
valid_generator = valid_datagen.flow_from_directory(
directory='./hot-dog-not-hot-dog/test',
target_size=(224, 224),
classes=['hot_dog','not_hot_dog'],
class_mode='categorical',
batch_size=batch_size,
shuffle=True,
seed=42)
# + colab={"base_uri": "https://localhost:8080/", "height": 434} colab_type="code" id="md-_KzmjXUwW" outputId="cbcca782-2fe2-418a-811a-6132c2128da9"
model.fit_generator(train_generator, steps_per_epoch=15,
epochs=16, validation_data=valid_generator,
validation_steps=15)
# + colab={} colab_type="code" id="0eto287aXUwa"
| notebooks/transfer_learning_in_tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 2D Isostatic gravity inversion - Initial Guess Model
# Este [IPython Notebook](http://ipython.org/videos.html#the-ipython-notebook) utiliza a biblioteca de código aberto [Fatiando a Terra](http://fatiando.org/)
# + active=""
# Initial Guess model of rifted margin. (model A)
# +
# #%matplotlib inline
import numpy as np
from scipy.misc import derivative
import scipy as spy
from scipy import interpolate
import matplotlib
matplotlib.use('TkAgg', force=True)
import matplotlib.pyplot as plt
import math
import cPickle as pickle
import datetime
import string as st
from scipy.misc import imread
from __future__ import division
from fatiando import gravmag, mesher, utils, gridder
from fatiando.mesher import Prism, Polygon
from fatiando.gravmag import prism
from fatiando.utils import ang2vec, si2nt, contaminate
from fatiando.gridder import regular
from fatiando.vis import mpl
from numpy.testing import assert_almost_equal
from numpy.testing import assert_array_almost_equal
from pytest import raises
plt.rc('font', size=16)
# -
import functions as fc
# ## Observation coordinates.
# +
# Model`s limits
ymin = 0.0
ymax = 195000.0
zmin = -1000.0
zmax = 37400.0
xmin = -100000.0
xmax = 100000.0
area = [ymin, ymax, zmax, zmin]
# -
ny = 150 # number of observation datas and number of prisms along the profile
# coordinates defining the horizontal boundaries of the
# adjacent columns along the profile
y = np.linspace(ymin, ymax, ny)
# coordinates of the center of the columns forming the
# interpretation model
n = ny - 1
dy = (ymax - ymin)/n
ycmin = ymin + 0.5*dy
ycmax = ymax - 0.5*dy
yc = np.reshape(np.linspace(ycmin, ycmax, n),(n,1))
x = np.zeros_like(yc)
z = np.zeros_like(yc)-150.0
## Edge extension (observation coordinates)
sigma = 2.0
edge = sigma*dy*n
# ## Model parameters
# +
# Model densities
# Indices and polygons relationship:
# cc = continental crust layer
# oc = ocean crust layer
# w = water layer
# s = sediment layer
# m = mantle layer
dw = np.array([1030.0])
ds = np.array([2600.0])
dcc = np.array([2790.0])
doc = np.array([2850.0])
dm = np.array([3200.0])
#dc = dcc
# coordinate defining the horizontal boundaries of the continent-ocean boundary
COT = 117000.0
# list defining crust density variance
dc = np.zeros_like(yc)
aux = yc <= COT
for i in range(len(yc[aux])):
dc[i] = dcc
for i in range(len(yc[aux]),n):
dc[i] = doc
# defining sediments layers density vector
ds = np.reshape(np.repeat(ds,n),(n,1))
# S0 => isostatic compensation surface (Airy's model)
# SR = S0+dS0 => reference Moho (Forward modeling)
S0 = np.array([29500.0])
dS0 = np.array([8500.0])
# -
# ## Synthetic data
gsyn = np.loadtxt('../data/A-model-rifted-margin-synthetic-gravity-data.txt')
# ## Water bottom
tw = np.reshape(np.loadtxt('../data/A-model-rifted-margin-bathymetry.txt'),(n,1))
# ## True surfaces
# +
true_basement = np.reshape(np.loadtxt('../data/A-model-rifted-margin-true-basement-surface.txt'),(n,1))
true_moho = np.reshape(np.loadtxt('../data/A-model-rifted-margin-true-moho-surface.txt'),(n,1))
# True reference moho surface (SR = S0+dS0)
true_S0 = np.array([29500.0])
true_dS0 = np.array([1500.0])
# -
# ## Known depths
# +
# Known values: basement and moho surfaces
base_known = np.loadtxt('../data/A-model-rifted-margin-basement-known-depths.txt', ndmin=2)
moho_known = np.loadtxt('../data/A-model-rifted-margin-moho-known-depths.txt', ndmin=2)
# -
# # Initial guess surfaces
# ### Basement surface
# + active=""
# mpl.close('all')
#
# mpl.subplot(2,1,1)
# mpl.title('Synthetic gravity disturbance', fontsize=14)
# mpl.paths([[ymin, 0.]], [[ymax, 0.]], style='--k', linewidth=1)
# mpl.plot(0.001*yc, gobs, label='obs')
# #mpl.ylim(-70.,5.)
# mpl.xlim(0.001*ymin, 0.001*ymax)
# mpl.ylabel('gravity disturbance (mGal)', fontsize=16)
# mpl.xticks(fontsize=12)
# mpl.yticks(fontsize=12)
# mpl.legend(loc='best')
#
# axes = mpl.subplot(2,1,2)
# mpl.ylim(zmax, zmin)
# mpl.xlim(ymin, ymax)
# mpl.xticks(fontsize=12)
# mpl.yticks(fontsize=12)
# mpl.xlabel('y (m)')
# mpl.ylabel('z (m)')
# mpl.paths([[ymin, 0.0]], [[ymax, 0.0]], style='-k', linewidth=1)
# mpl.plot(yc, tw, '-b', linewidth=1)
# mpl.plot(yc, true_basement, '-b', linewidth=1)
# mpl.plot(base_known[:,0], base_known[:,1], '*g', linewidth=1)
# mpl.plot(moho_known[:,0], moho_known[:,1], '*b', linewidth=1)
# mpl.m2km()
#
# sed_picks = mpl.draw_polygon(area, axes, color='r')
# + active=""
# sed_picks
# -
sed_picks = np.array([[ 393.14516129, 6456.53905054],
[ 22212.7016129 , 6792.57055349],
[ 32434.47580645, 10992.96434041],
[ 194213.70967742, 10992.96434041]])
# change the coordinates of the extremum points in order to
# avoid problems for constructing the interpolator
sed_picks[0,0] = ymin
sed_picks[-1,0] = ymax
basement = fc.surface_interpolate_function(sed_picks,yc)
for i in range(len(basement)):
if basement[i] < tw[i]:
basement[i] = tw[i]
# layer sediments thickness
ts = basement - tw
# +
#np.savetxt('../data/A-model-rifted-margin-initial-basement-surface.txt', basement, fmt='%.18f')
# -
# ### Moho surface
# + active=""
# mpl.close('all')
#
# mpl.subplot(2,1,1)
# mpl.title('Synthetic gravity disturbance', fontsize=14)
# mpl.paths([[ymin, 0.]], [[ymax, 0.]], style='--k', linewidth=1)
# mpl.plot(0.001*yc, gobs, label='obs')
# mpl.xlim(0.001*ymin, 0.001*ymax)
# mpl.ylabel('gravity disturbance (mGal)', fontsize=16)
# mpl.xticks(fontsize=12)
# mpl.yticks(fontsize=12)
# mpl.legend(loc='best')
#
# axes = mpl.subplot(2,1,2)
# mpl.ylim(zmax, zmin)
# mpl.xlim(ymin, ymax)
# mpl.xticks(fontsize=12)
# mpl.yticks(fontsize=12)
# mpl.xlabel('y (m)')
# mpl.ylabel('z (m)')
# mpl.paths([[ymin, 0.0]], [[ymax, 0.0]], style='-k', linewidth=1)
# mpl.plot(yc, tw, '-b', linewidth=1)
# mpl.plot(yc, basement, '-b', linewidth=1)
# mpl.plot(yc, true_basement, '--r', linewidth=1)
# mpl.plot(yc, true_moho, '--r', linewidth=1)
# mpl.plot(base_known[:,0], base_known[:,1], '*g', linewidth=1)
# mpl.plot(moho_known[:,0], moho_known[:,1], '*b', linewidth=1)
# mpl.m2km()
#
# moho_picks = mpl.draw_polygon(area, axes, color='r')
# + active=""
# moho_picks
# -
moho_picks = np.array([[ 1572.58064516, 25440. ],
[ 36562.5 , 24068.57142857],
[ 51108.87096774, 22268.57142857],
[ 193034.27419355, 21125.71428571]])
# change the coordinates of the extremum points in order to
# avoid problems for constructing the interpolator
moho_picks[0,0] = ymin
moho_picks[-1,0] = ymax
moho = fc.surface_interpolate_function(moho_picks,yc)
for i in range(len(moho)):
if moho[i] < basement[i]:
moho[i] = basement[i]
# +
# layer mantle thickness
tm = S0 - moho
# layer crust thickness
toc = moho - tw - ts
# +
#np.savetxt('../data/A-model-rifted-margin-initial-moho-surface.txt', moho, fmt='%.18f')
# -
# ## Initial guess data
# initial guess parameters vector
p0= []
p0 = np.vstack((ts, tm, dS0))
# prisms calculation by <NAME>
prism_w = fc.prism_w_function(xmax,xmin,dy,edge,dw,dcc,tw,yc)
prism_s = fc.prism_s_function(xmax,xmin,dy,edge,ds,dcc,tw,p0,yc)
prism_c = fc.prism_c_function(xmax,xmin,dy,edge,S0,dcc,dc,tw,p0,yc)
prism_m = fc.prism_m_function(xmax,xmin,dy,edge,S0,dcc,dm,p0,yc)
# +
# z component of gravity calculation by <NAME>
gzw = prism.gz(np.reshape(x,(n,)),np.reshape(yc,(n,)),np.reshape(z,(n,)),prism_w)
gzs = prism.gz(np.reshape(x,(n,)),np.reshape(yc,(n,)),np.reshape(z,(n,)),prism_s[0])
gzc = prism.gz(np.reshape(x,(n,)),np.reshape(yc,(n,)),np.reshape(z,(n,)),prism_c)
gzm = prism.gz(np.reshape(x,(n,)),np.reshape(yc,(n,)),np.reshape(z,(n,)),prism_m)
#Observed data calculation:
#g0 = fc.g_function(x,yc,z,gzw,prism_s,prism_c,prism_m)
g0 = gzw + gzs + gzc + gzm
# +
#np.savetxt('../data/A-model-rifted-margin-initial-guess-gravity-data.txt', g0, fmt='%.18f')
# -
# ## Model plot
# +
polygons_water = []
for (yi, twi) in zip(yc, tw):
y1 = yi - 0.5*dy
y2 = yi + 0.5*dy
polygons_water.append(Polygon(np.array([[y1, y2, y2, y1],
[0.0, 0.0, twi, twi]]).T,
props={'density': dw - dcc}))
polygons_sediments = []
for (yi, twi, si, dsi) in zip(yc, np.reshape(tw,(n,)), np.reshape(basement,(n,)), ds):
y1 = yi - 0.5*dy
y2 = yi + 0.5*dy
polygons_sediments.append(Polygon(np.array([[y1, y2, y2, y1],
[twi, twi, si, si]]).T,
props={'density': ds - dcc}))
polygons_crust = []
for (yi, si, Si, dci) in zip(yc, np.reshape(basement,(n,)), np.reshape(moho,(n,)), dc):
y1 = yi - 0.5*dy
y2 = yi + 0.5*dy
polygons_crust.append(Polygon(np.array([[y1, y2, y2, y1],
[si, si, Si, Si]]).T,
props={'density': dci - dcc}))
polygons_mantle = []
for (yi, Si) in zip(yc, np.reshape(moho,(n,))):
y1 = yi - 0.5*dy
y2 = yi + 0.5*dy
polygons_mantle.append(Polygon(np.array([[y1, y2, y2, y1],
[Si, Si, S0+dS0, S0+dS0]]).T,
props={'density': dm - dcc}))
# +
# %matplotlib inline
plt.close('all')
fig = plt.figure(figsize=(12,13))
import matplotlib.gridspec as gridspec
heights = [8, 8, 1]
gs = gridspec.GridSpec(3, 1, height_ratios=heights)
ax3 = plt.subplot(gs[0])
ax4 = plt.subplot(gs[1])
ax5 = plt.subplot(gs[2])
ax3.axhline(y=0.0, xmin=ymin, xmax=ymax, color='k', linestyle='--', linewidth=1)
ax3.plot(0.001*yc, gsyn, '-g', linewidth=2, label='simulated data')
ax3.plot(0.001*yc, g0, '-b', linewidth=2, label='initial guess data')
ax3.set_xlim(0.001*ymin, 0.001*ymax)
ax3.set_ylabel('gravity disturbance (mGal)', fontsize=16)
ax3.set_xticklabels(['%g'% (l) for l in ax3.get_xticks()], fontsize=14)
ax3.set_yticklabels(['%g'% (l) for l in ax3.get_yticks()], fontsize=14)
ax3.legend(loc='best', fontsize=14, facecolor='silver')
ax4.axhline(y=0.0, xmin=ymin, xmax=ymax, color='k', linestyle='-', linewidth=1)
aux = yc <= COT
for (pwi) in (polygons_water):
tmpx = [x for x in pwi.x]
tmpx.append(pwi.x[0])
tmpy = [y for y in pwi.y]
tmpy.append(pwi.y[0])
ax4.plot(tmpx, tmpy, linestyle='None')
ax4.fill(tmpx, tmpy, color='lightskyblue')
for (psi) in (polygons_sediments):
tmpx = [x for x in psi.x]
tmpx.append(psi.x[0])
tmpy = [y for y in psi.y]
tmpy.append(psi.y[0])
ax4.plot(tmpx, tmpy, linestyle='None')
ax4.fill(tmpx, tmpy, color='tan')
for (pci) in (polygons_crust[:len(yc[aux])]):
tmpx = [x for x in pci.x]
tmpx.append(pci.x[0])
tmpy = [y for y in pci.y]
tmpy.append(pci.y[0])
ax4.plot(tmpx, tmpy, linestyle='None')
ax4.fill(tmpx, tmpy, color='orange')
for (pcoi) in (polygons_crust[len(yc[aux]):n]):
tmpx = [x for x in pcoi.x]
tmpx.append(pcoi.x[0])
tmpy = [y for y in pcoi.y]
tmpy.append(pcoi.y[0])
ax4.plot(tmpx, tmpy, linestyle='None')
ax4.fill(tmpx, tmpy, color='olive')
for (pmi) in (polygons_mantle):
tmpx = [x for x in pmi.x]
tmpx.append(pmi.x[0])
tmpy = [y for y in pmi.y]
tmpy.append(pmi.y[0])
ax4.plot(tmpx, tmpy, linestyle='None')
ax4.fill(tmpx, tmpy, color='pink')
#ax4.axhline(y=S0, xmin=ymin, xmax=ymax, color='w', linestyle='--', linewidth=3)
ax4.plot(yc, tw, '-k', linewidth=3)
ax4.plot(yc, true_basement, '-k', linewidth=3, label='true surfaces')
ax4.plot(yc, true_moho, '-k', linewidth=3)
ax4.plot(yc, basement, '-.b', linewidth=3, label='initial guess surfaces')
ax4.plot(yc, moho, '-.b', linewidth=3)
ax4.axhline(y=true_S0+true_dS0, xmin=ymin, xmax=ymax, color='k', linestyle='-', linewidth=3)
ax4.axhline(y=S0+dS0, xmin=ymin, xmax=ymax, color='b', linestyle='-.', linewidth=3)
ax4.plot(base_known[:,0], base_known[:,1], 'v', color = 'yellow', markersize=15, label='known depths (basement)')
ax4.plot(moho_known[:,0], moho_known[:,1], 'D', color = 'lime', markersize=15, label='known depths (moho)')
#ax4.set_ylim((S0+dS0), zmin)
ax4.set_ylim((39000.0), zmin)
ax4.set_xlim(ymin, ymax)
ax4.set_xlabel('y (km)', fontsize=16)
ax4.set_ylabel('z (km)', fontsize=16)
ax4.set_xticklabels(['%g'% (0.001*l) for l in ax4.get_xticks()], fontsize=14)
ax4.set_yticklabels(['%g'% (0.001*l) for l in ax4.get_yticks()], fontsize=14)
ax4.legend(loc='lower right', fontsize=14, facecolor='silver')
X, Y = fig.get_dpi()*fig.get_size_inches()
plt.title('Density contrast (kg/m$^{3}$)', fontsize=18)
ax5.axis('off')
layers_list1 = ['water', 'sediment', 'continental', 'oceanic', 'mantle']
layers_list2 = ['', '', 'crust', 'crust', '']
colors_list = ['lightskyblue', 'tan', 'orange', 'olive', 'pink']
density_list = ['-1760', '-190', '0', '60', '410']
ncols = len(colors_list)
nrows = 1
h = Y / nrows
w = X / (ncols + 1)
i=ncols-1
for color, density, layers1, layers2 in zip(colors_list, density_list, layers_list1, layers_list2):
col = i // nrows
row = i % nrows
x = X - (col*w) - w
yi_line = Y
yf_line = Y - Y*0.15
yi_text1 = Y - Y*0.2
yi_text2 = Y - Y*0.27
yi_text3 = Y - Y*0.08
i-=1
poly = Polygon(np.array([[x, x+w*0.75, x+w*0.75, x], [yi_line, yi_line, yf_line, yf_line]]).T)
tmpx = [x for x in poly.x]
tmpx.append(poly.x[0])
tmpy = [y for y in poly.y]
tmpy.append(poly.y[0])
ax5.plot(tmpx, tmpy, linestyle='-', color='k', linewidth=1)
ax5.fill(tmpx, tmpy, color=color)
ax5.text(x+w*0.375, yi_text1, layers1, fontsize=(w*0.14), horizontalalignment='center', verticalalignment='top')
ax5.text(x+w*0.375, yi_text2, layers2, fontsize=(w*0.14), horizontalalignment='center', verticalalignment='top')
ax5.text(x+w*0.375, yi_text3, density, fontsize=(w*0.14), horizontalalignment='center', verticalalignment='center')
plt.tight_layout()
#mpl.savefig('../manuscript/figures/A-model-rifted-margin-initial-guess-model-grafics.png', dpi='figure', bbox_inches='tight')
plt.show()
# +
# %matplotlib inline
plt.close('all')
fig = plt.figure(figsize=(12,7))
import matplotlib.gridspec as gridspec
heights = [8, 1]
gs = gridspec.GridSpec(2, 1, height_ratios=heights)
ax1 = plt.subplot(gs[0])
ax2 = plt.subplot(gs[1])
ax1.axhline(y=0.0, xmin=ymin, xmax=ymax, color='k', linestyle='-', linewidth=1)
aux = yc <= COT
for (pwi) in (polygons_water):
tmpx = [x for x in pwi.x]
tmpx.append(pwi.x[0])
tmpy = [y for y in pwi.y]
tmpy.append(pwi.y[0])
ax1.plot(tmpx, tmpy, linestyle='None')
ax1.fill(tmpx, tmpy, color='lightskyblue')
for (psi) in (polygons_sediments):
tmpx = [x for x in psi.x]
tmpx.append(psi.x[0])
tmpy = [y for y in psi.y]
tmpy.append(psi.y[0])
ax1.plot(tmpx, tmpy, linestyle='None')
ax1.fill(tmpx, tmpy, color='tan')
for (pci) in (polygons_crust[:len(yc[aux])]):
tmpx = [x for x in pci.x]
tmpx.append(pci.x[0])
tmpy = [y for y in pci.y]
tmpy.append(pci.y[0])
ax1.plot(tmpx, tmpy, linestyle='None')
ax1.fill(tmpx, tmpy, color='orange')
for (pcoi) in (polygons_crust[len(yc[aux]):n]):
tmpx = [x for x in pcoi.x]
tmpx.append(pcoi.x[0])
tmpy = [y for y in pcoi.y]
tmpy.append(pcoi.y[0])
ax1.plot(tmpx, tmpy, linestyle='None')
ax1.fill(tmpx, tmpy, color='olive')
for (pmi) in (polygons_mantle):
tmpx = [x for x in pmi.x]
tmpx.append(pmi.x[0])
tmpy = [y for y in pmi.y]
tmpy.append(pmi.y[0])
ax1.plot(tmpx, tmpy, linestyle='None')
ax1.fill(tmpx, tmpy, color='pink')
#ax1.axhline(y=S0, xmin=ymin, xmax=ymax, color='w', linestyle='--', linewidth=3)
ax1.plot(yc, tw, '-k', linewidth=3)
ax1.plot(yc, true_basement, '-k', linewidth=3, label='true surfaces')
ax1.plot(yc, true_moho, '-k', linewidth=3)
ax1.plot(yc, basement, '-.b', linewidth=3, label='initial guess surfaces')
ax1.plot(yc, moho, '-.b', linewidth=3)
ax1.axhline(y=true_S0+true_dS0, xmin=ymin, xmax=ymax, color='k', linestyle='-', linewidth=3)
ax1.axhline(y=S0+dS0, xmin=ymin, xmax=ymax, color='b', linestyle='-.', linewidth=3)
ax1.plot(base_known[:,0], base_known[:,1], 'v', color = 'yellow', markersize=15, label='known depths (basement)')
ax1.plot(moho_known[:,0], moho_known[:,1], 'D', color = 'lime', markersize=15, label='known depths (moho)')
#ax1.set_ylim((S0+dS0), zmin)
ax1.set_ylim((39000.0), zmin)
ax1.set_xlim(ymin, ymax)
ax1.set_xlabel('y (km)', fontsize=16)
ax1.set_ylabel('z (km)', fontsize=16)
ax1.set_xticklabels(['%g'% (0.001*l) for l in ax1.get_xticks()], fontsize=14)
ax1.set_yticklabels(['%g'% (0.001*l) for l in ax1.get_yticks()], fontsize=14)
ax1.legend(loc='lower right', fontsize=14, facecolor='silver')
X, Y = fig.get_dpi()*fig.get_size_inches()
plt.title('Density contrast (kg/m$^{3}$)', fontsize=18)
ax2.axis('off')
layers_list1 = ['water', 'sediment', 'continental', 'oceanic', 'mantle']
layers_list2 = ['', '', 'crust', 'crust', '']
colors_list = ['lightskyblue', 'tan', 'orange', 'olive', 'pink']
density_list = ['-1760', '-190', '0', '60', '410']
ncols = len(colors_list)
nrows = 1
h = Y / nrows
w = X / (ncols + 1)
i=ncols-1
for color, density, layers1, layers2 in zip(colors_list, density_list, layers_list1, layers_list2):
col = i // nrows
row = i % nrows
x = X - (col*w) - w
yi_line = Y
yf_line = Y - Y*0.15
yi_text1 = Y - Y*0.2
yi_text2 = Y - Y*0.27
yi_text3 = Y - Y*0.08
i-=1
poly = Polygon(np.array([[x, x+w*0.75, x+w*0.75, x], [yi_line, yi_line, yf_line, yf_line]]).T)
tmpx = [x for x in poly.x]
tmpx.append(poly.x[0])
tmpy = [y for y in poly.y]
tmpy.append(poly.y[0])
ax2.plot(tmpx, tmpy, linestyle='-', color='k', linewidth=1)
ax2.fill(tmpx, tmpy, color=color)
ax2.text(x+w*0.375, yi_text1, layers1, fontsize=(w*0.14), horizontalalignment='center', verticalalignment='top')
ax2.text(x+w*0.375, yi_text2, layers2, fontsize=(w*0.14), horizontalalignment='center', verticalalignment='top')
ax2.text(x+w*0.375, yi_text3, density, fontsize=(w*0.14), horizontalalignment='center', verticalalignment='center')
plt.tight_layout()
#mpl.savefig('../manuscript/figures/A-model-rifted-margin-initial-guess-model.png', dpi='figure', bbox_inches='tight')
plt.show()
# -
| code/A-model-rifted-margin-model-(initial-guess).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Assignment 2a - Convolutional nets
# The purpose of this assignment is to help the student get started with convolutional image recogition models for the course project. This assignment will explore how convolutional nets work in depth.
import numpy as np
import os
import glob
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
date="05092019" #defining the date for saving files later
wd= os.getcwd()
allyes=glob.glob("../data/raw/yes/"+'*.[pjJ][npP][gG]') #grabs all jpg and png files
allno=glob.glob("../data/raw/no/"+'*.[pjJ][npP][gG]')
#can load with:
X=np.load("../data/processed/%s_X.npy"%(date))
y=np.load("../data/processed/%s_y.npy"%(date))
X.shape
# +
# X.shape
# +
# import cv2
# imlist=[]
# #filename = 'your_image.jpg'
# W = 224.
# #oriimg = cv2.imread(filename,cv2.CV_LOAD_IMAGE_COLOR)
# for image in X:
# height, width, depth = image.shape
# imgScale = W/width
# newX,newY = image.shape[1]*imgScale, image.shape[0]*imgScale
# newimg = cv2.resize(image,(int(newX),int(newY)))
# imlist.append(newimg)
# X=np.array(imlist)
# y = y.astype('float32')
# +
# resX=[]
# xres = 224
# yres = 224
# resizeval = (xres,yres)
# # resize and convert to grayscale
# for img in X:
# img = cv2.resize(img,resizeval)
# # gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# # img_expanded = gray[:, :, np.newaxis]
# # #print(img_expanded.shape)
# # print(gray.shape)
# # gray = img.reshape((len(gray), -1)).T
# # print(gray.shape)
# resX.append(img)
# resX = np.array(resX)
# #resX = resX.reshape((len(resX), -1)).T
# #resX = resX.reshape(xres,yres, 1)
# X = resX
# #y = y.T
# y = y.astype('float32')
# +
resX=[]
xres = 224
yres = 224
resizeval = (xres,yres)
# resize and convert to grayscale
for img in X:
img = cv2.resize(img,resizeval)
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
img_expanded = gray[:, :, np.newaxis]
#print(img_expanded.shape)
# print(gray.shape)
#gray = img_expanded.reshape((len(X), -1)).T
# print(gray.shape)
resX.append(img_expanded)
resX = np.array(resX)
#resX = resX.reshape((len(resX), -1)).T
#resX = resX.reshape(xres,yres, 1)
X = resX
#y = y.T
y = y.astype('float32')
# -
X.shape
224*224
pixels = X.flatten().reshape(253, 50176)
pixels.shape
y_tindex = y[:,0]==True
y_findex = y[:,1]==True
y2=np.zeros(len(y))
y2[y_tindex]=1
# Northwestern MSiA 432 Deep learning
# Assignment #2 starter code, Spring 2019.
#
# This code demonstrates the use of transfer learning to speed up
# the training process for a convolutional neural network.
#
# Notes:
# - Heatmaps may appear black in the first few epochs. Wait until accuracy improves.
# - The native image size is 224x224 for VGG, resize/crop your images to match
# - Filter visualization is slow, change vizfilt_timeout for more speed or accuracy
# - Be sure to rename/delete the basepath when changing model parameters, e.g. layers or random labels
#see which pid you are am
import os
os.getpid()
# !nvidia-smi
# +
# ##change gpus
# ###NOTE: this needs to be run before importing tensorflow.
# import os
# os.environ["CUDA_VISIBLE_DEVICES"]="0"
# +
# #%% ------ CPU/GPU memory fix -------
# import tensorflow as tf, keras.backend.tensorflow_backend as ktf
# def get_session(gpu_fraction=0.45):
# gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=gpu_fraction, allow_growth=True)
# return tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
# ktf.set_session(get_session())
# -
# Show devices
from tensorflow.python.client import device_lib
local_device_protos = device_lib.list_local_devices()
print([x.name for x in local_device_protos])
# +
import tensorflow as tf
from tensorflow.python.client import device_lib
def get_available_gpus():
local_device_protos = device_lib.list_local_devices()
return [x.name for x in local_device_protos if x.device_type == 'GPU']
print("Tensorflow Version:", tf.__version__)
print(tf.keras.__version__)
print(get_available_gpus())
# +
# ##change gpus
# import os
# os.environ["CUDA_VISIBLE_DEVICES"]="1"
# -
# # imports and function definitions
# +
# #may need to install a older scipy version as some functions require it
# # !pip install --ignore-installed --user scipy==1.2.1
# +
# Obligatory imports
from IPython.display import display
from PIL import Image
import imageio
import os, time, numpy as np, scipy, random, pandas as pd, socket, warnings
import matplotlib.pyplot as plt
from keras.optimizers import Adam, SGD, RMSprop, Adagrad
os.environ["KMP_DUPLICATE_LIB_OK"] = "TRUE" # Workaround for Mac
# %matplotlib inline
# +
# Set project configuration
def change_basepath(folder_name=None):
"""
GE added, makes it easy to specify a folder extention that will be created onto basepath. if folder_name==None, then basepath will be reset to it's origional value.
"""
basepath= "../data"
if folder_name != None:
address = os.path.join(basepath, folder_name)
else:
address = basepath
if not os.path.exists(address):
os.makedirs(address)
return(address)
basepath= change_basepath()
# -
# VGG net definition starts here. Change the vggblocks to set how many blocks to transfer
def make_vgg():
from keras.models import Model
from keras.regularizers import l1_l2
from keras.layers import Flatten, Dense, Input, Convolution2D, MaxPooling2D, BatchNormalization
from keras.layers.advanced_activations import LeakyReLU
img_input = Input(shape=tsize)
if vggblocks == 0: x = img_input
if vggblocks >= 1: # Block 1
x = LeakyReLU(alpha)(Convolution2D(64, (3, 3), padding='same', name='block1_conv1')(img_input))
x = LeakyReLU(alpha)(Convolution2D(64, (3, 3), padding='same', name='block1_conv2')(x))
x = MaxPooling2D((2, 2), strides=(2, 2), name='block1_pool')(x)
if batchnorm: x = BatchNormalization()(x)
if vggblocks >= 2: # Block 2
x = LeakyReLU(alpha)(Convolution2D(128, (3, 3), padding='same', name='block2_conv1')(x))
x = LeakyReLU(alpha)(Convolution2D(128, (3, 3), padding='same', name='block2_conv2')(x))
x = MaxPooling2D((2, 2), strides=(2, 2), name='block2_pool')(x)
if batchnorm: x = BatchNormalization()(x)
if vggblocks >= 3: # Block 3
x = LeakyReLU(alpha)(Convolution2D(256, (3, 3), padding='same', name='block3_conv1')(x))
x = LeakyReLU(alpha)(Convolution2D(256, (3, 3), padding='same', name='block3_conv2')(x))
x = LeakyReLU(alpha)(Convolution2D(256, (3, 3), padding='same', name='block3_conv3')(x))
x = MaxPooling2D((2, 2), strides=(2, 2), name='block3_pool')(x)
if batchnorm: x = BatchNormalization()(x)
if vggblocks >= 4: # Block 4
x = LeakyReLU(alpha)(Convolution2D(512, (3, 3), padding='same', name='block4_conv1')(x))
x = LeakyReLU(alpha)(Convolution2D(512, (3, 3), padding='same', name='block4_conv2')(x))
x = LeakyReLU(alpha)(Convolution2D(512, (3, 3), padding='same', name='block4_conv3')(x))
x = MaxPooling2D((2, 2), strides=(2, 2), name='block4_pool')(x)
if batchnorm: x = BatchNormalization()(x)
if vggblocks >= 5: # Block 5
x = LeakyReLU(alpha)(Convolution2D(512, (3, 3), padding='same', name='block5_conv1')(x))
x = LeakyReLU(alpha)(Convolution2D(512, (3, 3), padding='same', name='block5_conv2')(x))
x = LeakyReLU(alpha)(Convolution2D(512, (3, 3), padding='same', name='block5_conv3')(x))
x = MaxPooling2D((2, 2), strides=(2, 2), name='block5_pool')(x)
if batchnorm: x = BatchNormalization()(x)
x = Flatten(name='flatten')(x)
for i in range(fclayers): x = LeakyReLU(alpha)(Dense(fclayersize, kernel_regularizer=l1_l2(l1_reg, l2_reg))(x))
x = Dense(len(obj_classes), activation='softmax', name='predictions')(x)
inputs = img_input
model = Model(inputs, x, name='vgg16')
# VGG Transfer weights
from keras.applications import vgg16
import keras.layers.convolutional
vgg16model = vgg16.VGG16(include_top=False)
modelconv = [l for l in model.layers if type(l) == keras.layers.convolutional.Conv2D]
vgg16conv = [l for l in vgg16model.layers if type(l) == keras.layers.convolutional.Conv2D]
for i, l in enumerate(modelconv):
if i > xferlearning: continue # Transfer only first n layers
print('**** Transferring layer %d: %s from VGG ****' % (i, l))
weights = vgg16conv[i].get_weights()
modelconv[i].set_weights(weights)
if freeze_conv: l.trainable = False
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
model.summary()
return model, img_input
#%% resnet50 definition
def make_resnet50():
#The default input size for this model is 224x224.
#This model and can be built both with 'channels_first' data format (channels, height, width)
#or 'channels_last' data format (height, width, channels).
import keras
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D, Dropout
#from keras_applications.resnet import ResNet50
from keras.applications import ResNet50
from keras.regularizers import l1
xmodel = ResNet50(include_top=False,
# weights='imagenet',
# input_tensor=None,
# input_shape=None,
# pooling=None,
classes=10)
#xmodel = keras.applications.Xception(include_top=False)
# xmodel =keras.applications.resnet50(include_top=False)
x = xmodel.output
#for layer in xmodel.layers: layer.trainable = False
x = GlobalAveragePooling2D()(x)
x = Dropout(0.5)(x)
x = Dense(256, kernel_regularizer=l1(1e-7))(x)
predictions = Dense(len(obj_classes), activation='softmax')(x)
model = Model(inputs=xmodel.input, outputs=predictions)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=Adam())
model.summary()
return model, xmodel.input
#%% Xception definition
def make_xception():
import keras
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D, Dropout
from keras.regularizers import l1
xmodel = keras.applications.Xception(include_top=False) #keras.applications.resnet50 to change it
x = xmodel.output
#for layer in xmodel.layers: layer.trainable = False
x = GlobalAveragePooling2D()(x)
x = Dropout(0.5)(x)
x = Dense(256, kernel_regularizer=l1(1e-7))(x)
predictions = Dense(len(obj_classes), activation='softmax')(x)
model = Model(inputs=xmodel.input, outputs=predictions)
model.compile(loss='categorical_crossentropy', metrics=['accuracy'], optimizer=Adam())
model.summary()
return model, xmodel.input
# # Image Import, Scaling, Batching options
# +
# from sklearn.model_selection import train_test_split
# X_train,X_test,y_train,y_test = train_test_split(X, y, test_size=0.2, random_state=13)
# train_X,valid_X,train_label,valid_label = train_test_split(X_train, y_train, test_size=0.2, random_state=13)
# +
######### CHANGE IMSIZE, BATCHSIZE HERE #########
imsize = (224, 224) # n x n square images, VGG default is 224x224. Remember to change this.
tsize = imsize + (3,) #for 3 color channels
batch_size, nb_epoch = 32, 10000 # Change for early stopping regularization.
#changed batchsize from #32 to 253
######### ****************************** #########
#image path specification
trainfolder = '../data/train'#os.path.join(basepath, 'train')
testfolder = '../data/test'#os.path.join(basepath, 'test')
#%% Create demo data
def makedata(basepath):
from keras.datasets import cifar10
from keras.utils.generic_utils import Progbar
(X_train, y_train), (X_test, y_test) = cifar10.load_data()
obj_classes = ['airplane', 'automobile', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck']
for (X_data, y_data, bp) in [(X_train, y_train, trainfolder), (X_test, y_test, testfolder)]:
if os.path.exists(bp): return
for c in obj_classes: os.makedirs(os.path.join(bp, c), exist_ok=True)
pb = Progbar(len(y_data), interval=1)
print('\nMaking data folder')
for i, (im, lbl) in enumerate(zip(X_data, y_data)):
pn = os.path.join(bp, obj_classes[int(lbl)], "%d.png" % i)
pb.update(i)
if not os.path.exists(pn): imageio.imwrite(pn, im)#scipy.misc.imsave(pn, im)
# +
# makedata(basepath) # Comment out this line to use your own data
# Load data
from keras.preprocessing.image import ImageDataGenerator
##middleground data aug
datagen = ImageDataGenerator(
rescale=1./255, # rescale data
shear_range=0.2,
zoom_range=0.2,
rotation_range=0.10, #0 # randomly rotate images in the range (degrees, 0 to 180)
width_shift_range=0.1, #0 # randomly shift images horizontally (fraction of total width)
height_shift_range=0.1, #0 # randomly shift images vertically (fraction of total height)
horizontal_flip=True, # randomly flip images
vertical_flip=False) # randomly flip images
#no data aug
# datagen = ImageDataGenerator(
# rescale=1./255, # rescale data
# shear_range=0.0,
# zoom_range=0.0,
# rotation_range=0.0, #0 # randomly rotate images in the range (degrees, 0 to 180)
# width_shift_range=0.0, #0 # randomly shift images horizontally (fraction of total width)
# height_shift_range=0.0, #0 # randomly shift images vertically (fraction of total height)
# horizontal_flip=False, # randomly flip images
# vertical_flip=False) # randomly flip images
# ##lots of data aug
# datagen = ImageDataGenerator(
# rescale=1./255, # rescale data
# shear_range=0.7,
# zoom_range=0.7,
# rotation_range=0.7, #0 # randomly rotate images in the range (degrees, 0 to 180)
# width_shift_range=0.7, #0 # randomly shift images horizontally (fraction of total width)
# height_shift_range=0.7, #0 # randomly shift images vertically (fraction of total height)
# horizontal_flip=True, # randomly flip images
# vertical_flip=True) # randomly flip images
train_generator = datagen.flow_from_directory(
trainfolder,
target_size=imsize,
batch_size=batch_size #changd
)
test_generator = datagen.flow_from_directory(
testfolder,
target_size=imsize,
batch_size=-1)
X_test, Y_test = test_generator.next()
obj_classes = sorted(train_generator.class_indices.keys())
class_to_idx = dict([(y, x) for (x,y) in enumerate(obj_classes)])
img_rows, img_cols, img_channels = X_test.shape[1:]
# -
obj_classes
# ## visualization functions
#%% Visualization code
def viz_losses(stats):
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(10,6))
epoch = len(stats)
convlayers = len([l.name for l in model.layers if 'conv' in l.name])
blocks = len(set([l.name.split('_')[0] for l in model.layers if 'block' in l.name]))
dense = len([l.name for l in model.layers if 'dense' in l.name])
fcsize = model.layers[-1].input_shape[1]
fig.suptitle("Training %s blocks=%d, conv=%d, dense=%d, fcsize=%d, epoch=%d" % (model.name, blocks, convlayers, dense, fcsize, epoch))
ax1.plot(stats['Train loss'].values, label='Train loss', color='blue')
ax1.plot(stats['Test loss'].values, label='Test loss', color='green')
ax1.set_yscale('log')
ax2.plot(stats['Accuracy'].values, label='Test accuracies', color='red')
ax2.plot(stats['Train accuracy'].values, label='Train accuracies', color='blue')
ax2.axhline(1.0/len(obj_classes), linestyle='dashed', color='gray')
dataset = pd.Series(train_generator.classes)
chance = dataset.value_counts().max() / dataset.value_counts().sum()
ax2.text(0, chance, 'Chance')
ax2.axhline(np.max(stats['Accuracy']), linestyle='dashed', color='red')
ax2.text(0, np.max(stats['Accuracy']), 'Best')
ax2.set_ylim([0, 1])
ax2.set_title('Accuracy: %0.2f%%' % (100.0*stats['Accuracy'].values[-1]))
ax1.legend(), ax2.legend()
plt.savefig(os.path.join(basepath, 'loss-%s.png' % modelarch))
plt.show()
plt.close()
# +
#%% Explanations
import skimage.exposure, skimage.filters
from skimage.color import gray2rgb
from keras import backend as K
def hide_axes(ax): ax.set_xticks([]), ax.set_yticks([])
class Heatmap:
def __init__(self, model, obj_classes):
self.obj_classes = obj_classes
self.nclasses = len(obj_classes)
self.model = model
def make_masks(self, im, n=8, maskval=0.1):
masks = []
xwidth, ywidth = int(np.ceil(im.shape[0]/n)), int(np.ceil(im.shape[1]/n))
for i in range(n):
for j in range(n):
mask = np.ones(im.shape[:2])
mask[(i*xwidth):((i+1)*xwidth), (j*ywidth):((j+1)*ywidth)] = maskval
mask = skimage.filters.gaussian(mask, 1) # Change this for local mask smoothing
masks.append(mask)
return np.array(masks)
def get_slice_masks(self, im, n_segments=16, blur=0.03):
from skimage.segmentation import slic
segments = slic(im, n_segments=n_segments, sigma=5)
masks = []
# loop over the unique segment values
for (i, segVal) in enumerate(np.unique(segments)):
# construct a mask for the segment
mask = np.zeros(im.shape[:2], dtype="float32")
mask[segments == segVal] = 1
mask = skimage.filters.gaussian(mask, im.shape[1]*blur) # Change this for local mask smoothing
masks.append(mask)
return np.array(masks), segments
def explain_prediction_heatmap(self, im, actual):
##ge added start##
global trainstats
##ge added end#
import skimage.color
def hsv_fn(im): return skimage.color.hsv2rgb(im) if hsv else im
plt.imshow(hsv_fn(im), interpolation='bilinear'), plt.xticks([]), plt.yticks([]), plt.title('Full image'), plt.show(), plt.close()
masks = np.concatenate([self.make_masks(im, n=i) for i in (9, 7, 5, 3, 2)])
#masks, segments = self.get_slice_masks(im)
masknorm = masks.sum(axis=0)
heatmaps = np.zeros((self.nclasses,) + im.shape[:2])
for m in masks:
prediction = self.model.predict(np.expand_dims(im*gray2rgb(m), 0))
for c in range(self.nclasses):
heatmaps[c] += (prediction[0][c]*m)
for h in heatmaps: h = h / masknorm
fig, axes = plt.subplots(2, self.nclasses + 1, figsize=(20, 5))
#axes[0,0].imshow(hsv(im)), axes[1,0].imshow(mark_boundaries(im, segments))
axes[0,0].imshow(hsv_fn(im)), axes[1,0].imshow(im)
axes[0,0].set_title(actual)
axes[1,0].set_title('HSV' if hsv else 'RGB')
hide_axes(axes[0,0]), hide_axes(axes[1,0])
predictions = np.sum(heatmaps, axis=(1,2,))
predictions /= predictions.max()
for n, i in enumerate(np.argsort(predictions)[::-1][:self.nclasses]):
h = ((255 * heatmaps[i])/heatmaps[i].max()).astype('uint16')
h = skimage.exposure.equalize_adapthist(h)
h = skimage.filters.gaussian(h, 1) # Change this for global mask smoothing
axes[0, n+1].imshow(gray2rgb(h))
axes[1, n+1].imshow(gray2rgb(h) * hsv_fn(im) * (0.5 + 0.5*predictions[i]))
hide_axes(axes[0, n+1]), hide_axes(axes[1, n+1])
axes[0, n+1].set_title(self.obj_classes[i] + ': %0.1f%%' % (100*predictions[i]/predictions.sum()))
fig.tight_layout()
##ge modify: added e to heatmapfilename and text inserted the len(trainstats)=epoch number ##
plt.savefig(os.path.join(basepath, 'heatmap-e%05d.png') % len(trainstats))#np.random.randint(0, 99999))
plt.show()
plt.close()
return heatmaps
# +
#layer_dict = dict([(layer.name, layer) for layer in model.layers])
def deprocess_image(x):
x -= x.mean()
x /= (x.std() + 1e-5)
x = x*0.1 + 0.5
x = np.clip(x, 0, 1) * 255
x = np.clip(x, 0, 255).astype('uint8')
return x
def viz_filter_max(layer_name, filter_index=0, max_steps=9999, timeout=3):
from keras.utils.generic_utils import Progbar
layer_output = layer_dict[layer_name].output
loss = K.mean(layer_output[:, :, :, filter_index])
grads = K.gradients(loss, img_input)[0]
grads /= (K.sqrt(K.mean(K.square(grads))) + 1e-5)
iterate = K.function([img_input], [loss, grads])
step = 1e-0
input_img_data = np.random.random((1, img_rows, img_cols, 3))
input_img_data = (input_img_data - 0.5) * 20 + 128
tm = time.time()
for i in range(max_steps):
loss_value, grads_value = iterate([input_img_data])
input_img_data += grads_value * step
if time.time() - tm > timeout:
plt.text(0.1, 0.1, "Filter viz timeout: %d" % timeout, color='red')
break
img = input_img_data[0]
img = deprocess_image(img)
fig = plt.imshow(img)
hide_axes(fig.axes)
return layer_output
def viz_filters(model, img_input, img_rows, img_cols, nbfilters=3, timeout=60):
tm = time.time()
print("Visualizing filters (CTRL-C to cancel)")
try:
for layer_name in sorted(layer_dict.keys()):
if time.time() - tm > timeout:
print("Filter visualization timed out: %d. Change timeout in viz_filters()." % timeout)
break
if not hasattr(layer_dict[layer_name], 'filters'): continue
nfilters = layer_dict[layer_name].filters
fig, ax = plt.subplots(1, nbfilters, figsize=(8, 4))
fig.suptitle("Layer %s has %d filters" % (layer_name, nfilters))
for j in range(nbfilters):
plt.subplot(1, nbfilters, j + 1)
viz_filter_max(layer_name, random.randint(0, nfilters-1), timeout=vizfilt_timeout)
fig.tight_layout()
plt.savefig(os.path.join(basepath, 'filters-%s-%s.png' % (modelarch, layer_name)))
plt.show(), plt.close()
except KeyboardInterrupt: return
# +
def test_prediction(im, y):
pred = model.predict(np.expand_dims(im, 0))
cls = np.argmax(y)
heatmap = Heatmap(model, obj_classes)
heatmap.explain_prediction_heatmap(im, obj_classes[cls])
print("Actual: %s(%d)" % (obj_classes[cls], cls))
for cls in list(reversed(np.argsort(pred)[0]))[:5]:
conf = float(pred[0, cls])/pred.sum()
print(" predicted: %010s(%d), confidence=%0.2f [%-10s]" % (obj_classes[cls], cls, conf, "*" * int(10*conf)))
return pred
def confusion_matrix(model, X, T, accpct, save=False):
##GE modified to save##
import seaborn
global trainstats
from sklearn.metrics import classification_report, confusion_matrix
Y_pred = model.predict(X)
y_pred = np.argmax(Y_pred, axis=1)
y_test = np.argmax(T, axis=1)
print('Confusion Matrix')
data = confusion_matrix(y_test, y_pred)
data = data / data.sum(axis=1)
#print('Classification Report')
#print(classification_report(y_test, y_pred, target_names=obj_classes))
seaborn.set_style("whitegrid", {'axes.grid' : False})
seaborn.heatmap(data, annot=data*100, fmt='0.0f', cmap='Wistia', xticklabels=obj_classes, yticklabels=obj_classes)
plt.xlabel('Predicted'), plt.ylabel('Actual'), plt.title('Confusion matrix (ACC %0.2f%%)' % (accpct*100))
if save==True and len(trainstats) in [1,epochs] :
plt.savefig(os.path.join(basepath, 'conf-%s.png' % len(trainstats)))
plt.show(), plt.close()
def tsne_viz(model, X, Y, accpct, n=500, save=False):
##GE modified to save##
global X_embedded, xx, yy, d, predictions
import sklearn.manifold, matplotlib.cm as cm
predictions = model.predict(X)
colors = iter(cm.rainbow(np.linspace(0, 1, len(obj_classes))))
X_embedded = sklearn.manifold.TSNE(n_components=2).fit_transform(predictions[:n])
for d in range(len(obj_classes)):
xx = X_embedded[Y[:n][:, d] == 1, 0]
yy = X_embedded[Y[:n][:, d] == 1, 1]
plt.scatter(xx, yy, c=[next(colors)], label=obj_classes[d])
t = plt.text(np.median(xx), np.median(yy), obj_classes[d], fontsize=24)
t.set_bbox({'facecolor': 'white', 'alpha': 0.75})
plt.title('T-SNE viz - Accuracy: %0.2f%%' % (accpct*100)), plt.legend()
if save==True and len(trainstats) in [1,epochs] :
plt.savefig(os.path.join(basepath, 'tsne-%s.png' % len(trainstats)))
# -
##grand training visualization function
from keras.callbacks import Callback
class VizTraining(Callback):
def on_epoch_end(self, epoch, logs={}):
clear_output(wait=True)
tacc = logs.get('val_acc')
trainstats.loc[len(trainstats)] = (logs.get('loss'), logs.get('val_loss'), tacc, logs.get('acc'))
confusion_matrix(model, X_test, Y_test, tacc, save=True)
tsne_viz(model, X_test, Y_test, tacc, save=False)
viz_losses(trainstats)
t_ind = random.randint(0, len(X_test) - 1)
test_prediction(X_test[t_ind], Y_test[t_ind])
if vizfilt_timeout > 0 and np.random.randint(0, 3) == 0:
viz_filters(model, img_input, img_rows, img_cols)
if checkpoint: model.save(modelid, overwrite=True)
print("Total training time: %0.2f min, epoch: %d" % ((time.time() - tm)/60.0, len(trainstats)))
print("Average time per epoch: %0.2f s" % ((time.time() - tm)/len(trainstats)))
# # example run:
# # ## copy these two cells and make appropriate changes for each model
# vgg with block=1, non-xfer learning, 25 epoch:
# +
basepath= change_basepath('xception_heavyaug') ####change
# Model settings
vggblocks = 3 # Number of VGG blocks to create, 0-5 blocks
xferlearning = 2 # Enable transfer learning up to layer n (max 12, -1 = off)
freeze_conv = False # Freeze convolutional layers
fclayersize = 64#128 # Size of fully connected (FC) layers
fclayers = 1 # Number of FC layers
fcdropout = 0.0 # Dropout regularization factor for FC layers
alpha = 0.0 # Leaky ReLU alpha
l1_reg = 0.0 # L1 regularization for FC
l2_reg = 0.0 # L2 regularization for FC
# Optimizer settings
optimizer = Adam()
# batch_size, nb_epoch = 32, 10000 # Change for early stopping regularization
batchnorm = True # Batch normalization
checkpoint = True # Checkpoint models to continue training
# Visualization settings
hsv = False # Convert images to Hue/Saturation/Value to be more robust to color variations
vizfilt_timeout = 0 # Decrease for speed, increase for better viz. 0 = off.####changed
# Model checkpointing/stats
# modeltype = 'xception'#'resnet50'#'vgg' # Use default VGG model
modeltype = 'xception'#'resnet50'#'vgg' # Use default VGG model
#modeltype = 'xception' # New Xception model, recommended imsize = (64, 64) or larger
modelarch = '%s%d-fcl%d-fcs%d-%s-%s' % (modeltype, vggblocks, fclayers, fclayersize, 'hsv' if hsv else 'rgb', socket.gethostname())
##GE added start##: adding quick script to make a model folder for modelid for easier saving
if not os.path.exists(os.path.join(basepath, "models/")):
os.makedirs(os.path.join(basepath, "models/"))
##GE added end##
modelid = os.path.join(basepath, 'models/xception_heavyaug.h5') ####change
##make model
if modeltype == 'vgg':
model, img_input = make_vgg()
elif modeltype == 'xception':
model, img_input = make_xception()
elif modeltype == 'resnet50':
model, img_input = make_resnet50()
elif modeltype == 'unet':
model, img_input = unet()
layer_dict = dict([(layer.name, layer) for layer in model.layers]) ####IMPORTANT FIX FOR VISUALIZATION FILTER
#loading model weights
if checkpoint and os.path.exists(modelid):
print("**** Loading existing model: %s ****" % modelid)
try:
model.load_weights(modelid)
except ValueError:
print("Model restore failed. Model topology must match to restore weights. Please delete weight checkpoint model.h5.")
except OSError:
print("Model checkpoint corrupted. Please delete.")
# +
#%% Training code
##transfer learning
from IPython.display import clear_output
tm = time.time()
trainstats = pd.DataFrame(columns=('Train loss', 'Test loss', 'Accuracy', 'Train accuracy'))
epochs=50 ####change
from keras.callbacks import Callback
loss = model.fit_generator(
train_generator,
steps_per_epoch=int(len(train_generator.filenames)/batch_size),
validation_data=(X_test, Y_test),
validation_steps=1,
verbose=1, epochs=epochs,
use_multiprocessing=True,
workers=4,
callbacks=[VizTraining()]
)
# -
# +
def target_category_loss(x, category_index, nb_classes):
return tf.multiply(x, K.one_hot([category_index], nb_classes))
def target_category_loss_output_shape(input_shape):
return input_shape
def normalize(x):
# utility function to normalize a tensor by its L2 norm
return x / (K.sqrt(K.mean(K.square(x))) + 1e-5)
def grad_cam(input_model, image, category_index, layer_name):
from keras.preprocessing import image
from keras.layers.core import Lambda
from keras.models import Sequential
from tensorflow.python.framework import ops
import keras.backend as K
import tensorflow as tf
import numpy as np
import keras
import sys
import cv2
model = Sequential()
model.add(input_model)
nb_classes = 2
target_layer = lambda x: target_category_loss(x, category_index, nb_classes)
print(target_layer)
model.add(Lambda(target_layer,
output_shape = target_category_loss_output_shape))
model.summary()
loss = K.sum(model.layers[-1].output)
conv_output = [l for l in model.layers[0].layers if l.name is layer_name][0].output
grads = K.gradients(loss, conv_output)[0]#normalize(K.gradients(loss, conv_output)[0])
gradient_function = K.function([model.layers[0].input], [conv_output, grads])
output, grads_val = gradient_function([image])
output, grads_val = output[0, :], grads_val[0, :, :, :]
weights = np.mean(grads_val, axis = (0, 1))
cam = np.ones(output.shape[0 : 2], dtype = np.float32)
for i, w in enumerate(weights):
cam += w * output[:, :, i]
cam = cv2.resize(cam, (224, 224))
cam = np.maximum(cam, 0)
heatmap = cam / np.max(cam)
#Return to BGR [0..255] from the preprocessed image
image = image[0, :]
image -= np.min(image)
image = np.minimum(image, 255)
cam = cv2.applyColorMap(np.uint8(255*heatmap), cv2.COLORMAP_JET)
cam = np.float32(cam) + np.float32(image)
cam = 255 * cam / np.max(cam)
return np.uint8(cam), heatmap
# -
K.gradients(loss, conv_output)[0]
# +
# ## testing adding this sequential layer
# from keras.models import Sequential
# from keras.layers.core import Lambda
# ##importing predictions
# def load_image(path):
# from keras.preprocessing import image
# import sys
# #img_path = sys.argv[1]
# img = image.load_img(img_path, target_size=(224, 224))
# x = image.img_to_array(img)
# x = np.expand_dims(x, axis=0)
# #x = preprocess_input(x)
# x= x/255
# return x
# img_path='../data/train/False/1.png'
# preprocessed_input= load_image(img_path)
# predictions = model.predict(preprocessed_input)
# #top_1 = decode_predictions(predictions)[0][0]
# predicted_class = np.argmax(predictions)
# ###testing fxn internals
# input_model, image, category_index, layer_name= model, preprocessed_input, predicted_class, "block14_sepconv2_act"
# model2 = Sequential()
# model2.add(input_model)
# nb_classes = 2
# target_layer = lambda x: target_category_loss(x, category_index, nb_classes)
# print(target_layer)
# model2.add(Lambda(target_layer,
# output_shape = target_category_loss_output_shape))
# model2.summary()
# loss = K.sum(model2.layers[-1].output)
# conv_output = [l for l in model2.layers[0].layers if l.name is layer_name][0].output
# grads = K.gradients(loss, conv_output)[0]#normalize(K.gradients(loss, conv_output)[0])
# #model2.inputs
# #gradient_function = K.function([model2.layers[0].input], [conv_output, grads])
# gradient_function = K.function([model2.inputs], [conv_output, grads])
# -
grads
K.gradients(loss, conv_output)[0]
grads
model2.inputs
model2.layers[0].layers[0]#.input
cam, heatmap = grad_cam(model, preprocessed_input, predicted_class, "block14_sepconv2_act")
loss
[model.layers[0].input]
visualize_cam(model, layer_idx, filter_indices, seed_input, penultimate_layer_idx=None, \
backprop_modifier=None, grad_modifier=None)
# +
def load_image(path):
from keras.preprocessing import image
import sys
#img_path = sys.argv[1]
img = image.load_img(img_path, target_size=(224, 224))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
#x = preprocess_input(x)
x= x/255
return x
img_path='../data/train/False/1.png'
preprocessed_input= load_image(img_path)
predictions = model.predict(preprocessed_input)
#top_1 = decode_predictions(predictions)[0][0]
# print('Predicted class:')
# print('%s (%s) with probability %.2f' % (top_1[1], top_1[0], top_1[2]))
predicted_class = np.argmax(predictions)
# +
predictions = model.predict(preprocessed_input)
#top_1 = decode_predictions(predictions)[0][0]
# print('Predicted class:')
# print('%s (%s) with probability %.2f' % (top_1[1], top_1[0], top_1[2]))
predicted_class = np.argmax(predictions)
cam, heatmap = grad_cam(model, preprocessed_input, predicted_class, "conv2d_4")
# -
model.layers[0].layers
preprocessed_input.shape
predictions
top_1
# +
predictions = model.predict(X_test)
top_1 = decode_predictions(predictions)[0][0]
def on_epoch_end(self, epoch, logs={}):
clear_output(wait=True)
tacc = logs.get('val_acc')
trainstats.loc[len(trainstats)] = (logs.get('loss'), logs.get('val_loss'), tacc, logs.get('acc'))
confusion_matrix(model, X_test, Y_test, tacc, save=False)
pred = model.predict(np.expand_dims(im, 0))
# print('Predicted class:')
# print('%s (%s) with probability %.2f' % (top_1[1], top_1[0], top_1[2]))
# predicted_class = np.argmax(predictions)
# -
cam, heatmap = grad_cam(model, preprocessed_input, predicted_class, "block5_conv3")
img_path
X_test.shape
# +
def load_image(path):
from keras.preprocessing import image
import sys
#img_path = sys.argv[1]
img = image.load_img(img_path, target_size=(224, 224))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
#x = preprocess_input(x)
x= x/255
return x
img_cam= load_image(img_path)
# -
def visualize_class_activation_map(model_path, img_path, output_path):
from keras.models import load_model
from keras import model
model = model#load_model(model_path)
original_img = cv2.imread(img_path, 1)
width, height, _ = original_img.shape
#Reshape to the network input shape (3, w, h).
img = np.array([np.transpose(np.float32(original_img), (2, 0, 1))])
#Get the 512 input weights to the softmax.
class_weights = model.layers[-1].get_weights()[0]
final_conv_layer = get_output_layer(model, "block14_sepconv2_act")#get_output_layer(model, "block14_sepconv2_act") #changed
get_output = K.function([model.layers[0].input], \
[final_conv_layer.output,
model.layers[-1].output])
[conv_outputs, predictions] = get_output([img])
#[conv_outputs, predictions] = model.output([img])#get_output([img])
conv_outputs = conv_outputs[0, :, :, :]
#Create the class activation map.
cam = np.zeros(dtype = np.float32, shape = conv_outputs.shape[1:3])
target_class = 1
for i, w in enumerate(class_weights[:, target_class]):
cam += w * conv_outputs[i, :, :]
# +
model= model
img_path='../data/train/False/1.png'
origional_img=cv2.imread(img_path,1)
original_img = cv2.imread(img_path, 1)
width, height, _ = original_img.shape
#Reshape to the network input shape (3, w, h).
img = np.array([np.transpose(np.float32(original_img), (2, 0, 1))]) #(224, 224, 3)
class_weights = model.layers[-1].get_weights()[0]
final_conv_layer = get_output_layer(model, "block14_sepconv2_act")#get_output_layer(model, "block14_sepconv2_act") #changed #<tf.Tensor 'block14_sepconv2_act/Relu:0' shape=(?, ?, ?, 2048) dtype=float32>
# -
get_output = K.function(
[model.layers[0].input], #[<tf.Tensor 'input_1:0' shape=(?, ?, ?, 3) dtype=float32>]
[final_conv_layer.output, #[<tf.Tensor 'block14_sepconv2_act/Relu:0' shape=(?, ?, ?, 2048) dtype=float32>
model.layers[-1].output] # <tf.Tensor 'dense_2/Softmax:0' shape=(?, 2) dtype=float32>
)
get_output([img])
img.shape
#get_output([3,])
#[model.layers[0].input]
final_conv_layer.output
model.layers[-1].output
def get_output_layer(model, layer_name):
# get the symbolic outputs of each "key" layer (we gave them unique names).
layer_dict = dict([(layer.name, layer) for layer in model.layers])
layer = layer_dict[layer_name]
return layer
final_conv_layer
[np.transpose(np.float32(original_img), (2, 0, 1))]
img
[conv_outputs, predictions] = model.output([img])
# +
model_save= model
class_weights = model.layers[-1].get_weights()[0]
final_conv_layer = get_output_layer(model, "block14_sepconv2_act")
get_output = K.function([model.layers[0].input], \
[final_conv_layer.output,
model.layers[-1].output])
convout1_f = theano.function([model.get_input(train=False)], convout1.get_output(train=False))
[conv_outputs, predictions] = get_output([img])
conv_outputs = conv_outputs[0, :, :, :]
# -
final_conv_layer
layer.output
final_conv_layer
def get_output_layer(model, layer_name):
# get the symbolic outputs of each "key" layer (we gave them unique names).
layer_dict = dict([(layer.name, layer) for layer in model.layers])
layer = layer_dict[layer_name]
return layer
test_img=cv2.imread('../data/train/False/1.png', 1)
test_img.shape
img = np.array([np.transpose(np.float32(test_img), (2, 0, 1))])
img.shape
visualize_class_activation_map(modelid, '../data/train/False/1.png', '../data/')
# +
#%% Training code
##transfer learning
from IPython.display import clear_output
tm = time.time()
trainstats = pd.DataFrame(columns=('Train loss', 'Test loss', 'Accuracy', 'Train accuracy'))
epochs=50 ####change
from keras.callbacks import Callback
loss = model.fit_generator(
train_generator,
steps_per_epoch=int(len(train_generator.filenames)/batch_size),
validation_data=(X_test, Y_test),
validation_steps=1,
verbose=1, epochs=epochs,
use_multiprocessing=True,
workers=4,
callbacks=[VizTraining()]
)
# +
# loss
# -
loss
# # working heatmap
# +
##### Author: <NAME>, <NAME>, <NAME>, <NAME> #####
import os
import cv2
import sys
import glob
import math
import tempfile
import argparse
import numpy as np
import pandas as pd
from PIL import Image
from PIL import ImageFile
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import tensorflow as tf
from tensorflow import keras
import tensorflow.keras.backend as K
from tensorflow.keras.models import *
from tensorflow.keras import activations
from tensorflow.keras import optimizers
from tensorflow.keras.callbacks import *
from tensorflow.keras.preprocessing import image
from tensorflow.keras.utils import to_categorical
from tensorflow.python.keras.optimizers import Adam
from tensorflow.keras.layers import Dense,Input,Flatten, Dropout
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.python.keras.applications.inception_resnet_v2 import InceptionResNetV2, preprocess_input, decode_predictions
ImageFile.LOAD_TRUNCATED_IMAGES = True
os.environ["KMP_DUPLICATE_LIB_OK"]="TRUE"
os.environ["CUDA_VISIBLE_DEVICES"]="-1" # do not use gpu
os.environ['TF_CPP_MIN_LOG_LEVEL'] ="3" # Supress tensorflow warning
# # Fix tensorflow GPU allocation
# #%% GPU memory fix
# def get_session(gpu_fraction=0.5):
# gpu_options = tf.GPUOptions(per_process_gpu_memory_fraction=gpu_fraction, allow_growth=True)
# return tf.Session(config=tf.ConfigProto(gpu_options=gpu_options))
# keras.backend.set_session(get_session())
# # Qucik check if it is using GPU
# from tensorflow.python.client import device_lib
# def get_available_gpus():
# local_device_protos = device_lib.list_local_devices()
# return [x.name for x in local_device_protos if x.device_type == 'GPU']
# print("Tensorflow Version:", tf.__version__)
# print(tf.keras.__version__)
# print(get_available_gpus())
def find_layer_idx(model, layer_name):
"""Looks up the layer index corresponding to `layer_name` from `model`.
Args:
model: The `keras.models.Model` instance.
layer_name: The name of the layer to lookup.
Returns:
The layer index if found. Raises an exception otherwise.
"""
layer_idx = None
for idx, layer in enumerate(model.layers):
if layer.name == layer_name:
layer_idx = idx
break
if layer_idx is None:
raise ValueError("No layer with name '{}' within the model".format(layer_name))
return layer_idx
def apply_modifications(model, custom_objects=None):
"""Applies modifications to the model layers to create a new Graph. For example, simply changing
`model.layers[idx].activation = new activation` does not change the graph. The entire graph needs to be updated
with modified inbound and outbound tensors because of change in layer building function.
Args:
model: The `keras.models.Model` instance.
Returns:
The modified model with changes applied. Does not mutate the original `model`.
"""
# The strategy is to save the modified model and load it back. This is done because setting the activation
# in a Keras layer doesnt actually change the graph. We have to iterate the entire graph and change the
# layer inbound and outbound nodes with modified tensors. This is doubly complicated in Keras 2.x since
# multiple inbound and outbound nodes are allowed with the Graph API.
model_path = os.path.join(tempfile.gettempdir(), next(tempfile._get_candidate_names()) + '.h5')
try:
model.save(model_path)
return load_model(model_path, custom_objects=custom_objects)
finally:
os.remove(model_path)
def convert_model(current_model):
# Utility to search for layer index by name.
# Alternatively we can specify this as -1 since it corresponds to the last layer.
layer_idx = find_layer_idx(current_model, 'dense_2')
# Swap softmax with linear
current_model.layers[layer_idx].activation = activations.linear
new_model = apply_modifications(current_model)
return new_model
def visualize_cam(model, img_path, conv_layer = 'conv_7b', size = (300,300), hif = .8, bw = False):
original_img = cv2.imread(img_path, 3)
img = image.load_img(img_path, target_size=size)
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
x = x/255
# Prediction
preds = model.predict(x)
print(preds)
argmax = np.argmax(preds[0])
output = model.output[:, argmax]
print("* Predicted class for color: ", argmax)
last_conv_layer = model.get_layer(conv_layer)
grads = K.gradients(output, last_conv_layer.output)[0]
pooled_grads = K.mean(grads, axis=(0, 1, 2))
iterate = K.function([model.input], [pooled_grads, last_conv_layer.output[0]])
pooled_grads_value, conv_layer_output_value = iterate([x])
for i in range(last_conv_layer.output_shape[3]):
conv_layer_output_value[:, :, i] *= pooled_grads_value[i]
heatmap = np.mean(conv_layer_output_value, axis=-1)
heatmap = np.maximum(heatmap, 0)
heatmap /= np.max(heatmap)
heatmap = cv2.resize(heatmap, (original_img.shape[1], original_img.shape[0]))
heatmap = np.uint8(255 * heatmap)
heatmap = cv2.applyColorMap(heatmap, cv2.COLORMAP_JET)
superimposed_img = heatmap * hif + original_img
if bw:
# grayscale
original_img_bw = cv2.imread(img_path, 0)
img_bw = cv2.resize(original_img_bw,(300,300))
img_bw = cv2.cvtColor(img_bw, cv2.COLOR_GRAY2RGB);
x_bw = image.img_to_array(img_bw)
x_bw = np.expand_dims(x_bw, axis=0)
x_bw = preprocess_input(x_bw)
x_bw = x_bw/255
# Prediction
preds_bw = model.predict(x_bw)
print(preds_bw)
argmax_bw = np.argmax(preds_bw[0])
output_bw = model.output[:, argmax_bw]
print("* Predicted class for bw: ", argmax_bw)
last_conv_layer = model.get_layer(conv_layer)
grads_bw = K.gradients(output_bw, last_conv_layer.output)[0]
pooled_grads_bw = K.mean(grads_bw, axis=(0, 1, 2))
iterate = K.function([model.input], [pooled_grads_bw, last_conv_layer.output[0]])
pooled_grads_value_bw, conv_layer_output_value_bw = iterate([x])
for i in range(last_conv_layer.output_shape[3]):
conv_layer_output_value_bw[:, :, i] *= pooled_grads_value_bw[i]
heatmap_bw = np.mean(conv_layer_output_value_bw, axis=-1)
heatmap_bw = np.maximum(heatmap_bw, 0)
heatmap_bw /= np.max(heatmap_bw)
heatmap_bw = cv2.resize(heatmap_bw, (original_img_bw.shape[1], original_img_bw.shape[0]))
heatmap_bw = np.uint8(255 * heatmap_bw)
heatmap_bw = cv2.applyColorMap(heatmap_bw, cv2.COLORMAP_BONE)
original_img_bw = cv2.cvtColor(original_img_bw, cv2.COLOR_GRAY2RGB);
superimposed_img_bw = heatmap_bw * hif + original_img_bw
return [original_img, superimposed_img, superimposed_img_bw]
else:
return [original_img, superimposed_img, 0]
def get_args():
parser = argparse.ArgumentParser()
parser.add_argument("--image_path", type = str, help = "Path of an image to run the network on")
parser.add_argument("--output_path", type = str, default = "./", help = "Output path + basename")
parser.add_argument("--model_path", type = str, default = "./model.h5", help = "Path of the trained (and converted) model")
parser.add_argument("--convert_model", type = bool, default = False, help = 'Convert model softmax layer to linear layer?')
parser.add_argument("--output_all", type = bool, default = False, help = 'Output all images (original+color+bw)?')
parser.add_argument("--conv_layer_name", type = str, default = 'conv', help = 'Name of last convolutional layer?')
parser.add_argument("--image_size", type = int, default = 300, help = 'Image size of model (an int)?')
parser.add_argument("--hif", type = float, default = .8, help = 'Heatmap factor?')
args = parser.parse_args()
return args
# -
cmodel = convert_model(model)
# +
# model_path=modelid
# model=model
# output_path="../misc/testingF"
# img_size=224
# image_path='../data/test/False/19.png' #'../data/train/False/1.png'
# conv_layer_name='block14_sepconv2_act'
# convert=True
# hif=0.6
# output_all=True
# # print('* Loading model from: ', model_path)
# # model = model#modload_model(model_path)
# img_size = img_size
# imgs = visualize_cam(cmodel, image_path, conv_layer = conv_layer_name, size = (img_size,img_size), hif = hif, bw = True)
# print('* Saving color heatmap at: ', output_path + '_heatmap_color.jpg')
# cv2.imwrite(output_path + '_heatmap_color.jpg', imgs[1])
# if output_all==True:
# print('* Saving original image at: ', output_path + '_original.jpg')
# cv2.imwrite(output_path + '_original.jpg', imgs[0])
# print('* Saving grayscale heatmap at: ', output_path + '_heatmap_bw.jpg')
# cv2.imwrite(output_path + '_heatmap_bw.jpg', imgs[2])
# +
model_path=modelid
model=model
# output_path="../misc/19"
img_size=224
# image_path='../data/test/True/19.png' #'../data/train/False/1.png'
conv_layer_name='block14_sepconv2_act'
convert=True
hif=0.7
output_all=True
for element in glob.glob("../data/test/True/"+"*"):
file_num=element.strip(".png").split('/')[-1]
image_path= element
output_path="../misc/no_aug_true/{}".format(file_num)
img_size = img_size
imgs = visualize_cam(cmodel, image_path, conv_layer = conv_layer_name, size = (img_size,img_size), hif = hif, bw = False)
print('* Saving color heatmap at: ', output_path + '_heatmap_color.jpg')
cv2.imwrite(output_path + '_heatmap_color.jpg', imgs[1])
if output_all==True:
print('* Saving original image at: ', output_path + '_original.jpg')
cv2.imwrite(output_path + '_original.jpg', imgs[0])
print('* Saving grayscale heatmap at: ', output_path + '_heatmap_bw.jpg')
cv2.imwrite(output_path + '_heatmap_bw.jpg', imgs[2])
# +
model_path=modelid
model=model
# output_path="../misc/19"
img_size=224
# image_path='../data/test/True/19.png' #'../data/train/False/1.png'
conv_layer_name='block14_sepconv2_act'
convert=True
hif=0.7
output_all=True
for element in glob.glob("../data/test/False/"+"*"):
file_num=element.strip(".png").split('/')[-1]
image_path= element
output_path="../misc/no_aug_false/{}".format(file_num)
img_size = img_size
imgs = visualize_cam(cmodel, image_path, conv_layer = conv_layer_name, size = (img_size,img_size), hif = hif, bw = False)
print('* Saving color heatmap at: ', output_path + '_heatmap_color.jpg')
cv2.imwrite(output_path + '_heatmap_color.jpg', imgs[1])
if output_all==True:
print('* Saving original image at: ', output_path + '_original.jpg')
cv2.imwrite(output_path + '_original.jpg', imgs[0])
print('* Saving grayscale heatmap at: ', output_path + '_heatmap_bw.jpg')
cv2.imwrite(output_path + '_heatmap_bw.jpg', imgs[2])
# -
# +
# str.strip?
| notebooks/cnn_server_template-vis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introductory Coding Packages
#
# This notebook includes several packages that can be used to support elementary coding acitivities within a Jupyter notebook context.
# ## A Simple Turtle
#
# The basic Python `turtle` package does not work simply with Jupyter notebooks, and the [`ipython-turtle-widget`](https://github.com/gkvoelkl/ipython-turtle-widget) seems to have stopped working with recent versions of the notebooks. (*It would be a Good Thing to create a standard IPython turtle with Jupyter notebook and Jupyterlab integration.*)
#
# The `calysto` package includes a simple canvas with Python turtle like commands.
# %%capture
# !pip install calysto
#via https://jupyter.brynmawr.edu/services/public/dblank/Experiments/Calysto%20Turtle%20Graphics.ipynb
from calysto.graphics import *
from calysto.display import display, clear_output
import time
import math
# There is some boilerplate that the basic Python `turtle` packages, but we can still write some turtle like programs:
# +
canvas = Canvas(size=(400, 400))
turtle1 = Turtle(canvas, (200, 200), 0)
turtle2 = Turtle(canvas, (197, 200), 180)
turtle2.stroke = Color(255, 0, 0)
for i in range(100):
turtle1.left(45)
turtle1.forward(1 + i)
turtle2.left(45)
turtle2.forward(1 + i)
clear_output(wait=True)
display(canvas)
time.sleep(.1)
# -
# We can start to fudge the calysto code to make it a bit simpler:
# +
#tree via https://github.com/psychemedia/showntell/blob/computing/index_computing.ipynb
canvas = Canvas(size=(150, 150))
canvas.clear()
turtle = Turtle(canvas, (100, 100), 0)
left = turtle.left
right = turtle.right
forward = turtle.forward
backward = turtle.backward
# -
# We can now call on these commands directly, although we still have to manage the canvas.
# +
def tree(length):
"""Draw a symmetric tree with a trunk of the given length."""
# don't draw very small trees
if length < 5:
return
# draw the trunk
forward(length)
# draw the left branch, which is just a smaller tree
left(20)
tree(0.8 * length)
right(20)
# draw the right branch, which is just a smaller tree
right(20)
tree(0.8 * length)
left(20)
# return the turtle to the start position
backward(length)
clear_output(wait=True)
display(canvas)
time.sleep(.2)
display(canvas)
left(90)
tree(20)
# -
# ## Blocks Style Programming with `Jigsaw`
#
# Many students are introduced to programming through blocks style interfaces. The `jigsaw` magic that is part of the `metakernel` python package provides
#magics
# !pip install metakernel
import metakernel
metakernel.register_ipython_magics()
# A newly created programme is saved to a local file related to the workspace name.
#
# Programmes are constructed by dragging programme blocks from the menu on the left onto the main canvas.
#
# (Use the + and - controls on the canvas to set the zoom level.)
#
# Run the programme by clicking the `Run` button just below the canvas.
#
# The `Generate Python Code` buttom will create a notebook code cell underneath the `jigsaw` cell containing python code equivalent to the programme constructed on the canvas.
#
# If several disconnected blocks are placed on the canvas, the code equivalents are generated in top down order.
# %jigsaw Python --workspace mynewProgram
# The blocks programme is saved as an .xml file. This means the notebook session can be closed and the blockly programme reopened in the same - or different - notebook at a later date: if an appropriate .xml file with the same name as the workspace exists, the corresponding programme will be loaded onto the canvas.
# ## Variable Inspector
#
#
# A variable inspector panel, from the[`varInspector`](../nbextensions/?nbextension=varInspector/main) extension, can be selected from the toolbar to pop up a display showing the variables that have been defined in the current Python session and their values.
#
# Enable the extension and then refresh this notebook webpage to see the toolbar button and pop open the floating variable inspector panel.
# ## `nbtutor`
#
# The [`nbtutor`](https://github.com/jorisvandenbossche/nbtutor) package provides a simple step tracer / debugger that can visualise the execution of a series of staments in a Jupyter notebook code cell.
# +
# %%capture
#https://github.com/lgpage/nbtutor
# !pip install nbtutor
# !jupyter nbextension install --overwrite --user --py nbtutor
# !jupyter nbextension enable --py nbtutor
# -
# %load_ext nbtutor
# When you run the following cell, a menu will appear in a toolbar that appears at the top of the cell. Select the `Memory` option and then use the `Next>` button to step through the command exeuction steps.
#
# Hide the cell toolbars from the notebook `View` menu: `View > Cell Toolbar > None`.
# +
# %%nbtutor -r -f
import math
def multi(x, y):
tmp=x * y
return y
xy=multi(3, 5)
print(xy)
| Getting Started With Notebooks/3.4.0 Introductory Coding Packages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 설치
# +
# #!pip install numpy
# -
import numpy as np
# + [markdown] tags=[]
# # 생성 및 shape 속성
# -
list1 = [1, 2, 3, 4]
a = np.array(list1)
print(a.shape) # (4, )
b = np.array([[1,2,3],[4,5,6]])
print(b.shape) # (2, 3)
# # numpy 슬라이싱
lst = [
[1, 2, 3],
[4, 5, 6],
[7, 8, 9]
]
arr = np.array(lst)
# 슬라이스
a = arr[0:2, 0:2]
print(a)
# 슬라이스
a = arr[1:3, 1:3]
print(a)
# 슬라이스
a = arr[:, 0:2]
print(a)
# 슬라이스
a = arr[0:2, :]
print(a)
# + [markdown] tags=[]
# # 배열 shape 바꾸기
# -
a = np.array([[51,55],[14,19],[0,4]])
print(a)
print(a.shape)
a = a.reshape(2,3)
print(a)
print(a.shape)
a = a.flatten() # 1차원으로 바꾸기
print(a)
print(a.shape)
a = a.reshape(-1,3)
print(a)
print(a.shape)
a = a.reshape(-1,2)
print(a)
print(a.shape)
a = a.reshape(2, -1)
print(a)
print(a.shape)
a = a.reshape(-1)
print(a)
print(a.shape)
# # numpy random
# - `numpy.random.randint`: 균일 분포의 정수 난수 1개 생성 (int)
# - `numpy.random.rand` : 0~1 사이의 균일 분포에서 난수 matrix 생성 (float)
# - `numpy.random.randn` : 가우시안 표준 정규분포에서 난수 matrix 생성 (float)
import numpy as np
# ## numpy.random.randint: 균일 분포의 정수 난수 1개 생성 (int)
np.random.randint(6) # 0 ~ 5 사이의 값중 1개
# np.randome.randint(0,6)
np.random.randint(1,20) # 1~19 까지 랜덤 숫자 1개
# ## numpy.random.rand : 0~1 사이의 균일 분포에서 난수 matrix 생성 (float)
np.random.rand(6)
np.random.rand(2,3)
# numpy.random.rand : 0~1 사이의 균일 분포에서 난수 matrix 생성 (float)
np.random.randn(6)
np.random.randn(3,2)
| practice/day4/numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Get the Data
import os
import tarfile
import urllib
# +
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/ageron/handson-ml2/master/"
HOUSING_PATH = os.path.join("datasets", "housing")
HOUSING_URL = DOWNLOAD_ROOT + 'datasets/housing/housing.tgz'
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
os.makedirs(housing_path, exist_ok=True)
tgz_path = os.path.join(housing_path, "housing.tgz")
urllib.request.urlretrieve(housing_url, tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
# +
import pandas as pd
fetch_housing_data()
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
# -
housing = load_housing_data()
#housing = pd.read_csv("housing.csv")
housing.head()
# # Explore the data
housing.info()
# +
# to find out what are the unique values for column ocean_proximity, in other words, what are the catergorical values
housing['ocean_proximity'].value_counts()
# -
housing.describe()
# +
import matplotlib.pyplot as plt
housing.hist(bins=50, figsize=(20,15))
plt.show()
# +
# let's set aside some test data before analysing any further to reduce the risk of biases
from sklearn.model_selection import train_test_split
train_set, test_set = train_test_split(housing, test_size=0.2, random_state=42)
# +
# It is very useful to have enough representation of all the categories in the training data for a model to learn properly
# So let's split the Dataframe to categorize the median income to represent the more normally distributed portion (1.5-6)
import numpy as np
housing['income_cat'] = pd.cut(housing['median_income'],
bins=[0., 1.5, 3.0, 4.5, 6., np.inf],
labels=[1, 2, 3, 4, 5])
housing['income_cat'].hist()
# +
# now we can create stratafied sameples of the Dataframe, essentially to create a more robust train and test set
from sklearn.model_selection import StratifiedShuffleSplit
split = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_index, test_index in split.split(housing, housing['income_cat']):
strat_train_set = housing.loc[train_index]
strat_test_set = housing.loc[test_index]
# +
#income category proportions in the stratified test set
strat_test_set['income_cat'].value_counts() / len(strat_test_set)
# +
#income category proportions in the random test set
test_set['income_cat'].value_counts() / len(test_set)
# +
#income category proportions in the overall set
# as in above, the stratified one closely matches with the overall set
housing['income_cat'].value_counts() / len(housing)
# +
# so not to confuse the ML algorithm with 'income_cat', let's drop that from the train and test sets
for set_ in (strat_train_set, strat_test_set):
set_.drop('income_cat', axis=1, inplace=True)
# -
housing_t = strat_train_set.copy()
housing_t.plot(kind='scatter', x='longitude', y='latitude')
# +
# Let's visualize the same but locate how dense the data points are around the entire state of California
housing_t.plot(kind='scatter', x='longitude', y='latitude', alpha=0.1) #this is for density
# +
# Let's plot one for housing prices with option cmap called jet
# defining districts population (option s) as the radius of each circle
# defining price with the colour, with blue representing low values and red representing high ones
housing_t.plot(kind='scatter', x='longitude', y='latitude', alpha=0.4,
s=housing_t['population']/100, label='population', figsize=(10,7),
c='median_house_value', cmap=plt.get_cmap('jet'), colorbar=True,
)
plt.legend()
# +
# Let's look at the correlations between median_house_value with each of the factors
corr_matrix = housing_t.corr()
corr_matrix['median_house_value'].sort_values(ascending=False)
# +
#scatterplot for the interesting ones
from pandas.plotting import scatter_matrix
attributes = ['median_house_value', 'median_income', 'total_rooms', 'housing_median_age']
scatter_matrix(housing_t[attributes], figsize=(12, 8))
# +
# looking at the median_income since it seems to be the most promising one
housing_t.plot(kind='scatter', x='median_income', y='median_house_value', alpha=0.2)
# +
# one interesting thing to notice is the horizontal lines along $450k and $350k and possibly $280k and some may be below that
# it might be worth discarding those districts to prevent the algorithms from learing to reproduce these data quirks
# +
# Let's create some useful and interesting attributes out of the no so useful individual ones
housing_t["rooms_per_household"] = housing_t["total_rooms"]/housing_t["households"]
housing_t["bedrooms_per_room"] = housing_t["total_bedrooms"]/housing_t["total_rooms"]
housing_t["population_per_household"]=housing_t["population"]/housing_t["households"]
# +
# and checkout their correlation to the 'median_house_value'
corr_matrix = housing_t.corr()
corr_matrix['median_house_value'].sort_values(ascending=False)
# -
# # Clean the data
housing = strat_train_set.drop('median_house_value', axis=1)
housing_labels = strat_train_set['median_house_value'].copy()
housing
# +
# lets deal with the missing values
# the following ways are the usuals
# housing.dropna(subset=["total_bedrooms"]) # gets rid of the corresponding districts
# housing.drop("total_bedrooms", axis=1) # gets rid of the whole attribute
# median = housing["total_bedrooms"].median() # sets the values to the median (or mean/zero etc.)
# housing["total_bedrooms"].fillna(median, inplace=True)”
# However let's use a scikit-learn transformer function to make it easier
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy='median')
# +
# Let's drop 'ocean_proximity' to only address the numerical ones for the median
housing_num = housing.drop('ocean_proximity', axis=1)
# +
# Let's fit imputer with the df
imputer.fit(housing_num)
# -
imputer.statistics_
housing_num.median().values
# +
# Now let's transform housing_num with imputer (replacing all the missing values with the learned medians)
X = imputer.transform(housing_num)
# -
X
# +
# let's put X back into a df
housing_tr = pd.DataFrame(X, columns=housing_num.columns,
index=housing_num.index)
# -
housing_tr
housing_tr.info()
# +
# Now let's deal with the ocean_proximity attribute by converting it's categorical values to a numerical value
housing_cat = housing[['ocean_proximity']]
housing_cat.head(10)
# +
# Let's use OrdinalEncoder (every category gets a number in no specific order)
from sklearn.preprocessing import OrdinalEncoder
ordinal_encoder = OrdinalEncoder()
housing_cat_encoded = ordinal_encoder.fit_transform(housing_cat)
housing_cat_encoded[0:10]
# +
# categories
ordinal_encoder.categories_
# +
# Let's try OneHotEncoder (every category gets a vector of 0 [cold] and 1 [hot])
from sklearn.preprocessing import OneHotEncoder
cat_encoder = OneHotEncoder()
housing_cat_1hot = cat_encoder.fit_transform(housing_cat)
housing_cat_1hot
# -
housing_cat_1hot.toarray()
# +
# Let's build a custom transformer for adding the combined attributes
# add_bedrooms_per_room is a hyperparameter
from sklearn.base import BaseEstimator, TransformerMixin
rooms_ix, bedrooms_ix, population_ix, households_ix = 3, 4, 5, 6
class CombinedAttributesAdder(BaseEstimator, TransformerMixin):
def __init__(self, add_bedrooms_per_room = True): # no *args or **kargs
self.add_bedrooms_per_room = add_bedrooms_per_room
def fit(self, X, y=None):
return self # nothing else to do
def transform(self, X, y=None):
rooms_per_household = X[:, rooms_ix] / X[:, households_ix]
population_per_household = X[:, population_ix] / X[:, households_ix]
if self.add_bedrooms_per_room:
bedrooms_per_room = X[:, bedrooms_ix] / X[:, rooms_ix]
return np.c_[X, rooms_per_household, population_per_household,
bedrooms_per_room]
else:
return np.c_[X, rooms_per_household, population_per_household]
attr_adder = CombinedAttributesAdder(add_bedrooms_per_room=False)
housing_extra_attribs = attr_adder.transform(housing.values)
# -
housing_extra_attribs
# +
# Transformer Pipelines (with functions from Scikit-learn)
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
num_pipeline = Pipeline([
('imputer', SimpleImputer(strategy="median")),
('attribs_adder', CombinedAttributesAdder()),
('std_scaler', StandardScaler()),
])
housing_num_tr = num_pipeline.fit_transform(housing_num)
# -
housing_num_tr
# +
# Now let's build the full pipeline that incorporates the num_pipeline along with categorical OnehotEncoder transformation
from sklearn.compose import ColumnTransformer
num_attribs = list(housing_num) #list of the columns
cat_attribs = ['ocean_proximity']
full_pipeline = ColumnTransformer([
('num', num_pipeline, num_attribs),
('cat', OneHotEncoder(), cat_attribs)
])
housing_prepared = full_pipeline.fit_transform(housing)
# -
housing_prepared
# # Select and Train a Model
# ### Linear Regression Model
# +
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(housing_prepared, housing_labels)
# +
some_data = housing.iloc[:5]
some_labels = housing_labels.iloc[:5]
some_data_prepared = full_pipeline.transform(some_data)
print("Predictionns:", lin_reg.predict(some_data_prepared))
print("Actuals:", list(some_labels))
# -
some_data_prepared
# +
# Let's measure the RMSE
from sklearn.metrics import mean_squared_error
housing_predictions = lin_reg.predict(housing_prepared)
lin_mse = mean_squared_error(housing_labels, housing_predictions) # squared=False would return RMSE as well
lin_rmse = np.sqrt(lin_mse)
lin_rmse
# -
# ### Decesion Tree Regressor
# +
from sklearn.tree import DecisionTreeRegressor
tree_reg = DecisionTreeRegressor()
tree_reg.fit(housing_prepared, housing_labels)
# -
print("Predictionns:", tree_reg.predict(some_data_prepared))
print("Actuals:", list(some_labels))
# +
# let’s evaluate it on the training set
housing_predictions = tree_reg.predict(housing_prepared)
tree_rmse = mean_squared_error(housing_labels, housing_predictions, squared=False)
tree_rmse
# +
# Since the above model is unbelivably accurate,, let's cross validate to find out any overfitting issue, if any
# Let's use Scikit-Learn's Kfold CV to split the training set into 10 folds and running the model on each of them
from sklearn.model_selection import cross_val_score
scores = cross_val_score(tree_reg, housing_prepared, housing_labels,
scoring='neg_mean_squared_error', cv=10)
tree_rmse_scores = np.sqrt(-scores) # '-' is used since values are in negative
# +
# Let's look at the result
def display_scores(scores):
print('Scores:', scores)
print('Mean:', scores.mean())
print('Standard deviation:', scores.std())
display_scores(tree_rmse_scores)
# +
# Let's run the cross validation on the Linear Regression Model
lin_scores = cross_val_score(lin_reg, housing_prepared, housing_labels,
scoring='neg_mean_squared_error', cv=10)
lin_rmse_scores = np.sqrt(-lin_scores)
display_scores(lin_rmse_scores)
# -
# ### Random Forest Regressor
# +
from sklearn.ensemble import RandomForestRegressor
forest_reg = RandomForestRegressor()
forest_reg.fit(housing_prepared, housing_labels)
# -
print("Predictionns:", forest_reg.predict(some_data_prepared))
print("Actuals:", list(some_labels))
# +
housing_predictions = forest_reg.predict(housing_prepared)
forest_rmse = mean_squared_error(housing_labels, housing_predictions, squared=False)
forest_rmse
# +
forest_scores = cross_val_score(forest_reg, housing_prepared, housing_labels,
scoring='neg_mean_squared_error', cv=10)
forest_rmse_scores = np.sqrt(-forest_scores)
display_scores(forest_rmse_scores)
# -
# # Fine-Tune your Model
# ### Grid Search
# +
from sklearn.model_selection import GridSearchCV
param_grid = [
{'n_estimators': [3, 10, 30, 40], 'max_features': [2, 4, 6, 8, 10]},
{'bootstrap': [False], 'n_estimators': [3, 10, 30, 40], 'max_features': [2, 3, 4, 8, 10]},
]
forest_reg = RandomForestRegressor()
grid_search = GridSearchCV(forest_reg, param_grid, cv=5,
scoring='neg_mean_squared_error',
return_train_score=True)
grid_search.fit(housing_prepared, housing_labels)
# +
# Let's use the best_params_ attribute to find out the best hyperparameters
grid_search.best_params_
# +
# to see the evaluations scores
cvres = grid_search.cv_results_
for mean_score, params in zip(cvres['mean_test_score'], cvres['params']):
print(np.sqrt(-mean_score), params)
# +
# Let's find out the values of every feature to assess how important they are
feature_importances = grid_search.best_estimator_.feature_importances_
feature_importances
# -
extra_attribs = ["rooms_per_hhold", "pop_per_hhold", "bedrooms_per_room"]
cat_encoder = full_pipeline.named_transformers_["cat"]
cat_one_hot_attribs = list(cat_encoder.categories_[0])
attributes = num_attribs + extra_attribs + cat_one_hot_attribs
sorted(zip(feature_importances, attributes), reverse=True)
# # Evaluation on the Test set
# +
final_model = grid_search.best_estimator_
X_test = strat_test_set.drop('median_house_value', axis=1)
y_test = strat_test_set['median_house_value'].copy()
X_test_prepared = full_pipeline.transform(X_test)
final_predictions = final_model.predict(X_test_prepared)
final_rmse = mean_squared_error(y_test, final_predictions, squared=False)
final_rmse
# +
# Let's compute a 95% confidence interval for the error
from scipy import stats
confidence = 0.95
squared_errors = (final_predictions - y_test) ** 2
np.sqrt(stats.t.interval(confidence, len(squared_errors) - 1,
loc=squared_errors.mean(),
scale=stats.sem(squared_errors)))
# -
# # Exercises
# ### Trying a Support Vector Machine Regressor
# +
from sklearn.svm import SVR
svr_reg = SVR()
svr_reg.fit(housing_prepared, housing_labels)
# -
print("Predictionns:", svr_reg.predict(some_data_prepared))
print("Actuals:", list(some_labels))
# +
housing_predictions = svr_reg.predict(housing_prepared)
svr_rmse = mean_squared_error(housing_labels, housing_predictions, squared=False)
svr_rmse
# +
svr_scores = cross_val_score(svr_reg, housing_prepared, housing_labels,
scoring='neg_mean_squared_error', cv=10)
svr_rmse_scores = np.sqrt(-svr_scores)
display_scores(svr_rmse_scores)
# +
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import expon, reciprocal
# Note: gamma is ignored when kernel is "linear"
param_distribs = {
"kernel": ['linear', 'rbf'],
'C': reciprocal (20, 200000),
'gamma': expon(scale=1.0),
}
svm_reg = SVR()
rnd_search = RandomizedSearchCV(svm_reg, param_distributions=param_distribs,
n_iter=50, cv=5, scoring='neg_mean_squared_error',
verbose=2, random_state=42)
rnd_search.fit (housing_prepared, housing_labels)
# -
negative_mse = rnd_search.best_score_
rmse = np.sqrt(-negative_mse)
rmse
rnd_search.best_params_
# ### Custom Transformer to select the most important attributes
# +
from sklearn.base import BaseEstimator, TransformerMixin
def indices_of_top_k(arr, k):
return np.sort(np.argpartition(np.array(arr), -k)[-k:])
class TopFeatureSelector(BaseEstimator, TransformerMixin):
def __init__(self, feature_importances, k):
self.feature_importances = feature_importances
self.k = k
def fit(self, X, y=None):
self.feature_indices_ = indices_of_top_k(self.feature_importances, self.k)
return self
def transform(self, X):
return X[:, self.feature_indices_]
# +
# top features from the randomforrest we found earlier
k = 5
top_k_feature_indices = indices_of_top_k(feature_importances, k)
top_k_feature_indices
# -
np.array(attributes)[top_k_feature_indices]
sorted(zip(feature_importances, attributes), reverse=True)[:k]
# +
# Let's create a new pipeline and integrate the top feature selection within it
preparation_and_feature_selection_pipeline = Pipeline([
('preparation', full_pipeline),
('feature_selection', TopFeatureSelector(feature_importances, k))
])
# +
# Let's run this on the housing_t dataframe
housing_prepared_top_k_features = preparation_and_feature_selection_pipeline.fit_transform(housing)
# -
housing_prepared_top_k_features[0:3]
# +
# Let's double check with the housing_prepared
housing_prepared[0:3, top_k_feature_indices]
# -
# ### Final pipeline incorporating all the steps
prepare_select_and_predict_pipeline_SVR = Pipeline([
('preparation', full_pipeline),
('feature_selection', TopFeatureSelector(feature_importances, k)),
('svm_reg', SVR(**rnd_search.best_params_))
])
prepare_select_and_predict_pipeline_SVR.fit(housing, housing_labels)
# +
some_data = housing.iloc[:4]
some_labels = housing_labels.iloc[:4]
print("Predictions:\t", prepare_select_and_predict_pipeline_SVR.predict(some_data))
print("Labels:\t\t", list(some_labels))
# -
prepare_select_and_predict_pipeline_forest = Pipeline([
('preparation', full_pipeline),
('feature_selection', TopFeatureSelector(feature_importances, k)),
('forest_reg', RandomForestRegressor(**{'bootstrap': False, 'max_features': 5, 'n_estimators': 40}))
])
prepare_select_and_predict_pipeline_forest.fit(housing, housing_labels)
print("Predictions:\t", prepare_select_and_predict_pipeline_forest.predict(some_data))
print("Labels:\t\t", list(some_labels))
# ### Automatically explore some data prep options using Grid Search
# +
# handle_unknow is set to ignore in order to ignore any errors related to the one item in the Island category in case it pops up in the test set
full_pipeline.named_transformers_["cat"].handle_unknown = 'ignore'
param_grid = [{
'preparation__num__imputer__strategy': ['mean', 'median', 'most_frequent'],
'feature_selection__k': list(range(1, len(feature_importances) + 1))
}]
grid_search_prep = GridSearchCV(prepare_select_and_predict_pipeline, param_grid, cv=5,
scoring='neg_mean_squared_error', verbose=2)
grid_search_prep.fit(housing, housing_labels)
# -
grid_search_prep.best_params_
| Chapter 2 - End to End Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={}
# # Submitting and Managing Jobs
#
# Launch this tutorial in a Jupyter Notebook on Binder:
# [](https://mybinder.org/v2/gh/htcondor/htcondor-python-bindings-tutorials/master?urlpath=lab/tree/Submitting-and-Managing-Jobs.ipynb)
# -
# ## What is HTCondor?
#
# An HTCondor pool provides a way for you (as a user) to submit units of work, called **jobs**, to be executed on a distributed network of computing resources.
# HTCondor provides tools to monitor your jobs as they run, and make certain kinds of changes to them after submission, which we call "managing" jobs.
#
# In this tutorial, we will learn how to submit and manage jobs *from Python*.
# We will see how to submit jobs with various toy executables, how to ask HTCondor for information about them, and how to tell HTCondor to do things with them.
# All of these things are possible from the command line as well, using tools like `condor_submit`, `condor_qedit`, and `condor_hold`.
# However, working from Python instead of the command line gives us access to the full power of Python to do things like generate jobs programmatically based on user input, pass information consistently from submission to management, or even expose an HTCondor pool to a web application.
# We start by importing the HTCondor Python bindings modules, which provide the functions we will need to talk to HTCondor.
# + pycharm={}
import htcondor # for submitting jobs, querying HTCondor daemons, etc.
import classad # for interacting with ClassAds, HTCondor's internal data format
# + [markdown] pycharm={}
# ## Submitting a Simple Job
#
# To submit a job, we must first describe it.
# A submit description is held in a `Submit` object.
# `Submit` objects consist of key-value pairs, and generally behave like Python dictionaries.
# If you're familiar with HTCondor's submit file syntax, you should think of each line in the submit file as a single key-value pair in the `Submit` object.
#
# Let's start by writing a `Submit` object that describes a job that executes the `hostname` command on an execute node, which prints out the "name" of the node.
# Since `hostname` prints its results to standard output (stdout), we will capture stdout and bring it back to the submit machine so we can see the name.
# + pycharm={}
hostname_job = htcondor.Submit({
"executable": "/bin/hostname", # the program to run on the execute node
"output": "hostname.out", # anything the job prints to standard output will end up in this file
"error": "hostname.err", # anything the job prints to standard error will end up in this file
"log": "hostname.log", # this file will contain a record of what happened to the job
"request_cpus": "1", # how many CPU cores we want
"request_memory": "128MB", # how much memory we want
"request_disk": "128MB", # how much disk space we want
})
print(hostname_job)
# + [markdown] pycharm={}
# The available descriptors are documented in the `condor_submit` [manual](https://htcondor.readthedocs.io/en/latest/man-pages/condor_submit.html).
# The keys of the Python dictionary you pass to `htcondor.Submit` should be the same as for the submit descriptors, and the values should be **strings containing exactly what would go on the right-hand side**.
#
# Note that we gave the `Submit` object several relative filepaths.
# These paths are relative to the directory containing this Jupyter notebook (or, more generally, the current working directory).
# When we run the job, you should see those files appear in the file browser on the left as HTCondor creates them.
#
# Now that we have a description, let's submit a job.
# To do so, we must ask the HTCondor scheduler to open a transaction.
# Once we have the transaction, we can "queue" (i.e., submit) a job via the `Submit` object.
# + pycharm={}
schedd = htcondor.Schedd() # get the Python representation of the scheduler
with schedd.transaction() as txn: # open a transaction, represented by `txn`
cluster_id = hostname_job.queue(txn) # queues one job in the current transaction; returns job's ClusterId
print(cluster_id)
# -
# The integer returned by the `queue` method is the `ClusterId` for the submission.
# It uniquely identifies this submission.
# Later in this module, we will use it to ask the HTCondor scheduler for information about our jobs.
#
# It isn't important to understand the transaction mechanics for now; think of it as boilerplate.
# (There are advanced use cases where it might be useful.)
#
# For now, our job will hopefully have finished running.
# You should be able to see the files in the file browser on the left.
# Try opening one of them and seeing what's inside.
#
# We can also look at the output from inside Python:
# +
import os
import time
output_path = "hostname.out"
# this is a crude way to wait for the job to finish
# see the Advanced tutorial "Scalable Job Tracking" for better methods!
while not os.path.exists(output_path):
print("Output file doesn't exist yet; sleeping for one second")
time.sleep(1)
with open(output_path, mode = "r") as f:
print(f.read())
# -
# If you got some text, it worked!
#
# If the file never shows up, it means your job didn't run.
# You might try looking at the `log` or `error` files specified in the submit description to see if there is any useful information in them about why the job failed.
# ## Submitting Multiple Jobs
# + [markdown] pycharm={}
# By default, each `queue` will submit a single job.
# A more common use case is to submit many jobs at once, often sharing some base submit description.
# Let's write a new submit description which runs `sleep`.
#
# When we have multiple **jobs** in a single **cluster**, each job will be identified not just by its **ClusterId** but also by a **ProcID**.
# We can use the ProcID to separate the output and error files for each individual job.
# Anything that looks like `$(...)` in a submit description is a **macro**, a placeholder which will be "expanded" later by HTCondor into a real value for that particular job.
# The ProcID expands to a series of incrementing integers, starting at 0.
# So the first job in a cluster will have ProcID 0, the next will have ProcID 1, etc.
# +
sleep_job = htcondor.Submit({
"executable": "/bin/sleep",
"arguments": "10s", # sleep for 10 seconds
"output": "sleep-$(ProcId).out", # output and error for each job, using the $(ProcId) macro
"error": "sleep-$(ProcId).err",
"log": "sleep.log", # we still send all of the HTCondor logs for every job to the same file (not split up!)
"request_cpus": "1",
"request_memory": "128MB",
"request_disk": "128MB",
})
print(sleep_job)
# -
# We will submit 10 of these jobs.
# All we need to change from our previous `queue` call is to add the `count` keyword argument.
schedd = htcondor.Schedd()
with schedd.transaction() as txn:
cluster_id = sleep_job.queue(txn, count=10) # submit 10 jobs
print(cluster_id)
# Now that we have a bunch of jobs in flight, we might want to check how they're doing.
# We can ask the HTCondor scheduler about jobs by using its `query` method.
# We give it a **constraint**, which tells it which jobs to look for, and a **projection**, which tells it what information to return.
schedd.query(
constraint=f"ClusterId == {cluster_id}",
projection=["ClusterId", "ProcId", "Out"],
)
# There are a few things to notice here:
# - Depending on how long it took you to run the cell, you may only get a few of your 10 jobs in the query. Jobs that have finished **leave the queue**, and will no longer show up in queries. To see those jobs, you must use the `history` method instead, which behaves like `query`, but **only** looks at jobs that have left the queue.
# - The results may not have come back in ProcID-sorted order. If you want to guarantee the order of the results, you must do so yourself.
# - Attributes are often renamed between the submit description and the actual job description in the queue. See [the manual](https://htcondor.readthedocs.io/en/latest/classad-attributes/job-classad-attributes.html) for a description of the job attribute names.
# - The objects returned by the query are instances of `ClassAd`. ClassAds are the common data exchange format used by HTCondor. In Python, they mostly behave like dictionaries.
# ## Using Itemdata to Vary Over Parameters
#
# By varying some part of the submit description using the ProcID, we can change how each individual job behaves.
# Perhaps it will use a different input file, or a different argument.
# However, we often want more flexibility than that.
# Perhaps our input files are named after different cities, or by timestamp, or some other naming scheme that already exists.
#
# To use such information in the submit description, we need to use **itemdata**.
# Itemdata lets us pass arbitrary extra information when we queue, which we can reference with macros inside the submit description.
# This lets use the full power of Python to generate the submit descriptions for our jobs.
#
# Let's mock this situation out by generating some files with randomly-chosen names.
# We'll also switch to using `pathlib.Path`, Python's more modern file path manipulation library.
# +
from pathlib import Path
import random
import string
import shutil
def random_string(length):
"""Produce a random lowercase ASCII string with the given length."""
return "".join(random.choices(string.ascii_lowercase, k = length))
# make a directory to hold the input files, clearing away any existing directory
input_dir = Path.cwd() / "inputs"
shutil.rmtree(input_dir, ignore_errors = True)
input_dir.mkdir()
# make 5 input files
for idx in range(5):
rs = random_string(5)
input_file = input_dir / "{}.txt".format(rs)
input_file.write_text("Hello from job {}".format(rs))
# -
# Now we'll get a list of all the files we just created in the input directory.
# This is precisely the kind of situation where Python affords us a great deal of flexibility over a submit file: we can use Python instead of the HTCondor submit language to generate and inspect the information we're going to put into the submit description.
# +
input_files = list(input_dir.glob("*.txt"))
for path in input_files:
print(path)
# -
# Now we'll make our submit description.
# Our goal is just to print out the text held in each file, which we can do using `cat`.
#
# We will tell HTCondor to transfer the input file to the execute location by including it in `transfer_input_files`.
# We also need to call `cat` on the right file via `arguments`.
# Keep in mind that HTCondor will move the files in `transfer_input_files` directly to the scratch directory on the execute machine, so instead of the full path, we just need the file's "name", the last component of its path.
# `pathlib` will make it easy to extract this information.
# +
cat_job = htcondor.Submit({
"executable": "/bin/cat",
"arguments": "$(input_file_name)", # we will pass in the value for this macro via itemdata
"transfer_input_files": "$(input_file)", # we also need HTCondor to move the file to the execute node
"should_transfer_files": "yes", # force HTCondor to transfer files even though we're running entirely inside a container (and it normally wouldn't need to)
"output": "cat-$(ProcId).out",
"error": "cat-$(ProcId).err",
"log": "cat.log",
"request_cpus": "1",
"request_memory": "128MB",
"request_disk": "128MB",
})
print(cat_job)
# -
# The itemdata should be passed as a list of dictionaries, where the keys are the macro names to replace in the submit description.
# In our case, the keys are `input_file` and `input_file_name`, so should have a list of 10 dictionaries, each with two entries.
# HTCondor expects the input file list to be a comma-separated list of POSIX-style paths, so we explicitly convert our `Path` to a POSIX string.
# +
itemdata = [{"input_file": path.as_posix(), "input_file_name": path.name} for path in input_files]
for item in itemdata:
print(item)
# -
# Now we'll submit the jobs, using `queue_with_itemdata` instead of `queue`:
# +
schedd = htcondor.Schedd()
with schedd.transaction() as txn:
submit_result = cat_job.queue_with_itemdata(txn, itemdata = iter(itemdata)) # submit one job for each item in the itemdata
print(submit_result.cluster())
# -
# Note that `queue_with_itemdata` returns a "submit result", not just the ClusterId.
# The ClusterId can be retreived from the submit result with its `cluster()` method.
#
# Let's do a query to make sure we got the itemdata right (these jobs run fast, so you might need to re-run the jobs if your first run has already left the queue):
schedd.query(
constraint=f"ClusterId == {submit_result.cluster()}",
projection=["ClusterId", "ProcId", "Out", "Args", "TransferInput"],
)
# And let's take a look at all the output:
# +
# again, this is very crude - see the advanced tutorials!
while not len(list(Path.cwd().glob("cat-*.out"))) == len(itemdata):
print("Not all output files exist yet; sleeping for one second")
time.sleep(1)
for output_file in Path.cwd().glob("cat-*.out"):
print(output_file, "->", output_file.read_text())
# + [markdown] pycharm={}
# ## Managing Jobs
#
# Once a job is in queue, the scheduler will try its best to execute it to completion.
# There are several cases where you may want to interrupt the normal flow of jobs.
# Perhaps the results are no longer needed; perhaps the job needs to be edited to correct a submission error.
# These actions fall under the purview of **job management**.
#
# There are two `Schedd` methods dedicated to job management:
#
# * `edit()`: Change an attribute for a set of jobs.
# * `act()`: Change the state of a job (remove it from the queue, hold it, suspend it, etc.).
#
# The `act` method takes an argument from the `JobAction` enum.
# Commonly-used values include:
#
# * `Hold`: put a job on hold, vacating a running job if necessary. A job will stay in the hold
# state until told otherwise.
# * `Release`: Release a job from the hold state, returning it to Idle.
# * `Remove`: Remove a job from the queue. If it is running, it will stop running.
# This requires the execute node to acknowledge it has successfully vacated the job, so ``Remove`` may
# not be instantaneous.
# * `Vacate`: Cause a running job to be killed on the remote resource and return to the Idle state. With
# `Vacate`, jobs may be given significant time to cleanly shut down.
#
# To play with this, let's bring back our sleep submit description, but increase the sleep time significantly so that we have time to interact with the jobs.
# +
long_sleep_job = htcondor.Submit({
"executable": "/bin/sleep",
"arguments": "10m", # sleep for 10 minutes
"output": "sleep-$(ProcId).out",
"error": "sleep-$(ProcId).err",
"log": "sleep.log",
"request_cpus": "1",
"request_memory": "128MB",
"request_disk": "128MB",
})
print(long_sleep_job)
# + pycharm={}
schedd = htcondor.Schedd()
with schedd.transaction() as txn:
cluster_id = long_sleep_job.queue(txn, 5)
# -
# As an experiment, let's set an arbitrary attribute on the jobs and check that it worked.
# When we're really working, we could do things like change the amount of memory a job has requested by editing its `RequestMemory` attribute.
# The job attributes that are built-in to HTCondor are described [here](https://htcondor.readthedocs.io/en/latest/classad-attributes/job-classad-attributes.html), but your site may specify additional, custom attributes as well.
# +
# sets attribute foo to the string "bar" for all of our jobs
# note the nested quotes around bar! The outer "" make it a Python string; the inner "" make it a ClassAd string.
schedd.edit(f"ClusterId == {cluster_id}", "foo", "\"bar\"")
# do a query to check the value of attribute foo
schedd.query(
constraint=f"ClusterId == {cluster_id}",
projection=["ClusterId", "ProcId", "JobStatus", "foo"],
)
# -
# Although the job status appears to be an attribute, we cannot `edit` it directly.
# As mentioned above, we must instead `act` on the job.
# Let's hold the first two jobs so that they stop running, but leave the others going.
# +
# hold the first two jobs
schedd.act(htcondor.JobAction.Hold, f"ClusterId == {cluster_id} && ProcID <= 1")
# check the status of the jobs
ads = schedd.query(
constraint=f"ClusterId == {cluster_id}",
projection=["ClusterId", "ProcId", "JobStatus"],
)
for ad in ads:
# the ClassAd objects returned by the query act like dictionaries, so we can extract individual values out of them using []
print(f"ProcID = {ad['ProcID']} has JobStatus = {ad['JobStatus']}")
# -
# The various job statuses are represented by numbers. `1` means `Idle`, `2` means `Running`, and `5` means `Held`. If you see `JobStatus = 5` above for `ProcID = 0` and `ProcID = 1`, then we succeeded!
#
# The opposite of `JobAction.Hold` is `JobAction.Release`.
# Let's release those jobs and let them go back to `Idle`.
# +
schedd.act(htcondor.JobAction.Release, f"ClusterId == {cluster_id}")
ads = schedd.query(
constraint=f"ClusterId == {cluster_id}",
projection=["ClusterId", "ProcId", "JobStatus"],
)
for ad in ads:
# the ClassAd objects returned by the query act like dictionaries, so we can extract individual values out of them using []
print(f"ProcID = {ad['ProcID']} has JobStatus = {ad['JobStatus']}")
# -
# Note that we simply released all the jobs in the cluster.
# Releasing a job that is not held doesn't do anything, so we don't have to be extremely careful.
#
# Finally, let's clean up after ourselves:
schedd.act(htcondor.JobAction.Remove, f"ClusterId == {cluster_id}")
# ## Exercises
#
# Now let's practice what we've learned.
#
# - In each exercise, you will be given a piece of code and a test that does not yet pass.
# - The exercises are vaguely in order of increasing difficulty.
# - Modify the code, or add new code to it, to pass the test. Do whatever it takes!
# - You can run the test by running the block it is in.
# - Feel free to look at the test for clues as to how to modify the code.
# - Many of the exercises can be solved either by using Python to generate inputs, or by using advanced features of the [ClassAd language](https://htcondor.readthedocs.io/en/latest/misc-concepts/classad-mechanism.html#htcondor-s-classad-mechanism). Either way is valid!
# - Don't modify the test. That's cheating!
# ### Exercise 1: Incrementing Sleeps
#
# Submit five jobs which sleep for `5`, `6`, `7`, `8`, and `9` seconds, respectively.
# +
# MODIFY OR ADD TO THIS BLOCK...
incrementing_sleep = htcondor.Submit({
"executable": "/bin/sleep",
"arguments": "1",
"output": "ex1-$(ProcId).out",
"error": "ex1-$(ProcId).err",
"log": "ex1.log",
"request_cpus": "1",
"request_memory": "128MB",
"request_disk": "128MB",
})
schedd = htcondor.Schedd()
with schedd.transaction() as txn:
cluster_id = incrementing_sleep.queue(txn, 5)
# +
# ... TO MAKE THIS TEST PASS
expected = [str(i) for i in range(5, 10)]
print("Expected ", expected)
ads = schedd.query(f"ClusterId == {cluster_id}", projection = ["Args"])
arguments = sorted(ad["Args"] for ad in ads)
print("Got ", arguments)
assert arguments == expected, "Arguments were not what we expected!"
print("The test passed. Good job!")
# -
# ### Exercise 2: Echo to Target
#
# Run a job that makes the text `Echo to Target` appear in a file named `ex3.txt`.
# +
# MODIFY OR ADD TO THIS BLOCK...
echo = htcondor.Submit({
"request_cpus": "1",
"request_memory": "128MB",
"request_disk": "128MB",
})
schedd = htcondor.Schedd()
with schedd.transaction() as txn:
cluster_id = echo.queue(txn, 1)
# +
# ... TO MAKE THIS TEST PASS
does_file_exist = os.path.exists("ex3.txt")
assert does_file_exist, "ex3.txt does not exist!"
expected = "Echo to Target"
print("Expected ", expected)
contents = open("ex3.txt", mode = "r").read().strip()
print("Got ", contents)
assert expected in contents, "Contents were not what we expected!"
print("The test passed. Good job!")
# -
# ### Exercise 3: Holding Odds
#
# Hold all of the odd-numbered jobs in this large cluster.
#
# - Note that the test block **removes all of the jobs you own** when it runs, to prevent these long-running jobs from corrupting other tests!
# +
# MODIFY OR ADD TO THIS BLOCK...
long_sleep = htcondor.Submit({
"executable": "/bin/sleep",
"arguments": "10m",
"output": "ex2-$(ProcId).out",
"error": "ex2-$(ProcId).err",
"log": "ex2.log",
"request_cpus": "1",
"request_memory": "128MB",
"request_disk": "128MB",
})
schedd = htcondor.Schedd()
with schedd.transaction() as txn:
cluster_id = long_sleep.queue(txn, 100)
# +
# ... TO MAKE THIS TEST PASS
import getpass
try:
ads = schedd.query(f"ClusterId == {cluster_id}", projection = ["ProcID", "JobStatus"])
proc_to_status = {int(ad["ProcID"]): ad["JobStatus"] for ad in sorted(ads, key = lambda ad: ad["ProcID"])}
for proc, status in proc_to_status.items():
print("Proc {} has status {}".format(proc, status))
assert len(proc_to_status) == 100, "Wrong number of jobs (perhaps you need to resubmit them?)."
assert all(status == 5 for proc, status in proc_to_status.items() if proc % 2 != 0), "Not all odd jobs were held."
assert all(status != 5 for proc, status in proc_to_status.items() if proc % 2 == 0), "An even job was held."
print("The test passed. Good job!")
finally:
schedd.act(htcondor.JobAction.Remove, f'Owner=="{getpass.getuser()}"')
| docs/apis/python-bindings/tutorials/Submitting-and-Managing-Jobs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Imports
import pandas as pd
import os
# ### Directories
root_dir = os.path.dirname(os.path.dirname(os.getcwd()))
data_dir = os.path.join(root_dir, 'data')
volume_path = os.path.join(data_dir, 'gx_volume.csv')
merged_months_path = os.path.join(data_dir, 'gx_merged_months.csv')
merged_lags_moths_path = os.path.join(data_dir, 'gx_merged_lags_months.csv')
train_path = os.path.join(data_dir, 'train_split.csv')
valid_split = os.path.join(data_dir, 'valid_split.csv')
submission_template_path = os.path.join(data_dir, 'submission_template.csv')
# ### Read data
volume = pd.read_csv(volume_path, index_col=0)
submission = pd.read_csv(submission_template_path)
merged = pd.read_csv(merged_months_path, index_col=0)
merged_lags = pd.read_csv(merged_lags_moths_path, index_col=0)
train = pd.read_csv(train_path)
valid = pd.read_csv(valid_split)
# ### EDA
print(volume.shape)
print(volume.head())
print(train.head())
print(merged.head())
print(merged_months.describe())
print(merged_months.head())
#Total number of medicaments
volume.brand.nunique()
# ### Plots
ind = (volume.brand == 'brand_20') & (volume.country == 'country_1')
brand_20 = volume.loc[ind, :]
brand_20.head()
brand_20.plot.line(x='month_num',y='volume',c='DarkBlue')
min_values = volume.groupby(['country', 'brand'], as_index=False)['month_num'].min()
print(min_values)
min_values.groupby(['brand'], as_index=False).count().sort_values(by='month_num', ascending=False)
mean_df = volume.groupby(['month_num'], as_index=False)['volume'].mean()
mean_df.plot.line(x='month_num',y='volume',c='DarkBlue')
# ### Checks
# Are all country, brands in test set also in volume?
keys_submission = submission.loc[:, ['country', 'brand']].drop_duplicates()
keys_volume = volume.loc[volume.month_num < 0, ['country', 'brand']].drop_duplicates()
inner_both = keys_submission.merge(keys_volume, 'left', on = ['country', 'brand'])
print(inner_both.shape)
print(keys_submission.shape)
volume
| eda/initial_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Visualization of the Hinge loss function
# #### Used mainly for Support Vector Machines (SVMs). If you want to find more information visit <a href='https://en.wikipedia.org/wiki/Hinge_loss'>Wikipedia</a>
#importing matplotlib library for plotting our results
import matplotlib.pyplot as plt
# %matplotlib inline
def hinge_loss(z, label):
"""
Input:
-> z - value for which we want to calculate hinge_loss()
(In Machine Learning usually the product of feature vector and parameters)
-> label - ground truth label of an example
"""
if label:
return max(0, 1-1*z)
else:
return max(0, 1+1*z)
# +
#creating a list of hypothetical z values
Z = [i for i in range(-10,11)]
#setting label to 1, it can significantly change the outcome of the hinge_loss()
label = 1
#creating a list of hinge_loss() function's output performed on Z list
y1 = [hinge_loss(z, label) for z in Z]
plt.plot(Z,y1)
plt.show()
# +
#setting label to 0, so that we can see the difference from label being equal to 1
label = 0
y2 = [hinge_loss(z, label) for z in Z]
plt.plot(Z,y2)
plt.show()
# -
#displaying both plots on one plot
#red plot corresponds to label equal to 0
#green plot corresponds to label equal to 1
plt.plot(Z,y1,'g')
plt.plot(Z,y2,'r')
plt.show()
| Hinge_loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practice Exercise - 03 - Solution
# ### Question 1:
#
# Convert the string given below into a list by taking each word separated by a comma as an element.
#
# languages = "Java, Python, C++, Scala, C"
#
# #### Expected Output:
#
# ['Java', ' Python', ' C++', ' Scala', ' C']
# +
languages = "Java, Python, C++, Scala, C"
languages.split(',')
# -
# ### Question 2:
#
# We have a list of numbers given below. Add the element '10' at the 3rd position of the list and finally print the list.
#
# num_list = [5, 6, 8, 7, 9]
#
# #### Expected Output:
#
# [5, 6, 8, 10, 7, 9]
# +
num_list = [5, 6, 8, 7, 9]
num_list.insert(3, 10)
print(num_list)
# -
# ### Question 3:
#
# Print all the multiples of 3 between 5 and 25.
#
# #### Expected Output:
#
# 6
# <br>9
# <br>12
# <br>15
# <br>18
# <br>21
# <br>24
for i in range(5, 25):
if i % 3 == 0:
print(i)
# ### Question 4:
#
# We have a list of numbers given below. Print the square of these numbers into another list using list comprehension.
#
# num = [2, 4, 6, 8]
#
# #### Expected Output:
#
# [4, 16, 36, 64]
# +
num = [2, 4, 6, 8]
square = [i**2 for i in num]
print(square)
# -
# ### Question 5:
#
# Print the first 10 natural numbers.
#
# #### Expected Output:
#
# 1
# <br>2
# <br>3
# <br>4
# <br>5
# <br>6
# <br>7
# <br>8
# <br>9
# <br>10
# +
num = 1
while num <= 10:
print(num)
num += 1
# -
# ### Question 6:
#
# We have a tuple of numbers given below. Remove the largest number from the tuple and print it in sorted order.
#
# num_tuple = (5, 8, 13, 2, 17, 1)
#
# #### Expected Output:
#
# [1, 2, 5, 8, 13]
# +
num_tuple = (5, 8, 13, 2, 17, 1)
sorted_num = sorted(num_tuple)
sorted_num.pop()
print(sorted_num)
# -
# ### Question 7:
#
# Convert the list given below into a string using a comma as a separator argument.
#
# myList = ['Lenovo', ' Dell', ' Acer', ' Asus', ' HP']
#
# #### Expected Output:
#
# 'Lenovo, Dell, Acer, Asus, HP'
# +
myList = ['Lenovo', ' Dell', ' Acer', ' Asus', ' HP']
myString = ','.join(myList)
myString
# -
# ### Question 8:
#
# Print all the even numbers between -10 and 0.
#
# #### Expected Output:
#
# -10
# <br>-8
# <br>-6
# <br>-4
# <br>-2
for i in range(-10, 0, 2):
print(i)
# ### Question 9:
#
# Reverse the integer given below.
#
# n = 5623
#
# #### Expected Output:
#
# 3265
# +
n = 5623
rev = 0
while n > 0:
rem = n % 10
rev = (rev * 10) + rem
n = n // 10
print(rev)
# -
# ### Question 10:
#
# We have a list of numbers given below. Print the number, square of the number, and the cube of the number, all in a single list.
#
# num = [2, 4, 6, 8]
#
# #### Expected Output:
#
# [(2, 4, 8), (4, 16, 64), (6, 36, 216), (8, 64, 512)]
# +
num = [2, 4, 6, 8]
result = [(x, x**2, x**3) for x in num]
print(result)
| practice-exercise-03-solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="C2Nqtj5Zr1Qa"
# # *ADMISSION PREDICTION*
# + id="2ClwXSCfFH8j" outputId="122739b4-1221-4a76-b73c-212dd8a8993f" colab={"base_uri": "https://localhost:8080/", "height": 195}
#import packages
import pandas as pd
import numpy as np
#to plot within notebook
import matplotlib.pyplot as plt
# %matplotlib inline
#setting figure size
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 20,10
#read the file
df = pd.read_csv('/content/college admission prediction.csv')
#print the head
df.head()
# + id="72VDhHpmYoAI" outputId="efcdb6f8-122b-4d89-b1d6-f960b680c1e1" colab={"base_uri": "https://localhost:8080/", "height": 346}
college=np.unique(df['College'])
print(college)
clg_code=[]
for i in range(len(college)):
clg_code.append(i+1)
# clg_code
df['College']=df['College'].replace(college,clg_code)
bak_college=np.array(df['College'])
df.head()
# + id="H_crkyJ9YXic" outputId="13602891-2aaf-444f-e3fa-29fd28a75876" colab={"base_uri": "https://localhost:8080/", "height": 168}
from sklearn import datasets, linear_model
from sklearn.model_selection import train_test_split
# Using only one feature
x = df.iloc[:, 4].values
y = df.iloc[:, 5].values
# Split the data
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size= 0.2, random_state=0)
print('Shape of X_train:: ', x_train.shape)
print('Shape of y_train:: ', y_train.shape)
print('Shape of X_test:: ', x_test.shape)
print('Shape of y_test:: ', y_test.shape)
x_train= x_train.reshape(-1, 1)
x_test = x_test.reshape(-1, 1)
print('\nShape of X_train:: ', x_train.shape)
print('Shape of y_train:: ', y_train.shape)
print('Shape of X_test:: ', x_test.shape)
print('Shape of y_test:: ', y_test.shape)
# + id="hmsySoxda7iu" outputId="d71077be-49a6-4246-a623-91522302f88f" colab={"base_uri": "https://localhost:8080/", "height": 628}
model = linear_model.LinearRegression()
model.fit(x_train, y_train)
y_pred = model.predict(x_test)
print('Coefficient:', model.coef_)
print("RMSE: %.2f" % np.sqrt(np.mean((model.predict(x_test) - y_test) ** 2)))
plt.figure(figsize=(20,10))
plt.scatter(x_test, y_test, color='black')
plt.plot(x_test, y_pred, color='blue', linewidth=3)
plt.xlabel("AIEEE Rank")
plt.ylabel("College Code")
plt.title("Admission Prediction")
plt.show()
# + id="Jo0NPOUQZyOd" outputId="8ee05794-922f-4048-83ca-931dcc8ddb0a" colab={"base_uri": "https://localhost:8080/", "height": 84}
col=df.columns.tolist()[4:5]
print(col)
usrip=[]
for i in col:
print("==================================================")
usrip.append(eval(input(i+": ")))
userpreddt=model.predict([usrip])
print("You may have change to get entrance in: ",college[clg_code.index(int(userpreddt[0]))])
# + id="VTaIFQ4-hdJv"
import tensorflow as tf
from tensorflow import lite
keras_file = "linear.h5"
tf.keras.models.save_model(model, keras_file)
# converter = lite.TFLiteConverter.from_keras_model(model)
# tfmodel = converter.convert()
# open("linear.tflite", "wb").write(tfmodel)
| ML/SIH_module1_Admission_LinearRegression (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="eTY9DyDRzLs2"
# # Model Output with min_freq = 1
# + [markdown] id="Yuh5BPAtButa"
# 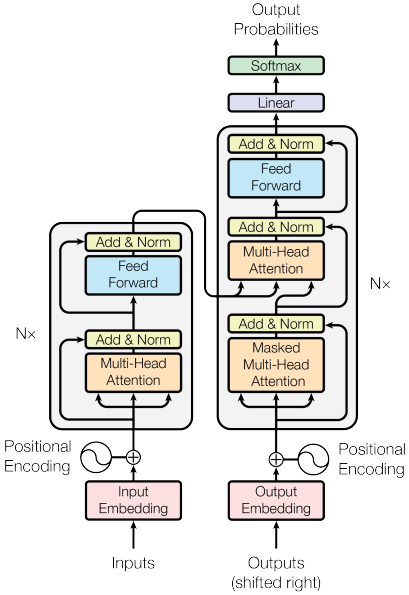
# + id="1-joZEP1d36h"
# # ! pip install datasets transformers
# + id="dJWBWb31mz4B"
from tokenize import tokenize, untokenize, NUMBER, STRING, NAME, OP
from io import BytesIO
# + colab={"base_uri": "https://localhost:8080/"} id="vBnYmQg3k_FD" outputId="9bfae1b5-3dba-4afb-e13b-1342bbb8dbf9"
from google.colab import drive
drive.mount('/content/drive')
# + id="9qtMB_iAzqas"
# ! cp "/content/drive/My Drive/NLP/english_python_data_modified.txt" english_python_data_modified.txt
# # ! cp '/content/drive/My Drive/NLP/cornell_movie_dialogs_corpus.zip' .
# + id="cgTxHeAvl3Er"
import torch
import torch.nn as nn
import torch.optim as optim
import torchtext
# from torchtext.data import Field, BucketIterator, TabularDataset
from torchtext.legacy.data import Field, BucketIterator, TabularDataset
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import spacy
import numpy as np
import random
import math
import time
# + id="KwMMe5CQCU5g"
SEED = 1234
random.seed(SEED)
np.random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.backends.cudnn.deterministic = True
# + [markdown] id="6NDqzSxYRmZI"
# ### Downloading the File
# + colab={"base_uri": "https://localhost:8080/"} id="BMgNsUwa-CgB" outputId="687015e9-dc48-4bfd-8f34-1f8d2758e247"
import requests
import os
import datetime
# !wget "https://drive.google.com/u/0/uc?id=1rHb0FQ5z5ZpaY2HpyFGY6CeyDG0kTLoO&export=download"
os.rename("uc?id=1rHb0FQ5z5ZpaY2HpyFGY6CeyDG0kTLoO&export=download","english_python_data.txt")
# + [markdown] id="co_t15woRtLB"
# ### Reading the File and Separating Out English Text and Python Code
# + id="-KL3_pcI4MUS" colab={"base_uri": "https://localhost:8080/"} outputId="6a65af00-b249-46d3-b473-6be750b4d503"
# https://stackoverflow.com/questions/31786823/print-lines-between-two-patterns-in-python/31787181
fastq_filename = "english_python_data_modified.txt"
fastq = open(fastq_filename) # fastq is the file object
for line in fastq:
if line.startswith("#") or line.isalpha():
print(line.replace("@", ">"))
# + id="uC6prHvc62MR"
def generate_df(filename):
with open(filename) as file_in:
newline = '\n'
lineno = 0
lines = []
Question = []
Answer = []
Question_Ind =-1
mystring = "NA"
revised_string = "NA"
Initial_Answer = False
# you may also want to remove whitespace characters like `\n` at the end of each line
for line in file_in:
lineno = lineno +1
if line in ['\n', '\r\n']:
pass
else:
linex = line.rstrip() # strip trailing spaces and newline
# if string[0].isdigit()
if linex.startswith('# '): ## to address question like " # write a python function to implement linear extrapolation"
if Initial_Answer:
Answer.append(revised_string)
revised_string = "NA"
mystring = "NA"
Initial_Answer = True
Question.append(linex.strip('# '))
# Question_Ind = Question_Ind +1
elif linex.startswith('#'): ## to address question like "#24. Python Program to Find Numbers Divisible by Another Number"
linex = linex.strip('#')
# print(linex)
# print(f"amit:{len(linex)}:LineNo:{lineno}")
if (linex[0].isdigit()): ## stripping first number which is 2
# print("Amit")
linex = linex.strip(linex[0])
if (linex[0].isdigit()): ## stripping 2nd number which is 4
linex = linex.strip(linex[0])
if (linex[0]=="."):
linex = linex.strip(linex[0])
if (linex[0].isspace()):
linex = linex.strip(linex[0]) ## stripping out empty space
if Initial_Answer:
Answer.append(revised_string)
revised_string = "NA"
mystring = "NA"
Initial_Answer = True
Question.append(linex)
else:
# linex = '\n'.join(linex)
if (mystring == "NA"):
mystring = f"{linex}{newline}"
revised_string = mystring
# print(f"I am here:{mystring}")
else:
mystring = f"{linex}{newline}"
if (revised_string == "NA"):
revised_string = mystring
# print(f"I am here revised_string:{revised_string}")
else:
revised_string = revised_string + mystring
# print(f"revised_string:{revised_string}")
# Answer.append(string)
lines.append(linex)
Answer.append(revised_string)
return Question, Answer
# + colab={"base_uri": "https://localhost:8080/"} id="UxHaS2XhNITT" outputId="7f1cb771-97ae-4816-ac81-b366ebd1ad6f"
Question, Answer = generate_df("english_python_data_modified.txt")
print(f"Length of Question:{len(Question)}")
print(f"Length of Answer:{len(Answer)}")
# + id="m62cKbnfP4Wp"
# Answer[0]
# num1 = 1.5\nnum2 = 6.3\nsum = num1 + num2\nprint(f'Sum: {sum}')\n\n\n
# + id="XT5_YaO_yV4h"
# with open("english_emp.txt") as file_in:
# newline = '\n'
# lines = []
# Question = []
# Answer = []
# Question_Ind =-1
# mystring = "NA"
# revised_string = "NA"
# Initial_Answer = False
# # you may also want to remove whitespace characters like `\n` at the end of each line
# for line in file_in:
# linex = line.rstrip() # strip trailing spaces and newline
# if linex.startswith('# '):
# if Initial_Answer:
# # print(f"Answer:{Answer}")
# Answer.append(revised_string)
# revised_string = "NA"
# mystring = "NA"
# Initial_Answer = True
# Question.append(linex.strip('# '))
# Question_Ind = Question_Ind +1
# else:
# # linex = '\n'.join(linex)
# if (mystring == "NA"):
# mystring = f"{linex}{newline}"
# revised_string = mystring
# # print(f"I am here:{mystring}")
# else:
# mystring = f"{linex}{newline}"
# if (revised_string == "NA"):
# revised_string = mystring
# # print(f"I am here revised_string:{revised_string}")
# else:
# revised_string = revised_string + mystring
# # print(f"revised_string:{revised_string}")
# # Answer.append(string)
# lines.append(linex)
# Answer.append(revised_string)
# print(Question[1])
# + colab={"base_uri": "https://localhost:8080/"} id="_2wa08maSzP0" outputId="43f0c403-24f2-4228-a1b7-537c3d330a8e"
## do some random check
print(f"Question[0]:\n{Question[0]}")
print(f"Answer[0]:\n{Answer[0]}")
# + colab={"base_uri": "https://localhost:8080/"} id="wBQA1KcPOMqP" outputId="c99526c3-b6c1-4e44-c103-e9f69cee5298"
## do some random check
print(f"Question[1]:\n {Question[1]}")
print(f"Answer[1]:\n {Answer[1]}")
# + colab={"base_uri": "https://localhost:8080/"} id="qKz_yoZdRNI5" outputId="98562ff9-122e-445b-9caf-a6aefa890ffe"
## do some random check
print(f"Question[4849]:\n{Question[4849]}")
print(f"Answer[4849]:\n{Answer[4849]}")
# + [markdown] id="fpqoN4b8R2ou"
# ### Converting into dataframe and dumping into CSV
# + id="mHUaiT42R5J0"
import pandas as pd
df_Question = pd.DataFrame(Question, columns =['Question'])
df_Answer = pd.DataFrame(Answer,columns =['Answer'])
frames = [df_Question, df_Answer]
combined_question_answer = pd.concat(frames,axis=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 106} id="FjmMf_vESo6g" outputId="6be20ba6-26ff-4b4a-fc51-4b4d358a0daa"
combined_question_answer.head(2)
# + id="ckmEM2Wsizlu"
combined_question_answer.to_csv("combined_question_answer_from_df.csv",index=False)
# + id="Se895tbRW5PA"
combined_question_answer['AnswerLen'] = combined_question_answer['Answer'].astype(str).map(len)
# + colab={"base_uri": "https://localhost:8080/"} id="eMO3nPltXy3v" outputId="baf90015-d113-4c66-be0e-8cecef347769"
combined_question_answer.size
# + colab={"base_uri": "https://localhost:8080/", "height": 106} id="-zimalfYXfud" outputId="7d7279b8-e844-465e-cf2c-5dad72a91ccd"
combined_question_answer.head(2)
# + colab={"base_uri": "https://localhost:8080/"} id="uiWsWRmVX9K7" outputId="a975d713-69db-4f0f-e06d-7605695d00f0"
combined_question_answer_df = combined_question_answer[combined_question_answer['AnswerLen'] < 495]
combined_question_answer_df.size
# + id="lHctGq43aRRo"
combined_question_answer_df = combined_question_answer_df.drop(['AnswerLen'], axis=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 106} id="M8D-QVkdaky4" outputId="9385df8c-4844-489f-f80a-53b99f3a0e46"
combined_question_answer_df.head(2)
# + id="-an664Rc29Wl"
from sklearn.model_selection import train_test_split
train_combined_question_answer, val_combined_question_answer = train_test_split(combined_question_answer_df, test_size=0.2)
train_combined_question_answer.to_csv("train_combined_question_answer.csv",index=False)
val_combined_question_answer.to_csv("val_combined_question_answer.csv",index=False)
# + [markdown] id="kQJDZHQjj0ra"
# ### Downloading spacy and tokenization
# + colab={"base_uri": "https://localhost:8080/"} id="xpNTE2GhCY2V" outputId="748bc6e4-4c8a-410e-afb5-e24ad69b75f1"
# !python -m spacy download en
# + id="j37U6a5OCi_M"
spacy_en = spacy.load('en')
# + [markdown] id="Oea17h5baJMU"
# ### Defining Iterator and Tokenization
# + id="15wydnnsCpH0"
def tokenize_en(text):
"""
Tokenizes English text from a string into a list of strings
"""
return [tok.text for tok in spacy_en.tokenizer(text)]
# + [markdown] id="pW_i6zCZzJeW"
#
# + id="UeiubrXM6pvC"
def tokenize_en_python(text):
"""
Tokenizes English text from a string into a list of strings
"""
return [tokenize(text)]
# + id="60O0YZ8eCrLd"
TEXT = Field(tokenize = tokenize_en,
eos_token = '<eos>',
init_token = '<sos>',
# lower = True,
batch_first = True)
fields = [("Question", TEXT), ("Answer", TEXT)]
# + colab={"base_uri": "https://localhost:8080/"} id="7lfgaRJs1-GT" outputId="8a65b056-57fa-45b8-8238-5375e5ecfd86"
# !wget https://raw.githubusercontent.com/pytorch/pytorch/master/torch/utils/collect_env.py
# !python collect_env.py
# !python -c "import torchtext; print(\"torchtext version is \", torchtext.__version__)"
# + [markdown] id="nMEB7kKDEtdR"
# An example article-title pair looks like this:
#
# **article**: the algerian cabinet chaired by president ab<NAME> on sunday adopted the #### finance bill predicated on an oil price of ## dollars a barrel and a growth rate of #.# percent , it was announced here .
#
# **title**: algeria adopts #### finance bill with oil put at ## dollars a barrel
# + id="i2Pd_P_ezou1"
train_data, valid_data = TabularDataset.splits(path=f'/content',
train='train_combined_question_answer.csv', validation='val_combined_question_answer.csv',
format='csv', skip_header=True, fields=fields)
# + colab={"base_uri": "https://localhost:8080/"} id="ljmYlUKesNkH" outputId="32ed4d8e-cc34-4607-88d6-e7a513d0e0fe"
print(f'Number of training examples: {len(train_data)}')
print(f'Number of validation examples: {len(valid_data)}')
#print(f'Number of testing examples: {len(test_data)}')
# + colab={"base_uri": "https://localhost:8080/"} id="rPB35AzZ54Cc" outputId="16d075dd-a1b2-400f-b8f1-627b367faec8"
# a sample of the preprocessed data
print(train_data[0].Question, train_data[0].Answer)
# + id="fETHzIIyCu7p"
TEXT.build_vocab(train_data, min_freq = 1)
# + colab={"base_uri": "https://localhost:8080/"} id="DmotHU_StV6D" outputId="97bd700f-715d-49a5-82a6-f106d8ef4b89"
print(f"Unique tokens in TEXT vocabulary: {len(TEXT.vocab)}")
# + colab={"base_uri": "https://localhost:8080/"} id="2XONz6UrCwuh" outputId="ce6bc48d-0e77-4645-c0aa-5fb223b6fb4a"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device
# + id="aBtCv6sACyZ8"
BATCH_SIZE = 32
train_iterator, valid_iterator = BucketIterator.splits(
(train_data, valid_data),
batch_size = BATCH_SIZE,
device = device,
sort_key=lambda x: len(x.Question),
shuffle=True,
sort_within_batch=False,
repeat=False)
# + [markdown] id="HgvD_MpkC2OS"
# 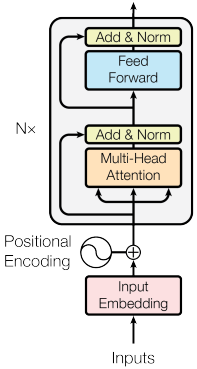
# + [markdown] id="u3JvQnHDaPgg"
# ### Transformer Model Architecture
# + id="NE6JimgOCz-w"
class Encoder(nn.Module):
def __init__(self,
input_dim,
hid_dim,
n_layers,
n_heads,
pf_dim,
dropout,
device,
max_length = 500):
super().__init__()
self.device = device
self.tok_embedding = nn.Embedding(input_dim, hid_dim)
self.pos_embedding = nn.Embedding(max_length, hid_dim)
self.layers = nn.ModuleList([EncoderLayer(hid_dim,
n_heads,
pf_dim,
dropout,
device)
for _ in range(n_layers)])
self.dropout = nn.Dropout(dropout)
self.scale = torch.sqrt(torch.FloatTensor([hid_dim])).to(device)
def forward(self, src, src_mask):
#src = [batch size, src len]
#src_mask = [batch size, 1, 1, src len]
batch_size = src.shape[0]
src_len = src.shape[1]
pos = torch.arange(0, src_len).unsqueeze(0).repeat(batch_size, 1).to(self.device)
#pos = [batch size, src len]
src = self.dropout((self.tok_embedding(src) * self.scale) + self.pos_embedding(pos))
#src = [batch size, src len, hid dim]
for layer in self.layers:
src = layer(src, src_mask)
#src = [batch size, src len, hid dim]
return src
# + id="2LheiXWVFDEg"
class EncoderLayer(nn.Module):
def __init__(self,
hid_dim,
n_heads,
pf_dim,
dropout,
device):
super().__init__()
self.self_attn_layer_norm = nn.LayerNorm(hid_dim)
self.ff_layer_norm = nn.LayerNorm(hid_dim)
self.self_attention = MultiHeadAttentionLayer(hid_dim, n_heads, dropout, device)
self.positionwise_feedforward = PositionwiseFeedforwardLayer(hid_dim,
pf_dim,
dropout)
self.dropout = nn.Dropout(dropout)
def forward(self, src, src_mask):
#src = [batch size, src len, hid dim]
#src_mask = [batch size, 1, 1, src len]
#self attention
_src, _ = self.self_attention(src, src, src, src_mask)
#dropout, residual connection and layer norm
src = self.self_attn_layer_norm(src + self.dropout(_src))
#src = [batch size, src len, hid dim]
#positionwise feedforward
_src = self.positionwise_feedforward(src)
#dropout, residual and layer norm
src = self.ff_layer_norm(src + self.dropout(_src))
#src = [batch size, src len, hid dim]
return src
# + [markdown] id="6h_Iqnk4Jg5k"
# 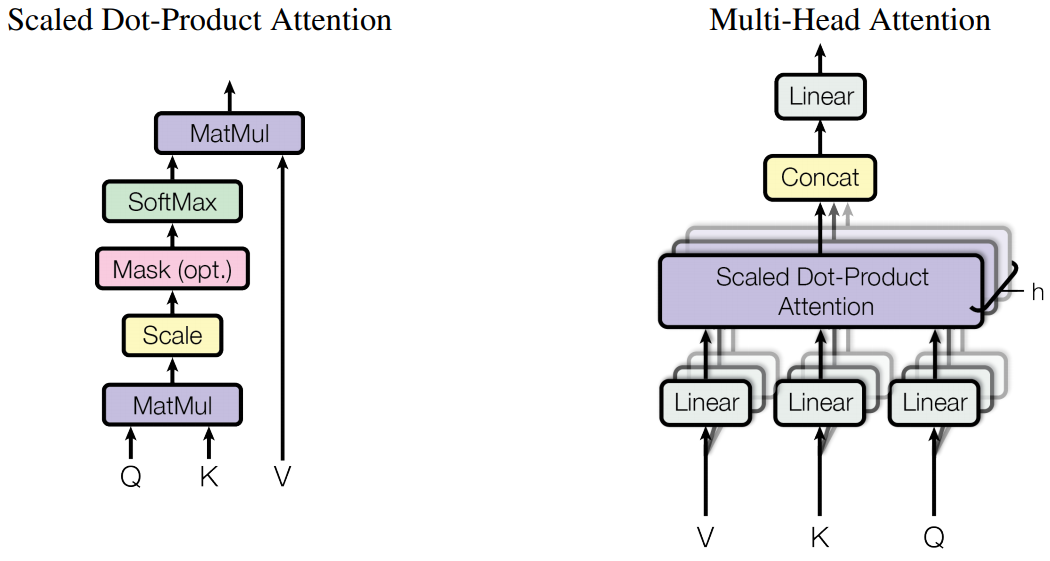
# + id="ZZmeHfGhGzkN"
class MultiHeadAttentionLayer(nn.Module):
def __init__(self, hid_dim, n_heads, dropout, device):
super().__init__()
assert hid_dim % n_heads == 0
self.hid_dim = hid_dim
self.n_heads = n_heads
self.head_dim = hid_dim // n_heads
self.fc_q = nn.Linear(hid_dim, hid_dim)
self.fc_k = nn.Linear(hid_dim, hid_dim)
self.fc_v = nn.Linear(hid_dim, hid_dim)
self.fc_o = nn.Linear(hid_dim, hid_dim)
self.dropout = nn.Dropout(dropout)
self.scale = torch.sqrt(torch.FloatTensor([self.head_dim])).to(device)
def forward(self, query, key, value, mask = None):
batch_size = query.shape[0]
#query = [batch size, query len, hid dim]
#key = [batch size, key len, hid dim]
#value = [batch size, value len, hid dim]
Q = self.fc_q(query)
K = self.fc_k(key)
V = self.fc_v(value)
#Q = [batch size, query len, hid dim]
#K = [batch size, key len, hid dim]
#V = [batch size, value len, hid dim]
Q = Q.view(batch_size, -1, self.n_heads, self.head_dim).permute(0, 2, 1, 3)
K = K.view(batch_size, -1, self.n_heads, self.head_dim).permute(0, 2, 1, 3)
V = V.view(batch_size, -1, self.n_heads, self.head_dim).permute(0, 2, 1, 3)
#Q = [batch size, n heads, query len, head dim]
#K = [batch size, n heads, key len, head dim]
#V = [batch size, n heads, value len, head dim]
energy = torch.matmul(Q, K.permute(0, 1, 3, 2)) / self.scale
#energy = [batch size, n heads, query len, key len]
if mask is not None:
energy = energy.masked_fill(mask == 0, -1e10)
attention = torch.softmax(energy, dim = -1)
#attention = [batch size, n heads, query len, key len]
x = torch.matmul(self.dropout(attention), V)
#x = [batch size, n heads, query len, head dim]
x = x.permute(0, 2, 1, 3).contiguous()
#x = [batch size, query len, n heads, head dim]
x = x.view(batch_size, -1, self.hid_dim)
#x = [batch size, query len, hid dim]
x = self.fc_o(x)
#x = [batch size, query len, hid dim]
return x, attention
# + id="R9w9xDUKL7LU"
class PositionwiseFeedforwardLayer(nn.Module):
def __init__(self, hid_dim, pf_dim, dropout):
super().__init__()
self.fc_1 = nn.Linear(hid_dim, pf_dim)
self.fc_2 = nn.Linear(pf_dim, hid_dim)
self.dropout = nn.Dropout(dropout)
def forward(self, x):
#x = [batch size, seq len, hid dim]
x = self.dropout(torch.relu(self.fc_1(x)))
#x = [batch size, seq len, pf dim]
x = self.fc_2(x)
#x = [batch size, seq len, hid dim]
return x
# + [markdown] id="YbTr7YPSMRpC"
# 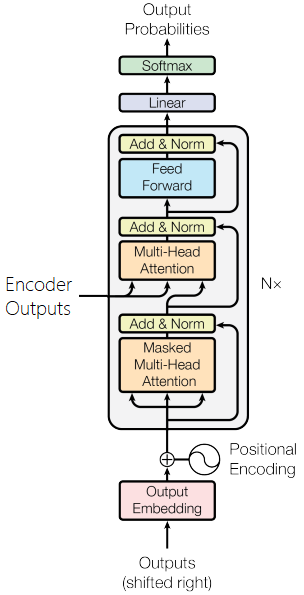
# + id="iWBMMF45MMNS"
class Decoder(nn.Module):
def __init__(self,
output_dim,
hid_dim,
n_layers,
n_heads,
pf_dim,
dropout,
device,
max_length = 500):
super().__init__()
self.device = device
self.tok_embedding = nn.Embedding(output_dim, hid_dim)
self.pos_embedding = nn.Embedding(max_length, hid_dim)
self.layers = nn.ModuleList([DecoderLayer(hid_dim,
n_heads,
pf_dim,
dropout,
device)
for _ in range(n_layers)])
self.fc_out = nn.Linear(hid_dim, output_dim)
self.dropout = nn.Dropout(dropout)
self.scale = torch.sqrt(torch.FloatTensor([hid_dim])).to(device)
def forward(self, trg, enc_src, trg_mask, src_mask):
#trg = [batch size, trg len]
#enc_src = [batch size, src len, hid dim]
#trg_mask = [batch size, 1, trg len, trg len]
#src_mask = [batch size, 1, 1, src len]
batch_size = trg.shape[0]
trg_len = trg.shape[1]
pos = torch.arange(0, trg_len).unsqueeze(0).repeat(batch_size, 1).to(self.device)
#pos = [batch size, trg len]
trg = self.dropout((self.tok_embedding(trg) * self.scale) + self.pos_embedding(pos))
#trg = [batch size, trg len, hid dim]
for layer in self.layers:
trg, attention = layer(trg, enc_src, trg_mask, src_mask)
#trg = [batch size, trg len, hid dim]
#attention = [batch size, n heads, trg len, src len]
output = self.fc_out(trg)
#output = [batch size, trg len, output dim]
return output, attention
# + id="CMEr1IFUMxco"
class DecoderLayer(nn.Module):
def __init__(self,
hid_dim,
n_heads,
pf_dim,
dropout,
device):
super().__init__()
self.self_attn_layer_norm = nn.LayerNorm(hid_dim)
self.enc_attn_layer_norm = nn.LayerNorm(hid_dim)
self.ff_layer_norm = nn.LayerNorm(hid_dim)
self.self_attention = MultiHeadAttentionLayer(hid_dim, n_heads, dropout, device)
self.encoder_attention = MultiHeadAttentionLayer(hid_dim, n_heads, dropout, device)
self.positionwise_feedforward = PositionwiseFeedforwardLayer(hid_dim,
pf_dim,
dropout)
self.dropout = nn.Dropout(dropout)
def forward(self, trg, enc_src, trg_mask, src_mask):
#trg = [batch size, trg len, hid dim]
#enc_src = [batch size, src len, hid dim]
#trg_mask = [batch size, 1, trg len, trg len]
#src_mask = [batch size, 1, 1, src len]
#self attention
_trg, _ = self.self_attention(trg, trg, trg, trg_mask)
#dropout, residual connection and layer norm
trg = self.self_attn_layer_norm(trg + self.dropout(_trg))
#trg = [batch size, trg len, hid dim]
#encoder attention
_trg, attention = self.encoder_attention(trg, enc_src, enc_src, src_mask)
# query, key, value
#dropout, residual connection and layer norm
trg = self.enc_attn_layer_norm(trg + self.dropout(_trg))
#trg = [batch size, trg len, hid dim]
#positionwise feedforward
_trg = self.positionwise_feedforward(trg)
#dropout, residual and layer norm
trg = self.ff_layer_norm(trg + self.dropout(_trg))
#trg = [batch size, trg len, hid dim]
#attention = [batch size, n heads, trg len, src len]
return trg, attention
# + [markdown] id="udpPhQ2UN8oQ"
# 10000
# 11000
# 11100
# 11100
# 11100
# + id="Dr3Mg8OGN6ul"
class Seq2Seq(nn.Module):
def __init__(self,
encoder,
decoder,
src_pad_idx,
trg_pad_idx,
device):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.src_pad_idx = src_pad_idx
self.trg_pad_idx = trg_pad_idx
self.device = device
def make_src_mask(self, src):
#src = [batch size, src len]
src_mask = (src != self.src_pad_idx).unsqueeze(1).unsqueeze(2)
#src_mask = [batch size, 1, 1, src len]
return src_mask
def make_trg_mask(self, trg):
#trg = [batch size, trg len]
trg_pad_mask = (trg != self.trg_pad_idx).unsqueeze(1).unsqueeze(2)
#trg_pad_mask = [batch size, 1, 1, trg len]
trg_len = trg.shape[1]
trg_sub_mask = torch.tril(torch.ones((trg_len, trg_len), device = self.device)).bool()
#trg_sub_mask = [trg len, trg len]
trg_mask = trg_pad_mask & trg_sub_mask
#trg_mask = [batch size, 1, trg len, trg len]
return trg_mask
def forward(self, src, trg):
#src = [batch size, src len]
#trg = [batch size, trg len]
src_mask = self.make_src_mask(src)
trg_mask = self.make_trg_mask(trg)
#src_mask = [batch size, 1, 1, src len]
#trg_mask = [batch size, 1, trg len, trg len]
enc_src = self.encoder(src, src_mask)
#enc_src = [batch size, src len, hid dim]
output, attention = self.decoder(trg, enc_src, trg_mask, src_mask)
#output = [batch size, trg len, output dim]
#attention = [batch size, n heads, trg len, src len]
return output, attention
# + id="4zsZjSSWOSHc"
INPUT_DIM = len(TEXT.vocab)
OUTPUT_DIM = len(TEXT.vocab)
HID_DIM = 256
ENC_LAYERS = 3
DEC_LAYERS = 3
ENC_HEADS = 8
DEC_HEADS = 8
ENC_PF_DIM = 512
DEC_PF_DIM = 512
ENC_DROPOUT = 0.1
DEC_DROPOUT = 0.1
enc = Encoder(INPUT_DIM,
HID_DIM,
ENC_LAYERS,
ENC_HEADS,
ENC_PF_DIM,
ENC_DROPOUT,
device)
dec = Decoder(OUTPUT_DIM,
HID_DIM,
DEC_LAYERS,
DEC_HEADS,
DEC_PF_DIM,
DEC_DROPOUT,
device)
# + id="iYVZYDVcOUGK"
SRC_PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token]
TRG_PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token]
model = Seq2Seq(enc, dec, SRC_PAD_IDX, TRG_PAD_IDX, device).to(device)
# + colab={"base_uri": "https://localhost:8080/"} id="Qd0ePzj0OzLa" outputId="1a8bbf76-701b-4613-d3e6-c8571c7c6e3e"
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
# + id="fmZ0hyo8O0vE"
def initialize_weights(m):
if hasattr(m, 'weight') and m.weight.dim() > 1:
nn.init.xavier_uniform_(m.weight.data)
# + id="NRtAM9Y4O2N2"
model.apply(initialize_weights);
# + id="NEpApG3YO3ZE"
LEARNING_RATE = 0.0005
optimizer = torch.optim.Adam(model.parameters(), lr = LEARNING_RATE)
# + id="n9Dy_wWrO46l"
criterion = nn.CrossEntropyLoss(ignore_index = TRG_PAD_IDX)
# + id="ycBBiEpuO6cG"
def train(model, iterator, optimizer, criterion, clip):
model.train()
epoch_loss = 0
for i, batch in enumerate(iterator):
Question = batch.Question
Answer = batch.Answer
optimizer.zero_grad()
output, _ = model(Question, Answer[:,:-1])
#output = [batch size, trg len - 1, output dim]
#trg = [batch size, trg len]
output_dim = output.shape[-1]
output = output.contiguous().view(-1, output_dim)
Answer = Answer[:,1:].contiguous().view(-1)
#output = [batch size * trg len - 1, output dim]
#trg = [batch size * trg len - 1]
loss = criterion(output, Answer)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), clip)
optimizer.step()
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + id="zi3Ev8gaO79_"
def evaluate(model, iterator, criterion):
model.eval()
epoch_loss = 0
with torch.no_grad():
for i, batch in enumerate(iterator):
Question = batch.Question
Answer = batch.Answer
output, _ = model(Question, Answer[:,:-1])
#output = [batch size, trg len - 1, output dim]
#trg = [batch size, trg len]
output_dim = output.shape[-1]
output = output.contiguous().view(-1, output_dim)
Answer = Answer[:,1:].contiguous().view(-1)
#output = [batch size * trg len - 1, output dim]
#trg = [batch size * trg len - 1]
loss = criterion(output, Answer)
epoch_loss += loss.item()
return epoch_loss / len(iterator)
# + id="JuB4JqQRO9Wg"
def epoch_time(start_time, end_time):
elapsed_time = end_time - start_time
elapsed_mins = int(elapsed_time / 60)
elapsed_secs = int(elapsed_time - (elapsed_mins * 60))
return elapsed_mins, elapsed_secs
# + [markdown] id="MlnSmXjkaYvk"
# ### Model Training
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="aax76Ie4O_Cr" outputId="4a136103-2aff-4a43-ae57-a0c7c4db6c1b"
N_EPOCHS = 35
CLIP = 1
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start_time = time.time()
train_loss = train(model, train_iterator, optimizer, criterion, CLIP)
valid_loss = evaluate(model, valid_iterator, criterion)
end_time = time.time()
epoch_mins, epoch_secs = epoch_time(start_time, end_time)
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), '/content/drive/My Drive/NLP/tut6-model.pt')
print(f'Epoch: {epoch+1:02} | Time: {epoch_mins}m {epoch_secs}s')
print(f'\tTrain Loss: {train_loss:.3f} | Train PPL: {math.exp(train_loss):7.3f}')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. PPL: {math.exp(valid_loss):7.3f}')
# + id="f3_aq7QTPBFc" colab={"base_uri": "https://localhost:8080/"} outputId="062484f8-30a9-4d27-8634-7d833ca2e028"
model.load_state_dict(torch.load('/content/drive/My Drive/NLP/tut6-model.pt'))
test_loss = evaluate(model, valid_iterator, criterion)
print(f'| Test Loss: {test_loss:.3f} | Test PPL: {math.exp(test_loss):7.3f} |')
# + [markdown] id="aZ3FJk4Raeug"
# ### Model Validations
# + id="ieIjql9uPKH1"
def translate_sentence(sentence, src_field, trg_field, model, device, max_len = 50):
model.eval()
if isinstance(sentence, str):
nlp = spacy.load('de')
tokens = [token.text.lower() for token in nlp(sentence)]
else:
tokens = [token.lower() for token in sentence]
tokens = [src_field.init_token] + tokens + [src_field.eos_token]
src_indexes = [src_field.vocab.stoi[token] for token in tokens]
src_tensor = torch.LongTensor(src_indexes).unsqueeze(0).to(device)
src_mask = model.make_src_mask(src_tensor)
with torch.no_grad():
enc_src = model.encoder(src_tensor, src_mask)
trg_indexes = [trg_field.vocab.stoi[trg_field.init_token]]
for i in range(max_len):
trg_tensor = torch.LongTensor(trg_indexes).unsqueeze(0).to(device)
trg_mask = model.make_trg_mask(trg_tensor)
with torch.no_grad():
output, attention = model.decoder(trg_tensor, enc_src, trg_mask, src_mask)
pred_token = output.argmax(2)[:,-1].item()
trg_indexes.append(pred_token)
if pred_token == trg_field.vocab.stoi[trg_field.eos_token]:
break
trg_tokens = [trg_field.vocab.itos[i] for i in trg_indexes]
return trg_tokens[1:], attention
# + id="WTBipndfPOlX"
def display_attention(sentence, translation, attention, n_heads = 8, n_rows = 4, n_cols = 2):
assert n_rows * n_cols == n_heads
fig = plt.figure(figsize=(15,25))
for i in range(n_heads):
ax = fig.add_subplot(n_rows, n_cols, i+1)
_attention = attention.squeeze(0)[i].cpu().detach().numpy()
cax = ax.matshow(_attention, cmap='bone')
ax.tick_params(labelsize=12)
ax.set_xticklabels(['']+['<sos>']+[t.lower() for t in sentence]+['<eos>'],
rotation=45)
ax.set_yticklabels(['']+translation)
ax.xaxis.set_major_locator(ticker.MultipleLocator(1))
ax.yaxis.set_major_locator(ticker.MultipleLocator(1))
plt.show()
# + id="TJHNiBOoPQhG" colab={"base_uri": "https://localhost:8080/"} outputId="53a9f93b-1042-46b5-a643-ed60f1a7fae5"
example_idx = 11
src = vars(train_data.examples[example_idx])['Question']
trg = vars(train_data.examples[example_idx])['Answer']
print(f'src = {src}')
print(f'trg = {trg}')
# + id="dZSai6VDPR3o"
# translation, attention = translate_sentence(Question, TEXT, TEXT, model, device)
# print(f'predicted trg = {translation}')
# + id="CHWqhmvtPTJv"
# display_attention(src, translation, attention)
# + id="0YyJFiJXPVjr" colab={"base_uri": "https://localhost:8080/"} outputId="a7558756-107c-445c-e84d-22c4bb64a0c7"
example_idx = 1
src = vars(valid_data.examples[example_idx])['Question']
trg = vars(valid_data.examples[example_idx])['Answer']
print(f'src = {src}')
print(f'trg = {trg}')
# + id="LDmsvAhoPWze" colab={"base_uri": "https://localhost:8080/"} outputId="78aa1d66-1d31-4590-a7e2-033b50fcdc6b"
translation, attention = translate_sentence(src, TEXT, TEXT, model, device)
print(f'predicted trg = {translation}')
# + colab={"base_uri": "https://localhost:8080/"} id="iQqz12eRrQY4" outputId="ffa586e4-7de5-4184-c504-1798b21c370e"
example_idx = 19
src = vars(valid_data.examples[example_idx])['Question']
trg = vars(valid_data.examples[example_idx])['Answer']
print(f'src = {src}')
print(f'trg = {trg}')
# + id="9k_YvNBPPX02" colab={"base_uri": "https://localhost:8080/"} outputId="111d7491-16d0-4fca-8925-d39f993ba1c3"
translation, attention = translate_sentence(src, TEXT, TEXT, model, device)
print(f'predicted trg = {translation}')
# + id="tBp2QYMzQM4F" colab={"base_uri": "https://localhost:8080/"} outputId="5c38d48d-6766-4835-85ae-b436b826696b"
example_idx = 39
src = vars(valid_data.examples[example_idx])['Question']
trg = vars(valid_data.examples[example_idx])['Answer']
# print(f'src = {src}')
listToStr = ' '.join([str(elem) for elem in src])
print(f'src: {listToStr}')
# print(f'trg = {trg}')
listToStr = ' '.join([str(elem) for elem in trg])
print(f'Target:\n{listToStr}')
# + colab={"base_uri": "https://localhost:8080/"} id="2q8wN9lhDY6n" outputId="f899efb9-3386-4ceb-8927-f479e867613c"
translation, attention = translate_sentence(src, TEXT, TEXT, model, device)
print(f'predicted trg = {translation}')
# + colab={"base_uri": "https://localhost:8080/"} id="6_wAScirOkPB" outputId="7da3ede0-6f0a-459a-f63c-e37b1ab264ce"
# # output = []
# for x in translation:
# output.append(x)
# # print(x)
# print(output)
listToStr = ' '.join([str(elem) for elem in translation])
print(listToStr)
# + [markdown] id="cDnyJ4qwakfc"
# ### Model Prediction to generate Python Code on 25 Random Python Question
# + colab={"base_uri": "https://localhost:8080/"} id="BOQolLBzWHGx" outputId="7dc176be-0c31-4631-cf7e-6d596adb9742"
import random
#Generate 5 random numbers between 10 and 30
randomlist = random.sample(range(0, len(valid_data)), 25)
for ele in randomlist:
example_idx = ele
src = vars(valid_data.examples[example_idx])['Question']
trg = vars(valid_data.examples[example_idx])['Answer']
# print(f'src = {src}')
listToStr = ' '.join([str(elem) for elem in src])
print(f'Question: {listToStr}')
listToStr = ' '.join([str(elem) for elem in trg])
print(f'Source Python:\n{listToStr}')
print(f'\n')
# print(f'\n')
# print(f'trg = {trg}')
listToStr = ' '.join([str(elem) for elem in trg])
translation, attention = translate_sentence(src, TEXT, TEXT, model, device)
listToStr = ' '.join([str(elem) for elem in translation])
listToStrx = listToStr.replace('<eos>', '')
print(f'Target Python:\n{listToStrx}')
print('#########################################################################################################')
print('#########################################################################################################')
# + colab={"base_uri": "https://localhost:8080/"} id="eUmHFhfEqYlI" outputId="a09550a0-1b36-41c7-9c78-c540f6ea8618"
import random
#Generate 5 random numbers between 10 and 30
randomlist = random.sample(range(0, len(valid_data)), 25)
for ele in randomlist:
example_idx = ele
src = vars(valid_data.examples[example_idx])['Question']
trg = vars(valid_data.examples[example_idx])['Answer']
# print(f'src = {src}')
listToStr = ' '.join([str(elem) for elem in src])
print(f'Question: {listToStr}')
listToStr = ' '.join([str(elem) for elem in trg])
print(f'Source Python:\n{listToStr}')
print(f'\n')
# print(f'\n')
# print(f'trg = {trg}')
listToStr = ' '.join([str(elem) for elem in trg])
translation, attention = translate_sentence(src, TEXT, TEXT, model, device)
listToStr = ' '.join([str(elem) for elem in translation])
listToStrx = listToStr.replace('<eos>', '')
print(f'Target Python:\n{listToStrx}')
print('#########################################################################################################')
print('#########################################################################################################')
# + colab={"base_uri": "https://localhost:8080/"} id="4a1ttZSFqeTM" outputId="886ba345-6c42-4cfa-c76c-7415768b0a13"
import random
#Generate 5 random numbers between 10 and 30
randomlist = random.sample(range(0, len(valid_data)), 25)
for ele in randomlist:
example_idx = ele
src = vars(valid_data.examples[example_idx])['Question']
trg = vars(valid_data.examples[example_idx])['Answer']
# print(f'src = {src}')
listToStr = ' '.join([str(elem) for elem in src])
print(f'Question: {listToStr}')
listToStr = ' '.join([str(elem) for elem in trg])
print(f'Source Python:\n{listToStr}')
print(f'\n')
# print(f'\n')
# print(f'trg = {trg}')
listToStr = ' '.join([str(elem) for elem in trg])
translation, attention = translate_sentence(src, TEXT, TEXT, model, device)
listToStr = ' '.join([str(elem) for elem in translation])
listToStrx = listToStr.replace('<eos>', '')
print(f'Target Python:\n{listToStrx}')
print('#########################################################################################################')
print('#########################################################################################################')
| transformer-based-model-python-code-generator/src/END_NLP_CAPSTONE_PROJECT_English_Python_Code_Transformer_3_0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Db2 Macros
# The `%sql` command also allows the use of macros. Macros are used to substitute text into SQL commands that you execute. Macros substitution is done before any SQL is executed. This allows you to create macros that include commonly used SQL commands or parameters rather than having to type them in. Before using any macros, we must make sure we have loaded the Db2 extensions.
# %run db2.ipynb
# ### Macro Basics
# A Macro command begins with a percent sign (`%` similar to the `%sql` magic command) and can be found anywhere within a `%sql` line or `%%sql` block. Macros must be separated from other text in the SQL with a space.
#
# To define a macro, the `%%sql macro <name>` command is used. The body of the macro is found in the cell below the definition of the macro. This simple macro called EMPTABLE will substitute a SELECT statement into a SQL block.
# + magic_args="macro emptable" language="sql"
# select * from employee
# -
# The name of the macro follows the `%%sql macro` command and is case sensitive. To use the macro, we can place it anywhere in the `%sql` block. This first example uses it by itself.
# %sql %emptable
# The actual SQL that is generated is not shown by default. If you do want to see the SQL that gets generated, you can use the `-e` (echo) option to display the final SQL statement. The following example will display the generated SQL. Note that the echo setting is only used to display results for the current cell that is executing.
# + magic_args="-e" language="sql"
# %emptable
# -
# Since we can use the `%emptable` anywhere in our SQL, we can add additional commands around it. In this example we add some logic to the select statement.
# + language="sql"
# %emptable
# where empno = '000010'
# -
# Macros can also have parameters supplied to them. The parameters are included after the name of the macro. Here is a simple macro which will use the first parameter as the name of the column we want returned from the EMPLOYEE table.
# + magic_args="macro emptable" language="sql"
# SELECT {1} FROM EMPLOYEE
# -
# This example illustrates two concepts. The `MACRO` command will replace any existing macro with the same name. Since we already have an emptable macro, the macro body will be replaced with this code. In addition, macros only exist for the duration of your notebook. If you create another Jupyter notebook, it will not contain any macros that you may have created. If there are macros that you want to share across notebooks, you should create a separate notebook and place all of the macro definitions in there. Then you can include these macros by executing the `%run` command using the name of the notebook that contains the macros.
#
# The following SQL shows the use of the macro with parameters.
# + language="sql"
# %emptable(lastname)
# -
# The remainder of this notebook will explore the advanced features of macros.
# ## Macro Parameters
# Macros can have up to 9 parameters supplied to them. The parameters are numbered from 1 to 9, left to right in the argument list for the macro. For instance, the following macro has 5 paramters:
# ```
# # %emptable(lastname,firstnme,salary,bonus,'000010')
# ```
# Parameters are separated by commas, and can contain strings as shown using single or double quotes. When the parameters are used within a macro, the quotes are not included as part of the string. If you do want to pass the quotes as part of the parameter, use square brackets **[]** around the string. For instance, the following parameter will not have quotes passed to the macro:
#
# ```python
# # %sql %abc('no quotes')
# ```
#
# To send the string with quotes, you could surround the parameter with other quotes **`"'hello'"`** or use the following technique if you use multiple quotes in your string:
#
# ```python
# # %sql %abc (['quotes'])
# ```
#
# To use a parameter within your macro, you enclose the parameter number with braces `{}`. The next command will illustrate the use of the five parameters.
# + magic_args="macro emptable" language="sql"
# display on
# SELECT {1},{2},{3},{4}
# FROM EMPLOYEE
# WHERE EMPNO = '{5}'
# -
# Note that the `EMPNO` field is a character field in the `EMPLOYEE` table. Even though the employee number was supplied as a string, the quotes are not included in the parameter. The macro places quotes around the parameter `{5}` so that it is properly used in the SQL statement. The other feature of this macro is that the display (on) command is part of the macro body so the generated SQL will always be displayed.
# %sql %emptable(lastname,firstnme,salary,bonus,'000010')
# We can modify the macro to assume that the parameters will include the quotes in the string.
# + magic_args="macro emptable" language="sql"
# SELECT {1},{2},{3},{4}
# FROM EMPLOYEE
# WHERE EMPNO = {5}
# -
# We just have to make sure that the quotes are part of the parameter now.
# %sql -e %emptable(lastname,firstnme,salary,bonus,"'000010'")
# We could use the square brackets as an alternative way of passing the parameter.
# %sql -e %emptable(lastname,firstnme,salary,bonus,['000010'])
# Parameters can also be named in a macro. To name an input value, the macro needs to use the format:
# ```
# field=value
# ```
# For instance, the following macro call will have 2 numbered parameters and one named parameter:
# ```
# # %showemp(firstnme,lastname,logic="WHERE EMPNO='000010'")
# ```
# From within the macro the parameter count would be 2 and the value for parameter 1 is `firstnme`, and the value for parameter 2 is `lastname`. Since we have a named parameter, it is not included in the list of numbered parameters. In fact, the following statement is equivalent since unnamed parameters are numbered in the order that they are found in the macro, ignoring any named parameters that are found:
# ```
# # %showemp(firstnme,logic="WHERE EMPNO='000010'",lastname)
# ```
# The following macro illustrates this feature.
# + magic_args="macro showemp" language="sql"
# SELECT {1},{2} FROM EMPLOYEE
# {logic}
# -
# %sql %showemp(firstnme,lastname,logic="WHERE EMPNO='000010'")
# %sql %showemp(firstnme,logic="WHERE EMPNO='000010'",lastname)
# Named parameters are useful when there are many options within the macro and you don't want to keep track of which position it is in. In addition, if you have a variable number of parameters, you should use named parameters for the fixed (required) parameters and numbered parameters for the optional ones.
# ## Macro Coding Overview
# Macros can contain any type of text, including SQL commands. In addition to the text, macros can also contain the following keywords:
#
# * echo - Display a message
# * exit - Exit the macro immediately
# * if/else/endif - Conditional logic
# * var - Set a variable
# * display - Turn the display of the final text on
#
# The only restriction with macros is that macros cannot be nested. This means I can't call a macro from within a macro. The sections below explain the use of each of these statement types.
# ### Echo Option
# The `-e` option will result in the final SQL being display after the macro substitution is done.
# ```
# # %%sql -e
# # %showemp(...)
# ```
# + magic_args="macro showdisplay" language="sql"
# SELECT * FROM EMPLOYEE FETCH FIRST ROW ONLY
# -
# Using the `-e` flag will display the final SQL that is run.
# %sql -e %showdisplay
# If we remove the `-e` option, the final SQL will not be shown.
# %sql %showdisplay
# ### Exit Command
# The `exit` command will terminate the processing within a macro and not run the generated SQL. You would use this when a condition is not met within the macro (like a missing parameter).
# + magic_args="macro showexit" language="sql"
# echo This message gets shown
# SELECT * FROM EMPLOYEE FETCH FIRST ROW ONLY
# exit
# echo This message does not get shown
# -
# The macro that was defined will not show the second statement, nor will it execute the SQL that was defined in the macro body.
# %sql %showexit
# ### Echo Command
# As you already noticed in the previous example, the `echo` command will display information on the screen. Any text following the command will have variables substituted and then displayed with a green box surrounding it. The following code illustates the use of the command.
# + magic_args="macro showecho" language="sql"
# echo Here is a message
# echo Two lines are shown
# -
# The echo command will show each line as a separate box.
# %sql %showecho
# If you want to have a message go across multiple lines use the `<br>` to start a new line.
# + magic_args="macro showecho" language="sql"
# echo Here is a paragraph. <br> And a final paragraph.
# -
# %sql %showecho
# ### Var Command
# The var (variable) command sets a macro variable to a value. A variable is referred to in the macro script using curly braces `{name}`. By default the arguments that are used in the macro call are assigned the variable names `{1}` to `{9}`. If you use a named argument (option="value") in the macro call, a variable called `{option}` will contain the value within the macro.
#
# To set a variable within a macro you would use the `var` command:
# ```
# var name value
# ```
# The variable name can be any name as long as it only includes letters, numbers, underscore `_` and `$`. Variable names are case sensitive so `{a}` and `{A}` are different. When the macro finishes executing, the contents of the variables will be lost. If you do want to keep a variable between macros, you should start the name of the variable with a `$` sign:
# ```
# var $name value
# ```
# This variable will persist between macro calls.
# + magic_args="macro initialize" language="sql"
# var $hello Hello There
# var hello You won't see this
# + magic_args="macro runit" language="sql"
# echo The value of hello is *{hello}*
# echo {$hello}
# -
# Calling `runit` will display the variable that was set in the first macro.
# %sql %initialize
# %sql %runit
# A variable can be converted to uppercase by placing the `^` beside the variable name or number.
# + magic_args="macro runit" language="sql"
# echo The first parameter is {^1}
# -
# %sql %runit(Hello There)
# The string following the variable name can include quotes and these will not be removed. Only quotes that are supplied in a parameter to a macro will have the quotes removed.
# + magic_args="macro runit" language="sql"
# var hello This is a long string without quotes
# var hello2 'This is a long string with quotes'
# echo {hello} <br> {hello2}
# -
# %sql %runit
# When passing parameters to a macro, the program will automatically create variables based on whether they are positional parameters (1, 2, ..., n) or named parameters. The following macro will be used to show how parameters are passed to the routine.
# + magic_args="macro showvar" language="sql"
# echo parm1={1} <br>parm2={2} <br>message={message}
# -
# Calling the macro will show how the variable names get assigned and used.
# %sql %showvar(parameter 1, another parameter,message="Hello World")
# If you pass an empty value (or if a variable does not exist), a "null" value will be shown.
# %sql %showvar(1,,message="Hello World")
# An empty string also returns a null value.
# %sql %showvar(1,2,message="")
# Finally, any string that is supplied to the macro will not include the quotes in the variable. The Hello World string will not have quotes when it is displayed:
# %sql %showvar(1,2,message="Hello World")
# You need to supply the quotes in the script or macro when using variables since quotes are stripped from any strings that are supplied.
# + magic_args="macro showvar" language="sql"
# echo parm1={1} <br>parm2={2} <br>message='{message}'
# -
# %sql %showvar(1,2,message="Hello World")
# The count of the total number of parameters passed is found in the `{argc}` variable. You can use this variable to decide whether or not the user has supplied the proper number of arguments or change which code should be executed.
# + magic_args="macro showvar" language="sql"
# echo The number of unnamed parameters is {argc}. The where clause is *{where}*.
# -
# Unnamed parameters are included in the count of arguments while named parameters are ignored.
# %sql %showvar(1,2,option=nothing,3,4,where=)
# ### If/Else/Endif Command
# If you need to add conditional logic to your macro then you should use the `if/else/endif` commands. The format of the `if` statement is:
# ```
# if variable condition value
# statements
# else
# statements
# endif
# ```
# The else portion is optional, but the block must be closed with the `endif` command. If statements can be nested up to 9 levels deep:
# ```
# if condition 1
# if condition 2
# statements
# else
# if condition 3
# statements
# end if
# endif
# endif
# ```
# If the condition in the if clause is true, then anything following the if statement will be executed and included in the final SQL statement. For instance, the following code will create a SQL statement based on the value of parameter 1:
# ```
# if {1} = null
# SELECT * FROM EMPLOYEE
# else
# SELECT {1} FROM EMPLOYEE
# endif
# ```
# #### Conditions
# The `if` statement requires a condition to determine whether or not the block should be executed. The condition uses the following format:
# ```
# if {variable} condition {variable} | constant | null
# ```
# `Variable` can be a number from 1 to 9 which represents the argument in the macro list. So `{1}` refers to the first argument. The variable can also be the name of a named parameter or global variable.
#
# The condition is one of the following comparison operators:
# - `=`, `==`: Equal to
# - `<`: Less than
# - `>`: Greater than
# - `<=`,`=<`: Less than or equal to
# - `>=`, `=>`: Greater than or equal to
# - `!=`, `<>` : Not equal to
#
# The variable or constant will have quotes stripped away before doing the comparison. If you are testing for the existence of a variable, or to check if a variable is empty, use the keyword `null`.
# + magic_args="macro showif" language="sql"
# if {argc} = 0
# echo No parameters supplied
# if {option} <> null
# echo The optional parameter option was set: {option}
# endif
# else
# if {argc} = "1"
# echo One parameter was supplied
# else
# echo More than one parameter was supplied: {argc}
# endif
# endif
# -
# Running the previous macro with no parameters will check to see if the option keyword was used.
# %sql %showif
# Now include the optional parameter.
# %sql %showif(option="Yes there is an option")
# Finally, issue the macro with multiple parameters.
# %sql %showif(Here,are,a,number,of,parameters)
# One additional option is available for variable substitution. If the first character of the variable name or parameter number is the `^` symbol, it will uppercase the entire string.
# + magic_args="macro showif" language="sql"
# if {option} <> null
# echo The optional parameter option was set: {^option}
# endif
# -
# %sql %showif(option="Yes there is an option")
# #### Credits: IBM 2018, <NAME> [<EMAIL>]
| Db2 Jupyter Macros.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# orphan: true
# ---
# (tune-lightgbm-example)=
#
# # Using LightGBM with Tune
#
# ```{image} /images/lightgbm_logo.png
# :align: center
# :alt: LightGBM Logo
# :height: 120px
# :target: https://lightgbm.readthedocs.io
# ```
#
# ```{contents}
# :backlinks: none
# :local: true
# ```
#
# ## Example
# + pycharm={"name": "#%%\n"}
import lightgbm as lgb
import numpy as np
import sklearn.datasets
import sklearn.metrics
from sklearn.model_selection import train_test_split
from ray import tune
from ray.tune.schedulers import ASHAScheduler
from ray.tune.integration.lightgbm import TuneReportCheckpointCallback
def train_breast_cancer(config):
data, target = sklearn.datasets.load_breast_cancer(return_X_y=True)
train_x, test_x, train_y, test_y = train_test_split(data, target, test_size=0.25)
train_set = lgb.Dataset(train_x, label=train_y)
test_set = lgb.Dataset(test_x, label=test_y)
gbm = lgb.train(
config,
train_set,
valid_sets=[test_set],
valid_names=["eval"],
verbose_eval=False,
callbacks=[
TuneReportCheckpointCallback(
{
"binary_error": "eval-binary_error",
"binary_logloss": "eval-binary_logloss",
}
)
],
)
preds = gbm.predict(test_x)
pred_labels = np.rint(preds)
tune.report(
mean_accuracy=sklearn.metrics.accuracy_score(test_y, pred_labels), done=True
)
if __name__ == "__main__":
import argparse
parser = argparse.ArgumentParser()
parser.add_argument(
"--server-address",
type=str,
default=None,
required=False,
help="The address of server to connect to if using " "Ray Client.",
)
args, _ = parser.parse_known_args()
if args.server_address:
import ray
ray.init(f"ray://{args.server_address}")
config = {
"objective": "binary",
"metric": ["binary_error", "binary_logloss"],
"verbose": -1,
"boosting_type": tune.grid_search(["gbdt", "dart"]),
"num_leaves": tune.randint(10, 1000),
"learning_rate": tune.loguniform(1e-8, 1e-1),
}
analysis = tune.run(
train_breast_cancer,
metric="binary_error",
mode="min",
config=config,
num_samples=2,
scheduler=ASHAScheduler(),
)
print("Best hyperparameters found were: ", analysis.best_config)
| doc/source/tune/examples/lightgbm_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
# Required only when running on Windows
# Add the latest files to the path
# %matplotlib inline
import sys
import pandas as pd
#import seaborn as sns
from sklearn import preprocessing
from matplotlib import pyplot as plt
#sys.path.insert(0, r'\\newwinsrc\sasgen\dev\mva-vb005\GTKWX6ND\misc\python')
# -
# ## Connect to CAS
from swat import *
cassession = CAS('sasserver.demo.sas.com', 5570, authinfo='~/.authinfo', caslib="casuser")
cassession.loadactionset(actionset="table")
# ## Check the server status
#
cassession.builtins.serverstatus()
# ----
# ## Load data set
# Show available caslibs
cassession.caslibinfo()
#result = cassession.upload('/home/viyauser/SamsViya/ViyaRace/hmeq.sas7bdat')
#result
indata_dir = 'data'
indata = 'hmeq'
result = cassession.loadTable(indata_dir + '/' + indata + '.sas7bdat', casout = indata)
# +
# Generate an ID column for the data set. Will allow us to uniquely identify rows
# NOTE: use of _N_ is not recommended, especially in MPP environments
# TODO: Replace with SQL when possible.
cassession.loadactionset('datastep')
cassession.datastep.runcode("""
DATA hmeq;
ID=_N_;
SET hmeq;
RUN;
""")
# +
# Save pointer to in-memory table in CAS
hmeq = cassession.CASTable('hmeq')
hmeq.head()
# Bad = 1 if applicat defaulted on the loan
# Loan = Requested loan amount
# MortDue = Amount due on existing mortgage
# Value = Value of current property
# Reason = DebtCon / HomeImp
# Job = Occupation category
# YoJ = Years at current job
# Derog = # of major derogatory reports
# Delinq = Number of delinquent credit lines
# CLAge = Age of oldest credit line in months
# NInq = # of recent credit inquiries
# CLNo = # of credit lines
# DebtInc = Debt:Income ratio
# -
# ----
# ## Load action sets
# +
# %%capture
# Instruct CAS to load action sets if not already loaded.
#cas.loadactionset('cardinality')
#cas.loadactionset('dataPreprocess')
#cas.loadactionset('varReduce')
#cas.loadactionset('clustering')
#cas.loadactionset('pca')
#cas.loadactionset('sampling')
#cas.loadactionset('decisionTree')
#cas.loadactionset('dataStep')
#cas.loadactionset('neuralNet')
#cas.loadactionset('svm')
#cas.loadactionset('astore')
#cas.loadactionset('fedsql')
#cas.loadactionset('percentile')
# create python list containing the required CAS Actionsets
casActionsets = ['cardinality','dataPreprocess','varReduce','clustering','pca','sampling','decisionTree','dataStep','neuralNet','svm','astore','fedsql','percentile']
# use list comprehension to loade these actionsets
[cassession.loadactionset(actionset=i) for i in casActionsets]
# list loaded actionsets
cassession.actionsetinfo(all=False)
# -
# ----
# ## Explore Data
# Calculate summary statistics for each variable
hmeq.summarize(cardinality=dict(name='hmeq_card', replace=True))
# +
# Link to results and filter to just variables with missing values
hmeq_card = cassession.CASTable('hmeq_card', where='_NMISS_ > 0')
# Pull the results from the server. Returned as child class of Pandas DF
df_hmeq_card = hmeq_card.fetch().Fetch
df_hmeq_card
# Note: What we just did is to calcualate summary statistics on the columns of the table in CAS, then return these
# results back to local Python ina pandas dataframe. We would NEVER do this with a large CAS table but this
# is very handy when dealing with small results. So we can go back and forth with the computation locality.
# +
# Calculate percentage of each variable that's missing
# Performed locally using Pandas
df_hmeq_card['Percent_Missing'] = df_hmeq_card._NMISS_ / df_hmeq_card._NOBS_ * 100
# Plot results
# Performed locally using Pandas Bar Plotting
df_hmeq_card.plot.bar('_VARNAME_','Percent_Missing', title='Percentage of Missing Values', legend=False)
# -
# ----
# ## Impute missing values
# +
# Note: We are now using a pointer to a cas table, so we will be executing in CAS, not locally. The following command
# is a python function that returns the "type" of the object - when demoing you can switch out between the two statements
# to show insight into computation locality (the print function forces each result to be shown, otherwise Jupyter will only
# show the last function result)
print(type(hmeq))
print(type(df_hmeq_card))
# -
# Preview data. Credit Line Age (CLAge) has missing values. The head function is a CAS function that mimics the python head
hmeq.head()
# +
# Impute missing values for a single column (clage)
result = hmeq.datapreprocess.impute(casout =dict(name='hmeq_prepped', replace=True),
allIdVars =True,
techForCont='MEAN',
vars ='clage')
hmeq_prepped = result.OutputCasTables.ix[0,'casTable']
result.ImputeInfo
# +
# Select min and max values for NInq column using SQL
ninq_range = cassession.fedsql.execdirect("SELECT _min_, _max_ FROM hmeq_card WHERE _varname_='NINQ'")['Result Set']
ninq_min = ninq_range['_MIN_'][0]
ninq_max = ninq_range['_MAX_'][0]
# Pipeline multiple imputation steps.
result = hmeq.datapreprocess.transform(casOut=dict(name='hmeq_prepped', replace=True),
allIdVars=True,
outVarsNameGlobalPrefix='IM',
reqPacks=[
dict(impute='mean', inputs='clage'),
dict(impute='median', inputs='delinq'),
dict(impute={'method':'random', 'minrandom':ninq_min, 'maxrandom':ninq_max},
inputs='ninq'),
dict(impute={'method':'value', 'valuesContinuous':[50,100]},
inputs=['debtinc','yoj'])
])
hmeq_prepped = result.OutputCasTables.ix[0,'casTable']
result.VarTransInfo
# -
print(type(hmeq_prepped))
hmeq_prepped.head()
# ----
# ## Dimensionality reduction
# +
model = dict(depvars='bad',
effects=[{'vars':['loan','mortdue','value','reason','job','yoj','derog','delinq',
'clage','ninq','clno','debtinc','im_clage','im_debtinc','im_delinq',
'im_ninq','im_yoj']}])
# Perform supervised variable selection using discriminant analysis. Return max of 8 features.
hmeq_prepped.varreduce.super(analysis='DSC', class_=[dict(vars=['bad','reason','job'])],
model=model,
maxeffects=8,
outputtables=dict(names=['SelectionSummary','SelectedEffects']))
# +
# Retrieve the selection metrics from CAS
summ = cassession.CASTable('SelectionSummary').fetch().Fetch
# Rescale the values using Sklearn so they can be plotted together
cols = ['VarExp', 'MSE','AIC','BIC','SSE','AICC']
summ[cols] = preprocessing.MinMaxScaler().fit_transform(summ[cols].values)
# Reshape using Pandas
summ = pd.melt(summ, id_vars=['Iteration','Variable','Parameter'])
# Plot using Seaborn
# sns.pointplot(x='Iteration', y='value', hue='variable', data=summ, scale=0.75)
# -
# ----
# ## K-Means clustering
# Use K-means to compute cluster assignments
hmeq_prepped.clustering.kclus(inputs=['loan','im_debtinc','im_clage',],
distance='EUCLIDEAN',
standardize='STD',
nclusters=6,
maxIters=50,
output=dict(casout=dict(name='hmeq_clusters', replace=True), copyvars='ALL'),
display=dict(names=['ModelInfo','ClusterSum','ClusterCenters']))
cassession.CASTable('hmeq_clusters').head()
# ----
# ## Partition data
# +
# Build list of retained features using results from varreduce
columns = ['BAD']
columns.extend(list(cassession.CASTable('SelectedEffects').fetch().Fetch['Variable']))
columns = ', '.join(['prep.{}'.format(x) for x in columns])
# Join tables and select feature set
# Using in-line FedSQL options to control CAS behavior
query = """CREATE TABLE hmeq_reduced {{options replace=true, replication=0}} AS (
SELECT {},
clust._cluster_id_ as cluster
FROM hmeq_prepped as prep
LEFT OUTER JOIN hmeq_clusters as clust ON prep.id=clust.id
)
""".format(columns)
cassession.fedsql.execdirect(query)
# -
# Split the data into training and crossval sets. 70/30 split, adding an indicator column
hmeq_part = cassession.sampling.stratified(table='hmeq_reduced',
sampPct=70,
partInd=True,
output=dict(casout=dict(name='hmeq_part', replace=True),
copyvars='ALL')).OutputCasTables.ix[0,'casTable']
hmeq_part.head()
# Define some commonly used inputs
train = cassession.CASTable(name='hmeq_part', where='_PartInd_ = 1')
crossval = cassession.CASTable(name='hmeq_part', where='_PartInd_ = 0')
features = ['delinq','derog','clage','job','ninq','clno','loan','cluster']
nominal_fields = ['bad', 'job']
# ----
# ## Gradient Boosting
# Train a gradient boosting model
hmeq_part.decisiontree.gbtreetrain(table=train, inputs=features, nominals=nominal_fields,
target='bad',
nTree=10,
nBins=20,
varImp=True,
missing='USEINSEARCH',
casOut=dict(name='gb_model', replace=True)
)
# +
# Score the model using the crossvalidation data
result = crossval.decisiontree.gbtreescore(model='gb_model',
casout=dict(name='hmeq_scored_gb', replace=True),
copyVars=['bad','_PartInd_'])
# Save a pointer to the output in CAS
hmeq_scored_gb = result.OutputCasTables.ix[0,'casTable']
# Display summary of results
result.ScoreInfo
# -
hmeq_scored_gb.head()
# +
# Use the Assess action on the scored crossvalidation data to generate ROC and Lift statistics
results2 = hmeq_scored_gb[hmeq_scored_gb['_PartInd_'] == 0].percentile.assess(
inputs='P_Bad1',
response='Bad',
event='1',
pVar='P_Bad0',
pEvent='0'
)
# Extract the ROC stats
roc2 = results2.ROCInfo
# -
# ---
# ## Neural Network
# +
# Fit a neural network to the data
result = train.neuralNet.annTrain(validTable=crossval,
inputs=features,
nominals=nominal_fields,
target='bad',
hiddens=2,
acts='TANH',
combs='LINEAR',
targetAct='SOFTMAX',
errorFunc='ENTROPY',
std='MIDRANGE',
randDist='UNIFORM',
scaleInit=1,
nloOpts=dict(
optmlOpt = dict(maxIters=100, fConv=1e-10),
lbfgsOpt = dict(numCorrections=6),
printOpt = dict(printLevel='printDetail'),
validate = dict(frequency=1)
),
casOut=dict(name='nn_model', replace=True)
)
# Plot the error by iteration
result.OptIterHistory[['Loss','Progress']].plot(x='Progress')
# +
# Use the model to score the training & crossvalidation data
result = cassession.neuralNet.annScore(
table=hmeq_part,
modelTable='nn_model',
casOut=dict(name='nn_scored', replace=True),
copyVars=['BAD', '_PartInd_']
)
nn_scored = cassession.CASTable('nn_scored')
nn_scored.head()
# +
#cassession.tableinfo()
#cassession.columninfo('nn_scored')
cassession.columninfo('HMEQ_SCORED_GB')
# -
# _NN_PredP_ is the probability associated with the predicted value
# Convert this into separate columns for the probability that Bad=0 and Bad=1
def prep_roc(ctable,predname,predp):
cassession.dataStep.runCode("""
DATA """ + ctable + """;
SET """ + ctable + """;
IF """ + predname + """=1 THEN do;
p_bad1=""" + predp + """;
p_bad0=1-p_bad1;
END;
IF """ + predname + """=0 THEN do;
p_bad0=""" + predp + """;
p_bad1=1-p_bad0;
END;
RUN;
""")
prep_roc("nn_scored","_NN_PredName_","_NN_PredP_")
prep_roc("HMEQ_SCORED_GB","_GBT_PredName_","_GBT_PredP_")
hmeq_scored_gb.head()
# +
# Use the Assess action on the scored crossvalidation data to generate ROC and Lift statistics
results1 = nn_scored[nn_scored['_PartInd_'] == 0].percentile.assess(
inputs='P_Bad1',
response='Bad',
event='1',
pVar='P_Bad0',
pEvent='0'
)
# Extract the ROC stats
roc1 = results1.ROCInfo
# +
# Plot the ROC curve
plt.figure()
plt.plot(roc1["FPR"], roc["Sensitivity"])
plt.plot(roc2["FPR"], roc["Sensitivity"])
plt.plot([0,1], [0,1], "k--")
plt.xlabel("False Positive Rate")
plt.ylabel("True Positive Rate")
plt.legend(['Neural','GD'],loc="best")
plt.title("ROC Curve (using validation data)")
plt.show()
# -
# Terminate the session
cassession.close()
| .ipynb_checkpoints/hmeq-intro-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Version Check
# Plotly's python package is updated frequently. Run `pip install plotly --upgrade` to use the latest version.
import plotly
plotly.__version__
# ### Adding Text to Data in Line and Scatter Plots
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace1 = go.Scatter(
x=[0, 1, 2],
y=[1, 1, 1],
mode='lines+markers+text',
name='Lines, Markers and Text',
text=['Text A', 'Text B', 'Text C'],
textposition='top center'
)
trace2 = go.Scatter(
x=[0, 1, 2],
y=[2, 2, 2],
mode='markers+text',
name='Markers and Text',
text=['Text D', 'Text E', 'Text F'],
textposition='bottom center'
)
trace3 = go.Scatter(
x=[0, 1, 2],
y=[3, 3, 3],
mode='lines+text',
name='Lines and Text',
text=['Text G', 'Text H', 'Text I'],
textposition='bottom center'
)
data = [trace1, trace2, trace3]
layout = go.Layout(
showlegend=False
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='text-chart-basic')
# -
# ### Adding Hover Text to Data in Line and Scatter Plots
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [
go.Scatter(
x=[0, 1, 2],
y=[1, 3, 2],
mode='markers',
text=['Text A', 'Text B', 'Text C']
)
]
layout = go.Layout(
title='Hover over the points to see the text'
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='hover-chart-basic')
# -
# ### Simple Annotation
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace1 = go.Scatter(
x=[0, 1, 2, 3, 4, 5, 6, 7, 8],
y=[0, 1, 3, 2, 4, 3, 4, 6, 5]
)
trace2 = go.Scatter(
x=[0, 1, 2, 3, 4, 5, 6, 7, 8],
y=[0, 4, 5, 1, 2, 2, 3, 4, 2]
)
data = [trace1, trace2]
layout = go.Layout(
showlegend=False,
annotations=[
dict(
x=2,
y=5,
xref='x',
yref='y',
text='dict Text',
showarrow=True,
arrowhead=7,
ax=0,
ay=-40
)
]
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='simple-annotation')
# -
# ### Multiple Annotations
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace1 = go.Scatter(
x=[0, 1, 2, 3, 4, 5, 6, 7, 8],
y=[0, 1, 3, 2, 4, 3, 4, 6, 5]
)
trace2 = go.Scatter(
x=[0, 1, 2, 3, 4, 5, 6, 7, 8],
y=[0, 4, 5, 1, 2, 2, 3, 4, 2]
)
data = [trace1, trace2]
layout = go.Layout(
showlegend=False,
annotations=[
dict(
x=2,
y=5,
xref='x',
yref='y',
text='dict Text',
showarrow=True,
arrowhead=7,
ax=0,
ay=-40
),
dict(
x=4,
y=4,
xref='x',
yref='y',
text='dict Text 2',
showarrow=True,
arrowhead=7,
ax=0,
ay=-40
)
]
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='multiple-annotation')
# -
# ### 3D Annotations
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [go.Scatter3d(
x = ["2017-01-01", "2017-02-10", "2017-03-20"],
y = ["A", "B", "C"],
z = [1, 1000, 100000],
name = "z",
)]
layout = go.Layout(
scene = dict(
aspectratio = dict(
x = 1,
y = 1,
z = 1
),
camera = dict(
center = dict(
x = 0,
y = 0,
z = 0
),
eye = dict(
x = 1.96903462608,
y = -1.09022831971,
z = 0.405345349304
),
up = dict(
x = 0,
y = 0,
z = 1
)
),
dragmode = "turntable",
xaxis = dict(
title = "",
type = "date"
),
yaxis = dict(
title = "",
type = "category"
),
zaxis = dict(
title = "",
type = "log"
),
annotations = [dict(
showarrow = False,
x = "2017-01-01",
y = "A",
z = 0,
text = "Point 1",
xanchor = "left",
xshift = 10,
opacity = 0.7
), dict(
x = "2017-02-10",
y = "B",
z = 4,
text = "Point 2",
textangle = 0,
ax = 0,
ay = -75,
font = dict(
color = "black",
size = 12
),
arrowcolor = "black",
arrowsize = 3,
arrowwidth = 1,
arrowhead = 1
), dict(
x = "2017-03-20",
y = "C",
z = 5,
ax = 50,
ay = 0,
text = "Point 3",
arrowhead = 1,
xanchor = "left",
yanchor = "bottom"
)]
),
xaxis = dict(title = "x"),
yaxis = dict(title = "y")
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename = "3d annotations")
# -
# ### Custom Text Color and Styling
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace1 = go.Scatter(
x=[0, 1, 2],
y=[1, 1, 1],
mode='lines+markers+text',
name='Lines, Markers and Text',
text=['Text A', 'Text B', 'Text C'],
textposition='top right',
textfont=dict(
family='sans serif',
size=18,
color='#1f77b4'
)
)
trace2 = go.Scatter(
x=[0, 1, 2],
y=[2, 2, 2],
mode='lines+markers+text',
name='Lines and Text',
text=['Text G', 'Text H', 'Text I'],
textposition='bottom center',
textfont=dict(
family='sans serif',
size=18,
color='#ff7f0e'
)
)
data = [trace1, trace2]
layout = go.Layout(
showlegend=False
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='text-chart-styling')
# -
# ### Styling and Coloring Annotations
# +
import plotly.plotly as py
import plotly.graph_objs as go
trace1 = go.Scatter(
x=[0, 1, 2, 3, 4, 5, 6, 7, 8],
y=[0, 1, 3, 2, 4, 3, 4, 6, 5]
)
trace2 = go.Scatter(
x=[0, 1, 2, 3, 4, 5, 6, 7, 8],
y=[0, 4, 5, 1, 2, 2, 3, 4, 2]
)
data = [trace1, trace2]
layout = go.Layout(
showlegend=False,
annotations=[
dict(
x=2,
y=5,
xref='x',
yref='y',
text='max=5',
showarrow=True,
font=dict(
family='Courier New, monospace',
size=16,
color='#ffffff'
),
align='center',
arrowhead=2,
arrowsize=1,
arrowwidth=2,
arrowcolor='#636363',
ax=20,
ay=-30,
bordercolor='#c7c7c7',
borderwidth=2,
borderpad=4,
bgcolor='#ff7f0e',
opacity=0.8
)
]
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='style-annotation')
# -
# ### Disabling Hover Text
# +
import plotly.plotly as py
trace = dict(
x=[1, 2, 3,],
y=[10, 30, 15],
type='scatter',
name='first trace',
hoverinfo='none'
)
py.iplot([trace], filename='hoverinfo=none')
# -
# ### Text Font as an Array - Styling Each Text Element
# +
import plotly.plotly as py
import plotly.graph_objs as go
fig = go.Figure(
data=[
go.Scattergeo(
lat=[45.5,43.4,49.13,51.1,53.34,45.24,44.64,48.25,49.89,50.45],
lon=[-73.57,-79.24,-123.06,-114.1,-113.28,-75.43,-63.57,-123.21,-97.13,-104.6],
marker={
"color": ["#bebada","#fdb462","#fb8072","#d9d9d9","#bc80bd","#b3de69","#8dd3c7","#80b1d3","#fccde5","#ffffb3"],
"line": {
"width": 1
},
"size": 10
},
mode="markers+text",
name="",
text=["Montreal","Toronto","Vancouver","Calgary","Edmonton","Ottawa","Halifax","Victoria","Winnepeg","Regina"],
textfont={
"color": ["#bebada","#fdb462","#fb8072","#d9d9d9","#bc80bd","#b3de69","#8dd3c7","#80b1d3","#fccde5","#ffffb3"],
"family": ["Arial, sans-serif","Balto, sans-serif","Courier New, monospace","Droid Sans, sans-serif","Droid Serif, serif","Droid Sans Mono, sans-serif","Gravitas One, cursive","Old Standard TT, serif","Open Sans, sans-serif","PT Sans Narrow, sans-serif","Raleway, sans-serif","Times New Roman, Times, serif"],
"size": [22,21,20,19,18,17,16,15,14,13]
},
textposition=["top center","middle left","top center","bottom center","top right","middle left","bottom right","bottom left","top right","top right"]
)
],
layout={
"title": "Canadian cities",
"geo": {
"lataxis": {
"range": [40, 70]
},
"lonaxis": {
"range": [-130, -55]
},
"scope": "north america"
}
}
)
py.iplot(fig, filename='Canadian Cities')
# -
# ### Adding Annotations with xref and yref as Paper
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [
go.Scatter(
x=[1, 2, 3],
y=[1, 2, 3],
name='y',
)
]
layout = go.Layout(
annotations=[
dict(
x=0.5004254919715793,
y=-0.16191064079952971,
showarrow=False,
text='Custom x-axis title',
xref='paper',
yref='paper'
),
dict(
x=-0.04944728761514841,
y=0.4714285714285711,
showarrow=False,
text='Custom y-axis title',
textangle=-90,
xref='paper',
yref='paper'
)
],
autosize=True,
margin=dict(
b=100
),
title='Plot Title',
xaxis=dict(
autorange=False,
range=[-0.05674507980728292, -0.0527310420933204],
type='linear'
),
yaxis=dict(
autorange=False,
range=[1.2876210047544652, 1.2977732997811402],
type='linear'
),
height=550,
width=1137
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
# -
# ### Dash Example
# [Dash](https://plotly.com/products/dash/) is an Open Source Python library which can help you convert plotly figures into a reactive, web-based application. Below is a simple example of a dashboard created using Dash. Its [source code](https://github.com/plotly/simple-example-chart-apps/tree/master/dash-text-annotationsplot) can easily be deployed to a PaaS.
from IPython.display import IFrame
IFrame(src= "https://dash-simple-apps.plotly.host/dash-text-annotationsplot/", width="100%", height="750px", frameBorder="0")
from IPython.display import IFrame
IFrame(src= "https://dash-simple-apps.plotly.host/dash-text-annotationsplot/code", width="100%", height=500, frameBorder="0")
# #### Reference
# See https://plotly.com/python/reference/#layout-annotations for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'text-and-annotations.ipynb', 'python/text-and-annotations/', 'Text and Annotations',
'How to add text labels and annotations to plots in python.',
title = 'Text and Annotations | plotly',
thumbnail='thumbnail/text-and-annotations.jpg', language='python',
has_thumbnail='true', display_as='file_settings', order=30,
ipynb='~notebook_demo/204', uses_plotly_offline=False)
| _posts/python-v3/fundamentals/annotations/text-and-annotations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="RGGFAk4ok3W2"
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="wfnC1bgpk43k"
# # Finite ConvNet Training with Conv-KIP distilled images
#
# This notebook demonstrate a simple example of finite neural network transfer using Conv-KIP distilled image.
#
# Training code is based off of [Lee et al., Finite Versus Infinite Neural Networks: an Empirical Study, NeurIPS 2020](https://arxiv.org/abs/2007.15801), as adapted in [Nguyen et al., Dataset Distillation with Infinitely Wide Convolutional Networks](https://arxiv.org/abs/2107.13034).
# + [markdown] id="tpXmXTFgpNKC"
# ## Imports
# + executionInfo={"elapsed": 60, "status": "ok", "timestamp": 1642198368728, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="_jZmTD5hpDkG" outputId="6d30b5f7-be66-482c-f225-a8f1b931e927"
# Install ml_collections and neural_tangents
# !pip install -q git+https://www.github.com/google/ml_collections
# !pip install -q git+https://www.github.com/google/neural-tangents
# + executionInfo={"elapsed": 7693, "status": "ok", "timestamp": 1642198376575, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="HCN_G3bA7WU_" outputId="5f609670-9109-490b-b73a-4e1fa96da309"
from absl import app
from absl import logging
import functools
import time
import operator as op
import jax
from jax.experimental import optimizers
from jax.experimental import stax as ostax
import jax.numpy as jnp
from jax.tree_util import tree_map
from jax.tree_util import tree_reduce
import ml_collections
import neural_tangents as nt
from neural_tangents import stax
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
# + [markdown] id="iaWIPFZvmtEE"
# ## Experiment Configs
# + executionInfo={"elapsed": 54, "status": "ok", "timestamp": 1642198376772, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="uIz0K4QEmwpU"
def get_config():
# Note that max_lr_factor and l2_regularization is found through grid search.
config = ml_collections.ConfigDict()
config.seed = 0
# Dataset
config.dataset = ml_collections.ConfigDict()
config.dataset.name = 'cifar10' # ['cifar10', 'cifar100', 'mnist', 'fashion_mnist']
config.preprocess_type = 'zca_normalize' # ['zca_normalize', 'standard']
config.zca_reg = 0.1
# Optimization
config.optimizer = 'momentum' # Note: In this notebook `run_trainer` optimizer is hard-coded to be `momentum`.
config.momentum = 0.9
config.batch_size = 100
config.eval_batch_size = 100
config.train_steps = 5_000
config.empirical_ntk_num_inputs = 50 # Number of samples to estimate max LR.
config.max_lr_factor = 1.0
config.l2_regularization = 0.001
# Network Architecture
config.width = 1024
config.depth = 3
config.use_gap = False
config.W_std = 2.0**0.5
config.b_std = 0.1
config.activation = 'relu' # ['relu', 'identity', 'gelu'', 'identity']
config.loss_type = 'mse' # ['mse', 'xent']
config.parameterization = 'standard' # ['standard', 'ntk']
config.kip = ml_collections.ConfigDict()
# Put any KIP / Label Solve checkpoint path here to use as training data."
config.kip.data_ckpt_path = (
'gs://kip-datasets/kip/cifar10/ConvNet_ssize100_zca_nol_noaug_ckpt1000.npz')
return config
# + [markdown] id="BwO9GQYkmWfI"
# ## Define Training Utilities
# + executionInfo={"elapsed": 66, "status": "ok", "timestamp": 1642198376973, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="SLBLnZz47uoW"
def load_kip_data(config):
"""Load and preprocess dataset with TFDS and KIP ckpt path."""
data = _load_dataset(config)
data = apply_preprocess(config, data)
logging.info('valid data: %s, %s',
data['valid']['images'].shape, data['valid']['labels'].shape)
logging.info('test data: %s, %s',
data['test']['images'].shape, data['test']['labels'].shape)
# Override training data
ckpt_name = config.kip.data_ckpt_path
with tf.io.gfile.GFile(ckpt_name, 'rb') as f:
loaded_data = jnp.load(f)
train_images = loaded_data['images']
train_labels = loaded_data['labels']
if config.loss_type == 'xent':
# Recover one-hot label even for the learned labels.
n_classes = train_labels.shape[-1]
train_labels = jnp.array(
jnp.argmax(train_labels, axis=-1)[:, None] == jnp.arange(n_classes),
dtype=train_labels.dtype)
# Override training
data['train'] = {'images': train_images, 'labels': train_labels}
logging.info('Overrriding train with ckpt %s, size: (%s, %s)', ckpt_name,
data['train']['images'].shape, data['train']['labels'].shape)
return data
def _load_dataset(config):
"""Get per channel normalized / one hot encoded data from TFDS."""
VALID_SIZE = 5000
dataset_name = config.dataset.name
ds_builder = tfds.builder(dataset_name)
ds_train, ds_test = tfds.as_numpy(
tfds.load(
dataset_name,
split=['train', 'test'],
batch_size=-1,
as_dataset_kwargs={'shuffle_files': False}))
train_images, train_labels, test_images, test_labels = (ds_train['image'],
ds_train['label'],
ds_test['image'],
ds_test['label'])
height, width, num_channels = ds_builder.info.features['image'].shape
num_classes = ds_builder.info.features['label'].num_classes
with config.dataset.unlocked():
config.dataset.height = height
config.dataset.width = width
config.dataset.num_channels = num_channels
config.dataset.num_classes = num_classes
# One hot encode
train_labels = jax.nn.one_hot(train_labels, num_classes)
test_labels = jax.nn.one_hot(test_labels, num_classes)
if config.get('loss_type', 'mse') == 'mse':
shift = (1. / num_classes if num_classes > 1 else 0.5)
train_labels -= shift
test_labels -= shift
# Normalize by precomputed per channel mean/std from training images
train_xs = (train_images - np.mean(train_images, axis=(0, 1, 2))) / np.std(
train_images, axis=(0, 1, 2))
test_xs = (test_images - np.mean(train_images, axis=(0, 1, 2))) / np.std(
train_images, axis=(0, 1, 2))
test_ys = test_labels
train_xs, valid_xs = train_xs[:-VALID_SIZE], train_xs[-VALID_SIZE:]
train_ys, valid_ys = train_labels[:-VALID_SIZE], train_labels[-VALID_SIZE:]
train = (train_xs, train_ys)
valid = (valid_xs, valid_ys)
test = (test_xs, test_ys)
data = {'train': {'images': train_xs, 'labels': train_ys},
'valid': {'images': valid_xs, 'labels': valid_ys},
'test': {'images': test_xs, 'labels': test_ys}
}
return data
# + cellView="form" executionInfo={"elapsed": 82, "status": "ok", "timestamp": 1642198377198, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="eUwrmFWxfuAf"
#@title Preprocess utilties
def apply_preprocess(config, data):
"""Apply ZCA preprocessing on the standard normalized data."""
x_train, y_train = data['train']['images'], data['train']['labels']
x_valid, y_valid = data['valid']['images'], data['valid']['labels']
x_test, y_test = data['test']['images'], data['test']['labels']
preprocess_type = config.get('preprocess_type', 'standard')
if preprocess_type == 'standard':
# Normalization is already done.
pass
else:
zca_reg = config.get('zca_reg', 0.0)
if preprocess_type == 'zca_normalize':
preprocess_op = _get_preprocess_op(
x_train,
layer_norm=True,
zca_reg=zca_reg,
zca_reg_absolute_scale=config.get('zca_reg_absolute_scale', False))
x_train = preprocess_op(x_train)
x_valid = preprocess_op(x_valid)
x_test = preprocess_op(x_test)
else:
NotImplementedError('Preprocess type %s is not implemented' %
preprocess_type)
return {'train': {'images': x_train, 'labels': y_train},
'valid': {'images': x_valid, 'labels': y_valid},
'test': {'images': x_test, 'labels': y_test}}
def _get_preprocess_op(x_train,
layer_norm=True,
zca_reg=1e-5,
zca_reg_absolute_scale=False,
on_cpu=False):
"""ZCA preprocessing function."""
whitening_transform = _get_whitening_transform(x_train, layer_norm, zca_reg,
zca_reg_absolute_scale,
on_cpu)
def _preprocess_op(images):
orig_shape = images.shape
images = images.reshape(orig_shape[0], -1)
if layer_norm:
# Zero mean every feature
images = images - jnp.mean(images, axis=1)[:, jnp.newaxis]
# Normalize
image_norms = jnp.linalg.norm(images, axis=1)
# Make features unit norm
images = images / image_norms[:, jnp.newaxis]
images = (images).dot(whitening_transform)
images = images.reshape(orig_shape)
return images
return _preprocess_op
def _get_whitening_transform(x_train,
layer_norm=True,
zca_reg=1e-5,
zca_reg_absolute_scale=False,
on_cpu=False):
"""Returns 2D matrix that performs whitening transform.
Whitening transform is a (d,d) matrix (d = number of features) which acts on
the right of a (n, d) batch of flattened data.
"""
orig_train_shape = x_train.shape
x_train = x_train.reshape(orig_train_shape[0], -1).astype('float64')
if on_cpu:
x_train = jax.device_put(x_train, jax.devices('cpu')[0])
n_train = x_train.shape[0]
if layer_norm:
logging.info('Performing layer norm preprocessing.')
# Zero mean every feature
x_train = x_train - jnp.mean(x_train, axis=1)[:, jnp.newaxis]
# Normalize
train_norms = jnp.linalg.norm(x_train, axis=1)
# Make features unit norm
x_train = x_train / train_norms[:, jnp.newaxis]
logging.info('Performing zca whitening preprocessing with reg: %.2e', zca_reg)
cov = 1.0 / n_train * x_train.T.dot(x_train)
if zca_reg_absolute_scale:
reg_amount = zca_reg
else:
reg_amount = zca_reg * jnp.trace(cov) / cov.shape[0]
logging.info('Raw zca regularization strength: %f', reg_amount)
u, s, _ = jnp.linalg.svd(cov + reg_amount * jnp.eye(cov.shape[0]))
inv_sqrt_zca_eigs = s**(-1 / 2)
# rank control
if n_train < x_train.shape[1]:
inv_sqrt_zca_eigs = inv_sqrt_zca_eigs.at[n_train:].set(
jnp.ones(inv_sqrt_zca_eigs[n_train:].shape[0]))
whitening_transform = jnp.einsum(
'ij,j,kj->ik', u, inv_sqrt_zca_eigs, u, optimize=True)
return whitening_transform
# + executionInfo={"elapsed": 55, "status": "ok", "timestamp": 1642198377389, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="StI0M1sd9pN0"
# Loss Definition.
cross_entropy = lambda y, y_hat: -np.mean(np.sum(y * y_hat, axis=1))
mse_loss = lambda y, y_hat: 0.5 * jnp.mean((y - y_hat)**2)
_l2_norm = lambda params: tree_map(lambda x: jnp.sum(x ** 2), params)
l2_regularization = lambda params: tree_reduce(op.add, _l2_norm(params))
def cosine_schedule(initial_learning_rate, training_steps):
def _cosine_schedule(t):
return initial_learning_rate * 0.5 * (
1 + jnp.cos(t / training_steps * jnp.pi))
return _cosine_schedule
def _epoch_from_step(step, train_size, batch_size):
if train_size == batch_size:
return step
else:
return float(step / train_size * batch_size)
# + [markdown] id="B2NJelTBpBL_"
# ## Define Networks
# + executionInfo={"elapsed": 53, "status": "ok", "timestamp": 1642198377601, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="5ehZ4P7BpCv2"
def _get_activation_fn(config):
if config.activation.lower() == 'relu':
activation_fn = stax.Relu()
elif config.activation.lower() == 'erf':
activation_fn = stax.Erf()
elif config.activation.lower() == 'identity':
activation_fn = stax.Identity()
elif config.activation.lower() == 'gelu':
activation_fn = stax.Gelu()
else:
raise ValueError('activation function %s not implemented' %
config.activation)
return activation_fn
def _get_norm_layer(normalization):
normalization = normalization.lower()
if 'layer' in normalization:
norm_layer = stax.LayerNorm(axis=(1, 2, 3))
elif 'instance' in normalization:
norm_layer = stax.LayerNorm(axis=(1, 2))
elif normalization == '':
norm_layer = stax.Identity()
else:
raise ValueError('normalization %s not implemented' % normalization)
return norm_layer
def _ConvNet(config):
return ConvNet(
depth=config.depth,
width=config.width,
use_gap=config.get('use_gap', False),
W_std=config.W_std,
b_std=config.b_std,
num_classes=config.dataset.num_classes,
parameterization=config.parameterization,
activation_fn=_get_activation_fn(config),
norm_layer=_get_norm_layer(config.get('normalization', '')),
image_format=config.get('image_format', 'NHWC'))
def ConvNet(
depth: int,
width: int,
prepend_conv_layer: bool = True,
use_gap: bool = False,
W_std=2**0.5,
b_std=0.1,
num_classes: int = 10,
parameterization: str = 'ntk',
activation_fn=stax.Relu(),
norm_layer=stax.Identity(),
image_format: str = 'NHWC'):
"""Adaptation of ConvNet baseline of Dataset Condensation.
Original architecture is based on (Gidaris & Komodakis, 2018)
and here we adapt version of Zhao et al., Dataset Condensation with Gradient
Matching, https://openreview.net/pdf?id=mSAKhLYLSsl
Implements depth-many blocks of convolution, activation, 2x2 avg pooling.
Normalization layer of corresponding finite-width neural network is omitted.
For the 'ConvNet' settings of Nguyen et al., Dataset Distillation with
Infinitely Wide Convolutional Networks, set depth=3 (width is immaterial for
'ntk' parameterization) and prepend_conv_layer=True. For 'ConvNet3' settings
closer to Zhao et al., set prepend_conv_layer=False.
Args:
depth: depth of network
width: width of network
prepend_conv_layer: if True, add an additional conv and relu layer before
main set of blocks
use_gap: if True, use global average pooling for preclassifier layer
W_std: standard deviation of weight matrix initialization
b_std: standard deviation of bias initialization
num_classes: number of classes for output layer
parameterization: 'ntk' or 'standard' for initializing network and NTK
activation_fn: NT activation function of network
norm_layer: NT normalization layer, default is Identity.
image_format: Image format 'NHWC', 'NCHW' etc.
Returns:
Corresponding neural_tangents stax model.
"""
layers = []
conv = functools.partial(
stax.Conv,
W_std=W_std,
b_std=b_std,
padding='SAME',
parameterization=parameterization)
if prepend_conv_layer:
layers += [
conv(width, (3, 3),
dimension_numbers=(image_format, 'HWIO', 'NHWC')),
activation_fn
]
# generate blocks of convolutions followed by average pooling
for _ in range(depth):
layers += [conv(width, (3, 3)), norm_layer,
activation_fn, stax.AvgPool((2, 2), strides=(2, 2))]
if use_gap:
layers.append(stax.GlobalAvgPool())
else:
layers.append(stax.Flatten())
layers.append(stax.Dense(num_classes, W_std, b_std,
parameterization=parameterization))
return stax.serial(*layers)
# + [markdown] id="iEd6CeN5npqo"
# ## Define Trainer
# + executionInfo={"elapsed": 300, "status": "ok", "timestamp": 1642198378063, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="nYEqYzzEkmyB"
def run_trainer(data, config):
"""Train a neural network."""
# Experiment Parameters.
batch_size = config.batch_size
eval_batch_size = config.get('eval_batch_size', config.batch_size)
train_size = data['train']['images'].shape[0]
steps_per_epoch = int(np.ceil(train_size / batch_size))
train_steps = int(config.train_steps)
train_epochs = int(np.ceil(config.train_steps / steps_per_epoch))
l2_lambda = config.l2_regularization
key = jax.random.PRNGKey(config.seed)
# Construct tf.data
train_ds = tf.data.Dataset.from_tensor_slices({
'images': data['train']['images'],
'labels': data['train']['labels'],
}).repeat().shuffle(
data['train']['images'].shape[0], seed=0).batch(batch_size).as_numpy_iterator()
# This is used for computing training metrics.
train_eval_ds = tf.data.Dataset.from_tensor_slices({
'images': data['train']['images'],
'labels': data['train']['labels'],
}).batch(eval_batch_size)
valid_ds = tf.data.Dataset.from_tensor_slices({
'images': data['valid']['images'][:1000], # Smaller validation set size for notebook
'labels': data['valid']['labels'][:1000],
}).batch(eval_batch_size)
test_ds = tf.data.Dataset.from_tensor_slices({
'images': data['test']['images'],
'labels': data['test']['labels'],
}).batch(eval_batch_size)
# Initialize Network.
network = _ConvNet(config)
init_f, f, _ = network
key, split = jax.random.split(key)
_, init_params = init_f(split, (-1,) + data['train']['images'].shape[1:])
# Estimate maximum learning rate
def logit_reduced_f(params, x):
out = f(params, x)
return jnp.sum(out, axis=-1) / out.shape[-1]**(1 / 2)
input_sample = data['valid']['images'][:config.empirical_ntk_num_inputs]
empirical_kernel_fn = lambda x1, x2, params: nt.empirical_ntk_fn(
logit_reduced_f, trace_axes=(), vmap_axes=0, implementation=1)(x1, x2,
params)
empirical_kernel_fn = nt.batch(empirical_kernel_fn, batch_size=10)
logging.info('input_sample shape: %s', input_sample.shape)
max_lr_estimate_start = time.time()
kernel = empirical_kernel_fn(input_sample, None, init_params)
logging.info('kernel shape: %s', kernel.shape)
y_train_size = kernel.shape[0] * config.dataset.num_classes
assert y_train_size == data['valid']['labels'][:config.empirical_ntk_num_inputs].size
max_lr = nt.predict.max_learning_rate(
ntk_train_train=kernel, y_train_size=y_train_size, eps=1e-12)
print('Max LR estimate took: %.2fs'%(time.time() - max_lr_estimate_start))
learning_rate = float(max_lr * config.max_lr_factor)
print('max LR: %f, current LR: %f'%(max_lr, learning_rate))
# Define Raw loss, Accuracy, and Optimizer.
@jax.jit
def raw_loss(params, batch):
"""Loss without weight decay."""
images, labels = batch['images'], batch['labels']
loss_type = config.get('loss_type', 'xent')
if loss_type == 'xent':
return cross_entropy(ostax.logsoftmax(f(params, images)), labels)
elif loss_type == 'mse':
return mse_loss(f(params, images), labels)
else:
raise NotImplementedError('Loss type %s not implemented:' % loss_type)
@jax.jit
def loss(params, batch):
l2_loss = 0.5 * l2_lambda * l2_regularization(params)
return raw_loss(params, batch) + l2_loss
grad_loss = jax.jit(jax.grad(loss))
@jax.jit
def accuracy(params, batch):
images, labels = batch['images'], batch['labels']
return jnp.mean(
jnp.array(
jnp.argmax(f(params, images), axis=1) == jnp.argmax(labels, axis=1),
dtype=np.float32))
learning_rate_fn = cosine_schedule(learning_rate, config.train_steps)
print('Using momentum optimizer.')
opt_init_fn, opt_apply_fn, get_params = optimizers.momentum(
learning_rate_fn, config.momentum)
opt_apply_fn = jax.jit(opt_apply_fn)
state = opt_init_fn(init_params)
del init_params # parameters obatined from optimizer state
# Define Update and Evaluate Function.
@jax.jit
def update(step, state, batch):
"""Training updates."""
params = get_params(state)
new_step = step
dparams = grad_loss(params, batch)
return new_step + 1, opt_apply_fn(step, dparams, state)
def dataset_evaluate(state, dataset):
"""Compute loss and accuracy metrics over entire dataset."""
params = get_params(state)
tot_metrics ={'raw_loss':0., 'loss': 0., 'correct': 0, 'count': 0}
for eval_batch in dataset.as_numpy_iterator():
eval_size = eval_batch['images'].shape[0]
tot_metrics['raw_loss'] += raw_loss(params, eval_batch) * eval_size
tot_metrics['loss'] += loss(params, eval_batch) * eval_size
tot_metrics['correct'] += accuracy(params, eval_batch) * eval_size
tot_metrics['count'] += eval_size
metric ={'raw_loss': tot_metrics['raw_loss'] / tot_metrics['count'],
'loss': tot_metrics['loss'] / tot_metrics['count'],
'accuracy': tot_metrics['correct'] / tot_metrics['count'] }
return metric
measurements = []
# Define logging steps.
log_max_steps = np.log10(train_steps)
log_steps = [0] + sorted(
list(set([int(10**t) for t in np.linspace(0.0, log_max_steps, 10)])))
start_time = time.time()
global_step = 0
step_time = 0
hparams_json = config.to_json_best_effort(indent=2)
print('hparams: %s', hparams_json)
print('Total training steps %d, steps_per_epoch %d' %
(train_steps, steps_per_epoch))
print('Step (Epoch)\tLearning Rate\tTrain Loss\tValid Loss\t'
'Train Acc\tValid Acc\tTime Elapsed\tEval Time')
while global_step <= train_steps:
i = int(global_step)
epoch = _epoch_from_step(i, train_size, batch_size)
if i in log_steps or i % 250 == 0 or i == train_steps:
eval_start_time = time.time()
train_metric = dataset_evaluate(state, train_eval_ds)
if not jnp.isfinite(train_metric['raw_loss']):
msg = 'NaN during Training! Terminating current trial.'
raise ValueError(msg)
valid_metric = dataset_evaluate(state, valid_ds)
eval_time = time.time() - eval_start_time
elapsed_time = float(time.time() - start_time)
lr = float(learning_rate_fn(i))
measurements.append([
i, epoch, lr,
train_metric['loss'],
valid_metric['loss'],
train_metric['accuracy'],
valid_metric['accuracy'], elapsed_time
])
print(
('{:06d}\t({:06.1f})\t' + ('{:.6e}\t' * 3) + ('{:.6f}\t' * 4)).format(
i, epoch, lr, train_metric['loss'], valid_metric['loss'],
train_metric['accuracy'], valid_metric['accuracy'],
elapsed_time, eval_time))
global_step, state = update(global_step, state, next(train_ds))
print('Training finished')
test_metric = dataset_evaluate(state, test_ds)
print('Step\tEpoch\tLearning Rate\tTrain Loss\tValid Loss\tTest Loss\t'
'Train Acc\tValid Acc\tTest Acc\tTime Elapsed')
print(('{:06d}\t({:06.1f})\t' + ('{:.6e}\t' * 4) + ('{:.6f}\t' * 4)).format(
i, epoch, lr, train_metric['loss'], valid_metric['loss'],
test_metric['loss'], train_metric['accuracy'],
valid_metric['accuracy'], test_metric['accuracy'],
time.time() - start_time))
return measurements
# + [markdown] id="pIPKnsUixAgs"
# ## Run an Experiment with Trainer
# + executionInfo={"elapsed": 2038363, "status": "ok", "timestamp": 1642200416564, "user": {"displayName": "", "photoUrl": "", "userId": ""}, "user_tz": 480} id="IMcqOycCo-Vu" outputId="27fd0ec7-ba4c-445e-aab2-aa6b5a8b084b"
tf.config.experimental.set_visible_devices([], 'GPU')
config = get_config()
data = load_kip_data(config)
measurements = run_trainer(data, config)
| kip/nn_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# 
# # Automated Machine Learning
# _**Classification with Deployment using a Bank Marketing Dataset**_
#
# ## Contents
# 1. [Introduction](#Introduction)
# 1. [Setup](#Setup)
# 1. [Train](#Train)
# 1. [Results](#Results)
# 1. [Deploy](#Deploy)
# 1. [Test](#Test)
# 1. [Acknowledgements](#Acknowledgements)
# ## Introduction
#
# In this example we use the UCI Bank Marketing dataset to showcase how you can use AutoML for a classification problem and deploy it to an Azure Container Instance (ACI). The classification goal is to predict if the client will subscribe to a term deposit with the bank.
#
# If you are using an Azure Machine Learning Compute Instance, you are all set. Otherwise, go through the [configuration](../../../configuration.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
#
# Please find the ONNX related documentations [here](https://github.com/onnx/onnx).
#
# In this notebook you will learn how to:
# 1. Create an experiment using an existing workspace.
# 2. Configure AutoML using `AutoMLConfig`.
# 3. Train the model using local compute with ONNX compatible config on.
# 4. Explore the results, featurization transparency options and save the ONNX model
# 5. Inference with the ONNX model.
# 6. Register the model.
# 7. Create a container image.
# 8. Create an Azure Container Instance (ACI) service.
# 9. Test the ACI service.
#
# In addition this notebook showcases the following features
# - **Blocking** certain pipelines
# - Specifying **target metrics** to indicate stopping criteria
# - Handling **missing data** in the input
# ## Setup
#
# As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
# +
import logging
from matplotlib import pyplot as plt
import pandas as pd
import os
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.core.dataset import Dataset
from azureml.train.automl import AutoMLConfig
from azureml.interpret import ExplanationClient
# -
# This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
print("This notebook was created using version 1.33.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
# Accessing the Azure ML workspace requires authentication with Azure.
#
# The default authentication is interactive authentication using the default tenant. Executing the `ws = Workspace.from_config()` line in the cell below will prompt for authentication the first time that it is run.
#
# If you have multiple Azure tenants, you can specify the tenant by replacing the `ws = Workspace.from_config()` line in the cell below with the following:
#
# ```
# from azureml.core.authentication import InteractiveLoginAuthentication
# auth = InteractiveLoginAuthentication(tenant_id = 'mytenantid')
# ws = Workspace.from_config(auth = auth)
# ```
#
# If you need to run in an environment where interactive login is not possible, you can use Service Principal authentication by replacing the `ws = Workspace.from_config()` line in the cell below with the following:
#
# ```
# from azureml.core.authentication import ServicePrincipalAuthentication
# auth = auth = ServicePrincipalAuthentication('mytenantid', 'myappid', '<PASSWORD>')
# ws = Workspace.from_config(auth = auth)
# ```
# For more details, see [aka.ms/aml-notebook-auth](http://aka.ms/aml-notebook-auth)
# +
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = 'automl-classification-bmarketing-all'
experiment=Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# -
# ## Create or Attach existing AmlCompute
# You will need to create a compute target for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.
#
# > Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
#
# #### Creation of AmlCompute takes approximately 5 minutes.
# If the AmlCompute with that name is already in your workspace this code will skip the creation process.
# As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
# +
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "cpu-cluster-4"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS12_V2',
max_nodes=6)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
# -
# # Data
# ### Load Data
#
# Leverage azure compute to load the bank marketing dataset as a Tabular Dataset into the dataset variable.
# ### Training Data
data = pd.read_csv("https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/bankmarketing_train.csv")
data.head()
# +
# Add missing values in 75% of the lines.
import numpy as np
missing_rate = 0.75
n_missing_samples = int(np.floor(data.shape[0] * missing_rate))
missing_samples = np.hstack((np.zeros(data.shape[0] - n_missing_samples, dtype=np.bool), np.ones(n_missing_samples, dtype=np.bool)))
rng = np.random.RandomState(0)
rng.shuffle(missing_samples)
missing_features = rng.randint(0, data.shape[1], n_missing_samples)
data.values[np.where(missing_samples)[0], missing_features] = np.nan
# +
if not os.path.isdir('data'):
os.mkdir('data')
# Save the train data to a csv to be uploaded to the datastore
pd.DataFrame(data).to_csv("data/train_data.csv", index=False)
ds = ws.get_default_datastore()
ds.upload(src_dir='./data', target_path='bankmarketing', overwrite=True, show_progress=True)
# Upload the training data as a tabular dataset for access during training on remote compute
train_data = Dataset.Tabular.from_delimited_files(path=ds.path('bankmarketing/train_data.csv'))
label = "y"
# -
# ### Validation Data
validation_data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/bankmarketing_validate.csv"
validation_dataset = Dataset.Tabular.from_delimited_files(validation_data)
# ### Test Data
test_data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/bankmarketing_test.csv"
test_dataset = Dataset.Tabular.from_delimited_files(test_data)
# ## Train
#
# Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
#
# |Property|Description|
# |-|-|
# |**task**|classification or regression or forecasting|
# |**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
# |**iteration_timeout_minutes**|Time limit in minutes for each iteration.|
# |**blocked_models** | *List* of *strings* indicating machine learning algorithms for AutoML to avoid in this run. <br><br> Allowed values for **Classification**<br><i>LogisticRegression</i><br><i>SGD</i><br><i>MultinomialNaiveBayes</i><br><i>BernoulliNaiveBayes</i><br><i>SVM</i><br><i>LinearSVM</i><br><i>KNN</i><br><i>DecisionTree</i><br><i>RandomForest</i><br><i>ExtremeRandomTrees</i><br><i>LightGBM</i><br><i>GradientBoosting</i><br><i>TensorFlowDNN</i><br><i>TensorFlowLinearClassifier</i><br><br>Allowed values for **Regression**<br><i>ElasticNet</i><br><i>GradientBoosting</i><br><i>DecisionTree</i><br><i>KNN</i><br><i>LassoLars</i><br><i>SGD</i><br><i>RandomForest</i><br><i>ExtremeRandomTrees</i><br><i>LightGBM</i><br><i>TensorFlowLinearRegressor</i><br><i>TensorFlowDNN</i><br><br>Allowed values for **Forecasting**<br><i>ElasticNet</i><br><i>GradientBoosting</i><br><i>DecisionTree</i><br><i>KNN</i><br><i>LassoLars</i><br><i>SGD</i><br><i>RandomForest</i><br><i>ExtremeRandomTrees</i><br><i>LightGBM</i><br><i>TensorFlowLinearRegressor</i><br><i>TensorFlowDNN</i><br><i>Arima</i><br><i>Prophet</i>|
# |**allowed_models** | *List* of *strings* indicating machine learning algorithms for AutoML to use in this run. Same values listed above for **blocked_models** allowed for **allowed_models**.|
# |**experiment_exit_score**| Value indicating the target for *primary_metric*. <br>Once the target is surpassed the run terminates.|
# |**experiment_timeout_hours**| Maximum amount of time in hours that all iterations combined can take before the experiment terminates.|
# |**enable_early_stopping**| Flag to enble early termination if the score is not improving in the short term.|
# |**featurization**| 'auto' / 'off' Indicator for whether featurization step should be done automatically or not. Note: If the input data is sparse, featurization cannot be turned on.|
# |**n_cross_validations**|Number of cross validation splits.|
# |**training_data**|Input dataset, containing both features and label column.|
# |**label_column_name**|The name of the label column.|
#
# **_You can find more information about primary metrics_** [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#primary-metric)
# +
automl_settings = {
"experiment_timeout_hours" : 0.3,
"enable_early_stopping" : True,
"iteration_timeout_minutes": 5,
"max_concurrent_iterations": 4,
"max_cores_per_iteration": -1,
#"n_cross_validations": 2,
"primary_metric": 'AUC_weighted',
"featurization": 'auto',
"verbosity": logging.INFO,
}
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
compute_target=compute_target,
experiment_exit_score = 0.9984,
blocked_models = ['KNN','LinearSVM'],
enable_onnx_compatible_models=True,
training_data = train_data,
label_column_name = label,
validation_data = validation_dataset,
**automl_settings
)
# -
# Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
remote_run = experiment.submit(automl_config, show_output = False)
# Run the following cell to access previous runs. Uncomment the cell below and update the run_id.
# +
#from azureml.train.automl.run import AutoMLRun
#remote_run = AutoMLRun(experiment=experiment, run_id='<run_ID_goes_here')
#remote_run
# -
# Wait for the remote run to complete
remote_run.wait_for_completion()
best_run_customized, fitted_model_customized = remote_run.get_output()
# ## Transparency
#
# View updated featurization summary
custom_featurizer = fitted_model_customized.named_steps['datatransformer']
df = custom_featurizer.get_featurization_summary()
pd.DataFrame(data=df)
# Set `is_user_friendly=False` to get a more detailed summary for the transforms being applied.
df = custom_featurizer.get_featurization_summary(is_user_friendly=False)
pd.DataFrame(data=df)
df = custom_featurizer.get_stats_feature_type_summary()
pd.DataFrame(data=df)
# ## Results
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
# ### Retrieve the Best Model's explanation
# Retrieve the explanation from the best_run which includes explanations for engineered features and raw features. Make sure that the run for generating explanations for the best model is completed.
# +
# Wait for the best model explanation run to complete
from azureml.core.run import Run
model_explainability_run_id = remote_run.id + "_" + "ModelExplain"
print(model_explainability_run_id)
model_explainability_run = Run(experiment=experiment, run_id=model_explainability_run_id)
model_explainability_run.wait_for_completion()
# Get the best run object
best_run, fitted_model = remote_run.get_output()
# -
# #### Download engineered feature importance from artifact store
# You can use ExplanationClient to download the engineered feature explanations from the artifact store of the best_run.
client = ExplanationClient.from_run(best_run)
engineered_explanations = client.download_model_explanation(raw=False)
exp_data = engineered_explanations.get_feature_importance_dict()
exp_data
# #### Download raw feature importance from artifact store
# You can use ExplanationClient to download the raw feature explanations from the artifact store of the best_run.
client = ExplanationClient.from_run(best_run)
engineered_explanations = client.download_model_explanation(raw=True)
exp_data = engineered_explanations.get_feature_importance_dict()
exp_data
# ### Retrieve the Best ONNX Model
#
# Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
#
# Set the parameter return_onnx_model=True to retrieve the best ONNX model, instead of the Python model.
best_run, onnx_mdl = remote_run.get_output(return_onnx_model=True)
# ### Save the best ONNX model
from azureml.automl.runtime.onnx_convert import OnnxConverter
onnx_fl_path = "./best_model.onnx"
OnnxConverter.save_onnx_model(onnx_mdl, onnx_fl_path)
# ### Predict with the ONNX model, using onnxruntime package
# +
import sys
import json
from azureml.automl.core.onnx_convert import OnnxConvertConstants
from azureml.train.automl import constants
from azureml.automl.runtime.onnx_convert import OnnxInferenceHelper
def get_onnx_res(run):
res_path = 'onnx_resource.json'
run.download_file(name=constants.MODEL_RESOURCE_PATH_ONNX, output_file_path=res_path)
with open(res_path) as f:
result = json.load(f)
return result
if sys.version_info < OnnxConvertConstants.OnnxIncompatiblePythonVersion:
test_df = test_dataset.to_pandas_dataframe()
mdl_bytes = onnx_mdl.SerializeToString()
onnx_result = get_onnx_res(best_run)
onnxrt_helper = OnnxInferenceHelper(mdl_bytes, onnx_result)
pred_onnx, pred_prob_onnx = onnxrt_helper.predict(test_df)
print(pred_onnx)
print(pred_prob_onnx)
else:
print('Please use Python version 3.6 or 3.7 to run the inference helper.')
# -
# ## Deploy
#
# ### Retrieve the Best Model
#
# Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
# #### Widget for Monitoring Runs
#
# The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
#
# **Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details
best_run, fitted_model = remote_run.get_output()
# +
model_name = best_run.properties['model_name']
script_file_name = 'inference/score.py'
best_run.download_file('outputs/scoring_file_v_1_0_0.py', 'inference/score.py')
# -
# ### Register the Fitted Model for Deployment
# If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered.
# +
description = 'AutoML Model trained on bank marketing data to predict if a client will subscribe to a term deposit'
tags = None
model = remote_run.register_model(model_name = model_name, description = description, tags = tags)
print(remote_run.model_id) # This will be written to the script file later in the notebook.
# -
# ### Deploy the model as a Web Service on Azure Container Instance
# +
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AciWebservice
from azureml.core.model import Model
inference_config = InferenceConfig(entry_script=script_file_name)
aciconfig = AciWebservice.deploy_configuration(cpu_cores = 2,
memory_gb = 2,
tags = {'area': "bmData", 'type': "automl_classification"},
description = 'sample service for Automl Classification')
aci_service_name = 'automl-sample-bankmarketing-all'
print(aci_service_name)
aci_service = Model.deploy(ws, aci_service_name, [model], inference_config, aciconfig)
aci_service.wait_for_deployment(True)
print(aci_service.state)
# -
# ### Get Logs from a Deployed Web Service
#
# Gets logs from a deployed web service.
# +
#aci_service.get_logs()
# -
# ## Test
#
# Now that the model is trained, run the test data through the trained model to get the predicted values. This calls the ACI web service to do the prediction.
#
# Note that the JSON passed to the ACI web service is an array of rows of data. Each row should either be an array of values in the same order that was used for training or a dictionary where the keys are the same as the column names used for training. The example below uses dictionary rows.
# Load the bank marketing datasets.
from numpy import array
X_test = test_dataset.drop_columns(columns=['y'])
y_test = test_dataset.keep_columns(columns=['y'], validate=True)
test_dataset.take(5).to_pandas_dataframe()
X_test = X_test.to_pandas_dataframe()
y_test = y_test.to_pandas_dataframe()
# +
import requests
X_test_json = X_test.to_json(orient='records')
data = "{\"data\": " + X_test_json +"}"
headers = {'Content-Type': 'application/json'}
resp = requests.post(aci_service.scoring_uri, data, headers=headers)
y_pred = json.loads(json.loads(resp.text))['result']
# -
actual = array(y_test)
actual = actual[:,0]
print(len(y_pred), " ", len(actual))
# ### Calculate metrics for the prediction
#
# Now visualize the data as a confusion matrix that compared the predicted values against the actual values.
#
# +
# %matplotlib notebook
from sklearn.metrics import confusion_matrix
import itertools
cf =confusion_matrix(actual,y_pred)
plt.imshow(cf,cmap=plt.cm.Blues,interpolation='nearest')
plt.colorbar()
plt.title('Confusion Matrix')
plt.xlabel('Predicted')
plt.ylabel('Actual')
class_labels = ['no','yes']
tick_marks = np.arange(len(class_labels))
plt.xticks(tick_marks,class_labels)
plt.yticks([-0.5,0,1,1.5],['','no','yes',''])
# plotting text value inside cells
thresh = cf.max() / 2.
for i,j in itertools.product(range(cf.shape[0]),range(cf.shape[1])):
plt.text(j,i,format(cf[i,j],'d'),horizontalalignment='center',color='white' if cf[i,j] >thresh else 'black')
plt.show()
# -
# ### Delete a Web Service
#
# Deletes the specified web service.
aci_service.delete()
# ## Acknowledgements
# This Bank Marketing dataset is made available under the Creative Commons (CCO: Public Domain) License: https://creativecommons.org/publicdomain/zero/1.0/. Any rights in individual contents of the database are licensed under the Database Contents License: https://creativecommons.org/publicdomain/zero/1.0/ and is available at: https://www.kaggle.com/janiobachmann/bank-marketing-dataset .
#
# _**Acknowledgements**_
# This data set is originally available within the UCI Machine Learning Database: https://archive.ics.uci.edu/ml/datasets/bank+marketing
#
# [Moro et al., 2014] <NAME>, <NAME> and <NAME>. A Data-Driven Approach to Predict the Success of Bank Telemarketing. Decision Support Systems, Elsevier, 62:22-31, June 2014
| how-to-use-azureml/automated-machine-learning/classification-bank-marketing-all-features/auto-ml-classification-bank-marketing-all-features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''r-py-test'': conda)'
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from notebook_utils.OpenKbcMSToolkit import ExtractionToolkit as exttoolkit
deg_path = "resultFiles/DEG_RRvsCIS_by_Jun/"
expr_path = "../data/counts_normalized/rawFiles/"
deg_df = pd.read_csv(deg_path+"CD4_DEG.result",sep=' ', index_col=0).dropna()
sig_df = deg_df.loc[(deg_df['pvalue']<0.05)]
sig_df = sig_df.loc[(sig_df['log2FoldChange'] > 0.58) | (sig_df['log2FoldChange'] < -0.58)]
expr_df = pd.read_csv(expr_path+"counts_norm_CD4.csv", index_col=0)
expr_df.loc[sig_df.index.tolist()]
expr_df.columns = [x.split('.')[0] for x in expr_df.columns.tolist()]
expr_df = expr_df.applymap(lambda x : np.log2(x+1))
expr_df = expr_df.subtract(expr_df.median(axis=1), axis=0)
meta_data = pd.read_csv('../data/annotation_metadata/EPIC_HCvB_metadata_baseline_updated-share.csv')
# +
sample_list, sample_category = exttoolkit.get_sample_name_by_category(dataframe=meta_data, sampleColumn='HCVB_ID', dataColname='DiseaseCourse')
sample_list[0] = list(set(expr_df.columns.tolist()).intersection(set(sample_list[0])))
sample_list[4] = list(set(expr_df.columns.tolist()).intersection(set(sample_list[4])))
ext_samples = sample_list[0] + sample_list[4] # RR + CIS
ext_category = [0]*len(sample_list[0])+[1]*len(sample_list[4])
expr_df = expr_df[ext_samples].loc[sig_df.index]
expr_df = expr_df.replace(0, np.nan).dropna(thresh=len(expr_df.columns)-2).replace(np.nan, 0)
# -
len(ext_samples)
X = expr_df.T.values
y = ext_category
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn import metrics
auc_arr = []
val_auc = []
for t in list(range(0,100)):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=t)
X_test, X_val, y_test, y_val = train_test_split(X_test, y_test, test_size=0.5, random_state=t)
#randomState = list(range(0,5))
clf = SVC(kernel="linear")
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
fpr, tpr, thresholds = metrics.roc_curve(y_test, y_pred, pos_label=1)
auc_arr.append([t, metrics.auc(fpr, tpr)])
y_val_pred = clf.predict(X_val)
fpr, tpr, thresholds = metrics.roc_curve(y_val, y_val_pred, pos_label=1)
val_auc.append([t, metrics.auc(fpr, tpr)])
auc_test_df = pd.DataFrame(data=auc_arr, columns=['state', 'auc']).set_index('state')
auc_val_df = pd.DataFrame(data=val_auc, columns=['state', 'auc']).set_index('state')
# -
auc_df = pd.concat([auc_test_df, auc_val_df], axis=1)
auc_df.columns = ['test_auc', 'val_auc']
auc_df['diff'] = auc_df['test_auc'] - auc_df['val_auc']
auc_df
import seaborn as sns
sns.distplot(auc_test_df['auc'].values.tolist())
sns.distplot(auc_val_df['auc'].values.tolist())
sns.distplot(auc_df['diff'].values.tolist())
| notebook/notebook_archive/Jun10192021/feature_test_with_DEG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# from urllib.request import urlopen
# from bs4 import BeautifulSoup
# html = urlopen('http://www.pythonscraping.com/pages/warandpeace.html')
# bs = BeautifulSoup(html, 'html.parser')
# print(bs)
# -
from urllib.request import urlopen
from bs4 import BeautifulSoup
html = urlopen('http://www.pythonscraping.com/pages/warandpeace.html')
bs = BeautifulSoup(html, "html.parser")
nameList = bs.findAll('span', {'class': 'green'})
for name in nameList:
print(name.get_text())
titles = bs.find_all(['h1', 'h2','h3','h4','h5','h6'])
print([title for title in titles])
allText = bs.find_all('span', {'class':{'green', 'red'}})
print([text for text in allText])
nameList = bs.find_all(text='the prince')
print(len(nameList))
title = bs.find_all(id='title', class_='text')
print([text for text in allText])
# +
from urllib.request import urlopen
from bs4 import BeautifulSoup
html = urlopen('http://www.pythonscraping.com/pages/page3.html')
bs = BeautifulSoup(html, 'html.parser')
for child in bs.find('table',{'id':'giftList'}).children:
print(child)
# +
from urllib.request import urlopen
from bs4 import BeautifulSoup
html = urlopen('http://www.pythonscraping.com/pages/page3.html')
bs = BeautifulSoup(html, 'html.parser')
for sibling in bs.find('table', {'id':'giftList'}).tr.next_siblings:
print(sibling)
# +
from urllib.request import urlopen
from bs4 import BeautifulSoup
html = urlopen('http://www.pythonscraping.com/pages/page3.html')
bs = BeautifulSoup(html, 'html.parser')
print(bs.find('img',
{'src':'../img/gifts/img1.jpg'})
.parent.previous_sibling.get_text())
# +
from urllib.request import urlopen
from bs4 import BeautifulSoup
import re
html = urlopen('http://www.pythonscraping.com/pages/page3.html')
bs = BeautifulSoup(html, 'html.parser')
images = bs.find_all('img', {'src':re.compile('\.\.\/img\/gifts/img.*\.jpg')})
for image in images:
print(image['src'])
# -
bs.find_all(lambda tag: len(tag.attrs) == 2)
bs.find_all(lambda tag: tag.get_text() == 'Or maybe he\'s only resting?')
bs.find_all('', text='Or maybe he\'s only resting?')
| Web_Scraping/Chapter02-AdvancedHTMLParsing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Name
# Data processing by creating a cluster in Cloud Dataproc
#
#
# # Label
# Cloud Dataproc, cluster, GCP, Cloud Storage, KubeFlow, Pipeline
#
#
# # Summary
# A Kubeflow Pipeline component to create a cluster in Cloud Dataproc.
#
# # Details
# ## Intended use
#
# Use this component at the start of a Kubeflow Pipeline to create a temporary Cloud Dataproc cluster to run Cloud Dataproc jobs as steps in the pipeline.
#
# ## Runtime arguments
#
# | Argument | Description | Optional | Data type | Accepted values | Default |
# |----------|-------------|----------|-----------|-----------------|---------|
# | project_id | The Google Cloud Platform (GCP) project ID that the cluster belongs to. | No | GCPProjectID | | |
# | region | The Cloud Dataproc region to create the cluster in. | No | GCPRegion | | |
# | name | The name of the cluster. Cluster names within a project must be unique. You can reuse the names of deleted clusters. | Yes | String | | None |
# | name_prefix | The prefix of the cluster name. | Yes | String | | None |
# | initialization_actions | A list of Cloud Storage URIs identifying executables to execute on each node after the configuration is completed. By default, executables are run on the master and all the worker nodes. | Yes | List | | None |
# | config_bucket | The Cloud Storage bucket to use to stage the job dependencies, the configuration files, and the job driver console’s output. | Yes | GCSPath | | None |
# | image_version | The version of the software inside the cluster. | Yes | String | | None |
# | cluster | The full [cluster configuration](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.clusters#Cluster). | Yes | Dict | | None |
# | wait_interval | The number of seconds to pause before polling the operation. | Yes | Integer | | 30 |
#
# ## Output
# Name | Description | Type
# :--- | :---------- | :---
# cluster_name | The name of the cluster. | String
#
# Note: You can recycle the cluster by using the [Dataproc delete cluster component](https://github.com/kubeflow/pipelines/tree/master/components/gcp/dataproc/delete_cluster).
#
#
# ## Cautions & requirements
#
# To use the component, you must:
# * Set up the GCP project by following these [steps](https://cloud.google.com/dataproc/docs/guides/setup-project).
# * The component can authenticate to GCP. Refer to [Authenticating Pipelines to GCP](https://www.kubeflow.org/docs/gke/authentication-pipelines/) for details.
# * Grant the following types of access to the Kubeflow user service account:
# * Read access to the Cloud Storage buckets which contains initialization action files.
# * The role, `roles/dataproc.editor` on the project.
#
# ## Detailed description
#
# This component creates a new Dataproc cluster by using the [Dataproc create cluster REST API](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.clusters/create).
#
# Follow these steps to use the component in a pipeline:
#
# 1. Install the Kubeflow Pipeline SDK:
#
# +
# %%capture --no-stderr
# !pip3 install kfp --upgrade
# -
# 2. Load the component using KFP SDK
# +
import kfp.components as comp
dataproc_create_cluster_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/pipelines/1.7.0-alpha.2/components/gcp/dataproc/create_cluster/component.yaml')
help(dataproc_create_cluster_op)
# -
# ### Sample
# Note: The following sample code works in an IPython notebook or directly in Python code. See the sample code below to learn how to execute the template.
#
# #### Set sample parameters
# + tags=["parameters"]
# Required Parameters
PROJECT_ID = '<Please put your project ID here>'
# Optional Parameters
EXPERIMENT_NAME = 'Dataproc - Create Cluster'
# -
# #### Example pipeline that uses the component
import kfp.dsl as dsl
import json
@dsl.pipeline(
name='Dataproc create cluster pipeline',
description='Dataproc create cluster pipeline'
)
def dataproc_create_cluster_pipeline(
project_id = PROJECT_ID,
region = 'us-central1',
name='',
name_prefix='',
initialization_actions='',
config_bucket='',
image_version='',
cluster='',
wait_interval='30'
):
dataproc_create_cluster_op(
project_id=project_id,
region=region,
name=name,
name_prefix=name_prefix,
initialization_actions=initialization_actions,
config_bucket=config_bucket,
image_version=image_version,
cluster=cluster,
wait_interval=wait_interval)
# #### Compile the pipeline
pipeline_func = dataproc_create_cluster_pipeline
pipeline_filename = pipeline_func.__name__ + '.zip'
import kfp.compiler as compiler
compiler.Compiler().compile(pipeline_func, pipeline_filename)
# #### Submit the pipeline for execution
# +
#Specify pipeline argument values
arguments = {}
#Get or create an experiment and submit a pipeline run
import kfp
client = kfp.Client()
experiment = client.create_experiment(EXPERIMENT_NAME)
#Submit a pipeline run
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id, run_name, pipeline_filename, arguments)
# -
# ## References
# * [Kubernetes Engine for Kubeflow](https://www.kubeflow.org/docs/started/getting-started-gke/#gcp-service-accounts)
# * [Component Python code](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/component_sdk/python/kfp_component/google/dataproc/_create_cluster.py)
# * [Component Docker file](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/Dockerfile)
# * [Sample notebook](https://github.com/kubeflow/pipelines/blob/master/components/gcp/dataproc/create_cluster/sample.ipynb)
# * [Dataproc create cluster REST API](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.clusters/create)
#
# ## License
# By deploying or using this software you agree to comply with the [AI Hub Terms of Service](https://aihub.cloud.google.com/u/0/aihub-tos) and the [Google APIs Terms of Service](https://developers.google.com/terms/). To the extent of a direct conflict of terms, the AI Hub Terms of Service will control.
| components/gcp/dataproc/create_cluster/sample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
filename = 'query_all_org_class_results2.xlsx'
outfile = 'output_class_to_orgs_20190501_v10.xlsx'
filepath = '/Volumes/backup_128G/z_repository/TBIO_data/RequestsFromTana/all_org_classes'
read_file_from = '{0}/{1}'.format(filepath, filename)
write_file_to = '{0}/{1}'.format(filepath, outfile)
# -
df = pd.read_excel(read_file_from)
df = df.fillna(0)
df.head()
print(df.count())
count = max(df.count())
print('max count = {0}'.format(count))
# +
count = df.groupby(['orgName']).count()['graph']
count = count.sort_values(ascending=False)
print(count)
maxCount = max(count)
# +
results = []
for idx in range(0, len(df)):
row = df.loc[idx]
# print(row)
graph = row['graph']
orgName = row['orgName']
className = row['class']
comments = ''
for cidx in range(3, len(row)):
if row[cidx] != 0:
comments += '{0};'.format(row[cidx])
result = {'graph': graph, 'org': orgName, 'class': className, 'comments': comments}
if result not in results:
results.append(result)
# print(graph, orgName, className, comments)
print(len(results))
print(results[0:5])
# +
# classResults = {}
# for res in results:
# className = res['class']
# graph = res['graph']
# orgName = res['org']
# comments = res['comments']
# if className not in classResults:
# classResults[className] = {'orgGra': {}, 'comments': ''}
# orgGraph = classResults[className]['orgGra'].copy()
# if orgName not in orgGraph:
# orgGraph[orgName] = []
# if graph not in orgGraph[orgName]:
# orgGraph[orgName].append(graph)
# classResults[className] = {'orgGra': orgGraph, 'comments': comments}
# print(len(results), len(classResults))
# -
def mapFunc(orgName, defaultVal, mapTable):
res = defaultVal;
for tKey in mapTable:
for tVale in mapTable[tKey]:
if tVale in orgName:
return tKey
return res
# ## Ownership mapping table
# Ownership mapping table
OwnershipMapping = {
'PrivateOrganization': ['PrivateOrganization'],
'StateOrganization': ['StateOrganization'],
'PublicOrganization': ['PublicOrganization']
}
UnknownOwnership = '';
# ## ModeOfEntry mapping table
# ModeOfEntry mapping table
ModeOfEntryMapping = {
'Appointed': ['總督府','街','庄','州','公學校','利組合','組合'],
'Elected': ['協議會']
}
UnknownModeOfEntry = '';
# ## Period mapping table
# +
## Period mapping table
# 總督府,街,庄,信託會社,州,公學校,株式會,株式会社,利組合,組合,國語講習所
PeriodMapping = {
'JapaneseColonialOrganization': ['總督府','街','庄','信託會社','州','公學校',
'株式會','株式会社','利組合','組合','國語講習所'],
'EarlyPostWarOrganization': ['縣議會','台灣省','台灣長官公署'],
'RepublicanOrganization': [],
}
UnknownPeriod = '';
## yjc 為 JapaneseColonialOrganization
# -
# ## LevelOfOperation mapping table
## LevelOfOperation mapping table
LevelOfOperationMapping = {
'CentralOrganization': [],
'LocalOrganization': ['庄','街','區','州','市','縣'],
'ProvincialOrganization': ['省','廳']
}
UnknownLevelOfOperation = '';
# ## Function mapping table
# +
## Function mapping table
func_filename = 'query_org_divisionContent_results.csv'
func_file_from = '{0}/{1}'.format(filepath, func_filename)
funcDf = pd.read_csv(func_file_from)
funcDf.head()
def mapOrgToFunction(orgName, defaultVal):
# print(orgName)
row = funcDf.loc[funcDf['sName'] == orgName]
if row.empty:
division = [defaultVal]
else:
division = row['orgName'].tolist()
return division
# UnknownFunction = 'UnknwownFunction';
# print(mapOrgToFunction('壯圍庄古亭笨第十五保第七保保甲', UnknownFunction))
# print(mapOrgToFunction('高雄州農會', UnknownFunction))
# print(mapOrgToFunction('在魚池庄經營製茶業', UnknownFunction))
# print(mapOrgToFunction('xxxxx', UnknownFunction))
# for idx in range(0, len(funcDf.index)):
# # for idx in range(16,23):
# # print(funcDf.loc[idx, 'sName'])
# print(mapOrgToFunction(funcDf.loc[idx, 'sName'], UnknownFunction))
# +
# Function mapping table
FunctionMapping = {
'ArmedForces': ['ArmedForces'],
'PrivateBusiness': ['AgriculturalOrganization', 'Industry', 'Business', 'Trade_FinancialOrganization'],
'PublicAdministration': ['SocialAffairs', 'AdministrativeOrganization', 'CommunicationPost',
'Hygiene_Sanitation', 'Canals_Irrigation', 'Transportation',
'JudicialOrganization'],
'Academia_Education': ['EducationalOrganization', 'Research_Investigation'],
'Hospital': ['HealthOrganization'],
'VoluntaryAssociation': ['ReligiousOrganization'],
'StateBusiness': ['MonopolyOrganization'],
'MassMedia': ['Media'],
'NeedToBeSolved': ['EntertainmentOrganization']
}
UnknownFunction = '';
# -
# ## Main generation
print(len(results))
# +
# orgList = []
# # [{'graph': 'chrj_pr', 'org': 'Organization', 'class': '大興紡織', 'comments': ''},
# for res in results:
# className = str(res['class'])
# graph = res['graph']
# orgName = res['org']
# orgArr = []
# # graph
# orgArr.append(graph)
# # Organization
# orgArr.append(className)
# # Ownership
# # orgArr.append(mapFunc(orgName, UnknownOwnership, OwnershipMapping))
# # ModeOfEntry
# # orgArr.append(mapFunc(className, UnknownModeOfEntry, ModeOfEntryMapping))
# # Period
# periodTitle = mapFunc(className, UnknownPeriod, PeriodMapping)
# if periodTitle == UnknownPeriod and graph == 'yjc':
# periodTitle = 'JapaneseColonialOrganization'
# orgArr.append(periodTitle)
# # LevelOfOperation
# # orgArr.append(mapFunc(className, UnknownLevelOfOperation, LevelOfOperationMapping))
# # Function
# funcArray = mapOrgToFunction(className, UnknownFunction)
# # orgArr.append(mapOrgToFunction(strClassName, UnknownFunction))
# # comments
# # comments = res['comments']
# for funcName in funcArray:
# finalOrg = []
# finalOrg = orgArr.copy()
# finalOrg.append(mapFunc(funcName, UnknownFunction, FunctionMapping))
# # finalOrg.append(comments)
# if finalOrg not in orgList:
# orgList.append(finalOrg)
# # else:
# # print("Duplicated", finalOrg)
# +
# print("results", len(results))
# print("orgList", len(orgList))
# print(orgList[0:5])
# -
# ## Merge what Meng-lun has done.
merge_file_name = 'output_class_to_orgs_20190501_v10_tana.xlsx'
merge_file = '{0}/{1}'.format(filepath, merge_file_name)
mergeDf = pd.read_excel(merge_file)
mergeDf = mergeDf.fillna(0)
mergeDf.head()
# +
mergeResults = []
for idx in range(0, len(mergeDf)):
row = mergeDf.loc[idx]
# print(row)
organization = str(row['Organization'])
# find the graph of the organization
graph = ''
for res in results:
resGraph = res['graph']
resOrg = res['class']
if resOrg == organization and resGraph == 'chrj_pr':
break
mergeResults.append([organization, graph])
# societalSector = str(row['SocietalSector'])
# period = str(row['Period'])
# corrName = str(row['Correct name'])
# if organization not in mergeResults:
# mergeResults[organization] = {'societalSector': societalSector, 'period': period, 'Correct': corrName}
# if societalSector != '0':
# mergeResults[organization]['societalSector'] = societalSector
# if period != '0':
# mergeResults[organization]['period'] = period
# if comments != '0':
# mergeResults[organization]['comments'] = comments
# mergeResults
# -
mergeResults[0:5]
print(len(mergeResults))
orgDf = pd.DataFrame(mergeResults, columns=['org', 'graph'])
orgDf.head()
write_orgGraph_file = '{0}/{1}'.format(filepath, 'orgGraphFile.xlsx')
orgDf.to_excel(write_orgGraph_file)
# +
# mergeList = []
# for org in orgList:
# orgName = org[0]
# periodSuggest = org[1]
# societalSuggest = org[2]
# merge = []
# merge.append(orgName)
# comments = ''
# if orgName in mergeResults:
# societal = mergeResults[orgName]['societalSector']
# if societal == '0':
# societal = ''
# merge.append(societal)
# period = mergeResults[orgName]['period']
# if period == '0':
# period = ''
# merge.append(period)
# comments = mergeResults[orgName]['comments']
# if comments == '0':
# comments = ''
# else:
# merge.append('')
# merge.append('')
# merge.append(societalSuggest)
# merge.append(periodSuggest)
# merge.append(comments)
# mergeList.append(merge)
# print(len(mergeList), mergeList[0:5])
# -
# ## Output file
# +
# def getMaxLen(orgList):
# maxEleLen = 0
# for org in orgList:
# orgLen = len(org)
# if maxEleLen < orgLen:
# maxEleLen = orgLen
# return maxEleLen
# print(getMaxLen(mergeList))
# # columns = ['Organization','Ownership','ModeOfEntry','Period','LevelOfOperation','SocietalSector','comments']
# columns = ['Organization','SocietalSector','Period','SocietalSectorSuggest','PeriodSuggest','comments']
# columns
# +
# orgDf = pd.DataFrame(mergeList, columns=columns)
# orgDf.drop_duplicates(subset='Organization', keep='first', inplace=True)
# orgDf.sort_values(by=['Organization', 'PeriodSuggest'], inplace=True)
# # orgDf = orgDf.fillna('')
# orgDf.head()
# +
# orgDf.to_excel(write_file_to, index=False)
# +
# # double check the duplicated
# duplicateRowsDF = orgDf[orgDf.duplicated(['Organization'], keep='last')]
# duplicateRowsDF
# -
| tbio/reorg_class_org.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Transformations
#
# One important piece of the visualization pipeline is **data transformation**.
#
# With Altair, you have two possible routes for data transformation; namely:
#
# 1. pre-transformation in Python
# 2. transformation in Altair/Vega-Lite
# +
import altair as alt
# Altair plots render by default in JupyterLab and nteract
# Uncomment/run this line to enable Altair in the classic notebook (not in JupyterLab)
# alt.renderers.enable('notebook')
# Uncomment/run this line to enable Altair in Colab
# alt.renderers.enable('colab')
# -
# ## Calculate Transform
#
# As an example, let's take a look at transforming some input data that is not encoded in the most intuitive manner.
# The ``population`` dataset lists aggregated US census data by year, sex, and age, but lists the sex as "1" and "2", which makes charts labels not particularly intuitive:
from vega_datasets import data
population = data.population()
population.head()
alt.Chart(population).mark_bar().encode(
x='year:O',
y='sum(people):Q',
color='sex:N'
)
# One way we could address this from Python is to use tools in Pandas to re-map these column names; for example:
(alt
.Chart(data=population.assign(sex = lambda fr: fr['sex'].map({1: 'Men', 2: 'Women'})))
.mark_bar()
.encode(
x='year:O',
y='sum(people):Q',
color='sex:N'
))
# But Altair is designed to be used with URL-based data as well, in which such pre-processing is not available.
# In these situations, it is better to make the transformation *part of the plot specification*.
# Here this can be done via the ``transform_calculate`` method, which accepts a [Vega Expression](), which is essentially a string that can contain a small subset of javascript operations:
(alt
.Chart(population)
.transform_calculate(men_women='datum.sex == 1 ? "Men" : "Women"')
.mark_bar()
.encode(x='year:O',
y='sum(people):Q',
color='men_women:N')
)
# The one potentially confusing piece is the presence of the word "datum": this is simply the convention by which Vega expressions refer to a row of the data.
#
# If you would prefer to build these expressions in Python, Altair provides a lightweight API to do so:
# +
from altair.expr import datum, if_
alt.Chart(population).mark_bar().encode(
x='year:O',
y='sum(people):Q',
color='men_women:N'
).transform_calculate(
men_women=if_(datum.sex == 1, "Men", "Women")
)
# -
# ## Filter Transform
#
# The filter transform is similar. For example, suppose you would like to create a chart consisting only of the male population from these census records.
# As above, this could be done from Pandas, but it is useful to have this operation available within the chart specification as well.
# It can be done via the ``transform_filter()`` method:
alt.Chart(population).mark_bar().encode(
x='year:O',
y='sum(people):Q',
).transform_filter(
"datum.sex == 1"
)
# We have seen this ``transform_filter`` method before, when we filtered based on the result of a selection.
# ## Other Transforms
#
# Other transform methods are available, and though we won't demonstrate them here, there are examples available in the Altair [Transform documentation]().
#
# Altair provides a number of other transforms. Some will be quite familiar:
#
# - ``transform_aggregate()``
# - ``transform_bin()``
# - ``transform_timeUnit()``
#
# These three transforms accomplish exactly the types of operations we discussed in [03-Binning-and-aggregation](03-Binning-and-aggregation.ipynb), with the distinction that they result in the creation of a new named value that can be referenced in multiple places throughout the chart.
#
# Two other transforms exist as well:
#
# - ``transform_lookup()``: this lets you perform one-sided joins of multiple datasets. It is used often, for example, in geographic visualizations where you join data (such as unemployment within states) to data about geographic regions used to represent that data
# - ``transform_window()``: this lets you perform aggregates across sliding windows, for example computing local means of data. It was recently added to Vega-Lite, and so the Altair API for this transform is not yet very convenient.
# ## Exercise
#
# Take the following data:
import pandas as pd
import numpy as np
x = pd.DataFrame({'x': np.linspace(-5, 5)})
# 1. Create a chart based on this data, and plot sine and cosine curves using Altair's ``transform_calculate`` API.
# 2. Use ``transform_filter`` on this chart, and remove the regions of the plot where the value of the cosine curve is less than the value of the sine curve.
| 04-altair/notebooks/07-Transformations.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.0
# language: julia
# name: julia-1.0
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # A Very Quick Introduction to Git/Github for Julia Users
#
# Julia's package system and Github are very closely intertwined:
#
# - Julia's package management system (METADATA) is a Github repository
# - The packages are hosted as Github repositories
# - Julia packages are normally referred to with the ending “.jl”
# - Repositories register to become part of the central package management by sending a pull request to METADATA.jl
# - The packages can be found / investigated at Github.com
# - Julia's error messages are hyperlinks to the page in Github
#
# Because of this, it's very useful for everyone using Julia to know a little bit about Git/Github.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Git Basics
#
# - Git is a common Version Control System (VCS)
# - A project is a **repository** (repos)
# - After one makes changes to a project, they **commit** the changes
# - Changes are **pulled** to the main repository hosted online
# - To download the code, you **clone** the repository
# - Instead of editing the main repository, one edits a **branch**
# - To get the changes of the main branch in yours, you **fetch**, **pull**, or **rebase**
# - One asks the owner of the repository to add their changes via a **pull request**
# - Stable versions are cut to **releases**
# + [markdown] slideshow={"slide_type": "slide"}
# ## Github Basics
#
# - The major online server for git repositories is Github
# - Github is a free service
# - Anyone can get a Github account
# - The code is hosted online, free for everyone to view
# - Users can open **Issues** to ask for features and give bug reports to developers
# - Many projects are brought together into **organizations** (JuliaMath, JuliaDiffEq, JuliaStats, etc.)
#
# An example Github repository for a Julia package is is DifferentialEquations.jl: https://github.com/JuliaDiffEq/DifferentialEquations.jl
# + [markdown] slideshow={"slide_type": "slide"}
# ## Basic through Advanced / Video Tutorial on Github
# - For the visually inclined, here is a video tutorial on basic and advanced github workflow for developing and editing Julia Packages, as well as for setting up Continuous Integration (CI).
#
# [](https://www.youtube.com/watch?v=tx8DRc7_c9I)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Examining a Github Repository
#
# 
#
# Components:
#
# - Top Left: Username/Repository name
# - Top Right: The stars. Click this button to show support for the developer!
# - Issues Tab: Go here to file a bug report
# - Files: These are the source files in the repository
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Examining A Github Repository
#
# 
#
# The badges on a Github repository show you the current state of the repo. From left to right:
#
# - Gitter Badge: Click this to go to a chatroom and directly chat with the developers
# - CI Build Badges: This tell you whether the CI tests pass. Click on them to see what versions of Julia the package works for.
# - Coverage: This tells you the percentage of the code that the tests cover. If coverage is low, parts of the package may not work even if the CI tests pass.
# - Docs Badges: Click on these to go to the package documentation.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Github Organizations
#
# - A (mostly complete) list of Julia organizations can be found at http://julialang.org/community/
# - Organizations manage large domains of packages and ensure that they work well together
# - Examples: JuliaStats, JuliaMath, JuliaDiffEq
# - Organizations are informal but have been "unusually effective"
# - Packages from the main Julia organizations can be considered official
# - Some functionality which used to be in the Base language now exists in organization packages
# + [markdown] slideshow={"slide_type": "slide"}
# ## Using Julia's Package Manager
#
# ### Adding a Package
#
# Julia's package manager functions are mirror the Git functions. Julia's package system is similar to R/Python in that a large number of packages are freely available. You search for them in places like [Julia Observer](https://juliaobserver.com/), or from the [Julia Package Listing](http://pkg.julialang.org/). Let's take a look at the [Plots.jl package by <NAME>](https://github.com/tbreloff/Plots.jl). To add a package, use `Pkg.add`
# + slideshow={"slide_type": "fragment"}
using Pkg
Pkg.update() # You may need to update your local packages first
Pkg.add("Plots")
# + [markdown] slideshow={"slide_type": "fragment"}
# This will install the package to your local system. However, this will only work for registered packages. To add a non-registered package, go to the Github repository to find the clone URL and use `Pkg.clone`. For example, to install the `ParameterizedFunctions` package, we can use:
# + slideshow={"slide_type": "fragment"}
Pkg.clone("https://github.com/JuliaDiffEq/ParameterizedFunctions.jl")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Importing a Package
#
# To use a package, you have to import the package. The `import` statement will import the package without exporting the functions to the namespace. (Note that the first time a package is run, it will precompile a lot of the functionality.) For example:
# + slideshow={"slide_type": "fragment"}
import Plots
Plots.plot(rand(4,4))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exporting Functionality
#
# To instead export the functions (of the developers choosing) to the namespace, we can use the `using` statement. Since Plots.jl exports the `plot` command, we can then use it without reference to the package that it came from:
# + slideshow={"slide_type": "fragment"}
using Plots
plot(rand(4,4))
# + [markdown] slideshow={"slide_type": "fragment"}
# What really makes this possible in Julia but not something like Python is that namespace clashes are usually avoided by multiple dispatch. Most packages will define their own types in order to use dispatches, and so when they export the functionality, the methods are only for their own types and thus do not clash with other packages. Therefore it's common in Julia for concise syntax like `plot` to be part of packages, all without fear of clashing.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Getting on the Latest Version
#
# Since Julia is currently under lots of development, you may wish to checkout newer versions. By default, `Pkg.add` is the "latest release", meaning the latest tagged version. However, the main version shown in the Github repository is usually the "master" branch. It's good development practice that the latest release is kept "stable", while the "master" branch is kept "working", and development takes place in another branch (many times labelled "dev"). You can choose which branch your local repository takes from. For example, to checkout the master branch, we can use:
# + slideshow={"slide_type": "fragment"}
Pkg.checkout("Plots")
# + [markdown] slideshow={"slide_type": "fragment"}
# This will usually gives us pretty up to date features (if you are using a "unreleased version of Julia" like building from the source of the Julia nightly, you may need to checkout master in order to get some packages working). However, to go to a specific branch we can give the branch as another argument:
# + slideshow={"slide_type": "fragment"}
Pkg.checkout("Plots","dev")
# + [markdown] slideshow={"slide_type": "fragment"}
# This is not advised if you don't know what you're doing (i.e. talk to the developer or read the pull requests (PR)), but this is common if you talk to a developer and they say "yes, I already implemented that. Checkout the dev branch and use `plot(...)`).
| Notebooks/GithubIntroduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Classifier
# Automatic image classifier using tensorflow
#
# To use it just import the Classifier (`from Classify import Classifier`)
#
# (You need the _Classify.py_ file from this repo in the same directory in order to do so)
#
# The Classifier has several inputs:
#
# - The first argument is `images` and those are the images for the training (required)
#
# - The second argument is `labels` and those are the labels for the training (required)
#
# - The third argument is `number_of_neurons` and this is the number of neurons in the second layer [default: 100]
#
# - The forth argument is `method` and this is the activation method of the second layer [default: relu]
#
# - The fifth argument is `optimizer` and this is the optimizer used for the training [default: Adam]
#
# - The last argument is `runs` and this is the number of times the model will be trained [default: 1]
#
# See the example below for more details:
# !python3 -m pip install tensorflow matplotlib --user
# %matplotlib inline
from Classify import Classifier
from tensorflow.keras.datasets import fashion_mnist
from matplotlib import pyplot
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
labels = ["Top", "Trouser", "Pullover", "Dress", "Coat", "Sandals", "Shirt", "Sneaker", "Bag", "Boot"]
model = Classifier(train_images, train_labels, runs = 3)
pyplot.imshow(test_images[0], cmap = pyplot.cm.binary)
pyplot.title(f"Prediction: {labels[model.predict_label(test_images[0])]}")
pyplot.show()
pyplot.imshow(test_images[1], cmap = pyplot.cm.binary)
pyplot.title(f"Prediction: {labels[model.predict_label(test_images[1])]}")
pyplot.show()
pyplot.imshow(test_images[2], cmap = pyplot.cm.binary)
pyplot.title(f"Prediction: {labels[model.predict_label(test_images[2])]}")
pyplot.show()
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collaborative filtering with side information
# ** *
# This IPython notebook illustrates the usage of the [cmfrec](https://github.com/david-cortes/cmfrec) Python package for collective matrix factorization using the [MovieLens-1M data](https://grouplens.org/datasets/movielens/1m/), consisting of ratings from users about movies + user demographic information, plus the [movie tag genome](https://grouplens.org/datasets/movielens/latest/).
#
# Collective matrix factorization is a technique for collaborative filtering with additional information about the users and items, based on low-rank joint factorization of different matrices with shared factors – for more details see the paper [_<NAME>., & <NAME>. (2008, August). Relational learning via collective matrix factorization. In Proceedings of the 14th ACM SIGKDD international conference on Knowledge discovery and data mining (pp. 650-658). ACM._](http://ra.adm.cs.cmu.edu/anon/usr/ftp/ml2008/CMU-ML-08-109.pdf).
#
# ** Small note: if the TOC here is not clickable or the math symbols don't show properly, try visualizing this same notebook from nbviewer following [this link](http://nbviewer.jupyter.org/github/david-cortes/cmfrec/blob/master/example/cmfrec_movielens_sideinfo.ipynb). **
# ** *
# ## Sections
#
#
# [1. Model description](#p1)
#
# [2. Loading the data](#p2)
# * [2.1 Ratings data](#p21)
# * [2.2 Creating a train and test split](#p22)
# * [2.3 Processing item tags](#p23)
# * [2.4 Processing user demographic info](#p24)
#
# [3. Basic model - only movie ratings](#p3)
# * [3.1 Fitting the model](#p31)
# * [3.2 Evaluating results](#p32)
#
# [4. Model with user side information](#p4)
# * [4.1 Original version](#p41)
# * [4.2 Offsets model](#p42)
#
# [5. Model with item side information](#p5)
# * [5.1 Original version](#p51)
# * [5.2 Offsets model](#p52)
#
# [6. Full model](#p6)
# * [6.1 Original version](#p61)
# * [6.2 Offsets model](#p62)
#
# [7. Examining some recomendations](#p7)
# ** *
# <a id="p1"></a>
# # 1. Model description
#
# The colective matrix mactorization model is an extension of the typical low-rank matrix factorization model to incorporate user and/or item side information. In its most basic form, low-rank matrix factorization tries to find an approximation of a matrix $X$ given by two lower-rank matrices $A$ and $B$, which in recommender systems would represent, respectively, a matrix of latent factors for users and items, which are determined by minimizing the squared differences between their product and $X$, i.e.:
#
# $$ argmin_{A, B} \lVert X - AB^T \lVert$$
#
# This basic formula can be improved by adding regularization on the $A$ and $B$ matrices, as well as by centering the matrix $X$ by substracting its global mean for each entry, adding user and item biases, and considering only the non-missing entries, i.e.:
#
# $$ argmin_{A, B, U_b, I_b} \lVert (X - \mu - U_b - I_b - AB^T)I_{x} \lVert^2 + \lambda (\lVert A\lVert^2 + \lVert B \lVert^2 + \lVert U_b \lVert^2 + \lVert I_b \lVert^2) $$
#
# Where:
# * $X$ is the ratings matrix (entry in row $\{ i,j\}$ contains the rating given by user $i$ to item $j$).
# * $A$ and $B$ are lower-dimensional matrices (model parameters).
# * $U_b$ is a column matrix of user biases, containing at each row a constant for its respective user.
# * $I_b$ is a row matrix of item biases, containing at each column a constant for its respective item.
# * $mu$ is the mean of the entries in $X$.
# * $I_x$ is an indicator matrix with entries in row $\{i,j\}$ equal to one when that same entry is present in the matrix $X$, and equal to zero when the corresponding entry is missing in $X$.
# * $\lambda$ is a regularization parameter.
#
# In collective matrix factorization, this model is further extended by also factorizing matrices of user and/or item side information (e.g. movie tags, user demographic info in tabular format, etc.)., e.g.:
#
# $$ argmin_{A, B, C, D, U_b, I_b} \lVert (X - \mu - U_b - I_b - AB^T)I_{x} \lVert^2 + \lVert U - AC^T \lVert^2 + \lVert I - BD^T \lVert^2 + \lambda (\lVert A\lVert^2 + \lVert B \lVert^2 + \lVert C \lVert^2 + \lVert D \lVert^2 + \lVert U_b \lVert^2 + \lVert I_b \lVert^2) $$
#
# Where, in addition to the previous model:
# * $U$ is the user side information matrix.
# * $I$ is the item side information matrix.
# * $C$ is a matrix of latent factors for user attributes (model parameters).
# * $D$ is a matrix of latent factors for item attributes (model parameters).
#
# (Other variations such as different weights for each factorization, different regularization for each paramter, and most notably, applying a sigmoid function on factorized values for binary variables, among others, are also possible to fit with this package).
#
# Intuitively, latent factors that also do a good job at explainin user/item attributes should generalize better to ratings data than latent factors that don't, even though it might adversely affect training error in the factorization of interest (the $X$ matrix).
#
# Alternatively, the package can also use a different formulation, in which the user and/or item attributes can be though of as the base of the factorization, with additional latent matrices acting as offsets for each user and item (deviations from its expected ratings according to the side information), e.g.:
#
# $$ argmin_{A, B, C, D, U_b, I_b} \lVert (X - \mu - U_b - I_b - (UC + A)(ID + B)\:)I_{x} \lVert^2 + \lambda (\lVert A\lVert^2 + \lVert B \lVert^2 + \lVert C \lVert^2 + \lVert D \lVert^2 + \lVert U_b \lVert^2 + \lVert I_b \lVert^2) $$
#
# Both of these models allow for making recommendations based only on user/item side information without ratings, in the first case by either training the model with extra users/items, or by computing the corresponding rows/columns of $A$ and $B$ by minimizing *only* the factorization of $U$ and $I$ (for a new user/item, there won't be any new entries in $C$ or $D$), which can be done in closed form; and in the second case, by setting the corresponding rows/columns of $A$ and $B$ to zero. As the ratings are centered, the expected value of both $U_b$ and $I_b$ are zero, which aids in cold-start recommendations.
#
# ** *
# <a id="p2"></a>
# # 2. Loading the data
#
# This example notebook uses the MovieLens 1M dataset, with movie tags taken from the latest MovieLens release, and user demographic information linked to the user information provided in the dataset, by taking a [publicly available table](http://federalgovernmentzipcodes.us/) mapping zip codes to states, [another one](http://www.fonz.net/blog/archives/2008/04/06/csv-of-states-and-state-abbreviations/) mapping state names to their abbreviations, and finally classifying the states into regions according to [usual definitions](https://www.infoplease.com/us/states/sizing-states).
#
# Unfortunately, later (bigger) release of the MovieLens dataset no longer include include user demographic information.
#
# <a id="p21"></a>
# ## 2.1 Ratings data
#
# The ratings come in the form of a table with columns UserId, ItemId, Rating, and Timestamp:
# +
import numpy as np, pandas as pd, time, re
from datetime import datetime
from cmfrec import CMF
ratings = pd.read_table('~/movielens/ml-1m/ratings.dat', sep='::',
engine='python', names=['UserId','ItemId','Rating','Timestamp'])
del ratings['Timestamp']
ratings.head()
# -
# <a id="p22"></a>
# ## 2.2 Creating a train and test split
#
# Usually, a good way to test recommender models is by temporal splits (splitting the data at some temporal cutoff point between train and test), but in this case, it's more desirable to make a distinction between warm start (predicting ratings strictly for users and items that were in the training data), and different forms of cold-start, i.e.: completely new users and items, new users with known items, and vice versa, which I'll try do here:
# +
np.random.seed(1)
user_ids = ratings.UserId.drop_duplicates().values
item_ids = ratings.ItemId.drop_duplicates().values
users_train = set(np.random.choice(user_ids, size=int(user_ids.shape[0] * .75), replace=False))
items_train = set(np.random.choice(item_ids, size=int(item_ids.shape[0] * .75), replace=False))
train = ratings.loc[ratings.UserId.isin(users_train) & ratings.ItemId.isin(items_train)].reset_index(drop=True)
np.random.seed(1)
train_ix = train.sample(frac=.85).index
test_ix = np.setdiff1d(train.index.values, train_ix)
test_warm_start = train.loc[test_ix].reset_index(drop=True)
train = train.loc[train_ix].reset_index(drop=True)
users_train = set(train.UserId)
items_train = set(train.ItemId)
test_warm_start = test_warm_start.loc[test_warm_start.UserId.isin(users_train) &
test_warm_start.ItemId.isin(items_train)].reset_index(drop=True)
test_cold_start = ratings.loc[~ratings.UserId.isin(users_train) & ~ratings.ItemId.isin(items_train)].reset_index(drop=True)
test_new_users = ratings.loc[(~ratings.UserId.isin(users_train)) & (ratings.ItemId.isin(items_train))].reset_index(drop=True)
test_new_items = ratings.loc[(ratings.UserId.isin(users_train)) & (~ratings.ItemId.isin(items_train))].reset_index(drop=True)
test_new_items_set = set(test_new_items.ItemId)
users_coldstart = set(test_cold_start.UserId)
items_coldstart = set(test_cold_start.ItemId)
print(train.shape)
print(test_warm_start.shape)
# -
print(len(users_train))
print(len(items_train))
# <a id="p23"></a>
# ## 2.3 Processing item tags
#
# Item tags were taken from the latest MovieLens release, and joined to the dataset by movie title, which is not a perfect match but does a reasonable job. They are unfortunately not available for all the movies for which there are ratings.
#
# For the second model, as the dimensionality of the tags is quite high, I'll also take a smaller transformation consisting of the first 50 principal components of these tags.
# +
movie_titles = pd.read_table('~/movielens/ml-1m/movies.dat',
sep='::', engine='python', header=None)
movie_titles.columns = ['ItemId', 'title', 'genres']
movie_titles = movie_titles[['ItemId', 'title']]
# will save the movie titles for later
movie_id_to_title = {i.ItemId:i.title for i in movie_titles.itertuples()}
movies = pd.read_csv('~/movielens/ml-latest/movies.csv')
movies = movies[['movieId', 'title']]
movies = pd.merge(movies, movie_titles)
movies = movies[['movieId', 'ItemId']]
tags = pd.read_csv('~/movielens/ml-latest/genome-scores.csv')
tags_wide = tags.pivot(index='movieId', columns='tagId', values='relevance')
tags_wide.columns=["tag"+str(i) for i in tags_wide.columns.values]
item_side_info = pd.merge(movies, tags_wide, how='inner', left_on='movieId', right_index=True)
del item_side_info['movieId']
items_w_sideinfo = set(item_side_info.ItemId)
test_new_items = test_new_items.loc[test_new_items.ItemId.isin(items_w_sideinfo)].reset_index(drop=True)
item_sideinfo_train = item_side_info.loc[item_side_info.ItemId.isin(items_train)].reset_index(drop=True)
item_sideinfo_testnew = item_side_info.loc[item_side_info.ItemId.isin(test_new_items_set)].reset_index(drop=True)
test_cold_start = test_cold_start.loc[test_cold_start.ItemId.isin(items_w_sideinfo)].reset_index(drop=True)
item_sideinfo_train.head()
# +
from sklearn.decomposition import PCA
pca_obj = PCA(n_components = 50)
item_sideinfo_reduced = item_side_info.copy()
del item_sideinfo_reduced['ItemId']
pca_obj.fit(item_sideinfo_reduced)
item_sideinfo_pca = pca_obj.transform(item_sideinfo_reduced)
item_sideinfo_pca = pd.DataFrame(item_sideinfo_pca)
item_sideinfo_pca.columns = ["pc"+str(i) for i in range(item_sideinfo_pca.shape[1])]
item_sideinfo_pca['ItemId'] = item_side_info.ItemId.values.copy()
item_sideinfo_pca_train = item_sideinfo_pca.loc[item_sideinfo_pca.ItemId.isin(items_train)].reset_index(drop=True)
item_sideinfo_pca_testnew = item_sideinfo_pca.loc[item_sideinfo_pca.ItemId.isin(test_new_items_set)].reset_index(drop=True)
item_sideinfo_pca_coldstart = item_sideinfo_pca.loc[item_sideinfo_pca.ItemId.isin(items_coldstart)].reset_index(drop=True)
item_sideinfo_pca_train.head()
# -
print(test_new_items.shape[0])
print(test_new_items.ItemId.drop_duplicates().shape[0])
# <a id="p24"></a>
# ## 2.4 Processing user demographic info
#
# The extra data is exaplained at the beginning. Joining all the data:
# +
zipcode_abbs = pd.read_csv("~/movielens/states.csv")
zipcode_abbs_dct = {z.State:z.Abbreviation for z in zipcode_abbs.itertuples()}
us_regs_table = [
('New England', 'Connecticut, Maine, Massachusetts, New Hampshire, Rhode Island, Vermont'),
('Middle Atlantic', 'Delaware, Maryland, New Jersey, New York, Pennsylvania'),
('South', 'Alabama, Arkansas, Florida, Georgia, Kentucky, Louisiana, Mississippi, Missouri, North Carolina, South Carolina, Tennessee, Virginia, West Virginia'),
('Midwest', 'Illinois, Indiana, Iowa, Kansas, Michigan, Minnesota, Nebraska, North Dakota, Ohio, South Dakota, Wisconsin'),
('Southwest', 'Arizona, New Mexico, Oklahoma, Texas'),
('West', 'Alaska, California, Colorado, Hawaii, Idaho, Montana, Nevada, Oregon, Utah, Washington, Wyoming')
]
us_regs_table = [(x[0], [i.strip() for i in x[1].split(",")]) for x in us_regs_table]
us_regs_dct = dict()
for r in us_regs_table:
for s in r[1]:
us_regs_dct[zipcode_abbs_dct[s]] = r[0]
zipcode_info = pd.read_csv("~/movielens/free-zipcode-database.csv")
zipcode_info = zipcode_info.groupby('Zipcode').first().reset_index()
zipcode_info['State'].loc[zipcode_info.Country != "US"] = 'UnknownOrNonUS'
zipcode_info['Region'] = zipcode_info['State'].copy()
zipcode_info['Region'].loc[zipcode_info.Country == "US"] = \
zipcode_info.Region\
.loc[zipcode_info.Country == "US"]\
.map(lambda x: us_regs_dct[x] if x in us_regs_dct else 'UsOther')
zipcode_info = zipcode_info[['Zipcode', 'Region']]
users = pd.read_table('~/movielens/ml-1m/users.dat',
sep='::', names=["UserId", "Gender", "Age", "Occupation", "Zipcode"], engine='python')
users["Zipcode"] = users.Zipcode.map(lambda x: np.int(re.sub("-.*","",x)))
users = pd.merge(users,zipcode_info,on='Zipcode',how='left')
users['Region'] = users.Region.fillna('UnknownOrNonUS')
users['Occupation'] = users.Occupation.map(lambda x: str(x))
users['Age'] = users.Age.map(lambda x: str(x))
user_side_info = pd.get_dummies(users[['UserId', 'Gender', 'Age', 'Occupation', 'Region']])
users_w_sideinfo = set(user_side_info.UserId)
test_new_users = test_new_users.loc[test_new_users.ItemId.isin(users_w_sideinfo)].reset_index(drop=True)
user_sideinfo_train = user_side_info.loc[user_side_info.UserId.isin(users_train)].reset_index(drop=True)
test_cold_start = test_cold_start.loc[test_cold_start.UserId.isin(users_w_sideinfo)].reset_index(drop=True)
user_sideinfo_train.head()
# -
print(test_new_users.shape[0])
print(test_new_users.UserId.drop_duplicates().shape[0])
print(test_cold_start.shape[0])
print(test_cold_start.UserId.unique().shape[0])
print(test_cold_start.ItemId.unique().shape[0])
# ** *
# <a id="p3"></a>
# # 3. Basic model - only movie ratings
#
# Non-collective factorization model - including user and item biases + regularization:
#
# <a id="p31"></a>
# ## 3.1 Fitting the model
# +
# %%time
from copy import deepcopy
from cmfrec import CMF
model_no_side_info = CMF(k=40, reg_param=1e-4, random_seed=1)
model_no_side_info.fit(deepcopy(train))
test_warm_start['Predicted'] = model_no_side_info.predict(test_warm_start.UserId, test_warm_start.ItemId)
# -
# <a id="p32"></a>
# ## 3.2 Evaluating results
#
# For this model and the ones that will follow, I will evaluate the recommendations by computing:
# * Root mean squared error (RMSE), i.e. sum( sqrt( (real - predicted)^2 ) ) - which can be though of the average star-rating error for each predicted rating. This is the most typical measure but has some drawbacks as it doesn't tend to be a good measure when ranking and can be substantially improved without changing the relative order of predictions.
# * Taking the average rating of the top-10 recommended movies for each user.
#
# There are other more appropriate evaluation criteria, but these are easy to understand and provide reasonable insights on model performance.
print("RMSE (no side info, warm start): ", np.sqrt(np.mean( (test_warm_start.Predicted - test_warm_start.Rating)**2) ))
# +
avg_ratings = train.groupby('ItemId')['Rating'].mean().to_frame().rename(columns={"Rating" : "AvgRating"})
test_ = pd.merge(test_warm_start, avg_ratings, left_on='ItemId', right_index=True, how='left')
print('Averge movie rating:', test_.groupby('UserId')['Rating'].mean().mean())
print('Average rating for top-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for top-10 recommendations of best-rated movies:', test_.sort_values(['UserId','AvgRating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('----------------------')
print('Average rating for top-10 recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 (non-)recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
# -
# ** *
# <a id="p4"></a>
# # 4. Model with user side information
#
# Now I'll add only the user information (without the movie tags). These are exclusively binary columns (only 0/1 values), so I'll apply a sigmoid function on the factorized values to be between zero and one.
#
# <a id="p41"></a>
# ## 4.1 Original version
# %%time
model_user_info = CMF(k=40, w_main=10.0, w_user=1.0, reg_param=1e-3, random_seed=1)
model_user_info.fit(deepcopy(train),
user_info=deepcopy(user_sideinfo_train),
cols_bin_user=[cl for cl in user_side_info.columns if cl!='UserId'])
test_warm_start['Predicted'] = model_user_info.predict(test_warm_start.UserId, test_warm_start.ItemId)
# In theory, new users can be incorporated into the model without refitting it entirely from scratch (a very slow procedure on larger datasets), but this is quite slow to do with many users. The following code would do it, but for time reasons it was not executed here:
# +
# # %%time
# for u in list(test_new_users.UserId.unique()):
# user_vec = user_side_info.loc[user_side_info.UserId == u]
# del user_vec['UserId']
# user_vec = user_vec.values.reshape((1, -1))
# model_user_info.add_user(new_id = u, attributes = user_vec)
# test_new_users['Predicted'] = model_user_info.predict(test_new_users.UserId, test_new_users.ItemId)
# -
# Side information from users which have no ratings can still be incorporated, and predictions can also be made for these users despite not having any ratings:
# %%time
model_user_info_all = CMF(k=40, w_main=10.0, w_user=1.0, reg_param=1e-3, random_seed=1)
model_user_info_all.fit(deepcopy(train),
user_info=deepcopy(user_side_info),
cols_bin_user=[cl for cl in user_side_info.columns if cl!='UserId'])
test_warm_start['PredictedAll'] = model_user_info_all.predict(test_warm_start.UserId, test_warm_start.ItemId)
test_new_users['PredictedAll'] = model_user_info_all.predict(test_new_users.UserId, test_new_users.ItemId)
# The evaluation metrics are the same as before plus simple correlation coefficient:
print("RMSE (user side info, warm start): ", np.sqrt(np.mean( (test_warm_start.Predicted - test_warm_start.Rating)**2) ))
print("RMSE (user side info, warm start, extra users): ", np.sqrt(np.mean( (test_warm_start.PredictedAll - test_warm_start.Rating)**2) ))
# print("RMSE (user side info, new users, added afterwards): ", np.sqrt(np.mean( (test_new_users.Predicted - test_new_users.Rating)**2) ))
print("RMSE (user side info, users trained without ratings): ", np.sqrt(np.mean( (test_new_users.PredictedAll - test_new_users.Rating)**2) ))
print("Rho (user side info, warm start): ", np.corrcoef(test_warm_start.Predicted, test_warm_start.Rating)[0][1])
print("Rho (user side info, warm start, extra users): ", np.corrcoef(test_warm_start.PredictedAll, test_warm_start.Rating)[0][1])
# print("RMSE (user side info, new users, added afterwards): ", np.corrcoef(test_new_users.Predicted, test_new_users.Rating)[0][1])
print("Rho (user side info, users trained without ratings): ", np.corrcoef(test_new_users.PredictedAll, test_new_users.Rating)[0][1])
# +
avg_ratings = train.groupby('ItemId')['Rating'].mean().to_frame().rename(columns={"Rating" : "AvgRating"})
test_ = pd.merge(test_warm_start, avg_ratings, left_on='ItemId', right_index=True, how='left')
print('Averge movie rating:', test_.groupby('UserId')['Rating'].mean().mean())
print('Average rating for top-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for top-10 recommendations of best-rated movies:', test_.sort_values(['UserId','AvgRating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('----------------------')
print('Average rating for top-10 recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 (non-)recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
# -
print('Average rating for top-10 recommendations (per user) from this model per configuration')
print('warm start: ', test_warm_start.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('warm start, extra users: ', test_warm_start.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# print('new users, added afterwards: ', test_new_users.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('users trained without ratings: ', test_new_users.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# This shows a slight improvement over not using user demographic information, and as shown, it gets a slight advantage by incorporating side information from more users than there are ratings from. It seems to perform surprisingly well on users trained without ratings too.
# <a id="p42"></a>
# ## 4.2 Offsets model
#
# Alternative formulation as explained in [section 1](#p1):
# %%time
model_user_info2 = CMF(k=40, reg_param=1e-4, offsets_model=True, random_seed=1)
model_user_info2.fit(deepcopy(train),
user_info = deepcopy(user_sideinfo_train))
test_warm_start['Predicted'] = model_user_info2.predict(test_warm_start.UserId, test_warm_start.ItemId)
# %%time
for u in list(test_new_users.UserId.unique()):
user_vec = deepcopy(user_side_info.loc[user_side_info.UserId == u])
del user_vec['UserId']
user_vec = user_vec.values.reshape((1, -1))
model_user_info2.add_user(new_id = u, attributes = user_vec)
test_new_users['Predicted'] = model_user_info2.predict(test_new_users.UserId, test_new_users.ItemId)
print("RMSE (user side info, warm start): ", np.sqrt(np.mean( (test_warm_start.Predicted - test_warm_start.Rating)**2) ))
print("RMSE (user side info, new users, added afterwards): ", np.sqrt(np.mean( (test_new_users.Predicted - test_new_users.Rating)**2) ))
print("Rho (user side info, warm start): ", np.corrcoef(test_warm_start.Predicted, test_warm_start.Rating)[0][1])
print("Rho (user side info, new users, added afterwards): ", np.corrcoef(test_new_users.Predicted, test_new_users.Rating)[0][1])
# +
avg_ratings = train.groupby('ItemId')['Rating'].mean().to_frame().rename(columns={"Rating" : "AvgRating"})
test_ = pd.merge(test_warm_start, avg_ratings, left_on='ItemId', right_index=True, how='left')
print('Averge movie rating:', test_.groupby('UserId')['Rating'].mean().mean())
print('Average rating for top-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for top-10 recommendations of best-rated movies:', test_.sort_values(['UserId','AvgRating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('----------------------')
print('Average rating for top-10 recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 (non-)recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
# -
print('Average rating for top-10 recommendations (per user) from this model per configuration')
print('warm start: ', test_warm_start.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('new users, added afterwards: ', test_new_users.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# Unfortunately, this formulation doesn't seem to perform as well as the previous one in either case, but it is much faster to add new users.
# <a id="p5"></a>
# # 5. Model with item side information
#
# Like before, now fitting the collective model incorporating movie tags, but not user information
#
# <a id="p51"></a>
# ## 5.1 Original version
# %%time
model_item_info = CMF(k=35, k_main=15, k_item=10, reg_param=1e-3, w_main=10.0, w_item=0.5, random_seed=1)
model_item_info.fit(deepcopy(train),
item_info = deepcopy(item_sideinfo_train))
test_warm_start['Predicted'] = model_item_info.predict(test_warm_start.UserId, test_warm_start.ItemId)
# Same as before, it's possible to add new items to the model without having to refit it entirely from scratch, but this is very slow to do with many items and was not run here for time reasons:
# +
# for i in test_new_items.ItemId.unique():
# item_vec = item_side_info.loc[item_side_info.ItemId == i]
# del user_vec['ItemId']
# item_vec = item_vec.values.reshape((1, -1))
# model_user_info.add_item(new_id = i, attributes = user_vec)
# test_new_items['Predicted'] = model_item_info.predict(test_new_items.UserId, test_new_items.ItemId)
# -
# %%time
model_item_info_all = CMF(k=35, k_main=15, k_item=10, reg_param=1e-3, w_main=10.0, w_item=0.5, random_seed=1)
model_item_info_all.fit(deepcopy(train), item_info = deepcopy(item_side_info))
test_warm_start['PredictedAll'] = model_item_info_all.predict(test_warm_start.UserId, test_warm_start.ItemId)
test_new_items['PredictedAll'] = model_item_info_all.predict(test_new_items.UserId, test_new_items.ItemId)
# This time, I will also try a version that puts more emphasis in correct factorization of the side information:
# %%time
model_item_info_diffweight = CMF(k=50, k_main=0, k_item=0, reg_param=1e-3, w_main=5.0, w_item=5.0, random_seed=1)
model_item_info_diffweight.fit(deepcopy(train), item_info = deepcopy(item_side_info))
test_warm_start['PredictedAll2'] = model_item_info_diffweight.predict(test_warm_start.UserId, test_warm_start.ItemId)
test_new_items['PredictedAll2'] = model_item_info_diffweight.predict(test_new_items.UserId, test_new_items.ItemId)
print("RMSE (item side info, warm start): ", np.sqrt(np.mean( (test_warm_start.Predicted - test_warm_start.Rating)**2) ))
print("RMSE (item side info, warm start, extra items): ", np.sqrt(np.mean( (test_warm_start.PredictedAll - test_warm_start.Rating)**2) ))
print("RMSE (item side info, warm start, extra items, diff. weighting): ", np.sqrt(np.mean( (test_warm_start.PredictedAll2 - test_warm_start.Rating)**2) ))
# print("RMSE (item side info, new items, added afterwards): ", np.sqrt(np.mean( (test_new_items.Predicted - test_new_items.Rating)**2) ))
print("RMSE (item side info, items trained without ratings): ", np.sqrt(np.mean( (test_new_items.PredictedAll - test_new_items.Rating)**2) ))
print("RMSE (item side info, items trained without ratings, diff. weighting): ", np.sqrt(np.mean( (test_new_items.PredictedAll2 - test_new_items.Rating)**2) ))
print("Rho (item side info, warm start): ", np.corrcoef(test_warm_start.Predicted, test_warm_start.Rating)[0][1])
print("Rho (item side info, warm start, extra items): ", np.corrcoef(test_warm_start.PredictedAll, test_warm_start.Rating)[0][1])
print("Rho (item side info, warm start, extra items, diff. weighting): ", np.corrcoef(test_warm_start.PredictedAll2, test_warm_start.Rating)[0][1])
# print("Rho (item side info, new items, added afterwards): ", np.corrcoef(test_new_items.Predicted, test_new_items.Rating)[0][1])
print("Rho (item side info, items trained without ratings): ", np.corrcoef(test_new_items.PredictedAll, test_new_items.Rating)[0][1])
print("Rho (item side info, items trained without ratings, diff. weighting): ", np.corrcoef(test_new_items.PredictedAll2, test_new_items.Rating)[0][1])
# +
avg_ratings = train.groupby('ItemId')['Rating'].mean().to_frame().rename(columns={"Rating" : "AvgRating"})
test_ = pd.merge(test_warm_start, avg_ratings, left_on='ItemId', right_index=True, how='left')
print('Averge movie rating:', test_.groupby('UserId')['Rating'].mean().mean())
print('Average rating for top-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for top-10 recommendations of best-rated movies:', test_.sort_values(['UserId','AvgRating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('----------------------')
print('Average rating for top-10 recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 (non-)recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
# -
print('Average rating for top-10 recommendations (per user) from this model per configuration')
print('warm start: ', test_warm_start.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('warm start, extra items: ', test_warm_start.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# print('new items, added afterwards: ', test_new_items.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('items trained without ratings: ', test_new_items.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('items trained without ratings, diff. weighting: ', test_new_items.sort_values(['UserId','PredictedAll2'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# The improvement is comparable to that form adding user side information in the warm-start case, but for new items, it seems the recommendations are not as good as for new users. Putting a heaver weight in the movie tags factorization didn't seem to make it perform better in cold-start according to ranking metrics, but it did bring a slight improvement in terms of RMSE.
# <a id="p52"></a>
# ## 5.2 Offsets model
# %%time
model_item_info2 = CMF(k=50, reg_param=1e-4, offsets_model=True, random_seed=1)
model_item_info2.fit(deepcopy(train), item_info = deepcopy(item_sideinfo_pca_train))
test_warm_start['Predicted'] = model_item_info2.predict(test_warm_start.UserId, test_warm_start.ItemId)
# %%time
for i in list(test_new_items.ItemId.unique()):
item_vec = deepcopy(item_sideinfo_pca.loc[item_sideinfo_pca.ItemId == i])
del item_vec['ItemId']
item_vec = item_vec.values.reshape((1, -1))
model_item_info2.add_item(new_id = i, attributes = item_vec)
test_new_items['Predicted'] = model_item_info2.predict(test_new_items.UserId, test_new_items.ItemId)
print("RMSE (item side info, warm start): ", np.sqrt(np.mean( (test_warm_start.Predicted - test_warm_start.Rating)**2) ))
print("RMSE (item side info, new items, added afterwards): ", np.sqrt(np.mean( (test_new_items.Predicted - test_new_items.Rating)**2) ))
print("Rho (item side info, warm start): ", np.corrcoef(test_warm_start.Predicted, test_warm_start.Rating)[0][1])
print("Rho (item side info, new items, added afterwards): ", np.corrcoef(test_new_items.Predicted, test_new_items.Rating)[0][1])
# +
avg_ratings = train.groupby('ItemId')['Rating'].mean().to_frame().rename(columns={"Rating" : "AvgRating"})
test_ = pd.merge(test_warm_start, avg_ratings, left_on='ItemId', right_index=True, how='left')
print('Averge movie rating:', test_.groupby('UserId')['Rating'].mean().mean())
print('Average rating for top-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 rated by each user:', test_.sort_values(['UserId','Rating'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for top-10 recommendations of best-rated movies:', test_.sort_values(['UserId','AvgRating'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('----------------------')
print('Average rating for top-10 recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('Average rating for bottom-10 (non-)recommendations from this model:', test_.sort_values(['UserId','Predicted'], ascending=True).groupby('UserId')['Rating'].head(10).mean())
# -
print('Average rating for top-10 recommendations (per user) from this model per configuration')
print('warm start: ', test_warm_start.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('new items, added afterwards: ', test_new_items.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# This alternative formulation doesn't perform as well for warm-start recommendations (in this regards, it seems even worse than when not incorporating it), but it performs significantly better for cold-start.
# <a id="p6"></a>
# # 6. Full model
#
# Now a model incorporating both user and item side information, fit to extra users and items without any ratings in the training set. Note that the hyperparameters of this model are a lot harder to tune.
# %%time
model_user_item_info = CMF(k=40, k_main=10, k_user=5, k_item=15,
w_main=1.0, w_user=2.0, w_item=0.05,
reg_param=5e-5, random_seed=1)
model_user_item_info.fit(deepcopy(train),
user_info=deepcopy(user_side_info),
item_info=deepcopy(item_side_info),
cols_bin_user=[cl for cl in user_side_info.columns if cl!='UserId'])
test_warm_start['PredictedAll'] = model_user_item_info.predict(test_warm_start.UserId, test_warm_start.ItemId)
test_cold_start['PredictedAll'] = model_user_item_info.predict(test_cold_start.UserId, test_cold_start.ItemId)
test_new_users['PredictedAll'] = model_user_item_info.predict(test_new_users.UserId, test_new_users.ItemId)
test_new_items['PredictedAll'] = model_user_item_info.predict(test_new_items.UserId, test_new_items.ItemId)
print("RMSE (user and item side info, warm start, extra users and items): ", np.sqrt(np.mean( (test_warm_start.PredictedAll - test_warm_start.Rating)**2) ))
print("RMSE (user and item side info, cold start): ", np.sqrt(np.mean( (test_cold_start.PredictedAll - test_cold_start.Rating)**2) ))
print("RMSE (user and item side info, users trained without ratings, extra items): ", np.sqrt(np.mean( (test_new_users.PredictedAll - test_new_users.Rating)**2) ))
print("RMSE (user and item side info, items trained without ratings, extra users): ", np.sqrt(np.mean( (test_new_items.PredictedAll - test_new_items.Rating)**2) ))
print("Rho (user and item side info, warm start, extra users and items): ", np.corrcoef(test_warm_start.PredictedAll, test_warm_start.Rating)[0][1])
print("Rho (user and item side info, cold start): ", np.corrcoef(test_cold_start.PredictedAll, test_cold_start.Rating)[0][1])
print("Rho (user and item side info, users trained without ratings, extra items): ", np.corrcoef(test_new_users.PredictedAll, test_new_users.Rating)[0][1])
print("Rho (user and item side info, items trained without ratings, extra users): ", np.corrcoef(test_new_items.PredictedAll, test_new_items.Rating)[0][1])
print('Average rating for top-10 recommendations (per user) from this model per configuration')
print('warm start, extra users and items: ', test_warm_start.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('cold start: ', test_cold_start.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('users trained without ratings: ', test_new_users.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('items trained without ratings: ', test_new_items.sort_values(['UserId','PredictedAll'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# The user and item side information didn't seem to improve upon the model without that uses only ratings for warm-start recomendations, but it seems to perform very well for cold-start - almost as good as for warm start in fact.
# ** *
# Alternative formulation with the "offsets" model:
# %%time
model_user_item_info2 = CMF(k=50, reg_param=5e-3, offsets_model=True, random_seed=1)
model_user_item_info2.fit(deepcopy(train),
user_info=deepcopy(user_sideinfo_train),
item_info=deepcopy(item_sideinfo_pca_train))
test_warm_start['Predicted'] = model_user_item_info2.predict(test_warm_start.UserId, test_warm_start.ItemId)
# +
# %%time
for u in list(np.unique(np.r_[test_new_users.UserId, test_cold_start.UserId])):
user_vec = deepcopy(user_side_info.loc[user_side_info.UserId == u])
del user_vec['UserId']
user_vec = user_vec.values.reshape((1, -1))
model_user_item_info2.add_user(new_id = u, attributes = user_vec)
for i in list(np.unique(np.r_[test_new_items.ItemId.unique(), test_cold_start.ItemId.unique()])):
item_vec = deepcopy(item_sideinfo_pca.loc[item_sideinfo_pca.ItemId == i])
if item_vec.shape[0] > 0:
del item_vec['ItemId']
item_vec = item_vec.values.reshape((1, -1))
model_user_item_info2.add_item(new_id = i, attributes = item_vec)
test_new_users['Predicted'] = model_user_item_info2.predict(test_new_users.UserId, test_new_users.ItemId)
test_new_items['Predicted'] = model_user_item_info2.predict(test_new_items.UserId, test_new_items.ItemId)
test_cold_start['Predicted'] = model_user_item_info2.predict(test_cold_start.UserId, test_cold_start.ItemId)
# -
print("RMSE (user and item side info, warm start, extra users and items): ", np.sqrt(np.mean( (test_warm_start.Predicted - test_warm_start.Rating)**2) ))
print("RMSE (user and item side info, cold start, users and items added afterwards): ", np.sqrt(np.mean( (test_cold_start.Predicted - test_cold_start.Rating)**2) ))
print("RMSE (user and item side info, users added afterwards): ", np.sqrt(np.mean( (test_new_users.Predicted - test_new_users.Rating)**2) ))
print("RMSE (user and item side info, items added afterwards): ", np.sqrt(np.mean( (test_new_items.Predicted - test_new_items.Rating)**2) ))
print("Rho (user and item side info, warm start, extra users and items): ", np.corrcoef(test_warm_start.Predicted, test_warm_start.Rating)[0][1])
print("Rho (user and item side info, cold start, users and items added afterwards): ", np.corrcoef(test_cold_start.Predicted, test_cold_start.Rating)[0][1])
print("Rho (user and item side info, users added afterwards): ", np.corrcoef(test_new_users.Predicted, test_new_users.Rating)[0][1])
print("Rho (user and item side info, items added afterwards): ", np.corrcoef(test_new_items.Predicted, test_new_items.Rating)[0][1])
print('Average rating for top-10 recommendations (per user) from this model per configuration')
print('warm start: ', test_warm_start.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('cold start: ', test_cold_start.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('users added afterwards: ', test_new_users.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
print('items added afterwards: ', test_new_items.sort_values(['UserId','Predicted'], ascending=False).groupby('UserId')['Rating'].head(10).mean())
# This model seems again to perform better for cold-start recommendations.
# <a id="p7"></a>
# # 7. Examining some recomendations
#
# Now I'll examine the Top-10 recommended movies for some random users under different models:
# +
from collections import defaultdict
# aggregate statistics
avg_movie_rating = defaultdict(lambda: 0)
num_ratings_per_movie = defaultdict(lambda: 0)
for i in train.groupby('ItemId')['Rating'].mean().to_frame().itertuples():
avg_movie_rating[i.Index] = i.Rating
for i in train.groupby('ItemId')['Rating'].agg(lambda x: len(tuple(x))).to_frame().itertuples():
num_ratings_per_movie[i.Index] = i.Rating
# function to print recommended lists more nicely
def print_reclist(reclist):
list_w_info = [str(m + 1) + ") - " + movie_id_to_title[reclist[m]] +\
" - Average Rating: " + str(np.round(avg_movie_rating[reclist[m]], 2))+\
" - Number of ratings: " + str(num_ratings_per_movie[reclist[m]])\
for m in range(len(reclist))]
print("\n".join(list_w_info))
# -
# User with ID = 948 - this user was in the training set:
# +
reclist1 = model_no_side_info.topN(user=948, n=10, exclude_seen=True)
reclist2 = model_user_info_all.topN(user=948, n=10, exclude_seen=True)
reclist3 = model_item_info_all.topN(user=948, n=10, exclude_seen=True)
reclist4 = model_user_item_info.topN(user=948, n=10, exclude_seen=True)
reclist5 = model_user_item_info2.topN(user=948, n=10, exclude_seen=True)
print('Recommendations from ratings-only model:')
print_reclist(reclist1)
print("------")
print('Recommendations from ratings + user demographics model:')
print_reclist(reclist2)
print("------")
print('Recommendations from ratings + movie tags model:')
print_reclist(reclist3)
print("------")
print('Recommendations from ratings + user demographics + movie tags model:')
print_reclist(reclist4)
print("------")
print('Recommendations from ratings + user demographics + movie tags model (alternative formulation):')
print_reclist(reclist5)
# -
# User with ID = 1 - this user was not in the training set:
# +
# reclist1 = model_no_side_info.topN(user=1, n=10) # not possible with this model
reclist2 = model_user_info_all.topN(user=1, n=10)
# reclist3 = model_item_info_all.topN(user=1, n=10) # not possible with this model
reclist4 = model_user_item_info.topN(user=1, n=10)
reclist5 = model_user_item_info2.topN(user=1, n=10)
# print('Recommendations from ratings-only model:')
# print_reclist(reclist1)
# print("------")
print('Recommendations from ratings + user demographics model:')
print_reclist(reclist2)
# print("------")
# print('Recommendations from ratings + movie tags model:')
# print_reclist(reclist3)
print("------")
print('Recommendations from ratings + user demographics + movie tags model:')
print_reclist(reclist4)
print("------")
print('Recommendations from ratings + user demographics + movie tags model (alternative formulation):')
print_reclist(reclist5)
# -
# As seen from these lists, the alternative formulation of the model tends to recommend more movies that were not in the training set, which in many contexts would be a desirable thing despite the slightly lower achieved metrics.
| example/cmfrec_movielens_sideinfo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import torch
import torch.nn as nn
import numpy as np
import matplotlib.pyplot as plt
from sklearn import datasets
% matplotlib inline
# ## Create toy data set
x_train, y_train = datasets.make_regression(n_features=1, n_informative=1, noise=20, bias=10)
x_train = x_train.astype(np.float32)
y_train = y_train[:,None].astype(np.float32)
plt.plot(x_train, y_train, 'v')
# ## Train a Linear Regression model
model = nn.Linear(1,1)
loss_fn = nn.MSELoss()
optimizer = torch.optim.SGD(model.parameters(), lr=0.005)
for epoch in range(1, 1001):
y_pred = model(torch.from_numpy(x_train))
optimizer.zero_grad()
loss = loss_fn(y_pred, torch.from_numpy(y_train))
loss.backward()
optimizer.step()
if epoch % 100 == 0:
print(f'Loss: {loss} at epoch {epoch}')
plt.plot(x_train, y_train, 'v', label='Original Data')
plt.plot(x_train, model(torch.from_numpy(x_train)).data.numpy(), label='Fitted Line')
plt.legend()
plt.show()
| backup/pytorch_basics_linear_model_20180515.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pickle
import os
def naive_init_dataset(Ninit, Flist, params, scores):
mf_Xtr_list = []
mf_ytr_list = []
mf_Xte_list = []
mf_yte_list = []
for fid in Flist:
buff_X = []
buff_y = []
for n in range(Ninit):
buff_X.append(params[n, :])
buff_y.append(scores[n, fid])
# buff_y.append(-scores[n, fid])
#
mf_Xtr_list.append(np.array(buff_X))
mf_ytr_list.append(np.array(buff_y).reshape([-1,1]))
#
mfdata = {}
mfdata['mf_Xtr_list'] = mf_Xtr_list
mfdata['mf_ytr_list'] = mf_ytr_list
mfdata['mf_Xte_list'] = mf_Xtr_list
mfdata['mf_yte_list'] = mf_ytr_list
return mfdata, mf_Xtr_list, mf_ytr_list
#
sobol=False
if sobol:
prefix = 'sobol_raw_'
else:
prefix = 'uniform_raw_'
domain='BurgersShock'
pickle_name = os.path.join('buff', domain, prefix+domain+'.pickle')
with open(pickle_name, 'rb') as handle:
raw = pickle.load(handle)
params = raw['X']
scores = raw['Y']
Ninit = 10
Flist = [0,1,2]
mfdata, mf_Xtr_list, mf_ytr_list = naive_init_dataset(Ninit, Flist, params, scores)
dump_path = 'preload'
if not os.path.exists(dump_path):
os.makedirs(dump_path)
dump_fname = domain+'.pickle'
with open(os.path.join(dump_path, dump_fname), 'wb') as handle:
pickle.dump(mfdata, handle, protocol=pickle.HIGHEST_PROTOCOL)
#
print(np.hstack([mf_ytr_list[0], mf_ytr_list[1], mf_ytr_list[2]]))
# -
| data/ProcessPredload.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Random Forest Example
#
# Implement Random Forest algorithm with TensorFlow, and apply it to classify
# handwritten digit images. This example is using the MNIST database of
# handwritten digits as training samples (http://yann.lecun.com/exdb/mnist/).
#
# - Author: <NAME>
# - Project: https://github.com/aymericdamien/TensorFlow-Examples/
# +
from __future__ import print_function
import tensorflow as tf
from tensorflow.python.ops import resources
from tensorflow.contrib.tensor_forest.python import tensor_forest
# Ignore all GPUs, tf random forest does not benefit from it.
import os
os.environ["CUDA_VISIBLE_DEVICES"] = ""
# -
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=False)
# +
# Parameters
num_steps = 500 # Total steps to train
batch_size = 1024 # The number of samples per batch
num_classes = 10 # The 10 digits
num_features = 784 # Each image is 28x28 pixels
num_trees = 10
max_nodes = 1000
# Input and Target data
X = tf.placeholder(tf.float32, shape=[None, num_features])
# For random forest, labels must be integers (the class id)
Y = tf.placeholder(tf.int32, shape=[None])
# Random Forest Parameters
hparams = tensor_forest.ForestHParams(num_classes=num_classes,
num_features=num_features,
num_trees=num_trees,
max_nodes=max_nodes).fill()
# +
# Build the Random Forest
forest_graph = tensor_forest.RandomForestGraphs(hparams)
# Get training graph and loss
train_op = forest_graph.training_graph(X, Y)
loss_op = forest_graph.training_loss(X, Y)
# Measure the accuracy
infer_op, _, _ = forest_graph.inference_graph(X)
correct_prediction = tf.equal(tf.argmax(infer_op, 1), tf.cast(Y, tf.int64))
accuracy_op = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# Initialize the variables (i.e. assign their default value) and forest resources
init_vars = tf.group(tf.global_variables_initializer(),
resources.initialize_resources(resources.shared_resources()))
# +
# Start TensorFlow session
sess = tf.train.MonitoredSession()
# Run the initializer
sess.run(init_vars)
# Training
for i in range(1, num_steps + 1):
# Prepare Data
# Get the next batch of MNIST data (only images are needed, not labels)
batch_x, batch_y = mnist.train.next_batch(batch_size)
_, l = sess.run([train_op, loss_op], feed_dict={X: batch_x, Y: batch_y})
if i % 50 == 0 or i == 1:
acc = sess.run(accuracy_op, feed_dict={X: batch_x, Y: batch_y})
print('Step %i, Loss: %f, Acc: %f' % (i, l, acc))
# Test Model
test_x, test_y = mnist.test.images, mnist.test.labels
print("Test Accuracy:", sess.run(accuracy_op, feed_dict={X: test_x, Y: test_y}))
| notebooks/2_BasicModels/random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import os.path
import fnmatch
import numpy as np
import matplotlib.pyplot as plt # after importting matplotlib, mayavi can not set api to 2: first run mayavi!!!!
from latexify import latexify
import scipy.io
from scipy import ndimage
from smoothn import smoothn
# %matplotlib notebook
# +
import matplotlib
'''To make the tex-style/mathtext text look like the regular text,
you need to set the mathtext font to Bitstream Vera Sans:'''
matplotlib.rcParams['mathtext.fontset'] = 'custom'
# set the font
plt.rcParams["font.family"] = 'arial' #'Bitstream Vera Sans'# 'arial' #
matplotlib.rcParams['mathtext.rm'] = 'arial' #'Bitstream Vera Sans'
matplotlib.rcParams['mathtext.it'] = 'arial:italic' #'Bitstream Vera Sans:italic'
matplotlib.rcParams['mathtext.bf'] = 'arial:bold' #'Bitstream Vera Sans:bold'
fig, ax = plt.subplots(figsize=(5, 4))
matplotlib.pyplot.title(r'ABC123 vs $ABC^2\mathrm{ABC123}^{123}\mathsf{ABCabc}$')
'''If you want the regular text to look like the mathtext text,
you can change everything to Stix. This will affect labels, titles, ticks, etc.'''
# matplotlib.rcParams['mathtext.fontset'] = 'stix'
# matplotlib.rcParams['font.family'] = 'arial' #'STIXGeneral'
# matplotlib.pyplot.text(0.,0.5,r'ABC123 vs $\mathrm{ABC123}^{123}\mathsf{ABCabc}$')
#plt.style.use(u'classic')#u'seaborn-paper') # Set back to default
# -
lam = np.array([0.248,0.196,0.177]) # \AA
lam/(2*np.sin(4.0/180.*3.14))
# +
def read_spe(fpath):
'''Read the spe inelastic neutron scttering data exported by MSlice
1. the empty grid is set to -1.00;
'''
with open(fpath, 'rU') as f:
i = 0
qs, es, data, error = [],[],[],[]
for line in f:
line = line.rstrip('\n')
if i==0:
shape = np.array(line.split(),dtype='int'); print('Data shape: ', shape)
i+=1
continue
#data, error = np.empty(shape), np.empty(shape)
if line==r'### |Q| (\AA!U-1!N)': # not use line.rstrip('\n').split()[1]=='\Q\' becasue stimes len(line)=1
readQ, readE, readI, readEr = True, False, False, False
continue
if line==r'### E (meV)':
readQ, readE, readI, readEr = False, True, False, False
continue
if line==r'### Intensity (arb. units)':
readQ, readE, readI, readEr = False, False, True, False
continue
if line==r'### Errors':
readQ, readE, readI, readEr = False, False, False, True
continue
if readQ:
#qs.append(np.array(line, dtype='f')); continue
qs.append(line.split()); continue
if readE:
es.append(line.split()); continue
if readI:
data.append(line.split()); continue
if readEr:
error.append(line.split()); continue
#return np.array(qs),np.array(es),np.array(data).reshape(shape),np.array(error).reshape(shape)
return np.array(np.concatenate(qs),dtype='f')[:-1], \
np.array(np.concatenate(es),dtype='f')[:-1], \
np.array(np.concatenate(data),dtype='f').reshape(shape), \
np.array(np.concatenate(error),dtype='f').reshape(shape)
# without the last 0.000
def binning2D(x,y,D, xybins):
'''Try to flatten the 2D data first and then to histogram2d
but nan weight is not take cared
The nb of elements in each bin at least larger than one (xybins<[len(x),len(y)])'''
X, Y = np.meshgrid(x,y)
xx, yy, dd = X.flatten(), Y.flatten(), D.T.flatten()
# use reshape([1,-1]) not working for it && must add '.T' (so that coresponding reightly)
#print xx, yy, dd
xbin_no_pts= np.histogram(x, xybins[0])[0] #the no of data points in every bin
ybin_no_pts= np.histogram(y, xybins[1])[0]
if 0 in np.concatenate([xbin_no_pts, ybin_no_pts]):
print("There are bins contanining 0 nb, desrease nb of bins;\nThe orginal data is returned")
return x, y, data
else:
binxy_no_pts = xbin_no_pts.reshape(xybins[0],1).dot(ybin_no_pts.reshape(1,xybins[1])) #2D: nb of point per xy bin
binx = np.histogram(x, bins=xybins[0],weights=x)[0]/ xbin_no_pts
biny = np.histogram(y, bins=xybins[1],weights=y)[0]/ ybin_no_pts
binD = np.histogram2d(xx,yy, bins=xybins, normed=False, weights=dd)[0]/binxy_no_pts
return binx, biny, binD
def binning2Dloop(x,y,D, xbins,ybins): # x and y are 1 by m or n arrays, D is m by n 2D data
'''do not take care of Nan wight!!!'''
xlen, ylen, ddim = len(x), len(y), D.shape
#print xlen, ylen, ddim
assert [xlen, ylen] == [ddim[0],ddim[1]]
xbin_no_pts= np.histogram(x, xbins)[0] #the no of data points in every bin
ybin_no_pts= np.histogram(y, ybins)[0]
#print "binning scheme:"; print xbin_no_pts, ybin_no_pts
binx = np.histogram(x,xbins,weights=x)[0] / xbin_no_pts
biny = np.histogram(y,ybins,weights=y)[0] / ybin_no_pts
Dbinx = np.array([ np.histogram(x, xbins, weights=D[:,i])[0] / xbin_no_pts for i in range(ddim[1])]) # shape:[ylen,xbins]
Dbiny = np.array([ np.histogram(y, ybins, weights=Dbinx[:,i])[0] / ybin_no_pts for i in range(xbins)]) #shape:[xbins,ybins]
# try to take care of nan: failed
# keep = ~np.isnan(D)
# Dbinx = np.array([ np.histogram(x[keep[:,i]], xbins, weights=D[keep[:,i],i])[0] / xbin_no_pts
# for i in range(ddim[1])]) # shape:[ylen,xbins]
# Dbiny = np.array([ np.histogram(y[keep[i,:]], ybins, weights=Dbinx[keep[i,:],i])[0] / ybin_no_pts
# for i in range(xbins)]) #shape:[xbins,ybins]
return binx, biny, Dbiny
from scipy import ndimage
def myGfilter(U, sigma, order=0, output=None, mode='reflect', cval=0.0, truncate=4.0,nanout=1):
#Gaussian filter with igonoring 'nan'
#https://stackoverflow.com/questions/18697532/gaussian-filtering-a-image-with-nan-in-python
nans = U!=U # positions of nan: nan is not equal to nan
V=U.copy()
V[nans]=0 # replace 'nan' by 'zero'
VV=ndimage.gaussian_filter(V, sigma, order=0, output=None, mode='reflect', cval=0.0, truncate=4.0)
W=0*U.copy()+1
W[nans]=0 # label 'nan' and values with '0' and '1' respectively
WW=ndimage.gaussian_filter(W, sigma, order=0, output=None, mode='reflect', cval=0.0, truncate=4.0)
output = VV/WW
if nanout:
output[nans] = np.nan
return output
# Test binnging2D()
N,M = 10,6
data = np.reshape(np.arange(N*M),(N,M))
x, y = np.arange(N),np.arange(M)
print(x, y)
print(data)
binning2D(x,y, data, [11,6])
# +
a = [[1,2],[2,[4,3]]]; b = [2,3]; c = [4,5]; d = [[1,2],[2,3]];
e = [np.array([1,2]),np.array([2,3,5])]
#print np.concatenate([c, b])
#print a+b,np.concatenate(a+b)
import itertools
list(itertools.chain(*d))
print(list(itertools.chain(*a)))
print(np.ravel(d), np.concatenate(d))
print(np.concatenate(e),np.ravel(e),np.hstack(e))
#np.array(a)
#print(sum(a,[]))
# +
# Test load and plot
fpath = r'D:\5_Neutron Scattering\1_US ARCS_oct2014\data_2d_for_plot/'
fname = r'nzo_50mev_5k.spe'
filename = os.path.join(fpath,fname)
qs, es, data, error = read_spe(filename)
print qs.shape,es.shape, data.shape, error.shape
#data = np.where(data==-1.00, np.nan, data)
xbins,ybins = 198, 138
#qs, es, data = binning2Dloop(qs, es, data, xbins,ybins)
qs, es, data = binning2D(qs, es, data, [xbins,ybins])
#print qs, es, data
X, Y = np.meshgrid(qs,es)
# -
fig = plt.figure(figsize=(6,4))
# pcolormesh actually draws individual rectangles which contains white lines
cmap = plt.cm.RdBu_r
#cmap = plt.cm.jet
cmap.set_bad('w',1.)
Zm = np.ma.masked_where(data==-1.00,data)# mask Nan values then plot in white color
pcol = plt.pcolormesh(X,Y, Zm.T,vmin=0,vmax=0.00025, cmap=cmap,linewidth=0,rasterized=True,shading='gouraud')# '_r' is reversed colormap
pcol.set_edgecolor('face') # remove the white lines in the plot
plt.show()
# +
# Get the data file path
fpath = r'D:\5_Neutron Scattering\1_US ARCS_oct2014\data_2d_for_plot/'
fname = r'*.spe'
ii=0 #index for different files
fnames = []
for file in os.listdir(fpath):
if fnmatch.fnmatch(file, fname):
print(file)
fnames.append(os.path.join(fpath,file))
print(fnames[0])
# +
# Load and plot
labels = ['150 meV', '400 meV', '50 meV']
x_lims = np.array([[0,16], [0,26], [0,9]])
y_lims = np.array([[-10,130],[-20,350],[-3,45]])
v_maxs = [1.,0.05,3.0]
texts = [r'La$_2$Zr$_2$O$_7$',r'La$_2$Zr$_2$O$_7$',r'La$_2$Zr$_2$O$_7$',
r'Nd$_2$Zr$_2$O$_7$',r'Nd$_2$Zr$_2$O$_7$',r'Nd$_2$Zr$_2$O$_7$']
cmap = plt.cm.RdBu_r
cmap = plt.cm.jet
#cmap.set_bad('w',1.)
nb_of_files = 6
smooth = 0
for i in np.arange(0,6,1):
print(fnames[i])
idx = np.remainder(i,3)
qs, es, data, error = read_spe(fnames[i])
X, Y = np.meshgrid(qs,es)
#data_view = np.where(data==-1.00,np.nan, data)# Set the empty point to nan for smoothing
#Z,s,exitflag,Wtot = smoothn(data_view,s=smooth) # But smooth does not fill the gap! So export good data!!
Zm = np.ma.masked_where(data==-1.00,data)# mask Nan values then plot in white color
fig = plt.figure(figsize=(6,4))
pcol = plt.pcolormesh(X,Y, Zm.T*10000,vmin=0,vmax=v_maxs[idx], cmap=cmap,linewidth=0,rasterized=True,shading='gouraud')# '_r' is reversed colormap
pcol.set_edgecolor('face') # remove the white lines in the plot
plt.text(0.77, 0.9, r'$E_\mathrm{i}=$'+'\n'+labels[idx] ,size=15,color='black', ha='left', va='center',transform=plt.gca().transAxes,
backgroundcolor='white',bbox=dict(facecolor='white', alpha=0, edgecolor='white', boxstyle='round'))
plt.text(0.14, 0.9, texts[i] ,size=15,color='black', ha='center', va='center',transform=plt.gca().transAxes,
backgroundcolor='white',bbox=dict(facecolor='white', alpha=0, edgecolor='white', boxstyle='round'))
cb = plt.colorbar(aspect=20,pad=0.05,orientation="vertical") # label='Intensity', ticks=range(0,100)
plt.minorticks_on()
plt.xticks( color='k', size=14)
plt.yticks( color='k', size=14)
plt.xlim(x_lims[idx,:])
plt.ylim(y_lims[idx,:])
plt.xlabel(r'$Q\ (\mathrm{\AA^{-1}})$',size=14)
plt.ylabel(r'$E$ (meV)',size=14)
#fig.savefig(fnames[i].replace("spe", "pdf"), bbox_inches="tight",verbose=True)
plt.show()
# -
# # Load Sm2Hf2O7 Merlin data exported form Matlab Mslice (.mat files)
# +
# Get the data file path
fpath = r'D:\5_Neutron Scattering\7_Merlin_Mar_2016_SmZrO\SmHfO_analysis/'
fname = r'*.mat'
ii=0 #index for different files
fnames = []
for file in os.listdir(fpath):
if fnmatch.fnmatch(file, fname):
print(file)
fnames.append(os.path.join(fpath,file))
print(fnames[0])
# +
# Load
labels = ['241 meV', '241 meV', '50 meV']
x_lims = np.array([[0,20], [0,20]])
y_lims = np.array([[-10,200],[-10,200]])
v_maxs = [10, 6]
texts0 = ['(b)','(a)']
texts1 = [r'La$_2$Hf$_2$O$_7$',r'Sm$_2$Hf$_2$O$_7$']
cmap = plt.cm.RdBu_r
cmap = plt.cm.PiYG_r
cmap = plt.cm.jet
cmap.set_bad('w',0.)
latexify()
nb_of_files = 2
smooth = 1
for i in np.arange(0,nb_of_files,1):
print(fnames[i])
idx = np.remainder(i,3)
data = scipy.io.loadmat(fnames[i])
X, Y, Z = data['X'], data['Y'], data['Z']
#data_view = np.where(data==-1.00,np.nan, data)# Set the empty point to nan for smoothing
#Z,s,exitflag,Wtot = smoothn(Z,s=smooth) # But smooth does not fill the gap! So export good data!!
#Z = ndimage.gaussian_filter(Z, [1,1], order=0, mode='nearest', cval=0.0, truncate=4.0)
#print(Z)
Z = myGfilter(Z, [2,2],)
#Zm = np.ma.masked_where(data==-1.00,data)# mask Nan values then plot in white color
fig = plt.figure(figsize=(6,4))
#pcol = plt.pcolormesh(X,Y, Z*1,vmin=0,vmax=v_maxs[idx], cmap=cmap,linewidth=0,rasterized=True,shading='gouraud')# '_r' is reversed colormap
pcol = plt.pcolor(X,Y, Z*1,vmin=0,vmax=v_maxs[idx], cmap=cmap,linewidth=0,rasterized=True)# here we used pcolor to avoid whitelines
pcol.set_edgecolor('face') # remove the white lines in the plot
# plt.text(0.77, 0.92, r'$E_\mathrm{i}=$'+'\n'+labels[idx] ,size=15,color='black', ha='left', va='center',transform=plt.gca().transAxes,
# backgroundcolor='white',bbox=dict(facecolor='white', alpha=0, edgecolor='white', boxstyle='round'))
# plt.text(0.14, 0.92, texts1[i] ,size=15,color='black', ha='center', va='center',transform=plt.gca().transAxes,
# backgroundcolor='white',bbox=dict(facecolor='white', alpha=0, edgecolor='white', boxstyle='round'))
plt.text(0.04, 0.92, texts0[i] ,size=16,color='black', ha='left', va='center',transform=plt.gca().transAxes,
backgroundcolor='white',bbox=dict(facecolor='white', alpha=0, edgecolor='white', boxstyle='round'))
plt.text(0.88, 0.92, texts1[i] ,size=13,color='black', ha='center', va='center',transform=plt.gca().transAxes,
backgroundcolor='white',bbox=dict(facecolor='white', alpha=0, edgecolor='white', boxstyle='round'))
if i==1:
plt.arrow(2,128,0.5,0,width=0.5,head_width=4,head_length=0.5,facecolor='k')
plt.arrow(3,156,0.5,0,width=0.5,head_width=4,head_length=0.5,facecolor='k')
plt.arrow(3.3,166,0.5,0,width=0.5,head_width=4,head_length=0.5,facecolor='k')
plt.arrow(4,183,0.5,0,width=0.5,head_width=4,head_length=0.5,facecolor='k')
cb = plt.colorbar(aspect=20,pad=0.05,orientation="vertical")#, ticks=range(0,100)
cb.ax.set_ylabel('Intensity (a.u.)',fontsize=15)
if i==0:
cb.ax.set_ylabel('Intensity (a.u.)',fontsize=15,labelpad=-5)# because '10' takes more space
cb.ax.tick_params(labelsize=15)
plt.minorticks_on()
plt.xticks(color='k', size=14)
plt.yticks(color='k', size=14)
plt.xlim(x_lims[idx,:])
plt.ylim(y_lims[idx,:])
plt.xlabel(r'Q $(\mathrm{\AA^{-1}})$',size=1)
plt.ylabel(r'E (meV)',size=15)
fig.savefig(fnames[i].replace("mat", "pdf"), bbox_inches="tight",pad_inches=0.01,verbose=True)
plt.show()
# -
# ## Below is trying to use RE to read SPE
# but the last data set cannot use a simple pattern to find
#
# Winexpect_call_spectra.py also used re.
#
# re tricks:
# 1. The special characters you should use \ in prefix: \\ for \, \! for !, \[,\( ...
# 2. The part in brackets of the pattern will be retured
# 3. Multiple-pattern matching (with | without spaces around it) give list and each element containning has the len of nb of patterns
# 4. "(B\([246],[036]\)[\ *c]?)\s+=\s+(-?\d+\.\d+E[-+]\d+)" matchs B20 2E-2
# Test finding words between 'start' and 'end'
START = 'i'
END = 'e'
test = "i0 1\n2 3\ne1 1\n1 1\ni2 2\n 3 3 \ne"
m = re.compile(r'%s.*?%s' % (START,END),re.S)
m1 = m.findall(test)
m1
# +
# Data: load in as a string and find all
fpath = r'D:\5_Neutron Scattering\1_US ARCS_oct2014\data_2d_for_plot/'
fname = r'nzo_50mev_5k1.spe'
fname = os.path.join(fpath,fname)
f = open(fname, 'r')
ftext = f.read()
f.close()
#print ftext
# Prapre the patterns (it is Tuple for formating latter)
StartEnd = ('### \|Q\| \(\\\\AA\!U\-1\!N\)\n', '### E \(meV\)\n', # for Qs: take care of the special chars with \
'### E \(meV\)\n', '### Intensity \(arb. units\)\n', # for Es
'### Intensity \(arb. units\)\n', '### Errors', # for intensity
'### Errors\n', '### Intensity') # for error
# Multiline (re.DOTALL!) match using | (no space around it!)
#m = re.compile(r'%s(.*?)%s|%s(.*?)%s|%s(.*?)%s|%s(.*?)%s' % StartEnd, re.DOTALL) # StartEnd must be a tuple not list!!!
#m1 = m.findall(ftext)
# Failed: above try to find all the data in one search but failed (not find Es and Errors, and retures tuples)
# Below find them sparately failed: the last block of error not find due to not match
StartEnd0 = ('### \|Q\| \(\\\\AA\!U\-1\!N\)\n', '0.00000\n### E \(meV\)\n') # for Qs: take care of the special chars with \
StartEnd1 = ('### E \(meV\)\n', '0.00000\n### Intensity \(arb. units\)\n') # for Es
StartEnd2 = ('### Intensity \(arb. units\)\n', '### Errors')
StartEnd3 = ('### Errors\n', '### Intensity') # the last bolck of error can not be found with it!
#StartEnd3 = ('### Errors\n', '\Z') # \A and \Z are the beginning and end of the string
m = re.compile(r'%s(.*?)%s' % StartEnd3, re.DOTALL)
m1 = m.findall(ftext)
m1
# m2 = [item.rstrip('\n').split() for item in m1]
#np.array(m1.rstrip('\n').split(),dtype='f')
#_.shape
# -
| scientific_data_analysis/crystal_field/Read_and_plot_spe_mat_CEF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Motifs analysis - Part 1: Motifs extraction
#
# Notebook to perform the discriminative motifs analysis. It requires a trained model but it is an independant analysis from the analysis of feature space and from the prototypes analysis.
#
# Motifs are extracted on the base of Class-Activation Maps (CAMs) which display the saliency of a class in a given input according to a model. CAMs towards any class can be computed regardless of the actual class of the input. This means that one can look for discriminative motifs of class B in an input of class A. However, for the sake of motif extraction, we don't use this feature of CAMs. Instead we produce CAMs towards the actual class of the input.
#
# The motif extraction procedure is as follow:
# 1. Select trajectories from which to extract motifs.
# 2. Compute CAM for each trajectory (saliency towards its own class).
# 3. Binarize each time point into 'relevant' and 'non-relevant' to recognize input class.
# 4. Optional but recommended, extend the 'relevant' regions to capture more context around the motifs and connect smaller adjacents motifs into a bigger one. Also filter for motif length.
# 5. Extract the longest 'relevant' stretches of time-points. These are the final motifs.
#
# In order to visualize these motifs, we propose to cluster them afterwards as follow:
# 1. Build a distance matrix between the motifs with dynamic time warping (dtw)
# 2. Cluster with hierarchical clustering.
# 3. Visualize dynamics captured by each cluster.
#
# This clustering can be run in 2 modes: either patterns from every class are pooled together, either a separate clustering is run indepently for each class. In the 1st case, this will reflect the diversity of patterns at the dataset level and can reveal dynamics overlap between classes. In the second case, the emphasis is put on the diversity of dynamics induced by each class.
#
#
# This notebook covers only the motif extraction part. It ends with the export of the motifs to a csv file. Go to the next one for computing DTW and clustering!
#
#
# ## Import libraries
# +
# Standard libraries
import torch
from torch.utils.data import DataLoader
from torchvision import transforms
import numpy as np
import pandas as pd
from skimage.filters import threshold_li, threshold_mean
import os
from itertools import chain
from tqdm import tqdm
import sys
# Custom functions/classes
path_to_module = '../source' # Path where all the .py files are, relative to the notebook folder
sys.path.append(path_to_module)
from load_data import DataProcesser
from results_model import top_confidence_perclass, least_correlated_set
from pattern_utils import extend_segments, create_cam, longest_segments, extract_pattern
from class_dataset import myDataset, ToTensor, RandomCrop
# For reproducibility
myseed = 7
torch.manual_seed(myseed)
torch.cuda.manual_seed(myseed)
np.random.seed(myseed)
cuda_available = torch.cuda.is_available()
# -
# ## Parameters
#
# Parameters for the motifs extraction:
# - selected_set: str one of ['all', 'training', 'validation', 'test'], from which set of trajectories should motifs be extracted? For this purprose, extracting from training data also makes sense.
# - n_series_perclass: int, maximum number of series, per class, on which motif extraction is attempted.
# - n_pattern_perseries: int, maximum number of motifs to extract out of a single trajectory.
# - mode_series_selection: str one of ['top_confidence', 'least_correlated']. Mode to select the trajectories from which to extract the motifs (see Prototype analysis). If top confidence, the motifs might be heavily biased towards a representative subpopulation of the class. Hence, the output might not reflect the whole diversity of motifs induced by the class.
# - extend_patt: int, by how many points to extend motifs? After binarization into 'relevant' and 'non-relevant time points', the motifs are usually fragmented because a few points in their middle are improperly classified as 'non-relevant'. This parameter allows to extend each fragment by a number of time points (in both time directions) before extracting the actual patterns.
# - min_len_patt/max_len_patt: int, set minimum/maximum size of a motif. **/!\ The size is given in number of time-points. This means that if the input has more than one channel, the actual length of the motifs will be divided across them.** For example, a motif that spans over 2 channels for 10 time points will be considered of length 20.
#
# Parameters for the groups of motifs:
# - export_perClass: bool, whether to run the motif clustering class per class.
# - export_allPooled: bool, whether to pool all motifs across classes for clustering.
# +
selected_set = 'all'
n_series_perclass = 50
n_pattern_perseries = 1
mode_series_selection = 'top_confidence'
# mode_series_selection = 'least_correlated'
thresh_confidence = 0.5 # used in least_correlated mode to choose set of series with minimal classification confidence
extend_patt = 0
min_len_patt = 0
max_len_patt = 200 # length to divide by nchannel
export_perClass = False
export_allPooled = True
assert selected_set in ['all', 'training', 'validation', 'test']
assert mode_series_selection in ['top_confidence', 'least_correlated']
# -
# ## Load model and data
#
# - Pay attention to the order of 'meas_var', should be the same as for training the model!
# - Pay attention to trajectories preprocessing.
# - Set batch_size as high as memory allows for speed up.
# +
data_file = '../sample_data/Synthetic_Univariate.zip'
model_file = 'models/FRST/sampleModel_Synthetic_Univariate.pytorch'
# data_file = '../sample_data/GrowthFactor_ErkAkt_Bivariate.zip'
# model_file = 'models/ERK_AKT/sampleModel_GrowthFactor_ErkAkt_Bivariate.pytorch'
out_dir = 'auto' # If 'auto' will automatically create a directory to save motifs tables
meas_var = None # Set to None for auto detection
start_time = None # Set to None for auto detection
end_time = None # Set to None for auto detection
batch_size = 32 # Set as high as memory allows for speed up
is_cuda = torch.cuda.is_available()
device = torch.device('cuda' if is_cuda else 'cpu')
model = torch.load(model_file) if cuda_available else torch.load(model_file, map_location='cpu')
model.eval()
model.double()
model.batch_size = batch_size
model = model.to(device)
# -
# Pay attention that **data.process() is already centering the data**, so don't do a second time when loading the data in the DataLoader. The **random crop** should be performed before passing the trajectories to the model to ensure that the same crop is used as input and for extracting the patterns.
# +
# Transformations to perform when loading data into the model
ls_transforms = transforms.Compose([RandomCrop(output_size=model.length, ignore_na_tails=True),
ToTensor()])
# Loading and PREPROCESSING
data = DataProcesser(data_file)
meas_var = data.detect_groups_times()['groups'] if meas_var is None else meas_var
start_time = data.detect_groups_times()['times'][0] if start_time is None else start_time
end_time = data.detect_groups_times()['times'][1] if end_time is None else end_time
# Path where to export tables with motifs
if out_dir == 'auto':
out_dir = 'output/' + '_'.join(meas_var) + '/local_motifs/'
if not os.path.exists(out_dir):
os.makedirs(out_dir)
data.subset(sel_groups=meas_var, start_time=start_time, end_time=end_time)
cols_to_check=data.dataset.columns.values[data.dataset.columns.str.startswith('FGF')]
cols_dict={k:'float64' for k in cols_to_check}
data.dataset=data.dataset.astype(cols_dict)
data.get_stats()
data.process(method='center_train', independent_groups=True) # do here and not in loader so can use in df
data.crop_random(model.length, ignore_na_tails=True)
data.split_sets(which='dataset')
classes = tuple(data.classes[data.col_classname])
dict_classes = data.classes[data.col_classname]
# Random crop before to keep the same in df as the ones passed in the model
if selected_set == 'validation':
selected_data = myDataset(dataset=data.validation_set, transform=ls_transforms)
df = data.validation_set
elif selected_set == 'training':
selected_data = myDataset(dataset=data.train_set, transform=ls_transforms)
df = data.train_set
elif selected_set == 'test':
selected_data = myDataset(dataset=data.test_set, transform=ls_transforms)
df = data.train_set
elif selected_set == 'all':
try:
selected_data = myDataset(dataset=data.dataset_cropped, transform=ls_transforms)
df = data.dataset_cropped
except:
selected_data = myDataset(dataset=data.dataset, transform=ls_transforms)
df = data.dataset
if batch_size > len(selected_data):
raise ValueError('Batch size ({}) must be smaller than the number of trajectories in the selected set ({}).'.format(batch_size, len(selected_data)))
data_loader = DataLoader(dataset=selected_data,
batch_size=batch_size,
shuffle=True,
num_workers=4)
# Dataframe used for retrieving trajectories. wide_to_long() instead of melt() because can do melting per group of columns
df = pd.wide_to_long(df, stubnames=meas_var, i=[data.col_id, data.col_class], j='Time', sep='_', suffix='\d+')
df = df.reset_index() # wide_to_long creates a multi-level Index, reset index to retrieve indexes in columns
df.rename(columns={data.col_id: 'ID', data.col_class: 'Class'}, inplace=True)
df['ID'] = df['ID'].astype('U32')
del data # free memory
# -
# ## Select trajectories from which to extract patterns
# +
if mode_series_selection == 'least_correlated':
set_trajectories = least_correlated_set(model, data_loader, threshold_confidence=thresh_confidence, device=device,
n=n_series_perclass, labels_classes=dict_classes)
elif mode_series_selection == 'top_confidence':
set_trajectories = top_confidence_perclass(model, data_loader, device=device, n=n_series_perclass,
labels_classes=dict_classes)
# free some memory by keeping only relevant series
selected_trajectories = set_trajectories['ID']
df = df[df['ID'].isin(selected_trajectories)]
# Make sure that class is an integer (especially when 0 or 1, could be read as boolean)
df['Class'] = df['Class'].astype('int32')
# -
# ## Extract patterns
#
# ### Extract, extend and filter patterns.
#
# Outputs a report of how many trajectories were filtered out by size.
# +
# Initialize dict to store the patterns and set progress bar
store_patts = {i:[] for i in classes}
model.batch_size = 1 # Leave it to 1!
report_filter = {'Total number of patterns': 0,
'Number of patterns above maximum length': 0,
'Number of patterns below minimum length': 0}
pbar = tqdm(total=len(selected_trajectories))
for id_trajectory in selected_trajectories:
# Read and format the trajectories to numpy
series_numpy = np.array(df.loc[df['ID'] == id_trajectory][meas_var]).astype('float').squeeze()
# Row: measurement; Col: time
if len(meas_var) >= 2:
series_numpy = series_numpy.transpose()
series_tensor = torch.tensor(series_numpy)
class_trajectory = df.loc[df['ID']==id_trajectory]['Class'].iloc[0] # repeated value through all series
class_label = classes[class_trajectory]
# Create and process the CAM for the trajectory
cam = create_cam(model, array_series=series_tensor, feature_layer='features',
device=device, clip=0, target_class=class_trajectory)
thresh = threshold_li(cam)
bincam = np.where(cam >= thresh, 1, 0)
bincam_ext = extend_segments(array=bincam, max_ext=extend_patt)
patterns = longest_segments(array=bincam_ext, k=n_pattern_perseries)
# Filter short/long patterns
report_filter['Total number of patterns'] += len(patterns)
report_filter['Number of patterns above maximum length'] += len([k for k in patterns.keys() if patterns[k] > max_len_patt])
report_filter['Number of patterns below minimum length'] += len([k for k in patterns.keys() if patterns[k] < min_len_patt])
patterns = {k: patterns[k] for k in patterns.keys() if (patterns[k] >= min_len_patt and
patterns[k] <= max_len_patt)}
if len(patterns) > 0:
for pattern_position in list(patterns.keys()):
store_patts[class_label].append(extract_pattern(series_numpy, pattern_position, NA_fill=False))
pbar.update(1)
print(report_filter)
# -
# ### Dump patterns into csv
# +
if export_allPooled:
concat_patts_allPooled = np.full((sum(map(len, store_patts.values())), len(meas_var) * max_len_patt), np.nan)
irow = 0
for classe in classes:
concat_patts = np.full((len(store_patts[classe]), len(meas_var) * max_len_patt), np.nan)
for i, patt in enumerate(store_patts[classe]):
if len(meas_var) == 1:
len_patt = len(patt)
concat_patts[i, 0:len_patt] = patt
if len(meas_var) >= 2:
len_patt = patt.shape[1]
for j in range(len(meas_var)):
offset = j*max_len_patt
concat_patts[i, (0+offset):(len_patt+offset)] = patt[j, :]
if len(meas_var) == 1:
headers = ','.join([meas_var[0] + '_' + str(k) for k in range(max_len_patt)])
fout_patt = out_dir + 'motif_{}.csv.gz'.format(classe)
if export_perClass:
np.savetxt(fout_patt, concat_patts,
delimiter=',', header=headers, comments='')
elif len(meas_var) >= 2:
headers = ','.join([meas + '_' + str(k) for meas in meas_var for k in range(max_len_patt)])
fout_patt = out_dir + 'motif_{}.csv.gz'.format(classe)
if export_perClass:
np.savetxt(fout_patt, concat_patts,
delimiter=',', header=headers, comments='')
if export_allPooled:
concat_patts_allPooled[irow:(irow+concat_patts.shape[0]), :] = concat_patts
irow += concat_patts.shape[0]
if export_allPooled:
concat_patts_allPooled = pd.DataFrame(concat_patts_allPooled)
concat_patts_allPooled.columns = headers.split(',')
pattID_col = [[classe] * len(store_patts[classe]) for classe in classes]
concat_patts_allPooled['pattID'] = [j+'_'+str(i) for i,j in enumerate(list(chain.from_iterable(pattID_col)))]
concat_patts_allPooled.set_index('pattID', inplace = True)
fout_patt = out_dir + 'motif_allPooled.csv.gz'.format(classe)
concat_patts_allPooled.to_csv(fout_patt, header=True, index=True, compression='gzip')
# -
# ### Build distance matrix between patterns with DTW
#
# This is done in R with the implementation of the *parallelDist* package. It is very efficient and has support for multivariate cases.
#
# Check next notebook.
| Notebooks/2_Motifs_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction To Ansible
#
# Ansible is a tool for automating system administration and operation tasks. It uses SSH to talk to a collection of servers and do tasks such as: configure aspects of the server, install or upgrade software, and pretty much any other task you could do at the command line.
#
# ## Basic Notions
#
# Ansible does not run any agents on the remote servers. Instead, it connects over SSH to each server and runs a (typically small) program to accomplish each task. Users write Ansible scripts to perform multiple tasks on a given set of servers. Users then execute these scripts from any machine that has access to Ansible and
#
# Make sure Ansible is installed on your VM:
# ```
# $ ansible --version
#
# ```
# If not, install it with:
# ```
# $ sudo apt-get install ansible
# ```
#
# ### Hosts Files
# Ansible has the notion of a hosts or inventory file. This is a plain text file that describes all of the servers you want Ansible to interact with. Each host listed in the file should provide the basic SSH connectivity information for Ansible to use to perform tasks on them. Hosts can be also be collected into groups.
#
# Here is an example hosts file with three hosts, two in the "web" group and one in the "databases" group.
#
# ```
# [web]
# web_1 ansible_ssh_host=192.168.127.12 ansible_ssh_private_key_file=~/web.key ansible_ssh_user=ubuntu
# web_2 ansible_ssh_host=192.168.3.11 ansible_ssh_private_key_file=~/web.key ansible_ssh_user=ubuntu
#
# [databases]
# mysql ansible_ssh_host=172.16.17.32 ansible_ssh_private_key_file=~/db.key ansible_ssh_user=centos
#
# ```
#
# Note that each line corresponding to a host begins with a name, which can be any identifier we want, and then provides two variables `ansible_ssh_host` and `ansible_ssh_private_key_file`.
#
# ```
# Exercise. Create a hosts file for your jupyter host. Don't worry about putting it in a group yet.
# ```
#
# Once we have a hosts file, we can already run basic commands against the host(s). To run a command on all hosts in a hosts file, use the following:
#
# ```
# $ ansible all -i <hosts_file> -a '<ommand>'
# ```
#
# For example, let's check the uptime of the server by running the "uptime" command:
# ```
# $ ansible all -i hosts -a 'uptime'
# ```
#
# ### Modules and Tasks
#
# Part of Ansible's power comes from its rich library of modules. Modules are configurable programs that can be used so accomplish specific tasks. For example, we have the "command" module for simply running arbitrary commands. But there are many other modules for describing tasks at a higher level. For example, Ansible provides the "copy" module for ensuring files from the Ansible host machine are present on the remote server.
#
# We can specify a module to use by passing the name of the module to the `-m` flag and then providing any required parameters of the module. For instance, the copy module requires `src` and `dest` parameters. Let's use the copy module to copy a file called "test.txt" in the current directory to `/root` in the remote server. We'll name it "remote.txt" on the remote:
# ```
# $ ansible all -m copy -a "src=test.txt dest=/root/remote.txt"
# 10.10.100.7 | SUCCESS => {
# "changed": true,
# "checksum": "da39a3ee5e6b4b0d3255bfef95601890afd80709",
# "dest": "/root/remote.txt",
# "gid": 0,
# "group": "root",
# "md5sum": "d41d8cd98f00b204e9800998ecf8427e",
# "mode": "0644",
# "owner": "root",
# "size": 0,
# "src": "/root/.ansible/tmp/ansible-tmp-1500597877.29-186213310577275/source",
# "state": "file",
# "uid": 0
# }
# ```
#
# After running that command we see some output (note the color). In particular, we see a `"changed": true` . Ansible detected that we it needed to actually change the host to ensure that the file was there. Let's try running the command again and see what we get:
# ```
# $ ansible all -m copy -a "src=test.txt dest=/root/remote.txt"
# 10.10.100.7 | SUCCESS => {
# "changed": false,
# "checksum": "da39a3ee5e6b4b0d3255bfef95601890afd80709",
# "dest": "/root/remote.txt",
# "gid": 0,
# "group": "root",
# "mode": "0644",
# "owner": "root",
# "path": "/root/remote.txt",
# "size": 0,
# "state": "file",
# "uid": 0
# }
# ```
# This time, the output is green and Ansible indicates to us that it didn't change anything on the remote server. Indeed, Ansible's copy module first checks if there is anything to do, and if the file already there, it doesn't do anything. This is a key part of Ansible's power which we explore in the next section. But first, let's change the contents of our test.txt and then re-run our command.
#
# ```
# # open test.txt in vi and change the contents:
# $ vi test.txt
# # . . .
# # now run the command:
# $ ansible all -m copy -a "src=test.txt dest=/root/remote.txt"
# 10.10.100.9 | SUCCESS => {
# "changed": true,
# "checksum": "4e1243bd22c66e76c2ba9eddc1f91394e57f9f83",
# "dest": "/root/remote.txt",
# "gid": 0,
# "group": "root",
# "md5sum": "d8e8fca2dc0f896fd7cb4cb0031ba249",
# "mode": "0644",
# "owner": "root",
# "size": 5,
# "src": "/root/.ansible/tmp/ansible-tmp-1500598336.43-267287598231809/source",
# "state": "file",
# "uid": 0
# }
# ```
#
# ### Idempotence
# Ansible's modules provide us with the power to ensure a given Ansible script we write is `idempotent`. A given task is said to be idempotent if performing the task once produces the same result as performing it repeadly, assuming no other intervening actions.
#
# Idempotence is very important for ensuring re-runability of your provisioning scripts. Suppose you wanted to automate maintenance of a database sever. You might have the following high level tasks:
#
# ```
# 1. Create a linux user account for the MySQL service to run under
# 2. Install the MySQL server package
# 3. Start the mysql daemon
# 4. Create the database
# 5. Run the latest migrations on the database to install the schema
# ```
#
# The problem is that, if we just use basic commands such as `useradd` and `mysql create database`, the script will run on a brand new server but it will fail every subsequent time. With idempotent tasks we can avoid this issue.
#
# ```
# 1. Ensure a linux user account exists for the MySQL service to run under
# 2. Ensure the MySQL server package is installed
# 3. Ensure the mysql daemon is running
# 4. Ensure the database has been created
# 5. Ensure the database has the latest migrations on the database to install the schema.
# ```
#
#
#
| notebooks/Introduction To Ansible.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:cv_py27_2]
# language: python
# name: conda-env-cv_py27_2-py
# ---
# NOTE: this should eventually be converted to a Python script once the algorithm to determin the sign point is finalised
import pandas as pd
import numpy as np
pd.set_option("display.notebook_repr_html", False)
df = pd.read_csv("../data/processed/170329/predictions/df_garmin_joined.csv")
df.shape
# initialise cluster related columns
df["cluster_id"] = -1 # means that this row has not been id'ed as a sign by model
df["is_cluster_main"] = 0 # means that this is the closest point to the centre
df["num_in_cluster"] = -1 # only for rows where is_cluster_main = 1
def get_main_cluster_point(cluster_members, df):
#find member closest to middle
middle = 1920/2 #middle pixel in x direction
closest_id = cluster_members[0]
closest_distance = np.abs( df.loc[ df["id"]==closest_id, "x"].values[0] - middle )
for id in cluster_members[1:]:
d = np.abs( df.loc[ df["id"]==id, "x" ].values[0] - middle)
#print "distance = %d" % d
if d < closest_distance:
closest_id = id
closest_distance = d
#print "closest distance = %d" % closest_distance
return closest_id
def lookahead(iterable):
"""Pass through all values from the given iterable, augmented by the
information if there are more values to come after the current one
(True), or if it is the last value (False).
"""
# Get an iterator and pull the first value.
it = iter(iterable)
last = next(it)
# Run the iterator to exhaustion (starting from the second value).
for val in it:
# Report the *previous* value (more to come).
yield last, True
last = val
# Report the last value.
yield last, False
import pprint
cluster_id = 1
am_mid_cluster = False
frames_between_clusters = 24 #threshold of empty frames before I start a new cluster
n_to_process = 1e6 #set to large number to process all frames
n_processed = 0
#base_frame_i = 18261
#base_frame_f = 19000
#for _, row in df[ df["base_frame"].between(base_frame_i, base_frame_f) ].iterrows():
for ((_, row), has_more) in lookahead(df.iterrows()):
#print "base_frame = {}".format(row["base_frame"])
base_frame = row["base_frame"]
is_sign = row["accum"] > 0.4 and row["score"] > 0
#is_sign = row["ymain"] == 1
#end the cluster if no sign seen for frame_between_clusters frames
#or sign in frame but this is last frame
end_cluster = (~is_sign and (base_frame-latest_cluster_frame > frames_between_clusters))\
or (is_sign and not has_more)
if not am_mid_cluster and is_sign:
#start of new cluster
print "new cluster starting at frame %d" % row["base_frame"]
latest_cluster_frame = base_frame
am_mid_cluster = True
cluster_members = [row["id"]]
elif not am_mid_cluster and not is_sign:
#not in a cluster and no sign in frame. Keep moving
pass
elif am_mid_cluster:
if is_sign:
print "found sign"
#sign in frame. Add to cluster members
cluster_members.append(row["id"])
latest_cluster_frame = base_frame
if end_cluster:
# This is the end of the cluster
print "end of cluster. cluster_members"
pprint.pprint(cluster_members)
df.loc[ df["id"].isin(cluster_members), "cluster_id" ] = cluster_id
cluster_id += 1
cluster_point_id = get_main_cluster_point(cluster_members, df)
df.loc[ df["id"]==cluster_point_id, "is_cluster_main"] = 1
df.loc[ df["id"]==cluster_point_id, "num_in_cluster"] = len(cluster_members)
#reset cluster variables
am_mid_cluster = False
else:
print "WTF"
print am_mid_cluster
print is_sign
n_processed += 1
if n_processed % 1000 == 0:
print n_processed
if n_processed > n_to_process:
break
df["cluster_id"].unique()
df.columns
df.head()
df["num_in_cluster"].unique()
df.to_csv("../data/processed/170329/predictions/df_garmin_joined_clustered.csv")
| notebooks/8.1-mmh-garmin_sign_clusters_to_point.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pierredumontel/Portfolio_management/blob/main/Notebook/Portfolio_management.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="aovY8_Egrhhe"
# # Portfolio Management
#
# Group :
# <NAME>,
# <NAME>,
# <NAME>,
# <NAME>,
# <NAME>,
# <NAME>
# + [markdown] id="ihupX3pbBGgP"
# #Installation des packages necessaires.
# + colab={"base_uri": "https://localhost:8080/"} id="Uv0RVIL2jeS-" outputId="4d4edcd1-d7a9-4a6c-e20e-77b1177212f1"
pip uninstall --quiet matplotlib
# + id="QtYnyongBWfe"
pip install --quiet requests
# + id="uQ3X4lK2CnO-" colab={"base_uri": "https://localhost:8080/"} outputId="be7952da-2b88-41eb-dbea-21bf520591f8"
pip install --quiet imgaug
# + colab={"base_uri": "https://localhost:8080/"} id="yAommv2cjh2k" outputId="866de143-68b7-46dc-ddbf-afc1c187b499"
pip install --quiet matplotlib matplotlib==3.1.3
# + id="Q5grKuS5YQml"
pip install --quiet yfinance
# + id="dhcUfphL2_V-" colab={"base_uri": "https://localhost:8080/"} outputId="801bb288-eb18-4e5f-985b-13feb3f66e23"
# !pip install --quiet riskfolio-lib
# + id="TS9qSH8ECYR5"
import numpy as np
import pandas as pd
import yfinance as yf
import matplotlib.pyplot as plt
import scipy.optimize as optimization
# + [markdown] id="lYGDyT21biMi"
# # Collection des données
# + id="hip3P-nJYK9M"
# Telechargement des prix depuis yahoo finance
assets = ['BDORY','STN','MKC',
'AWK','CNI','AY','CSCO','OC','ESALY','CADNF','BXBLY','IBDRY','VWDRY',
'VWSYF','CRTSF','SMAWF','TT','AKZOY','IGIFF','HPE','ACXIF','ABB','NVZMY',
'JCI','AOMFF','ADSK','TCLAF','BNPQY','BMO','BLL','ALIZF','HPQ','CMA','TU','DASTY','ISNPY','SMSMY',
'INTC', 'ACN','SNYNF', 'VLEEF', 'CRZBY','CGEAF','SLF','XRX','TKPHF','AEM','ADI',
'ADDDF','PLD','LNVGF','UL','ORKLY','AZN','SHG','SAP','NRDBY','ERIC','GOOG','TECK',
'KKPNF','WDAY','TSLA','NVO','CDNAF','NVDA','^GSPC']
start_date = '2016-06-08'
end_date = '2021-12-01'
def download_data():
stock_data = {}
for stock in assets:
ticker = yf.Ticker(stock)
stock_data[stock] = ticker.history(start=start_date, end=end_date)['Close']
return pd.DataFrame(stock_data)
stock_data = download_data()
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="b_g97ROVYTow" outputId="3e3f710d-6fcd-4fff-e537-51abec6edf72"
stock_data = stock_data.drop(stock_data.index[0])
stock_data.head()
# + [markdown] id="uzPeWFDD3fIn"
# # Traitement des données <br/> <br/>
# - Calcul des rendements
# - Partition des données en ensemble d'apprentissage et de validation
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="30xj6x7DYiEa" outputId="76166686-2207-45e3-cab9-6ee2fc884344"
# Obtention des rendements à partir des prix
returns = stock_data.pct_change()
returns = returns.drop(returns.index[:2])
returns.head()
# + colab={"base_uri": "https://localhost:8080/"} id="NfKMEAEGYoWS" outputId="c1fdce12-3bf3-4678-c993-9e6b17c36a3c"
train_size_stock = int(len(stock_data) * 0.46032)
train_stock, test_stock = stock_data[0:train_size_stock], stock_data[train_size_stock:len(stock_data)]
train_size_rets = int(len(returns) * 0.46032)
train_rets, test_rets = returns[0:train_size_rets], returns[train_size_rets:len(returns)]
print('Observations totales: %d' % (len(stock_data)))
print('Observations dans le trainset : %d' % (len(train_rets)))
print('Observations dans le testset: %d' % (len(test_rets)))
# + [markdown] id="Vi68hF7cbpRE"
# #Calcul des poids des actifs <br/> <br/>
# Pour calculer les poids accordés à chaque actif, nous allons nous servir de la librairie riskfolio. Elle permet de calculer les poids via différents moyens :
# - Minimisation du risque
# - Maximisation des rendements
# - etc.
# https://riskfolio-lib.readthedocs.io/en/latest/index.html
# + [markdown] id="YNCMK0owF1qB"
# Dans la cellule de code ci-dessous, nous allons calculer le poids optimal de notre portefeuille(sur le trainset) en instaurant une à la fois une contrainte sur le nombre minimal d'actifs non significatif (actif avec un poids très négligeable 0.0x%) et sur le nombre d'actif total dans le portefeuille. Pour obtenir les poids nous allons maximiser le ratio de sharpe<br/> <br/>
#
# mesure de risque utilisée : variance <br/>
# fonction objective: ratio de sharpe <br/>
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="xi2uet7XY8Py" outputId="22241617-4a43-45bb-a25c-d9c95715338f"
import riskfolio as rp
port = rp.Portfolio(returns=train_rets)
method_mu='hist'
method_cov='hist'
port.assets_stats(method_mu=method_mu, method_cov=method_cov, d=0.94)
model='Classic'
rm = 'MV'
obj = 'Sharpe' #Fonction objective
hist = True
rf = 0
l = 0
port.card = None
w_sr= {}
data = {}
var = {}
std = {}
ret = {}
SR = {}
stats_sr = {}
for nb_stocks, port.nea in zip(range(27,34),range(27,34)):
w_sr[port.nea] = port.optimization(model=model, rm=rm, obj=obj, rf=rf, l=l, hist=hist)
data[nb_stocks] = pd.DataFrame(w_sr[port.nea])
data[nb_stocks] = data[nb_stocks].T
data[nb_stocks] = data[nb_stocks].reset_index()
data[nb_stocks] = data[nb_stocks].drop(columns='index')
data[nb_stocks] = data[nb_stocks].T
var[nb_stocks] = data[nb_stocks] * (train_rets.cov() @ data[nb_stocks]) * 252
var[nb_stocks] = var[nb_stocks].sum().to_frame().T #variance
std[nb_stocks] = np.sqrt(var[nb_stocks])
ret[nb_stocks] = train_rets.mean().to_frame().T @ data[nb_stocks] * 252
SR[nb_stocks] = (ret[nb_stocks] - 0.0176)/std[nb_stocks] #Sharpe ratio
stats_sr[nb_stocks] = pd.concat([ret[nb_stocks], std[nb_stocks], var[nb_stocks], SR[nb_stocks]], axis=0)
stats_sr[nb_stocks].index = ['Return', 'Std. Dev.', 'Variance', 'Sharpe Ratio']
#Résulats pour différents nombres d'actifs dans le portefeuille (27,28...33)
stats = pd.concat([stats_sr[27],stats_sr[28],stats_sr[29],stats_sr[30],stats_sr[31],stats_sr[32],stats_sr[33]],axis=1)
stats = stats.set_axis(['Max Sharpe 27','Max Sharpe 28', 'Max Sharpe 29', 'Max Sharpe 30','Max Sharpe 31', 'Max Sharpe 32','Max Sharpe 33'], axis=1)
stats
# + id="p73Zn2wE_eQ-"
#Création d'une fonction pour calculer le poids optimal
#On restreint les poids très négligeable à zéro. Seuil de négligeabilité 0.1% :
def calcule_portefeuille_optimal (return_data) :
port = rp.Portfolio(returns=return_data)
port.nea = 30 #On veut au minimum 30 actifs dans le portefeuille
method_mu='hist'
method_cov='hist'
port.assets_stats(method_mu=method_mu, method_cov=method_cov, d=0.94)
model='Classic'
rm = 'MV'
obj = 'Sharpe'
hist = True
rf = 0
l = 0
port.card = None # First we need to delete the cardinality constraint
opti_portfolio = port.optimization(model=model, rm=rm, obj=obj, rf=rf, l=l, hist=hist)*100 #Caclule des poids en pourcentage
opti_portfolio = opti_portfolio.reset_index().rename(columns={"index": "Actif", "weights": "Poids"})
#Contrainte pour mettre à zéro les poids très négligeable : poids < 0.1%
opti_portfolio.loc[(opti_portfolio.Poids < 0.1 ), 'Poids'] = 0
return opti_portfolio
# + id="nVGBZSXQWnrv" colab={"base_uri": "https://localhost:8080/", "height": 144} outputId="fe4addcd-f554-4cc2-c3a3-b703a593a012"
#Exemple
poids = data[30].applymap(lambda x: "{0:.4f}".format(x*100))
poids = poids.T
poids
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="vEU0HMR6cAf7" outputId="85a8d259-9506-4f88-8669-2e9e34d79fbc"
#Performance du portefeuille plus haut sur le testset : Exemple
var_t = data[30] * (test_rets.cov() @ data[30]) * 252
var_t = var_t.sum().to_frame().T
std_t = np.sqrt(var_t)
ret_t = test_rets.mean().to_frame().T @ data[30] * 252
SR_t = (ret_t - 0.0176)/std_t
stats_sr_t = pd.concat([ret_t, std_t, SR_t], axis=0)
stats_sr_tt = stats_sr_t
stats_sr_t.index = ['Rendement', 'Volatilité', 'Sharpe ratio']
stats_sr_t = stats_sr_t.T
stats_sr_t[["Rendement","Volatilité"]] = stats_sr_t[["Rendement","Volatilité"]].applymap(lambda x: "{0:.1f}%".format(x*100))
stats_sr_t[["Sharpe ratio"]] = stats_sr_t[["Sharpe ratio"]].applymap(lambda x: "{0:.2f}".format(x))
display(stats_sr_t)
# + [markdown] id="hVkEgtUWbvlt"
# # Frontière efficiente
# + colab={"base_uri": "https://localhost:8080/", "height": 357} id="7fMcz9Y0ZOOf" outputId="d6649fc8-bad9-46e7-a83a-edd6e6c34081"
port.nea =30
points = 50
frontier = port.efficient_frontier(model=model, rm=rm, points=points, rf=rf, hist=hist)
display(frontier.T.head())
# + colab={"base_uri": "https://localhost:8080/", "height": 441} id="NnDqv49BZbWA" outputId="6f57e2d6-3c6b-4efc-8f70-d70d36590965"
# Plotting the efficient frontier
import riskfolio.PlotFunctions as plf
label = 'Max Risk Adjusted Return Portfolio' # Title of plot
mu = port.mu # Expected returns
cov = port.cov # Covariance matrix
ret = port.returns # Returns of the assets
ax = plf.plot_frontier(w_frontier=frontier, mu=mu, cov=cov, returns=ret, rm=rm, rf=rf, alpha=0.05, cmap='viridis',
w=data[30], label=label, marker='*', s=16, c='r', height=6, width=10, ax=None)
# + [markdown] id="Sas-DeYrZl2_"
# #Strategie de test 1 : Long/short strategy <br/>
#
# + [markdown] id="_qpam1tJMAxH"
# - Etape 1 : traitement des données pour obtenir les signaux à chaque date et pour chaque stocks
# + id="0tapep4KZpbC"
tr_sto = stock_data
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="ayRN5_T0ZepZ" outputId="86fae257-5646-4b17-d9a2-b13ed472dddb"
SMA_50 = []
SMA_200 = []
for x in tr_sto:
SMA_50.append(tr_sto[x].rolling(window = 50, min_periods = 1).mean())
SMA_200.append(tr_sto[x].rolling(window = 200, min_periods = 1).mean())
SMA_200 = pd.DataFrame(SMA_200).T
SMA_50 = pd.DataFrame(SMA_50).T
SMA_200 = SMA_200.drop(SMA_200.index[0])
SMA_50 = SMA_50.drop(SMA_50.index[0])
SMA_50.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="1mQcadFgZ3a9" outputId="b2766113-fef0-4247-a9b7-32126cf3fded"
SMA_50.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="uy2hUuMQZ63s" outputId="01e53684-4581-406c-d992-0b10ba332129"
SMA_200.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="g6LW0JuFZ9Ps" outputId="90555add-ff19-41fb-a38f-cf365dc53bff"
df_signal = []
for x in tr_sto:
df_signal.append(tr_sto[x].rolling(window = 50, min_periods = 1).mean())
df_signal = pd.DataFrame(df_signal)
df_signal = df_signal.T
df_signal[df_signal > 0] = 0
df_signal = df_signal.drop(df_signal.index[0])
df_signal.head()
# + id="4HywjWKoaQi4"
for stock in list(df_signal.columns):
df_signal[stock] = np.where(SMA_200[stock] > SMA_50[stock], -1.0, 1.0)
# + id="uqzGGrstahvZ"
pd.set_option('display.max_rows', None)
df_signal.iloc[200:500]
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="oO0iMTsfa5mT" outputId="f10960f8-2798-4eb2-d1fb-31325df97c32"
df_signal.head()
# + id="vze5Kp_kboAE"
M = []
for stock in list(df_signal.columns):
L = []
d = {}
for i in range(len(df_signal[stock])-1):
if df_signal[stock][i] < df_signal[stock][i+1]:
L.append((list(df_signal.index)[i+1], "achat"))
elif df_signal[stock][i] > df_signal[stock][i+1]:
L.append((list(df_signal.index)[i+1], "vente"))
d[stock] = L
M.append(d.copy())
stock_name = []
data = []
for i in range(len(M)) :
for j in range(len(list(M[i].values())[0])) :
stock_name.append(list(M[i].keys())[0])
data.extend(list(M[i].values())[0])
data_signaux = pd.DataFrame(data, columns = ["Date","Signal"])
data_signaux["Stocks"] = stock_name
data_signaux = data_signaux.sort_values(by='Date').reset_index().drop('index',axis = 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="wokGuuaUopye" outputId="43379544-7681-4ceb-a48e-89d0cd3f10a6"
# Restriction des signaux à l'ensemble d'entrainement.
data_signaux = data_signaux[data_signaux.Date.isin(test_stock.index)].reset_index().drop('index',axis = 1)
data_signaux.sample()
# + [markdown] id="Za_1BlkO28Dx"
# # Etape 2 : backtest avec la stratégie en question
# + id="X-NNrjgl28Dy"
portefeuille_intial = { "Actif":list(poids.columns) , "Poids":list(poids.iloc[0,:].values) }
portefeuille_intial_df = pd.DataFrame(portefeuille_intial)
portefeuille_intial_df["Poids"] = portefeuille_intial_df["Poids"].astype('float64')
#portefeuille = portefeuille_intial_df[portefeuille_intial_df['Poids']>0].reset_index().drop('index',axis=1) #Portefeuille contenant tous les actifs
portefeuille = portefeuille_intial_df
# + id="Q49hasKfhcGh"
#On récupère les périodes de calcul des poids les poids sont calculés utilisant les data des 252 jours précédent.
jours = 252
dates_signaux = data_signaux.Date
dates_intervalle = [ train_rets.index[-jours:] ]
for i in range(len(dates_signaux)) :
dates_intervalle.append(returns[ returns.index < dates_signaux[i] ].index[-jours:] )
# + id="037mMj1O28D3" colab={"base_uri": "https://localhost:8080/"} outputId="5dd0eca8-0abf-4b3b-b16c-2dcbb484d7a9"
#On back test uniquement sur le testset
dico_strat = {"Date":[], "Portfolio":[],'Signal':[],"Actif":[],'Rendement':[],'Volatilité':[],"Sharpe ratio":[]}
portefeuille_initial = portefeuille
dates_début = [ test_rets.index[0] ] #Date de début entre signaux afin de calculer les rendements
for i in range(len(data_signaux)) : #Pour chaque date
print("chargement : ", 100*(i+1)/len(data_signaux),'/100' )
date_de_signal = data_signaux['Date'][i] #date du signal
actif_concerné = data_signaux['Stocks'][i] #actif qui emet le signal
type_signal = data_signaux['Signal'][i] #type de signal : achat ou vente
data_returns = returns[returns.index.isin(dates_intervalle[i]) ] # Période sur laquelle on va ajuster les poids
data_returns2 = returns[returns.index.isin(dates_intervalle[i+1]) ] #Periode sur laquelle on va calculer les rendements
if type_signal == 'achat' :
if actif_concerné in portefeuille.Actif.values : #Si Actif déja présent dans le portefeuille
portefeuille = calcule_portefeuille_optimal(data_returns) #Recalculer le poids du portefeuille en ajustant la période
else : #Si Actif pas présent dans le portefeuille
portefeuille = calcule_portefeuille_optimal(data_returns) #Recalculer le poids du portefeuille en ajustant la période
if type_signal == 'vente' :
if actif_concerné in portefeuille.Actif.values : #Si Actif déja présent dans le portefeuille
data_returns2 = data_returns2.drop(actif_concerné,axis=1) #On le vire
portefeuille = calcule_portefeuille_optimal(data_returns.drop(actif_concerné,axis=1) ) #on recalcule le poids (Sans l'actif)
portefeuille = portefeuille[portefeuille.Actif != actif_concerné ]
#else : #Actif pas présent dans le portefeuille on ne fait rien
dates_début.append(date_de_signal) #Ajouter les dates de début pour savoir quand on rentre dans le portefeuille
#Calcul des metrics : rendement et volatilité
#r_i = data_returns[ (data_returns.index >= dates_début[i] ) & (data_returns.index <= dates_début[i+1] ) ]
r_i = data_returns2
w_i = (1/100) * portefeuille[["Poids"]]
#volatility : calcule des rendement
var_p = w_i.values.reshape(-1,1) *( r_i.cov() @ w_i.values.reshape(-1,1) ) * 252
var_p = var_p.sum()
std_p = np.sqrt(var_p)
#Returns
r_p = r_i.mean().to_frame().T @ w_i.values.reshape(-1,1) * 252
r_p
#Sharpe
SR_p = (r_p - 0.0176 )/std_p #Rendre le rf rate journalier
#On enregistre la composition de chaque portefeuille pour chaque date
dico_strat["Date"].append(date_de_signal)
dico_strat["Portfolio"].append(portefeuille)
dico_strat['Signal'].append(type_signal)
dico_strat["Actif"].append(actif_concerné)
dico_strat['Rendement'].append(r_p.values[0][0] )
dico_strat['Volatilité'].append(std_p[0] )
dico_strat["Sharpe ratio"].append(SR_p.values[0][0] )
# + id="COIjNhlBP1tZ"
#Créer une base de donnée pour stocker les résultats
test = pd.DataFrame(dico_strat)
resultat = {'Date':[],'Signal':[],'Emetteurs':[],'Portfolio':[], 'Rendement':[],'Volatilité':[],'Sharpe ratio':[] }
date_list = test.Date.unique()
#for date in date_list :
#Checker si le signal d'achat apparait 2 fois
for i in range(len(date_list)) :
#signaux d'achat
if len(test[ (test.Date==date_list[i] ) & (test.Signal=="achat") ]) != 0 :
signaux_achat = test[ (test.Date==date_list[i] ) & (test.Signal=="achat") ].reset_index()
resultat["Date"].append(date_list[i] )
resultat['Signal'].append("achat")
resultat['Emetteurs'].append(signaux_achat.Actif.values)
if len(signaux_achat) == 1 :
resultat['Portfolio'].append( signaux_achat.loc[0,"Portfolio"].to_dict('records') )
else :
resultat['Portfolio'].append(signaux_achat.Portfolio[0].to_dict('records'))
resultat['Rendement'].append(signaux_achat.Rendement[0] )
resultat['Volatilité'].append(signaux_achat['Volatilité'][0])
resultat["Sharpe ratio"].append( signaux_achat['Sharpe ratio'][0] )
#signaux de vente
if len(test[ (test.Date==date_list[i] ) & (test.Signal=="vente") ]) != 0 :
signaux_vente = test[ (test.Date==date_list[i] ) & (test.Signal=="vente") ].reset_index()
resultat["Date"].append(date_list[i] )
resultat['Signal'].append("vente")
resultat['Emetteurs'].append(signaux_vente.Actif.values)
if len(signaux_vente) == 1 :
resultat['Portfolio'].append( signaux_vente.loc[0,"Portfolio"].to_dict('records') )
else :
resultat['Portfolio'].append(signaux_vente.Portfolio[0].to_dict('records'))
resultat['Rendement'].append(signaux_vente.Rendement[0] )
resultat['Volatilité'].append(signaux_vente['Volatilité'][0])
resultat["Sharpe ratio"].append( signaux_vente['Sharpe ratio'][0] )
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="0wZXaHF428D4" outputId="85a09c0c-3bae-4fa3-d090-1bb0e20ecc19"
strat_1_res = pd.DataFrame(resultat)
strat_1_res = strat_1_res.drop_duplicates(subset = "Date",keep="last")
strat_1_res[['Rendement','Volatilité']] = strat_1_res[['Rendement','Volatilité']].applymap(lambda x: "{0:.1f}%".format(x*100))
strat_1_res
# + id="SLhnE22jz-BS"
nombre_signaux = len(strat_1_res.Date.unique())
periode_test = len(test_rets)
signaux_par_periode = nombre_signaux/periode_test
# + colab={"base_uri": "https://localhost:8080/", "height": 248} id="0rbYKmuq_k4Z" outputId="153eb322-c6c0-4c5d-8085-de36abd07525"
#Diagrame signaux/jours
import seaborn as sns
colors = sns.color_palette('bright')[0:2]
explode = (0.1, 0.0)
plt.pie([periode_test,nombre_signaux],labels = ["Nombre de jours dans le set de test","Nombre de signaux dans le set de test"], colors = colors, autopct='%.0f%%',explode = explode)
#plt.legend()
plt.show()
# + id="FnmMCykg28D6"
#Exemple : composition du portefeuille au premier signal
portf = pd.DataFrame(strat_1_res.iloc[0,:]['Portfolio'])
em_signaux = strat_1_res.iloc[0,:]['Emetteurs']
# + colab={"base_uri": "https://localhost:8080/", "height": 999} id="FLpQv8n2Dzmb" outputId="4c343a10-9e4d-41e7-d0ca-2e26e5eccd67"
import seaborn as sns
w = list(portf[portf.Poids>0].Poids)
stocks_names = list(portf[portf.Poids>0].Actif)
colors = sns.color_palette('bright')[0:len(portf)]
#explode = np.arange()
plt.figure(figsize=(25, 16), dpi=80)
plt.pie(w ,labels = stocks_names , colors = colors, autopct='%.0f%%')
#plt.legend()
plt.show()
#
# + colab={"base_uri": "https://localhost:8080/"} id="wmxIgEydJTJM" outputId="d8e3e8d4-8836-40d5-eb38-ef9b45e2cb9f"
em_signaux
# + id="ZPrStA67AGRp" colab={"base_uri": "https://localhost:8080/", "height": 81} outputId="96cc5b88-f8bd-48aa-ca44-e949056897ff"
#Performance moyenne du portefeuille sur l'horizon de test :
strat_1_res = pd.DataFrame(resultat)
strat_1_res = strat_1_res.drop_duplicates(subset = "Date",keep="last")
resultat_1 = {"Rendement":[], "Volatilité":[], 'Sharpe ratio':[]}
resultat_1["Rendement"].append(strat_1_res.Rendement.mean())
resultat_1["Volatilité"].append(strat_1_res.Volatilité.mean())
resultat_1["Sharpe ratio"].append(strat_1_res["Sharpe ratio"].mean() )
res_f = pd.DataFrame(resultat_1)
res_f[["Rendement","Volatilité"]] = res_f[["Rendement","Volatilité"]].applymap(lambda x: "{0:.1f}%".format(x*100))
res_f
# + [markdown] id="q5rGoETnyua9"
# #Strategie de test 2 : Poids fixe <br/> <br/>
# On calcule le poids sur l'échantillon de train et on garde les mêmes poids pour le back test sur l'échantillon de test
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="RaOzmTCE9cIR" outputId="db4f7e9b-40f4-404c-8362-bdd308f7b0a4"
#Etape 1 calculer les poids sur la périodes de train :
poids_1 = calcule_portefeuille_optimal(train_rets)
#Etape 2 : calculer les rendements sur la période de test
r_i = test_rets
w_i = (1/100) * poids_1[["Poids"]]
#volatility : calcul des rendement
var_p = w_i.values.reshape(-1,1) *( r_i.cov() @ w_i.values.reshape(-1,1) ) * 252
var_p = var_p.sum()
std_p = np.sqrt(var_p)
#Returns
r_p = r_i.mean().to_frame().T @ w_i.values.reshape(-1,1) * 252
#Sharpe
SR_p = (r_p - 0.0176)/std_p
portefeuille_2 = poids_1[poids_1.Poids>0.01].reset_index().drop('index',axis = 1)
resultat_2 = {"Portefeuille":[] , 'Rendement':[],'Volatilité':[],'Sharpe ratio':[] }
resultat_2["Portefeuille"].append( portefeuille_2.to_dict('records') )
resultat_2["Rendement"].append( r_p[0][0] )
resultat_2["Volatilité"].append( std_p[0] )
resultat_2["Sharpe ratio"].append( SR_p[0][0])
res_f2 = pd.DataFrame(resultat_2)
res_f2[["Rendement","Volatilité"]] = res_f2[["Rendement","Volatilité"]] .applymap(lambda x: "{0:.1f}%".format(x*100))
res_f2
# + [markdown] id="l5taJ6nOzDUS"
# #Strategie de test 3 : Poids dynamique <br/> <br/>
# On réactualise le poids chaque 252 jours sur l'échantillon de test
# + id="No4hwq2a90OI"
from datetime import timedelta
jours = 252
nbr_années_test = 3 #On décide de splitter sur 3 années
periode_annuelles = [test_rets.index[0] + timedelta(days=jours)] #Date à laquelle on attribue de nouveau poids
resultat_3 = {'Date':[] , "Portefeuille":[] , 'Rendement':[],'Volatilité':[],'Sharpe ratio':[] }
dates_intervalle = [ train_rets.index[-jours:] ] #dates des périodes de calcul des poids.
#On récupère les périodes de calcul des poids et les dates auxquelles on attribue de nouveaux poids
for i in range(1,nbr_années_test+1) :
periode_annuelles.append( periode_annuelles[i-1] + timedelta(days=jours) )
dates_intervalle.append(test_rets[ test_rets.index < periode_annuelles[i-1] ].index[-jours:] )
#Back test :
for i in range(1,len(periode_annuelles) ) :
data_poids = returns[returns.index.isin(dates_intervalle[i-1]) ]
poids_2 = calcule_portefeuille_optimal(data_poids) #On calcule le poids sur les data dispo avant notre période de rotation
#On calcule les rendements sur la période actuelle
r_i = returns[returns.index.isin(dates_intervalle[i]) ]
w_i = (1/100) * poids_2[["Poids"]]
#volatility : calcule des rendement
var_p = w_i.values.reshape(-1,1) *( r_i.cov() @ w_i.values.reshape(-1,1) ) * 252
var_p = var_p.sum()
std_p = np.sqrt(var_p)
#Returns
r_p = r_i.mean().to_frame().T @ w_i.values.reshape(-1,1) * 252
#Sharpe
SR_p = (r_p - 0.0176)/std_p
portefeuille_3 = poids_2[poids_2.Poids>0.01].reset_index().drop('index',axis = 1)
resultat_3["Date"].append( periode_annuelles[i] )
resultat_3["Portefeuille"].append( portefeuille_3.to_dict('records') )
resultat_3["Rendement"].append( r_p[0][0] )
resultat_3["Volatilité"].append( std_p[0] )
resultat_3["Sharpe ratio"].append( SR_p[0][0])
# + colab={"base_uri": "https://localhost:8080/", "height": 144} id="OX5GTZQkwyro" outputId="7951abbe-799d-48a5-fcae-90aa625d7c62"
strat_2_res = pd.DataFrame(resultat_3)
strat_2_res = pd.DataFrame(resultat_3)
strat_2_res[["Rendement","Volatilité"]] = strat_2_res [["Rendement","Volatilité"]] .applymap(lambda x: "{0:.1f}%".format(x*100))
strat_2_res
# + colab={"base_uri": "https://localhost:8080/", "height": 81} id="IBDx-BDx0_96" outputId="7d9ef6b3-0ddb-4b9b-f61c-6a222bc1ccad"
#Resultat moyen des portefeuille sur la période de test.
strat_2_res = pd.DataFrame(resultat_3)
resultat_2 = {"Rendement":[], "Volatilité":[], 'Sharpe ratio':[]}
resultat_2["Rendement"].append(strat_2_res.Rendement.mean())
resultat_2["Volatilité"].append(strat_2_res.Volatilité.mean())
resultat_2["Sharpe ratio"].append(strat_2_res["Sharpe ratio"].mean() )
res_f3 = pd.DataFrame(resultat_2)
res_f3[["Rendement","Volatilité"]] = res_f3[["Rendement","Volatilité"]].applymap(lambda x: "{0:.1f}%".format(x*100))
res_f3
# + id="eVPU1rf1gYvN"
# + id="H6rvYoksJc2I"
| Notebook/Portfolio_management.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.9 64-bit (''cudaEnv'': venv)'
# language: python
# name: python3
# ---
import numpy as np
from tqdm import tqdm
import torch.nn as nn
import torch
import time
from SparsityAnalysis import extract_patterns, SparseConvArrays
# ### Load Actual Weights from Pattern-Pruned ResNet-34
# +
path = 'resnet34_6_pattern_connectivity_pruning.pt'
state_dict = torch.load(path, map_location=torch.device('cpu'))
# residual_conv_dict = {k:v.cpu().numpy() for (k,v) in state_dict.items() if "layer" in k and "conv" in k}
residual_convs = [v.cpu().numpy() for (k, v) in state_dict.items() if "layer" in k and "conv" in k]
data_shapes = [
[1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32],
[1, 64, 32, 32], [1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16],
[1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8],
[1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8],
[1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 512, 4, 4], [1, 512, 4, 4], [1, 512, 4, 4],
[1, 512, 4, 4], [1, 512, 4, 4],
]
# -
# ### Time Cost w/o memory transfer - nnpack
# +
from conv_naive import Convolution
conv = Convolution()
cuda0 = torch.device('cuda:0')
cpu = torch.device('cpu')
total_time = 0
time_nnpack = []
repeat = 10
for idx in tqdm(range(len(residual_convs[:]))):
input_data = np.ones(data_shapes[idx]).astype(np.float32)
conv_mask = residual_convs[idx].astype(np.float32)
input_data_g = torch.tensor(input_data, device = cuda0)
conv_mask_g = torch.tensor(conv_mask, device = cuda0)
cnt_total = 0
for r in range(repeat):
torch.cuda.synchronize()
start = time.time()
output_gt = nn.functional.conv2d(input_data_g, conv_mask_g,padding=1)
#output_gt = nn.functional.conv2d(torch.tensor(input_data), torch.tensor(conv_mask),padding=1)
torch.cuda.synchronize()
end = time.time()
cnt_total += end - start
total_time += cnt_total/repeat
time_nnpack.append(cnt_total/repeat)
print(f'{round(total_time,3)}s')
# -
# ### Time Cost w/o memory transfer - normal conv
# +
from conv_naive import Convolution
import numpy as np
from tqdm import tqdm
import time
conv = Convolution()
cuda0 = torch.device('cuda:0')
total_time = 0
time_normal_conv = []
for idx in tqdm(range(len(residual_convs[:]))):
input_data = np.ones(data_shapes[idx]).astype(np.float32)
conv_mask = residual_convs[idx].astype(np.float32)
cnt_total = 0
for r in range(repeat):
output_1, time_= conv.conv_multiple_filters(input_data, conv_mask)
cnt_total += time_
total_time += cnt_total/repeat
time_normal_conv.append(cnt_total/repeat)
print(f'{round(total_time,3)}s')
# -
# ### Time Cost w/o memory transfer - sparse conv
# +
from execution_time import SparseConvolution
from tqdm import tqdm
conv = SparseConvolution()
path = 'resnet34_6_pattern_connectivity_pruning.pt'
state_dict = torch.load(path, map_location=torch.device('cpu'))
residual_convs = [v.cpu().numpy() for (k, v) in state_dict.items() if "layer" in k and "conv" in k]
data_shapes = [
[1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32], [1, 64, 32, 32],
[1, 64, 32, 32], [1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16],
[1, 128, 16, 16], [1, 128, 16, 16], [1, 128, 16, 16], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8],
[1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8],
[1, 256, 8, 8], [1, 256, 8, 8], [1, 256, 8, 8], [1, 512, 4, 4], [1, 512, 4, 4], [1, 512, 4, 4],
[1, 512, 4, 4], [1, 512, 4, 4]]
time_sparse_naive = []
time_sparse_shared = []
time_sparse_constant = []
total_time_naive = 0
total_time_shared = 0
total_time_constant = 0
for i in tqdm(range(len(residual_convs[:]))):
input_data = np.float32(np.ones(data_shapes[i]))
if i == len(residual_convs) - 1:
output_data = np.float32(np.zeros(data_shapes[i]))
else:
output_data = np.float32(np.zeros(data_shapes[i + 1]))
conv_layer_weight = residual_convs[i].astype(np.float32)
patterns = np.array(extract_patterns(conv_layer_weight))
sparse_conv_arrays = SparseConvArrays(conv_layer_weight, patterns)
offset = sparse_conv_arrays.offset
reorder = sparse_conv_arrays.reorder
index = sparse_conv_arrays.index
stride = sparse_conv_arrays.stride
sparse_weight = sparse_conv_arrays.weight
ptset = np.float32(sparse_conv_arrays.ptset)
# step 卷积步长
if i == len(residual_convs) - 1:
step = 1
else:
step = int(data_shapes[i][2] / data_shapes[i + 1][2])
cnt_total_naive = 0; cnt_total_shared = 0
for r in range(repeat):
output_naive, time_without_mem_naive, time_include_mem_naive = conv.conv_sparse_naive(input_data, offset, reorder, index, stride, sparse_weight, ptset, step, output_data)
output_shared, time_without_mem_shared, time_include_mem_shared = conv.conv_sparse_shared_mem(input_data, offset, reorder, index, stride, sparse_weight, ptset, step, output_data)
cnt_total_naive += time_without_mem_naive
cnt_total_shared += time_without_mem_shared
time_sparse_naive.append(cnt_total_naive/repeat)
time_sparse_shared.append(cnt_total_shared/repeat)
total_time_naive += cnt_total_naive/repeat
total_time_shared += cnt_total_shared/repeat
#constant memory limit
if sparse_weight.shape[0] <= 16384:
cnt_total_constant= 0;
for r in range(repeat):
output_constant, time_without_mem_constant, time_include_mem_constant = conv.conv_sparse_shared_constant_mem(input_data, offset, reorder, index, stride, sparse_weight, ptset, step, output_data)
cnt_total_constant += time_without_mem_constant
time_sparse_constant.append(cnt_total_constant/repeat)
total_time_constant += cnt_total_constant/repeat
print(f'naive: {round(total_time_naive,3)}s')
print(f'shared: {round(total_time_shared,3)}s')
print(f'constant: {round(total_time_constant,3)}s')
# -
# ### Time Cost w/o memory transfer - sparse conv - vs datasize
# +
from execution_time import SparseConvolution
from tqdm import tqdm
conv = SparseConvolution()
path = 'resnet34_6_pattern_connectivity_pruning.pt'
state_dict = torch.load(path, map_location=torch.device('cpu'))
residual_convs = [v.cpu().numpy() for (k, v) in state_dict.items() if "layer" in k and "conv" in k]
data_shapes = [[1, 64, 32, 32], [1, 64, 128, 128], [1, 64, 256, 256], [1, 64, 512, 512], [1, 64, 1024, 1024]]
time_without_mem_list_naive = []
time_include_mem_list_naive = []
time_without_mem_list_shared = []
time_include_mem_list_shared = []
time_without_mem_list_constant = []
time_include_mem_list_constant = []
time_wo_naive = 0
time_wo_shared = 0
time_wo_constant = 0
conv_layer_weight = residual_convs[i].astype(np.float32)
patterns = np.array(extract_patterns(conv_layer_weight))
sparse_conv_arrays = SparseConvArrays(conv_layer_weight, patterns)
offset = sparse_conv_arrays.offset
reorder = sparse_conv_arrays.reorder
index = sparse_conv_arrays.index
stride = sparse_conv_arrays.stride
sparse_weight = sparse_conv_arrays.weight
ptset = np.float32(sparse_conv_arrays.ptset)
for i in tqdm(range(len(data_shapes))):
input_data = np.float32(np.ones(data_shapes[i]))
if i == len(residual_convs) - 1:
output_data = np.float32(np.zeros(data_shapes[i]))
else:
output_data = np.float32(np.zeros(data_shapes[i + 1]))
conv_layer_weight = residual_convs[i].astype(np.float32)
patterns = np.array(extract_patterns(conv_layer_weight))
sparse_conv_arrays = SparseConvArrays(conv_layer_weight, patterns)
offset = sparse_conv_arrays.offset
reorder = sparse_conv_arrays.reorder
index = sparse_conv_arrays.index
stride = sparse_conv_arrays.stride
sparse_weight = sparse_conv_arrays.weight
ptset = np.float32(sparse_conv_arrays.ptset)
# step 卷积步长
if i == len(residual_convs) - 1:
step = 1
else:
step = int(data_shapes[i][2] / data_shapes[i + 1][2])
output_naive, time_without_mem_naive, time_include_mem_naive = conv.conv_sparse_naive(input_data, offset, reorder, index, stride, sparse_weight, ptset, step, output_data)
output_shared, time_without_mem_shared, time_include_mem_shared = conv.conv_sparse_shared_mem(input_data, offset, reorder, index, stride, sparse_weight, ptset, step, output_data)
time_without_mem_list_naive.append(time_without_mem_naive)
time_include_mem_list_naive.append(time_include_mem_naive)
time_without_mem_list_shared.append(time_without_mem_shared)
time_include_mem_list_shared.append(time_include_mem_shared)
time_wo_naive += time_without_mem_naive
time_wo_shared += time_without_mem_shared
#constant memory limit
if sparse_weight.shape[0] <= 16384:
output_constant, time_without_mem_constant, time_include_mem_constant = conv.conv_sparse_shared_constant_mem(input_data, offset, reorder, index, stride, sparse_weight, ptset, step, output_data)
time_without_mem_list_constant.append(time_without_mem_constant)
time_include_mem_list_constant.append(time_include_mem_constant)
print(time_wo_naive)
print(time_wo_shared)
print(time_without_mem_list_naive)
print(time_without_mem_list_shared)
# -
# ### Plot Generation
import matplotlib.pyplot as plt
time_nnpack = np.array(time_nnpack)
time_normal_conv = np.array(time_normal_conv)
time_sparse_naive = np.array(time_sparse_naive)
time_sparse_shared = np.array(time_sparse_shared)
time_sparse_constant = np.array(time_sparse_constant)
# +
avg_nnpack = np.mean(time_nnpack)*1000
avg_normal = np.mean(time_normal_conv)*1000
avg_sparse_naive = np.mean(time_sparse_naive)*1000
avg_sparse_shared = np.mean(time_sparse_shared)*1000
avg_sparse_constant = np.mean(time_sparse_constant)*1000
labels = [ 'normal\nconv', 'sparse\nnaive', 'sparse\nshared', 'sparse\nconstant','nnpack']
data = [ avg_normal, avg_sparse_naive, avg_sparse_shared, avg_sparse_constant, avg_nnpack]
def autolabel(rects):
for rect in rects:
height = rect.get_height()
plt.text(rect.get_x()+rect.get_width()/2-0.22, height + 0.01, '%.2f' % height)
plt.figure(figsize=(5, 4), dpi=150)
a = plt.bar(range(len(data)), data, tick_label=labels,color=['tomato','turquoise','turquoise', 'turquoise', 'mediumorchid'], alpha=0.9)
autolabel(a)
plt.title('Average Inference Time Per Layer')
plt.ylabel('time (ms)')
plt.savefig('Average Inference Time Per Layer.png')
# +
#plt.plot(time_normal_conv)
from numpy import polyfit, poly1d
plt.figure(figsize=(5, 4), dpi=150)
layer_id = [i for i in range(26)]
f_sparse_constant = poly1d(polyfit(layer_id,time_sparse_constant * 1000,12))
#f_sparse_naive = poly1d(polyfit(layer_id,time_sparse_naive[:26] * 1000,12))
f_nnpack = poly1d(polyfit(layer_id,time_nnpack[:26] * 1000,12))
#plt.plot(layer_id, f_sparse_naive(layer_id),label = 'naive', color = 'turquoise')
plt.plot(layer_id, f_sparse_constant(layer_id),label = 'sparse', color = 'turquoise')
plt.plot(layer_id, f_nnpack(layer_id), label = 'nnpack', color = 'mediumorchid')
plt.title('Average Inference Time of Each Layer')
plt.xlabel('conv layer id')
plt.ylabel('time (ms)')
plt.legend()
plt.savefig('Average Inference Time of Each Layer')
| Evalutaions.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4 ('base')
# language: python
# name: python3
# ---
# # "Recommendation systems and Collaborative Filtering"
#
# > "Demo and Discussion of Collaborative filtering using FastAI"
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [Recommendation systems, Collaborative filtering, FastAI, Python, Deep learning]
# + colab={"base_uri": "https://localhost:8080/"} id="GQVLtEKPd2Wy" outputId="185f01be-4729-4a12-efe0-563566e18704"
#hide
# !pip install -Uqq fastbook
import fastbook
fastbook.setup_book()
from fastbook import *
# -
# <img src= ../images/Recommend.JPG height=500>
# - One very common problem to solve is when you have a number of users and a number of products, and you want to recommend which products are most likely to be useful for which users.
# There are many variations of this: for example, recommending movies (such as on Netflix), figuring out what to highlight for a user on a home page, deciding what stories to show in a social media feed, and so forth.
# Most internet products we use today are powered by recommender systems. YouTube, Netflix, Amazon, Pinterest, and long list of other internet products all rely on recommender systems to filter millions of content and make personalized recommendations to their users.
#
# - A Recommender/Recommendation system is an information filtering system that seeks to predict the "rating" or "preference" a user would give to an item. These systems typically produce a list of recommendations and there are few ways in which it can be done. Two of the most popular ways are – through collaborative filtering or through content-based filtering.
#
#
#
# <img src= ../images/cf_types.JPG>
# + [markdown] id="PYhld5U8H10j"
# **Content based**: This approach utilizes a series of discrete characteristics of an item in order to recommend additional items with similar properties.
# Content-based filtering methods are based on a description of the item and a profile of the user's preferences.
# for eg, it will suggest you similar movies based on the movie we give (movie name would be the input) or based on all of the movies watched by a user (user is the input). It extracts features of a item and it can also look at the user's history to make the suggestions.
#
# **Collaborative based filtering** works like this: look at what products the current user has used or liked, find other users that have used or liked similar products, and then recommend other products that those users have used or liked.
#
# For example, on Netflix you may have watched lots of movies that are science fiction, full of action, and were made in the 1970s. Netflix may not know these particular properties of the films you have watched, but it will be able to see that other people that have watched the same movies that you watched also tended to watch other movies that are science fiction, full of action, and were made in the 1970s. In other words, to use this approach we don't necessarily need to know anything about the movies, except who like to watch them.
#
# There is actually a more general class of problems that this approach can solve, not necessarily involving users and products. Indeed, for collaborative filtering we more commonly refer to *items*, rather than *products*. Items could be links that people click on, diagnoses that are selected for patients, and so forth.
# The key foundational idea is that of *latent factors*. In the Netflix example, we started with the assumption that you like old, action-packed sci-fi movies. But you never actually told Netflix that you like these kinds of movies. And Netflix never actually needed to add columns to its movies table saying which movies are of these types. Still, there must be some underlying concept of sci-fi, action, and movie age, and these concepts must be relevant for at least some people's movie watching decisions.
# + [markdown] id="jpYdUW3ZH10m"
# ## A First Look at the Data
# + [markdown] id="RQkJCx-sH10n"
# Looking at the movie recommendation problem using MovieLens dataset (https://grouplens.org/datasets/movielens/). This dataset contains tens of millions of movie rankings (a combination of a movie ID, a user ID, and a numeric rating), although we will just use a subset of 100,000 of them for our study.
# + colab={"base_uri": "https://localhost:8080/", "height": 55} id="lutU-l3Mi6kw" outputId="0fac9436-5943-4469-c0fb-174caab48aaf"
#hide-output
from fastai.collab import *
from fastai.tabular.all import *
path = untar_data(URLs.ML_100k)
path
# + colab={"base_uri": "https://localhost:8080/"} id="0iJhBIFZkAS5" outputId="78f7accc-248f-4928-a056-83d09ed9b70e"
#hide
Path.BASE_PATH = path
path.ls()
# + [markdown] id="ZmTeeHgUH10t"
# According to the *README*, the main table is in the file *u.data*. It is tab-separated and the columns are, respectively user, movie, rating, and timestamp.
# + colab={"base_uri": "https://localhost:8080/", "height": 237} id="8pH9L3gxmWgp" outputId="2e536b6e-c9d0-48b6-dc90-7b1a7c62a581"
#collapse-input
ratings = pd.read_csv(path/'u.data', delimiter='\t', header=None,
names=['user', 'movie', 'rating', 'timestamp'])
ratings.set_index('timestamp', inplace=True, drop=True)
ratings.head()
# -
# <img src = ../images/Crosstab.JPG height=600>
# + [markdown] id="4iXCE-QtH10w"
# We have selected just a few of the most popular movies, and users who watch the most movies, for this crosstab example. The empty cells in this table are the things that we would like our model to learn to fill in via prediction. Those are the places where a user has not reviewed the movie yet, presumably because they have not watched it. For each user, we would like to figure out which of those movies they might be most likely to enjoy and recommend it to them.
#
# If we knew for each user to what degree they liked each important category that a movie might fall into, such as genre, age, preferred directors and actors, and so forth, and we knew the same information about each movie, then a simple way to fill in this table would be to multiply this information together for each movie and use a combination. For instance, assuming these factors range between -1 and +1, with positive numbers indicating stronger matches and negative numbers weaker ones, and the categories are science-fiction, action, and old movies, then we could represent the movie *The Last Skywalker* as:
# + id="bdFvy83dH10x"
last_skywalker = np.array([0.98,0.9,-0.9])
# + [markdown] id="TQe0NYr5H10x"
# Here, for instance, we are scoring *very science-fiction* as 0.98, *very action* as 0.9, and *very not old* as -0.9. We could represent a user who likes modern sci-fi action movies as:
# + id="EH3NRrKfH10y"
user1 = np.array([0.9,0.8,-0.6])
# + [markdown] id="ecQTf41SH10y"
# and we can now calculate the match between this combination:
# + colab={"base_uri": "https://localhost:8080/"} id="RMP8YBpPH10y" outputId="1e46c024-3f14-44ae-c424-c4e6c801e65a"
(user1*last_skywalker).sum()
# + [markdown] id="XHI73rttH106"
# When we multiply two vectors together and add up the results, this is known as the *dot product*. It is used a lot in machine learning, and forms the basis of matrix multiplication.
# + [markdown] id="1FFmbhg3H108"
# On the other hand, we might represent the movie *Casablanca* as:
# + colab={"base_uri": "https://localhost:8080/"} id="wB-POsh9H108" outputId="3dde5252-ca1d-4595-b3cd-3d71f623159e"
casablanca = np.array([-0.99,-0.3,0.8])
(user1*casablanca).sum()
# + [markdown] id="ENZ7DWYCH10-"
# Since we don't know what the latent factors actually are, and we don't know how to score them for each user and movie, we should learn them.
# + [markdown] id="g6Jm2p_EH10-"
# ## Learning the Latent Factors
# + [markdown] id="Px_CNJ1aH10_"
# Step 1 of this approach is to randomly initialize some parameters. These parameters will be a set of latent factors for each user and movie. We will have to decide how many to use. For illustrative purposes let's use 5 for now. Because each user will have a set of these factors and each movie will have a set of these factors, we can show these randomly initialized values right next to the users and movies in our crosstab, and we can then fill in the dot products for each of these combinations in the middle. For example, below shows what it looks like in Microsoft Excel, with the top-left cell formula displayed as an example.
# + [markdown] id="NDntL5RCH10_"
# <img src="../images/Crosstab2.JPG">
# + [markdown] id="Irfyck-3H11A"
# - Step 2 of this approach is to calculate our predictions. We can do this by simply taking the dot product of each movie with each user. If, for instance, the first latent user factor represents how much the user likes action movies and the first latent movie factor represents if the movie has a lot of action or not, the product of those will be particularly high if either the user likes action movies and the movie has a lot of action in it or the user doesn't like action movies and the movie doesn't have any action in it. On the other hand, if we have a mismatch (a user loves action movies but the movie isn't an action film, or the user doesn't like action movies and it is one), the product will be very low.
#
# - Step 3 is to calculate our loss. We can use any loss function that we wish; using mean squared error for now, since that is one reasonable way to represent the accuracy of a prediction.
#
# That's all we need. With this in place, we can optimize our parameters (that is, the latent factors) using stochastic gradient descent, such as to minimize the loss. At each step, the stochastic gradient descent optimizer will calculate the match between each movie and each user using the dot product, and will compare it to the actual rating that each user gave to each movie. It will then calculate the derivative of this value and will step the weights by multiplying this by the learning rate. After doing this lots of times, the loss will get better and better, and the recommendations will also get better and better.
# + [markdown] id="hFceSDpgH11B"
# To use the usual `Learner.fit` function we will need to get our data into a `DataLoaders`, so let's focus on that now.
# + [markdown] id="gFrrWdVFH11C"
# ## Creating the DataLoaders
# + [markdown] id="bDD9sNuLH11C"
# When showing the data, we would rather see movie titles than their IDs. The table `u.item` contains the correspondence of IDs to titles:
# + colab={"base_uri": "https://localhost:8080/"} id="XkoGBvtwb7IK" outputId="1c909e51-08b6-4a8d-b990-d83c42d2bf00"
movies = pd.read_csv(path/'u.item', encoding='latin-1', delimiter='|',
usecols=(0,1), header=None, names=["Movie_ID", "Title"])
movies.head(), movies.shape
# + [markdown] id="jH0RDZhTH11H"
# We can merge this with our `ratings` table to get the user ratings by title:
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="LWJhLcEWihn-" outputId="cccd961f-8ecf-43e4-e2bc-5e64a859b5bd"
ratings = ratings.merge(movies, left_on="movie" , right_on="Movie_ID").drop("movie", axis=1)
ratings.head()
# + [markdown] id="pOC_ZZmfH11J"
# We can then build a `DataLoaders` object from this table. By default, it takes the first column for the user, the second column for the item (here our movies), and the third column for the ratings. We need to change the value of `item_name` in our case to use the titles instead of the IDs:
# + colab={"base_uri": "https://localhost:8080/", "height": 363} id="rtx3lr9LmJKd" outputId="71d15f07-d1d4-483c-ce05-2803c06ce899"
dls = CollabDataLoaders.from_df(ratings, item_name="Title", rating_name="rating", bs=64)
dls.show_batch()
# + [markdown] id="rh1GpvKBRueV"
# ## Embeddings
# + [markdown] id="ashVEXu6H11L"
# To represent collaborative filtering in PyTorch we can't just use the crosstab representation directly, especially if we want it to fit into our deep learning framework. We can represent our movie and user latent factor tables as simple matrices:
# + colab={"base_uri": "https://localhost:8080/"} id="VNonaEFuakC4" outputId="aec4f07c-8ef0-47a8-e082-112ae33ca31e"
dls.classes
# + id="meG2_Z2hH11L"
n_users = len(dls.classes['user'])
n_movies = len(dls.classes['Title'])
n_factors = 5
user_factors = torch.randn(n_users, n_factors)
movie_factors = torch.randn(n_movies, n_factors)
# + [markdown] id="x9rUFEffH11N"
# To calculate the result for a particular movie and user combination, we have to look up the index of the movie in our movie latent factor matrix and the index of the user in our user latent factor matrix; then we can do our dot product between the two latent factor vectors. But *look up in an index* is not an operation our deep learning models know how to do. They know how to do matrix products, and activation functions.
#
# Fortunately, it turns out that we can represent *look up in an index* as a matrix product. The trick is to replace our indices with one-hot-encoded vectors. Here is an example of what happens if we multiply a vector by a one-hot-encoded vector representing the index 3:
# + colab={"base_uri": "https://localhost:8080/"} id="MhKrRCWAH11N" outputId="d5db0512-c8ce-4013-c834-56a18cb9cd92"
one_hot_3 = one_hot(3, n_users).float()
user_factors.t() @ one_hot_3
# + [markdown] id="Pz6VSk2TH11d"
# It gives us the same vector as the one at index 3 in the matrix:
# + colab={"base_uri": "https://localhost:8080/"} id="K0NvL3i7H11e" outputId="e328949d-6043-4a73-e40a-54e3d31ea346"
user_factors[3], user_factors.shape
# + [markdown] id="SpJvBf3AH11f"
# If we do that for a few indices at once, we will have a matrix of one-hot-encoded vectors, and that operation will be a matrix multiplication! This would be a perfectly acceptable way to build models using this kind of architecture, except that it would use a lot more memory and time than necessary. We know that there is no real underlying reason to store the one-hot-encoded vector, or to search through it to find the occurrence of the number one—we should just be able to index into an array directly with an integer. Therefore, most deep learning libraries, including PyTorch, include a special layer that does just this; it allows indexing into a vector using an integer, but we can also compute gradients of it, identical to what it would have been if it had done a matrix multiplication with a one-hot-encoded vector. This is called an *embedding*.
#
# Embedding: Multiplying by a one-hot-encoded matrix, using the computational shortcut that it can be implemented by simply indexing directly and having gradients. The thing that you multiply the one-hot-encoded matrix by (or, using the computational shortcut, index into directly) is called the _embedding matrix_.
# + [markdown] id="2EGId_dLH11j"
# In computer vision, we have a very easy way to get all the information of a pixel through its RGB values: each pixel in a colored image is represented by three numbers. Those three numbers give us the redness, the greenness and the blueness, which is enough to get our model to work afterward.
#
# For the problem at hand, we don't have the same easy way to characterize a user or a movie. There are probably relations with genres: if a given user likes romance, they are likely to give higher scores to romance movies. Other factors might be whether the movie is more action-oriented versus heavy on dialogue, or the presence of a specific actor that a user might particularly like.
#
# How do we determine numbers to characterize those? The answer is, we don't. We will let our model *learn* them. By analyzing the existing relations between users and movies, our model can figure out itself the features that seem important or not.
#
# This is what embeddings are. We will attribute to each of our users and each of our movies a random vector of a certain length (here, `n_factors=5`), and we will make those learnable parameters. That means that at each step, when we compute the loss by comparing our predictions to our targets, we will compute the gradients of the loss with respect to those embedding vectors and update them with the rules of SGD (or another optimizer).
#
# At the beginning, those numbers don't mean anything since we have chosen them randomly, but by the end of training, they will. By learning on existing data about the relations between users and movies, without having any other information, we will see that they still get some important features, and can isolate blockbusters from independent cinema, action movies from romance, and so on.
# + [markdown] id="8aLEfPB5H11k"
# ## Collaborative Filtering from Scratch
# + [markdown] id="0ZBEHbMoH11m"
# In PyTorch, all models are written as classes. Here is an example of a simple class:
# + id="Iiz9KucUDcuA"
class Example:
def __init__(self, a): self.a = a
def say(self, x): return f'Hello {self.a}, {x}.'
# + [markdown] id="nGOMoNtpH11n"
# The most important piece of this is the special method called `__init__` (pronounced *dunder init*). In Python, any method surrounded in double underscores like this is considered special. It indicates that there is some extra behavior associated with this method name. In the case of `__init__`, this is the method Python will call when your new object is created. So, this is where you can set up any state that needs to be initialized upon object creation. Any parameters included when the user constructs an instance of your class will be passed to the `__init__` method as parameters. Note that the first parameter to any method defined inside a class is `self`, so you can use this to set and get any attributes that you will need:
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="_RiXzJD9DxwM" outputId="4e8b119f-08ab-4e71-d69a-455619bf2a61"
ex = Example('Ashish')
ex.say('What\'s up?')
# + [markdown] id="j8TQ1xzEH11q"
# Also note that creating a new PyTorch module requires inheriting from `Module` class, which provides some basic foundations that we want to build on. So, we add the name of this *superclass* after the name of the class that we are defining, as shown below.
#
# The final thing that you need to know to create a new PyTorch module is that when your module is called, PyTorch will call a method in your class called `forward`, and will pass along to that any parameters that are included in the call. Here is the class defining our simple dot product model:
# + id="HwBBxHtJGsxt"
class DotProduct(Module):
def __init__(self, n_users, n_movies, n_factors):
self.user_factors = Embedding(n_users, n_factors)
self.movie_factors = Embedding(n_movies, n_factors)
def forward(self, x):
users = self.user_factors(x[:,0])
movies = self.movie_factors(x[:,1])
return (users * movies).sum(dim=1)
# + [markdown] id="V0IV7uTJH11r"
# Note that the input of the model is a tensor of shape `batch_size x 2`, where the first column (`x[:, 0]`) contains the user IDs and the second column (`x[:, 1]`) contains the movie IDs. We use the *embedding* layers to represent our matrices of user and movie latent factors:
# + colab={"base_uri": "https://localhost:8080/"} id="POPhxewNhT9l" outputId="87398ae9-7d97-4676-adaf-991b97895086"
x,y = dls.one_batch()
x.shape
# + [markdown] id="du-8tnuzH11t"
# Now that we have defined our architecture, and created our parameter matrices, we need to create a `Learner` to optimize our model. Since we are doing things from scratch here, we will use the plain `Learner` class:
# + id="9NS4m7mSmtWN"
model = DotProduct(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="2n4qJqFkzGQX" outputId="1448014d-0973-4178-ae4b-b454a3b86266"
learn.fit_one_cycle(5, 5e-3)
# + [markdown] id="8ATi0M6pH11x"
# The first thing we can do to make this model a little bit better is to force those predictions to be between 0 and 5. For this, we just need to use `sigmoid_range`. One thing discovered empirically is that it's better to have the range go a little bit over 5, so we use `(0, 5.5)`:
# + id="djrHRVNCH11x"
class DotProduct(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0, 5.5)):
self.user_factors = Embedding(n_users, n_factors)
self.movie_factors = Embedding(n_movies, n_factors)
self.y_range = y_range
def forward(self, x):
users = self.user_factors(x[:,0])
movies = self.movie_factors(x[:,1])
return sigmoid_range((users * movies).sum(dim=1), *self.y_range)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="yU8hHV7uH11x" outputId="6cec83d2-45bd-4207-ddd6-5f727aad579e"
model = DotProduct(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3)
# + [markdown] id="yjZlTg4uH11y"
# Not much difference, we need to make some changes to the architecture.
# One obvious missing piece is that some users are just more positive or negative in their recommendations than others, and some movies are just plain better or worse than others irrespective of the rating from the user. But in our dot product representation we do not have any way to encode either of these things. If all you can say about a movie is, for instance, that it is very sci-fi, very action-oriented, and very not old, then you don't really have any way to say whether most people like it.
#
# That's because at this point we only have weights; we do not have biases. If we have a single number for each user that we can add to our scores, and ditto for each movie, that will handle this missing piece very nicely. So first of all, let's adjust our model architecture:
# + id="UMfLL0uixsJP"
class DotProductBias(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0, 5.5)):
self.user_factors = Embedding(n_users, n_factors)
self.user_bias = Embedding(n_users, 1)
self.movie_factors = Embedding(n_movies, n_factors)
self.movie_bias = Embedding(n_movies, 1)
self.y_range = y_range
def forward(self, x):
users = self.user_factors(x[:, 0])
movies = self.movie_factors(x[:, 1])
res = (users * movies).sum(dim=1, keepdim=True)
res += self.user_bias(x[:, 0 ]) + self.movie_bias(x[:, 1])
return sigmoid_range(res, *self.y_range)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="aCU0E94c1gmg" outputId="a750f00e-69a4-4779-dec0-b976b88f1aee"
model = DotProductBias(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3)
# + [markdown] id="OApfNYEEH110"
# Instead of being better, it ends up being worse (at least at the end of training). If we look at both trainings carefully, we can see the validation loss stopped improving in the middle and started to get worse, which is a clear indication of overfitting.
# + [markdown] id="8T6oCn3uH111"
# ### Regularising the model - Weight Decay
# + [markdown] id="gmmHQ7ACH111"
# Weight decay, or *L2 regularization*, consists in adding to your loss function the sum of all the weights squared. When we compute the gradients, it will add a contribution to them that will encourage the weights to be as small as possible.
#
# The idea is that the larger the coefficients are, the sharper canyons we will have in the loss function,which we don't want since it means that small changes to the input results in big changes to the loss. If we take the basic example of a parabola, `y = a * (x**2)`, the larger `a` is, the more *narrow* the parabola is.
# + colab={"base_uri": "https://localhost:8080/", "height": 377} id="PZElrPeNH112" outputId="a9ac305b-5b65-4cee-87c9-5de4d3cfe1ba"
#hide_input
#id parabolas
x = np.linspace(-2,2,100)
a_s = [1,2,5,10,50]
ys = [a * x**2 for a in a_s]
_,ax = plt.subplots(figsize=(8,6))
for a,y in zip(a_s,ys): ax.plot(x,y, label=f'a={a}')
ax.set_ylim([0,5])
ax.legend();
# + [markdown] id="o4DV2RZCH114"
# So, letting our model learn high parameters might cause it to fit all the data points in the training set with an overcomplex function that has very sharp changes, which will lead to overfitting.
#
# Limiting our weights from growing too much is going to hinder the training of the model, but it will yield a state where it generalizes better.
#
# Weight decay (`wd`) is a parameter that controls that sum of squares we add to our loss (assuming `parameters` is a tensor of all parameters):
#
# ``` python
# loss_with_wd = loss + wd * (parameters**2).sum()
# ```
#
# In practice, though, it would be very inefficient to compute that big sum and add it to the loss, instead the derivative of `p**2` with respect to `p` is `2*p`, so adding that big sum to our loss is exactly the same as doing:
#
# ``` python
# parameters.grad += wd * 2 * parameters
# ```
#
# In practice, since `wd` is a parameter that we choose, we can just make it twice as big, so we don't even need the `*2` in this equation.
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="myfnEFfW0EXU" outputId="816c459d-edbf-4b61-f388-50e737b9c538"
model = DotProductBias(n_users, n_movies, 50)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3, wd=0.1)
# + [markdown] id="Br4nr-6eH116"
# ### Creating Our Own Embedding Module
# + [markdown] id="a2PqaiTgH116"
# Let's re-create `DotProductBias` *without* using the Embedding class, we'll need a randomly initialized weight matrix for each of the embeddings.
# Optimizers require that they can get all the parameters of a module from the module's `parameters` method. However, this does not happen fully automatically. If we just add a tensor as an attribute to a `Module`, it will not be included in `parameters`:
# + colab={"base_uri": "https://localhost:8080/"} id="x25IjdasILub" outputId="cc658b33-4b14-4ae7-bd66-401c1d81cd25"
class T(Module):
def __init__(self): self.a = torch.ones(3)
L(T().parameters())
# + [markdown] id="UCqc7SVYH117"
# To tell `Module` that we want to treat a tensor as a parameter, we have to wrap it in the `nn.Parameter` class. This class doesn't actually add any functionality (other than automatically calling `requires_grad_` for us). It's only used as a "marker" to show what to include in `parameters`:
# + colab={"base_uri": "https://localhost:8080/"} id="vBCOVB2TJQQl" outputId="019c816b-bcde-4d81-f716-ee803d8bee06"
class T(Module):
def __init__(self): self.a = nn.Parameter(torch.ones(3))
L(T().parameters())
# + [markdown] id="pEcncs0VH118"
# All PyTorch modules use `nn.Parameter` for any trainable parameters, which is why we haven't needed to explicitly use this wrapper up until now:
# + colab={"base_uri": "https://localhost:8080/"} id="9qGCHxUuLQIF" outputId="dc41b8d8-8635-4966-d8fc-bb1060fb0cbf"
class T(Module):
def __init__(self): self.a = nn.Linear(1, 3, bias=False)
t = T()
L(t.parameters())
# + colab={"base_uri": "https://localhost:8080/"} id="g60FQG5XH11-" outputId="ea6a03d0-be0b-4c02-afbd-623446c208cb"
type(t.a.weight)
# + [markdown] id="6OXfPUD3H11-"
# Creating a tensor as a parameter, with random initialization and
# using it to create `DotProductBias` again, but without `Embedding`:
# + id="e8oJpQyjOTsi"
def create_params(size):
return nn.Parameter(torch.zeros(*size).normal_(0, 0.01))
# + id="oOctM9aMaw6r"
class DotProductBias(Module):
def __init__(self, n_users, n_movies, n_factors, y_range=(0, 5.5)):
self.user_factors = create_params([n_users, n_factors])
self.movie_factors = create_params([n_movies, n_factors])
self.user_bias = create_params([n_users])
self.movie_bias = create_params([n_movies])
self.y_range = y_range
def forward(self, x):
users = self.user_factors[x[:, 0]]
movies = self.movie_factors[x[:, 1]]
res = (users * movies).sum(dim=1)
res += self.user_bias[x[:, 0]] + self.movie_bias[x[:, 1]]
return sigmoid_range(res, *self.y_range)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="6f8fVM2XUc2R" outputId="2a96ec6a-5ab8-434e-8b37-99e7a7ed3546"
model = DotProductBias(n_users, n_movies, 50)
learn = Learner(dls, model, MSELossFlat())
learn.fit_one_cycle(5, 5e-3, wd=0.1)
# + [markdown] id="kRnZU8xwH12A"
# ## Interpreting Embeddings and Biases
# + [markdown] id="1WpTPW6fH12A"
# The movies with the lowest values in the bias vector:
# + colab={"base_uri": "https://localhost:8080/"} id="sj_ts5k-d5EF" outputId="f4c7fb59-7dce-4578-bb02-270542275a34"
movie_bias = learn.model.movie_bias.squeeze()
idxs = movie_bias.argsort()[:5]
[dls.classes['Title'][i] for i in idxs]
# + [markdown] id="TjChdNDRH12B"
# - For each of these movies, even when a user is very well matched to its latent factors ( things like level of action, age of movie, and so forth), they still generally don't like it.
# - We could have simply sorted the movies directly by their average rating, but looking at the learned bias tells us something much more interesting. It tells us not just whether a movie is of a kind that people tend not to enjoy watching, but that people tend not to like watching it even if it is of a kind that they would otherwise enjoy!
#
# By the same token, here are the movies with the highest bias:
# + colab={"base_uri": "https://localhost:8080/"} id="TXowwU3rjC-6" outputId="5e072a44-acb7-4bde-ed5e-cad2eb2f3ced"
idxs = movie_bias.argsort(descending=True)[:5]
[dls.classes["Title"][i] for i in idxs]
# + [markdown] id="PrhjVSxfH12B"
# So, for instance, even if you don't normally enjoy drama movies, you might enjoy *Shawshank Redemption*!
#
# It is not quite so easy to directly interpret the embedding matrices, there are just too many factors. Using PCA to pull out the most important underlying *directions* from the matrix.
# Below shows what our movies look like based on two of the strongest PCA components.
# + id="RdbsDlHHH12B" outputId="cbe32bb3-f7cf-405b-845e-e5f3c8f99d91"
g = ratings.groupby('title')['rating'].count()
top_movies = g.sort_values(ascending=False).index.values[:1000]
top_idxs = tensor([learn.dls.classes['title'].o2i[m] for m in top_movies])
movie_w = learn.model.movie_factors[top_idxs].cpu().detach()
movie_pca = movie_w.pca(3)
fac0,fac1,fac2 = movie_pca.t()
idxs = list(range(50))
X = fac0[idxs]
Y = fac2[idxs]
plt.figure(figsize=(12,12))
plt.scatter(X, Y)
for i, x, y in zip(top_movies[idxs], X, Y):
plt.text(x,y,i, color=np.random.rand(3)*0.7, fontsize=11)
plt.show()
# + [markdown] id="2WBHZtz5H12C"
# We can see here that the model seems to have discovered a concept of *classic* versus *pop culture* movies, or perhaps it is *critically acclaimed* that is represented here.
# + [markdown] id="PPi5MprTH12D"
# ### Using fastai.collab
# + [markdown] id="nuK8b0piH12D"
# We can create and train a collaborative filtering model using the exact structure shown earlier by using fastai's `collab_learner`:
# + id="h9CAEJtrPcBH"
learn = collab_learner(dls, n_factors=50, y_range=(0, 5.5))
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="tVTLGurcPlqQ" outputId="785c6417-5baa-45eb-edf7-9c5c2f5a1863"
learn.fit_one_cycle(5, 5e-3, wd=0.1)
# + [markdown] id="oAseYRiEH12E"
# The names of the layers can be seen by printing the model:
# + id="evjkma2EH12E" outputId="a9f132c8-5bff-4796-fa79-6238e20b6588"
learn.model
# + [markdown] id="udc_Ja28H12I"
# We can use these to replicate any of the analyses we did in the previous section—for instance:
# + id="CqfmtEFkH12I" outputId="65540f02-9169-4482-d959-fd448245d658"
movie_bias = learn.model.i_bias.weight.squeeze()
idxs = movie_bias.argsort(descending=True)[:5]
[dls.classes['title'][i] for i in idxs]
# + [markdown] id="K7xixgagH12J"
# Another interesting thing we can do with these learned embeddings is to look at _distance_.
# + [markdown] id="A_x0RoxLH12R"
# ### Embedding Distance
# + [markdown] id="nJxktbkHH12R"
# On a two-dimensional map we can calculate the distance between two coordinates using the formula of Pythagoras: $\sqrt{x^{2}+y^{2}}$ (assuming that *x* and *y* are the distances between the coordinates on each axis). For a 50-dimensional embedding we can do exactly the same thing, except that we add up the squares of all 50 of the coordinate distances.
#
# If there were two movies that were nearly identical, then their embedding vectors(and distance from origin) would also have to be nearly identical, because the users that would like them would be nearly exactly the same. There is a more general idea here: movie similarity can be defined by the similarity of users that like those movies.
# The distance between two movies' embedding vectors can define the similarity between them. We can use this to find the most similar movie to *Silence of the Lambs*:
# + colab={"base_uri": "https://localhost:8080/", "height": 36} id="CBlBOIxMahsm" outputId="8f888333-b808-4d51-e100-5e02a6f0b4a7"
movie_factors = learn.model.i_weight.weight
idx = dls.classes['Title'].o2i['Silence of the Lambs, The (1991)']
distances = nn.CosineSimilarity(dim=1)(movie_factors, movie_factors[idx][None])
idx2 = distances.argsort(descending=True)[1]
dls.classes["Title"][idx2]
# + [markdown] id="2qYnD1gtt9pj"
# Now that we have succesfully trained a model, let's see how to deal with the situation where we have no data for a user. How can we make recommendations to new users?
# + [markdown] id="72UVGmluH12S"
# ## Bootstrapping a Collaborative Filtering Model
# + [markdown] id="Ba0X9NN7H12S"
# The biggest challenge with using collaborative filtering models in practice is the *bootstrapping problem*. The most extreme version of this problem is when you have no users, and therefore no history to learn from. What products do you recommend to your very first user?
#
# But even if you are a well-established company with a long history of user transactions, you still have the question: what do you do when a new user signs up? And indeed, what do you do when you add a new product to your portfolio? There is no magic solution to this problem, and really the solutions are just variations of *use your common sense*. You could assign new users the mean of all of the embedding vectors of your other users, but this has the problem that the particular combination of latent factors may be not at all common (for instance, the average for the science-fiction factor may be high, and the average for the action factor may be low, but it is not that common to find people who like science-fiction without action). Better would probably be to pick some particular user to represent *average taste*.
#
# Better still is to use a tabular model based on user meta data to construct your initial embedding vector. When a user signs up, think about what questions you could ask them that could help you to understand their tastes. Then you can create a model where the dependent variable is a user's embedding vector, and the independent variables are the results of the questions that you ask them, along with their signup metadata. (You may have noticed that when you sign up for services such as Pandora and Netflix, they tend to ask you a few questions about what genres of movie or music you like; this is how they come up with your initial collaborative filtering recommendations.)
# + [markdown] id="vASyOVMTH12T"
# One thing to be careful of is that a small number of extremely enthusiastic users may end up effectively setting the recommendations for your whole user base. This is a very common problem, for instance, in movie recommendation systems. People that watch anime tend to watch a whole lot of it, and don't watch very much else, and spend a lot of time putting their ratings on websites. As a result, anime tends to be heavily overrepresented in a lot of *best ever movies* lists. In this particular case, it can be fairly obvious that you have a problem of representation bias, but if the bias is occurring in the latent factors then it may not be obvious at all.
#
# Such a problem can change the entire makeup of your user base, and the behavior of your system. This is particularly true because of positive feedback loops. If a small number of your users tend to set the direction of your recommendation system, then they are naturally going to end up attracting more people like them to your system. And that will, of course, amplify the original representation bias. This type of bias has a natural tendency to be amplified exponentially. You may have seen examples of company executives expressing surprise at how their online platforms rapidly deteriorated in such a way that they expressed values at odds with the values of the founders. In the presence of these kinds of feedback loops, it is easy to see how such a divergence can happen both quickly and in a way that is hidden until it is too late.
#
# In a self-reinforcing system like this, we should probably expect these kinds of feedback loops to be the norm, not the exception. Therefore, we should assume that we will see them, plan for that, and identify up front how you will deal with these issues.
# Try to think about all of the ways in which feedback loops may be represented in the system, and how you might be able to identify them in your data.
# In the end, when rolling out any kind of machine learning system. It's all about ensuring that there are humans in the loop; that there is careful monitoring, and a gradual and thoughtful rollout.
# + [markdown] id="dVcV5It5H12U"
# This approach to collaborative filtering is known as *probabilistic matrix factorization* (PMF). Another approach, which generally works similarly well given the same data, is deep learning.
# + [markdown] id="6qp_AmosH12U"
# ## Deep Learning for Collaborative Filtering
# + [markdown] id="fDJYqizHH12V"
# To turn our architecture into a deep learning model, the first step is to take the results of the embedding lookup and concatenate those activations together. This gives us a matrix which we can then pass through linear layers and nonlinearities in the usual way.
#
# Since we'll be concatenating the embeddings, rather than taking their dot product, the two embedding matrices can have different sizes (i.e., different numbers of latent factors).
# + colab={"base_uri": "https://localhost:8080/"} id="ac94_t-7qMRz" outputId="35cd3fab-d707-4268-991e-96da8e42970a"
embs = get_emb_sz(dls)
embs
# + id="iAIXst-F3zar"
class CollabNN(Module):
def __init__(self, user_sz, item_sz, y_range=(0,5.5), n_act=100):
self.user_factors = Embedding(*user_sz)
self.item_factors = Embedding(*item_sz)
self.layers = nn.Sequential(
nn.Linear(user_sz[1] + item_sz[1], n_act),
nn.ReLU(),
nn.Linear(n_act, 1))
self.y_range = y_range
def forward(self, x):
embs = self.user_factors(x[:, 0]), self.item_factors(x[:, 1])
x = self.layers(torch.cat(embs, dim=1))
return sigmoid_range(x, *self.y_range)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="G8yzYWJJH12a" outputId="36301ba0-c744-460e-8c29-3a3296509dff"
model = CollabNN(*embs)
learn = Learner(dls, model, loss_func=MSELossFlat())
learn.fit_one_cycle(5, 5e-3, wd=0.1)
# + [markdown] id="zoEOZ905H12a"
# `CollabNN` creates our `Embedding` layers in the same way as previous classes , except that we now use the `embs` sizes. `self.layers` is a mini-neural net. Then, in `forward`, we apply the embeddings, concatenate the results, and pass this through the mini-neural net. Finally, we apply `sigmoid_range` as we have in previous models.
# + [markdown] id="02Ah884xH12c"
# fastai provides this model in `fastai.collab` if you pass `use_nn=True` in your call to `collab_learner` (including automatically calling `get_emb_sz`), and it lets you easily create more layers. For instance, here we're creating two hidden layers, of size 100 and 50, respectively:
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="i_YXxCEhDvBT" outputId="3b109025-3dbb-4be7-da7f-183b4c448115"
learn = collab_learner(dls, use_nn=True, y_range=(0, 5.5), layers=[100,50])
learn.fit_one_cycle(5, 5e-3, wd=0.1)
# + colab={"base_uri": "https://localhost:8080/"} id="EaiMprUOKDMU" outputId="82f3012e-d32d-4e48-b1e7-f7d266cdf480"
learn.model
# + id="3A-xfYCiH12d"
@delegates(TabularModel)
class EmbeddingNN(TabularModel):
def __init__(self, emb_szs, layers, **kwargs):
super().__init__(emb_szs, layers=layers, n_cont=0, out_sz=1, **kwargs)
# + [markdown] id="79hs4Nt-H12e"
# ### kwargs and Delegates
#
# `learn.model` is an object of type `EmbeddingNN`. This class *inherits* from `TabularModel`, which is where it gets all its functionality from. In `__init__` it calls the same method in `TabularModel`, passing `n_cont=0` and `out_sz=1`; other than that, it only passes along whatever arguments it received.
#
# `EmbeddingNN` includes `**kwargs` as a parameter to `__init__`. In Python `**kwargs` in a parameter list means "put any additional keyword arguments into a dict called `kwargs`. And `**kwargs` in an argument list means "insert all key/value pairs in the `kwargs` dict as named arguments here". This approach is used in many popular libraries, such as `matplotlib`, in which the main `plot` function simply has the signature `plot(*args, **kwargs)`. The [`plot` documentation](https://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot) says "The `kwargs` are `Line2D` properties" and then lists those properties.
#
# `**kwargs` in `EmbeddingNN` used to avoid having to write all the arguments to `TabularModel` a second time, and keep them in sync. However, this makes the API quite difficult to work with, because now Jupyter doesn't know what parameters are available. Consequently things like tab completion of parameter names and pop-up lists of signatures won't work.
#
# fastai resolves this by providing a special `@delegates` decorator, which automatically changes the signature of the class or function (`EmbeddingNN` in this case) to insert all of its keyword arguments into the signature.
# + [markdown] id="5JS0B2hiH12f"
# ### Benefits of using a NN
# + [markdown] id="l6mbcmLRH12f"
# Although the results of `EmbeddingNN` are a bit worse than the dot product approach (which shows the power of carefully constructing an architecture for a domain), it does allow us to do something very important: we can now directly incorporate other user and movie information, date and time information, or any other information that may be relevant to the recommendation and use a `TabularModel`.
# + [markdown] id="g5QSSx8DH12g"
# ## Questionnaire
# - Done in clean nb
# + [markdown] id="mPYGAYcEH12g"
# 1. What problem does collaborative filtering solve?
# 1. How does it solve it?
# 1. Why might a collaborative filtering predictive model fail to be a very useful recommendation system?
# 1. What does a crosstab representation of collaborative filtering data look like?
# 1. What is a latent factor? Why is it "latent"?
# 1. What is a dot product? Calculate a dot product manually using pure Python with lists.
# 1. What does `pandas.DataFrame.merge` do?
# 1. What is an embedding matrix?
# 1. What is the relationship between an embedding and a matrix of one-hot-encoded vectors?
# 1. Why do we need `Embedding` if we could use one-hot-encoded vectors for the same thing?
# 1. What does an embedding contain before we start training (assuming we're not using a pretained model)?
# 1. Create a class (without peeking, if possible!) and use it.
# 1. What does `x[:,0]` return?
# 1. Rewrite the `DotProduct` class (without peeking, if possible!) and train a model with it.
# 1. What is a good loss function to use for MovieLens? Why?
# 1. What would happen if we used cross-entropy loss with MovieLens? How would we need to change the model?
# 1. What is the use of bias in a dot product model?
# 1. What is another name for weight decay?
# 1. Write the equation for weight decay (without peeking!).
# 1. Write the equation for the gradient of weight decay. Why does it help reduce weights?
# 1. Why does reducing weights lead to better generalization?
# 1. What does `argsort` do in PyTorch?
# 1. Does sorting the movie biases give the same result as averaging overall movie ratings by movie? Why/why not?
# 1. How do you print the names and details of the layers in a model?
# 1. What is the "bootstrapping problem" in collaborative filtering?
# 1. How could you deal with the bootstrapping problem for new users? For new movies?
# 1. How can feedback loops impact collaborative filtering systems?
# 1. When using a neural network in collaborative filtering, why can we have different numbers of factors for movies and users?
# 1. Why is there an `nn.Sequential` in the `CollabNN` model?
# 1. What kind of model should we use if we want to add metadata about users and items, or information such as date and time, to a collaborative filtering model?
# + [markdown] id="wgRHr2OHH12h"
# ### Further Research
# 1. Take a look at all the differences between the `Embedding` version of `DotProductBias` and the `create_params` version, and try to understand why each of those changes is required. If you're not sure, try reverting each change to see what happens. (NB: even the type of brackets used in `forward` has changed!)
# 1. Find three other areas where collaborative filtering is being used, and find out what the pros and cons of this approach are in those areas.
# 1. Complete this notebook using the full MovieLens dataset, and compare your results to online benchmarks. See if you can improve your accuracy. Look on the book's website and the fast.ai forum for ideas. Note that there are more columns in the full dataset—see if you can use those too (the next chapter might give you ideas).
# Done in separate notebook.
# 1. Create a model for MovieLens that works with cross-entropy loss, and compare it to the model in this chapter.
# - Study PCA, svd
# - Distance b/w 2 coordinates using pythogrus and cosine sim on khan acadeky
# - learn about yrange in the previous lesson
# + id="d7Wi1GbnG5he"
| _notebooks/2022-06-01-CF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import Image
from IPython.core.display import HTML
from IPython.display import IFrame
# # JODA Luento 4
# Luentomuistion valmisteli [Arho Suominen](https://www.tuni.fi/fi/ajankohtaista/kun-teknologia-muuttuu-yrityksen-taytyy-loytaa-keinot-sopeutua-muutokseen). Vuonna 2022 JODAa luennoi [<NAME>](https://www.tuni.fi/fi/jukka-huhtamaki) ([@jnkka](https://twitter.com/jnkka?lang=fi)).
#
# <hr/>
#
# <!-- <NAME><br />
# Senior Scientist, D.Sc. (Tech)<br />
# Adjunct Professor in Technology Forecasting and Analytics at Tampere University of Technology <br />
# VTT TECHNICAL RESEARCH CENTRE OF FINLAND<br />
# Innovations, Economy, and Policy<br />
# Vuorimiehentie 3, P.O. Box 1000, 02044 Espoo, Finland<br />
# Tel. +358 50 5050 354<br />
# www.vtt.fi, <EMAIL><br />
# https://www.linkedin.com/in/arhosuominen<br />
# Twitter: @ArhoSuominen <br /> -->
# ## Oppimistavoitteet
#
# * Harjoitustyöhön syventyminen
# * Datatiedeprosessin käytäntöihin tutustuminen
# * Erilaisiin suunnittelumalleihin (prosessimallit, mahdollisuuskehikko, analytiikkakehikko) perehtyminen
#
# ## Mitä prosessia käyttäisi tehdessäsi harjoitustyötä?
# [CRISP-DM](https://www.kdnuggets.com/polls/2014/analytics-data-mining-data-science-methodology.html)-malli on osoittautunut monessa projektissa hyödylliseksi jäsennykseksi ja ajattelun välineeksi. Katso myös [Houston Analyticsin projektimalli](https://www.houston-analytics.com/project-methodology) ja Microsoftin laajempi [tiimidatiedekehikko](https://docs.microsoft.com/en-us/azure/architecture/data-science-process/overview). Tänään tehdäänkin asioita käytännössä.
#
# Kevään mittaan olemme jo käyneet läpi useammankin aineiston. Olemme katsoneet huollon vikailmoituksia, kuluttajadataa finanssisektorilta ja tämän päivän luentoon valmistauduttiin tutustumalla [AirBnB-esimerkkianalyysiin](https://github.com/InfoTUNI/joda2022/blob/master/koodiesimerkit/airnbn/python_scikit_airbnb.ipynb).
#
# Tänään on tarkoituksena käydä keskustellen läpi CRISP-DM -prosessia (tai ainakin sen ensimmäisiä askeleita). Tavoitteena on korostaa sitä, että todellisen maailman datatiedeprojekteissa lähtökohtana on **oletusarvoisesti liiketoimintaongelma tai muu kiinnostava kysymys, ei data**.
#
# Tässä prosessissa käytetään hyödyksi <NAME>in esittelemää [kehikkoa mahdollisuuksien tunnistamiseen](https://www.jpattonassociates.com/opportunity-canvas/). Mahdollisuuskehikko on kehitetty liiketoiminta- ja Lean-kehikkojen pohjalta ja sen tarkoituksena on mahdollistaa keskustelu ratkaisusta jota olemme ideoimassa. Tehtävän tavoitteena on tukea harjoitustyöhön orientoitumista ongelmalähtöisesti.
#
# <NAME> [kuvaa hyvin miten mahdollisuuskehikkoa](https://www.stephenson.dk/opportunity-canvas/) voi hyödyntää ja me käytämme sitä tänään samalla prosessilla. Mahdollisuuskehikon käyttö on yksinkertaista ja se käydään alla läpi lyhyesti.
IFrame("https://www.dropbox.com/s/a7jydl9n7zplqwg/opportunity_canvas.pdf?dl=0", width=1000, height=700)
# <ol>
# <li>Aloita kohdasta 1. Ongelma. Mahdollisuuskehikko on suunniteltu niin, että voit aloittaa joko ratkaisusta tai ongelmasta ja päätyä molempia reittejä takaisin ratkaisuun tai ongelmaan. JODAn puitteissa lähdetään liikkeelle siitä että kuvaat ongelman, eli kysymyksen minkä haluat ratkaista. Vastaakin kysymykseen, mikä on se ongelma johon haluan tuottaa ratkaisun?</li>
# <li>Kuka käyttää ratkaisua? Kuten yleensä, on huono ratkaisu todeta että tätä käyttää kaikki vaan pikemminkin kuvata ne segmentit jotka voisivat käyttää palvelua ja kuvata mitkä ovat näiden segmenttien erot.</li>
# <li>Kuvaa miten ongelma on ratkaistu tänään. Tämä antaa kuvan siitä miten voit mitata oman ratkaisusi "suorituskykyä" verrattuna nykyiseen ratkaisuun. Ratkaisuna voi olla myös jokin epätyypillinen kiertotie ongelman ratkaisemiseen.</li>
# <li>Kohdassa neljä määritellään liiketoimintaongelma, tai pikemminkin se, miten tarjoamasi ongelman ratkaisu vaikuttaa liiketoimintaan. Asiaa voi ajatella myös käänteisesti, miten ongelman jättäminen ratkaisematta vaikuttaa liiketoimintaan.</li>
# <li>Miten käyttäjät käyttävät ratkaisua? Miten käyttäjän prosessi muuttuu siitä että käyttää ratkaisua ja mikä on heidän saama hyöty?</li>
# <li>Miten voisit mitata sitä että käyttäjät testaavat, käyttävät, sisäistävät ratkaisun? Miten mittaat käyttäjän ratkaisusta saamaa hyötyä?</li>
# <li><a href="https://en.wikipedia.org/wiki/Technology_acceptance_model">Teknologian hyväksyntä</a> voidaan käytännössä jakaa kahteen osaan: subjektiivisesti koettu hyöty ja helppokäyttöisyys. Hyöty määritellään siten, että kuinka paljon toteutettu ratkaisu voi helpottaa henkilön työprosessia. Helppokäyttöisyys kuvaa sitä kuinka pienen työn järjestelmän käyttöönotto vaatii.</li>
# <li>Miten voisit mitata ratkaisun vaikutusta yrityksesi liiketoimintaan?</li>
# <li>Budjetti, miten tämä realisoituu käytännössä.</li>
# </ol>
# Lähdetään liikkeelle tärkeimmästä.
# ### Ymmärrä liiketoimintasi
# Tämä vaihe on keskeinen projektisi onnistumiselle. Jotta dataprojektisi lähtisi liikkeelle ja tulisi käyttöönotetuksi, on sen tuotettava selkeää arvo yrityksen liiketoimintaprosessille. Kuten minkä tahansa tuotteen määrittelyssä, on keskeistä tuoda esille se miten jokin yrityksen prosessi tai tuote muuttuu paremmaksi kun siihen yhdistetään jokin datakomponentti.
# <!-- Alla oleva tapauskuvaus on lainaus [Tekoälyaikaraportista](https://www.tekoalyaika.fi/raportit/edellakavijana-tekoalyaikaan/2-kansainvaliset-tekoalyn-asiantuntijat-kohti-tekoalyn-kolmatta-aaltoa/).-->
# <!-- #### Case K-ryhmä: Tekoäly tietää paremmin kuin sinä, mitä haluat syödä -->
# <!--<i>Ruokakauppojen kanta-asiakasjärjestelmät ovat jo pitkään tunteneet asiakkaansa läpikotaisin. Ostosdatan perusteella on helppo tehdä päätelmiä asiakkaan ostoskäyttäytymisestä. Lisäksi tiedämme, että ihmiset tuppaavat ostamaan ruokakaupasta suunnilleen samankaltaisia asioita.
#
# K-RYHMÄSSÄ päätettiin muutama vuosi sitten ryhtyä rakentamaan tästä datasta huomattavasti nykyistä älykkäämpää järjestelmää, joka suosittelisi ihmisille reseptejä heidän puolestaan ja siten helpottaisi heidän arkeaan. Järjestelmä otettiin käyttöön vuonna 2017. Yksinkertaisimmillaan se tarkoittaa sitä, että kun hakukenttään kirjoittaa ”maito”, hakukone antaa sisäänkirjautuneelle käyttäjälle automaattisesti ostoshistoriaan perustuvan suosikkimaidon.
#
# Pian kuitenkin järjestelmää opetettiin älykkäämmäksi ja hakua laajennettiin antamaan myös reseptisuosituksia. Se päättelee ihmisten hakutuloksista, minkälaisia ruokia ostavat ihmiset käyttävät minkäkinlaisia reseptejä, ja suosittelee näitä automaattisesti uusille käyttäjille.
#
# Vuoden alusta järjestelmää on opetettu entistä älykkäämmäksi ja se on laajennettu koskemaan ostolistoja. Käytännössä sisäänkirjautunut käyttäjä voi nyt yhdellä napinpainalluksella saada ensi viikon ostoslistan valmiina itselleen. Harkinnan siitä, mitä ruokakassiin tulisi laittaa, on tehnyt käyttäjän puolesta tekoäly pohjautuen aiempaan ostoslistaan. Se ei siis suosittele ainoastaan aiemmin ostettuja tuotteita, vaan päättelee, mitä käyttäjä haluaisi ensi viikolla mahdollisesti syödä. Mitä enemmän ostoksia ja listoja käyttäjä tekee, sitä paremmin järjestelmä oppii tuntemaan – ja suosittelemaan myös uusia.
#
# Tällöin voi käydä lopulta niin, että tekoäly oppii ihmistä paremmaksi mielitekojen asiantuntijaksi ja ihmisten aikaa vapautuu johonkin muuhun kuin sen miettimiseen, mitä seuraavaksi tekisi mieli.
#
# Ruokaostosdataa on monista asiakkaista jo hyvin pitkän ajan takaa, joten ruokaostoksia ja reseptejä suositteleva tekoäly lähtee liikkeelle varsin hyvästä lähtökohdasta. Dataa on runsaasti ja se on laadukasta.
#
# Sen sijaan vaikeampaa tekoälylle on ollut opettaa henkilökohtaisten mieltymysten ja sesonkivaihtelujen eroa. Eli vaikka käyttäjä kuinka rakastaisi runebergintorttuja, hän tuskin haluaa niistä suosituksia marraskuussa. Haasteena on ollut datan saavutettavuus, laatu ja datamassojen laskentatehojen riittävyys, joissa on vasta viime vuosina saavutettu sellaisia teknisiä harppauksia, että relevanttien suositusten antaminen on mahdollista.
#
# Seuraavana tavoitteena on nivoa resepti- ja ostoslistasuosittelu eheäksi kokonaisuudeksi, joka muuttaa ruokakaupassa käymisen kokonaan. Silloin ruokaostosten valinta ja hankkiminen tapahtuvat käytännössä muutamalla napinpainalluksella. Tähän tapaan:
#
# Järjestelmä kysyy käyttäjältä, haluaako tämä kokata ensi viikolla makaronilaatikkoa ja kasvispastaa, johon käyttäjä vastaa kyllä. Tämän jälkeen järjestelmä ehdottaa ostoslistalle tuotteita, jotka käyttäjä hyväksyy. Tämän jälkeen järjestelmä kysyy, haluatko ostaa tuotteet, jolloin käyttäjä hyväksyy ja maksaa ostokset – ja saa ne hetken päästä toimitettuna kotiovellensa.
#
# Tämä voi olla mahdollista jo vuoden sisällä.
#
# K-ryhmän reseptisuositukset on esimerkki digitaalisten palveluiden kehittämisestä. Kyseessä on ihmisten arkeen vahvasti vaikuttava tekijä: käytämme merkittävän osan ajastamme ruokaostosten hankkimiseen ja sen miettimiseen, mitä ruuaksi laittaisimme tai haluaisimme laittaa. Näin ollen tekoäly voi mullistaa käytännön arkeamme minimoimalla tuon ajan ja oppimalla älykkääksi arvioimaan puolestamme sen, mitä olemme syöneet ja mitä meidän kannattaisi seuraavaksi syödä.</i>-->
# Käy läpi Airbnb:n tiedote mukavuuksista joita [majoittajan kannattaa juuri nyt vierailleen tarjota](https://www.airbnb.fi/resources/hosting-homes/a/the-best-amenities-to-offer-right-now-203).
#
# <!-- majoitus- ja elämysjohtaja [Catherine Powellin viikkokatsaus syksyltä 2020](https://www.airbnb.fi/resources/hosting-homes/a/introducing-airbnbs-covid-19-safety-practices-274). -->
# <b>Keskustellaan pienryhmissä:</b>
# <ol>
# <li>Minkä ongelman Airbnb haluaa ratkaista?</li>
# <li>Listatkaa muutamia mahdollisia analyysejä joiden pohjalta viikkokatsauksen teemat ja esitetyt ohjeistukset ovat muodostuneet.</li>
# </ol>
#
# Aikaa kymmenen minuuttia, varautukaa kertomaan vastaukset myös muille.
# <!-- Etukäteinen perustelu ei kuitenkaan riitä, vaan on luotava myös jonkinlainen arviointikehikko sille miten tiedät datan tuoneen arvoa liiketoimintaprosessiin.
#
# <NAME>ver kuvaa sitä miten voit [mitata analytiikan hyötyjä](https://web.archive.org/web/20201020003032/https://www.ibmbigdatahub.com/blog/top-3-ways-measure-success-your-analytics-investment). Hän nostaa esille kolme tekijää: laatu, nopeus ja robustisuus. -->
# <!-- **Jatketaan vielä pienryhmissä:**
#
# Miten näkisitte että Airbnb tapauskuvauksessa voisi yllä mainittu mittakehikko toimia?
#
# Miten laatu, nopeus ja robustisuus voisi muuttua datan avulla?
#
# Aikaa kymmenen minuuttia, varautukaa kertomaan vastaukset myös muille -->
# Vaikka harjoitustyössä ei olekaan (välttämättä) kyse yrityksen liiketoiminnasta, on yhtä olennaista päättää jokin selkeä kysymys johon voit vastata datan avulla ja vasta sitten pohtia onko kysymykseen vastaamiseksi saatavilla dataa.
# #### Harjoitustyön aloittaminen
# [Mahdollisuuskehikko](https://www.dropbox.com/s/vzzz7nvjvobhcka/opportunity_canvas_editable.pptx?dl=0) ([katso keskustelu suomenkielisestä käsitteestä](https://twitter.com/jnkka/status/1376788470962790401)) on tarjolla verkossa. Meillä on käytössä [Miro-versio kehikosta](https://miro.com/welcomeonboard/dEhDUFB3QmJXUDFtd1A0Q3VtM2pZZjY2ZVh5aDJ6aExIU0xBZ3FmRFVKTk1lcWI0VGRDa3BOa05NUUpXQnNZd3wzMDc0NDU3MzQ3NDA3MzExNDI5?invite_link_id=922037219634).
#
# <ol>
# <li>Pohdi itse minkä ongelman haluisit harjoitustyössä ratkaista. Vaikka <a href="https://infotuni.github.io/joda2022/harjoitustyo/">harjoitustyökuvauksessa</a> keskitytään AirBnB-dataan, olet vapaa miettimään itsellesi mielenkiintoista aihetta. Hahmottele itsellesi mielenkiintoisia ideoita mahdollisuuskehikon lokeroon 1. Näitä voi olla useita, joten ideoi vapaasti.</li>
# <li>Listaa ongelmia <a href="https://flinga.fi/s/FD3AJBT">Flingaan</a>. Mikä ongelmista on oikeasti olemassa? Suodatetaan kiinnostavimmat ideat tykkäämällä.</li>
# </ol>
#
# Listataan ideoita 10 minuuttia.
#
# <ol>
# <li>Jatketaan kiinnostavimpien ideoiden kehittelyä <a href="https://miro.com/app/board/o9J_lMv9Ko8=/">Mirossa</a>. Ota kopio aihekuvauspohjasta tai täydennä olemassa olevaa aihetta.</li>
# <li>Kuka ehdotettua ratkaisua voisi käyttää? Miten voit rajata tämän joukon kaikista muista?</li>
# <li>Mitkä ovat nykyisin käytössä olevat ratkaisut? Kuvaa ne lyhyesti.</li>
# </ol>
#
# Aikaa käytössä on 20 minuuttia. <!--, jonka jälkeen lisää kanvaasisi omaksi kalvokseen.--> <!-- Kun jatkamme eteenpäin, käy lisäämässä oma ideasi ketjuun muiden ideoita lisäämällä oma kanvaasisi [muiden jatkoksi](https://docs.google.com/presentation/d/12UU5E6DXWBuZuFHKZ3uBell8MyyyofyF619D_k1h9Ek/edit?usp=sharing)-->
# ### Hanki data
# Kun kysymys on lukittu voidaan pohtia mistä on mahdollista saada dataa kysymykseen vastaamiseksi. Esittelyssä ovat jo olleet Kaggle-datat, mutta vaihtoehtoja on myös muita. Tässä muutamia:
# <ol>
# <li>Onko käytössäsi organisaation itse luomaa dataa? Jos mahdollista, tulee arvioida mitä datalähteitä organisaatiolla itsellään on käytössä. Vaikka osa aineistosta ei alunperin olisi suunniteltu data analyysiin, voi yrityksen omissa strukturoiduissa tai epästrukturoiduissa datalähteissä olla merkittäviä tietovarantoja.</li>
# <li>Käytä API-rajapintoja: rajapintojen takaa löytyy mitä moninaisempia tietolähteitä joilla voi olla suoraa hyödynnettävyyttä omassa prosessissasi. Esimerkkinä rajapinnoista löytyy vaikka <a href="https://www.programmableweb.com/">ProgrammableWeb-alustalta</a>, johon on koottuu erilaisia rajapintoja, tällä hetkellä yli 20 000 eri rajapintaa.</li>
# <li>Etsi avointa dataa: Varsinkin julkiset toimijat ovat lähteneet hyvin aktiivisesti julkaisemaan aineistojaan avoimen rajapinnan kautta. Esimerkkejä on vaikka Euroopan Unionin ODP, jonka kautta on saatavilla monenlaista aineistoa.</li>
# </ol>
# Alla yksinkertainen esimerkki [Europe PMC rajapinnasta](https://europepmc.org/RestfulWebService).
import requests, json
search="nanorobot%20or%20nanobot%20or%20nanomachine%20or%20nanomotors%20or%20nanoid%20or%20nanite%20or%20nanomite"
r = requests.get('https://www.ebi.ac.uk/europepmc/webservices/rest/search?query='+search+'&format=json&resulttype=core')
query_data = json.loads(r.text)
records=query_data['hitCount']
print("Rajapinnan kautta löytyi yhteensä "+str(records)+" julkaisua")
if records > 1000:
pageSize=1000
pages=int((records/1000)+1)
else:
pageSize=records
pages=1
# ### Tarkastele ja käsittele dataa
# Kun olet valinnut aineiston jota käytät ja hankkinut käsiisi datan (tai ainakin osan siitä), aloita tarkastelemalla mitä datassa on. Esimerkiksi lainojen hyväksyntää kuvaava aineisto oli täynnä virheellisiä arvoja, puuttujia lukuja ja emme voineet olla täysin varmoja, että kaikki tapaukset kuvautuvat datassa alkujaankaan.
#
# Pohdi tässä kohtaa alkuperäistä kysymystäsi. Mitä nyt käytössä olevat muutujat oikestaan kertovat? Mitkä ovat muuttujien asteikot, ovatko muuttujat vinoja ja vain osa kategorisista luokista on edustettuina. Tee itsellesi kuva siitä mihin muuttujat voivat vastata ja ymmärrä myös se mihin ne eivät voi vastata.
#
# Kun olet ymmärtänyt mitä datassa on, voi aloittaa datan jalostamisen. Onko aineistossasi tekstikenttiä joissa on [kirjoitusvirheitä](https://twitter.com/jnkka/status/1110226356217044992?s=19)? Onko jokin muu osa ainestoa käsin syötettyä ja mahdollisesti osittain virheellistä? Pitääkö muuttujia muuntaa? Käy jokainen muuttuja läpi ja varmista että muuttujan arvot on varmasti siivottu.
# ### Rikasta aineistoa
# Jos olet tehnyt edelliset vaiheet hyvin, on tämä vaihe helpompi. Pyri tässä vaiheessa löytämään keino yhdistää eri lähteistä tulevat data yhdeksi. Vastaa kysymyksiin mikä datoja yhdistää? Ovatko muuttujat yhteismitallisia vai tuleeko minun asettaa rajoitteita siihen miten aineistoja voidaan myöhemmin analysoida yhdessä.
#
# Muunna data muotoon jossa tunnistat alkuperäiselle kysymykselle arvokkaat piirteet. Kuvaile mistä eri piirteet ovat tulleet (lähde), miten ne muuttuvat sekä mikä on niiden merkitys alkuperäiselle kysymykselle. Tarvittaessa pyri myös muuntaamaan muuttujia muotoon jossa ne palvelevat paremmin kysymyksen asettelua. Onko sinun analyysissä aikatieto relevantti minuutteina, tunteina vai vuosina?
#
# Hyvän esimerkin siitä miten dataa voi ymmärtää tarjoaa [analytiikkasuunnittelukehikko](https://reader.elsevier.com/reader/sd/pii/S2212827118301549?token=<PASSWORD>).
# +
# IFrame("https://doi.org/10.1016/j.procir.2018.02.031", width=1000, height=1200)
# -
# ### Tee visualisointeja
# Erityisesti jos sinulla on käsissäsi huomattava määrä muuttujia ja aineisto on laaja, tulee kriittisesti arvioida mitkä visualisoinnit ovat arvokkaita loppukäyttäjälle. Vaikka jonkin visualisoinnin tekeminen on mahdollista, sitä ei välttämättä kannata tehdä.
#
# Visualisointien tekemiseen on useita hyviä työkaluja kuten Bokeh, Plotly ja Matplotlib, mutta näistä ei ole erityisemmin hyötyä mikäli et pysty selittämään miksi jokin kuvaaja on arvokas asiaa analysoivalle loppukäyttäjälle. Visualisointien tekemisessä on itseasiassa merkittävä valinta tilanne, kun analyytikko päättää esimerkiksi visualisoida kaksi muuttujaa, mutta taustalle voi jäädä kymmeniä muuttujia. Miksi valitsit siis juuri nämä muuttujat?
# 
#
# Datan kartoittamiseen palataan tulevilla luentoviikoilla.
# ### Mallinna
# Tässä vaiheessa voit huoletta mallintaa dataasi. Mikä olikaan alkuperäinen kysymyksesi ja miten se kääntyy kysymykseksi joka ratkeaa ohjelmallisesti?
# 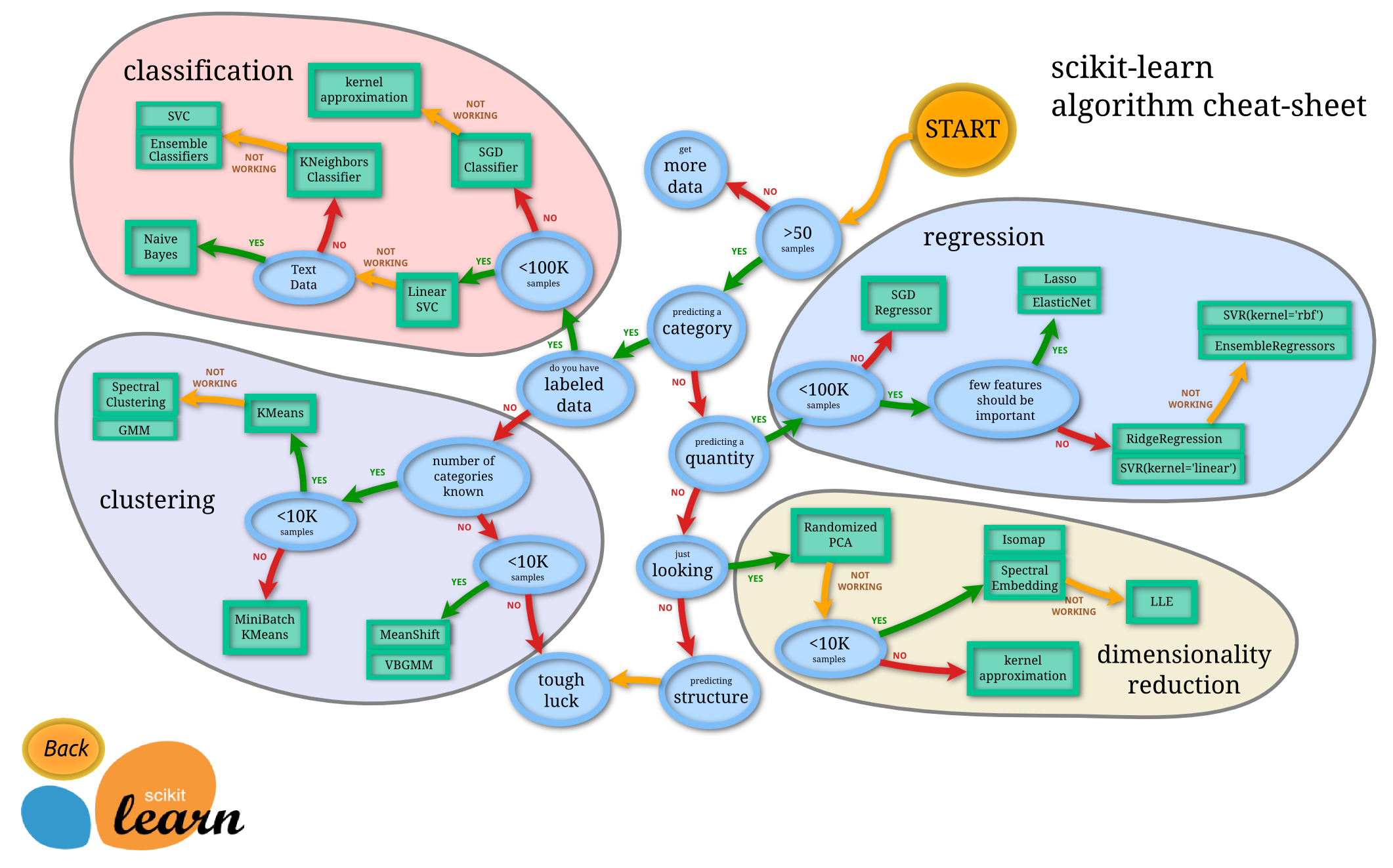
# ### Iteroi
# Työt tuskin loppuivat tähän vaan nyt iteroit. Data analyysin mielenkiintoisin osa on se, että yleensä siinä vaiheessa kun olet käynyt prosessin läpi ensimmäisen kerran on sinulla jo hyvä käsitys siitä miten kyseistä aineistoa voisi edellen täydenttä uudella datalla, rikastaa, tai mallintaa toisella tavalla. Parhaimmillaan huomaat että mieleen on tullut täysin uusi kysymys, mutta jos näin on päässyt käymään, arvioi kriittisesti vastaako nyt kerätty aineisto uuteen kysymykseen.
| luentomuistio/luento04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="AI1Rq7g5ev6R" colab_type="text"
# # LM算法 (Levenberg-Marquardt algorithm)
# + id="XxpVyw8_ergn" colab_type="code" colab={}
#@title LM算法 { display-mode: "both" }
# 本程序实现通过 Levenberg-Marquardt 算法寻找 Rosenbrock 函数的极小值点
# 可选 Rosenbrock 函数由公式 (a-x)**2+b*(y-x**2)**2 决定,代码内设置: a=1., b=1.
# 在未知最优值的情况下,初始 lam 应设置较小,LM算法接近牛顿法,收敛加速,但在最优值附近会出现震荡
# coding: utf-8
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits.mplot3d import Axes3D
import time
# + id="Qcu73hZ33fzO" colab_type="code" colab={}
#@markdown - **计时装饰器**
def timer(func):
def wrapper(*args, **kwargs):
start_time = time.time()
result = func(*args, **kwargs)
end_time = time.time()
t = end_time - start_time
print(' Running time is: {:.4f} s.'.format(t))
return result
return wrapper
# + [markdown] id="whgnp7wIgYd1" colab_type="text"
# ## 类的定义
# + id="onvYhoAAfPAb" colab_type="code" colab={}
#@markdown - **LM算法类**
class Strategy:
def __init__(self, name='Levenberg-Marquardt-algorithm', opt=True, lam=5e-2, max_iters=None, tol=1e-6):
self.name = name
self.opt = opt
self.lam = lam
self.max_iters = max_iters
self.iters = 0
self.tol = tol
def rosenbrock(self, x, y, a=1., b=1.): # 原函数
return (a - x)**2 + b * (y - x**2)**2
def rosenbrock_grad(self, x, y, a=1., b=1.): # 梯度计算式
x_d = 2 * (x - a) - 4 * b * x * (y - x**2)
y_d = 2 * b * (y - x**2)
return np.array([x_d, y_d])
#@markdown - **根据路径画出 3D 示意图**
def draw_chart(self, path, ax, x_lim=[-2., 2], y_lim=[-2, 2], d_off=-1.5):
x, y = np.meshgrid(np.arange(x_lim[0], x_lim[1], 0.1),
np.arange(y_lim[0], y_lim[1], 0.1))
z = self.rosenbrock(x, y)
ax.plot_surface(x, y, z, rstride=2, cstride=2, alpha=0.6, cmap=cm.jet)
# ax.contourf(x, y, z,zdir='z', offset=d_off, cmap=cm.coolwarm) # 显示等高平面
ax.set_xlabel('X', fontsize=14)
ax.set_ylabel('Y', fontsize=14)
ax.set_zlabel('Z', fontsize=14)
# ax.set_zlim([-800.0,2500.0])
ax.view_init(elev=27, azim=65)
# z_labels = self.rosenbrock(np.array(path[0]), np.array(path[1]))
z_path = self.rosenbrock(np.array(path[0]), np.array(path[1]))
if path is not None:
ax.plot(path[0], path[1], z_path, c="#b22222", linewidth=1.)
# ax.plot(path[0], path[1], z_labels, c="#b22222", linewidth=1.)
ax.scatter(path[0][0], path[1][0], z_path[0], c='r', s=30, marker='o')
ax.scatter(path[0][-1], path[1][-1], z_path[-1], c='r', s=30, marker='*')
ax.set_xlim(x_lim), ax.set_ylim(y_lim)
ax.set_xticks(np.linspace(-2., 2., 5, endpoint=True))
ax.set_yticks(np.linspace(-2., 2., 5, endpoint=True))
ax.tick_params(labelsize=14)
#@markdown - **Levenberg-Marquardt算法**
@timer
def LM_algor(self, init_position):
x = [init_position[0]]
y = [init_position[1]]
x_real, y_real = 1., 1. # 理想情况
while True:
cx = x[-1]
cy = y[-1]
grad = self.rosenbrock_grad(cx, cy)
Jacob_coe = np.mat(grad).T * np.mat(grad) + self.lam * np.eye(2) # 步长 delta = -(J.T * J + lam * I)^-1 * J.T * z(x,y)
if self.opt:
err_xy = np.array([abs(x_real - cx), abs(y_real - cy)]) # 已知最小值的坐标
step = np.array(-Jacob_coe.I * np.mat(grad).T) * err_xy.reshape(-1, 1) # 已知最小值情况下加速收敛
else: step = np.array(-Jacob_coe.I * np.mat(grad).T) * self.rosenbrock(cx, cy) # 实际公式,此时需将 lam 设置较小,initial_lam = 8e-8
step = step.flatten()
x.append(cx + step[0])
y.append(cy + step[1])
self.iters += 1
# magnitude = np.sqrt(np.dot(grad, grad))
magnitude = abs((1 - cx) * (1 - cy)) # 误差选择与真实值差的积
if magnitude < self.tol or (self.max_iters is not None and self.iters >= self.max_iters):
break
return {'final_pos': [x[-1], y[-1]], 'iters': self.iters, 'final_grad': grad, 'path': [x, y]}
# + [markdown] id="EFKfeiaF5OU9" colab_type="text"
# ## 已知最优值的情况
# + id="VX71Q1vjvKEW" colab_type="code" outputId="bd97e4a0-843c-4618-afe8-8442e0d37de3" executionInfo={"status": "ok", "timestamp": 1560540846575, "user_tz": -480, "elapsed": 806, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 118}
# 作出 3D 路径及 x-y 平面投影图
init_position = [-1.5, -1.5] # 初始点位置
name = 'Levenberg-Marquardt-algorithm'
s = Strategy(name)
s.__dict__
# + id="DusRuauiv6wZ" colab_type="code" outputId="aae42681-bfcc-46a1-8182-638af271bbb0" executionInfo={"status": "ok", "timestamp": 1560540847614, "user_tz": -480, "elapsed": 1831, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 575}
# 作出 3D 路径及 x-y 平面投影图
fig = plt.figure(1, figsize=(18, 8))
ax1 = fig.add_subplot(121, projection='3d')
print(name.capitalize())
result = s.LM_algor(init_position)
s.draw_chart(result['path'], ax1)
ax1.set_title('{}, loops: {:,}'.format(name, result['iters']), fontsize=14)
x_loc,y_loc = result['final_pos']
print(' Location of the final point: \n x={:.4f}, y={:.4f}'.format(x_loc, y_loc))
ax2 = fig.add_subplot(122)
x_path,y_path = result['path']
ax2.plot(x_path, y_path,'r')
ax2.scatter(x_path[0], y_path[0], c='r', s=30, marker='o')
ax2.scatter(x_path[-1], y_path[-1], c='r', s=30, marker='*')
ax2.set_xlabel('X', fontsize=14), ax2.set_ylabel('Y', fontsize=14)
ax2.tick_params(labelsize=14)
ax2.grid(b=True)
plt.show()
# + [markdown] id="jXKPDCPR_WRj" colab_type="text"
# ## 未知最优值的情况
# + id="oiHQLcwx_Vz8" colab_type="code" outputId="ae189f18-26e8-4514-f9ec-5627790f9ac1" executionInfo={"status": "ok", "timestamp": 1560540847616, "user_tz": -480, "elapsed": 1825, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 118}
# 作出 3D 路径及 x-y 平面投影图
# 初始 lam 较小,LM算法接近牛顿法,收敛加速,但在最优值附近会出现震荡
init_position = [-1.5, -1.5] # 初始点位置
name = 'Levenberg-Marquardt-algorithm'
s = Strategy(name, opt=False, lam=8e-8)
s.__dict__
# + id="S2XCgZeH_i_a" colab_type="code" outputId="496c552f-d248-4a8e-fc91-8c79177013a8" executionInfo={"status": "ok", "timestamp": 1560540848282, "user_tz": -480, "elapsed": 2481, "user": {"displayName": "\u041b\u044f\u043d\u043f\u044d\u043d \u041a", "photoUrl": "https://lh6.googleusercontent.com/-GXVG-PbMfAw/AAAAAAAAAAI/AAAAAAAAADo/wvm2q-yqQzs/s64/photo.jpg", "userId": "04289897042674768581"}} colab={"base_uri": "https://localhost:8080/", "height": 575}
# 作出 3D 路径及 x-y 平面投影图
fig = plt.figure(1, figsize=(18, 8))
ax1 = fig.add_subplot(121, projection='3d')
print(name.capitalize())
result = s.LM_algor(init_position)
s.draw_chart(result['path'], ax1)
ax1.set_title('{}, loops: {:,}'.format(name, result['iters']), fontsize=14)
x_loc,y_loc = result['final_pos']
print(' Location of the final point: \n x={:.4f}, y={:.4f}'.format(x_loc, y_loc))
ax2 = fig.add_subplot(122)
x_path,y_path = result['path']
ax2.plot(x_path, y_path,'r')
ax2.scatter(x_path[0], y_path[0], c='r', s=30, marker='o')
ax2.scatter(x_path[-1], y_path[-1], c='r', s=30, marker='*')
ax2.set_xlabel('X', fontsize=14), ax2.set_ylabel('Y', fontsize=14)
ax2.tick_params(labelsize=14)
ax2.grid(b=True)
plt.show()
| notebooks/LM_algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # List assignments 1
# +
#Question:1
#1. Let us say your expense for every month are listed below,
#1. January - 2200
#2. February - 2350
#3. March - 2600
#4. April - 2130
#5. May - 2190
Month=["jan","Feb","March","April","May"]
exp = [2200,2350,2600,2130,2190]
# -
#In Feb, how many dollars you spent extra compare to January?
Spend=exp[1]-exp[0]
print("how many dollars you spent extra compare to January=",Spend)
#Find out your total expense in first quarter (first three months) of the year.
Spend=exp[0]+exp[1]+exp[2]
print("total expense in first quarter=",Spend)
#Find out if you spent exactly 2000 dollars in any month
2000 in exp
#June month just finished and your expense is 1980 dollar. Add this item to our monthly expense list
Month=["jan","Feb","March","April","May"]
exp = [2200,2350,2600,2130,2190]
exp.append(1980)
print("Added this item to our monthly expense lis=",exp)
#You returned an item that you bought in a month of April and got a refund of 200$.
#Make a correction to your monthly expense list based on this
Month=["jan","Feb","March","April","May"]
exp = [2200,2350,2600,2130,2190]
exp[3]=exp[3]-200
print("correction to your monthly expense list=",exp)
# +
#Question:2
#You have a list of your favourite marvel super heros.
#heros=['spider man','thor','hulk','iron man','captain america']
#Length of the list
heros=['spider man','thor','hulk','iron man','captain america']
print(len(heros))
# -
#Add 'black panther' at the end of this list
heros=['spider man','thor','hulk','iron man','captain america']
heros.append("black panther")
print("Added 'black panther in the list=",heros)
#You realize that you need to add 'black panther' after 'hulk',so remove it from the list first and then add it after 'hulk'
heros=['spider man', 'thor', 'hulk', 'iron man', 'captain america']
heros.insert(3,"black panther")
print(heros)
#4. Now you don't like thor and hulk because they get angry easily :)
#So you want to remove thor and hulk from list and replace them with doctor strange (because he is cool).
#Do that with one line of code.
heros=['spider man', 'thor', 'hulk', 'iron man', 'captain america']
heros[1:3]=["doctor strange"]
print(heros)
#5 Sort the heros list in alphabetical order (Hint. Use dir() functions to list down all functions available in list)
heros=['spider man', 'thor', 'hulk', 'iron man', 'captain america']
heros.sort()
print(heros)
# add 'black panther' after 'hulk', then remove it from the list first and then add it after 'spider man'
heros=['spider man','thor','hulk','iron man','captain america']
heros[3]="black panther"
print(heros)
heros.remove("black panther")
print(heros)
heros[1]="black panther"
print(heros)
| List assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from skimage.io import imread, imsave
import matplotlib.pyplot as plt
from skimage.measure import label
image = imread("Gold_nooverlay.png",as_gray=True)
fig = plt.figure(figsize=(5,5))
plt.imshow(label(image>0.3),cmap='nipy_spectral')
plt.axis('off')
plt.show()
fig = plt.figure(figsize=(10,10))
plt.imshow(image>0.3,cmap='nipy_spectral')
plt.axis('off')
plt.show()
from scipy.ndimage import distance_transform_edt
dst = distance_transform_edt(image>0.3)
fig = plt.figure(figsize=(10,10))
plt.imshow(dst,cmap='nipy_spectral')
plt.axis('off')
plt.show()
from skimage.feature import peak_local_max
peaks = peak_local_max(dst,min_distance=7,exclude_border=True)
fig = plt.figure(figsize=(10,10))
plt.imshow(label(image>0.3),cmap='nipy_spectral')
plt.scatter(peaks[:,1],peaks[:,0])
plt.axis('off')
plt.show()
peaks = peak_local_max(dst,min_distance=1,exclude_border=True)
fig = plt.figure(figsize=(10,10))
plt.imshow(image>0.3,cmap='nipy_spectral')
plt.scatter(peaks[:,1],peaks[:,0])
plt.axis('off')
plt.show()
| pages/particle_separation_with_clustering/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
file = open('../data/df-ac-measurements.csv', 'r')
lines = file.readlines()
# +
filename = '../data/df-ac-measurements-fixed.csv'
outfile = open(filename, "w")
outfile.write(lines[0][1:]) # write header
for i in range(1, 11): # first 10 lines
outfile.write(lines[i][2:])
for i in range(11, 20):
outfile.write(lines[19][3:])
for i in range(20, 22):
line = lines[i][0:]
fields = line.split(',')
device = fields[1]
date = fields[2]
fields[1] = date
fields[2] = device
line = ','.join(fields[0:606])
line = line + '\n'
outfile.write(line)
outfile.close()
# -
file = open('../data/df-ac-measurements-fixed.csv', 'r')
lines = file.readlines()
lines[20]
| notebooks/.ipynb_checkpoints/Fix-dataset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Looking at Muse EEG data
# +
# %matplotlib inline
import mne
from scipy.signal import periodogram
import numpy as np
import pandas as pd
# +
dat = mne.io.read_raw_fif('219b3f487156461e9c37f2eb7f2aaba7_raw.fif')
df_dat = dat.to_data_frame()
# -
df_dat.index
# +
per_freqs,per_dat = periodogram(df_dat.values.T,fs = 256) # 1.*4) # 1000./4)
df_per = pd.DataFrame(per_dat,columns=per_freqs,index=dat.ch_names).T
df_per.index.names = ['Hz']
# -
df_per.plot(logx=True,logy=True,alpha=0.3,figsize=(12,3))
df_per.loc[1:100].plot(logx=True,logy=True,figsize=(12,3),alpha=0.3)
df_per['ch1'].loc[1:100].plot(logx=True,logy=True,figsize=(12,3),alpha=0.5,c='k')
# +
# now:
# bandpass filter
# etc.
# as with eeg notebooks
| scratch/looking_at_muse_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="http://cocl.us/pytorch_link_top">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/Pytochtop.png" width="750" alt="IBM Product " />
# </a>
#
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/cc-logo-square.png" width="200" alt="cognitiveclass.ai logo" />
#
# <h1>Linear Regression Multiple Outputs</h1>
#
# <h2>Objective</h2><ul><li> How to create a complicated models using pytorch build in functions.</li></ul>
#
# <h2>Table of Contents</h2>
# In this lab, you will create a model the Pytroch way. This will help you as models get more complicated.
#
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <li><a href="#ref0">Make Some Data</a></li>
# <li><a href="#ref1">Create the Model and Cost Function the Pytorch way</a></li>
# <li><a href="#ref2">Train the Model: Batch Gradient Descent</a></li>
# <li><a href="#ref3">Practice Questions </a></li>
# <br>
# <p></p>
# Estimated Time Needed: <strong>20 min</strong>
# </div>
#
# <hr>
#
# Import the following libraries:
#
import torch
import numpy as np
import matplotlib.pyplot as plt
from torch import nn,optim
from mpl_toolkits.mplot3d import Axes3D
from torch.utils.data import Dataset, DataLoader
import torchvision.transforms as transforms
# Set the random seed:
#
torch.manual_seed(1)
# <a id="ref0"></a>
#
# <h2 align=center>Make Some Data </h2>
# Create a dataset class with two-dimensional features and two targets:
#
from torch.utils.data import Dataset, DataLoader
class Data(Dataset):
def __init__(self):
self.x=torch.zeros(20,2)
self.x[:,0]=torch.arange(-1,1,0.1)
self.x[:,1]=torch.arange(-1,1,0.1)
self.w=torch.tensor([ [1.0,-1.0],[1.0,3.0]])
self.b=torch.tensor([[1.0,-1.0]])
self.f=torch.mm(self.x,self.w)+self.b
self.y=self.f+0.001*torch.randn((self.x.shape[0],1))
self.len=self.x.shape[0]
def __getitem__(self,index):
return self.x[index],self.y[index]
def __len__(self):
return self.len
# create a dataset object
#
data_set=Data()
# <a id="ref1"></a>
#
# <h2 align=center>Create the Model, Optimizer, and Total Loss Function (cost)</h2>
#
# Create a custom module:
#
class linear_regression(nn.Module):
def __init__(self,input_size,output_size):
super(linear_regression,self).__init__()
self.linear=nn.Linear(input_size,output_size)
def forward(self,x):
yhat=self.linear(x)
return yhat
# Create an optimizer object and set the learning rate to 0.1. **Don't forget to enter the model parameters in the constructor.**
#
model=linear_regression(2,2)
# Create an optimizer object and set the learning rate to 0.1. **Don't forget to enter the model parameters in the constructor.**
#
# <img src = "https://ibm.box.com/shared/static/f8hskuwrnctjg21agud69ddla0jkbef5.png" width = 100, align = "center">
#
optimizer = optim.SGD(model.parameters(), lr = 0.1)
# Create the criterion function that calculates the total loss or cost:
#
criterion = nn.MSELoss()
# Create a data loader object and set the batch_size to 5:
#
train_loader=DataLoader(dataset=data_set,batch_size=5)
# <a id="ref2"></a>
#
# <h2 align=center>Train the Model via Mini-Batch Gradient Descent </h2>
#
# Run 100 epochs of Mini-Batch Gradient Descent and store the total loss or cost for every iteration. Remember that this is an approximation of the true total loss or cost.
#
# +
LOSS=[]
epochs=100
for epoch in range(epochs):
for x,y in train_loader:
#make a prediction
yhat=model(x)
#calculate the loss
loss=criterion(yhat,y)
#store loss/cost
LOSS.append(loss.item())
#clear gradient
optimizer.zero_grad()
#Backward pass: compute gradient of the loss with respect to all the learnable parameters
loss.backward()
#the step function on an Optimizer makes an update to its parameters
optimizer.step()
# -
# Plot the cost:
#
plt.plot(LOSS)
plt.xlabel("iterations ")
plt.ylabel("Cost/total loss ")
plt.show()
# <a href="http://cocl.us/pytorch_link_bottom">
# <img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/notebook_bottom%20.png" width="750" alt="PyTorch Bottom" />
# </a>
#
# ### About the Authors:
#
# [<NAME>](https://www.linkedin.com/in/joseph-s-50398b136?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork-20647811&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ&cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork-20647811&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ) has a PhD in Electrical Engineering. His research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition.
#
# Other contributors: [<NAME>](https://www.linkedin.com/in/michelleccarey?cm_mmc=Email_Newsletter-_-Developer_Ed%2BTech-_-WW_WW-_-SkillsNetwork-Courses-IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork-20647811&cm_mmca1=000026UJ&cm_mmca2=10006555&cm_mmca3=M12345678&cvosrc=email.Newsletter.M12345678&cvo_campaign=000026UJ)
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ---------- | ----------------------------------------------------------- |
# | 2020-09-23 | 2.0 | Shubham | Migrated Lab to Markdown and added to course repo in GitLab |
#
# Copyright © 2018 <a href="cognitiveclass.ai?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu">cognitiveclass.ai</a>. This notebook and its source code are released under the terms of the <a href="https://bigdatauniversity.com/mit-license/">MIT License</a>.
#
# v
#
| 4.4.training_multiple_output_linear_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook contains my replication of [this](https://jakevdp.github.io/blog/2014/06/10/is-seattle-really-seeing-an-uptick-in-cycling/) blog post by [<NAME>](http://vanderplas.com/) on using data from bicycle traffic across Seattle's Fremont Bridge to learn about commuting patterns.
# +
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('seaborn')
from seattlecycling.data import get_fremont_data
from seattlecycling.toolbox import hours_of_daylight
from seattlecycling.toolbox import print_rms
# +
# Load data
start = '1 Oct 2012'
end = '15 May 2014'
data = get_fremont_data()
data = data.loc[start:end]
data.head(3)
# +
# Resample data into daily and weekly totals
daily = data.resample('d').sum()
weekly = data.resample('w').sum()
# +
# A first look at the data
weekly.plot();
plt.ylabel('Weekly rides');
# +
# Look at rolling weekly mean to smooth out short-term variation
data.resample('d').sum().rolling(30, center=True).mean().plot();
# -
# Blog post points out that 2014 has seen increased cycle traffic across the bridge. Below we're modelling seasonal variation based on what we think influences peoples' decision whether or not to ride a bike.
# # Accounting for hours of daylight
# +
# Hours of daylight
weekly['daylight'] = list(map(hours_of_daylight, weekly.index))
daily['daylight'] = list(map(hours_of_daylight, daily.index))
weekly['daylight'].plot()
plt.ylabel('Hours of daylight (Seattle)');
# +
# Relationship between daylight and cycle traffic
plt.scatter(weekly.daylight, weekly.total)
plt.xlabel('Hours of daylight')
plt.ylabel('Weekly bicycle traffic');
# +
# Adding a linear trend
X = weekly[['daylight']]
y = weekly['total']
clf = LinearRegression(fit_intercept=True).fit(X, y)
weekly['daylight_trend'] = clf.predict(X)
weekly['daylight_corrected_total'] = weekly.total - weekly.daylight_trend + np.mean(weekly.daylight_trend)
xfit = np.linspace(7, 17)
yfit = clf.predict(xfit[:, None])
plt.scatter(weekly.daylight, weekly.total)
plt.plot(xfit, yfit, '-k')
plt.title('Bycicle traffic through the year')
plt.xlabel('Hours of daylight')
plt.ylabel('Weekly bicycle traffic');
# -
clf.coef_[0]
# +
# Plot detrended data
trend = clf.predict(weekly[['daylight']].values)
plt.scatter(weekly.daylight, weekly.total - trend + np.mean(trend))
plt.plot(xfit, np.mean(trend) + 0 * yfit, '-k')
plt.title('Weekly traffic through the year (detrended)')
plt.xlabel('Hours of daylight')
plt.ylabel('Adjusted weekly bicycle traffic');
# -
# In the graph above, we have removed the number of riders per week that correlate with the number of hours of daylight, so that we can think of what is shown of the number of rides per week we'd expect to see if daylight was not an issue.
# +
fix, ax = plt.subplots(1, 2, figsize=(15,5))
weekly[['total', 'daylight_trend']].plot(ax=ax[0])
weekly['daylight_corrected_total'].plot(ax=ax[1])
ax[0].set_ylabel('Weekly crossing')
ax[0].set_title('Total weekly crossings and trend')
ax[1].set_ylabel('Adjusted weekly crossings')
ax[1].set_title('Detrended weekly crossings')
print_rms(weekly['daylight_corrected_total'])
# -
# # Accounting for day of the week
# +
# Plot average number of trips by weekday
days = ['Mon', 'Tue', 'Wed', 'Thu', 'Fri', 'Sat', 'Sun']
daily['dayofweek'] = daily['total'].index.dayofweek
grouped = daily.groupby('dayofweek')['total'].mean()
grouped.index = days
grouped.plot()
plt.title('Average crossings by weekday')
plt.ylabel('Average daily crossings');
# +
# Account for hours of daylight and day of week simultaneously
# Add one-hot indicators of weekdays
for i in range(7):
daily[days[i]] = (daily.index.dayofweek == i).astype(float)
# Detrend on days of week and daylight together
X = daily[days + ['daylight']]
y = daily['total']
clf = LinearRegression().fit(X, y)
daily['dayofweek_trend'] = clf.predict(X)
daily['dayofweek_corrected'] = daily['total'] - daily['dayofweek_trend'] + daily['dayofweek_trend'].mean()
# +
# Plot crossings and trend, and detrended data
fix, ax = plt.subplots(1, 2, figsize=(15,5))
daily[['total', 'dayofweek_trend']].plot(ax=ax[0])
daily['dayofweek_corrected'].plot(ax=ax[1])
ax[0].set_ylabel('Daily crossing')
ax[0].set_title('Total daily crossings and trend')
ax[1].set_ylabel('Adjusted daily crossings')
ax[1].set_title('Detrended daily crossings')
print_rms(daily['dayofweek_corrected'])
# -
# # Accounting for rainfall and temparature
# +
# Read in weather data
weather = pd.read_csv('SeaTacWeather.csv', index_col='DATE',
parse_dates=True, usecols=[2, 5, 9, 10])
weather = weather.loc[start:end]
weather.columns = map(str.lower, weather.columns)
# Temparatures are in 1/10 deg F; convert to deg C
weather['tmax'] = (weather['tmax'] - 32) * 5/9
weather['tmin'] = (weather['tmin'] -32) * 5/9
# Rainfall is in inches; convert to mm
weather['prcp'] *= 25.4
weather['tmax'].resample('w').max().plot()
weather['tmin'].resample('w').min().plot()
plt.title('Temperature extremes in Seattle')
plt.ylabel('Weekly temperature extremes (C)');
# -
weather['prcp'].resample('w').sum().plot()
plt.title('Precipitation in Seattle')
plt.ylabel('Weekly precipitation in Seattle (mm)');
# +
# Combine daily and weather dataset
daily = daily.join(weather)
# +
# Detrend data including weather information
columns = days + ['daylight', 'tmax', 'tmin', 'prcp']
X = daily[columns]
y = daily['total']
clf = LinearRegression().fit(X, y)
daily['overall_trend'] = clf.predict(X)
daily['overall_corrected'] = daily['total'] - daily['overall_trend'] + daily['overall_trend'].mean()
# +
# Plot crossings and trend, and detrended data
fix, ax = plt.subplots(1, 2, figsize=(15,5))
daily[['total', 'overall_trend']].plot(ax=ax[0])
daily['overall_corrected'].plot(ax=ax[1])
ax[0].set_ylabel('Daily crossing')
ax[0].set_title('Total daily crossings and trend')
ax[1].set_ylabel('Adjusted daily crossings')
ax[1].set_title('Detrended daily crossings')
print_rms(daily['overall_corrected'])
# +
# Plot rolling 30 day average
daily['overall_corrected'].rolling(30, center=True).mean().plot();
plt.title('1-month moving average');
# -
# # Accounting for a steady increase in riders
# +
daily['daycount'] = np.arange(len(daily))
columns = days + ['daycount', 'daylight', 'tmax', 'tmin', 'prcp']
X = daily[columns]
y = daily['total']
final_model = LinearRegression().fit(X, y)
daily['final_trend'] = final_model.predict(X)
daily['final_corrected'] = daily['total'] - daily['final_trend'] + daily['final_trend'].mean()
# +
# Plot crossings and trend, and detrended data
fix, ax = plt.subplots(1, 2, figsize=(15,5))
daily[['total', 'final_trend']].plot(ax=ax[0])
daily['final_corrected'].plot(ax=ax[1])
ax[0].set_ylabel('Daily crossing')
ax[0].set_title('Total daily crossings and trend')
ax[1].set_ylabel('Adjusted daily crossings')
ax[1].set_title('Detrended daily crossings')
print_rms(daily['final_corrected'])
# -
# # What can the final model tell us?
# +
# Compute error variance
vy = np.sum((y - daily['final_trend']) ** 2) / len(y)
X2 = np.hstack([X, np.ones((X.shape[0], 1))])
C = vy * np.linalg.inv(np.dot(X2.T, X2))
var = C.diagonal()
# -
# ### How does rain affect ridership?
ind = columns.index('prcp')
slope = final_model.coef_[ind]
error = np.sqrt(var[ind])
print('{0: .0f} +/- {1: .0f} daily crossings lost per cm of rain'.format(-slope * 10, error * 10))
# The model shows that for every cm of rain, about 300 cyclists stay home or use another mode of transport.
# ### How does temparature affect ridership?
ind1, ind2 = columns.index('tmin'), columns.index('tmax')
slope = final_model.coef_[ind1] + final_model.coef_[ind2]
error = np.sqrt(var[ind1] + var[ind2])
print('{0:.0f} +/- {1:.0f} riders per ten degrees Celsius'.format(10 * slope, 10 * error))
# ### How does daylight affect ridership?
ind = columns.index('daylight')
slopt = final_model.coef_[ind]
error = np.sqrt(var[ind])
print('{0:.0f} +/- {1:.0f} riders per hour of daylight'.format(slope, error))
# ### Is ridership increasing?
ind = columns.index('daycount')
slope = final_model.coef_[ind]
error = np.sqrt(var[ind])
print("{0:.2f} +/- {1:.2f} new riders per day".format(slope, error))
print("{0:.1f} +/- {1:.1f} new riders per week".format(7 * slope, 7 * error))
print("annual change: ({0:.0f} +/- {1:.0f})%".format(100 * 365 * slope / daily['total'].mean(),
100 * 365 * error / daily['total'].mean()))
| seattle_supervised_modeling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
# ## Bubble Sort
# Bubble sort is described in detail on [Wikipedia](https://en.wikipedia.org/wiki/Bubble_sort).
#
# Its complexity is $O(n^2)$.
def bubble_sort(array, verbose=False):
'''The bubble sort algorithm.
INPUT: array or list
OUTPUT: sorted array or list
'''
# setup
unsorted = True
list_length = len(array) - 1
# main logic
while unsorted:
changes = 0
for i in range(list_length):
if mylist[i] > mylist[i+1]:
mylist[i], mylist[i+1] = mylist[i+1], mylist[i]
changes += 1
if not changes:
unsorted = False
if verbose:
print(mylist)
mylist = np.random.randint(0, 50, 25)
mylist
bubble_sort(mylist, verbose=True)
| notebooks/Python/Programming_Problems/Bubble_Sort.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: drlnd
# language: python
# name: drlnd
# ---
# # Deep Deterministic Policy Gradients (DDPG)
# ---
# In this notebook, we train DDPG with OpenAI Gym's BipedalWalker-v2 environment.
#
# ### 1. Import the Necessary Packages
# +
import gym
import random
import torch
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
# %matplotlib inline
from agent import Agent
# -
# ### 2. Instantiate the Environment and Agent
env = gym.make('BipedalWalker-v2')
env.seed(10)
agent = Agent(state_size=env.observation_space.shape[0], action_size=env.action_space.shape[0], random_seed=10)
next(agent.actor_target.parameters()).is_cuda
# ### 3. Train the Agent with DDPG
#
# Run the code cell below to train the agent from scratch. Alternatively, you can skip to the next code cell to load the pre-trained weights from file.
# +
def ddpg(n_episodes=2000, max_t=700):
scores_deque = deque(maxlen=100)
scores = []
max_score = -np.Inf
for i_episode in range(1, n_episodes+1):
state = env.reset()
agent.reset()
score = 0
for t in range(max_t):
action = agent.act(state)
next_state, reward, done, _ = env.step(action)
agent.step(state, action, reward, next_state, done)
state = next_state
score += reward
if done:
break
scores_deque.append(score)
scores.append(score)
print('\rEpisode {}\tAverage Score: {:.2f}\tScore: {:.2f}'.format(i_episode, np.mean(scores_deque), score), end="")
if i_episode % 100 == 0:
torch.save(agent.actor_local.state_dict(), 'checkpoint_actor.pth')
torch.save(agent.critic_local.state_dict(), 'checkpoint_critic.pth')
print('\rEpisode {}\tAverage Score: {:.2f}'.format(i_episode, np.mean(scores_deque)))
return scores
scores = ddpg()
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# -
# ### 4. Watch a Smart Agent!
#
# In the next code cell, you will load the trained weights from file to watch a smart agent!
# +
agent.actor_local.load_state_dict(torch.load('checkpoint_actor.pth'))
agent.critic_local.load_state_dict(torch.load('checkpoint_critic.pth'))
state = env.reset()
agent.reset()
while True:
action = agent.act(state)
env.render()
next_state, reward, done, _ = env.step(action)
state = next_state
if done:
break
env.close()
# -
# ### 5. Explore
#
# In this exercise, we have provided a sample DDPG agent and demonstrated how to use it to solve an OpenAI Gym environment. To continue your learning, you are encouraged to complete any (or all!) of the following tasks:
# - Amend the various hyperparameters and network architecture to see if you can get your agent to solve the environment faster than this benchmark implementation. Once you build intuition for the hyperparameters that work well with this environment, try solving a different OpenAI Gym task!
# - Write your own DDPG implementation. Use this code as reference only when needed -- try as much as you can to write your own algorithm from scratch.
# - You may also like to implement prioritized experience replay, to see if it speeds learning.
# - The current implementation adds Ornsetein-Uhlenbeck noise to the action space. However, it has [been shown](https://blog.openai.com/better-exploration-with-parameter-noise/) that adding noise to the parameters of the neural network policy can improve performance. Make this change to the code, to verify it for yourself!
# - Write a blog post explaining the intuition behind the DDPG algorithm and demonstrating how to use it to solve an RL environment of your choosing.
| ddpg-bipedal/my/DDPG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
# This magic just sets up matplotlib's interactive mode
# %matplotlib inline
# So you have to explicitely import the module into the namespace
import matplotlib.pyplot as plt
def exact_solution(x0, v0, dt, m, t):
pass
def ordinary_verlet(xt, xt_1, dt, m):
return 2*xt - xt_1 + (dt**2)*(xt - xt**3)/m + (dt**4)/12*(-6*xt)/m, xt
def OV_operator_SHO(xt, xt_1, xt_2, xt_3, dt, m):
dt2_12= (( 12*xt - 24*xt_1 + 12*xt_2)*(-6*xt_2/m) - ( 12*xt_1 - 24*xt_2 + 12*xt_3 )*(-6*xt_1/m))/((xt_1-xt_1**3)/m*(-6*xt_2/m) - (xt_2-xt_2**3)/m*(-6*xt_1/m))
temp= (24*xt - 12*xt_1)*(-6*xt_1/m)+ ( 12*xt - 24*xt_1 + 12*xt_2 )*(-6*xt/m) + dt2_12*((xt-xt**3)/m*(-6*xt_1/m) - (xt_1-xt_1**3)/m*(-6*xt/m))
x_new= temp/12.0/(-6*xt_1/m)
# 12*xtss1*(-6*xt_1/m)
#x_new = 2*xt - xt_1 + (xt - xt**3)*(xt-2*xt_1+xt_2)/(xt_1-xt_1**3)
vt = (x_new-xt_1)/2/dt
te = 0.5 * m * vt**2 + (((xt **4)/4) - ((xt **2)/2))
return x_new, xt, xt_1, xt_2, te
def velocity_verlet(xt, vt, dt, m):
#xt - xt**3
xt_1 = xt + dt*vt + (dt**2)/2*(xt - xt**3)/m
vt_1 = vt + dt/2*( xt - xt**3 + xt_1 - xt_1**3)/m
te = 0.5 * m * vt_1**2 + (((xt_1 **4)/4) - ((xt_1 **2)/2))
return xt_1, vt_1, te
# -
selected_time
# +
dt=0.1
x0=-2.0
v0=0.0
m=1.0
total_time= 20
########ground truth##########
ground_truth_dt=0.001
time_ground = np.arange(0, total_time, ground_truth_dt)
iterations = int(total_time/ground_truth_dt)
x0_=x0
v0_=v0
vv_array_ground = np.array([x0_])
te = 0.5 * m * v0_**2 + (((x0_ **4)/4) - ((x0_ **2)/2))
vv_energy_ground = np.array([te])
vv_array_ground = exact_solution(x0_, v0_, time_ground, m)
selected_time = time_ground[::int(dt/ground_truth_dt)]
seleted_ground_truth = vv_array_ground[::int(dt/ground_truth_dt)]
####################################
time = np.arange(0, total_time, dt)
iterations = int(total_time/dt)
x0_=x0
v0_=v0
vv_array = np.array([x0_])
te = 0.5 * m * v0_**2 + (((x0_ **4)/4) - ((x0_ **2)/2))
vv_energy = np.array([te])
for i in range(iterations-1):
x0_, v0_, te = velocity_verlet(x0_, v0_, dt, m)
vv_array= np.append(vv_array,x0_)
vv_energy= np.append(vv_energy,te)
x0_=x0
v0_=v0
x1_=seleted_ground_truth[1]
ov_array = np.array([x0_, x1_])
for i in range(iterations-2):
x1_, x0_ = ordinary_verlet(x1_, x0_, dt, m)
ov_array= np.append(ov_array, x1_)
x0_=x0
v0_=v0
x1_=seleted_ground_truth[1]
x2_=seleted_ground_truth[2]
x3_=seleted_ground_truth[3]
ov_op_array = np.array([x0_, x1_, x2_, x3_])
ov_op_energy = np.array([vv_energy[0], vv_energy[1], vv_energy[2]])
for i in range(iterations-4):
x3_, x2_, x1_, x0_, te = OV_operator_SHO(x3_, x2_, x1_, x0_, dt, m)
ov_op_array= np.append(ov_op_array,x3_)
ov_op_energy= np.append(ov_op_energy,te)
fig, ax = plt.subplots(figsize=(20, 10))
plt.plot(time_ground, vv_array_ground, label='VV_ground')
plt.plot(time, vv_array, label='VV',marker='v', ms=5, markerfacecolor="None",linestyle='None')
plt.plot(time, ov_array, label='OV',marker='x', ms=5, markerfacecolor="None",linestyle='None')
plt.plot(time,ov_op_array, label='OV operator',marker='s', ms=5, markerfacecolor="None",linestyle='None')
#abs(vv_energy-vv_energy[0])/vv_energy[0]
#plt.plot(time, vv_energy, label='VV',marker='v', ms=5, markerfacecolor="None",linestyle='None')
#plt.plot(time[0:-1], ov_op_energy, label='OV',marker='v', ms=5, markerfacecolor="None",linestyle='None')
#plt.ylim(-5,5)
#plt.xlim(950,1000)
plt.legend()
print("OV error: "+str(np.mean(np.abs(ov_array-seleted_ground_truth))))
print("OV_operator error: "+str(np.mean(np.abs(ov_op_array-seleted_ground_truth))))
# +
fig, ax = plt.subplots(figsize=(20, 10))
plt.plot(abs(ov_array-vv_array), label='VV',marker='v', ms=5, markerfacecolor="None",linestyle='None')
#plt.plot(abs(ov_op_array-vv_array), label='VV',marker='v', ms=5, markerfacecolor="None",linestyle='None')
# -
print("VV error: "+str(np.sum((vv_array-exact_result)**2)))
print("OV error: "+str(np.sum((ov_array-exact_result)**2)))
print("OV_operator error: "+str(np.sum((ov_op_array-exact_result)**2)))
| src/md-codes/Voperator_Example-DW-Higher_order.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="-5ZlL0fO_Nme"
# # Documentation
#
#
# + [markdown] colab_type="text" id="XnX1dhtN_TUh"
# ## Introduction
#
# Differentiation is ubiquitous in almost all aspects of computer science, mathematics, and physics. It is used for numeric root-finding as in Newton's Method, and used for optimization with different forms of gradient descent.
# However, calculating analytic derivatives is difficult and can lead to exponentially growing abstract syntax trees, which makes finding the derivative infeasible in many cases. Calculating the derivative numerically using the limit definition runs into numeric instabilities due to limited machine precision. Automatic differentiation addresses both of these issues - it uses the chain rule and the fact that computers calculate any function as a sequence of elementary operations to find the derivative.
#
# The package can also evaluate the Hessian of multivariable functions and higher order derivatives of scalar functions. This is particularly useful for optimization problems, where convergence on a function minimum is often significantly faster when Hessian information is included in the root finding algorithm. This is useful for Control Theory applications, as well as many important algorithms used in robotics. Calculating the Hessian is useful for path planning, as well as for inverse kinematics. Since it is relatively difficult to get a closed form solution for the Hessian in complicated inverse kinematics problems, it is useful to be able to use Automatic Differentiation for the Hessian.
# + [markdown] colab_type="text" id="2L2ci5Tw_ZJF"
# ## Installation and Usage
#
#
# ### Installing from Source
#
# If you want the latest nightly version of AutoDiffX, clone from our github
# repository and install the latest version directly.
#
# ```bash
# git clone https://github.com/CS207-Project-Team-1/cs207-FinalProject autodiffx
# # # cd autodiffx
# pip install -r requirements.txt
# python3 setup.py install
# ```
#
# If you are working on a python virtual environment or Mac OSX or your user's
# python distribution, this should work. If editing the system python, you may
# need to run the last command with root permissions by adding `sudo`.
#
# ### Installing from pip
#
# For the stable version, you can install our package from PyPI.
#
# ```bash
# pip install autodiffx
# ```
#
# ## Testing
#
# All of the tests are run using pytest. To run pytest, you want to be in the
# root directory of the repository. To ensure that `pytest` gets the imports
# correct, you want to run it such that it adds the current path to `PYTHONPATH`.
# The easiest way to do so is:
#
# ```bash
# python -m pytest
# ```
#
# This should run all of the tests for the package.
#
# Currently, our only module that we actually use lives in the `ad/` folder. The tests can be found in the different test files in `ad/tests`.
#
#
#
# + [markdown] colab_type="text" id="QSxx1_h9_UTc"
# ## Background
#
# **Automatic Differentiation**
#
# Automatic differentiation relies heavily on the principles of chain rule differentiation. A graph of elementary functions is built to calculate the values of more complex functions. Using the chain rule on the graph of elementary functions, the value of the derivative at each node can also be calculated. This gives us the ability to calculate the values of functions and their derivatives, no matter how complex, to near machine precision (a significant advantage compared to alternatives such as finite differences).
#
# The chain rule tells us that:
# \begin{align}
# \frac{d}{dx}(f(g(x))) &= f'(g(x))g'(x)\\
# \end{align}
#
# Since each step in our graph is just a combination of linear operations, we can find the derivative at a node by considering the value and derivative of the expressions at the previous node. By starting with an initial 'seed' vector for the derivative (often set to 1), we can find the derivative in any desired search direction.
#
# Below is an example of constructing a graph to find the exact values of a function and its derivative. The function we used was:
#
# $$f\left(x, y, z\right) = \dfrac{1}{xyz} + \sin\left(\dfrac{1}{x} + \dfrac{1}{y} + \dfrac{1}{z}\right)$$
#
# We worked through this, starting with trace elements $x_1$ for $x$, $x_2$ for $y$ and $x_3$ for $z$. We wanted to solve this function at $(x, y, z) = (1, 2, 3)$.
#
# | Trace | Elementary Function | Current Value | Elementary Function Derivative | $\nabla_{x}$ Value | $\nabla_{y}$ Value | $\nabla_{z}$ Value |
# | :---: | :-----------------: | :-----------: | :----------------------------: | :-----------------: | :-----------------: | :-----------------: |
# | $x_{1}$ | $x$ | 1 | $\dot{x}$ | 1 | 0 | 0 |
# | $x_{2}$ | $y$ | 2 | $\dot{y}$ | 0 | 1 | 0 |
# | $x_{3}$ | $z$ | 3 | $\dot{z}$ | 0 | 0 | 1 |
# | $x_{4}$ | $1/x_{1}$ | 1 | $-\dot{x}_{1}/x_{1}^{2}$ | $-1$ | $0$ | $0$ |
# | $x_{5}$ | $1/x_{2}$ | 1/2 | $-\dot{x}_{2}/x_{2}^{2}$ | $0$ | $-1/4$ | $0$ |
# | $x_{6}$ | $1/x_{3}$ | 1/3 | $-\dot{x}_{3}/x_{3}^{2}$ | $0$ | $0$ | $-1/9$ |
# | $x_{7}$ | $x_{4} + x_{5} + x_{6}$ | 11/6 |$\dot{x}_{4} + \dot{x}_{5} + \dot{x}_{6}$ | -1 | -0.25 | -0.11 |
# | $x_{8}$ | $sin(x_{7})$ | 0.966 |$\dot{x}_{7}cos(x_{7})$ | 0.260 | 0.065 | 0.029 |
# | $x_{9}$ | $x_{4}x_{5}x_{6}$| 1/6 |$\dot{x}_{4}x_{5}x_{6} + \dot{x}_{5}x_{4}x_{6} + \dot{x}_{6}x_{4}x_{5} $ |-0.167 | -0.083 | -0.056 |
# | $x_{10}$ | $x_{8} + x_{9}$ | 1.132 |$\dot{x}_{8} + \dot{x}_{9}$ | 0.093| -0.018 | -0.027 |
#
# This isn't a very complicated function, but it shows how we can use the most basic of functions to create a graph allowing us to find exact values and gradients.
#
# **Higher-Order Derivatives**
#
# An effective approach to high-order automatic differention can be obtained by considering the calculus of a Taylor series. If we define $f_k$ as follows:
#
# $$f_k = f_{k}(x_0) = \dfrac{f^{(k)}(x_0)}{k!}$$
#
# We can show that basic arithmetic operations are as follow, where $f$ and $g$ are separate functions with the same input variable. Full derivations of these basic operations can be found [here](https://www.sintef.no/globalassets/project/evitameeting/2010/ad2010.pdf):
#
# $$(f + g)_k = f_k + g_k$$
#
# $$(f - g)_k = f_k - g_k$$
#
# $$(f \times g)_k = \sum_{i = 0}^{k} f_{i}g_{k-i} $$
#
# $$(f \div g)_k = \dfrac{1}{g_0} \left(f_k - \sum_{i = 0}^{k - 1} (f \div g)_{i} g_{k-i}\right) $$
#
# $$ (e^g)_k = \dfrac{1}{k} \sum_{i = 1}^{k} ig_{i}(e^g)_{k-i}$$
#
# **Hessian**
#
#
# Our package implements the Hessian for functions of the form
# $$f: \mathbb{R}^m \to \mathbb{R}$$
# In this case, the Hessian should be a square matrix. Since the Hessian will be a square matrix with entries of the form
# $$\frac{\partial^2 f}{\partial x_i \partial x_j}$$
# for some $i, j$, we return a dictionary of dictionaries for the Hessian. This makes it easy to index the Hessian as ```hessian[x][y]``` to get the second order derivative with respect to x and y.
# + [markdown] colab_type="text" id="z13vUmhoQFdB"
# ## Simple Usage
#
# Our package supports scalar functions with scalar or multivariable inputs.
#
# **Scalar Input**
#
# Suppose that we wanted to find the derivative of the following function, $f(x)$:
#
# $$f(x) = x \exp(\cos(x) + x^2)$$
#
# First, we have to allocate a Variable for our input variable $x$. We can do that, and give it a string identifier `"x"` for convenience later.
#
# ```python
# import ad
# x = ad.Variable('x')
# ```
# Now, we have to define our function. Our module `ad` has a lot of built in functions, and all of the regular operators `+`, `-`, `*`, `/`, `**` should also work. We can make our function $f(x)$ by just writing it out in code.
#
# ```python
# f = x * ad.Exp(ad.Cos(x) + x ** 2)
# ```
#
# Now, we've defined our function in terms of our input variable $x$. Now, in order to evaluate our function or evaluate the derivative at a specific point we actually have to provide a value for the input. Since we later plan on handling multiple variable inputs, we pass the input in as a dictionary. To evaluate the function, we use the function ```eval```. For the derivative at a point, we use ```d```. Suppose that we wanted to evaluate the function $f$ and its derivative at $x = 0$ and $x = 1$. We can just run:
#
# ```python
# >>> f.eval({x: 0})
# 2.718281828459045
#
# >>> f.d({x: 0})
# 1.0
#
# >>> f.eval({x: 1})
# 5.666000617166735
#
# >>> f.d({x: 1})
# 6.405697099891925
# ```
#
# It is also possible to evaluate higher order derivatives of functions with scalar inputs. This can be done using ```f.d_n(n, val)``` where n and val are arguments expressing the desired order of differentiation and the value at which the derivative should be evaluated. We can run:
#
# ```python
# >>> f.d_n(n = 3, val = 2)
# 1902.7256925837773
# ```
#
# **Multi-Variable Input**
#
# Suppose that we wanted to find the derivative of the following function, $f(x, y)$:
#
# $$f(x) = x \exp(\cos(y) + x^2)$$
#
# We have to allocate a variables for our inputs $x$ and $y$. We can do that, and give them string identifiers `"x"` and `"y"` for convenience later.
#
# ```python
# import ad
# x = ad.Variable('x')
# y = ad.Variable('y')
# ```
# We now define our function.
#
# ```python
# f = x * ad.Exp(ad.Cos(y) + x ** 2)
# ```
#
# We can evaluate our function and it's derivative exactly as we evaluated our scalar input example earlier. Suppose that we wanted to evaluate the function $f$ and its derivative at $x = 0$ and $y = 1$. We can just run:
#
# ```python
# >>> f.eval({x: 0, y : 1})
# 1.7165256995489035
#
# >>> f.d({x: 0, y : 1})
# {y: -1.4444065708474794, x: 1.0}
# ```
#
# Using the Hessian is very similar to calling the derivative of a multivariable function. If the function depends on $n$ input variables, the package will return an $n \times n$ matrix in the form of a dictionary of dictionaries, where each sub-dictionary refers to one of the rows in the matrix.
#
# Hence, for our multivariable example above, $f(x, y)$:
#
# $$f(x) = x \exp(\cos(y) + x^2)$$
#
# We can evaluate the Hessian by calling:
#
# ```python
# >>> f.hessian({x: 0, y : 1})
# {y: {y: 0.28798342608583105, x: 0.0}, x: {y: 0.0, x: 3.433051399097807}}
# ```
# If we want the specific second order derivatives, we can evaluate specific elements from the Hessian by calling the keys. For example, if we wanted to find the second order derivative of our function with respect to $x$, we would call:
#
# ```python
# >>> f.hessian({x: 0, y : 1})[x][x]
# 3.433051399097807
# ```
#
# More complicated demonstrations (scalar input Newton's Method, multivariable input Newton's Method with Hessian) can be found in Jupyter notebooks at the top level directory.
# + [markdown] colab_type="text" id="1cfu_jYk_iK2"
# ## Software Organization
# + [markdown] colab_type="text" id="vF3Ni2RGPJTt"
# ## Directory structure
#
# ```bash
# cs207project
# ├── LICENSE
# ├── README
# ├── docs
# │ ├── Documentation
# │ ├── Milestone1.ipynb
# │ ├── Milestone2.ipynb
# │ └── SETUP.md
# ├── ad
# │ ├── __init__.py
# │ ├── ad.py
# │ ├── simple_ops.py
# │ ├── activation_ops.py.../
# │ └── tests
# │ ├── test_complex_ops.py
# │ ├── test_d_expr.py
# │ ├── test_expression.py
# │ ├── test_high_order.py
# │ ├── test_multivar_hessian.py
# │ ├── test_simple_hessian.py
# │ ├── test_simple_ops.py
# │ └── test_vector.py
# ├── demos
# │ ├── Newton_Method_Demonstration.ipynb
# │ ├── Hessian_Demonstration.ipynb
# │ ├── Higher_Order_Demonstration.ipynb
# ├── requirements.txt
# └── setup.py
# ```
#
# ## Modules
#
# `ad`: Implementation of core structures used in our graph-based automatic differentiation library. +, -, *, /, and exponentiation is also implemented here to support simple operator overloading.
#
# `simple_ops`: Unary operations including Sin(x), Cos(x), Tan(x), Sinh(x), Cosh(x), Tanh(x), Exp(x), Log(x), Arcsin(x), Arccos(x), Arctan(x), Logistic(x).
#
# ## Test
#
# See Testing.
#
# ## Installment
#
# See Installation and Usage
#
#
# + [markdown] colab_type="text" id="iUlwF_YJ_mBU"
# ## Implementation Details
#
# We implemented our automatic differentiation using a graph based structure. First, the user will build up their function in terms of elementary operations. Then, the user will be able to feed a value for their input variables, and the package will calculate the derivatives at each step.
# + [markdown] colab_type="text" id="dcrJbb9E_4ZC"
# ### Automatic Differentiation Basics
# + [markdown] colab_type="text" id="tvtimCjqAAdL"
# Our core data structure is the computational graph. Every node in the computational graph would be an "Expression", which is our core class.
#
# Based on "Expression", we have "Variable(Expression)", "Constant(Expression)", "Unop(Expression)" and "Biop(Expression)". "Unop" is for unary operations such as log and power. "Biop" is for binary operations such as addition. For "Expression" and its subclasses, there are two important attributes: "grad", a boolean variable indicating whether we want to calculate the derivative or not; children, a list of nodes pointing to current one, the number of children is one for "Unop" and the number of chilren for "Biop" is two.
#
# The elementary functions we support are:
#
# Unary operations:
#
# * Sin(x)
# * Cos(x)
# * Tan(x)
# * Sinh(x)
# * Cosh(x)
# * Tanh(x)
# * Exp(x)
# * Log(x)
# * Arcsin(x)
# * Arccos(x)
# * Arctan(x)
# * Logistic(x)
#
# Binary operations:
#
# * Addition (+)
# * Substraction (-)
# * Multiplication (*)
# * Division (/)
# * Power(x, n)
#
# We will mainly be using `numpy` as an external dependency for mathematical and vector operations.
#
#
#
# + [markdown] colab_type="text" id="yPdt0eOf_7xP"
# ### Multivariate Inputs
# + [markdown] colab_type="text" id="V6K5gmNpFFhL"
# We want to also support getting the Jacobian when we have more than one input for a scalar function. We are able to support functions of the form
# $$f: \mathbb{R}^m \to \mathbb{R}$$
# Our implementation for multivariate inputs is by using a `dict` to hold the different partial derivatives. We can go through an example. Suppose that a user wanted the Jacobian of a function:
#
# $$f(x, y, z) = x + y \cos(yz)$$
#
# Then, we are able to find the Jacobian in code using the following syntax.
#
# ```python
# >>> x = ad.Variable('x')
# >>> y = ad.Variable('y')
# >>> z = ad.Variable('z')
#
# >>> f = x + y * ad.Cos(y * z)
#
# >>> f.eval({x:6, y:1, z:0})
# 7
# >>> f.d({x:6, y:1, z:0})
# {x: 1, y: 1, z: 0}
# ```
#
# -
# ### Extension Feature - Higher Order Derivatives and Hessian
# + [markdown] colab_type="text" id="4pZ4XasAJm8_"
# **Higher Order Derivatives**
#
# See background.
#
# **Hessian**
#
# See background.
#
# An example function and calculated Hessian will be look as follows.
#
# $$f: \mathbb{R}^3 \to \mathbb{R}$$
# $$f(x, y, z) = xy + z$$
#
# The code uses the function `hessian()` to calculate the Hessian at a certain point.
#
# ```python
# >>> x = ad.Variable('x')
# >>> y = ad.Variable('y')
# >>> z = ad.Variable('z')
#
# >>> f = x * y + z
#
# >>> f.eval({x:1, y:1, z:1})
# 2
#
# >>> f.hessian({x:1, y:1, z:1})
# {x: {x: 0, y: 1, z:} y: {x: 1, y: 0, z: 0} z: {x: 0, y: 0, z: 0}}
# ```
# -
# ### Future Extensions
# **Multivariate Outputs**
#
# Currently, our package only fully supports scalar functions with scalar or multivariate input.
#
# However, the next step after supporting multivariate inputs is to also support multivariate outputs. We will create a wrapper class `ad.Vector` that essentially wraps multiple different `ad.Expression` instances together. This will allow us to combine multiple scalar functions into a multivariate function. For example, suppose that a user wanted to find the Jacobian of a function in the form:
# $$f: \mathbb{R}^3 \to \mathbb{R}^3$$
# and suppose that the function was in the form:
# $$f(x, y, z) =
# \begin{pmatrix}
# x + y + z \\
# \cos(yz) \\
# \exp(x - y - z)
# \end{pmatrix}
# $$
# This could be represented in code as follows:
#
# ```python
# >>> x = ad.Variable('x')
# >>> y = ad.Variable('y')
# >>> z = ad.Variable('z')
#
# >>> f1 = x + y + z
# >>> f2 = ad.Cos(y * z)
# >>> f3 = ad.Exp(x - y - z)
#
# >>> f = ad.Vector(f1, f2, f3)
# >>> f.eval({x:0, y:0, z:0})
# [0, 1, 1]
#
# >>> f.d({x:0, y:0, z:0})
# {x: [1, 0, 1], y: [1, 0, -1], z: [1, 0, -1]}
# ```
# ## Future
# An automatic differentiation package with Hessian and higher-order scalar derivative function will have many potential scientific applications. Root finding algorithms with relatively rapid convergence properties rely on higher order derivative information being available. These algorithms are ubiquitous across many fields, from economics to physics, and our package makes it significantly easier for users to work with these algorithms, since the big work of calculating the Hessian (often the most complex part of implementing a method such as the Newton Method) is done automatically.
#
# **Robotics**
#
# In particular, we are interested in its potential use in Robotics, where being able to calculate the Hessian to machine precision quickly is critically important for path planning and inverse kinematics. Recent research work has shown that using the exact Hessian in these systems gives a significant performance boost when compared with using approximate methods (such as BFGS). The research focussed on a variety of fields beyond Robotics, stretching to motion capture, character animation and computer graphics. The [paper](http://image.diku.dk/kenny/download/erleben.andrews.17.pdf) (Erleben and Andrews) showed that 'using exact Hessians can give performance advantages and higher accuracy compared to standard numerical methods used for solving these problems.'
#
# **Control Theory**
#
# For many control theory problems, particularly problems requiring online predictions, having the ability to quickly compute the Hessian of the control function is very important. Researchers at Warwick University [found](https://warwick.ac.uk/fac/sci/physics/research/condensedmatt/imr_cdt/students/david_goodwin/publications/imrcdt_southampton_mar16.pdf) that using Hessian information during the control of magnetic resonance imaging led to increased accuracy in the resulting images. Similar uses of rapid Hessian calculation are possible across many different control applications.
#
# **Computational Cost**
#
# Calculating the Hessian becomes very expensive as the number of variables used by a function increases. For larger problems, calculating the full Hessian stops making sense, since the benefit derived from having full accuracy are outweighed by the cost of achieving such a result. For larger problems, we would like to implement approximate Hessian algorithms (such as BFGS), to make this package universally useful for researchers needing Hessian information.
| docs/Documentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Formatting and Positioning Images as Logos
# +
import plotly.plotly as py
import plotly.graph_objs as go
data = [
go.Bar(
x=['-35.3', '-15.9', '-15.8', '-15.6', '-11.1',
'-9.6', '-9.2', '-3.5', '-1.9', '-0.9',
'1.0', '1.4', '1.7', '2.0', '2.8', '6.2',
'8.1', '8.5', '8.5', '8.6', '11.4', '12.5',
'13.3', '13.7', '14.4', '17.5', '17.7',
'18.9', '25.1', '28.9', '41.4'],
y=['Designers, musicians, artists, etc.',
'Secretaries and administrative assistants',
'Waiters and servers', 'Archivists, curators, and librarians',
'Sales and related', 'Childcare workers, home car workers, etc.',
'Food preparation occupations', 'Janitors, maids, etc.',
'Healthcare technicians, assistants. and aides',
'Counselors, social and religious workers',
'Physical, life and social scientists', 'Construction',
'Factory assembly workers', 'Machinists, repairmen, etc.',
'Media and communications workers', 'Teachers',
'Mechanics, repairmen, etc.', 'Financial analysts and advisers',
'Farming, fishing and forestry workers',
'Truck drivers, heavy equipment operator, etc.','Accountants and auditors',
'Human resources, management analysts, etc.', 'Managers',
'Lawyers and judges', 'Engineers, architects and surveyors',
'Nurses', 'Legal support workers',
'Computer programmers and system admin.', 'Police officers and firefighters',
'Chief executives', 'Doctors, dentists and surgeons'],
marker=dict(
color='rgb(253, 240, 54)',
line=dict(color='rgb(0, 0, 0)',
width=2)
),
orientation='h',
)
]
layout = go.Layout(
images=[dict(
source="https://raw.githubusercontent.com/cldougl/plot_images/add_r_img/vox.png",
xref="paper", yref="paper",
x=1, y=1.05,
sizex=0.2, sizey=0.2,
xanchor="right", yanchor="bottom"
)],
autosize=False, height=800, width=700,
bargap=0.15, bargroupgap=0.1,
barmode='stack', hovermode='x',
margin=dict(r=20, l=300,
b=75, t=125),
title='Moving Up, Moving Down<br><i>Percentile change in income between childhood and adulthood</i>',
xaxis=dict(
dtick=10, nticks=0,
gridcolor='rgba(102, 102, 102, 0.4)',
linecolor='#000', linewidth=1,
mirror=True,
showticklabels=True, tick0=0, tickwidth=1,
title='<i>Change in percentile</i>',
),
yaxis=dict(
anchor='x',
gridcolor='rgba(102, 102, 102, 0.4)', gridwidth=1,
linecolor='#000', linewidth=1,
mirror=True, showgrid=False,
showline=True, zeroline=False,
showticklabels=True, tick0=0,
type='category',
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
# +
import plotly.plotly as py
import plotly.graph_objs as go
fig = py.get_figure('https://plotly.com/~Dreamshot/8152/', raw=True)
fig['layout']['yaxis']['tickangle'] = 0
fig = go.Figure(fig)
fig.layout.images = [dict(
source="https://raw.githubusercontent.com/cldougl/plot_images/add_r_img/accuweather.jpeg",
xref="paper", yref="paper",
x=0.1, y=1.05,
sizex=0.4, sizey=0.4,
xanchor="center", yanchor="bottom"
)]
py.iplot(fig, fileopt='overwrite', filename='Logos/Florida_Rainfall_AccuWeather')
# +
import plotly.plotly as py
import plotly.plotly as py
import plotly.graph_objs as go
fig = py.get_figure('https://plotly.com/~Dreamshot/8160/', raw=True)
for j in range(len(fig['data'])):
del fig['data'][j]['autobinx']
del fig['data'][j]['autobiny']
fig = go.Figure(fig)
fig.layout.images = [dict(
source="https://raw.githubusercontent.com/cldougl/plot_images/add_r_img/bleacherreport.png",
xref="paper", yref="paper",
x=0.5, y=-0.35,
sizex=0.3, sizey=0.3,
xanchor="center", yanchor="top"
)]
py.iplot(fig, fileopt='overwrite', filename='Logos/Top_Earners_BleacherReport')
# +
import plotly.plotly as py
fig = py.get_figure('https://plotly.com/~Dreamshot/8158/')
fig.layout.images = [dict(
source="https://raw.githubusercontent.com/cldougl/plot_images/add_r_img/theverge.png",
xref="paper", yref="paper",
x=0.1, y=1.0,
sizex=0.2, sizey=0.3,
xanchor="center", yanchor="bottom"
)]
fig.layout.legend.orientation = 'h'
py.iplot(fig, fileopt='overwrite', filename='Logos/Apple_Labor_Violations_TheVerge')
# +
import plotly.plotly as py
fig = py.get_figure('https://plotly.com/~Dreamshot/8155/')
fig.layout.images = [dict(
source="https://raw.githubusercontent.com/cldougl/plot_images/add_r_img/politico.png",
xref="paper", yref="paper",
x=0.1, y=-0.2,
sizex=0.4, sizey=0.4,
xanchor="center", yanchor="bottom"
)]
py.iplot(fig, fileopt='overwrite', filename='Logos/Foreign_Policy_Politico')
# -
# #### Reference
# See https://plotly.com/python/images/ for more examples of adding images<br>
# and https://plotly.com/python/reference/#layout-images for more information and chart attribute options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'logos.ipynb', 'python/logos/', 'Add Logos to Charts',
'How to add images as logos to Plotly charts.',
title = 'Add Logos to Charts | plotly',
name = 'Logos',
has_thumbnail='false', thumbnail='thumbnail/your-tutorial-chart.jpg',
language='python', page_type='example_index',
display_as='style_opt', order=6,
ipynb= '~notebook_demo/92')
# -
| _posts/python-v3/advanced/logos/logos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
# # MPSGE.jl
#
# ## <NAME>, UC Berkeley, <EMAIL>
# This is the demo that I gave during my talk at the WiNDC Annual Meeting 2021. Please note that the software I show here is very much work in progress, so things will likely change significantly going forward.
# First we load the MPSGE.jl package.
using MPSGE
# Then we create a new model and store it in the global variable `m`:
m = Model()
# Now we start to add things to this new model. The first addition is a parameter named `endow`, with a value of `1`:
@parameter(m, endow, 1.0)
# Then we add three sectors to the model:
@sector(m, X)
@sector(m, Y)
@sector(m, U)
# And five commodities:
@commodity(m, PX)
@commodity(m, PY)
@commodity(m, PU)
@commodity(m, PL)
@commodity(m, PK)
# Next, we specify one consumer, including a benchmark value:
@consumer(m, RA, benchmark=150.)
# Then we add three production functions:
@production(m, X, 1, PX, 100, [Input(PL, 50), Input(PK, 50)])
@production(m, Y, 1, PY, 50, [Input(PL, 20), Input(PK, 30)])
@production(m, U, 1, PU, 150, [Input(PX, 100), Input(PY, 50)])
# And finally, we add one demand function:
@demand(m, RA, PU, [Endowment(PL, :(70 * $endow)), Endowment(PK, 80.)])
# At this point we can view a nice algebraic version of our model, just to check what we have:
algebraic_version(m)
# As a first step, we will check whether the benchmark values we provied are actually a model solution:
solve!(m, cumulative_iteration_limit=0)
# Next we change the value of the parameter of our model, and then resolve the model:
set_value(endow, 1.1)
solve!(m)
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Pre-processing amplicon data and creating community trajectory figure
# ## (1) Pre-processing data
# **Housekeeping**
library(tidyverse)
library(reshape2)
library(vegan)
# **Read in community composition data**
species_composition = read.table("../../../data/amplicon/strain_table.txt",
sep = "\t",
header = T,
row.names = 1)
# Inspect data:
dim(species_composition)
head(species_composition)
# There should be 193 (192 samples plus stock --> 193) but there are only 191 samples. This indicates that two samples were lost during bioinformatic analysis due to poor quality sequence data.
# **Remove samples with very low number of reads (< 100)**
x = colSums(species_composition) >= 100
ncol(species_composition)
species_composition = species_composition[,x]
ncol(species_composition) # all samples ok
# **Convert read counts to relative abundance of species**
# +
species_composition = as.data.frame(decostand(t(species_composition), method = "total"))
head(species_composition)
# Check
head(rowSums(species_composition)) # ok
# -
# **Create metadata file**
# Extract metadata:
# +
# extract treatment string
species_composition$TREATMENT = as.character(rownames(species_composition))
# split treatment into subtreatments separated by underline
metadata = data.frame(do.call('rbind', strsplit(as.character(species_composition$TREATMENT),'_',fixed=TRUE)))
# name columns by subtreatment
colnames(metadata) = c("Time_point", "Immigration", "Streptomycin", "Replicate")
# inspect
head(metadata)
# -
# Edit metadata:
# +
# convert subtreatments into character type to allow downstream grepl and gsub commands
metadata$Time_point = as.character(metadata$Time_point)
metadata$Immigration = as.character(metadata$Immigration)
metadata$Streptomycin = as.character(metadata$Streptomycin)
metadata$Replicate = as.character(metadata$Replicate)
# mark all subtreatment columns as "stock" for stock community
metadata$Time_point = ifelse(grepl("STOCK", metadata$Time_point), "stock", metadata$Time_point)
metadata$Immigration = ifelse(grepl("stock", metadata$Time_point), "stock", metadata$Immigration)
metadata$Streptomycin = ifelse(grepl("stock", metadata$Time_point), "stock", metadata$Streptomycin)
metadata$Replicate = ifelse(grepl("stock", metadata$Time_point), "stock", metadata$Replicate)
# remove subtreatment-specifying characters from subtreatment fields
metadata$Time_point = gsub("T", "", metadata$Time_point)
metadata$Immigration = gsub("I", "", metadata$Immigration)
metadata$Streptomycin = gsub("AB", "", metadata$Streptomycin)
metadata$Replicate = gsub("REP", "", metadata$Replicate)
# give metadata table same rownames as community composition table
metadata$TREATMENT = rownames(species_composition)
# inspect
head(metadata)
# -
# Convert time point from transfers (every four days) to days
# +
# create day column and convert transfers to days
metadata$Day = metadata$Time_point
metadata$Day = gsub("4", 16, metadata$Day)
metadata$Day = gsub("8", 32, metadata$Day)
metadata$Day = gsub("12", 48, metadata$Day)
# remove old time point column
metadata = metadata[,-1]
# reorder columns
metadata = metadata[,c("Immigration", "Streptomycin", "Replicate", "Day", "TREATMENT")]
# inspect
head(metadata)
# -
# **Merge species composition with metadata**
# +
comm_data = merge(species_composition, metadata)
# inspect
head(comm_data)
# -
# **Write out relative abundance data and metadata**
# +
# write out relative abundance data leaving out stock community
write.table(comm_data[comm_data$TREATMENT != "STOCK", 1:30],
"../../../data/amplicon/rel_abund.txt",
row.names = F,
sep = "\t")
# write out metadata leaving out stock community
write.table(comm_data[comm_data$TREATMENT != "STOCK", -c(2:30)],
"../../../data/amplicon/meta.txt",
row.names = F,
sep = "\t")
# -
# ## (2) Create community trajectory figure
# **For visualization purposes alone, add same initial community composition (stock) to all communities over time**
# +
# replicate stock (for day zero) for all treatment combinations
stock = comm_data[comm_data$Immigration == "stock",]
stock$TREATMENT = "stock"
stock = rbind(stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock,
stock, stock, stock, stock, stock, stock, stock, stock)
stock$Replicate = rep(c(1, 2, 3, 4, 5, 6, 7, 8), 8)
stock$Immigration = rep(c(0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 1, 1, 1, 1), 4)
stock$Streptomycin = c(0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0,
4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4,
16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16, 16,
128, 128, 128, 128, 128, 128, 128, 128, 128, 128, 128, 128, 128, 128, 128, 128)
# inspect
head(stock)
dim(stock)
# -
# **Add initial composition data to later species composition data over time**
# +
comm_data = merge(comm_data[comm_data$TREATMENT != "STOCK",], stock, all = T)
# inspect
head(comm_data)
dim(comm_data)
# -
# **For visualization purposes alone, collapse rare species into one species group entitled "Others"**
# +
# compute sum of rare species
# create function
colMax = function(data) sapply(data, max, na.rm = TRUE)
# extract species that fail to reach 5 % abundance in at least one community and time point
rare_species = colMax(comm_data[,2:30]) < 0.05
# inspect number of species left
length(rare_species[rare_species == F])
# there 18 species that reach 5 % abundance at least once
# aggregate frequencies of rare species into new "Others" column
comm_data2 = comm_data[,2:30]
comm_data$Others = rowSums(comm_data2[,colnames(comm_data2) %in% names(rare_species[rare_species == T])])
# remove individual entries for aggregated rare species
comm_data = comm_data[,!(colnames(comm_data) %in% names(rare_species[rare_species == T]))]
# inspect
dim(comm_data)
head(comm_data)
# -
# **For visualization purposes alone, sqrt transform and rescale (0-1) data**
# +
# separate data from metadata
meta = comm_data[,c(1,(ncol(comm_data)-4):(ncol(comm_data)-1))]
data = comm_data[,-c(1,(ncol(comm_data)-4):(ncol(comm_data)-1))]
# sqrt transform and rescale
data = sqrt(data)
data = decostand(data, method = "total")
# re-merge
comm_data = cbind(meta[,1, drop = F], data, meta[,-1])
# inspect
dim(comm_data)
head(comm_data)
# -
# **Convert table into narrow format required for plotting with ggplot**
# +
comm_data_melt = melt(comm_data,
id.vars = c("TREATMENT", "Replicate", "Streptomycin", "Immigration", "Day"))
# give molten columns descriptive names
colnames(comm_data_melt) = c("TREATMENT", "Replicate", "Streptomycin", "Immigration", "Day", "Species", "Abundance")
# inspect
head(comm_data_melt)
# -
# **Assign subtreatments into correct type for plotting**
# +
# first convert "stock" into day 0
comm_data_melt$Day = gsub("stock", 0, comm_data_melt$Day)
# assign correct types
comm_data_melt$Day = as.numeric(comm_data_melt$Day)
comm_data_melt$Immigration = as.numeric(comm_data_melt$Immigration)
comm_data_melt$Streptomycin = as.numeric(comm_data_melt$Streptomycin)
# -
# **Assign species order by decreasing abundance**
# +
# Define species order by abundance
x = names(sort(colSums(comm_data[,c(2:19)]), decreasing = T))
comm_data_melt$Species = factor(comm_data_melt$Species, levels = c(x, "Others"))
# -
# **Create colour palette for species**
levels(comm_data_melt$Species)
mypalette = c("#006884",
"#CF97D7",
"#FFD08D",
"#FA9D00",
"#ED0026",
"#89DBEC",
"#6E006C",
"#00909E",
"#B00051",
"#7570B3",
"#E7298A",
"#5B5B5B",
"midnightblue",
"#7FB005",
"#CCFF00",
"#CCCC00",
"#CCEBC5",
"purple",
"#000000")
# **Rename immigration and antibiotic level treatments**
# +
comm_data_melt$Immigration = factor(comm_data_melt$Immigration,
levels = c(0, 1),
labels = c("No immigration",
"Immigration"))
comm_data_melt$Streptomycin = factor(comm_data_melt$Streptomycin,
levels = c(0, 4, 16, 128),
labels = c("No\nantibiotic",
"Low\nantibiotic",
"Intermediate\nantibiotic",
"High\nantibiotic"))
# -
# **Plot code**
# +
# Plot code
p1 = ggplot() +
geom_area(data = comm_data_melt, aes(x = Day, y = Abundance, fill = Species)) +
facet_grid(Immigration*Streptomycin~Replicate) +
scale_y_continuous(expand = c(0, 0), breaks = c(0, 0.2, 0.4, 0.6, 0.8, 1.0)) +
scale_fill_manual(values = c(mypalette), guide =
guide_legend(label.theme = element_text(angle = 0, face = "italic"), ncol = 1),
labels = c(gsub("_", " ", levels(comm_data_melt$Species)))) +
geom_rect(data = comm_data_melt[comm_data_melt$Streptomycin == "No\nantibiotic",], xmin=16, xmax=32, ymin=0, ymax=Inf, alpha=0.015, fill = "#D3D3D3") +
geom_rect(data = comm_data_melt[comm_data_melt$Streptomycin == "Low\nantibiotic",], xmin=16, xmax=32, ymin=0, ymax=Inf, alpha=0.015, fill = "#cd6090") +
geom_rect(data = comm_data_melt[comm_data_melt$Streptomycin == "Intermediate\nantibiotic",], xmin=16, xmax=32, ymin=0, ymax=Inf, alpha=0.015, fill = "#8f4364") +
geom_rect(data = comm_data_melt[comm_data_melt$Streptomycin == "High\nantibiotic",], xmin=16, xmax=32, ymin=0, ymax=Inf, alpha=0.015, fill = "#522639") +
scale_x_continuous(breaks = c(0, 16, 32, 48), expand = c(0, 0)) +
theme_classic() +
ylab("Species abundance (sqrt transformed and rescaled 0-1)") +
theme(panel.spacing = unit(1, "lines"),
legend.title=element_blank(),
strip.background = element_blank())
p1
# -
# **Save figure**
# +
# ggsave("../../../manuscript/figures/fig2_community_trajectories.pdf",
# width = 14,
# height = 10)
| src/notebooks/amplicon_data/1_pre_processing_amplicon_data_and_community_trajectory_Figure_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import argparse
import sys
import os
import time
import copy
import tensorflow as tf
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import dataset
from models.conv2_dense2_dropout import Model
#from models.dense3 import Model
from helpers.os_utils import os_info
from helpers.history import ExpHistory
from helpers.estimator_utils import create_model_fn, split_datasource
# -
# You may want to rerun and should close the session, if one is open.
try:
sess.close()
except NameError:
print("Don't worry. Need to ignore this error once")
sess = tf.InteractiveSession()
# ### Get the history and the runtime context
# +
tf.logging.set_verbosity(tf.logging.INFO)
HIST_FILE_NAME = 'experiment_history.csv'
history = ExpHistory(HIST_FILE_NAME)
localtime = time.asctime(time.localtime(time.time()))
user = os.environ.get('USER', os.environ.get('USERNAME', 'anonymous'))
print("\n\n")
print("Welcome, %s, it's %s, and you'll be working with Tensorflow version %s" % (user, localtime, tf.__version__))
rt=os_info()
this_os = rt['os']
this_node = rt['node']
this_machine = rt['machine']
this_cuda = rt['cuda']
print("Your current runtime: \n node: %s, \n os: %s, \n machine: %s, \n cuda: %s" % (this_node, this_os, this_machine, this_cuda))
print("\n")
columns=[
'node',
#'os',
#'machine',
'cuda',
'multi_gpu',
'model',
'batch_size',
'data_dir',
#'model_dir',
'train_epochs',
#'user',
#'time_stamp',
'localtime',
'steps',
'accuracy',
'duration'
]
history.experiments.tail(10)[columns]
# -
# ### Want to start with the most recent record from this platform?
hparams=history.suggest_from_history()
#hparams=history.copy_from_record(18)
hparams
# ### Use as new hyper-parameter record, with adaptations
# +
#DATA_SET = 'DIGITS'
#hparams.data_dir = '/var/ellie/data/mnist'
DATA_SET = 'FASHION'
hparams.data_dir = '/var/ellie/data/mnist_fashion'
hparams.train_epochs = 200
hparams.batch_size = 1024
hparams.multi_gpu = True
hparams.model = Model.id
hparams
# -
# ### Always have a quick peek at your input data!
samples = dataset.training_dataset(hparams.data_dir, DATA_SET).batch(10).make_one_shot_iterator().get_next()
samples = sess.run(samples)
f, arr = plt.subplots(2,5)
for row in (0, 1):
for col in range(5):
i = 5 * row + col
img = samples[0][i].reshape([28,28])
arr[row, col].imshow(img)
samples[1][:10]
# # Get to work!
# For the sake of this tutorial, we always start from scratch
# !rm -rf /tmp/mnist_model
# ### The model function constructs the computational graphs for training, eval and test
# Note that the actual construction takes place within the Estimator. Thus, none of the the constructing code should be explicitly called from the API client. The Estimator will complain that parts that have been constructed prior to those that itself constructs, don't belong to the same graph.
model_function = create_model_fn(
lambda params: Model(params),
tf.train.AdamOptimizer(),
tf.losses.sparse_softmax_cross_entropy,
hparams)
#
# Performance depends on the data format, and differs between CPU and GPU computations
data_format = ('channels_first' if tf.test.is_built_with_cuda() else 'channels_last')
# ### The Estimator is the center piece of Tensorflow's new API
mnist_classifier = tf.estimator.Estimator(
model_fn=model_function,
model_dir=hparams.model_dir,
params={
'data_format': data_format,
'multi_gpu': hparams.multi_gpu
})
# ##### ```input_fn``` functions are a factories for ```DataSet```s
# ### Split the training dataset into training and evaluation sets
# +
def train_input_fn():
ds_tr = dataset.training_dataset(hparams.data_dir, DATA_SET)
ds_tr_tr, _ = split_datasource(ds_tr, 60000, 0.95)
ds1 = ds_tr_tr.cache().shuffle(buffer_size=57000).\
repeat(hparams.train_epochs).\
batch(hparams.batch_size)
return ds1
def eval_input_fn():
ds_tr = dataset.training_dataset(hparams.data_dir, DATA_SET)
_, ds_tr_ev = split_datasource(ds_tr, 60000, 0.95)
ds2 = ds_tr_ev.batch(hparams.batch_size)
return ds2
# -
# ### Logging hooks
tensors_to_log = {'train_accuracy': 'train_accuracy'}
logging_hook = tf.train.LoggingTensorHook(tensors=tensors_to_log, every_n_iter=1000)
# ### Run the training and report the new hyper-parameters
# +
# Train
start_time=time.time()
mnist_classifier.train(input_fn=train_input_fn, hooks=[logging_hook])
duration=time.time() - start_time
# Evaluate
eval_results = mnist_classifier.evaluate(input_fn=eval_input_fn)
hparams.accuracy = eval_results['accuracy']
hparams.steps = eval_results['global_step']
hparams.duration = int(duration)
# Report!
history.report_experiment(hparams)
print('Evaluation results:\n\t%s' % eval_results)
hparams
# -
| experiments/mnist_sota/run_experiment_1.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import splat
import wisps
import matplotlib.pyplot as plt
from wisps.data_analysis import selection_criteria as sel_crt
from wisps.simulations import selection_function as slf
import numpy as np
import pandas as pd
import numba
import matplotlib as mpl
mpl.rcParams['font.size'] = 18
from itertools import combinations
# #%%capture output
import itertools
from tqdm import tqdm
import seaborn as sns
cmap=sns.light_palette((260, 75, 60), input="husl", as_cmap=True)
from tensorflow import keras
from scipy import stats
# -
rfdict=pd.read_pickle(wisps.OUTPUT_FILES+'/random_forest_classifier.pkl') #the classifier
neural_network= keras.models.load_model(wisps.OUTPUT_FILES+'/deep_model_september23.h5')
features=rfdict['feats']
#probs= neural_network
#labels=np.logical_or.reduce((probs[:, 2] > .95, probs[:,3] >.95 , probs[:,4] >0.8))
# +
#cands[features]
# -
# %matplotlib inline
#read in some data
sf=pd.read_pickle(wisps.LIBRARIES+'/selection_function.pkl.gz') #the simulated spectral data
#selection objects
rfdict=pd.read_pickle(wisps.OUTPUT_FILES+'/random_forest_classifier.pkl') #the classifier
indices_to_use= pd.read_pickle(wisps.OUTPUT_FILES+'/best_indices_to_use.pkl')
neural_network= keras.models.load_model(wisps.OUTPUT_FILES+'/deep_model_september23.h5')
#some formatting
sampled_data=pd.DataFrame.from_records(pd.DataFrame(sf).values.flatten())
sampled_data['sp_old']=np.vstack(sampled_data.sp_old.values)[:,0]
sampled_data['spt_new']=np.vstack(sampled_data.spt_new.values)[:,0]
#things that missed their classification
sampled_data['missed_label']=sampled_data['sp_old'].apply(wisps.make_spt_number) != sampled_data['spt_new'].apply(wisps.make_spt_number)
sampled_data['missed_label']=1-sampled_data.missed_label.apply(int).apply(float)
sampled_data['Names']=['spctr'+ str(idx) for idx in sampled_data.index]
sampled_data['spt']=sampled_data['sp_old'].apply(wisps.make_spt_number)
#selection criteria
slc_crts=sel_crt.crts_from_file()
indices_to_use
# +
#define a number of selectors
#each selection should a column of zeros and ones corresponding
#to where objects were selected
#each selector input is the simulated df
def f_test_fx(x, df1, df2):
return stats.f.cdf(x, df1, df2)
def select_by_indices(df, idx, spt_range):
print(spt_range)
bs=idx.shapes
bx=[x for x in bs if x.shape_name==spt_range][0]
_, bools= bx._select(np.array([df[idx.xkey].values, df[idx.ykey].values]))
return bools
def apply_scale(x):
##remove nans
##this is the same scaling used
## same scaling used for
#replace nans
y=x
if np.isnan(y) or np.isinf(y) or abs(y) > 1e10:
y=-99
return y
def select_by_random_forest(df):
#use the classification given by my rf classifier
rf=rfdict['classifier']
#min_max_scaler=rfdict['sclr']
features=rfdict['feats']
#apply logs to problematic features the same way I did on my classification
pred_df=df.copy()
for c in features:
if c not in ['spt', 'f_test', 'x']:
pred_df.assign(c=np.log10(pred_df[c].apply(apply_scale)))
else:
pred_df.assign(c=pred_df[c].apply(apply_scale))
pred_df[features]=pred_df[features].applymap(apply_scale)
#make predictions
probs=rf.predict_proba(pred_df[features].values)
labels=np.logical_or.reduce((
probs[:,2] > .8, \
probs[:,3] >.8 ,\
probs[:,4] >0.8))
#labels=np.logical_or.reduce([ probs[:, 0]<0.05, labels ])
#labels=rf.predict(pred_df[features].values)
return {'probs': probs, 'labels': labels}
def select_by_neuralnet(df):
#define features (start with indices alone)
#apply logs to problematic features the same way I did on my classification
features=rfdict['feats']
pred_df=df.copy()
for c in features:
if c not in ['spt']:
pred_df.assign(c=np.log10(pred_df[c].apply(apply_scale)))
else:
pred_df.assign(c=pred_df[c].apply(apply_scale))
pred_df[features]=pred_df[features].applymap(apply_scale)
#probs= neural_network.predict( pred_df[features].values)
#need to reshape
#probs=neural_network.predict( pred_df[features].values.reshape(-1, len(features), 1))
#my cuts
#labels=np.logical_or.reduce((probs[:, 2] > .7, probs[:,3] >.5 , probs[:,4] >0.5))
#labels=probs[:,0] <0.5
#labels=neural_network.predict_classes( pred_df[features].values.reshape(-1, len(features), 1))
#labels=neural_network.predict( pred_df[features].values.reshape(-1, len(features), 1))
probs= neural_network( pred_df[features].values, training=False)
labels=np.logical_or.reduce((
probs[:, 2] > .8, \
probs[:,3] >.8 ,\
probs[:,4] >0.8))
#labels=np.logical_or.reduce([probs[:, 1]>0.9, labels ])
#labels=neural_network.predict_classes( pred_df[features].values)
#labels=np.logical_or.reduce([ probs[:, 0]<0.05, labels ])
return {'probs': probs, 'labels': labels}
# -
df=wisps.Annotator.reformat_table(sampled_data)
#indices
for idxk, k in indices_to_use:
idx=slc_crts[idxk]
df['selected_by_{}'.format(k)]= select_by_indices(df, idx, k)
df['x']=df.spex_chi/df.line_chi
df['f_test']= f_test_fx(df.x, df.dof-1, df.dof-2)
plt.plot(df.spex_chi/df.line_chi, df.f_test, '.')
df['f_test_label']=np.logical_and.reduce([df.f_test<0.02, df.x <0.5, df.snr1>=3.])
df
select_by_random_forest(df)
# +
#machine learning
df['rf_label']=select_by_random_forest(df)['labels']
df['neural_net_label']=select_by_neuralnet(df)['labels']
df['rf_label']=np.logical_and(df['rf_label'], df.snr1>=3. ).apply(int)
df['neural_net_label']=np.logical_and(df['neural_net_label'], df.snr1>=3. ).apply(int)
# +
#indices and total
df.f_test_label=(df['f_test_label']).apply(int)
df['index_label']=np.logical_or.reduce([df['selected_by_{}'.format(x)].values for x in np.vstack(indices_to_use)[:,1]]).astype(int)
df['idx_ft_label']=np.logical_and(df['index_label'].apply(bool), df['f_test_label'].apply(bool) ).apply(int)
df['tot_label']=np.logical_or.reduce((df['idx_ft_label'].apply(bool), df['rf_label'].apply(bool), df['neural_net_label'].apply(bool)))
df.tot_label=np.logical_and(df.tot_label.values, (df.snr1>=3.).values).astype(int)
#put things on log-scale
df['logsnr']=df['snr1'].apply(np.log10)
# -
df_small=(df[['logsnr', 'spt','tot_label']]).reset_index(drop=True).dropna().values
# +
#x, y= np.meshgrid(df_small[:,0], df_small[:,1])
x= df_small[:,0]
y= df_small[:,1]
z= df_small[:,2]
xx, yy, zz = np.meshgrid(x, y,z, indexing='ij',sparse=True)
# -
xx.shape, yy.shape, zz.shape
df=df[np.logical_and(df.logsnr.between(0.36, 2), df.spt.between(16, 40))].reset_index(drop=True)
np.ptp(df.spt)
import matplotlib.colors as mcolors
import matplotlib
#mymap=mcolors.LinearSegmentedColormap.from_list('my_colormap', colors)
cmap='cubehelix'
# +
fig, ax=plt.subplots(ncols=2, nrows=2, figsize=(5.5*2, 5*2),
sharex=False, sharey=True)
wisps.plot_annotated_heatmap(ax[0][0], df, int(np.ptp(df.spt)), ['logsnr', 'spt', 'idx_ft_label'], cmap=cmap)
wisps.plot_annotated_heatmap(ax[0][1], df, int(np.ptp(df.spt)), ['logsnr', 'spt', 'rf_label'], cmap=cmap)
wisps.plot_annotated_heatmap(ax[1][0], df, int(np.ptp(df.spt)), ['logsnr', 'spt', 'neural_net_label'], cmap=cmap)
wisps.plot_annotated_heatmap(ax[1][1], df, int(np.ptp(df.spt)), ['logsnr', 'spt', 'tot_label'], cmap=cmap)
#df.plot.hexbin(x='logsnr', y='spt', C='idx_ft_label', reduce_C_function=np.nanmean, gridsize=50, cmap=cmap, ax=ax[0][0])
#df.plot.hexbin(x='logsnr', y='spt', C='rf_label', reduce_C_function=np.nanmean, gridsize=50, cmap=cmap, ax=ax[0][1])
#df.plot.hexbin(x='logsnr', y='spt', C='neural_net_label', reduce_C_function=np.nanmean, gridsize=50, cmap=cmap, ax=ax[1][0])
#df.plot.hexbin(x='logsnr', y='spt', C='tot_label', reduce_C_function=np.nanmean, gridsize=50, cmap=cmap, ax=ax[1][1])
#ax[0][0].scatter( sf.data.snr1.apply(np.log10), sf.data.spt, marker='+', color='#111111', alpha=.05)
ax[0][0].set_title('Indices, F-test ', fontsize=18)
ax[0][1].set_title('Random Forest', fontsize=18)
ax[1][0].set_title('Neural Network', fontsize=18)
ax[1][1].set_title('Total (or) ', fontsize=18)
for a in np.concatenate(ax):
a.set_xlabel('Log SNR-J', fontsize=18)
a.set_ylabel('SpT', fontsize=18)
a.axvline(np.log10(3), linestyle='--', color='#111111')
a.tick_params(which='major',direction='inout')
a.tick_params(which='minor',direction='out')
a.minorticks_on()
#a.set_yticks(np.arange(17, 42), minor=True)
a.set_yticks([17, 20, 25, 30, 35, 40], minor=False)
a.set_yticklabels(['M7', 'L0', 'L5', 'T0', 'T5', 'Y0'], minor=False)
#a.set_xlim([0., 2.3])
#a.set_ylim([17., 42.])
plt.tight_layout()
cax = fig.add_axes([1.01, 0.06, .03, 0.9])
norm= matplotlib.colors.Normalize(vmin=0.0,vmax=1.0)
mp=matplotlib.cm.ScalarMappable(norm=norm, cmap=cmap)
cbar=plt.colorbar(mp, cax=cax, orientation='vertical')
cbar.ax.set_ylabel(r'Selection Probability', fontsize=18)
plt.savefig(wisps.OUTPUT_FIGURES+'/selection_function_samples.pdf', bbox_inches='tight', dpi=200)
# -
#save partial of the data to use for my selection function calculations
df2=(df[['logsnr', 'tot_label', 'spt']])
df2.logsnr=df2.logsnr.apply(lambda x: np.round(x, 1))
# +
#df2.groupby(['spt', 'logsnr'])['tot_label'].mean().plot()
# -
df2.to_pickle(wisps.OUTPUT_FILES+'/selection_function_lookup_table.pkl')
stats.f.cdf(1, 30000, 100)
| notebooks/selection function2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# ## _Track Building (Reco.) from GNN Score_
#
# - after GNN Stage, one has evaluation (_**i.e.** edge score_) of GNN on test data.
# - the GNN evaluation data (_**i.e.** edge score_) is stored in _`run/gnn_evaluation/test`_.
# - use _`trkx_from_gnn.py`_ to reconstruct tracks saved to _`run/trkx_from_gnn`_ folder.
#
# Following the breakdown of _`trkx_from_gnn.py`_ script.
# -
import glob, os, sys, yaml
import numpy as np
import scipy as sp
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
import time
from sklearn.cluster import DBSCAN
from multiprocessing import Pool
from functools import partial
# select a device
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# functions from draw_utils for drawing
from LightningModules.Processing.utils.draw_utils import draw_proc_event, cylindrical_to_cartesian
# functions from tracks_from_gnn.py scripts
from trkx_from_gnn import tracks_from_gnn
# ### _(1) Processed Data_
inputdir="run/feature_store/test"
proc_files = sorted(glob.glob(os.path.join(inputdir, "*")))
proc_files[:5]
feature_data = torch.load(proc_files[1], map_location=device)
print("Length of Data: {}".format(len(feature_data)))
# event ID
event_id = int(os.path.basename(feature_data.event_file)[-10:])
print("Event ID: {}".format(event_id))
# number of tracks
track_ids = np.unique(feature_data.pid)
print("Track IDs: {}".format(track_ids))
# ### _(2) GNN Evaluation Data_
inputdir="run/gnn_evaluation/test"
gnn_eval_files = sorted(glob.glob(os.path.join(inputdir, "*")))
gnn_eval_files[:5]
gnn_eval_data = torch.load(gnn_eval_files[1], map_location=device)
print("Length of Data: {}".format(len(gnn_eval_data)))
# event ID
event_id = int(os.path.basename(gnn_eval_data.event_file)[-10:])
print("Event ID: {}".format(event_id))
# number of tracks
track_ids = np.unique(gnn_eval_data.pid)
print("Track IDs: {}".format(track_ids))
# evaluation score (only first 5 edge scores)
scores = gnn_eval_data.scores[:5].numpy()
print("Evaluation/Edge Score: {}".format(scores))
# ### _(3) Tracks from GNN_
#
# - We have everything in _`run/gnn_evaluation/test`_, _**i.e.**_ input feature data (from Processing Stage) and evaluation score (from GNN Stage).
# - The _score_ from GNN Stage is also called the _edge score_ or _evaluation score_, etc.
#
# Here is breakdown of _`tracks_from_gnn.py`_ script.
# Input dir from GNN Evaluation (GNN Test Step)
inputdir="run/gnn_evaluation/test"
# Output dir for Track Building/Reco
outputdir = "run/trkx_from_gnn"
os.makedirs(outputdir, exist_ok=True)
# +
# GNN Evaluation Data
# use os.listdir to fetch files
# all_events = os.listdir(inputdir) # get a list of files
# all_events = sorted([os.path.join(inputdir, event) for event in all_events]) # list-comprehension to join path with files & sort
# all_events[:10]
# +
# GNN Evaluation Data
# use glob.glob to fetch files (Note: glob.glob is wrapper around os.listdir)
gnn_eval_files = sorted(glob.glob(os.path.join(inputdir, "*")))
gnn_eval_files[:10]
# -
gnn_eval_data = torch.load(gnn_eval_files[1], map_location=device)
print("Length of Data: {}".format(len(gnn_eval_data)))
# input to GNN (processed data)
feature_data
# output from GNN (evaluated data)
gnn_eval_data
gnn_eval_data.edge_index
gnn_eval_data.edge_index.flip(0)
gnn_eval_data.scores
# process(): input params
max_evts = 100
n_tot_files = len(gnn_eval_files)
max_evts = max_evts if max_evts > 0 and max_evts <= n_tot_files else n_tot_files
# process(): prepare data for tracks_from_gnn()
score = gnn_eval_data.scores[:gnn_eval_data.edge_index.shape[1]]
senders = gnn_eval_data.edge_index[0]
receivers = gnn_eval_data.edge_index[1]
hit_id = gnn_eval_data.hid
score.shape
senders.shape
receivers.shape
hit_id.shape
# ### _Plotting Events_
# +
# plotting input_edges
plt.close('all')
# init subplots
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10, 10))
# detector layout
det = pd.read_csv("src/stt.csv")
skw = det.query('skewed==0')
nkw = det.query('skewed==1') # one may look for +ve/-ve polarity
plt.scatter(skw.x.values, skw.y.values, s=20, facecolors='none', edgecolors='lightgreen')
plt.scatter(nkw.x.values, nkw.y.values, s=20, facecolors='none', edgecolors='coral')
# feature data
x,y,_ = cylindrical_to_cartesian(r=gnn_eval_data.x[:, 0].detach().numpy(),
phi=gnn_eval_data.x[:, 1].detach().numpy(),
z=gnn_eval_data.x[:, 2].detach().numpy())
# particle track(s)
pids = np.unique(gnn_eval_data.pid)
for pid in pids:
idx = gnn_eval_data.pid == pid
ax.scatter(x[idx], y[idx], label='particle_id: %d' %pid)
# plotting params
ax.set_title('Event ID # %d' % event_id)
ax.set_xlabel('x [cm]', fontsize=10)
ax.set_ylabel('y [cm]', fontsize=10)
ax.set_xlim(-41, 41)
ax.set_ylim(-41, 41)
ax.grid(False)
ax.legend(fontsize=10, loc='best')
fig.tight_layout()
# fig.savefig("input_edges.png")
# -
# predicted/reco tracks using DBSCAN
reco_tracks = tracks_from_gnn(hit_id, score, senders, receivers, edge_score_cut=0.25, epsilon=0.25, min_samples=2)
# let fetch hit_ids of a track, e.g. track_id ==
reco_tracks.query("track_id==-1")
# +
# plotting input_edges
plt.close('all')
# init subplots
fig, ax = plt.subplots(nrows=1, ncols=1, figsize=(10, 10))
# detector layout
det = pd.read_csv("src/stt.csv")
skw = det.query('skewed==0')
nkw = det.query('skewed==1') # one may look for +ve/-ve polarity
plt.scatter(skw.x.values, skw.y.values, s=20, facecolors='none', edgecolors='lightgreen')
plt.scatter(nkw.x.values, nkw.y.values, s=20, facecolors='none', edgecolors='coral')
# feature data
x,y,_ = cylindrical_to_cartesian(r=gnn_eval_data.x[:, 0].detach().numpy(),
phi=gnn_eval_data.x[:, 1].detach().numpy(),
z=gnn_eval_data.x[:, 2].detach().numpy())
# reco track(s)
pids = np.unique(reco_tracks.track_id)
for pid in pids:
print("pid: ", pid)
idx = gnn_eval_data.pid == (pid+1)
if pid >= 0:
ax.scatter(x[idx], y[idx], s=30, label='particle_id: %d' %(pid+1))
# Missed hits
missed_hids= reco_tracks.query("track_id==-1")["hit_id"].values
hids = gnn_eval_data.hid.numpy()
idx = np.where(np.isin(hids, missed_hids))[0]
ax.scatter(x[idx], y[idx], facecolors='none', edgecolors='red', s=100, linewidth=1, label='missed')
# plotting params
ax.set_title('Event ID # %d' % event_id)
ax.set_xlabel('x [cm]', fontsize=10)
ax.set_ylabel('y [cm]', fontsize=10)
ax.set_xlim(-41, 41)
ax.set_ylim(-41, 41)
ax.grid(False)
ax.legend(fontsize=10, loc='best')
fig.tight_layout()
# fig.savefig("input_edges.png")
# -
np.unique(reco_tracks.track_id.values)
missed_hid = reco_tracks.query("track_id==-1")["hit_id"]
missed_hids = missed_hid.values
missed_hids
gnn_eval_data.hid
hids = gnn_eval_data.hid.numpy()
# let get indices of missed_hids in hids
idx = np.where(np.isin(hids, missed_hids))[0]
# ## Fixing the Srcipt using above EDA
#
# NOTE: The script `tracks_from_gnn.py` is taken from the `exatrkx-iml2020/exatrkx/scripts` repository. It needs to be fixed according to `exatrkx-hsf` repo.
#
# Above disection of this script provides info on how to make it compatible with the `exatrkx-hsf` pipeline.
#
# - **_Keep_**: _`tracks_from_gnn()`_
# - **_Modify_**: _`process()`_
# - **_Modify_**: _`__main__`_
# +
# tracks_from_gnn() declared above
# -
# Input/Output Data. Get Data from test Folder.
inputdir="run/gnn_evaluation/test"
outputdir = "run/trkx_from_gnn"
os.makedirs(outputdir, exist_ok=True)
# use os.listdir(path) to fetch files in arbitrary order
all_events = os.listdir(inputdir) # only list of files in arbitrary order
all_events = [os.path.join(inputdir, event) for event in all_events] # join path+files as a list
all_events = sorted(all_events) # sorted() over list/tuple iterator
all_events[:10]
# OR, use glob.glob to fetch files (Note: glob.glob is wrapper around os.listdir)
all_files = glob.glob(os.path.join(inputdir, "*")) # list of files with path in arbitrary order
all_files = sorted(all_files) # sorted() over list/tuple iterator
all_files[:10]
max_evts = 100
n_tot_files = len(all_files)
max_evts = max_evts if max_evts > 0 and max_evts <= n_tot_files else n_tot_files
# Let Peep into a Event
fname = all_files[0]
int(os.path.basename(fname))
evtid = int(os.path.basename(fname))
print("event_id: {}".format(evtid))
# Load Event
gnn_data = torch.load(fname)
print("Length of Data: {}".format(len(gnn_data)))
gnn_data.scores.shape[0]/2
# Get score, edge pair (sender, receiver) and hit_id from the Event
score = gnn_data.scores[:gnn_data.edge_index.shape[1]]
senders = gnn_data.edge_index[0]
receivers = gnn_data.edge_index[1]
hit_id = gnn_data.hid
def process(filename, outdir, score_name, **kwargs):
"""prepare a multiprocessing function for track building"""
# get the event_id from the filename
#evtid = int(os.path.basename(filename)) # [:-4] was to skip .npz extension, skipped in my case.
evtid = os.path.basename(filename)
# gnn_prcessed data by GNNBuilder Callback
gnn_data = torch.load(filename)
score = gnn_data.scores[:gnn_data.edge_index.shape[1]] # scores has twice the size of edge_index (flip(0) was used)
senders = gnn_data.edge_index[0]
receivers = gnn_data.edge_index[1]
hit_id = gnn_data.hid
# predicted tracks from the GNN stage
predicted_tracks = tracks_from_gnn(hit_id, score, senders, receivers, **kwargs)
# save reconstructed tracks into a file
# PyTorch convention is to save tensors using .pt file extension
# See https://pytorch.org/docs/stable/notes/serialization.html#preserve-storage-sharing
torch.save(predicted_tracks, os.path.join(outdir, "{}.pt".format(evtid)))
# after success move to gnn_trkx.py
process(fname, outputdir, "scores")
| eda/trkx_from_gnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis of Contact Tracing Applications
# +
import re
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from tracking import config, utils
# %load_ext autoreload
# %autoreload 2
# -
sns.set(font_scale=1.5)
# ## Loading data
df_apps = pd.read_csv(config.data / 'project-data - all-apps.csv')
df_apps
df_apps = (df_apps.fillna({
'open_source': 'no',
'quarantine_enforcement': 'no',
'government': 'no'
})
.fillna('unknown'))
df_apps['notes'] = df_apps['notes'].replace('unknown', '')
utils.display_all_cols(df_apps)
# ## Descriptive stats
df_apps['name'].nunique()
df_apps['country'].nunique()
df_apps['country'].value_counts()
df_apps['protocol'].value_counts()
df_apps[df_apps['protocol'] == 'unknown']['country'].value_counts()
# ## Plotting
cols_to_plot = [
'data_type',
'centralized',
'status',
'data_storage',
'protocol',
'data_persistence_days',
'government',
'opt_in',
'open_source',
'covid_positive_verification',
'quarantine_enforcement'
]
ncols = 1
width = 8
height = 4
fig, axs = plt.subplots(int(np.ceil(len(cols_to_plot) / ncols)), ncols, figsize=(ncols * width, len(cols_to_plot) / ncols * height))
for col, ax in zip(cols_to_plot, axs.flat):
subset = df_apps.query(f'{col} != "unknown"')
sns.countplot(x=col, data=subset, ax=ax)
plt.show()
# Looking at government vs. different variables
day_order = ['unknown', '14', '21', '30', 'unlimited']
cols = [
'centralized',
'open_source',
'data_persistence_days'
]
fig, axs = plt.subplots(1, 3, figsize=(20, 8))
for col, ax in zip(cols, axs):
if col == 'data_persistence_days':
sns.countplot(x=col, ax=ax, hue='government', order=day_order, data=df_apps)
else:
sns.countplot(x=col, ax=ax, hue='government', data=df_apps)
plt.savefig(config.figs / 'gov_bars.png')
plt.show()
df_apps[df_apps['quarantine_enforcement'] == 'yes']
df_apps['data_persistence_days'].value_counts()
| notebooks/1_contact_tracing_applications.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # name Table
#
# <a href="https://colab.research.google.com/github/source-foundry/opentype-notes/blob/master/notebooks/tables/name.ipynb">
# <img style="margin-left:0;margin-top:15px" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/>
# </a>
#
# ## Description
#
# The name table includes platform-specific localized records of font metadata. These records are organized by platform ID, platform endcoding ID, and language ID. There are 26 defined name record fields.
#
# ## Documentation
#
# - [Apple Specification](https://developer.apple.com/fonts/TrueType-Reference-Manual/RM06/Chap6name.html)
# - [Microsoft Specification](https://docs.microsoft.com/en-us/typography/opentype/spec/name)
#
# + [markdown] pycharm={"metadata": false}
# ## Source
#
# + [markdown] pycharm={"metadata": false}
# ### Settings
#
# Change the paths below to view the table in a different font.
#
# + pycharm={"is_executing": false, "metadata": false, "name": "#%%\n"}
FONT_URL = "https://github.com/source-foundry/opentype-notes/raw/master/assets/fonts/roboto/Roboto-Regular.ttf"
FONT_PATH = "Roboto-Regular.ttf"
# + [markdown] pycharm={"metadata": false}
# ### Setup
#
# + pycharm={"metadata": false, "name": "#%%\n"}
import os
try:
import fontTools
except ImportError:
# !pip install fontTools
if not os.path.exists(FONT_PATH):
# !curl -L -O {FONT_URL}
# + [markdown] pycharm={"metadata": false}
# ### View Table
#
# + pycharm={"metadata": false, "name": "#%%\n"}
# !ttx -t name -o - {FONT_PATH}
# + [markdown] pycharm={"metadata": false}
# ### Read/Write Access to Table
#
# - [fontTools `_n_a_m_e.py` module](https://github.com/fonttools/fonttools/blob/master/Lib/fontTools/ttLib/tables/_n_a_m_e.py)
#
# + pycharm={"metadata": false, "name": "#%%\n"}
import inspect
from fontTools.ttLib import TTFont
# instantiate table object
tt = TTFont(FONT_PATH)
table = tt["name"]
# print table methods
print("Printing methods of {}:".format(table))
methods = inspect.getmembers(table, predicate=inspect.ismethod)
methods_list = [method[0] for method in methods]
for x in sorted(methods_list):
print(x)
# + [markdown] pycharm={"metadata": false}
# ### Cleanup
#
# + pycharm={"metadata": false, "name": "#%%\n"}
# !rm {FONT_PATH}
| notebooks/tables/name.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>In-class transcript from Lecture 6, January 28, 2019</h2>
#
# # Imports and defs for lecture
# +
# These are the standard imports for CS 111.
# This list may change as the quarter goes on.
import os
import time
import math
import numpy as np
import numpy.linalg as npla
import scipy
from scipy import sparse
from scipy import linalg
import scipy.sparse.linalg as spla
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits.mplot3d import axes3d
# %matplotlib tk
# -
def Usolve(U, y, unit_diag = False):
"""Backward solve an upper triangular system Ux = y for x
Parameters:
U: the matrix, must be square, upper triangular, with nonzeros on the diagonal
y: the right-hand side vector
unit_diag = False: if true, assume the diagonal is all ones
Output:
x: the solution vector to U @ x == y
"""
# Check the input
m, n = U.shape
assert m == n, "matrix must be square"
assert np.all(np.triu(U) == U), "matrix U must be upper triangular"
if unit_diag:
assert np.all(np.diag(U) == 1), "matrix U must have ones on the diagonal"
yn, = y.shape
assert yn == n, "rhs vector must be same size as U"
# Make a copy of y that we will transform into the solution
x = y.astype(np.float64).copy()
# Back solve
for col in reversed(range(n)):
if not unit_diag:
x[col] /= U[col, col]
x[:col] -= x[col] * U[:col, col]
return x
# # Lecture starts here
A = np.round(20*np.random.rand(4,4))
A
Q, R = linalg.qr(A)
# +
print('Q:', Q.shape); print(Q)
print('\nR:', R.shape); print(R)
npla.norm(Q @ R - A )
# -
Q.T @ Q
b = np.random.rand(4)
b
x = Usolve(R, Q.T @ b)
x
npla.norm(b - A @ x) / npla.norm(b)
A = np.round(20*np.random.rand(10,4))
A
Q, R = linalg.qr(A)
npla.norm(Q.T @ Q - np.eye(10))
R
npla.norm(Q @ R - A )
| 01.28/Class_transcript_01_28_QR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
np.random.seed(1337) # for reproducibility
from keras.datasets import mnist
from keras.utils import np_utils
from keras.models import Sequential
from keras.layers import Dense, Activation, Convolution2D, MaxPooling2D, Flatten
from keras.optimizers import Adam
# +
# download the mnist to the path '~/.keras/datasets/' if it is the first time to be called
# training X shape (60000, 28x28), Y shape (60000, ). test X shape (10000, 28x28), Y shape (10000, )
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# data pre-processing
X_train = X_train.reshape(-1, 1,28, 28)/255.
X_test = X_test.reshape(-1, 1,28, 28)/255.
y_train = np_utils.to_categorical(y_train, num_classes=10)
y_test = np_utils.to_categorical(y_test, num_classes=10)
# +
# Another way to build your CNN
model = Sequential()
# Conv layer 1 output shape (32, 28, 28)
model.add(Convolution2D(
batch_input_shape=(None, 1, 28, 28),
filters=32,
kernel_size=5,
strides=1,
padding='same', # Padding method
data_format='channels_first',
))
model.add(Activation('relu'))
# Pooling layer 1 (max pooling) output shape (32, 14, 14)
model.add(MaxPooling2D(
pool_size=2,
strides=2,
padding='same', # Padding method
data_format='channels_first',
))
# Conv layer 2 output shape (64, 14, 14)
model.add(Convolution2D(64, 5, strides=1, padding='same', data_format='channels_first'))
model.add(Activation('relu'))
# Pooling layer 2 (max pooling) output shape (64, 7, 7)
model.add(MaxPooling2D(2, 2, 'same', data_format='channels_first'))
# Fully connected layer 1 input shape (64 * 7 * 7) = (3136), output shape (1024)
model.add(Flatten())
model.add(Dense(1024))
model.add(Activation('relu'))
# Fully connected layer 2 to shape (10) for 10 classes
model.add(Dense(10))
model.add(Activation('softmax'))
# +
# Another way to define your optimizer
adam = Adam(lr=1e-4)
# We add metrics to get more results you want to see
model.compile(optimizer=adam,
loss='categorical_crossentropy',
metrics=['accuracy'])
# +
print('Training ------------')
# Another way to train the model
model.fit(X_train, y_train, epochs=1, batch_size=64,)
print('\nTesting ------------')
# Evaluate the model with the metrics we defined earlier
loss, accuracy = model.evaluate(X_test, y_test)
print('\ntest loss: ', loss)
print('\ntest accuracy: ', accuracy)
# -
| notebook/keras_cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Regression
from IPython.display import Image
# %matplotlib inline
# +
import pandas as pd
df = pd.read_csv('https://raw.githubusercontent.com/rasbt/'
'python-machine-learning-book-3rd-edition/'
'master/ch10/housing.data.txt',
header=None,
sep='\s+')
df.columns = ['CRIM', 'ZN', 'INDUS', 'CHAS',
'NOX', 'RM', 'AGE', 'DIS', 'RAD',
'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
df.head()
# -
import matplotlib.pyplot as plt
from mlxtend.plotting import scatterplotmatrix
# +
cols = ['LSTAT', 'INDUS', 'NOX', 'RM', 'MEDV']
scatterplotmatrix(df[cols].values, figsize=(10, 8),
names=cols, alpha=0.5)
plt.tight_layout()
plt.show()
# -
import numpy as np
from mlxtend.plotting import heatmap
cm = np.corrcoef(df[cols].values.T)
hm = heatmap(cm, row_names=cols, column_names=cols)
plt.show()
# ## Simple regression gradient descent
class LinearRegressionGD(object):
def __init__(self, eta=0.001, n_iter=20):
self.eta = eta
self.n_iter = n_iter
def fit(self, X, y):
self.w_ = np.zeros(1 + X.shape[1])
self.cost_ = []
for i in range(self.n_iter):
output = self.net_input(X)
errors = (y - output)
self.w_[1:] += self.eta * X.T.dot(errors)
self.w_[0] += self.eta * errors.sum()
cost = (errors**2).sum() / 2.0
self.cost_.append(cost)
return self
def net_input(self, X):
return np.dot(X, self.w_[1:]) + self.w_[0]
def predict(self, X):
return self.net_input(X)
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X = df[['RM']].values
y = df['MEDV'].values
scx=StandardScaler()
scy=StandardScaler()
Xstd=scx.fit_transform(X)
ystd=scy.fit_transform(y[: ,np.newaxis]).flatten()
X_train, X_test, y_train, y_test = train_test_split(Xstd, ystd, test_size=0.33, random_state=42)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
print(y_train[:5])
print(y_test[:5])
plt.plot(X_train,y_train,'ro')
lr = LinearRegressionGD()
lr.fit(X_train, y_train)
print("intercept:",lr.w_[0])
print("slope:",lr.w_[1])
#prediction
from sklearn.metrics import accuracy_score , mean_squared_error
ytrue=y_train
ypred = lr.predict(X_train)
print('Mean Squared Error:',mean_squared_error(ytrue,ypred))
# # multi linear regression
from sklearn.model_selection import train_test_split
X = df[['RM','LSTAT','INDUS','NOX']].values
y = df['MEDV'].values
X_train, X_test, y_train, y_test = train_test_split(Xstd, ystd, test_size=0.33, random_state=42)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
lr = LinearRegressionGD()
lr.fit(X_train, y_train)
print("intercept:",lr.w_[0])
print("slope:",lr.w_[1])
#prediction
from sklearn.metrics import accuracy_score , mean_squared_error
ytrue=y_train
ypred = lr.predict(X_train)
print('Mean Squared Error:',mean_squared_error(ytrue,ypred))
# # polynomial regression
# +
X = np.array([258.0, 270.0, 294.0,
320.0, 342.0, 368.0,
396.0, 446.0, 480.0, 586.0])\
[:, np.newaxis]
y = np.array([236.4, 234.4, 252.8,
298.6, 314.2, 342.2,
360.8, 368.0, 391.2,
390.8])
# -
from sklearn.preprocessing import PolynomialFeatures
lr = LinearRegressionGD()
pr = LinearRegressionGD()
quadratic = PolynomialFeatures(degree=2)
X_quad = quadratic.fit_transform(X)
# +
# fit linear features
lr.fit(X, y)
X_fit = np.arange(250, 600, 10)[:, np.newaxis]
y_lin_fit = lr.predict(X_fit)
# fit quadratic features
pr.fit(X_quad, y)
y_quad_fit = pr.predict(quadratic.fit_transform(X_fit))
# plot results
plt.scatter(X, y, label='Training points')
plt.plot(X_fit, y_lin_fit, label='Linear fit', linestyle='--')
plt.plot(X_fit, y_quad_fit, label='Quadratic fit')
plt.xlabel('Explanatory variable')
plt.ylabel('Predicted or known target values')
plt.legend(loc='upper left')
plt.tight_layout()
plt.show()
# -
y_lin_pred = lr.predict(X)
y_quad_pred = pr.predict(X_quad)
print(y_quad_pred)
# ## poly reg on real dataset
# +
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
X = df[['LSTAT', 'INDUS', 'NOX', 'RM']].values
y = df['MEDV'].values
scx=StandardScaler()
scy=StandardScaler()
Xstd=scx.fit_transform(X)
ystd=scy.fit_transform(y[: ,np.newaxis]).flatten()
# -
X_train, X_test, y_train, y_test = train_test_split(Xstd, ystd, test_size=0.33, random_state=42)
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
quadratic = PolynomialFeatures(degree=2)
X_quad_train = quadratic.fit_transform(X_train)
X_quad_train
preg = LinearRegressionGD()
pr.fit(X_quad_train, y_train)
y_pred_quad_train = pr.predict(X_quad_train)
y_pred_quad_train[:10]
plt.figure()
#plt.plot(X_train,y_train ,'bo' , label="original data no quad")
plt.plot(X_quad_train,y_train,'r+' ) #quad features vs true label
plt.plot(X_quad_train,pr.predict(X_quad_train),'go') #quad features vs predictions
pr.w_
mean_squared_error(y_train,pr.predict(X_quad_train)) #for training
# ## testing
plt.plot(X_quad_train,y_train,'bo' )
plt.legend(['Origial data','Predicted data'])
y_pred_train_quad = pr.predict(X_quad_train)
plt.plot(X_quad_train, y_pred_train_quad ,'r+')
xtest_quad = quadratic.fit_transform(X_test) #pick test x and make quad out ofit
ytest_pred = pr.predict(xtest_quad) #pred ion test quad x
plt.plot(X_test,y_test,'bo')
plt.plot(xtest_quad,ytest_pred,'r+')
mean_squared_error(y_test,ytest_pred)
# Transforming the dataset:
# +
X = df[['LSTAT']].values
y = df['MEDV'].values
# transform features
X_log = np.log(X)
y_sqrt = np.sqrt(y)
# fit features
X_fit = np.arange(X_log.min()-1, X_log.max()+1, 1)[:, np.newaxis]
regr = regr.fit(X_log, y_sqrt)
y_lin_fit = regr.predict(X_fit)
linear_r2 = r2_score(y_sqrt, regr.predict(X_log))
# plot results
plt.scatter(X_log, y_sqrt, label='Training points', color='lightgray')
plt.plot(X_fit, y_lin_fit,
label='Linear (d=1), $R^2=%.2f$' % linear_r2,
color='blue',
lw=2)
plt.xlabel('log(% lower status of the population [LSTAT])')
plt.ylabel('$\sqrt{Price \; in \; \$1000s \; [MEDV]}$')
plt.legend(loc='lower left')
plt.tight_layout()
plt.show()
# -
# <br>
# <br>
| 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # WeatherPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
import country_converter as coco
# Import API key
from api_keys import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
# api.openweathermap.org/data/2.5/box/city?bbox={bbox}&appid={API key}
# bbox required Bounding box [lon-left,lat-bottom,lon-right,lat-top,zoom]
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
# minimum 500 cities
len(cities)
#Cities is a list
# cities
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
# loop through cities, create list of responseL
# Lists for the dataframe
locations = []
clouds = []
humidity = []
lat = []
lon = []
max_temp = []
wind = []
country = []
dates = []
allcitydata = []
# Initial counter
counter = 0
# Did not used this - for some crazy reason - when using this format, the exception statement will not print
# url = "http://api.openweathermap.org/data/2.5/weather?"
# query_url = url + "appid=" + weather_api_key + "&q=" + city + "&units=imperial"
query_url = f"http://api.openweathermap.org/data/2.5/weather?appid="+ weather_api_key +"&units=imperial&q="
for city in cities:
try:
response = requests.get(query_url + city.replace(" ","&")).json()
clouds.append(response['clouds']['all'])
humidity.append(response['main']['humidity'])
lat.append(response['coord']['lat'])
lon.append(response['coord']['lon'])
max_temp.append(response['main']['temp_max'])
wind.append(response['wind']['speed'])
country.append(response['sys']['country'])
locations.append(response['name'])
dates.append(response['dt'])
allcitydata.append(response)
counter = counter + 1
print(f"Counter : {counter}, City : {city}")
except Exception:
print("weather data not available")
# +
# allcitydata
# clouds
# dates
# humidity
# lat
# lon
# max_temp
# wind
# country
# locations
# -
#Convert country abbreviations to full name
full_cnames = coco.convert(names=country, to='name_short')
# full_cnames
# +
# dates.dtype
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
# +
# Weather data (wd) dataframe
wd_df = pd.DataFrame({"City" : locations, "Country" : full_cnames,
"Latitude" : lat, "Longitude" : lon,
"Max_temp_F" : max_temp, "Humidity_Percent" : humidity,
"Cloudy_Percent" : clouds, "Wind_Speed_mph" : wind,
"Date" : dates
})
wd_df
# -
wd_df.dtypes
# wd_df['Date'] = pd.to_datetime(wd_df['Date'],unit='s')
# wd_df['Date'] = pd.to_datetime(wd_df['Date'],format = "%d/%m/%Y")
wd_df['Date'] = pd.to_datetime(wd_df['Date'])
wd_df['Date'].dt.minute
wd_df
wd_df.dtypes
wd_df.to_csv('../output_data/cities.csv', index=False)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
# +
# Humidity check
humg100 = (wd_df["Humidity_Percent"] > 100)
humg100
humg100 = humg100.to_frame('Hum>100%')
wdHumg100_df = wd_df.merge(humg100, how = "outer", left_index=True, right_index=True)
wdHumg100_df["Hum>100%"] = wdHumg100_df["Hum>100%"]*1
wdHumg100_df.head()
print(wdHumg100_df.shape)
# -
# Get the cities that have humidity over 100%.
wdHumg100_df.loc[wdHumg100_df["Hum>100%"] == 1]
# +
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
clean_city_data = wdHumg100_df.drop(wdHumg100_df[wdHumg100_df['Hum>100%'] == 1].index, inplace = False)
print(clean_city_data.shape)
# -
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
clean_city_data.plot.scatter(x = "Latitude", y = "Max_temp_F", c="DarkBlue")
# day = clean_city_data["Date"].astype(str)
# plt.title ("City Latitude vs. Max Temperature" + (day))
plt.savefig("../Images/scatterLvT.png")
# plt.show()
plt.xlabel("Latitude")
plt.ylabel("Max Temperature (F)")
plt.title (f"City Latitude vs Max Temperature ({day})")
# -
# ## Latitude vs. Humidity Plot
clean_city_data.plot.scatter(x = "Latitude", y = "Humidity_Percent", c="DarkBlue")
plt.xlabel("Latitude")
plt.ylabel("Humidity (%)")
plt.title (f"City Latitude vs Humidity ({day})")
plt.savefig("../Images/scatterLvH.png")
# ## Latitude vs. Cloudiness Plot
clean_city_data.plot.scatter(x = "Latitude", y = "Cloudy_Percent", c="DarkBlue")
plt.xlabel("Latitude")
plt.ylabel("Cloudiness (%)")
plt.title (f"City Latitude vs Cloudiness ({day})")
plt.savefig("../Images/scatterLvC.png")
# ## Latitude vs. Wind Speed Plot
clean_city_data.plot.scatter(x = "Latitude", y = "Wind_Speed_mph", c="DarkBlue")
plt.xlabel("Latitude")
plt.ylabel("Wind Speed (mph)")
plt.title (f"City Latitude vs Wind Speeed ({day})")
plt.savefig("../Images/scatterLvWS.png")
# ## Linear Regression
nhclean_city_data = clean_city_data.loc[clean_city_data['Latitude'] >= 0]
shclean_city_data = clean_city_data.loc[clean_city_data['Latitude'] < 0]
# shclean_city_data
print(shclean_city_data.shape)
print(nhclean_city_data.shape)
print(clean_city_data.shape)
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
x_values = nhclean_city_data['Latitude']
y_values = nhclean_city_data['Max_temp_F']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("North Hemisphere Latitude")
plt.ylabel("Max Temperature (F)")
plt.title (f"City Latitude vs Max Temperature ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scatternhLvTlr.png")
plt.show()
# -
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
x_values = shclean_city_data['Latitude']
y_values = shclean_city_data['Max_temp_F']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("South Hemisphere Latitude")
plt.ylabel("Max Temperature (F)")
plt.title (f"City Latitude vs Max Temperature ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scattershLvTlr.png")
plt.show()
# -
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
x_values = nhclean_city_data['Latitude']
y_values = nhclean_city_data['Humidity_Percent']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("Northern Hemisphere Latitude")
plt.ylabel("Humidity (%)")
plt.title (f"City Latitude vs Humidity ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scatternhLvHlr.png")
plt.show()
# -
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
x_values = shclean_city_data['Latitude']
y_values = shclean_city_data['Humidity_Percent']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("Southern Hemisphere Latitude")
plt.ylabel("Humidity (%)")
plt.title (f"City Latitude vs Humidity ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scattershLvHlr.png")
plt.show()
# -
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
x_values = nhclean_city_data['Latitude']
y_values = nhclean_city_data['Cloudy_Percent']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("Northern Hemisphere Latitude")
plt.ylabel("Cloudiness (%)")
plt.title (f"City Latitude vs Cloudiness ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scatternhLvClr.png")
plt.show()
# -
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
x_values = shclean_city_data['Latitude']
y_values = shclean_city_data['Cloudy_Percent']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("Southern Hemisphere Latitude")
plt.ylabel("Cloudiness (%)")
plt.title (f"City Latitude vs Cloudiness ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scattershLvClr.png")
plt.show()
# -
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
x_values = nhclean_city_data['Latitude']
y_values = nhclean_city_data['Wind_Speed_mph']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("Northern Hemisphere Latitude")
plt.ylabel("Wind Speed (mph)")
plt.title (f"City Latitude vs Wind Speed ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scatternhLvWSlr.png")
plt.show()
# -
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
x_values = shclean_city_data['Latitude']
y_values = shclean_city_data['Wind_Speed_mph']
#need to grab a single day
# day = nhclean_city_data["Date"]
day = 1
(slope, intercept, rvalue, pvalue, stderr) = linregress(x_values, y_values)
regress_values = x_values * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
plt.scatter(x_values,y_values)
plt.plot(x_values,regress_values,"r-")
plt.annotate(line_eq,(2, -10),fontsize=15,color="red")
plt.xlabel("Southern Hemisphere Latitude")
plt.ylabel("Wind Speed (mph)")
plt.title (f"City Latitude vs Wind Speed ({day})")
# plt.annotate(f"Correlation coefficient is {round(correlation[0],2)}",(20,36),fontsize=10,color="red")
# plt.xlim(14.5,25.5)
plt.grid()
plt.savefig("../Images/scattershLvWSlr.png")
plt.show()
# -
| WeatherPy/working_code/hw6_wea_Thu5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Parameters
CLUSTER_ALGO = "KMeans"
NC = 21
CLUSTER_STD = 1.0
N_P_CLUSTERS = [3,30, 100, 300, 3000]
OUTER_FOLD = 5
INNER_FOLD =10
# + [markdown] heading_collapsed=true
# ## IMPORTS
# + hidden=true
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
from sklearn.datasets import load_breast_cancer
from sklearn.datasets import load_iris
from sklearn.datasets import make_blobs
from sklearn.datasets import make_moons
# + hidden=true
# %load_ext autoreload
# %autoreload 2
# packages = !conda list
packages
# + hidden=true
# !pwd
# + [markdown] heading_collapsed=true
# ## Output registry
# + hidden=true
from __future__ import print_function
import sys, os
# old__file__ = !pwd
# __file__ = !cd ../photon ;pwd
# #__file__ = !pwd
__file__ = __file__[0]
__file__
sys.path.append(__file__)
print(sys.path)
os.chdir(old__file__[0])
# !pwd
old__file__[0]
# + hidden=true
import seaborn as sns; sns.set() # for plot styling
import numpy as np
import pandas as pd
from math import ceil
from sklearn.model_selection import KFold
from sklearn.manifold import TSNE
import itertools
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
#set font size of labels on matplotlib plots
plt.rc('font', size=16)
#set style of plots
sns.set_style('white')
#define a custom palette
customPalette = ['#40111D', '#DCD5E4', '#E7CC74'
,'#39C8C6', '#AC5583', '#D3500C'
,'#FFB139', '#98ADA7', '#AD989E'
,'#708090','#6C8570','#3E534D'
,'#0B8FD3','#0B47D3','#96D30B'
,'#630C3A','#F1D0AF','#64788B'
,'#8B7764','#7A3C5D','#77648B'
,'#eaff39','#39ff4e','#4e39ff'
,'#ff4e39','#87ff39','#ff3987', ]
sns.set_palette(customPalette)
sns.palplot(customPalette)
from clusim.clustering import Clustering, remap2match
import clusim.sim as sim
from photonai.base import Hyperpipe, PipelineElement, Preprocessing, OutputSettings
from photonai.optimization import FloatRange, Categorical, IntegerRange
from photonai.base.photon_elements import PhotonRegistry
from photonai.visual.graphics import plot_cm
#from photonai.base.registry.registry import PhotonRegistry
# + hidden=true
def results_to_df(results):
ll = []
for obj in results:
ll.append([obj.operation,
obj.value,
obj.metric_name])
_results=pd.DataFrame(ll).pivot(index=2, columns=0, values=1)
_results.columns=['Mean','STD']
return(_results)
# + hidden=true
def cluster_plot(my_pipe, data_X, customPalette):
y_pred= my_pipe.predict(data_X)
data = pd.DataFrame(data_X[:, 0],columns=['x'])
data['y'] = data_X[:, 1]
data['labels'] = y_pred
facet = sns.lmplot(data=data, x='x', y='y', hue='labels',
aspect= 1.0, height=7,
fit_reg=False, legend=True, legend_out=True)
for i, label in enumerate( np.sort(data['labels'].unique())):
plt.annotate(label,
data.loc[data['labels']==label,['x','y']].mean(),
horizontalalignment='center',
verticalalignment='center',
size=10, weight='bold',
color='white',
backgroundcolor=customPalette[i])
plt.show()
return y_pred
# + hidden=true
__file__ = "exp1.log"
base_folder = os.path.dirname(os.path.abspath(''))
custom_elements_folder = os.path.join(base_folder, 'custom_elements')
custom_elements_folder
# + hidden=true
registry = PhotonRegistry(custom_elements_folder=custom_elements_folder)
registry.activate()
registry.PHOTON_REGISTRIES,PhotonRegistry.PHOTON_REGISTRIES
# + hidden=true
registry.activate()
registry.list_available_elements()
# take off last name
# -
# ## KMedoids yield_parameters_ellipse
registry.info(CLUSTER_ALGO)
def yield_parameters_ellipse(n_p_clusters):
n_cluster = NC
cluster_std = CLUSTER_STD
for n_p_cluster in n_p_clusters:
n_cluster_std = [cluster_std for k in range(n_cluster)]
n_samples = [n_p_cluster for k in range(n_cluster)]
data_X, data_y = make_blobs(n_samples=n_samples,
cluster_std=n_cluster_std, random_state=0)
transformation = [[0.6, -0.6], [-0.4, 0.8]]
X_ellipse = np.dot(data_X, transformation)
yield [X_ellipse, data_y]
def hyper_cluster(cluster_name):
n_p_clusters = N_P_CLUSTERS
for data_X, data_y in yield_parameters_ellipse(n_p_clusters):
print('n_points:', len(data_y))
"""
Example script for KMedoids hopt
"""
X = data_X.copy(); y = data_y.copy()
# DESIGN YOUR PIPELINE
settings = OutputSettings(project_folder='./tmp/')
my_pipe = Hyperpipe('batching',
optimizer='sk_opt',
# optimizer_params={'n_configurations': 25},
metrics=['ARI', 'MI', 'HCV', 'FM'],
best_config_metric='ARI',
outer_cv=KFold(n_splits= OUTER_FOLD),
inner_cv=KFold(n_splits=INNER_FOLD),
verbosity=0,
output_settings=settings)
my_pipe += PipelineElement(cluster_name, hyperparameters={
'n_clusters': IntegerRange(1, ceil(NC*1.2)),
},random_state=777)
# NOW TRAIN YOUR PIPELINE
my_pipe.fit(X, y)
debug = True
#------------------------------plot
y_pred=cluster_plot(my_pipe, X, customPalette)
#--------------------------------- best
print(pd.DataFrame(my_pipe.best_config.items()
,columns=['n_clusters', 'k']))
#------------------------------
print('train','\n'
,results_to_df(my_pipe.results.metrics_train))
print('test','\n'
,results_to_df(my_pipe.results.metrics_test))
#------------------------------
# turn the ground-truth labels into a clusim Clustering
true_clustering = Clustering().from_membership_list(y)
kmeans_clustering = Clustering().from_membership_list(y_pred) # lets see how similar the predicted k-means clustering is to the true clustering
#------------------------------
# using all available similar measures!
row_format2 ="{:>25}" * (2)
for simfunc in sim.available_similarity_measures:
print(row_format2.format(simfunc, eval('sim.' + simfunc+'(true_clustering, kmeans_clustering)')))
#------------------------------# The element-centric similarity is particularly useful for understanding
# how a clustering method performed
# Let's start with the single similarity value:
elsim = sim.element_sim(true_clustering, kmeans_clustering)
print("Element-centric similarity: {}".format(elsim))
hyper_cluster(CLUSTER_ALGO)
| Cluster/kmeans-kmedoids/KMeans-ellipse-21-1.0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Detection
#
# Classification can tell you what object you are seeing, but you may also need to answer:
# * Where is an object?
# * How many are there?
#
# We generally call this object detection
#
# # Outline
#
#
#
# * Our Goal for this lecture is to identify signs of Malaria in human blood smears
#
#
#
# ## Challenge Data Set
#
# *P. vivax* (malaria) infected human blood smears
#
# Accession number BBBC041 · Version 1
#
#
#
# From the Broad Institute https://data.broadinstitute.org/bbbc/BBBC041/
#
#
# <img src=https://data.broadinstitute.org/bbbc/BBBC041/BBBC041_example1.png>
#
# Malaria is a disease caused by *Plasmodium* parasites that remains a major threat in global health, affecting 200 million people and causing 400,000 deaths a year. The main species of malaria that affect humans are *Plasmodium falciparum* and *Plasmodium vivax*.
#
# For malaria as well as other microbial infections, manual inspection of thick and thin blood smears by trained microscopists remains the gold standard for parasite detection and stage determination because of its low reagent and instrument cost and high flexibility. Despite manual inspection being extremely low throughput and susceptible to human bias, automatic counting software remains largely unused because of the wide range of variations in brightfield microscopy images. However, a robust automatic counting and cell classification solution would provide enormous benefits due to faster and more accurate quantitative results without human variability; researchers and medical professionals could better characterize stage-specific drug targets and better quantify patient reactions to drugs.
#
# Previous attempts to automate the process of identifying and quantifying malaria have not gained major traction partly due to difficulty of replication, comparison, and extension. Authors also rarely make their image sets available, which precludes replication of results and assessment of potential improvements. The lack of a standard set of images nor standard set of metrics used to report results has impeded the field.
#
#
# # Coordinates
#
# We are going to use the index coordinate system
#
# <img src=../assets/index_coords.png>
#
# * **Packages often have different systems so be careful!**
#
#
# ## Bounding Boxes
#
# Data is often labeled by bounding boxes, which also can be describe in a number of different ways. We'll use
# Upper left corner (note this is the smallest y-coordinate), the length and the width.
#
# <img src=../assets/BBox.png>
#
# Each bounding box also has a category attached to it. For example moon or cloud.
# +
import os
import numpy as np
import matplotlib.pyplot as plt
from bbtoydata import BBToyData
import matplotlib.patches as patches
import tensorflow as tf
# +
#Lets Practice
# +
x_pixel=0
y_pixel=1
img=np.zeros((3,3))
img[y_pixel,x_pixel]=1.0## Notice y,x here
fig,ax = plt.subplots(1)
ax.imshow(img,cmap='gray')
plt.xlabel(r" X -0.5 $\rightarrow$ 3.5 ")
plt.ylabel(r" 2.5 $\leftarrow$ -0.5 Y ")
print(img)
##Draw a bounding box
rec_corner=((x_pixel-0.5,y_pixel-0.5)) #half a pixel Width
width=1
height=1
# We'll uses pat
rec=patches.Rectangle(rec_corner, width,height,edgecolor='r',facecolor='none')
ax.add_patch(rec)
plt.show()
# -
# We'll add a lot of bounding boxes so let's write a function to do it
# ## Quick Test
# Draw a 2x2 box in the lower left hand corner of the image
# +
# We'll be drawing a lot of boxes so lets make it a function
# +
def add_bbox(ax,bbox,color='r'):
corner_x,corner_y,width,height=bbox
rec=patches.Rectangle((corner_x-0.5,corner_y-0.5), width,height,edgecolor=color,facecolor='none')
ax.add_patch(rec)
# -
# ## Identifying where something is with a neural network
#
# Again, we'll use a toy example that's easy to under and visualize.
#
# Start by finding just one object.
#
# * 3-Object Classes
# * Square
# * Circle
# * Triangle
# * 1 - Background Class
#
# * 4 - bbox x, y, width, height
#
# We need a classification network to identify the shape, and a new network go predict the bounding box
#
# * We'll use two prediction networks that share features!
#
# <img src=../assets/network_diagrams/1_object_identifier.png>
#
#
# # Toy Data
#
# We'll use a set of images that contain either a square, circle, triangle, or nothing.
#
# * Goal will be to identify the object, and draw a bounding box around it
#
#
#
bb=BBToyData()
# +
def plot_example(image,labels):
colors=['r','g','b','y','#000000']
fig,ax = plt.subplots(1,figsize=(10,10))
ax.imshow(image,cmap='gray')
for cat,bbox in labels:
add_bbox(ax,bbox,color=colors[cat])
plt.show()
for i in range(10):
index=np.random.randint(bb.X_train.shape[0])
print(bb.Y_train[index])
plot_example(np.squeeze(bb.X_train[index]),(bb.Y_train[index]))
# -
# ## Build our Convolutional Feature Finders
# +
cnn_input=tf.keras.layers.Input( shape=bb.X_train.shape[1:] ) # Shape here does not including the batch size
cnn_layer1=tf.keras.layers.Convolution2D(64, (4,4),strides=2,padding='same')(cnn_input)
cnn_activation=tf.keras.layers.LeakyReLU()(cnn_layer1)
cnn_layer2=tf.keras.layers.Convolution2D(64, (4,4),strides=2,padding='same')(cnn_activation)
cnn_activation=tf.keras.layers.LeakyReLU()(cnn_layer2)
#cnn_layer3=tf.keras.layers.Convolution2D(64, (4,4),strides=2,padding='same')(cnn_activation)
#cnn_activation=tf.keras.layers.LeakyReLU()(cnn_layer3)
flat=tf.keras.layers.Flatten()(cnn_activation)
flat=tf.keras.layers.Dropout(0.5)(flat)
# +
#Make Our Prediction layer
cat_output_layer=tf.keras.layers.Dense(4)(flat)
cat_output=tf.keras.layers.Activation('softmax')(cat_output_layer)
#Make our bounding box regressor
bbox_output=tf.keras.layers.Dense(50)(flat)
bbox_output=tf.keras.layers.LeakyReLU()(bbox_output)
bbox_output=tf.keras.layers.Dense(50)(bbox_output)
bbox_output=tf.keras.layers.LeakyReLU()(bbox_output)
bbox_output=tf.keras.layers.Dense(4)(bbox_output)
# +
model=tf.keras.models.Model(cnn_input,[cat_output,bbox_output])
##Two!! loss fuctions one for each output, the total loss is the sum of each of these
model.compile(loss=['categorical_crossentropy','mae'],
optimizer='adam',
metrics=[])
model.summary()
# -
# # Make our Data
# We need to transform labels from a list to a one hot encoding
#
# +
def get_single_object_labels(labels):
Y_bbox=[]
Y_cat=[]
for l in labels:
if len(l)==0:
Y_cat.append(3) #Background
Y_bbox.append(np.array([[0,0,1,1]])) #Divide by our image size so everything is between 0-1
else:
cat,bbox=l[0]
Y_cat.append(cat)
Y_bbox.append(np.expand_dims(bbox,0)/50,) #Divide by our image size so everything is between 0-1
Y_cat=tf.keras.utils.to_categorical(Y_cat, num_classes=4)
Y_bbox=np.concatenate(Y_bbox)
return [Y_cat,Y_bbox]
Y_train=get_single_object_labels(bb.Y_train)
Y_develop=get_single_object_labels(bb.Y_develop)
Y_test=get_single_object_labels(bb.Y_test)
# +
history=model.fit(bb.X_train, Y_train,
batch_size=32, epochs=25, verbose=1,
validation_data=(bb.X_develop,Y_develop)
)
# -
cat_pred,bbox_pred=model.predict(bb.X_develop)
for i in range(20):
cat=np.argmax(cat_pred[i])
label=[ [cat,bbox_pred[i]*50]]
plot_example(np.squeeze(bb.X_develop[i]),label)
print(cat,bbox_pred[i]*100)
print(bb.Y_develop[i])
# ## Identifying more than one thing at a time
#
# **Run the next several cells to get your model training before we get started**
#
#
# That worked pretty well, but we have another problem - what if there is more than one object in the image?
# * There are several ways to handle this:
# * Scan the image (like we did with the cancer example)
# * This works but can be expensive
# * Use multiple detectors
# * Strategry used by algorithms like SSD (single shot detector) or YOLO (you only look once)
# * Faster and better suited for devices
# * Will discuss using multiple detectors
# * The algorithm above had 1 detector, and predicted one bounding box and one class label
# * We can add more of them to detect several objects
# * Challenges
# * How do we efficiently add detectors?
# * How do we assign objects to detectors?
#
# * Convolutional Detectors
# * One way to add a lot of detectors quickly is to not use a dense layer at all
# * We replace the dense network above with two convolutional layers
# * Each layer will have 4 filters (so the output will have 4 channels)
# * We can apply a softmax to each pixel in one layer to get a category prediction
# * We can use the second layer for bounding box predictions
#
#
#
# <img src=../assets/network_diagrams/multi_object_identifier.png>
#
# In the case below our output CNN layers will have a shape of 5 pixels by 5 pixels,
# so we we'll assign each of these pixels to watch a box in the original input image.
# The area each pixel is watching for is called a prior box or an anchor.
#
# If the center of an object in in the box we assign it to that detector, otherwise a detector is assigned a background class.
#
# <img src=../assets/network_diagrams/Detectors.png>
#
# Another detail is instead of directly predicting the bounding box with make a prediction with respect to the prior box, we predict the x,y offsets to the new bounding box and the length and width scales.
#
#
#
#
# # Build It
# Same problem now with the possiblity of multiple objects
bb_multi=BBToyData(multi_object= True)
# +
plt.imshow(np.squeeze(bb_multi.X_train[10]))
# +
cnn_input=tf.keras.layers.Input( shape=bb_multi.X_train.shape[1:] ) # Shape here does not including the batch size
cnn_layer1=tf.keras.layers.Convolution2D(16, (4,4),strides=2,padding='same')(cnn_input) #50x50
cnn_activation=tf.keras.layers.LeakyReLU()(cnn_layer1)
#cnn_activation=tf.keras.layers.BatchNormalization()(cnn_activation)
#cnn_activation=tf.keras.layers.Dropout(0.5)(cnn_activation)
cnn_layer2=tf.keras.layers.Convolution2D(32, (4,4),strides=2,padding='same')(cnn_activation) #25x25
cnn_activation=tf.keras.layers.LeakyReLU()(cnn_layer2)
#cnn_activation=tf.keras.layers.Dropout(0.5)(cnn_activation)
#cnn_activation=tf.keras.layers.BatchNormalization()(cnn_activation)
cnn_layer3=tf.keras.layers.Convolution2D(64, (10,10),strides=5,padding='same')(cnn_activation) #5x5
cnn_activation=tf.keras.layers.LeakyReLU()(cnn_layer3)
#cnn_activation=tf.keras.layers.BatchNormlization()(cnn_activation)
#cnn_activation=tf.keras.layers.Dropout(0.5)(cnn_activation)
###Prediction Layers
prediction=tf.keras.layers.Convolution2D(4,(5,5),padding='same',activation='softmax')(cnn_activation)
#Softmax is applied to the last
bbox_offset=tf.keras.layers.Convolution2D(4,(5,5),padding='same')(cnn_activation)
model=tf.keras.models.Model(cnn_input,[prediction,bbox_offset])
model.compile(loss=['categorical_crossentropy','mse'],
optimizer='adam',
metrics=[])
model.summary()
test,test1=model.predict(bb_multi.X_train[0:1])
# +
def get_labels(input_labels):
Y_train=[[],[]]
for image_labels in input_labels:
class_features=np.zeros((5,5,4))
class_features[:,:,3]=1. #Defaults to no class
bbox_features=np.zeros((5,5,4))
bbox_features[:,:,2:]=1. #Scale =1
detected=[]
for i,(cat,bbox) in enumerate(image_labels):
bbox=[i/100. for i in bbox] # Normalize
bbox_center_x=int((bbox[0]+bbox[2]/2.)//0.2) #priorbox bin
bbox_center_y=int((bbox[1]+bbox[3]/2.)//0.2) #priorbox bin
bbox_offset_x=(bbox[0]+bbox[2]/2.)/0.2-(bbox_center_x+0.5)#priorbox bin
bbox_offset_y=(bbox[1]+bbox[3]/2.)/0.2-(bbox_center_y+0.5) #priorbox bin
bbox_scale_x=bbox[2]/0.2
bbox_scale_y=bbox[3]/0.2
# Y comes first here
class_features[bbox_center_y,bbox_center_x,cat]=1.
class_features[bbox_center_y,bbox_center_x,3]=0
bbox_features[bbox_center_y,bbox_center_x,:]=[bbox_offset_x,bbox_offset_y,bbox_scale_x,bbox_scale_y]
Y_train[0].append(np.expand_dims(class_features,0))
Y_train[1].append(np.expand_dims(bbox_features,0))
Y_train[0]=np.concatenate(Y_train[0])
Y_train[1]=np.concatenate(Y_train[1])
return Y_train
Y_train=get_labels(bb_multi.Y_train)
Y_develop=get_labels(bb_multi.Y_develop)
Y_test=get_labels(bb_multi.Y_test)
# +
#plt.imshow(np.argmax(Y_train[0][0],axis=-1))
bbox=Y_train[1][0]
plt.show()
for i in range(10):
index=np.random.randint(len(bb_multi.X_train))
plot_example(np.squeeze(bb_multi.X_train[index]),bb_multi.Y_train[index])
# -
print(len(bb_multi.X_train))
history=model.fit(bb_multi.X_train, Y_train,
batch_size=32, epochs=15, verbose=1,
validation_data=(bb_multi.X_develop,Y_develop)
)
plt.plot(history.history['loss']),plt.plot(history.history['val_loss'])
# +
def predictions_to_labels(image,cat_map,bbox_map,show_background=False):
# Each map is a 5x5 image
labels=[]
for y,r in enumerate(bbox_map):
for x,v in enumerate(r):
cat=np.argmax(cat_map[y,x,:])
# if cat_map[y,x,cat] < 0.9:continue
if not show_background and cat==3:continue
#Center - l/2
width=.2*bbox_map[y,x,2]
height=.2*bbox_map[y,x,3]
if cat !=3:
print(bbox_map[y,x,:])
print(cat_map[y,x,:])
x_start=(x+bbox_map[y,x,0]+.5)*.2 -width/2.
y_start=(y+bbox_map[y,x,1]+.5)*.2 -height/2.
labels.append([cat,[(x_start)*100,y_start*100,width*100,height*100]])
return labels
cat_map,bbox_map=model.predict(bb_multi.X_develop)
# +
for i in range(10):
labels=predictions_to_labels(np.squeeze(bb_multi.X_develop[i]),cat_map[i],bbox_map[i],show_background=True)
# labels=predictions_to_labels(np.squeeze(bb_multi.X_develop[i]),Y_develop[0][i],Y_develop[1][i],show_background=True)
plot_example(np.squeeze(bb_multi.X_develop[i]),labels)
print('Predicted Labels')
plt.imshow(np.hstack([cat_map[i,:,:,n] for n in range(4)]))
plt.show()
print('True Labels')
plt.imshow(np.hstack([Y_develop[0][i,:,:,n] for n in range(4)]))
plt.show()
# -
# ## A Fullly Functional Algorigthm
#
# There are still problems with the simple model above
# * What happens if more than one object matches a box?
# * You might see problems if the objects are really big
# * You might also see problems if the objects are really small
#
# The best object detection algorithm use several prior boxes per location: i.e SSD
#
# <img src="https://miro.medium.com/max/974/1*51joMGlhxvftTxGtA4lA7Q.png">
#
# * We used one convolutional map that was 5x5
# * These algorithms use several maps with different sizes (for smaller and larger objects)
# * We only used square prior boxes
# * These algorithms use several aspect ratio bounding boxes per point
#
# **These are well engineered, and can take some time to reproduce, so we'll use an existing implementation**
#
# Using this model is a good example of how everything we've talked about can be rolled to a one analysis
# # Don't be a hero part2
#
# Use an existing package from github
# https://github.com/pierluigiferrari/ssd_keras
#
# * Fairly Normal Open Source Package
# * Limited Docs
# * Some Examples
# * Not Super Easy to use
# * I'll try to point out where what you learned above will help you
#
# !git clone https://github.com/pierluigiferrari/ssd_keras.git
os.chdir('./ssd_keras')
# # Back to the Goal
# We are trying to design a system to help with malaria diagnosis
# * We have blood smear slides labeled with bounding boxes
# * Each bounding box identifies an infected cell and its stage of development
# +
cat_dict={}
cat_dict["background"]=0
cat_dict["ring"]=1
cat_dict['trophozoite']=2
cat_dict['schizont']=3
cat_dict['gametocyte']=4
int_2_cat={}
for i,v in cat_dict.items():
int_2_cat[v]=i
# -
# ## Ring stage example
# <img src="https://www.mcdinternational.org/trainings/malaria/english/DPDx5/images/ParasiteImages/M-R/Malaria/falciparum/Pfal-rings-atlasdx.JPG">
#
#
# # Our Annotations come in a json format
# * These generally look like dictionaries to python users
#
#
#
#Data is stored on Talapas
images_dir="/projects/bgmp/shared/2019_ML_workshop/datasets/BBBC041o/"
annotations_train="/projects/bgmp/shared/2019_ML_workshop/datasets/BBBC041o/train.json"
annotations_develop="/projects/bgmp/shared/2019_ML_workshop/datasets/BBBC041o/develop.json"
annotations_test="/projects/bgmp/shared/2019_ML_workshop/datasets/BBBC041o/test.json"
# +
import json
train_label=json.load(open(annotations_train))
print(train_label.keys())
print("Image Data",train_label['images'][0])
print("Annotations 0",train_label['annotations'][0])
print("Annotations 1",train_label['annotations'][1])
print("Annotations 2",train_label['annotations'][2])
print("Annotations 3",train_label['annotations'][3])
# -
# In the above, the list of annotations is matched to images using the tag image_id
# * This is informational, but our package knows how to read these files
# # Practice Working With Bounding Boxes
# Use the lists above to answer the following questions:
# # Question: How Many Annotations Are in the Train Set?
#
# # What Box Has The Largest Area?
# +
## Important data checks
all_bboxs=[ anno['bbox'] for anno in train_label['annotations']]
all_classes=[ anno['category_id'] for anno in train_label['annotations']]
for select in range(1,5):
widths=[bb[2] for bb,cat in zip(all_bboxs,all_classes) if cat == select ]
heights=[bb[3] for bb,cat in zip(all_bboxs,all_classes) if cat == select ]
areas=[bb[3]*bb[2] for bb,cat in zip(all_bboxs,all_classes) if cat == select ]
print( len(widths), int_2_cat[select], " in training set" )
print("Average Width", np.mean(widths),'pixels')
print("Average Height", np.mean(heights),'pixels')
print("Max Width", np.max(widths),'pixels')
print("Max Height", np.max(heights),'pixels')
print("Min Width", np.min(widths),'pixels')
print("Min Height", np.min(heights),'pixels')
print("__________________")
# -
# * Our targets are mostly trophozites
# * Our targets are fairly square
# * Our targets are ~70-255 pixels in size
# # Building Our Model - Part 1 Data
# Lots of imports for our package
# +
import h5py
import keras
from keras.optimizers import Adam
from keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau, TerminateOnNaN, CSVLogger
from keras import backend as K
from keras.models import load_model
from math import ceil
import numpy as np
from matplotlib import pyplot as plt
from ssd_encoder_decoder.ssd_input_encoder import SSDInputEncoder
from ssd_encoder_decoder.ssd_output_decoder import decode_detections, decode_detections_fast
from data_generator.object_detection_2d_data_generator import DataGenerator
from data_generator.object_detection_2d_misc_utils import apply_inverse_transforms
from data_generator.data_augmentation_chain_variable_input_size import DataAugmentationVariableInputSize
from data_generator.data_augmentation_chain_constant_input_size import DataAugmentationConstantInputSize
from data_generator.data_augmentation_chain_original_ssd import SSDDataAugmentation
#from models.keras_ssd7 import build_model
from models.keras_ssd300 import ssd_300 as build_model
from keras_loss_function.keras_ssd_loss import SSDLoss
from keras_layers.keras_layer_AnchorBoxes import AnchorBoxes
from keras_layers.keras_layer_DecodeDetections import DecodeDetections
from keras_layers.keras_layer_DecodeDetectionsFast import DecodeDetectionsFast
import matplotlib.patches as patches
# -
# ## Remember how we needed to properly preprocess the data for existing packages?
# This one has custom data generators for that
# +
img_height = 1200 # Height of the input images
img_width = 1600 # Width of the input images
img_channels = 3 # Number of color channels of the input images
train_dataset = DataGenerator(load_images_into_memory=True, hdf5_dataset_path=None)
image_augmentation=SSDDataAugmentation(img_height,img_width)
train_dataset.parse_json( images_dirs=images_dir,
annotations_filenames=[annotations_train],
ground_truth_available=True,
include_classes='all'
)
develop_dataset = DataGenerator(load_images_into_memory=True, hdf5_dataset_path=None)
develop_dataset.parse_json( images_dirs=images_dir,
annotations_filenames=[annotations_develop],
ground_truth_available=True,
include_classes='all'
)
test_dataset = DataGenerator(load_images_into_memory=True, hdf5_dataset_path=None)
test_dataset.parse_json( images_dirs=images_dir,
annotations_filenames=[annotations_test],
ground_truth_available=True,
include_classes='all'
)
# +
plot_generator = train_dataset.generate(batch_size=10,
shuffle=False,
label_encoder=None,
returns={'processed_images',
'processed_labels',
'filenames'},
keep_images_without_gt=False)
# -
# Plot and draw example data
# +
#Grab a batch of images
batch_images, batch_labels, batch_filenames = next(plot_generator)
print(len(batch_images))
#These labels are in corner representation (xmin,ymin) (xmax,ymax)
# Need to convert to (xmin,ymin),(lenth,width)
def convert_ssd_labels(label):
if len(label)==5:
return [int(label[0]),[label[1],label[2],label[3]-label[1],label[4]-label[2]]]
if len(label)==6: #This are labels that also include a prediction
return [int(label[0]),[label[2],label[3],label[4]-label[2],label[5]-label[3]]]
for i in range(10):
labels=[convert_ssd_labels(l) for l in batch_labels[i]]
print(labels)
plot_example(batch_images[i],labels)
# -
# # Build the Model - Part 2 The Model
# * Step one: Panic!
# * Step two: stay calm and follow the default configurations (I pulled these from a notebook example)
#
# * Important things you need to know (from above)
# * Your CNN layers make as many predictions as there output maps
# * Your training annotations needed to be properly encoded
# * You eventually need to decode the training labels
# * This is the same thing you did before, but now with more layers
#
# **The Really Important Part**: All these settings, scales, aspect, ratios, etc., Must match. The setting for the Model must be the same as the encoder and the decoder
#
#
#
#
# +
n_classes=4
scales = [0.1, 0.2, 0.37, 0.54, 0.71, 0.88, 1.05]
aspect_ratios = [[1.0, 2.0, 0.5],
[1.0, 2.0, 0.5, 3.0, 1.0/3.0],
[1.0, 2.0, 0.5, 3.0, 1.0/3.0],
[1.0, 2.0, 0.5, 3.0, 1.0/3.0],
[1.0, 2.0, 0.5],
[1.0, 2.0, 0.5]] # The anchor box aspect ratios used in the original SSD300; the order matters
two_boxes_for_ar1 = True
steps = [8, 16, 32, 64, 100, 300] # The space between two adjacent anchor box center points for each predictor layer.
offsets = [0.5, 0.5, 0.5, 0.5, 0.5, 0.5] # The offsets of the first anchor box center points from the top and left borders of the image as a fraction of the step size for each predictor layer.
clip_boxes = False # Whether or not to clip the anchor boxes to lie entirely within the image boundaries
variances = [0.1, 0.1, 0.2, 0.2] # The variances by which the encoded target coordinates are divided as in the original implementation
normalize_coords = True
mean_color = [123, 117, 104] # The per-channel mean of the images in the dataset. Do not change this value if you're using any of the pre-trained weights.
model = build_model(image_size=(img_height, img_width, img_channels),
n_classes=n_classes,
mode='training',
l2_regularization=0.0005,
scales=scales,
aspect_ratios_per_layer=aspect_ratios,
two_boxes_for_ar1=two_boxes_for_ar1,
steps=steps,
offsets=offsets,
clip_boxes=clip_boxes,
variances=variances,
normalize_coords=normalize_coords,
subtract_mean=mean_color)
predictor_sizes = [model.get_layer('conv4_3_norm_mbox_conf').output_shape[1:3],
model.get_layer('fc7_mbox_conf').output_shape[1:3],
model.get_layer('conv6_2_mbox_conf').output_shape[1:3],
model.get_layer('conv7_2_mbox_conf').output_shape[1:3],
model.get_layer('conv8_2_mbox_conf').output_shape[1:3],
model.get_layer('conv9_2_mbox_conf').output_shape[1:3]]
ssd_input_encoder = SSDInputEncoder(img_height=img_height,
img_width=img_width,
n_classes=n_classes,
predictor_sizes=predictor_sizes,
scales=scales,
aspect_ratios_per_layer=aspect_ratios,
two_boxes_for_ar1=two_boxes_for_ar1,
steps=steps,
offsets=offsets,
clip_boxes=clip_boxes,
variances=variances,
matching_type='multi',
pos_iou_threshold=0.5,
neg_iou_limit=0.5,
normalize_coords=normalize_coords)
# +
batch_size=5
train_generator = train_dataset.generate(batch_size=batch_size,
shuffle=True,
transformations=[image_augmentation],
label_encoder=ssd_input_encoder,
returns={'processed_images',
'encoded_labels'},
keep_images_without_gt=False)
develop_generator = develop_dataset.generate(batch_size=batch_size,
shuffle=False,
transformations=[],
label_encoder=ssd_input_encoder,
returns={'processed_images',
'encoded_labels'},
keep_images_without_gt=False)
adam = Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
ssd_loss = SSDLoss(neg_pos_ratio=3, alpha=1.0)
model.compile(optimizer=adam, loss=ssd_loss.compute_loss)
# +
model.summary()
_1,_2=next(train_generator)
print(_2.shape)
# -
# ```
# conv4_3_norm_mbox_conf (Conv2D) (None, 150, 200, 20) 92180 conv4_3_norm[0][0]
# __________________________________________________________________________________________________
# fc7_mbox_conf (Conv2D) (None, 75, 100, 30) 276510 fc7[0][0]
# __________________________________________________________________________________________________
# conv6_2_mbox_conf (Conv2D) (None, 38, 50, 30) 138270 conv6_2[0][0]
# __________________________________________________________________________________________________
# conv7_2_mbox_conf (Conv2D) (None, 19, 25, 30) 69150 conv7_2[0][0]
# __________________________________________________________________________________________________
# conv8_2_mbox_conf (Conv2D) (None, 17, 23, 20) 46100 conv8_2[0][0]
# __________________________________________________________________________________________________
# conv9_2_mbox_conf (Conv2D) (None, 15, 21, 20) 46100 conv9_2[0][0]
# _________________________________________________________________________________________________
# input_1 (InputLayer) (None, 1200, 1600, 3) 0
# ```
# ## Detectors at different resolution
# ### Boxes layer size
#
# 4 boxes per pixel * 4 predictions per box = 16 measurements
# ### Boxes layer size
#
# 5 (4+1 background) classes per box = 4*5=20
#
# | Map Size |Pixels in original image per prediction feature pixel|
# | ------------- |-------------|
# |150x200 | 8x8 |
# |75x100 | 16x16 |
# |38x50 | ~32x32 |
# |19x25 | ~64x64 |
# |17x23 | ~64x64 |
# |15x21 | ~64x64 |
#
#
# **One thing to notice immedialte is that our 'biggest' detector starts out at 64x64 pixels**
# * What was the average size of our objects?
# +
model.load_weights("/projects/bgmp/shared/2019_ML_workshop/models/ssd300.h5")
if False:
final_epoch = 500
history = model.fit_generator(generator=train_generator,
epochs=final_epoch,
steps_per_epoch=len(train_dataset.images)//batch_size,
validation_data=develop_generator,
validation_steps=len(develop_dataset.images)//batch_size,
)
#model.save_weights("/projects/bgmp/shared/2019_ML_workshop/models/ssd300.h5")
# +
example_generator = develop_dataset.generate(batch_size=batch_size,
shuffle=False,
transformations=[],
label_encoder=ssd_input_encoder,
returns={'processed_images',
'processed_labels'},
keep_images_without_gt=False)
# +
batch_images,batch_labels=next(example_generator)
y_pred = model.predict(batch_images)
y_pred_decoded = decode_detections(y_pred,
confidence_thresh=0.3,
iou_threshold=0.1,
top_k=200,
normalize_coords=True,#normalize_coords,
img_height=img_height,
img_width=img_width)
for i in range(batch_size):
labels=[convert_ssd_labels(l) for l in y_pred_decoded[i]]
plot_example(batch_images[i],labels)
# -
# ## Lets evaluate the model and do some experiments
#
# We are far from perfect, but let's evaluate the model
#
# ## Testing Our Model
#
# How good is it? An important part of these algorithms is evaluating the quality of matches.
#
# ## Intersect Over Union
# Intersect over union
# <img src=https://www.pyimagesearch.com/wp-content/uploads/2016/09/iou_equation.png>
#
# This is a very useful quanity
#
# * 1 if the agreement is perfect, 0 if the boxes don't overlap
# * Some entries may have several overlapping boxes
# * IOU can be a measure of how similar they are
# * If IOU > 0.1 only use the highest confidence bounding box
# (iou_threshold)
# * Does a Truth Box have a match
# * Need to define how good a match counts as a match
# * If IOU > .5 a box matches the truth
#
#
#
#
#
#
#
#
# +
develop_for_pred_gen = develop_dataset.generate(batch_size=1,
shuffle=False,
transformations=[],
label_encoder=ssd_input_encoder,
returns={'processed_images',
},
keep_images_without_gt=False)
true_labels=develop_dataset.labels
y_pred=model.predict_generator(develop_for_pred_gen,len(develop_dataset.images))
print("Done with Prediction")
# -
print("Decode First Five Images")
y_pred_decoded = decode_detections(y_pred[0:5],
confidence_thresh=0.0, ##Not cofidence cut
iou_threshold=0.1, # Overlap removal
top_k=200, #Only the best 200 boxes
normalize_coords=True,#normalize_coords,
img_height=img_height,
img_width=img_width)
# # Lets Look at One Image
# +
image_index=2
develop_images=develop_dataset.images
plt.imshow(develop_dataset.images[image_index])
plt.show()
print("labels",y_pred_decoded[image_index])
print("Truth", develop_dataset.labels[image_index])
# -
# # Decoded Labels for prediction are a list
# each element is in the form [best category, confidence, x,y,length,width]
# sorted with best result last
#
# +
all_categories=[l[0] for l in y_pred_decoded[image_index] ]
all_confidences=[l[1] for l in y_pred_decoded[image_index]]
plt.hist(all_categories)
plt.xlabel("Classes")
plt.show()
plt.hist(all_confidences)
plt.show("All Confidences")
# +
# Not very confident, put lets look at the best two boxes
plot_labels=[convert_ssd_labels(l) for l in y_pred_decoded[image_index][-10:]]
print("Best 10 Boxes")
plot_example(develop_images[image_index],plot_labels)
print("Truth Boxes")
plot_labels=[convert_ssd_labels(l) for l in true_labels[image_index]]
plot_example(develop_images[image_index],plot_labels)
# -
# # Not Bad, Lets make a more Statistical Statment
#
# Lets start by deciding anything with a confidence greater than 0.4 is a match (I encourage you to play with this)
# +
y_pred_decoded = decode_detections(y_pred,
confidence_thresh=0.4, ##Add a Confidence Cut
iou_threshold=0.1,
top_k=200,
normalize_coords=True,
img_height=img_height,
img_width=img_width)
# -
# # Lets Look at a Bunch of Distributions
#
#
# +
##
def area_minmax(bbox):
xmin,ymin,xmax,ymax=bbox
return(xmax-xmin)*(ymax-ymin)
def intersect_minmax(bbox1,bbox2):
xmin1,ymin1,xmax1,ymax1=bbox1
xmin2,ymin2,xmax2,ymax2=bbox2
xintmin=max(xmin1,xmin2)
yintmin=max(ymin1,ymin2)
xintmax=min(xmax1,xmax2)
yintmax=min(ymax1,ymax2)
int_width=(xintmax-xintmin)
int_height=(yintmax-yintmin)
if int_width <0 or int_height <0 :return 0
return int_width*int_height
def iou(bbox1,bbox2):
intarea=intersect_minmax(bbox1,bbox2)
u=area_minmax(bbox1)+area_minmax(bbox2)-intarea
return intarea/u
def flatten_labels(labels):
values=[]
for l in labels:
for n in l:
values.append(n)
return values
t_all=flatten_labels(true_labels)
p_all=flatten_labels(y_pred_decoded)
true_area=[area_minmax(l[1:5]) for l in t_all]
pred_area=[area_minmax(l[2:6]) for l in p_all]
true_width=[l[3]-l[1] for l in t_all]
pred_width=[l[4]-l[2] for l in p_all]
true_x=[l[1] for l in t_all]
pred_x=[l[2] for l in p_all]
true_y=[l[2] for l in t_all]
pred_y=[l[3] for l in p_all]
true_cat=[l[0] for l in t_all]
pred_cat=[l[0] for l in p_all]
plt.hist(true_area,range=(50*50,200*200),bins=10,label='true',histtype='step')
plt.hist(pred_area,range=(50*50,200*200),bins=10,label='pred',histtype='step')
plt.xlabel("Area")
plt.legend()
plt.show()
plt.hist(true_width,range=(50,200),bins=10,label='true',histtype='step')
plt.hist(pred_width,range=(50,200),bins=10,label='pred',histtype='step')
plt.xlabel("Width")
plt.legend()
plt.show()
plt.hist(true_cat,range=(0,10),bins=10,label='true',histtype='step')
plt.hist(pred_cat,range=(0,10),bins=10,label='pred',histtype='step')
plt.xlabel("Category")
plt.legend()
plt.show()
plt.hist(true_x,range=(0,1600),bins=10,label='true',histtype='step')
plt.hist(pred_x,range=(0,1600),bins=10,label='pred',histtype='step')
plt.xlabel("X")
plt.legend()
plt.show()
plt.hist(true_x,range=(0,1200),bins=10,label='true',histtype='step')
plt.hist(pred_x,range=(0,1200),bins=10,label='pred',histtype='step')
plt.xlabel("Y")
plt.legend()
plt.show()
# -
# # What do you see?
# # How well do the results match the truth
#
# Make another choice if a has an IOU > 0.1 it has a match (I also encourage you to play with this, remember we're creating algorithms and the often means making choices to be more or less conservative )
# +
#match best box
def match(truth,pred):
best=[]
for t in truth:
matches=[ [iou(t[1:],p[2:]),i] for i,p in enumerate(pred)]
matches.sort()
if matches!=[]:
best.append(matches[-1]) # largest iou
else:
best.append([0,-1])
return best
matches=[match(t,p) for t,p in zip(true_labels,y_pred_decoded)]
n_no_matches=0
n_matches=0
correct=0
for index,image in enumerate(matches):
for li,m in enumerate(image):
if m[0] < .1 or m[1]==-1:n_no_matches+=1 # no match
else:
n_matches+=1 # no match
y_pred_decoded[index][m[1]][0]==true_labels[index][li][0]
correct+=1
print(n_matches/(n_no_matches+n_matches)*100,"% True Boxes Matched")
print(correct/(n_no_matches+n_matches)*100,"% True Boxes Matched and identified")
print(len(p_all)-n_matches," wrong boxes", correct,"correct boxes" )
# -
# # That was a Lot of Code, Some Take Aways:
#
# ## Multi-Box Detectors can be used to find objects in images
# * You can write them yourself, but a lot of prexisting packages are around
# * The magic that makes this work is using fully convolutional predictors (no dense layers at the end)
# * This also means you can use any size image
# * You have to encode annotations, and decode them
# * How exactly this is done will depend on the exact algorithm
# * There are a lot of ways to mess this up, so don't get discouraged if it dosen't work at first
# * One of the first things you can tune with these models is to pick better anchor box sizes and aspect ratios!
#
# # Other ways to do this
# * Yolo (you only look once)
# * Recursive CNNS (similar to scaning an image like we did last lecture)
#
# # If you finish early, I encourage you to play around with this or the previous datasets for practice, and we will wrap up at the end
#
#
#
#
#
| notebooks/8-Malaria_and_SSD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''pt1-env'': conda)'
# name: python3
# ---
# **Examples of Collaborative Filtering based Recommendation Systems**
import sys, os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn.metrics as metrics
import numpy as np
from sklearn.neighbors import NearestNeighbors
from scipy.spatial.distance import correlation, cosine
import ipywidgets as widgets
from IPython.display import display, clear_output
from sklearn.metrics import pairwise_distances
from sklearn.metrics import mean_squared_error
from math import sqrt
from contextlib import contextmanager
# +
# M 是 user-item 评分矩阵, 其中评分是1-10的整数
M = np.asarray([[3, 7, 4, 9, 9, 7],
[7, 0, 5, 3, 8, 8],
[7, 5, 5, 0, 8, 4],
[5, 6, 8, 5, 9, 8],
[5, 8, 8, 8, 10, 9],
[7, 7, 0, 4, 7, 8]])
M = pd.DataFrame(M)
# 定义全局变量 k, metric 后续可以被user 改变
global k, metric
k = 4
# 相关性相似度
metric = 'cosine'
# -
M
from sklearn.metrics.pairwise import cosine_similarity
u_0 = [3, 7, 4, 9, 9, 7]
u_1 = [7, 0, 5, 3, 8, 8]
cosine_similarity(np.array([u_0]), np.array([u_1]))
# **User-based Recommendation Systems**
#get cosine similarities for ratings matrix M; pairwise_distances returns the distances between ratings and hence
#similarities are obtained by subtracting distances from 1
cosine_sim = 1 - pairwise_distances(M, metric="cosine")
#Cosine similarity matrix
pd.DataFrame(cosine_sim)
M
#get pearson similarities for ratings matrix M
pearson_sim = 1 - pairwise_distances(M, metric="correlation")
#Pearson correlation similarity matrix
pd.DataFrame(pearson_sim)
# This function finds k similar users given the user_id and ratings matrix M
# Note that the similarities are same as obtained via using pairwise_distances
def findksimilarusers(user_id, ratings, metric=metric, k=k):
similarities=[]
indices=[]
model_knn = NearestNeighbors(metric = metric, algorithm = 'brute')
model_knn.fit(ratings)
distances, indices = model_knn.kneighbors(ratings.iloc[user_id-1, :].values.reshape(1, -1), n_neighbors = k+1)
similarities = 1 - distances.flatten()
print('{0} most similar users for User {1}:\n'.format(k,user_id))
for i in range(0, len(indices.flatten())):
if indices.flatten()[i]+1 == user_id:
continue
else:
print('{0}: User {1}, with similarity of {2}'.format(i, indices.flatten()[i]+1, similarities.flatten()[i]))
return similarities, indices
M
similarities, indices = findksimilarusers(1, M, metric='cosine')
similarities,indices = findksimilarusers(1, M, metric='correlation')
#This function predicts rating for specified user-item combination based on user-based approach
def predict_userbased(user_id, item_id, ratings, metric = metric, k=k):
prediction=0
similarities, indices=findksimilarusers(user_id, ratings,metric, k) #similar users based on cosine similarity
mean_rating = ratings.loc[user_id-1,:].mean() #to adjust for zero based indexing
sum_wt = np.sum(similarities)-1
product=1
wtd_sum = 0
for i in range(0, len(indices.flatten())):
if indices.flatten()[i]+1 == user_id:
continue
else:
ratings_diff = ratings.iloc[indices.flatten()[i],item_id-1]-np.mean(ratings.iloc[indices.flatten()[i],:])
product = ratings_diff * (similarities[i])
wtd_sum = wtd_sum + product
prediction = int(round(mean_rating + (wtd_sum/sum_wt)))
print('\nPredicted rating for user {0} -> item {1}: {2}'.format(user_id,item_id,prediction))
return prediction
predict_userbased(3, 4, M);
# **Item-based Recommendation Systems**
# +
#This function finds k similar items given the item_id and ratings matrix M
def findksimilaritems(item_id, ratings, metric=metric, k=k):
similarities = []
indices = []
ratings = ratings.T
model_knn = NearestNeighbors(metric = metric, algorithm = 'brute')
model_knn.fit(ratings)
distances, indices = model_knn.kneighbors(ratings.iloc[item_id-1, :].values.reshape(1, -1), n_neighbors = k+1)
similarities = 1 - distances.flatten()
print('{0} most similar items for item {1}:\n'.format(k,item_id))
for i in range(0, len(indices.flatten())):
if indices.flatten()[i]+1 == item_id:
continue;
else:
print('{0}: Item {1} :, with similarity of {2}'.format(i,indices.flatten()[i]+1, similarities.flatten()[i]))
return similarities,indices
# -
similarities, indices = findksimilaritems(3, M)
#This function predicts the rating for specified user-item combination based on item-based approach
def predict_itembased(user_id, item_id, ratings, metric = metric, k=k):
prediction = wtd_sum =0
similarities, indices=findksimilaritems(item_id, ratings) #similar users based on correlation coefficients
sum_wt = np.sum(similarities)-1
product=1
for i in range(0, len(indices.flatten())):
if indices.flatten()[i] + 1 == item_id:
continue;
else:
product = ratings.iloc[user_id-1,indices.flatten()[i]] * (similarities[i])
wtd_sum = wtd_sum + product
prediction = int(round(wtd_sum/sum_wt))
print('\nPredicted rating for user {0} -> item {1}: {2}'.format(user_id,item_id,prediction))
return prediction
prediction = predict_itembased(1, 3, M)
#This function is used to compute adjusted cosine similarity matrix for items
def computeAdjCosSim(M):
sim_matrix = np.zeros((M.shape[1], M.shape[1]))
M_u = M.mean(axis=1) # means
for i in range(M.shape[1]):
for j in range(M.shape[1]):
if i == j:
sim_matrix[i][j] = 1
else:
if i<j:
sum_num = sum_den1 = sum_den2 = 0
for k,row in M.loc[:,[i,j]].iterrows():
if ((M.loc[k,i] != 0) & (M.loc[k,j] != 0)):
num = (M[i][k]-M_u[k]) * (M[j][k]-M_u[k])
den1= (M[i][k]-M_u[k]) ** 2
den2= (M[j][k]-M_u[k]) ** 2
sum_num = sum_num + num
sum_den1 = sum_den1 + den1
sum_den2 = sum_den2 + den2
else:
continue
den=(sum_den1**0.5)*(sum_den2 ** 0.5)
if den!=0:
sim_matrix[i][j] = sum_num/den
else:
sim_matrix[i][j] = 0
else:
sim_matrix[i][j] = sim_matrix[j][i]
return pd.DataFrame(sim_matrix)
adjcos_sim = computeAdjCosSim(M)
adjcos_sim
# This function finds k similar items given the item_id and ratings matrix M
def findksimilaritems_adjcos(item_id, ratings, k=k):
sim_matrix = computeAdjCosSim(ratings)
similarities = sim_matrix[item_id-1].sort_values(ascending=False)[:k+1].values
indices = sim_matrix[item_id-1].sort_values(ascending=False)[:k+1].index
print('{0} most similar items for item {1}:\n'.format(k,item_id))
for i in range(0, len(indices)):
if indices[i]+1 == item_id:
continue
else:
print('{0}: Item {1} :, with similarity of {2}'.format(i,indices[i]+1, similarities[i]))
return similarities, indices
similarities, indices = findksimilaritems_adjcos(3,M)
#This function predicts the rating for specified user-item combination for adjusted cosine item-based approach
#As the adjusted cosine similarities range from -1,+1, sometimes the predicted rating can be negative or greater than max value
#Hack to deal with this: Rating is set to min if prediction is negative, Rating is set to max if prediction is above max
def predict_itembased_adjcos(user_id, item_id, ratings):
prediction=0
similarities, indices = findksimilaritems_adjcos(item_id, ratings) #similar users based on correlation coefficients
sum_wt = np.sum(similarities) - 1
product = 1
wtd_sum = 0
for i in range(0, len(indices)):
if indices[i]+1 == item_id:
continue
else:
product = ratings.iloc[user_id-1,indices[i]] * (similarities[i])
wtd_sum = wtd_sum + product
prediction = int(round(wtd_sum/sum_wt))
if prediction < 0:
prediction = 1
elif prediction >10:
prediction = 10
print('\nPredicted rating for user {0} -> item {1}: {2}'.format(user_id,item_id,prediction))
return prediction
prediction=predict_itembased_adjcos(3, 4, M)
adjcos_sim
#This function utilizes above function to recommend items for selected approach. Recommendations are made if the predicted
#rating for an item is greater than or equal to 6, and the items has not been rated already
def recommendItem(user_id, item_id, ratings):
if user_id < 1 or user_id > 6 or type(user_id) is not int:
print('Userid does not exist. Enter numbers from 1-6')
else:
ids = ['User-based CF (cosine)','User-based CF (correlation)','Item-based CF (cosine)',
'Item-based CF (adjusted cosine)']
approach = widgets.Dropdown(options=ids, value=ids[0],
description='Select Approach', width='500px')
def on_change(change):
prediction = 0
clear_output(wait=True)
if change['type'] == 'change' and change['name'] == 'value':
if (approach.value == 'User-based CF (cosine)'):
metric = 'cosine'
prediction = predict_userbased(user_id, item_id, ratings, metric)
elif (approach.value == 'User-based CF (correlation)') :
metric = 'correlation'
prediction = predict_userbased(user_id, item_id, ratings, metric)
elif (approach.value == 'Item-based CF (cosine)'):
prediction = predict_itembased(user_id, item_id, ratings)
else:
prediction = predict_itembased_adjcos(user_id,item_id,ratings)
if ratings[item_id-1][user_id-1] != 0:
print('Item already rated')
else:
if prediction>=6:
print('\nItem recommended')
else:
print('Item not recommended')
approach.observe(on_change)
display(approach)
#check for incorrect entries
recommendItem(-1, 3, M)
recommendItem(3, 4, M)
recommendItem(3, 4, M)
recommendItem(3, 4, M)
recommendItem(3, 4, M)
# if the item is already rated, it is not recommended
recommendItem(2, 1, M)
# This is a quick way to temporarily suppress stdout in particular code section
@contextmanager
def suppress_stdout():
with open(os.devnull, "w") as devnull:
old_stdout = sys.stdout
sys.stdout = devnull
try:
yield
finally:
sys.stdout = old_stdout
#This is final function to evaluate the performance of selected recommendation approach and the metric used here is RMSE
#suppress_stdout function is used to suppress the print outputs of all the functions inside this function. It will only print
#RMSE values
def evaluateRS(ratings):
ids = ['User-based CF (cosine)','User-based CF (correlation)','Item-based CF (cosine)','Item-based CF (adjusted cosine)']
approach = widgets.Dropdown(options=ids, value=ids[0],description='Select Approach', width='500px')
n_users = ratings.shape[0]
n_items = ratings.shape[1]
prediction = np.zeros((n_users, n_items))
prediction= pd.DataFrame(prediction)
def on_change(change):
clear_output(wait=True)
with suppress_stdout():
if change['type'] == 'change' and change['name'] == 'value':
if (approach.value == 'User-based CF (cosine)'):
metric = 'cosine'
for i in range(n_users):
for j in range(n_items):
prediction[i][j] = predict_userbased(i+1, j+1, ratings, metric)
elif (approach.value == 'User-based CF (correlation)') :
metric = 'correlation'
for i in range(n_users):
for j in range(n_items):
prediction[i][j] = predict_userbased(i+1, j+1, ratings, metric)
elif (approach.value == 'Item-based CF (cosine)'):
for i in range(n_users):
for j in range(n_items):
prediction[i][j] = predict_userbased(i+1, j+1, ratings)
else:
for i in range(n_users):
for j in range(n_items):
prediction[i][j] = predict_userbased(i+1, j+1, ratings)
MSE = mean_squared_error(prediction, ratings)
RMSE = round(sqrt(MSE),3)
print("RMSE using {0} approach is: {1}".format(approach.value,RMSE))
approach.observe(on_change)
display(approach)
evaluateRS(M)
evaluateRS(M)
# **Thanks for reading this notebook**
| CF/cf_examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Exploring the Lorenz System of Differential Equations
# In this Notebook we explore the Lorenz system of differential equations:
#
# $$
# \begin{aligned}
# \dot{x} & = \sigma(y-x) \\
# \dot{y} & = \rho x - y - xz \\
# \dot{z} & = -\beta z + xy
# \end{aligned}
# $$
#
# This is one of the classic systems in non-linear differential equations. It exhibits a range of different behaviors as the parameters ($\sigma$, $\beta$, $\rho$) are varied.
# ## Imports
# First, we import the needed things from IPython, NumPy, Matplotlib and SciPy.
# %matplotlib inline
from ipywidgets import interact, interactive
from IPython.display import clear_output, display, HTML
# +
import numpy as np
from scipy import integrate
from matplotlib import pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib.colors import cnames
from matplotlib import animation
# -
# ## Computing the trajectories and plotting the result
# We define a function that can integrate the differential equations numerically and then plot the solutions. This function has arguments that control the parameters of the differential equation ($\sigma$, $\beta$, $\rho$), the numerical integration (`N`, `max_time`) and the visualization (`angle`).
def solve_lorenz(N=10, angle=0.0, max_time=4.0, sigma=10.0, beta=8./3, rho=28.0):
fig = plt.figure()
ax = fig.add_axes([0, 0, 1, 1], projection='3d')
ax.axis('off')
# prepare the axes limits
ax.set_xlim((-25, 25))
ax.set_ylim((-35, 35))
ax.set_zlim((5, 55))
def lorenz_deriv(x_y_z, t0, sigma=sigma, beta=beta, rho=rho):
"""Compute the time-derivative of a Lorenz system."""
x, y, z = x_y_z
return [sigma * (y - x), x * (rho - z) - y, x * y - beta * z]
# Choose random starting points, uniformly distributed from -15 to 15
np.random.seed(1)
x0 = -15 + 30 * np.random.random((N, 3))
# Solve for the trajectories
t = np.linspace(0, max_time, int(250*max_time))
x_t = np.asarray([integrate.odeint(lorenz_deriv, x0i, t)
for x0i in x0])
# choose a different color for each trajectory
colors = plt.cm.jet(np.linspace(0, 1, N))
for i in range(N):
x, y, z = x_t[i,:,:].T
lines = ax.plot(x, y, z, '-', c=colors[i])
plt.setp(lines, linewidth=2)
ax.view_init(30, angle)
plt.show()
return t, x_t
# Let's call the function once to view the solutions. For this set of parameters, we see the trajectories swirling around two points, called attractors.
t, x_t = solve_lorenz(angle=0, N=10)
# Using IPython's `interactive` function, we can explore how the trajectories behave as we change the various parameters.
w = interactive(solve_lorenz, angle=(0.,360.), N=(0,50), sigma=(0.0,50.0), rho=(0.0,50.0))
display(w)
# The object returned by `interactive` is a `Widget` object and it has attributes that contain the current result and arguments:
t, x_t = w.result
w.kwargs
# After interacting with the system, we can take the result and perform further computations. In this case, we compute the average positions in $x$, $y$ and $z$.
xyz_avg = x_t.mean(axis=1)
xyz_avg.shape
# Creating histograms of the average positions (across different trajectories) show that on average the trajectories swirl about the attractors.
plt.hist(xyz_avg[:,0])
plt.title('Average $x(t)$')
plt.hist(xyz_avg[:,1])
plt.title('Average $y(t)$')
| examples/notebooks/widgets/widgets_lorenz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
import pandas as pd
import numpy as np
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Dense, Embedding, LSTM, SpatialDropout1D, Bidirectional, Activation
from sklearn.model_selection import train_test_split
from keras.utils.np_utils import to_categorical
from keras.callbacks import EarlyStopping
from keras.layers import Dropout
# load numpy array from csv file
from numpy import loadtxt
# load array
X_train = loadtxt('x_train_small.csv', delimiter=',')
X_train_hyp = loadtxt('x_train_hyp_small.csv', delimiter=',')
Y_train = loadtxt('y_train_small.csv', delimiter=',')
# print the array
X_train
X_train_hyp
Y_train
# only for testing
X_train_hyp = X_train_hyp[:, :100]
VOCAB_SIZE = 1254
INPUT_LENGTH = 100 #3000
EMBEDDING_DIM = 128
# model
def build_model(vocab_size, embedding_dim, input_length):
model = Sequential()
model.add(Embedding(vocab_size, embedding_dim, input_length=input_length))
model.add(SpatialDropout1D(0.2))
model.add(Bidirectional(LSTM(128, dropout=0.2, recurrent_dropout=0.2, return_sequences=True)))
model.add((LSTM(41)))
model.add(Activation('softmax'))
return model
# +
model = build_model(VOCAB_SIZE, EMBEDDING_DIM, INPUT_LENGTH)
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
# +
epochs = 2
batch_size = 64
history = model.fit(X_train_hyp, Y_train, epochs=epochs, batch_size=batch_size,validation_split=0.1,callbacks=[EarlyStopping(monitor='val_loss', patience=3, min_delta=0.0001)])
# -
example_x = X_train_hyp[0]
print(np.shape(example_x))
temp = model.predict(X_train_hyp)
print(np.sum(temp[0]))
# print(len(temp)), temp
print(temp[0])
for i in temp:
print(np.argmax(i))
| deepmath/deephol/train/B_Skeleton_Architectures/bilstm_goals_hyp_rnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Qiskit 사용법 강의
# 안녕하세요,
# 이 노트북은 기본적인 Qiskit 사용법을 익히기 위한 노트북입니다. 간단한 양자회로를 Qiskit 라이브러리를 이용해 구성하고 사용하는 방법을 배우고 이전에 Johri 박사님과 함께 다루었던 GHZ 상태를 만드는 양자회로를 다시 한 번 복습할 것입니다.
# ### Qiskit 과 Backend 설치
# Python 환경에 익숙하지 않은 분들은 다음 셀을 실행해서 Qiskit과 IonQ backend 라이브러리를 설치하실 수 있습니다.
## hit shift+enter to run this cell
# IonQ backend installation
# !curl https://static.ionq.co/c501c899-8b6a-4aa3-8c51-1c4193943954/qiskit_ionq_provider-0.0.1.dev0%2Bd5a56c7-py3-none-any.whl -O -J
# !pip install qiskit_ionq_provider-0.0.1.dev0%2Bd5a56c7-py3-none-any.whl
# Qiskit installation
# !pip install qiskit
# Matplotlib, pylatexenc installation - for QuantumCircuit Visualization
# !pip install matplotlib
# !pip install pylatexenc
# for qsphere visualization
# !pip install seaborn
token_ionq = '94fROwsIfZ2W5r6e3JbECmeiridtFWIG'
# ## 양자회로 QuantumCircuit
import qiskit
qiskit.__qiskit_version__
# Qiskit에서 양자회로는 `QuantumCircuit` 클래스로 표현됩니다. 회로 초기화, 양자 게이트 추가 및 측정을 다뤄보겠습니다.
# +
# QuantumCircuit을 import합니다.
from qiskit import QuantumCircuit
# Quantum Circuit을 작성
qc = QuantumCircuit(3) # 3 qubit 양자회로를 만듭니다.
# 주어진 양자회로를 그려봅니다.
qc.draw()
# +
# 양자 게이트들을 추가해보겠습니다.
qc.h(0) # 0번 큐비트에 hadamard 게이트를 가합니다.
qc.cx(0,1) # CNOT 게이트를 가합니다. Control 큐비트 0번 , Target 큐비트 1번
qc.cx(0,2) # CNOT 게이트를 가합니다. Control 큐비트 0번 , Target 큐비트 2번
qc.draw()
# +
# 측정은 다음과 같이 합니다.
qc.measure_all()
qc.draw()
# -
# matplotlib을 이용하면 좀 더 이쁜 QuantumCircuit을 그릴수 있습니다.
qc.draw('mpl')
# ### 알아두면 좋은 것들
# +
# 처음 Quantum Circuit을 만들때 Classical registre를 할당하면 직접 어떤 큐빗의 정보가 어떤 레지스터에 저장될지 할당할 수 있습니다.
qc = QuantumCircuit(3, 3) # 3 qubit + 3 classcal register 양자회로를 만듭니다.
# 양자 게이트들을 추가해보겠습니다.
qc.h(0) # 0번 큐비트에 hadamard 게이트를 가합니다.
qc.cx(0,1) # CNOT 게이트를 가합니다. Control 큐비트 0번 , Target 큐비트 1번
qc.cx(0,2) # CNOT 게이트를 가합니다. Control 큐비트 0번 , Target 큐비트 2번
qc.measure((0,1,2), (0,1,2)) # i번 큐비트의 정보를 i번째 레지스터에 측정하여 저장합니다.
qc.draw('mpl')
# +
# QuantumCircuit은 서로 더할 수 있습니다.
qc1 = QuantumCircuit(3)
qc1.h(0)
qc1.barrier()
qc2 = QuantumCircuit(3)
qc2.cx(0,1)
qc2.cy(0,2)
qc = qc1 + qc2
qc.draw('mpl')
# -
# 양자게이트의 개수는 다음 함수로 불러올수 있습니다.
qc.size()
# 큐비트를 개별적으로 초기화할 수도 있습니다.
qc.reset(0)
qc.draw('mpl')
# 임의의 큐비트 백터 상태로 초기화하고 싶을 때는 `initialize` 함수를 이용합니다.
import numpy as np
qc = QuantumCircuit(3)
qc.initialize([1,0,0,0,0,0,0,1]/np.sqrt(2), [0,1,2])
qc.draw('mpl')
# ## Backend 와 Job
# 작성한 양자회로는 Backend를 이용해서 구동합니다. Backend는 양자회로를 구동할 수 있는 하드웨어나 시뮬레이터 소프트웨어를 말합니다. 주어진 양자회로를 수행하고 결과값을 출력해줍니다. IonQ 하드웨어를 사용하려면 IonQ 백엔드를 사용하고, 그 외에 하드웨어나 시뮬레이터를 사용하려면 해당하는 백엔드를 불러와서 사용하면 됩니다.
#
# Qiskit은 기본적으로 여러가지의 양자회로 시뮬레이터를 Backend로 제공합니다. Qiskit 라이브러리의 여러 시뮬레이터를 먼저 알아보고 IonQ의 Backend를 활용하는 법을 배워보겠습니다.
# +
# qiskit.Aer 라이브러리를 불러옵니다.
from qiskit import Aer
Aer.backends()
# -
# `qiskit.Aer`에서 기본적으로 제공하는 시뮬레이터는 총 4가지가 있습니다. 기본적으로 사용자의 컴퓨터에서 로컬하게 계산을 하기 때문에 간단한 연산확인에 유용합니다.
#
# * `QasmSimulator`는 실제 하드웨어의 작동을 모사합니다. 측정을 포함한 양자회로의 연산을 여러번 수행한뒤 측정값의 통계분포를 결과값으로 출력합니다.
# * `StatevectorSimulator`는 양자상태를 벡터형태로 기술하여 양자회로를 계산합니다.
# * `UnitarySimulator`는 유니터리 연산자 형태로 전체 양자회로를 계산합니다. 측정/큐비트 리셋은 사용할 수 없습니다.
# * `PulseSimulator`는 qiskit.pulse를 위해 사용됩니다. 오늘 강의에서는 다루지 않습니다.
#
# 무엇을 골라야 할지 잘 모르겠으면 `QasmSimulator`를 이용하면 될 것 같습니다. 오늘은 `QasmSimulator`만 짧게 다루고 넘어가겠습니다.
# GHZ 상태를 만드는 양자회로를 반환하는 함수를 미리 정의하겠습니다.
def ghz_circuit(n: int):
assert isinstance(n, int) and n > 0
qc = QuantumCircuit(n)
qc.h(0)
if n>1:
qc.cx(0, range(1,n))
qc.measure_all()
return qc
# 백엔드에서 양자회로를 수행하기 위해서는 두 가지 방법이 있습니다. `execute`를 사용하거나 직접 `backend.run` 함수를 사용하는 방법입니다.
#
# `execute` 함수는 백엔드의 설정에 맞춰서 transpile을 자동적으로 수행합니다. `QuantumCircuit` 클래스로 양자 회로를 만들어서 실행하면 백엔드에 맞추어서 트랜스파일을 한 뒤 실행해줍니다.
# +
# 회로를 실행하기 위한 `execute`함수를 불러옵니다.
from qiskit import execute
# Qubit 3개짜리 GHZ 상태를 만드는 양자회로를 만듭니다.
qc = ghz_circuit(3)
# QasmSimulator 백엔드를 불러옵니다.
backend = Aer.get_backend('qasm_simulator')
# 양자회로는 다음과 같이 수행합니다. `execute` 함수는 Job 오브젝트를 반환합니다.
job = execute(qc, backend)
# -
# 백엔드는 `Job` 오브젝트를 반환하여 연산의 수행상태와 결과를 확인할 수 있습니다. 반환되는 `job` 오브젝트를 꼭 저장해두는 것이 좋습니다.
# 연산의 진행상태는 다음 함수로 확인가능합니다.
job.status()
# +
# 연산 결과는 `result` 함수로 확인가능합니다.
result = job.result()
# 측정 결과를 히스토그램으로 그려보겠습니다.
from qiskit.visualization import plot_histogram
plot_histogram(result.get_counts())
# +
# excute 함수는 여러 run_config 변수들을 받습니다. 이 변수들을 활용해서 연산설정을 변경할 수 있습니다.
# 측정을 10만번을 수행하고, statevector simulation 방법을 사용하고, single (16bit) 정확도로 연산을 수행하는 옵션입니다.
job = execute(qc, backend, shots = 10e5, method = 'statevector', precision = 'single')
plot_histogram(job.result().get_counts())
# -
# `job` 오브젝트는 진행상태를 확인하기 위한 다양한 함수를 제공합니다.
# +
# job id는 다음과 같이 확인합니다.
job.job_id()
# job 상태는 다음 함수로 확인합니다.
print(job.status())
# job 상태를 확인할때 유용한 함수들입니다.
print('job.done: ', job.done())
print('job.running: ', job.running())
print('job.in_final_state: ', job.in_final_state())
# -
# Job이 너무 오래 걸리거나 회로에 버그가 있을때는 다음과 같이 취소할 수 있습니다.
job = execute(qc, backend, shots = 10e5)
job.cancel()
job.status()
# `Result` 오브젝트는 연산 결과에 대한 정보를 담고 있습니다. `QasmSimulator`의 경우 측정 카운트에 대한 정보만 담고 있지만 백엔드 시뮬레이터의 종류에 따라 보다 자세한 정보를 포함하고 있을때도 있습니다.
# Bell 상태를 만드는 양자회로를 생각해봅시다. (측정 X)
qc = QuantumCircuit(2)
qc.h(0)
qc.cx(0,1)
qc.draw()
# +
# vector의 density matrix를 확인하고 싶을 때 사용하는 visualization 툴입니다.
from qiskit.visualization import plot_state_city
# statevector simulator를 사용하면 `get_statevector` 함수를 이용할 수 있습니다.
backend = Aer.get_backend('statevector_simulator')
job = execute(qc, backend)
result = job.result()
vector = result.get_statevector()
plot_state_city(vector)
# +
# unitary simulator를 이용하면 `get_unitary` 함수를 사용할 수 있습니다.
backend = Aer.get_backend('unitary_simulator')
job = execute(qc, backend)
result = job.result()
unitary = result.get_unitary()
print(unitary)
# -
plot_state_city(unitary)
# 직접 스케쥴을 만들어서 실행을 하고 싶을 경우 `backend.run()` 함수를 이용할 수 있습니다. 백엔드가 `BaseBackend`일 경우 `assemble` 함수를 이용해서 양자회로를 한번 변환하는 절차가 필요합니다.
# Aer 기본 시뮬레이터들은 BaseBackend class에 해당합니다.
from qiskit.providers import Backend, BaseBackend
backend = Aer.get_backend('qasm_simulator')
print('Backend?: ', isinstance(backend, Backend))
print('BaseBackend?: ', isinstance(backend, BaseBackend))
# +
# 트랜스파일, 어샘블을 직접 해주어야합니다.
from qiskit import transpile, assemble
qc = ghz_circuit(3)
t_qc = transpile(qc)
qobj = assemble(t_qc)
# 회로를 backend에서 수행합니다.
job = backend.run(qobj)
# -
# 결과 출력 방법은 동일합니다.
result = job.result()
counts = result.get_counts()
plot_histogram(counts)
# ### IonQ Backend
# IonQ backend를 사용하려면 인증 토큰이 필요합니다. backend를 불러온 뒤에는 위에서 본 시뮬레이터와 비슷하게 사용하면 됩니다.
# +
from qiskit_ionq_provider import IonQProvider
# Provider를 불러옵니다.
provider = IonQProvider(token=token_ionq)
# -
provider.backends()
# IonQ backend는 시뮬레이터 백엔드 ('ionq_simulator')와 QPU 하드웨어 백엔드 ('ionq_qpu') 두 가지가 제공됩니다. Qiskit Aer 시뮬레이터와 다르게 ionq_simulator는 ionq 서버에서 연산을 수행하고 결과를 네트워크로 돌려줍니다.
backend = provider.get_backend('ionq_simulator')
# +
qc = ghz_circuit(3)
job = backend.run(qc, shots=1000)
job.wait_for_final_state() # job이 끝날때까지 기다립니다.
result = job.result()
plot_histogram(result.get_counts())
# -
backend = provider.get_backend('ionq_qpu')
# +
qc = ghz_circuit(3)
job = backend.run(qc, shots=1000)
job.wait_for_final_state() # job이 끝날때까지 기다립니다.
result = job.result()
plot_histogram(result.get_counts())
# -
# ## 양자회로의 Visualization
# Qiskit은 양자회로의 다양한 시각화 툴을 제공합니다.
#
# 앞서 본 `plot_histogram`, `plot_state_city` 외에 어떤 시각화 툴이 있는지 알아보겠습니다.
# +
# Bloch vector의 시각화입니다.
from qiskit.visualization import plot_bloch_vector
vec = [np.pi/2, 0, 1] # theta, pi, radius
plot_bloch_vector(vec)
# +
# 양자회로의 vector 상태를 이용하여 bloch vector를 그리려면 `plot_bloch_multivector`를 사용합니다.
from qiskit.visualization import plot_bloch_multivector
backend = Aer.get_backend('statevector_simulator')
qc = QuantumCircuit(1)
qc.h(0)
job = execute(qc, backend)
vec = job.result().get_statevector()
plot_bloch_multivector(vec)
# +
# Qsphere도 유용하게 사용됩니다.
from qiskit.visualization import plot_state_qsphere
plot_state_qsphere(vec)
# +
## Qsphere는 multi-qubit entangled state에서 강점이 있습니다.
qc = QuantumCircuit(2)
qc.h(0)
qc.cx(0,1)
# 다음 함수도 양자회로에서 State vector를 얻어내는데에 사용가능합니다.
from qiskit.quantum_info import Statevector
vec = Statevector.from_instruction(qc)
# 두 개 이상의 큐비트 상태는 Bloch vector로 표현하면 그 의미가 모호합니다.
plot_bloch_multivector(vec)
# -
# QSphere는 보다 직관적인 이해를 도와줍니다.
plot_state_qsphere(vec)
# ## Cat state (GHZ state) 만들어보기
# 위에서도 살펴보았고, 지난번 Sonika 박사님과도 실습해보았던 고양이 상태 또는 GHZ 상태를 만들어 봅시다.
#
# Hadamard 게이트는 $|0\rangle$ 상태를 $(|0\rangle + |1\rangle)/\sqrt{2}$ 의 중첩상태로 바꿔줍니다. 이 Hadamard gate와 CNOT 게이트들을 활용해서 임의의 $n$ 큐비트에 대해서 $(|00...0\rangle + |11....1\rangle)/\sqrt{2}$ 상태를 만들수 있습니다. 이런 상태를 GHZ (Greenberger–Horne–Zeilinger) 상태 또는 거시적 양자 중첩상태이기 때문에 Cat 상태라고도 합니다.
# +
# GHZ 상태를 만드는 양자회로(측정 x)를 반환하는 함수를 미리 정의하겠습니다.
def ghz_circuit_wo_measure(n: int):
assert isinstance(n, int) and n > 1
qc = QuantumCircuit(n)
qc.h(0)
qc.cx(0, range(1,n))
return qc
# 큐비트 갯수를 정합니다.
qubit_number = 5
qc = ghz_circuit_wo_measure(qubit_number)
qc.draw('mpl')
# -
vec = Statevector.from_instruction(qc)
plot_state_qsphere(vec)
backend = provider.get_backend('ionq_simulator')
# +
# 큐비트 숫자를 바꿔가면서 회로를 만들어서 실제 하드웨어에서 확인해 볼 수 있습니다.
qubit_number = 1
qc = ghz_circuit(qubit_number)
job = backend.run(qc, shots=1000)
job.wait_until_final_state()
result = job.result()
plot_histogram(result.get_counts(), title = f'qubit number = {qubit_number:d}')
# +
result = job.result()
plot_histogram(result.get_counts(), title = f'qubit number = {qubit_number:d}')
# -
| 20210329_Qiskit_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Pandas - Data Analysis Library for Python - Part 1 Basics
#
# Original by <NAME>
#
# Modified by <NAME> (<EMAIL>)
#
# `Pandas` is one of the most important libraries available for data analysts. It is extremely valuable when processing time-series output data
# %matplotlib inline
from IPython.core.display import Image
Image(filename='pandaslogo.jpg')
# ## Pandas video tutorial
# <NAME> created the Pandas library and this notebook was used in a live tutorial which can be found on youtube:
from IPython.display import YouTubeVideo
YouTubeVideo("w26x-z-BdWQ")
from IPython.core.display import Image
Image(filename='pandasbook.jpg')
# The first step in any analysis is to load the necessary libraries -- in this case `pandas` and `numpy`
# +
from pandas import *
import pandas
import numpy as np
# plt.rc('figure', figsize=(10, 6))
# pandas.set_option('notebook_repr_html',False)
# -
# Pandas Series Object: 1 dimensional data container
# ======
#
# This object is a data container for vectors -- incorporating an index and string search functions
s = Series(np.random.randn(5))
s
labels = ['a', 'b', 'c', 'd', 'e']
s = Series(np.random.randn(5), index = labels)
s
'b' in s
s['b']
s
mapping = s.to_dict()
mapping
s = Series(mapping)
s
s[:3]
s.index
# DataFrame: 2D collection of Series
# ==================================
df = DataFrame({'a': np.random.randn(6),
'b': ['foo', 'bar'] * 3,
'c': np.random.randn(6)})
df.info()
df.index
df
df = DataFrame({'a': np.random.randn(6),
'b': ['foo', 'bar'] * 3,
'c': np.random.randn(6)},
index = date_range('1/1/2000', periods=6))
df
df = DataFrame({'a': np.random.randn(6),
'b': ['foo', 'bar'] * 3,
'c': np.random.randn(6)},
columns=['a', 'b', 'c', 'd'])
df
# Creation from nested dicts
# --------------------------
#
# These arise naturally in Python code
data = {}
for col in ['foo', 'bar', 'baz']:
for row in ['a', 'b', 'c', 'd']:
data.setdefault(col, {})[row] = np.random.randn()
data
DataFrame(data)
# Data alignment
# ==============
close_px = read_csv('stock_data.csv', index_col=0, parse_dates=True)
close_px
s1 = close_px['AAPL'][-20:]
s2 = close_px['AAPL'][-25:-10]
s1
s2
s1 + s2
df = close_px.iloc[-10:, :3]
df
b, c = s1.align(s2, join='inner')
b
c
b, c = s1.align(s2, join='outer')
b
b, c = s1.align(s2, join='right')
df = close_px.ix[-10:, ['AAPL', 'IBM', 'MSFT']]
df
df2 = df.ix[::2, ['IBM', 'MSFT']]
df2
df + df2
b, c = df.align(df2, join='inner')
# ## Truncation - clipping a datetime indexed object
df
df.truncate(before='2011-10-05')
df.truncate(before='2011-10-05',after='2011-10-12')
# ## Resampling - Useful time series aggregation
df
df.resample('M').mean()
df.resample('5D').mean()
df.resample('5D').max()
# ## Missing Data - Filling in the Gaps
dfgaps = df.resample('D').mean()
dfgaps
dfgaps.dropna()
dfgaps.fillna(method = 'bfill')
# Transposing: no copy if all columns are same type
# -------------------------------------------------
df[:5].T
# Columns can be any type
# -----------------------
n = 10
foo = DataFrame(index=range(n))
foo['floats'] = np.random.randn(n)
foo['ints'] = np.arange(n)
foo['strings'] = ['foo', 'bar'] * (n / 2)
foo['bools'] = foo['floats'] > 0
foo['objects'] = date_range('1/1/2000', periods=n)
foo
foo.dtypes
# N.B. transposing is not roundtrippable in this case (column-oriented data structure)
foo.T.T
foo.T.T.dtypes
# ## Function application
#
# You can apply arbitrary functions to the rows or columns of a DataFrame
df.apply(np.mean)
df.apply(np.mean, axis=1)
# You can get as fancy as you want
close_px
df.apply(lambda x: x.max() - x.min()) # np.ptp
np.log(close_px)
# ## Plotting
#
# Some basic plotting integration with matplotlib in Series / DataFrame
close_px[['AAPL', 'IBM', 'MSFT', 'XOM']].plot();
# Hierarchical indexing
# ---------------------
index = MultiIndex(levels=[['foo', 'bar', 'baz', 'qux'],
['one', 'two', 'three']],
labels=[[0, 0, 0, 1, 1, 2, 2, 3, 3, 3],
[0, 1, 2, 0, 1, 1, 2, 0, 1, 2]])
hdf = DataFrame(np.random.randn(10, 3), index=index,
columns=['A', 'B', 'C'])
hdf
hdf.loc['foo']
hdf.loc['foo'] = 0
hdf
hdf.loc['foo', 'three']
# Stacking and unstacking
# -----------------------
tuples = zip(*[['bar', 'bar', 'baz', 'baz',
'foo', 'foo', 'qux', 'qux'],
['one', 'two', 'one', 'two',
'one', 'two', 'one', 'two']])
index = MultiIndex.from_tuples(tuples)
columns = MultiIndex.from_tuples([('A', 'cat'), ('B', 'dog'),
('B', 'cat'), ('A', 'dog')])
df = DataFrame(np.random.randn(8, 4), index=index, columns=columns)
df
df2 = df.iloc[[0, 1, 2, 4, 5, 7]]
df2
df.unstack()['B']
# ## GroupBy
#
df = DataFrame({'A' : ['foo', 'bar', 'foo', 'bar',
'foo', 'bar', 'foo', 'foo'],
'B' : ['one', 'one', 'two', 'three',
'two', 'two', 'one', 'three'],
'C' : np.random.randn(8),
'D' : np.random.randn(8)})
df
for key, group in df.groupby('A'):
print key
print group
df.groupby('A')['C'].describe().T
df.groupby('A').mean()
for key, group in df.groupby('A'):
print key
print group
df.groupby(['A', 'B']).mean()
df.groupby(['A', 'B'], as_index=False).mean()
# GroupBy with hierarchical indexing
# ----------------------------------
tuples = zip(*[['bar', 'bar', 'baz', 'baz',
'foo', 'foo', 'qux', 'qux'],
['one', 'two', 'one', 'two',
'one', 'two', 'one', 'two']])
index = MultiIndex.from_tuples(tuples)
columns = MultiIndex.from_tuples([('A', 'cat'), ('B', 'dog'),
('B', 'cat'), ('A', 'dog')])
df = DataFrame(np.random.randn(8, 4), index=index, columns=columns)
df
df.groupby(level=0, axis=0).mean()
df.stack()
df.stack().mean(1).unstack()
# could also have done
df.groupby(level=1, axis=1).mean()
| 1_PandasTutorial/Pandas_DataAnalysisLibrary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="-tk5ROC6mw0f"
# # Text Summarization of Amazon reviews
#
# This notebook implements the seq2seq model for text summerizer
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="B_ULQF2smw0h"
import os
import pandas as pd
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
from collections import Counter
import Summarizer
import summarizer_data_utils
import summarizer_model_utils
# -
print(tf.__version__)
# + [markdown] colab_type="text" id="-XBIbvs4mw0m"
# ## The data
#
#
# The data we will be using with is a dataset from Kaggle, the Amazon Fine Food Reviews dataset.
# It contains, as the name suggests, 570.000 reviews of fine foods from Amazon and summaries of those reviews.
# Our aim is to input a review (Text column) and automatically create a summary (Summary colum) for it.
#
#
# https://www.kaggle.com/snap/amazon-fine-food-reviews/data
# + [markdown] colab_type="text" id="8NYVncFKmw0n"
# ### Reading and exploring
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 4894, "status": "ok", "timestamp": 1526227108183, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="2OiSxpApmw0o" outputId="98a255ad-248e-4f1e-b43f-1c8d384fd453"
# load csv file using pandas.
file_path = './Reviews.csv'
data = pd.read_csv(file_path)
data.shape
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="rElWMbT2mw0t" outputId="08f4b9ee-8c78-4f3b-c986-f7f3dd2ff248"
# we will only use the last two columns Summary (target) and Text (input).
data.head()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 1055, "status": "ok", "timestamp": 1526227113630, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="N9bHztjpmw0x" outputId="aa4ce93d-efde-4ebb-bc3e-7b03d477a5e4"
# check for missings --> got some in summary drop those.
# 26 are missing, so we will drop those!
data.isnull().sum()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="boMCgsgTmw00"
# drop row, if values in Summary is missing.
data.dropna(subset=['Summary'],inplace = True)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 734, "status": "ok", "timestamp": 1526227125421, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="ESv4XLgQmw03" outputId="0ca5d3f7-7efc-46dc-bf8e-bb5803c6376c"
# only summary and text are useful for us.
data = data[['Summary', 'Text']]
data.head()
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="BjmIGbXtmw08"
# we will not use all of them, only short ones and ones of similar size.
# choosing the ones that are of similar length makes it easier for the model to learn.
raw_texts = []
raw_summaries = []
for text, summary in zip(data.Text, data.Summary):
if 100< len(text) < 150:
raw_texts.append(text)
raw_summaries.append(summary)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1560, "status": "ok", "timestamp": 1526227148045, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="t5JBoyqKmw0_" outputId="150af52c-a0af-4eec-f0ff-399ec33e434d"
len(raw_texts), len(raw_summaries)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="tMsDeec4mw1F" outputId="de46147e-d2c4-40e5-9a0a-22f27ac360fd"
for t, s in zip(raw_texts[:5], raw_summaries[:5]):
print('Text:\n', t)
print('Summary:\n', s, '\n\n')
# -
import nltk
nltk.download('punkt')
# + [markdown] colab_type="text" id="Y-CyKX1gmw1J"
# ### Clean and prepare the data
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 47800, "status": "ok", "timestamp": 1526227313932, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="4CLoTqyzmw1K" outputId="2fcb8327-e714-48e2-fca1-6bf39b8e6411"
# the function gives us the option to keep_most of the characters inisde the texts and summaries, meaning
# punctuation, question marks, slashes...
# or we can set it to False, meaning we only want to keep letters and numbers like here.
processed_texts, processed_summaries, words_counted = summarizer_data_utils.preprocess_texts_and_summaries(
raw_texts,
raw_summaries,
keep_most=False
)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="yquphHYJmw1R" outputId="cd9ad917-33da-4294-eb41-f3b0abb967e9"
for t,s in zip(processed_texts[:5], processed_summaries[:5]):
print('Text\n:', t, '\n')
print('Summary:\n', s, '\n\n\n')
# + [markdown] colab_type="text" id="34eUnqVQmw1c"
# ### Create lookup dicts
#
# We cannot feed our network actual words, but numbers. So we first have to create our lookup dicts, where each words gets and int value (high or low, depending on its frequency in our corpus). Those help us to later convert the texts into numbers.
#
# We also add special tokens. EndOfSentence and StartOfSentence are crucial for the Seq2Seq model we later use.
# Pad token, because all summaries and texts in a batch need to have the same length, pad token helps us do that.
#
# So we need 2 lookup dicts:
# - From word to index
# - from index to word.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 716, "status": "ok", "timestamp": 1526227336251, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="zwqbTP8jmw1d" outputId="3788ccc5-bec0-4d32-8885-0698b45d7164"
specials = ["<EOS>", "<SOS>","<PAD>","<UNK>"]
word2ind, ind2word, missing_words = summarizer_data_utils.create_word_inds_dicts(words_counted,
specials = specials)
print(len(word2ind), len(ind2word), len(missing_words))
# + [markdown] colab_type="text" id="EVZ1Qmk9mw1j"
# ### Pretrained embeddings
#
# Optionally we can use pretrained word embeddings. Those have proved to increase training speed and accuracy.
# Here I used two different options. Either we use glove embeddings or embeddings from tf_hub.
# The ones from tf_hub worked better, so we use those.
# -
glove_embeddings_path = './glove.6B.300d.txt'
embedding_matrix_save_path = './embeddings/my_embedding_github.npy'
emb = summarizer_data_utils.create_and_save_embedding_matrix(word2ind, glove_embeddings_path, embedding_matrix_save_path)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 105} colab_type="code" executionInfo={"elapsed": 61662, "status": "ok", "timestamp": 1526227413054, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="ObE6ggfAmw1o" outputId="e12b1f22-0fad-4a3d-e934-5fc480b83788"
# the embeddings from tf_hub.
#embed = hub.load("https://tfhub.dev/google/nnlm-en-dim128/2")
#embed = hub.load("https://tfhub.dev/google/Wiki-words-250/2")
#emb = embed([key for key in word2ind.keys()])
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
sess.run(tf.tables_initializer())
embedding = sess.run(emb)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 687, "status": "ok", "timestamp": 1526227413774, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="ayXi9D7Umw1u" outputId="7b5b5522-8c21-4c70-a5be-46f6ed2cdbd2"
embedding.shape
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="QoGa9EWdmw11"
np.save('./tf_hub_embedding.npy', embedding)
# + [markdown] colab_type="text" id="QV1HB3zzmw12"
# ### Convert text and summaries
#
# As I said before we cannot feed the words directly to our network, we have to convert them to numbers first of all. This is what we do here. And we also append the SOS and EOS tokens.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="NjudfxFPmw13"
# converts words in texts and summaries to indices
# it looks like we have to set eos here to False
converted_texts, unknown_words_in_texts = summarizer_data_utils.convert_to_inds(processed_texts,
word2ind,
eos = False)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="1dFsLoAqmw16"
converted_summaries, unknown_words_in_summaries = summarizer_data_utils.convert_to_inds(processed_summaries,
word2ind,
eos = True,
sos = True)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 476} colab_type="code" executionInfo={"elapsed": 2143, "status": "ok", "timestamp": 1526227545460, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="ghATcyE4mw2A" outputId="06d3f934-7c97-49e2-c358-21bd7552689d"
converted_texts[0]
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 54} colab_type="code" executionInfo={"elapsed": 1124, "status": "ok", "timestamp": 1526227550694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="pSTzMURHmw2E" outputId="4dc8b405-f811-49f2-b7b3-83495dbf7640"
# seems to have worked well.
print( summarizer_data_utils.convert_inds_to_text(converted_texts[0], ind2word),
summarizer_data_utils.convert_inds_to_text(converted_summaries[0], ind2word))
# + [markdown] colab_type="text" id="a8b9Nd0zmw2H"
# ## The model
#
# Now we can build and train our model. First we define the hyperparameters we want to use. Then we create our Summarizer and call the function .build_graph(), which as the name suggests, builds the computation graph.
# Then we can train the model using .train()
#
# After training we can try our model using .infer()
# + [markdown] colab_type="text" id="L2z9xOKzmw2I"
# ### Training
#
# We can optionally use a cyclic learning rate, which we do here.
# I trained the model for 20 epochs and the loss was low then, but we could train it longer and would probably get better results.
#
# Unfortunately I do not have the resources to find the perfect (or right) hyperparameters, but these do pretty well.
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="tEItjpP4mw2J"
# model hyperparametes
num_layers_encoder = 4
num_layers_decoder = 4
rnn_size_encoder = 512
rnn_size_decoder = 512
batch_size = 256
epochs = 200
clip = 5
keep_probability = 0.5
learning_rate = 0.0005
max_lr=0.005
learning_rate_decay_steps = 700
learning_rate_decay = 0.90
pretrained_embeddings_path = './tf_hub_embedding.npy'
summary_dir = os.path.join('./tensorboard', str('Nn_' + str(rnn_size_encoder) + '_Lr_' + str(learning_rate)))
use_cyclic_lr = True
inference_targets=True
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1464, "status": "ok", "timestamp": 1526234914336, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="u8lJ_OI5mw2Q" outputId="1c06bc51-01eb-4a68-b4ca-38d56a4a2a76"
len(converted_summaries)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1026, "status": "ok", "timestamp": 1526234915582, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="w_VDuiHyQK84" outputId="9bc17a2e-837b-41bd-a40d-0f116e143d8f"
round(78862*0.9)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85881} colab_type="code" executionInfo={"elapsed": 8531236, "status": "error", "timestamp": 1526243447242, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="E0BX6Z7Kmw2T" outputId="e734411e-2fbc-4960-e2aa-fb98108c4576"
# build graph and train the model
summarizer_model_utils.reset_graph()
summarizer = Summarizer.Summarizer(word2ind,
ind2word,
save_path='./models/amazon/my_model',
mode='TRAIN',
num_layers_encoder = num_layers_encoder,
num_layers_decoder = num_layers_decoder,
rnn_size_encoder = rnn_size_encoder,
rnn_size_decoder = rnn_size_decoder,
batch_size = batch_size,
clip = clip,
keep_probability = keep_probability,
learning_rate = learning_rate,
max_lr=max_lr,
learning_rate_decay_steps = learning_rate_decay_steps,
learning_rate_decay = learning_rate_decay,
epochs = epochs,
pretrained_embeddings_path = pretrained_embeddings_path,
use_cyclic_lr = use_cyclic_lr,
summary_dir = summary_dir)
summarizer.build_graph()
summarizer.train(converted_texts[:70976],
converted_summaries[:70976],
validation_inputs=converted_texts[70976:],
validation_targets=converted_summaries[70976:])
# hidden training output.
# both train and validation loss decrease nicely.
# + [markdown] colab_type="text" id="U5Hqzvocmw2W"
# ### Inference
# Now we can use our trained model to create summaries.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 85} colab_type="code" executionInfo={"elapsed": 4607, "status": "ok", "timestamp": 1526243454761, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="ljN9a1hemw2Y" outputId="f60102af-44f0-4c45-8ba3-2548e5af0a4c"
summarizer_model_utils.reset_graph()
summarizer = Summarizer.Summarizer(word2ind,
ind2word,
'./models/amazon/my_model',
'INFER',
num_layers_encoder = num_layers_encoder,
num_layers_decoder = num_layers_decoder,
batch_size = len(converted_texts[:50]),
clip = clip,
keep_probability = 1.0,
learning_rate = 0.0,
beam_width = 5,
rnn_size_encoder = rnn_size_encoder,
rnn_size_decoder = rnn_size_decoder,
inference_targets = True,
pretrained_embeddings_path = pretrained_embeddings_path)
summarizer.build_graph()
preds = summarizer.infer(converted_texts[:50],
restore_path = './models/amazon/my_model',
targets = converted_summaries[:50])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 11917} colab_type="code" executionInfo={"elapsed": 833, "status": "ok", "timestamp": 1526243456128, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "102636220151368904258"}, "user_tz": -120} id="JtB2kNIWmw2j" outputId="b2b34d18-062a-4ea0-e48f-29ed6c3fe123"
# show results
summarizer_model_utils.sample_results(preds,
ind2word,
word2ind,
converted_summaries[:50],
converted_texts[:50])
# + [markdown] colab_type="text" id="j_bcG5CPmw2m"
# # Conclusion
#
# Generally I am really impressed by how well the model works.
# We only used a limited amount of data, trained it for a limited amount of time and used nearly random hyperparameters and it still delivers good results.
#
# However, we are clearly overfitting the training data and the model does not perfectly generalize.
# Sometimes the summaries the model creates are good, sometimes bad, sometimes they are better than the original ones and sometimes they are just really funny.
#
#
# Therefore it would be really interesting to scale it up and see how it performs.
#
# To sum up, I am impressed by seq2seq models, they perform great on many different tasks and I look foward to exploring more possible applications.
# (speech recognition...)
| summarizer_amazon_reviews.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# from PyQuantum.JC.Cavity import Cavity
# cv = Cavity(wc=-1, wa=1, g=1, n_atoms=1)
print(1/0)
# print(__name__)
# cv.info()
# import pandas as pd
# data = [[str(cv.wc) + ' Hz'], [str(cv.wa) + ' Hz'], [str(cv.g) + ' Hz'], [], [cv.n_atoms], [cv.n_levels]]
# pd.DataFrame(data, columns=None, index=['wc','wa', 'g', '', 'n_atoms', 'n_levels'])
# data2 = [[cv.n_atoms], [cv.n_levels]]
# pd.DataFrame(data2, columns=None, index=['n_atoms', 'n_levels'])
# -
| .ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %config InlineBackend.figure_formats = ['svg']
# %matplotlib inline
# Alternative plotting backend for interative data exploration
# # %matplotlib notebook
from coronavirus import overview, fetch_deaths, fetch_cases
# If you want to edit the source in the notebook, try "%load coronavirus.py"
# and comment out the import statement above.
import ipywidgets as widgets
from IPython.display import HTML
import numpy as np
import matplotlib.pyplot as plt
# -
EXPLANATION = """\
<div class="app-sidebar">
<p><em>Coronavirus Numbers per Country.</em><p>
<p>Select which country to plot the coronavirus numbers for.</p>
<p>Code used from <a href="https://github.com/fangohr/coronavirus-2020/blob/master/index.ipynb">
Hans Fangohr's coronavirus-2020 repository</a>.</p>
</div>
"""
HTML("""\
<style>
.app-subtitle {
font-size: 1.5em;
}
.app-subtitle a {
color: #106ba3;
}
.app-subtitle a:hover {
text-decoration: underline;
}
.app-sidebar p {
margin-bottom: 1em;
line-height: 1.7;
}
.app-sidebar a {
color: #106ba3;
}
.app-sidebar a:hover {
text-decoration: underline;
}
</style>
""")
class App:
def __init__(self):
self._deaths = fetch_deaths()
self._cases = fetch_cases()
available_countries = np.unique(self._cases.index.values)
self._country_dropdown = self._create_dropdown(available_countries, np.argwhere(available_countries =="Germany")[0][0])
self._plot_container = widgets.Output()
_app_container = widgets.VBox([
widgets.HBox([self._country_dropdown]),
self._plot_container
], layout=widgets.Layout(align_items='center', flex='2 0 auto'))
self.container = widgets.VBox([
widgets.HTML(
(
'<h1>Coronavirus Country Status</h1>'
# '<h2 class="app-subtitle"><a href="https://github.com/pbugnion/voila-gallery/blob/master/country-indicators/index.ipynb">Link to code</a></h2>'
),
layout=widgets.Layout(margin='0 0 5em 0')
),
widgets.HBox([
_app_container,
widgets.HTML(EXPLANATION, layout=widgets.Layout(margin='0 0 0 2em'))
])
], layout=widgets.Layout(flex='1 1 auto', margin='0 auto 0 auto', max_width='1024px'))
self._update_app()
def _create_dropdown(self, indicators, initial_index):
dropdown = widgets.Dropdown(options=indicators, value=indicators[initial_index])
dropdown.observe(self._on_change, names=['value'])
return dropdown
def _create_plot(self, country):
self._overview(country)
def _on_change(self, _):
self._update_app()
def _overview(self, country):
return overview(country)
def _update_app(self):
country = self._country_dropdown.value
self._plot_container.clear_output(wait=True)
with self._plot_container:
self._create_plot(country)
plt.show()
# +
app = App()
app.container
# -
| notebooks/dashboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: fe_test
# language: python
# name: fe_test
# ---
# ## Complete Case Analysis
#
#
# Complete-case analysis (CCA), also called "list-wise deletion" of cases, consists in **discarding** observations where values in **any** of the variables are missing. Complete Case Analysis means literally analysing only those observations for which there is information in **all** of the variables in the dataset.
#
# ### Which variables can I impute with CCA?
#
# CCA can be applied to both categorical and numerical variables.
#
#
# ### Assumptions
#
# CCA works well when the data are missing completely at random (MCAR). In fact, we should use CCA if we have reasons to believe that data is missing at random, and not otherwise. When data is MCAR, excluding observations with missing information is in essence the same as randomly excluding some observations from the dataset. Therefore the dataset after CCA is a fair representation of the original dataset.
#
#
# ### Advantages
#
# - Easy to implement
# - No data manipulation required
# - Preserves variable distribution (if data is MCAR, then the distribution of the variables of the reduced dataset should match the distribution in the original dataset)
#
# ### Disadvantages
#
# - It can exclude a large fraction of the original dataset (if missing data is abundant)
# - Excluded observations could be informative for the analysis (if data is not missing at random)
# - CCA will create a biased dataset if the complete cases differ from the original data (e.g., when missing information is in fact MAR or NMAR and not missing at random).
# - When using our models in production, the model will not know how to handle missing data
#
# ### When to use CCA
#
# - Data is missing completely at random
# - No more than 5% of the total dataset contains missing data
#
# In practice, CCA may be an acceptable method when the amount of missing information is small. Unfortunately, there is no rule of thumb to determine how much missing data is small or negligible. However, as general guidance, if the total amount of missing data is ~5% of the original dataset or less, CCA is a viable option.
#
# In many real life datasets, the amount of missing data is never small, and therefore CCA is typically never an option.
#
# ### CCA and models in production
#
# When using CCA, we remove all observations that contain missing information. However, the data that we want to score with our model, may indeed contain missing information. This will pose a problem when using our model in live systems, or as we call it, when putting or models into production: when an observation contains missing data, the model will not be able to handle it.
#
# In order to avoid this problem, when putting models into production we need to do 1 of 2 things: either we do not score observations with missing data, or we replace the missing values by another number. We can choose any from the imputation techniques that we will discuss in the following lectures to replace NA in the data to be scored.
#
# ## In this demo:
#
# We will use the House Prices dataset to demonstrate how to perform Complete Case Analysis.
#
# - For instructions on how to download the dataset, please refer to the lecture **Datasets** in **Section 1** of the course.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# to show all the columns of the dataframe in the notebook
pd.set_option('display.max_columns', None)
# +
# let's load the House Prices dataset
# and explore its shape (rows and columns)
data = pd.read_csv('../houseprice.csv')
data.shape
# -
# let's visualise the dataset
data.head()
# +
# find the variables with missing observations
vars_with_na = [var for var in data.columns if data[var].isnull().mean() > 0]
vars_with_na
# -
# let's find out whether they are numerical or categorical
data[vars_with_na].dtypes
# There are both numerical and categorical variables with missing observations. We can see from the variable types that some are float and some are object.
# +
# let's have a look at the values of the variables with
# missing data
data[vars_with_na].head(10)
# +
# let's find out the percentage of observations missing per variable
# calculate the percentage of missing (as we did in section 3)
# using the isnull() and mean() methods from pandas
data_na = data[vars_with_na].isnull().mean()
# transform the array into a dataframe
data_na = pd.DataFrame(data_na.reset_index())
# add column names to the dataframe
data_na.columns = ['variable', 'na_percentage']
# order the dataframe according to percentage of na per variable
data_na.sort_values(by='na_percentage', ascending=False, inplace=True)
# show
data_na
# -
# The first 6 variables contain a lot of missing information. So we can't use CCA if we consider those variables, as most of the observations in the dataset will be discarded. We could otherwise use CCA if we omit using those variables with a lot of NA.
#
# For this demo, I will ignore the first 6 variables with a lot of missing data, and proceed with CCA in the remaining of the dataset.
# +
# capture variables with no or less than 5% NA
vars_cca = [var for var in data.columns if data[var].isnull().mean() < 0.05]
vars_cca
# +
# calculate percentage of observations with complete
# cases: i.e., with values for all the variables
# the method dropna(), discards the observations that contain
# na in any of the rows / columns
len(data[vars_cca].dropna()) / len(data)
# +
# create the complete case dataset
# in other words, remove observations with na in any variable
data_cca = data[vars_cca].dropna()
data.shape, data_cca.shape
# +
# plot the histograms for all numerival variables in the complete
# case dataset (as we did in section 3)
data_cca.hist(bins=50, density=True, figsize=(12, 12))
plt.show()
# +
## let's check the distribution of a few variables before and after
# cca: histogram
fig = plt.figure()
ax = fig.add_subplot(111)
# original data
data['GrLivArea'].hist(bins=50, ax=ax, density=True, color='red')
# data after cca, the argument alpha makes the color transparent, so we can
# see the overlay of the 2 distributions
data_cca['GrLivArea'].hist(bins=50, ax=ax, color='blue', density=True, alpha=0.8)
# +
## let's check the distribution of a few variables before and after
# cca: density plot
fig = plt.figure()
ax = fig.add_subplot(111)
# original data
data['GrLivArea'].plot.density(color='red')
# data after cca
data_cca['GrLivArea'].plot.density(color='blue')
# +
## let's check the distribution of a few variables before and after
# cca: histogram
fig = plt.figure()
ax = fig.add_subplot(111)
# original data
data['BsmtFinSF1'].hist(bins=50, ax=ax, density=True, color='red')
# data after cca, the argument alpha makes the color transparent, so we can
# see the overlay of the 2 distributions
data_cca['BsmtFinSF1'].hist(bins=50, ax=ax, color='blue', density=True, alpha=0.8)
# +
## let's check the distribution of a few variables before and after
# cca: density plot
fig = plt.figure()
ax = fig.add_subplot(111)
# original data
data['BsmtFinSF1'].plot.density(color='red')
# data after cca
data_cca['BsmtFinSF1'].plot.density(color='blue')
# -
# As we can see from the above plots, the distribution of the selected numerical variables in the original and complete case dataset is very similar, which is what we expect from CCA if data is missing at random and only for a small proportion of the observations.
#
# In the next cells I will explore the distribution of categorical variables. To do so, I will evaluate the percentage of observations that show each of the unique categories, as we did in sections 2 and 3 of the course.
# +
# the following function captures the percentage of observations
# for each category in the original and complete case dataset
# and puts them together in a new dataframe
def categorical_distribution(df, df_cca, variable):
tmp = pd.concat(
[
# percentage of observations per category, original data
df[variable].value_counts() / len(df),
# percentage of observations per category, cca data
df_cca[variable].value_counts() / len(df_cca)
],
axis=1)
# add column names
tmp.columns = ['original', 'cca']
return tmp
# -
# run the function in a categorical variable
categorical_distribution(data, data_cca, 'BsmtQual')
categorical_distribution(data, data_cca, 'MasVnrType')
categorical_distribution(data, data_cca, 'SaleCondition')
# As we can see from the output of the cells above, the distribution of houses in each of the categories, is very similar in the original and complete case dataset, which again, is what is expected if the data is missing completely at random, and the percentage of missing data is small.
| Section-04-Missing-Data-Imputation/04.01-Complete-Case-Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training vs validation loss
#
# [](https://colab.research.google.com/github/parrt/fundamentals-of-deep-learning/blob/main/notebooks/3.train-test-diabetes.ipynb)
#
# By [<NAME>](https://explained.ai).
#
# This notebook explores how to use a validation set to estimate how well a model generalizes from its training data to unknown test vectors. We will see that deep learning models often have so many parameters that we can drive training loss to zero, but unfortunately the validation loss grows as the model overfits. We will also compare how deep learning performs compared to a random forest model as a baseline. Instead of the cars data set, we will use the [diabetes data set](https://scikit-learn.org/stable/datasets/toy_dataset.html#diabetes-dataset) loaded via sklearn.
# ## Support code
# +
import os
import sys
import torch
import copy
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_diabetes
from sklearn.ensemble import RandomForestRegressor
import matplotlib.pyplot as plt
from matplotlib import colors
import colour
# %config InlineBackend.figure_format = 'retina'
import tsensor
# -
def plot_history(history, ax=None, maxy=None, file=None):
if ax is None:
fig, ax = plt.subplots(1,1, figsize=(3.5,3))
ax.set_ylabel("Loss")
ax.set_xlabel("Epochs")
loss = history[:,0]
val_loss = history[:,1]
if maxy:
ax.set_ylim(0,maxy)
else:
ax.set_ylim(0,torch.max(val_loss))
ax.spines['top'].set_visible(False) # turns off the top "spine" completely
ax.spines['right'].set_visible(False)
ax.spines['left'].set_linewidth(.5)
ax.spines['bottom'].set_linewidth(.5)
ax.plot(loss, label='train_loss')
ax.plot(val_loss, label='val_loss')
ax.legend(loc='upper right')
plt.tight_layout()
if file:
plt.savefig(f"/Users/{os.environ['USER']}/Desktop/{file}.pdf")
# ## Load diabetes data set
#
# From [sklearn diabetes data set](https://scikit-learn.org/stable/datasets/toy_dataset.html#diabetes-dataset):
# "<i>Ten baseline variables, age, sex, body mass index, average blood pressure, and six blood serum measurements were obtained for each of n = 442 diabetes patients, as well as the response of interest, a quantitative measure of disease progression one year after baseline.</i>"
#
# So, the goal is to predict disease progression based upon all of these features.
d = load_diabetes()
len(d.data)
df = pd.DataFrame(d.data, columns=d.feature_names)
df['disease'] = d.target # "quantitative measure of disease progression one year after baseline"
df.head(3)
# ## Split data into train, validation sets
#
# Any sufficiently powerful model is able to effectively drive down the training loss (error). What we really care about, though, is how well the model generalizes. That means we have to look at the validation or test error, computed from records the model was not trained on. (We'll use "test" as shorthand for "validation" but technically they are not the same.) For non-time-sensitive data sets, we can simply randomize and hold out 20% as our validation set:
np.random.seed(1) # set a random seed for consistency across runs
n = len(df)
X = df.drop('disease',axis=1).values
y = df['disease'].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20) # hold out 20%
# Let's also make sure to normalize the data to make training easier:
m = np.mean(X_train,axis=0)
std = np.std(X_train,axis=0)
X_train = (X_train-m)/std
X_test = (X_test-m)/std # use training data only when prepping test sets
# ## Baseline with random forest
#
# When building machine learning models, it's always important to ask how good your model is. One of the best ways is to choose a baseline model, such as a random forest or a linear regression model, and compare your new model to make sure it can beat the old model. Random forests are easy to use, understand, and train so they are a good baseline. Don't worry about the details of using sklearn to train the random forest regressor, just assume it is a powerful and efficient model. Training the model is as simple as calling `fit()`:
rf = RandomForestRegressor(n_estimators=500)#, min_samples_leaf=2, max_features=1)
rf.fit(X_train, y_train.reshape(-1))
# To evaluate our models, let's compute the mean squared error (MSE) for both training and validation sets:
# +
y_pred = rf.predict(X_train)
mse = np.mean((y_pred - y_train.reshape(-1))**2)
y_pred_test = rf.predict(X_test)
mse_test = np.mean((y_pred_test - y_test.reshape(-1))**2)
print(f"Training MSE {mse:.2f} validation MSE {mse_test:.2f}")
# -
# #### Exercise
#
# Why is the validation error much larger than the training error?
#
# <details>
# <summary>Solution</summary>
# Because the model was trained on the training set, one would expect it to generally perform better on it than any other data set. The more the validation error diverges from the training error, the less general you should assume your model is.
# </details>
# ## Train network with increasingly sophisticated training method
#
# Ok, so now we have a baseline and an understanding of how well a decent model performs on this data set. Let's see if we can beat that baseline with a neural network. First we will see how easy it is to drive the training error down and then show how the validation error is not usually very good in that case. We will finish by considering ways to get better validation errors, which means more general models.
# ### Most basic network training
#
# A basic training loop for a neural network model simply measures and tracks the training loss or error/metric. (In this case, our loss and metric are the same.) The following function embodies such a training loop:
def train0(model, X_train, X_test, y_train, y_test,
learning_rate = .5, nepochs=2000):
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
for epoch in range(nepochs+1):
y_pred = model(X_train)
loss = torch.mean((y_pred - y_train)**2)
if epoch % (nepochs//10) == 0:
print(f"Epoch {epoch:4d} MSE train loss {loss:12.3f}")
optimizer.zero_grad()
loss.backward() # autograd computes w1.grad, b1.grad, ...
optimizer.step()
# To use this method, we have to convert the training and validation data sets to pytorch tensors from numpy (they are already normalized):
X_train = torch.tensor(X_train).float()
X_test = torch.tensor(X_test).float()
y_train = torch.tensor(y_train).float().reshape(-1,1) # column vector
y_test = torch.tensor(y_test).float().reshape(-1,1)
# Let's create a model with one hidden layer and an output layer, glued together with a ReLU nonlinearity. There is an implied input layer which is really just the input vector of features. The output layer takes the output of the hidden layer and generates a single output, our $\hat{y}$:
# +
ncols = X.shape[1]
n_neurons = 150
model = nn.Sequential(
nn.Linear(ncols, n_neurons), # hidden layer
nn.ReLU(), # nonlinearity
nn.Linear(n_neurons, 1) # output layer
)
train0(model, X_train, X_test, y_train, y_test, learning_rate=.08, nepochs=5000)
# -
# Run this a few times and you'll see that we can drive the training error very close to zero with 150 neurons and many iterations (epochs).
# #### Exercise
#
# Why does the training loss sometimes pop up and then go back down? Why is it not monotonically decreasing?
#
# <details>
# <summary>Solution</summary>
# The only source of randomness is the initialization of the model parameters, but that does not explain the lack of monotonicity. In this situation, it is likely that the learning rate is too high and therefore, as we approach the minimum of the lost function, our steps are too big. We are jumping back and forth across the location of the minimum in parameter space.
# </details>
# #### Exercise
#
# Change the learning rate from 0.08 to 0.001 and rerun the example. What happens to the training loss? Is it better or worse than the baseline random forest and the model trained with learning rate 0.08?
#
# <details>
# <summary>Solution</summary>
# The training loss continues to decrease but much lower than before and stops long before reaching a loss near zero. On the other hand, it is better than the training error from the baseline random forest.
# </details>
# ## Reducing the learning rate to zero in on the minimum
#
# In one of the above exercises we discussed that the learning rate was probably too high in the vicinity of the lost function minimum. There are ways to throttle the learning rate down as we approach the minimum, but we are using a fixed learning rate here. In order to get a smooth, monotonic reduction in loss function let's start with a smaller learning rate, but that means increasing the number of epochs:
# +
ncols = X.shape[1]
n_neurons = 150
model = nn.Sequential(
nn.Linear(ncols, n_neurons), # hidden layer
nn.ReLU(), # nonlinearity
nn.Linear(n_neurons, 1) # output layer
)
train0(model, X_train, X_test, y_train, y_test, learning_rate=.02, nepochs=8000)
# -
# Notice now that we can reliably drive that training error down to zero without bouncing around, although it takes longer with the smaller learning rate.
# ### Tracking validation loss
#
# A low training error doesn't really tell us that much, other than the model is able to capture the relationship between the features and the target variable. What we really want is a general model, which means evaluating the model's performance on a validation set. We have both sets, so let's now track the training and validation error in the loop. We will see that our model performs much worse on the records in the validation set (on which the model was not trained).
def train1(model, X_train, X_test, y_train, y_test,
learning_rate = .5, nepochs=2000):
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
history = [] # track training and validation loss
for epoch in range(nepochs+1):
y_pred = model(X_train)
loss = torch.mean((y_pred - y_train)**2)
y_pred_test = model(X_test)
loss_test = torch.mean((y_pred_test - y_test)**2)
history.append((loss, loss_test))
if epoch % (nepochs//10) == 0:
print(f"Epoch {epoch:4d} MSE train loss {loss:12.3f} test loss {loss_test:12.3f}")
optimizer.zero_grad()
loss.backward() # autograd computes w1.grad, b1.grad, ...
optimizer.step()
return torch.tensor(history)
# Let's create the exact same model that we had before:
# +
ncols = X.shape[1]
n_neurons = 150
model = nn.Sequential(
nn.Linear(ncols, n_neurons),
nn.ReLU(),
nn.Linear(n_neurons, 1)
)
history = train1(model, X_train, X_test, y_train, y_test,
learning_rate=.02, nepochs=8000)
plot_history(torch.clamp(history, 0, 12000), file="train-test")
# -
# Wow. The validation error is much much worse than the training error, which is almost 0. That tells us that the model is severely overfit to the training data and is not general at all. Well, the validation error actually makes a lot of progress initially but then after a few thousand epochs immediately starts to grow (we'll use this fact later). Unless we do something fancier, the best solution can be obtained by selecting the model parameters that gives us the lowest validation loss.
# ### Track best loss and choose best model
#
# We saw in the previous section that the most general model appears fairly soon in the training cycle. So, despite being able to drive the training error to zero if we keep going long enough, the most general model actually is known very early in the training process. This is not always the case, but it certainly is here for this data. Let's exploit this by tracking the best model, the one with the lowest validation error. There some indication that a good approach is to (sometimes crank up the power of the model and then) just stop early, or at least pick the model with the lowest validation error. The following function embodies that by making a copy of our neural net model when it finds an improved version.
def train2(model, X_train, X_test, y_train, y_test,
learning_rate = .5, nepochs=2000, weight_decay=0):
optimizer = torch.optim.Adam(model.parameters(),
lr=learning_rate, weight_decay=weight_decay)
history = [] # track training and validation loss
best_loss = 1e10
best_model = None
for epoch in range(nepochs+1):
y_pred = model(X_train)
loss = torch.mean((y_pred - y_train)**2)
y_pred_test = model(X_test)
loss_test = torch.mean((y_pred_test - y_test)**2)
history.append((loss, loss_test))
if loss_test < best_loss:
best_loss = loss_test
best_model = copy.deepcopy(model)
best_epoch = epoch
if epoch % (nepochs//10) == 0:
print(f"Epoch {epoch:4d} MSE train loss {loss:12.3f} test loss {loss_test:12.3f}")
optimizer.zero_grad()
loss.backward() # autograd computes w1.grad, b1.grad, ...
optimizer.step()
print(f"BEST MSE test loss {best_loss:.3f} at epoch {best_epoch}")
return torch.tensor(history), best_model
# Let's use the exact same model and learning rate with no weight decay and see what happens.
# +
ncols = X.shape[1]
n_neurons = 150
model = nn.Sequential(
nn.Linear(ncols, n_neurons),
nn.ReLU(),
nn.Linear(n_neurons, 1)
)
history, best_model = train2(model, X_train, X_test, y_train, y_test,
learning_rate=.02, nepochs=1000,
weight_decay=0)
# verify we got the best model out
y_pred = best_model(X_test)
loss_test = torch.mean((y_pred - y_test)**2)
plot_history(torch.clamp(history, 0, 12000))
# -
# The best MSE bounces around a loss value of 3000 from run to run, a bit above it or a bit below, depending on the run. And this decent result occurs without having to understand or use weight decay (more on this next). You might find this article interesting: [Why Deep Learning Works Even Though It Shouldn’t](https://moultano.wordpress.com/2020/10/18/why-deep-learning-works-even-though-it-shouldnt/).
# ### Weight decay to reduce overfitting
#
# Other than stopping early, one of the most common ways to reduce model overfitting is to use weight decay, otherwise known as L2 (Ridge) regression, to constrain the model parameters. Without constraints, model parameters can get very large, which typically leads to a lack of generality. Using the `Adam` optimizer, we turn on weight decay with parameter `weight_decay`, but otherwise the training loop is the same:
def train3(model, X_train, X_test, y_train, y_test,
learning_rate = .5, nepochs=2000, weight_decay=0, trace=True):
optimizer = torch.optim.Adam(model.parameters(),
lr=learning_rate, weight_decay=weight_decay)
history = [] # track training and validation loss
for epoch in range(nepochs+1):
y_pred = model(X_train)
loss = torch.mean((y_pred - y_train)**2)
y_pred_test = model(X_test)
loss_test = torch.mean((y_pred_test - y_test)**2)
history.append((loss, loss_test))
if trace and epoch % (nepochs//10) == 0:
print(f"Epoch {epoch:4d} MSE train loss {loss:12.3f} test loss {loss_test:12.3f}")
optimizer.zero_grad()
loss.backward() # autograd computes w1.grad, b1.grad, ...
optimizer.step()
return torch.tensor(history)
# How do we know what the right value of the weight decay is? Typically we try a variety of weight decay values and then see which one gives us the best validation error, so let's do that using a grid of images. The following loop uses the same network and learning rate for each run but varies the weight decay:
# +
ncols = X.shape[1]
n_neurons = 150
fig, axes = plt.subplots(1, 4, figsize=(12.5,2.5))
for wd,ax in zip([0,.3,.6,1.5],axes):
model = nn.Sequential(
nn.Linear(ncols, n_neurons),
nn.ReLU(),
nn.Linear(n_neurons, 1)
)
history = train3(model, X_train, X_test, y_train, y_test,
learning_rate=.05, nepochs=1000, weight_decay=wd,
trace=False)
mse_valid = history[-1][1]
ax.set_title(f"wd={wd:.1f}, valid MSE {mse_valid:.0f}")
plot_history(torch.clamp(history, 0, 10000), ax=ax, maxy=10_000)
plt.tight_layout()
plt.show()
# -
# From this experiment, we can conclude that a weight decay of 1.5 gives the best final mean squared error. But, the experiment is reporting the final MSE all the way on the right side of the graph.
#
# The minimum MSE in the above experiment (of four side-by-side graphs), however, appears before the right edge and the validation error simply gets worse after that. That tells us that we should not pick the parameters simply as the parameters where the training leaves off. We should pick the model parameters that give the minimum loss, as we did before.
# #### Exercise
#
# Set the weight decay to something huge like 100. What do you observe about the training and validation curves?
#
# <details>
# <summary>Solution</summary>
#
# The two curves are flat, and about the same level. The minimum validation error is about 6000 so much worse than with more reasonable weight decay. We have seriously biased the model because we cannot even drive the training error downwards. The bias comes from the extreme constraint we've placed on the model parameters.
#
# <pre>
# model = nn.Sequential(
# nn.Linear(ncols, n_neurons),
# nn.ReLU(),
# nn.Linear(n_neurons, 1)
# )
# history = train2(model, X_train, X_test, y_train, y_test,
# learning_rate=.05, nepochs=1000, weight_decay=100,
# trace=False)
# mse_valid = history[-1][1]
# ax.set_title(f"wd={wd:.1f}, valid MSE {mse_valid:.0f}")
# plot_history(torch.clamp(history, 0, 10000), ax=ax, maxy=10_000)
# </pre>
# </details>
| notebooks/3.train-test-diabetes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Specviz Demonstration Notebook
# This notebook demonstrates the Specviz API in the Notebook setting. UI equivalents for these actions, as well as additional documentation about Specviz, can be found here: https://jdaviz.readthedocs.io/en/latest/specviz/
#
# ## Setup
import astropy.units as u
# +
# Suppress warnings
import warnings
with warnings.catch_warnings():
warnings.simplefilter("ignore")
# -
# ## Create Specviz via Helper
from jdaviz import Specviz
specviz = Specviz()
# ## Display Specviz
specviz.app
# ## Load a file to Specviz
from astropy.utils.data import download_file
fn = download_file('https://data.sdss.org/sas/dr14/sdss/spectro/redux/26/spectra/0751/spec-0751-52251-0160.fits', cache=True)
specviz.load_spectrum(fn, "myfile", format="SDSS-III/IV spec")
# ## Subset Retrieval
# User Task: Select a subset in the viewer above
#
# After defining regions in your spectra, you can retrieve your subsets through different means. To retrieve your subset by name:
# Returns a version of the whole spectra, with a mask applied
specviz.get_spectra('Subset 1')
# Or, if you've defined multiple regions, you can retrieve all defined regions/subsets via:
specviz.app.get_subsets_from_viewer('spectrum-viewer')
# ## Screenshot Saving
specviz.app.get_viewer("spectrum-viewer").figure.save_png()
# ## Panning/Zooming in Specviz
# ### Limit Methods
# You can use the methods x_limits() and y_limits() to modify the field of view of Specviz. You can provide it a scalar (which assumes the units of the loaded spectra), an Astropy Quantity, or 'auto' to automatically scale.
specviz.x_limits()
specviz.x_limits(650*u.nm,750*u.nm)
specviz.y_limits('auto', 110.0)
# ### Disable Scientific Notation
# Scientific notation is used in the axes of the viewers by default. To deactivate it, run the following code:
# axis 1 corresponds to the Y-axis and axis 0 to the X-axis
# fmt can be '0.1e' to set scientific notation or '0.2f' to turn it off
specviz.set_spectrum_tick_format(fmt="0.2f", axis=1)
# ### Autoscale Methods
# You can also quickly return to the default zoom using autoscale_x() and autoscale_y()
specviz.autoscale_x()
specviz.autoscale_y()
# ## Extracting Models
# If models were added using the model fitting plugin, they can be extracted using the following property
specviz.fitted_models
# If the name of a particular model is known, it can be extracted from the fitted_models property
specviz.fitted_models['Model']
# Alternatively, the following getter can be used
models = specviz.get_models()
models
| notebooks/SpecvizExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Example Notebook for the tunneling Fermions
#
# This Notebook is based on the following [paper](https://journals.aps.org/prl/abstract/10.1103/PhysRevLett.114.080402) from the Jochim group. In these experiments two fermions of different spins are put into a single tweezer and then coupled to a second tweezer. The dynamics is then controlled by two competing effects. The interactions and the tunneling.
#
# Let us first start by looking at the data, then look how the can be described in the Hamiltonian language and finally in the gate language.
# +
import pennylane as qml
from pennylane import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# +
data_murmann_no_int = pd.read_csv("Data/Murmann_No_Int.csv", names=["time", "nR"])
data_murmann_with_int = pd.read_csv("Data/Murmann_With_Int.csv", names=["time", "nR"])
# plt.figure(dpi=96)
f, (ax1, ax2) = plt.subplots(2, 1, sharex=True, sharey=True)
ax1.plot(
data_murmann_no_int.time, data_murmann_no_int.nR, "ro", label="U = 0", markersize=4
)
ax2.plot(
data_murmann_with_int.time,
data_murmann_with_int.nR,
"bo",
label="U = J",
markersize=4,
)
ax1.set_ylabel(r"atoms in right valley")
ax2.set_ylabel(r"atoms in right valley")
ax2.set_xlabel(r"time (ms)")
ax1.legend()
ax2.legend()
# -
# ## Analytical prediction
#
# For the two atoms the Hamiltonian can be written down in the basis $\{LL, LR, RL, RR\}$ as:
#
# $$
# H = \left(\begin{array}{cccc}
# U & -J & -J & 0\\
# -J & 0 & 0 &-J\\
# -J & 0 & 0 &-J\\
# 0 & -J & -J & U
# \end{array}
# \right)
# $$
#
# And we start out in the basis state $|LL\rangle$. So we can write
from scipy.sparse.linalg import expm
J = np.pi * 134
# in units of hbar
U = 0.7 * J;
Nt_an = 50
t_analytical = np.linspace(0, 20, Nt_an) * 1e-3
H_With_Int = np.array([[U, -J, -J, 0], [-J, 0, 0, -J], [-J, 0, 0, -J], [0, -J, -J, U]])
H_Wo_Int = np.array([[0, -J, -J, 0], [-J, 0, 0, -J], [-J, 0, 0, -J], [0, -J, -J, 0]])
psi0 = np.zeros(4) * 1j
psi0[0] = 1.0 + 0j
print(psi0)
# +
psis_wo_int = 1j * np.zeros((4, Nt_an))
psis_w_int = 1j * np.zeros((4, Nt_an))
for ii in np.arange(Nt_an):
U_wo = expm(-1j * t_analytical[ii] * H_Wo_Int)
psis_wo_int[:, ii] = np.dot(U_wo, psi0)
U_w = expm(-1j * t_analytical[ii] * H_With_Int)
psis_w_int[:, ii] = np.dot(U_w, psi0)
ps_wo = np.abs(psis_wo_int) ** 2
ps_w = np.abs(psis_w_int) ** 2
# -
nR_wo = ps_wo[1, :] + ps_wo[2, :] + 2 * ps_wo[3, :]
nR_w = ps_w[1, :] + ps_w[2, :] + 2 * ps_w[3, :];
# +
f, (ax1, ax2) = plt.subplots(2, 1, sharex=True, sharey=True)
ax1.plot(t_analytical * 1e3, nR_wo, "r-", label="U = 0", linewidth=4, alpha=0.5)
ax1.plot(
data_murmann_no_int.time, data_murmann_no_int.nR, "ro", label="U = 0", markersize=4
)
ax2.plot(t_analytical * 1e3, nR_w, "b-", label="U = 0", linewidth=4, alpha=0.5)
ax2.plot(
data_murmann_with_int.time,
data_murmann_with_int.nR,
"bo",
label="U = J",
markersize=4,
)
ax1.set_ylabel(r"atoms in right valley")
ax2.set_ylabel(r"atoms in right valley")
ax2.set_xlabel(r"time (ms)")
ax2.set_xlim(0, 20)
ax1.legend()
ax2.legend()
# -
# ## Pennylane
#
# And now we also compare to the pennylane simulation. Make sure that you followed the necessary steps for obtaining the credentials as desribed in the [introduction](https://synqs.github.io/pennylane-ls/intro.html).
from pennylane_ls import *
from credentials import username, password
FermionDevice = qml.device("synqs.fs", shots=500, username=username, password=password)
# In the experiments two Fermions are loaded onto the right side, i.e. into the wire 0 and 1.
# ## No interaction
# In a first set of experiments there are no interactions and the two atoms are simply allowed to hop. The experiment is then described by the following very simple circuit.
@qml.qnode(FermionDevice)
def simple_hopping(theta=0):
"""
The circuit that simulates the experiments.
theta ... angle of the hopping
"""
# load atoms
FermionOps.Load(wires=0)
FermionOps.Load(wires=1)
# let them hop
FermionOps.Hop(theta, wires=[0, 1, 2, 3])
# measure the occupation on the right side
obs = FermionOps.ParticleNumber([2, 3])
return qml.expval(obs)
simple_hopping(0)
print(simple_hopping.draw())
# now let us simulate the time evolution
Ntimes = 15
times = np.linspace(0, 20, Ntimes) * 1e-3
means = np.zeros(Ntimes)
for i in range(Ntimes):
if i % 10 == 0:
print("step", i)
# Calculate the resulting states after each rotation
means[i] = simple_hopping(-2 * J * times[i])
# and compare to the data
f, ax1 = plt.subplots(1, 1, sharex=True, sharey=True)
ax1.plot(times * 1e3, means, "r-", label="U = 0", linewidth=4, alpha=0.5)
ax1.plot(
data_murmann_no_int.time, data_murmann_no_int.nR, "ro", label="U = 0", markersize=4
)
ax1.set_xlim(0, 20)
# ## Hopping with interactions
#
# In a next step the atoms are interacting. The circuit description of the experiment is the application of the hopping gate and the interaction gate. It can be written as
@qml.qnode(FermionDevice)
def correlated_hopping(theta=0, gamma=0, Ntrott=15):
"""
The circuit that simulates the experiments.
theta ... angle of the hopping
gamma ... angle of the interaction
"""
# load atoms
FermionOps.Load(wires=0)
FermionOps.Load(wires=1)
# let them hop
# evolution under the Hamiltonian
for ii in range(Ntrott):
FermionOps.Hop(theta / Ntrott, wires=[0, 1, 2, 3])
FermionOps.Inter(gamma / Ntrott, wires=[0, 1, 2, 3, 4, 5, 6, 7])
# measure the occupation on the right side
obs = FermionOps.ParticleNumber([2, 3])
return qml.expval(obs)
Ntimes = 15
times = np.linspace(0, 20, Ntimes) * 1e-3
means_int = np.zeros(Ntimes)
for i in range(Ntimes):
if i % 10 == 0:
print("step", i)
means_int[i] = correlated_hopping(-2 * J * times[i], U * times[i])
# And we compare to the data to obtain
# +
f, ax2 = plt.subplots(1, 1, sharex=True, sharey=True)
ax2.plot(times * 1e3, means_int, "b-", label="simulation", linewidth=4, alpha=0.5)
ax2.plot(
data_murmann_with_int.time,
data_murmann_with_int.nR,
"bo",
label="U = J",
markersize=4,
)
ax2.set_ylabel(r"atoms in right valley")
ax2.set_xlabel(r"time (ms)")
ax2.legend()
ax2.set_xlim(0, 20)
# -
# ## Summary
#
# And finally we can compare the experimental data with all the descriptions.
# +
f, (ax1, ax2) = plt.subplots(2, 1, sharex=True, sharey=True)
ax1.plot(times * 1e3, means, "r-", label="pennylane", linewidth=4, alpha=0.5)
ax1.plot(t_analytical * 1e3, nR_wo, "r-.", label="analytical", linewidth=4, alpha=0.5)
ax1.plot(
data_murmann_no_int.time,
data_murmann_no_int.nR,
"ro",
label="experiment",
markersize=4,
)
ax2.plot(times * 1e3, means_int, "b-", label="pennylane", linewidth=4, alpha=0.5)
ax2.plot(t_analytical * 1e3, nR_w, "b-.", label="analytical", linewidth=4, alpha=0.5)
ax2.plot(
data_murmann_with_int.time,
data_murmann_with_int.nR,
"bo",
label="experiment",
markersize=4,
)
ax1.set_ylabel(r"atoms in right valley")
ax2.set_ylabel(r"atoms in right valley")
ax2.set_xlabel(r"time (ms)")
ax1.legend(loc="upper right")
ax2.legend(loc="upper right")
ax1.set_xlim(-1, 20)
| examples_before_PR_accept_by_RPB/Fermions_in_double_well.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kyle-gao/DNN_from_scratch/blob/master/MLP_From_Scratch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Y2ibPqjy5nLK"
# Copyright 2021 <NAME>(Kyle) Gao
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 .
# + id="TCWHpmITF0L8"
import numpy as np
import matplotlib.pyplot as plt
from keras.datasets import mnist
# + id="4amYo8XgF3Vw"
(train_X, train_Y), (test_X, test_Y) = mnist.load_data()
train_X = train_X/255.0
test_X = test_X/255.0
# + id="EGoS4qKXMAXN"
def sigmoid(Z):
#sigmoid function
return 1/(1+np.exp(-Z))
def d_sigmoid(Z):
#sigmoid derivative
return sigmoid(Z)*(1-sigmoid(Z))
# + id="sLTtqxxHMnbE"
def relu(Z):
return Z*(Z>0) #(Z>0) is an array of true and false, but np can convert into array of 0 and 1s.
def d_relu(Z):
return 1.0 *(Z>0)
# + id="-0Gaoc2zMrDW"
def softmax(Z,eps=1e-10):
"""Numerically stable softmax
Given (m,n) input, returns softmax over the last dimension"""
shiftZ = Z - np.max(Z,axis=-1,keepdims=True)
expZ=np.exp(shiftZ)
total=np.sum(expZ,axis=-1,keepdims=True)+eps
return expZ/total
def d_softmax(Z):
#Note, this is not the softmax derivative
#instead, we absorb the softmax derivative into the loss function derivative, which simplifies the math.
#so we just pass the identity function.
return 1 #turns out we don't need
# + id="L8raAFy8Om8n"
def one_hot(Y,n_classes):
"""
Returns the one hot encoding of Y shape (n_batch, n_classes)
args:
Y: 1-d array of integers from 0 to n_classes of length n_batch
n_classes: number of classes
"""
m = Y.shape[0]
O_h = np.zeros((m,n_classes))
O_h[range(m),Y] = 1 #first element, at Yth entry = 1 etc...
return O_h
# + id="hIRApgfdYAK4"
def get_minibatch(X,Y,minibatch_size = 64):
"""
Returns a list of (batch_X,batch_Y)
Args:
X -- array of shape (num_samples, img_height, img_width)
Y -- array of shape (num_samples)
minibatch_size -- integer
"""
num_batches = X.shape[0]//minibatch_size
minibatches = []
for b in range(num_batches):
batch_X = X[b*minibatch_size:(b+1)*minibatch_size,:,:]
batch_Y = Y[b*minibatch_size:(b+1)*minibatch_size]
minibatches.append((batch_X,batch_Y))
return minibatches
# + id="g-rNV_PzcJP2"
def categorical_cross_entropy(Y,Yhat, eps=1e-8):
num_samples,dim_y = np.shape(Y)
Yhat = np.clip(Yhat, eps, 1-eps)
J = - np.sum(Y*np.log(Yhat),axis=-1)
return np.mean(J)
def sparse_categorical_cross_entropy(Y,Yhat,n_classes=10, eps=1e-8):
Y_oh = one_hot(Y,n_classes)
return categorical_cross_entropy(Y_oh, Yhat, eps = eps)
# + id="JciU7OPcqL7C"
def sgd(Ws,Bs,dWs,dBs,lr):
L = len(Ws)
Bs=[Bs[i]-lr*dBs[i] for i in range(L)]
Ws=[Ws[i]-lr*dWs[i] for i in range(L)]
return Ws, Bs
# + id="eT-sBz2KQnVb"
class MLP():
def __init__(self,layers):
self.layers = layers
self.num_classes = layers[-1]
self.Ws = None #list of 2D numpy arrays
self.Bs = None #list of 1D numpy arrays
self.As = None #list of 1D numpy arrays
self.Zs = None #list of 1D numpy arrays
def initialize_parameters(self):
layers = self.layers
L=len(layers)
Ws=[]
Bs=[]
for l in range(1,L):
wl=np.random.randn(layers[l-1],layers[l]) *np.sqrt(2/(layers[l-1])) # he initialization
bl=np.zeros((layers[l],1))
Ws.append(wl)
Bs.append(bl)
self.Ws = Ws
self.Bs = Bs
def update_parameters(self,Ws,Bs):
self.Ws = Ws
self.Bs = Bs
def forward_1(self,A_prev,W,b,activation = relu):
#A_prev -- (n_classes,dim_a_prev)
#W -- (dim_a,dim_a_prev)
#b -- (dim_a,1)
#activation -- a function (relu,sigmoid,softmax)
Z = np.dot(A_prev,W) + b.T # this is (n_classes, dim_a) + (1,dim_a)
A = activation(Z)
return A,Z
def forward(self, X, middle_activation = relu, final_activation = softmax,flatten = True):
As = []
Zs = []
Ws = self.Ws
Bs = self.Bs
L = len(Ws)
X = np.reshape(X,(-1,28*28)) #ANN take in vector input, need to flatten image.
As.append(X)
for i in range(L-1):
A,Z = self.forward_1(As[i],Ws[i],Bs[i],middle_activation)
As.append(A)
Zs.append(Z)
#final layer
A,Z = self.forward_1(A,Ws[-1],Bs[-1],final_activation)
As.append(A)
Zs.append(Z)
self.As = As
self.Zs = Zs
return A
def backward_1 (self, dA, W, A_prev, Z, d_activation = d_sigmoid):
m = A_prev.shape[0]
dZ = dA*d_activation(Z)
dA_prev = np.dot(dZ,W.T) #this is the delta in slide 25
dW = np.dot(A_prev.T,dZ)/m #gradient wrt to W, dW is a 2D array with same shape as the W of the same layer averaged over the samples.
dB = np.mean(dZ,axis=0) #gradient wrt to B, dB is a 1D array with same shape as the B of the same layer averaged over the samples.
dB = dB=np.reshape(dB,(len(dB),1)) #reshape so the 1D array is of shape (n_neurons, 1)
return dA_prev, dW, dB
def backward(self, Y, middle_d_activation = d_relu, final_d_activation = d_softmax):
"""
We will assume we are doing a classification task, in which we can absorb the softmax derivative into the Loss function derivative
"""
if len(Y.shape) == 1:
Y = one_hot(Y, self.layers[-1])
dWs = []
dBs = []
L = len(self.Ws)
dA = self.As[-1]-Y
dA,dW,dB = self.backward_1(dA, self.Ws[L-1], self.As[L-1], self.Zs[-1], final_d_activation)
dWs.append(dW)
dBs.append(dB)
for i in range(L-2,-1,-1):
dA_prev, dW, dB=self.backward_1(dA,self.Ws[i],self.As[i],self.Zs[i],middle_d_activation)
dWs.append(dW)
dBs.append(dB)
dA=dA_prev
dWs.reverse()
dBs.reverse()
return dWs,dBs
def predict(self,X):
A = self.forward(X)
return np.argmax(A,axis=-1)
def evaluate(self,X,Y):
predictions = self.predict(X)
acc = np.count_nonzero(predictions==Y)/len(Y)
return acc
# + id="zCo9LBRzYbJG"
mlp = MLP([784,64,10])
mlp.initialize_parameters()
# + id="1vvKKvDdaXc2"
train_batches = get_minibatch(train_X,train_Y,64)
test_batches = get_minibatch(test_X,test_Y,64)
lr = 1e-1
n_epochs = 20 # an epoch is a training over the entire training set once
# + colab={"base_uri": "https://localhost:8080/"} id="KDuqhFdeAOwd" outputId="cf104569-786a-46c6-d372-27280362b5f0"
train_losses = []
test_losses = []
for epoch in range(n_epochs):
train_loss = 0
test_loss = 0
for b in train_batches:
X_b = b[0]
Y_b = b[1]
Y_hat_b = mlp.forward(X_b)
train_loss = train_loss + sparse_categorical_cross_entropy(Y_b,Y_hat_b)
Ws = mlp.Ws
Bs = mlp.Bs
dWs,dBs = mlp.backward(Y_b)
Ws, Bs = sgd(Ws,Bs,dWs,dBs,lr)
mlp.update_parameters(Ws,Bs)
for b in test_batches:
X_b = b[0]
Y_b = b[1]
Y_hat_b = mlp.forward(X_b)
test_loss = test_loss + sparse_categorical_cross_entropy(Y_b,Y_hat_b)
train_loss = train_loss/len(train_batches)
test_loss = test_loss/len(test_batches)
print("Train loss = {}, Test loss = {}".format(train_loss,test_loss))
train_losses.append(train_loss)
test_losses.append(test_loss)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="szbeGXj6OKI9" outputId="b9eec114-189c-493d-c316-915d3746257e"
plt.plot(train_losses)
plt.plot(test_losses)
# + colab={"base_uri": "https://localhost:8080/"} id="fEXc1MyEZlIR" outputId="e2194d4a-7a2a-4c62-f23d-d5033a3c758e"
mlp.evaluate(test_X,test_Y)
# + id="wznEiRsfcJl0"
start_idx = 1000
examples = test_X[start_idx:start_idx+5]
predictions = mlp.predict(examples)
# + colab={"base_uri": "https://localhost:8080/"} id="NGsaNbwdctGF" outputId="45d61136-9a38-4601-8665-31d6c2389a19"
print(predictions)
# + colab={"base_uri": "https://localhost:8080/", "height": 194} id="xda_qSF_cucx" outputId="d47d8292-d199-4dd8-85cb-270bd9627f94"
fig, axs = plt.subplots(1, 5,figsize=(15,15))
for idx,a in enumerate(axs):
a.imshow(test_X[start_idx+idx,:,:])
| MLP_From_Scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import altair as alt
import os as os
alt.data_transformers.enable('json')
os.getcwd()
df = pd.read_csv('../data/crimedata_csv_all_years.csv')
df['DATE'] = pd.to_datetime({'year':df['YEAR'],
'month':df['MONTH'],
'day':df['DAY'],
'hour':df['HOUR']})
dofw = pd.DataFrame({'day_of_week': ["Monday", "Tuesday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"],
'day_index': [1,2,3,4,5,6,7]})
df['day_of_week'] = pd.DatetimeIndex(df['DATE']).day_name()
df = pd.merge(df, dofw, how="left", on="day_of_week")
(df.groupby(['TYPE', 'NEIGHBOURHOOD']).count()/df.groupby(['TYPE']).count())[['DAY']].max()
df_line = df.query('TYPE == "Break and Enter Commercial"').groupby(['YEAR']).count().reset_index()
df_line.head()
df['TYPE'].unique()
# +
time_scale = "HOUR"
crime = "Theft of Bicycle"
neighbourhood = "ALL"
if neighbourhood != "ALL":
if crime != "ALL":
df_line = df.query('TYPE == @crime & NEIGHBOURHOOD == @neighbourhood').groupby([time_scale]).count().reset_index()
else:
df_line = df.query('NEIGHBOURHOOD == @neighbourhood').groupby([time_scale]).count().reset_index()
else:
if crime != "ALL":
df_line = df.query('TYPE == @crime').groupby([time_scale]).count().reset_index()
else:
df_line = df.groupby([time_scale]).count().reset_index()
alt.Chart(df_line).mark_line().encode(
alt.X(time_scale+':N'),
alt.Y('TYPE:Q', title='Number of Crimes'),
alt.Color(value="blue")
).configure(
background='#f7e0bc' #HEX color code
).configure_axisX(
labelAngle=45,
grid=True
).configure_axis(
labelFontSize=12,
titleFontSize=15
).configure_title(
fontSize=15
).properties(
height=300,
width=500,
title=crime
)
# -
| scr/altair-plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="41c10e35" colab={"base_uri": "https://localhost:8080/"} outputId="90b3d86a-9227-4b2e-e270-78570ef1194a"
#importing libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import json
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.ensemble import GradientBoostingClassifier, HistGradientBoostingClassifier, RandomForestClassifier
from sklearn.tree import DecisionTreeClassifier
from xgboost import XGBClassifier
from sklearn.linear_model import LogisticRegression
from google.colab import drive
drive.mount('/content/drive')
# + id="Knj_RKEHisAZ" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="68524da8-2321-46b8-ec7f-a8e7d78ce945"
# loading data
data = pd.read_csv("/content/drive/MyDrive/AML_category_identification/USvideos.csv") #40949 × 16
data
# + [markdown] id="LSfFzPFk8pN4"
# ### **Preprocessing Performed**
#
# Video ID: Label Encoding
#
# Trending_Date: Make new columns "day", "month", "year"
#
# Title: use top 50 words and see if each video has it
#
# Channel_title: Label Encoding
#
# Publish time: Adding column of "published quarters" and Label Encoding
#
# Tags: Label Encoding
#
# Views: Sector grouping
#
# Likes: Sector grouping
#
# Dislikes: Sector grouping
#
# Comments: Sector grouping
#
# Comments Disabled: Label Encoding
#
# Ratings Disabled: Label Encoding
#
# Video Error or Removed: Label Encoding
#
# Thumbnail_link: Drop
#
# Description: Label Encoding
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="FVlgrttmj4MB" outputId="e8eef2aa-556c-4998-99e9-05c471af9267"
# outputting json file details
with open('/content/drive/MyDrive/AML_category_identification/US_category_id.json') as f:
jj = json.load(f)
category_data = {}
for item in jj['items']:
category_data[int(item['id'])] = item['snippet']['title']
category_data
# + id="2H9yKQ9DkVWR"
# defining functions to use for preprocessing
def my_label_encoding(original_feature , new_feature) :
'''Performs label encoding on old feature from dataset and assigns it as a new feature. Drops the old feature.
Parameters
----------
original_feature: name of original feature
new_feature: name of new feature
Returns
-------
None
'''
enc = LabelEncoder()
enc.fit(data[original_feature])
data[new_feature] = enc.transform(data[original_feature])
data.drop([original_feature],axis=1, inplace=True)
def my_countplot(feature) :
'''Plotting category distribution on a countplot for a specific feature.
Parameters
----------
feature : name of the feature
Returns
-------
None
'''
sns.countplot(x=feature, data=data,facecolor=(0, 0, 0, 0),linewidth=5,edgecolor=sns.color_palette("prism", 5))
# + colab={"base_uri": "https://localhost:8080/"} id="pXzEtlkWkKiI" outputId="ff8adbc4-60e6-4a80-a541-b3adb0f4b534"
# checking unique values for each feature to determine which preprocessing method to take
for items in data.columns :
print('Unique values in "{0}": {1}'.format(items ,len(data[items].unique())))
# + [markdown] id="zRAigS5S8K4T"
# **Video ID**
# + id="zSYq_COj8AVO"
# label encoding
my_label_encoding('video_id', 'video_id_enc')
# + [markdown] id="aKF0zFVc8xJM"
# **Trending Date**
# + id="KtJ9dI_uo_vB"
# dividing into unique year, month, day
year_list = [] #17, 18
month_list = [] #01, 02, 03, 04, 05, 06, 11, 12
day_list = [] #01 - 31
for x in range(data.shape[0]) :
year_list.append(data['trending_date'][x][:2])
month_list.append(data['trending_date'][x][6:])
day_list.append(data['trending_date'][x][3:5])
# adding columns to the data
data.insert(16,'Year',year_list)
data.insert(17,'Month',month_list)
data.insert(18,'Day',day_list)
# + id="hZeHI4IHpysP"
# year encoding and deleting "trending_date" column
data.drop(['trending_date'],axis=1, inplace=True)
year_dict = {'17': 0,'18':1}
data['Year'] = data['Year'].map(year_dict)
# + [markdown] id="xrtZfCRV-usA"
# **Title**
# + colab={"base_uri": "https://localhost:8080/"} id="HG37-LY8-tMG" outputId="aed9719a-50e4-4de5-c6fc-dba15a2a520a"
# splitting all words and counting them
all_words = []
for x in range(data.shape[0]) :
all_words = all_words + data['title'][x].split()
print('Number of total words in title:', len(all_words))
# + colab={"base_uri": "https://localhost:8080/"} id="wGtPLndxsJu4" outputId="056646d9-df4e-4fd3-9aa8-8b20d7542083"
# finding top 50 words in "title"
all_words_serie = pd.Series(all_words)
top_50words = all_words_serie.value_counts()[:50]
top_50words = list(top_50words.index)
top_50words
# + id="zFzQiutXsQzW"
# removing words that give no info
top_50words.remove('-')
top_50words.remove('|')
top_50words.remove('The')
top_50words.remove('the')
top_50words.remove('a')
top_50words.remove('to')
top_50words.remove('of')
top_50words.remove('In')
top_50words.remove('with')
top_50words.remove('A')
top_50words.remove('&')
top_50words.remove('and')
top_50words.remove('in')
top_50words.remove('on')
top_50words.remove('To')
top_50words.remove('Is')
top_50words.remove('With')
top_50words.remove('at')
top_50words.remove('What')
top_50words.remove('is')
top_50words.remove('On')
top_50words.remove('This')
top_50words.remove('THE')
top_50words.remove('TO')
top_50words.remove('Of')
# + colab={"base_uri": "https://localhost:8080/"} id="d2c9fb77siUJ" outputId="c536dfbd-a63f-4e94-c379-041b45600242"
top_50words
# + id="tQWqxd4bslI9"
# assigning 1 if one of top 50 words exist for each "title" else 0
w1 = []
w2 = []
w3 = []
w4 = []
w5 = []
w6 = []
w7 = []
w8 = []
w9 = []
w10 = []
w11 = []
w12 = []
w13 = []
w14 = []
w15 = []
w16 = []
w17 = []
w18 = []
w19 = []
w20 = []
this_list = []
for x in range(data.shape[0]) :
this_list = data['title'][x].split()
if ('Video)' in this_list) or ('Video]' in this_list):
w1.append(1)
else:
w1.append(0)
if ('(Official' in this_list) or ( 'Official'in this_list) or ( '[Official' in this_list):
w2.append(1)
else:
w2.append(0)
if 'Trailer' in this_list:
w3.append(1)
else:
w3.append(0)
if 'You' in this_list:
w4.append(1)
else:
w4.append(0)
if '2' in this_list:
w5.append(1)
else:
w5.append(0)
if '2017' in this_list:
w6.append(1)
else:
w6.append(0)
if ('My' in this_list) or ( 'MY'in this_list) in this_list:
w7.append(1)
else:
w7.append(0)
if 'Me' in this_list:
w8.append(1)
else:
w8.append(0)
if 'I' in this_list:
w9.append(1)
else:
w9.append(0)
if ('for' in this_list) or ('For' in this_list):
w10.append(1)
else:
w10.append(0)
if '2018' in this_list:
w11.append(1)
else:
w11.append(0)
if 'Music' in this_list:
w12.append(1)
else:
w12.append(0)
if 'ft.' in this_list:
w13.append(1)
else:
w13.append(0)
if 'How' in this_list:
w14.append(1)
else:
w14.append(0)
if 'Why' in this_list:
w15.append(1)
else:
w15.append(0)
if 'New' in this_list:
w16.append(1)
else:
w16.append(0)
if ('from' in this_list) or ('From' in this_list):
w17.append(1)
else:
w17.append(0)
if 'it' in this_list:
w18.append(1)
else:
w18.append(0)
if 'We' in this_list:
w19.append(1)
else:
w19.append(0)
if 'Game' in this_list:
w20.append(1)
else:
w20.append(0)
# + id="lzeTKn_btUZU"
# defining the top words as new columns
data.insert(18,'word 1',w1)
data.insert(19,'word 2',w2)
data.insert(20,'word 3',w3)
data.insert(21,'word 4',w4)
data.insert(22,'word 5',w5)
data.insert(23,'word 6',w6)
data.insert(24,'word 7',w7)
data.insert(25,'word 8',w8)
data.insert(26,'word 9',w9)
data.insert(27,'word 10',w10)
data.insert(28,'word 11',w11)
data.insert(29,'word 12',w12)
data.insert(30,'word 13',w13)
data.insert(31,'word 14',w14)
data.insert(32,'word 15',w15)
data.insert(33,'word 16',w16)
data.insert(34,'word 17',w17)
data.insert(35,'word 18',w18)
data.insert(36,'word 19',w19)
data.insert(37,'word 20',w20)
# + id="XsoYEPv8ta_u"
# dropping title column
data.drop(['title'],axis=1, inplace=True)
# + [markdown] id="tcJDzr2IMiUd"
# **Channel Title**
# + id="fI8BdK2etdXs"
# label encoding channel title
my_label_encoding('channel_title' , 'channel_title_enc')
# + [markdown] id="GxpSwh7VNPYN"
# **Publish Time**
# + id="UPxAWiT8t4Fz"
# dividing published times into quarters (early morning, morning, afternoon, evening/night)
publish_quarter_periods = []
for x in range(data.shape[0]):
hour = int(str(data['publish_time'][x])[11:13])
if hour >=0 and hour < 6 :
publish_quarter_periods.append(1)
elif hour >=6 and hour < 12 :
publish_quarter_periods.append(2)
elif hour >=12 and hour < 18 :
publish_quarter_periods.append(3)
else:
publish_quarter_periods.append(4)
# + id="KqNhXOvUuDN2"
# inserting published quarter and label encoding published time
data.insert(37,'Publish Quarter',publish_quarter_periods)
my_label_encoding('publish_time' , 'publish_time_enc')
# + [markdown] id="3oo3fnE9N_ye"
# **Tags**
# + id="KixwUjWLuR1I"
# label encoding tags
my_label_encoding('tags' , 'tags_enc')
# + [markdown] id="W6fTABuwOiWc"
# **Views**
# + colab={"base_uri": "https://localhost:8080/"} id="dzmcbikbKMTk" outputId="55635fd2-fa2e-4c47-ec35-54cb15a83260"
# finding the max and min number of views
print('The minimum number of views:', data['views'].min())
print('The maximum number of views:', data['views'].max())
# + id="mG3uieGALDTM"
# defining function that splits feature numbers into groups
def feature_counter(data , max_value , feature , new_feature):
'''Splits a feature value into 5 groups.
Parameters
----------
data : dataset to be used
max_value : value to be considered when performing the feature magnitude splits
feature : feature name of dataset
new_feature : new feature name
Returns
-------
None
'''
data_max=data[feature].max()
data_min=data[feature].min()
rate = (max_value- data_min)/5
feature_val_list = []
for i in range(data.shape[0]) :
if data[feature][i] <= (data_min + rate):
feature_val_list.append(1)
elif data[feature][i] <= (data_min + (2*rate)):
feature_val_list.append(2)
elif data[feature][i] <= (data_min + (3*rate)):
feature_val_list.append(3)
elif data[feature][i] <= (data_min + (4*rate)):
feature_val_list.append(4)
else:
feature_val_list.append(5)
data.insert(data.shape[1], new_feature , feature_val_list)
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="qbnqvBFywgbf" outputId="a171788a-159f-4dd0-d718-c9aff97402b7"
# Creating countplot to visualize the group distibution and defining a "view" column for that
feature_counter(data ,10000000 , 'views' , 'views_group')
data.drop(['views'],axis=1, inplace=True)
my_countplot('views_group')
# + [markdown] id="rChn_iYjpLKj"
# **Likes**
# + colab={"base_uri": "https://localhost:8080/"} id="YDfq8EhRpT92" outputId="71a8101c-ac62-44bd-a7ff-8b3e1f31982e"
# finding the max and min number of likes
print('The minimum number of views:', data['likes'].min())
print('The maximum number of views:', data['likes'].max())
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="tO9MHve-xHbk" outputId="fe199494-2ceb-4ffb-9cd3-927834946381"
# Creating countplot to visualize the group distibution and defining a "likes" column for that
feature_counter(data ,500000 , 'likes' , 'likes_group')
data.drop(['likes'],axis=1, inplace=True)
my_countplot('likes_group')
# + [markdown] id="WxfYJkKjptyi"
# **Dislikes**
# + colab={"base_uri": "https://localhost:8080/"} id="hTkRAuhKpwMX" outputId="29a380c8-45e2-4804-80fd-bfe3a037604a"
# finding the max and min number of likes
print('The minimum number of views:', data['dislikes'].min())
print('The maximum number of views:', data['dislikes'].max())
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="2bcnAnQsTNes" outputId="7d06e5b7-81d2-4d3e-ec47-9697dad0603c"
# Creating countplot to visualize the group distibution and defining a "dislikes" column for that
feature_counter(data ,50000 , 'dislikes' , 'dislikes_group')
data.drop(['dislikes'],axis=1, inplace=True)
my_countplot('dislikes_group')
# + [markdown] id="xwRg_ScWQYfd"
# **Comments**
# + colab={"base_uri": "https://localhost:8080/", "height": 280} id="SBHeG7ydxd_a" outputId="ccd4b162-487f-40ab-c8b6-ec7c25a0ef16"
# Creating countplot to visualize the group distibution and defining a "comment_count" column for that
feature_counter(data ,10000 , 'comment_count' , 'comment_count_group')
data.drop(['comment_count'],axis=1, inplace=True)
my_countplot('comment_count_group')
# + [markdown] id="M9PuQdFmVUX1"
# **Thumbnail Link**
# + id="wsTURRVMVThH"
# dropping thumbnail link as no information is retreived
data.drop(['thumbnail_link'],axis=1, inplace=True)
# + [markdown] id="Yn35pJdfQiQz"
# **Comments disabled**
# + id="3tCyxMvUxrZM"
# label encoding "comments_disabled" column
my_label_encoding('comments_disabled','comments_disabled_enc')
# + [markdown] id="HBvp1U8nVujQ"
# **Ratings Disabled**
# + id="QxW9KFNOVooN"
# label encoding "ratings disabled" column
my_label_encoding('ratings_disabled','ratings_disabled_enc')
# + [markdown] id="3dXMaBZyWCWc"
# **Video Error or Removed**
# + id="sJM03JJsWKtV"
# label encoding "video error or removed" column
my_label_encoding('video_error_or_removed','video_error_or_removed_enc')
# + [markdown] id="X36hPwQEWWym"
# **Description**
# + id="QSyjmlXbWWBF"
# label encoding "description" column
my_label_encoding('description','description_enc')
# + [markdown] id="WytZ9I6cc70m"
# **Defining Datasets**
# + id="xwdromRSyA_Y" colab={"base_uri": "https://localhost:8080/"} outputId="c8209a26-1be8-4c79-e6e0-2cc078408e24"
# defining input matrix and labels
X = data.drop(['category_id'], axis=1, inplace=False)
y = data['category_id']
# splitting datasets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=33, shuffle =True, stratify=y)
# outputting shapes of each
print('X_train shape is ' , X_train.shape)
print('X_test shape is ' , X_test.shape)
print('y_train shape is ' , y_train.shape)
print('y_test shape is ' , y_test.shape)
# + [markdown] id="CLsts2SVdeNi"
# **Evaluating performances on Model (Only the optimal parameters are chosen and evaluated here)**
# + colab={"base_uri": "https://localhost:8080/"} id="rTswdawCyMom" outputId="1579b5a3-7bc5-4ddb-99f0-50f637392149"
# training and evaluating on gradient boosting classifier
GBCModel = GradientBoostingClassifier(n_estimators=100,max_depth=25,random_state=33)
GBCModel.fit(X_train, y_train)
print('GBCModel Test Score: ' , GBCModel.score(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="BLbyOIp3d3z5" outputId="a4d271d1-be88-4f86-84d1-af98bb5e946c"
# training and evaluating on histogram gradient boosting classifier
HGBCModel = HistGradientBoostingClassifier(max_leaf_nodes=100,max_depth=25,random_state=33)
HGBCModel.fit(X_train, y_train)
print('HGBCModel Test Score: ' , HGBCModel.score(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="77n0PxJueIwB" outputId="344689d2-3c53-4b75-c815-6ec98ea7d0cd"
# training and evaluating on xgboost classifier
X_train['Month'] = X_train['Month'].astype(str).astype(int)
X_train['Day'] = X_train['Day'].astype(str).astype(int)
X_test['Month'] = X_test['Month'].astype(str).astype(int)
X_test['Day'] = X_test['Day'].astype(str).astype(int)
XGBModel = XGBClassifier(n_estimators=100,max_depth=20,random_state=33)
XGBModel.fit(X_train, y_train)
print('XGBModel Test Score: ' , XGBModel.score(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="om8wGI__yYam" outputId="21efe6cc-7fab-42c2-923c-4fd28c655b37"
# training and evaluating on decision tree classifier
DecisionTreeClassifierModel = DecisionTreeClassifier(criterion='entropy',max_depth=30,random_state=33) #criterion can be entropy
DecisionTreeClassifierModel.fit(X_train, y_train)
#Calculating Details
print('DecisionTreeClassifierModel Train Score is : ' , DecisionTreeClassifierModel.score(X_train, y_train))
print('DecisionTreeClassifierModel Test Score is : ' , DecisionTreeClassifierModel.score(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="9-zG0xSVdYqg" outputId="3be6d858-4f2f-4d80-acfa-3817b75894df"
# training and evaluating on random forest classifier
RFCModel = RandomForestClassifier(max_leaf_nodes=2000, max_depth=50, random_state=33)
RFCModel.fit(X_train, y_train)
print('RandomForestClassifierModel Train Score is : ' , RFCModel.score(X_train, y_train))
print('RandomForestClassifierModel Test Score is : ' , RFCModel.score(X_test, y_test))
# + [markdown] id="Or4xJZBbwfIP"
# **Comparing Predictions and Actual values using our best model**
# + colab={"base_uri": "https://localhost:8080/"} id="aAO5OBUzq9sS" outputId="f120a01a-269e-4cd6-bcdf-7d6c4b23f4ca"
print('The first 5 category prediction: ', XGBModel.predict(X_test)[:5])
print('The first 5 categories: ', list(y_test[:5]))
# + [markdown] id="_waky_t-wuZ1"
# **Final Dataframe**
# + colab={"base_uri": "https://localhost:8080/", "height": 461} id="uPfL9QjEGJi_" outputId="dc74bb6d-07b9-4ecf-da0d-b311deeae456"
# dataframe used for preprocessing
data
# + id="39rShEh6vzIs"
| notebooks/category_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import importlib
#importlib.reload(_prepare)
# -
import warnings
warnings.filterwarnings(action='once')
# Original idea from: https://www.machinelearningplus.com/nlp/topic-modeling-visualization-how-to-present-results-lda-models/
#
#
# In this post, we discuss techniques to visualize the output and results from topic model (LDA) based on the gensim package. I will be using a portion of the 20 Newsgroups dataset since the focus is more on approaches to visualizing the results.
#
# Let’s begin by importing the packages and the 20 News Groups dataset.
# +
import sys
# # !{sys.executable} -m spacy download en
import re, numpy as np, pandas as pd
from pprint import pprint
# Gensim
import gensim, spacy, logging, warnings
import gensim.corpora as corpora
from gensim.utils import lemmatize, simple_preprocess
from gensim.models import CoherenceModel
import matplotlib.pyplot as plt
# NLTK Stop words
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
stop_words.extend(['linkremoved',' <link removed>','usernameremoved','<usernameremoved>','<linkremoved>','usernameremoved_usernameremoved','linkremoved_linkremoved'])
# %matplotlib inline
warnings.filterwarnings("ignore",category=DeprecationWarning)
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.ERROR)
# +
#Instructions about how to install mallet are available here: http://mallet.cs.umass.edu/download.php
'''
Windows installation: After unzipping MALLET, set the environment variable %MALLET_HOME% to point to the MALLET directory.
In all command line examples, substitute bin\mallet for bin/mallet.
'''
import os
from gensim.models.wrappers import LdaMallet
path_to_mallet_binary = "C:\\mallet-2.0.8\\bin\\mallet"
os.environ.update({'MALLET_HOME':r'C:\mallet-2.0.8'}) #OJO!, por alguna razon mallet solo puede estar disponible en esa carpeta
# -
# ## Import Cambridge Analytica datasets
#
#this libraries are for this dataset
import pandas as pd
file_name_1 = 'english_europe_tweets_20190411.csv'
file_name_2 = 'english_northamerica_tweets_20190411.csv'
df_1 = pd.read_csv('data/cambridge_analytica/regional_datasets/'+file_name_1)
df_2 = pd.read_csv('data/cambridge_analytica/regional_datasets/'+file_name_2)
# +
#df_1 = df_1.sample(1000) #remove this line
#df_2 = df_2.sample(1000) #remove this line
# -
df_1.head()
df_2.head()
# ## Tokenize Sentences and Clean
# Removing the emails, new line characters, single quotes and finally split the sentence into a list of words using gensim’s simple_preprocess(). Setting the deacc=True option removes punctuations.
# +
def sent_to_words(sentences):
for sent in sentences:
sent = re.sub('\S*@\S*\s?', '', sent) # remove emails
sent = re.sub('\s+', ' ', sent) # remove newline chars
sent = re.sub("\'", "", sent) # remove single quotes
sent = gensim.utils.simple_preprocess(str(sent), deacc=True)
yield(sent)
# +
# Convert to list
data_1 = df_1.texto_completo.values.tolist()
data_words_1 = list(sent_to_words(data_1))
print(data_words_1[:1])
data_2 = df_2.texto_completo.values.tolist()
data_words_2 = list(sent_to_words(data_2))
print(data_words_2[:2])
# -
# ## 4. Build the Bigram, Trigram Models and Lemmatize
#
# Let’s form the bigram and trigrams using the Phrases model. This is passed to Phraser() for efficiency in speed of execution.
#
# Next, lemmatize each word to its root form, keeping only nouns, adjectives, verbs and adverbs.
#
# We keep only these POS tags because they are the ones contributing the most to the meaning of the sentences. Here, I use spacy for lemmatization.
# +
#In case you haven't installed yet
# #!python -m spacy download en_core_web_sm
# +
# Build the bigram and trigram models
bigram_1 = gensim.models.Phrases(data_words_1, min_count=5, threshold=100) # higher threshold fewer phrases.
trigram_1 = gensim.models.Phrases(bigram_1[data_words_1], threshold=100)
bigram_mod_1 = gensim.models.phrases.Phraser(bigram_1)
trigram_mod_1 = gensim.models.phrases.Phraser(trigram_1)
bigram_2 = gensim.models.Phrases(data_words_2, min_count=5, threshold=100) # higher threshold fewer phrases.
trigram_2 = gensim.models.Phrases(bigram_2[data_words_2], threshold=100)
bigram_mod_2 = gensim.models.phrases.Phraser(bigram_2)
trigram_mod_2 = gensim.models.phrases.Phraser(trigram_2)
# +
# # !python3 -m spacy download en # run in terminal once
"""Remove Stopwords, Form Bigrams, Trigrams and Lemmatization"""
def process_words(texts, bigram_mod, trigram_mod, stop_words=stop_words, allowed_postags=['NOUN', 'ADJ', 'VERB', 'ADV']):
texts = [[word for word in simple_preprocess(str(doc)) if word not in stop_words] for doc in texts]
texts = [bigram_mod[doc] for doc in texts]
texts = [trigram_mod[bigram_mod[doc]] for doc in texts]
texts_out = []
nlp = spacy.load('en_core_web_sm', disable=['parser', 'ner'])
for sent in texts:
doc = nlp(" ".join(sent))
texts_out.append([token.lemma_ for token in doc if token.pos_ in allowed_postags])
# remove stopwords once more after lemmatization
texts_out = [[word for word in simple_preprocess(str(doc)) if word not in stop_words] for doc in texts_out]
return texts_out
# -
data_ready_1 = process_words(data_words_1, bigram_mod_1, trigram_mod_1) # processed Text Data!
data_ready_2 = process_words(data_words_2, bigram_mod_2, trigram_mod_2) # processed Text Data!
# # Build the topic model
# To build the LDA topic model using LdaModel(), you need the corpus and the dictionary. Let’s create them first and then build the model. The trained topics (keywords and weights) are printed below as well.
#
#
# ## topic modeling europe dataset
#with genim and mallet
number_topics_1 = 11
# +
# Create Dictionary
id2word_1 = corpora.Dictionary(data_ready_1)
# Create Corpus: Term Document Frequency
corpus_1 = [id2word_1.doc2bow(text) for text in data_ready_1]
# -
lda_model_1 = LdaMallet(path_to_mallet_binary, corpus=corpus_1, num_topics=number_topics_1, id2word=id2word_1)
#convert lda mallet model to lda gensim model
lda_model_1 = gensim.models.wrappers.ldamallet.malletmodel2ldamodel(lda_model_1)
pprint(lda_model_1.print_topics())
# ## topic modeling north america dataset
#with genim and mallet
number_topics_2 = 11
# +
# Create Dictionary
id2word_2 = corpora.Dictionary(data_ready_2)
# Create Corpus: Term Document Frequency
corpus_2 = [id2word_2.doc2bow(text) for text in data_ready_2]
# -
lda_model_2 = LdaMallet(path_to_mallet_binary, corpus=corpus_2, num_topics=number_topics_2, id2word=id2word_2)
#convert lda mallet model to lda gensim model
lda_model_2 = gensim.models.wrappers.ldamallet.malletmodel2ldamodel(lda_model_2)
pprint(lda_model_2.print_topics())
# ## Get most relevant documents - Europe dataset
# +
#should I use text or text_completo column? in the first, username and link are not removed
# -
'''
See the discusion here:
https://stackoverflow.com/questions/23509699/understanding-lda-transformed-corpus-in-gensim/37708396?noredirect=1#comment77429460_37708396
https://stackoverflow.com/questions/45310925/how-to-get-a-complete-topic-distribution-for-a-document-using-gensim-lda
'''
#with this code we get the full matrix of topic-documents contribution
matrix_documents_topic_contribution_1, _ = lda_model_1.inference(corpus_1)
matrix_documents_topic_contribution_1 /= matrix_documents_topic_contribution_1.sum(axis=1)[:, None]
matrix_documents_topic_contribution_1 = pd.DataFrame(matrix_documents_topic_contribution_1)
matrix_documents_topic_contribution_1.head()
contents_1 = pd.Series(df_1['text']).reset_index(drop=True)
matrix_documents_topic_contribution_1 = pd.concat([matrix_documents_topic_contribution_1, contents_1], axis=1)
matrix_documents_topic_contribution_1.head()
# ## Get most relevant documents - North america dataset
#with this code we get the full matrix of topic-documents contribution
matrix_documents_topic_contribution_2, _ = lda_model_2.inference(corpus_2)
matrix_documents_topic_contribution_2 /= matrix_documents_topic_contribution_2.sum(axis=1)[:, None]
matrix_documents_topic_contribution_2 = pd.DataFrame(matrix_documents_topic_contribution_2)
matrix_documents_topic_contribution_2.head()
contents_2 = pd.Series(df_2['text']).reset_index(drop=True)
matrix_documents_topic_contribution_2 = pd.concat([matrix_documents_topic_contribution_2, contents_2], axis=1)
matrix_documents_topic_contribution_2.head()
# # Topic similarity metric
# +
# Choose the # top keywords and # top documents a considerar en la metrica
topn_terms = 20
topk_documents = 20
relevance_lambda = 0.6
ruta_word_embedding = 'data/embedding_english_europe_northamerica_word2vec_300dimensions_cbow_trim3_epoch50.model'
word_embedding_model = gensim.models.Word2Vec.load(ruta_word_embedding)
# -
import topicvisexplorer
import importlib
importlib.reload(topicvisexplorer)
warnings.filterwarnings('ignore')
vis = topicvisexplorer.TopicVisExplorer("borrar_nombre")
topic_similarity_matrix_multicorpora = vis.calculate_topic_similarity_on_multi_corpora(word_embedding_model, lda_model_1,lda_model_2, corpus_1,corpus_2, id2word_1,id2word_2, matrix_documents_topic_contribution_1,matrix_documents_topic_contribution_2, topn_terms, topk_documents, relevance_lambda)
# ### Show visualization - Multi corpora
# +
#import topicvisexplorer
#import importlib
importlib.reload(topicvisexplorer)
vis = topicvisexplorer.TopicVisExplorer("borrar_nombre")
vis.prepare_multi_corpora( lda_model_1,lda_model_2, corpus_1, corpus_2, id2word_1,id2word_2, matrix_documents_topic_contribution_1, matrix_documents_topic_contribution_2, topic_similarity_matrix_multicorpora)
# -
#save data
vis.save_multi_corpora_data("models_output/multi_corpora_data_europe_northamerica_ca_lda_mallet_gensim.pkl")
'''
import topicvisexplorer
import importlib
importlib.reload(topicvisexplorer)
vis = topicvisexplorer.TopicVisExplorer("borrar_nombre")
vis.load_multi_corpora_data("multi_corpora_data_europe_northamerica_ca_lda_mallet_gensim.pkl")
'''
'''
vis.run()
'''
# # Topic similarity metric baseline
#
# +
# Choose the # top keywords and # top documents a considerar en la metrica
topn_terms = 20
topk_documents = 20
relevance_lambda = 0.6
ruta_word_embedding = 'data/embedding_english_europe_northamerica_word2vec_300dimensions_cbow_trim3_epoch50.model'
word_embedding_model = gensim.models.Word2Vec.load(ruta_word_embedding)
# +
import topicvisexplorer
import importlib
importlib.reload(topicvisexplorer)
warnings.filterwarnings('ignore')
vis = topicvisexplorer.TopicVisExplorer("borrar_nombre")
topic_similarity_matrix_multicorpora_metric_baseline = vis.calculate_topic_similarity_on_multi_corpora_metric_baseline(word_embedding_model, lda_model_1,lda_model_2, corpus_1, corpus_2, id2word_1,id2word_2, relevance_lambda, topn_terms)
# +
#import topicvisexplorer
#import importlib
importlib.reload(topicvisexplorer)
vis = topicvisexplorer.TopicVisExplorer("borrar_nombre")
vis.prepare_multi_corpora( lda_model_1,lda_model_2, corpus_1, corpus_2, id2word_1,id2word_2, matrix_documents_topic_contribution_1, matrix_documents_topic_contribution_2, topic_similarity_matrix_multicorpora_metric_baseline)
# -
#save data
vis.save_multi_corpora_data("models_output/multi_corpora_data_europe_northamerica_ca_lda_mallet_gensim_topic_similarity_baseline.pkl")
| official_notebooks/[User study] Multicorpora - A topic model for the Cambridge Analytica - Europe - Northamerica - Mallet and LDA Gensim.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decomposing unitary matrix into quantum gates
#
# This tool is useful when you have $2^n \times 2^n$ matrix representing a untary operator acting on register of $n$ bits and want to implement this operator in Q#.
#
# This notebook demonstrates how to use it.
# ## Tl;DR
import numpy, quantum_decomp
SWAP = numpy.array([[1,0,0,0],[0,0,1,0],[0,1,0,0], [0,0,0,1]])
print(quantum_decomp.matrix_to_qsharp(SWAP, op_name='Swap'))
# ## Example
#
# Consider following matrix:
#
# $$A = \frac{1}{\sqrt{3}}
# \begin{pmatrix}
# 1 & 1 & 1 & 0 \\
# 1 & e^{\frac{2\pi i}{3}} & e^{\frac{4 \pi i}{3}} & 0 \\
# 1 & e^{\frac{4\pi i}{3}} & e^{\frac{2 \pi i}{3}} & 0 \\
# 0 & 0 & 0 & -i \sqrt{3}
# \end{pmatrix}$$
#
# This is $3\times 3$ [DFT matrix](https://en.wikipedia.org/wiki/DFT_matrix), padded to have shape $4 \times 4$. Implementing such matrix was one way to solve problem B2 in [Microsoft Q# Coding Contest - Winter 2019](https://codeforces.com/blog/entry/65579).
# [Here](https://assets.codeforces.com/rounds/1116/contest-editorial.pdf) you can find another approach to implementing this matrix, but let's see how we can implement it using our tool and Q#.
#
# First, let's construct this matrix:
import numpy as np
w = np.exp((2j / 3) * np.pi)
A = np.array([[1, 1, 1, 0],
[1, w, w * w, 0],
[1, w * w, w, 0],
[0, 0, 0, -1j*np.sqrt(3)]]) / np.sqrt(3)
print(A)
# Now, let's use quantum_decomp library to construct Q# code.
import quantum_decomp as qd
print(qd.matrix_to_qsharp(A))
# As you can see from code in qsharp/ directory of this repository, this code indeed implements given unitary matrix.
# Also you can get the same sequence of operations as sequence of gates, where each gate is instance of GateFC or GateSingle, which are internal classes implementing fully controlled gate or gate acting on single qubit.
gates = qd.matrix_to_gates(A)
print('\n'.join(map(str, gates)))
# This can be represented by a quantum circuit (made with [Q-cirquit](http://physics.unm.edu/CQuIC/Qcircuit/)):
#
# <img src="res/circuit1.png">
# This is how you can view decomposition of matrix into 2-level gates, which is used to build sequence of gates.
print('\n'.join(map(str,qd.two_level_decompose_gray(A))))
# Those matrices are ordered in order they are applied, so to write them as a matrix product, we have to reverse them. This product can be written as follows:
#
# $$A =
# \begin{pmatrix} 0 & -i \\ -i & 0 \end{pmatrix}_{2,3}
# \begin{pmatrix} -\frac{\sqrt{2}}{2} & -\frac{\sqrt{2}}{2}i \\ -\frac{\sqrt{2}}{2}i & -\frac{\sqrt{2}}{2} \end{pmatrix}_{1,3}
# \begin{pmatrix} \sqrt{\frac{1}{3}} & \sqrt{\frac{2}{3}} \\ -\sqrt{\frac{2}{3}} & \sqrt{\frac{1}{3}} \end{pmatrix}_{0,1}
# \begin{pmatrix} \frac{\sqrt{2}}{2} & \frac{\sqrt{2}}{2} \\ -\frac{\sqrt{2}}{2} & \frac{\sqrt{2}}{2} \end{pmatrix}_{1,3}
# \begin{pmatrix} 0 & 1 \\ 1 & 0 \end{pmatrix}_{2,3}
# $$
#
# Or, in full form:
#
# $$A =
# \begin{pmatrix} 1 & 0 & 0 & 0 \\0& 1 & 0& 0 \\ 0 & 0 & 0 & -i \\ 0 & 0 & -i & 0 \end{pmatrix}
# \begin{pmatrix} 1 & 0 & 0 & 0 \\
# 0 & -\frac{\sqrt{2}}{2} & 0 & -\frac{\sqrt{2}}{2}i \\
# 0 & 0 & 1 & 0 \\
# 0 & -\frac{\sqrt{2}}{2}i & 0 & -\frac{\sqrt{2}}{2} \end{pmatrix}
# \begin{pmatrix} \sqrt{\frac{1}{3}} & \sqrt{\frac{2}{3}} & 0 & 0 \\
# -\sqrt{\frac{2}{3}} & \sqrt{\frac{1}{3}} & 0 & 0 \\
# 0 & 0 & 1 & 0 \\
# 0 & 0 & 0 & 1 \end{pmatrix}
# \begin{pmatrix} 1 & 0 & 0 & 0 \\
# 0 & \frac{\sqrt{2}}{2} & 0 & \frac{\sqrt{2}}{2} \\
# 0 & 0 & 1 & 0 \\
# 0 & -\frac{\sqrt{2}}{2} & 0 & \frac{\sqrt{2}}{2} \end{pmatrix}
# \begin{pmatrix} 1 & 0 & 0 & 0 \\
# 0 & 1 & 0 & 0 \\
# 0 & 0 & 0 & 1 \\
# 0 & 0 & 1 & 0 \end{pmatrix}
# $$
#
# ## Output size
#
# Number of Q# commands this tool produces is proportional to number of elements in matrix, which is $O(4^n)$, where $n$ is number of qubits in a register. More accurately, it's asymtotically $2 \cdot 4^n$. As it grows very fast, unfortunately this tool is useful only for small values of $n$.
#
# See detailed experimental complexity analysis of this tool in [this notebook](https://github.com/fedimser/quantum_decomp/blob/master/complexity.ipynb).
# ## Implementation
#
# Implementation is based on:
#
# * Article ["Decomposition of unitary matrices and quantum gates"](https://arxiv.org/pdf/1210.7366.pdf) by <NAME> and <NAME>;
# * Book "Quantum Computing: From Linear Algebra to Physical Implementations" (chapter 4) by <NAME> and <NAME>.
#
# It consists of following steps:
#
# 1. Decomposing matrix into 2-level unitary matrices;
# 2. Using Gray code to transform those matrices into matrices acting on states whose index differ only in one bit;
# 3. Implementing those matrices as fully controled single-qubit gates;
# 4. Implementing single-gate qubits as Rx, Ry and R1 gates;
# 5. Optimizations: cancelling X gates and removing identity gates.
# ## Paper
# Algorithm used in this tool is in detail outlined in this [paper](https://github.com/fedimser/quantum_decomp/blob/master/res/Fedoriaka2019Decomposition.pdf).
# ## Updates
#
# ### Optimized algorithm for 4x4 unitaries (Dec 2019)
#
# In case of 4x4 unitary one can implement it in much more effective way. Generic algorithm described above will produce 18 contolled gates, each of which should be implemented with at least 2 CNOTs and 3 single-qubit gates.
#
# As proven in [this paper](https://arxiv.org/pdf/quant-ph/0308006.pdf), it's possible to implement any 4x4 unitary using not more than 3 CNOT gates and 15 elementary single-qubit Ry and Rz gates.
#
# Algorithm for such optimal decomposition is now implemented in this library. To use it, pass `optimize=True` to functions performing decomposition.
#
# This example shows optimized decomposition for matrix A defined above.
qd.matrix_to_gates(A, optimize=True)
print(qd.matrix_to_qsharp(A, optimize=True))
# ### Circ support (Dec 2019)
#
# Now it's possible to convert unitary matrix to [Cirq](https://github.com/quantumlib/Cirq) circquit.
#
# You don't need to install Cirq to use the library, unless you want to have output as Cirq cirquit.
#
# See examples below.
print(qd.matrix_to_cirq_circuit(SWAP))
qd.matrix_to_cirq_circuit(A)
# To verify it's correct, let's convert random unitary to Cirq circuit, and then convert circuit back to matrix, and make sure we get the same matrix.
from scipy.stats import unitary_group
U = unitary_group.rvs(16)
np.linalg.norm(U - qd.matrix_to_cirq_circuit(U).unitary())
# ### Qiskit support (Dec 2020)
#
# *Feature added by [<NAME>](https://github.com/rvanasa).*
print(qd.matrix_to_qiskit_circuit(SWAP))
A_qiskit = qd.matrix_to_qiskit_circuit(A)
print(A_qiskit)
# Verify correctness of decompostion.
import qiskit.quantum_info as qi
np.linalg.norm(qi.Operator(A_qiskit).data - A)
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is one VERY simple example how to use RVmod as a fitting library.
# The RVmod options however, are a lot more. Reminder: the Exo-Striker
# GUI interface is warped around the RVmod.
#
#
# This example script delas with the Eta Ceti system (the usual demo in
# the Exo-Striker tool) and demonstrates how to fit Keplerian
# and Dynamical models to RV data.
#
#
# 1. We add the RV data
# 2. We find the RV offsets
# 3. We apply approx. parameters (to be taken from a GLS, for example.)
# 4. We fit to get the best two-planet Keplerian model
# 5. We adopt the best Keplerian fit and we include the dynamics into the modeling.
# 6. We make a simple plot showing the deviation between Keplerian and N-body models.
#
# There are some (commented) examples how one can run mcmc and/or nested sampling
# to get errors and/or posterior distributions.
#
# More detailed examples of how to use the RVmod will be provided
# as Jupyter notebooks in future.
#
#
# Created on Sun Jun 2 09:30:02 2019
#
# @author: <NAME>
# +
# # %load RVmod_as_py_lib_kep_vs_dyn_example.py
# #!/usr/bin/env python3
import sys
sys.path.append('../lib/') #RV_mod directory must be in your path
import RV_mod as rv
# Lets create the RVmod object
fit=rv.signal_fit('Eta Ceti demo',readinputfile=False);
fit.cwd = '../' # it is also important that the ES current working directory (cwd)
# point to the "lib" directory. This will be fixed in future releases
# add the stellar mass
fit.params.stellar_mass = 1.7 # In M sol.
# -
# Lets add the RV data
# +
fit.add_dataset("hip5364_lick", "../datafiles/hip5364.vels",0.0,0.0) # the last two entries are initial offset and jitter
fit.add_dataset("hip5364_VLT", "../datafiles/hip5364_crires.vels",0.0,0.0)
# Lets not fit for jitters now, i.e. keep at the initial value of 0 m/s
fit.use.use_jitters[0] = False
fit.use.use_jitters[1] = False
# Run it once to find the RV offsets, no planets yet.
fit.fitting(outputfiles=[1,1,1], doGP=False, minimize_fortran=True, minimize_loglik=False, amoeba_starts=20, print_stat=False)
#lets print the best fit params
print("Loglik = %s"%fit.loglik)
fit.print_info() #this is an obsolete function call, will be replaced!
# -
# Add the planetary initial estimates:
# Adding planets can be done automatically using the GLS first, but this will be covered in another intro.
# Since we know aprrox. the Keplerian parameters lets dirrectly add them as initial params.
# +
fit.add_planet(50, 400, 0.10, 200, 230, 90.0, 0.0) # K,P,e,omega,M0,i,Omega of planet 1
fit.add_planet(50, 770, 0.10, 170, 170, 90.0, 0.0) # K,P,e,omega,M0,i,Omega of planet 2
# lets fix the eccentricities first, this is advisable if you don't know well the system you are fitting.
fit.use.update_use_planet_params_one_planet(0,True,True,False,True,True,False,False)
fit.use.update_use_planet_params_one_planet(1,True,True,False,True,True,False,False)
# alternativly one can apply priors on the eccentricity, but this will be another exercise....
# one must get familiar with the priors
# for example:
#fit.e_norm_pr[0] = [0.0,0.1, True] first is \mu, second is \sigma, the Boolean is weather to use or not the prior
#fit.e_norm_pr[1] = [0.0,0.1, True]
#also if you apply priors with the fitting you must select minimize_fortran=False to use the SciPy wrapper.
# -
# Lets fit again with jitter fixed, and then again with jitter optimized
# +
fit.fitting(outputfiles=[1,1,1], doGP=False, minimize_fortran=True, minimize_loglik=False, amoeba_starts=20, print_stat=False)
#lets print the best fit params
print("Loglik = %s"%fit.loglik)
fit.print_info() #this is an obsolete function call, will be replaced!
# We can now relax the eccentricities and jitters
fit.use.update_use_planet_params_one_planet(0,True,True,True,True,True,False,False)
fit.use.update_use_planet_params_one_planet(1,True,True,True,True,True,False,False)
fit.use.use_jitters[0] = True
fit.use.use_jitters[1] = True
fit.fitting(outputfiles=[1,1,1], doGP=False, minimize_fortran=True, minimize_loglik=True, amoeba_starts=20, print_stat=False)
#lets print the best fit params
print("Loglik = %s"%fit.loglik)
fit.print_info() #this is an obsolete function call, will be relaced!
# -
# Now lets fit a dynamical model starting from the best Keplerian derived above
# +
#first lets copy the Keplerian object, we will need it later for plotting
import dill
kep_fit = dill.copy(fit)
fit.mod_dynamical=True
fit.fitting(outputfiles=[1,1,1], doGP=False, minimize_fortran=True, minimize_loglik=True, amoeba_starts=20, print_stat=False, eps=1000, dt=10, npoints=6000, model_max=0, model_min=0)
#lets print the best fit params
print("Loglik = %s"%fit.loglik)
fit.print_info() #this is an obsolete function call, will be replaced!
#lets copy the fit object as dyn_fit, we will need it later for plotting
dyn_fit = dill.copy(fit)
# -
# Lets make some basic plots with the "fit" object results
# +
################# Plotting #############################
import matplotlib.pyplot as plt
from matplotlib import gridspec
import matplotlib as mpl
import numpy as np
###### For nice plotting ##############
mpl.rcParams['axes.linewidth'] = 2.0 #set the value globally
mpl.rcParams['xtick.major.pad']='8'
mpl.rcParams['ytick.major.pad']='2'
# set tick width
mpl.rcParams['xtick.major.size'] = 8
mpl.rcParams['xtick.major.width'] = 2
mpl.rcParams['xtick.minor.size'] = 5
mpl.rcParams['xtick.minor.width'] = 2
mpl.rcParams['ytick.major.size'] = 8
mpl.rcParams['ytick.major.width'] = 2
mpl.rcParams['ytick.minor.size'] = 5
mpl.rcParams['ytick.minor.width'] = 2
mpl.rc('text',usetex=True)
font = {'family' : 'normal','weight' : 'bold','size' : 18,'serif':['Helvetica']}
mpl.rc('font', **font)
################## Time series plotting ###############
f = plt.figure(0, figsize=(8,6.5))
plt.subplots_adjust(hspace=0.005)
format_im = 'pdf'
dpi = 300
gs = gridspec.GridSpec(2, 1, height_ratios=[3, 1])
#gs.update( wspace=0.05)
ax1 = plt.subplot(gs[:-1, -1])
ax2 = plt.subplot(gs[-1, -1])
color = ['b', 'r', 'g', 'r']
symbol = ['o', 'o', 'o', 'o']
markersize = [6, 6, 6, 6]
alpha = [1, 1, 1, 1]
model_color = 'k'
model_lw = '1.0'
#### Get the time series (these below are self explanatory) ########
jd = kep_fit.fit_results.rv_model.jd
rvs = kep_fit.fit_results.rv_model.rvs
rv_err = kep_fit.fit_results.rv_model.rv_err
o_c = kep_fit.fit_results.rv_model.o_c
data_set = kep_fit.filelist.idset
# we can add the jitter
add_jitter = True
if add_jitter == True:
rv_err = np.array([np.sqrt(rv_err[i]**2 + kep_fit.params.jitters[ii]**2) for i,ii in enumerate(data_set)])
# Kep model time series #
kep_model_x = kep_fit.fit_results.model_jd
kep_model_y = kep_fit.fit_results.model
# Dyn model time series #
dyn_model_x = dyn_fit.fit_results.model_jd
dyn_model_y = dyn_fit.fit_results.model
###################################################################
zero_point_T = range((int(min(jd))-10),(int(max(jd))-10),10)
zero_point = np.zeros(len(zero_point_T))
ax1.plot(kep_model_x, kep_model_y, '-', linewidth=model_lw, color=model_color)
ax2.plot(zero_point_T,zero_point,'-', linewidth=model_lw, color=model_color)
overplot_dyn = True
if overplot_dyn == True:
ax1.plot(dyn_model_x, dyn_model_y, '--', linewidth=1.5, color='r')
for i in range(len(data_set)):
ax1.errorbar(jd[i],rvs[i], yerr=rv_err[i], alpha=alpha[int(data_set[i])], fmt=symbol[int(data_set[i])], linestyle='None', markersize = markersize[int(data_set[i])], color=color[int(data_set[i])], capsize = 0, elinewidth=1,mew=0.1)
ax2.errorbar(jd[i],o_c[i], yerr=rv_err[i], alpha=alpha[int(data_set[i])], fmt=symbol[int(data_set[i])], linestyle='None', markersize = markersize[int(data_set[i])],color=color[int(data_set[i])], capsize = 0, elinewidth=1,mew=0.1)
ax1.set_ylabel(r'RV [m/s]',fontsize=16, rotation = 'vertical')
ax1.set_xlim(min(jd),max(jd))
ax2.set_xlabel(r'BJD [day]',fontsize=16)
ax2.set_ylabel(r'o$-$c [m/s]',fontsize=16, rotation = 'vertical')
ax2.set_xlim(min(jd),max(jd))
ax2.locator_params(axis="x", nbins=9)
plt.setp( ax2.get_yticklabels(), fontsize=15,weight='bold')
plt.setp( ax2.get_xticklabels(), fontsize=15,weight='bold')
# Fine-tune figure; make subplots close to each other and hide x ticks for
# all but bottom plot.
plt.setp([a.get_xticklabels() for a in f.axes[:-1]], visible=False)
plt.show()
#plt.savefig('RV_plot_example.%s'%(format_im), format=format_im,dpi=dpi, bbox_inches='tight' )
ax1.cla()
ax2.cla()
# + active=""
# more options:
# +
#####################
# Run MCMC
#fit = rv.run_mcmc(fit, burning_ph=1000, mcmc_ph=5000, threads=30, output=False, fileoutput=True,save_means=False, save_mode=True, save_maxlnL=False)
# Run Nested sampling
#fit = rv.run_nestsamp(fit, threads=30, std_output=False, stop_crit = 0.0001, Dynamic_nest = False, live_points = 500, fileoutput=True, save_means=False, save_mode=False, save_maxlnL=True)
# WARNING! setup the bounds/prioirs first. Usually these are wide open and if you dont set them up
# it my take forever for the Nest. Samp. to finish. Unfortunatly I have to provide another example how to work with the RVmod priors/
# Work in progress....
#if you already have a session saved you may try:
#import dill
#file = open("session.ses", 'rb')
#fit = dill.load(file)
#file.close()
# and then for example
#fit = rv.run_mcmc(fit, burning_ph=1000, mcmc_ph=5000, threads=30, output=False, fileoutput=True, save_means=False, save_mode=True, save_maxlnL=False)
# -
| Notebook_and_script_examples/RVmod_as_py_lib_kep_vs_dyn_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Sbbarse787/Chances-of-Admit-through-GREscore-using-Linear-regression-model/blob/master/Linear_Regression_Model_.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="lH6SWhuslgKl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 142} outputId="971ce314-bb9f-43e3-d77b-ccb58c731f89"
from google.colab import drive
drive.mount('/gdrive')
# %cd /gdrive
# + [markdown] id="mUw6Tp61_7Ob" colab_type="text"
# # Lets Get Started
#
# + id="VqYaaZOGEj3p" colab_type="code" colab={}
# ls
# + id="bOdkoscblzww" colab_type="code" outputId="76da3049-6141-4b8d-c980-3a0d1591c65c" colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd /gdrive/My Drive/Colab Notebooks/SAS
# + id="GOfStSEq70tp" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + id="GhRvHS1G79RF" colab_type="code" outputId="fb106eaf-780b-4841-b171-807c3265b07a" colab={"base_uri": "https://localhost:8080/", "height": 194}
#import data set
dataset= pd.read_csv('Admission.csv') # Admission.csv ?? you have to deploy this dataset on your google drive
X= dataset.iloc[:,:-1].values
Y= dataset.iloc[:,1].values
dataset.head()
# + id="ZOoB5VIR7-KV" colab_type="code" outputId="a472c760-048a-4b54-db41-6127762e7fe8" colab={"base_uri": "https://localhost:8080/", "height": 34}
#Splitting the data
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test= train_test_split(X,Y,test_size= 1/3)
#Fitting Simple Linear Regression ipynb
#This is called Model
from sklearn.linear_model import LinearRegression
regressor= LinearRegression()
regressor.fit(X_train,Y_train)
# + id="IyGu8yVO8DbZ" colab_type="code" outputId="cc9d7167-e49e-4f01-b8dd-3ad4d2cbbe7d" colab={"base_uri": "https://localhost:8080/", "height": 573}
##Predicting the test results
Y_pred= regressor.predict(X_test)
#Visualising the training set Results
plt.scatter(X_train, Y_train, color='red')
plt.plot(X_train, regressor.predict(X_train), color='blue')
plt.title('GRE Score VS Chance Of Admit')
plt.xlabel('GRE Score')
plt.ylabel('Chance Of Admit')
plt.show()
plt.scatter(X_test, Y_test, color='red')
plt.plot(X_test, regressor.predict(X_test), color='blue')
plt.title('GRE Score Vs. Chance Of Admit')
plt.xlabel('Gre Score')
plt.ylabel('Chance of Admit')
plt.show()
# + id="P0tTbPSioHlN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="aa22f408-7ee9-4e4b-c6e9-e5d90e02834c"
a=int(input("What is the GRE Score? "))
print('The Value is', regressor.predict([[a]]))
| Linear_Regression_Model_.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Jupyter (formerly ipython) notebook intro
# * this document is a jupyter notebook
# * flexible tool to create readable documents that embeds:
# * code
# * images
# * comments
# * formulas
# * plots
# * jupyter supports notebooks form many different languages (including python, julia and R)
# * each document runs a computational engine that executes the code written in the cells
#
# Built-in magic commands in jupyter
#
# jupyter embeds special commands (called magics) that enables to run different kind of code.
# magic inline commands start with "%"
# magic multiple line commands start with "%%"
# you can list magic commands running %lsmagic
# you can pop-up the magic command documentation (if any) adding "?" after the command with no space
# few magics you might find useful:
# # ! runs inline shell commands
# # %%bash to run bash program (same syntax for other languages)
# embed latex inline between $$
# # %%latex to render a latex block
# ...
# # %reset to remove all names defined by the user
#
# %lsmagic
# + language="bash"
# for i in $(ls)
# do
# echo $i
# done
# -
# !ls | grep ipy
# ## Python interpreter
# * Compile source to bytecode then execute it on a virtual machine
# * Survives any execution error, even in case of syntax errors
# * in python also indentation is syntax
# * expression is code that returns an object
# * if no error, the prompt (">>>") automatically:
# * prints the result on screen
# * assigns the result to "_"
# expression that returns something
12345
# that something was assigned to "_"
_
# Error example: division by 0
1/0
print = 7
print("hello world")
# %reset
print("hello world")
| lectures/lectures/python/01_intro/01_jupyter_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prefilter Enviroment Map II
import random
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# 下面我们来处理第二部分:
# $$
# \int_{H} f(l, v) \cos \theta_l \mathrm{d} l = F_0 \int_{H} \frac{f(l, v)}{F(v, h)} \left(1-(1-v \cdot h)^{5}\right) \cos \theta_l \mathrm{d} l
# +
# \int_{H} \frac{f(l, v)}{F(v, h)}(1-v \cdot h)^{5} \cos \theta_l \mathrm{d} l
# $$
#
# - 也就是对 $F_0$ 的一个缩放系数和一个偏移量: $F_{0} w + b$
#
# 回顾我们的 BRDF:
# $$
# f(l, v) = \frac{D(h) F(v, h) G(l, v, h)}{4(n \cdot l)(n \cdot v)}
# $$
#
# 把它带入到上述公式的话,$F(v, h)$ 就消掉了。我们最终需预计算的也就是下面这两个积分:
# $$
# f^{\prime}(l, v) = \frac{D(h) G(l, v, h)}{4(n \cdot l)(n \cdot v)} \\
# w = \int_{H} f^{\prime}(l, v) \left(1-(1-v \cdot h)^{5}\right) \cos \theta_l \mathrm{d} l \\
# b = \int_{H} f^{\prime}(l, v) (1-v \cdot h)^{5} \cos \theta_l \mathrm{d} l
# $$
#
# +
v = np.random.rand(3)
v /= np.linalg.norm(v)
print(np.linalg.norm(v))
np.array([1,2,3])
# -
def integrate_brdf(roughness, NoV):
N = np.array([0,0,1])
V = np.zeros(3)
V[0] = NoV # cos
V[1] = np.sqrt(1-NoV**2) # sin
pdf = 2*np.pi
samples = 10
for i in range(samples):
R = roughness
G = NoV
return [R,G,0]
# +
img_size = 256
img = np.zeros((img_size,img_size,3)) # 生成一个两位数组,数组的每个元素是[R,G,B]
for y in range(img_size):
ny = 1-y/img_size
for x in range(img_size):
nx = x/img_size
img[y][x] = integrate_brdf(nx, ny)
plt.figure(figsize=[10,10])
plt.imshow(img)
plt.xlabel("$\\cos \\theta$")
plt.ylabel("roughness")
plt.show()
# -
# ## 参考资料
#
# - [SIGGRAPH 2013, Real Shading in Unreal Engine 4 (<NAME>)](https://blog.selfshadow.com/publications/s2013-shading-course/)
| docs/part-3/PrefilterEnvMap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## THE GOLDEN RULES
#
# | **Category** | **Rule** |
# | :--- | :--- |
# | 0. **PLAN BEFORE YOU CODE** | A. "Pseudo code" is writing out the broad steps in plain language. I often (almost always for complicated tasks) do this on paper, then translate it to code as an outline (in the code's comments). <br> <br> Maybe planning sounds boring and like a waste of time. I get it; I also want to [shoot first](https://youtu.be/la7uuFsCIrg?t=43) like [Han did...](https://youtu.be/93pXrmCdlI0?t=26) but coders like Han often [end up looking like this guy](https://youtu.be/mLyOj_QD4a4?t=67)... |
# | | B. Break the problem into chunks/smaller problems. This dovetails with rule 5.B below nicely. |
# | 1. Automation | A. Automate everything that can be automated, don't do point-and-click analysis! |
# | | B. Write a single script that executes all code from beginning to end |
# | 2. Version control | A. Store code and data under version control. |
# | | B. **Before checking the directory back in, clear all outputs and temp files and then run the whole directory!** (Check: Did it work right?) |
# | 3. [Directories/folders](10_Golden_6) | A. Separate directories/folders by function |
# | | B. Put input files into an input folder and outputs into a different one |
# | | A + B = your folders and files will be largely self documenting |
# | | C. Make directories portable - they should run on any computer, or if you move them to another place on your computer |
# | | D [**Use RELATIVE FILE PATHS, not absolute file paths**](10_Golden_7) |
# | 4. Keys / Units | A. Store cleaned data in tables with unique, non-missing "keys" |
# | | B. Keep data [**normalized**](10_Golden_4) as far into your code pipeline as you can |
# | 5. Abstraction - fncs/classes | A. Abstract to eliminate redundancy |
# | | B. Abstract to improve clarity |
# | | C. Otherwise, don't abstract |
# | | D. **Unit test your functions!** |
# | | E. Don't use magic numbers, define once as variables and refer as needed |
# | 6. Documentation | A. Is good... to a point |
# | | B. Don't write documentation you will not maintain |
# | | C. Code is better off when it is self-documenting |
# | 7. Look at your data/objects | As discussed [here](../01/07_debugging.html#seriously-print-your-data-and-objects-often) |
#
| content/02/10_Golden_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import ee
import geemap
import pandas as pd
Map = geemap.Map()
df = pd.read_excel("comunas.xlsx")
comunas = df["COD_COM"].unique().tolist()
def registros(cod, cant):
diccionario = {}
diccionario["COMUNA"] = cod
diccionario[year] = cant
return diccionario
# +
# startDate = ['2000-01-01', '2001-01-01', '2002-01-01', '2003-01-01', '2004-01-01', '2005-01-01', '2006-01-01', '2007-01-01', '2008-01-01', '2009-01-01', '2010-01-01', '2011-01-01', '2012-01-01', '2013-01-01', '2014-01-01', '2015-01-01', '2016-01-01', '2017-01-01', '2018-01-01', '2019-01-01', '2020-01-01']
# endDate = ['2001-01-01', '2002-01-01', '2003-01-01', '2004-01-01', '2005-01-01', '2006-01-01', '2007-01-01', '2008-01-01', '2009-01-01', '2010-01-01', '2011-01-01', '2012-01-01', '2013-01-01', '2014-01-01', '2015-01-01', '2016-01-01', '2017-01-01', '2018-01-01', '2019-01-01', '2020-01-01', '2021-01-01']
startDate = ['2004-01-01', '2005-01-01', '2006-01-01', '2007-01-01', '2008-01-01', '2009-01-01', '2010-01-01', '2011-01-01', '2012-01-01', '2013-01-01', '2014-01-01', '2015-01-01', '2016-01-01', '2017-01-01', '2018-01-01', '2019-01-01', '2020-01-01']
endDate = ['2005-01-01', '2006-01-01', '2007-01-01', '2008-01-01', '2009-01-01', '2010-01-01', '2011-01-01', '2012-01-01', '2013-01-01', '2014-01-01', '2015-01-01', '2016-01-01', '2017-01-01', '2018-01-01', '2019-01-01', '2020-01-01', '2021-01-01']
# -
for i in zip(startDate,endDate):
Fecha_inicial = str(i[0])
Fecha_final = str(i[1])
year = Fecha_inicial[:4]
puntos = []
for i in comunas:
studyArea = geemap.shp_to_ee("data/comunas/Lim_comunas.shp")
studyArea = studyArea.filterMetadata("COMUNA","equals", str(i))
FIRMS_colection2 = ee.ImageCollection('FIRMS')
FIRMS_colection = ee.ImageCollection('FIRMS')
FIRMS4 =FIRMS_colection2 \
.select(['T21']) \
.filterDate(Fecha_inicial,Fecha_final) \
.filterBounds(studyArea)
FIRMS =FIRMS_colection \
.select(['T21']) \
.filterDate(Fecha_inicial,Fecha_final) \
.filterBounds(studyArea)
FIRMScount4 = ee.Image(FIRMS4.count()).clip(studyArea)
FIRMSbinary4 = FIRMScount4.eq(FIRMScount4).rename('FIRMS_binary_alert_3')
project_crs = ee.Image(FIRMS.first()).projection().crs()
scale = ee.Image(FIRMS.first()).projection().nominalScale()
FIRMSpoint4 = FIRMSbinary4.reduceToVectors(
geometry = studyArea,
eightConnected=True,
labelProperty='modis_fire',
maxPixels=1e16,
crs=project_crs,
scale=scale,
geometryType= 'centroid',
bestEffort= True,
tileScale= 16
)
numero_PI = ee.FeatureCollection(FIRMSpoint4).filterBounds(studyArea)
cantidad_PI = numero_PI.size()
cantidad = cantidad_PI.getInfo()
diccionario = registros(i, cantidad)
puntos.append(diccionario.copy())
data = pd.DataFrame(puntos)
data.to_excel(str(year) + ".xlsx", index=False)
| algoritmos/hectareas_quemadas/descarga_comuna_anual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Software Engineering for Data Scientists
#
# ## *Sophisticated Data Manipulation*
# ## DATA 515 A
# ## 1. Python's Data Science Ecosystem
#
# With this simple Python computation experience under our belt, we can now move to doing some more interesting analysis.
# ### Python's Data Science Ecosystem
#
# In addition to Python's built-in modules like the ``math`` module we explored above, there are also many often-used third-party modules that are core tools for doing data science with Python.
# Some of the most important ones are:
#
# #### [``numpy``](http://numpy.org/): Numerical Python
#
# Numpy is short for "Numerical Python", and contains tools for efficient manipulation of arrays of data.
# If you have used other computational tools like IDL or MatLab, Numpy should feel very familiar.
#
# #### [``scipy``](http://scipy.org/): Scientific Python
#
# Scipy is short for "Scientific Python", and contains a wide range of functionality for accomplishing common scientific tasks, such as optimization/minimization, numerical integration, interpolation, and much more.
# We will not look closely at Scipy today, but we will use its functionality later in the course.
#
# #### [``pandas``](http://pandas.pydata.org/): Labeled Data Manipulation in Python
#
# Pandas is short for "Panel Data", and contains tools for doing more advanced manipulation of labeled data in Python, in particular with a columnar data structure called a *Data Frame*.
# If you've used the [R](http://rstats.org) statistical language (and in particular the so-called "Hadley Stack"), much of the functionality in Pandas should feel very familiar.
#
# #### [``matplotlib``](http://matplotlib.org): Visualization in Python
#
# Matplotlib started out as a Matlab plotting clone in Python, and has grown from there in the 15 years since its creation. It is the most popular data visualization tool currently in the Python data world (though other recent packages are starting to encroach on its monopoly).
# # 2. Installation
# ### Installing Pandas & friends
#
# Because the above packages are not included in Python itself, you need to install them separately. While it is possible to install these from source (compiling the C and/or Fortran code that does the heavy lifting under the hood) it is much easier to use a package manager like ``conda``. All it takes is to run
#
# ```
# $ conda install numpy scipy pandas matplotlib
# ```
#
# and (so long as your conda setup is working) the packages will be downloaded and installed on your system.
# # 3. Arrays and slicing in Numpy
import numpy as np
# ### Lists in native Python
#
# Let's create a **list**, a native Python object that we've used earlier today.
my_list = [2, 5, 7, 8]
my_list
type(my_list)
# This list is one-dimensional, let's make it multidimensional!
multi_list = [[1, 2, 3], [4, 5, 6]]
# How do we access the *6* element in the second row, third column for native Python list?
# +
#
# -
# ### Converting to numpy Arrays
my_array = np.array(my_list)
type(my_array)
my_array.dtype
multi_array.shape
multi_array = np.array([[1, 2, 3], [4, 5, 6]], np.int32)
# How do we access the *6* element in the second row, third column for numpy array?
# +
#
# -
# How do we retrieve a slice of the array, `array([[1, 2], [4, 5]])`?
# +
#
# -
# How do we retrieve the second column of the array?
# +
#
# -
# ## 4. Introduction to Pandas DataFrames
# What are the elements of a table?
# Pandas DataFrames as table elements
import pandas as pd
# What operations do we perform on tables?
df = pd.DataFrame({'A': [1,2,3], 'B': [2, 4, 6], 'ccc': [1.0, 33, 4]})
df
sub_df = df[['A', 'ccc']]
sub_df
df['A'] + 2*df['B']
# # Operations on a Pandas DataFrame
# ## 5. Manipulating Data with DataFrames
# ### Downloading the data
#
# Shell commands can be run from the notebook by preceding them with an exclamation point:
# !ls
# uncomment this to download the data:
# !curl -o pronto.csv https://data.seattle.gov/api/views/tw7j-dfaw/rows.csv?accessType=DOWNLOAD
# ### Loading Data into a DataFrame
# Because we'll use it so much, we often import under a shortened name using the ``import ... as ...`` pattern:
import pandas as pd
df = pd.read_csv('pronto.csv')
type(df)
len(df)
# Now we can use the ``read_csv`` command to read the comma-separated-value data:
# *Note: strings in Python can be defined either with double quotes or single quotes*
# ### Viewing Pandas Dataframes
# The ``head()`` and ``tail()`` methods show us the first and last rows of the data
df.head()
df.columns
df.index
smaller_df = df.loc[[1,4,6,7,9,34],:]
smaller_df.index
# The ``shape`` attribute shows us the number of elements:
df.shape
# The ``columns`` attribute gives us the column names
# The ``index`` attribute gives us the index names
# The ``dtypes`` attribute gives the data types of each column:
df.dtypes
# ### Sophisticated Data Manipulation
#
# Here we'll cover some key features of manipulating data with pandas
# Access columns by name using square-bracket indexing:
df_small = df['stoptime']
type(df_small)
df_small.tolist()
# Mathematical operations on columns happen *element-wise*:
trip_duration_hours = df['tripduration']/3600
trip_duration_hours[:2]
trip_duration_hours.head()
df['trip_duration_hours'] = df['tripduration']/3600
del df['trip_duration_hours']
df.head()
df.loc[[0,1],:]
df_long_trips = df[df['tripduration'] >10000]
sel = df['tripduration'] > 10000
df_long_trips = df[sel]
df_long_trips
df[sel].shape
# Make a copy of a slice
df_subset = df[['starttime', 'stoptime']].copy()
df_subset['trip_hours'] = df['tripduration']/3600
# Columns can be created (or overwritten) with the assignment operator.
# Let's create a *tripminutes* column with the number of minutes for each trip
# More complicated mathematical operations can be done with tools in the ``numpy`` package:
# ### Working with Times
# One trick to know when working with columns of times is that Pandas ``DateTimeIndex`` provides a nice interface for working with columns of times.
#
# For a dataset of this size, using ``pd.to_datetime`` and specifying the date format can make things much faster (from the [strftime reference](http://strftime.org/), we see that the pronto data has format ``"%m/%d/%Y %I:%M:%S %p"``
# (Note: you can also use ``infer_datetime_format=True`` in most cases to automatically infer the correct format, though due to a bug it doesn't work when AM/PM are present)
# With it, we can extract, the hour of the day, the day of the week, the month, and a wide range of other views of the time:
# ### Simple Grouping of Data
#
# The real power of Pandas comes in its tools for grouping and aggregating data. Here we'll look at *value counts* and the basics of *group-by* operations.
# #### Value Counts
# Pandas includes an array of useful functionality for manipulating and analyzing tabular data.
# We'll take a look at two of these here.
# The ``pandas.value_counts`` returns statistics on the unique values within each column.
#
# We can use it, for example, to break down rides by gender:
pd.value_counts(df["gender"])
# Or to break down rides by age:
pd.value_counts(2019 - df["birthyear"])
# By default, the values rather than the index are sorted. Use ``sort=False`` to turn this behavior off:
pd.value_counts(df["birthyear"], sort=False)
# We can explore other things as well: day of week, hour of day, etc.
# +
#
# -
# ### Group-by Operation
#
# One of the killer features of the Pandas dataframe is the ability to do group-by operations.
# You can visualize the group-by like this (image borrowed from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do))
df.head()
df_count = df.groupby(['from_station_id']).count()
df_count.head()
df_mean = df.groupby(['from_station_id']).mean()
df_mean.head()
dfgroup = df.groupby(['from_station_id'])
dfgroup.groups
# The simplest version of a groupby looks like this, and you can use almost any aggregation function you wish (mean, median, sum, minimum, maximum, standard deviation, count, etc.)
#
# ```
# <data object>.groupby(<grouping values>).<aggregate>()
# ```
#
# for example, we can group by gender and find the average of all numerical columns:
df.groupby(gender).mean()
# It's also possible to index the grouped object like it is a dataframe:
# You can even group by multiple values: for example we can look at the trip duration by time of day and by gender:
# The ``unstack()`` operation can help make sense of this type of multiply-grouped data. What this technically does is split a multiple-valued index into an index plus columns:
# ### Visualizing data with ``pandas``
#
# Of course, looking at tables of data is not very intuitive.
# Fortunately Pandas has many useful plotting functions built-in, all of which make use of the ``matplotlib`` library to generate plots.
#
# Whenever you do plotting in the IPython notebook, you will want to first run this *magic command* which configures the notebook to work well with plots:
# %matplotlib inline
# Now we can simply call the ``plot()`` method of any series or dataframe to get a reasonable view of the data:
import matplotlib.pyplot as plt
df['tripduration'].hist()
# ### Adjusting the Plot Style
#
# Matplotlib has a number of plot styles you can use. For example, if you like R you might use the ggplot style:
plt.style.use("ggplot")
# ### Other plot types
#
# Pandas supports a range of other plotting types; you can find these by using the <TAB> autocomplete on the ``plot`` method:
plt.
# For example, we can create a histogram of trip durations:
# If you'd like to adjust the x and y limits of the plot, you can use the ``set_xlim()`` and ``set_ylim()`` method of the resulting object:
# ## Breakout: Exploring the Data
#
# Make a plot of the total number of rides as a function of month of the year (You'll need to extract the month, use a ``groupby``, and find the appropriate aggregation to count the number in each group).
# Split this plot by gender. Do you see any seasonal ridership patterns by gender?
# Split this plot by user type. Do you see any seasonal ridership patterns by usertype?
# Repeat the above three steps, counting the number of rides by time of day rather that by month.
# Are there any other interesting insights you can discover in the data using these tools?
# ### Using Files
# - Writing and running python modules
# - Using python modules in your Jupyter Notebook
# A script for creating a dataframe with counts of the occurrence of a columns' values
df_count = df.groupby('from_station_id').count()
df_count1 = df_count[['trip_id']]
df_count2 = df_count1.rename(columns={'trip_id': 'count'})
df_count2.head()
def make_table_count(df_arg, groupby_column):
df_count = df_arg.groupby(groupby_column).count()
column_name = df.columns[0]
df_count1 = df_count[[column_name]]
df_count2 = df_count1.rename(columns={column_name: 'count'})
return df_count2
dff = make_table_count(df, 'from_station_id')
dff.head()
| Spring2019/02_Procedural_Python/Sophisticated Data Manipulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import calendar
import geopandas as gpd
import matplotlib.dates as mdates
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import Markdown
from pyspark.sql import SparkSession
from pyspark.sql.functions import udf
from pyspark.sql.types import *
from shapely.geometry import Point
palette = sns.color_palette("colorblind", 20)
spark = (
SparkSession.builder
.master("local")
.appName("TFL Notebook")
.config('spark.executor.memory', '8G')
.config('spark.driver.memory', '16G')
.config('spark.driver.maxResultSize', '10G')
.config("spark.sql.crossJoin.enabled", "true")
.getOrCreate()
)
trips = spark.read.parquet("../data/parquet_trip")
trips.createOrReplaceTempView("trips")
df = spark.sql("""
select
count(distinct(bike_id)) as bike_count,
count(1) as trip_count,
start_year,
start_month,
start_year || '-' || start_month as month
from trips
group by start_year, start_month
order by start_year asc, start_month asc
""")
def month_label(year, month):
return "{0} {1}".format(calendar.month_name[int(month)], year)
month_label_udf = udf(month_label, StringType())
df = df.withColumn("month_label", month_label_udf("start_year", "start_month"))
df.createOrReplaceTempView("monthly_counts")
monthly_counts = df.toPandas()
monthly_counts.head(10)
# +
fig, ax = plt.subplots(figsize=(15, 8))
monthly_counts.plot(ax=ax, kind='line', x='month_label', xticks=range(0, len(monthly_counts)), y='bike_count', linewidth=4.0)
plt.title("Number of bikes used per month", fontsize=18)
plt.xlabel('')
#plt.ylim(bottom=0, top=13000)
plt.xticks(rotation=90)
plt.ylabel('Number of Bikes', fontsize=12)
plt.legend(loc='best')
plt.show()
# +
df = spark.sql("""
select
count(distinct(t.bike_id)) as bike_count,
t.start_year,
t.start_month,
t.start_day
from trips t
group by t.start_year, t.start_month, t.start_day
""")
df.createOrReplaceTempView("daily_counts")
df = spark.sql("""
select
dc.bike_count as bikes_used,
mc.bike_count as bikes_available,
mc.bike_count - dc.bike_count as unused_bikes,
dc.start_year,
dc.start_month,
dc.start_day
from daily_counts dc
join monthly_counts mc on (dc.start_year = mc.start_year and dc.start_month = mc.start_month)
""")
df.createOrReplaceTempView("daily_usage")
df = spark.sql("""
select
min(bikes_used) as bikes_used_min,
max(bikes_used) as bikes_used_max,
avg(bikes_used) as bikes_used_avg,
max(bikes_available) as bikes_available,
min(unused_bikes) as unused_bikes_min,
max(unused_bikes) as unused_bikes_max,
avg(unused_bikes) as unused_bikes_avg,
start_year,
start_month
from daily_usage
group by start_year, start_month
order by start_year asc, start_month asc
""")
df = df.withColumn("month_label", month_label_udf("start_year", "start_month"))
utilisation = df.toPandas()
utilisation.head(10)
# +
df = spark.sql("""
select
min(bikes_used) as bikes_used_min,
max(bikes_used) as bikes_used_max,
avg(bikes_used) as bikes_used_avg,
max(bikes_available) as bikes_available,
min(unused_bikes) as unused_bikes_min,
max(unused_bikes) as unused_bikes_max,
avg(unused_bikes) as unused_bikes_avg,
start_year,
start_month
from daily_usage
group by start_year, start_month
order by start_year asc, start_month asc
""")
fig, ax = plt.subplots(figsize=(15, 8))
ax.plot(utilisation['month_label'], utilisation['unused_bikes_avg'], linewidth=4.0, label="Unused bikes: min, max, avg")
ax.fill_between(utilisation['month_label'], utilisation['unused_bikes_min'], utilisation['unused_bikes_max'], alpha=0.5)
ax.plot(utilisation['month_label'], utilisation['bikes_available'], linewidth=4.0, label="Available bikes")
plt.title("TFL Cycle Utilization", fontsize=18)
plt.xlabel('')
plt.ylim(bottom=0, top=13000)
plt.xticks(rotation=90)
plt.ylabel('Number of Bikes', fontsize=12)
plt.legend(loc='lower right')
plt.show()
# -
| notebooks/bike_utilisation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ingetsing larger catalogs: PanStarrs DR1 Mean/Thin Objects
#
# This notebook follows up on *insert_example*, be sure to go through that one first.
#
# Here we show a possible way in which the catalog ingestion step can be sped up though the use of mulitprocessing. In particular, we will try to import some of the files from a subset and compilation of PS1 DR1 mean and thin object tables. The entire subsample contains 1.92B sources, selected requiring nDetections>2.
#
# Reference: https://panstarrs.stsci.edu/
#
# The test files we will be using for this test can be downloaded from:
#
# https://desycloud.desy.de/index.php/s/stCkA6uJ8ayKvjI
# +
from extcats import CatalogPusher
import pandas as pd
import numpy as np
import concurrent.futures
from healpy import ang2pix
import importlib
importlib.reload(CatalogPusher)
# build the pusher object and point it to the raw files.
ps1p = CatalogPusher.CatalogPusher(
catalog_name = 'ps1_test', # short name of the catalog
data_source = '../testdata/PS1DR1_test/', # where to find the data (other options are possible)
file_type = '*.csv.gz' # filter files (there is col definition file in data_source)
)
# define the reader for the raw files (import column names from file.)
headfile = '../testdata/PS1DR1_test/column_headings.csv'
with open(headfile, 'r') as header:
catcols=[c.strip() for c in header.readline().split(',')]
# skimm out some columns
bad = ['projectionID', 'skyCellID']
usecols = [c for c in catcols if (not c in bad) or ('gNpt' in c)]
# specify some data types to save up on the storage
# See https://outerspace.stsci.edu/display/PANSTARRS/PS1+MeanObject+table+fields
types = {}
for c in usecols:
types[c] = np.float16
if c == 'objID':
types[c] = np.int32
if 'Flags' in c:
types[c] = np.int16
if ('ra' in c) or ('dec' in c):
types[c] = np.float32
ps1p.assign_file_reader(
reader_func = pd.read_csv, # callable to use to read the raw_files.
read_chunks = True, # weather or not the reader process each file into smaller chunks.
names=catcols, # All other arguments are passed directly to this function.
usecols=usecols,
dtype = types,
na_values = -999,
chunksize=50000,
engine='c')
# define modifier. This time the healpix grid is finer (an orer 16 corresponds to 3")
hp_nside16=2**16
def ps1_modifier(srcdict):
srcdict['_id'] = srcdict.pop('objID')
srcdict['hpxid_16']=int(
ang2pix(hp_nside16, srcdict['raMean'], srcdict['decMean'], lonlat = True, nest = True))
return srcdict
ps1p.assign_dict_modifier(ps1_modifier)
# wrap up the file pushing function so that we can
# use multiprocessing to speed up the catalog ingestion
def pushfiles(filerange):
# push stuff
ps1p.push_to_db(
coll_name = 'srcs',
index_on = ['hpxid_16'],
filerange = filerange,
overwrite_coll = False,
dry = False,
fillna_val = None)
# add metadata to direct queries
ps1p.healpix_meta(
healpix_id_key = '<KEY>',
order = 16, is_indexed = True, nest = True)
# each job will run on a subgroup of all the files
file_groups = ps1p.file_groups(group_size=1)
with concurrent.futures.ProcessPoolExecutor(max_workers = 2) as executor:
executor.map(pushfiles, file_groups)
print ("done! Enjoy your PS1_test database.")
# -
| notebooks/example_ingest_multiproc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
class Node:
def __init__(self,data):
self.data = data
self.next = None
class Node:
def __init__(self,data):
self.data = data
self.next = None
def take_input():
inputList = list(map(int, input().split()))
head = None
for curr_data in inputList:
if curr_data == -1:
break
newNode = Node(curr_data)
if head is None:
head = newNode
last = head
else:
last.next = newNode
last = newNode
printLL(head)
return head
def lengthLL(head):
l = 0
while head is not None:
l += 1
head = head.next
return l
def insertatI(head, i, data):
if i < 0 or i > lengthLL(head):
return head
count = 0
prev = None
curr = head
while count < i:
prev = curr
curr = curr.next
count += 1
newNode = Node(data)
if prev is not None:
prev.next = newNode
else:
head = newNode
newNode.next = curr
return head
def printLL(head):
while head is not None:
print(str(head.data) + "=>" , end = "")
head = head.next
print(None)
return
def reverseR1(head):
prev = None
curr = head
while curr is not None:
next = curr.next
curr.next = prev
prev = curr
curr = next
head = prev
return head
def makeLL():
inputList = [i for i in range(14)]
head = None
for curr_data in inputList:
if curr_data == -1:
break
newNode = Node(curr_data)
if head is None:
head = newNode
last = head
else:
last.next = newNode
last = newNode
printLL(head)
return head
# -
def reverseT(head):
prev = None
curr = head
while curr is not None:
next = curr.next
curr.next = prev
prev = curr
curr = next
return prev, head
# +
def kReverse(head,k):
if head is None:
return None
h1 = head
t1 = head
count = 1
while count < k and t1 is not None:
t1 = t1.next
count += 1
if t1 is None:
a, b = reverseT(h1)
return a
h2 = t1.next
t1.next = None
hR, tR = reverseT(h1)
smallHead = kReverse(h2, k)
tR.next = smallHead
return hR
# -
head = makeLL()
head = kReverse(head,3)
printLL(head)
def reverse(head, k):
if head == None:
return None
current = head
next = None
prev = None
count = 0
# Reverse first k nodes of the linked list
while(current is not None and count < k):
next = current.next
current.next = prev
prev = current
current = next
count += 1
# next is now a pointer to (k+1)th node
# recursively call for the list starting
# from current. And make rest of the list as
# next of first node
if next is not None:
head.next = reverse(next, k)
# prev is new head of the input list
return prev
head = makeLL()
head = reverse(head,3)
printLL(head)
def bubbleSort(head):
swap = 0
while True:
swap = 0
temp = head
while temp.next is not None:
if temp.data > temp.next.data:
# make a swap
swap += 1
p = temp.data
temp.data = temp.next.data
temp.next.data = p
temp = temp.next
else:
temp = temp.next
if swap == 0:
break
return head
head = take_input()
head = bubbleSort(head)
printLL(head)
class Node:
def __init__(data,next):
self.data = data
self.next = next
| CN DSA/LL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (dl)
# language: python
# name: dl
# ---
# ## Compare 3D Convolutional Neural Networks
# One approach when dealing with multiple frames is to employ 3D convolutional neural networks. In addition to convolving over the height and width of an image, we convolve through time (the depth dimension). We're hoping that this will allow us to detect changes in the video over time such as flickers or jitter.
import torch
import numpy as np
from fastai.core import *
from fastai.vision import *
import matplotlib.pyplot as plt
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import math
from functools import partial
__all__ = [
'ResNet', 'resnet10', 'resnet18', 'resnet34', 'resnet50', 'resnet101',
'resnet152', 'resnet200'
]
def conv3x3x3(in_planes, out_planes, stride=1):
# 3x3x3 convolution with padding
return nn.Conv3d(
in_planes,
out_planes,
kernel_size=3,
stride=stride,
padding=1,
bias=False)
def downsample_basic_block(x, planes, stride):
out = F.avg_pool3d(x, kernel_size=1, stride=stride)
zero_pads = torch.Tensor(
out.size(0), planes - out.size(1), out.size(2), out.size(3),
out.size(4)).zero_()
if isinstance(out.data, torch.cuda.FloatTensor):
zero_pads = zero_pads.cuda()
out = Variable(torch.cat([out.data, zero_pads], dim=1))
return out
class BasicBlock(nn.Module):
expansion = 1
def __init__(self, inplanes, planes, stride=1, downsample=None):
super(BasicBlock, self).__init__()
self.conv1 = conv3x3x3(inplanes, planes, stride)
self.bn1 = nn.BatchNorm3d(planes)
self.relu = nn.ReLU(inplace=True)
self.conv2 = conv3x3x3(planes, planes)
self.bn2 = nn.BatchNorm3d(planes)
self.downsample = downsample
self.stride = stride
def forward(self, x):
residual = x
out = self.conv1(x)
out = self.bn1(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn2(out)
if self.downsample is not None:
residual = self.downsample(x)
out += residual
out = self.relu(out)
return out
class Bottleneck(nn.Module):
expansion = 4
def __init__(self, inplanes, planes, stride=1, downsample=None):
super(Bottleneck, self).__init__()
self.conv1 = nn.Conv3d(inplanes, planes, kernel_size=1, bias=False)
self.bn1 = nn.BatchNorm3d(planes)
self.conv2 = nn.Conv3d(
planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn2 = nn.BatchNorm3d(planes)
self.conv3 = nn.Conv3d(planes, planes * 4, kernel_size=1, bias=False)
self.bn3 = nn.BatchNorm3d(planes * 4)
self.relu = nn.ReLU(inplace=True)
self.downsample = downsample
self.stride = stride
def forward(self, x):
residual = x
out = self.conv1(x)
out = self.bn1(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn2(out)
out = self.relu(out)
out = self.conv3(out)
out = self.bn3(out)
if self.downsample is not None:
residual = self.downsample(x)
out += residual
out = self.relu(out)
return out
class ResNet(nn.Module):
def __init__(self,
block,
layers,
sample_size,
sample_duration,
shortcut_type='B',
num_classes=400):
self.inplanes = 64
super(ResNet, self).__init__()
self.sample_duration = sample_duration
self.conv1 = nn.Conv3d(
3,
64,
kernel_size=7,
stride=(1, 2, 2),
padding=(3, 3, 3),
bias=False)
self.bn1 = nn.BatchNorm3d(64)
self.relu = nn.ReLU(inplace=True)
self.maxpool = nn.MaxPool3d(kernel_size=(3, 3, 3), stride=2, padding=1)
self.layer1 = self._make_layer(block, 64, layers[0], shortcut_type)
self.layer2 = self._make_layer(
block, 128, layers[1], shortcut_type, stride=2)
self.layer3 = self._make_layer(
block, 256, layers[2], shortcut_type, stride=2)
self.layer4 = self._make_layer(
block, 512, layers[3], shortcut_type, stride=2)
last_duration = int(math.ceil(sample_duration / 16))
last_size = int(math.ceil(sample_size / 32))
self.avgpool = nn.AvgPool3d(
(last_duration, last_size, last_size), stride=1)
self.fc = nn.Linear(512 * block.expansion, num_classes)
for m in self.modules():
if isinstance(m, nn.Conv3d):
m.weight = nn.init.kaiming_normal_(m.weight, mode='fan_out')
elif isinstance(m, nn.BatchNorm3d):
m.weight.data.fill_(1)
m.bias.data.zero_()
def _make_layer(self, block, planes, blocks, shortcut_type, stride=1):
downsample = None
if stride != 1 or self.inplanes != planes * block.expansion:
if shortcut_type == 'A':
downsample = partial(
downsample_basic_block,
planes=planes * block.expansion,
stride=stride)
else:
downsample = nn.Sequential(
nn.Conv3d(
self.inplanes,
planes * block.expansion,
kernel_size=1,
stride=stride,
bias=False), nn.BatchNorm3d(planes * block.expansion))
layers = []
layers.append(block(self.inplanes, planes, stride, downsample))
self.inplanes = planes * block.expansion
for i in range(1, blocks):
layers.append(block(self.inplanes, planes))
return nn.Sequential(*layers)
def forward(self, x):
batch_size, stacked_size, height, width = x.shape
rgb_channels = 3
# fastai requires that inputs be of shape:
# (BATCH, CHANNELS * NUM_FRAMES, HEIGHT, WIDTH)
# PyTorch's 3D convolution operations require that inputs be of shape:
# (BATCH, CHANNELS, NUM_FRAMES, HEIGHT, WIDTH)
x = x.view(batch_size,10, rgb_channels, height, width).permute(0, 2,1,3,4)
# print(x.shape) #torch.Size([64, 3, 10, 128, 128])
# plt.imshow(x[0,:,0,:,:].permute(1,2,0))
# plt.show()
# Reshaping and permuting puts x back on the CPU, so we'll move it back to the GPU.
x = x.cuda()
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.maxpool(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.layer4(x)
x = self.avgpool(x)
x = x.view(x.size(0), -1)
x = self.fc(x)
return x
def get_fine_tuning_parameters(model, ft_begin_index):
if ft_begin_index == 0:
return model.parameters()
ft_module_names = []
for i in range(ft_begin_index, 5):
ft_module_names.append('layer{}'.format(i))
ft_module_names.append('fc')
parameters = []
for k, v in model.named_parameters():
for ft_module in ft_module_names:
if ft_module in k:
parameters.append({'params': v})
break
else:
parameters.append({'paramsbatch_size, height, width, stacked_size = input.shape': v, 'lr': 0.0})
return parameters
def resnet10(**kwargs):
"""Constructs a ResNet-18 model.
"""
model = ResNet(BasicBlock, [1, 1, 1, 1], **kwargs)
return model
def resnet18(**kwargs):
"""Constructs a ResNet-18 model.
"""
model = ResNet(BasicBlock, [2, 2, 2, 2], **kwargs)
return model
def resnet34(**kwargs):
"""Constructs a ResNet-34 model.
"""
model = ResNet(BasicBlock, [3, 4, 6, 3], **kwargs)
return model
def resnet50(**kwargs):
"""Constructs a ResNet-50 model.
"""
model = ResNet(Bottleneck, [3, 4, 6, 3], **kwargs)
return model
def resnet101(**kwargs):
"""Constructs a ResNet-101 model.
"""
model = ResNet(Bottleneck, [3, 4, 23, 3], **kwargs)
return model
def resnet152(**kwargs):
"""Constructs a ResNet-101 model.
"""
model = ResNet(Bottleneck, [3, 8, 36, 3], **kwargs)
return model
def resnet200(**kwargs):
"""Constructs a ResNet-101 model.
"""
model = ResNet(Bottleneck, [3, 24, 36, 3], **kwargs)
return model
# -
path = Path('../data/hard_video_frames')
# +
def open_video_frames_as_image(self, filename):
# Open frames
frames = np.load(filename)
# Convert to tensor and normalize
frames_tensor = torch.from_numpy(frames).float()
frames_tensor.div_(255)
frames_tensor = frames_tensor.permute(2, 0, 1)
return Image(frames_tensor)
ImageList.open = open_video_frames_as_image
# -
src = ImageList.from_folder(path, extensions='.npy').split_by_folder(train='train', valid='val')
src
def get_data(bs,size):
data = (src.label_from_re('([A-Z]+).npy$')
.transform(get_transforms(max_warp=0, max_zoom=1), size=size)
.databunch(bs=bs))
#TODO: Normalize somehow
return data
bs, sz = 32, 256
data = get_data(bs, sz)
# 10 frames in each sample
sample_duration = 10
def run_resnet(model_name):
if model_name == "resnet18":
model = resnet18(sample_size=sz, sample_duration=sample_duration, num_classes=400, shortcut_type='A')
model = model.cuda()
model = nn.DataParallel(model, device_ids=None)
saved_state = torch.load('3DCNNs/resnet-18-kinetics.pth')
state_dict = saved_state['state_dict']
model.load_state_dict(state_dict)
# Adjust the last layer to our problems classification task
model.module.fc = torch.nn.Linear(in_features=512, out_features=2, bias=True)
elif model_name == "resnet34":
model = resnet34(sample_size=sz, sample_duration=sample_duration, num_classes=400, shortcut_type='A')
model = model.cuda()
model = nn.DataParallel(model, device_ids=None)
saved_state = torch.load('3DCNNs/resnet-34-kinetics.pth')
state_dict = saved_state['state_dict']
model.load_state_dict(state_dict)
# Adjust the last layer to our problems classification task
model.module.fc = torch.nn.Linear(in_features=512, out_features=2, bias=True)
elif model_name == "resnet50":
bs, sz = 16, 256
data = get_data(bs, sz)
# 10 frames in each sample
sample_duration = 10
model = resnet50(sample_size=sz, sample_duration=sample_duration, num_classes=400, shortcut_type='B')
model = model.cuda()
model = nn.DataParallel(model, device_ids=None)
saved_state = torch.load('3DCNNs/resnet-50-kinetics.pth')
state_dict = saved_state['state_dict']
model.load_state_dict(state_dict)
# Adjust the last layer to our problems classification task
model.module.fc = torch.nn.Linear(in_features=2048, out_features=2, bias=True)
# Create a learner and group parameters
learner = Learner(data, model, metrics=[accuracy])
learner.fit_one_cycle(20, 1e-4)
# Post-Training Review
learner.recorder.plot_losses()
interp = ClassificationInterpretation.from_learner(learner)
interp.plot_confusion_matrix()
#interp.plot_top_losses(9, figsize=(10,10))
run_resnet('resnet18')
run_resnet('resnet34')
run_resnet('resnet50')
# ## Conclusions
# They all work poorly compared to looking at individual frames. RIP.
# |Network | Pretrained | Discriminitive | Final Accuracy % | Peak Accuracy %| Time for 1 Epoch (s) |
# |----------------|----------------|----------------|------------------|----------------|----------------------|
# |`resnet18-B` | True | False | 67.5 | 68.5 | 36 |
# |`resnet34-B` | True | False | 66.0 | 69.0 | 50 |
# |`resnet50-A` | True | False | **68.5** | **71.0** | 47 |
| face_detection/11_Compare3DConvNets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Generate intial word embedding for headlines and description
#
# 为标题和说明生成初始字嵌入
# The embedding is limited to a fixed vocabulary size (`vocab_size`) but a vocabulary of all the words that appeared in the data is built.
#
# 嵌入仅限于固定的词汇大小(`vocab_size`),但构建了数据中出现的所有单词的词汇表。
FN = 'vocabulary-embedding'
seed=42
vocab_size = 40000
embedding_dim = 100
lower = False # dont lower case the text
# # read tokenized headlines and descriptions
import cPickle as pickle
FN0 = 'tokens' # this is the name of the data file which I assume you already have
with open('data/%s.pkl'%FN0, 'rb') as fp:
heads, desc, keywords = pickle.load(fp) # keywords are not used in this project
if lower:
heads = [h.lower() for h in heads]
if lower:
desc = [h.lower() for h in desc]
i=0
heads[i]
desc[i]
keywords[i]
len(heads),len(set(heads))
len(desc),len(set(desc))
# # build vocabulary
from collections import Counter
from itertools import chain
def get_vocab(lst):
vocabcount = Counter(w for txt in lst for w in txt.split())
vocab = map(lambda x: x[0], sorted(vocabcount.items(), key=lambda x: -x[1]))
return vocab, vocabcount
vocab, vocabcount = get_vocab(heads+desc)
# most popular tokens
print vocab[:50]
print '...',len(vocab)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot([vocabcount[w] for w in vocab]);
plt.gca().set_xscale("log", nonposx='clip')
plt.gca().set_yscale("log", nonposy='clip')
plt.title('word distribution in headlines and discription')
plt.xlabel('rank')
plt.ylabel('total appearances');
# always nice to see [Zipf's law](https://en.wikipedia.org/wiki/Zipf%27s_law)
# # Index words
empty = 0 # RNN mask of no data
eos = 1 # end of sentence
start_idx = eos+1 # first real word
def get_idx(vocab, vocabcount):
word2idx = dict((word, idx+start_idx) for idx,word in enumerate(vocab))
word2idx['<empty>'] = empty
word2idx['<eos>'] = eos
idx2word = dict((idx,word) for word,idx in word2idx.iteritems())
return word2idx, idx2word
word2idx, idx2word = get_idx(vocab, vocabcount)
# # Word Embedding
# ## read GloVe
fname = 'glove.6B.%dd.txt'%embedding_dim
import os
datadir_base = os.path.expanduser(os.path.join('~', '.keras'))
if not os.access(datadir_base, os.W_OK):
datadir_base = os.path.join('/tmp', '.keras')
datadir = os.path.join(datadir_base, 'datasets')
glove_name = os.path.join(datadir, fname)
if not os.path.exists(glove_name):
path = 'glove.6B.zip'
path = get_file(path, origin="http://nlp.stanford.edu/data/glove.6B.zip")
# !unzip {datadir}/{path}
# glove_n_symbols = !wc -l {glove_name}
glove_n_symbols = int(glove_n_symbols[0].split()[0])
glove_n_symbols
glove_index_dict = {}
glove_embedding_weights = np.empty((glove_n_symbols, embedding_dim))
globale_scale=.1
with open(glove_name, 'r') as fp:
i = 0
for l in fp:
l = l.strip().split()
w = l[0]
glove_index_dict[w] = i
glove_embedding_weights[i,:] = map(float,l[1:])
i += 1
glove_embedding_weights *= globale_scale
glove_embedding_weights.std()
for w,i in glove_index_dict.iteritems():
w = w.lower()
if w not in glove_index_dict:
glove_index_dict[w] = i
# ## embedding matrix
# use GloVe to initialize embedding matrix
# +
import numpy as np
# generate random embedding with same scale as glove
np.random.seed(seed)
shape = (vocab_size, embedding_dim)
scale = glove_embedding_weights.std()*np.sqrt(12)/2 # uniform and not normal
embedding = np.random.uniform(low=-scale, high=scale, size=shape)
print 'random-embedding/glove scale', scale, 'std', embedding.std()
# # copy from glove weights of words that appear in our short vocabulary (idx2word)
c = 0
for i in range(vocab_size):
w = idx2word[i]
g = glove_index_dict.get(w, glove_index_dict.get(w.lower()))
if g is None and w.startswith('#'): # glove has no hastags (I think...)
w = w[1:]
g = glove_index_dict.get(w, glove_index_dict.get(w.lower()))
if g is not None:
embedding[i,:] = glove_embedding_weights[g,:]
c+=1
print 'number of tokens, in small vocab, found in glove and copied to embedding', c,c/float(vocab_size)
# -
# lots of word in the full vocabulary (word2idx) are outside `vocab_size`.
# Build an alterantive which will map them to their closest match in glove but only if the match
# is good enough (cos distance above `glove_thr`)
glove_thr = 0.5
word2glove = {}
for w in word2idx:
if w in glove_index_dict:
g = w
elif w.lower() in glove_index_dict:
g = w.lower()
elif w.startswith('#') and w[1:] in glove_index_dict:
g = w[1:]
elif w.startswith('#') and w[1:].lower() in glove_index_dict:
g = w[1:].lower()
else:
continue
word2glove[w] = g
# for every word outside the embedding matrix find the closest word inside the mebedding matrix.
# Use cos distance of GloVe vectors.
#
# Allow for the last `nb_unknown_words` words inside the embedding matrix to be considered to be outside.
# Dont accept distances below `glove_thr`
# +
normed_embedding = embedding/np.array([np.sqrt(np.dot(gweight,gweight)) for gweight in embedding])[:,None]
nb_unknown_words = 100
glove_match = []
for w,idx in word2idx.iteritems():
if idx >= vocab_size-nb_unknown_words and w.isalpha() and w in word2glove:
gidx = glove_index_dict[word2glove[w]]
gweight = glove_embedding_weights[gidx,:].copy()
# find row in embedding that has the highest cos score with gweight
gweight /= np.sqrt(np.dot(gweight,gweight))
score = np.dot(normed_embedding[:vocab_size-nb_unknown_words], gweight)
while True:
embedding_idx = score.argmax()
s = score[embedding_idx]
if s < glove_thr:
break
if idx2word[embedding_idx] in word2glove :
glove_match.append((w, embedding_idx, s))
break
score[embedding_idx] = -1
glove_match.sort(key = lambda x: -x[2])
print '# of glove substitutes found', len(glove_match)
# -
# manually check that the worst substitutions we are going to do are good enough
for orig, sub, score in glove_match[-10:]:
print score, orig,'=>', idx2word[sub]
# build a lookup table of index of outside words to index of inside words
glove_idx2idx = dict((word2idx[w],embedding_idx) for w, embedding_idx, _ in glove_match)
# # Data
Y = [[word2idx[token] for token in headline.split()] for headline in heads]
len(Y)
plt.hist(map(len,Y),bins=50);
X = [[word2idx[token] for token in d.split()] for d in desc]
len(X)
plt.hist(map(len,X),bins=50);
import cPickle as pickle
with open('data/%s.pkl'%FN,'wb') as fp:
pickle.dump((embedding, idx2word, word2idx, glove_idx2idx),fp,-1)
import cPickle as pickle
with open('data/%s.data.pkl'%FN,'wb') as fp:
pickle.dump((X,Y),fp,-1)
| vocabulary-embedding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load the Dataset
# These datasets can be found at: https://support.10xgenomics.com/single-cell-gene-expression/datasets under the heading **Single Cell 3' Paper: Zheng et al. 2017**. Please replace ``path_prefix`` to the location where you have downloaded the files.
# +
import numpy as np, h5py, os
import matplotlib.pyplot as plt
from operator import itemgetter
from scipy.sparse import vstack, coo_matrix, csc_matrix, isspmatrix_csc
# %matplotlib inline
fnames = ['293t', #0
'aml027_post', #1
'aml027_pre', #2
'aml035_post', #3
'aml035_pre', #4
'b', #5
'bmmc_healthy_1', #6
'bmmc_healthy_2', #7
]
path_prefix = 'C:/Users/aabid/OneDrive/' #Replace with your own path
path_suffix = '/filtered_matrices_mex/hg19/matrix.mtx'
# -
# For faster access, save all of the files as h5py files.
# +
def gen_path(fname):
return path_prefix + fname + path_suffix
for fname in fnames:
if not(os.path.isfile(path_prefix+fname+'.h5')): # returns whether the h5 file exists or not
data = np.genfromtxt(gen_path(fname),delimiter=' ',skip_header=3,filling_values=0)
row = data[:,1]-1 #1-indexed
col = data[:,0]-1 #1-indexed
values = data[:,2]
print('Filename read:',fname)
with h5py.File(path_prefix+fname+'.h5', 'w') as hf:
hf.create_dataset("filtered_matrix", data=data)
print('Filename written:',fname)
# -
# # Preprocess to Reduce Num of Genes
# This is a helper class that loads the files, preprocesses them, and then separates them into target and background datasets. This will be useful, since we are running PCA and cPCA, on multiple sets of target files. The Dataset class contains methods to perform standard and contrastive PCA.
# +
from utils import Dataset
# %matplotlib inline
class SingleCell(Dataset):
def __init__(self, active_files, background_file, N_GENES = 500, to_standardize=True, verbose=True):
self.active = vstack([self.file_to_features(fname) for fname in active_files])
self.bg = vstack([self.file_to_features(fname) for fname in background_file])
self.reduce_features(N_GENES)
self.data = np.concatenate((self.active, self.bg),axis=0)
self.active_labels = np.concatenate([self.file_to_labels(fname, l) for l, fname in enumerate(active_files)])
# Pre-processing - done in main class
if (verbose):
print("Data size\t\t", self.data.shape)
print("Active dataset size: \t", self.active.shape)
print("Background dataset size:", self.bg.shape)
super(self.__class__, self).__init__(to_standardize=to_standardize)
self.pca_active()
def description():
print("To Add")
def file_to_features(self, fname):
with h5py.File(path_prefix+fname+'.h5', 'r') as hf:
data = hf['filtered_matrix'][:]
row = data[:,1]-1 #1-indexed
col = data[:,0]-1 #1-indexed
values = data[:,2]
c = csc_matrix((values, (row, col)), shape=(row.max()+1, col.max()+1))
return c
def reduce_features(self, N_GENES):
n_active = self.active.shape[0]
n_bg = self.bg.shape[0]
c = vstack((self.active, self.bg), format="csc")
nonzero_idx = np.where(np.amax(c, axis=0).toarray().flatten()>0)[0]
c = c[:,nonzero_idx]
c = c.toarray()
total_dispersion = np.var(c,axis=0)/np.mean(c,axis=0)
ind = np.argpartition(total_dispersion, -N_GENES)[-N_GENES:].flatten()
c = c[:,ind]
self.active = c[:n_active]
self.bg = c[-n_bg:]
def file_to_labels(self, fname, l):
with h5py.File(path_prefix+fname+'.h5', 'r') as hf:
data = hf['filtered_matrix'][:]
row = data[:,1]-1 #1-indexed
col = data[:,0]-1 #1-indexed
values = data[:,2]
c = coo_matrix((values, (row, col)), shape=(row.max()+1, col.max()+1))
c = c.toarray()
num_cells = c.shape[0]
labels = np.repeat([l], num_cells)
return labels
# -
# # Run Standard and Contrastive PCA (2 Groups)
# +
import matplotlib
active_file_idx = [1,2]
dataset = SingleCell(itemgetter(*active_file_idx)(fnames), [fnames[6]])
colors = ['#1f77b4','#d62728', '#2ca02c', '#ff7f0e']
projected_data, alphas = dataset.automated_cpca(max_log_alpha=3)
active_labels = dataset.get_active_labels()
# -
plt.figure(figsize=[28,8])
for j, (fg,bg) in enumerate(projected_data):
plt.subplot(1,4,j+1)
if (j==0):
plt.title('PCA')
plt.xlabel('PC1')
plt.ylabel('PC2')
else:
plt.title('cPCA')
plt.xlabel('cPC1')
plt.ylabel('cPC2')
if (j==1 or j==2):
fg[:,0] = -fg[:,0]
for i, l in enumerate((np.sort(np.unique(active_labels)))):
idx = np.where(active_labels==l)[0]
plt.scatter(fg[idx,0],fg[idx,1], color=colors[i], alpha=0.5, s=25)
plt.title(r'$\alpha=$' +str(np.round(alphas[j],1)))
matplotlib.rcParams.update({'font.size': 36})
plt.locator_params(nbins=4, axis='x')
plt.locator_params(nbins=6, axis='y')
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
if (j==0):
plt.xlim([-100, 25])
plt.ylim([-40, 42])
if (j==2):
plt.xlim([-40, 45])
plt.ylim([-2, 1.75])
plt.tight_layout()
# Note that this code can be extended to more than two, as is done in Fig 3(c) and 3(d) of the paper. Here, for illustration, we run the same analysis with three groups of cells.
# # Run Standard and Contrastive PCA (3 Groups)
# +
import matplotlib
active_file_idx = [1,2,3]
dataset = SingleCell(itemgetter(*active_file_idx)(fnames), [fnames[6]])
colors = ['#1f77b4','#d62728', '#2ca02c', '#ff7f0e']
projected_data, alphas = dataset.automated_cpca(max_log_alpha=3)
active_labels = dataset.get_active_labels()
# -
for j, (fg,bg) in enumerate(projected_data):
plt.figure(figsize=[3.5,3.5])
if (j==0):
plt.title('PCA')
plt.xlabel('PC1')
plt.ylabel('PC2')
else:
plt.title('cPCA')
plt.xlabel('cPC1')
plt.ylabel('cPC2')
for i, l in enumerate(np.sort(np.unique(active_labels))):
idx = np.where(active_labels==l)[0]
plt.scatter(fg[idx,0],fg[idx,1], color=colors[i], alpha=0.5, s=5)
matplotlib.rcParams.update({'font.size': 14})
plt.locator_params(nbins=4, axis='x')
plt.locator_params(nbins=6, axis='y')
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
| experiments/Single-Cell RNA-seq (Figure 3).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# #### High level:
# This notebook shows all the inconsistencies of field that were produced with dictionaries (and have `hebrew` in the name) with their respective numeric values for the `markers_hebrew` table.
#
# The specific analysis below is based on data from `2019-11-16_views_and_main_tables` folder from Nov 16, 2019 that can be found here: https://drive.google.com/drive/folders/1StZkyR7KG_cfPpk8xMj5es3HGkIA00C9?usp=sharing
#
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set()
pd.options.display.max_rows = 200
pd.options.display.max_columns = 100
markers_raw = pd.read_csv('../../views_and_main_tables_2019_11/markers_hebrew.csv')
m_all = markers_raw[markers_raw['accident_year'] < 2019]
m_all.info()
#
# ### Helper functions
def calc_diff_counts_hebrew(data, feat_name):
data = data[(data[feat_name].isnull() == False) & (data[feat_name + '_hebrew'].isnull() == False)]
print(f'Shape of data: {data.shape}')
return data[feat_name].value_counts().reset_index(drop=True) - \
data[feat_name + '_hebrew'].value_counts().reset_index(drop=True)
def merge_with_hebrew(data, feat_name):
nums_df = data[feat_name].value_counts().reset_index()
nums_df.columns = ['index_' + feat_name, 'count']
hebrew_df = data[feat_name + '_hebrew'].value_counts().reset_index()
hebrew_df.columns = ['index_' + feat_name + '_hebrew', 'count']
return pd.merge(nums_df, hebrew_df, how='outer', on='count')
# ### road_shape and road_shape_hebrew
# road_shape 720541 non-null int64
# road_shape_hebrew 720538 non-null object
m_all[(m_all['road_shape'].isnull() == False) & (m_all['road_shape_hebrew'].isnull() == True)]
m_all[(m_all['road_shape']==0) & (m_all['accident_year']==2008)]
m_all[m_all['road_shape']==0]
m_all[(m_all['road_shape']==15) & (m_all['accident_year']==2012)]
m_all[m_all['road_shape']==15]
# **Null Conclusion**: seems that all `road_shape` 0 and 15 don't have translation into `road_shape_hebrew`
# **Specific values mistmatch investigations:**
m_all[(m_all['road_shape']!=0) & (m_all['road_shape']!=15)]['road_shape'].value_counts()
m_all[(m_all['road_shape']!=0) & (m_all['road_shape']!=15)]['road_shape_hebrew'].value_counts()
calc_diff_counts_hebrew(m_all[(m_all['road_shape']!=0) & (m_all['road_shape']!=15)], 'road_shape')
m_all[m_all['road_shape_hebrew'] == 'על גשר מנהרה'].describe()
m_all[m_all['road_shape_hebrew'] == 'על גשר/ בתוך מנהרה'].describe()
# **Specific values conclusions:**
# - Till 2011 (including) `road_shape` 3 was called `על גשר מנהרה` that was later changed to `על גשר/ בתוך מנהרה`
#
# ### one_lane and one_lane_hebrew
# one_lane 720541 non-null int64
# one_lane_hebrew 657395 non-null object
m_all[(m_all['one_lane'].isnull() == False) & (m_all['one_lane_hebrew'].isnull() == True)].describe()
m_all[(m_all['one_lane'].isnull() == False) & (m_all['one_lane_hebrew'].isnull() == True)].shape
m_all[m_all['one_lane'] == 0].shape
# **Null Conclusion:** `one_lane` == 0 of all years is missing
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[m_all['one_lane'] != 0], 'one_lane')
# **Specific values conclusion:** no issues
#
# ### multi_lane and multi_lane_hebrew
# multi_lane 720541 non-null int64
# multi_lane_hebrew 63177 non-null object
m_all[(m_all['multi_lane'].isnull() == False) & (m_all['multi_lane_hebrew'].isnull() == True)].shape
m_all[(m_all['multi_lane'].isnull() == False) & (m_all['multi_lane_hebrew'].isnull() == True)].describe()
m_all[m_all['multi_lane'] == 0].shape
# **Null Conclusion:** `multi_lane` == 0 of all years is missing
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[m_all['multi_lane'] != 0], 'multi_lane')
# **Specific values conclusion:** no issues
#
# ### speed_limit and speed_limit_hebrew
# speed_limit 720541 non-null int64
# speed_limit_hebrew 720540 non-null object
m_all[(m_all['speed_limit'].isnull() == False) & (m_all['speed_limit_hebrew'].isnull() == True)]
m_all[(m_all['speed_limit'].isnull() == False) & (m_all['speed_limit_hebrew'].isnull() == True)]['speed_limit']
m_all['speed_limit'].value_counts()
# **Null Conclusion:** Speed limit 3383 has wrong information
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[m_all['speed_limit'] != 3383], 'speed_limit')
# **Specific values conclusion:** no issues
#
# ### road_control and road_control_hebrew
# road_control 80906 non-null float64
# road_control_hebrew 80904 non-null object
m_all[(m_all['road_control'].isnull() == False) & (m_all['road_control_hebrew'].isnull() == True)]
m_all[m_all['road_control'] == 0]
# **Null Conclusion:** `road_control` == 0 of all years is missing, all in year 2010
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[(m_all['road_control'] != 0) | (m_all['accident_year'] != 2010)], 'road_control')
# **Specific values conclusion:** no issues
#
# ### object_distance and object_distance_hebrew
# object_distance 720541 non-null int64
# object_distance_hebrew 720477 non-null object
m_all[(m_all['object_distance'].isnull() == False) & (m_all['object_distance_hebrew'].isnull() == True)].shape
m_all[(m_all['object_distance'].isnull() == False) & (m_all['object_distance_hebrew'].isnull() == True)].describe()
m_all[(m_all['object_distance'].isnull() == False) & (m_all['object_distance_hebrew'].isnull() == True)]['object_distance'].value_counts()
m_all['object_distance'].value_counts()
m_all[(m_all['object_distance']==5) & (m_all['object_distance'].isnull() == False) & (m_all['object_distance_hebrew'].isnull() == True)].shape
m_all[(m_all['object_distance']==5) & (m_all['object_distance'].isnull() == False) & (m_all['object_distance_hebrew'].isnull() == True)].describe()
m_all[(m_all['object_distance']==5) & ((m_all['accident_year'] == 2012) | (m_all['accident_year'] == 2013) | (m_all['accident_year'] == 2014))].shape
m_all[(m_all['object_distance']==1462) & (m_all['object_distance'].isnull() == False) & (m_all['object_distance_hebrew'].isnull() == True)].shape
# **Null Conclusion:**
# - *all* instances of `object_distance` == 1462
# - *all* instances of `object_distance` == 5 during years 2012-2014. Rest of the years `object_distance` == 5 is OK
# **Specific values mistmatch investigations:**
temp = m_all[(m_all['object_distance'] != 1462) & \
((m_all['object_distance'] !=5) | \
((m_all['accident_year'] != 2012) & (m_all['accident_year'] != 2013) & (m_all['accident_year'] != 2014)))]
calc_diff_counts_hebrew(temp, 'object_distance')
# **Specific values conclusion:** no issues
#
# ### didnt_cross and didnt_cross_hebrew
# didnt_cross 720541 non-null int64
# didnt_cross_hebrew 720538 non-null object
m_all[(m_all['didnt_cross'].isnull() == False) & (m_all['didnt_cross_hebrew'].isnull() == True)]
m_all[m_all['didnt_cross'] == 0]
# **Null Conclusion**: all instances of `didnt_cross` == 0 are missing translation
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[m_all['didnt_cross'] != 0], 'didnt_cross')
# **Specific values conclusion:** no issues
#
# ### cross_mode and cross_mode_hebrew
# cross_mode 720541 non-null int64
# cross_mode_hebrew 33365 non-null object
m_all[(m_all['cross_mode'].isnull() == False) & (m_all['cross_mode_hebrew'].isnull() == True)].shape
m_all[(m_all['cross_mode'].isnull() == False) & (m_all['cross_mode_hebrew'].isnull() == True)].describe()
m_all[(m_all['cross_mode'].isnull() == False) & (m_all['cross_mode_hebrew'].isnull() == True)]['cross_mode'].value_counts()
m_all[m_all['cross_mode']==0].shape
m_all[m_all['cross_mode']==0]['accident_year'].value_counts()
m_all[m_all['cross_mode']==9].shape
m_all[(m_all['cross_mode']==9) & (m_all['cross_mode'].isnull() == False) & (m_all['cross_mode_hebrew'].isnull() == True)]
m_all[m_all['cross_mode']==9]['accident_year'].value_counts()
# **Null Conclusions**:
# - All `cross_mode` == 0 that's spread over all years, translation is missing
# - `cross_mode` == 9 from year 2008 is problematic (other years are OK)
# **Specific values mistmatch investigations:**
temp = m_all[(m_all['cross_mode'] !=0) & ((m_all['cross_mode'] != 9) | (m_all['accident_year'] != 2008))]
calc_diff_counts_hebrew(temp, 'cross_mode')
# **Specific values conclusion:** no issues
#
# ### cross_location and cross_location_hebrew
# cross_location 720541 non-null int64
# cross_location_hebrew 28925 non-null object
m_all[(m_all['cross_location'].isnull() == False) & (m_all['cross_location_hebrew'].isnull() == True)].shape
m_all[(m_all['cross_location'].isnull() == False) & (m_all['cross_location_hebrew'].isnull() == True)].describe()
m_all[m_all['cross_location']==0].shape
# **Null Conclusion:** all `cross_location` == 0 spread over numerous years has no translation
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[m_all['cross_location'] != 0], 'cross_location')
# **Specific values conclusion:** no issues
#
# ### region and region_hebrew
# region 605597 non-null float64
# region_hebrew 565389 non-null object
m_all[(m_all['region'].isnull() == False) & (m_all['region_hebrew'].isnull() == True)].shape
m_all[(m_all['region'].isnull() == False) & (m_all['region_hebrew'].isnull() == True)].describe()
m_all[(m_all['region'] == 9].shape
m_all[(m_all['region'] == 9) & (m_all['region_hebrew'].isnull())]['accident_year'].value_counts()
m_all[(m_all['region'] == 9) & (m_all['region_hebrew'].isnull() == False)]['accident_year'].value_counts()
m_all[(m_all['region'] == 9) & (m_all['accident_year'] != 2018)].shape
# **Null Conclusion:** all `region` == 9 spread over numerous years missing translation, 2018 it was already fixed
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[(m_all['region'] != 9) | (m_all['accident_year'] == 2018)], 'region')
# **Specific values conclusion:** no issues
#
# ### district and district_hebrew
# district 720541 non-null int64
# district_hebrew 561538 non-null object
m_all[(m_all['district'].isnull() == False) & (m_all['district_hebrew'].isnull() == True)].shape
m_all[(m_all['district'].isnull() == False) & (m_all['district_hebrew'].isnull() == True)].describe()
m_all[m_all['district']==99].shape
# **Null Conclusion:** all `district` == 99 spread over numerous years missing translation
# **Specific values mistmatch investigations:**
calc_diff_counts_hebrew(m_all[m_all['district'] != 99], 'district')
# **Specific values conclusion:** no issues
#
# ### natural_area and natural_area_hebrew
# natural_area 605597 non-null float64
# natural_area_hebrew 557164 non-null object
m_all[(m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)].shape
m_all[(m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)].describe()
m_all[(m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)]['natural_area'].value_counts()
for area in [999, 432, 777, 877]:
print(f'{area} shape: {m_all[m_all["natural_area"]==area].shape}')
m_all[(m_all['natural_area']==432) & (m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)].describe()
m_all[(m_all['natural_area']==432) & ((m_all['accident_year']==2013) | (m_all['accident_year']==2014))].shape
m_all[m_all['natural_area']==432]['accident_year'].value_counts()
m_all[(m_all['natural_area']==999) & (m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)].describe()
m_all[(m_all['natural_area']==777) & (m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)].describe()
m_all[(m_all['natural_area']==877) & (m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == True)]
# **Null Conclusion:** Following `natural_area` had translation issues:
# - 999 - all values missing, spread over various years
# - 777 - all values missing, spread over various years
# - 877 - all values (it's actually 1 value in 2017).
# - 432 - years 2013, 2014 are problematic, rest of the years are OK
# **Specific values mistmatch investigations:**
temp = m_all[(m_all['natural_area'].isnull() == False) & (m_all['natural_area_hebrew'].isnull() == False)]
calc_diff_counts_hebrew(temp, 'natural_area')
merge_with_hebrew(temp, 'natural_area')
temp[temp['natural_area_hebrew'] == 'אזור לוד'].shape
temp[temp['natural_area'] == 431].shape
temp[temp['natural_area'] == 431].shape[0] - temp[temp['natural_area_hebrew'] == 'אזור לוד'].shape[0]
temp[temp['natural_area_hebrew'] == 'אזור מודיעין'].describe()
temp[temp['natural_area_hebrew'] == 'אזור לוד'].describe()
# **Specific values conclusion:**
# Seems that `natural_area` 431 was called `אזור לוד` until and including 2014, and `אזור מודיעין` since
#
# ### municipal_status and municipal_status_hebrew
# municipal_status 604357 non-null float64
# municipal_status_hebrew 604314 non-null object
m_all[(m_all['municipal_status'].isnull() == False) & (m_all['municipal_status_hebrew'].isnull() == True)].shape
m_all[(m_all['municipal_status'].isnull() == False) & (m_all['municipal_status_hebrew'].isnull() == True)].describe()
m_all[m_all['municipal_status'] == 999].shape
# **Null Conclusion:** `municipal_status` == 999 is problematic, spread over numerous years
# **Specific values mistmatch investigations:**
# +
temp = m_all[m_all['municipal_status_hebrew'].isnull()==False]
merge_with_hebrew(temp, 'municipal_status')
# -
temp[temp['municipal_status_hebrew'] == 'בקעת בית שאן']['accident_year'].describe()
temp[temp['municipal_status_hebrew'] == 'עמק המעיינות']['accident_year'].describe()
# **Specific values conclusion:**
# `municipal_status` 7 was called `בקעת בית שאן` until and including 2014, and `עמק המעיינות` since
#
# ## yishuv_shape and yishuv_shape_hebrew
# yishuv_shape 605597 non-null float64
# yishuv_shape_hebrew 561532 non-null object
m_all[(m_all['yishuv_shape'].isnull() == False) & (m_all['yishuv_shape_hebrew'].isnull() == True)].shape
m_all[(m_all['yishuv_shape'].isnull() == False) & (m_all['yishuv_shape_hebrew'].isnull() == True)].describe()
m_all[(m_all['yishuv_shape'].isnull() == False) & (m_all['yishuv_shape_hebrew'].isnull() == True)]['yishuv_shape'].value_counts()
m_all[m_all['yishuv_shape']==99].shape
m_all[(m_all['yishuv_shape']==99) & (m_all['yishuv_shape'].isnull() == False) & (m_all['yishuv_shape_hebrew'].isnull() == True)]['accident_year'].value_counts()
m_all[m_all['yishuv_shape']==53].shape
m_all[(m_all['yishuv_shape']==53) & (m_all['yishuv_shape'].isnull() == False) & (m_all['yishuv_shape_hebrew'].isnull() == True)]['accident_year'].value_counts()
m_all[m_all['yishuv_shape']==53]['accident_year'].value_counts()
# **Null Conclusion:** `yishuv_shape` translation issues:
# - 99 spread over numerous years
# - 53 for years 2008-2010, rest of the years there is 53 without problems
# **Specific values mistmatch investigations:**
temp = m_all[(m_all['yishuv_shape'].isnull() == False) & (m_all['yishuv_shape_hebrew'].isnull() == False)]
temp.shape
merge_with_hebrew(temp, 'yishuv_shape')
calc_diff_counts_hebrew(temp, 'yishuv_shape')
# **Specific values conclusion:** a big mess, worth investing more time to understand
#
# ### street1 and street1_hebrew
# street1 553240 non-null float64
# street1_hebrew 389936 non-null object
m_all[(m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)].shape
m_all[(m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)].describe()
m_all['street1'].nunique()
m_all[(m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)]['street1'].value_counts()
m_all[(m_all['street1'] != 0) & (m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)].shape
num_total = []
for street in m_all[(m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)]['street1'].value_counts().index:
num_total.append(m_all[m_all['street1'] == street].shape[0])
num_total - m_all[(m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)]['street1'].value_counts()
temp = m_all[(m_all['street1'].isnull()==False) & (m_all['street1_hebrew'].isnull()==False)]
temp.shape
m_all[(m_all['street1'] != 0) & (m_all['street1'].isnull() == False) & (m_all['street1_hebrew'].isnull() == True)]['street1'].value_counts()
# **Null Conclusion:** 160902 out of 163304 that are missing are `street1` == 0, other 2402 miscellenious.
# Some of the miscellenious all the index translation is missing, some only part of the instances translation missing.
# See details above.
# **Specific values mistmatch investigations:**
# Didn't do detailed investigation, especially due to all the differences already in the nulls
# **Specific values conclusion:**
# See above
#
# ## street2 and street2_hebrew
# street2 71838 non-null float64
# street2_hebrew 70713 non-null object
m_all[(m_all['street2'].isnull() == False) & (m_all['street2_hebrew'].isnull() == True)].shape
m_all[(m_all['street2'].isnull() == False) & (m_all['street2_hebrew'].isnull() == True)]['street2'].value_counts()
num_total = []
for street in m_all[(m_all['street2'].isnull() == False) & (m_all['street2_hebrew'].isnull() == True)]['street2'].value_counts().index:
num_total.append(m_all[m_all['street2'] == street].shape[0])
num_total - m_all[(m_all['street2'].isnull() == False) & (m_all['street2_hebrew'].isnull() == True)]['street2'].value_counts()
# **Null Conclusion:** 1092 out of 1125 that are missing are `street2` == 0, other 33 miscellenious.
# Some of the miscellenious all the index translation is missing, some only part of the instances translation missing.
# See details above.
# **Specific values mistmatch investigations:** see conclusions for `street1` above
# **Specific values conclusion:** see conclusions for `street1` above
#
# ## non_urban_intersection and non_urban_intersection_hebrew
# non_urban_intersection 34382 non-null float64
# non_urban_intersection_hebrew 33018 non-null object
m_all[(m_all['non_urban_intersection'].isnull() == False) & (m_all['non_urban_intersection_hebrew'].isnull() == True)].shape
m_all['non_urban_intersection'].nunique()
m_all[(m_all['non_urban_intersection'].isnull() == False) & (m_all['non_urban_intersection_hebrew'].isnull() == True)].describe()
m_all[(m_all['non_urban_intersection'].isnull() == False) & (m_all['non_urban_intersection_hebrew'].isnull() == True)]['non_urban_intersection'].value_counts()
num_total = []
for inter in m_all[(m_all['non_urban_intersection'].isnull() == False) & (m_all['non_urban_intersection_hebrew'].isnull() == True)]['non_urban_intersection'].value_counts().index:
num_total.append(m_all[m_all['non_urban_intersection'] == inter].shape[0])
num_total - m_all[(m_all['non_urban_intersection'].isnull() == False) & (m_all['non_urban_intersection_hebrew'].isnull() == True)]['non_urban_intersection'].value_counts()
# **Null Conclusion:** A lot of `non_urban_intersection` are missing, most of them only part of the instances translation missing. See details above.
# **Specific values mistmatch investigations:** because of all the issues above, is it worth to investigate?
# **Specific values conclusion:** because of all the issues above, not investigated
#
# ## accident_hour_raw and accident_hour_raw_hebrew
# accident_hour_raw 720541 non-null int64
# accident_hour_raw_hebrew 322739 non-null object
m_all[(m_all['accident_hour_raw'].isnull() == False) & (m_all['accident_hour_raw_hebrew'].isnull() == True)].shape
m_all[(m_all['accident_hour_raw'].isnull() == False) & (m_all['accident_hour_raw_hebrew'].isnull() == True)].describe()
m_all[(m_all['accident_hour_raw'].isnull() == False) & (m_all['accident_hour_raw_hebrew'].isnull() == True)]['accident_hour_raw'].value_counts()
# **Null Conclusion:** Half the values of missing, a lot of different views. See details above
# **Specific values mistmatch investigations:** not investigated because of all the issues above
# **Specific values conclusion:** not investigated because of all the issues above
| analysis_notebooks/markers_hebrew_mismatches.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Think Bayes: Chapter 3
#
# This notebook presents example code and exercise solutions for Think Bayes.
#
# Copyright 2016 <NAME>
#
# MIT License: https://opensource.org/licenses/MIT
# +
from __future__ import print_function, division
% matplotlib inline
import thinkplot
from thinkbayes2 import Hist, Pmf, Suite, Cdf
# -
# ## The Dice problem
#
# Suppose I have a box of dice that contains a 4-sided die, a 6-sided
# die, an 8-sided die, a 12-sided die, and a 20-sided die.
#
# Suppose I select a die from the box at random, roll it, and get a 6.
# What is the probability that I rolled each die?
#
# The `Dice` class inherits `Update` and provides `Likelihood`
class Dice(Suite):
def Likelihood(self, data, hypo):
if hypo < data:
return 0
else:
return 1/hypo
# Here's what the update looks like:
suite = Dice([4, 6, 8, 12, 20])
suite.Update(6)
suite.Print()
# And here's what it looks like after more data:
# +
for roll in [6, 8, 7, 7, 5, 4]:
suite.Update(roll)
suite.Print()
# -
# ## The train problem
#
# The Train problem has the same likelihood as the Dice problem.
class Train(Suite):
def Likelihood(self, data, hypo):
if hypo < data:
return 0
else:
return 1/hypo
# But there are many more hypotheses
hypos = xrange(1, 1001)
suite = Train(hypos)
suite.Update(60)
# Here's what the posterior looks like
thinkplot.Pdf(suite)
# And here's how we can compute the posterior mean
# +
def Mean(suite):
total = 0
for hypo, prob in suite.Items():
total += hypo * prob
return total
Mean(suite)
# -
# Or we can just use the method
suite.Mean()
# ## Sensitivity to the prior
#
# Here's a function that solves the train problem for different priors and data
def MakePosterior(high, dataset, constructor=Train):
"""Solves the train problem.
high: int maximum number of trains
dataset: sequence of observed train numbers
constructor: function used to construct the Train object
returns: Train object representing the posterior suite
"""
hypos = range(1, high+1)
suite = constructor(hypos)
for data in dataset:
suite.Update(data)
return suite
# Let's run it with the same dataset and several uniform priors
# +
dataset = [30, 60, 90]
for high in [500, 1000, 2000]:
suite = MakePosterior(high, dataset)
print(high, suite.Mean())
# -
# The results are quite sensitive to the prior, even with several observations.
# ## Power law prior
#
# Now let's try it with a power law prior.
class Train2(Train):
def __init__(self, hypos, alpha=1.0):
Pmf.__init__(self)
for hypo in hypos:
self[hypo] = hypo**(-alpha)
self.Normalize()
# Here's what a power law prior looks like, compared to a uniform prior
high = 100
hypos = range(1, high+1)
suite1 = Train(hypos)
suite2 = Train2(hypos)
thinkplot.Pdf(suite1)
thinkplot.Pdf(suite2)
# Now let's see what the posteriors look like after observing one train.
# +
dataset = [60]
high = 1000
thinkplot.PrePlot(num=2)
constructors = [Train, Train2]
labels = ['uniform', 'power law']
for constructor, label in zip(constructors, labels):
suite = MakePosterior(high, dataset, constructor)
suite.label = label
thinkplot.Pmf(suite)
thinkplot.Config(xlabel='Number of trains',
ylabel='Probability')
# -
# The power law gives less prior probability to high values, which yields lower posterior means, and less sensitivity to the upper bound.
# +
dataset = [30, 60, 90]
for high in [500, 1000, 2000]:
suite = MakePosterior(high, dataset, Train2)
print(high, suite.Mean())
# -
# ## Credible intervals
#
# To compute credible intervals, we can use the `Percentile` method on the posterior.
# +
hypos = xrange(1, 1001)
suite = Train(hypos)
suite.Update(60)
suite.Percentile(5), suite.Percentile(95)
# -
# If you have to compute more than a few percentiles, it is more efficient to compute a CDF.
#
# Also, a CDF can be a better way to visualize distributions.
cdf = Cdf(suite)
thinkplot.Cdf(cdf)
thinkplot.Config(xlabel='Number of trains',
ylabel='Cumulative Probability',
legend=False)
# `Cdf` also provides `Percentile`
cdf.Percentile(5), cdf.Percentile(95)
# ## Exercises
# **Exercise:** To write a likelihood function for the locomotive problem, we had
# to answer this question: "If the railroad has `N` locomotives, what
# is the probability that we see number 60?"
#
# The answer depends on what sampling process we use when we observe the
# locomotive. In this chapter, I resolved the ambiguity by specifying
# that there is only one train-operating company (or only one that we
# care about).
#
# But suppose instead that there are many companies with different
# numbers of trains. And suppose that you are equally likely to see any
# train operated by any company.
# In that case, the likelihood function is different because you
# are more likely to see a train operated by a large company.
#
# As an exercise, implement the likelihood function for this variation
# of the locomotive problem, and compare the results.
# +
class Train(Suite):
def Likelihood(self,data, hypo):
if data>hypo:
likelihood = 0
else:
likelihood = hypo * (1/hypo)
return likelihood
primer = {i:1/i for i in range(1, 1001)}
pmf = Pmf(primer)
train = Train(pmf)
# -
train.Update(60)
thinkplot.Pdf(train)
# **Exercise:** Suppose I capture and tag 10 rock hyraxes. Some time later, I capture another 10 hyraxes and find that two of them are already tagged. How many hyraxes are there in this environment?
#
# As always with problems like this, we have to make some modeling assumptions.
#
# 1) For simplicity, you can assume that the environment is reasonably isolated, so the number of hyraxes does not change between observations.
#
# 2) And you can assume that each hyrax is equally likely to be captured during each phase of the experiment, regardless of whether it has been tagged. In reality, it is possible that tagged animals would avoid traps in the future, or possible that the same behavior that got them caught the first time makes them more likely to be caught again. But let's start simple.
#
# I suggest the following notation:
#
# * `N`: total population of hyraxes
# * `K`: number of hyraxes tagged in the first round
# * `n`: number of hyraxes caught in the second round
# * `k`: number of hyraxes in the second round that had been tagged
#
# So `N` is the hypothesis and `(K, n, k)` make up the data. The probability of the data, given the hypothesis, is the probability of finding `k` tagged hyraxes out of `n` if (in the population) `K` out of `N` are tagged.
#
# If you are familiar with the hypergeometric distribution, you can use the hypergeometric PMF to compute the likelihood function. Otherwise, you can figure it out using combinatorics.
# +
# Solution goes here
from itertools import combinations
from scipy.special import binom
class Hyrax(Suite):
def Likelihood(self, data, hypo):
N = hypo
K, n, k = data
if N >= K >= k and N >= n >= k:
likelihood = binom(N-K, n-k) / binom(N, n)
else:
likelihood = 0
return likelihood
# -
# Solution goes here
hyrax = Hyrax(range(1,1000))
thinkplot.Pdf(hyrax)
# Solution goes here
hyrax.Update((10,10,2))
thinkplot.Pdf(hyrax)
# Solution goes here
# 后验概率平均值
# 最大后验概率估计
# 90置信区间估计
print(hyrax.Mean())
print(hyrax.MaximumLikelihood())
print(hyrax.CredibleInterval(90))
# +
# Solution goes here
# -
| code/chap03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>Quiz 1 : Hello World to Python</h2>
#
# Print Say Hello World to Python
print("Hello World Python")
# Expected Output :
#
# Hello World Python
# <h2>Quiz 2 : Aritmatika di Python </h2>
#
# - Buat statement pertambahan antara 2 number di Python
# - Buat statement perkurangan antara 2 number di Python
# - Buat statement perkalian antara 2 number di Python
# - Buat statement pembagian antara 2 number di Python
jml = 1 + 2
print(jml)
krg = 3 - 4
print(krg)
kli = 5*6
print(kli)
div = 8/4
print(div)
# <h2>Quiz 3 : Assign Variable dan Tipe Data Integer, Float </h2>
#
# - Buat suatu variabel a dan b, dimana a dan b adalah nilai bertipe data numeric
# - Berikan suatu nilai bertipe data integer, hasil pembagian dari a dengan b
# - Berikan suatu nilai bertipe data float, hasil pembagian dari a dengan b
a=15
b=4
jumlahBagiInt= int(a/b)
print(jumlahBagiInt)
jumlahBagiFloat = (a/b)
print(jumlahBagiFloat)
# <h2>Quiz 4 : String Operation </h2>
#
# - masukan nama depan kamu kedalam suatu variable firstname
# - masukan nama belakang kamu kedalam suatu variable lasname
# - tampilkan suatu kalimat 'Hello sanbercode, saya firstname lastname! saya siap belajar python data science.'
firstName = "Ardana"
lastName = "Rizky"
print("hello sanbercode, saya",firstName,lastName+"! saya siap belajar python data science")
# Expected Output :
#
# Hello sanbercode, saya fauzan taufik! saya siap belajar python data science.
# <h2>Quiz 5 : Tipe Data</h2>
#
# Lengkapi code di bawah ini untuk menghasilkan output yang sesuai
p = 9.99999
q = 'the number : '
print(q + str(p))
# Expected Output :
#
# the number : 9.99999
| JupyterNote Data Science/Tugas Day 1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
os.add_dll_directory(os.path.join(os.environ['CUDA_PATH'], 'bin'))
import cv2
import numpy as np
net = cv2.dnn.readNetFromONNX('../../helper/dat/vgg_model_keras_weights.onnx')
blob[0].shape
# +
vc = cv2.VideoCapture(0);
while True:
ret, img = vc.read();
h, w = img.shape[:2];
#shape=(None, 224, 224, 3)
blob = cv2.dnn.blobFromImage(cv2.resize(img, (224, 224)), 1.0, (224, 224), (104.0, 177.0, 123.0))
net.setInput(blob[:2])
faces = net.forward()
for i in range(faces.shape[2]):
confidence = faces[0, 0, i, 2]
if confidence > 0.72:
box = faces[0, 0, i, 3:7] * np.array([w, h, w, h])
(x, y, x1, y1) = box.astype("int")
cv2.rectangle(img, (x, y), (x1, y1), (0, 0, 255), 2)
# Oluşan çerçeveyi ekrana yansıt
cv2.imshow('Video', img)
# Çıkış için 'q'
if cv2.waitKey(1) & 0xFF == ord('q'):
break
#end
#end
#Kamerayı kapat
vc.release();
cv2.destroyAllWindows();
# -
vc.release()
cv2.destroyAllWindows()
| Face/onnx/faceD__cv2_VGG_ONNX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
from pyvista import set_plot_theme
set_plot_theme('document')
# Colormap Choices
# ================
#
# Use a Matplotlib, Colorcet, cmocean, or custom colormap when plotting
# scalar values.
#
from pyvista import examples
import pyvista as pv
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
import numpy as np
# Any colormap built for `matplotlib`, `colorcet`, or `cmocean` is fully
# compatible with PyVista. Colormaps are typically specified by passing
# the string name of the colormap to the plotting routine via the `cmap`
# argument.
#
# See [Matplotlib\'s complete list of available
# colormaps](https://matplotlib.org/tutorials/colors/colormaps.html),
# [Colorcet\'s complete
# list](https://colorcet.holoviz.org/user_guide/index.html), and
# [cmocean\'s complete list](https://matplotlib.org/cmocean/).
#
# Custom Made Colormaps
# =====================
#
# To get started using a custom colormap, download some data with scalar
# values to plot.
#
mesh = examples.download_st_helens().warp_by_scalar()
# Add scalar array with range (0, 100) that correlates with elevation
mesh['values'] = pv.plotting.normalize(mesh['Elevation']) * 100
# Build a custom colormap - here we make a colormap with 5 discrete colors
# and we specify the ranges where those colors fall:
#
# +
# Define the colors we want to use
blue = np.array([12/256, 238/256, 246/256, 1])
black = np.array([11/256, 11/256, 11/256, 1])
grey = np.array([189/256, 189/256, 189/256, 1])
yellow = np.array([255/256, 247/256, 0/256, 1])
red = np.array([1, 0, 0, 1])
mapping = np.linspace(mesh['values'].min(), mesh['values'].max(), 256)
newcolors = np.empty((256, 4))
newcolors[mapping >= 80] = red
newcolors[mapping < 80] = grey
newcolors[mapping < 55] = yellow
newcolors[mapping < 30] = blue
newcolors[mapping < 1] = black
# Make the colormap from the listed colors
my_colormap = ListedColormap(newcolors)
# -
# Simply pass the colormap to the plotting routine!
#
mesh.plot(scalars='values', cmap=my_colormap)
# Or you could make a simple colormap\... any Matplotlib colormap can be
# passed to PyVista!
#
boring_cmap = plt.cm.get_cmap("viridis", 5)
mesh.plot(scalars='values', cmap=boring_cmap)
# You can also pass a list of color strings to the color map. This
# approach divides up the colormap into 5 equal parts.
#
mesh.plot(scalars=mesh['values'], cmap=['black', 'blue', 'yellow', 'grey', 'red'])
# If you still wish to have control of the separation of values, you can
# do this by creating a scalar array and passing that to the plotter along
# with the the colormap
#
# +
scalars = np.empty(mesh.n_points)
scalars[mesh['values'] >= 80] = 4 # red
scalars[mesh['values'] < 80] = 3 # grey
scalars[mesh['values'] < 55] = 2 # yellow
scalars[mesh['values'] < 30] = 1 # blue
scalars[mesh['values'] < 1] = 0 # black
mesh.plot(scalars=scalars, cmap=['black', 'blue', 'yellow', 'grey', 'red'])
# -
# Matplotlib vs. Colorcet
# =======================
#
# Let\'s compare Colorcet\'s perceptually uniform \"fire\" colormap to
# Matplotlib\'s \"hot\" colormap much like the example on the [first page
# of Colorcet\'s docs](https://colorcet.holoviz.org/index.html).
#
# The \"hot\" version washes out detail at the high end, as if the image
# is overexposed, while \"fire\" makes detail visible throughout the data
# range.
#
# Please note that in order to use Colorcet\'s colormaps including
# \"fire\", you must have Colorcet installed in your Python environment:
# `pip install colorcet`
#
# +
p = pv.Plotter(shape=(2, 2), border=False)
p.subplot(0, 0)
p.add_mesh(mesh, scalars='Elevation', cmap="fire",
lighting=True, scalar_bar_args={'title': "Colorcet Fire"})
p.subplot(0, 1)
p.add_mesh(mesh, scalars='Elevation', cmap="fire",
lighting=False, scalar_bar_args={'title': "Colorcet Fire (No Lighting)"})
p.subplot(1, 0)
p.add_mesh(mesh, scalars='Elevation', cmap="hot",
lighting=True, scalar_bar_args={'title': "Matplotlib Hot"})
p.subplot(1, 1)
p.add_mesh(mesh, scalars='Elevation', cmap="hot",
lighting=False, scalar_bar_args={'title': "Matplotlib Hot (No Lighting)"})
p.show()
| examples/02-plot/cmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # MadMiner particle physics tutorial
#
# # Part 3c: Training a likelihood estimator
#
# <NAME>, <NAME>, <NAME>, and <NAME> 2018-2019
# In part 3c of this tutorial we will train a third neural estimator: this time of the likelihood function itself (rather than its ratio). We assume that you have run part 1 and 2a of this tutorial. If, instead of 2a, you have run part 2b, you just have to load a different filename later.
# ## Preparations
# +
from __future__ import absolute_import, division, print_function, unicode_literals
import logging
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
# %matplotlib inline
from madminer.sampling import SampleAugmenter
from madminer import sampling
from madminer.ml import LikelihoodEstimator
# +
# MadMiner output
logging.basicConfig(
format='%(asctime)-5.5s %(name)-20.20s %(levelname)-7.7s %(message)s',
datefmt='%H:%M',
level=logging.INFO
)
# Output of all other modules (e.g. matplotlib)
for key in logging.Logger.manager.loggerDict:
if "madminer" not in key:
logging.getLogger(key).setLevel(logging.WARNING)
# -
# ## 1. Make (unweighted) training and test samples with augmented data
# At this point, we have all the information we need from the simulations. But the data is not quite ready to be used for machine learning. The `madminer.sampling` class `SampleAugmenter` will take care of the remaining book-keeping steps before we can train our estimators:
#
# First, it unweights the samples, i.e. for a given parameter vector `theta` (or a distribution `p(theta)`) it picks events `x` such that their distribution follows `p(x|theta)`. The selected samples will all come from the event file we have so far, but their frequency is changed -- some events will appear multiple times, some will disappear.
#
# Second, `SampleAugmenter` calculates all the augmented data ("gold") that is the key to our new inference methods. Depending on the specific technique, these are the joint likelihood ratio and / or the joint score. It saves all these pieces of information for the selected events in a set of numpy files that can easily be used in any machine learning framework.
sampler = SampleAugmenter('data/delphes_data_shuffled.h5')
# The `SampleAugmenter` class defines five different high-level functions to generate train or test samples:
# - `sample_train_plain()`, which only saves observations x, for instance for histograms or ABC;
# - `sample_train_local()` for methods like SALLY and SALLINO, which will be demonstrated in the second part of the tutorial;
# - `sample_train_density()` for neural density estimation techniques like MAF or SCANDAL;
# - `sample_train_ratio()` for techniques like CARL, ROLR, CASCAL, and RASCAL, when only theta0 is parameterized;
# - `sample_train_more_ratios()` for the same techniques, but with both theta0 and theta1 parameterized;
# - `sample_test()` for the evaluation of any method.
#
# For the arguments `theta`, `theta0`, or `theta1`, you can (and should!) use the helper functions `benchmark()`, `benchmarks()`, `morphing_point()`, `morphing_points()`, and `random_morphing_points()`, all defined in the `madminer.sampling` module.
#
# Here we'll train a likelihood estimator with the SCANDAL method, so we focus on the `extract_samples_train_density()` function. We'll sample the numerator hypothesis in the likelihood ratio with 1000 points drawn from a Gaussian prior, and fix the denominator hypothesis to the SM.
#
# Note the keyword `sample_only_from_closest_benchmark=True`, which makes sure that for each parameter point we only use the events that were originally (in MG) generated from the closest benchmark. This reduces the statistical fluctuations in the outcome quite a bit.
# ## 3. Evaluate likelihood estimator
# `estimator.evaluate_log_likelihood(theta,x)` estimated the log likelihood for all combination between the given phase-space points `x` and parameters `theta`. That is, if given 100 events `x` and a grid of 25 `theta` points, it will return 25\*100 estimates for the log likelihood, indexed by `[i_theta,i_x]`.
theta_each = np.linspace(0.,2.,31)
#theta0, theta1 = np.meshgrid(theta_each, theta_each)
theta0 = np.meshgrid(theta_each)[0]
#theta_grid = np.vstack((theta0.flatten())).T # doesn't work
theta_grid = np.vstack((theta0.flatten()))
np.save('data/samples/theta_grid.npy', theta_grid)
# +
estimator = LikelihoodEstimator(
n_mades=3,
n_hidden=(300,),
activation="tanh"
)
estimator.load('models/scandal')
log_p_hat, _ = estimator.evaluate_log_likelihood(
theta='data/samples/theta_grid.npy',
x='data/samples/x_test.npy',
evaluate_score=False
)
# -
# Let's look at the result:
# +
bin_size = theta_each[1] - theta_each[0]
#edges = np.linspace(theta_each[0] - bin_size/2, theta_each[-1] + bin_size/2, len(theta_each)+1)
fig = plt.figure(figsize=(6,5))
#ax = plt.gca()
expected_llr = np.mean(log_p_hat,axis=1)
best_fit = theta_grid[np.argmin(-2.*expected_llr)]
plt.scatter(theta_grid**4, -2.* expected_llr)
plt.scatter(best_fit**4, (-2.* expected_llr).min(), s=80., color='black', marker='*', label="Best Fit")
#plt.xlabel(r'$\theta_0$')
plt.xlabel(r'$\mu$')
plt.ylabel(r'-2* expected llr Scandal')
plt.tight_layout()
plt.legend()
plt.show()
# -
theta_grid**4, best_fit**4
# Note that in this tutorial our sample size was very small, and the network might not really have a chance to converge to the correct likelihood function. So don't worry if you find a minimum that is not at the right point (the SM, i.e. the origin in this plot). Feel free to dial up the event numbers in the run card as well as the training samples and see what happens then!
| examples/tutorial_h4l/3c_plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"></ul></div>
# +
# default_exp trainer.mellotron
# -
# +
# export
import os
from pathlib import Path
from pprint import pprint
import torch
from torch.cuda.amp import autocast, GradScaler
import torch.distributed as dist
from torch.nn.parallel import DistributedDataParallel as DDP
from tensorboardX import SummaryWriter
import time
from torch.utils.data import DataLoader
from random import choice
from uberduck_ml_dev.models.common import MelSTFT
from uberduck_ml_dev.utils.plot import (
plot_attention,
plot_gate_outputs,
plot_spectrogram,
save_figure_to_numpy,
)
from uberduck_ml_dev.text.util import text_to_sequence, random_utterance
from uberduck_ml_dev.trainer.tacotron2 import Tacotron2Trainer, Tacotron2Loss
from uberduck_ml_dev.models.mellotron import Mellotron
from uberduck_ml_dev.data_loader import TextMelDataset, TextMelCollate
class MellotronTrainer(Tacotron2Trainer):
REQUIRED_HPARAMS = [
"audiopaths_and_text",
"checkpoint_name",
"checkpoint_path",
"epochs",
"mel_fmax",
"mel_fmin",
"n_mel_channels",
"text_cleaners",
"pos_weight",
]
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.n_frames_per_step_current = self.n_frames_per_step_initial
self.reduction_window_idx = 0
def adjust_frames_per_step(
self,
model: Mellotron,
train_loader: DataLoader,
sampler,
collate_fn: TextMelCollate,
):
"""If necessary, adjust model and loader's n_frames_per_step."""
if not self.reduction_window_schedule:
return train_loader, sampler, collate_fn
settings = self.reduction_window_schedule[self.reduction_window_idx]
if settings["until_step"] is None:
return train_loader, sampler, collate_fn
if self.global_step < settings["until_step"]:
return train_loader, sampler, collate_fn
old_fps = settings["n_frames_per_step"]
while settings["until_step"] and self.global_step >= settings["until_step"]:
self.reduction_window_idx += 1
settings = self.reduction_window_schedule[self.reduction_window_idx]
fps = settings["n_frames_per_step"]
bs = settings["batch_size"]
print(f"Adjusting frames per step from {old_fps} to {fps}")
self.batch_size = bs
model.set_current_frames_per_step(fps)
self.n_frames_per_step_current = fps
_, _, train_loader, sampler, collate_fn = self.initialize_loader(
n_frames_per_step_current=self.n_frames_per_step_current,
include_f0=self.include_f0,
)
return train_loader, sampler, collate_fn
def validate(self, **kwargs):
model = kwargs["model"]
val_set = kwargs["val_set"]
collate_fn = kwargs["collate_fn"]
criterion = kwargs["criterion"]
sampler = DistributedSampler(val_set) if self.distributed_run else None
(
total_loss,
total_mel_loss,
total_gate_loss,
total_mel_loss_val,
total_gate_loss_val,
) = (0, 0, 0, 0, 0)
total_steps = 0
model.eval()
speakers_val = []
total_mel_loss_val = []
total_gate_loss_val = []
with torch.no_grad():
val_loader = DataLoader(
val_set,
sampler=sampler,
shuffle=False,
batch_size=self.batch_size,
collate_fn=collate_fn,
)
for batch in val_loader:
total_steps += 1
if self.distributed_run:
X, y = model.module.parse_batch(batch)
speakers_val.append(X[5])
else:
X, y = model.parse_batch(batch)
speakers_val.append(X[5])
y_pred = model(X)
mel_loss, gate_loss, mel_loss_batch, gate_loss_batch = criterion(
y_pred, y
)
if self.distributed_run:
reduced_mel_loss = reduce_tensor(mel_loss, self.world_size).item()
reduced_gate_loss = reduce_tensor(gate_loss, self.world_size).item()
reduced_val_loss = reduced_mel_loss + reduced_gate_loss
else:
reduced_mel_loss = mel_loss.item()
reduced_gate_loss = gate_loss.item()
reduced_mel_loss_val = mel_loss_batch.detach()
reduced_gate_loss_val = gate_loss_batch.detach()
total_mel_loss_val.append(reduced_mel_loss_val)
total_gate_loss_val.append(reduced_gate_loss_val)
reduced_val_loss = reduced_mel_loss + reduced_gate_loss
total_mel_loss += reduced_mel_loss
total_gate_loss += reduced_gate_loss
total_loss += reduced_val_loss
mean_mel_loss = total_mel_loss / total_steps
mean_gate_loss = total_gate_loss / total_steps
mean_loss = total_loss / total_steps
total_mel_loss_val = torch.hstack(total_mel_loss_val)
total_gate_loss_val = torch.hstack(total_gate_loss_val)
speakers_val = torch.hstack(speakers_val)
self.log_validation(
X,
y_pred,
y,
mean_loss,
mean_mel_loss,
mean_gate_loss,
total_mel_loss_val,
total_gate_loss_val,
speakers_val,
)
model.train()
def sample_inference(self, model):
if self.rank is not None and self.rank != 0:
return
# Generate an audio sample
with torch.no_grad():
utterance = torch.LongTensor(
text_to_sequence(random_utterance(), self.text_cleaners, self.p_arpabet)
)[None].cuda()
speaker_id = (
choice(self.sample_inference_speaker_ids)
if self.sample_inference_speaker_ids
else randint(0, self.n_speakers - 1)
)
input_ = [utterance, 0, torch.LongTensor([speaker_id]).cuda()]
if self.include_f0:
input_.append(torch.zeros([1, 1, 200], device=self.device))
# 200 can be changed
model.eval()
_, mel, gate, attn = model.inference(input_)
model.train()
try:
audio = self.sample(mel[0])
self.log("SampleInference", self.global_step, audio=audio)
except Exception as e:
print(f"Exception raised while doing sample inference: {e}")
print("Mel shape: ", mel[0].shape)
self.log(
"Attention/sample_inference",
self.global_step,
image=save_figure_to_numpy(
plot_attention(attn[0].data.cpu().transpose(0, 1))
),
)
self.log(
"MelPredicted/sample_inference",
self.global_step,
image=save_figure_to_numpy(plot_spectrogram(mel[0].data.cpu())),
)
self.log(
"Gate/sample_inference",
self.global_step,
image=save_figure_to_numpy(
plot_gate_outputs(gate_outputs=gate[0].data.cpu())
),
)
@property
def training_dataset_args(self):
return [
*super().training_dataset_args,
# self.training_audiopaths_and_text,
# self.text_cleaners,
# self.p_arpabet,
# audio params
# self.n_mel_channels,
# self.sampling_rate,
# self.mel_fmin,
# self.mel_fmax,
# self.filter_length,
# self.hop_length,
# self.win_length,
# self.max_wav_value,
self.include_f0,
# self.pos_weight,
]
# def warm_start(self, model, optimizer, start_epoch=0):
# print("Starting warm_start", time.perf_counter())
# checkpoint = self.load_checkpoint()
# # TODO(zach): Once we are no longer using checkpoints of the old format, remove the conditional and use checkpoint["model"] only.
# model_state_dict = (
# checkpoint["model"] if "model" in checkpoint else checkpoint["state_dict"]
# )
# model.from_pretrained(
# model_dict=model_state_dict,
# device=self.device,
# ignore_layers=self.ignore_layers,
# )
# if "optimizer" in checkpoint and len(self.ignore_layers) == 0:
# optimizer.load_state_dict(checkpoint["optimizer"])
# if "iteration" in checkpoint:
# start_epoch = checkpoint["iteration"]
# if "learning_rate" in checkpoint:
# optimizer.param_groups[0]["lr"] = checkpoint["learning_rate"]
# self.learning_rate = checkpoint["learning_rate"]
# if "global_step" in checkpoint:
# self.global_step = checkpoint["global_step"]
# print(f"Adjusted global step to {self.global_step}")
# print("Ending warm_start", time.perf_counter())
# return model, optimizer, start_epoch
def train(self):
print("start train", time.perf_counter())
train_set, val_set, train_loader, sampler, collate_fn = self.initialize_loader(
include_f0=self.include_f0
)
criterion = Tacotron2Loss(
pos_weight=self.pos_weight
) # keep higher than 5 to make clips not stretch on
model = Mellotron(self.hparams)
# move to TTSTrainer class
if self.device == "cuda":
model = model.cuda()
if self.distributed_run:
model = DDP(model, device_ids=[self.rank])
optimizer = torch.optim.Adam(
model.parameters(),
lr=self.learning_rate,
weight_decay=self.weight_decay,
)
start_epoch = 0
if self.warm_start_name:
model, optimizer, start_epoch = self.warm_start(model, optimizer)
if self.fp16_run:
scaler = GradScaler()
# main training loop
for epoch in range(start_epoch, self.epochs):
train_loader, sampler, collate_fn = self.adjust_frames_per_step(
model, train_loader, sampler, collate_fn
)
if self.distributed_run:
sampler.set_epoch(epoch)
for batch in train_loader:
start_time = time.perf_counter()
self.global_step += 1
model.zero_grad()
if self.distributed_run:
X, y = model.module.parse_batch(batch)
else:
X, y = model.parse_batch(batch)
if self.fp16_run:
with autocast():
y_pred = model(X)
(
mel_loss,
gate_loss,
mel_loss_batch,
gate_loss_batch,
) = criterion(y_pred, y)
loss = mel_loss + gate_loss
loss_batch = mel_loss_batch + gate_loss_batch
else:
y_pred = model(X)
mel_loss, gate_loss, mel_loss_batch, gate_loss_batch = criterion(
y_pred, y
)
loss = mel_loss + gate_loss
loss_batch = mel_loss_batch + gate_loss_batch
if self.distributed_run:
reduced_mel_loss = reduce_tensor(mel_loss, self.world_size).item()
reduced_gate_loss = reduce_tensor(gate_loss, self.world_size).item()
reduced_loss = reduce_mel_loss + reduced_gate_loss
else:
reduced_mel_loss = mel_loss.item()
reduced_gate_loss = gate_loss.item()
reduced_gate_loss_batch = gate_loss_batch.detach()
reduced_mel_loss_batch = mel_loss_batch.detach()
reduced_loss = reduced_mel_loss + reduced_gate_loss
reduced_loss_batch = reduced_gate_loss_batch + reduced_mel_loss_batch
if self.fp16_run:
scaler.scale(loss).backward()
scaler.unscale_(optimizer)
grad_norm = torch.nn.utils.clip_grad_norm(
model.parameters(), self.grad_clip_thresh
)
scaler.step(optimizer)
scaler.update()
else:
loss.backward()
grad_norm = torch.nn.utils.clip_grad_norm(
model.parameters(), self.grad_clip_thresh
)
optimizer.step()
step_duration_seconds = time.perf_counter() - start_time
self.log_training(
model,
X,
y_pred,
y,
reduced_loss,
reduced_mel_loss,
reduced_gate_loss,
reduced_mel_loss_batch,
reduced_gate_loss_batch,
grad_norm,
step_duration_seconds,
)
if epoch % self.epochs_per_checkpoint == 0:
self.save_checkpoint(
f"{self.checkpoint_name}_{epoch}",
model=model,
optimizer=optimizer,
iteration=epoch,
learning_rate=self.learning_rate,
global_step=self.global_step,
)
# There's no need to validate in debug mode since we're not really training.
if self.debug:
continue
self.validate(
model=model,
val_set=val_set,
collate_fn=collate_fn,
criterion=criterion,
)
# -
| nbs/trainer.mellotron.ipynb |