code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (data)
# language: python
# name: data
# ---
# https://leetcode.com/problems/short-encoding-of-words/
# +
class Solution:
def minimumLengthEncoding(self, words) -> int:
words.sort(key=lambda x: len(x))
words_len = len(words)
pop_idxs = []
for i,w1 in enumerate(words):
l = len(w1)
for j,w2 in enumerate(words[i+1:]):
if w1 == w2[-l:]:
pop_idxs.append(i)
break
res = 0
for i,w in enumerate(words):
if pop_idxs and i == pop_idxs[0]:
pop_idxs.pop(0)
else:
res += len(w) + 1
return res
# -
# prefix search
# https://leetcode.com/problems/short-encoding-of-words/discuss/734845/Python%3A-Two-methods.-(Prefix-search-and-Trie-with-suffix)
good = set(words)
for word in words:
for k in range(1, len(word)):
good.discard(word[k:])
return sum(len(word)+1 for word in good)
| lt820_short encoding of words prefix search trie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
from multiinstance.agglomerative_clustering import AgglomerativeClustering
from multiinstance.ward_clustering import WardClustering
# +
# AgglomerativeClustering??
# +
from multiinstance.utils import *
from multiinstance.distanceApproaches import *
from multiinstance.data.syntheticData import buildDataset,getBag
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
import scipy.stats as ss
from tqdm.notebook import tqdm
# +
# AgglomerativeClustering??
# -
for rep in range(10):
dsi = buildDataset(10,alphaDistr=lambda: np.random.uniform(.01,.25),
nP=5,nU=10)
dsi = addTransformScores(dsi)
dsi.alphaHats,dsi.curves = getBagAlphaHats(dsi,numbootstraps=10)
agg0 = AgglomerativeClustering(dsi, 0.65)
agg0.cluster()
fig,ax=plt.subplots(2,1,sharex=True)
ax[0].plot(agg0.meanAbsErrs,label="mae")
# ax[1].plot(agg0.bagEstimateVariances,label="var")
ax[0].legend()
ward = WardClustering(dsi)
ward.cluster()
ward.meanAbsErrs
ax[1].plot(ward.meanAbsErrs,label="ward")
ax[1].legend()
plt.savefig("figs/nb_08/fig_{}.pdf".format(rep),
format="pdf")
| 08_compare_ks_agglomerative_to_ward_clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Evaluating Classifiers
import pandas as pd
# * **The first model** predicts a high risk of cancer for 800 out of 1000 patients’ images in the test set.
# * Of these 800 images 50 actually show signs of skin cancer.
# * Hence, all problematic images are correctly identified.
# +
TP = 50
TN = 200
FN = 0
FP = 750
acc = (TP + TN) / 1000
acc # not such a great metric when there is class imbalance (in y_train)
# +
# confusion matrix
data = [[TN, FN],
[FP, TP]]
pd.DataFrame(data, columns=['real 0', 'real 1'], index=['predict 0', 'predict 1'])
# +
# precision and recall
# precision: how many of our positive predictions (cancers) are really positives
# 0.0 .. 1.0
precision = TP / (TP + FP)
precision
# -
# recall: how many of the real positives (cancers) do we detect?
# 0.0 .. 1.0
recall = TP / (TP + FN)
recall
# * **The second classifier** categorizes 100 out of 1000 images into the high risk group.
# * 40 of the 100 images show real signs of cancer.
# * 10 images are not identified and falsely classified as low-risk.
# +
TP = 40
TN = 890
FP = 60
FN = 10
acc = (TP + TN) / (TP + TN + FP + FN)
acc
# -
# accuracy = #correct predictions / #total predictions
# +
# the titanic data is class imbalanced
dead = 550
alive = 350
dead / (alive + dead)
# -
# ### Challenge (easy):
#
# * look up functions for confusion matrix, precision and recall in scikit
# ### Challenge (difficult):
#
# * plot an ROC curve
| week2/EvaluatingClassifiers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Autograph: Intuitive Data-Driven Control At Last
# Continuing our blogs, we shift our focus to yet another new feature in TF, the `autograph` functionality.
#
# Previously, and as far as intuitive expression of code was concerned, "graph ops" efficiently solved complex calculations while failed at simple, sequential control.
#
# By generating on-demand Python code now, `autograph` transparently patches all the necessary graph ops together and packages the result into a "python op".
#
# While the generated new ops are potentially faster than the code before them, in this blog we are more interested in the new expressive powers of the `autograph` package.
#
# Specifically, we look at what becomes possible when decorating our functions with the new `tf.function` decorator, as doing this would by default invoke the `autograph` functionality.
#
# Our objective is to ultimately arrive at a model as represented by the [graph](./autograph.pdf).
#
# Just as before, we need to prep our environment to run any meaningful code:
import numpy as np
import tensorflow as tf
import dataset as qd
import custom as qc
import layers as ql
ks = tf.keras
kl = ks.layers
# Next, we borrow the `pos_timing` function from our previous blogs, and override it to return a constant "timing signal" tensor, depending on the `width` and `depth` arguments.
#
# As our first task is to implement a "python branch" in our new `Embed` op, we will be using two different "timing" tensors, one for the `encode` input and the other for the `decode` input.
def pos_timing(width, depth):
t = ql.pos_timing(width, depth)
t = tf.constant(t, dtype=tf.float32)
return t
# The `Embed` layer will thus create the two constant tensors to be sourced in the subsequent `call` methods.
#
# Our model will call the shared `Embed` instance for both of our stacks. As we have decorated its `call` method with `tf.function`, we can use familiar and intuitive Python comparisons to branch on the value of tensors on-the-fly, during graph execution.
#
# Clearly, our two stacks, while having the same `depth`s, have different `width`s. Also the constant "timing" tensors have different `width`s as well.
#
# Yet we are still able to pick-and-match the otherwise incompatible tensors and successfully add them together, all depending on the actual `width` of our "current" input tensor:
class Embed(qc.Embed):
def __init__(self, ps):
super().__init__(ps)
self.enc_p = pos_timing(ps.width_enc, ps.dim_hidden)
self.dec_p = pos_timing(ps.width_dec, ps.dim_hidden)
@tf.function(input_signature=[[
tf.TensorSpec(shape=[None, None], dtype=tf.int32),
tf.TensorSpec(shape=[None], dtype=tf.int32)
]])
def call(self, x):
y, lens = x
y = tf.nn.embedding_lookup(self.emb, y)
s = tf.shape(y)
if s[-2] == self.ps.width_enc:
y += tf.broadcast_to(self.enc_p, s)
elif s[-2] == self.ps.width_dec:
y += tf.broadcast_to(self.dec_p, s)
else:
pass
y *= tf.cast(s[-1], tf.float32)**0.5
return [y, lens]
# Next we demonstrate how on-the-fly "python ops" can also provide insights into inner processes and data flows.
#
# We borrow our `Frames` layer from the previous blog and override its `call` method with a `tf.function` decorated new version that, besides calling `super().call()`, also calls a new `print_row` Python function on every row in our batch.
#
# Yes, we are calling a Python function and printing its results in a TF graph op while never leaving our intuitive and familiar Python environment! Isn't that great?
#
# The `print_row` function itself is simple, it iterates through the tokens of the "row", it does a lookup of each in our `vocab` "table" for the actual character representing the token and then it "joins" all the characters and prints out the resulting string.
#
# And, if we scroll down to the listing of our training session, we can actually see the "sliding context" of our samples as they fly by during our training.
#
# Needless to say, the listing confirms that our `Frames` layer does a good job concatenating the varied length sample inputs, the target results, as well as the necessary separators.
#
# As a result, a simple Python function, usable during graph ops, provides us invaluable insights deep into our inner processes and data flow.
class Frames(qc.Frames):
@tf.function
def call(self, x):
y = super().call.python_function(x)
tf.print()
def print_row(r):
tf.print(
tf.numpy_function(
lambda ts: ''.join([qd.vocab[t] for t in ts]),
[r],
Tout=[tf.string],
))
return r
tf.map_fn(print_row, self.prev)
return y
# Our next new layer is the partial `Deduce` layer, showcasing how control is intuitive at last from data-driven branching to searching.
#
# This layer will be used in the next group of blogs as a replacement for our previous `Debed` layer. It contains a tensor-dependent `for` loop to iteratively replace our masked characters with "deduced" ones.
#
# The future `Probe` layer, building on the `Deduce` scheme, implements an approximation of "Beam Search", see [paper](https://arxiv.org/pdf/1702.01806.pdf).
#
# It effectively iterates through the hidden dimensions of the output, and based on parallel `topk` searches, comparing various choices for "debeding" the output, it settles on an "optimal" debedding and thus final token output for our `decoder`.
#
# Without `autograph` the data-driven looping/branching graph ops would have to be expressed in a much more convoluted manner:
"""
class Deduce(Layer):
@tf.function
def call(self, x):
toks, *x = x
if self.cfg.runtime.print_toks:
qu.print_toks(toks, qd.vocab)
y = self.deduce([toks] + x)
n = tf.shape(y)[1]
p = tf.shape(toks)[1] - n
for i in tf.range(n):
t = toks[:, :n]
m = tf.equal(t, qd.MSK)
if tf.equal(tf.reduce_any(m), True):
t = self.update(t, m, y)
if self.cfg.runtime.print_toks:
qu.print_toks(t, qd.vocab)
toks = tf.pad(t, [[0, 0], [0, p]])
y = self.deduce([toks] + x)
else:
e = tf.equal(t, qd.EOS)
e = tf.math.count_nonzero(e, axis=1)
if tf.equal(tf.reduce_any(tf.not_equal(e, 1)), False):
break
return y
"""
class Probe(ql.Layer):
def __init__(self, ps):
super().__init__(ps)
self.dbd = qc.Dense(self, 'dbd', [ps.dim_hidden, ps.dim_vocab])
@tf.function
def call(self, x):
y, lens = x
s = tf.shape(y)
y = tf.reshape(y, [s[0] * s[1], -1])
y = self.dbd(y)
y = tf.reshape(y, [s[0], s[1], -1])
y = y[:, :tf.math.reduce_max(lens), :]
return y
# Our model needs to be updated as well to use the newly defined components.
#
# Other than that, we are ready to start training:
def model_for(ps):
x = [ks.Input(shape=(), dtype='int32'), ks.Input(shape=(), dtype='int64')]
x += [ks.Input(shape=(), dtype='int32'), ks.Input(shape=(), dtype='int64')]
x += [ks.Input(shape=(), dtype='int32'), ks.Input(shape=(), dtype='int64')]
y = qc.ToRagged()(x)
y = Frames(ps)(y)
embed = Embed(ps)
ye = qc.Encode(ps)(embed(y[:2]))
yd = qc.Decode(ps)(embed(y[2:]) + [ye[0]])
y = Probe(ps)(yd)
m = ks.Model(inputs=x, outputs=y)
m.compile(optimizer=ps.optimizer, loss=ps.loss, metrics=[ps.metric])
print(m.summary())
return m
# By firing up our training session, we can confirm the model's layers and connections. The listing of a short session follows.
#
# We can easily adjust the parameters to tailor the length of the sessions to our objectives.
ps = qd.Params(**qc.params)
ps.num_epochs = 1
import masking as qm
qm.main_graph(ps, qc.dset_for(ps).take(10), model_for(ps))
# With our TensorBoard `callback` in place, the model's `fit` method will generate the standard summaries that TB can conveniently visualize.
#
# If you haven't run the code below, an already generated graph is [here](./autograph.pdf).
# +
# #%load_ext tensorboard
# #%tensorboard --logdir /tmp/q/logs
# -
# This concludes our blog, please see how to use customize the losses and metrics driving the training by clicking on the next blog.
| qnarre.com/static/pybooks/autograph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
with open('exported-discussion.txt') as f:
text = f.read()
# + pycharm={"name": "#%%\n"}
import nltk
import nltk.corpus as corpus
nltk.download("stopwords")
# + pycharm={"name": "#%%\n"}
import numpy as np
x, y = np.ogrid[:1000, :1000]
mask = (x - 500) ** 2 + (y - 500) ** 2 > 400 ** 2
mask = 255 * mask.astype(int)
# + pycharm={"name": "#%%\n"}
from wordcloud import WordCloud, STOPWORDS
current_stopwords = set(STOPWORDS)
current_stopwords.update(corpus.stopwords.words("german"))
current_stopwords.update(["mal", "ja", "aber", "schon", "wäre", "gibt"])
wordcloud = WordCloud(background_color="white", mask=mask, width=1000, height=1000, stopwords=current_stopwords).generate(text)
wordcloud.to_file("wordcloud.png")
# + pycharm={"name": "#%%\n"}
import matplotlib.pyplot as plt
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.show()
| wordcloud.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exploring Data in Jupyter Notebook
# Importing the data
import pandas
df = pandas.read_excel('house_price.xlsx')
df[:10]
# A quick look of the data...
df.describe()
# We can also create histograms to see the distributions of our data...
df.hist(figsize=(20,20))
# Or we can also create scatter plots to see the relationships among all columns..
from pandas.plotting import scatter_matrix
scatter_matrix(df,figsize=(20, 20), diagonal='kde')
# # Exploring the data.. we want to see how many different types of houses are in the record:
df.groupby('house_type').count()
# Let's compare the average prices and views for each type of houses.
df.groupby('house_type').mean()
# We can also visualize the difference by creating histograms for each type of houses.
df.groupby('house_type').hist(figsize=(12,4),column=['price','views'])
# Or if we want to see how the ages of houses determine the prices. Here we use a scatterplot.
df.plot.scatter(x='built_in',y='price')
# Since house prices are largely determined by sizes, ages and other factors, let's see how the house ages determine the house unite price, i.e., price/size. Let's calculate the unit price first, and then add the results to a new column called unit_price
df['unit_price']=df['price']/df['area']
# For each year, we calculate the average unite price, and assign the result to a new dataframe
mean_price =df.groupby('built_in').mean()
# A Line chart is a good choice for visualizing trends. Let's use a line chart to see how the unite prices is determined by the house ages.
mean_price.plot.line(y='unit_price')
# This is how we visualize and read files in Jupyter Notebook with Excel.
| Reading Files .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 1.linear
def linear_search(data, target):
for idx in range(len(data)):
if data[idx] == target:
return idx
return None
# -
if __name__ =="__main__":
data = [i ** 2 for i in range(1, 11)]
target = 9
idx = linear_search(data, target)
if idx == None:
print("{} 이 존재하지 않습니다".format(target))
else:
print("찾는 데이터의 인덱스는 {} 이고 데이터는 {}입니다".format(idx, data[idx]))
def linear_search(data, target):
for i in range(len(data)):
if data[i] == target:
print("index is {}, data is {}".format(i, data[i]))
return i
return None
if __name__ == "__main__":
data = [i **2 for i in range(1, 11)]
target = 9
linear_search(data, target)
# +
#2.Binary Search
def binary_search(data, target):
# 먼저 정렬합니다.
data.sort()
start = 0
end = len(data) - 1
while start <= end:
mid = (start + end) // 2
if data[mid] == target:
return mid
elif data[mid] > target:
end = mid -1
else:
start = mid + 1
return None
# -
if __name__ =="__main__":
data = [i ** 2 for i in range(1, 11)]
target = 9
idx = binary_search(data, target)
if idx == None:
print("{} 이 존재하지 않습니다".format(target))
else:
print("찾는 데이터의 인덱스는 {} 이고 데이터는 {}입니다".format(idx, data[idx]))
def binary_search(data, target):
data.sort()
start = 0
end = len(data) - 1
while start <= end:
mid = (start+end) // 2
if data[mid] == target:
print("index is {}, data is {}".format(mid, data[mid]))
return mid
elif target < data[mid]:
end = mid -1
else:
start = mid +1
return None
if __name__ == "__main__":
data = [i **2 for i in range(1, 11)]
target = 9
binary_search(data, target)
#3 Bubble sort
def bubble_sort(data):
data_len = len(data)
for i in range(data_len - 1):
for j in range(data_len -1 -i):
if data[j] > data[j+1]:
data[j], data[j+1] = data[j+1], data[j]
if __name__ == "__main__":
li = [2, 3, 5, 2, 3, 8, 6, 7, 10, 8, 1, 4]
bubble_sort(li)
print(li)
# +
#bubble sort
def bubble_sort(data):
for i in range(len(data) - 1):
for j in range(len(data) -1 -i):
if data[j] > data[j+1]:
data[j], data[j+1] = data[j+1], data[j]
print(data)
return data
# -
if __name__ == "__main__":
li = [2, 3, 5, 2, 3, 8, 6, 7, 10, 8, 1, 4]
bubble_sort(li)
# +
#4 Quick sort
def quick_sort(data, start, end):
if start >= end:
return
left = start
right = end
pivot = data[(start + end) // 2]
# right = index의 마지막이다
while left <= right:
while data[left] < pivot:
left +=1
while data[right] > pivot:
right -= 1
#left 와 right가 교차하지 않았다면 교환
if left <= right:
data[left], data[right] = data[right], data[left]
left += 1
right -=1
quick_sort(data, start, right)
quick_sort(data, left, end)
# -
if __name__ == "__main__":
data = [2, 5, 4, 1, 8, 10, 5, 3, 6, 6, 5, 7, 9, 12, 11]
quick_sort(data, 0, len(data) -1)
print(data)
| fun_coding/basic concepts/basic_algorithms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Probabilistic view of least squares
import numpy as np
import matplotlib.pyplot as plt
# Assume a diagonal covariance matrix $\Sigma = \sigma^2I$. The density is:
#
# $$p(y|\mu,\sigma^2) = \frac{1}{(2\pi\sigma^2)^{\frac{n}{2}}}exp\left(-\frac{1}{2\sigma^2}(y-\mu)^T(y-\mu)\right)$$
#
# if we set $\mu = Xw$ and find maximum likelihood for w:
#
# $$w_{ML} = \arg \underset{w}{\max} \ln p(y|\mu = Xw,\sigma^2)$$
#
# $$w_{ML} = \arg \underset{w}{\max} -\frac{1}{2\sigma^2}||y-Xw||^2 - \frac{n}{2}\ln(2\pi\sigma^2)$$
#
# we remove the last bit because it doesn't involve w
#
# $$w_{ML} = \arg \underset{w}{\max} -\frac{1}{2\sigma^2}||y-Xw||^2$$
#
# Using least squares:
#
# $$w_{ML} = \arg \underset{w}{\min}||y − Xw||^2$$
#
# These are both the same, so:
#
# $$w_{ML} = \arg \underset{w}{\max} -\frac{1}{2\sigma^2}||y-Xw||^2 \Leftrightarrow \arg \underset{w}{\min}||y − Xw||^2$$
# we are making an independent Gaussian noise assumption about error:
#
# $\epsilon_i = y_i - x^T_iw$
#
# or other ways to say same thing:
#
# $y_i = x^T_iw + \epsilon_i$, $\epsilon_i\overset{iid}{\tilde{}} N(0, \sigma^2)$, for $i = 1,...,n$
#
# $y_i\overset{ind}{\tilde{}} N(x^T_iw, \sigma^2)$, for $i = 1,...,n$
#
# $y \tilde{} N(Xw, \sigma^2I)$
# ### our model assumption is $y \tilde{} N(Xw, \sigma^2I)$
#
# $\mathbb{E}[w_{ML}] = \mathbb{E}[(X^TX)^{-1} X^Ty]$
#
# or $\mathbb{E}[w_{ML}] = \int_{}[(X^TX)^{-1} X^Ty] p(y|X, w)dy = \mu$
#
# $\mathbb{E}[w_{ML}] = (X^TX)^{-1} X^T\mathbb{E}[y]$
#
# $\mathbb{E}[w_{ML}] = (X^TX)^{-1} X^TXw$
#
# $\mathbb{E}[w_{ML}] = w$
#
# $w_{ML}$ is an un-biased estimate of w
#
# the expected solution = the correct one
#
# but if the variance is too high, we cant expect a value near the solution
#
# ### covariance
#
# $Var[y] = \mathbb{E}[(y - \mathbb{E}[y])(y - \mathbb{E}[y])^T] = \Sigma$
#
# plugging in $\mathbb{E}[y] = \mu$
#
# $Var[y] = \mathbb{E}[(y - \mu)(y - \mu)^T]$
#
# $= \mathbb{E}[yy^T - y\mu^T - \mu y^T + \mu\mu^T]$
#
# $= \mathbb{E}[yy^T] - \mu\mu^T = \Sigma$
#
# $= \mathbb{E}[yy^T] = \Sigma + \mu\mu^T$
#
# ### Variance of solution
#
# with least squares regression, we need to find:
#
# $Var[w_{ML}] = \mathbb{E}[(w_{ML} - \mathbb{E}[w_{ML}])(w_{ML} - \mathbb{E}[w_{ML}])^T]$
#
# $= \mathbb{E}[w_{ML}w^T_{ML}] - \mathbb{E}[w_{ML}]\mathbb{E}[w_{ML}]^T$
#
# plugging in our previous equalities:
#
# $Var[w_{ML}] = \mathbb{E}[(X^TX)^{-1} X^Tyy^TX (X^TX)^{-1}] - ww^T$
#
# $= (X^TX)^{-1} X^T\mathbb{E}[yy^T]X (X^TX)^{-1} - ww^T$
#
# $= (X^TX)^{-1} X^T(\sigma^2I + Xww^TX^T)X (X^TX)^{-1} - ww^T$
#
# $= (X^TX)^{-1} X^T\sigma^2IX (X^TX)^{-1} + (X^TX)^{-1} X^TXww^TX^TX (X^TX)^{-1} - ww^T$
#
# $= \sigma^2(X^TX)^{-1}$
# so with model $y \tilde{} N(Xw, \sigma^2I)$:
#
# $\mathbb{E}[w_{ML}] = w$
#
# $Var[w_{ML}] = \sigma^2(X^TX)^{-1}$
#
# this means that if there are very large values in the variance $\sigma^2(X^TX)^{-1}$ the values of $w_{ML}$ are very sensitive to the measured data y
#
# this is bad if we want to use $w_{ML}$ to predict and analyze because we aren't confident in the maximum likelihood of w
# ## Ridge Regression
#
# the values in $w_{ML}$ can be huge so we constrain the model parameters
#
# were using form:
#
# $w_{OPT} = arg \underset{w}{\min}||y - Xw||^2 + \lambda g(w)$
#
# - $\lambda > 0$: a regularization parameter
# - $g(w) < 0$: a penalty function which encourages a desired property/properties from w
#
# ridge regression uses the squared penalty $g(w) = ||w||^2$
#
# $w_{RR} = arg \underset{w}{\min}||y - Xw||^2 + \lambda||w||^2$
#
# this penalizes large values of w
#
# the trade off between main term and penalty term is controlled by $\lambda$
#
# - Case $\lambda \to 0 : w_{RR} \to w_{LS}$
# - Case $\lambda \to \infty : w_{RR} \to \vec{0}$
# ### Solving ridge regression
#
# same procedure as least squares
#
# $L = ||y - Xw||^2 + \lambda||w||^2$
#
# $L = (y - Xw)^T(y - Xw) + \lambda w^Tw$
#
# set gradient of loss to 0
#
# $\triangledown_w L = -2X^Ty + 2X^tXw + 2\lambda w = 0$
#
# $w_{RR} = (\lambda I + X^TX)^{-1}X^Ty$
#
# There is a trade off between the squared error and the penalty
#
# we write each term as level sets, Curves where function evaluation gives the same number
#
# The sum of these gives a new set of levels with a unique minimum
#
# $||y−Xw||^2 +\lambda ||w||^2 = (w−w_{LS})^T(X^TX)(w−w_{LS})+\lambda w^Tw+ (const. w.r.t. w)$
# ### Preprocessing
#
# Because the weights are limited,
#
# We assume this preprocessing has been applied before:
#
# we take away the mean:
#
# $y \gets y - \frac{1}{n}\sum^n_{i=1}y_i$
#
# we standardize the dimensions of $x_i$:
#
# $x_{ij} \gets (x_{ij} - \bar{x}_{\cdot j})/\hat{\sigma}_j$
#
# $\hat{\sigma}_j = \sqrt{\frac{1}{n}\sum^n_{i=1}(x_{ij} - \bar{x}_{\cdot j})^2}$
#
# this subtracts the empirical mean and divides by the empirical standard
# deviation for each dimension
#
# this means there is no need for the dimension of 1's
# ### Analysis of ridge regression
#
# $w_{RR} = (\lambda(X^TX)^{-1} + I)w_{LS}$
#
# $||w_{RR}||_2 \le ||w_{LS}||_2$
#
# $w_{RR}= V(\lambda S^{−2} + I)^{−1}V^Tw_{LS}$
#
# $w_{RR}= VMV^Tw_{LS}$
#
# $M_{ii} = \frac{s^2_{ii}}{\lambda + s^2_{ii}}$
#
# can add $\sqrt{\lambda}$ diagonal matrix to X
# +
a = np.arange(5).reshape(-1)
print np.square(a)
print np.matmul(np.transpose(a), a)
# -
| lectures/Lecture_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import datetime
import json
import uuid
import xml.etree.ElementTree as ET
from collections import OrderedDict
import os
import pandas as pd
import numpy as np
import argparse
import xmltodict
from cassandra.cluster import Cluster
# +
def unix_time(dt):
epoch = datetime.datetime.utcfromtimestamp(0)
delta = dt - epoch
return delta.total_seconds()
def unix_time_millis(dt):
return int(unix_time(dt) * 1000.0)
def generate_stmt(session, table, columns):
col_exp = ', '.join(columns)
val_exp = ', '.join(['?' for _ in range(len(columns))])
stmt = """INSERT INTO {table} ({columns}) VALUES({values})""".format(table=table, columns=col_exp, values=val_exp)
return session.prepare(stmt)
# -
def parse_meta_data(raw_meta_data):
tree = ET.parse(raw_meta_data) # element tree
root = tree.getroot()
meta_data = OrderedDict()
mess_cols = []
json_data = []
for i, child in enumerate(root):
tag = child.tag
if tag == 'KOPF' or tag == 'ZEIT':
for subchild in child:
value = subchild.text
if subchild.tag in ['ANFANG', 'ENDE']:
value = unix_time_millis(datetime.datetime.strptime(value, '%Y-%m-%d %H:%M:%S'))
meta_data[subchild.tag] = value
elif tag == 'DATEN':
for subchild in child:
if subchild.tag == 'SPALTE':
col = subchild.text
index = subchild.attrib.get('messstelle', '0')
mess_cols.append('.'.join([col, index]))
else:
json_data.append(xmltodict.parse(ET.tostring(child, encoding='utf-8', method='xml')))
meta_data['json'] = json.dumps(json_data, ensure_ascii=False, separators=(';', ':')).replace(',', '.').replace(';', ',')
return meta_data, mess_cols
meta, mess_cols = parse_meta_data('data/Maschine 77 (1045670479)/main.xml')
mess_cols
data = pd.read_csv('data/Maschine 77 (1045670479)/mess.txt', sep='\t', header=0, names=mess_cols)
data.head()
data.shape
data = data.rename(columns={'LAENGE.1': 'LAENGE'})
df = data.melt(id_vars=['LAENGE'], var_name='VARIABLE', value_name='VAL')
df[df.VAL == '-999;-999;-999;-999;-999;-999;-999;-999;-999;-999'].head()
# %time df.LAENGE = df.LAENGE.str.replace(',', '.')
# %time df.VAL = df.VAL.str.replace(',', '.')
def column_to_rows(df, column, sep):
def _duplicate(new_row, value):
copy = new_row.copy()
copy[column] = value
return copy
def _cell_to_rows(row, new_rows):
split_row = row[column].split(sep)
new_row = row.to_dict()
new_rows += [_duplicate(new_row, value) for value in split_row]
new_rows = []
df.apply(lambda row: _cell_to_rows(row, new_rows), axis=1)
new_df = pd.DataFrame(new_rows)
return new_df
# %time df2 = column_to_rows(df, 'VAL', ';')
df2.head()
df2.dtypes
# %time df2.LAENGE = pd.to_numeric(df2.LAENGE, errors='coerce')
# %time df2.VAL = pd.to_numeric(df2.VAL, errors='coerce')
df2.dtypes
df2.shape
split = df2.VARIABLE.str.rsplit('.')
df2['VARIABLE'] = split.str.get(0)
df2['MESSSTELLE'] = split.str.get(1).astype(np.int8)
df2.head()
df2[(df2.VARIABLE=='DURCH_MITTEL') & (df2.LAENGE==10.0)]
df2.dtypes
tuple(df2.head().VARIABLE.values)
| hierarchy-data/specs-scraper/Ingest_Data_To_Casandra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center/><h3/><font color='green'/>Bibliotecas
# +
#Biblioteca necessária
#pip install selenium
# -
#espera algo
from selenium.webdriver.support.ui import WebDriverWait
# Configurações
from selenium.webdriver.firefox.options import Options
#envia comandos
from selenium.webdriver.common.keys import Keys
from selenium.webdriver import Firefox
from selenium import webdriver
import getpass
import time
# <center/><h3/><font color='green'/>Pega Usuário e Senha
usuario = input("Diga seu usuário: ")
senha = getpass.getpass("Digite sua senha: ")
# <center/><h3/><font color='green'/>Seta as configurações e Abre a página
# +
# Grab content from URL (Pegar conteúdo HTML a partir da URL)
url = "https://www.instagram.com"
# todas as opções disponíveis
option = Options()
#Sete False para aparecer a págia web
option.headless = False
#Abre a aba do navegador
driver = webdriver.Firefox(options=option)
#Define o tamanho da tela do navegador
driver.set_window_size(448,708)
#Define a posição da janela do navegador
driver.set_window_position(800,0)
#Envia o url do instagram para o navegador acessar
driver.get(url)
# -
# <center/><h3/><font color='green'/>Seta Usuário e Senha
# +
def esperar_campo(firefox):
return driver.find_element_by_name("username")
carregando = WebDriverWait(driver, 5).until(esperar_campo)
# Insere usuário no campo
name_campo = driver.find_element_by_name("username")
name_campo.clear()
name_campo.send_keys(usuario)
# Insere senha no campo
senha_campo = driver.find_element_by_name("password")
senha_campo.clear()
senha_campo.send_keys(<PASSWORD>ha)
# Aperta ENTER
senha_campo.send_keys(Keys.ENTER)
# -
# <center/><h3/><font color='green'/>Tira caixa de diálogo
# +
#Variáveis que vou precisar, elas trazem informações do código da página!
dialog_box = "//div[@class='piCib']"
button_dialog_box = "//button[@class='aOOlW HoLwm ']"
# Funcao para esperar caso o que foi pedido ainda não ser encontrado, espera 5s se for
def espera_dialog(firefox):
return driver.find_element_by_xpath(dialog_box)
# Espera janela de dialogo inicial
esperando_jane_dialog = WebDriverWait(driver, 10).until(espera_dialog)
# Aperta para fechar caixa de dialogo
driver.find_element_by_xpath(button_dialog_box).click()
# -
#
# <center/><h3/><font color='green'/>Visualiza Story's
# <center><font size='2'>Periodicamente o nome das classes dos botões são trocados </font></center>
#
# +
#Variáveis do tamanho da tela
window_big = "//button[@class='jZyv1 H-yu6']"
window_little = "//button[@class='OE3OK ']"
# Procedimento: Aperta para abrir um story
def open_story ():
try:
#Tela Reduzida
driver.find_element_by_xpath(window_little).click()
except:
#Tela Maximizada
driver.find_element_by_xpath(window_big).click()
# -
#Abre o story
open_story()
# <center/>Passa as janelas do story para a direita
# +
#window_story: Diz se os story's ainda estão abertos
window_story ="//section[@class='_8XqED carul']"
#button_story_pass: É o endereço do botão para passar o story
button_story_pass ="//button[@class='ow3u_']"
#Abre os story's e vai passando
while(True):
# Enquanto ainda tiver story, tente apertar no botão, se der erro, espere
try:
while(driver.find_element_by_xpath(window_story)):
driver.find_element_by_xpath(button_story_pass).click()
time.sleep(1)
except Exception as e:
try:
time.sleep(1.5)
driver.find_element_by_xpath(button_story_pass).click()
except Exception as e:
# Normalmente, o código vem pra cá quando acabam os story e precisamos reabrir
# Porém, se fizermos isso sem atualizar a página antes, não mudará nada.
# Sendo assim, coloquei um tempo de espera de 8s e um refresh para atualizar o navegador.
driver.refresh()
time.sleep(8)
open_story()
# -
#
# <center/><h3/><font color='green'/>Curte Publicações
#
def curte_publicacoes():
a = 0
try:
driver.find_element_by_css_selector("article._8Rm4L:nth-child(1n) > div:nth-child(3) > section:nth-child(1) > span:nth-child(1) > button:nth-child(1) > svg:nth-child(1)[aria-label='Curtir']").click()
except Exception as e:
#print(driver.find_element_by_css_selector("article._8Rm4L:nth-child(1n) > div:nth-child(3) > section:nth-child(1) > span:nth-child(1) > button:nth-child(1) > svg:nth-child(1)[aria-label='Curtir']"))
#a+= 1
#print(a)
pass
#Desde página
aux = 1
while(True):
aux += 100
driver.execute_script(f'window.scrollTo(0,{aux})')
curte_publicacoes()
#
# <center/><h3/><font color='green'/>Finaliza as Operações
#
driver.quit()
| webScraping-instagram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/THargreaves/beginners-python/blob/master/session_three/session_three_solutions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# <center>Spotted a mistake? Report it <a href="https://github.com/THargreaves/beginners-python/issues/new">here</a></center>
# # Beginner's Python—Session Three Homework Solutions
# ## Comparisons and Boolean Arithmetic
# Confirm that applying `or` to two `True` values does indeed return `True`
True or True
# _Note: Creating interesting questions for Boolean variables is difficult since there's not much to them, and so the following questions are a bit more abstract in nature. The first gives a useful application of comparisons for data analysis, and the second offers a way of dealing with the shortcomings of how computers store numbers. Feel free to skip to the next section if you only want to know the bare bones of Python._
# Create a list of seven numbers called `nums` and find their mean (sum divided by length), saving this in the variable `mean`
nums = [1, 1, 3, 4, 8, 10, 14]
mean = sum(nums) / len(nums)
print(mean)
# Run the code below which will calculate the median value of the list (the middle value of the list when they are placed in order)
median = sorted(nums)[3]
print(median)
# Use a Boolean expression to see which of the mean and median is larger
median < mean
# A rule-of-thumb for estimating how skewed a collection of numbers is to compare the mean and the median. When the mean is larger than the median then your data is likely positively skewed. For the reverse, a negative skew is more likely. Is your data more positively or negatively skewed?
# 
# This data appears to have a positively skewed. An example of a real distribution that is positively skewed is wealth.
# Computers are not able to store decimal numbers with an infinite number of decimal places. This means that as we manipulate decimal numbers, small rounding errors can occur. For example:
print(1 / 49 * 49)
print(1 / 49 * 49 == 1)
# This can be incredibly annoying. We know $\frac{1}{49} \times 49$ and $1$ are the same thing yet "computer says no". We can get around this by checking that two numbers are near each other, rather than exactly equal. We typically do this by taking the difference of two numbers, making this difference positive using the `abs()` function, and then checking that this postive difference is less than some small tolerance that we set. To warm up, start my using `abs()` on the numbers $1$ and $-1$ to see that it does indeed make any value you pass into it positive
print(abs(1))
print(abs(-1))
# Now, use the tolerance defined below to check if $\frac{1}{49}\times 49$ and $1$ are indeed the same value (within said tolerance)
tolerance = 10 ** (-8) # 0.000000001 - i.e. very small tolerance
abs( (1 / 49 * 49) - 1 ) < tolerance
# ## Control Flow
# Ask for user input. If it lies between 8 and 12, print "Approximately 10" (if you want to show off, you can do this with one condition using `abs()`)
# +
num = float(input("Enter a number: "))
# method one
if num > 8 and num < 12:
print("Approximately 10")
# method two
if abs(10 - num) < 2: # i.e. distance to 10 is less than two
print("Approximately 10")
# -
# Ask the user whether it will rain, requesting that they answer either "yes" or "no". If their answer is "yes" print "Better bring an umbrella", and if it's no print "That's great to hear". If they answer anything but these two options, print "I don't understand"
rain = input("Will it rain? (y/n) ")
if rain == "y":
print("Better bring an umbrella")
elif rain == "n":
print("That's great to hear")
else:
print("I don't understand")
# Use `or` twice to check whether a number inputted by the user is one of $6$, $28$, or $496$. If it is, print "Perfect..."
num = int(input("Enter a number: "))
if num == 6 or num == 28 or num == 496:
print("Perfect...")
# Python has a useful command `in`, which lets you check if a value is in a list. For example, `5 in [4, 5, 6]` will return `True`. Use this to simplify the previous solution
num = int(input("Enter a number: "))
if num in [6, 28, 496]:
print("Perfect...")
# Ask the user for a number and store this as a float called `num`. Check whether the number could be stored as an integer by seeing if `num` and `int(num)` have the same value. If so, convert `num` to an integer type. (Recall: running `int()` on a decimal number will round it down to the nearest integer)
num = float(input("Enter a number: "))
if num == int(num):
num = int(num)
print(type(num))
# ## While Loops
# Print the first ten squared numbers (you can either times a number by itself to get a square or use `x ** 2`)
n = 1
while n <= 10:
print(n ** 2)
n = n + 1 # or n += 1
# Define a string `password` containing a password (not a real one, obviously!). Use a while loop to allow a user, 3 guesses at this password. You might want to keep the number of guesses in a variable called `guesses`, initial set to $0$. You can also use a variable `correct` to keep track of whether they guessed correctly. This should most likely default to `False` and then get set to `True` if they guess correctly
# +
password = "<PASSWORD>"
guesses = 0
correct = False
while guesses < 3 and not correct:
guess = input("What is the password? ")
if guess == password:
correct = True
else:
guesses = guesses + 1
if correct:
print("Access granted")
else:
print("Access denied")
# -
# The game FizzBuzz goes as follows:
#
# * Count up from one
# * Replace each multiple of 3 with the word "Fizz"
# * Replace each multiple of 5 with the word "Buzz"
# * Replace each multiple of 15 with the word "FizzBuzz"
# * Otherwise, print the number
#
# For example, the sequence begins:
#
# ```
# 1
# 2
# Fizz
# 4
# Buzz
# Fizz
# 7
# ...
# 14
# FizzBuzz
# 16
# ...
# ```
#
# Ask for a user inputted number and place FizzBuzz up to that number. Note you can check if `x` is divible by `y` using `x % y == 0`. For example `5 % 2 == 0` is `False` but `9 % 3` is `True`
final_number = int(input("Play FizzBuzz until: "))
n = 1
while n <= final_number:
if n % 15 == 0:
print("FizzBuzz")
elif n % 5 == 0:
print("Buzz")
elif n % 3 == 0:
print("Fizz")
else:
print(n)
n = n + 1
# The Collatz conjecture goes as follows
#
# > Take a number $n$. If it is even, halve it, if it is odd, times it by $3$ and add $1$. Repeat this process. You will eventually reach the number $1$
#
# We have not proved that this is definitely true but we have good reason to believe it is. You Python to take any starting number and print out the sequence of numbers generated by the Collatz conjecture, stopping when you reach one
# +
n = int(input("Choose a starting number: "))
while n != 1:
print(n)
if n % 2 == 0:
n = n // 2 # https://www.geeksforgeeks.org/division-operator-in-python/
else:
n = 3 * n + 1
print(n)
# -
# What is the first power of two to surpass one million? Use a while loop to find out (Hint: start with a variable `n = 1` and set `n = n * 2` at each iteration, stopping when this goes above one million)
n = 1
while n <= 10 ** 6:
n = n * 2
print(n)
| session_three/session_three_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Text provided under a Creative Commons Attribution license, CC-BY. All code is made available under the FSF-approved BSD-3 license. (c) <NAME>, <NAME> 2017. Thanks to NSF for support via CAREER award #1149784.
# -
# [@LorenaABarba](https://twitter.com/LorenaABarba)
# 12 steps to Navier–Stokes
# =====
# ***
# The final two steps in this interactive module teaching beginning [CFD with Python](https://bitbucket.org/cfdpython/cfd-python-class) will both solve the Navier–Stokes equations in two dimensions, but with different boundary conditions.
#
# The momentum equation in vector form for a velocity field $\vec{v}$ is:
#
# $$\frac{\partial \vec{v}}{\partial t}+(\vec{v}\cdot\nabla)\vec{v}=-\frac{1}{\rho}\nabla p + \nu \nabla^2\vec{v}$$
#
# This represents three scalar equations, one for each velocity component $(u,v,w)$. But we will solve it in two dimensions, so there will be two scalar equations.
#
# Remember the continuity equation? This is where the [Poisson equation](./13_Step_10.ipynb) for pressure comes in!
# Step 11: Cavity Flow with Navier–Stokes
# ----
# ***
# Here is the system of differential equations: two equations for the velocity components $u,v$ and one equation for pressure:
# $$\frac{\partial u}{\partial t}+u\frac{\partial u}{\partial x}+v\frac{\partial u}{\partial y} = -\frac{1}{\rho}\frac{\partial p}{\partial x}+\nu \left(\frac{\partial^2 u}{\partial x^2}+\frac{\partial^2 u}{\partial y^2} \right) $$
#
#
# $$\frac{\partial v}{\partial t}+u\frac{\partial v}{\partial x}+v\frac{\partial v}{\partial y} = -\frac{1}{\rho}\frac{\partial p}{\partial y}+\nu\left(\frac{\partial^2 v}{\partial x^2}+\frac{\partial^2 v}{\partial y^2}\right) $$
#
# $$\frac{\partial^2 p}{\partial x^2}+\frac{\partial^2 p}{\partial y^2} = -\rho\left(\frac{\partial u}{\partial x}\frac{\partial u}{\partial x}+2\frac{\partial u}{\partial y}\frac{\partial v}{\partial x}+\frac{\partial v}{\partial y}\frac{\partial v}{\partial y} \right)$$
# From the previous steps, we already know how to discretize all these terms. Only the last equation is a little unfamiliar. But with a little patience, it will not be hard!
# ### Discretized equations
# First, let's discretize the $u$-momentum equation, as follows:
# $$
# \begin{split}
# & \frac{u_{i,j}^{n+1}-u_{i,j}^{n}}{\Delta t}+u_{i,j}^{n}\frac{u_{i,j}^{n}-u_{i-1,j}^{n}}{\Delta x}+v_{i,j}^{n}\frac{u_{i,j}^{n}-u_{i,j-1}^{n}}{\Delta y} = \\
# & \qquad -\frac{1}{\rho}\frac{p_{i+1,j}^{n}-p_{i-1,j}^{n}}{2\Delta x}+\nu\left(\frac{u_{i+1,j}^{n}-2u_{i,j}^{n}+u_{i-1,j}^{n}}{\Delta x^2}+\frac{u_{i,j+1}^{n}-2u_{i,j}^{n}+u_{i,j-1}^{n}}{\Delta y^2}\right)
# \end{split}
# $$
# Similarly for the $v$-momentum equation:
# $$
# \begin{split}
# &\frac{v_{i,j}^{n+1}-v_{i,j}^{n}}{\Delta t}+u_{i,j}^{n}\frac{v_{i,j}^{n}-v_{i-1,j}^{n}}{\Delta x}+v_{i,j}^{n}\frac{v_{i,j}^{n}-v_{i,j-1}^{n}}{\Delta y} = \\
# & \qquad -\frac{1}{\rho}\frac{p_{i,j+1}^{n}-p_{i,j-1}^{n}}{2\Delta y}
# +\nu\left(\frac{v_{i+1,j}^{n}-2v_{i,j}^{n}+v_{i-1,j}^{n}}{\Delta x^2}+\frac{v_{i,j+1}^{n}-2v_{i,j}^{n}+v_{i,j-1}^{n}}{\Delta y^2}\right)
# \end{split}
# $$
# Finally, the discretized pressure-Poisson equation can be written thus:
# $$
# \begin{split}
# & \frac{p_{i+1,j}^{n}-2p_{i,j}^{n}+p_{i-1,j}^{n}}{\Delta x^2}+\frac{p_{i,j+1}^{n}-2p_{i,j}^{n}+p_{i,j-1}^{n}}{\Delta y^2} = \\
# & \qquad \rho \left[ \frac{1}{\Delta t}\left(\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}+\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right) -\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x} - 2\frac{u_{i,j+1}-u_{i,j-1}}{2\Delta y}\frac{v_{i+1,j}-v_{i-1,j}}{2\Delta x} - \frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right]
# \end{split}
# $$
# You should write these equations down on your own notes, by hand, following each term mentally as you write it.
#
# As before, let's rearrange the equations in the way that the iterations need to proceed in the code. First, the momentum equations for the velocity at the next time step.
#
# The momentum equation in the $u$ direction:
#
# $$
# \begin{split}
# u_{i,j}^{n+1} = u_{i,j}^{n} & - u_{i,j}^{n} \frac{\Delta t}{\Delta x} \left(u_{i,j}^{n}-u_{i-1,j}^{n}\right) - v_{i,j}^{n} \frac{\Delta t}{\Delta y} \left(u_{i,j}^{n}-u_{i,j-1}^{n}\right) \\
# & - \frac{\Delta t}{\rho 2\Delta x} \left(p_{i+1,j}^{n}-p_{i-1,j}^{n}\right) \\
# & + \nu \left(\frac{\Delta t}{\Delta x^2} \left(u_{i+1,j}^{n}-2u_{i,j}^{n}+u_{i-1,j}^{n}\right) + \frac{\Delta t}{\Delta y^2} \left(u_{i,j+1}^{n}-2u_{i,j}^{n}+u_{i,j-1}^{n}\right)\right)
# \end{split}
# $$
#
# The momentum equation in the $v$ direction:
#
# $$
# \begin{split}
# v_{i,j}^{n+1} = v_{i,j}^{n} & - u_{i,j}^{n} \frac{\Delta t}{\Delta x} \left(v_{i,j}^{n}-v_{i-1,j}^{n}\right) - v_{i,j}^{n} \frac{\Delta t}{\Delta y} \left(v_{i,j}^{n}-v_{i,j-1}^{n})\right) \\
# & - \frac{\Delta t}{\rho 2\Delta y} \left(p_{i,j+1}^{n}-p_{i,j-1}^{n}\right) \\
# & + \nu \left(\frac{\Delta t}{\Delta x^2} \left(v_{i+1,j}^{n}-2v_{i,j}^{n}+v_{i-1,j}^{n}\right) + \frac{\Delta t}{\Delta y^2} \left(v_{i,j+1}^{n}-2v_{i,j}^{n}+v_{i,j-1}^{n}\right)\right)
# \end{split}
# $$
# Almost there! Now, we rearrange the pressure-Poisson equation:
#
# $$
# \begin{split}
# p_{i,j}^{n} = & \frac{\left(p_{i+1,j}^{n}+p_{i-1,j}^{n}\right) \Delta y^2 + \left(p_{i,j+1}^{n}+p_{i,j-1}^{n}\right) \Delta x^2}{2\left(\Delta x^2+\Delta y^2\right)} \\
# & -\frac{\rho\Delta x^2\Delta y^2}{2\left(\Delta x^2+\Delta y^2\right)} \\
# & \times \left[\frac{1}{\Delta t}\left(\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}+\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right)-\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x} -2\frac{u_{i,j+1}-u_{i,j-1}}{2\Delta y}\frac{v_{i+1,j}-v_{i-1,j}}{2\Delta x}-\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right]
# \end{split}
# $$
# The initial condition is $u, v, p = 0$ everywhere, and the boundary conditions are:
#
# $u=1$ at $y=2$ (the "lid");
#
# $u, v=0$ on the other boundaries;
#
# $\frac{\partial p}{\partial y}=0$ at $y=0$;
#
# $p=0$ at $y=2$
#
# $\frac{\partial p}{\partial x}=0$ at $x=0,2$
#
# Implementing Cavity Flow
# ----
#
import numpy
from matplotlib import pyplot, cm
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib inline
# +
nx = 41
ny = 41
nt = 500
nit = 50
c = 1
dx = 2 / (nx - 1)
dy = 2 / (ny - 1)
x = numpy.linspace(0, 2, nx)
y = numpy.linspace(0, 2, ny)
X, Y = numpy.meshgrid(x, y)
rho = 1
nu = .1
dt = .001
u = numpy.zeros((ny, nx))
v = numpy.zeros((ny, nx))
p = numpy.zeros((ny, nx))
b = numpy.zeros((ny, nx))
# -
# The pressure Poisson equation that's written above can be hard to write out without typos. The function `build_up_b` below represents the contents of the square brackets, so that the entirety of the PPE is slightly more manageable.
def build_up_b(b, rho, dt, u, v, dx, dy):
b[1:-1, 1:-1] = (rho * (1 / dt *
((u[1:-1, 2:] - u[1:-1, 0:-2]) /
(2 * dx) + (v[2:, 1:-1] - v[0:-2, 1:-1]) / (2 * dy)) -
((u[1:-1, 2:] - u[1:-1, 0:-2]) / (2 * dx))**2 -
2 * ((u[2:, 1:-1] - u[0:-2, 1:-1]) / (2 * dy) *
(v[1:-1, 2:] - v[1:-1, 0:-2]) / (2 * dx))-
((v[2:, 1:-1] - v[0:-2, 1:-1]) / (2 * dy))**2))
return b
# The function `pressure_poisson` is also defined to help segregate the different rounds of calculations. Note the presence of the pseudo-time variable `nit`. This sub-iteration in the Poisson calculation helps ensure a divergence-free field.
def pressure_poisson(p, dx, dy, b):
pn = numpy.empty_like(p)
pn = p.copy()
for q in range(nit):
pn = p.copy()
p[1:-1, 1:-1] = (((pn[1:-1, 2:] + pn[1:-1, 0:-2]) * dy**2 +
(pn[2:, 1:-1] + pn[0:-2, 1:-1]) * dx**2) /
(2 * (dx**2 + dy**2)) -
dx**2 * dy**2 / (2 * (dx**2 + dy**2)) *
b[1:-1,1:-1])
p[:, -1] = p[:, -2] # dp/dx = 0 at x = 2
p[0, :] = p[1, :] # dp/dy = 0 at y = 0
p[:, 0] = p[:, 1] # dp/dx = 0 at x = 0
p[-1, :] = 0 # p = 0 at y = 2
return p
# Finally, the rest of the cavity flow equations are wrapped inside the function `cavity_flow`, allowing us to easily plot the results of the cavity flow solver for different lengths of time.
def cavity_flow(nt, u, v, dt, dx, dy, p, rho, nu):
un = numpy.empty_like(u)
vn = numpy.empty_like(v)
b = numpy.zeros((ny, nx))
for n in range(nt):
un = u.copy()
vn = v.copy()
b = build_up_b(b, rho, dt, u, v, dx, dy)
p = pressure_poisson(p, dx, dy, b)
u[1:-1, 1:-1] = (un[1:-1, 1:-1]-
un[1:-1, 1:-1] * dt / dx *
(un[1:-1, 1:-1] - un[1:-1, 0:-2]) -
vn[1:-1, 1:-1] * dt / dy *
(un[1:-1, 1:-1] - un[0:-2, 1:-1]) -
dt / (2 * rho * dx) * (p[1:-1, 2:] - p[1:-1, 0:-2]) +
nu * (dt / dx**2 *
(un[1:-1, 2:] - 2 * un[1:-1, 1:-1] + un[1:-1, 0:-2]) +
dt / dy**2 *
(un[2:, 1:-1] - 2 * un[1:-1, 1:-1] + un[0:-2, 1:-1])))
v[1:-1,1:-1] = (vn[1:-1, 1:-1] -
un[1:-1, 1:-1] * dt / dx *
(vn[1:-1, 1:-1] - vn[1:-1, 0:-2]) -
vn[1:-1, 1:-1] * dt / dy *
(vn[1:-1, 1:-1] - vn[0:-2, 1:-1]) -
dt / (2 * rho * dy) * (p[2:, 1:-1] - p[0:-2, 1:-1]) +
nu * (dt / dx**2 *
(vn[1:-1, 2:] - 2 * vn[1:-1, 1:-1] + vn[1:-1, 0:-2]) +
dt / dy**2 *
(vn[2:, 1:-1] - 2 * vn[1:-1, 1:-1] + vn[0:-2, 1:-1])))
u[0, :] = 0
u[:, 0] = 0
u[:, -1] = 0
u[-1, :] = 1 # set velocity on cavity lid equal to 1
v[0, :] = 0
v[-1, :] = 0
v[:, 0] = 0
v[:, -1] = 0
return u, v, p
# Let's start with `nt = 100` and see what the solver gives us:
u = numpy.zeros((ny, nx))
v = numpy.zeros((ny, nx))
p = numpy.zeros((ny, nx))
b = numpy.zeros((ny, nx))
nt = 3000
u, v, p = cavity_flow(nt, u, v, dt, dx, dy, p, rho, nu)
# +
# -
# You can see that two distinct pressure zones are forming and that the spiral pattern expected from lid-driven cavity flow is beginning to form. Experiment with different values of `nt` to see how long the system takes to stabilize.
u = numpy.zeros((ny, nx))
v = numpy.zeros((ny, nx))
p = numpy.zeros((ny, nx))
b = numpy.zeros((ny, nx))
nt = 700
u, v, p = cavity_flow(nt, u, v, dt, dx, dy, p, rho, nu)
fig = pyplot.figure(figsize=(11, 7), dpi=100)
pyplot.contourf(X, Y, p, alpha=0.5, cmap=cm.viridis)
pyplot.colorbar()
pyplot.contour(X, Y, p, cmap=cm.viridis)
pyplot.quiver(X[::2, fc00:db20:35b:7399::5], Y[::2, fc00:db20:35b:7399::5], u[::2, fc00:db20:35b:7399::5], v[::2, ::2])
pyplot.xlabel('X')
pyplot.ylabel('Y');
# The quiver plot shows the magnitude of the velocity at the discrete points in the mesh grid we created.
# (We're actually only showing half of the points because otherwise it's a bit of a mess. The `X[::2, ::2]` syntax above is a convenient way to ask for every other point.)
#
# Another way to visualize the flow in the cavity is to use a `streamplot`:
fig = pyplot.figure(figsize=(11, 7), dpi=100)
pyplot.contourf(X, Y, p, alpha=0.5, cmap=cm.viridis)
pyplot.colorbar()
pyplot.contour(X, Y, p, cmap=cm.viridis)
pyplot.streamplot(X, Y, u, v)
pyplot.xlabel('X')
pyplot.ylabel('Y');
# ## Learn More
# The interactive module **12 steps to Navier–Stokes** is one of several components of the Computational Fluid Dynamics class taught by Prof. <NAME> in Boston University between 2009 and 2013.
#
# For a sample of what the other components of this class are, you can explore the **Resources** section of the Spring 2013 version of [the course's Piazza site](https://piazza.com/bu/spring2013/me702/resources).
#
# ***
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
# > (The cell above executes the style for this notebook.)
| lessons/14_Step_11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="iExk8GQEl-0m" colab_type="text"
# In this tutorial, you will customize a `pre-trained` model with `feature extraction` and `fine-tune`.
# + id="981CQegDjql0" colab_type="code" colab={}
# # !pip install -q tf-nightly
# + id="Pp3dn5JOj_5L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="7a83fa8d-3660-436b-f0df-7d3a46dcc5ab" executionInfo={"status": "ok", "timestamp": 1581062562454, "user_tz": -480, "elapsed": 3582, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
import tensorflow as tf
import matplotlib.pyplot as plt
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
print("Tensorflow Version: {}".format(tf.__version__))
print("GPU {} available.".format("is" if tf.config.experimental.list_physical_devices("GPU") else "not"))
# + [markdown] id="Ce028wYSmuwI" colab_type="text"
# # Data Preprocessing
# + [markdown] id="TamYuPIDsMM-" colab_type="text"
# ## Download Datasets
# + [markdown] id="hM4KTTSUnkPe" colab_type="text"
# Here we are going to use the `cats_vs_dogs` datasets via `tfds.load()` method. However, the dataset didn't be split before so we have to split the dataset into three subsets.
# + id="vHhGMdSAkerF" colab_type="code" colab={}
(raw_train, raw_validation, raw_test), metadata = tfds.load(
name="cats_vs_dogs",
split=['train[:80%]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True)
# + id="dMmk5P1IoIx1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 493} outputId="b33856a3-f2c9-4ffb-84ee-700fd3a9fc79" executionInfo={"status": "ok", "timestamp": 1581062563735, "user_tz": -480, "elapsed": 4746, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
raw_train, metadata
# + id="7ZftZDVzq-4t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b1b1ab6c-6f1b-4cd3-a853-594ff4c3c2bc" executionInfo={"status": "ok", "timestamp": 1581062563736, "user_tz": -480, "elapsed": 4639, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
# a method
get_label_name = metadata.features['label'].int2str
get_label_name
# + id="qZhgH_gMrPZP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 758} outputId="051b6c80-24ff-48b4-f78a-85bd7b67f904" executionInfo={"status": "ok", "timestamp": 1581062564478, "user_tz": -480, "elapsed": 5364, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
for image, label in raw_train.take(3):
plt.imshow(image)
plt.title(get_label_name(label) + ", label: {}".format(label))
plt.axis("off")
plt.show()
# + [markdown] id="PeubNbT7sOgs" colab_type="text"
# ## Data Generators
# + id="M581SMfer6cS" colab_type="code" colab={}
IMG_SIZE = 160
def resize_normalize(image, label):
img = tf.cast(image, tf.float32)
img = tf.image.resize(img, size=(IMG_SIZE, IMG_SIZE))
img = (img / 127.5) - 1.0
return img, label
# + [markdown] id="dNspWRX5s40J" colab_type="text"
# Apply the preprocessing function to the datasets.
# + id="1zZtTK8uszMc" colab_type="code" colab={}
train = raw_train.map(resize_normalize)
validation = raw_validation.map(resize_normalize)
test = raw_test.map(resize_normalize)
# + [markdown] id="GpedwZJItJFw" colab_type="text"
# Shuffle and batch the datasets.
# + id="TgTewdDNtHDV" colab_type="code" colab={}
BATCH_SIZE = 32
BUFFER_SIZE = 1000
# + id="Q-hv26TFtZl6" colab_type="code" colab={}
train_batches = train.shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
validation_batches = validation.batch(BATCH_SIZE)
test_batches = test.batch(BATCH_SIZE)
# + [markdown] id="HLAlO2sTtqdh" colab_type="text"
# Inspect the train dataset.
# + id="PlfeGjpUtob7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="e40307a3-3a1b-4bb9-b4f1-253b92001ce9" executionInfo={"status": "ok", "timestamp": 1581062565342, "user_tz": -480, "elapsed": 6076, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
for batch_imgs, batch_labels in train_batches.take(1):
print(batch_imgs.shape, batch_labels.shape)
print("Batch Data: {}".format(batch_labels))
# + [markdown] id="PvshyxrWuj38" colab_type="text"
# # Create the Base Model From a Pre-Trained Model
# + [markdown] id="8Lf93eJlwx1d" colab_type="text"
# Here we create a base model from a pretrained model named MobileNetV2, which was created by Google and was trained on a large image dataset ImageNets.
#
# First, you need to decide which layer of MobileNetV2 is used for feature extraction. The very last layer or the top layer of the MobileNetV2 is the classification layer, which outputs the probabilities of 1000 categories. We don't need the very last layer, instead, we need the feature extraction layer which is the previous one before the classification layer, or more specifically, the layer before the flatten operation. This layer is also called the `bottleneck layer`.
#
# In `TF.keras`, you can simply load the model without the classification layers using a parameter `include_top` which is set to `False` if you don't want to load the final layer.
# + id="ajw6O7Jet6uz" colab_type="code" colab={}
IMG_SHAPE = (IMG_SIZE, IMG_SIZE, 3)
# load the pre-trained model MobileNetV2
base_model = tf.keras.applications.MobileNetV2(
input_shape=IMG_SHAPE,
include_top=False,
weights='imagenet')
# + [markdown] id="QYIV6TfPxfT6" colab_type="text"
# Such a feature extraction layer converts each `160x160x3` image into a `5x5x1280` block of features.
# + id="Oe4vCwVqxMNr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="672b11ff-dcea-423c-86cb-0186771db149" executionInfo={"status": "ok", "timestamp": 1581062569079, "user_tz": -480, "elapsed": 9772, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
feature_batch = base_model(batch_imgs)
feature_batch.shape
# + [markdown] id="Mk3iUkpdxr4P" colab_type="text"
# # Feature Extraction
# + [markdown] id="nO-TZCaAx6qK" colab_type="text"
# ## Freeze the Convolutional Base
# + [markdown] id="crE-9ktUxuly" colab_type="text"
# You can freeze the model (not to train it) and extract the features using it.
# + id="iYQJ0_qixjod" colab_type="code" colab={}
base_model.trainable = False
# + id="nWM3kHWPyD7n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="fd3c9338-2f2c-4a3d-b6ab-bdc57e2e1561" executionInfo={"status": "ok", "timestamp": 1581062569083, "user_tz": -480, "elapsed": 9740, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
base_model.summary()
# + [markdown] id="b3qQGw-OyKwJ" colab_type="text"
# ## Add a Classification Layer
# + [markdown] id="voMmwsYAzDKt" colab_type="text"
# To generate a prediction from a block of features, you can average over the spatial `5x5` locations, using a `tf.keras.layers.GlobalAveragePooling2D` layer to convert the features into a 1280-size vector.
# + id="M03cGwW3yHRG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="48c29ab7-0107-4831-8fb6-b594b3171224" executionInfo={"status": "ok", "timestamp": 1581062569084, "user_tz": -480, "elapsed": 9698, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
global_average_layer = tf.keras.layers.GlobalAveragePooling2D()
feature_batch_average = global_average_layer(feature_batch)
feature_batch_average.shape
# + [markdown] id="bjPKqdtfb0Lm" colab_type="text"
# Here the activation function could not be set, the result is the logit number which is positive for class 1 and negative for class 2.
# + id="byigkAJfzVpJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f8037cf1-3164-4ce6-f66e-47fbe816a6a1" executionInfo={"status": "ok", "timestamp": 1581062569085, "user_tz": -480, "elapsed": 9681, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
prediction_layer = tf.keras.layers.Dense(units=1)
prediction_batch = prediction_layer(feature_batch_average)
prediction_batch.shape
# + [markdown] id="D586ebPdb4Hs" colab_type="text"
# Stack the partial network into a bigger one.
# + id="emHP2LRWzy0R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="788bc0a2-45f8-49ff-9b25-09cde4abb0b7" executionInfo={"status": "ok", "timestamp": 1581062570662, "user_tz": -480, "elapsed": 11240, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
def build_model(inputs):
fe = base_model(inputs)
gv = global_average_layer(fe)
new_cls_res = prediction_layer(gv)
return new_cls_res
inputs = tf.keras.Input(shape=IMG_SHAPE)
outputs = build_model(inputs)
model = tf.keras.Model(inputs, outputs)
model.summary()
# + [markdown] id="nV0RlUhwdkby" colab_type="text"
# Only 1281 variables are available for training and the main architecture body 2.5M ones are frozen.
# + id="-p39ZouGd0N6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="700d20fc-cb40-486c-acc8-483a9ee27efb" executionInfo={"status": "ok", "timestamp": 1581062570663, "user_tz": -480, "elapsed": 11223, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
len(model.trainable_variables)
# + id="7cmQzsdId8eT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="445c3aac-6e7f-40d1-9abb-ffc6653e0920" executionInfo={"status": "ok", "timestamp": 1581062570664, "user_tz": -480, "elapsed": 11202, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
for v in model.trainable_variables:
print(v.name)
# + [markdown] id="F22_RT8weEjg" colab_type="text"
# As you see, the trainable variables are only related to the final dense layer.
# + [markdown] id="ExvBVkTWdAAW" colab_type="text"
# ## Compile the Model
# + id="vZvICisscnM4" colab_type="code" colab={}
model.compile(loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.Adam(learning_rate=1e-4),
metrics=[tf.keras.metrics.BinaryAccuracy()])
# + [markdown] id="0vNnVpmQeUL4" colab_type="text"
# ## Train the Model
# + id="nDd-cz7JdcLb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="b0946507-87b9-4d1b-b35a-c299ca81b81c" executionInfo={"status": "ok", "timestamp": 1581062995646, "user_tz": -480, "elapsed": 436089, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
history = model.fit(train_batches, epochs=10, validation_data=validation_batches)
# + id="knmsbqNAfDW6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="60640a89-4bb7-4858-d376-40b9799950e0" executionInfo={"status": "ok", "timestamp": 1581062995651, "user_tz": -480, "elapsed": 436068, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
history.history.keys()
# + id="v8-3_J15hcst" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="b13badb2-7300-47bd-cd8f-e3306ba839ab" executionInfo={"status": "ok", "timestamp": 1581063516340, "user_tz": -480, "elapsed": 964, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
plt.figure(figsize=(12, 6))
plt.subplot(1, 2, 1)
plt.plot(history.history['binary_accuracy'], label="Training")
plt.plot(history.history['val_binary_accuracy'], label='Validation')
plt.title("Accuracy")
plt.legend(loc="lower right")
plt.subplot(1, 2, 2)
plt.plot(history.history['loss'], label="Training")
plt.plot(history.history['val_loss'], label='Validation')
plt.title("Loss")
plt.legend(loc="upper right")
plt.show()
# + [markdown] id="Vv9PcT5SiPJm" colab_type="text"
# # Fine Tuning
# + [markdown] id="CJEc9UMalaBU" colab_type="text"
# In the previous training, adding a classifier layer on the top of a base model and then train the model only on the newly added layer. Such training can't update the weights of the base model. However, the performance of the model can be further improved when training the base model. Next, we are going to train the whole model include the newly added layer on the specific dataset. Such an operation is called fine-tune. But remember fine-tuning must be after the previous job, which training the newly added layer is finished, otherwise, the gradient would be influenced too much in the pretrained base model. It might lose the generalization ability of the base model.
#
# In general, the fine-tuning is only available for the last several layers. The higher layer goes, it learns from more specific to the purpose. On the contrary, the lower layer is much generalized for any kind of task.
# + id="XMJQ-1R5lejh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="85c1b95a-84b6-4ad6-987e-537b09a10556" executionInfo={"status": "ok", "timestamp": 1581062996127, "user_tz": -480, "elapsed": 436503, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
base_model.trainable = True
# look at how many layers in the base model
print("Number of Layers in the base model: {}.".format(len(base_model.layers)))
# fine-tune from this layers
fine_tune_at = 100
# freeze all the layers before the `fine_tune_at` layer
for layer in base_model.layers[:fine_tune_at]:
layer.trainable = False
# + [markdown] id="F88invZImGwL" colab_type="text"
# ## Compile and Train the Model
# + id="hm3-oPoJlp4U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 289} outputId="57c4a9fc-3ea7-4ced-929d-86aea73d3580" executionInfo={"status": "ok", "timestamp": 1581062996128, "user_tz": -480, "elapsed": 436488, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
model.compile(loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.Adam(learning_rate=1e-5),
metrics=[tf.keras.metrics.BinaryAccuracy()])
model.summary()
# + id="CUNIzQa4mcq7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d17063bd-1f0b-4071-8e16-6c577fbce181" executionInfo={"status": "ok", "timestamp": 1581062996537, "user_tz": -480, "elapsed": 436880, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
len(model.trainable_variables)
# + [markdown] id="HUDEQPXtmgte" colab_type="text"
# As you see, the trainable variables are increasing.
# + id="Sc3aDTyzmf78" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="9114f555-9d11-4bf0-dd34-5f25be54dd7b" executionInfo={"status": "ok", "timestamp": 1581063471550, "user_tz": -480, "elapsed": 911877, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
history_fine = model.fit(train_batches, epochs=20,
initial_epoch=history.epoch[-1],
validation_data=validation_batches)
# + [markdown] id="t7hnVTkJpFBK" colab_type="text"
# ## Visualize the Training
# + id="z1Juvz_JmzQr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a54e604c-523f-4fc4-a9fb-0507c7f075c2" executionInfo={"status": "ok", "timestamp": 1581063471555, "user_tz": -480, "elapsed": 911864, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
history_fine.history.keys()
# + id="ODgkwvq0pS7X" colab_type="code" colab={}
acc = history.history['binary_accuracy'] + history_fine.history['binary_accuracy']
val_acc = history.history['val_binary_accuracy'] + history_fine.history['val_binary_accuracy']
loss = history.history['loss'] + history_fine.history['loss']
val_loss = history.history['val_loss'] + history_fine.history['val_loss']
# + id="xeI2OubZpyx-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="8edd0565-fb12-4a3e-b941-39e820e18f44" executionInfo={"status": "ok", "timestamp": 1581063523925, "user_tz": -480, "elapsed": 1239, "user": {"displayName": "\u738bDevOps", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAzLB0C3whTHAdHpq24UrEWqGtbhJElQxTU5_b_4g=s64", "userId": "04300517850278510646"}}
plt.figure(figsize=(12, 6))
plt.subplot(1, 2, 1)
plt.plot(acc, label="Training")
plt.plot(val_acc, label="Validation")
plt.ylim([0.5, 1])
plt.plot([9, 9], plt.ylim(), label='Start Fine-Tuning')
plt.title("Accuracy")
plt.legend(loc="lower right")
plt.subplot(1, 2, 2)
plt.plot(loss, label="Training")
plt.plot(val_loss, label="Validation")
plt.ylim([0., 1.])
plt.plot([9, 9], plt.ylim(), label="Start Fine-Tuning")
plt.title("Loss")
plt.legend(loc="upper right")
plt.show()
# + id="-VHcYlOsqvqn" colab_type="code" colab={}
| deep_learning/space_image/TF2Keras_Transfer_Learning_PreTrained_Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import scipy, os, glob, sys
ft_path = '/Users/jvanbaar/Dropbox (Brown)/Python'
sys.path.append(ft_path)
import FigureTools
blockDat = pd.read_csv('/Users/jvanbaar/Dropbox (Brown)/Postdoc FHL/JEROEN/SOC_STRUCT_LEARN/Study2_EyeTracking'
+'/Data/Cleaned/blockDat.csv',index_col = 0, dtype={ 'subID':str})
subIDs = blockDat['subID'].unique()
print(len(subIDs))
blockDat.head()
combs = []
for model1 in ['0','1']:
for model2 in ['0','1']:
for model3 in ['0','1']:
for model4 in ['0','1']:
combs.append(model1+model2+model3+model4)
combs = combs[1:]
# combs
niter = 5
results_Feature_RL = pd.DataFrame()
for subID in subIDs:
print(subID, end=',')
filename = glob.glob('FeatureRL/Results/results_06-Sep-2019/subInd-*_subID-*%s_niter-%i_results.csv'%(
subID, niter))[0]
# print(filename)
results = pd.read_csv(filename,header=0, index_col=None, dtype = {'subID':str})
results['subID'] = subID
results_Feature_RL = results_Feature_RL.append(results.iloc[:119,:])
results_Feature_RL = results_Feature_RL.reset_index(drop=True)
# ##### Parse feature combination
results_Feature_RL['combInd'].unique()
# ## Fix error in feature combination indices (combInd): mapping is off
mapping = dict(zip(np.arange(1,9),[4, 5, 6, 7, 12, 13, 14, 15]))
mapping
results_Feature_RL['combInd_fixed'] = results_Feature_RL['combInd'].apply(lambda x: mapping[x])
results_Feature_RL['comb'] = [combs[combInd-1] for combInd in results_Feature_RL['combInd_fixed'].values]
results_Feature_RL.head()
paramCols = []
cols = list(results_Feature_RL.columns.values)
for ni,name in enumerate(cols):
if 'param' in name:
paramCols.append(ni)
paramCols = [cols[pc] for pc in paramCols]
def penalizedModelFit(NLL,nParams, whichOne = 'both'):
AIC = 2*nParams - 2 * -NLL
BIC = nParams * np.log(64) - 2 * -NLL
# see https://en.wikipedia.org/wiki/Bayesian_information_criterion and
# https://en.wikipedia.org/wiki/Akaike_information_criterion
if whichOne == 'both':
return AIC, BIC
elif whichOne == 'AIC':
return AIC
elif whichOne == 'BIC':
return BIC
else:
ValueError('invalid value for argument whichOne')
for i in results_Feature_RL.index:
# fitToNParams = (results_all.loc[i,'fitTo']-1)*3
# combNParams = combs[(results_all.loc[i,'combInd']-1)].count('1')
# asymm_LRNParams = (results_all.loc[i,'asymm_LR'])
# nParams = 2 + combNParams + asymm_LRNParams + fitToNParams
nParams = sum(~np.isnan(results_Feature_RL.loc[i,paramCols].values.astype(float)))
# print(nParams)
AIC, BIC = penalizedModelFit(results_Feature_RL.loc[i,'bestNLL'],nParams)
# print(BIC)
results_Feature_RL.loc[i,'AIC'] = AIC
results_Feature_RL.loc[i,'BIC'] = BIC
results_Feature_RL['bestModel'] = False
for subID in results_Feature_RL['subID'].unique():
subDat = results_Feature_RL.query('subID == @subID'
).sort_values(by='BIC',ascending=True).copy()
bestModel = subDat.iloc[0]
bestBIC = bestModel['BIC']
results_Feature_RL.loc[(results_Feature_RL['subID']==subID) &
(results_Feature_RL['BIC']==bestBIC),'bestModel'] = True
# ##### 50 subjects * 8 combs * asymm_LR * bounded_weights = 1600:
results_Feature_RL.shape
results_Feature_RL.head()
results_Feature_RL.to_csv('FeatureRL/Results/results_06-Sep-2019/results_all.csv')
# ## Single subject
subID = '5005'
results_Feature_RL.query('subID == @subID & comb == "1111"')
# ## Model comparison overall
# ##### All models – BIC
plotDat = results_Feature_RL.query('fitTo == 1 & asymm_LR == 0').copy()
means = plotDat[['comb','bounded_weights','BIC']].groupby(
['comb','bounded_weights']).mean().sort_values(by='BIC',ascending=True).reset_index()
combOrder = means['comb'].unique()
print(combOrder)
g = sns.catplot(kind='bar',data=results_Feature_RL,x='comb',y='BIC',col = 'asymm_LR',
hue = 'bounded_weights', order = combOrder, row_order = [1,2])
g.set_xticklabels(rotation=45);
ax1, ax2 = g.axes[0]
ax1.axhline(means.loc[0,'BIC'], color='k',ls=':')
ax2.axhline(means.loc[0,'BIC'], color='k',ls=':');
# ##### All models - AIC
plotDat = results_Feature_RL.query('fitTo == 1 & asymm_LR == 0').copy()
means = plotDat[['comb','bounded_weights','AIC']].groupby(
['comb','bounded_weights']).mean().sort_values(by='AIC',ascending=True).reset_index()
combOrder = means['comb'].unique()
print(combOrder)
g = sns.catplot(kind='bar',data=results_Feature_RL,
x='comb',y='AIC',col = 'asymm_LR',
hue = 'bounded_weights', order = combOrder, row_order = [1,2])
g.set_xticklabels(rotation=45);
ax1, ax2 = g.axes[0]
ax1.axhline(means.loc[0,'AIC'], color='k',ls=':')
ax2.axhline(means.loc[0,'AIC'], color='k',ls=':');
# ##### Pretty plot for choice-only
plotDat = results_Feature_RL.query('fitTo == 1 & asymm_LR == 0').copy()
means = plotDat[['comb','asymm_LR','bounded_weights','BIC']].groupby(
['comb','asymm_LR','bounded_weights']).mean().sort_values(
by='BIC',ascending=True).reset_index()
combOrder = means['comb'].unique()
means.head()
sns.set_context('notebook')
fig, ax = plt.subplots(nrows = 1, ncols = 1, figsize = [10,5]);
sns.barplot(data = plotDat, x = 'comb', y = 'BIC', hue = 'bounded_weights', ax = ax,
alpha = 0.5, errwidth = 0, zorder = 1, order = combOrder);
[hand, lab] = ax.get_legend_handles_labels();
sns.swarmplot(data = plotDat, x = 'comb', y = 'BIC', hue = 'bounded_weights', dodge = True, ax = ax,
alpha = 1, size = 2, zorder = 2, order = combOrder);
sns.barplot(data = plotDat, x = 'comb', y = 'BIC', hue = 'bounded_weights', ax = ax,
alpha = 0, errwidth = 1.5, capsize = .2, errcolor = 'k', zorder = 3, order = combOrder);
plt.plot([0,15],[np.min(means.BIC), np.min(means.BIC)], 'k:', lw=1);
plt.xticks(rotation=45);
# plt.ylim([0,140]);
plt.legend(hand, ['Weights can be >= 0 and < 0','Weights strictly >= 0'], loc = [0.01, 0.83]);
ax.set(xlabel = 'Combination of features', title =
'Model comparison - symmetric learning rate - sort by BIC');
# plt.savefig('/Users/jvanbaar/Desktop/modelCompBIC.pdf',bbox_inches='tight');
# ## Plot with bounded_LR = 0 and asymm_LR = 0, BIC - split by features only
import re
plotDat.head()
# +
plotDat = results_Feature_RL.query('fitTo == 1 & asymm_LR == 0 & bounded_weights == 0').copy()
letter_list = ['Coop.','Greed','Risk','Regret']
plotDat['comb_letters'] = plotDat['comb'].apply(lambda x:
',\n'.join(letter_list[k] for k in [i.span()[0] for i in re.finditer('1',x)]))
means = plotDat[['comb_letters','BIC']].groupby('comb_letters').mean().sort_values(by='BIC',ascending=True).reset_index()
combOrder = means['comb_letters'].unique()
means.head()
sns.set_context('talk')
fig, ax = plt.subplots(nrows = 1, ncols = 1, figsize = [8,5]);
sns.barplot(data = plotDat, x = 'comb_letters', y = 'BIC', color = sns.color_palette('tab10')[0], ax = ax,
alpha = 0.3, errwidth = 0, zorder = 1, order = combOrder, edgecolor = 'k', linewidth = 1);
[hand, lab] = ax.get_legend_handles_labels();
sns.swarmplot(data = plotDat, x = 'comb_letters', y = 'BIC', ax = ax, color = sns.color_palette('tab10')[0],
alpha = .8, size = 4, zorder = 2, order = combOrder);
sns.barplot(data = plotDat, x = 'comb_letters', y = 'BIC', ax = ax,
alpha = 0, errwidth = 2, capsize = .2, errcolor = 'k', zorder = 3, order = combOrder);
plt.plot([0,15],[np.min(means.BIC), np.min(means.BIC)], 'k:', lw=1);
plt.xticks(rotation=0);
# plt.ylim([0,120]);
# plt.legend(hand, ['Weights can be >= 0 and < 0','Weights strictly >= 0'], loc = [0.01, 0.83]);
ax.set(xlabel = 'Motives included in model', title = 'Group-level model fit');
plt.savefig('/Users/jvanbaar/Desktop/modelCompBIC_motiveLabels_study2.pdf',bbox_inches='tight',
transparent = True);
# -
# ## Model comparison per subject - counts (based on BIC)
bestPerSubject = pd.DataFrame()
for sub in results_Feature_RL['subID'].unique():
subDat = results_Feature_RL.loc[results_Feature_RL['subID']==sub,:].copy()
subDat = subDat.sort_values(by=['BIC','AIC'], ascending=True)
# bestBIC = np.min(subDat.BIC)
# bestModel = subDat.loc[subDat['BIC']==bestBIC,
# ['subID','comb','asymm_LR','bounded_weights']].copy()
bestPerSubject = bestPerSubject.append(subDat.iloc[0])
bestPerSubject = bestPerSubject.reset_index(drop=True)
bestModelCounts = pd.DataFrame(pd.value_counts(bestPerSubject.comb)).reset_index()
bestModelCounts.columns = ['comb','count']
bestModelCounts = bestModelCounts.sort_values(by='count', ascending=False)
bestPerSubject['nDim'] = [sum([char=='1' for
char in bestPerSubject.comb.values[i]]) for i in np.arange(len(bestPerSubject))]
bestModelComplexityCounts = pd.DataFrame(pd.value_counts(bestPerSubject.nDim)).reset_index()
bestModelComplexityCounts.columns = ['nDim','count']
bestModelComplexityCounts = bestModelComplexityCounts.sort_values(by='count', ascending=False)
bestModelComplexityCounts
bestPerSubject.head()
label_list = ['Coop.','Greed','Risk','Regret']
bestModelCounts['comb_labels'] = bestModelCounts['comb'].apply(lambda x:
',\n'.join(label_list[k] for k in [i.span()[0] for i in re.finditer('1',x)]))
bestModelCounts['pct'] = bestModelCounts['count']/150*100
bestModelComplexityCounts['pct'] = bestModelComplexityCounts['count']/150*100
bestModelCounts_trunc = bestModelCounts.iloc[:6,:]
bmc_other = pd.DataFrame(bestModelCounts.iloc[6:,:].sum()).T[['count','comb_labels','pct']]
bmc_other.loc[0,'comb_labels'] = 'other'
bestModelCounts_aggTrunc = bestModelCounts.iloc[:6,:].append(bmc_other, sort = False)
bestModelCounts_aggTrunc
sns.set_context('talk')
fig, ax = plt.subplots(nrows = 1, ncols = 2, figsize = [14,6], gridspec_kw = {'width_ratios':[2,1]});
sns.barplot(data = bestModelCounts_aggTrunc, x='comb_labels', y='pct',
order = bestModelCounts_aggTrunc.comb_labels, ax = ax[0],
palette = 'tab10')
ax[0].set(xlabel = 'Motives included in model', ylabel = 'Frequency (%%)',
title = 'Best model per subject');
ax[0].set_xticklabels(ax[0].get_xticklabels(),rotation=0)
sns.barplot(data = bestModelComplexityCounts, x='nDim', y='pct', order = [1,2,3,4], ax = ax[1])
ax[1].set(xlabel = 'Number of motives in model', ylabel = 'Frequency (%%)',
title = 'Model complexity per subject');
plt.tight_layout()
plt.savefig('/Users/jvanbaar/Desktop/modelComp_perSub_labels_pct_study2.pdf',bbox_inches='tight',
transparent = True)
bestPerSubject_features = bestPerSubject.copy()
label_list = ['Coop.','Greed','Risk','Regret']
bestPerSubject_features['comb_labels'] = bestPerSubject_features['comb'].apply(lambda x:
',\n'.join(label_list[k] for k in [i.span()[0] for i in re.finditer('1',x)]))
for feature in ['Coop','Greed','Risk','Regret']:
bestPerSubject_features[feature] = bestPerSubject_features['comb_labels'].apply(lambda x: feature in x)
bestPerSubject_features.head()
featureCounts = pd.DataFrame(bestPerSubject_features[['Coop','Greed','Risk','Regret']].sum()).reset_index()
featureCounts.columns = ['Motive','Count']
featureCounts['pct'] = featureCounts['Count']/150
featureCounts.sort_values(by='Count', ascending = False, inplace=True)
featureCounts
sns.set_context('talk')
fig, ax = plt.subplots(nrows = 1, ncols = 2, figsize = [14,6], gridspec_kw = {'width_ratios':[2,1]});
sns.barplot(data = bestModelCounts_aggTrunc, x='comb_labels', y='pct',
order = bestModelCounts_aggTrunc.comb_labels, ax = ax[0],
palette = 'tab10')
ax[0].set(xlabel = 'Motives included in model', ylabel = 'Frequency (%%)',
title = 'Best model per subject');
ax[0].set_xticklabels(ax[0].get_xticklabels(),rotation=0)
sns.barplot(data = featureCounts, x='Motive', y='pct', ax = ax[1])
ax[1].set(xlabel = 'Motive', ylabel = 'Frequency (%%)',
title = 'Popularity of inferred motives');
plt.tight_layout()
plt.savefig('/Users/jvanbaar/Desktop/modelComp_perSub_labels_pct_withFeatureCounts_study2.pdf',bbox_inches='tight',
transparent = True)
# ## Store features per subject
bestPerSubject_features[['subID','Coop','Greed','Risk','Regret','comb','comb_labels']].to_csv(
'/Users/jvanbaar/Dropbox (Brown)/Postdoc FHL/JEROEN/SOC_STRUCT_LEARN/Study2_EyeTracking/'+
'Data/Cleaned/ModelFeaturesPerSubject.csv')
# ## Relationship between model features/complexity and performance
blockDat = blockDat.merge(bestPerSubject[['subID','nDim','comb']],on='subID')
blockDat.head()
# ##### Merge data based on asymm_LR = 0, bounded_weights = 0, BIC (dataframe 'bestPerSubject')
totalDat = blockDat[['subID','Confidence','Score']].groupby(['subID']).mean().reset_index()
totalDat = totalDat.merge(bestPerSubject[['subID','nDim','comb']],on='subID')
totalDat.head()
means = totalDat[['comb','nDim','Score']].groupby(['comb','nDim']).mean().sort_values(
by='Score',ascending=False).reset_index()
combOrder = list(means['comb'].values)
print(combOrder)
nDimOrder = list(means[['nDim','comb']].sort_values(by='nDim',ascending=False)['comb'].values)
print(nDimOrder)
fig, ax = plt.subplots(nrows = 1, ncols = 1, figsize = [12,4]);
sns.set_palette('tab10')
sns.barplot(data = totalDat, x = 'comb', y = 'Score', ax=ax, order = combOrder,
alpha = .4, errwidth = 0, zorder = 0, palette = 'tab20');
[hand, lab] = ax.get_legend_handles_labels();
sns.stripplot(data = totalDat, x = 'comb', y = 'Score', ax=ax, order = combOrder, dodge=True,
alpha = 0.8, size = 10, jitter = .2, zorder = 1, palette = 'tab20');
sns.barplot(data = totalDat, x = 'comb', y = 'Score', ax=ax, order = combOrder,
alpha = 0, errwidth = 1, capsize = .2, errcolor = 'k', zorder = 2, palette = 'tab20');
ax.set(xlabel = 'Model features available to subject', ylabel = 'Mean accuracy overall',
title = 'Task performance by model features');
# plt.savefig('/Users/jvanbaar/Desktop/PerformanceByFeatureSet.pdf',bbox_inches='tight',
# transparent = True)
# +
nCombsPernDim = [len(totalDat.query('nDim == @nDim')['comb'].unique()) for nDim in [4,3,2,1]]
nCombsPernDim.append(6)
fig, ax = plt.subplots(nrows = 1, ncols = 5, figsize = [12,4], sharey=True,
gridspec_kw={'width_ratios':nCombsPernDim});
nPast = 0
for ndi,nDim in enumerate([4,3,2,1]):
nCombs = len(totalDat.query('nDim == @nDim')['comb'].unique())
if nCombs > 1:
means = totalDat.query('nDim == @nDim')[['comb','nDim','Score']].groupby(
['comb','nDim']).mean().sort_values(by='Score',ascending=False).reset_index()
combOrder = list(means['comb'].values)
else:
combOrder = ['1111']
sns.barplot(data = totalDat.query('nDim == @nDim'), y = 'Score', x='comb', ax=ax[ndi],
alpha = .6, errwidth = 0, zorder = 0, order = combOrder,
palette = sns.color_palette('tab20')[nPast:(nPast+nCombs)]);
sns.stripplot(data = totalDat.query('nDim == @nDim'), y = 'Score', x='comb', ax=ax[ndi],
alpha = 1, zorder = 1, order = combOrder, size = 5, jitter = .2,
palette = sns.color_palette('tab20')[nPast:(nPast+nCombs)]);
sns.barplot(data = totalDat.query('nDim == @nDim'), y = 'Score', x='comb', ax=ax[ndi],
alpha = 0, errwidth = 1, errcolor = 'k', capsize = .1, zorder = 2, order = combOrder,
palette = sns.color_palette('tab20')[nPast:(nPast+nCombs)]);
nPast = nPast + nCombs
ax[0].set(ylabel = 'Mean accuracy across 64 trials')
[ax[i].set(ylabel = '', xlabel = 'Features') for i in [1,2,3]]
sns.barplot(data = totalDat, x = 'nDim', y = 'Score', order = [4,3,2,1], ax = ax[4],
alpha = .6, errwidth = 0, zorder = 0)
sns.stripplot(data = totalDat, x = 'nDim', y = 'Score', order = [4,3,2,1], ax = ax[4],
alpha = 1, size = 5, jitter = .2, zorder = 1)
sns.barplot(data = totalDat, x = 'nDim', y = 'Score', order = [4,3,2,1], ax = ax[4],
alpha = 0, errwidth = 1, errcolor = 'k', capsize = .2, zorder = 3)
[ax[i].set(title = '%i features'%(4-i)) for i in [0,1,2]]
ax[3].set(title = '1 feature')
ax[4].set(title = 'Aggregate', ylabel = '', xlabel = 'Number of features')
plt.tight_layout()
# plt.savefig('/Users/jvanbaar/Desktop/PerformanceByFeatureCount.pdf',bbox_inches='tight',
# transparent = True)
# -
print(scipy.stats.spearmanr(totalDat['nDim'],totalDat['Score']))
print(scipy.stats.f_oneway(totalDat.query('nDim == 1')['Score'],
totalDat.query('nDim == 2')['Score'],
totalDat.query('nDim == 3')['Score'],
totalDat.query('nDim == 4')['Score']))
# ##### Exclude single subject in ndim = 4
print(scipy.stats.spearmanr(totalDat.query('nDim < 4')['nDim'],
totalDat.query('nDim < 4')['Score']))
print(scipy.stats.f_oneway(totalDat.query('nDim == 2')['Score'],
totalDat.query('nDim == 3')['Score'],
totalDat.query('nDim == 4')['Score']))
# ##### Best subject
totalDat.query('nDim == 4')['subID'].unique()
gameDat.query('subID == "2133"')[['Type_Total','SelfReport']].drop_duplicates()
# ##### Performance by Opt/Pess
relevant_feature_dat = blockDat[['subID','Confidence','Score','Type_Total']].copy()
relevant_feature_dat = relevant_feature_dat.merge(bestPerSubject[['subID','nDim','comb']],on='subID')
relevant_feature_dat['has_greed'] = relevant_feature_dat['comb'].apply(lambda x: x[1] == '1')
relevant_feature_dat['has_risk'] = relevant_feature_dat['comb'].apply(lambda x: x[2] == '1')
relevant_feature_dat.head()
# +
fig, ax = plt.subplots(nrows = 1, ncols = 2, figsize = [14,6]);
sns.set_palette('tab10')
sns.barplot(data = relevant_feature_dat, hue = 'has_greed', y = 'Score', x = 'Type_Total',
order = ['opt_nat','pess_nat'],
ax=ax[0], alpha = .4, errwidth = 0, zorder = 0);
[hand, lab] = ax[0].get_legend_handles_labels();
sns.stripplot(data = relevant_feature_dat, hue = 'has_greed', y = 'Score', x = 'Type_Total', ax=ax[0],
order = ['opt_nat','pess_nat'],
dodge=True, alpha = 0.8, size = 10, jitter = .2, zorder = 1);
sns.barplot(data = relevant_feature_dat, hue = 'has_greed', y = 'Score', ax=ax[0], x = 'Type_Total',
order = ['opt_nat','pess_nat'],
alpha = 0, errwidth = 1, capsize = .2, errcolor = 'k', zorder = 2, palette = 'tab20');
ax[0].legend(hand,lab,title = 'Has Greed motive')
sns.barplot(data = relevant_feature_dat, hue = 'has_risk', y = 'Score', x = 'Type_Total',
order = ['opt_nat','pess_nat'],
ax=ax[1], alpha = .4, errwidth = 0, zorder = 0);
[hand, lab] = ax[1].get_legend_handles_labels();
sns.stripplot(data = relevant_feature_dat, hue = 'has_risk', y = 'Score', x = 'Type_Total', ax=ax[1],
order = ['opt_nat','pess_nat'],
dodge=True, alpha = 0.8, size = 10, jitter = .2, zorder = 1);
sns.barplot(data = relevant_feature_dat, hue = 'has_risk', y = 'Score', ax=ax[1], x = 'Type_Total',
order = ['opt_nat','pess_nat'],
alpha = 0, errwidth = 1, capsize = .2, errcolor = 'k', zorder = 2, palette = 'tab20');
ax[1].legend(hand,lab,title = 'Has Risk motive')
# plt.legend(hand,lab, loc = [1.1,.5])
# ax.set(xlabel = 'Model features available to subject', ylabel = 'Mean accuracy overall',
# title = 'Task performance by model features');
# # plt.savefig('/Users/jvanbaar/Desktop/PerformanceByFeatureSet.pdf',bbox_inches='tight',
# # transparent = True)
# -
fig, axes = plt.subplots(nrows = 1, ncols = 2, figsize = [14,6]);
sns.set_palette('tab10')
player_types = ['opt_nat','pess_nat']
lf_player_types = ['Optimist','Pessimist']
motive_columns = ['has_greed','has_risk']
lf_motive_columns = ['Greed','Risk']
colors = sns.color_palette('tab10')[:2]
for ai, ax in enumerate(axes.ravel()):
player_type = player_types[ai]
motive_column = motive_columns[ai]
tmp = relevant_feature_dat.query('Type_Total == @player_type').copy()
sns.barplot(data = tmp,
x = motive_column, y = 'Score', ax=ax, alpha = .4, errwidth = 0, zorder = 0,
color = colors[ai]);
# [hand, lab] = ax[0].get_legend_handles_labels();
sns.stripplot(data = tmp,
x = motive_column, y = 'Score', ax=ax, alpha = 0.8, size = 10, jitter = .2, zorder = 1,
color = colors[ai]);
sns.barplot(data = tmp,
x = motive_column, y = 'Score', ax=ax,
alpha = 0, errwidth = 2, capsize = .1, errcolor = 'k', zorder = 2,
color = colors[ai]);
ax.set(title = 'Predicting %s'%lf_player_types[ai], ylabel = 'Accuracy',
xticklabels = ['Motive not\nconsidered', 'Motive\nconsidered'],
xlabel = lf_motive_columns[ai])
ttest_results = scipy.stats.ttest_ind(tmp.loc[tmp[motive_column],'Score'],
tmp.loc[~tmp[motive_column],'Score'])
pval = ttest_results[1]
FigureTools.add_sig_markers(ax, relationships = [[0,1,pval]])
plt.tight_layout()
plt.savefig('/Users/jvanbaar/Desktop/PerformanceByMotive.pdf',bbox_inches='tight',
transparent = True)
# ## Relationship between model features/complexity and response time
gameDat = pd.read_csv('FeatureRL/gameDat.csv',index_col = 0, dtype={ 'subID':str})
gameDat.head()
RTdat = gameDat[['subID','Type_Total','RT_radio','RT_submit']
].groupby(['subID','Type_Total']).sum().reset_index()
RTdat.head()
RTdat = RTdat.merge(bestPerSubject[['subID','nDim','comb']],on='subID')
RTdat.head()
# ##### Total game time
RTdatOverall = RTdat[['subID','RT_radio','RT_submit','nDim','comb']].groupby(['subID','nDim','comb']
).sum().reset_index()
RTdatOverall.head()
RTdatOverall_unitS = RTdatOverall.copy()
RTdatOverall_unitS['RT_radio'] = RTdatOverall_unitS['RT_radio']/1000
RTdatOverall_unitS['RT_submit'] = RTdatOverall_unitS['RT_submit']/1000
RTdatOverall_unitS.head()
# ##### Remove outliers
RTdatOverall_unitS = RTdatOverall_unitS.query('RT_submit < 2000').copy()
# ##### Plot
# +
plotDat = RTdatOverall_unitS.copy()
RTchoice = 'RT_submit' # Can be 'RT_radio' or 'RT_submit'
nCombsPernDim = [len(plotDat.query('nDim == @nDim')['comb'].unique()) for nDim in [4,3,2,1]]
nCombsPernDim.append(6)
fig, ax = plt.subplots(nrows = 1, ncols = 5, figsize = [12,4], sharey=True,
gridspec_kw={'width_ratios':nCombsPernDim});
nPast = 0
for ndi,nDim in enumerate([4,3,2,1]):
nCombs = len(plotDat.query('nDim == @nDim')['comb'].unique())
if nCombs > 1:
means = plotDat.query('nDim == @nDim')[['comb','nDim',RTchoice]].groupby(
['comb','nDim']).mean().sort_values(by=RTchoice,ascending=True).reset_index()
combOrder = list(means['comb'].values)
else:
combOrder = ['1111']
sns.barplot(data = plotDat.query('nDim == @nDim'), y = RTchoice, x='comb', ax=ax[ndi],
alpha = .8, errwidth = 0, zorder = 0, order = combOrder,
palette = sns.color_palette('tab20')[nPast:(nPast+nCombs)]);
sns.stripplot(data = plotDat.query('nDim == @nDim'), y = RTchoice, x='comb', ax=ax[ndi],
alpha = 1, zorder = 1, order = combOrder, size = 5, jitter = .2,
edgecolor = 'k', linewidth=.5, palette = sns.color_palette('tab20')[nPast:(nPast+nCombs)]);
sns.barplot(data = plotDat.query('nDim == @nDim'), y = RTchoice, x='comb', ax=ax[ndi],
alpha = 0, errwidth = 1, errcolor = 'k', capsize = .1, zorder = 2, order = combOrder,
palette = sns.color_palette('tab20')[nPast:(nPast+nCombs)]);
nPast = nPast + nCombs
ax[0].set(ylabel = 'Total %s (64 trials)'%RTchoice)
[ax[i].set(ylabel = '', xlabel = 'Features') for i in [1,2,3]]
sns.barplot(data = plotDat, x = 'nDim', y = RTchoice, order = [4,3,2,1], ax = ax[4],
alpha = .8, errwidth = 0, zorder = 0)
sns.stripplot(data = plotDat, x = 'nDim', y = RTchoice, order = [4,3,2,1], ax = ax[4],
alpha = 1, size = 5, jitter = .2, zorder = 1, edgecolor = 'k', linewidth=.5)
sns.barplot(data = plotDat, x = 'nDim', y = RTchoice, order = [4,3,2,1], ax = ax[4],
alpha = 0, errwidth = 1, errcolor = 'k', capsize = .2, zorder = 3)
[ax[i].set(title = '%i features'%(4-i)) for i in [0,1,2]]
ax[3].set(title = '1 feature')
ax[4].set(title = 'Aggregate', ylabel = '', xlabel = 'Number of features')
plt.tight_layout()
plt.savefig('/Users/jvanbaar/Desktop/RTByFeatureCount.pdf',bbox_inches='tight',
transparent = True)
# -
print(scipy.stats.pearsonr(RTdatOverall_unitS['nDim'],RTdatOverall_unitS['RT_submit']))
print(scipy.stats.f_oneway(RTdatOverall_unitS.query('nDim == 1')['RT_submit'],
RTdatOverall_unitS.query('nDim == 2')['RT_submit'],
RTdatOverall_unitS.query('nDim == 3')['RT_submit'],
RTdatOverall_unitS.query('nDim == 4')['RT_submit']))
# ## Which dimensions are used by people from the Coop, Opt, and Pess groups (self-report)?
baseDir = '/Users/jvanbaar/Dropbox (Brown)/Postdoc FHL/JEROEN/SOC_STRUCT_LEARN/'
DIYquiz = pd.read_csv(baseDir+'Study1_MTurk/Data/Cleaned/DIYquiz.csv',
dtype = {'subID':str}, index_col=0)
DIYquiz.head()
type(DIYquiz['subID'].values[0])
DIYquiz = DIYquiz.merge(bestPerSubject[['subID','comb','nDim']], on='subID')
DIYquiz.head()
combs = DIYquiz['comb'].unique()
combs.sort()
combs
combCountsAll = pd.DataFrame(columns=combs, index=DIYquiz['selfType'].unique())
combCountsAll[:] = 0
combCountsAll
for st in DIYquiz['selfType'].unique():
combCounts = DIYquiz.loc[DIYquiz['selfType']==st,'comb'].value_counts(dropna=False)
for comb in combCounts.index.values:
combCountsAll.loc[st, comb] = combCounts[comb]
combCountsAll
# combCountsAll = combCountsAll.iloc[[0,4,1],:]
combCountsAll_other = pd.DataFrame(combCountsAll.iloc[[2,3,5],:].sum(axis=0))
combCountsAll_other.columns = ['other']
combCountsAll_other = combCountsAll_other.T
# combCountsAll_other
combCountsAll = combCountsAll.iloc[[0,4,1],:].append(combCountsAll_other)
combCountsAll
# fbg_red = fbg_red.append(mean_other)
combCountsAllShare = combCountsAll.copy()
for st in combCountsAllShare.index:
# vals = combCountsAllShare.loc[st,:].values
combCountsAllShare.loc[st,:] = np.divide(combCountsAllShare.loc[st,:],
np.sum(combCountsAllShare.loc[st,:].values))
combCountsAllShare
sns.set_context('talk')
fig, ax = plt.subplots(nrows = 1, ncols = 1);
sns.heatmap(combCountsAllShare, ax=ax)
ax.set(xlabel = 'Combination of features', ylabel = 'Participant''s own strategy',
title = 'Does it take one to know one?');
# plt.savefig('/Users/jvanbaar/Desktop/featuresByOwnType.pdf',bbox_inches='tight');
combs_considered = combCountsAllShare.columns
featuresByGroup = pd.DataFrame(columns=[0,1,2,3], index = combCountsAllShare.index)
featuresByGroup[:] = 0
for feature in [0,1,2,3]:
columnsToCheck = []
for comb in combs_considered:
if comb[feature]=='1':
columnsToCheck.append(comb)
for st in combCountsAllShare.index:
featuresByGroup.loc[st, feature] = np.sum(combCountsAllShare.loc[st, columnsToCheck])
featuresByGroup = featuresByGroup*100
featuresByGroup
# +
# fbg_red = featuresByGroup.iloc[[0,4,1],:]
# mean_other = pd.DataFrame(featuresByGroup.iloc[[2,3,5],:].mean(axis=0))
# mean_other.columns = ['other']
# mean_other = mean_other.T
# fbg_red = fbg_red.append(mean_other)
# fbg_red
# -
sns.set_context('talk')
fig, ax = plt.subplots(nrows = 1, ncols = 1, figsize = [6,5]);
sns.heatmap(featuresByGroup, ax=ax, cbar_kws = {'label':'Motive included in best model\n(%)'})
ax.set(xlabel = 'Motive', ylabel = 'Participant\'s own strategy',
# title = 'Frequency of motives inferred',
xticklabels = ['Coop','Greed','Risk','Regret'],
yticklabels = ['Optimist','Pessimist','Trustful','Other']
);
plt.yticks(rotation = 0);
plt.savefig('/Users/jvanbaar/Desktop/featuresByOwnType_reduced_pct.pdf',bbox_inches='tight',
transparent = True);
| Study1_MTurk/Analysis_scripts/2.Modeling/Matlab_cluster_modules/PlotResults_study2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building steel coils
#
# This tutorial includes everything you need to set up decision optimization engines, build constraint programming models.
#
#
# When you finish this tutorial, you'll have a foundational knowledge of _Prescriptive Analytics_.
#
# >This notebook is part of **[Prescriptive Analytics for Python](http://ibmdecisionoptimization.github.io/docplex-doc/)**
# >
# >It requires either an [installation of CPLEX Optimizers](http://ibmdecisionoptimization.github.io/docplex-doc/getting_started.html) or it can be run on [IBM Watson Studio Cloud](https://www.ibm.com/cloud/watson-studio/>) (Sign up for a [free IBM Cloud account](https://dataplatform.cloud.ibm.com/registration/stepone?context=wdp&apps=all>)
# and you can start using Watson Studio Cloud right away).
#
# Table of contents:
#
# - [Describe the business problem](#Describe-the-business-problem)
# * [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
# * [Use decision optimization](#Use-decision-optimization)
# * [Step 1: Download the library](#Step-1:-Download-the-library)
# * [Step 2: Model the Data](#Step-2:-Model-the-data)
# * [Step 3: Set up the prescriptive model](#Step-3:-Set-up-the-prescriptive-model)
# * [Define the decision variables](#Define-the-decision-variables)
# * [Express the business constraints](#Express-the-business-constraints)
# * [Express the objective](#Express-the-objective)
# * [Solve with Decision Optimization solve service](#Solve-with-Decision-Optimization-solve-service)
# * [Step 4: Investigate the solution and run an example analysis](#Step-4:-Investigate-the-solution-and-then-run-an-example-analysis)
# * [Summary](#Summary)
# ****
# ### Describe the business problem
#
# * The problem is to build steel coils from slabs that are available in a work-in-process inventory of semi-finished products. There is no limitation in the number of slabs that can be requested, but only a finite number of slab sizes is available (sizes 11, 13, 16, 17, 19, 20, 23, 24, 25, 26, 27, 28, 29, 30, 33, 34, 40, 43, 45).
# * The problem is to select a number of slabs to build the coil orders, and to satisfy the following constraints:
# * A coil order can be built from only one slab.
# * Each coil order requires a specific process to build it from a slab. This process is encoded by a color.
# * Several coil orders can be built from the same slab. But a slab can be used to produce at most two different "colors" of coils.
# * The sum of the sizes of each coil order built from a slab must not exceed the slab size.
#
#
# * Finally, the production plan should minimize the unused capacity of the selected slabs.
#
#
# * This problem is based on **"prob038: Steel mill slab design problem" from CSPLib (www.csplib.org). It is a simplification of an industrial problem described in <NAME>, <NAME>, <NAME>, <NAME>. "Inventory Matching Problems in the Steel Industry," IBM Research Report RC 21171, 1998**.
#
# * Please refer to documentation for appropriate setup of solving configuration.
# *****
# ## How decision optimization can help
# * Prescriptive analytics technology recommends actions based on desired outcomes, taking into account specific scenarios, resources, and knowledge of past and current events. This insight can help your organization make better decisions and have greater control of business outcomes.
#
# * Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
#
# * Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
# <br/>
#
# + For example:
# + Automate complex decisions and trade-offs to better manage limited resources.
# + Take advantage of a future opportunity or mitigate a future risk.
# + Proactively update recommendations based on changing events.
# + Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
#
# ## Use decision optimization
# ### Step 1: Download the library
#
# Run the following code to install Decision Optimization CPLEX Modeling library. The *DOcplex* library contains the two modeling packages, Mathematical Programming and Constraint Programming, referred to earlier.
import sys
try:
import docplex.cp
except:
if hasattr(sys, 'real_prefix'):
#we are in a virtual env.
# !pip install docplex
else:
# !pip install --user docplex
# Note that the more global package <i>docplex</i> contains another subpackage <i>docplex.mp</i> that is dedicated to Mathematical Programming, another branch of optimization.
# ### Step 2: Model the data
from docplex.cp.model import *
# Set model parameter
# +
from collections import namedtuple
##############################################################################
# Model configuration
##############################################################################
# The number of coils to produce
TUPLE_ORDER = namedtuple("TUPLE_ORDER", ["index", "weight", "color"])
orders = [ TUPLE_ORDER(1, 22, 5),
TUPLE_ORDER(2, 9, 3),
TUPLE_ORDER(3, 9, 4),
TUPLE_ORDER(4, 8, 5),
TUPLE_ORDER(5, 8, 7),
TUPLE_ORDER(6, 6, 3),
TUPLE_ORDER(7, 5, 6),
TUPLE_ORDER(8, 3, 0),
TUPLE_ORDER(9, 3, 2),
TUPLE_ORDER(10, 3, 3),
TUPLE_ORDER(11, 2, 1),
TUPLE_ORDER(12, 2, 5)
]
NB_SLABS = 12
MAX_COLOR_PER_SLAB = 2
# The total number of slabs available. In theory this can be unlimited,
# but we impose a reasonable upper bound in order to produce a practical
# optimization model.
# The different slab weights available.
slab_weights = [ 0, 11, 13, 16, 17, 19, 20, 23, 24, 25,
26, 27, 28, 29, 30, 33, 34, 40, 43, 45 ]
# +
nb_orders = len(orders)
slabs = range(NB_SLABS)
allcolors = set([ o.color for o in orders ])
# CPO needs lists for pack constraint
order_weights = [ o.weight for o in orders ]
# The heaviest slab
max_slab_weight = max(slab_weights)
# The amount of loss incurred for different amounts of slab use
# The loss will depend on how much less steel is used than the slab
# just large enough to produce the coils.
loss = [ min([sw-use for sw in slab_weights if sw >= use]) for use in range(max_slab_weight+1)]
# -
# ### Step 3: Set up the prescriptive model
# Create CPO model
mdl = CpoModel(name="trucks")
# #### Define the decision variables
# +
# Which slab is used to produce each coil
production_slab = integer_var_dict(orders, 0, NB_SLABS-1, "production_slab")
# How much of each slab is used
slab_use = integer_var_list(NB_SLABS, 0, max_slab_weight, "slab_use")
# -
# #### Express the business constraints
# +
# The total loss is
total_loss = sum([element(slab_use[s], loss) for s in slabs])
# The orders are allocated to the slabs with capacity
mdl.add(pack(slab_use, [production_slab[o] for o in orders], order_weights))
# At most MAX_COLOR_PER_SLAB colors per slab
for s in slabs:
su = 0
for c in allcolors:
lo = False
for o in orders:
if o.color==c:
lo = (production_slab[o] == s) | lo
su += lo
mdl.add(su <= MAX_COLOR_PER_SLAB)
# -
# #### Express the objective
# Add minimization objective
mdl.add(minimize(total_loss))
# #### Solve the model
# +
print("\nSolving model....")
# Search strategy
mdl.set_search_phases([search_phase([production_slab[o] for o in orders])])
msol = mdl.solve(FailLimit=100000, TimeLimit=10)
# -
# ### Step 4: Investigate the solution and then run an example analysis
# Print solution
if msol:
print("Solution: ")
from_slabs = [set([o.index for o in orders if msol[production_slab[o]]== s])for s in slabs]
slab_colors = [set([o.color for o in orders if o.index in from_slabs[s]])for s in slabs]
for s in slabs:
if len(from_slabs[s]) > 0:
print("Slab = " + str(s))
print("\tLoss = " + str(loss[msol[slab_use[s]]]))
print("\tcolors = " + str(slab_colors[s]))
print("\tOrders = " + str(from_slabs[s]) + "\n")
else:
print("No solution found")
# ## Summary
#
# You learned how to set up and use the IBM Decision Optimization CPLEX Modeling for Python to formulate and solve a Constraint Programming model.
# #### References
# * [CPLEX Modeling for Python documentation](https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html)
# * [Decision Optimization on Cloud](https://developer.ibm.com/docloud/)
# * Need help with DOcplex or to report a bug? Please go [here](https://stackoverflow.com/questions/tagged/docplex)
# * Contact us at <EMAIL>
# Copyright © 2017, 2018 IBM. IPLA licensed Sample Materials.
| examples/cp/jupyter/SteelMill.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
data = pd.read_csv("driver-data.csv", index_col="id")
data.head()
data['profit'] = np.random.randint(6,20, size=len(data))
x = data.iloc[:, [0,1,2]].values
print(x)
from sklearn.cluster import KMeans
#To check optimum number of clusters
Error =[]
for i in range(1, 11):
kmeans = KMeans(n_clusters = i).fit(x)
kmeans.fit(x)
Error.append(kmeans.inertia_)
import matplotlib.pyplot as plt
plt.plot(range(1, 11), Error)
plt.title('Elbow method')
plt.xlabel('No of clusters')
plt.ylabel('Error')
plt.show()
kmeans = KMeans(n_clusters=4)
labels = kmeans.fit_predict(x)
# +
unique, counts = np.unique(labels, return_counts=True)
dict_data = dict(zip(unique, counts))
dict_data
# -
data["cluster"] = labels
print(data)
import seaborn as sns
sns.lmplot('mean_dist_day', 'mean_over_speed_perc', data=data, hue='cluster', palette='coolwarm', size=6, aspect=1, fit_reg=False)
# +
#For normalising
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.preprocessing import StandardScaler
data2 = data
# print(data2)
scaler = StandardScaler()
scaler.fit(data2.drop('cluster',axis=1))
scaled_features = scaler.transform(data2.drop('cluster',axis=1))
df_feat = pd.DataFrame(scaled_features,columns=data2.columns[:-1])
df_feat.head()
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(scaled_features,data2['cluster'],test_size=0.30)
# +
#To check optimum number of k neighbours
from sklearn.model_selection import cross_val_score
accuracy_rate = []
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
score=cross_val_score(knn,df_feat,data2['cluster'],cv=10)
accuracy_rate.append(score.mean())
plt.figure(figsize=(10,6))
plt.plot(range(1,40),accuracy_rate,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Accuracy Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Accuracy Rate')
# -
#As from above plot we can see the accuracy rate is stable when k=17
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=17)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
#Classification Report
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,pred))
print(classification_report(y_test,pred))
| mainfile.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: s1-enumerator
# language: python
# name: s1-enumerator
# ---
# This is to demonstrate how to use the `s1-enumerator` to get a full time series of GUNWs.
#
# We are going basically take each month in acceptable date range and increment by a month and make sure the temporal window is large enough to ensure connectivity across data gaps.
# # Parameters
#
# This is what the operator is going to have to change. Will provide some comments.
# +
# toggle user-controlled parameters here
import datetime
# product cutline
aoi_shapefile = 'aois/LaPalma_pathNumber169.geojson'
### Spatial coverage constraint parameter 'azimuth_mismatch'
# The merged SLC area over the AOI is allowed to be smaller by 'azimuth_mismatch' x swath width (i.e. 250km)
azimuth_mismatch = 5
# Define job-name
job_name = 'LaPalma_T169'
# product directory
prod_dir = 'LaPalma_T169'
# Specify deployment URL
#deploy_url = 'https://hyp3-tibet.asf.alaska.edu' #for Tibet
deploy_url = 'https://hyp3-isce.asf.alaska.edu' #for access
# Number of nearest neighbors
num_neighbors = 2
#set temporal parameters
today = datetime.datetime.now()
# Earliest year for reference frames
START_YEAR = 2014
# Latest year for reference frames
END_YEAR = today.year
# Adjust depending on seasonality
# For annual IFGs, select a single months of interest and you will get what you want.
MONTHS_OF_INTEREST = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12]
# -
from s1_enumerator import get_aoi_dataframe, distill_all_pairs, enumerate_ifgs, get_s1_coverage_tiles, enumerate_ifgs_from_stack, get_s1_stack_by_dataframe
import concurrent
from rasterio.crs import CRS
from s1_enumerator import duplicate_gunw_found
from tqdm import tqdm
from shapely.geometry import Point, shape
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import numpy as np
from dateutil.relativedelta import relativedelta
import networkx as nx
import boto3
def shapefile_area(file_bbox,
bounds = False):
"""Compute km\u00b2 area of shapefile."""
# import dependencies
from pyproj import Proj
# loop through polygons
shape_area = 0
# pass single polygon as list
if file_bbox.type == 'Polygon': file_bbox = [file_bbox]
for polyobj in file_bbox:
#first check if empty
if polyobj.is_empty:
shape_area += 0
continue
# get coords
if bounds:
# Pass coordinates of bounds as opposed to cutline
# Necessary for estimating DEM/mask footprints
WSEN = polyobj.bounds
lon = np.array([WSEN[0],WSEN[0],WSEN[2],WSEN[2],WSEN[0]])
lat = np.array([WSEN[1],WSEN[3],WSEN[3],WSEN[1],WSEN[1]])
else:
lon, lat = polyobj.exterior.coords.xy
# use equal area projection centered on/bracketing AOI
pa = Proj("+proj=aea +lat_1={} +lat_2={} +lat_0={} +lon_0={}". \
format(min(lat), max(lat), (max(lat)+min(lat))/2, \
(max(lon)+min(lon))/2))
x, y = pa(lon, lat)
cop = {"type": "Polygon", "coordinates": [zip(x, y)]}
shape_area += shape(cop).area/1e6 # area in km^2
return shape_area
def continuous_time(product_df, iter_id='fileID'):
"""
Split the products into spatiotemporally continuous groups.
Split products by individual, continuous interferograms.
Input must be already sorted by pair and start-time to fit
the logic scheme below.
Using their time-tags, this function determines whether or not
successive products are in the same orbit.
If in the same orbit, the program determines whether or not they
overlap in time and are therefore spatially contiguous,
and rejects/reports cases for which there is no temporal overlap
and therefore a spatial gap.
"""
from shapely.ops import unary_union
# pass scenes that have no gaps
sorted_products = []
track_rejected_inds = []
pair_dict = {}
product_df_dict = product_df.to_dict('records')
# Check for (and remove) duplicate products
# If multiple pairs in list, cycle through
# and evaluate temporal connectivity.
for i in enumerate(product_df_dict[:-1]):
# Parse the first frame's metadata
scene_start = i[1]['startTime']
scene_end = i[1]['stopTime']
first_frame_ind = i[1]['ind_col']
first_frame = datetime.datetime.strptime( \
i[1]['fileID'][17:25], "%Y%m%d")
# Parse the second frame's metadata
new_scene_start = product_df_dict[i[0]+1]['startTime']
new_scene_end = product_df_dict[i[0]+1]['stopTime']
next_frame_ind = product_df_dict[i[0]+1]['ind_col']
next_frame = datetime.datetime.strptime( \
product_df_dict[i[0]+1]['fileID'][17:25], "%Y%m%d")
# Determine if next product in time is in same orbit AND overlaps
# AND corresponds to same scene
# If it is within same orbit cycle, try to append scene.
# This accounts for day change.
if abs(new_scene_end-scene_end) <= \
datetime.timedelta(minutes=100) \
and abs(next_frame-first_frame) <= \
datetime.timedelta(days=1):
# Don't export product if it is already tracked
# as a rejected scene
if first_frame_ind in track_rejected_inds or \
next_frame_ind in track_rejected_inds:
track_rejected_inds.append(first_frame_ind)
track_rejected_inds.append(next_frame_ind)
# Only pass scene if it temporally overlaps with reference scene
elif ((scene_end <= new_scene_start) and \
(new_scene_end <= scene_start)) or \
((scene_end >= new_scene_start) and \
(new_scene_end >= scene_start)):
# Check if dictionary for scene already exists,
# and if it does then append values
try:
dict_ind = sorted_products.index(next(item for item \
in sorted_products if i[1][iter_id] \
in item[iter_id]))
sorted_products[dict_ind] = {key: np.hstack([value] + \
[product_df_dict[i[0]+1][key]]).tolist() \
for key, value in sorted_products[dict_ind].items()}
# Match corresponding to scene NOT found,
# so initialize dictionary for new scene
except:
sorted_products.extend([dict(zip(i[1].keys(), \
[list(a) for a in zip(i[1].values(), \
product_df_dict[i[0]+1].values())]))])
# Else if scene doesn't overlap, this means there is a gap.
# Reject date from product list,
# and keep track of all failed dates
else:
track_rejected_inds.append(first_frame_ind)
track_rejected_inds.append(next_frame_ind)
# Products correspond to different dates,
# So pass both as separate scenes.
else:
# Check if dictionary for corresponding scene already exists.
if [item for item in sorted_products if i[1][iter_id] in \
item[iter_id]]==[] and i[1]['ind_col'] not in \
track_rejected_inds:
sorted_products.extend([dict(zip(i[1].keys(), \
[list(a) for a in zip(i[1].values())]))])
# Initiate new scene
if [item for item in sorted_products if \
product_df_dict[i[0]+1][iter_id] in item[iter_id]]==[] \
and next_frame_ind not in track_rejected_inds:
sorted_products.extend([dict(zip( \
product_df_dict[i[0]+1].keys(), \
[list(a) for a in \
zip(product_df_dict[i[0]+1].values())]))])
if first_frame_ind in track_rejected_inds:
track_rejected_inds.append(first_frame_ind)
if next_frame_ind in track_rejected_inds:
track_rejected_inds.append(next_frame_ind)
# Remove duplicate dates
track_rejected_inds = list(set(track_rejected_inds))
if len(track_rejected_inds) > 0:
print("{}/{} scenes rejected as stitched IFGs have gaps".format( \
len(track_rejected_inds), len(product_df)))
# Provide report of which files were kept vs. which were not.
print("Specifically, the following scenes were rejected:")
for item in product_df_dict:
if item['ind_col'] in track_rejected_inds:
print(item['fileID'])
else:
print("All {} scenes are spatially continuous.".format( \
len(sorted_products)))
# pass scenes that have no gaps
sorted_products = [item for item in sorted_products \
if not (any(x in track_rejected_inds for x in item['ind_col']))]
# Report dictionaries for all valid products
if sorted_products == []: #Check if pairs were successfully selected
raise Exception('No scenes meet spatial criteria'
'due to gaps and/or invalid input.'
'Nothing to export.')
# Combine polygons
for i in enumerate(sorted_products):
sorted_products[i[0]]['geometry'] = unary_union(i[1]['geometry'])
# combine and record scenes with gaps
track_kept_inds = pd.DataFrame(sorted_products)['ind_col'].to_list()
track_kept_inds = [item for sublist in track_kept_inds for item in sublist]
temp_gap_scenes_dict = [item for item in product_df_dict \
if not item['ind_col'] in track_kept_inds]
gap_scenes_dict = []
for i in enumerate(temp_gap_scenes_dict[:-1]):
# Parse the first frame's metadata
first_frame_ind = i[1]['ind_col']
first_frame = datetime.datetime.strptime( \
i[1]['fileID'][17:25], "%Y%m%d")
# Parse the second frame's metadata
next_frame_ind = temp_gap_scenes_dict[i[0]+1]['ind_col']
next_frame = datetime.datetime.strptime( \
temp_gap_scenes_dict[i[0]+1]['fileID'][17:25], "%Y%m%d")
# Determine if next product in time is in same orbit
# If it is within same orbit cycle, try to append scene.
# This accounts for day change.
if abs(next_frame-first_frame) <= \
datetime.timedelta(days=1):
# Check if dictionary for scene already exists,
# and if it does then append values
try:
dict_ind = gap_scenes_dict.index(next(item for item \
in gap_scenes_dict if i[1][iter_id] \
in item[iter_id]))
gap_scenes_dict[dict_ind] = {key: np.hstack([value] + \
[temp_gap_scenes_dict[i[0]+1][key]]).tolist() \
for key, value in gap_scenes_dict[dict_ind].items()}
# Match corresponding to scene NOT found,
# so initialize dictionary for new scene
except:
gap_scenes_dict.extend([dict(zip(i[1].keys(), \
[list(a) for a in zip(i[1].values(), \
temp_gap_scenes_dict[i[0]+1].values())]))])
# Products correspond to different dates,
# So pass both as separate scenes.
else:
# Check if dictionary for corresponding scene already exists.
if [item for item in gap_scenes_dict if i[1][iter_id] in \
item[iter_id]]==[]:
gap_scenes_dict.extend([dict(zip(i[1].keys(), \
[list(a) for a in zip(i[1].values())]))])
# Initiate new scene
if [item for item in gap_scenes_dict if \
temp_gap_scenes_dict[i[0]+1][iter_id] in item[iter_id]]==[]:
gap_scenes_dict.extend([dict(zip( \
temp_gap_scenes_dict[i[0]+1].keys(), \
[list(a) for a in \
zip(temp_gap_scenes_dict[i[0]+1].values())]))])
# there may be some extra missed pairs with gaps
if gap_scenes_dict != []:
extra_track_rejected_inds = pd.DataFrame(gap_scenes_dict)['ind_col'].to_list()
extra_track_rejected_inds = [item for sublist in extra_track_rejected_inds for item in sublist]
track_rejected_inds.extend(extra_track_rejected_inds)
return sorted_products, track_rejected_inds, gap_scenes_dict
def minimum_overlap_query(tiles,
aoi,
azimuth_mismatch=0.01,
iter_id='fileID'):
"""
Master function managing checks for SAR scene spatiotemporal contiguity
and filtering out scenes based off of user-defined spatial coverage threshold
"""
# initiate dataframe
tiles = tiles.sort_values(['startTime'])
updated_tiles = tiles.copy()
# Drop scenes that don't intersect with AOI at all
orig_len = updated_tiles.shape[0]
for index, row in tiles.iterrows():
intersection_area = aoi.intersection(row['geometry'])
overlap_area = shapefile_area(intersection_area)
aoi_area = shapefile_area(aoi)
percentage_coverage = (overlap_area/aoi_area)*100
if percentage_coverage == 0:
drop_ind = updated_tiles[updated_tiles['fileID'] == row['fileID']].index
updated_tiles = updated_tiles.drop(index=drop_ind)
updated_tiles = updated_tiles.reset_index(drop=True)
print("{}/{} scenes rejected for not intersecting with the AOI".format( \
orig_len-updated_tiles.shape[0], orig_len))
# group IFGs spatiotemporally
updated_tiles['ind_col'] = range(0, len(updated_tiles))
updated_tiles_dict, dropped_indices, gap_scenes_dict = continuous_time(updated_tiles, iter_id)
for i in dropped_indices:
drop_ind = updated_tiles.index[updated_tiles['ind_col'] == i]
updated_tiles.drop(drop_ind, inplace=True)
updated_tiles = updated_tiles.reset_index(drop=True)
# Kick out scenes that do not meet user-defined spatial threshold
aoi_area = shapefile_area(aoi)
orig_len = updated_tiles.shape[0]
track_rejected_inds = []
minimum_overlap_threshold = aoi_area - (250 * azimuth_mismatch)
print("")
print("AOI coverage: {}".format(aoi_area))
print("Allowable area of miscoverage: {}".format(250 * azimuth_mismatch))
print("minimum_overlap_threshold: {}".format(minimum_overlap_threshold))
print("")
if minimum_overlap_threshold < 0:
raise Exception('WARNING: user-defined mismatch of {}km\u00b2 too large relative to specified AOI'.format(azimuth_mismatch))
for i in enumerate(updated_tiles_dict):
intersection_area = aoi.intersection(i[1]['geometry'])
overlap_area = shapefile_area(intersection_area)
# Kick out scenes below specified overlap threshold
if minimum_overlap_threshold > overlap_area:
for iter_ind in enumerate(i[1]['ind_col']):
track_rejected_inds.append(iter_ind[1])
print("Rejected scene {} has only {}km\u00b2 overlap with AOI".format( \
i[1]['fileID'][iter_ind[0]], int(overlap_area)))
drop_ind = updated_tiles[updated_tiles['ind_col'] == iter_ind[1]].index
updated_tiles = updated_tiles.drop(index=drop_ind)
updated_tiles = updated_tiles.reset_index(drop=True)
print("{}/{} scenes rejected for not meeting defined spatial criteria".format( \
orig_len-updated_tiles.shape[0], orig_len))
# record rejected scenes separately
rejected_scenes_dict = [item for item in updated_tiles_dict \
if (any(x in track_rejected_inds for x in item['ind_col']))]
# pass scenes that are not tracked as rejected
updated_tiles_dict = [item for item in updated_tiles_dict \
if not (any(x in track_rejected_inds for x in item['ind_col']))]
return updated_tiles, pd.DataFrame(updated_tiles_dict), pd.DataFrame(gap_scenes_dict), pd.DataFrame(rejected_scenes_dict)
def pair_spatial_check(tiles,
aoi,
azimuth_mismatch=0.01,
iter_id='fileID'):
"""
Santity check function to confirm selected pairs meet user-defined spatial coverage threshold
"""
tiles['ind_col'] = range(0, len(tiles))
tiles = tiles.drop(columns=['reference', 'secondary'])
tiles_dict, dropped_pairs, gap_scenes_dict = continuous_time(tiles, iter_id='ind_col')
# Kick out scenes that do not meet user-defined spatial threshold
aoi_area = shapefile_area(aoi)
orig_len = tiles.shape[0]
minimum_overlap_threshold = aoi_area - (250 * azimuth_mismatch)
if minimum_overlap_threshold < 0:
raise Exception('WARNING: user-defined mismatch of {}km\u00b2 too large relative to specified AOI'.format(azimuth_mismatch))
for i in enumerate(tiles_dict):
intersection_area = aoi.intersection(i[1]['geometry'])
overlap_area = shapefile_area(intersection_area)
# Kick out scenes below specified overlap threshold
if minimum_overlap_threshold > overlap_area:
for iter_ind in enumerate(i[1]['ind_col']):
print("Rejected pair {} has only {}km\u00b2 overlap with AOI {}ID {}Ind".format( \
i[1]['reference_date'][iter_ind[0]].replace('-', '') + '_' + \
i[1]['secondary_date'][iter_ind[0]].replace('-', ''), \
overlap_area, iter_ind[1], i[0]))
drop_ind = tiles[tiles['ind_col'] == iter_ind[1]].index
tiles = tiles.drop(index=drop_ind)
tiles = tiles.reset_index(drop=True)
print("{}/{} scenes rejected for not meeting defined spatial criteria".format( \
orig_len-tiles.shape[0], orig_len))
return pd.DataFrame(tiles_dict)
df_aoi = gpd.read_file(aoi_shapefile)
aoi = df_aoi.geometry.unary_union
aoi
df_aoi
# Currently, there is a lot of data in each of the rows above. We really only need the AOI `geometry` and the `path_number`.
path_numbers = df_aoi.path_number.unique().tolist()
path_numbers
# # Generate a stack
#
# Using all the tiles that are needed to cover the AOI we make a geometric query based on the frame. We now include only the path we are interested in.
path_dict = {}
path_dict['pathNumber'] = str(path_numbers[0])
aoi_geometry = pd.DataFrame([path_dict])
aoi_geometry = gpd.GeoDataFrame(aoi_geometry, geometry=[shape(aoi)], crs=CRS.from_epsg(4326))
aoi_geometry['pathNumber'] = aoi_geometry['pathNumber'].astype(int)
df_stack = get_s1_stack_by_dataframe(aoi_geometry,
path_numbers=path_numbers)
f'We have {df_stack.shape[0]} frames in our stack'
# +
fig, ax = plt.subplots()
df_stack.plot(ax=ax, alpha=.5, color='green', label='Frames interesecting tile')
df_aoi.exterior.plot(color='black', ax=ax, label='AOI')
plt.legend()
# -
# Note, we now see the frames cover the entire AOI as we expect.
# First remove all scenes that do not produce spatiotemporally contiguous pairs and not meet specified intersection threshold
df_stack, df_stack_dict, gap_scenes_dict, rejected_scenes_dict = minimum_overlap_query(df_stack, aoi, azimuth_mismatch=azimuth_mismatch)
f'We have {df_stack.shape[0]} frames in our stack'
# Plot acquisitions that aren't continuous (i.e. have gaps)
if not gap_scenes_dict.empty:
gap_scenes_dict = gap_scenes_dict.sort_values(by=['start_date'])
for index, row in gap_scenes_dict.iterrows():
fig, ax = plt.subplots()
p = gpd.GeoSeries(row['geometry'])
p.exterior.plot(color='black', ax=ax, label=row['start_date_str'][0])
df_aoi.exterior.plot(color='red', ax=ax, label='AOI')
plt.legend()
plt.show
# Plot all mosaicked acquisitions that were rejected for not meeting user-specified spatial constraints
if not rejected_scenes_dict.empty:
rejected_scenes_dict = rejected_scenes_dict.sort_values(by=['start_date'])
fig, ax = plt.subplots()
for index, row in rejected_scenes_dict.iterrows():
p = gpd.GeoSeries(row['geometry'])
p.exterior.plot(color='black', ax=ax)
df_aoi.exterior.plot(color='red', ax=ax, label='AOI')
plt.legend()
# Plot each individual mosaicked acquisitions that were rejected for not meeting user-specified spatial constraints
if not rejected_scenes_dict.empty:
for index, row in rejected_scenes_dict.iterrows():
fig, ax = plt.subplots()
p = gpd.GeoSeries(row['geometry'])
p.exterior.plot(color='black', ax=ax, label=row['start_date_str'][0])
df_aoi.exterior.plot(color='red', ax=ax, label='AOI')
plt.legend()
plt.show
# Plot all mosaicked acquisitions that meet user-defined spatial coverage
# +
fig, ax = plt.subplots()
for index, row in df_stack_dict.iterrows():
p = gpd.GeoSeries(row['geometry'])
p.exterior.plot(color='black', ax=ax)
df_aoi.exterior.plot(color='red', ax=ax, label='AOI')
plt.legend()
# -
# Next, we filter the stack by month to ensure we only have SLCs we need.
df_stack_month = df_stack[df_stack.start_date.dt.month.isin(MONTHS_OF_INTEREST)]
# We will create a list of ```min_reference_dates``` by taking the most recent date from the SLC stack ```df_stack_month``` as the start date. A range of dates with interval of ```-6 days``` is then generated until ```(START_YEAR, MONTHS_OF_INTEREST[0], 1)```.
latestSLC_date = df_stack_month.iloc[-1].start_date
min_reference_dates = [latestSLC_date - datetime.timedelta(days=i) for i in range(0, (latestSLC_date-datetime.datetime(START_YEAR,MONTHS_OF_INTEREST[0],1)).days, 6)]
# We can now enumerate the SLC pairs that will produce the interferograms (GUNWs) based on initially defined parameters that are exposed at the top-level of this jupyter notebook.
# +
ifg_pairs = []
for min_ref_date in tqdm(min_reference_dates):
temporal_window_days = 365*3
temp = enumerate_ifgs_from_stack(df_stack_month,
aoi,
min_ref_date,
enumeration_type='tile', # options are 'tile' and 'path'. 'path' processes multiple references simultaneously
min_days_backward=0,
num_neighbors_ref=1,
num_neighbors_sec=num_neighbors,
temporal_window_days=temporal_window_days,
min_tile_aoi_overlap_km2=.1,#Minimum reference tile overlap of AOI in km2
min_ref_tile_overlap_perc=.1,#Relative overlap of secondary frames over reference frame
minimum_ifg_area_km2=0.1,#The minimum overlap of reference and secondary in km2
minimum_path_intersection_km2=.1,#Overlap of common track union with respect to AOI in km2
)
ifg_pairs += temp
# -
f'The number of GUNWs (likely lots of duplicates) is {len(ifg_pairs)}'
# # Get Dataframe
df_pairs = distill_all_pairs(ifg_pairs)
f"# of GUNWs: ' {df_pairs.shape[0]}"
# As a sanity check, confirm all IFG pairs meet user-defined spatial coverage
# Check if there are any gaps in the mosaicked IFGs, or if any are rejected for not meeting user-specified spatial constraints
#
# *NOTE: No products should be rejected at this stage. If any are, there is a problem either due to a:
# 1) Loose constraint on `azimuth_mismatch` variable whereby scenes not encompassing the entire AOI are getting passed
# 2) User-driven error
# 3) Bug in the code
df_pairs_dict = pair_spatial_check(df_pairs, aoi, azimuth_mismatch=azimuth_mismatch)
# +
fig, ax = plt.subplots()
for index, row in df_pairs_dict.iterrows():
p = gpd.GeoSeries(row['geometry'])
p.exterior.plot(color='black', ax=ax)
df_aoi.exterior.plot(color='red', ax=ax, label='AOI')
plt.legend()
# -
# # Deduplication Pt. 1
#
# A `GUNW` is uniquely determined by the reference and secondary IDs. We contanenate these sorted lists and generate a lossy hash to deduplicate products we may have introduced from the enumeration above.
# +
import hashlib
import json
def get_gunw_hash_id(reference_ids: list, secondary_ids: list) -> str:
all_ids = json.dumps([' '.join(sorted(reference_ids)),
' '.join(sorted(secondary_ids))
]).encode('utf8')
hash_id = hashlib.md5(all_ids).hexdigest()
return hash_id
# +
def hasher(row):
return get_gunw_hash_id(row['reference'], row['secondary'])
df_pairs['hash_id'] = df_pairs.apply(hasher, axis=1)
df_pairs.head(5)
# -
f"# of duplicated entries: {df_pairs.duplicated(subset=['hash_id']).sum()}"
df_pairs = df_pairs.drop_duplicates(subset=['hash_id']).reset_index(drop=True)
f"# of UNIQUE GUNWs: {df_pairs.shape[0]}"
# # Viewing GUNW pairs
# +
# start index
M = 0
# number of pairs to view
N = 5
for J in range(M, M + N):
pair = ifg_pairs[J]
fig, axs = plt.subplots(1, 2, sharey=True, sharex=True)
df_ref_plot = pair['reference']
df_sec_plot = pair['secondary']
df_ref_plot.plot(column='start_date_str',
legend=True,
ax=axs[0], alpha=.15)
df_aoi.exterior.plot(ax=axs[0], alpha=.5, color='black')
axs[0].set_title('Reference')
df_sec_plot.plot(column='start_date_str',
legend=True,
ax=axs[1], alpha=.15)
df_aoi.exterior.plot(ax=axs[1], alpha=.5, color='black')
axs[0].set_title(f'Reference {J}')
axs[1].set_title('Secondary')
# -
# # Update types for Graphical Analysis
#
# We want to do some basic visualization to support the understanding if we traverse time correctly. We do some simple standard pandas manipulation.
df_pairs['reference_date'] = pd.to_datetime(df_pairs['reference_date'])
df_pairs['secondary_date'] = pd.to_datetime(df_pairs['secondary_date'])
df_pairs.head()
# # Visualize a Date Graph from Time Series
#
# We can put this into a network Directed Graph and use some simple network functions to check connectivity.
#
# We are going to use just dates for nodes, though you could use `(ref_date, hash_id)` for nodes and then inspect connected components. That is for another notebook.
list(zip(df_pairs.reference_date, df_pairs.secondary_date))[:15]
unique_dates = df_pairs.reference_date.tolist() + df_pairs.secondary_date.tolist()
unique_dates = sorted(list(set(unique_dates)))
unique_dates[:5]
date2node = {date: k for (k, date) in enumerate(unique_dates)}
node2date = {k: date for (date, k) in date2node.items()}
# +
# %matplotlib widget
G = nx.DiGraph()
edges = [(date2node[ref_date], date2node[sec_date])
for (ref_date, sec_date) in zip(df_pairs.reference_date, df_pairs.secondary_date)]
G.add_edges_from(edges)
# -
nx.draw(G)
# This function checks there is a path from the first date to the last one. The y-axis is created purely for display so doesn't really indicated anything but flow by month.
nx.has_path(G,
target=date2node[unique_dates[0]],
source=date2node[unique_dates[-1]])
# Ensure that the result above returns a ```True``` value to be able to produce a time-series.
# +
fig, ax = plt.subplots(figsize=(15, 5))
increment = [date.month + date.day for date in unique_dates]
# source: https://stackoverflow.com/a/27852570
scat = ax.scatter(unique_dates, increment)
position = scat.get_offsets().data
pos = {date2node[date]: position[k] for (k, date) in enumerate(unique_dates)}
nx.draw_networkx_edges(G, pos=pos, ax=ax)
ax.grid('on')
ax.tick_params(axis='x',
which='major',
labelbottom=True,
labelleft=True)
ymin, ymax = ax.get_ylim()
# for y in range(2014, 2022):
# label = 'June to Oct' if y == 2016 else None
# ax.fill_between([datetime.datetime(y, 6, 1), datetime.datetime(y, 11, 1)],
# ymin, ymax,
# alpha=.5, color='green', zorder=0, label=label)
# plt.legend()
# -
# Observe there is a gap in 2018 over are area of interest. This is where our 3 year "temporal_window_days" parameter in our enumeration was essential.
# # Deduplication Pt. 2
#
# This is to ensure that previous processing hasn't generate any of the products we have just enumerated.
#
#
# # Check CMR
#
# This function checks the ASF DAAC if there are GUNWs with the same spatial extent and same date pairs as the ones created. At some point, we will be able to check the input SLC ids from CMR, but currently that is not possible.
#
# If you are processing a new AOI whose products have not been delivered, you can ignore this step. It is a bit time consuming as the queries are done product by product.
# +
from s1_enumerator import duplicate_gunw_found
import concurrent
from tqdm import tqdm
n = df_pairs.shape[0]
with concurrent.futures.ThreadPoolExecutor(max_workers=15) as executor:
results = list(tqdm(executor.map(duplicate_gunw_found, df_pairs.to_dict('records')), total=n))
# -
df_pairs['existing_gunw'] = [r != '' for r in results]
df_pairs['existing_gunw_id'] = results
total_existing_gunws = df_pairs['existing_gunw'].sum()
print('existing_gunws: ', total_existing_gunws)
print('Total pairs', df_pairs.shape[0])
df_pairs_filtered = df_pairs[~df_pairs['existing_gunw']].reset_index(drop=True)
# df_pairs_filtered.drop_duplicates(subset=['hash_id'], inplace=True)
print('after filtering, total pairs: ', df_pairs_filtered.shape[0])
# # Check Hyp3 Account
#
# We are now going to check
#
# 1. check products in the open s3 bucket
# 2. check running/pending jobs
#
# Notes:
#
# 1. Above, to accomplish step 1., there is some verbose code (see below). Once we automate delivery, this step will be obsolete. However, until we have delivery, we have to make sure that there are no existing products. Additionally, if we are using a separate (non-operational account), then would be good to use this.
# 2. If we are debugging products and some of our previously generated products were made incorrectly, we will want to ignore this step.
# +
import hyp3_sdk
# uses .netrc; add `prompt=True` to prompt for credentials;
hyp3_isce = hyp3_sdk.HyP3(deploy_url)
pending_jobs = hyp3_isce.find_jobs(status_code='PENDING') + hyp3_isce.find_jobs(status_code='RUNNING')
all_jobs = hyp3_isce.find_jobs()
# -
print(all_jobs)
# ## 1. Get existing products in s3 bucket
job_data = [j.to_dict() for j in all_jobs]
job_data[0]
# Get bucket (there is only one)
job_data_s3 = list(filter(lambda job: 'files' in job.keys(), job_data))
bucket = job_data_s3[0]['files'][0]['s3']['bucket']
# Get all keys
job_keys = [job['files'][0]['s3']['key'] for job in job_data_s3]
# +
from botocore import UNSIGNED
from botocore.config import Config
s3 = boto3.resource('s3',config=Config(signature_version=UNSIGNED))
prod_bucket = s3.Bucket(bucket)
objects = list(prod_bucket.objects.all())
ncs = list(filter(lambda x: x.key.endswith('.nc'), objects))
#ncs[:10]
# -
# Need to physically check if the products are not there (could have been deleted!)
nc_keys = [nc_ob.key for nc_ob in ncs]
jobs_with_prods_in_s3 = [job for (k, job) in enumerate(job_data_s3) if job_keys[k] in nc_keys]
slcs = [(job['job_parameters']['granules'],
job['job_parameters']['secondary_granules'])
for job in jobs_with_prods_in_s3]
slcs[:2]
hash_ids_of_prods_in_s3 = [get_gunw_hash_id(*slc) for slc in slcs]
f"We are removing {df_pairs_filtered['hash_id'].isin(hash_ids_of_prods_in_s3).sum()} GUNWs for submission"
items = hash_ids_of_prods_in_s3
df_pairs_filtered = df_pairs_filtered[~df_pairs_filtered['hash_id'].isin(items)].reset_index(drop=True)
f"Current # of GUNWs: {df_pairs_filtered.shape[0]}"
# ## 2. Running or Pending Jobs
pending_job_data = [j.to_dict() for j in pending_jobs]
pending_slcs = [(job['job_parameters']['granules'],
job['job_parameters']['secondary_granules'])
for job in pending_job_data]
hash_ids_of_pending_jobs = [get_gunw_hash_id(*slc) for slc in pending_slcs]
hash_ids_of_pending_jobs[:4]
items = hash_ids_of_pending_jobs
f"We are removing {df_pairs_filtered['hash_id'].isin(items).sum()} GUNWs for submission"
items = hash_ids_of_pending_jobs
df_pairs_filtered = df_pairs_filtered[~df_pairs_filtered['hash_id'].isin(items)].reset_index(drop=True)
f"Current # of GUNWs: {df_pairs_filtered.shape[0]}"
# # Submit jobs to Hyp3
records_to_submit = df_pairs_filtered.to_dict('records')
records_to_submit[0]
# The below puts the records in a format that we can submit to the Hyp3 API.
#
# **Note 1**: there is an index in the records to submit to ensure we don't over submit jobs for generating GUNWs. \
# **Note 2**: uncomment the code to *actually* submit the jobs.
# +
import hyp3_sdk
# uses .netrc; add `prompt=True` to prompt for credentials;
hyp3_isce = hyp3_sdk.HyP3(deploy_url)
job_dicts = [{'name': job_name,
# NOTE: we are still using the `dev` branch. Change this to "INSAR_ISCE" to use the `main` branch.
'job_type': 'INSAR_ISCE_TEST',
'job_parameters': {'granules': r['reference'],
'secondary_granules': r['secondary']}}
# NOTE THERE IS AN INDEX - this is to submit only a subset of Jobs
for r in records_to_submit]
# -
#UNCOMMENT TO SUBMIT
prepared_jobs = job_dicts
submitted_jobs = hyp3_sdk.Batch()
for batch in hyp3_sdk.util.chunk(prepared_jobs):
submitted_jobs += hyp3_isce.submit_prepared_jobs(batch)
# Query all jobs on the server
jobs = hyp3_isce.find_jobs()
print(jobs)
# Query your particular job
jobs = hyp3_isce.find_jobs(name=job_name)
print(jobs)
# +
# # create clean directory to deposit products in
if os.path.exists(prod_dir):
os.remove(prod_dir)
os.mkdir(prod_dir)
# -
# Below, we show how to download files. The multi-threading example will download products in parallel much faster than `jobs.download_files()`.
# +
jobs = hyp3_isce.find_jobs(name=job_name)
print(jobs)
import concurrent.futures
from tqdm import tqdm
with concurrent.futures.ThreadPoolExecutor(max_workers=10) as executor:
results = list(tqdm(executor.map(lambda job: job.download_files(), jobs), total=len(jobs)))
| LaPalma/0_LaPalma_Enumeration_T169.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ctKdQ8Hw9Sfi"
# # HackNITP Winter '21 AI Challenge
# + colab={"base_uri": "https://localhost:8080/"} id="4EAPyLVr9Sfv" outputId="cd383040-c030-454b-ec8b-b008d3439f79"
# Download the data
# !wget -O "hacknitp_winter_'21_ai_challenge-dataset.zip" "https://dockship-job-models.s3.ap-south-1.amazonaws.com/be39200da389544f8ecef9fc9b43781d?X-Amz-Algorithm=AWS4-HMAC-SHA256&X-Amz-Credential=AKIAIDOPTEUZ2LEOQEGQ%2F20210117%2Fap-south-1%2Fs3%2Faws4_request&X-Amz-Date=20210117T130233Z&X-Amz-Expires=1800&X-Amz-Signature=315045dec29bf229baffd1758ffba9a5926a621cf1d91df93d341c4410c8cbca&X-Amz-SignedHeaders=host&response-content-disposition=attachment%3B%20filename%3D%22hacknitp_winter_%2721_ai_challenge-dataset.zip%22"
# + id="l1GHbuJv9c2p"
# Required modules
import os
import cv2
import glob
import numpy as np
import pandas as pd
import librosa as lr
import tensorflow as tf
from zipfile import ZipFile
from matplotlib import pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (12, 7)
# + id="mN4i_3u6BuHP"
# Extracting the dataset
if not os.path.isfile('TRAIN.csv'):
with ZipFile('hacknitp_winter_\'21_ai_challenge-dataset.zip', 'r') as zf:
zf.extractall('./')
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="OYjKEnYC9c78" outputId="057b94ee-8702-4351-e591-06b3400d0df8"
# Loading the train meta-data
data = pd.read_csv('TRAIN.csv')
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 331} id="ATn011MF9c-S" outputId="21748553-1659-4b9e-8e42-be578df56fd1"
# Inspecting the data
data.info()
data.describe()
# + id="LXj587N4LJP3"
# Helper Functions
def fix_audio_length(fname, duration=2.97, sr=22050):
in_len = sr * duration
audio_data, sr = lr.load(fname, sr=sr, duration=duration)
dur = lr.get_duration(audio_data)
if round(dur) < duration:
audio_data = lr.util.fix_length(audio_data, round(in_len))
spec = lr.feature.melspectrogram(audio_data, sr=sr)
return spec
# + id="yT7MgPPbrWBV"
# Directory paths
train_dir = './TRAIN/'
test_dir = './TEST/'
# + id="KJuIcpGTs6DG"
# Constants
MAPPER = {'Positive': 0, 'Neutral': 1, 'Negative': 2}
REVERSE_MAPPER = {0: 'Positive', 1: 'Neutral', 2: 'Negative'}
# + id="nc-dQe4Nr-cC"
# Training data
X = []
y = []
for fname in os.listdir(train_dir):
spectrogram = fix_audio_length(train_dir+fname)
if spectrogram.shape != (128, 128):
spectrogram = cv2.resize(spectrogram, (128, 128))
label = MAPPER[data[data['Filename'] == fname]['Class'].tolist()[0]]
X.append(np.expand_dims(spectrogram, 2))
y.append([0 if i != label else 1 for i in range(3)])
X = np.array(X)
y = np.array(y)
# + id="D0gQ8TCvvbDl"
# Testing data
X_test = []
for fname in os.listdir(test_dir):
spectrogram = fix_audio_length(test_dir+fname)
if spectrogram.shape != (128, 128):
spectrogram = cv2.resize(spectrogram, (128, 128))
X_test.append(np.expand_dims(spectrogram, 2))
X_test = np.array(X_test)
# + id="KfZwv1sGBiB8"
# Defining the model
input_shape = (128, 128, 1)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(16, (5, 5), strides=(1, 1), input_shape=input_shape),
tf.keras.layers.MaxPooling2D((4, 2), strides=(4, 2)),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Conv2D(32, (5, 5), padding="valid"),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(3),
tf.keras.layers.Activation('softmax')
])
# + id="BVU1Fqu9CK9y"
# Compiling the model
optim = tf.keras.optimizers.Adam()
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="XuBH5YsiRzSf" outputId="612b23c4-c9d4-453b-d3f3-5a848ac34ab5"
# Model Summary
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="MoyLDeXkCN4H" outputId="c0b905e5-72df-42e2-810d-c9cab9496fb4"
# Fitting the model
history = model.fit(X, np.array(y), epochs=50, batch_size=16)
# + id="b-DkuOQLCUc_" colab={"base_uri": "https://localhost:8080/", "height": 630} outputId="23e89678-4790-4305-c19c-30891a719eed"
# Plotting the accuracy
plt.plot(history.history['accuracy'], label='accuracy')
plt.plot(history.history['val_accuracy'], label = 'val_accuracy')
plt.legend()
# + colab={"base_uri": "https://localhost:8080/", "height": 630} id="9jpYsakxdVY1" outputId="82f3ab78-3de6-4a6c-c650-fb4cb3831f04"
# Plotting the loss
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label = 'val_loss')
plt.legend()
# + id="F-LB8L1_Utpf"
# Prediction on the test set
pred = np.argmax(model.predict(np.array(X_test)), axis=1)
pred = [REVERSE_MAPPER[p] for p in pred]
# + id="LJYn2S7TAdZh"
# Submission
submission = pd.DataFrame({'Filename': os.listdir('./TEST'), 'Class': pred})
submission.to_csv('output.csv', index=False)
| HackNITP Winter '21 AI Challenge 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # City Weather vs. Latitude Coordinates
# ----
#
# In this analysis, we utilized Python code in conjunction with open access API systems to curate weather and geographic coordinates of 500+ cities located around the globe. By cleaning and analyzing data gathered from OpenWeather API, some conclusions could be made on the basis of city weather in relation to the location of the equator:
#
# 1. There were distinctive differences in the relationship of maximum ambient temperature and latitude coordinates between the northern and southern hemispheres. The northern hemisphere had a defined negative relationship as latitude increased, with the correlation coefficient equaling -1.42, meaning that as the latitude increased and cities were located further away from the equator, the temperatures dropped substantially. This would make sense as this data was collected in February 2021 and the northern hemisphere is in it's winter solstice during that time. The opposite can be observed with the analysis of temperature and latitude in the southern hemisphere, which had a slightly positive relationship with a positive correlation coefficient of 0.28. As the latitude is further away from the equator maximum temperature is lower, however there is a weaker coefficient for the southern hemisphere in February 2021 as it is currently within it's summer solstice so on average the temperatures are higher.
#
# 2. For both hemispheres, there seems to be minimal influence or correlations between percent humidity and a city's latitude coordinates. With very low correlation coefficients (less than 0.5) displayed in the linear regression analysis, observing relationships based on temperatures would be more accurate as humidity is dependent on the temperature levels.
#
# 3. Given that wind speed is affected by changes in air pressure in relation to temperature differences in the atmosphere, it makes sense that the southern hemisphere would have a slightly negative assocation with wind speed and latitude. As cities are located closer to the equator the air temperature is naturally higher, so there is less cool air dropping as warm air rises. The further away a city is from the equator the ambient temperature is generally cooler, so as warm air rises there is a greater change in air flow as cool air descends and increases overall wind speed.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
import scipy.stats as st
from scipy.stats import linregress
# Import API key
from api_keys import weather_api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
# -
# ## Generate Cities List
# +
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
# +
#debugging only
# cities = cities[0:10]
# cities
# -
# ### Perform API Calls
# * Perform a weather check on each city using a series of successive API calls.
# * Include a print log of each city as it'sbeing processed (with the city number and city name).
#
# +
#create query url to scan API
base_url = "http://api.openweathermap.org/data/2.5/weather?"
units = "imperial"
query_url = f"{base_url}&appid={weather_api_key}&units={units}&q="
# -
#list for analysis parameters
city_name = []
city_lat = []
city_lng = []
city_country =[]
city_date =[]
city_temp = []
city_humidity = []
city_cloud_cover = []
city_wind_speed = []
# +
print("Retrieving Desired City Data")
print(f"----------------------------------")
records = 0
records_set = 1
#for loop to go through each city from the API with try/except block so code doesn't break
for city in cities:
city_url = f"{query_url}{city}"
records = records + 1
try:
response = requests.get(city_url).json()
time.sleep(0.5)
print(f"Processing city number {records}")
print(response)
print()
print()
city_name.append(response["name"])
city_lat.append(response["coord"]["lat"])
city_lng.append(response["coord"]["lon"])
city_country.append(response["sys"]["country"])
city_date.append(response["dt"])
city_temp.append(response["main"]["temp_max"])
city_humidity.append(response["main"]["humidity"])
city_cloud_cover.append(response["clouds"]["all"])
city_wind_speed.append(response["wind"]["speed"])
#Conditional for group city outputs
if records > 50:
records_set += 1
records = 1
except:
print(f"City not found")
print(f"----------------------------------")
print(f"End of Data Retrieval Process")
print(f"----------------------------------")
# -
# ### Convert Raw Data to DataFrame
# * Export the city data into a .csv.
# * Display the DataFrame
city_weather_df = pd.DataFrame({"City":city_name, "Latitude":city_lat, "Longitude":city_lng, "Country":city_country, "Date":city_date, "Max Temperature":city_temp, "Humidity":city_humidity, "Cloudiness":city_cloud_cover, "Wind Speed":city_wind_speed})
city_weather_df.head()
city_weather_df.to_csv("City_Weather.csv", index=False, header=True)
# ## Inspect the data and remove the cities where the humidity > 100%.
# ----
# Skip this step if there are no cities that have humidity > 100%.
city_weather_df["Humidity"].max()
# ## Plotting the Data
# * Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
# * Save the plotted figures as .pngs.
# ## Latitude vs. Temperature Plot
# +
#Assign data to new variables
latitude = city_weather_df["Latitude"]
temperature = city_weather_df["Max Temperature"]
#Plot scatter plot with x and y values
plt.figure(figsize = (18,10))
plt.scatter(latitude, temperature)
#create x- and y-axis labels and a chart title
plt.title(f"City Latitude vs. Maximum Temperature (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Maximum Temperature (F)", fontsize = 15)
plt.savefig("../Images/Latitude_vs_Max_Temp_Plot.png")
plt.show()
# -
# This code is visualizing the maximum temperatures (F) of the 600+ cities found in comparison to their latitude coordinates. This scatter plot is showing that as the latitude moves further away from 0 (the equator), the maximum temperature declines for both positive and negative latitudes.
# ## Latitude vs. Humidity Plot
# +
#Assign new variables
latitude = city_weather_df["Latitude"]
humidity = city_weather_df["Humidity"]
#Plot figure
plt.figure(figsize = (20, 10))
plt.scatter(latitude, humidity)
#chart labels and save plot image
plt.title(f"City Latitude vs. Percent Humidity (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize=15)
plt.ylabel("Humidity (%)", fontsize = 15)
plt.savefig("../Images/Latitude_vs_Humidity_Plot.png")
plt.show()
# -
# This code is visualizing the percent humidity measurements of the 600+ random cities in comparison to their latitude coordinates. This scatter plot is indicating that humidity may not be dependent on latitude, as there are high and low humidity percentages for both city coordinates closer the the equator and those that are located further away.
# ## Latitude vs. Cloudiness Plot
# +
#define variables
latitude = city_weather_df["Latitude"]
cloudiness = city_weather_df["Cloudiness"]
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(latitude, cloudiness)
#designate labels and save as png file
plt.title(f"City Latitude vs. Percent Cloudiness(%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Cloudiness (%)", fontsize = 15)
plt.savefig("../Images/Latitude_vs_Cloudiness_Plot.png")
plt.show()
# -
# This code is visualizing the percent cloudiness of 600+ randomly selected cities around the world in relation to their latitude coordinates. This scatter plot is indicating that there may not be a strong assocation between percent cloud cover and latitude coordinates, as the points are widely distributed across the graph with high and low percentages at coordinates closer and further away from the equator.
# ## Latitude vs. Wind Speed Plot
# +
#define variables
latitude = city_weather_df["Latitude"]
wind_speed = city_weather_df["Wind Speed"]
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(latitude, wind_speed)
#assign labels and save to png file
plt.title(f"City Latitude vs. Wind Speed (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Wind Speed (MPH)", fontsize = 15)
plt.savefig("../Images/Latitude_vs_Wind_Speed.png")
plt.show()
# -
# This code is visualizing the relationship between the latitude coordinates of 600+ randomly selected cities around the globe and their calculated wind speeds, measured in mph. This scatter plot is indicating that generally speaking there are few outliers of high wind speeds across all latitude coordinates, meaning that wind speed may be attributed to different factors beyond latitude.
# ## Linear Regression
# #### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
#Use .loc() function to filter for city latitudes above the equator
n_lats = city_weather_df.loc[(city_weather_df["Latitude"] > 0)]
#define variables
north_latitude = n_lats["Latitude"]
north_max_temp = n_lats["Max Temperature"]
#designate linear regression between latitude and max temp
temp_lat_slope, temp_lat_int, temp_lat_r, temp_lat_p, temp_lat_std_err = st.linregress(north_latitude, north_max_temp)
#create slope intercept equation
temp_lat_best_fit = temp_lat_slope * north_latitude + temp_lat_int
#convert to y=mx+b format for graph
north_temp_equation = "y=" + str(round(temp_lat_slope, 2)) + "x+" + str(round(temp_lat_int, 2))
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(north_latitude, north_max_temp)
#Plot linear regression
plt.plot(north_latitude, temp_lat_best_fit, "--", color = "red")
#plot y=mx+b equation on chart
plt.annotate(north_temp_equation, (0, 50), fontsize = 15, color="red")
#assign labels and save to png file
plt.title(f"Northern Hemisphere Max Temperature vs Latitude (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Maximum Temperatures (F)", fontsize = 15)
#include r-value in output
print(f"r=value: {temp_lat_r}")
plt.savefig("../Images/North_Hem_Max_Temp_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between maximum temperature of 600+ random cities around the globe and their associated latitude coordinates in the northern hemisphere. This scatter plot indicates that as cities are located further away from the equator, their maximum temperature decreases which is expected.
# #### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
# +
#.loc() for cities below equator and define variables
s_lats = city_weather_df.loc[city_weather_df["Latitude"] <0]
south_latitude = s_lats["Latitude"]
south_max_temp = s_lats["Max Temperature"]
#linear regression/slope-intercept
s_lat_slope, s_lat_int, s_lat_r, s_lat_p, s_lat_std_err = st.linregress(south_latitude, south_max_temp)
s_lat_fit = s_lat_slope * south_latitude + s_lat_int
#y=mx+b equation
s_lat_equation = "y=" + str(round(s_lat_slope, 2)) + "x+" + str(round(s_lat_int, 2))
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(south_latitude, south_max_temp)
#Plot linear regression
plt.plot(south_latitude, s_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(s_lat_equation, (-45,80), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Southern Hemisphere Max Temperatures vs Latitude (%s)" % time.strftime("%x"), fontsize = 20, )
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Maximum Temperatures (F)", fontsize = 15)
#designate r-value
print(f"r-value: {s_lat_r}")
plt.savefig("../Images/South_Hem_Max_Temp_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between maximum temperature of 600+ random cities around the globe and their associated latitude coordinates in the southern hemisphere. This scatter plot indicates that as cities move closer to the equator, there is a positive relationship between temperature and latitude, i.e. they experience higher temperatures which is expected.
# #### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
#Define variables
north_latitude = n_lats["Latitude"]
north_humidity = n_lats["Humidity"]
#linear regression/slope intercept
n_lat_slope, n_lat_int, n_lat_r, n_lat_p, n_lat_std_err = st.linregress(north_latitude, north_humidity)
n_lat_fit = n_lat_slope * north_latitude + n_lat_int
#y=mx+b equation
n_lat_equation = "y=" + str(round(n_lat_slope, 2)) + "x+" + str(round(n_lat_int, 2))
#plot figure
plt.figure(figsize = (15,10))
plt.scatter(north_latitude, north_humidity)
#Plot linear regression
plt.plot(north_latitude, n_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(n_lat_equation, (0,50), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Northern Hemisphere Humidity vs Latitude (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Humidity (%)", fontsize = 15)
#designate r-value
print(f"r-value: {n_lat_r}")
plt.savefig("../Images/North_Hem_Humidity_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between percent humidity of 600+ random cities around the globe and their associated latitude coordinates in the northern hemisphere. This scatter plot indicates that there is a minimal positive relationship between humidity levels and latitude coordinates, as some locations closer to the equator have lower humidity percentage and some locations further from the equator experience higher humidity percentages. Given that there is a wide array of city points beyond the best fit line of this linear regression, there may not be any definitive conclusions to be made on the association of latitude and humidity.
# #### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
# +
#Define variables
south_latitude = s_lats["Latitude"]
south_humidity = s_lats["Humidity"]
#linear regression/slope-intercept
s_lat_slope, s_lat_int, s_lat_r, s_lat_p, s_lat_std_err = st.linregress(south_latitude, south_humidity)
s_lat_fit = s_lat_slope * south_latitude + s_lat_int
#y=mx+b equation
s_lat_equation = "y=" + str(round(s_lat_slope, 2)) + "x+" + str(round(s_lat_int, 2))
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(south_latitude, south_humidity)
#Plot linear regression
plt.plot(south_latitude, s_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(s_lat_equation, (-45,80), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Southern Hemisphere Humidity vs Latitude (%s)" % time.strftime("%x"), fontsize = 20, )
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Humidity (%)", fontsize = 15)
#designate r-value
print(f"r-value: {s_lat_r}")
plt.savefig("../Images/South_Hem_Humidity_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between percent humidity of 600+ random cities around the globe and their associated latitude coordinates in the southern hemisphere. This scatter plot indicates that there is a minimal positive relationship between humidity levels and latitude coordinates, as some locations closer to the equator have lower humidity percentage and some locations further from the equator experience higher humidity percentages. Given that there is a wide array of city points beyond the best fit line of this linear regression, there may not be any definitive conclusions to be made on the association of latitude and humidity.
# #### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
#Define variables
north_latitude = n_lats["Latitude"]
north_cloudiness = n_lats["Cloudiness"]
#linear regression/slope intercept
n_lat_slope, n_lat_int, n_lat_r, n_lat_p, n_lat_std_err = st.linregress(north_latitude, north_cloudiness)
n_lat_fit = n_lat_slope * north_latitude + n_lat_int
#y=mx+b equation
n_lat_equation = "y=" + str(round(n_lat_slope, 2)) + "x+" + str(round(n_lat_int, 2))
#plot figure
plt.figure(figsize = (15,10))
plt.scatter(north_latitude, north_cloudiness)
#Plot linear regression
plt.plot(north_latitude, n_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(n_lat_equation, (35,60), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Northern Hemisphere Cloudiness vs Latitude (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Cloudiness (%)", fontsize = 15)
#designate r-value
print(f"r-value: {n_lat_r}")
plt.savefig("../Images/North_Hem_Cloud_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between percent cloudiness of 600+ random cities around the globe and their associated latitude coordinates in the northern hemisphere. This scatter plot indicates that there is a minimal positive relationship between humidity levels and latitude coordinates, as some locations closer to the equator have lower cloudiness percentage and some locations further from the equator experience higher cloudiness percentages. Given that there is a wide array of city points beyond the best fit line of this linear regression, there may not be any definitive conclusions to be made on the association of latitude and cloudiness.
# #### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
# +
#Define variables
south_latitude = s_lats["Latitude"]
south_cloudiness = s_lats["Cloudiness"]
#linear regression/slope-intercept
s_lat_slope, s_lat_int, s_lat_r, s_lat_p, s_lat_std_err = st.linregress(south_latitude, south_cloudiness)
s_lat_fit = s_lat_slope * south_latitude + s_lat_int
#y=mx+b equation
s_lat_equation = "y=" + str(round(s_lat_slope, 2)) + "x+" + str(round(s_lat_int, 2))
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(south_latitude, south_cloudiness)
#Plot linear regression
plt.plot(south_latitude, s_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(s_lat_equation, (-42,55), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Southern Hemisphere Cloudiness vs Latitude (%s)" % time.strftime("%x"), fontsize = 20, )
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Cloudiness (%)", fontsize = 15)
#designate r-value
print(f"r-value: {s_lat_r}")
plt.savefig("../Images/South_Hem_Cloud_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between percent cloudiness of 600+ random cities around the globe and their associated latitude coordinates in the southern hemisphere. This scatter plot indicates that there is a minimal positive relationship between humidity levels and latitude coordinates, as some locations closer to the equator have lower cloudiness percentage and some locations further from the equator experience higher cloudiness percentages. Given that there is a wide array of city points beyond the best fit line of this linear regression, there may not be any definitive conclusions to be made on the association of latitude and cloudiness.
# #### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
#Define variables
north_latitude = n_lats["Latitude"]
north_wind_speed = n_lats["Wind Speed"]
#linear regression/slope intercept
n_lat_slope, n_lat_int, n_lat_r, n_lat_p, n_lat_std_err = st.linregress(north_latitude, north_wind_speed)
n_lat_fit = n_lat_slope * north_latitude + n_lat_int
#y=mx+b equation
n_lat_equation = "y=" + str(round(n_lat_slope, 2)) + "x+" + str(round(n_lat_int, 2))
#plot figure
plt.figure(figsize = (15,10))
plt.scatter(north_latitude, north_wind_speed)
#Plot linear regression
plt.plot(north_latitude, n_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(n_lat_equation, (35,25), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Northern Hemisphere Wind Speed vs Latitude (%s)" % time.strftime("%x"), fontsize = 20)
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Wind Speed (mph)", fontsize = 15)
#designate r-value
print(f"r-value: {n_lat_r}")
plt.savefig("../Images/North_Hem_Wind_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between wind speed (mph) of 600+ random cities around the globe and their associated latitude coordinates in the northern hemisphere. This scatter plot indicates that there is a minimal positive relationship between humidity levels and latitude coordinates, as some locations closer to the equator have lower wind speeds and some locations further from the equator experience higher wind speeds. Given that the slope of the best fit line is as low as 0.04, it is indicative of a weaker association between latitude coordinates and average wind speeds.
# #### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
# +
#Define variables
south_latitude = s_lats["Latitude"]
south_wind_speed = s_lats["Wind Speed"]
#linear regression/slope-intercept
s_lat_slope, s_lat_int, s_lat_r, s_lat_p, s_lat_std_err = st.linregress(south_latitude, south_wind_speed)
s_lat_fit = s_lat_slope * south_latitude + s_lat_int
#y=mx+b equation
s_lat_equation = "y=" + str(round(s_lat_slope, 2)) + "x+" + str(round(s_lat_int, 2))
#plot figure
plt.figure(figsize = (20,10))
plt.scatter(south_latitude, south_wind_speed)
#Plot linear regression
plt.plot(south_latitude, s_lat_fit, "--", color="red")
#Add y=mx+b to chart
plt.annotate(s_lat_equation, (-50,12), color="red", fontsize = 15)
#Assign labels and save as png file
plt.title(f"Southern Hemisphere Wind Speed vs Latitude (%s)" % time.strftime("%x"), fontsize = 20, )
plt.xlabel("Latitude", fontsize = 15)
plt.ylabel("Wind Speed (mph)", fontsize = 15)
#designate r-value
print(f"r-value: {s_lat_r}")
plt.savefig("../Images/South_Wind_vs_Lat_Plot.png")
plt.show()
# -
# This code is visualizing the relationship between wind speeds of 600+ random cities around the globe and their associated latitude coordinates in the northern hemisphere. This scatter plot indicates that there is a minimal negative relationship between average wind speeds and latitude coordinates, as some locations closer to the equator have lower wind speeds and some locations further from the equator experience higher wind speeds. Given that there is a wide array of city points beyond the best fit line of this linear regression, and that the slope is as minimal as 0.06, there may not be any definitive conclusions to be made on the association of latitude and average wind speeds.
| WeatherPy/WeatherPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Procedural programming in python
#
# ## Topics
# * Tuples, lists and dictionaries
# * Flow control, part 1
# * If
# * For
# * range() function
# * Some hacky hack time
# * Flow control, part 2
# * Functions
# <hr>
# ### Tuples
#
# Let's begin by creating a tuple called `my_tuple` that contains three elements.
my_tuple = ('I', 'like', 'cake')
my_tuple
# Tuples are simple containers for data. They are ordered, meaining the order the elements are in when the tuple is created are preserved. We can get values from our tuple by using array indexing, similar to what we were doing with pandas.
my_tuple[0]
# Recall that Python indexes start at 0. So the first element in a tuple is 0 and the last is array length - 1. You can also address from the `end` to the `front` by using negative (`-`) indexes, e.g.
my_tuple[-1]
# You can also access a range of elements, e.g. the first two, the first three, by using the `:` to expand a range. This is called ``slicing``.
my_tuple[0:2]
my_tuple[0:3]
# What do you notice about how the upper bound is referenced?
#
# Without either end, the ``:`` expands to the entire list.
my_tuple[1:]
my_tuple[:-1]
my_tuple[:]
# Tuples have a key feature that distinguishes them from other types of object containers in Python. They are _immutable_. This means that once the values are set, they cannot change.
my_tuple[2]
# So what happens if I decide that I really prefer pie over cake?
# +
#my_tuple[2] = 'pie'
# -
# Facts about tuples:
# * You can't add elements to a tuple. Tuples have no append or extend method.
# * You can't remove elements from a tuple. Tuples have no remove or pop method.
# * You can also use the in operator to check if an element exists in the tuple.
#
# So then, what are the use cases of tuples?
# * Speed
# * `Write-protects` data that other pieces of code should not alter
# You can alter the value of a tuple variable, e.g. change the tuple it holds, but you can't modify it.
my_tuple
my_tuple = ('I', 'love', 'pie')
my_tuple
# There is a really handy operator ``in`` that can be used with tuples that will return `True` if an element is present in a tuple and `False` otherwise.
'love' in my_tuple
# Finally, tuples can contain different types of data, not just strings.
import math
my_second_tuple = (42, 'Elephants', 'ate', math.pi)
my_second_tuple
# Numerical operators work... Sort of. What happens when you add?
#
# ``my_second_tuple + 'plus'``
# Not what you expects? What about adding two tuples?
my_second_tuple + my_tuple
# Other operators: -, /, *
# ### Questions about tuples before we move on?
# <hr>
#
# ### Lists
#
# Let's begin by creating a list called `my_list` that contains three elements.
my_list = ['I', 'like', 'cake']
my_list
# At first glance, tuples and lists look pretty similar. Notice the lists use '[' and ']' instead of '(' and ')'. But indexing and refering to the first entry as 0 and the last as -1 still works the same.
my_list[0]
my_list[-1]
my_list[0:3]
# Lists, however, unlike tuples, are mutable.
my_list[2] = 'pie'
my_list
# Multiple elements in the list can even be changed at once!
my_list[1:] = ['love', 'puppies']
my_list
# You can still use the `in` operator.
'puppies' in my_list
'kittens' in my_list
# So when to use a tuple and when to use a list?
#
# * Use a list when you will modify it after it is created?
#
# Ways to modify a list? You have already seen by index. Let's start with an empty list.
my_new_list = []
my_new_list
# We can add to the list using the append method on it.
my_new_list.append('Now')
my_new_list
# We can use the `+` operator to create a longer list by adding the contents of two lists together.
my_new_list + my_list
# One of the useful things to know about a list how many elements are in it. This can be found with the `len` function.
len(my_list)
# Some other handy functions with lists:
# * max
# * min
# * cmp
# Sometimes you have a tuple and you need to make it a list. You can `cast` the tuple to a list with ``list(my_tuple)``
list(my_tuple)
# What in the above told us it was a list?
#
# You can also use the ``type`` function to figure out the type.
type(tuple)
type(list(my_tuple))
# There are other useful methods on lists, including:
#
# | methods | description |
# |---|---|
# | list.append(obj) | Appends object obj to list |
# | list.count(obj)| Returns count of how many times obj occurs in list |
# | list.extend(seq) | Appends the contents of seq to list |
# | list.index(obj) | Returns the lowest index in list that obj appears |
# | list.insert(index, obj) | Inserts object obj into list at offset index |
# | list.pop(obj=list[-1]) | Removes and returns last object or obj from list |
# | list.remove(obj) | Removes object obj from list |
# | list.reverse() | Reverses objects of list in place |
# | list.sort([func]) | Sort objects of list, use compare func, if given |
#
# Try some of them now.
#
# ```
# my_list.count('I')
# my_list
#
# my_list.append('I')
# my_list
#
# my_list.count('I')
# my_list
#
# #my_list.index(42)
#
# my_list.index('puppies')
# my_list
#
# my_list.insert(my_list.index('puppies'), 'furry')
# my_list
# ```
# +
my_list.count('I')
my_list
my_list.append('I')
my_list
my_list.count('I')
my_list
#my_list.index(42)
my_list.index('puppies')
my_list
my_list.insert(my_list.index('puppies'), 'furry')
my_list
my_list.pop()
my_list
my_list.remove('puppies')
my_list
my_list.append('cabbages')
my_list
# -
# ### Any questions about lists before we move on?
# <hr>
# ### Dictionaries
#
# Dictionaries are similar to tuples and lists in that they hold a collection of objects. Dictionaries, however, allow an additional indexing mode: keys. Think of a real dictionary where the elements in it are the definitions of the words and the keys to retrieve the entries are the words themselves.
#
# | word | definition |
# |------|------------|
# | tuple | An immutable collection of ordered objects |
# | list | A mutable collection of ordered objects |
# | dictionary | A mutable collection of named objects |
#
# Let's create this data structure now. Dictionaries, like tuples and elements use a unique referencing method, '{' and its evil twin '}'.
my_dict = { 'tuple' : 'An immutable collection of ordered objects',
'list' : 'A mutable collection of ordered objects',
'dictionary' : 'A mutable collection of objects' }
my_dict
# We access items in the dictionary by name, e.g.
my_dict['dictionary']
# Since the dictionary is mutable, you can change the entries.
my_dict['dictionary'] = 'A mutable collection of named objects'
my_dict
# Notice that ordering is not preserved!
#
# And we can add new items to the list.
my_dict['cabbage'] = 'Green leafy plant in the Brassica family'
my_dict
# To delete an entry, we can't just set it to ``None``
my_dict['cabbage'] = None
my_dict
# To delete it propery, we need to pop that specific entry.
my_dict.pop('cabbage', None)
my_dict
# You can use other objects as names, but that is a topic for another time. You can mix and match key types, e.g.
my_new_dict = {}
my_new_dict[1] = 'One'
my_new_dict['42'] = 42
my_new_dict
# You can get a list of keys in the dictionary by using the ``keys`` method.
my_dict.keys()
# Similarly the contents of the dictionary with the ``items`` method.
my_dict.items()
# We can use the keys list for fun stuff, e.g. with the ``in`` operator.
'dictionary' in my_dict.keys()
# This is a synonym for `in my_dict`
'dictionary' in my_dict
# Notice, it doesn't work for elements.
'A mutable collection of ordered objects' in my_dict
# Other dictionary methods:
#
# | methods | description |
# |---|---|
# | dict.clear() | Removes all elements from dict |
# | dict.get(key, default=None) | For ``key`` key, returns value or ``default`` if key doesn't exist in dict |
# | dict.items() | Returns a list of dicts (key, value) tuple pairs |
# | dict.keys() | Returns a list of dictionary keys |
# | dict.setdefault(key, default=None) | Similar to get, but set the value of key if it doesn't exist in dict |
# | dict.update(dict2) | Add the key / value pairs in dict2 to dict |
# | dict.values | Returns a list of dictionary values|
#
# Feel free to experiment...
# <hr>
# ## Flow control
#
# <img src="https://docs.oracle.com/cd/B19306_01/appdev.102/b14261/lnpls008.gif">Flow control figure</img>
#
# Flow control refers how to programs do loops, conditional execution, and order of functional operations. Let's start with conditionals, or the venerable ``if`` statement.
#
# Let's start with a simple list of instructors for these classes.
instructors = ['Dave', 'Jim', '<NAME>']
instructors
# ### If
# If statements can be use to execute some lines or block of code if a particular condition is satisfied. E.g. Let's print something based on the entries in the list.
if 'Dorkus the Clown' in instructors:
print('#fakeinstructor')
# Usually we want conditional logic on both sides of a binary condition, e.g. some action when ``True`` and some when ``False``
if 'Dorkus the Clown' in instructors:
print('There are fake names for class instructors in your list!')
else:
print("Nothing to see here")
# There is a special do nothing word: `pass` that skips over some arm of a conditional, e.g.
if 'Jim' in instructors:
print("Congratulations! Jim is teaching, your class won't stink!")
else:
pass
# _Note_: what have you noticed in this session about quotes? What is the difference between ``'`` and ``"``?
#
#
# Another simple example:
if True is False:
print("I'm so confused")
else:
print("Everything is right with the world")
# It is always good practice to handle all cases explicity. `Conditional fall through` is a common source of bugs.
#
# Sometimes we wish to test multiple conditions. Use `if`, `elif`, and `else`.
# +
my_favorite = 'pie'
if my_favorite is 'cake':
print("He likes cake! I'll start making a double chocolate velvet cake right now!")
elif my_favorite is 'pie':
print("He likes pie! I'll start making a cherry pie right now!")
else:
print("He likes " + my_favorite + ". I don't know how to make that.")
# -
# Conditionals can take ``and`` and ``or`` and ``not``. E.g.
# +
my_favorite = 'pie'
if my_favorite is 'cake' or my_favorite is 'pie':
print(my_favorite + " : I have a recipe for that!")
else:
print("Ew! Who eats that?")
# -
# ## For
#
# For loops are the standard loop, though `while` is also common. For has the general form:
# ```
# for items in list:
# do stuff
# ```
#
# For loops and collections like tuples, lists and dictionaries are natural friends.
for instructor in instructors:
print(instructor)
# You can combine loops and conditionals:
for instructor in instructors:
if instructor.endswith('Clown'):
print(instructor + " doesn't sound like a real instructor name!")
else:
print(instructor + " is so smart... all those gooey brains!")
# Dictionaries can use the `keys` method for iterating.
for key in my_dict.keys():
if len(key) > 5:
print(my_dict[key])
# ### range()
#
# Since for operates over lists, it is common to want to do something like:
# ```
# NOTE: C-like
# for (i = 0; i < 3; ++i) {
# print(i);
# }
# ```
#
# The Python equivalent is:
#
# ```
# for i in [0, 1, 2]:
# do something with i
# ```
#
# What happens when the range you want to sample is big, e.g.
# ```
# NOTE: C-like
# for (i = 0; i < 1000000000; ++i) {
# print(i);
# }
# ```
#
# That would be a real pain in the rear to have to write out the entire list from 1 to 1000000000.
#
# Enter, the `range()` function. E.g.
# ```range(3) is [0, 1, 2]```
range(3)
# Notice that Python (in the newest versions, e.g. 3+) has an object type that is a range. This saves memory and speeds up calculations vs. an explicit representation of a range as a list - but it can be automagically converted to a list on the fly by Python. To show the contents as a `list` we can use the type case like with the tuple above.
#
# Sometimes, in older Python docs, you will see `xrange`. This used the range object back in Python 2 and `range` returned an actual list. Beware of this!
list(range(3))
# Remember earlier with slicing, the syntax `:3` meant `[0, 1, 2]`? Well, the same upper bound philosophy applies here.
#
for index in range(3):
instructor = instructors[index]
if instructor.endswith('Clown'):
print(instructor + " doesn't sound like a real instructor name!")
else:
print(instructor + " is so smart... all those gooey brains!")
# This would probably be better written as
for index in range(len(instructors)):
instructor = instructors[index]
if instructor.endswith('Clown'):
print(instructor + " doesn't sound like a real instructor name!")
else:
print(instructor + " is so smart... all those gooey brains!")
# But in all, it isn't very Pythonesque to use indexes like that (unless you have another reason in the loop) and you would opt instead for the `instructor in instructors` form.
#
# More often, you are doing something with the numbers that requires them to be integers, e.g. math.
sum = 0
for i in range(10):
sum += i
print(sum)
# #### For loops can be nested
#
# _Note_: for more on formatting strings, see: [https://pyformat.info](https://pyformat.info)
for i in range(1, 4):
for j in range(1, 4):
print('%d * %d = %d' % (i, j, i*j)) # Note string formatting here, %d means an integer
# #### You can exit loops early if a condition is met:
for i in range(10):
if i == 4:
break
i
# #### You can skip stuff in a loop with `continue`
sum = 0
for i in range(10):
if (i == 5):
continue
else:
sum += i
print(sum)
# #### There is a unique language feature call ``for...else``
sum = 0
for i in range(10):
sum += i
else:
print('final i = %d, and sum = %d' % (i, sum))
# #### You can iterate over letters in a string
my_string = "DIRECT"
for c in my_string:
print(c)
# <hr>
# ## Hacky Hack Time with Ifs, Fors, Lists, and imports!
#
# Objective: Replace the `bash magic` bits for downloading the HCEPDB data and uncompressing it with Python code. Since the download is big, check if the zip file exists first before downloading it again. Then load it into a pandas dataframe.
#
# Notes:
# * The `os` package has tools for checking if a file exists: ``os.path.exists``
# ```
# import os
# filename = 'HCEPDB_moldata.zip'
# if os.path.exists(filename):
# print("wahoo!")
# ```
# * Use the `requests` package to get the file given a url (got this from the requests docs)
# ```
# import requests
# url = 'http://faculty.washington.edu/dacb/HCEPDB_moldata.zip'
# req = requests.get(url)
# assert req.status_code == 200 # if the download failed, this line will generate an error
# with open(filename, 'wb') as f:
# f.write(req.content)
# ```
# * Use the `zipfile` package to decompress the file while reading it into `pandas`
# ```
# import pandas as pd
# import zipfile
# csv_filename = 'HCEPDB_moldata.csv'
# zf = zipfile.ZipFile(filename)
# data = pd.read_csv(zf.open(csv_filename))
# ```
#
# ### Now, use your code from above for the following URLs and filenames
#
# | URL | filename | csv_filename |
# |-----|----------|--------------|
# | http://faculty.washington.edu/dacb/HCEPDB_moldata_set1.zip | HCEPDB_moldata_set1.zip | HCEPDB_moldata_set1.csv |
# | http://faculty.washington.edu/dacb/HCEPDB_moldata_set2.zip | HCEPDB_moldata_set2.zip | HCEPDB_moldata_set2.csv |
# | http://faculty.washington.edu/dacb/HCEPDB_moldata_set3.zip | HCEPDB_moldata_set3.zip | HCEPDB_moldata_set3.csv |
#
# What pieces of the data structures and flow control that we talked about earlier can you use?
# How did you solve this problem?
#
# <hr>
# ### Functions
#
# For loops let you repeat some code for every item in a list. Functions are similar in that they run the same lines of code for new values of some variable. They are different in that functions are not limited to looping over items.
#
# Functions are a critical part of writing easy to read, reusable code.
#
# Create a function like:
# ```
# def function_name (parameters):
# """
# optional docstring
# """
# function expressions
# return [variable]
# ```
#
# _Note:_ Sometimes I use the word argument in place of parameter.
#
# Here is a simple example. It prints a string that was passed in and returns nothing.
def print_string(str):
"""This prints out a string passed as the parameter."""
print(str)
return
# To call the function, use:
# ```
# print_string("Dave is awesome!")
# ```
#
# _Note:_ The function has to be defined before you can call it!
print_string("Dave is awesome!")
# If you don't provide an argument or too many, you get an error.
# Parameters (or arguments) in Python are all passed by reference. This means that if you modify the parameters in the function, they are modified outside of the function.
#
# See the following example:
#
# ```
# def change_list(my_list):
# """This changes a passed list into this function"""
# my_list.append('four');
# print('list inside the function: ', my_list)
# return
#
# my_list = [1, 2, 3];
# print('list before the function: ', my_list)
# change_list(my_list);
# print('list after the function: ', my_list)
# ```
# +
def change_list(my_list):
"""This changes a passed list into this function"""
my_list.append('four');
print('list inside the function: ', my_list)
return
my_list = [1, 2, 3];
print('list before the function: ', my_list)
change_list(my_list);
print('list after the function: ', my_list)
# -
# Variables have scope: `global` and `local`
#
# In a function, new variables that you create are not saved when the function returns - these are `local` variables. Variables defined outside of the function can be accessed but not changed - these are `global` variables, _Note_ there is a way to do this with the `global` keyword. Generally, the use of `global` variables is not encouraged, instead use parameters.
#
# ```
# my_global_1 = 'bad idea'
# my_global_2 = 'another bad one'
# my_global_3 = 'better idea'
#
# def my_function():
# print(my_global)
# my_global_2 = 'broke your global, man!'
# global my_global_3
# my_global_3 = 'still a better idea'
# return
#
# my_function()
# print(my_global_2)
# print(my_global_3)
# ```
# In general, you want to use parameters to provide data to a function and return a result with the `return`. E.g.
#
# ```
# def sum(x, y):
# my_sum = x + y
# return my_sum
# ```
#
# If you are going to return multiple objects, what data structure that we talked about can be used? Give and example below.
# ### Parameters have four different types:
#
# | type | behavior |
# |------|----------|
# | required | positional, must be present or error, e.g. `my_func(first_name, last_name)` |
# | keyword | position independent, e.g. `my_func(first_name, last_name)` can be called `my_func(first_name='Dave', last_name='Beck')` or `my_func(last_name='Beck', first_name='Dave')` |
# | default | keyword params that default to a value if not provided |
#
def print_name(first, last='<NAME>'):
print('Your name is %s %s' % (first, last))
return
# Play around with the above function.
# Functions can contain any code that you put anywhere else including:
# * if...elif...else
# * for...else
# * while
# * other function calls
def print_name_age(first, last, age):
print_name(first, last)
print('Your age is %d' % (age))
if age > 35:
print('You are really old.')
return
print_name_age(age=40, last='Beck', first='Dave')
# #### How would you functionalize the above code for downloading, unzipping, and making a dataframe?
# Once you have some code that is functionalized and not going to change, you can move it to a file that ends in `.py`, check it into version control, import it into your notebook and use it!
#
#
# Homework:
# Save your functions to `hcepdb_utils.py`. Import the functions and use them to rewrite HW1. This will be laid out in the homework repo for HW2. Check the website.
| L5.Procedural_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from BurstCube.simGenerator import configurator
conf = configurator('config.yaml')
from BurstCube.bcSim import simFiles
sfs = simFiles('config.yaml')
print(sfs.sims[0].simFile)
type(sfs.sims[0].simFile)# fileN = 'FIN_1000.000keV_15.00ze_0.00az.inc1.id1.sim'
simulation = 0; detectOne = 0; detectTwo = 0; detectThree = 0; detectFour = 0
percentOne = 0; percentTwo = 0; percentThree = 0; percentFour = 0
# +
for file in sfs.sims:
simulation = simulation + 1
# print(file.simFile)
import re
counterXX = 0
counterNXX = 0
counterNXY = 0
counterXY = 0
# count = 0
prevLine = 'random'
# counter = 0
with open(file.simFile) as sim:
for i, line in enumerate(sim):
if line.startswith('ID'):
ID = line.split()
for i in range(len(ID)):
trigger = ID[1]
if line.startswith('HTsim'):
data = line.split(";")
data = [w.strip(' ') for w in data]
# counter = counter + 1
for i in range(len(data)):
if data[i] == '5.52500' and data[i+1] == '5.52500':
counterXX += 1
elif data[i] == '5.52500' and data[i+1] == '-5.52500':
counterXY += 1
elif data[i] == '-5.52500' and data[i+1] == '5.52500':
counterNXX += 1
# print(counterNXX)
elif data[i] == '-5.52500' and data[i+1] == '-5.52500':
counterNXY += 1
# print('****', counterNXY)
if prevLine.startswith('HTsim') and line.startswith('HTsim'):
print('**For trigger', trigger, 'there are 2 hits!!**')
prevLine = line
# print('***', trigger)
# vars = re.split('[_ .]', file.simFile)
# print("----------------------------------------------------------------------------")
# # print('Simulation', simulation, '--', trigger, ': Energy ({} keV), Azimuth ({} degrees), Zenith ({} degrees)'.format(vars[2], vars[4], vars[6]))
# print("----------------------------------------------------------------------------")
# print('Detector 1 has', counterXX , 'hits.')
# print('Detector 2 has', counterNXX , 'hits.')
# print('Detector 3 has', counterNXY , 'hits.')
# print('Detector 4 has', counterXY , 'hits.')
detectOne = counterXX + detectOne
detectTwo = counterNXX + detectTwo
detectThree = counterNXY + detectThree
detectFour = counterXY + detectFour
# +
total = detectOne + detectTwo + detectThree + detectFour
percentOne = (detectOne // total) * 100
percentTwo = (detectTwo // total) * 100
percentThree = (detectThree // total) * 100
percentFour = (detectFour // total) * 100
# -
| lepalmer/localCodes/local_test/1000_15_c/Localization Code Azimuth Changing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="_Pkmfqg8zWco" colab_type="code" colab={}
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# import data from the github page of the book
data = pd.read_csv('https://raw.githubusercontent.com/PacktWorkshops/The-Data-Analysis-Workshop/master/Chapter02/data/Absenteeism_at_work.csv', sep=";")
# + id="iSLL4gk5zWct" colab_type="code" outputId="f0d50c8b-ba9e-4aa1-ea10-21e869f7aa9c" colab={"base_uri": "https://localhost:8080/", "height": 391}
# print dimensionality of the data, columns, types and missing values
print(f"Data dimension: {data.shape}")
for col in data.columns:
print(f"Column: {col:35} | type: {str(data[col].dtype):7} | missing values: {data[col].isna().sum():3d}")
# + id="10xzWXruzWcx" colab_type="code" outputId="cc22ea2d-0cda-4793-e7d5-067455db58d3" colab={"base_uri": "https://localhost:8080/", "height": 700}
# compute statistics on numerical features
data.describe().T
# + id="KM11JNkzzWc0" colab_type="code" colab={}
# define encoding dictionaries
month_encoding = {1: "January", 2: "February", 3: "March", 4: "April",
5: "May", 6: "June", 7: "July", 8: "August",
9: "September", 10: "October", 11: "November", 12: "December", 0: "Unknown"}
dow_encoding = {2: "Monday", 3: "Tuesday", 4: "Wednesday", 5: "Thursday", 6: "Friday"}
season_encoding = {1: "Spring", 2: "Summer", 3: "Fall", 4: "Winter"}
education_encoding = {1: "high_school", 2: "graduate", 3: "postgraduate", 4: "master_phd"}
yes_no_encoding = {0: "No", 1: "Yes"}
# backtransform numerical variables to categorical
preprocessed_data = data.copy()
preprocessed_data["Month of absence"] = preprocessed_data["Month of absence"]\
.apply(lambda x: month_encoding[x])
preprocessed_data["Day of the week"] = preprocessed_data["Day of the week"]\
.apply(lambda x: dow_encoding[x])
preprocessed_data["Seasons"] = preprocessed_data["Seasons"]\
.apply(lambda x: season_encoding[x])
preprocessed_data["Education"] = preprocessed_data["Education"]\
.apply(lambda x: education_encoding[x])
preprocessed_data["Disciplinary failure"] = preprocessed_data["Disciplinary failure"]\
.apply(lambda x: yes_no_encoding[x])
preprocessed_data["Social drinker"] = preprocessed_data["Social drinker"]\
.apply(lambda x: yes_no_encoding[x])
preprocessed_data["Social smoker"] = preprocessed_data["Social smoker"]\
.apply(lambda x: yes_no_encoding[x])
# + id="E7G0sSqtzWc3" colab_type="code" outputId="99286e6c-6570-4fb1-a9d4-45f747b27dd2" colab={"base_uri": "https://localhost:8080/", "height": 700}
# transform columns
preprocessed_data.head().T
# + id="bcSvhsJNzWc8" colab_type="code" outputId="936a58df-3570-4ba1-8ca7-6fda96b34006" colab={"base_uri": "https://localhost:8080/", "height": 499}
# define function, which checks if the provided integer value
# is contained in the ICD or not
def in_icd(val):
return "Yes" if val >= 1 and val <= 21 else "No"
# add Disease column
preprocessed_data["Disease"] = preprocessed_data["Reason for absence"]\
.apply(in_icd)
# plot value counts
plt.figure(figsize=(10, 8))
sns.countplot(data=preprocessed_data, x='Disease')
plt.savefig('figs/disease_plot.png', format='png', dpi=300)
# + [markdown] id="WIN2F996zWc_" colab_type="text"
# # Initial analysis on the reason for absence
# + id="6sZCKBV2zWdB" colab_type="code" outputId="6e2062ae-4c51-45e3-a423-6b63699b47ab" colab={"base_uri": "https://localhost:8080/", "height": 334}
# get the number of entries for each reason for absence
plt.figure(figsize=(10, 5))
ax = sns.countplot(data=preprocessed_data, x="Reason for absence")
ax.set_ylabel("Number of entries per reason of absence")
plt.savefig('figs/absence_reasons_distribution.png', format='png', dpi=300)
# + [markdown] id="K1BuNyIvzWdE" colab_type="text"
# #### Social drinkers and smokers analysis
# + id="3zoXqQdbzWdF" colab_type="code" outputId="3e5f0ff5-41f6-4f47-9a7e-1d1dc5b5c5c5" colab={"base_uri": "https://localhost:8080/", "height": 759}
# plot reasons for absence against being a social drinker/smoker
plt.figure(figsize=(8, 6))
sns.countplot(data=preprocessed_data, x="Reason for absence",
hue="Social drinker", hue_order=["Yes", "No"])
plt.savefig('figs/absence_reasons_drinkers.png', format='png', dpi=300)
plt.figure(figsize=(8, 6))
sns.countplot(data=preprocessed_data, x="Reason for absence",
hue="Social smoker", hue_order=["Yes", "No"])
plt.savefig('figs/absence_reasons_smokers.png', format='png', dpi=300)
# + id="ohk2cUInzWdH" colab_type="code" outputId="9405b4fb-6522-45fe-dcb5-d93dceef377f" colab={"base_uri": "https://localhost:8080/", "height": 119}
print(preprocessed_data["Social drinker"].value_counts(normalize=True))
print(preprocessed_data["Social smoker"].value_counts(normalize=True))
# + id="FNKxOsvyzWdL" colab_type="code" outputId="96a8b827-1dec-47e1-e8ae-f376efd7348e" colab={"base_uri": "https://localhost:8080/", "height": 34}
# computation of conditional probability
sample_space = set(["BB", "BG", "GB", "GG"])
event_a = set(["BB"])
event_b = set(["BB", "BG", "GB"])
cond_prob = (0.25*len(event_a.intersection(event_b))) / (0.25*len(event_b))
print(round(cond_prob, 4))
# + id="1wZsM3KNzWdP" colab_type="code" outputId="6b76d43f-ee02-428a-fef0-c4fc8287a410" colab={"base_uri": "https://localhost:8080/", "height": 411}
# compute probabilities of being a drinker and smoker
drinker_prob = preprocessed_data["Social drinker"]\
.value_counts(normalize=True)["Yes"]
smoker_prob = preprocessed_data["Social smoker"]\
.value_counts(normalize=True)["Yes"]
print(f"P(social drinker) = {drinker_prob:.3f} | P(social smoker) = {smoker_prob:.3f}")
# create mask for social drinkers/smokers
drinker_mask = preprocessed_data["Social drinker"] == "Yes"
smoker_mask = preprocessed_data["Social smoker"] == "Yes"
# compute probabilities of absence reasons and being a social drinker/smoker
total_entries = preprocessed_data.shape[0]
absence_drinker_prob = preprocessed_data["Reason for absence"]\
[drinker_mask].value_counts()/total_entries
absence_smoker_prob = preprocessed_data["Reason for absence"]\
[smoker_mask].value_counts()/total_entries
# compute conditional probabilities
cond_prob = pd.DataFrame(index=range(0,29))
cond_prob["P(Absence | social drinker)"] = absence_drinker_prob/drinker_prob
cond_prob["P(Absence | social smoker)"] = absence_smoker_prob/smoker_prob
# plot probabilities
plt.figure()
ax = cond_prob.plot.bar(figsize=(10,6))
ax.set_ylabel("Conditional probability")
plt.savefig('figs/conditional_probabilities.png', format='png', dpi=300)
# + id="ATVXzS-4zWdT" colab_type="code" colab={}
# compute reason for absence probabilities
absence_prob = preprocessed_data["Reason for absence"].value_counts(normalize=True)
# + id="yu3cg0hQzWdV" colab_type="code" outputId="35a499fc-ee13-45cb-c0e1-3bccae3543aa" colab={"base_uri": "https://localhost:8080/", "height": 394}
# compute conditional probabilities for drinker/smoker
cond_prob_drinker_smoker = pd.DataFrame(index=range(0,29))
cond_prob_drinker_smoker["P(social drinker | Absence)"] = \
cond_prob["P(Absence | social drinker)"]*drinker_prob/absence_prob
cond_prob_drinker_smoker["P(social smoker | Absence)"] = \
cond_prob["P(Absence | social smoker)"]*smoker_prob/absence_prob
plt.figure()
ax = cond_prob_drinker_smoker.plot.bar(figsize=(10,6))
ax.set_ylabel("Conditional probability")
plt.savefig('figs/conditional_probabilities_drinker_smoker.png', format='png', dpi=300)
# + id="dWE-efohzWdY" colab_type="code" outputId="b5e1bf59-3fe8-4030-edb5-3f20ebe77af2" colab={"base_uri": "https://localhost:8080/", "height": 759}
# create violin plots of the absenteeism time in hours
plt.figure(figsize=(8,6))
sns.violinplot(x="Social drinker", y="Absenteeism time in hours", \
data=preprocessed_data, order=["No", "Yes"])
plt.savefig('figs/drinkers_hour_distribution.png', format='png', dpi=300)
plt.figure(figsize=(8,6))
sns.violinplot(x="Social smoker", y="Absenteeism time in hours", \
data=preprocessed_data, order=["No", "Yes"])
plt.savefig('figs/smokers_hour_distribution.png', format='png', dpi=300)
# + id="-6p-u2ZOzWdb" colab_type="code" outputId="873db75c-0fc4-4f47-f679-ad4404676f1f" colab={"base_uri": "https://localhost:8080/", "height": 51}
from scipy.stats import ttest_ind
hours_col = "Absenteeism time in hours"
# test mean absenteeism time for drinkers
drinkers_mask = preprocessed_data["Social drinker"] == "Yes"
hours_drinkers = preprocessed_data.loc[drinker_mask, hours_col]
hours_non_drinkers = preprocessed_data.loc[~drinker_mask, hours_col]
drinkers_test = ttest_ind(hours_drinkers, hours_non_drinkers)
print(f"Statistic value: {drinkers_test[0]}, p-value: {drinkers_test[1]}")
# test mean absenteeism time for smokers
smokers_mask = preprocessed_data["Social smoker"] == "Yes"
hours_smokers = preprocessed_data.loc[smokers_mask, hours_col]
hours_non_smokers = preprocessed_data.loc[~smokers_mask, hours_col]
smokers_test = ttest_ind(hours_smokers, hours_non_smokers)
print(f"Statistic value: {smokers_test[0]}, p-value: {smokers_test[1]}")
# + id="cN_vcYW0zWde" colab_type="code" outputId="8890d22b-93ef-4a4a-d37f-34735c8a259c" colab={"base_uri": "https://localhost:8080/", "height": 51}
# perform Kolmogorov-Smirnov test for comparing the distributions
from scipy.stats import ks_2samp
ks_drinkers = ks_2samp(hours_drinkers, hours_non_drinkers)
ks_smokers = ks_2samp(hours_smokers, hours_non_smokers)
print(f"Drinkers comparison: statistics={ks_drinkers[0]:.3f}, pvalue={ks_drinkers[1]:.3f}")
print(f"Smokers comparison: statistics={ks_smokers[0]:.3f}, pvalue={ks_smokers[1]:.3f}")
# + [markdown] id="9gjalGkdzWdh" colab_type="text"
# ### Body Mass Index
# + id="mwk6L2dOzWdi" colab_type="code" colab={}
# define function for computing the BMI category, based on BMI value
def get_bmi_category(bmi):
if bmi < 18.5:
category = "underweight"
elif bmi >= 18.5 and bmi < 25:
category = "healthy weight"
elif bmi >= 25 and bmi < 30:
category = "overweight"
elif bmi >= 30:
category = "obese"
return category
# compute BMI category
preprocessed_data["BMI category"] = preprocessed_data["Body mass index"]\
.apply(get_bmi_category)
# + id="nTfW7N_GzWdk" colab_type="code" outputId="ddd30b08-a741-43de-8444-dd1312fad9ce" colab={"base_uri": "https://localhost:8080/", "height": 334}
#### plot number of entries for each category
plt.figure(figsize=(10, 5))
sns.countplot(data=preprocessed_data, x='BMI category',
order=["underweight", "healthy weight", "overweight", "obese"],
palette="Set2")
plt.savefig('figs/bmi_categories.png', format='png', dpi=300)
# + id="byz6oUZXzWdm" colab_type="code" outputId="3045e400-8244-4c2b-844c-3efcab176d3f" colab={"base_uri": "https://localhost:8080/", "height": 932}
# plot BMI categories vs Reason for absence
plt.figure(figsize=(10, 16))
ax = sns.countplot(data=preprocessed_data, y="Reason for absence", hue="BMI category",
hue_order=["underweight", "healthy weight", "overweight", "obese"],
palette="Set2")
ax.set_xlabel("Number of employees")
plt.savefig('figs/reasons_bmi.png', format='png', dpi=300)
# + id="pKVvjIcGzWdq" colab_type="code" outputId="b500ffb2-4ef7-43b5-db27-f088dc5fb983" colab={"base_uri": "https://localhost:8080/", "height": 388}
# plot distribution of absence time, based on BMI category
plt.figure(figsize=(8,6))
sns.violinplot(x="BMI category", y="Absenteeism time in hours", \
data=preprocessed_data, order=["healthy weight", "overweight", "obese"])
plt.savefig('figs/bmi_hour_distribution.png', format='png')
# + [markdown] id="JPkpVxvUzWdt" colab_type="text"
# ### Age and education factors
# + id="wpD-i-aazWdu" colab_type="code" outputId="c16e422a-7adb-4de3-e36e-fc6997b6c14c" colab={"base_uri": "https://localhost:8080/", "height": 404}
from scipy.stats import pearsonr
# compute Pearson's correlation coefficient and p-value
pearson_test = pearsonr(preprocessed_data["Age"], \
preprocessed_data["Absenteeism time in hours"])
# create regression plot and add correlation coefficient in the title
plt.figure(figsize=(10, 6))
ax = sns.regplot(x="Age", y="Absenteeism time in hours",
data=preprocessed_data, scatter_kws={"alpha":0.5})
ax.set_title(f"Correlation={pearson_test[0]:.03f} | p-value={pearson_test[1]:.03f}");
plt.savefig('figs/correlation_age_hours.png', format='png', dpi=300)
# + [markdown] id="CvfG9lwOzWd2" colab_type="text"
# ### Exercise 2.04: Investigate Age impact on Reason for absence
# + id="qrF-ftBezWd3" colab_type="code" outputId="8f1e7eb2-f805-4d60-c850-bf0f9c64a681" colab={"base_uri": "https://localhost:8080/", "height": 388}
# create violin plot between the Age and Disease columns
plt.figure(figsize=(8,6))
sns.violinplot(x="Disease", y="Age", data=preprocessed_data)
plt.savefig('figs/exercise_204_age_disease.png', format='png', dpi=300)
# + id="s2Hz5wZ6zWd6" colab_type="code" outputId="f4697187-3e1c-48e8-d613-bf690f092840" colab={"base_uri": "https://localhost:8080/", "height": 51}
# get Age entries for employees with Disease == Yes and Disease == No
disease_mask = preprocessed_data["Disease"] == "Yes"
disease_ages = preprocessed_data["Age"][disease_mask]
no_disease_ages = preprocessed_data["Age"][~disease_mask]
# perform hypothesis test for equality of means
test_res = ttest_ind(disease_ages, no_disease_ages)
print(f"Test for equality of means: statistic={test_res[0]:0.3f}, pvalue={test_res[1]:0.3f}")
# test equality of distributions via Kolmogorov-Smirnov test
ks_res = ks_2samp(disease_ages, no_disease_ages)
print(f"KS test for equality of distributions: statistic={ks_res[0]:0.3f}, pvalue={ks_res[1]:0.3f}")
# + id="5Sq9EKXlzWd9" colab_type="code" outputId="ed11922c-de5b-49ab-e55e-5397d5111d52" colab={"base_uri": "https://localhost:8080/", "height": 497}
# violin plot of reason for absence vs age
plt.figure(figsize=(20,8))
sns.violinplot(x="Reason for absence", y="Age", data=preprocessed_data)
plt.savefig('figs/exercise_204_age_reason.png', format='png')
| Chapter02/Exercise2.04/Exercise2.04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (dlenv)
# language: python
# name: dlenv
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/shangeth/Google-ML-Academy/blob/master/1-Intro-to-Deep-Learning/1_2_1_Assignment_Polynomial_Regression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="MTd7W7TxCV7q"
# <hr>
# <h1 align="center"><a href='https://shangeth.com/courses/'>Deep Learning - Beginners Track</a></h1>
# <h3 align="center">Instructor: <a href='https://shangeth.com/'><NAME></a></h3>
# <hr>
# + [markdown] colab_type="text" id="es9QfSb6J8za"
# # Task-1 : Linear Regression on Non-Linear Data
#
# - Get X and y from dataset() function
# - Train a Linear Regression model for this dataset.
# - Visualize the model prediction
# + [markdown] colab_type="text" id="z-IlItpAKWBl"
# ## Dataset
#
# Call ```dataset()``` function to get X, y
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="_Qjqkxe7KXkh" outputId="e3648df2-f581-4cfe-fd9b-e9c6d257d088"
import numpy as np
import matplotlib.pyplot as plt
def dataset(show=True):
X = np.arange(-25, 25, 0.1)
# Try changing y to a different function
y = X**3 + 20 + np.random.randn(500)*1000
if show:
plt.scatter(X, y)
plt.show()
return X, y
X, y = dataset()
# + [markdown] colab_type="text" id="tKRpy65MNUlw"
# ## Scaling Dataset
#
# The maximum value of y in the dataset goes upto 15000 and the minimum values is less than -15000. The range of y is very large which makes the convergence/loss reduction slower. So will we scale the data, scaling the data will help the model converge faster. If all the features and target are in same range, there will be symmetry in the curve of Loss vs weights/bias, which makes the convergence faster.
#
# We will do a very simple type of scaling, we will divide all the values of the data with the maximum values for X and y respectively.
# + colab={} colab_type="code" id="QzPpSAUaNTpi"
X, y = dataset()
print(max(X), max(y), min(X), min(y))
# TODO: divide array X and y by its max values
X = #
y = #
print(max(X), max(y), min(X), min(y))
# + [markdown] colab_type="text" id="BqvlVbQDOoSd"
# This is not a great scaling method, but good to start. We will see many more scaling/normalizing methods later.
#
# **Try training the model with and without scaling and see the difference yourself.**
# + [markdown] colab_type="text" id="jpzO5Eb207_i"
# ## Linear Regression in TensorFlow
#
# If you are not sure about TF model definition check the [documentation](https://www.tensorflow.org/api_docs/python/tf/keras/Model)
#
# Refer the previous notebook, code is almost the same or you can even check the next notebook(solutions).
# + colab={} colab_type="code" id="ioav6Ec6rpbJ"
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
X, y = dataset(show=False)
X_scaled = # Scaled X value
y_scaled = # Scaled y value
model = # TF Dense Model
# check how to define a optimizer here https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
optimizer =
# train the model
# compile the model
tf_history = # fit the model
plt.plot(tf_history.history['loss'])
plt.xlabel('Epochs')
plt.ylabel('MSE Loss')
plt.show()
mse = tf_history.history['loss'][-1]
y_hat = model.predict(X_scaled)
plt.figure(figsize=(12,7))
plt.title('TensorFlow Model')
plt.scatter(X_scaled, y_scaled, label='Data $(X, y)$')
plt.plot(X_scaled, y_hat, color='red', label='Predicted Line $y = f(X)$',linewidth=4.0)
plt.xlabel('$X$', fontsize=20)
plt.ylabel('$y$', fontsize=20)
plt.text(0,0.70,'MSE = {:.3f}'.format(mse), fontsize=20)
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
# + [markdown] colab_type="text" id="M_yyuTj-RFin"
# # Polynomial Regression
#
# So when the dataset is not linear, linear regression cannot learn the dataset and make good predictions.
#
# So we need a polynomial model which consideres the polynomial terms as well. So we need terms like $x^2$, $x^3$, ..., $x^n$ for the model to learn a polynomial of $n^{th}$ degree.
#
# $\hat{y} = w_0 + w_1x + w_2x^2 + ... + w_nx^n$
#
# One down side of this model is that, We will have to decide the value of n. But this is better than a linear regression model. We can get an idea of the value of n by visualizing a dataset, but for multi variable dataset, we will have to try different values of n and check which is better.
# + [markdown] colab_type="text" id="x8u4rPzJTCJJ"
# ## Polynomial Features
#
# you can calculate the polynomial features for each feature by programming it or you can try ```sklearn.preprocessing.PolynomialFeatures``` which allows us to make polynomial terms of our data.
#
# We will try degree 2, 3 and 4
# + colab={} colab_type="code" id="oZbYgRFcT9q6"
X, y = dataset(show=False)
X_scaled = X/max(X)
y_scaled = y/max(y)
# + colab={} colab_type="code" id="gDHF3FLBRBTK"
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(degree=2)
X_2 = poly.fit_transform(X_scaled.reshape(-1,1))
print(X_2.shape)
print(X_2[0])
# + colab={} colab_type="code" id="Tp0jMLxPTxs2"
# TODO
# get the 3rd degree terms
poly = # poly features
X_3 = # 3rd degree terms
print(X_3.shape)
print(X_3[0])
# + colab={} colab_type="code" id="j0V5pFUWT4kg"
# TODO
# get the 4th degree terms
poly = # poly features
X_4 = # 4th degree terms
print(X_4.shape)
print(X_4[0])
# + [markdown] colab_type="text" id="q7rzpVB3VLN7"
# The PolynomialFeatures returns $[1, x, x^2, x^3,...]$.
# + [markdown] colab_type="text" id="mcdv2RDhUF7C"
# # Task - 2
#
# - Train a model with polynomial terms in the dataset.
# - Visualize the prediction of the model
#
#
# The code remains the same except, the no of input features will be 3, 4, 5 respectively.
# + [markdown] colab_type="text" id="WCVRwEynVxYL"
# ## Tensorflow Model with 2nd Degree
# + colab={} colab_type="code" id="lJXuJi6UUBZO"
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
model = # TF Dense Model
# check how to define a optimizer here https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
optimizer =
# train the model
# compile the model
tf_history = # fit the model
plt.plot(tf_history.history['loss'])
plt.xlabel('Epochs')
plt.ylabel('MSE Loss')
plt.show()
mse = tf_history.history['loss'][-1]
y_hat = model.predict(X_2)
plt.figure(figsize=(12,7))
plt.title('TensorFlow Model')
plt.scatter(X_2[:, 1], y_scaled, label='Data $(X, y)$')
plt.plot(X_2[:, 1], y_hat, color='red', label='Predicted Line $y = f(X)$',linewidth=4.0)
plt.xlabel('$X$', fontsize=20)
plt.ylabel('$y$', fontsize=20)
plt.text(0,0.70,'MSE = {:.3f}'.format(mse), fontsize=20)
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
# + [markdown] colab_type="text" id="ovvrgy0_fcRM"
# Why is the polynomial regression with 2-d features look like a straight line?
#
# Well because the model thinks that a straight line(look like, we can't be sure if its a straight like, it can a parabola as well) better fits the dataset than a parabola. If you train the model for less epochs you can notice the model tries to fit the data with a parabola(2-d) but it improves as it moves to a line.
#
# Train the same model for may be 50 epochs.
# + colab={} colab_type="code" id="wlJYRqnMVjCu"
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
model = # TF Dense Model
# check how to define a optimizer here https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
optimizer =
# train the model for 50 epochs
# compile the model
tf_history = # fit the model
plt.plot(tf_history.history['loss'])
plt.xlabel('Epochs')
plt.ylabel('MSE Loss')
plt.show()
mse = tf_history.history['loss'][-1]
y_hat = model.predict(X_2)
plt.figure(figsize=(12,7))
plt.title('TensorFlow Model')
plt.scatter(X_2[:, 1], y_scaled, label='Data $(X, y)$')
plt.plot(X_2[:, 1], y_hat, color='red', label='Predicted Line $y = f(X)$',linewidth=4.0)
plt.xlabel('$X$', fontsize=20)
plt.ylabel('$y$', fontsize=20)
plt.text(0,0.70,'MSE = {:.3f}'.format(mse), fontsize=20)
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
# + [markdown] colab_type="text" id="PN99ty_GER0Z"
# You can clearly see that the model tries to fit the data with a parabola which doen't seem to fit well, so it changes the parameters to fit a line.
# + [markdown] colab_type="text" id="gV2fxw9rV3aA"
# ## Tensorflow Model with 3rd Degree
# + colab={} colab_type="code" id="xVY5HcC7V3aO"
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
model = # TF Dense Model
# check how to define a optimizer here https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
optimizer =
# train the model
# compile the model
tf_history = # fit the model
plt.plot(tf_history.history['loss'])
plt.xlabel('Epochs')
plt.ylabel('MSE Loss')
plt.show()
mse = tf_history.history['loss'][-1]
y_hat = model.predict(X_3)
plt.figure(figsize=(12,7))
plt.title('TensorFlow Model')
plt.scatter(X_3[:, 1], y_scaled, label='Data $(X, y)$')
plt.plot(X_3[:, 1], y_hat, color='red', label='Predicted Line $y = f(X)$',linewidth=4.0)
plt.xlabel('$X$', fontsize=20)
plt.ylabel('$y$', fontsize=20)
plt.text(0,0.70,'MSE = {:.3f}'.format(mse), fontsize=20)
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
# + [markdown] colab_type="text" id="djQgZARjgYfI"
# 3-D features perfectly fit the data with a 3rd degree polynomial as expected.
# + [markdown] colab_type="text" id="zppsoHcaV4qK"
# # Tensorflow Model with 4th Degree
# + colab={} colab_type="code" id="oYMBxfHnV4qO"
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
model = # TF Dense Model
# check how to define a optimizer here https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
optimizer =
# train the model
# compile the model
tf_history = # fit the model
plt.plot(tf_history.history['loss'])
plt.xlabel('Epochs')
plt.ylabel('MSE Loss')
plt.show()
mse = tf_history.history['loss'][-1]
y_hat = model.predict(X_4)
plt.figure(figsize=(12,7))
plt.title('TensorFlow Model')
plt.scatter(X_4[:, 1], y_scaled, label='Data $(X, y)$')
plt.plot(X_4[:, 1], y_hat, color='red', label='Predicted Line $y = f(X)$',linewidth=4.0)
plt.xlabel('$X$', fontsize=20)
plt.ylabel('$y$', fontsize=20)
plt.text(0,0.70,'MSE = {:.3f}'.format(mse), fontsize=20)
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
# + [markdown] colab_type="text" id="bHIymsWZ_kxw"
# 4th degree poly-regression also did a good job in fitting the data as it also have the 3rd degree terms.
# + colab={} colab_type="code" id="BbS6DGuU_mpi"
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
model = # TF Dense Model
# check how to define a optimizer here https://www.tensorflow.org/api_docs/python/tf/keras/optimizers
optimizer =
# train the model for 50 epochs
# compile the model
tf_history = # fit the model
plt.plot(tf_history.history['loss'])
plt.xlabel('Epochs')
plt.ylabel('MSE Loss')
plt.show()
mse = tf_history.history['loss'][-1]
y_hat = model.predict(X_4)
plt.figure(figsize=(12,7))
plt.title('TensorFlow Model')
plt.scatter(X_4[:, 1], y_scaled, label='Data $(X, y)$')
plt.plot(X_4[:, 1], y_hat, color='red', label='Predicted Line $y = f(X)$',linewidth=4.0)
plt.xlabel('$X$', fontsize=20)
plt.ylabel('$y$', fontsize=20)
plt.text(0,0.70,'MSE = {:.3f}'.format(mse), fontsize=20)
plt.grid(True)
plt.legend(fontsize=20)
plt.show()
# + [markdown] colab_type="text" id="H5aMu5aQ_8y_"
# If you run the 4th degree poly-regression for fewer epochs, you can notice, the model tries to fit a 4th(or higher than 3rd) degree polynomial but as the loss is high, the model changes it parameters to set the 4th degree terms to almost 0 and thus giving a 3rd degree polynomial as you train for more epochs.
# + [markdown] colab_type="text" id="VC4GQ-bQXD9v"
#
#
# This is polynomial regression. Yes, its easy. But one issue, as this was a toy dataset we know its a 3rd degree data, so we tried 2,3,4. But when the data is multi dimensional we cannot visualize the dataset, so its difficult to decide the degree. This is why you will see Neural Networks are awesome. They are End-End, they do not need several feature extraction from our side, they can extract necessary features of their own.
#
#
# **Make a Higher degree (4th/5th degree) data and try polynomial regression on it. Also try different functions like exponents, trignometric..etc.**
| 1-Intro-to-Deep-Learning/.ipynb_checkpoints/1_2_1_Assignment_Polynomial_Regression-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (tensorflow)
# language: python
# name: tensorflow
# ---
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_10_3_text_generation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# # T81-558: Applications of Deep Neural Networks
# **Module 10: Time Series in Keras**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 10 Material
#
# * Part 10.1: Time Series Data Encoding for Deep Learning [[Video]](https://www.youtube.com/watch?v=dMUmHsktl04&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_1_timeseries.ipynb)
# * Part 10.2: Programming LSTM with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=wY0dyFgNCgY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_2_lstm.ipynb)
# * **Part 10.3: Text Generation with Keras and TensorFlow** [[Video]](https://www.youtube.com/watch?v=6ORnRAz3gnA&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_3_text_generation.ipynb)
# * Part 10.4: Image Captioning with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=NmoW_AYWkb4&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_4_captioning.ipynb)
# * Part 10.5: Temporal CNN in Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=i390g8acZwk&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_10_5_temporal_cnn.ipynb)
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
try:
# %tensorflow_version 2.x
COLAB = True
print("Note: using Google CoLab")
except:
print("Note: not using Google CoLab")
COLAB = False
# # Part 10.3: Text Generation with LSTM
#
# Recurrent neural networks are also known for their ability to generate text. This can allow the output of the neural network to be free-form text. In this part we will see how an LSTM can be trained on a textual document, such as classic literature, and learn to output new text that appears to be of the same form as the training material. If you train your LSTM on [Shakespeare](https://en.wikipedia.org/wiki/William_Shakespeare), then it will learn to crank out new prose that is similar to what Shakespeare had written.
#
# Don't get your hopes up. Your not going to each your deep neural network to write the next [Pulitzer Prize for Fiction](https://en.wikipedia.org/wiki/Pulitzer_Prize_for_Fiction). The prose generated by your neural network will be nonsensical. However, it will usually be nearly grammatically and of a similar style as the source training documents.
#
# A neural network generating nonsensical text based on literature may not seem terribly useful at first glance. However, the reason that this technology gets so much interest is that it forms the foundation for many more advanced technologies. The fact that the LSTM will typically learn human grammar from the source document opens a wide range of possibilities. Similar technology can be used to complete sentences when a user is entering text. Simply the ability to output free-form text becomes the foundation of many other technologies. In the next part, we will make use of this technique to create a neural network that can write captions for images to describe what is going on in the image.
#
# ### Additional Information
#
# The following are some of the articles that I found useful putting this section together.
#
# * [The Unreasonable Effectiveness of Recurrent Neural Networks](http://karpathy.github.io/2015/05/21/rnn-effectiveness/)
# * [Keras LSTM Generation Example](https://keras.io/examples/lstm_text_generation/)
#
# ### Character-Level Text Generation
#
# There are a number of different approaches to teaching a neural network to output free-form text. The most basic question is if you wish the neural network to learn at the word or character level. In many ways, learning at the character level is the more interesting of the two. The LSTM is learning construct its own words without even being shown what a word is. We will begin with character-level text generation. In the next module, we will see how we can use nearly the same technique to operate at the word level. The automatic captioning that will be implemented in the next module is at the word level.
#
# We begin by importing the needed Python packages and defining the sequence length, named **maxlen**. Time-series neural networks always accept their input as a fixed length array. Not all of the sequence might be used, it is common to fill extra elements with zeros. The text will be divided into sequences of this length and the neural network will be trained to predict what comes after this sequence.
from tensorflow.keras.callbacks import LambdaCallback
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LSTM
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.utils import get_file
import numpy as np
import random
import sys
import io
import requests
import re
# For this simple example we will train the neural network on the classic children's book [Treasure Island](https://en.wikipedia.org/wiki/Treasure_Island). We begin by loading this text into a Python string and displaying the first 1,000 characters.
r = requests.get("https://data.heatonresearch.com/data/t81-558/text/treasure_island.txt")
raw_text = r.text
print(raw_text[0:1000])
# We will extract all unique characters from the text and sort them. This allows us to assign a unique ID to each character. Because the characters are sorted, these IDs should remain the same. If new characters were added to the original text, then the IDs would change. We build up two dictionaries. The first **char2idx** is used to convert a character into its ID. The second **idx2char** converts an ID back into its character.
processed_text = raw_text.lower()
processed_text = re.sub(r'[^\x00-\x7f]',r'', processed_text)
# +
print('corpus length:', len(processed_text))
chars = sorted(list(set(processed_text)))
print('total chars:', len(chars))
char_indices = dict((c, i) for i, c in enumerate(chars))
indices_char = dict((i, c) for i, c in enumerate(chars))
# -
# We are now ready to build the actual sequences. Just like previous neural networks there will be an $x$ and $y$. However, for the LSTM, $x$ and $y$ will both be sequences. The $x$ input will specify the sequences where $y$ are the expected output. The following code generates all possible sequences.
# cut the text in semi-redundant sequences of maxlen characters
maxlen = 40
step = 3
sentences = []
next_chars = []
for i in range(0, len(processed_text) - maxlen, step):
sentences.append(processed_text[i: i + maxlen])
next_chars.append(processed_text[i + maxlen])
print('nb sequences:', len(sentences))
sentences
print('Vectorization...')
x = np.zeros((len(sentences), maxlen, len(chars)), dtype=np.bool)
y = np.zeros((len(sentences), len(chars)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, char in enumerate(sentence):
x[i, t, char_indices[char]] = 1
y[i, char_indices[next_chars[i]]] = 1
x.shape
y.shape
# The dummy variables for $y$ are shown below.
y[0:10]
# Next, we create the neural network. The primary feature of this neural network is the LSTM layer. This allows the sequences to be processed.
# +
# build the model: a single LSTM
print('Build model...')
model = Sequential()
model.add(LSTM(128, input_shape=(maxlen, len(chars))))
model.add(Dense(len(chars), activation='softmax'))
optimizer = RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer)
# -
model.summary()
# The LSTM will produce new text character by character. We will need to sample the correct letter from the LSTM predictions each time. The **sample** function accepts the following two parameters:
#
# * **preds** - The output neurons.
# * **temperature** - 1.0 is the most conservative, 0.0 is the most confident (willing to make spelling and other errors).
#
# The sample function below is essentially performing a [softmax]() on the neural network predictions. This causes each output neuron to become a probability of its particular letter.
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def on_epoch_end(epoch, _):
# Function invoked at end of each epoch. Prints generated text.
print("****************************************************************************")
print('----- Generating text after Epoch: %d' % epoch)
start_index = random.randint(0, len(processed_text) - maxlen - 1)
for temperature in [0.2, 0.5, 1.0, 1.2]:
print('----- temperature:', temperature)
generated = ''
sentence = processed_text[start_index: start_index + maxlen]
generated += sentence
print('----- Generating with seed: "' + sentence + '"')
sys.stdout.write(generated)
for i in range(400):
x_pred = np.zeros((1, maxlen, len(chars)))
for t, char in enumerate(sentence):
x_pred[0, t, char_indices[char]] = 1.
preds = model.predict(x_pred, verbose=0)[0]
next_index = sample(preds, temperature)
next_char = indices_char[next_index]
generated += next_char
sentence = sentence[1:] + next_char
sys.stdout.write(next_char)
sys.stdout.flush()
print()
# We are now ready to train. It can take up to an hour to train this network, depending on how fast your computer is. If you have a GPU available, please make sure to use it.
# +
# Ignore useless W0819 warnings generated by TensorFlow 2.0. Hopefully can remove this ignore in the future.
# See https://github.com/tensorflow/tensorflow/issues/31308
import logging, os
logging.disable(logging.WARNING)
os.environ["TF_CPP_MIN_LOG_LEVEL"] = "3"
# Fit the model
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y,
batch_size=128,
epochs=60,
callbacks=[print_callback])
# -
| t81_558_class_10_3_text_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lagrangian mechanics in generalized coordinates
#
# > <NAME>
# > Laboratory of Biomechanics and Motor Control ([http://demotu.org/](http://demotu.org/))
# > Federal University of ABC, Brazil
# ## Generalized coordinates
#
# The direct application of Newton's laws to mechanical systems results in a set of equations of motion in terms of Cartesian coordinates of each of the particles that make up the system. In many cases, this is not the most convenient coordinate system to solve the problem or describe the movement of the system. For example, in problems involving many particles, it may be convenient to choose a system that includes the coordinate of the center of mass. Another example is a serial chain of rigid links, such as a member of the human body or from a robot manipulator, it may be simpler to describe the positions of each link by the angles between links.
#
# Coordinate systems such as these are referred as [generalized coordinates](https://en.wikipedia.org/wiki/Generalized_coordinates). Generalized coordinates uniquely specify the positions of the particles in a system. Although there may be several generalized coordinates to describe a system, usually a judicious choice of generalized coordinates provides the minimum number of independent coordinates that define the configuration of a system (which is the number of <a href="https://en.wikipedia.org/wiki/Degrees_of_freedom_(mechanics)">degrees of freedom</a> of the system), turning the problem simpler to solve.
#
# Being a little more technical, according to [Wikipedia](https://en.wikipedia.org/wiki/Configuration_space_(physics)):
# "In classical mechanics, the parameters that define the configuration of a system are called generalized coordinates, and the vector space defined by these coordinates is called the configuration space of the physical system. It is often the case that these parameters satisfy mathematical constraints, such that the set of actual configurations of the system is a manifold in the space of generalized coordinates. This manifold is called the configuration manifold of the system."
#
# In problems where it is desired to use generalized coordinates, one can write Newton's equations of motion in terms of Cartesian coordinates and then transform them into generalized coordinates. However, it would be desirable and convenient to have a general method that would directly establish the equations of motion in terms of a set of convenient generalized coordinates. In addition, general methods for writing, and perhaps solving, the equations of motion in terms of any coordinate system would also be desirable. The [Lagrangian mechanics](https://en.wikipedia.org/wiki/Lagrangian_mechanics) is such a method.
#
# When describing a system of particles using any set of generalized coordinates, $q_1,\dotsc,q_{3N}$, these are related to, for example, the Cartesian coordinates by:
#
# \begin{equation} \begin{array}{rcl}
# q_i =q_i (x_1,\dotsc,x_{3N} ) \quad i=1,\dotsc,3N \\
# x_i =x_i (q_1,\dotsc,q_{3N} ) \quad i=1,\dotsc,3N
# \label{eq24}
# \end{array} \end{equation}
#
# The Cartesian components of velocity as a function of generalized coordinates are:
#
# \begin{equation}
# \dot{x}_i =\frac{\mathrm d x_i (q_1, q_2,\dotsc,q_{3N}
# )}{\mathrm d t}=\sum\limits_{j=1}^{3N} {\frac{\partial x_i }{\partial q_j }}
# \frac{\mathrm d q_j }{\mathrm d t}
# \label{eq26}
# \end{equation}
#
# where for simplicity we omitted the explicit mention of the temporal dependence of each coordinate.
#
# That is, any Cartesian component of the particle velocity as a function of generalized coordinates is a function of all the components of position and velocity in the generalized coordinates:
#
# \begin{equation}
# \dot{x}_i = \dot{x}_i (q_1,\dotsc,q_{3N} ,\dot{q}_1,\dotsc,\dot{q}_{3N} ) \quad i=1,\dotsc,3N
# \label{eq27}
# \end{equation}
# ## Lagrange's equation
#
# In analogy to Newtonian mechanics, one can think that the equations of motion can be obtained by equating the generalized force, $F_i$, to the temporal rate of change of each generalized momentum, $p_i$:
#
# \begin{equation}
# F_i =\frac{\partial p_i }{\partial t}
# \label{eq28}
# \end{equation}
#
# In the formula above, let's substitute the quantity $p_i$ by its definition in terms of the kinetic energy:
#
# \begin{equation}
# \frac{\partial p_i }{\partial t} =\frac{\partial }{\partial t}\left( {\frac{\partial T}{\partial
# \dot{q}_i }} \right)=\frac{\partial }{\partial t}\left(
# {\sum\limits_{j=1}^{3N} {m_j \dot{x}_j \frac{\partial \dot{x}_j
# }{\partial \dot{q}_i }} } \right)
# \label{eq29}
# \end{equation}
#
# where we used:
#
# \begin{equation}
# \frac{\partial T}{\partial \dot{q}_i }=\sum\limits_{j=1}^{3N}
# {\frac{\partial T}{\partial \dot{x}_j }\frac{\partial \dot{x}_j
# }{\partial \dot{q}_i }}
# \label{eq30}
# \end{equation}
#
# Using the [product rule](https://en.wikipedia.org/wiki/Product_rule), the derivative of the product in Eq. (\ref{eq29}) is:
#
# \begin{equation}
# \frac{\partial p_i }{\partial t}=\sum\limits_{j=1}^{3N} {m_j
# \ddot{x}_j \frac{\partial \dot{x}_j }{\partial \dot{q}_i }}
# +\sum\limits_{j=1}^{3N} {m_j \dot{x}_j \frac{\mathrm d }{\mathrm d t}\left(
# {\frac{\partial \dot{x}_j }{\partial \dot{q}_i }} \right)}
# \label{eq31}
# \end{equation}
#
# But:
#
# \begin{equation}
# \frac{\partial \dot{x}_i }{\partial \dot{q}_j }=\frac{\partial x_i
# }{\partial q_j } \quad because \quad \frac{\partial
# \dot{x}_i }{\partial \dot{q}_j }=\frac{\partial x_i }{\partial
# t}\frac{\partial t}{\partial q_j }=\frac{\partial x_i }{\partial q_j}
# \label{eq32}
# \end{equation}
#
# Then:
#
# \begin{equation}
# \frac{\partial p_i }{\partial t}=\sum\limits_{j=1}^{3N} {m_j
# \ddot{x}_j \frac{\partial x_j }{\partial q_i }}
# +\sum\limits_{j=1}^{3N} {m_j \dot{x}_j \frac{\mathrm d }{\mathrm d t}\left(
# {\frac{\partial x_j }{\partial q_i }} \right)}
# \label{eq33}
# \end{equation}
# The first term on the right side of the equation above is proportional to $m_j
# \ddot{x}_j$ and we will define as the generalized force, $Q_i$. But, different from Newtonian mechanics, the temporal variation of the generalized momentum is equal to the generalized force plus another term, which will investigate now. The last part of this second term can be derived as:
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial x_j }{\partial q_i }} \right) =
# \sum\limits_{k=1}^{3N} {\frac{\mathrm d }{\mathrm d q_k }\left( {\frac{\partial
# x_j }{\partial q_i }} \right)\frac{\mathrm d q_k }{\mathrm d t}} =\sum\limits_{k=1}^{3N}
# {\frac{\partial^2 x_j }{\partial q_k \partial q_i }\dot{q}_k }
# \label{eq34}
# \end{equation}
#
# where we used the [chain rule](https://en.wikipedia.org/wiki/Chain_rule) for the differentiation:
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\Big( {f\big({g(t)}\big)}\Big) = \frac{\partial f}{\partial g}\frac{\partial g}{\partial t}
# \label{eq35}
# \end{equation}
#
# But if we look at Eq. (\ref{eq26}) we see that the term at the right side of the Eq. (\ref{eq34}) can be obtained by:
#
# \begin{equation}
# \frac{\partial \dot{x}_j }{\partial q_i } = \frac{\partial }{\partial q_i }\left(\sum\limits_{k=1}^{3N} \frac{\partial
# x_j }{\partial q_i }\dot{q}_k \right) = \sum\limits_{k=1}^{3N}
# {\frac{\partial^2 x_j }{\partial q_k \partial q_i }\dot{q}_k }
# \label{eq36}
# \end{equation}
#
# Comparing the Eq. (\ref{eq34}) and Eq. (\ref{eq36}) we have:
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial x_j }{\partial q_i }} \right) =
# \frac{\mathrm d }{\mathrm d q_i}\left( {\frac{\partial x_j }{\partial t }} \right)
# \label{eq37}
# \end{equation}
# On the other hand, it is possible to relate the term $\partial \dot{x}_j / \partial q_i$ to the derivative of kinetic energy with respect to the coordinate $q_i$:
#
# \begin{equation}
# \frac{\partial T}{\partial q_i }=\frac{\partial }{\partial q_i }\left(
# {\sum\limits_{j=1}^{3N} {\frac{1}{2}m_j \dot{x}_j^2} }
# \right)=\sum\limits_{j=1}^{3N} {m_j \dot{x}_j } \frac{\partial
# \dot{x}_j }{\partial q_i }
# \label{eq38}
# \end{equation}
#
# where once again we used the chain rule for the differentiation.
#
# Using Eq. (\ref{eq37}), Eq. (\ref{eq38}) becomes
#
# \begin{equation}
# \frac{\partial T}{\partial q_i }=\sum\limits_{j=1}^{3N} {m_j
# \dot{x}_j } \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial x_j }{\partial q_i }}
# \right)
# \label{eq39}
# \end{equation}
#
# Returning to Eq. (\ref{eq33}), it can be rewritten as:
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial T}{\partial \dot{q}_i }} \right) = Q_i + \frac{\partial T}{\partial q_i }
# \label{eq40}
# \end{equation}
#
# and
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial T}{\partial \dot{q}_i }} \right) - \frac{\partial T}{\partial q_i } = Q_i
# \label{eq41}
# \end{equation}
# Now let's look at $Q_i$, the generalized force. It can be decomposed into two terms:
#
# The first term, composed of the conservative forces, i.e. forces that can be written as potential gradients:
#
# \begin{equation}
# Q_C =-\frac{\partial V}{\partial q_i } \quad , \quad V=V\left( {q_1,\dotsc,q_{3N} } \right)
# \label{eq42}
# \end{equation}
#
# An example of conservative force is the gravitational force.
#
# And the second term, encompassing all non-conservative forces, such as a frictional force, $Q_{NC}$.
#
# Then:
#
# \begin{equation} Q_i =-\frac{\partial V}{\partial q_i }+Q_{NCi} \quad , \quad V=V\left( {q_1,\dotsc,q_{3N} } \right) \end{equation}
#
# The Eq. (\ref{eq41}) becomes
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial T}{\partial \dot{q}_i }}
# \right)-\frac{\partial T}{\partial q_i }=-\frac{\partial V}{\partial q_i} + Q_{NCi}
# \label{eq43}
# \end{equation}
#
# Rearranging, we have:
#
# \begin{equation} \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial \left( {T-V} \right)}{\partial
# \dot{q}_i }} \right)-\frac{\partial \left( {T-V} \right)}{\partial q_i} = Q_{NCi}
# \label{eq44}
# \end{equation}
#
# This is possible because:
#
# \begin{equation}
# \frac{\partial V}{\partial \dot{q}_i} = 0
# \label{eq45}
# \end{equation}
#
# Defining:
#
# \begin{equation}
# \mathcal{L} \equiv \mathcal{L}(q_1,\dotsc,q_{3N} ,\dot{q}_1,\dotsc,\dot{q}_{3N} ) = T - V
# \label{eq46}
# \end{equation}
#
# as the Lagrange or Lagrangian function, we have the Lagrange's equation:
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial \mathcal{L}}{\partial \dot{q}_i }}
# \right)-\frac{\partial \mathcal{L}}{\partial q_i } = Q_{NCi} \quad i=1,\dotsc,3N
# \label{eq47}
# \end{equation}
#
# Once all derivatives of the Lagrangian function are calculated, this equation will be the equation of motion for each particle. If there are $N$ independent particles in a three-dimensional space, there will be $3N$ equations for the system.
#
# The set of equations above for a system are known as Euler–Lagrange equations, or Lagrange's equations of the second kind.
# ### Constraints
#
# An important class of problems in mechanics, in which the Lagrangian equations are particularly useful, are composed of constrained systems. A constraint is a restriction on the freedom of movement of a particle or a system of particles (a constraint decreases the number of degrees of freedom of a system). A rigid body, or the movement of a pendulum, are examples of constrained systems. It can be shown, in a similar way, that the Lagrange equation, deduced here for a system of free particles, is also valid for a system of particles under the action of constraints. The Lagrange's equation, for a system of $3N$ particles and with $k$ constraints, is then defined as:
#
# \begin{equation}
# \frac{\mathrm d }{\mathrm d t}\left( {\frac{\partial \mathcal{L}}{\partial \dot{q}_i}} \right)-\frac{\partial \mathcal{L}}{\partial q_i } = Q_{NCi} \quad i=1,\dotsc,3N-k
# \label{eq48}
# \end{equation}
# ## References
#
# - <NAME> (1980) [Classical Mechanics](https://books.google.com.br/books?id=tJCuQgAACAAJ), 3rd ed., Addison-Wesley.
# - <NAME> (1970) [Classical Dynamics of particles and systems](https://books.google.com.br/books?id=Ss43BQAAQBAJ), 2nd ed., Academic Press.
# - <NAME> (1949) [Principles of Mechanics](https://books.google.com.br/books?id=qsYfENCRG5QC), 2nd ed., McGraw-hill.
# - <NAME> (2005) [Classical Mechanics](https://archive.org/details/JohnTaylorClassicalMechanics). University Science Books.
| notebooks/lagrangian_mechanics_generalized.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Support Vector Machines
# ## CS/DSA 5970
# +
import pandas as pd
import numpy as np
import re
import matplotlib.pyplot as plt
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import mean_squared_error
from sklearn.metrics import confusion_matrix
from sklearn.metrics import log_loss
from sklearn.metrics import roc_curve, auc
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.linear_model import SGDClassifier, LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.svm import SVC
import pickle as pkl
##################
# Default parameters
FIGURESIZE=(10,6)
FONTSIZE=18
plt.rcParams['figure.figsize'] = FIGURESIZE
plt.rcParams['font.size'] = FONTSIZE
plt.rcParams['xtick.labelsize'] = 18
plt.rcParams['ytick.labelsize'] = 18
# -
def scatter_plot(ins, pred):
'''
Generate a scatter plot of the input samples, colored by label. This
label can be the true or predicted labels.
'''
elems_true = np.where(pred == 1)[0]
elems_false = np.where(pred == 0)[0]
fig, ax = plt.subplots(figsize=FIGURESIZE)
ax.plot(ins[elems_true,0], ins[elems_true,1], 'r.')
ax.plot(ins[elems_false,0], ins[elems_false,1], 'g.')
fig.legend(['Positive', 'Negative'], fontsize=18)
def plot_probs(outs, proba):
'''
Compute TPR/FPR statistics and show the TPR/FPR and ROC plots
'''
# Show confusion matrix assuming a neutral cut-off
pred = proba[:,0] >= 0.5
confusion = confusion_matrix(outs, pred)
print("Confusion:", confusion)
# Evaluate
print("log loss: ", log_loss(outs, proba))
# TPR/FPR plot
fpr, tpr, thresholds = roc_curve(outs, proba[:,0])
fig, ax = plt.subplots(figsize=FIGURESIZE)
ax.plot(thresholds, tpr, color='b')
ax.plot(thresholds, fpr, color='r')
ax.plot(thresholds, tpr - fpr, color='g')
ax.invert_xaxis()
ax.set_xlabel('threshold', fontsize=FONTSIZE)
ax.set_ylabel('fraction', fontsize=FONTSIZE)
ax.legend(['TPR', 'FPR', 'distance'], fontsize=FONTSIZE)
# ROC plot
fig, ax = plt.subplots(figsize=FIGURESIZE)
ax.plot(fpr, tpr, color='b')
ax.plot([0,1], [0,1], 'r--')
ax.set_xlabel('FPR', fontsize=FONTSIZE)
ax.set_ylabel('TPR', fontsize=FONTSIZE)
ax.set_aspect('equal', 'box')
print("AUC:", auc(fpr, tpr))
# ## Load data
fname = '../ml_practices/imports/datasets/misc/classification_data.pkl'
fp = open(fname, 'rb')
ins = pkl.load(fp)
outs = pkl.load(fp)
fp.close()
# ## SVC classifier: Linear
# ## Polynomial kernel
# ## Gaussian (RBF) Kernel
| skeletons/svm_script_skel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="exoz57mj2FzI"
# # Tutorial on Information Theory in neuroscience
# Aims of the tutorial:
# - Understand the concept of entropy, conditional entropy, and mutual information
# - Discuss the reasons for binning a continuous signal, the implications introduce some binning strategies and discuss their pros/cons
# - Introsuce the problem of limited-sampling bias: origin, implications and mitigation strategies
# - Discuss the effect of correlated firing in a neuronal population on information
#
# Suggested reading for a deeper understanding of the topics covered in this tutorial:
# 1. Timme, <NAME>., and <NAME>. "A tutorial for information theory in neuroscience." eneuro 5.3 (2018).
# 1. <NAME>., & <NAME>. (2009). Extracting information from neuronal populations: information theory and decoding approaches. Nature Reviews Neuroscience, 10(3), 173-185.
# 1. <NAME>., <NAME>. & <NAME>. Neural correlations, population coding and computation. Nat Rev Neurosci 7, 358–366 (2006). https://doi.org/10.1038/nrn1888 and references.
# + [markdown] id="HkWtYk-98bih"
# ---
# # Setup
# + id="0BrM9Dwl1fNH" colab={"base_uri": "https://localhost:8080/"} outputId="519ead28-9e47-4758-aa94-66bb4c52fb9f" executionInfo={"status": "ok", "timestamp": 1639631713548, "user_tz": -60, "elapsed": 25947, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} cellView="form"
#@title Import
# !pip install pydataset
import pydataset
from pydataset import data
import numpy as np
import random as rnd
from matplotlib import pyplot as plt
from scipy.stats import poisson
import warnings
import os, requests
rnd.seed(2020)
warnings.filterwarnings('ignore')
fname = []
for j in range(3):
fname.append('steinmetz_part%d.npz'%j)
url = ["https://osf.io/agvxh/download"]
url.append("https://osf.io/uv3mw/download")
url.append("https://osf.io/ehmw2/download")
for j in range(len(url)):
if not os.path.isfile(fname[j]):
try:
r = requests.get(url[j])
except requests.ConnectionError:
print("!!! Failed to download data !!!")
else:
if r.status_code != requests.codes.ok:
print("!!! Failed to download data !!!")
else:
with open(fname[j], "wb") as fid:
fid.write(r.content)
# + id="-_kx3sSw7X4Z" cellView="form"
#@title Figure settings
# %config InlineBackend.figure_format = 'retina'
plt.style.use("https://raw.githubusercontent.com/rmaffulli/MI_tutorial/main/figures.mplstyle")
# + id="4XKt93sJQoGm" cellView="form"
#@title Helper functions
def bin_data(data,nbins,mode):
if mode == 'eqspace':
bins = np.linspace(np.min(data)-0.1,np.max(data)+0.1,nbins+1)
data_binned = np.digitize(data,bins,right=True)
return data_binned
# + [markdown] id="MRFZegVu-REj"
# ---
# # Generation of Poisson spikes
# We know that spikes are binary discrete events.
# It is often assumed, in first approximation, that each spike is independent of the previous/following one. This implies that the probability of having a spike at a certain time is not influenced by the probability that a spike has occurred previously. This *model* (beware! this is a model the behaviour of a neuron) is consistent with a spike event to be drawn from a Poisson distribution.
#
# *Food for toughts: what are the mechanisms that invalidate the Poisson assumption? Can we think of some reasons to doubt that a spiking neuron is Poissonian?*
#
# In the cell below you can explore the function generating poissonian spike trains.
# + id="wZuBIU_C-YoV" cellView="form"
#@title Spike generation function
def poisson_spike(t,rate):
# t: time points
# rate: spiking rate in Hz
dt = t[1] - t[0];
sp = np.zeros_like(t)
for i,tp in enumerate(t):
if rnd.uniform(0,1) <= dt*rate:
sp[i] = 1
return sp
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="zbRoVLSNhPCO" cellView="form" outputId="179eff98-f804-420c-d11f-6d98cef1aa18" executionInfo={"status": "ok", "timestamp": 1639631714561, "user_tz": -60, "elapsed": 1031, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}}
#@title The effect of the number of trials on estimation of probabiltiy mass function { run: "auto" }
trials = 82 #@param {type:"slider", min:1, max:1000, step:1}
rate = 9.6 #@param {type:"slider", min:0.1, max:10, step:0.1}
spike_cnt = np.zeros((trials,))
t = np.linspace(0,10,1000)
for tr in range(trials):
sp = poisson_spike(t,rate)
spike_cnt[tr-1] = sum(sp)
_ = plt.hist(spike_cnt,
range=(0, 2*int(spike_cnt.max())),
bins=int(spike_cnt.max() + 1),
density=True, histtype='stepfilled',
color='teal', edgecolor='black')
x_poisson = np.linspace(1,2*int(spike_cnt.max()),2*int(spike_cnt.max()))
_ = plt.plot(x_poisson, poisson.pmf(x_poisson, rate*10),
'-o', label='Analytical Poisson PMF',
color='goldenrod')
plt.xlabel('Number of spikes in 10 seconds [-]')
plt.ylabel('Probability of N spikes in 10 seconds [-]')
_ = plt.legend()
# + [markdown] id="BMiR1HwQwckF"
# ---
# # Entropy of a random variable
# In this section we will:
# - Learn how to calculate the entropy of a random variable
# - Understand how the probability distribution of such random variable influences its entropy
#
# Given a Random Variable (RV) $R$, its entropy is defined as:
#
# $H(R) = \sum\limits_{r\in X}p(x)log_2\frac{1}{p(x)}$
#
# in the section below we will implement the following functions:
#
# - `def marginal_p(x):` to calculate the marginal probability of a RV $X$
# - `def entropy(x):` to calculate the entropy of $(X)$
# + id="1SVSoLmlPOnR" colab={"base_uri": "https://localhost:8080/"} outputId="9ef7f811-8f46-439e-9500-4abe9577758b" executionInfo={"status": "ok", "timestamp": 1639631714563, "user_tz": -60, "elapsed": 18, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}}
def marginal_p(x):
# returns marginal probabiltiy of random variable X based on observations x
# x is a column vector: rows are the trials
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# find unique values of x and count the number of occurrences across trials
# ( ... )
# calculate probabilities of each occurrences
# ( ... )
# return probability vector
return p_x
def entropy(x):
# returns entropy of random variable X
# x is a column vector: rows are the trials
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# initialize entropy
ent = 0
# calculate marginal probability
p_x = marginal_p(x)
# calculate entropy
# ( ... )
# return entropy
return ent
x = np.asarray([1, 1, 0, 0])
print(entropy(x))
if entropy(x) == 1:
print("Correct! the value of entropy(x) is: " + str(entropy(x)))
else:
raise NotImplementedError("Wrong values of entropy, double check the implementation of entropy and marginal functions")
# + colab={"base_uri": "https://localhost:8080/", "height": 445} id="dBIntCZD-nQC" outputId="ee151844-e630-4e7a-8c3e-8fa22277b69a" executionInfo={"status": "ok", "timestamp": 1639631715105, "user_tz": -60, "elapsed": 552, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} cellView="form"
#@title Entropy of a Gaussian random variable { run: "auto" }
mean = 0
std = 2.5 #@param {type:"slider", min:0.5, max:20, step:0.5}
trials = 99910 #@param {type:"slider", min:10, max:100000, step:100}
x = np.random.normal(mean,std,trials)
bins = np.linspace(mean-100,mean+100,1000)
x_binned = np.zeros_like(bins)
for i in np.digitize(x,bins,right=True):
x_binned[i-1] += 1
p_x = x_binned/sum(x_binned)
_ = plt.plot(bins,p_x,color='teal')
plt.xlabel('x [-]')
plt.ylabel('p(x) [-]')
print("Entropy: " + str(entropy(x_binned)) + " [bits]")
# + [markdown] id="UocSwXQCBFGz"
# ## Discussion
# - What is the effect of the standard deviation on entropy? Why? What pdf do you expect to carry maximum entropy?
# - What is the effect of the mean?
# - What is the effect of the number of trials?
# + [markdown] id="o0tbK43mws0T"
# ---
# # Conditional entropy of two random variables
# In this section we will:
# - Learn how to calculate the conditional entropy of one random variable X given a second variable Y
#
# In the section below we will implement the following functions:
#
# - `def joint_p(x,y):` to calculate the joint probability of $(X;Y)$
# - `def conditional_p(x,y):` to calculate the conditional probability of a $(X|Y)$
# - `def conditional_entropy(x,y):` to calculate the conditional entropy of $(X|Y)$
# + colab={"base_uri": "https://localhost:8080/"} id="38UmSKKWC85o" outputId="129363d3-ce42-413e-c840-b1424311d797" executionInfo={"status": "ok", "timestamp": 1639631715107, "user_tz": -60, "elapsed": 25, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}}
def joint_p(x,y):
# returns joint probabiltiy of random variables (X;Y) based on observations x,y
# x is a column vector: rows are the trials
# y is a column vector: rows are the trials
# the output value p_xy should be a matrix with rows number equal to the unique values of x and columns number equal to the unique values of y
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# find unique values of x and count the number of occurrences across trials
x_values, p_x = np.unique(x, axis=0, return_counts=True)
y_values, p_y = np.unique(y, axis=0, return_counts=True)
# initialize joint probability matrix
p_xy = np.zeros((x_values.size, y_values.size))
# calculate probabilities of each occurrences
for x_r,y_r in zip(x,y):
# ( ...)
p_xy = p_xy/np.shape(x)[0]
# return probability matrix
return p_xy
def conditional_p(x,y):
# returns conditional probabiltiy (X|Y) based on observations x,y
# x is a column vector: rows are the trials
# x is a column vector: rows are the trials
# HINT: use Bayes' formula!!!
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
p_xy = joint_p(x,y)
p_y = marginal_p(y)
p_x_cond_y = np.zeros_like(p_xy)
# return conditional probability matrix
# ( ... )
return p_x_cond_y
def conditional_entropy(x,y):
# returns conditional entropy (X|Y) based on observations x,y
# x is a column vector: rows are the trials
# y is a column vector: rows are the trials
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# calculate joint probability
p_xy = joint_p(x,y)
# calculate conditional probability
p_x_cond_y = conditional_p(x,y)
# calculate entropy
# ( ... )
# return entropy
return cond_entr
x = np.asarray([1, 2, 3, 4])
y = np.asarray([1, 1, 2, 2])
if conditional_entropy(x,y) == 1:
print("Correct! the value of entropy(x) is: " + str(conditional_entropy(x,y)))
else:
raise NotImplementedError("Wrong values of entropy, double check the implementation of entropy and marginal functions")
# + [markdown] id="Zu8yi9jbDJSR"
# ## Calculate the conditional entropy of real world data
# We will now play a bit with real data. We will import a dataset of property prices in the suburbs of Boston and investigate the conditional entropy between a few variables in this dataset.
#
# Source: The Boston house-price data of <NAME>. and <NAME>. 'Hedonic prices and the demand for clean air' J. Environ. Economics & Management, vol.5, 81-102, 1978.
# + id="pFWRLTXURBrY" cellView="form" executionInfo={"status": "ok", "timestamp": 1639631715823, "user_tz": -60, "elapsed": 731, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} outputId="44cc5723-c6d9-4b4b-dced-782b76827aa1" colab={"base_uri": "https://localhost:8080/", "height": 481}
#@title {run: "auto" }
x_var = "crim" #@param ["crim", "nox", "ptratio", "age", "medv"]
y_var = "nox" #@param ["crim", "nox", "ptratio", "age", "medv"]
axis_labels = {
"crim":"Per capita crime rate by town",
"nox":"Nitric oxides concentration (parts per 10 million)",
"ptratio":"Pupil-teacher ratio by town",
"age":"Proportion of owner-occupied units built prior to 1940",
"medv":"Median value of owner-occupied homes in $1000's"
}
# Import data
imported_data = data('Boston')
x = imported_data[x_var]
y = imported_data[y_var]
# bin data
x_binned = bin_data(x,10,'eqspace')
y_binned = bin_data(y,10,'eqspace')
plt.scatter(x,y,color='teal')
plt.xlabel(axis_labels[x_var])
plt.ylabel(axis_labels[y_var])
print("Entropy x = ", entropy(x_binned))
print("Entropy y = ", entropy(y_binned))
print("Conditional entropy (x|y)) = "
+ str(conditional_entropy(x_binned,y_binned)))
# + [markdown] id="RCyxUHwFU2m8"
# ## A note
# - Never forget that marginal, joint and conditional probability aare related through Bayes rule: $$ p(X|Y) = \frac{p(X,Y)}{p(Y)} $$
# so you normally have just to calculate two of them and use Bayes rule to obtain the third one!
# + [markdown] id="Yfi9_4lEcCnn"
# ## Discussion
# - Can you see a relation between the distributions of X, Y and the conditional entropy $H(X|Y)$?
# - Try to see what happens with `'crim'` as $X$ and `'nox'` as $Y$
# - What happens with the inverse (i.e. `'crim'` as $Y$ and `'nox'` as $X$)? Is conditional entropy invariant to swapping $X$ and $Y$?
# - What is the conditional entropy of a variable with itself?
# - What is the conditional entropy of `'age'` as $X$ and `'medv'` as $Y$?
# - Do you think that conditional entropy alone is enough to characterize the dependence between two variables? How do you define *high* vs *low* values of conditional entropy?
# + [markdown] id="qEfcI2OSwyc2"
# ---
# # Mutual Information
# Conditional entropy is an indication of how much entropy is left in a variable once we have observed another one. It is as such an indicator of the dependence between two variables but cannot tell us how informative is a variable about another one as it does not give us a comparison term to define how *high* or *low* this the information content that one variable has with respect to the other one. This is precisely the role of mutual information.
#
# Mutual information is defined as:
# $$MI(X;Y) = H(X) - H(X|Y)$$
# and it quantifies how much uncertainty is left in $X$ once we have observed variable $Y$. In this way it relates the conditional entropy $H(X|Y)$ with the total entropy $H(X)$.
#
# In this section we will:
# - Learn how to calculate mutual information between two random variables
# - Calculate MI for the Boston dataset analyzed above
# - Introduce MI for a spike-rate coding Poisson spiking neuron
#
# In the section below we will implement the following functions:
#
# - `def mi(x,y):` to calculate the mutual information of $(X;Y)$
# + id="9KShOHCk4RwI"
def mi(x,y):
# returns mutual information between (X;Y) based on observations x,y
# x is a column vector: rows are the trials
# y is a column vector: rows are the trials
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# calculate mutual information
# (...)
return info
# + colab={"base_uri": "https://localhost:8080/", "height": 506} id="GMW0ZyRZ78X1" cellView="form" outputId="02366afd-2241-41d0-c999-e3e291c51713" executionInfo={"status": "ok", "timestamp": 1639631716496, "user_tz": -60, "elapsed": 691, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}}
#@title {run: "auto" }
x_var = "crim" #@param ["crim", "nox", "ptratio", "age", "medv"]
y_var = "medv" #@param ["crim", "nox", "ptratio", "age", "medv"]
axis_labels = {
"crim":"Per capita crime rate by town",
"nox":"Nitric oxides concentration (parts per 10 million)",
"ptratio":"Pupil-teacher ratio by town",
"age":"Proportion of owner-occupied units built prior to 1940",
"medv":"Median value of owner-occupied homes in $1000's"
}
# Import data
imported_data = data('Boston')
x = imported_data[x_var]
y = imported_data[y_var]
# bin data
x_binned = bin_data(x,10,'eqspace')
y_binned = bin_data(y,10,'eqspace')
plt.scatter(x,y,color='coral')
plt.xlabel(axis_labels[x_var])
plt.ylabel(axis_labels[y_var])
print("Entropy x = ", entropy(x_binned))
print("Entropy y = ", entropy(y_binned))
print("Conditional entropy (x|y)) = "
+ str(conditional_entropy(x_binned,y_binned)))
print("Mutual information (x;y)) = "
+ str(mi(x_binned,y_binned)))
# + [markdown] id="Xy4JqPwE-Nqe"
# ## Discussion
# - Can you see a relation between the distributions of X, Y, the conditional entropy $H(X|Y)$ and the mutual information?
# - Try to see what happens with `'crim'` as $X$ and `'nox'` as $Y$
# - What happens with the inverse (i.e. `'crim'` as $Y$ and `'nox'` as $X$)? Is mutual information invariant to swapping $X$ and $Y$?
# - What is the MI of a variable with itself?
# - What is the MI of `'age'` as $X$ and `'medv'` as $Y$?
# - What is the MI of `'nox'` as $X$ and `'ptratio'` as $Y$?
# + [markdown] id="sodc5PpSw1R1"
# ---
# # A digression on binning
# In this section we will:
# - Discuss the effects of number of bins on the calculation of MI
# - Consider the difference between binning strategies
#
# As we have seen, calculating entropies and mutual information requires us to know (or better, estimate) probabilities. Broadly speaking, there are two main strategies to achieve this:
#
# 1. by fitting a probability distribution to our data (either by assuming its distribution or not)
# 1. by using our data to create a discrete histogram of the probability distribution (frequently referred to as *plug-in* or *direct* estimation)
#
# The methods of the first class are normally far more computationally expensive but, if done correctly, ensure more reliable results. In a plug-in estimation, instead, we calculate the probabilities using a frequentist approach on the data that we have available. This is the approach that we have followed so far and that we'll follow for the rest of the tutorial.
# + [markdown] id="MuIgpIHWcO8P"
# ## Discussion
# - Can you think of what are the consequences of using a plug-in estimation?
# - Can a plug-in estimation be applied directly to continuous data?
# + [markdown] id="8GiB-VRqcTe5"
# ## Binning strategies: uniform-width vs uniform-size
# - In uniform-width binning (the approach we followed so far) we divide the range between min and max observed values of the RV of interest in $N$ equally-wide bins.
# - In uniform-size (with size here we refer to the population size) bins we split the range bewteen min and max observed values in $N$ bins, all containing the same number of samples.
#
# These are **not** the only strategies used but are the most frequently used. In the sectiob below you will implement binning through equal sizing.
#
# In the section below we will implement the following function:
#
# - `defbin_data(data,nbins,mode):` to bin data using either equally spaced or equally populated bins
# + id="0YHmDhAaRmyh"
def bin_data(data,nbins,mode):
# returns binned version of data
# data is a column vector to be binned: rows are the trials
# nbins is the number of bins that is going to be used
# mode is the binning mode, either 'eqspace' for equally spaced bins or 'eqsize' for equally populated bins
# first implement 'eqspace' mode
if mode == 'eqspace':
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
#()...)#
# then implement equally sized mode
elif mode == 'eqsize':
data_binned = np.zeros_like(data)
sorted_idxs = np.argsort(data)
split_idxs = np.array_split(sorted_idxs, nbins)
bin = 1
for s_i in split_idxs:
for i in s_i:
data_binned[i] = bin
bin += 1
else:
raise ValueError("Wrong binning mode given as input")
# return data_binned
return data_binned
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="VJp90PcDhHpu" cellView="form" outputId="31936138-8931-4f01-e402-5d912e07e3e9" executionInfo={"status": "ok", "timestamp": 1639631717252, "user_tz": -60, "elapsed": 769, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}}
#@title Histogram resulting from different binning strategies {run: "auto" }
trials = 16 #@param {type:"slider", min:1, max:1000, step:1}
rate = 4 #@param {type:"slider", min:1, max:10, step:1}
n_bins = 5 #@param {type:"slider", min:2, max:50, step:1}
tm = np.linspace(0,10,2000)
sp_cnt = np.zeros((trials,))
for i,_ in enumerate(sp_cnt):
sp_cnt[i] = np.sum(poisson_spike(tm,rate))
binned_sp_cnt_esp = bin_data(sp_cnt,n_bins,'eqspace')
binned_sp_cnt_esi = bin_data(sp_cnt,n_bins,'eqsize')
f, axs = plt.subplots(1,3)
_ = axs[0].hist(sp_cnt,bins=np.arange(np.min(sp_cnt),np.max(sp_cnt)),
color='teal',edgecolor='black')
_ = axs[0].set_title('Original distrubition',
fontsize = 15)
_ = axs[1].hist(binned_sp_cnt_esp, bins=n_bins,
color='teal',edgecolor='black')
_ = axs[1].set_title('Equally spaced bins',
fontsize = 15)
_ = axs[2].hist(binned_sp_cnt_esi, bins=n_bins,
color='teal',edgecolor='black')
_ = axs[2].set_title('Equally sized bins',
fontsize = 15)
# + [markdown] id="7aLdxqnRZO_-"
# ## Discussion
# - What you can immediately see from the different histograms?
# - What do you think it's the favourable property of equal width binning?
# - Why instead you may prefer equally sized bins?
# + [markdown] id="wmsXDccKEYAX"
# ## Rules of thumb for estimation of the number of bins
# The estimation of the number of bins belongs to the category of the *\"(ubiquitous) things in science that are fundamental for something but having no exact rule to follow for their determination\"*. Much is left to the experience of the user.
#
# There are, however some established rules of thumb that can help:
# 1. **The Freedman-Diaconis rule** (<NAME>., <NAME>. On the histogram as a density estimator:L 2 theory. Z. Wahrscheinlichkeitstheorie verw Gebiete 57, 453–476 (1981). https://doi.org/10.1007/BF01025868):
# $$N = 2\times IQR\times n^{-1/3}$$
# where $IQR$ is the inter-quartile range and $n$ is the number of samples.
# 1. **Scott's rule (normality assumption)** (<NAME>., On optimal and data-based histograms, Biometrika, Volume 66, Issue 3, December 1979, Pages 605–610, https://doi.org/10.1093/biomet/66.3.605):
# $$N = 3.5\times \sigma\times n^{-1/3}$$
# where $\sigma$ is the standard deviation range and $n$ is the number of samples.
# 1.**Sturges rule (used only for very large number of samples [practically never the case in neuroscience])**(Sturges, <NAME>. "The choice of a class interval." Journal of the american statistical association 21.153 (1926): 65-66):
# $$N = 1 + log_2n$$
#
# Another issue with using a plug-in estimate is due to bias. We'll cover this in the next section.
# + [markdown] id="SD4dMqAHxKwY"
# ---
# # Bias and bias correction strategies
# Estimating the probability distribution from a limited size data sample results inevitably in errors. Mutual information is a positive quantity and errors in the probability estimation will **always (on average)** introduce a positive bias.
#
# We have seen that (always on average) limited sampling introduces a negative bias on entropy. Given that the conditional entropy $H(X|Y)$ is estimated on a smaller sample than the marginal entropy $H(X)$ (as it is estimated on a subset of the whole available data) it is going to be affected by a higher negative bias than $H(X)$. As such the MI will be affected by a positive bias.
#
# In this section we will:
# - Demonstrate the effect of bias on the calculation of MI
# - Implement quadratic extrapolation for bias correction
# + colab={"base_uri": "https://localhost:8080/", "height": 428} id="ic8keyBEauoF" cellView="form" outputId="15c7ee85-4641-4bf7-968f-74c7119fa523" executionInfo={"status": "ok", "timestamp": 1639631728988, "user_tz": -60, "elapsed": 11751, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}}
#@title {run: "auto" }
rate_1 = 3 #@param {type:"slider", min:1, max:10, step:0.5}
rate_2 = 3 #@param {type:"slider", min:1, max:10, step:0.5}
n_bins = 50 #@param {type:"slider", min:2, max:50, step:1}
n_rep_stats = 20
n_trials_to_sample = 6
rate = [rate_1, rate_2]
trials = np.round(np.linspace(2,100,n_trials_to_sample))
tm = np.linspace(0,2,400)
MI = np.zeros((n_trials_to_sample,n_rep_stats))
for t,trials_n in enumerate(trials):
for r in range(n_rep_stats):
trials_n = int(trials_n)
sp_cnt = np.zeros((2*trials_n,))
# build stimuli array
stimuli = np.block([1+np.zeros((trials_n,)), 2+np.zeros((trials_n,))])
# build response array
for s in [0,1]:
for i in range(0,trials_n):
sp_cnt[s*trials_n+i] = np.sum(poisson_spike(tm,rate[s]))
# bin responses
binned_sp_cnt = bin_data(sp_cnt,n_bins,'eqspace')
# calculate MI
MI[t,r] = mi(stimuli,binned_sp_cnt)
# extract mean values and std for error bars
MI_mn = np.mean(MI,axis=1)
MI_std = np.std(MI,axis=1)
_ = plt.errorbar(trials, MI_mn, yerr=MI_std, fmt='o', color='teal',
ecolor='lightgray', elinewidth=3, capsize=0);
_ = plt.xlabel("Number of trials per stimulus")
_ = plt.ylabel("MI [bits]")
# + [markdown] id="TqMM0YxJ0H83"
# ## Discussion
# - What is the effect of bias?
# - How does it change as one changes the number of trials/rates difference/number of bins? Why?
#
# ## Bias correction through quadratic extrapolation
# We have observed that, given a finite number of samples $N$, the MI will be always positively biased. One way to correct for finite sample size bias is to use quadratic extrapolation.
#
# The idea is to calculate $MI_N$, $MI_{N/2}$, $MI_{N/4}$ (the values of MI calculated using respectively $N$, $N/2$, $N/4$ samples. One can then simply infer $MI_{\infty}$ (value of MI with infinite samples) by quadratic extrapolation of $MI_N$, $MI_{N/2}$, $MI_{N/4}$.
#
# In the section below we will implement the following function:
#
# - `def bias_corr_mi(x,y):` to perform bias corrected predictions of MI using quadratic extrapolation
# + id="gRlvdA-GzN3M"
def bias_corr_mi(x,y):
# returns bias corrected mutual information between (X;Y) based on observations x,y
# x is a column vector: rows are the trials
# y is a column vector: rows are the trials
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# perform n_stats_runs different calculations for a reliable estimation of the the MI using a subset of the original samples
n_stats_runs = 50
n_trials = x.size
# store in MI_n the values of MI obtained with 100%, 50% and 25% of the trials
MI_n = np.zeros((3,))
subset_size = np.asarray([n_trials, round(n_trials/2), round(n_trials/4)])
# calculate MI for N, N/2 and N/4 and store it in MI_n
for i in range(2):
MI_tmp = np.empty((n_stats_runs,))
MI_tmp[:] = np.nan
for r in range(n_stats_runs):
subset_indices = np.random.choice(n_trials, subset_size[i])
MI_tmp[r] = mi(x[subset_indices],y[subset_indices])
if i==0:
break
MI_n[i] = np.nanmean(MI_tmp)
# perform quadratic extrapolation
# ( ... )
# return output bc_MI
return bc_MI
# + id="c0LsPghpCJnO" colab={"base_uri": "https://localhost:8080/", "height": 428} outputId="0eeb5bd8-8320-437d-b182-24bb8cab3b48" executionInfo={"status": "ok", "timestamp": 1639631757724, "user_tz": -60, "elapsed": 28747, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} cellView="form"
#@title {run: "auto" }
rate_1 = 10 #@param {type:"slider", min:1, max:10, step:0.5}
rate_2 = 9 #@param {type:"slider", min:1, max:10, step:0.5}
n_bins = 10 #@param {type:"slider", min:2, max:10, step:1}
n_rep_stats = 20
n_trials_to_sample = 8
rate = [rate_1, rate_2]
trials = np.round(np.linspace(2,50,n_trials_to_sample))
tm = np.linspace(0,2,400)
MI = np.zeros((n_trials_to_sample,n_rep_stats))
MI_bc = np.zeros((n_trials_to_sample,n_rep_stats))
for t,trials_n in enumerate(trials):
for r in range(n_rep_stats):
trials_n = int(trials_n)
sp_cnt = np.zeros((2*trials_n,))
# build stimuli array
stimuli = np.block([1+np.zeros((trials_n,)), 2+np.zeros((trials_n,))])
# build response array
for s in [0,1]:
for i in range(0,trials_n):
sp_cnt[s*trials_n+i] = np.sum(poisson_spike(tm,rate[s]))
# bin responses
binned_sp_cnt = bin_data(sp_cnt,n_bins,'eqspace')
# calculate MI
MI[t,r] = mi(stimuli,binned_sp_cnt)
MI_bc[t,r] = bias_corr_mi(stimuli,binned_sp_cnt)
# extract mean values and std for error bars
MI_mn = np.mean(MI,axis=1)
MI_std = np.std(MI,axis=1)
MI_bc_mn = np.mean(MI_bc,axis=1)
MI_bc_std = np.std(MI_bc,axis=1)
_ = plt.errorbar(trials, MI_mn, yerr=MI_std, fmt='o', color='teal',
ecolor='lightgray', elinewidth=3, capsize=0,
label='Naive estimate');
_ = plt.errorbar(trials+1, MI_bc_mn, yerr=MI_bc_std, fmt='o', color='coral',
ecolor='lightgray', elinewidth=3, capsize=0,
label='Bias corrected');
_ = plt.xlabel("Number of trials per stimulus")
_ = plt.ylabel("MI [bits]")
_ = plt.legend()
# + [markdown] id="BeY6k6ChIHJs"
# ---
# # Effect of noise correlations in population coding
# In this section we will:
# - Define the response of 2 neurons to 2 different stimuli and see how noise correlations affect their response:
# - Can we build an hypothesis about the effect on noise correlations on information in neuronal populations?
# - Discuss the problems arising from looking at multi-dimensional responses
# - Confirm/discard our hypothesis on the effect of noise correlations on information in neuronal populations through numerical calculations
#
# + id="ZpzGxBfIfAmO" executionInfo={"status": "ok", "timestamp": 1639631759068, "user_tz": -60, "elapsed": 1351, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} outputId="39829e58-2fc2-4a42-d74c-ac2c15e1ebe7" colab={"base_uri": "https://localhost:8080/", "height": 463} cellView="form"
#@title Noise correlation and MI {run: "auto" }
case = "Case B" #@param ["Case A", "Case B"]
if case == "Case A":
n1_rate_1 = 10
n1_rate_2 = 5
n2_rate_1 = 10
n2_rate_2 = 5
sign = [-1, -1]
else:
d = 0.2
n1_rate_1 = 12
n1_rate_2 = 6
n2_rate_1 = 22
n2_rate_2 = 16
sign = [1, -1]
alpha = 1 #@param {type:"slider", min:0, max:1, step:0.1}
n_trials = 200
sigma = 0.5
n_bins = 2
tm = np.linspace(0,5,1000)
rates = np.array([[n1_rate_1, n2_rate_1],
[n1_rate_2, n2_rate_2]])
sp_cnt_1 = np.zeros((2*n_trials,))
sp_cnt_2 = np.zeros((2*n_trials,))
sp_cnt = np.zeros((2*n_trials,2))
therates = np.zeros((2*n_trials,2))
binned_sp_cnt = np.zeros_like(sp_cnt)
# build stimuli array
stimuli = np.zeros((2*n_trials,))
# build response array
for s in range(2):
for i in range(0,n_trials):
noise_s = np.random.normal(2,1)
for n in range(2):
stimuli[s*n_trials+i] = s
therates[s*n_trials+i,n] = rates[s,n] - sigma*(rates[0,n] - rates[1,n])*\
(alpha*noise_s+np.sqrt(1-alpha**2)*np.random.normal(0,1))
sp_cnt[s*n_trials+i,n] = np.sum(poisson_spike(tm,
rates[s,n] +
sign[n]*sigma*(rates[0,n] - rates[1,n])*\
(alpha*noise_s+np.sqrt(1.0-alpha**2.0)*np.random.normal(0,1))))
# bin responses (independently for each neuron)
for n in [0,1]:
binned_sp_cnt[:,n] = bin_data(sp_cnt[:,n],n_bins,'eqspace')
# transform response to 1D
one_d_resps = np.zeros((binned_sp_cnt.shape[0],))
unique_resps, p_x = np.unique(binned_sp_cnt, axis=0, return_counts=True)
for i in range(binned_sp_cnt.shape[0]):
unique_resp_id = np.where(np.all(unique_resps == binned_sp_cnt[i,:],axis=1))[0]
one_d_resps[i] = unique_resp_id
# plot responses
scattercols = ['teal','coral']
legends = ['Stim 1', 'Stim 2']
for s in [0,1]:
_ = plt.plot(sp_cnt[s*n_trials:s*n_trials+n_trials-1,0] , sp_cnt[s*n_trials:s*n_trials+n_trials-1,1] , 'o',
markersize=3, color=scattercols[s], label=legends[s])
_ = plt.xlabel("Spike count neuron 1 [-]")
_ = plt.ylabel("Spike count neuron 2 [-]")
_ = plt.legend()
# calculate MI
MI = mi(stimuli,one_d_resps)
print("Mutual Information = " + str(MI))
print("Correlation coefficient = " + str(np.corrcoef(sp_cnt,rowvar=False)[0,1]))
# + [markdown] id="aFkwlJfz-4Ow"
# ## Discussion
# - Is *Case A* a case in which correlation limits or enhances information? Why is this the case?
# - What about *Case B*?
# + [markdown] id="MhQ6bXQZE7Ob"
# # Significance testing
# So far we have only looked at how to calculate values of mutual information but we have not considered how to assess how significative they are. Namely, what is the probabilty that the same value of $MI(X;Y)$ can be obtained from the same data if the $X$ and $Y$ actually contain no significative relation?
#
# We can then compare $MI(X;Y)$ with the values of $MI(X;Y)$ obtained by shuffling randomly the elements in $X$ or $Y$. The shuffling operation destroys the relation between $X$ or $Y$ and returns the value of $MI(X;Y)$ when the relation between the two variables is destroyed.
#
# In this section we will:
# - Build a new version of the `mi` function allowing us to calculate also the *p-value* of the estimated mutual information
# + id="JTnfU-z3G3BU"
def mi_btsp(x,y,n_btsp):
# returns mutual information between (X;Y) based on observations x,y
# x is a column vector: rows are the trials
# y is a column vector: rows are the trials
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
mutual_info = mi(x,y)
mutual_info_btsp = np.empty(n_btsp,)
# calculate bootstrap estimates of mutual information
# ( ... )
if n_btsp > 0:
p_val = sum(mutual_info_btsp >= mutual_info)/n_btsp
else:
p_val = 0
return mutual_info, p_val
# + [markdown] id="UMarRnUuwkYv"
# # Application to real data
# In this section we are going to use the tools developed in the previous parts of this tutorial to real neural recordings. For full details on the experimental dataset refer to:
#
# > <NAME>., <NAME>., <NAME>. et al. Distributed coding of choice, action and engagement across the mouse brain. Nature 576, 266–273 (2019). https://doi.org/10.1038/s41586-019-1787-x ]
#
# ## Brief descriptio of the experimental protocol
# 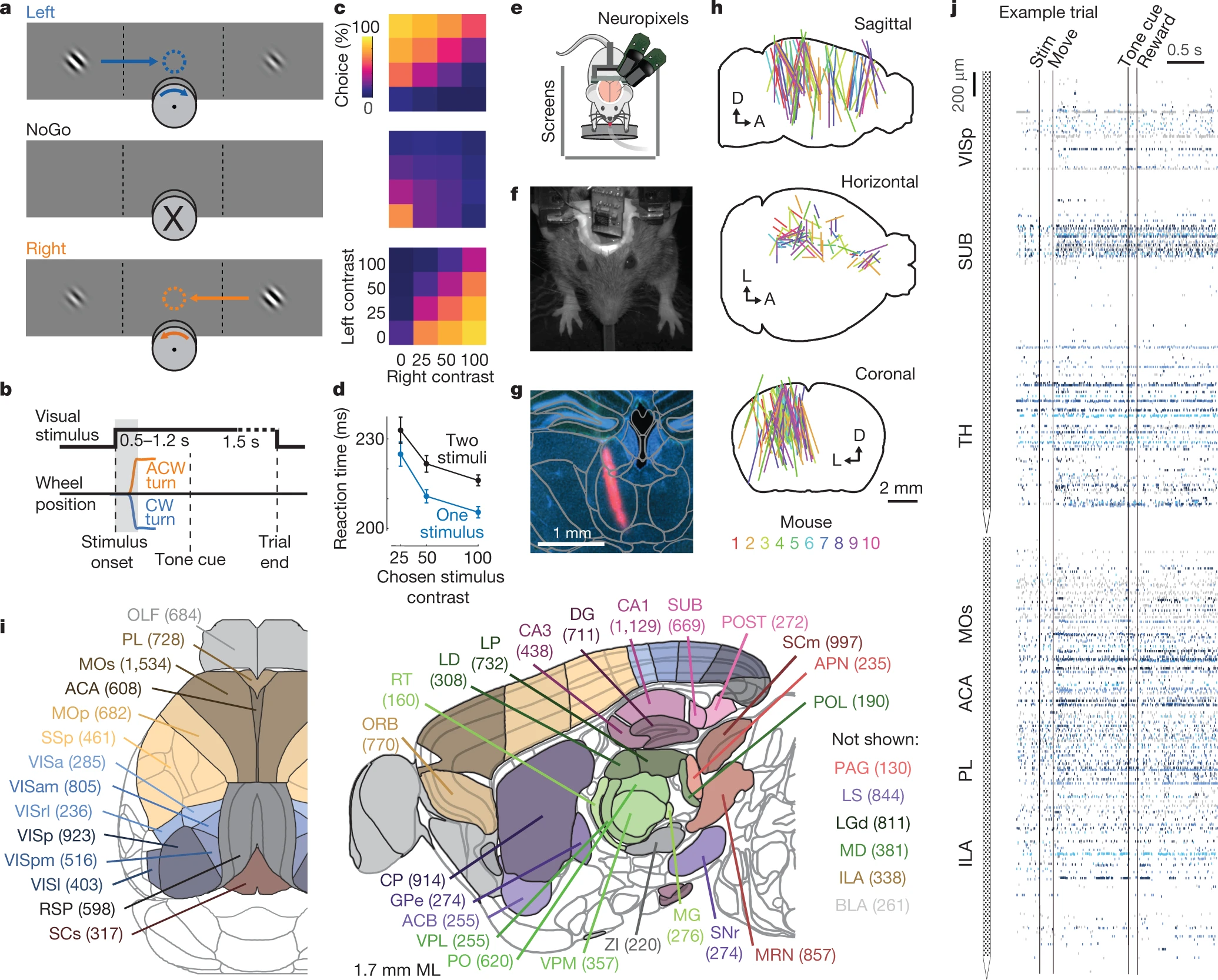
#
# In this section we will:
# - Calculate the stimulus-related MI in visual cortex
# - Compare it with the stimulus related MI in motor cortex
# - Compare it with the choice related MI in motor cortex
# + id="sffzC_hyLgWZ" cellView="form"
#@title Load dataset
import numpy as np
alldat = np.array([])
for j in range(len(fname)):
alldat = np.hstack((alldat, np.load('steinmetz_part%d.npz'%j, allow_pickle=True)['dat']))
# + id="E3-3wRLra7q3" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1639632087338, "user_tz": -60, "elapsed": 278, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} outputId="c8c75446-9812-4cb7-ec9b-a4d1caa7fc89" cellView="form"
#@title Extracting used data
session = 7
vis_area_neurons = alldat[session]['brain_area'] == "VISp"
mos_area_neurons = alldat[session]['brain_area'] == "MOs"
n_vis_area_neurons = sum(vis_area_neurons == True)
n_mos_area_neurons = sum(mos_area_neurons == True)
print("Number of VISp neurons:", n_vis_area_neurons)
print("Number of MOs neurons:", n_mos_area_neurons)
stim_i = 50
gocue_i = np.min(np.round(alldat[session]['gocue']/0.01+stim_i).astype(int))
# extract spike trains (neurons x trials x time points)
spikes_vis = alldat[session]['spks'][vis_area_neurons,:,stim_i:gocue_i]
spikes_mos = alldat[session]['spks'][mos_area_neurons,:,gocue_i:]
# sum spike count to obtain rate code
spikes_vis = np.sum(spikes_vis,axis=2)
spikes_mos = np.sum(spikes_mos,axis=2)
contrast_controlateral = alldat[session]['contrast_right']
choice = alldat[session]['response']
# + [markdown] id="8_jfYBWDKegL"
# The following data have been imported:
# - `spikes_vis`: array of spike count in Visual Cortex for one recording session (*nNeurons x nTrials*)
# - `spikes_mos`: array of spike count in Motor Cortex for one recording session (*nNeurons x nTrials*)
# - `choice`: behavioral outcome of each trial (*nTrials x 1*)
# - `contrast_controlateral`: visual stimulus (*nTrials x 1*)
# + [markdown] id="YLUw64G4sfDU"
# # Choice- and Stimulus-related Mutual Information in cortical areas
# In this section we will calculate, using the data above the following quantities:
# - $MI(Stim; Resp)$ in visual cortex
# - $MI(Choice; Resp)$ in motor cortex
# - $MI(Stim; Resp)$ in motor cortex
#
# together with their respective p-values. We will use those quantities to report how many neurons carrying significative MI are in each cortical region in the specific recording session we loaded, as well as their level of relative MI.
#
# In this exercise we will be comparing levels of MI obtained from different neurons, each binned separately using equally spaced binning.
# - Are absolute values of MI comparable betweenn different neurons in this case?
# - How can we normalize the values of MI?
# + id="zyg2_w7MyEJj" executionInfo={"status": "error", "timestamp": 1639636041854, "user_tz": -60, "elapsed": 335, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhT8N_rvNzzgK5762VMS_vSoimcJLfzYid4nPtIWg=s64", "userId": "00165796825529292911"}} colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="059076c7-440f-476b-b451-00f10d3d2633"
n_bins = 4
stim_info_vis = np.empty(n_vis_area_neurons,)
choice_info_mos = np.empty(n_mos_area_neurons,)
stim_info_mos = np.empty(n_mos_area_neurons,)
entr_resp_vis = np.empty(n_vis_area_neurons,)
entr_resp_mos = np.empty(n_mos_area_neurons,)
p_val_stim_info_vis = np.empty(n_vis_area_neurons,)
p_val_choice_info_mos = np.empty(n_mos_area_neurons,)
p_val_stim_info_mos = np.empty(n_mos_area_neurons,)
# calculate stimulus and choice entropies
entr_stim = entropy(contrast_controlateral)
entr_choice = entropy(choice)
#############################################################################
# Students: Fill in missing code (...) and comment or remove the next line
raise NotImplementedError("Function not yet implemented")
#############################################################################
# calculate MI between neural response and contrast difference in visual cortex
for n in range(n_vis_area_neurons):
responses_vis = spikes_vis[n,:]
# binned_responses_vis = ( ... )
# stim_info_vis[n], p_val_stim_info_vis[n] = ( ... )
# entr_resp_vis[n] = ( ... )
# calculate MI between neural response and contrast difference in motor cortex
for n in range(n_mos_area_neurons):
responses_mos = spikes_mos[n,:]
# binned_responses_mos = ( ... )
# stim_info_mos[n], p_val_stim_info_mos[n] = ( ... )
# entr_resp_mos[n] = ( ... )
# calculate MI between neural response and choice in motor cortex
for n in range(n_mos_area_neurons):
responses_mos = spikes_mos[n,:]
# binned_responses_mos = ( ... )
# choice_info_mos[n], p_val_choice_info_mos[n] = ( ... )
# entr_resp_mos[n] = ( ... )
# + colab={"base_uri": "https://localhost:8080/", "height": 480} id="u3qvcQIlXwDk" executionInfo={"status": "ok", "timestamp": 1639634210405, "user_tz": -60, "elapsed": 645, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/<KEY>", "userId": "00165796825529292911"}} outputId="d6fadb83-30b3-4c4f-a81e-f3a6c19b678e" cellView="form"
#@title Plot results
# filter only significant neurons
sig_vis_neurons = stim_info_vis[p_val_stim_info_vis <= 0.05]/entr_resp_vis[p_val_stim_info_vis <= 0.05]
sig_mos_neurons_choice = choice_info_mos[p_val_choice_info_mos <= 0.05]/entr_resp_mos[p_val_choice_info_mos <= 0.05]
sig_mos_neurons_stim = stim_info_mos[p_val_stim_info_mos <= 0.05]/entr_resp_mos[p_val_stim_info_mos <= 0.05]
sig_vis_neurons = sig_vis_neurons[~np.isnan(sig_vis_neurons)]
sig_mos_neurons_choice = sig_mos_neurons_choice[~np.isnan(sig_mos_neurons_choice)]
sig_mos_neurons_stim = sig_mos_neurons_stim[~np.isnan(sig_mos_neurons_stim)]
fig, ax = plt.subplots()
for i,arr in enumerate([sig_vis_neurons, sig_mos_neurons_choice, sig_mos_neurons_stim]):
ax.violinplot(dataset=arr,positions=[i],showmedians=True)
ax.set_xticks([0, 1, 2])
ax.set_xticklabels(["MI(Stim;Resp) VISa", "MI(Choice;Resp) MOs", "MI(Stim;Resp) MOs"])
plt.ylabel("Relative MI [AU]")
print("Percentage of neurons informative about stimulus in VISa:", len(sig_vis_neurons)/n_vis_area_neurons*100)
print("Percentage of neurons informative about choice in MOs:", len(sig_mos_neurons_choice)/n_mos_area_neurons*100)
print("Percentage of neurons informative about stimulus in MOs:", len(sig_mos_neurons_stim)/n_mos_area_neurons*100)
# + id="iMdASRfM8W2p"
| MI_tutorial_student.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Basic Probability
# -
import mxnet as mx
from mxnet import nd
# %matplotlib inline
from matplotlib import pyplot as plt
from IPython import display
display.set_matplotlib_formats('svg')
# Let's define a discrete distribution over 6 outcomes and sample from it.
probabilities = nd.ones(6) / 6
nd.random.multinomial(probabilities)
# + [markdown] slideshow={"slide_type": "slide"}
# Let's draw from it multiple times.
# -
print(nd.random.multinomial(probabilities, shape=(10)))
print(nd.random.multinomial(probabilities, shape=(5,10)))
# + [markdown] slideshow={"slide_type": "slide"}
# Let's see what happens for 1000 samples.
# -
rolls = nd.random.multinomial(probabilities, shape=(1000))
counts = nd.zeros((6,1000))
totals = nd.zeros(6)
for i, roll in enumerate(rolls):
totals[int(roll.asscalar())] += 1
counts[:, i] = totals
# To start, we can inspect the final tally at the end of $1000$ rolls.
totals / 1000
# + [markdown] slideshow={"slide_type": "slide"}
# Let's look at the counts.
# -
counts
# + [markdown] slideshow={"slide_type": "slide"}
# Normalizing by the number of tosses, we get:
# -
x = nd.arange(1000).reshape((1,1000)) + 1
estimates = counts / x
print(estimates[:,0])
print(estimates[:,1])
print(estimates[:,100])
# + slideshow={"slide_type": "slide"}
plt.figure(figsize=(8, 6))
for i in range(6):
plt.plot(estimates[i, :].asnumpy(), label=("P(die=" + str(i) +")"))
plt.axhline(y=0.16666, color='black', linestyle='dashed')
plt.legend()
plt.show()
| slides/1_24/probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="qR2zERyblg4v" colab_type="text"
# # Download data and have a quick look
# + id="eCEweBqDQlnO" colab_type="code" outputId="fb5ad7c1-e1a6-48a2-bfe3-3846d8be8812" colab={"base_uri": "https://localhost:8080/", "height": 104}
# !unzip data.zip
# + id="zJ0G660IQ_T4" colab_type="code" outputId="ab310428-dd6b-4daf-aefe-427cfe7a9cc5" colab={"base_uri": "https://localhost:8080/", "height": 233}
import pandas as pd
from pathlib import Path
DATA_PATH = Path('/content/Dataset')
train_data = pd.read_csv(DATA_PATH/'Train.csv', index_col=0, infer_datetime_format=True, converters={'DATE':pd.to_datetime})
train_data.head()
# + [markdown] id="HUas52IG7MHD" colab_type="text"
# Since there are no named features we cannot do some domain specific feature engineering.
# + id="2TOoufGeRQoM" colab_type="code" outputId="4bbda672-2a58-4e3a-f527-ccabfa272f74" colab={"base_uri": "https://localhost:8080/", "height": 433}
train_data.info()
# + [markdown] id="RaSoIJIq7aBC" colab_type="text"
# There are some null or nan values in `X_12`
# + [markdown] id="K6Xm3XIVlmJT" colab_type="text"
# # Correlation of variables
# + id="2aRadIQLS5Rg" colab_type="code" outputId="998b9295-890b-4d08-ae15-4094fcf9b03b" colab={"base_uri": "https://localhost:8080/", "height": 336}
corr = train_data.corr()
corr.style.background_gradient(cmap='coolwarm')
# + [markdown] id="1sUkwjhx6bzM" colab_type="text"
# The highest **negative** correlation for the target variable is `X_10` and the higihest **positive** correlation is with `X_15`.
# + id="A8wS-sP2Y6sE" colab_type="code" outputId="a5951a7b-9af2-4254-9bd3-df2e6a5ddf47" colab={"base_uri": "https://localhost:8080/", "height": 386}
mul_off = train_data['MULTIPLE_OFFENSE']
corr = train_data.corrwith(mul_off)
corr.plot(kind='bar', title='Target Correlation')
# + [markdown] id="txCmUxxnIi5C" colab_type="text"
# # Target Distribution
# + id="3pWaVg51Ilap" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 280} outputId="d7427fd4-fc3b-4f62-d05e-c9ac5aa3cc91"
train_data['MULTIPLE_OFFENSE'].value_counts().plot(kind='bar')
# + [markdown] id="aHA4In5jJBcX" colab_type="text"
# We have quite a skewed distribution, maybe now we can have make an assumption regarding what the classes of `MULTIPLE_OFFENSE` may mean, the problem title was **Predict if the server will be hacked**, assuming some normal conditions
#
# - `1` must be the times when the server was not attacked and running smoothly
# - `0` must be the times when the server was attacked and malfunctioning(maybe)
# + [markdown] id="xMz10YOsHv4_" colab_type="text"
# To understand the feature interaction we can try to build a simple decision tree classifier and plot the decision tree to view interaction visually. We should also consider this problem as a time-series classification problem.
# + [markdown] id="xpCrJSGyKPlc" colab_type="text"
# # Model setup
# + id="ldcCfYRs5eao" colab_type="code" colab={}
# since we may need the model to work on data with proper sequence
train_data.sort_values(by='DATE', inplace=True)
# + id="-Cklee7o0w2w" colab_type="code" colab={}
import numpy as np
seed=0
np.random.seed(seed=seed)
# + id="ShMj64aFZ1GB" colab_type="code" colab={}
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
# form a pipeline with scaler and decision tree
pipe = Pipeline([
('median_imputer', SimpleImputer(strategy='median')),
('scaler', StandardScaler()),
('tree', DecisionTreeClassifier(max_depth=4, random_state=seed))
])
# + [markdown] id="mf5ZUox5KRB2" colab_type="text"
# # Data Preprocessing
# + id="GVnzgU2KKTHK" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
# We will try not to use dates now
x_train, x_test, y_train, y_test = train_test_split(train_data.drop(['DATE', 'MULTIPLE_OFFENSE'], axis='columns'),
train_data['MULTIPLE_OFFENSE'],
random_state=seed,
# Stratified split
stratify=train_data['MULTIPLE_OFFENSE'].values)
# + id="QkKG57iJLiS2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4a232769-efbf-450d-e3fb-e5a94250d510"
x_train.shape, y_train.shape, x_test.shape, y_test.shape
# + [markdown] id="M5SFN2fqMb_j" colab_type="text"
# # Model Training
# + id="ZNATPyRaLrCG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="cd4cd47c-dab2-4491-81c2-d01a4a816644"
pipe.fit(x_train, y_train)
# + [markdown] id="RXR3CzLrMfhn" colab_type="text"
# # Model Evaluation
# + id="7DbkbMg-2V9F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9e332c72-9eed-4d76-fd3d-c347c066147e"
pipe.score(x_test, y_test)
# + [markdown] id="OOkLpCzMPDfu" colab_type="text"
# This might be misleading because our target is heavily skewed towards `1`
# + id="BUAkst1dTVw1" colab_type="code" colab={}
y_pred = pipe.predict(x_test)
# + id="ic3CoXwQS66P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 296} outputId="a00d10c5-e2b4-4b2e-afe9-7129a64a7e50"
from sklearn.metrics import plot_confusion_matrix
plot_confusion_matrix(pipe, x_test, y_test)
# + id="2iMEhMUU1IiN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="751984b3-a1c9-4a8b-9719-600c507234fa"
tn, fp, fn, tp = confusion_matrix(y_test, y_pred).ravel()
print(f'TP = {tp} \nFP = {fp}\nTN = {tn}\nFN = {fn}')
# + id="cxZ3TMjOUSrk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 173} outputId="60c1028d-e593-4636-ee7a-3a8fb358d916"
from sklearn.metrics import classification_report
print(classification_report(y_test, y_pred, target_names = ['class 0', 'class 1']))
# + [markdown] id="RuWVefc1VMiW" colab_type="text"
# Not bad at all, but based on our assumption if we assume that the classes represent
#
# - `1` must be the times when the server was not attacked and running smoothly
# - `0` must be the times when the server was attacked and malfunctioning(maybe)
#
# The model will be more useful if it could predict the malfunction correctly most of the times because it should be able to detect the fault.
#
# + [markdown] id="HaZdbaCpyrfh" colab_type="text"
# The `FP (False Positives)` *(ie)* the number of cases where the server was malfunctioning but the model predicted as **not** is **`78`**, not bad actually.
# + [markdown] id="fzVE8tBWyp3I" colab_type="text"
#
# If we look at the value of `recall` for `class 0` it is `0.71` which is okay for a simple decision tree classifier, with no feature engineering.
# + [markdown] id="B11MW0XqP6zo" colab_type="text"
# Let's try to plot the tree
# + id="4d9QLVH0QqY6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 777} outputId="32c3c49e-7bdc-4016-dfd4-bcdea474a4bc"
from sklearn import tree
import graphviz
# tree.plot_tree(pipe['tree'])
dot_data = tree.export_graphviz(pipe['tree'], out_file=None,
feature_names=[f'X_{i}' for i in range(1, 16)],
class_names=['0', '1'],
filled=True, rounded=True,
special_characters=True)
graph = graphviz.Source(dot_data)
graph
# + id="DEAYSL3vYtyf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e84cbc69-2b2b-462c-d926-3a9557f22f11"
graph.render("server_mal")
# + [markdown] id="TLfU8DFDQw41" colab_type="text"
# According the decision tree the most important features are `X_10`, `X_11`, `X_12` and `X_15`.
# + [markdown] id="dQ-lD0b26Yka" colab_type="text"
# ## Prediction on test data
# + id="Qs-evUC06dPp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 233} outputId="470a363d-3689-4fc7-dae2-aa756fb3a94c"
test_data = pd.read_csv(DATA_PATH/'Test.csv', index_col=0)
test_data.head()
# + [markdown] id="wZK3HEOB6pGU" colab_type="text"
# We used only the logging parameters of the server to model the decision tree so we don't need other variables.
# + id="pifPhcoG68IX" colab_type="code" colab={}
test_data.drop('DATE', axis='columns', inplace=True)
# + id="nITFEZjn7D_r" colab_type="code" colab={}
y_pred = pipe.predict(test_data)
# + id="MTLUmNyZ7LWe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="5c1bba85-1248-4d27-e0af-601cb0b49e61"
sub_df = pd.DataFrame(
{'INCIDENT_ID':test_data.index,
'MULTIPLE_OFFENSE':y_pred},
)
sub_df.head()
# + id="k7cI_Q_x8JsN" colab_type="code" colab={}
sub_df.to_csv('submission_df.csv', index=False)
# + [markdown] id="qu1PhPGV8eKI" colab_type="text"
# This submission scored a decent `85` (recall) on the test set, but it can be improved further.
# + [markdown] id="FzX-yGYMYYVe" colab_type="text"
# ## Model Persistence
# + id="pDrWaFv-Y03Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="931f9d8e-9637-4250-919c-e0c2570f7a99"
# !pip list | egrep 'scikit-learn|pandas'
# + id="yWZxhQaBS3re" colab_type="code" colab={}
import joblib
with open('desc_pipeline.joblib', 'wb') as f:joblib.dump(pipe, f)
# + id="CUDyNxkK0RjI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="d839facd-695e-48b3-efbc-95954295e169"
# !zip eda_model_interp.zip server_mal.pdf desc_pipeline.joblib
# + id="owXgyp6-9XPz" colab_type="code" colab={}
| EDA/Novarits_data_competition_eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Naming Conventions and Restrictions
#
# - repo names:
# - lower case letters and non-duplicated dashes (-) not at either end
# - environment and pipeline step names:
# - lower case letters and non-duplicated underscores (_) not at either end
# - prefix names:
# - lower case letters, at least 2
#
# ### Metadata
#
# - `[a-b]*_table` table names based on singular form of entity
# - see all in dogshow standard
| notebooks/doc-002-conventions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib widget
import os
import sys
sys.path.insert(0, os.getenv('HOME')+'/pycode/MscThesis/')
import pandas as pd
from amftrack.util import get_dates_datetime, get_dirname, get_plate_number, get_postion_number
import ast
from amftrack.plotutil import plot_t_tp1
from scipy import sparse
from datetime import datetime
from amftrack.pipeline.functions.node_id import orient
import pickle
import scipy.io as sio
from pymatreader import read_mat
from matplotlib import colors
import cv2
import imageio
import matplotlib.pyplot as plt
import numpy as np
from skimage.filters import frangi
from skimage import filters
from random import choice
import scipy.sparse
import os
from amftrack.pipeline.functions.extract_graph import from_sparse_to_graph, generate_nx_graph, sparse_to_doc
from skimage.feature import hessian_matrix_det
from amftrack.pipeline.functions.experiment_class_surf import Experiment
from amftrack.pipeline.paths.directory import run_parallel, find_state, directory_scratch, directory_project, run_parallel_stitch
# -
# ***Chose plate number and directory of interest where the folders with images are***
plate_number = 731
plate = get_postion_number(plate_number)
plate = 15
print("Plate #",plate_number)
print("Position ",plate)
directory = directory_scratch
# directory = directory_project
listdir = os.listdir(directory)
list_dir_interest = [name for name in listdir if name.split('_')[-1]==f'Plate{0 if plate<10 else ""}{plate}']
dates_datetime = get_dates_datetime(directory,plate)
listLength = len(list_dir_interest)
#get_dirname(dates_datetime[0], plate)
begin = 0
end = listLength - 1 #begin + 30
print('begin =', dates_datetime[begin], '\n end =', dates_datetime[end])
# ***Stitching***
num_parallel = 4
time = '20:00'
run_parallel_stitch(plate,directory,begin,end,num_parallel,time)
plate_number = get_plate_number(plate,dates_datetime[0])
print(0,plate_number)
for i in range(len(list_dir_interest)):
new_plate_number = get_plate_number(plate,dates_datetime[i])
if plate_number!=new_plate_number:
plate_number=new_plate_number
print(i,plate_number)
# ***Chose timestep to begin with and folder to end with***
# + jupyter={"outputs_hidden": true}
find_state(plate, begin, end, directory, True)
# -
# ***Skeletonization***
# - Only 4 skeletonization processes can be run in parallel on one node
num_parallel = 8
time = '3:00:00'
low = 30
high = 80
extend = 30
args=[plate, low, high, extend,directory]
run_parallel('extract_skel.py',args,begin,end,num_parallel,time,'skeletonization')
# ***Compress raw image***
num_parallel = 4
time = '2:00'
args = [plate, directory]
run_parallel('compress_image.py',args, begin, end, num_parallel, time, 'compress')
# ***Check Skeletonization***
# - The first cell loads the skeleton and the compressed raw image
# - The second cell shows the overlap of the skeleton and the raw image
start = begin+40
finish = start +1
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen=dates_datetime[start:finish+1]
dates = dates_datetime_chosen
skels = []
ims = []
kernel = np.ones((5,5),np.uint8)
itera = 1
for date in dates:
directory_name = get_dirname(date,plate)
path_snap=directory+directory_name
skel_info = read_mat(path_snap+'/Analysis/skeleton_compressed.mat')
skel = skel_info['skeleton']
skels.append(cv2.dilate(skel.astype(np.uint8),kernel,iterations = itera))
im = read_mat(path_snap+'/Analysis/raw_image.mat')['raw']
ims.append(im)
# - Chose start and finish to display the overlap of the skeleton and the raw image : no more than 10 at a time or jupyter will crash.
# - For display purpose, everything is compressed so connectivity may be lost
# - colors are just a feature of compression
plt.close('all')
start = 0
finish = start + 1
for i in range(start,finish):
plot_t_tp1([], [], None, None, skels[i], ims[i])
# ***Check specific image***
# - If something wrong is noticed in one of the skeletons one can chose to look closer at one of the images and the skeletonization process
# - chose ***i*** equal to the timestep where something wrong has been noticed
# +
#chose i equal to the timestep where something wrong has been noticed
i = 200
dates_datetime = get_dates_datetime(directory,plate)
dates = dates_datetime
date =dates [i]
directory_name = get_dirname(date,plate)
path_snap=directory+directory_name
path_tile=path_snap+'/Img/TileConfiguration.txt.registered'
try:
tileconfig = pd.read_table(path_tile,sep=';',skiprows=4,header=None,converters={2 : ast.literal_eval},skipinitialspace=True)
except:
print('error_name')
path_tile=path_snap+'/Img/TileConfiguration.registered.txt'
tileconfig = pd.read_table(path_tile,sep=';',skiprows=4,header=None,converters={2 : ast.literal_eval},skipinitialspace=True)
xs =[c[0] for c in tileconfig[2]]
ys =[c[1] for c in tileconfig[2]]
dim = (int(np.max(ys)-np.min(ys))+4096,int(np.max(xs)-np.min(xs))+4096)
ims = []
for name in tileconfig[0]:
imname = '/Img/'+name.split('/')[-1]
ims.append(imageio.imread(directory+directory_name+imname))
# -
# - Chose a x,y position where you want to see how the skeletonization process went (x is the scale on the left on the images and y is the bottom scale)
# - You can chose to display different part of the filter par commenting/uncommenting
# + jupyter={"outputs_hidden": true, "source_hidden": true}
plt.close('all')
#chose a spot where to look closer at
linex = 4000
liney = 7000
low = 30
high = 80
extend = 30
dist = extend
shape = (3000,4096)
linex *= 5
liney *= 5
for index,im in enumerate(ims):
boundaries = int(tileconfig[2][index][0]-np.min(xs)),int(tileconfig[2][index][1]-np.min(ys))
if boundaries[1]<=linex< boundaries[1]+shape[0] and boundaries[0]<=liney< boundaries[0]+shape[1]:
print(index)
im_cropped = im
# im_blurred =cv2.GaussianBlur(im_cropped, (201, 201),50)
im_blurred =cv2.blur(im_cropped, (200, 200))
im_back_rem = (im_cropped+1)/(im_blurred+1)*120
im_back_rem[im_back_rem>=130]=130
# im_back_rem = im_cropped*1.0
# # im_back_rem = cv2.normalize(im_back_rem, None, 0, 255, cv2.NORM_MINMAX)
frangised = frangi(im_back_rem,sigmas=range(1,20,4))*255
# # frangised = cv2.normalize(frangised, None, 0, 255, cv2.NORM_MINMAX)
hessian = hessian_matrix_det(im_back_rem,sigma = 20)
blur_hessian = cv2.blur(abs(hessian), (20, 20))
# transformed = (frangised+cv2.normalize(blur_hessian, None, 0, 255, cv2.NORM_MINMAX)-im_back_rem+120)*(im_blurred>=35)
# transformed = (frangised+cv2.normalize(abs(hessian), None, 0, 255, cv2.NORM_MINMAX)-im_back_rem+120)*(im_blurred>=35)
transformed = (frangised-im_back_rem+120)*(im_blurred>=35)
lowt = (transformed > low).astype(int)
hight = (transformed > high).astype(int)
hyst = filters.apply_hysteresis_threshold(transformed, low, high)
kernel = np.ones((3,3),np.uint8)
dilation = cv2.dilate(hyst.astype(np.uint8) * 255,kernel,iterations = 1)
for i in range(3):
dilation=cv2.erode(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
dilation = cv2.dilate(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
dilated = dilation>0
nb_components, output, stats, centroids = cv2.connectedComponentsWithStats(dilated.astype(np.uint8), connectivity=8)
#connectedComponentswithStats yields every seperated component with information on each of them, such as size
#the following part is just taking out the background which is also considered a component, but most of the time we don't want that.
sizes = stats[1:, -1]; nb_components = nb_components - 1
# minimum size of particles we want to keep (number of pixels)
#here, it's a fixed value, but you can set it as you want, eg the mean of the sizes or whatever
min_size = 4000
#your answer image
img2 = np.zeros((dilated.shape))
#for every component in the image, you keep it only if it's above min_size
for i in range(0, nb_components):
if sizes[i] >= min_size:
img2[output == i + 1] = 1
skeletonized = cv2.ximgproc.thinning(np.array(255*img2,dtype=np.uint8))
nx_g = generate_nx_graph(from_sparse_to_graph(scipy.sparse.dok_matrix(skeletonized)))
g,pos= nx_g
tips = [node for node in g.nodes if g.degree(node)==1]
dilated_bis = np.copy(img2)
for tip in tips:
branch = np.array(orient(g.get_edge_data(*list(g.edges(tip))[0])['pixel_list'],pos[tip]))
orientation = branch[0]-branch[min(branch.shape[0]-1,20)]
orientation = orientation/(np.linalg.norm(orientation))
window = 20
x,y = pos[tip][0],pos[tip][1]
if x-window>=0 and x+window< dilated.shape[0] and y-window>=0 and y+window< dilated.shape[1]:
shape_tip = dilated[x-window:x+window,y-window:y+window]
# dist = 20
for i in range(dist):
pixel = (pos[tip]+orientation*i).astype(int)
xp,yp = pixel[0],pixel[1]
if xp-window>=0 and xp+window< dilated.shape[0] and yp-window>=0 and yp+window< dilated.shape[1]:
dilated_bis[xp-window:xp+window,yp-window:yp+window]+=shape_tip
dilation = cv2.dilate(dilated_bis.astype(np.uint8) * 255,kernel,iterations = 1)
for i in range(3):
dilation=cv2.erode(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
dilation = cv2.dilate(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
skeletonized = cv2.ximgproc.thinning(np.array(255*(dilation>0),dtype=np.uint8))
print('Raw image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(-im,cmap = 'Greys')
print('Raw image with background removed')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(-im_back_rem,cmap = 'Greys')
print('frangised image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(frangised,cmap = 'Greys')
print('final transformed image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(transformed,cmap = 'Greys')
print('threhsolded image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(dilated>0,cmap = 'Greys')
print('threhsolded image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(img2>0,cmap = 'Greys')
print('extended tips')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(dilated_bis>0,cmap = 'Greys')
print('final_skel')
# fig=plt.figure(figsize=(10,9))
# ax = fig.add_subplot(111)
# ax.imshow(cv2.normalize(abs(hessian), None, 0, 255, cv2.NORM_MINMAX)-255*dilated)
plot_t_tp1([],[],None,None,skeletonized,im_back_rem)
# + jupyter={"outputs_hidden": true, "source_hidden": true}
plt.close('all')
#chose a spot where to look closer at
linex = 3300
liney = 3800
shape = (3000,4096)
linex *= 5
liney *= 5
for index,im in enumerate(ims):
boundaries = int(tileconfig[2][index][0]-np.min(xs)),int(tileconfig[2][index][1]-np.min(ys))
if boundaries[1]<=linex< boundaries[1]+shape[0] and boundaries[0]<=liney< boundaries[0]+shape[1]:
print(index)
im_cropped = im
im_blurred =cv2.blur(im_cropped, (200, 200))
im_back_rem = (im_cropped+1)/(im_blurred+1)*120
im_back_rem[im_back_rem>=130]=130
# # im_back_rem = im_cropped*1.0
# # # im_back_rem = cv2.normalize(im_back_rem, None, 0, 255, cv2.NORM_MINMAX)
frangised = frangi(im_back_rem,sigmas=range(1,20,4))*255
# # frangised = cv2.normalize(frangised, None, 0, 255, cv2.NORM_MINMAX)
hessian = hessian_matrix_det(im_back_rem,sigma = 20)
blur_hessian = cv2.blur(abs(hessian), (20, 20))
# transformed = (frangised+cv2.normalize(blur_hessian, None, 0, 255, cv2.NORM_MINMAX)-im_back_rem+120)*(im_blurred>=35)
# transformed = (frangised+cv2.normalize(abs(hessian), None, 0, 255, cv2.NORM_MINMAX)-im_back_rem+120)*(im_blurred>=35)
transformed = (frangised-im_back_rem+120)*(im_blurred>=35)
low = 40
high = 80
lowt = (transformed > low).astype(int)
hight = (transformed > high).astype(int)
hyst = filters.apply_hysteresis_threshold(transformed, low, high)
kernel = np.ones((3,3),np.uint8)
dilation = cv2.dilate(hyst.astype(np.uint8) * 255,kernel,iterations = 1)
for i in range(3):
dilation=cv2.erode(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
dilation = cv2.dilate(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
dilated = dilation>0
nb_components, output, stats, centroids = cv2.connectedComponentsWithStats(dilated.astype(np.uint8), connectivity=8)
#connectedComponentswithStats yields every seperated component with information on each of them, such as size
#the following part is just taking out the background which is also considered a component, but most of the time we don't want that.
sizes = stats[1:, -1]; nb_components = nb_components - 1
# minimum size of particles we want to keep (number of pixels)
#here, it's a fixed value, but you can set it as you want, eg the mean of the sizes or whatever
min_size = 4000
#your answer image
img2 = np.zeros((dilated.shape))
#for every component in the image, you keep it only if it's above min_size
for i in range(0, nb_components):
if sizes[i] >= min_size:
img2[output == i + 1] = 1
skeletonized = cv2.ximgproc.thinning(np.array(255*img2,dtype=np.uint8))
nx_g = generate_nx_graph(from_sparse_to_graph(scipy.sparse.dok_matrix(skeletonized)))
g,pos= nx_g
tips = [node for node in g.nodes if g.degree(node)==1]
dilated_bis = np.copy(img2)
for tip in tips:
branch = np.array(orient(g.get_edge_data(*list(g.edges(tip))[0])['pixel_list'],pos[tip]))
orientation = branch[0]-branch[min(branch.shape[0]-1,20)]
orientation = orientation/(np.linalg.norm(orientation))
window = 20
x,y = pos[tip][0],pos[tip][1]
if x-window>=0 and x+window< dilated.shape[0] and y-window>=0 and y+window< dilated.shape[1]:
shape_tip = dilated[x-window:x+window,y-window:y+window]
dist = 20
for i in range(dist):
pixel = (pos[tip]+orientation*i).astype(int)
xp,yp = pixel[0],pixel[1]
if xp-window>=0 and xp+window< dilated.shape[0] and yp-window>=0 and yp+window< dilated.shape[1]:
dilated_bis[xp-window:xp+window,yp-window:yp+window]+=shape_tip
dilation = cv2.dilate(dilated_bis.astype(np.uint8) * 255,kernel,iterations = 1)
for i in range(3):
dilation=cv2.erode(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
dilation = cv2.dilate(dilation.astype(np.uint8) * 255,kernel,iterations = 1)
skeletonized = cv2.ximgproc.thinning(np.array(255*(dilation>0),dtype=np.uint8))
print('Raw image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(im)
print('Raw image with background removed')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(im_back_rem)
print('frangised image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(frangised)
print('final transformed image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(transformed)
print('threhsolded image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(dilated>0)
print('threhsolded image')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(img2>0)
print('extended tips')
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(dilated_bis)
print('final_skel')
# fig=plt.figure(figsize=(10,9))
# ax = fig.add_subplot(111)
# ax.imshow(cv2.normalize(abs(hessian), None, 0, 255, cv2.NORM_MINMAX)-255*dilated)
plot_t_tp1([],[],None,None,skeletonized,im_back_rem)
# -
# ***Mask baits and border of the petri dish***
num_parallel = 4
time = '5:00'
thresh = 50
args=[plate,thresh, directory]
run_parallel('mask_skel.py',args,begin,end,num_parallel,time,'mask')
# ***Check Masking***
start = begin +40
finish = start +1
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen=dates_datetime[start:finish+1]
dates = dates_datetime_chosen
skels = []
ims = []
masks = []
kernel = np.ones((5,5),np.uint8)
itera = 1
for date in dates:
directory_name = get_dirname(date,plate)
path_snap = directory + directory_name
skel_info = read_mat(path_snap+'/Analysis/skeleton_masked_compressed.mat')
skel = skel_info['skeleton']
skels.append(cv2.dilate(skel.astype(np.uint8),kernel,iterations = itera))
mask_info = read_mat(path_snap+'/Analysis/mask.mat')
mask = mask_info['mask']
masks.append(mask)
im = read_mat(path_snap+'/Analysis/raw_image.mat')['raw']
ims.append(im)
plt.close('all')
start = 0
finish = start + 1
for i in range(start,finish):
plot_t_tp1([], [], None, None,masks[i], ims[i])
# ***Prune Graph***
num_parallel = 20
time = '50:00'
threshold = 1
args = [plate,threshold, directory]
run_parallel('prune_skel.py',args,begin,end,num_parallel,time,'prune_graph')
# ***Check Pruned Graphs***
start = begin + 40
finish = start +2
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen=dates_datetime[start:finish+1]
dates = dates_datetime_chosen
skels = []
ims = []
kernel = np.ones((5,5),np.uint8)
itera = 1
for date in dates:
directory_name = get_dirname(date,plate)
path_snap=directory+directory_name
skel_info = read_mat(path_snap+'/Analysis/skeleton_pruned_compressed.mat')
skel = skel_info['skeleton']
skels.append(cv2.dilate(skel.astype(np.uint8),kernel,iterations = itera))
im = read_mat(path_snap+'/Analysis/raw_image.mat')['raw']
ims.append(im)
plt.close('all')
start = 0
finish = start +2
for i in range(start,finish):
plot_t_tp1([],[],None,None,skels[i],ims[i])
# + jupyter={"source_hidden": true}
plt.close('all')
kernel = np.ones((5,5),np.uint8)
for i in range(len(compressed)-1):
plot_t_tp1([],[],None,None,cv2.dilate(compressed[i].astype(np.uint8),kernel,iterations = 2),cv2.dilate(compressed[i+1].astype(np.uint8),kernel,iterations = 2))
# -
# ***Realign***
num_parallel =16
time = '1:00:00'
thresh = 10000
args=[plate, thresh, directory]
run_parallel('final_alignment.py',args,begin,end,num_parallel,time,'realign')
# ***Check Alignment***
plt.close('all')
start = 18
finish = start +2
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen=dates_datetime[start:finish+1]
dates = dates_datetime_chosen
dilateds=[]
skels = []
skel_docs = []
Rs=[]
ts=[]
for date in dates[1:]:
directory_name = get_dirname(date,plate)
path_snap=directory+directory_name
skel_info = read_mat(path_snap+'/Analysis/skeleton.mat')
skel = skel_info['skeleton']
skels.append(skel)
skel_doc = sparse_to_doc(skel)
skel_docs.append(skel_doc)
transform = sio.loadmat(path_snap+'/Analysis/transform.mat')
R,t = transform['R'],transform['t']
Rs.append(R)
ts.append(t)
# start = 0
# for j in range(start,start + 5):
# print(dates[j],j+begin)
# skeleton1,skeleton2 = skel_docs[j],skel_docs[j+1]
# R,t = Rs[j],ts[j]
# skelet_pos = np.array(list(skeleton1.keys()))
# samples = np.random.choice(skelet_pos.shape[0],20000)
# X = np.transpose(skelet_pos[samples,:])
# skelet_pos = np.array(list(skeleton2.keys()))
# samples = np.random.choice(skelet_pos.shape[0],20000)
# Y = np.transpose(skelet_pos[samples,:])
# fig=plt.figure(figsize=(10,9))
# ax = fig.add_subplot(111)
# Yrep=np.transpose(np.transpose(np.dot(R,X))+t)
# ax.scatter(np.transpose(Yrep)[:,0],np.transpose(Yrep)[:,1])
# ax.scatter(np.transpose(Y)[:,0],np.transpose(Y)[:,1])
# ***Create realigned Skeleton***
num_parallel = 12
time = '1:00:00'
args = [plate, begin, end, directory]
run_parallel('realign.py', args, begin, end, num_parallel, time, 'create_realign')
# ***Check Fine Alignment***
start = begin+40
finish = start +2
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen=dates_datetime[start:finish+1]
dates = dates_datetime_chosen
skels = []
ims = []
kernel = np.ones((5, 5), np.uint8)
itera = 1
for date in dates:
directory_name = get_dirname(date,plate)
path_snap = directory + directory_name
skel_info = read_mat(path_snap+'/Analysis/skeleton_realigned_compressed.mat')
skel = skel_info['skeleton']
# skels.append(skel)
skels.append(cv2.dilate(skel.astype(np.uint8),kernel,iterations = itera))
im = read_mat(path_snap+'/Analysis/raw_image.mat')['raw']
ims.append(im)
plt.close('all')
start = 0
finish = start + 1
for i in range(start, finish):
plot_t_tp1([],[],None,None,skels[i],skels[i+1])
# + jupyter={"source_hidden": true}
plt.close('all')
directory = "/scratch/shared/mrozemul/Fiji.app/"
listdir=os.listdir(directory)
list_dir_interest=[name for name in listdir if name.split('_')[-1]==f'Plate{0 if plate<10 else ""}{plate}']
ss=[name.split('_')[0] for name in list_dir_interest]
ff=[name.split('_')[1] for name in list_dir_interest]
dates_datetime=[datetime(year=int(ss[i][:4]),month=int(ss[i][4:6]),day=int(ss[i][6:8]),hour=int(ff[i][0:2]),minute=int(ff[i][2:4])) for i in range(len(list_dir_interest))]
dates_datetime.sort()
begin = 0
end = 20
dates_datetime_chosen=dates_datetime[begin:end]
dates = [f'{0 if date.month<10 else ""}{date.month}{0 if date.day<10 else ""}{date.day}_{0 if date.hour<10 else ""}{date.hour}{0 if date.minute<10 else ""}{date.minute}' for date in dates_datetime_chosen]
zone = (13000,13000+5000+3000,20000,20000+5000+4096)
skels_aligned = []
for i,date in enumerate(dates):
directory_name=f'2020{dates[i]}_Plate{0 if plate<10 else ""}{plate}'
path_snap='/scratch/shared/mrozemul/Fiji.app/'+directory_name
skels_aligned.append(sio.loadmat(path_snap+'/Analysis/skeleton_realigned.mat')['skeleton'])
for i in range(11,13):
plot_t_tp1([],[],None,None,skels_aligned[i][zone[0]:zone[1],zone[2]:zone[3]].todense(),skels_aligned[i+1][zone[0]:zone[1],zone[2]:zone[3]].todense())
# + jupyter={"source_hidden": true}
plt.close('all')
zone = (6000,13000,12000,22000)
fig=plt.figure(figsize=(10,9))
ax = fig.add_subplot(111)
ax.imshow(skels_aligned[11][zone[0]:zone[1],zone[2]:zone[3]].todense())
# -
# ***Create graphs***
num_parallel = 5
time = '30:00'
args=[plate, directory]
run_parallel('extract_nx_graph.py',args,begin,end,num_parallel,time,'extract_nx')
# ***Extract Width***
num_parallel = 16
time = '1:00:00'
args = [plate,directory]
run_parallel('extract_width.py',args,begin,end,num_parallel,time,'extract_width')
# ***Identify Nodes***
num_parallel = 1
time = '12:00:00'
args=[plate,begin,end, directory]
run_parallel('extract_nodes.py',args,0,0,num_parallel,time,'node_id')
# ***Check Node Id***
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen = dates_datetime[begin:end+1]
dates = dates_datetime_chosen
exp = Experiment(plate)
exp.load(dates)
plt.close('all')
t = 2
nodes = np.random.choice(exp.nx_graph[t].nodes, 100)
# exp.plot([t,t+1,t+2],[list(nodes)]*3)
exp.plot([t,t+1,t+2],[nodes]*3)
# ***Hyphae extraction***
num_parallel = 1
time = '2:00:00'
args = [plate, begin, end, directory]
run_parallel('hyphae_extraction.py',args,0,0,num_parallel,time,'hyphae')
# ***Check Hyphae***
dates_datetime = get_dates_datetime(directory,plate)
dates_datetime_chosen=dates_datetime[begin:end+1]
dates = dates_datetime
exp = pickle.load( open(f'{directory}Analysis_Plate{plate}_{dates[0]}_{dates[-1]}/experiment_{plate}.pick', "rb" ) )
hyph = choice(exp.hyphaes)
hyph.ts
plt.close('all')
hyph.end.show_source_image(hyph.ts[-1],hyph.ts[-1])
plt.close('all')
exp.plot([0,hyph.ts[-2],hyph.ts[-1]],[[hyph.root.label,hyph.end.label]]*3)
| amftrack/pipeline/control/Data_pipeline_Loreto.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.svm import SVR
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn import linear_model
output=pd.read_csv("submission.csv")
# +
training_data = pd.read_csv('train.csv')
test_data=pd.read_csv('test.csv')
X=training_data
X=X.drop(['Close'], axis = 1)
X=X.drop(['Date'], axis = 1)
y=training_data['Close']
m=len(y)
X_test=test_data
X_test=X_test.drop(['Close'], axis = 1)
X_test=X_test.drop(['Date'], axis = 1)
# +
# multi linear regression model
model_ols = linear_model.LinearRegression(normalize=True)
model_ols.fit(X,y)
predictedclose = pd.DataFrame(model_ols.predict(X), columns=['Predicted close']) # Create new dataframe of column'Predicted Price'
actualclose = pd.DataFrame(y, columns=['Close'])
actualclose = actualclose.reset_index(drop=True) # Drop the index so that we can concat it, to create new dataframe
df_actual_vs_predicted = pd.concat([actualclose,predictedclose],axis =1)
df_actual_vs_predicted.T
# +
#results of linear model
price = model_ols.predict(X_test)
print(price)
#to csv file
output=pd.read_csv("submission.csv")
output
output['Close'] = price
output.to_csv("multivariatewodate.csv",index=False)
# +
#ridge model
model_r = linear_model.Ridge(normalize= True, alpha= 0.001)
model_r.fit(X,y)
priceridge = model_r.predict(X_test)
print(priceridge)
#output to csv
output=pd.read_csv("submission.csv")
output
output['Close'] = priceridge
output
output.to_csv("ridgeresults.csv",index=False)
# +
# svr polynomial model
model_svr_regr = make_pipeline(StandardScaler(), SVR(kernel="poly", C=100, gamma="auto", degree=3, epsilon=0.1, coef0=1))
model_svr_regr.fit(X, y)
price_svr=model_svr_regr.predict(X_test)
print(price_svr)
output=pd.read_csv("submission.csv")
output
output['Close'] = price_svr
output
output.to_csv("svrresults.csv",index=False)
# -
# # parameter tuning
#
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn import linear_model
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report, confusion_matrix
# +
training_data1 = pd.read_csv('train.csv')
target_data1 = training_data1['Close']
training_data1 = training_data1.drop(['Close'],axis=1)
training_data1 = training_data1.drop(['Date'],axis=1)
print(training_data1)
# +
#print(target_data)
df_feat = training_data1
df_target = target_data1
print(df_feat.head(),"\n\n",df_target.head())
# +
from sklearn.model_selection import train_test_split
X1_train, X1_test, y1_train, y1_test = train_test_split(
df_feat, df_target,
test_size = 0.30, random_state = 101)
# -
scaler = StandardScaler()
X1_train = scaler.fit_transform(X1_train)
X1_test = scaler.transform(X1_test)
model = SVR()
model.fit(X1_train, y1_train)
#baseline without tuning
print("[INFO] evaluating...")
print("R2: {:.2f}".format(model.score(X1_test, y1_test)))
# +
from scipy.stats import loguniform
model = SVR(gamma='auto')
kernel = [ "poly"]
tolerance = loguniform(1e-6, 1e-3)
C = [1, 1.5, 2, 2.5, 3,7,7.5,8,8.5,9,9.5,10]
coef0=[0,1]
grid = dict(kernel=kernel, tol=tolerance, C=C,coef0=coef0)
# +
# initialize a cross-validation fold and perform a randomized-search
# to tune the hyperparameters
from sklearn.model_selection import RandomizedSearchCV
from sklearn.model_selection import RepeatedKFold
print("[INFO] grid searching over the hyperparameters...")
cvFold = RepeatedKFold(n_splits=10, n_repeats=3, random_state=1)
randomSearch = RandomizedSearchCV(estimator=model, n_jobs=-1,
cv=cvFold, param_distributions=grid,
scoring="neg_mean_squared_error")
searchResults = randomSearch.fit(X1_train, y1_train)
# extract the best model and evaluate it
print("[INFO] evaluating...")
bestModel = searchResults.best_estimator_
print("R2: {:.2f}".format(bestModel.score(X1_test, y1_test)))
# -
bestModel
# +
model_svr_regr_tune = make_pipeline(StandardScaler(), bestModel)
model_svr_regr_tune.fit(X, y)
price_svr=model_svr_regr_tune.predict(X_test)
print(price_svr)
output=pd.read_csv("submission.csv")
output
output['Close'] = price_svr
output
output.to_csv("svrresultsparameterstuned.csv",index=False)
# +
import plotly.express as px
import plotly.graph_objects as go
mesh_size = 100
margin = 0
df = pd.read_csv('train.csv')
X = df[['Open', 'Volume']]
y = df['Close']
X=X.iloc[:10,:]
y=y.iloc[:10,]
print(X,y)
# Condition the model on sepal width and length, predict the petal width
model = bestModel
model.fit(X, y)
# Create a mesh grid on which we will run our model
x_min, x_max = X.Open.min() - margin, X.Open.max() + margin
y_min, y_max = X.Volume.min() - margin, X.Volume.max() + margin
xrange = np.arange(x_min, x_max, mesh_size)
yrange = np.arange(y_min, y_max, mesh_size)
xx, yy = np.meshgrid(xrange, yrange)
# Run model
pred = model.predict(np.c_[xx.ravel(), yy.ravel()])
pred = pred.reshape(xx.shape)
# Generate the plot
fig = px.scatter_3d(df, x='Open', y='Volume', z='Close')
fig.update_traces(marker=dict(size=5))
fig.add_traces(go.Surface(x=xrange, y=yrange, z=pred, name='pred_surface'))
fig.show()
# -
| input/.ipynb_checkpoints/stock_final_regression -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="UiNxsd4_q9wq" colab_type="text"
# ### What-If Tool toxicity text model comparison
#
# Copyright 2019 Google LLC.
# SPDX-License-Identifier: Apache-2.0
#
# This notebook shows use of the [What-If Tool](https://pair-code.github.io/what-if-tool) to compare two text models that determine sentence toxicity, one of which has had some debiasing performed during training.
#
# This notebook loads two pretrained toxicity models from [ConversationAI](https://github.com/conversationai/unintended-ml-bias-analysis) and compares them on the [wikipedia comments dataset](https://figshare.com/articles/Wikipedia_Talk_Labels_Toxicity/4563973).
#
# This notebook also shows how the What-If Tool can be used on non-TensorFlow models. In this case, these models are keras models that do not use tensorflow Examples as an input format. These models can be analyzed in the What-If Tool by supplying a custom prediction function to WitWidget.
#
# ##WARNING: Some text examples in this notebook include profanity, offensive statments, and offensive statments involving identity terms. Please feel free to avoid using this notebook.
#
# + id="qqB2tjOMETmr" colab_type="code" colab={}
#@title Install the What-If Tool widget if running in colab {display-mode: "form"}
# If running in colab then pip install, otherwise no need.
try:
import google.colab
# !pip install --upgrade witwidget
except Exception:
pass
# + id="EBOHfrOP7Iy5" colab_type="code" cellView="form" colab={}
#@title Download the pretrained keras model files
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/cnn_wiki_tox_v3_model.h5 -o ./cnn_wiki_tox_v3_model.h5
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/cnn_wiki_tox_v3_hparams.h5 -o ./cnn_wiki_tox_v3_hparams.h5
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/cnn_wiki_tox_v3_tokenizer.pkl -o ./cnn_wiki_tox_v3_tokenizer.pkl
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/cnn_debias_tox_v3_model.h5 -o ./cnn_debias_tox_v3_model.h5
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/cnn_debias_tox_v3_hparams.h5 -o ./cnn_debias_tox_v3_hparams.h5
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/cnn_debias_tox_v3_tokenizer.pkl -o ./cnn_debias_tox_v3_tokenizer.pkl
# !curl -L https://storage.googleapis.com/what-if-tool-resources/computefest2019/wiki_test.csv -o ./wiki_test.csv
# + id="zZR3i6UZlZ96" colab_type="code" cellView="both" colab={}
#@title Load the keras models
import sys
from keras.models import load_model
from six.moves import cPickle as pkl
def pkl_load(f):
return pkl.load(f) if sys.version_info < (3, 0) else pkl.load(
f, encoding='latin1')
model1 = load_model('cnn_wiki_tox_v3_model.h5')
with open('cnn_wiki_tox_v3_tokenizer.pkl', 'rb') as f:
tokenizer1 = pkl_load(f)
tokenizer1.oov_token = None # quick fix for version issues
model2 = load_model('cnn_debias_tox_v3_model.h5')
with open('cnn_debias_tox_v3_tokenizer.pkl', 'rb') as f:
tokenizer2 = pkl_load(f)
tokenizer2.oov_token = None # quick fix for version issues
# + id="nStoYhqT80WH" colab_type="code" cellView="both" colab={}
#@title Define custom prediction functions so that WIT infers using keras models
from keras.preprocessing.sequence import pad_sequences
# Set up model helper functions:
PADDING_LEN = 250
# Get raw string out of tf.Example and prepare it for keras model input
def examples_to_model_in(examples, tokenizer):
texts = [ex.features.feature['comment'].bytes_list.value[0] for ex in examples]
if sys.version_info >= (3, 0):
texts = [t.decode('utf-8') for t in texts]
# Tokenize string into fixed length sequence of integer based on tokenizer
# and model padding
text_sequences = tokenizer.texts_to_sequences(texts)
model_ins = pad_sequences(text_sequences, maxlen=PADDING_LEN)
return model_ins
# WIT predict functions:
def custom_predict_1(examples_to_infer):
model_ins = examples_to_model_in(examples_to_infer, tokenizer1)
preds = model1.predict(model_ins)
return preds
def custom_predict_2(examples_to_infer):
model_ins = examples_to_model_in(examples_to_infer, tokenizer2)
preds = model2.predict(model_ins)
return preds
# + id="NXaUORW0DVhg" colab_type="code" cellView="form" colab={}
#@title Define helper functions for dataset conversion from csv to tf.Examples
import numpy as np
import tensorflow as tf
# Converts a dataframe into a list of tf.Example protos.
def df_to_examples(df, columns=None):
examples = []
if columns == None:
columns = df.columns.values.tolist()
for index, row in df.iterrows():
example = tf.train.Example()
for col in columns:
if df[col].dtype is np.dtype(np.int64):
example.features.feature[col].int64_list.value.append(int(row[col]))
elif df[col].dtype is np.dtype(np.float64):
example.features.feature[col].float_list.value.append(row[col])
elif row[col] == row[col]:
example.features.feature[col].bytes_list.value.append(row[col].encode('utf-8'))
examples.append(example)
return examples
# Converts a dataframe column into a column of 0's and 1's based on the provided test.
# Used to force label columns to be numeric for binary classification using a TF estimator.
def make_label_column_numeric(df, label_column, test):
df[label_column] = np.where(test(df[label_column]), 1, 0)
# + id="nu398ARdeuxe" colab_type="code" colab={}
#@title Read the dataset from CSV and process it for model {display-mode: "form"}
import pandas as pd
# Set the path to the CSV containing the dataset to train on.
csv_path = 'wiki_test.csv'
# Set the column names for the columns in the CSV. If the CSV's first line is a header line containing
# the column names, then set this to None.
csv_columns = None
# Read the dataset from the provided CSV and print out information about it.
df = pd.read_csv(csv_path, names=csv_columns, skipinitialspace=True)
df = df[['is_toxic', 'comment']]
# Remove non ascii characters
comments = df['comment'].values
proc_comments = []
for c in comments:
try:
if sys.version_info >= (3, 0):
c = bytes(c, 'utf-8')
c = c.decode('unicode_escape')
if sys.version_info < (3, 0):
c = c.encode('ascii', 'ignore')
proc_comments.append(c.strip())
except:
proc_comments.append('')
df = df.assign(comment=proc_comments)
label_column = 'is_toxic'
make_label_column_numeric(df, label_column, lambda val: val)
examples = df_to_examples(df)
# + id="UwiWGrLlSWGh" colab_type="code" colab={}
#@title Invoke What-If Tool for the data and two models (Note that this step may take a while due to prediction speed of the toxicity model){display-mode: "form"}
from witwidget.notebook.visualization import WitWidget, WitConfigBuilder
num_datapoints = 1000 #@param {type: "number"}
tool_height_in_px = 720 #@param {type: "number"}
# Setup the tool with the test examples and the trained classifier
config_builder = WitConfigBuilder(examples[:num_datapoints]).set_custom_predict_fn(
custom_predict_1).set_compare_custom_predict_fn(custom_predict_2)
wv = WitWidget(config_builder, height=tool_height_in_px)
# + [markdown] id="A1s1_SiOyS0l" colab_type="text"
# #### Exploration ideas
#
# - Organize datapoints by setting X-axis scatter to "inference score 1" and Y-axis scatter to "inference score 2" to see how each datapoint differs in score between the original model (1) and debiased model (2). Points off the diagonal have differences in results between the two models.
# - Are there patterns of which datapoints don't agree between the two models?
# - If you set the ground truth feature dropdown in the "Performance + Fairness" tab to "is_toxic", then you can color or bin the datapoints by "inference correct 1" or "inference correct 2". Are there patterns of which datapoints are incorrect for model 1? For model 2?
#
# You may want to focus on terms listed [here](https://github.com/conversationai/unintended-ml-bias-analysis/blob/master/unintended_ml_bias/bias_madlibs_data/adjectives_people.txt)
# + id="QuZjEn5qOHFH" colab_type="code" cellView="both" colab={}
#@title Add a feature column for each identity term to indicate if it exists in the comment
# !wget https://raw.githubusercontent.com/conversationai/unintended-ml-bias-analysis/master/unintended_ml_bias/bias_madlibs_data/adjectives_people.txt
import re
with open('adjectives_people.txt', 'r') as f:
segments = f.read().strip().split('\n')
print(segments)
# Tag every sentence with an identity term
comments = df['comment'].values
seg_anns = {}
selected_segments = segments
for s in selected_segments:
is_seg = []
for c in comments:
if re.search(s, c):
is_seg.append(1)
else:
is_seg.append(0)
seg_anns[s] = is_seg
for seg_key, seg_ann in seg_anns.iteritems():
df[seg_key] = pd.Series(seg_ann, index=df.index)
examples = df_to_examples(df)
| WIT_Toxicity_Text_Model_Comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''ko-ecom'': conda)'
# language: python
# name: python3
# ---
# ### Build Dataframe from list
# +
import pandas as pd
value1 = list(range(0, 5))
value2 = [i for i in "ABCDE"]
print(value1)
print(value2)
df = pd.DataFrame({"Column1": value1, "Column2": value2})
df.head()
# -
"HELLO WORLD".split(" ")
"ABDDE".split("")
| notebooks/pandas/list2df.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cluster
import random, pylab, numpy
class Patient(cluster.Example):
pass
def scaleAttrs(vals):
vals = pylab.array(vals)
mean = sum(vals)/len(vals)
sd = numpy.std(vals)
vals = vals - mean
return vals/sd
def getData(toScale = False):
#read in data
hrList, stElevList, ageList, prevACSList, classList = [],[],[],[],[]
cardiacData = open('cardiacData.txt', 'r')
for l in cardiacData:
l = l.split(',')
hrList.append(int(l[0]))
stElevList.append(int(l[1]))
ageList.append(int(l[2]))
prevACSList.append(int(l[3]))
classList.append(int(l[4]))
if toScale:
hrList = scaleAttrs(hrList)
stElevList = scaleAttrs(stElevList)
ageList = scaleAttrs(ageList)
prevACSList = scaleAttrs(prevACSList)
#Build points
points = []
for i in range(len(hrList)):
features = pylab.array([hrList[i], prevACSList[i],\
stElevList[i], ageList[i]])
pIndex = str(i)
points.append(Patient('P'+ pIndex, features, classList[i]))
return points
def kmeans(examples, k, verbose = False):
#Get k randomly chosen initial centroids, create cluster for each
initialCentroids = random.sample(examples, k)
clusters = []
for e in initialCentroids:
clusters.append(cluster.Cluster([e]))
#Iterate until centroids do not change
converged = False
numIterations = 0
while not converged:
numIterations += 1
#Create a list containing k distinct empty lists
newClusters = []
for i in range(k):
newClusters.append([])
#Associate each example with closest centroid
for e in examples:
#Find the centroid closest to e
smallestDistance = e.distance(clusters[0].getCentroid())
index = 0
for i in range(1, k):
distance = e.distance(clusters[i].getCentroid())
if distance < smallestDistance:
smallestDistance = distance
index = i
#Add e to the list of examples for appropriate cluster
newClusters[index].append(e)
for c in newClusters: #Avoid having empty clusters
if len(c) == 0:
raise ValueError('Empty Cluster')
#Update each cluster; check if a centroid has changed
converged = True
for i in range(k):
if clusters[i].update(newClusters[i]) > 0.0:
converged = False
if verbose:
print('Iteration #' + str(numIterations))
for c in clusters:
print(c)
print('') #add blank line
return clusters
def trykmeans(examples, numClusters, numTrials, verbose = False):
"""Calls kmeans numTrials times and returns the result with the
lowest dissimilarity"""
best = kmeans(examples, numClusters, verbose)
minDissimilarity = cluster.dissimilarity(best)
trial = 1
while trial < numTrials:
try:
clusters = kmeans(examples, numClusters, verbose)
except ValueError:
continue #If failed, try again
currDissimilarity = cluster.dissimilarity(clusters)
if currDissimilarity < minDissimilarity:
best = clusters
minDissimilarity = currDissimilarity
trial += 1
return best
def printClustering(clustering):
"""Assumes: clustering is a sequence of clusters
Prints information about each cluster
Returns list of fraction of pos cases in each cluster"""
posFracs = []
for c in clustering:
numPts = 0
numPos = 0
for p in c.members():
numPts += 1
if p.getLabel() == 1:
numPos += 1
fracPos = numPos/numPts
posFracs.append(fracPos)
print('Cluster of size', numPts, 'with fraction of positives =',
round(fracPos, 4))
return pylab.array(posFracs)
def testClustering(patients, numClusters, seed = 0, numTrials = 5):
random.seed(seed)
bestClustering = trykmeans(patients, numClusters, numTrials)
posFracs = printClustering(bestClustering)
return posFracs
patients = getData()
for k in (2,):
print('\n Test k-means (k = ' + str(k) + ')')
posFracs = testClustering(patients, k, 2)
numPos = 0
for p in patients:
if p.getLabel() == 1:
numPos += 1
print('Total number of positive patients =', numPos)
| models/nearest_neighbors/k_means.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Excitation Signals for Room Impulse Response Measurement
#
# ### Criteria
# - Sufficient signal energy over the entire frequency range of interest
# - Dynamic range
# - Creast factor (peak-to-RMS value)
# - Noise rejection (repetition and average, longer duration)
# - Measurement duration
# - Time variance
# - Nonlinear distortion
#
# #### _References_
# * <NAME>, and <NAME>. "Transfer-function measurement with sweeps." Journal of the Audio Engineering Society 49.6 (2001): 443-471.
# [link](http://www.aes.org/e-lib/browse.cfm?elib=10189)
#
# * <NAME>. "Simultaneous measurement of impulse response and distortion with a swept-sine technique." Audio Engineering Society Convention 108. Audio Engineering Society, 2000.
# [link](http://www.aes.org/e-lib/browse.cfm?elib=10211)
#
# * <NAME>. "Advancements in impulse response measurements by sine sweeps." Audio Engineering Society Convention 122. Audio Engineering Society, 2007.
# [link](http://www.aes.org/e-lib/browse.cfm?elib=14106)
#
import tools
import numpy as np
from scipy.signal import chirp, max_len_seq, freqz, fftconvolve, resample
import matplotlib.pyplot as plt
import sounddevice as sd
# %matplotlib inline
# +
def crest_factor(x):
"""Peak-to-RMS value (crest factor) of the signal x
Parameter
---------
x : array_like
signal
"""
return np.max(np.abs(x)) / np.sqrt(np.mean(x**2))
def circular_convolve(x, y, outlen):
"""Circular convolution of x and y
Parameters
----------
x : array_like
Real-valued signal
y : array_like
Real-valued signal
outlen : int
Length of the output
"""
return np.fft.irfft(np.fft.rfft(x, n=outlen) * np.fft.rfft(y, n=outlen), n=outlen)
def plot_time_domain(x, fs=44100, ms=False):
time = np.arange(len(x)) / fs
timeunit = 's'
if ms:
time *= 1000
timeunit = 'ms'
fig = plt.figure()
plt.plot(time, x)
plt.xlabel('Time / {}'.format(timeunit))
return
def plot_freq_domain(x, fs=44100, khz=False):
Nf = len(x) // 2 + 1
freq = np.arange(Nf) / Nf * fs / 2
frequnit = 'Hz'
if khz:
freq /= 1000
frequnit = 'kHz'
fig = plt.figure()
plt.plot(freq, db(np.fft.rfft(x)))
plt.xscale('log')
plt.xlabel('Frequency / {}'.format(frequnit))
plt.ylabel('Magnitude / dB')
return
def compare_irs(h1, h2, ms=False):
t1 = np.arange(len(h1)) / fs
t2 = np.arange(len(h2)) / fs
timeunit = 's'
if ms:
t1 *= 1000
t2 *= 1000
timeunit = 'ms'
fig = plt.figure()
plt.plot(t1, h1, t2, h2)
plt.xlabel('Time / {}'.format(timeunit))
return
def compare_tfs(h1, h2, khz=False):
n1 = len(h1) // 2 + 1
n2 = len(h2) // 2 + 1
f1 = np.arange(n1) / n1 * fs / 2
f2 = np.arange(n2) / n2 * fs / 2
frequnit = 'Hz'
if khz:
freq /= 1000
frequnit = 'khz'
fig = plt.figure()
plt.plot(f1, db(np.fft.rfft(h1)), f2, db(np.fft.rfft(h2)))
plt.xscale('log')
plt.xlabel('Frequency / {}'.format(frequnit))
plt.ylabel('Magnitude / dB')
return
def pad_zeros(x, nzeros):
"""Append zeros at the end of the input sequence
"""
return np.pad(x, (0, nzeros), mode='constant', constant_values=0)
# -
# ## Parameters
fs = 44100
dur = 1
L = int(np.ceil(dur * fs))
time = np.arange(L) / fs
# ## White Noise
# Generate a random signal with normal (Gaussian) amplitude distribution. Use `numpy.random.randn` and normalize the amplitude with `tools.normalize`.
# Let's listen to it.
# Plot the signal in the time domain and in the frequency domain.
# Is the signal really white?
# What is the crest factor of a white noise?
# Now feed the white noise to an unkown system `tools.blackbox` and save the output signal.
# How do you think we can extract the impulse response of the system?
# Try to compute the impulse response from the output signal.
# Compare it with the actual impulse response which can be obtained by feeding an ideal impulse to `tools.blackbox`.
# ## Maximum Length Sequence
#
# > Maximum-length sequences (MLSs) are binary sequences that can be generated very easily with an N-staged shift register and an XOR gate (with up to four inputs) connected with the shift register in such a way that all possible 2N states, minus the case "all 0," are run through. This can be accomplished by hardware with very few simple TTL ICs or by software with less than 20 lines of assembly code.
#
# (Müller 2001)
nbit = int(np.ceil(np.log2(L)))
mls, _ = max_len_seq(nbit) # sequence of 0 and 1
mls = 2*mls - 1 # sequence of -1 and 1
# Take a look at the signal in the time domain.
# Examine the properties of the MLS
# * frequency response
# * crest factor
# * simulate the impulse response measurement of `tools.blackbox`
# * evaluate the obtained impulse response
# In practive, the (digital) signal has to be converted into an analog signal by an audio interface?
# Here, the process is simulated by oversampling the signal by a factor of 10.
# Pay attention to the crest factor before and after upsampling.
# +
upsample = 10
mls_up = resample(mls, num=len(mls) * upsample)
time = np.arange(len(mls)) / fs
time_up = np.arange(len(mls_up)) / fs / upsample
plt.figure(figsize=(10, 4))
plt.plot(time_up, mls_up, '-', label='Analog')
plt.plot(time, mls, '-', label='Digital')
plt.legend(loc='best')
plt.xlabel('Time / s')
plt.title('Crest factor {:.1f} -> {:.1f} dB'.format(db(crest_factor(mls)), db(crest_factor(mls_up))))
plt.figure(figsize=(10, 4))
plt.plot(time_up, mls_up, '-', label='Analog')
plt.plot(time, mls, 'o', label='Ditigal')
plt.xlim(0, 0.0025)
plt.legend(loc='best')
plt.xlabel('Time / s')
plt.title('Crest factor {:.1f} -> {:.1f} dB'.format(db(crest_factor(mls)), db(crest_factor(mls_up))));
# -
# ## Linear Sweep
# Generate a linear sweep with `lin_sweep`.
def lin_sweep(fstart, fstop, duration, fs):
"""Generation of a linear sweep signal.
Parameters
----------
fstart : int
Start frequency in Hz
fstop : int
Stop frequency in Hz
duration : float
Total length of signal in s
fs : int
Sampling frequency in Hz
Returns
-------
array_like
generated signal vector
Note that the stop frequency must not be greater than half the
sampling frequency (Nyquist-Shannon sampling theorem).
"""
if fstop > fs / 2:
raise ValueError("fstop must not be greater than fs/2")
t = np.arange(0, duration, 1 / fs)
excitation = np.sin(
2 * np.pi * ((fstop - fstart) /
(2 * duration) * t ** 2 + fstart * t))
# excitation = excitation - np.mean(excitation) # remove direct component
return excitation
# +
fs = 44100
fstart =
fstop =
duration =
lsweep =
# -
# Examine the properties of linear sweeps
# * spectrogram (Use `pyplot.specgram` with `NFFT=512` and `Fs=44100`)
# * frequency response
# * crest factor
# * simulate the impulse response measurement of `tools.blackbox`
# * evaluate the obtained impulse response
# ## Exponential Sweep
# Generate a exponential sweep with `exp_sweep`.
def exp_sweep(fstart, fstop, duration, fs):
"""Generation of a exponential sweep signal.
Parameters
----------
fstart : int
Start frequency in Hz
fstop : int
Stop frequency
duration : float
Total length of signal in s
fs : int
Sampling frequency in Hz
Returns
-------
array_like
Generated signal vector
Note that the stop frequency must not be greater than half the
sampling frequency (Nyquist-Shannon sampling theorem).
"""
if fstop > fs / 2:
raise ValueError("fstop must not be greater than fs/2")
t = np.arange(0, duration, 1 / fs)
excitation = np.sin(2 * np.pi * duration *
fstart / np.log(fstop / fstart) *
(np.exp(t / duration * np.log(fstop / fstart)) - 1))
# excitation = excitation - np.mean(excitation) # remove direct component
return excitation
# +
fs = 44100
fstart =
fstop =
duration =
esweep =
# -
# Examine the properties of linear sweeps
# * spectrogram (Use `pyplot.specgram` with `NFFT=512` and `Fs=44100`)
# * frequency response
# * crest factor
# * simulate the impulse response measurement of `tools.blackbox`
# * evaluate the obtained impulse response
| excitation-signal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="7d_M53KCRuqo" outputId="093df1ac-a2ce-4853-fb41-7bb67813cd23"
x=int(input('enter the first number '))
y=int(input('enter the second number '))
# + colab={"base_uri": "https://localhost:8080/"} id="F28r1tBxR1_z" outputId="f368ff15-5c0d-4bdd-cce7-ce819230e3b5"
10000*5*8/100
# + colab={"base_uri": "https://localhost:8080/"} id="bTUknH5RTa_P" outputId="b774f0e3-8f70-481d-e49f-3e3d0d955fb3"
P=int(input('enter the principal amount:'))
N=int(input('enter the no of years:'))
R=int(input('enter the rate:'))
I=P*N*R/100
T=P+I
print('interst amount:',I)
print('Total amount',T)
# + colab={"base_uri": "https://localhost:8080/"} id="gEk_SRGZUXAR" outputId="85387bcb-d5cb-4d63-9dbf-5edf911096bc"
x=int(input('enter your mark'))
if x>=50:
print('pass')
if x<50:
print('fail')
# + colab={"base_uri": "https://localhost:8080/"} id="LlkQ5Lf0YUuw" outputId="9715dfa6-4148-47dd-eb4c-93a105f082e1"
S=int(input('enter the salary'))
if S>=250000:
print('you are taxable')
if S<250000:
print('you are not taxable')
# + colab={"base_uri": "https://localhost:8080/"} id="miB6iK3wa9lW" outputId="ceba4619-1abb-4788-905b-22559775098e"
S=int(input('enter the salary'))
if S>=250000:
print('you are taxable')
T=print('Tax amount',S*10/100)
if S<250000:
print('you are not taxable')
# + [markdown] id="rLTLuvYSdCqO"
#
# + colab={"base_uri": "https://localhost:8080/"} id="0Y4EClRVcE4_" outputId="92e68b2c-a73e-4643-f856-067a8ad3c76b"
S=int(input('enter the salary'))
x=250000
if S>=250000:
print('you are taxable')
T=print('Tax amount',(S-x)*10/100)
if S<250000:
print('you are not taxable')
# + id="xJwKMqSXj3sX"
# + id="c614n5m5eXBG" colab={"base_uri": "https://localhost:8080/"} outputId="fb6184fa-ee48-40e1-f927-114b35403ede"
S=int(input('enter the salary'))
x=250000
if S>=250000 and S<500000:
print('taxable')
T=print('Tax amount',(S-x)*10/100)
if S>500000:
print('taxable')
T=print('tax amount',(S-x)*20/100)
if S<250000:
print('you are not taxable')
# + id="FJlSjrqflUOH"
| Pythontax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import time
from time import time
import os
import os
import gensim
from gensim.models import Word2Vec, KeyedVectors
from nltk.corpus import stopwords
from nltk import download
stop_words = stopwords.words('english')
from nltk import word_tokenize
download('punkt') # Download data for tokenizer.
from gensim.similarities import WmdSimilarity
start_nb = time()
#https://markroxor.github.io/gensim/static/notebooks/WMD_tutorial.html
# +
# Pre-processing a document.
def preprocess(doc):
doc = doc.lower() # Lower the text.
doc = word_tokenize(doc) # Split into words.
doc = [w for w in doc if not w in stop_words] # Remove stopwords.
doc = [w for w in doc if w.isalpha()] # Remove numbers and punctuation.
return doc
# -
relPath = 'data/1.0-billTitleSponsors.csv'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
#data_dir = '/Users/whs/Documents/GitHub/gpo-ai.com/data/processed/1.0-billTitleSponsors.csv'
#data_dir = 'C:\\Users\\wsolomon\\Desktop\\DataScience\\projects\\loc_challenge-master\\1.0-billTitleSponsors.csv'
relPath = 'data/1.0-billTitleSponsors.csv'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
df = pd.read_csv(absPath)
df.head()
# +
start = time()
relPath = 'model/GoogleNews-vectors-negative300.bin.gz'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
#path = '/Users/whs/Documents/GitHub/gpo-ai.com/data/processed/GoogleNews-vectors-negative300.bin.gz'
if not os.path.exists(absPath):
raise ValueError("SKIP: You need to download the google news model")
model = gensim.models.KeyedVectors.load_word2vec_format(absPath, binary=True)
print('Cell took %.2f seconds to run.' % (time() - start))
# -
doc1 = df['texts'][3]
doc2 = df['texts'][4]
distance = model.wmdistance(doc1,doc2)
print('distance = %.4f' % distance)
print(df.shape)
df = df.dropna()
df.shape
#preprocess(df.texts.values[1000])
df.texts.values[1000]
# +
start = time()
wmd_corpus = [preprocess(doc) for doc in df.texts.values]
print('Docs in Corpus: ',len(wmd_corpus))
num_best = 4
instance = WmdSimilarity(wmd_corpus, model, num_best=num_best)
print('Cell took %.2f seconds to run.' % (time() - start))
# +
start = time()
sent = 'guns'
query = preprocess(sent)
sims = instance[query] # A query is simply a "look-up" in the similarity class.
print(sims)
#print('Cell took %.2f seconds to run.' % (time() - start))
# Print the query and the retrieved documents, together with their similarities.
print('Query:')
print(sent)
for i in range(num_best):
print()
print('sim = %.4f' % sims[i][1])
print(df.title.values[sims[i][0]])
print(df.sponsers.values[sims[i][0]])
# +
relPath = 'model/1.0-whs-W2VModel'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
model.save(absPath)
# +
relPath = 'model/1.1-whs-Sim_Instance'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
instance.save(absPath)
# -
# +
w2v_relPath = 'model/GoogleNews-vectors-negative300.bin.gz'
w2v_absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
relPath = 'model/1.0-whs-W2VModel'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
#modelX = KeyedVectors.load_word2vec_format(w2v_absPath, binary=True) #.load(absPath)
relPath = 'model/1.1-whs-Sim_Instance'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
instanceX = WmdSimilarity.load(absPath)
relPath = 'data/1.0-billTitleSponsors.csv'
absPath = os.path.join(os.path.dirname(os.getcwd()), relPath)
df = pd.read_csv(absPath)
# +
start = time()
sent = 'I like sports'
query = preprocess(sent)
sims = instanceX[query] # A query is simply a "look-up" in the similarity class.
print(sims)
#print('Cell took %.2f seconds to run.' % (time() - start))
# Print the query and the retrieved documents, together with their similarities.
print('Query:')
print(sent)
for i in range(num_best):
print()
print('sim = %.4f' % sims[i][1])
print(df.title.values[sims[i][0]])
print(df.sponsers.values[sims[i][0]])
print('Cell took %.2f seconds to run.' % (time() - start))
# +
start = time()
modelX = Word2Vec.load(path)
from gensim.similarities import WmdSimilarity
wmd_corpus = [preprocess(doc) for doc in df.title.values]
print(len(wmd_corpus))
num_best = 10
instanceX = WmdSimilarity(wmd_corpus, modelX, num_best=10)
print('Cell took %.2f seconds to run.' % (time() - start))
# +
start = time()
#model = Word2Vec.load(fname)
path = '/Users/whs/Documents/GitHub/gpo-ai.com/models/1.1-whs-Sim_Instance'
instanceX = WmdSimilarity.load(path)
sent = 'guns'
query = preprocess(sent)
sims = instanceX[query] # A query is simply a "look-up" in the similarity class.
print(sims)
#print('Cell took %.2f seconds to run.' % (time() - start))
# Print the query and the retrieved documents, together with their similarities.
print('Query:')
print(sent)
for i in range(num_best):
print()
print('sim = %.4f' % sims[i][1])
print(df.title.values[sims[i][0]])
print(df.sponsers.values[sims[i][0]])
# -
| notebooks/2.0-whs-BuildModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Vascular, Extracellular and Restricted Diffusion for Cytometry in Tumors (VERDICT)
# Panagiotaki et al. (2014) proposed a multi-compartment model called VERDICT to characterize the composition of tumorous tissues. VERDICT models the diffusion in tumor cells, the extra-cellular space and surrounding bloodvessels as a restricted Sphere, an isotropic Gaussian Ball and a Stick compartment, respectively. VERDICT's design is as follows:
# \begin{equation}
# E_{\textrm{VERDICT}}= \underbrace{f_{\textrm{Tumor}}\overbrace{E(D|\lambda_{\textrm{intra}})}^{\textrm{Sphere}}}_{\textrm{Tumor Cells}} + \underbrace{f_{\textrm{extra}}\overbrace{E(\cdot|\lambda_{iso})}^{\textrm{Ball}}}_{\textrm{Hindered Extra-Cellular}}+\underbrace{f_{blood}\overbrace{E(\lambda_\parallel, \boldsymbol{\mu})}^{\textrm{Stick}}}_{\textrm{Vascular}}
# \end{equation}
# where $D$ is the sphere's diameter. VERDICT uses the Gaussian Phase approximation to model the sphere *(Balinov et al. 1993)*, which accounts to changes in gradient pulse duration $\delta$ and separation $\Delta$. Furthermore, some particular parameter constraints are imposed:
# - The intra-cellular (intra-spherical) diffusivity is fixed to 0.9e-9 m^2/s
# - The extra-cellular Gaussian diffusivity is also fixed to 0.9e-9 m^2/s
# - The optimization range for $\lambda_\parallel$ of the Vascular Stick is set between (3.05-10)e-9 m^2/s
#
# We can define the VERDICT model in the following lines of code:
# # Using Dmipy to set up the VERDICT model
# We first load the necessary modules to import the Sphere, Ball, and Stick models.
from dmipy.signal_models import sphere_models, cylinder_models, gaussian_models
# The Sphere model must be initiated with VERDICT's setting for intra-spherical diffusivity, while the Ball and Stick model are regularly imported.
sphere = sphere_models.S4SphereGaussianPhaseApproximation(diffusion_constant=0.9e-9)
ball = gaussian_models.G1Ball()
stick = cylinder_models.C1Stick()
# We then assemble the three models into a multi-compartment model:
from dmipy.core.modeling_framework import MultiCompartmentModel
verdict_mod = MultiCompartmentModel(models=[sphere, ball, stick])
verdict_mod.parameter_names
# Visualize the model:
from IPython.display import Image
verdict_mod.visualize_model_setup(view=False, cleanup=False, with_parameters=True)
Image('Model Setup.png')
# We then fix the Ball's diffusivity, and adjust the Stick's optimization range for $\lambda_\parallel$ for the optimization.
verdict_mod.set_fixed_parameter('G1Ball_1_lambda_iso', 0.9e-9)
verdict_mod.set_parameter_optimization_bounds('C1Stick_1_lambda_par', [3.05e-9, 10e-9])
# We are now ready to fit the data.
# # Fitting VERDICT to UCL Example Data
# As illustration data we use the voxel of VERDICT data that is freely available at the UCL website at http://camino.cs.ucl.ac.uk/index.php?n=Tutorials.VERDICTcol. We prepared the script to easily import the acquisition scheme and example data from there:
from dmipy.data import saved_data
scheme, data = saved_data.panagiotaki_verdict()
scheme.print_acquisition_info
# We can see that the VERDICT acquisition scheme is very particular, having many shells with different diffusion times and often only 3 perpendicular measurements per shell, with the exception of one DTI shell with 42 measurements.
#
# To fit the data we use the MIX algorithm *(Farooq et al. 2016)*, which is efficient for finding the global minimum in models with many compartments. We set parallel processing to False since there is only one voxel.
verdict_fit = verdict_mod.fit(scheme, data, solver='mix', use_parallel_processing=False)
# # Visualize results
# To illustrate that Dmipy's VERDICT implementation is correct, we show that we can produce a very similar signal fitting plots as the one shown on the UCL website. On the left we show their original graph, and on the right we show Dmipy's predicted signal together with the measured signal attenuation.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import matplotlib.image as mpimg
img=mpimg.imread("http://camino.cs.ucl.ac.uk/uploads/Tutorials/VCLSsynth.png")
mask_nonzero = scheme.gradient_strengths>0.
G_nonzero = scheme.gradient_strengths[mask_nonzero]
Delta_nonzero = scheme.Delta[mask_nonzero]
delta_nonzero = scheme.delta[mask_nonzero]
predicted_data = verdict_fit.predict()[0]
predicted_data_nonzero = predicted_data[mask_nonzero]
data_nonzero = data[mask_nonzero]
fig, axs = plt.subplots(1, 2, figsize=[30, 10])
axs = axs.ravel()
axs[0].imshow(img)
axs[0].set_title('UCL Verdict Results', fontsize=30)
axs[0].axis('off')
for delta_, Delta_ in np.unique(np.c_[scheme.shell_delta, scheme.shell_Delta], axis=0):
mask = np.all([Delta_nonzero == Delta_, delta_nonzero == delta_], axis=0)
axs[1].plot(G_nonzero[mask], predicted_data_nonzero[mask])
axs[1].scatter(G_nonzero[mask], data_nonzero[mask], s=3., marker='o')
axs[1].set_title('Dmipy Verdict Results', fontsize=30)
axs[1].set_xlabel('G(T/m)', fontsize=20)
axs[1].set_ylabel('S', fontsize=20);
# -
# While we didn't exactly duplicate their graph style, we can see that both the plots and signal fitting are very similar between Dmipy's and UCL's implementation.
#
# Showing the fitting parameters, we can also see that Dmipy's estimate of the tumor cell diameter is 1.54e-5m, i.e. 15.4$\mu m$, which falls exactly within the ranges that Panagiotaki et al. reports.
verdict_fit.fitted_parameters
# ## References
# - Panagiotaki, Eletheria, et al. "Noninvasive quantification of solid tumor microstructure using VERDICT MRI." Cancer research 74.7 (2014): 1902-1912.
# - <NAME>, et al. "The NMR self-diffusion method applied to restricted diffusion. Simulation of echo attenuation from molecules in spheres and between planes." Journal of Magnetic Resonance, Series A 104.1 (1993): 17-25.
# - <NAME>, et al. "Microstructure Imaging of Crossing (MIX) White Matter Fibers from diffusion MRI." Nature Scientific reports 6 (2016).
| examples/example_verdict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="NTp7EUU_kT1x" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="807ee578-43ac-4dec-cdff-4d3720914b9e"
# !pip install torch===1.4.0 torchvision===0.5.0 -f https://download.pytorch.org/whl/torch_stable.html
# + id="YL2WMyUhfkyX" colab={"base_uri": "https://localhost:8080/", "height": 523} outputId="8e920dd2-14bd-45c0-a943-e8939ad89fd6"
# !pip install pytorch_lightning
# + id="8aOFSycYfmFf"
# used fo TPU
# Uncomment if you want to use TPU
# ------------------------------------------------------------------
# import collections
# from datetime import datetime, timedelta
# import os
# import requests
# import threading
# _VersionConfig = collections.namedtuple('_VersionConfig', 'wheels,server')
# VERSION = "xrt==1.15.0" # @param ["xrt==1.15.0", "torch_xla==nightly"]
# CONFIG = {
# 'xrt==1.15.0': _VersionConfig('1.15', '1.15.0'),
# 'torch_xla==nightly': _VersionConfig('nightly', 'XRT-dev{}'.format(
# (datetime.today() - timedelta(1)).strftime('%Y%m%d'))),
# }[VERSION]
# DIST_BUCKET = 'gs://tpu-pytorch/wheels'
# TORCH_WHEEL = 'torch-{}-cp36-cp36m-linux_x86_64.whl'.format(CONFIG.wheels)
# TORCH_XLA_WHEEL = 'torch_xla-{}-cp36-cp36m-linux_x86_64.whl'.format(CONFIG.wheels)
# TORCHVISION_WHEEL = 'torchvision-{}-cp36-cp36m-linux_x86_64.whl'.format(CONFIG.wheels)
# # Update TPU XRT version
# def update_server_xrt():
# print('Updating server-side XRT to {} ...'.format(CONFIG.server))
# url = 'http://{TPU_ADDRESS}:8475/requestversion/{XRT_VERSION}'.format(
# TPU_ADDRESS=os.environ['COLAB_TPU_ADDR'].split(':')[0],
# XRT_VERSION=CONFIG.server,
# )
# print('Done updating server-side XRT: {}'.format(requests.post(url)))
# update = threading.Thread(target=update_server_xrt)
# update.start()
# + id="LHLEBnnSfosm"
import torch
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import DataLoader
import pytorch_lightning as pl
from pytorch_lightning import loggers
from numpy.random import choice
import os
from pathlib import Path
import shutil
from collections import OrderedDict
# + id="yjjEWS1Lfthl"
# lets hope this fixes the bug that my pc crashes after reconnecting
from IPython.display import clear_output
# + id="KmTaKo7mfwxc"
# custom weights initialization called on netG and netD
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
nn.init.normal_(m.weight.data, 0.0, 0.02)
elif classname.find('BatchNorm') != -1:
nn.init.normal_(m.weight.data, 1.0, 0.02)
nn.init.constant_(m.bias.data, 0)
# + id="Sx70EX9wfx1D"
# randomly flip some labels
def noisy_labels(y, p_flip=0.05): # # flip labels with 5% probability
# determine the number of labels to flip
n_select = int(p_flip * y.shape[0])
# choose labels to flip
flip_ix = choice([i for i in range(y.shape[0])], size=n_select)
# invert the labels in place
y[flip_ix] = 1 - y[flip_ix]
return y
# + id="noCnaCJZfzHm"
class AddGaussianNoise(object):
def __init__(self, mean=0.0, std=0.1):
self.std = std
self.mean = mean
def __call__(self, tensor):
tensor = tensor.cuda()
return tensor + (torch.randn(tensor.size()) * self.std + self.mean).cuda()
def __repr__(self):
return self.__class__.__name__ + '(mean={0}, std={1})'.format(self.mean, self.std)
# + id="XzAjUIW5f0kd"
def resize2d(img, size):
return (F.adaptive_avg_pool2d(img, size).data).cuda()
# + id="EptHLAyIf3xB"
def get_valid_labels(img):
return ((0.8 - 1.1) * torch.rand(img.shape[0], 1, 1, 1) + 1.1).cuda() # soft labels
# + id="0v-p-xCFf5HG"
def get_unvalid_labels(img):
return noisy_labels((0.0 - 0.3) * torch.rand(img.shape[0], 1, 1, 1) + 0.3).cuda() # soft labels
# + id="2amoqoiFf6WO"
class Generator(pl.LightningModule):
def __init__(self, ngf, nc, latent_dim):
super(Generator, self).__init__()
self.ngf = ngf
self.latent_dim = latent_dim
self.nc = nc
self.fc0 = nn.Sequential(
# input is Z, going into a convolution
nn.utils.spectral_norm(nn.ConvTranspose2d(latent_dim, ngf * 16, 4, 1, 0, bias=False)),
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ngf * 16)
)
self.fc1 = nn.Sequential(
# state size. (ngf*8) x 4 x 4
nn.utils.spectral_norm(nn.ConvTranspose2d(ngf * 16, ngf * 8, 4, 2, 1, bias=False)),
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ngf * 8)
)
self.fc2 = nn.Sequential(
# state size. (ngf*4) x 8 x 8
nn.utils.spectral_norm(nn.ConvTranspose2d(ngf * 8, ngf * 4, 4, 2, 1, bias=False)),
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ngf * 4)
)
self.fc3 = nn.Sequential(
# state size. (ngf*2) x 16 x 16
nn.utils.spectral_norm(nn.ConvTranspose2d(ngf * 4, ngf * 2, 4, 2, 1, bias=False)),
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ngf * 2)
)
self.fc4 = nn.Sequential(
# state size. (ngf) x 32 x 32
nn.utils.spectral_norm(nn.ConvTranspose2d(ngf * 2, ngf, 4, 2, 1, bias=False)),
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ngf)
)
self.fc5 = nn.Sequential(
# state size. (nc) x 64 x 64
nn.utils.spectral_norm(nn.ConvTranspose2d(ngf, nc, 4, 2, 1, bias=False)),
nn.Tanh()
)
# state size. (nc) x 128 x 128
# For Multi-Scale Gradient
# Converting the intermediate layers into images
self.fc0_r = nn.Conv2d(ngf * 16, self.nc, 1)
self.fc1_r = nn.Conv2d(ngf * 8, self.nc, 1)
self.fc2_r = nn.Conv2d(ngf * 4, self.nc, 1)
self.fc3_r = nn.Conv2d(ngf * 2, self.nc, 1)
self.fc4_r = nn.Conv2d(ngf, self.nc, 1)
def forward(self, input):
x_0 = self.fc0(input)
x_1 = self.fc1(x_0)
x_2 = self.fc2(x_1)
x_3 = self.fc3(x_2)
x_4 = self.fc4(x_3)
x_5 = self.fc5(x_4)
# For Multi-Scale Gradient
# Converting the intermediate layers into images
x_0_r = self.fc0_r(x_0)
x_1_r = self.fc1_r(x_1)
x_2_r = self.fc2_r(x_2)
x_3_r = self.fc3_r(x_3)
x_4_r = self.fc4_r(x_4)
return x_5, x_0_r, x_1_r, x_2_r, x_3_r, x_4_r
# + id="4PFMmdgzf8Xx"
class Discriminator(pl.LightningModule):
def __init__(self, ndf, nc):
super(Discriminator, self).__init__()
self.nc = nc
self.ndf = ndf
self.fc0 = nn.Sequential(
# input is (nc) x 128 x 128
nn.utils.spectral_norm(nn.Conv2d(nc, ndf, 4, 2, 1, bias=False)),
nn.LeakyReLU(0.2, inplace=True)
)
self.fc1 = nn.Sequential(
# state size. (ndf) x 64 x 64
nn.utils.spectral_norm(nn.Conv2d(ndf + nc, ndf * 2, 4, 2, 1, bias=False)),
# "+ nc" because of multi scale gradient
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ndf * 2)
)
self.fc2 = nn.Sequential(
# state size. (ndf*2) x 32 x 32
nn.utils.spectral_norm(nn.Conv2d(ndf * 2 + nc, ndf * 4, 4, 2, 1, bias=False)),
# "+ nc" because of multi scale gradient
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ndf * 4)
)
self.fc3 = nn.Sequential(
# state size. (ndf*4) x 16 x 16e
nn.utils.spectral_norm(nn.Conv2d(ndf * 4 + nc, ndf * 8, 4, 2, 1, bias=False)),
# "+ nc" because of multi scale gradient
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ndf * 8),
)
self.fc4 = nn.Sequential(
# state size. (ndf*8) x 8 x 8
nn.utils.spectral_norm(nn.Conv2d(ndf * 8 + nc, ndf * 16, 4, 2, 1, bias=False)),
nn.LeakyReLU(0.2, inplace=True),
nn.BatchNorm2d(ndf * 16)
)
self.fc5 = nn.Sequential(
# state size. (ndf*8) x 4 x 4
nn.utils.spectral_norm(nn.Conv2d(ndf * 16 + nc, 1, 4, 1, 0, bias=False)),
nn.Sigmoid()
)
# state size. 1 x 1 x 1
def forward(self, input, detach_or_not):
# When we train i ncombination with generator we use multi scale gradient.
x, x_0_r, x_1_r, x_2_r, x_3_r, x_4_r = input
if detach_or_not:
x = x.detach()
x_0 = self.fc0(x)
x_0 = torch.cat((x_0, x_4_r), dim=1) # Concat Multi-Scale Gradient
x_1 = self.fc1(x_0)
x_1 = torch.cat((x_1, x_3_r), dim=1) # Concat Multi-Scale Gradient
x_2 = self.fc2(x_1)
x_2 = torch.cat((x_2, x_2_r), dim=1) # Concat Multi-Scale Gradient
x_3 = self.fc3(x_2)
x_3 = torch.cat((x_3, x_1_r), dim=1) # Concat Multi-Scale Gradient
x_4 = self.fc4(x_3)
x_4 = torch.cat((x_4, x_0_r), dim=1) # Concat Multi-Scale Gradient
x_5 = self.fc5(x_4)
return x_5
# + id="lIPAhNKVf-oF"
class DCGAN(pl.LightningModule):
def __init__(self, hparams, checkpoint_folder, experiment_name):
super().__init__()
self.hparams = hparams
self.checkpoint_folder = checkpoint_folder
self.experiment_name = experiment_name
# networks
self.generator = Generator(ngf=hparams.ngf, nc=hparams.nc, latent_dim=hparams.latent_dim)
self.discriminator = Discriminator(ndf=hparams.ndf, nc=hparams.nc)
self.generator.apply(weights_init)
self.discriminator.apply(weights_init)
# cache for generated images
self.generated_imgs = None
self.last_imgs = None
# For experience replay
self.exp_replay_dis = torch.tensor([])
def forward(self, z):
return self.generator(z)
def adversarial_loss(self, y_hat, y):
return F.binary_cross_entropy(y_hat, y)
def training_step(self, batch, batch_nb, optimizer_idx):
clear_output(wait=True)
# For adding Instance noise for more visit: https://www.inference.vc/instance-noise-a-trick-for-stabilising-gan-training/
std_gaussian = max(0, self.hparams.level_of_noise - (
(self.hparams.level_of_noise * 2) * (self.current_epoch / self.hparams.epochs)))
AddGaussianNoiseInst = AddGaussianNoise(std=std_gaussian) # the noise decays over time
imgs, _ = batch
imgs = AddGaussianNoiseInst(imgs) # Adding instance noise to real images
self.last_imgs = imgs
# train generator
if optimizer_idx == 0:
# sample noise
z = torch.randn(imgs.shape[0], self.hparams.latent_dim, 1, 1).cuda()
# generate images
self.generated_imgs = self(z)
# ground truth result (ie: all fake)
g_loss = self.adversarial_loss(self.discriminator(self.generated_imgs, False), get_valid_labels(
self.generated_imgs[0])) # adversarial loss is binary cross-entropy; [0] is the image of the last layer
tqdm_dict = {'g_loss': g_loss}
log = {'g_loss': g_loss, "std_gaussian": std_gaussian}
output = OrderedDict({
'loss': g_loss,
'progress_bar': tqdm_dict,
'log': log
})
return output
# train discriminator
if optimizer_idx == 1:
# Measure discriminator's ability to classify real from generated samples
# how well can it label as real?
real_loss = self.adversarial_loss(
self.discriminator([imgs, resize2d(imgs, 4), resize2d(imgs, 8), resize2d(imgs, 16), resize2d(imgs, 32), resize2d(imgs, 64)],
False), get_valid_labels(imgs))
fake_loss = self.adversarial_loss(self.discriminator(self.generated_imgs, True), get_unvalid_labels(
self.generated_imgs[0])) # how well can it label as fake?; [0] is the image of the last layer
# discriminator loss is the average of these
d_loss = (real_loss + fake_loss) / 2
tqdm_dict = {'d_loss': d_loss}
log = {'d_loss': d_loss, "std_gaussian": std_gaussian}
output = OrderedDict({
'loss': d_loss,
'progress_bar': tqdm_dict,
'log': log
})
return output
def configure_optimizers(self):
lr_gen = self.hparams.lr_gen
lr_dis = self.hparams.lr_dis
b1 = self.hparams.b1
b2 = self.hparams.b2
opt_g = torch.optim.Adam(self.generator.parameters(), lr=lr_gen, betas=(b1, b2))
opt_d = torch.optim.Adam(self.discriminator.parameters(), lr=lr_dis, betas=(b1, b2))
return [opt_g, opt_d], []
def backward(self, trainer, loss, optimizer, optimizer_idx: int) -> None:
loss.backward(retain_graph=True)
def train_dataloader(self):
# transform = transforms.Compose([transforms.Resize((self.hparams.image_size, self.hparams.image_size)),
# transforms.ToTensor(),
# transforms.Normalize([0.5], [0.5])])
# dataset = torchvision.datasets.MNIST(os.getcwd(), train=False, download=True, transform=transform)
# return DataLoader(dataset, batch_size=self.hparams.batch_size)
# transform = transforms.Compose([transforms.Resize((self.hparams.image_size, self.hparams.image_size)),
# transforms.ToTensor(),
# transforms.Normalize([0.5], [0.5])
# ])
# train_dataset = torchvision.datasets.ImageFolder(
# root="./drive/My Drive/datasets/flower_dataset/",
# # root="./drive/My Drive/datasets/ghibli_dataset_small_overfit/",
# transform=transform
# )
# return DataLoader(train_dataset, num_workers=self.hparams.num_workers, shuffle=True,
# batch_size=self.hparams.batch_size)
transform = transforms.Compose([transforms.Resize((self.hparams.image_size, self.hparams.image_size)),
transforms.ToTensor(),
transforms.Normalize([0.5], [0.5])
])
train_dataset = torchvision.datasets.ImageFolder(
root="./drive/My Drive/datasets/ghibli_dataset_small_overfit/",
transform=transform
)
return DataLoader(train_dataset, num_workers=self.hparams.num_workers, shuffle=True,
batch_size=self.hparams.batch_size)
def on_epoch_end(self):
z = torch.randn(4, self.hparams.latent_dim, 1, 1).cuda()
# match gpu device (or keep as cpu)
if self.on_gpu:
z = z.cuda(self.last_imgs.device.index)
# log sampled images
sample_imgs = self.generator(z)[0]
sample_imgs = sample_imgs.view(-1, self.hparams.nc, self.hparams.image_size, self.hparams.image_size)
grid = torchvision.utils.make_grid(sample_imgs, nrow=2)
grid = grid.permute(1, 2, 0)
torchvision.utils.save_image(grid, f'generated_images_epoch{self.current_epoch}.png')
# save model
if self.current_epoch % self.hparams.save_model_every_epoch == 0:
trainer.save_checkpoint(
self.checkpoint_folder + "/" + self.experiment_name + "_epoch_" + str(self.current_epoch) + ".ckpt")
# + id="cpLirisygB7J"
from argparse import Namespace
args = {
'batch_size': 128, # batch size
'lr_gen': 0.0003, # TTUR;learnin rate of both networks; tested value: 0.0002
'lr_dis': 0.0003, # TTUR;learnin rate of both networks; tested value: 0.0002
'b1': 0.5, # Momentum for adam; tested value(dcgan paper): 0.5
'b2': 0.999, # Momentum for adam; tested value(dcgan paper): 0.999
'latent_dim': 256, # tested value which worked(in V4_1): 100
'nc': 3, # number of color channels
'ndf': 32, # number of discriminator features
'ngf': 32, # number of generator features
'epochs': 4, # the maxima lamount of epochs the algorith should run
'save_model_every_epoch': 1, # how often we save our model
'image_size': 128, # size of the image
'num_workers': 3,
'level_of_noise': 0.1, # how much instance noise we introduce(std; tested value: 0.15 and 0.1
'experience_save_per_batch': 1, # this value should be very low; tested value which works: 1
'experience_batch_size': 50 # this value shouldnt be too high; tested value which works: 50
}
hparams = Namespace(**args)
# + id="YZiLF4R3gDc1"
# Parameters
experiment_name = "DCGAN_6_2_MNIST_128px"
dataset_name = "mnist"
checkpoint_folder = "DCGAN/"
tags = ["DCGAN", "128x128"]
dirpath = Path(checkpoint_folder)
# + id="tWAfRXsVgGTX"
# defining net
net = DCGAN(hparams, checkpoint_folder, experiment_name)
# + id="4D_VasOmgJ1N" colab={"base_uri": "https://localhost:8080/", "height": 1000, "referenced_widgets": ["bc0c05a05ff84229a6ab8eb32783e09f", "465a6f8aff6a4924aff55f808536a0b8", "c88273c3c0af4485aa796ecacfc5c21d", "cca4bbb1f9bb49f082c74edb50c29fc9", "e2b295e333c848ccb33feb8bda6cd7a2", "524796e3ee2b47e881d6c435a244a727", "c29b6f0a22ed41809d110459ef585b61", "4a58e2fccb6e49c193caea3f88236bc9"]} outputId="6b2f91bb-ec5c-4362-e3b5-aaf2ae38ca29"
torch.autograd.set_detect_anomaly(True)
trainer = pl.Trainer( # resume_from_checkpoint="DCGAN_V4_2_GHIBLI_epoch_999.ckpt",
max_epochs=args["epochs"],
gpus=1
)
trainer.fit(net)
# + id="NE7Q9vDQfGn6"
| Code/GAN_V6_2_128px_without_cometml.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# # Mass-univariate twoway repeated measures ANOVA on single trial power
#
#
# This script shows how to conduct a mass-univariate repeated measures
# ANOVA. As the model to be fitted assumes two fully crossed factors,
# we will study the interplay between perceptual modality
# (auditory VS visual) and the location of stimulus presentation
# (left VS right). Here we use single trials as replications
# (subjects) while iterating over time slices plus frequency bands
# for to fit our mass-univariate model. For the sake of simplicity we
# will confine this analysis to one single channel of which we know
# that it exposes a strong induced response. We will then visualize
# each effect by creating a corresponding mass-univariate effect
# image. We conclude with accounting for multiple comparisons by
# performing a permutation clustering test using the ANOVA as
# clustering function. The results final will be compared to
# multiple comparisons using False Discovery Rate correction.
#
# +
# Authors: <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
# <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import numpy as np
import matplotlib.pyplot as plt
import mne
from mne.time_frequency import tfr_morlet
from mne.stats import f_threshold_mway_rm, f_mway_rm, fdr_correction
from mne.datasets import sample
print(__doc__)
# -
# Set parameters
# --------------
#
#
# +
data_path = sample.data_path()
raw_fname = data_path + '/MEG/sample/sample_audvis_raw.fif'
event_fname = data_path + '/MEG/sample/sample_audvis_raw-eve.fif'
tmin, tmax = -0.2, 0.5
# Setup for reading the raw data
raw = mne.io.read_raw_fif(raw_fname)
events = mne.read_events(event_fname)
include = []
raw.info['bads'] += ['MEG 2443'] # bads
# picks MEG gradiometers
picks = mne.pick_types(raw.info, meg='grad', eeg=False, eog=True,
stim=False, include=include, exclude='bads')
ch_name = 'MEG 1332'
# Load conditions
reject = dict(grad=4000e-13, eog=150e-6)
event_id = dict(aud_l=1, aud_r=2, vis_l=3, vis_r=4)
epochs = mne.Epochs(raw, events, event_id, tmin, tmax,
picks=picks, baseline=(None, 0), preload=True,
reject=reject)
epochs.pick_channels([ch_name]) # restrict example to one channel
# -
# We have to make sure all conditions have the same counts, as the ANOVA
# expects a fully balanced data matrix and does not forgive imbalances that
# generously (risk of type-I error).
#
#
# +
epochs.equalize_event_counts(event_id)
# Factor to down-sample the temporal dimension of the TFR computed by
# tfr_morlet.
decim = 2
freqs = np.arange(7, 30, 3) # define frequencies of interest
n_cycles = freqs / freqs[0]
zero_mean = False # don't correct morlet wavelet to be of mean zero
# To have a true wavelet zero_mean should be True but here for illustration
# purposes it helps to spot the evoked response.
# -
# Create TFR representations for all conditions
# ---------------------------------------------
#
#
epochs_power = list()
for condition in [epochs[k] for k in event_id]:
this_tfr = tfr_morlet(condition, freqs, n_cycles=n_cycles,
decim=decim, average=False, zero_mean=zero_mean,
return_itc=False)
this_tfr.apply_baseline(mode='ratio', baseline=(None, 0))
this_power = this_tfr.data[:, 0, :, :] # we only have one channel.
epochs_power.append(this_power)
# Setup repeated measures ANOVA
# -----------------------------
#
# We will tell the ANOVA how to interpret the data matrix in terms of factors.
# This is done via the factor levels argument which is a list of the number
# factor levels for each factor.
#
#
# +
n_conditions = len(epochs.event_id)
n_replications = epochs.events.shape[0] // n_conditions
factor_levels = [2, 2] # number of levels in each factor
effects = 'A*B' # this is the default signature for computing all effects
# Other possible options are 'A' or 'B' for the corresponding main effects
# or 'A:B' for the interaction effect only (this notation is borrowed from the
# R formula language)
n_freqs = len(freqs)
times = 1e3 * epochs.times[::decim]
n_times = len(times)
# -
# Now we'll assemble the data matrix and swap axes so the trial replications
# are the first dimension and the conditions are the second dimension.
#
#
# +
data = np.swapaxes(np.asarray(epochs_power), 1, 0)
# reshape last two dimensions in one mass-univariate observation-vector
data = data.reshape(n_replications, n_conditions, n_freqs * n_times)
# so we have replications * conditions * observations:
print(data.shape)
# -
# While the iteration scheme used above for assembling the data matrix
# makes sure the first two dimensions are organized as expected (with A =
# modality and B = location):
#
# .. table:: Sample data layout
#
# ===== ==== ==== ==== ====
# trial A1B1 A1B2 A2B1 B2B2
# ===== ==== ==== ==== ====
# 1 1.34 2.53 0.97 1.74
# ... ... ... ... ...
# 56 2.45 7.90 3.09 4.76
# ===== ==== ==== ==== ====
#
# Now we're ready to run our repeated measures ANOVA.
#
# Note. As we treat trials as subjects, the test only accounts for
# time locked responses despite the 'induced' approach.
# For analysis for induced power at the group level averaged TRFs
# are required.
#
#
# +
fvals, pvals = f_mway_rm(data, factor_levels, effects=effects)
effect_labels = ['modality', 'location', 'modality by location']
# let's visualize our effects by computing f-images
for effect, sig, effect_label in zip(fvals, pvals, effect_labels):
plt.figure()
# show naive F-values in gray
plt.imshow(effect.reshape(8, 211), cmap=plt.cm.gray, extent=[times[0],
times[-1], freqs[0], freqs[-1]], aspect='auto',
origin='lower')
# create mask for significant Time-frequency locations
effect = np.ma.masked_array(effect, [sig > .05])
plt.imshow(effect.reshape(8, 211), cmap='RdBu_r', extent=[times[0],
times[-1], freqs[0], freqs[-1]], aspect='auto',
origin='lower')
plt.colorbar()
plt.xlabel('Time (ms)')
plt.ylabel('Frequency (Hz)')
plt.title(r"Time-locked response for '%s' (%s)" % (effect_label, ch_name))
plt.show()
# -
# Account for multiple comparisons using FDR versus permutation clustering test
# -----------------------------------------------------------------------------
#
# First we need to slightly modify the ANOVA function to be suitable for
# the clustering procedure. Also want to set some defaults.
# Let's first override effects to confine the analysis to the interaction
#
#
effects = 'A:B'
# A stat_fun must deal with a variable number of input arguments.
# Inside the clustering function each condition will be passed as flattened
# array, necessitated by the clustering procedure. The ANOVA however expects an
# input array of dimensions: subjects X conditions X observations (optional).
# The following function catches the list input and swaps the first and
# the second dimension and finally calls the ANOVA function.
#
#
# +
def stat_fun(*args):
return f_mway_rm(np.swapaxes(args, 1, 0), factor_levels=factor_levels,
effects=effects, return_pvals=False)[0]
# The ANOVA returns a tuple f-values and p-values, we will pick the former.
pthresh = 0.001 # set threshold rather high to save some time
f_thresh = f_threshold_mway_rm(n_replications, factor_levels, effects,
pthresh)
tail = 1 # f-test, so tail > 0
n_permutations = 256 # Save some time (the test won't be too sensitive ...)
T_obs, clusters, cluster_p_values, h0 = mne.stats.permutation_cluster_test(
epochs_power, stat_fun=stat_fun, threshold=f_thresh, tail=tail, n_jobs=1,
n_permutations=n_permutations, buffer_size=None)
# -
# Create new stats image with only significant clusters:
#
#
# +
good_clusters = np.where(cluster_p_values < .05)[0]
T_obs_plot = np.ma.masked_array(T_obs,
np.invert(clusters[np.squeeze(good_clusters)]))
plt.figure()
for f_image, cmap in zip([T_obs, T_obs_plot], [plt.cm.gray, 'RdBu_r']):
plt.imshow(f_image, cmap=cmap, extent=[times[0], times[-1],
freqs[0], freqs[-1]], aspect='auto',
origin='lower')
plt.xlabel('Time (ms)')
plt.ylabel('Frequency (Hz)')
plt.title("Time-locked response for 'modality by location' (%s)\n"
" cluster-level corrected (p <= 0.05)" % ch_name)
plt.show()
# -
# Now using FDR:
#
#
# +
mask, _ = fdr_correction(pvals[2])
T_obs_plot2 = np.ma.masked_array(T_obs, np.invert(mask))
plt.figure()
for f_image, cmap in zip([T_obs, T_obs_plot2], [plt.cm.gray, 'RdBu_r']):
plt.imshow(f_image, cmap=cmap, extent=[times[0], times[-1],
freqs[0], freqs[-1]], aspect='auto',
origin='lower')
plt.xlabel('Time (ms)')
plt.ylabel('Frequency (Hz)')
plt.title("Time-locked response for 'modality by location' (%s)\n"
" FDR corrected (p <= 0.05)" % ch_name)
plt.show()
# -
# Both cluster level and FDR correction help get rid of
# putatively spots we saw in the naive f-images.
#
#
| stable/_downloads/d9e7f23ac267ddfa6023c7da2df2a984/plot_stats_cluster_time_frequency_repeated_measures_anova.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example Logistic Regression and Linear SVC
# +
import mglearn
import matplotlib.pyplot as plt
from sklearn.linear_model import LogisticRegression
from sklearn.svm import LinearSVC
# -
X, y = mglearn.datasets.make_forge()
# +
fig, axes = plt.subplots(1, 2, figsize=(10, 3))
for model, ax in zip([LinearSVC(), LogisticRegression()], axes):
clf = model.fit(X, y)
mglearn.plots.plot_2d_separator(clf, X, fill=False, eps=0.5,
ax=ax, alpha=0.7)
mglearn.discrete_scatter(X[:, 0], X[:, 1], y, ax=ax)
ax.set_title("{}".format(clf.__class__.__name__))
ax.set_xlabel("Feature 0")
ax.set_ylabel("Feature 1")
axes[0].legend()
# -
mglearn.plots.plot_linear_svc_regularization()
# Decision bounderies of a linear SVM on the forge dataset for different values of C
| Coding_exercices/Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <집합 자료형>
# ### - 집합 (set)
# list나 tuple 과는 다르게, 순서(order)가 없고 중복이 허용되지 않는다.<br>
# 리스트를 이용해 set() 메소드를 호출하면 된다.
# <b>집합 이름 = set( 리스트 )</b>로 사용한다.
# +
a = [1, 5, 3, 2, 1]
b = set(a)
print(b)
print(type(b))
# -
# ### - 교집합, 합집합, 차집합
#
# set은 list나 tuple 과는 다르게 순서가 없기 때문에 indexing이나 slicing을 할 수 없다.<br>대신 집합의 연산에 최적화 되어있다.
#
# 교집합 : <b>s1 & s2</b> or <b>s1.intersection(s2)</b><br>
# 합집합 : <b>s1 | s2</b> or <b>s1.union(s2)</b><br>
# 차집합 : <b>s1 - s2</b> or <b>s1.difference(s2)</b><br>
# +
s1 = set([1, 2, 3])
s2 = set([2, 4, 5])
print(s1 & s2)
print(s1.intersection(s2))
print(s1 | s2)
print(s1.union(s2))
print(s1 - s2)
print(s1.difference(s2))
# -
# ### - 요소 추가, 삭제
# 추가 : .add( 추가할 요소 )<br>
# 추가 - (리스트 형태로) : .update( 추가할 리스트 )<br>
# 제거 : .remove( 찾아서 삭제할 요소 )<br>
# +
s1 = set([1, 2, 3])
s1.add(4)
print(s1)
s1.update([5, 6])
print(s1)
s1.remove(1)
print(s1)
| python/Lecture05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Command line interface
# This interface is useful to debug, test connections and features. It can be used for start acquisition or simply connect to an existing one, stream markers, and storage data.
$ openbci_cli -h
# ## Endpoints
# There are 4 endpoints: `serial`, `wifi`, `stream` and `marker`
# ### `serial` and `wifi`
#
# The difference between `serial` and `wifi` are the options `--port` and `--ip` respectively, the first is used to select the serial port and the second is used for select the IP of the WiFi module. Additionally, we can use the option `--output` for specifying a file to write the acquired data in format `hdf5` and `--command` to send commands based on the [SDK](https://docs.openbci.com/docs/02Cyton/CytonSDK) documentation.
$ openbci_cli serial -h
# For example if we wan to start a stream from serial device in port `/dev/ttyUSB0` with daisy module attached, streaming 250 samples per second:
$ openbci_cli serial --port /dev/ttyUSB0 --daisy --streaming_package_size 250
# or start a stream from wifi device in IP `192.168.1.113` in marker mode without daisy module, streaming 2000 samples per second:
$ openbci_cli wifi --ip 192.168.1.113 -s 1000 -c ~4
# the option `~4` according to the [SDK](https://docs.openbci.com/docs/02Cyton/CytonSDK#sample-rate) is for configure the sample rate in 1000 samples per second, `SAMPLE_RATE_2KSPS` is valid too:
$ openbci_cli wifi --ip 192.168.1.113 -s 1000 -c SAMPLE_RATE_2KSPS
# ### `stream`
#
# With this option, we access to a real-time debugger to view the streaming status. We can use the option `--output` for specifying a file to write the acquired data in format `hdf5`.
$ openbci_cli stream -h
# ### `marker`
#
# With this option, we enter into an interactive terminal to create and stream markers.
$ openbci_cli marker
>>>
# ## Remote host
# All interfaces and commands here explained can be executed from a [remote host](A4-configure_remote_host) with the option `--host`:
$ openbci_cli serial --host 192.168.1.113 --port /dev/ttyUSB0 --daisy --streaming_package_size 250
$ openbci_cli stream -h --host 192.168.1.113
$ openbci_cli marker --host 192.168.1.113
| docs/source/notebooks/.ipynb_checkpoints/06-command_line_interface-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day4_Eigenvalues and SVD
import scipy
from scipy import linalg, matrix
import numpy as np
from sympy.solvers import solve
from sympy import Symbol
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Equation Solver
x = Symbol('x')
solve(x**2 - 1, x)
# ## Eigenvalues with Equation
A = matrix([[1, 2], [3, -4]])
A
lam = Symbol('lam')
A_lam = A - lam*np.asmatrix(np.identity(2))
A_lam
equation = A_lam[0,0]*A_lam[1,1] - A_lam[0,1]*A_lam[1,0]
equation
solve(equation, lam)
# ## Eigenvalues and Eigenvectors with Package
eigenvalue, eigenvector = linalg.eig(A)
eigenvalue
eigenvector
# ## Eigen Value Decomposition
eigenvalue, eigenvector = linalg.eig(A)
eigenvalue.shape[0]
L = np.identity(eigenvalue.shape[0])
for i in range(eigenvalue.shape[0]) :
L[i, i] = eigenvalue[i]
L
S= np.asmatrix(eigenvector)
S
A*S
S*L
A*S==S*L
np.allclose(A*S, S*L)
# ## SVD
A = matrix([[3, 1, 1], [-1, 3, 1]])
A
U, s, V = linalg.svd(A, full_matrices=True)
U = np.asmatrix(U)
U
s = np.asmatrix(s)
s
V = np.asmatrix(V)
V
list(A.shape)
np.min(list(A.shape))
S = np.zeros((A.shape))
for i in range(np.min(list(A.shape))) :
S[i, i] = s[0,i]
S
U*S*V
# ## Image Compression with SVD
# https://github.com/rameshputalapattu/jupyterexplore/blob/master/jupyter_interactive_environment_exploration.ipynb
# +
import matplotlib.image as mpimg
img = mpimg.imread('sample.png')
# -
plt.imshow(img)
# +
from skimage.color import rgb2gray
from skimage import img_as_ubyte, img_as_float
gray_images = {
"Pierrot":rgb2gray(img_as_float(img))
}
# -
def compress_svd(image, k):
U, s, V = linalg.svd(image,full_matrices=False)
reconst_matrix = np.dot(U[:,:k],np.dot(np.diag(s[:k]),V[:k,:]))
return reconst_matrix, s
reconst_matrix, s = compress_svd(rgb2gray(img_as_float(img)),50)
s[:5]
plt.plot(s[:5])
def compress_show_gray_images(img_name,k):
image=gray_images[img_name]
original_shape = image.shape
reconst_img,s = compress_svd(image, k)
fig,axes = plt.subplots(1,2,figsize=(8,5))
axes[0].plot(s)
compression_ratio =100.0* (k*(original_shape[0] + original_shape[1])+k)/(original_shape[0]*original_shape[1])
axes[1].set_title("compression ratio={:.2f}".format(compression_ratio)+"%")
axes[1].imshow(reconst_img,cmap='gray')
axes[1].axis('off')
fig.tight_layout()
from ipywidgets import interact,interactive,interact_manual
interact(compress_show_gray_images,img_name=list(gray_images.keys()),k=(1,100));
# ## Applications
# Data projection - pca
#
# Data quantization - spectral clustering methods
#
# Feature selection - apply svd keep high singular value dimensions
# https://github.com/bwcho75/dataanalyticsandML/blob/master/Clustering/1.%20KMeans%20clustering-IRIS%202%20feature.ipynb
# @ Iris Data
#
# Data Set Information:
#
# This is perhaps the best known database to be found in the pattern recognition literature. Fisher's paper is a classic in the field and is referenced frequently to this day. (See Duda & Hart, for example.) The data set contains 3 classes of 50 instances each, where each class refers to a type of iris plant. One class is linearly separable from the other 2; the latter are NOT linearly separable from each other.
#
# Predicted attribute: class of iris plant.
#
# This is an exceedingly simple domain.
#
# This data differs from the data presented in Fishers article (identified by <NAME>, spchadwick '@' espeedaz.net ). The 35th sample should be: 4.9,3.1,1.5,0.2,"Iris-setosa" where the error is in the fourth feature. The 38th sample: 4.9,3.6,1.4,0.1,"Iris-setosa" where the errors are in the second and third features.
#
# Attribute Information:
#
# sepal length in cm
# sepal width in cm
# petal length in cm
# petal width in cm
# class: -- Iris Setosa -- Iris Versicolour -- Iris Virginica
# +
from sklearn import datasets
import pandas as pd
iris = datasets.load_iris()
labels = pd.DataFrame(iris.target)
labels.columns=['labels']
data = pd.DataFrame(iris.data)
data.columns=['Sepal length','Sepal width','Petal length','Petal width']
data = pd.concat([data,labels],axis=1)
data.head()
# -
feature = data[['Sepal length','Sepal width']]
feature.head()
# +
from sklearn.cluster import SpectralClustering
import matplotlib.pyplot as plt
import seaborn as sns
model = SpectralClustering(n_clusters=3)
model.fit(feature)
predict = pd.DataFrame(model.fit_predict(feature))
predict.columns=['predict']
r = pd.concat([feature,predict],axis=1)
# -
plt.scatter(r['Sepal length'],r['Sepal width'],c=data['labels'],alpha=0.7)
plt.title("Real")
plt.show()
plt.scatter(r['Sepal length'],r['Sepal width'],c=r['predict'],alpha=0.7)
plt.title("Predict")
plt.show()
| blog/MachineLearning/src/Day4_Eigenvalues and SVD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
import numpy as np
import scipy.stats as stats
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="whitegrid")
# -
# # Applications
#
# Although the techniques we discussed are very general, it turns out that *most* of the time, there are actually only a very few types of problems encountered regularly in Statistical Inference. The following section describes how to handle them using the Bootstrap for Bayesian inference.
#
# 1. **Compare two rates, usually for Bernoulli trials** For example, the purchase rate for A and the purchase rate for B or the cure rate for A and the cure rate for B). Very often, what we are interested in for this case is the difference in rates or *lift*.
# 2. **Compare two real valued parameters, usually means** For example, average purchase amount for A and the average purchase amount for B or the average weight of A versus the average rate of B.
# 3. **Compare a single rate against some proposed or analytical value** For example, we might believe or have heard that our open rate is 10%. We might need to do a study to find out if that is true.
# 4. **Compare a single real valued parameter against some proposed or analytical value** For example, we might have a machine with specs for a tolerance of $\frac{1}{10}$ of an inch. We might need to do a study to find out if that is true.
#
# We will cover each of these in turn using the Bayesian Bootstrap.
# +
import random
def resample( data):
n = len( data)
return [ data[ i] for i in [random.randint(0, n - 1) for _ in range( 0, n)]]
# -
# ## Comparing the $p$ of two experiments.
#
# This sort of comparison--between two rates--happens a lot which we will see later in the semester. During the course of your exploratory data analysis (EDA) you may find what appears to be a relationship between purchases by women as opposed to men. Say, woman purchase at a rate of 1.7% and men purchase at a rate of 1.3%.
#
# There are two things to ask yourself. Is the difference *meaningful*? Does it make a difference to your business, research, outreach, organizational goal that the women's rate is 30.7% higher than the men's? The second question is, how *believable* is it? You might have three years of data. Is your calculation from all three years of data? The last month? There's a trade-off between using a lot of data to calculate a parameter and the parameter having been changing over the course of time. After all, this analysis is situated in data that has come from a process that has been evolving over time.
#
# Statistical inference deals only with the believability of a result. It is up to the stakeholders to decide if the result is meaningful. As Spock said, "a difference that makes no difference is no difference."
#
# The examples that follow are very stock statistical inference problems but we will be solving them using the Bootstrap. We could easily solve them using the Grid or Monte Carlo methods.
#
# Suppose we have a drug, Smokestopperin, that we want to test. We randomly assign pack-a-day smokers to two groups. The control group receives a placebo and the treatment group receives 10mg capsules of the drug. After 90 days, we compare the results (We will talk about experimental design later in the semester).
smoking = {}
smoking[ "control"] = np.array([1] * 25 + [0] * 35)
smoking[ "treatment"] = np.array([1] * 35 + [0] * 15)
print("control", np.mean( smoking[ "control"]))
print("treatment", np.mean( smoking[ "treatment"]))
# It would appear that the treatment is very effective. The difference is *meaningful* but is it believable? We will now turn to that question.
#
# We're going to start by resampling each of the groups data and calculating the *rate* which is just the average of the booleans for each resampling.
smoking[ "control_resample"] = np.array([np.mean( data) for data in [resample( smoking[ "control"]) for i in range( 0, 10000)]])
smoking[ "treatment_resample"] = np.array([np.mean( data) for data in [resample( smoking[ "treatment"]) for i in range( 0, 10000)]])
# +
ys, bins = np.histogram( smoking[ "control_resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 2, 1)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([0, 1])
axes.set_xlabel( r"$\theta_1$")
axes.set_title( "Control")
ys, bins = np.histogram( smoking[ "treatment_resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
axes = figure.add_subplot(1, 2, 2)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([0, 1])
axes.set_xlabel( r"$\theta_2$")
axes.set_title( "Treatment")
plt.show()
plt.close()
# -
# The plots above show the posterior distributions for the two parameters, the smoking cessation rates of the placebo and Smokestopperin but what we really want to see is the *difference*. No problem! In the course of the Bootstrap, we generated many samples and means from those samples. We can simply pair up the samples from A and B and calculate the differences in the rates and plot them.
# +
smoking[ "difference"] = smoking[ "treatment_resample"] - smoking[ "control_resample"]
ys, bins = np.histogram( smoking[ "control_resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
figure = plt.figure(figsize=(20,6))
axes = figure.add_subplot(1, 3, 1)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([0, 1])
axes.set_xlabel( r"$\theta_1$")
axes.set_title( "Control")
ys, bins = np.histogram( smoking[ "treatment_resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
axes = figure.add_subplot(1, 3, 2)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([0, 1])
axes.set_xlabel( r"$\theta_2$")
axes.set_title( "Treatment")
ys, bins = np.histogram( smoking[ "difference"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
axes = figure.add_subplot(1, 3, 3)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([0, 1])
axes.set_xlabel( r"$\theta_2 - \theta_1$")
axes.set_title( "Difference")
plt.show()
plt.close()
# -
# Ideally, we would have defined what we meant by a "meaningful" difference when we'd set up the experiment and this would have influence our report to the stakeholders. If we decided that a meaningful difference would be at least 20 points, then we can report on the probability that the difference is greater than 20 points:
print( "P(difference > 20 points)", np.mean( smoking[ "difference"] > 0.20))
# One calculation that is commonly reported is the 95% CI (credible interval) or BCI (Bayesian Confidence Interval). The BCI is what you *think* the Frequentist *confidence interval* means. Bayesian statistics eschews reporting point estimates but they can be shown as well. There are a lot more options with Bayesian statistics. For example, depending on the cost of Stopsmokerin and the cost of not quitting smoking we might be interested in other intervals as well as expected values. If the cost of Stopsmokerin is low and the cost of not quitting is very high, we could even be interested in an "even bet" such as the 50% CI.
print( "95% CI", stats.mstats.mquantiles( smoking[ "difference"], [0.025, 0.975]))
# ## Comparing two real valued $\theta$s.
#
# Another common problem in statistical inference involves the two (or more) *real valued* parameters, $\theta$s. Interestingly enough while other approaches to Bayesian (and Frequentist) statistics would require you to use a different tool from the boolean approach, the Bootstrap does not. The process is the same because we are not required to specify either a test (Frequentist) or a distribution (Bayesian). We use the empirical distribution.
#
# Imagine a biologist is trying to discover whether the bacteria *acidophilus* or *bulgarius* is a better starter culture for yogurt measured in hours to ripen.
# +
yogurt = {}
yogurt[ "acidophilus"] = np.array( [6.8, 6.3, 7.4, 6.1, 8.2, 7.3, 6.9])
yogurt[ "bulgarius"] = np.array([6.1, 6.4, 5.7, 5.5, 6.9, 6.3, 6.7])
print( "acidophilus", np.mean( yogurt[ "acidophilus"]))
print( "bulgarius", np.mean( yogurt[ "bulgarius"]))
# -
# Again, in general, we need to have asked ourselves what difference is meaningful *in advance* of the experiment or collection of data or analysis of the data. Once we have the data, if we find a meaningful difference, we can ask ourselves if it is believable...but these are made up problems. Let's assume that when you're making thousands of gallons of yogurt at a time, an hour is meaningful.
#
# Currently, we see that the difference is less than hour. Is it believable? We thought that bulgarius would at least do as well as acidophilus and maybe even better (otherwise, why do the study?). Because we've already developed the individual steps, I'm going to do the graphing portion of the analysis all at once:
# +
yogurt[ "acidophilus_resample"] = np.array([np.mean( data) for data in [resample( yogurt[ "acidophilus"]) for i in range( 0, 10000)]])
yogurt[ "bulgarius_resample"] = np.array([np.mean( data) for data in [resample( yogurt[ "bulgarius"]) for i in range( 0, 10000)]])
yogurt[ "difference"] = yogurt[ "bulgarius_resample"] - yogurt[ "acidophilus_resample"]
ys, bins = np.histogram( yogurt[ "acidophilus_resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
figure = plt.figure(figsize=(20,6))
axes = figure.add_subplot(1, 3, 1)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([5, 9])
axes.set_xlabel( r"$\theta_1$")
axes.set_title( "Acidophilus")
ys, bins = np.histogram( yogurt[ "bulgarius_resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
axes = figure.add_subplot(1, 3, 2)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([5, 9])
axes.set_xlabel( r"$\theta_2$")
axes.set_title( "Bulgarius")
ys, bins = np.histogram( yogurt[ "difference"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
axes = figure.add_subplot(1, 3, 3)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([-5.0, 5.0])
axes.set_xlabel( r"$\theta_2 - \theta_1$")
axes.set_title( "Difference")
plt.show()
plt.close()
# -
# We can now report on the distribution of differences as before.
print("P(difference > 1 hour)", np.mean( yogurt[ "difference"] < -1.))
print("95% CI", stats.mstats.mquantiles( yogurt[ "difference"], [0.025, 0.975]))
# You have to be careful of the signs here. *Bulgarius* ripens quicker so a larger negative number is *better*.
#
# One of the disadvantages of the Bootstrap is that you will sometimes come up with a zero probability for some events (or certainty for others). These should be interpreted with caution. While the probability of negative height is truly zero, we should make sure that any event that is shown to have a zero probability is truly impossible.
#
# Finally, as this is a made up problem, it's okay but in general, we need 10-30 observations at least for the Bootstrap. Still, I'm surprised at how well it works.
# ## Comparing a boolean $\theta$ with a hypothesized or analytical value
#
# This case and the following case are often the ones that statistics textbooks start with. I think this is because statistics as we generally know it, especially Frequentist statistics, started with industrial processes. For example, the specs say this machine should have a defect rate of 1.2%. For the last run, based on a random sample, we saw a defect rate of 1.7%. Should we stop the line and fix the machine? This is *definitely* the kind of problem that Neyman/Pearson had in mind.
#
# But I generally find that these examples completely confuse many students. Where did the 1.2% come from? How did we get 1.7%?
#
# Another way of looking at this kind of problem is that someone made a claim about the world: that the machine had a 1.2% defect rate, and we want to verify it. This could just as easily be a marketing person saying our company has a 1.1% purchase rate or a drug company saying their drug is 73% effective. We may want to look at the data and determine if this is true.
#
# There are a number of ways this can be handled in Bayesian Statistics. For example, we might use our beliefs about the hypothesized value as a prior. Since we're using the Bootstrap method, we'll just report our results relative to this value.
#
# Suppose we're told that a drug is 73% effective at treating some condition in 21 days or less. We start using it in our practice. Over the course of a few years, we notice that while on paper this drug has the highest effectiveness, we often have to switch to a different drug for our patients. We go back through our charts since the drug was introduced and compile the following data:
drug = {}
drug[ "data"] = np.array( [1] * 139 + [0] * 67)
print("effectiveness", np.mean( drug[ "data"]))
# Hmm, 67% isn't *that* far off from 73%. It could be that we're just remembering all the the times we switched drugs because we had a 2nd visit from the patient, had to write a 2nd prescription, etc. Whether this difference is meaningful would depend on whether or not there are other treatments, their cost, their effectiveness, what happens if the condition is not treated (does it just become chronic? can you be treated multiple times)?
#
# We'll side step these here but they are the *most* important questions.
#
# Here's a basic Bootstrapping and summary of our findings:
# +
drug[ "resample"] = np.array([np.mean( data) for data in [resample( drug[ "data"]) for i in range( 0, 10000)]])
ys, bins = np.histogram( drug[ "resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 1, 1)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([0, 1])
axes.set_xlabel( r"$\theta$")
axes.set_title( "Drug Effectiveness")
plt.show()
plt.close()
# -
print("P(effectiveness > 73%)", np.mean( drug[ "resample"] >= 0.73))
print("P(70% <= effectiveness <= 76%)", np.mean((0.7 <= drug[ "resample"]) & (drug[ "resample"] <= 0.76)))
print("P(effective > 70%)", np.mean( 0.7 <= drug[ "resample"]))
print("P(effective < 70%)", np.mean( drug[ "resample"] <= 0.7))
print("95% CI", stats.mstats.mquantiles( drug[ "resample"], [0.025, 0.975]))
# If we take 73% as our benchmark, we can see that there's only a 4.2% probability that the drug is at least 73% effective based our data but that's pretty stringent. If we want to look "around" 73% say, 70-76%, then there is only a 20% our drug is effective "around" 73% of the time. If we look more closely, though, we can see that there's a 79.4% probability the drug is 70% effective or better. The 95% CI is 61.2-73.8% effective.
#
# This problem illustrates the richness of Bayesian over Frequentists approaches to statistical inference.
# ## Comparing a real valued $\theta$ with a hypothesized or analytical value
#
# As one might imagine, the above situation arises for real valued measurements as well as for boolean outcomes.
#
# A company advertises the breaking strength of the wire it manufacturers as 75N (Newtons). A customer takes 9 different rolls of wire and tests their breaking strength.
wire = {}
wire[ "data"] = np.array([ 72.1, 74.5, 72.8, 75, 73.4, 75.4, 76.1, 73.5, 74.1])
np.mean( wire[ "data"])
# Did the customer get a bad batch? Is the manufacturer misrepresenting its product?
#
# Here we need a bit of domain knowledge--which I don't have--how many Newtons are important? You need to start by identifying the meaningful difference. Now let's see if it's believable, based on the data.
# +
wire[ "resample"] = np.array([np.mean( data) for data in [resample( wire[ "data"]) for i in range( 0, 10000)]])
ys, bins = np.histogram( wire[ "resample"], bins=10, density=True)
width = 0.7*(bins[ 1] - bins[ 0])
center = (bins[ :-1]+bins[1:])/2
figure = plt.figure(figsize=(10,6))
axes = figure.add_subplot(1, 1, 1)
axes.bar( center, ys, align='center', width=width, color="dimgray", alpha=0.8)
axes.set_xlim([70.0, 80.0])
axes.set_xlabel( r"$\theta$")
axes.set_title( "Wire Strength (Newtons)")
plt.show()
plt.close()
# -
# What is the summary?
print("P(strength >= 75N)", np.mean( wire[ "resample"] >= 75))
print("95% CI", stats.mstats.mquantiles( wire[ "resample"], [0.025, 0.975]))
# Again, it depends on the use whether the difference is meaningful (and thus the comparison is valid) but if we assume that very small values in Newtons are important, there is a very, very low probability, based on the data, that the average strength is 75N.
# ## Summary
#
# You can use these as templates for asking similar questions. If you find a problem that isn't here, refer back to the section on the Bootstrap in general. We will definitely look at the Bootstrap in the context of other modeling techniques.
#
# Make sure you understand each of these canonical problems. Try to think of similar problems and see if you can solve them. If you work with synthetic data, as you learned in the chapter on Mathematical distributions, you'll have no end of data to work with as well as the ability to know what the correct answer should be.
#
# Remember. This is inference. It isn't going to be right 100% of the time.
| fundamentals_2018.9/inference/applications.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Enamel Thickness Boxplot
#
# The plot that will be reproduced here is a grouped boxplot of molar enamel thickness for humans, fossil hominins, and the extant great apes from Skinner et al. (2015) in JHE (doi: [10.1016/j.jhevol.2015.03.012](http://dx.doi.org/10.1016/j.jhevol.2015.03.012)). I'm a little unsure of some of the color and formatting choices in the published plot, but it's a good dataset and I want an example plot to go along with each of the entries in my bioanth datasets repository.
#
# 
#
# The data underlying this plot are included in the supplementary material of the paper and have been extracted and made available in my bioanth datasets repository.
#
# ## Preliminaries
#
# First, I always change the R option for importing strings (I find that the default convert to factor causes more problems than it solves).
options(stringsAsFactors=F)
# You can safely ignore this next option. The default image rendering option for the system I am using here (Jupyter Notebook) does not show italic annotations on plots properly; this option change fixes that. Standard R on your system will work as it should without this option change.
options(jupyter.plot_mimetypes = 'image/png')
# Next, import the data. The resulting data frame has nine variables: accession, tooth, enamel_area (mm$^2$), dentine_area (mm$^2$), edj_length (enamel-dentine junction, mm), bcd (bi-cervical diameter, mm), aet (average enamel thickness, mm), and ret (relative enamel thickness).
dset = read.csv(url("https://raw.githubusercontent.com/ryanraaum/bioanth-datasets/master/raw/enamelthickness.csv"))
str(dset)
# The full dataset includes some specimens with "Uncertain" taxonomic affiliation that are not included in the targeted boxplot and *A. aethiopicus* is also dropped. So it's easiest to subset out those samples for the plotting going forward.
dset2 = subset(dset, !(taxon %in% c("Uncertain", "A. aethiopicus")))
nrow(dset)
nrow(dset2)
# For the plot, the chimpanzee species need to be merged and an ordered factor will be useful to have everything plot in the same order. It will also be useful to have the tooth variable as a factor.
# +
# make up a mapping vector to get new labels
taxa = c("A. afarensis", "A. africanus", "A. anamensis", "Gorilla sp.",
"Homo sapiens ", "Pan paniscus", "Pan troglodytes", "Pongo sp.",
"Homo erectus", "Homo sp. indet.", "A. boisei", "A. robustus")
map_labels = c("Aafar", "Aafri", "Aana", "Gorilla",
"Hsap", "Pan", "Pan", "Pongo",
"Here", "Hsp", "Aboi", "Arob")
names(map_labels) = taxa
# new group labels for all specimens in the reduced data set
newgroups = map_labels[dset2$taxon]
# set up the desired order
desiredorder = c("Aana", "Aafar", "Aafri", "Aboi", "Arob", "Hsp",
"Here", "Hsap", "Pongo", "Gorilla", "Pan")
# make the new groups into an ordered factor
newgroups = factor(newgroups, levels=desiredorder, ordered=T)
# add it to the data frame
dset2$otaxon = newgroups
# make the tooth variable a factor
dset2$tooth = factor(dset2$tooth)
# -
# Set up colors, font styles, legend labels, and calculate positions for taxon labels on the plot.
# +
barcols = c("steelblue4", "forestgreen", "lightgoldenrod3")
keynames = c("First molar", "Second molar", "Third molar")
fontstyle = c(1,1,1,1,1,1,1,1,3,3,3) # for base R & lattice (1=normal, 3=italic)
fontformat = c(rep("plain", 8), rep("italic", 3)) # for ggplot2
# pull out group labels and calculate number in preparation for plotting each
groups = levels(dset2$otaxon)
ngroups = length(groups)
# calculate locations for annotations
xloc = 1:ngroups
yloc = aggregate(aet ~ otaxon, min, data=dset2, simplify=T)$aet-0.1
# -
# ## Base R
#
# Base R plotting does not readily support this kind of plot, but it can be done with some trickery. Specifically, each taxon will be plotted in a separate panel of a multi-panel plot, but the axes will be left off and it will appear to all be a single plot. The resulting plot looks right here, but is probably a little fragile to changing plot sizes and dimensions.
# +
# set up plot parameters, saving the defaults as 'oldpar'
oldpar = par(mfrow=c(1,ngroups), oma=c(2,8,0,1), mar=rep(0.5,4))
# need to use the same y-limits for each of the per-group plots
ylim = c(0, 3)
# plot out each group, adding an axis for the first one
for (i in 1:ngroups) {
gdata = subset(dset2, otaxon==groups[i])
boxplot(aet ~ tooth, data=gdata, ylim=ylim, axes=F, col=barcols, outline=F, lty=1)
text(2,min(gdata$aet)-0.1,groups[i], font=fontstyle[i])
if (i == 1){
axis(2, line=2, las=1, cex.axis=1.5)
mtext("Average Enamel Thickness (mm)", 2, 6, cex=1.25)
}
}
# reset plotting parameters back to saved defaults
par(oldpar)
# Getting the legend in place also requires some trickery
# It is necessary to place a new blank plot over the top of the
# existing barplots and then place the legend on that
# I also take advantage of this trickery to draw a box around the prior plotting area
# set up a new figure area with
# x- and y-limits from 0 to 1 (fig)
# outer margins set to bottom 2, left 3.8, top 0, right 0 (oma)
# - some trial and error to get those number to get the bounding box to plot right
# spacing around sub-figures set to 0.5 (mar)
# overplot on existing plot (new=TRUE)
oldpar = par(fig=c(0, 1, 0, 1), oma=c(2, 3.8, 0, 0), mar=rep(0.5,4), new=TRUE)
# add empty plot with no points and no axes, but draw bounding box
plot(0, 0, type='n', xaxt='n', yaxt='n')
# add the legend with no box (bty='n'), inset from the edge by 0.5
# and slightly more than usual spacing between the lines
legend("topright", fill=barcols, legend=keynames,
bty="n", inset=0.05, y.intersp=1.2)
# reset to saved default plotting parameters
par(oldpar)
# -
# ## ggplot2
#
# This plot is probably easiest in ggplot2, which supports it pretty seamlessly. Even with all the extra fiddling to attempt to exactly match the published plot, the code is pretty short.
# +
library(ggplot2)
# this part is probably overkill..
# ggplot2 uses only the necessary decimal points on axis labels,
# but the target plot has 2 decimal places, so if we want to exactly match that
# we need a formatting function
fmt = function(x) { format(x, nsmall=2) }
# start the plot, identify the data and variables
plt = ggplot(dset2, aes(x=otaxon, y=aet)) +
# split each x group out by tooth, with extra spacing (width), and no outliers
geom_boxplot(aes(fill=tooth), width=0.5, outlier.shape=NA) +
# manually set the colors to use and legend labels
scale_fill_manual(values=barcols,
labels=keynames) +
# configure the y axis with
# - custom title
# - set the limits
# - and use the formatter created above to ensure 2 decimal places on tick labels
scale_y_continuous(name="Average Enamel Thickness (mm)",
limits=c(0,3),
labels=fmt) +
# add the group names directly below each set of box-and-whiskers
annotate("text", x=xloc, y=yloc, label=groups, fontface=fontformat) +
# change to simple theme (theme_bw = theme black & white)
theme_bw() +
# make more appearance changes
# get rid of the grid
theme(panel.grid=element_blank(),
# change the size and add some spacing to the right of the y axis title
axis.title.y=element_text(size=14, margin=margin(0,15,0,0)),
# change the size of the y axis tick labels
axis.text.y=element_text(size=12),
# get rid of the x axis ticks
axis.ticks.x=element_blank(),
# get rid of the x axis tick labels
axis.text.x=element_blank(),
# get rid of the x axis title
axis.title.x=element_blank(),
# put the legend inside the plot area
legend.position=c(0.8,0.9),
# get rid of the legend title
legend.title=element_blank(),
# get rid of the boxes around the legend elements
legend.key=element_blank())
plt
# -
# ## lattice
#
# lattice also supports this kind of plot pretty seamlessly, but the way that it allows it is a little hard to wrap one's mind around. Basically, what lattice would generally like to do for this situation is do a series of smaller panels for each taxon or each tooth, but it is possible to tell it to instead layer those separate panels on top of each other (superpose).
# +
library(lattice)
# specify the information for the key
key = list(points = list(
pch = 22, # square with separate border and fill colors
cex = 1.5, # make it a bit bigger than default
col = "black", # square border color
fill=barcols), # square fill color
text = list(text=keynames, # legend labels
cex=0.8), # make them a bit smaller than default
corner=c(0.9,0.95), # put the key in the upper right corner
padding=2) # add a little extra space between entries
# start plotting, separating out aet values by taxon from the data
bwplot(aet~otaxon, data=dset2,
# break down each taxon by tooth
groups=tooth,
# set the y axis label
ylab=list(label="Average Enamel Thickness (mm)"),
# set the y axis limits
ylim=c(0,3),
# don't show the outliers
do.out=F,
# use a line for the median marker in the bar rather than a point
pch="|",
# make each bar 1/5 the default width
# - this is necessary to fit 3 bars and some intergroup spacing into
# the space where one bar would normally fit
box.width=1/5,
# do some customization of the plotting area
panel = function(x,y,...) {
# superpose indicates that we want to plot the different groups in the
# same panel, rather than splitting them across panels
# also set the fill for the different groups (teeth)
panel.superpose(x,y,fill=barcols,...)
# then, for each taxon
for (i in 1:ngroups) {
# add the group name below where the bars for that group will be
panel.text(xloc[i]-0.1, yloc[i], groups[i], cex=0.8, font=fontstyle[i])
}
},
# customize the plotting of each tooth group
panel.groups = function(x, y,..., group.number) {
# boxplots for each tooth, with the x values offset a bit so they don't overlap
panel.bwplot(x + (group.number-2.5)/5, y, ...)
},
# change the appearance of the box-and-whiskers to have
# - black outlines on the box
# - black, solid line whiskers
par.settings = list(box.rectangle=list(col="black"),
box.umbrella=list(col="black", lty=1)),
# don't draw x axis ticks, labels, or title
scales = list(x=list(draw=FALSE)),
# add the previously specified key
key = key
)
| Enamel Thickness in R.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **PYTHON**
# Python is a high-level programming language invented by <NAME>. It was published in 1991.
#
# Merits of the Python Programming language:
# Uses...:
#
# 1. Data Analysis and Data Visualization
# 2. Web Scrapping
# 3. Creating Database
# 4. Web development (server-side),
# 5. Software development,
# 6. Mathematics,
# 7. System scripting.
#
# 8. Used in data science for Machine learning, Artificial intelligence,and Data Modelling
#
# + active=""
# What can Python do?
#
# Python can be used on a server to create web applications.
# Python can be used alongside software to create workflows.
# Python can connect to database systems. It can also read and modify files.
# Python can be used to handle big data and perform complex mathematics.
# Python can be used for rapid prototyping, or for production-ready software development
# .
# Why Python?
#
# -Highly demanded
#
# -Easy to read and understand
#
# -Clean code syntax
#
# Python works on different platforms (Windows, Mac, Linux, Raspberry Pi, etc).
# Python has a simple syntax similar to the English language.
# Python has syntax that allows developers to write programs with fewer lines than some other programming languages.
# Python runs on an interpreter system, meaning that code can be executed as soon as it is written. This means that prototyping can be very quick.
# Python can be treated in a procedural way, an object-orientated way or a functional way.
#
#
# Python was designed for readability, and has some similarities to the English language with influence from mathematics.
# Python uses new lines to complete a command, as opposed to other programming languages which often use semicolons or parentheses.
# Python relies on indentation, using whitespace, to define scope; such as the scope of loops, functions and classes. Other programming languages often use curly-brackets for this purpose.
#
# -
# **Companies Using Python**
# <br>Google, <br>YouTube,<br> Pinterest,<br> Netflix,<br> Spotify,<br> Instagram,<br> DropBox ,<br> Facebook etc.....
| Python_Programming_language.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn import datasets
from sklearn.naive_bayes import MultinomialNB, GaussianNB
from sklearn.model_selection import train_test_split
data = datasets.load_wine()
# Checking the available data items
dir(data)
data.data[:3]
data.feature_names
data.target_names
data.target[:3]
# Converting the dataset into a dataframe
wine_df = pd.DataFrame(data.data, columns=data.feature_names)
wine_df.head()
wine_df['target'] = data.target
wine_df.head()
X_train, X_test, y_train, y_test = train_test_split(data.data, data.target, test_size=0.3, random_state=101)
model = GaussianNB()
model.fit(X_train, y_train)
model.score(X_test, y_test)
model = MultinomialNB()
model.fit(X_train, y_train)
model.score(X_test, y_test)
| Supervised/Classification/Wine_Category_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Qvexyww7a-0o"
# <div>
# <img src="https://drive.google.com/uc?export=view&id=1vK33e_EqaHgBHcbRV_m38hx6IkG0blK_" width="350"/>
# </div>
#
# #**Artificial Intelligence - MSc**
# CS6501 - MACHINE LEARNING AND APPLICATIONS
# #**Business Analytics - MSc**
# ET5003 - MACHINE LEARNING APPLICATIONS
# ##***Annual Repeat***
# ###Instructor: <NAME>
#
# ###RepMLA_Etivity-2
#
# + cellView="form" id="LqXD_IwUQuBF"
#@title Current Date
Today = '2021-06-28' #@param {type:"date"}
# + cellView="form" id="uzDKau31OjVO"
#@markdown ---
#@markdown ### Enter your details here:
Student_ID = "" #@param {type:"string"}
Student_full_name = "" #@param {type:"string"}
#@markdown ---
# + cellView="form" id="r39xGZckTpKx"
#@title Notebook information
Notebook_type = 'Example' #@param ["Example", "Lab", "Practice", "Etivity", "Assignment", "Exam"]
Version = 'Draft' #@param ["Draft", "Final"] {type:"raw"}
Submission = False #@param {type:"boolean"}
# + [markdown] id="1vrJoNOO5w0f"
# # Etivity-2
# + [markdown] id="vw_00xBJ5c6S"
# ## Introduction
# + [markdown] id="ZMPM1RVc6PeW"
# ### Guidelines
# Use this cell as a guideline and remove it when submitting the Etivity
#
# - ID and full name at the top of the file.
# - Explanation of the problem addressed.
# - Use mathematical notation and images.
# - Add any links to the references.
# - Write your work taking in consideration the reader.
# - Your work must be easy to understand and replicate.
# + [markdown] id="hbOf8jmJ51jQ"
# ## Dataset
# + [markdown] id="8Nzipx1Y8wGx"
#
# ### Guidelines
# Use this cell as a guideline and remove it when submitting the Etivity
#
# - Explain the dataset and its features.
# - Give the link or information to get the dataset.
# - Add comments about any preprocessing you used if any.
# - Explain what technique was used to split the dataset into train and test sets.
# - Show bar graphs and/or plots to graphically explain the dataset.
# + [markdown] id="WaH8zkcQ55cF"
# ## Method
# + [markdown] id="ZACjApsa9QFt"
# ### Guidelines
# Use this cell as a guideline and remove it when submitting the Etivity
#
# - Explain the method used to solve the given problem.
# - Use mathematical notation and images.
# - Add any links to the references.
# + [markdown] id="N-hBEkbO58ed"
# ## Summary
# + [markdown] id="xm0Ix3VL9RrS"
# ### Guidelines
# Use this cell as a guideline and remove it when submitting the Etivity
#
# - This must be the final section in your notebook right before the references.
# - Give a summary of the work you did.
# - It is very important to use your own words to write this section.
# - Use the structure to reference any section in your notebook.
# - Add the discussion you had with your peers about any of the topics.
# - Explain the pros and cons of the datasets, techniques, and methods used.
# - List all your findings and give any conclusion related to your insights.
# - Highlight in your work any hint, tip, issue or any useful comment to improve the work.
# + [markdown] id="mFdAvJpp6AYL"
# ## References
# + [markdown] id="JPGT1d7O9TH6"
# ### Guidelines
# Use this cell as a guideline and remove it when submitting the Etivity
#
# - Use the American Psychological Association (APA) citation style.
# - References of books
# - References of articles
# - References of url (websites)
| WEEK_2/RepMLA_Etivity_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from os import path
import numpy as np
all_acting_path = '../../data/imbd_data/all_acting.csv'
all_acting_clean_path = '../../data/imbd_data/all_acting_clean.csv'
# +
def load_dataset(filepath):
df = pd.read_csv(filepath)
return df
def write_to_csv(df,filepath):
'''
input: df - a pandas DataFrame
filepath - an output filepath as a string
writes to a csv file
in same diretory as this script
returns: nothing
'''
# if no csv exists
if not path.exists(filepath):
df.to_csv(filepath,index=False)
else:
df.to_csv(filepath, mode='a', header=False,index=False)
# -
# load dataset
df = load_dataset(all_acting_path)
print(df.head(1))
print(len(df[df.ceremony.notna()]))
# +
categories_interest = ['ACTOR',
'ACTRESS',
'ACTOR IN A LEADING ROLE',
'ACTRESS IN A LEADING ROLE',
'ACTOR IN A SUPPORTING ROLE',
'ACTRESS IN A SUPPORTING ROLE',float("NAN")]
# only include actors/actresses
df = df[df['category'].isin(categories_interest)]
print(len(df[df.ceremony.notna()]))
# -
print(min(df.age))
print(max(df.age))
#-61
#1955
# because of errors in database we will use the age range from the 8 to 88 years old
df = df[df.age >=1]
df = df[df.deathYear >= df.film_year]
print(min(df.deathYear))
print(max(df.age))
print(len(df.nconst.unique()))
print(len(df[df.ceremony.notna()]))
#remove people who have movies after death
print(min(df.num_times_nominated))
print(min(df.num_times_nominated))
print(min(df.age))
print(max(df.age))
write_to_csv(df,all_acting_clean_path)
print(len(df[df.ceremony.notna()]))
| code/analysis/.ipynb_checkpoints/all_acting_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression algorithms for predicting stock market price
# ### Authors:
# <NAME> <br>
# <NAME>
# ## Objective:
# Examine different regression algorithms to predict future stock prices based on the historical data for 5 companies.
# ## First step:
# Finding a proper dataset and performing data processing and cleaning.
# Data was extracted from Yahoo Finance dataset. We selected 5 preferred companies and extracted their historical daily data from 01/01/1990 until 10/11/2020 into different json files.
# +
# #!/usr/bin/env python
import requests
import json
try:
# For Python 3.0 and later
from urllib.request import urlopen
except ImportError:
# Fall back to Python 2's urllib2
from urllib2 import urlopen
# Create list of companies we are interested in
names = ['AAPL','GOOG','MSFT','INTC','AMZN']
# For each company extract the historical data from API
for i in names:
url = "https://financialmodelingprep.com"
search_query_format = "{}/{}&apikey={}"
key = "<KEY>"
url2 = "/api/v3/historical-price-full/"+i+"?from=1990-01-01&to=2020-11-10"
query = search_query_format.format(url, url2, key )
r = requests.get(query)
# Create a json file with the data of each company
json_string = r.json()
with open("data_file_"+i+".json", "w") as write_file:
json.dump(json_string, write_file)
print("data_file_"+i+".json", "was created")
# -
# **In order to use the data with our Regression algorithm the data from the json files had to be converted to "csv" and "libsvm" formats**
# +
# Convert JSON file to CSV
import json
import csv
names = ['AAPL','GOOG','MSFT','INTC','AMZN']
# For each company a csv file is created
for i in names:
# Opening JSON file and loading the data into the variable data
with open('data_file_'+i+'.json') as json_file:
data = json.load(json_file)
company_data = data['historical']
# Open a file for writing
data_file = open('data_file_'+i+'.csv', 'w')
# Create the csv writer object
csv_writer = csv.writer(data_file)
# Counter variable used for writing
# headers to the CSV file
count = 0
for company in company_data:
if count == 0:
# Writing headers of CSV file
header = company.keys()
csv_writer.writerow(header)
count += 1
# Writing data of CSV file
csv_writer.writerow(company.values())
print('data_file_'+i+'.csv', 'was created')
data_file.close()
# +
# Convert JSON to LIBSVM
names = ["AAPL","GOOG",'MSFT','INTC',"AMZN"]
for i in names:
with open("data_file_"+i+".json") as file:
data = json.load(file)
with open("data_file_"+i+".txt", "w") as file:
for item in data['historical']:
file.write(str(item['close']) + ' 1:' + str(item['high']) + ' 2:' + str(item['low'])+ ' 3:' + str(item['open'])+ ' 4:' + str(item['adjClose'])
+ ' 5:' + str(item['volume']) + ' 6:' + str(item['unadjustedVolume']) + ' 7:' + str(item['change']) + ' 8:' + str(item['changePercent']) + ' 9:' + str(item['vwap'])
+ ' 10:' + str(item['changeOverTime']) +'\n')
# Convert JSON to CSV
with open("data_file_"+i+".csv", "w") as file:
header = 'label,high,low,open,adjClose,volume,unadjustedVolume,change,changePercent,vwap,changeOverTime'
file.write(header+'\n')
for item in data['historical']:
file.write(str(item['close']) + ',' + str(item['high']) + ',' + str(item['low'])+ ',' + str(item['open'])+ ',' + str(item['adjClose'])
+ ',' + str(item['volume']) + ',' + str(item['unadjustedVolume']) + ',' + str(item['change']) + ',' + str(item['changePercent']) + ',' + str(item['vwap'])
+ ',' + str(item['changeOverTime'])+'\n')
# -
# ## Data cleaning
# +
from pyspark.ml import Pipeline
from pyspark.ml.regression import RandomForestRegressor
from pyspark.ml.feature import VectorIndexer
from pyspark.ml.feature import VectorAssembler
from pyspark.sql import SparkSession
from pyspark.ml.regression import LinearRegression
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml.regression import GBTRegressor
from pyspark.ml.tuning import CrossValidator
from pyspark.ml.evaluation import RegressionEvaluator
from pyspark.ml.tuning import ParamGridBuilder
import numpy as np
import matplotlib.pyplot as plt
from pyspark.ml.stat import Correlation
import pandas as pd
import seaborn as sns
#Create Spark session
if __name__ == "__main__":
spark_session = SparkSession \
.builder \
.appName("Spark Regression") \
.getOrCreate()
# Loads data in libsvm format
names = ["AAPL","GOOG"]
for i in names:
dataset = spark_session \
.read \
.format("libsvm") \
.load("data_file_"+i+".txt")
#Print the number of entries in the dataset
print("Number of entries in dataset", dataset.count())
# delete duplicate rows
dataset = dataset.distinct()
#dropping rows that have empty values
dataset = dataset.dropna()
print("Number of entries after cleaning", dataset.count())
# -
# ### Print the correlation between the features in the dataset using Pearson and Spearman correlation. One difference between them is that the Pearson coefficient works with a linear relationship between the variables whereas the Spearman Coefficient works with monotonic relationships as well.
#
# +
features = dataset.select("features")
np.set_printoptions(linewidth=np.inf)
head_values = [["open","high","low","adjClose","volume","unadjustedVolume","change","changePercent","vwap","changeOverTime"]]
# Correlation for libsvm data
r1 = Correlation.corr(features, "features").head()
print("Pearson correlation matrix:\n" )
matrix = head_values
matrix.append(r1[0].toArray())
#r1.collect()[0]["pearson({})".format("features")].values
for row in matrix:
print(row)
print()
###########
df_correlation_matrix = pd.DataFrame(r1[0].toArray(),columns=head_values[0],index=head_values[0])
fig, ax = plt.subplots(figsize=(10,10))
sns.heatmap(df_correlation_matrix.abs(), annot = True)
plt.show()
###########
r2 = Correlation.corr(features, "features", "spearman").head()
print("Spearman correlation matrix:\n")# + str(r2[0]))
#spearman_matrix = head_values
##spearman_matrix = r2[0].toArray()
for row in spearman_matrix:
print(row)
print()
###########
df_correlation_matrix_2 = pd.DataFrame(r2[0].toArray(),columns=head_values[0],index=head_values[0])
fig, ax = plt.subplots(figsize=(10,10))
sns.heatmap(df_correlation_matrix_2.abs(), annot = True)
plt.show()
###########
# -
df_2 = df_correlation_matrix_2.abs()
df_2 > 0.8
# ## Second step:
# Implementing the regression algorithms
# ### Implementing Linear Regression and Gradient Boosting algorithms trained on a the processed dataset
dataset.printSchema()
dataset.show()
# Split the data into training and test sets (30% held out for testing)
(trainingData, testData) = dataset.randomSplit([0.7, 0.3])
#Create linear regression model
lr = LinearRegression(maxIter=10, regParam=0.3, elasticNetParam=0.8)
# Fit the model
lrModel = lr.fit(trainingData)
# Print the coefficients and intercept for linear regression
print("Coefficients: %s" % str(lrModel.coefficients))
print("Intercept: %s" % str(lrModel.intercept))
# Summarize the model over the training set and print out some metrics
trainingSummary = lrModel.summary
print("numIterations: %d" % trainingSummary.totalIterations)
print("objectiveHistory: %s" % str(trainingSummary.objectiveHistory))
trainingSummary.residuals.show()
print("RMSE: %f" % trainingSummary.rootMeanSquaredError)
print("r2: %f" % trainingSummary.r2)
lr_predictions = lrModel.transform(testData)
lr_predictions.select("prediction", "label", "features").show(5)
lr_evaluator = RegressionEvaluator(predictionCol="prediction", metricName="r2")
print("R Squared (R2) on test data = %g" % lr_evaluator.evaluate(lr_predictions))
print("Using Gradient-boosted tree regression")
gbt = GBTRegressor(featuresCol='features', labelCol='label', maxIter=10)
gbt_model = gbt.fit(trainingData)
gbt_predictions = gbt_model.transform(testData)
gbt_predictions.select('prediction', 'label', 'features').show(5)
lr_evaluator = RegressionEvaluator(predictionCol="prediction", metricName="r2")
print("R Squared (R2) on test data = %g" % lr_evaluator.evaluate(gbt_predictions))
spark_session.stop()
| notebooks/Classification/Prototype_Documentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 임포트
# +
import pandas as pd
from glob import glob
import os
from shutil import copyfile
from torch.utils.data import Dataset
from PIL import Image
import numpy as np
from numpy.random import permutation
import matplotlib.pyplot as plt
from torchvision import transforms
from torchvision.datasets import ImageFolder
from torchvision.models import resnet18,resnet34,densenet121
from torchvision.models.inception import inception_v3
from torch.utils.data import DataLoader
from torch.autograd import Variable
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import pickle
# %matplotlib inline
# -
is_cuda = torch.cuda.is_available()
is_cuda
# ## 유틸리티 함수
# +
def imshow(inp,cmap=None):
"""Imshow for Tensor."""
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.imshow(inp,cmap)
class FeaturesDataset(Dataset):
def __init__(self,featlst1,featlst2,featlst3,labellst):
self.featlst1 = featlst1
self.featlst2 = featlst2
self.featlst3 = featlst3
self.labellst = labellst
def __getitem__(self,index):
return (self.featlst1[index],self.featlst2[index],self.featlst3[index],self.labellst[index])
def __len__(self):
return len(self.labellst)
def fit(epoch,model,data_loader,phase='training',volatile=False):
if phase == 'training':
model.train()
if phase == 'validation':
model.eval()
volatile=True
running_loss = 0.0
running_correct = 0
for batch_idx , (data1,data2,data3,target) in enumerate(data_loader):
if is_cuda:
data1,data2,data3,target = data1.cuda(),data2.cuda(),data3.cuda(),target.cuda()
data1,data2,data3,target = Variable(data1,volatile),Variable(data2,volatile),Variable(data3,volatile),Variable(target)
if phase == 'training':
optimizer.zero_grad()
output = model(data1,data2,data3)
loss = F.cross_entropy(output,target)
running_loss += F.cross_entropy(output,target,size_average=False).data
preds = output.data.max(dim=1,keepdim=True)[1]
running_correct += preds.eq(target.data.view_as(preds)).cpu().sum()
if phase == 'training':
loss.backward()
optimizer.step()
loss = running_loss/len(data_loader.dataset)
accuracy = 100. * running_correct.item()/len(data_loader.dataset)
print(f'{phase} loss is {loss:{5}.{2}} and {phase} accuracy is {running_correct}/{len(data_loader.dataset)}{accuracy:{10}.{4}}')
return loss,accuracy
class LayerActivations():
features=[]
def __init__(self,model):
self.features = []
self.hook = model.register_forward_hook(self.hook_fn)
def hook_fn(self,module,input,output):
#out = F.avg_pool2d(output, kernel_size=8)
self.features.extend(output.view(output.size(0),-1).cpu().data)
def remove(self):
self.hook.remove()
# -
# ## PyTorch 데이터셋 생성
data_transform = transforms.Compose([
transforms.Resize((299,299)),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
# For Dogs & Cats dataset
train_dset = ImageFolder('../../Chapter03/dogsandcats/train/',transform=data_transform)
val_dset = ImageFolder('../../Chapter03/dogsandcats/valid/',transform=data_transform)
classes=2
imshow(train_dset[150][0])
# ## 학습과 검증 데이터셋을 위한 데이터 로더 생성
train_loader = DataLoader(train_dset,batch_size=32,shuffle=False,num_workers=3)
val_loader = DataLoader(val_dset,batch_size=32,shuffle=False,num_workers=3)
# ## 모델 생성
# +
# ResNet 모델 생성
my_resnet = resnet34(pretrained=True)
if is_cuda:
my_resnet = my_resnet.cuda()
my_resnet = nn.Sequential(*list(my_resnet.children())[:-1])
for p in my_resnet.parameters():
p.requires_grad = False
# Inception 모델 생성
my_inception = inception_v3(pretrained=True)
my_inception.aux_logits = False
if is_cuda:
my_inception = my_inception.cuda()
for p in my_inception.parameters():
p.requires_grad = False
# 덴스넷 모델 생성
my_densenet = densenet121(pretrained=True).features
if is_cuda:
my_densenet = my_densenet.cuda()
for p in my_densenet.parameters():
p.requires_grad = False
# -
# ## ResNet , Inception에서 DenseNet 컨볼루션 피처 추출
# +
### ResNet
trn_labels = []
trn_resnet_features = []
for d,la in train_loader:
o = my_resnet(Variable(d.cuda()))
o = o.view(o.size(0),-1)
trn_labels.extend(la)
trn_resnet_features.extend(o.cpu().data)
val_labels = []
val_resnet_features = []
for d,la in val_loader:
o = my_resnet(Variable(d.cuda()))
o = o.view(o.size(0),-1)
val_labels.extend(la)
val_resnet_features.extend(o.cpu().data)
### Inception
trn_inception_features = LayerActivations(my_inception.Mixed_7c)
for da,la in train_loader:
_ = my_inception(Variable(da.cuda()))
trn_inception_features.remove()
val_inception_features = LayerActivations(my_inception.Mixed_7c)
for da,la in val_loader:
_ = my_inception(Variable(da.cuda()))
val_inception_features.remove()
### Densenet(덴스넷)
trn_densenet_features = []
for d,la in train_loader:
o = my_densenet(Variable(d.cuda()))
o = o.view(o.size(0),-1)
trn_densenet_features.extend(o.cpu().data)
val_densenet_features = []
for d,la in val_loader:
o = my_densenet(Variable(d.cuda()))
o = o.view(o.size(0),-1)
val_densenet_features.extend(o.cpu().data)
# -
# ## 학습과 검증 데이터셋 생성
trn_feat_dset = FeaturesDataset(trn_resnet_features,trn_inception_features.features,trn_densenet_features,trn_labels)
val_feat_dset = FeaturesDataset(val_resnet_features,val_inception_features.features,val_densenet_features,val_labels)
# ## 학습과 검증 데이터로드 생성
trn_feat_loader = DataLoader(trn_feat_dset,batch_size=64,shuffle=True)
val_feat_loader = DataLoader(val_feat_dset,batch_size=64)
# ## 앙상블 모델 생성
class EnsembleModel(nn.Module):
def __init__(self,out_size,training=True):
super().__init__()
self.fc1 = nn.Linear(8192,512)
self.fc2 = nn.Linear(131072,512)
self.fc3 = nn.Linear(82944,512)
self.fc4 = nn.Linear(512,out_size)
def forward(self,inp1,inp2,inp3):
out1 = self.fc1(F.dropout(inp1,training=self.training))
out2 = self.fc2(F.dropout(inp2,training=self.training))
out3 = self.fc3(F.dropout(inp3,training=self.training))
out = out1 + out2 + out3
out = self.fc4(F.dropout(out,training=self.training))
return out
em = EnsembleModel(2)
if is_cuda:
em = em.cuda()
optimizer = optim.Adam(em.parameters(),lr=0.01)
# ## 앙상블 모델 학습
train_losses , train_accuracy = [],[]
val_losses , val_accuracy = [],[]
for epoch in range(1,10):
epoch_loss, epoch_accuracy = fit(epoch,em,trn_feat_loader,phase='training')
val_epoch_loss , val_epoch_accuracy = fit(epoch,em,val_feat_loader,phase='validation')
train_losses.append(epoch_loss)
train_accuracy.append(epoch_accuracy)
val_losses.append(val_epoch_loss)
val_accuracy.append(val_epoch_accuracy)
| Chapter08/ensembles/Ensemble_Dogs_and_cats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W3D4_ReinforcementLearning/W3D4_Intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a> <a href="https://kaggle.com/kernels/welcome?src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W3D4_ReinforcementLearning/W3D4_Intro.ipynb" target="_parent"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" alt="Open in Kaggle"/></a>
# + [markdown] pycharm={"name": "#%% md\n"}
# # Intro
# -
# **Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
#
# <p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Overview
#
# Reinforcement Learning (RL) is a framework for defining and solving a learning problem where an animal or agent knows or infers the state of the world and then learns the value of the states and actions that can be taken in them, by receiving a reward signal. Importantly, reinforcement learning provides formal, optimal descriptions of learning first derived from studies of animal behavior and then validated when the formal quantities used in the model were observed in the brain in humans and animals. It is probably one of the most widely used computational approaches in neuroscience.
#
# In the following tutorials, you will learn the core concepts of reinforcement learning and explore some of the most widely used models. In tutorial 1, you will learn about how we learn the value of future states from our experience. In tutorial 2, you will learn about how to make and learn from actions and the explore-exploit dilemma. In tutorial 3, you will explore how we can efficiently learn the future value of actions from experience. Finally, in tutorial 4 you will learn how having a model of the world’s dynamics can help you to learn and act.
#
# Reinforcement learning is a broad framework and it has deep connections to many topics covered in NMA, but the core reinforcement learning approaches define the world as a Markov Decision Problem, which is built on Hidden Dynamics and Optimal Control. Reinforcement learning, more broadly, can be seen as a framework that allows us to bring in many ideas and formalisms from other areas like economics, psychology, computer science, artificial intelligence, etc. to define algorithms or models that can solve large, complex problems with only a simple reward signal.
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Video
# + cellView="form" pycharm={"name": "#%%\n"}
# @markdown
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV1MC4y1b7hq", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"fz5T2QhUjbY", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Slides
# + cellView="form" pycharm={"name": "#%%\n"}
# @markdown
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/cpxqn/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
| tutorials/W3D4_ReinforcementLearning/W3D4_Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Jupyter Example 2 for HERMES: PiZero and Dragon2D
# +
from pyhermes import *
from pyhermes.units import TeV, deg, kpc, pc, radian2degree
import astropy.units as u
import numpy as np
import healpy
import matplotlib.pyplot as plt
# -
nside = 512
sun_pos = Vector3QLength(8.0*kpc, 0*pc, 0*pc)
Egamma = 0.1*TeV
dragon2D_proton = cosmicrays.Dragon2D(Proton)
dragon2D_helium = cosmicrays.Dragon2D(Helium)
cr_list = [dragon2D_proton, dragon2D_helium]
kamae_crosssection = interactions.Kamae06Gamma()
neutral_gas_HI = neutralgas.RingModel(neutralgas.RingType.HI)
integrator = PiZeroIntegrator(cr_list, neutral_gas_HI, kamae_crosssection)
integrator.setupCacheTable(120, 120, 24)
integrator.setSunPosition(sun_pos)
skymap = GammaSkymap(nside=nside, Egamma=Egamma)
top_left_edge = [5*deg, 20*deg]
bottom_right_edge = [-5*deg, 60*deg]
mask = RectangularWindow(top_left_edge, bottom_right_edge)
skymap.setMask(mask)
skymap.setIntegrator(integrator)
skymap.compute()
hermes_map = np.array(skymap)
#use_units = skymap.getUnits() # default units for GammaSkymap (GeV^-1 m^-2 s^-1 sr^-1)
use_units = "TeV^-1 cm^-2 s^-1 sr^-1"
skymap_units = u.Quantity(1, use_units)
base_units = skymap_units.unit.si.scale
healpy.mollview(hermes_map, cbar=False, title=r"$\pi^0$ Gamma Skymap")
# +
fig = plt.figure(figsize=(7,3))
nside = skymap.getNside()
mask_edges = [top_left_edge, bottom_right_edge]
mask_lon_in_deg = [float(radian2degree(a)) for a in [mask_edges[0][1], mask_edges[1][1]]]
mask_lat_in_deg = [float(radian2degree(a)) for a in [mask_edges[1][0], mask_edges[0][0]]]
projector = healpy.projector.CartesianProj(flipconv="geo")
projector.set_proj_plane_info(xsize=1000, ysize=None,
lonra=mask_lon_in_deg,
latra=mask_lat_in_deg
)
projected_map = projector.projmap(
hermes_map,
lambda x, y, z: healpy.pixelfunc.vec2pix(nside, x, y, z, nest=False)
)
w = ~(np.isnan(projected_map) | np.isinf(projected_map) | healpy.pixelfunc.mask_bad(projected_map, badval=UNSEEN))
projected_map[w] *= base_units
vmin = projected_map[w].min()
vmax = projected_map[w].max()
img = plt.imshow(projected_map, extent=mask_lon_in_deg + mask_lat_in_deg, cmap='jet', interpolation="nearest", origin="lower",
vmin=vmin, vmax=vmax)
plt.title(r"$\pi^0$, HI, $E_\gamma=${Egamma}, nside={nside}".format(
nside=nside, Egamma=np.around(Egamma.toAstroPy().to("TeV"), decimals=2).to_string(format="latex")))
plt.xlabel(r'${\rm Longitude} \; l \; {\rm [deg]}$')
plt.ylabel(r'${\rm Latitude} \; b \; {\rm [deg]}$')
cb = fig.colorbar(img, orientation='horizontal', shrink=.5, pad=0.25, format='%.0e')
cb.ax.xaxis.set_label_text(skymap_units.unit.to_string(format='latex_inline'))
cb.solids.set_edgecolor("face")
# -
| jupyter/e2-pizero-dragon2d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Custom Header Routing with Seldon and Ambassador
#
# This notebook shows how you can deploy Seldon Deployments which can have custom routing via Ambassador's custom header routing.
#
# ## Prerequistes
#
# You will need
#
# - [Git clone of Seldon Core](https://github.com/SeldonIO/seldon-core) running this notebook
# - A running Kubernetes cluster with kubectl authenticated
# - [Helm client](https://helm.sh/)
# - Seldon Core Python Module : `pip install seldon-core`
# ### Creating a Kubernetes Cluster
#
# Follow the [Kubernetes documentation to create a cluster](https://kubernetes.io/docs/setup/).
#
# Once created ensure ```kubectl``` is authenticated against the running cluster.
# ## Setup
# !kubectl create namespace seldon
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
# !kubectl create clusterrolebinding kube-system-cluster-admin --clusterrole=cluster-admin --serviceaccount=kube-system:default
# ## Install Helm
# !kubectl -n kube-system create sa tiller
# !kubectl create clusterrolebinding tiller --clusterrole cluster-admin --serviceaccount=kube-system:tiller
# !helm init --service-account tiller
# !kubectl rollout status deploy/tiller-deploy -n kube-system
# ## Start seldon-core
# !helm install ../../../helm-charts/seldon-core-operator --name seldon-core --set usageMetrics.enabled=true --namespace seldon-system
# !kubectl rollout status statefulset.apps/seldon-operator-controller-manager -n seldon-system
# ## Setup Ingress
# There are gRPC issues with the latest Ambassador, so we rewcommend 0.40.2 until these are fixed.
# !helm install stable/ambassador --name ambassador --set crds.keep=false
# !kubectl rollout status deployment.apps/ambassador
# ### Port Forward to Ambassador
#
# ```
# kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080
# ```
# ## Launch main model
#
# We will create a very simple Seldon Deployment with a dummy model image `seldonio/mock_classifier:1.0`. This deployment is named `example`.
# !pygmentize model.json
# !kubectl create -f model.json
# !kubectl rollout status deploy/production-model-single-7cd068f
# ### Get predictions
from seldon_core.seldon_client import SeldonClient
sc = SeldonClient(deployment_name="example",namespace="seldon")
# #### REST Request
r = sc.predict(gateway="ambassador",transport="rest")
print(r)
# #### gRPC Request
r = sc.predict(gateway="ambassador",transport="grpc")
print(r)
# ## Launch Model with Custom Routing
#
# We will now create a new graph for our Canary with a new model `seldonio/mock_classifier_rest:1.1`. To make it a canary of the original `example` deployment we add two annotations
#
# ```
# "annotations": {
# "seldon.io/ambassador-header":"location:london"
# "seldon.io/ambassador-service-name":"example"
# },
# ```
#
# The first annotation says we want to route traffic that has the header `location:london`. The second says we want to use `example` as our service endpoint rather than the default which would be our deployment name - in this case `example-canary`. This will ensure that this Ambassador setting will apply to the same prefix as the previous one.
# !pygmentize model_with_header.json
# !kubectl create -f model_with_header.json
# !kubectl rollout status deploy/header-model-single-4c8805f
# Check a request without a header goes to the existing model.
r = sc.predict(gateway="ambassador",transport="rest")
print(r)
# Check a REST request with the required header gets routed to the new model.
r = sc.predict(gateway="ambassador",transport="rest",headers={"location":"london"})
print(r)
# Now do the same checks with gRPC
r = sc.predict(gateway="ambassador",transport="grpc")
print(r)
r = sc.predict(gateway="ambassador",transport="grpc",headers={"location":"london"})
print(r)
| examples/ambassador/headers/ambassador_headers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting Started
# This notebook is for a short demonstration of Python for statistics.
#
# The notebook covers the following points:
# - Generating random numbers
# - Probability distributions
# - Curve fitting
# - Simple statistical anlysis with NetCDF files
# ## Import Python modules
#
# Special '%matplotlib inline' command should go first, before the matplotlib import
# %matplotlib inline
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
import time
from datetime import datetime, timedelta
from pathlib import Path
# ## Random numbers
# Get a random integer in the specified range
np.random.randint(0, 10)
# Get a random element from a sequence
np.random.choice([2, 4, 6, 9])
# Get a random real number between 0 and 1
np.random.random()
# Get a random real number in the specified range
np.random.uniform(0, 10)
# ## Probability distributions
# https://docs.scipy.org/doc/scipy/reference/tutorial/stats/continuous.html
#
# https://docs.scipy.org/doc/numpy-1.14.0/reference/routines.random.html
# +
np.random.seed(122) # Set the seed to an arbitrary value Set the
[np.random.uniform(0,10) for x in range(6)]
# -
# gaussian distribution with mean = 2.0, and a standard deviation of 1.0:
gaussian = stats.norm(loc=4.0, scale=1.0)
x = np.linspace(0.0, 8.0, 100)
y = gaussian.pdf(x)
plt.plot(x,y);
# Cumulative distribution
z = gaussian.cdf(x)
plt.plot(x,z);
# For hypothesis testing, one often needs the p-value. For example, for the given gaussian distribution above, what would be the x-value so that P(X <= x) = 0.95?
gaussian.ppf(0.95)
x = gaussian.rvs(1000);
# density --> If True, the first element of the return tuple will be the counts normalized to form a probability density,
# i.e., the area (or integral) under the histogram will sum to 1.
plt.hist(x, bins=50, density=True);
# ## Curve fitting
# +
# Seed the random number generator for reproducibility
np.random.seed(0)
x_data = np.linspace(-5, 5, num=50)
y_data = 2.9 * np.sin(1.5 * x_data) + np.random.normal(size=50)
# And plot it
plt.figure(figsize=(6, 4));
plt.scatter(x_data, y_data);
# +
from scipy import optimize
def test_func(x, a, b):
return a * np.sin(b * x)
params, params_covariance = optimize.curve_fit(test_func, x_data, y_data, p0=[2, 2])
print(params); # param[0] and param[1] are the mean and the standard deviation
# +
plt.figure(figsize=(6, 4))
plt.scatter(x_data, y_data, label='Data')
plt.plot(x_data, test_func(x_data, params[0], params[1]),
label='Fit function', color='r')
plt.legend(loc='best')
plt.show()
# +
from scipy.stats import norm
# picking 500 of from a normal distrubution
# with mean 0 and standard deviation 1
samp = norm.rvs(loc=0, scale=True, size=500) # rvs --> Random variates
param = norm.fit(samp) # distribution fitting
# generate 100 numbers between -5 and 5
x = np.linspace(-5, 5, 100)
# fitted distribution
pdf_fitted = norm.pdf(x, loc=param[0], scale=param[1])
# original distribution
pdf = norm.pdf(x)
plt.figure(figsize=(8, 6))
plt.title('Normal distribution')
plt.plot(x, pdf_fitted, 'r-', label='fitted distribution')
plt.plot(x, pdf, 'b-', label='original distribution')
plt.hist(samp, density=True, alpha=.3, label='sample')
plt.legend(loc='best', frameon=False)
plt.show()
# -
# # Exploring NetCDF data
import netCDF4
# # Read dataset
#
# This uses the pathlib.Path class, which can make working with filenames, directories and paths overall slightly more convenient
directory = Path("../../_data")
directory /= "cmip5/rcp85/Amon/pr/HadGEM2-ES/r1i1p1"
path = directory / "pr_Amon_HadGEM2-ES_rcp85_r1i1p1_200512-203011.nc"
dataset = netCDF4.Dataset(path, "r")
# # Discover the data
#
# A dataset when listed at the end of a cell already shows a lot of information.
dataset
# Below, we access some variables separately
pr = dataset['pr']
lat = dataset['lat']
lon = dataset['lon']
time = netCDF4.num2date(dataset['time'][:], dataset['time'].units)
time[0]
# Shows a tuple of a 1-element tuples, one for each shape
time.shape, lat.shape, lon.shape
# pr_Amon_HadGEM2-ES_rcp85_r1i1p1_200512-203011.nc
# 2005 December to 2030 November
pr.shape, pr.size, "Number of months = " + str((2030-2005)*12)
# # Getting the actual data
# To access the actual data from a dataset as a NumPy array, we need to actually access and copy the data, using the `[:]` operation.
# All variables above, `pr`, `lat`, `lon` and `time`, are still netCDF *variables*, not arrays. This has the convenience that we can directly access their netCDF attributes, and we avoid reading the full data into memory. The disadvantage is that we can't use them fully as a NumPy array. The code below tries to show the difference.
pr.ncattrs()
pr.units
# Trying to use it as a NumPy array will not work:
try:
pr.max()
except AttributeError as exc:
print(exc)
# But, if we, or the function, *requires* the data to be an array, it does work, since `np.max` takes an array.
# This also forces the data to be read into memory. (Under the hood, the data will be copied somewhere.)
#
# Usually, leaving `pr` and friends as a netCDF variable works for a lot of functionality.
np.max(pr)
# `pr` itself is still a netCDF variable
pr
# We now *permanently* turn `pr` into a NumPy array, by forcing a copy. Note that `[:]` also works for multi-dimensional variables.
# We lose all the netCDF variables attributes and methods for it, but gain the NumPy ones.
pr = pr[:]
# `pr` is now a NumPy array. A masked array, since there may be missing values.
pr
# There is one problem with the above `pr = pr[:]` assignment: if we nog go back a few cells in our notebook, where `pr` was still a netCDF variable, these cells won't work (apart from some functionality that NumPy arrays and netCDF variables have in common): our `pr` Python variable has changed type (and value, essentially). We would have to reassign `pr` to `dataset.variables['pr']` again for things to work normally.
# So be careful with reassigning variable names in notebooks in general: it will likely make your notebook less
# interactive, and more (forced) linear.
#
# Let's reset `pr` to be a variable, and use that instead of the NumPy array in our further analysis
pr = dataset['pr']
# ### Getting percentile values
np.percentile(pr, 99), np.max(pr), np.percentile(pr, 1), np.min(pr)
# ## Plotting the data
import matplotlib.pyplot as plt
import matplotlib.colors as colors
# +
# import cartopy.crs as ccrs
# import cartopy.io.shapereader as shpreader
# from cartopy.mpl.ticker import LongitudeFormatter, LatitudeFormatter
# -
# Find an interesting date
time[178]
plt.figure(figsize=(20, 10))
plt.imshow(pr[178,:,:]);
# ## Fit 2D gaussian
print(np.ndim(pr[178,:,:]))
pr[178,0:5,0:5]
# +
# curvefit with non linear least squares (curve_fit function)
from scipy.optimize import curve_fit
def func(x, a, b, c):
return a * np.sin(x[0]) + b * np.cos(x[1]) + c
limits = [0, 2*np.pi, 0, 2*np.pi] # [x1_min, x1_max, x2_min, x2_max]
side_x = np.linspace(limits[0], limits[1], 100)
side_y = np.linspace(limits[2], limits[3], 100)
X1, X2 = np.meshgrid(side_x, side_y)
size = X1.shape
x1_1d = X1.reshape((1, np.prod(size)))
x2_1d = X2.reshape((1, np.prod(size)))
xdata = np.vstack((x1_1d, x2_1d))
original = (3, 1, 0.5)
z = func(xdata, *original)
Z = z.reshape(size)
z_noise = z + .2*np.random.randn(len(z))
Z_noise = z_noise.reshape(size)
ydata = z_noise
popt, pcov = curve_fit(func, xdata, ydata)
# Python 3.6 & later f-strings: insert variables directly inside a string with {variable_name}
print (f"original: {original} \nfitted: {popt}")
z_fit = func(xdata, *popt)
Z_fit = z_fit.reshape(size)
# Plot it
plt.figure(figsize=(20, 5))
plt.subplot(1, 3, 1)
plt.title("Real Function")
plt.pcolormesh(X1, X2, Z)
plt.axis(limits)
plt.colorbar()
plt.subplot(1, 3, 2)
plt.title("Function w/ Noise")
plt.pcolormesh(X1, X2, Z_noise)
plt.axis(limits)
plt.colorbar()
plt.subplot(1, 3, 3)
plt.title("Fitted Function from Noisy One")
plt.pcolormesh(X1, X2, Z_fit)
plt.axis(limits)
plt.colorbar()
plt.show()
# -
# ## Extreme cases
# +
indices = pr > np.percentile(pr, 99.999)
# Note: need pr[:] here, since indices will be 3-dimensional,
# which does not work for a netCDF variable
extreme_values = pr[:][indices]
ind_dates = np.ma.where(pr > np.percentile(pr, 99.999))[0]
extreme_dates = time[ind_dates]
plt.figure(figsize=(20, 10))
plt.plot(extreme_dates, extreme_values);
# -
# ## Change over the years
fig = plt.figure(figsize=(16, 6), dpi= 80, facecolor='w', edgecolor='k')
plt.plot(time, pr[:, 50, 3])
plt.show();
# # Save and close
dataset.close()
| examples/statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:resc]
# language: python
# name: conda-env-resc-py
# ---
# %%HTML
<style>
div.prompt {display:none}
</style>
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import numpy as np
from netCDF4 import Dataset
from mpl_toolkits.basemap import Basemap
from matplotlib import pyplot as plt
import sys, os
import PySCRIP as scrip
from PySCRIP.config import PySCRIPConfig
a = PySCRIPConfig()
# +
ncfile = Dataset("/Users/dchandan/Volumes/Scinet/reserved/cesm/cesmL2data/PlioMIP_Eoi400_v2/ocn/PlioMIP_Eoi400_v2.tseries.SSH.nc", "r")
src_data = ncfile.variables["SSH"][100,:,:]
ncfile.close()
dest_data = scrip.remap(src_data, "/Users/dchandan/Research/CESM/mapping/pyscrip_PlioMIP_Eoi400_v2_gx1_to_ll1_conserv.nc")
# -
plt.imshow(np.flipud(dest_data))
plt.colorbar()
plt.imshow(np.flipud(src_data))
plt.colorbar()
plt.show()
# +
ncfile = Dataset("/Users/dchandan/Volumes/Scinet/reserved/cesm/cesmL2data/PlioMIP_Eoi400_v2/atm/PlioMIP_Eoi400_v2.tseries.FSNS.nc", "r")
src_data = ncfile.variables["FSNS"][100,:,:]
ncfile.close()
dest_data = scrip.remap(src_data, "/Users/dchandan/Research/CESM/mapping/pyscrip_PlioMIP_Eoi400_v2_fv1_to_ll1_conserv.nc")
# -
plt.imshow(np.flipud(dest_data))
plt.colorbar()
plt.imshow(np.flipud(src_data))
plt.colorbar()
dest_data.max()
src_data.max()
| test/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## this is a modified copy of my origional that i will be exploring what the data look like if we include +/short patients and convert them to -/short. 1/7/19
# the purpose of this notebook is to take the big dataframe created in 07.01-baseline_data_merging, and make an aggregate we can use for our baseline "worst case" scenario model. This will then be fed into R, where we will use the MICE package to impute data
# changelog:
#
# * 4/16/19: added newagg
# * 4/17/19: reformatted the ordering of how code runs, and variable names. added aggregation #3.
# * 4/19/19: changed the standardizing so that log(x+1) is now applied prior to standardization. also removed ordinal variables from standardizing algorithm and concat them in later with median 0 and iqr 1 so standardize value is either 0 or 1. values from ordinal are not log transformed.
#
# 06/14/19:
# # prior to this point my pipeline:
# 1. first median standardized
# 2. aggregated
# 3. converted to 2class
# 4. train/test split
# 5. imputed
#
#
# # a big change will happen in this notebook, I will first:
# 1. convert to two class (c-/abshort &c+/ablong)
# 2. split the train and test set
# 3. median standardize
# 4. aggregate
# 5. impute
#
# i do 1-3 BEFORE I AGGREGATE and median standardize, as some information may be leaking from the train/test set as i had it previously.
# +
import pandas as pd
import matplotlib.pyplot as plt
import os
from pathlib import Path
import seaborn as sns
import numpy as np
import glob
from sklearn.externals.joblib import Memory
from sklearn.model_selection import train_test_split
memory = Memory(cachedir='/tmp', verbose=0)
<EMAIL> above any def fxn.
# %matplotlib inline
plt.style.use('ggplot')
from notebook.services.config import ConfigManager
cm = ConfigManager()
cm.update('livereveal', {
'width': 1024,
'height': 768,
'scroll': True,
})
# %load_ext autotime
# +
#patients of interest from rotation_cohort_generation
from parameters import final_pt_df_v, date, repository_path, categorical
#patients of interest from rotation_cohort_generation
final_pt_df2 = final_pt_df_v #pd.read_csv('/Users/geickelb1/Documents/GitHub/mimiciii-antibiotics-modeling/data/raw/csv/%s_final_pt_df2.csv'%(most_updated_patient_df), index_col=0)
del(final_pt_df_v)
patients= list(final_pt_df2['subject_id'].unique())
hadm_id= list(final_pt_df2['hadm_id'].unique())
icustay_id= list(final_pt_df2['icustay_id'].unique())
icustay_id= [int(x) for x in icustay_id]
# -
len(patients)
# +
def save_df(df, df_name='default', add_subfolder=False):
#uses the date and supplied df name and saves to the savepath specified above.
save_path= str(repository_path)+'/data/cleaned_merged_agg/'
if df_name == 'default':
df_name= "%s"%(df)
address=save_path+'%s/'%(folder)
if not os.path.exists(address):
print(address)
os.makedirs(address)
pd.DataFrame(df).to_csv(Path(address+'%s_%s_cleaned_merged_agg.csv' %(date, df_name)))
# -
from parameters import lower_window, upper_window, folder, date, time_col, time_var, patient_df
# +
#importing cleaned_merged big_df
allFiles = glob.glob(str(repository_path)+ '/data/cleaned_merged/%s/'%("24_hr_window") + "{}_*.csv".format(date))
df_list=[]
for element in allFiles:
df_list.append(element.split('{}_'.format(date))[1].split('_prepped.csv')[0]) #making an list of all my dataframes in order they appear in file
i=0
for name in df_list:
big_df = pd.read_csv(allFiles[i], index_col=0)
# -
#quick housekeeping addition to accomidate older generated data
if len(big_df.loc[big_df['label']=="pao2/fio2",'label'])>1:
big_df.loc[big_df['label']=="pao2/fio2",'label']="pao2fio2ratio"
big_df.head()
#overview of all variables and formats
big_df.groupby('label')['value'].describe().sort_values('unique')
# ## initial data prep
# * convert to two class
final_pt_df2['final_bin'].unique()
#convert to two class
final_pt_df2['final_bin'].unique()
two_classes=['C_neg/A_partial','C_pos/A_full','C_pos/A_partial']
two_class_icu=final_pt_df2.loc[final_pt_df2.loc[:,"final_bin"].isin(two_classes),['icustay_id','subject_id','final_bin']]
big_df=big_df.loc[big_df['icustay_id'].isin(list(two_class_icu['icustay_id'])),:].copy()
len(big_df)
big_df['icustay_id'].nunique()
# ### roundabout way of sampling train/test set so that each subject is only in either train or test:
# * To ensure a single patient did not end up with samples in both training and testing sets, individual patients (subject_id's) are kept together when performing the stratified train and test split
# label each subject_id with the max of the two classes.
two_class_pt=two_class_icu.copy()
two_class_pt['final_bin']=pd.factorize(two_class_pt['final_bin'])[0]
two_class_maxsub=two_class_pt.loc[two_class_pt.groupby('subject_id')['final_bin'].idxmax(),:]
# +
# 70/30 train/test set split with 12345=seed, splitting on max final bin of each SUBJECT_ID
train, test = train_test_split(two_class_maxsub, test_size=0.3, random_state=12345,
stratify=two_class_maxsub['final_bin'])
# generate list of each SUBJECT_ID in each split
train_subject=list(train['subject_id'])
test_subject=list(test['subject_id'])
#filtering big_df on train subjects and test subjects to get my train/test splits.
big_df_train= big_df.loc[big_df.loc[:,'subject_id'].isin(train_subject),:].copy()
big_df_test= big_df.loc[big_df.loc[:,'subject_id'].isin(test_subject),:].copy()
del big_df
# -
# ## calc median/iqr of 'healthiest patients' for standardization
# take all non-categorical variables for HEALTHY PATIENTS and calculate the median and IQR for them. then will use this to make z scores via:
# $$Z=\frac{(X-\widetilde{X}_{-/short})}{(IQR_{-/short})}$$ where $\widetilde{X}_{-/short}$ is the median value of the patients with negative SSC and short duration EAT.
#
big_df_train[big_df_train['label']=='vent_recieved']['icustay_id'].nunique()
big_df_train['icustay_id'].nunique()
from parameters import continuous, onetime
# +
def median_label_fxn(big_df):
""" grabs the median and IQR values for all continuous variables based upon the values present in
the last 24 hours of the c_neg/A_partial patients (AKA the healthiest patients at the healthiest timepoints) """
global final_pt_df2
healthy_pt=list(final_pt_df2[final_pt_df2['final_bin']=="C_neg/A_partial"]['icustay_id'])
#filter to only healthy patients filter
##splitting big_df, making a copy and restricting it to all values that will be usd in standardizing
healthy_df=big_df[big_df['label'].isin(continuous+onetime)].copy()
healthy_df['value']= pd.to_numeric(healthy_df['value']) #converting to numeric
healthy_df=healthy_df[healthy_df['icustay_id'].isin(healthy_pt)].copy() #only numerical values for cneg/ab partial pt
#finding the last 24 hours of each healthy patient.
healthy_pt_end=pd.DataFrame(healthy_df.groupby("icustay_id")['delta'].max())
healthy_pt_end["start"]=healthy_pt_end['delta']- pd.to_timedelta("1 day 00:00:00")
healthy_pt_end=healthy_pt_end.rename(columns={"delta":"end"}).reset_index()
healthy_df= pd.merge(healthy_df,healthy_pt_end, left_on="icustay_id", right_on="icustay_id", how="left" ) #now have the last 24 hours annotated for each patient as start and end.
#calculating medians and iqr for each label based on healthy patient's last 24 hours in icu
median_label=pd.DataFrame((healthy_df.groupby("label")['value'].median())).reset_index()
median_label=median_label.rename(columns={'value':"median"})
iqr_label=pd.DataFrame((healthy_df.groupby("label")['value'].quantile(0.75)-healthy_df.groupby("label")['value'].quantile(0.25))).reset_index()
iqr_label=iqr_label.rename(columns={'value':"iqr"})
median_label=pd.merge(median_label,iqr_label) #final median df
return(median_label)
def standardization_fxn(big_df):
global continuous, onetime, vaso_active, ordinal, categorical
median_label=median_label_fxn(big_df)
### dataformatting: convert all dtypes to a numeric type that pereserves nan.
#splitting categorical, ordinal and continuous
big_categorical= big_df.loc[big_df.loc[:,'label'].isin(categorical),:].copy()
#continuous and ordinal variables
big_noCat= big_df.loc[big_df.loc[:,'label'].isin(continuous),:].copy()
big_noCat['value']= big_noCat['value'].apply(pd.to_numeric, args=('coerce',)) #instead of convert to float, may preserve nan's better.
### adding a standardized value (x-median)/iqr where median is of the last 24 hours in time window for culture neg/ ab partial patients
big_noCat=pd.merge(big_noCat, median_label, how="left")
#loging values
big_noCat['median']= np.log(big_noCat['median']+1.0)
big_noCat['iqr']= np.log(big_noCat['iqr']+1.0)
big_noCat['raw_value']=big_noCat['value']
big_noCat['value']=np.log(big_noCat['value']+1.0)
big_noCat['standardize']=((big_noCat['value']-big_noCat['median'])/big_noCat['iqr']).fillna(0) #standardize is log standardized
return(big_noCat, big_categorical)
# -
noCat_df_train, cat_df_train = standardization_fxn(big_df_train)
noCat_df_test, cat_df_test = standardization_fxn(big_df_test)
# +
#####
# -
#should be same number of col as len(categorical)
cat_df_train[cat_df_train['icustay_id']==200012.0].groupby('label')['value'].value_counts()#.head(10)
noCat_df_train[noCat_df_train['label']=='lactate']
# # aggregation1:
# ### clincally guided min/max
# +
from parameters import low_value, hi_value, both_value
important_onetime=['yearsold','weight'] #pco2 and bands are now in categorical 12-12-19
# -
noCat_df_train['label'].unique()#noCat_df_train['label']=='any_vasoactives'
# ### running the min/max aggregations.
#
#
def clin_agg(big_noCat, big_cat, big_df, values="standardize"):
"""
clincally guided aggregations.
values= choose here if want to use standardization or raw values.
note: getting two minor errors, could use some cleaning up at later date.
"""
global hi_value, low_value, both_value, important_onetime
#max aggregation for selected variables
big_max= big_noCat.loc[big_noCat.loc[:,'label'].isin(hi_value),:]
table = pd.pivot_table(big_max, values=values, columns='label', index=['icustay_id'],aggfunc=max, dropna=False)
#min aggregation for selected variables
big_min= big_noCat.loc[big_noCat.loc[:,'label'].isin(low_value),:]
table2 = pd.pivot_table(big_min, values=values, columns='label', index=['icustay_id'],aggfunc=min, dropna=False)
#max&min aggregation for selected variables
big_both= big_noCat.loc[big_noCat.loc[:,'label'].isin(both_value),:]
table3 = pd.pivot_table(big_both, values=values, columns='label', index=['icustay_id'],aggfunc=[max,min], dropna=False)
#first left join all different continuous aggregations together.
worst_df=pd.merge(table.reset_index(), table2.reset_index(), how='left')
worst_df=pd.merge(worst_df, table3.reset_index(), left_on='icustay_id', right_on='icustay_id',how='left')
### formatting categorical to wide format to match the tables/worst_df
big_cat= big_cat.pivot(
index='icustay_id',
values='value',
columns='label').reset_index() #need to convert to wide format. should be one row per icustay per time.
## merging the categorical and aggregated dataframes together.
worst_df=pd.merge(worst_df, big_cat, left_on='icustay_id', right_on='icustay_id',how='left') #using max/min aggregates
#adding important one_time values to final aggregated
agg_remaining= big_df.loc[big_df.loc[:,'label'].isin(important_onetime),:]
agg_table2 = pd.pivot_table(agg_remaining, values='value', columns='label', index=['icustay_id'],aggfunc=[max], dropna=False)
agg_table2.columns = agg_table2.columns.get_level_values(1)
agg_table2=agg_table2.reset_index()
agg_table2.head()#.rename(columns={})
worst_df=pd.merge(worst_df, agg_table2, how='left')
return(worst_df)
worst_df_train=clin_agg(noCat_df_train, cat_df_train,big_df_train, values="standardize")
worst_df_test=clin_agg(noCat_df_test, cat_df_test, big_df_test, values="standardize")
worst_df_train.head()
worst_df_train.head()
worst_df_train['pao2fio2ratio'].describe()
worst_df_train['vasopressin'].value_counts()
# +
#worst_df_train['vasopressin'].value_counts()
# -
worst_df_train['bands'].value_counts()
worst_df_train['pco2'].value_counts()
save_df(worst_df_train, 'train')
save_df(worst_df_test, 'test')
del worst_df_train, worst_df_test
| notebooks/outcomes_and_alternative_calcs/06-feature_aggregation-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from dwave.system import DWaveCliqueSampler, LeapHybridSampler
from helpers import *
# # Test integer instances from Burkardt
# ## taken from https://people.math.sc.edu/Burkardt/datasets/datasets.html
# ## at first we create our models and check the problems classically
qps = createModelsFromDir('BURKARDT_DATA/INT/')
for qp in qps.values():
print(qp.model.solve())
# # Solve with DWave Leap and CliqueSampler
solvers = [createSolver(LeapHybridSampler()), createSolver(DWaveCliqueSampler(),1000)]
results = {}
for qp_name in qps.keys():
print(qp_name)
qp_name = "testprob"
results[qp_name] = [optimizer.solve(qps[qp_name]) for optimizer in solvers]
# problem ids :
# Leap db6c6b80-8882-4636-9879-a18c3b57c980
# Advantage system 0b80397c-22a2-41e1-9dce-2c6cffca3ff3
for res in results[qp_name]:
print(res)
# ## So we found the solution of LP testprob with both
# ## now test a quadratic constraint
qp_name = "three_vars_quadratic"
results[qp_name] = [optimizer.solve(qps[qp_name]) for optimizer in solvers]
# ## quadratic constraints are not supported yet
| comparison/Ocean/LinearProgramming/DWave_Qiskit_Plugin_Test/External/Integer_LP_BURKARDT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import Dependency
import pandas as pd
import numpy as np
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error, mean_absolute_error
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
import matplotlib.pyplot as plt
from matplotlib import style
from sklearn.model_selection import cross_validate
# ### Data Import
df=pd.read_csv('sps3.csv')
print(df)
df.columns
X=df.drop(['Hardness', 'FT','FTT'],axis=1).values
y=df['Hardness'].values
# ### Scaling and Splitting
scaler = StandardScaler()
scaler.fit(X)
X = scaler.transform(X)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=30)
# ## Model training
#random State=0
from sklearn.inspection import permutation_importance
from sklearn import datasets, ensemble
params = {'n_estimators': 18,
'max_depth': 4,
'min_samples_split': 5,
'learning_rate': 0.55,
'loss': 'ls'}
reg = ensemble.GradientBoostingRegressor(**params)
reg.fit(X_train, y_train)
#y_pred=reg.predict(X_test)
#random State=30
from sklearn.inspection import permutation_importance
from sklearn import datasets, ensemble
params = {'n_estimators': 100,
'max_depth': 100,
'min_samples_split': 3,
'learning_rate': 0.05,
'loss': 'ls'}
reg = ensemble.GradientBoostingRegressor(**params)
reg.fit(X_train, y_train)
# ### Accuracy, MSE & MAE
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.9, random_state=25)
y_pred=reg.predict(X_test)
accuracy=r2_score(y_test, y_pred)
mse=mean_squared_error(y_test, y_pred)
mae = mean_absolute_error(y_test, y_pred)
print(accuracy,mse,mae)
# ### Feature Importance
feature_list = list(['Temperature', 'RD', 'Grain_size'])
feature_imp = pd.Series(reg.feature_importances_, index=feature_list).sort_values(ascending=False)
print(feature_imp)
# +
feature_importance = reg.feature_importances_
sorted_idx = np.argsort(feature_importance)
pos = np.arange(sorted_idx.shape[0]) + .5
fig = plt.figure(figsize=(12, 6))
plt.subplot(1, 2, 1)
plt.barh(pos, feature_importance[sorted_idx], align='center')
plt.yticks(pos, np.array(feature_list)[sorted_idx])
plt.title('Feature Importance (MDI)')
result = permutation_importance(reg, X_test, y_test, n_repeats=10,
random_state=42, n_jobs=2)
sorted_idx = result.importances_mean.argsort()
plt.subplot(1, 2, 2)
plt.boxplot(result.importances[sorted_idx].T,
vert=False, labels=np.array(feature_list)[sorted_idx])
plt.title("Permutation Importance (test set)")
fig.tight_layout()
plt.show()
# +
scoring = "neg_mean_absolute_percentage_error"
dropped_result = cross_validate(hist_dropped, X, y, cv=3, scoring=scoring)
one_hot_result = cross_validate(hist_one_hot, X, y, cv=3, scoring=scoring)
ordinal_result = cross_validate(hist_ordinal, X, y, cv=3, scoring=scoring)
native_result = cross_validate(hist_native, X, y, cv=3, scoring=scoring)
def plot_results(figure_title):
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 8))
plot_info = [('fit_time', 'Fit times (s)', ax1, None),
('test_score', 'Mean Absolute Percentage Error', ax2,
(0, 0.20))]
x, width = np.arange(4), 0.9
for key, title, ax, y_limit in plot_info:
items = [dropped_result[key], one_hot_result[key], ordinal_result[key],
native_result[key]]
ax.bar(x, [np.mean(np.abs(item)) for item in items],
width, yerr=[np.std(item) for item in items],
color=['C0', 'C1', 'C2', 'C3'])
ax.set(xlabel='Model', title=title, xticks=x,
xticklabels=["Dropped", "One Hot", "Ordinal", "Native"],
ylim=y_limit)
fig.suptitle(figure_title)
plot_results("Gradient Boosting on Adult Census")
# -
# ### Plots
plt.figure(figsize=(10,6))
plt.style.use('bmh')
plt.scatter( y_test, y_pred)
plt.xlabel('Practical')
plt.ylabel('Predction')
plt.title('Hardness Practical Vs Predction')
# +
test_score = np.zeros((params['n_estimators'],), dtype=np.float64)
for i, y_pred in enumerate(reg.staged_predict(X_test)):
test_score[i] = reg.loss_(y_test, y_pred)
fig = plt.figure(figsize=(12, 6))
plt.subplot(1, 1, 1)
plt.title('Deviance')
plt.plot(np.arange(params['n_estimators']) + 1, reg.train_score_, 'b-',
label='Training Set Deviance')
plt.plot(np.arange(params['n_estimators']) + 1, test_score, 'r-',
label='Test Set Deviance')
plt.legend(loc='upper right')
plt.xlabel('Boosting Iterations')
plt.ylabel('Deviance')
fig.tight_layout()
plt.show()
# -
print(plt.style.available)
# +
#Temperature Vs Hardness
temp=[row[0] for row in X_test]
dp={}
dt={}
for i in range(len(temp)):
dp[temp[i]]=y_pred[i]
dt[temp[i]]=y_test[i]
otemp=sorted(temp)
oy_pred=[]
oy_test=[]
for k in otemp:
oy_pred.append(dp[k])
oy_test.append(dt[k])
#Plot
plt.figure(figsize=(12,6))
plt.style.use('tableau-colorblind10')
plt.plot(otemp,oy_pred, label='Prediction')
plt.scatter(otemp,oy_pred)
plt.plot(otemp,oy_test, label='Test')
plt.scatter(otemp,oy_test)
plt.legend(loc='upper right')
plt.xlabel('Temperature')
plt.ylabel('Hardness')
plt.title('Temperature Vs Hardness')
plt.show()
# +
#Relative Density Vs Hardness
temp=[row[1] for row in X_test]
dp={}
dt={}
for i in range(len(temp)):
dp[temp[i]]=y_pred[i]
dt[temp[i]]=y_test[i]
otemp=sorted(temp)
oy_pred=[]
oy_test=[]
for k in otemp:
oy_pred.append(dp[k])
oy_test.append(dt[k])
#Plot
plt.figure(figsize=(12,6))
plt.style.use('seaborn')
plt.plot(otemp,oy_pred, label='Prediction')
plt.scatter(otemp,oy_pred)
plt.plot(otemp,oy_test, label='Test')
plt.scatter(otemp,oy_test)
plt.legend(loc='upper left')
plt.xlabel('Relative Density')
plt.ylabel('Hardness')
plt.title('Relative Density Vs Hardness')
plt.show()
# +
#Grain Size Vs Hardness
temp=[row[2] for row in X_test]
dp={}
dt={}
for i in range(len(temp)):
dp[temp[i]]=y_pred[i]
dt[temp[i]]=y_test[i]
otemp=sorted(temp)
oy_pred=[]
oy_test=[]
for k in otemp:
oy_pred.append(dp[k])
oy_test.append(dt[k])
#Plot
plt.figure(figsize=(12,6))
plt.style.use('ggplot')
plt.plot(otemp,oy_pred, label='Prediction')
plt.scatter(otemp,oy_pred)
plt.plot(otemp,oy_test, label='Test')
plt.scatter(otemp,oy_test)
plt.legend(loc='upper right')
plt.xlabel('Grain Size')
plt.ylabel('Hardness')
plt.title('Grain Size Vs Hardness')
plt.show()
| sps_GBR_Hardness.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# ##### Authors:
# - <NAME>
# - <NAME>
#
# ##### Date: 2/6/2019
#
# ##### Version: 3.0
#
# ##### Environment: Python 3.6.1 and Jupyter notebook
# # Table of contents
# ### 1. [Importing libraries](#library)
# ### 2. [Initialization](#initialisation)
# ### 3. [Read training and label](#read_train)
# ### 4. [Data pre-processing](#preprocess)
# ### 5. [Feature generation](#feature)
# - #### 5.1 [Dimention reduction technique(Chi-squared)](#dimension)
# - #### 5-2 [Multinomial logistic regression](#model)
# - #### 5-3 [Cross-validation](#cv)
#
# ### 6. [Predict on test data](#test)
# ## 1. Importing libraries <a name="library"></a>
import pandas as pd
import numpy as np
from tqdm import tqdm
from pattern.en import parse
from nltk.corpus import stopwords
import string
import re
import nltk
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score
from sklearn import svm
import swifter
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.svm import LinearSVC
from sklearn import metrics
import seaborn as sns
from wordcloud import WordCloud
import matplotlib.pyplot as plt
from sklearn.feature_selection import SelectKBest, chi2
import warnings
warnings.filterwarnings("ignore")
# ## 2. Initialization<a name="initialisation"></a>
#
# ### Creating a custom dictionary to expand all the decontract words
# +
#initialising the lemmatizer.
wn = nltk.WordNetLemmatizer()
# Creating a custom dictionary to expand all the decontract words
appos = {
"aren't" : "are not", "can't" : "cannot", "couldn't" : "could not", "didn't" : "did not", "doesn't" : "does not",
"don't" : "do not", "hadn't" : "had not", "hasn't" : "has not", "haven't" : "have not",
"he'd" : "he would", "he'll" : "he will", "he's" : "he is", "i'd" : "I would",
"i'd" : "I had", "i'll" : "I will", "i'm" : "I am", "isn't" : "is not",
"it's" : "it is", "it'll":"it will", "i've" : "I have", "let's" : "let us",
"mightn't" : "might not", "mustn't" : "must not", "shan't" : "shall not", "she'd" : "she would",
"she'll" : "she will", "she's" : "she is", "shouldn't" : "should not", "that's" : "that is",
"there's" : "there is", "they'd" : "they would", "they'll" : "they will", "they're" : "they are",
"they've" : "they have", "we'd" : "we would", "we're" : "we are", "weren't" : "were not",
"we've" : "we have", "what'll" : "what will", "what're" : "what are", "what's" : "what is",
"what've" : "what have", "where's" : "where is", "who'd" : "who would", "who'll" : "who will",
"who're" : "who are", "who's" : "who is", "who've" : "who have", "won't" : "will not",
"wouldn't" : "would not", "you'd" : "you would", "you'll" : "you will","you're" : "you are",
"you've" : "you have", "'re": " are", "wasn't": "was not", "we'll":" will","didn't": "did not"
}
#reference[1]
# -
# ## 3. Reading the training data and labels <a name="read_train"></a>
#
# ### merging both of them
data = pd.read_csv("train_data.csv", sep=',') # read training data
data_labels = pd.read_csv("train_label.csv", sep=',') # read training labels
df=pd.merge(data,data_labels,on='trn_id',how='left') # merging both of them
# ## 4. Data pre-processing <a name="preprocess"></a>
# +
#--------------------------
# Data pre-processing step
#--------------------------
def pre_process(text):
"""
Takes in a string of text, then performs the following:
1. converts to lower
2. Splits the sentence into tokens
3. Decontract the words. For example: "won't" --> "will not"
4. Lemmatization, reduces words to their base word
5. Returns the sentence of the cleaned text
"""
text = "".join([word.lower() for word in text])
tokens = text.split(" ")
tokens = [appos[word] if word in appos else word for word in tokens]
text = " ".join([wn.lemmatize(word) for word in tokens])
return text
#--------------------------
# execute pre-processing
#--------------------------
df['text']=df.swifter.apply(lambda x:pre_process(x['text']),axis=1)
# -
# ## 5. Feature generation <a name="feature"></a>
# ### 5.1- Dimension reduction technique (Chi-square)<a name="dimension"></a>
# +
#--------------------------------------
#dimension reduction using chi-square
#--------------------------------------
x_train, x_validation, y_train, y_validation = train_test_split(df['text'], df['label'], test_size=.02)
tvec = TfidfVectorizer(max_features=100000,ngram_range=(1, 3))
x_train_tfidf = tvec.fit_transform(x_train)
x_validation_tfidf = tvec.transform(x_validation)
#reference[2]
# -
# ### 5-2 Multinomial logistic regression<a name="model"></a>
# +
ch = SelectKBest(chi2, k=40000)
x_train_feature_selected=ch.fit_transform(x_train_tfidf, y_train)
x_test_chi_selected = ch.transform(x_validation_tfidf)
from sklearn import linear_model
clf = linear_model.LogisticRegression(multi_class='multinomial',solver = 'newton-cg')
clf.fit(x_train_feature_selected, y_train)
score = clf.score(x_test_chi_selected, y_validation)
score
# -
# ### 5-3 Cross-validation <a name="cv"></a>
# +
from sklearn.model_selection import KFold, cross_val_score
#rf = RandomForestClassifier(n_jobs=-1)
k_fold = KFold(n_splits=3)
cross_val_score(clf, x_train_chi2_selected, y_train, cv=k_fold, scoring='accuracy', n_jobs=-1)
# -
# --------------------------------
# # 6.Prediction on test data<a name="test"></a>
# +
#--------------------------------------
## Reading the test file into dataframe
#--------------------------------------
test=pd.read_csv("test_data.csv", sep=',')
# +
#--------------------------------------------------------------------
## Cleaning the test data as per the cleaning technique of train data
#--------------------------------------------------------------------
test['text']=test.swifter.apply(lambda x:pre_process(x['text']),axis=1)
# +
#--------------------------------------------------------------------
## Transforming the text into vector tfidf vectorizer with chi-sqaure
#--------------------------------------------------------------------
test_matrix= tvec.transform(test['text'])
test_matrix = ch.transform(test_matrix)
# +
#---------------------------------------------------------------------
## predicting the labels, storing it as label column in test dataframe
#---------------------------------------------------------------------
test['label'] = pd.DataFrame(clf.predict(test_matrix))
# +
#-----------------------------------------------------------
## dropping all other columns keeping only test_id and label
#-----------------------------------------------------------
test=test[['test_id','label']]
############################################################
#--------------------------------
#Converting the dataframe to csv
#--------------------------------
test.to_csv('predict_label.csv',index=False)
# -
# # References
# .[1] https://drive.google.com/file/d/0B1yuv8YaUVlZZ1RzMFJmc1ZsQmM/view
# [2] https://github.com/tthustla/twitter_sentiment_analysis_part8/blob/master/Capstone_part4-Copy6.ipynb
| NLP_python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computational Structural Design II <br/> Assignment II: Convex Polygon Blocks
# In this assignment, we will generate polygon blocks for the form-found barrel vault.
#
# <table>
# <tr>
# <td style='text-align:center;'>
# <img src="https://github.com/BlockResearchGroup/CSD2_2022/blob/main/3_Materialization/Tutorial7/img/brick3.png?raw=true" width="570" style="margin-left:auto; margin-right:auto"/><b> Barrel Vault Quad Mesh </b><img>
# </td>
# <td>
# <img src="https://github.com/BlockResearchGroup/CSD2_2022/blob/main/3_Materialization/Tutorial7/img/hex0.png?raw=true" width="500" style="margin-left:auto; margin-right:auto"/><b> Tessellation Pattern </b><img>
# </td>
# </tr>
# </table>
# <br>
#
# ### Steps:
# - **A. Load Mesh**
# - **B. Compute Tessellation Pattern** </br>
# The input mesh is a quad mesh, A hexagonal polygon can be generated with the vertices around two adjacent quad faces. You can modify the vertex coordinates in the quadmesh. Serialize the modified quad mesh. </br>
# Secondly, find the correct vertices in each block. Create a list,`block_faces`, and save the vertices on each block as a list in `block_faces`. Serialize the `block_faces`. </br>
# Visualize the blocks as `Polygon` in the viewer. </br>
#
# - **C. Generate Blocks** </br>
# Create a function `generate_block`. The input parameter is **the modified quad mesh**, **vertices on one block**, and **thickness of the block**. The function should return a 3D block, which has a planar top surface. </br>
# Call the function to generate all the blocks for the barrel vault. Serialize the blocks and visualize them in the viewer.
#
#
# <br>
#
# ---
# <br><br>
# +
# % pip install compas_notebook
# -
# ## A. Load Mesh
# +
from compas.datastructures import Mesh
from compas_notebook.app import App
mesh= Mesh.from_obj("https://raw.githubusercontent.com/BlockResearchGroup/CSD2_2022/main/3_Materialization/Tutorial6/data/barrel_vault.obj")
viewer = App()
viewer.add(mesh)
viewer.show()
# + [markdown] tags=[]
# # B. Compute Tessellation Pattern
#
# ## Hint:
# Here is one solution to compute the tessellation pattern. You can also use your own way to achieve the same result.
# <table>
# <tr>
# <td style='text-align:center;'>
# <img src="https://github.com/BlockResearchGroup/CSD2_2022/blob/main/3_Materialization/Tutorial7/img/hint1.png?raw=true" width="500" style="margin-left:auto; margin-right:auto"/><img>
# </td>
# <td>
# <img src="https://github.com/BlockResearchGroup/CSD2_2022/blob/main/3_Materialization/Tutorial7/img/hint2.png?raw=true" width="500" style="margin-left:auto; margin-right:auto"/><img>
# </td>
# <td style='text-align:center;'>
# <img src="https://github.com/BlockResearchGroup/CSD2_2022/blob/main/3_Materialization/Tutorial7/img/hint3.png?raw=true" width="500" style="margin-left:auto; margin-right:auto"/><img>
# </td>
# </tr>
# <tr>
# <td style='text-align:center;'>
# <b> 1. Find the staggered strip </b>
# </td>
# <td style='text-align:center;'>
# <b> 2. Modify the vertes coordinates of the strips </b>
# </td>
# <td style='text-align:center;'>
# <b> 3. Visualize the 2D blocks </b>
# </td>
# </tr>
# </table>
# <br>
#
# -
# ## B1. Modify the Quad Mesh
# Keep the topology of the mesh and modify the coordinates of the vertices in the mesh.
# +
import os
import compas
from compas.datastructures import Mesh
from compas_notebook.app import App
mesh= Mesh.from_obj("https://raw.githubusercontent.com/BlockResearchGroup/CSD2_2022/main/3_Materialization/Tutorial6/data/barrel_vault.obj")
# modify the quad mesh...
# export modified mesh data to a new file
dirname = '/content/drive/My Drive/Colab Notebooks'
file_out_name = '01_modified_barrel_vault.json'
file_out_path = os.path.join(dirname, file_out_name)
compas.json_dump(mesh, file_out_path, pretty=True)
# visualization
viewer = App()
viewer.add(mesh)
viewer.show()
# -
# ## B2. 2D Block
# +
import os
import compas
from compas.datastructures import Mesh
from compas_notebook.app import App
from random import random
from compas.utilities import i_to_rgb
# folder location
dirname = '/content/drive/My Drive/Colab Notebooks'
# load modified mesh from step B1
file_in_name = '01_modified_barrel_vault.json'
file_in_path = os.path.join(dirname, file_in_name)
mesh: Mesh = compas.json_load(file_in_path)
# vertices on the block
block_faces = []
# your code here...
# export block_faces
compas.json_dump(block_faces, os.path.join(dirname, "02_block_faces.json"))
# visualization
viewer = App()
# visualize the blocks
for block_face in block_faces:
v_xyz = [mesh.vertex_coordinates(vkey) for vkey in block_face]
viewer.add(Polygon(v_xyz), facecolor = i_to_rgb(random(), normalize=True))
viewer.show()
# -
# # C. Generate Blocks
def generate_block(mesh, block_face, thickness):
# your code here...
pass
# +
import os
import compas
from compas.datastructures import Mesh
from compas_notebook.app import App
# folder location
dirname = '/content/drive/My Drive/Colab Notebooks'
# load modified mesh from step B1
file_in_name = '01_modified_barrel_vault.json'
file_in_path = os.path.join(dirname, file_in_name)
mesh: Mesh = compas.json_load(file_in_path)
# load block faces
block_faces = compas.json_load(os.path.join(here, "02_block_faces.json"))
# generate blocks...
blocks = []
# export blocks to a new file...
# visualization
viewer = App()
for block in blocks:
viewer.add(block)
viewer.show()
| 3_Materialization/Tutorial7/assignment2.ipynb |
# +
import numpy as np
import matplotlib.pyplot as plt
try:
import probml_utils as pml
except ModuleNotFoundError:
# %pip install git+https://github.com/probml/probml-utils.git
import probml_utils as pml
x = np.linspace(-1, 1, 100)
y = np.power(x, 2)
plt.figure()
plt.plot(x, y, "-", lw=3)
plt.title("Smooth function")
pml.savefig("smooth-fn.pdf")
y = np.abs(x)
plt.figure()
plt.plot(x, y, "-", lw=3)
plt.title("Non-smooth function")
pml.savefig("nonsmooth-fn.pdf")
plt.show()
| notebooks/book1/08/smooth-vs-nonsmooth-1d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={}
# # This is a notebook for PMT gain estimation from calibration events
# - Calculate pedestal RMS
# - Load pedestal data and perform R0 correction
# - Check pedestal distribution and get RMS
# - Analyze flat-fielding events
# - Choose data to be used
# - Briefly look at an event.
# - Check waveforms after R0 correction
# - Calibration from R1 to DL1
# - Check waveform and integration window
# - Look at a camera image of the event
# - Estimate the number of p.e. from flat-fielding events
# - Extract charge from events and store them in an array
# - Check charge distribution in a pixel
# - Estimate the number of p.e. using excess noise factor method
# - See consistency between the number of p.e. estimated from high and low gain
# - Look at camera display of the estimated number of p.e.
# + [markdown] pycharm={}
# ### Data location
# You can download the data used in this notebook from http://www.icrr.u-tokyo.ac.jp/~yukihok/LSTdata/<p>
# You need user name and password to access the web page.<p>
# They are completely same as those needed for LST1 wiki http://www.lst1.iac.es/wiki/index.php/Main_Page/
# + [markdown] pycharm={}
# ### Import modules
# + pycharm={}
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from pylab import rcParams
from ctapipe.calib import CameraCalibrator
from ctapipe.instrument import CameraGeometry
from ctapipe.visualization import CameraDisplay
from ctapipe.io import EventSource
from traitlets.config.loader import Config
import sys
sys.path.append('../../ctapipe_io_lst') #add path to ctapipe_io_lst
from lstchain.calib.camera.r0 import LSTR0Corrections
# + [markdown] pycharm={}
# ### Prepare the class for R0 correction
# + pycharm={}
config = Config({
"LSTR0Corrections": {
"pedestal_path": "/home/yukihok/Work/CTA/Analysis/calibration/pedestal_run97_new.fits",
"offset": 300,
"tel_id": 0, # for EVB3 will be 1
"r1_sample_start": None,
"r1_sample_end": None
}
})
lst_r0 = LSTR0Corrections(config=config)
# + [markdown] pycharm={}
# ## Calculate pedestal RMS
# ### Load pedestal data and perform R0 correction
# + pycharm={}
ped_input_url = '/disk/gamma/cta/work/yukiho/RealData/LST-1.1.Run00158.0000.fits.fz'
pedestal_source = EventSource(input_url=ped_input_url, max_events = 1000)
# + pycharm={}
tel_id = 0
pedestal = []
window_start = 12 # define window position for pedestal calculation
window_width = 7
for event in pedestal_source:
if (event.count + 1) % 1000 == 0:
print('{} events have been read.'.format(event.count + 1))
#lst_r0.calibrate(event)
lst_r0.subtract_pedestal(event)
#lst_r0.time_lapse_corr(event)
#lst_r0.interpolate_spikes(event)
event.r1.tel[tel_id].waveform = event.r1.tel[tel_id].waveform - lst_r0.offset
pedestal.append(np.sum(event.r1.tel[tel_id].waveform[:, :, window_start:window_start + window_width], axis=2))
# + [markdown] pycharm={}
# ### Check pedestal distribution and get RMS
# + pycharm={}
pedestal = np.array(pedestal)
pedestal_var = np.var(pedestal, axis=0) # used for gain estimation later
print('the number of used pedestal events:', pedestal.shape[0])
pix = 0
plt.subplots_adjust(wspace=40, hspace=0.6)
rcParams['figure.figsize'] = 10, 6
rcParams['font.size'] = 14
fig, ax = plt.subplots(1, 2)
ax[0].hist(pedestal[:, 0, pix], bins=np.arange(-600, 1000, 50))
ax[1].hist(pedestal[:, 1, pix], bins=np.arange(-150, 200, 10))
ax[0].set_title('integrated pedestal (HG)')
ax[1].set_title('integrated pedestal (LG)')
ax[0].set_xlabel('ADC count')
ax[1].set_xlabel('ADC count')
plt.show()
print('high gain: average {}, std {} '.format(np.mean(pedestal[:, 0, pix]), np.std(pedestal[:, 0, pix])))
print('low gain : average {}, std {} '.format(np.mean(pedestal[:, 1, pix]), np.std(pedestal[:, 1, pix])))
# + [markdown] pycharm={}
# # Analyze flat-fielding events
# ## The flat-fielding data to be used
# + pycharm={}
ff_input_url = '/disk/gamma/cta/store/ZFITS2/CalibPulse/LST-1.1.Run00170.0000.fits.fz' # path to the flat-fielding data
# + [markdown] pycharm={}
# ## First, let's look at an event briefly
# + pycharm={}
ff_source = EventSource(input_url=ff_input_url, max_events=2) # The first event should be excluded from the analysis.
for event in ff_source:
print('event id: {}'.format(event.count))
# + [markdown] pycharm={}
# ### Check waveforms after R0 correction
# + pycharm={}
t = np.arange(0, 40, 1)
pix = 1
fig, ax = plt.subplots(1, 2)
ax[0].step(t, event.r0.tel[tel_id].waveform[0, pix], color="blue", label="R0")
ax[1].step(t, event.r0.tel[tel_id].waveform[1, pix], color="blue", label="R0")
#lst_r0.calibrate(event)
lst_r0.subtract_pedestal(event)
event.r1.tel[tel_id].waveform = event.r1.tel[tel_id].waveform - lst_r0.offset
#lst_r0.time_lapse_corr(event)
#lst_r0.interpolate_spikes(event)
ax[0].step(t, event.r1.tel[tel_id].waveform[0, pix], color="red", label="R1")
ax[1].step(t, event.r1.tel[tel_id].waveform[1, pix], color="red", label="R1")
axs = plt.gcf().get_axes()
for ax in axs:
plt.sca(ax)
plt.xlabel("time sample [ns]")
plt.ylabel("count [ADC]")
plt.legend(loc='upper left')
plt.grid()
plt.show()
# + [markdown] pycharm={}
# ### Calibration from R1 to DL1
# + pycharm={}
calibrator = CameraCalibrator(extractor_product = 'LocalPeakIntegrator', eventsource=ff_source)
event.r1.tel[tel_id].waveform[:, :, :2] = 0 # to avoid the edge of ROI being chosen as a peak.
event.r1.tel[tel_id].waveform[:, :, 38:] = 0
#calibrator.r1.calibrate(event)
calibrator.dl0.reduce(event)
calibrator.dl1.calibrate(event)
# + [markdown] pycharm={}
# ### Check waveform and integration window
# + pycharm={}
pix = 0
t = np.arange(2, 38, 1)
fig, ax = plt.subplots(1, 2)
ax[0].step(t, event.dl0.tel[tel_id].waveform[0, pix, 2:38], color="blue", label="waveform")
ax[1].step(t, event.dl0.tel[tel_id].waveform[1, pix, 2:38], color="blue", label="waveform")
ax[0].step(
t,
event.dl0.tel[tel_id].waveform[0, pix, 2:38]*event.dl1.tel[tel_id].extracted_samples[0, pix, 2:38],
color="red",
label="extracted"
)
ax[1].step(
t,
event.dl0.tel[tel_id].waveform[1, pix, 2:38]*event.dl1.tel[tel_id].extracted_samples[1, pix, 2:38],
color="red",
label="extracted"
)
axs = plt.gcf().get_axes()
for ax in axs:
plt.sca(ax)
plt.xlabel("time sample [ns]")
plt.ylabel("count [ADC]")
plt.legend(loc='upper left')
plt.grid()
plt.show()
# + [markdown] pycharm={}
# ### Look at a camera image of the event
# + pycharm={}
plt.rcParams['figure.figsize'] = (18, 6)
camera = CameraGeometry.from_name("LSTCam", 2)
plt.subplot(1,2,1)
display = CameraDisplay(camera)
display.image = event.dl1.tel[tel_id].image[0]
display.add_colorbar()
display.colorbar.set_label('ADC [count]')
#display.set_limits_minmax(0, 100)
plt.title('high gain')
plt.subplot(1,2,2)
display = CameraDisplay(camera)
display.image = event.dl1.tel[tel_id].image[1]
display.add_colorbar()
display.colorbar.set_label('ADC [count]')
plt.title('low gain')
plt.show()
# + [markdown] pycharm={}
# ## Estimate the number of p.e. from flat-fielding events
# ### Extract charge from events and store them in an array
# + pycharm={}
max_events = 4000
threshold = 1000
ff_source = EventSource(input_url = ff_input_url, max_events = max_events)
calibrator = CameraCalibrator(extractor_product = 'LocalPeakIntegrator', eventsource=ff_source)
hg_charge = []
lg_charge = []
num_events = 0
print ('analyzing', ff_input_url)
for event in ff_source:
if event.count == 0: # We don't use the first event.
continue;
if (event.count + 1) % 1000 == 0:
print('{} events have been read.'.format(event.count + 1))
#lst_r0.calibrate(event)
lst_r0.subtract_pedestal(event)
#lst_r0.time_lapse_corr(event)
#lst_r0.interpolate_spikes(event)
event.r1.tel[tel_id].waveform = event.r1.tel[tel_id].waveform - lst_r0.offset
event.r1.tel[tel_id].waveform[:, :, :2] = 0 # to avoid the edge of ROI being chosen as a peak.
event.r1.tel[tel_id].waveform[:, :, 38:] = 0
calibrator.dl0.reduce(event)
calibrator.dl1.calibrate(event)
if np.mean(event.dl1.tel[tel_id].image[0, :]) > threshold: # to exclude events triggered by NSB/muon/shower.
hg_charge.append(event.dl1.tel[tel_id].image[0, :])
lg_charge.append(event.dl1.tel[tel_id].image[1, :])
num_events = num_events + 1
num_gains = 2
num_pixels = 1855
charge = np.zeros((num_gains, num_events, num_pixels)) # This array contains all the charge information.
charge[0] = np.array(hg_charge)
charge[1] = np.array(lg_charge)
print('')
print(num_events, 'events are used for gain estimation.')
# + [markdown] pycharm={}
# ### Check charge distribution in a pixel
# + pycharm={}
pix = 0
rcParams['figure.figsize'] = 10, 6
fig, ax = plt.subplots(1, 2)
ax[0].hist(charge[0, :, pix], bins=40)
#ax[0].set_yscale('log')
ax[0].set_title('charge distribution (high gain)')
ax[0].set_xlabel('ADC count')
ax[1].hist(charge[1, :, pix], bins=40)
#ax[1].set_yscale('log')
ax[1].set_title('charge distribution (low gain)')
ax[1].set_xlabel('ADC count')
plt.show()
print('HG:', 'average', np.mean(charge[0, :, pix]), ' std', np.std(charge[0, :, pix]))
print('LG:', 'average', np.mean(charge[1, :, pix]),' std', np.std(charge[1, :, pix]))
# + [markdown] pycharm={}
# ### Estimate the number of p.e. using excess noise factor method
# + pycharm={}
mean_charge = np.mean(charge, axis=1)
var_charge = np.var(charge, axis=1)
F_square = 1.2 # square of ENF based on measurement
phe = mean_charge*mean_charge/(var_charge-pedestal_var)*F_square
# + pycharm={}
fig, ax = plt.subplots(1, 2)
ax[0].hist(phe[0], bins=np.arange(0, 120, 5))
ax[0].set_title('# of estimated p.e. (high gain)')
ax[0].set_xlabel('p.e.')
ax[1].hist(phe[1], bins=np.arange(0, 120, 5))
ax[1].set_title('# of estimated p.e. (low gain)')
ax[1].set_xlabel('p.e.')
plt.show()
print('HG:', 'average', np.mean(phe[0]), ' std', np.std(phe[0]))
print('LG:', 'average', np.mean(phe[1]), ' std', np.std(phe[1]))
# + [markdown] pycharm={}
# ### See consistency between the number of p.e. estimated from high and low gain
# + pycharm={}
t = np.linspace(0, 200, 200)
phe_range = [60, 100]
tick = phe_range[1] - phe_range[0]
plt.rcParams['figure.figsize'] = (8, 6)
plt.figure()
plt.hist2d(phe[0], phe[1], range=[phe_range, phe_range], bins = [tick, tick])
plt.plot(t, t, color='r')
plt.xlabel('# of estimated p.e. (HG)')
plt.ylabel('# of estimated p.e. (LG)')
plt.title('# of estimated p.e. in each pixel')
cb = plt.colorbar()
cb.set_label('pixels')
plt.show()
# + [markdown] pycharm={}
# ### Display the estimated number of p.e. on the camera
# + pycharm={}
camera = CameraGeometry.from_name("LSTCam", 2)
plt.rcParams['figure.figsize'] = (14, 6)
plt.subplot(1,2,1)
display = CameraDisplay(camera)
display.image = phe[0]
display.add_colorbar()
display.colorbar.set_label('p.e.')
#display.set_limits_minmax(0, 100)
plt.title('high gain')
plt.subplot(1,2,2)
display = CameraDisplay(camera)
display.image = phe[1]
display.add_colorbar()
display.colorbar.set_label('p.e.')
#display.set_limits_minmax(0, 100)
plt.title('low gain')
plt.show()
# + pycharm={}
| notebooks/gain_estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # 3D wireframe plots in one direction
#
#
# Demonstrates that setting rstride or cstride to 0 causes wires to not be
# generated in the corresponding direction.
#
# +
from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
fig, [ax1, ax2] = plt.subplots(2, 1, figsize=(8, 12), subplot_kw={'projection': '3d'})
# Get the test data
X, Y, Z = axes3d.get_test_data(0.05)
# Give the first plot only wireframes of the type y = c
ax1.plot_wireframe(X, Y, Z, rstride=10, cstride=0)
ax1.set_title("Column (x) stride set to 0")
# Give the second plot only wireframes of the type x = c
ax2.plot_wireframe(X, Y, Z, rstride=0, cstride=10)
ax2.set_title("Row (y) stride set to 0")
plt.tight_layout()
plt.show()
| matplotlib/gallery_jupyter/mplot3d/wire3d_zero_stride.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python implementations
# %matplotlib notebook
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split, KFold
from sklearn import linear_model
from sklearn.metrics import mean_squared_error
import sys
sys.path.insert(0, '../src/')
import turbo
from pathlib import Path
# +
fig_width_pt = 233.1 # Get this from LaTeX using \showthe\columnwidth
inches_per_pt = 1.0/72.27 # Convert pt to inches
golden_mean = (np.sqrt(5)-1.0)/2.0 # Aesthetic ratio
fig_width = fig_width_pt*inches_per_pt # width in inches
fig_height =fig_width*golden_mean # height in inches
fig_size = [fig_width,fig_height]
fig_path = Path("../latex/figures")
plt.rc('text', usetex=True)
fig_opts = {'font.size' : 10,
#'font.family': 'monospace',
'axes.labelsize' : 10,
'font.size' : 10,
'text.fontsize' : 10,
'legend.fontsize': 10,
'xtick.labelsize' : 8,
'ytick.labelsize' : 8,
}
plt.rc('font', family='serif')
plt.rc(fig_opts)
# -
# ### Construct the dataset
# Code is copied from Mehta et al.
# +
import numpy as np
import scipy.sparse as sp
np.random.seed(12)
import warnings
# Comment this to turn on warnings
warnings.filterwarnings('ignore')
# system size
L=40
# create 10000 random Ising states
states_raw=np.random.choice([-1, 1], size=(10000,L))
def ising_energies(states):
"""
This function calculates the energies of the states in the nn Ising Hamiltonian
"""
L = states.shape[1]
J = np.zeros((L, L),)
for i in range(L):
J[i,(i+1)%L]=-1.0 # interaction between nearest-neighbors
# compute energies
E = np.einsum('...i,ij,...j->...',states,J,states)
return E
# calculate Ising energies
energies=ising_energies(states_raw)
# reshape Ising states into RL samples: S_iS_j --> X_p
states=np.einsum('...i,...j->...ij', states_raw, states_raw)
shape=states.shape
states=states.reshape((shape[0],shape[1]*shape[2]))
# build final data set
Data=[states,energies]
# -
Xtrain, Xtest, ytrain, ytest = train_test_split(states, energies, test_size=0.3, random_state=42)
np.save("1D_xtrain.npy", Xtrain)
np.save("1D_xtest.npy", Xtest)
np.save("1D_ytrain.npy", ytrain)
np.save("1D_ytest.npy", ytest)
# +
from dataclasses import dataclass, field
@dataclass
class CVRun:
coeffs: list = field(default_factory=list)
r2_train: list = field(default_factory=list)
r2_test: list = field(default_factory=list)
mse_train: list = field(default_factory=list)
mse_test: list = field(default_factory=list)
parameter: float = 0.0
def store(self, model, X_train, y_train, X_test, y_test):
self.coeffs.append(np.reshape(model.coef_, (L, L)))
self.r2_train.append(model.score(X_train, y_train))
self.r2_test.append(model.score(X_test, y_test))
self.mse_train.append(mean_squared_error(y_train, model.predict(X_train)))
self.mse_test.append(mean_squared_error(y_test, model.predict(X_test)))
def average(self):
print(len(self.coeffs))
mean = np.mean(self.coeffs, axis=0)
print("--", mean.shape)
std = np.std(self.coeffs, axis=0)
return mean, std
class RegressionRun:
def __init__(self, Xtrain, ytrain, Xtest, ytest):
self.coefficients = []
self.parameters = []
self.test_mse = []
self.train_mse = []
self.test_se_mse = []
self.train_se_mse = []
self.test_r2 = []
self.train_r2 = []
self.test_se_r2 = []
self.train_se_r2 = []
self.Xtrain, self.ytrain = Xtrain, ytrain
self.Xtest, self.ytest = Xtest, ytest
def update(self, cv):
self.coefficients.append(cv.coeffs[0])
self.parameters.append(cv.parameter)
self.train_mse.append(np.mean(cv.mse_train))
self.train_se_mse.append(np.std(cv.r2_train))
self.train_r2.append(np.mean(cv.r2_train))
self.train_se_r2.append(np.std(cv.r2_train))
self.test_se_mse.append(np.std(cv.mse_test))
self.test_se_r2.append(np.std(cv.r2_test))
self.test_mse.append(np.mean(cv.mse_test))
self.test_r2.append(np.mean(cv.r2_test))
# -
kf = KFold(n_splits=5)
cv_results = CVRun()
lmmodel = linear_model.LinearRegression()
for train, test in kf.split(Xtrain):
X_train, y_train = Xtrain[train], ytrain[train]
X_test, y_test = Xtrain[test], ytrain[test]
lmmodel.fit(X_train, y_train)
cv_results.store(lmmodel, X_train, y_train, X_test, y_test)
# +
def grid_search(regressor, parameter_space):
#results = []
result = RegressionRun(Xtrain, ytrain, Xtest, ytest)
for α in parameter_space:
print(f"Working on α = {α}")
regressor.set_params(alpha=α)
cv_results = CVRun(parameter=α)
for train, test in kf.split(Xtrain):
X_train, y_train = Xtrain[train], ytrain[train]
X_test, y_test = Xtrain[test], ytrain[test]
regressor.fit(X_train, y_train)
cv_results.store(regressor, X_train, y_train, X_test, y_test)
#results.append(cv_results)
result.update(cv_results)
return result
alphas = np.logspace(-5, 5, 40)
#alphas = [10**-5, 10**-2, 1]
ridge = linear_model.Ridge()
ridge = grid_search(ridge, alphas)
lambdas = np.logspace(-5, 1, 20)
#lambdas = alphas
lasso = linear_model.Lasso()
lasso = grid_search(lasso, lambdas)
# -
def coeff_plot(regruns):
fig, axes = plt.subplots(nrows=3, figsize=(fig_width, 3*fig_height))
cmap_args = {'vmin': -1, 'vmax':1, 'cmap': "RdBu_r"}
for i in range(3):
im = axes[i].matshow(regruns.coefficients[i], **cmap_args)
axes[i].axis('off')
axes[i].set_title(r"$\lambda = 10^{" + f"{np.log10(regruns.parameters[i])}" +r"}$")
fig.subplots_adjust(right=0.8)
#cax = fig.add_axes([0.85, 0.15, 0.05, 0.7])
cax = fig.colorbar(im, ax=axes.ravel().tolist(), aspect=50)
cax.outline.set_linewidth(0)
return fig, axes
with plt.style.context('rapport'):
fig, axes = coeff_plot(lasso)
fig.savefig(fig_path / "lasso_coeff.png", dpi=300, transparent=True, bbox_inches='tight')
fig, axes = coeff_plot(ridge)
fig.savefig(fig_path / "ridge_coeff.png", dpi=300, transparent=True, bbox_inches='tight')
with plt.style.context('rapport'):
fig, ax = plt.subplots()
cmap_args = {'vmin': -1, 'vmax':1, 'cmap': "RdBu_r"}
im = ax.matshow(cv_results.coeffs[0], **cmap_args)
ax.axis('off')
cax = fig.colorbar(im, ax=ax)
cax.outline.set_linewidth(0)
fig.savefig(fig_path / "lm_coeff.png", dpi=300, transparent=True, bbox_inches='tight')
def plot_best(models):
means = np.asarray([np.mean(model.mse_test) for model in models])
best = means.argmin()
model = models[best]
mean, std = model.average()
fig, ax = plt.subplots(ncols=2)
cmap_args = {'vmin': -1, 'vmax':1, 'cmap': "RdBu_r"}
im = ax[0].matshow(mean, **cmap_args)
ax[1].matshow(std, **cmap_args)
fig.colorbar(im)
return fig, ax
def mse_evolution(regrun, ax=None):
if ax is None:
fig, ax = plt.subplots()
else:
fig = ax.figure()
#ax.plot(params, means)
#ax.fill_between(params, means-se, means+se, alpha=0.3)
ax.errorbar(regrun.parameters, regrun.train_mse, yerr=regrun.train_se_mse,
fmt='.-', ms=3, linewidth=0.5,
label=r'MSE Train')
ax.errorbar(regrun.parameters, regrun.test_mse, yerr=regrun.test_se_mse,
fmt='.-', ms=3, linewidth=0.5,
label=r'MSE Test')
ax.set_ylabel('MSE')
ax.set_xlabel('Regularization parameter')
ax.set_xscale('log')
ax.set_yscale('log')
ax2 = ax.twinx()
#ax2.plot(params, means)
#ax2.fill_between(params, means-se, means+se, alpha=0.3)
#ax2.errorbar([1], [1])
ax2.errorbar(regrun.parameters, regrun.train_r2, yerr=regrun.train_se_r2,
fmt='v-', ms=2, linewidth=0.5,
label=f'$R^2$ Train')
ax2.errorbar(regrun.parameters, regrun.test_r2, yerr=regrun.test_se_r2,
fmt='v-', ms=2, linewidth=0.5,
label=f'$R^2$ Test')
ax2.set_ylabel(r'$R^2$')
fig.legend(loc=9, ncol=2, frameon=False)
fig.tight_layout()
fig.subplots_adjust(top=0.75)
return fig, ax
with plt.style.context('rapport'):
fig, ax = mse_evolution(lasso)
fig.savefig(fig_path / "lasso_reg.png", dpi=300, transparent=True, bbox_inches='tight')
fig, ax = mse_evolution(ridge)
fig.savefig(fig_path / "ridge_reg.png", dpi=300, transparent=True, bbox_inches='tight')
#plt.show()
| projects/project2/notebook/Linear Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Boolean
# +
literal = ['A','B','C','Y']
for i in literal:
print(i,end='\t')
print()
for a in [0,1]:
for b in [0,1]:
for c in [0,1]:
print(a,end='\t')
print(b,end='\t')
print(c,end='\t')
print(a and b,end='\t')
print()
# -
def truthtable(y=0):
literal=['A','B','C','Y']
for i in literal:
print(i, end ="\t")
print()
for a in [0,1]:
for b in [0,1]:
for c in [0,1]:
print(a, end ="\t")
print(b, end ="\t")
print(c, end ="\t")
print(int(y(a,b,c)), end ="\t")
print()
# +
def y(a,b,c):
return a and b
truthtable(y)
# -
y = lambda a,b,c : a and b or not a and b
truthtable(y)
# # Example kmap
#example kmap long
y = lambda a,b,c : (not a and not b and not c) or (not a and not b and c)
truthtable(y)
#example kmap shot
y = lambda a,b,c : (not a and not b)
truthtable(y)
# # Examplr Gennext
#example gennext long
y = lambda a,b,c : (not a and not b) or (a and b and not c) or (a and not b)
truthtable(y)
#example gennext short
y = lambda a,b,c : (a and not c) or (not b)
truthtable(y)
# # Example Bob
# example Bob
y = lambda a,b,c : (a and c) or (not b and c) or (not a and b and c)
truthtable(y)
y = lambda a,b,c: c
truthtable(y)
# # Set
import numpy as np
N=20
np.random.seed(0)
index=np.arange(N)
A=np.random.randint(low=0,high=2,size=N)
B=np.random.randint(low=0,high=2,size=N)
C=np.random.randint(low=0,high=2,size=N)
for r in zip(index,A,B,C):
print(r)
print('index\tA\tB\tC\tY')
for r in zip(index,A,B,C):
i = r[0]
a = r[1]
b = r[2]
c = r[3]
print(i, end='\t')
print(a, end='\t')
print(b, end='\t')
print(c, end='\t')
print(int(a and not b ))
np.logical_and(A,np.logical_not(B))
index[np.logical_and(A,np.logical_not(B))]
#N(A^B)
np.count_nonzero(
np.logical_and(A,B)
)
#N(A)
np.count_nonzero(A)
#N(B)
np.count_nonzero(B)
#N(AUB)
np.count_nonzero(
np.logical_or(A,B)
)
# +
#N(AUB) = N(A) + N(B) - N(A^B)
#16 = 11 + 12 -7
# -
| dsi200_demo/_extra boolean.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
import re
import os
from time import sleep
import random
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from datetime import datetime
import sqlite3
from sqlite3 import Error
import pandas as pd
#Name of Database File that Will be Created
database = "transcripts1.db"
#Create SQL Lite Connection
def create_connection(db_file):
""" create a database connection to the SQLite database
specified by db_file
:param db_file: database file
:return: Connection object or None
"""
conn = None
try:
conn = sqlite3.connect(db_file)
return conn
except Error as e:
print(e)
return conn
#Connect and Create Table Transcripts if the Table Doesn't Exist
conn = create_connection(database)
c = conn.cursor()
c.execute('''CREATE TABLE IF NOT EXISTS TRANSCRIPTS ([TRANSCRIPTS] text PRIMARY KEY, [TICKER] text, [DATE] date,[TITLE] text)''')
conn.commit()
#Create list of tickers grouped up by exhange
nsdq_ticks = "data/nasdaqlisted.txt"
nyse_ticks = "data/companylist.csv"
nyse = pd.read_csv(nyse_ticks)
nsdq = pd.read_csv(nsdq_ticks,sep="|")
nsdq = nsdq[nsdq['Security Name'].str.contains("Common Stock",na=False)]
nsdq = nsdq[nsdq['ETF'].str.contains("N",na=False)]
nyse_tickers = list(nyse['Symbol'])
nsdq_tickers = list(nsdq['Symbol'])
nsdq_tickers[192]
# +
#Search through tickers in nasdaq list, load firefox browser and downloaded earning transcript text
transcripts = []
ticks = []
for i in nsdq_tickers[194:]:
try:
for j in range(1,8):
link ='https://www.fool.com/quote/nasdaq/apple/{}/earnings-call-transcripts'.format(i)
print(link)
driver = webdriver.Firefox()
driver.get(link)
sleep(3)
element = driver.find_element_by_xpath("/html/body/div[1]/div[2]/div/div/div[3]/div[2]/div[6]/div/div/article[{}]/div[2]/h4/a".format(j))
element.click()
soup = BeautifulSoup(driver.page_source, 'html.parser')
if "Earnings" in soup.find('h1').getText():
title = soup.find('h1').getText()
transcript_soup = soup.find_all('p')[3:]
try:
date = soup.find("span", id="date").getText()
except:
date = soup.find('h2').getText()
transcript = [i.getText() for i in transcript_soup]
c.execute('INSERT INTO TRANSCRIPTS VALUES (?,?,?,?)',[str(transcript),str(i),str(date),str(title)])
conn.commit()
driver.close()
else:
pass
except:
driver.close()
pass
# -
#Search through tickers in nyse list, load firefox browser and downloaded earning transcript text
for i in nyse_tickers[1501:]:
try:
for j in range(1,8):
link ='https://www.fool.com/quote/nyse/apple/{}/earnings-call-transcripts'.format(i)
driver = webdriver.Firefox()
driver.get(link)
sleep(2)
element = driver.find_element_by_xpath("/html/body/div[1]/div[2]/div/div/div[3]/div[2]/div[6]/div/div/article[{}]/div[2]/h4/a".format(j))
element.click()
soup = BeautifulSoup(driver.page_source, 'html.parser')
if "Earnings" in soup.find('h1').getText():
title = soup.find('h1').getText()
transcript_soup = soup.find_all('p')[3:]
try:
date = soup.find("span", id="date").getText()
except:
date = soup.find('h2').getText()
transcript = [i.getText() for i in transcript_soup]
c.execute('INSERT INTO TRANSCRIPTS VALUES (?,?,?,?)',[str(transcript),str(i),str(date),str(title)])
conn.commit()
driver.close()
else:
pass
except:
driver.close()
pass
| WebScraping/NLP_Web_Scraping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import sys
import shutil
from alpenglow.experiments import BatchFactorExperiment, ExternalModelExperiment
from alpenglow.evaluation import DcgScore
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
data = pd.read_csv('http://info.ilab.sztaki.hu/~fbobee/alpenglow/tutorial_dataset.csv', header=None, names=['time', 'user', 'item'])
# # alpenglow model
exp = ExternalModelExperiment(
period_length=60 * 60 * 24 * 7 * 4,
in_name_base="batches/batch",
mode="read",
)
res = exp.run(data)
res['dcg'] = DcgScore(res)
res['dcg'].mean()
# # lightfm
exp = ExternalModelExperiment(
period_length=60 * 60 * 24 * 7 * 4,
in_name_base="../lightfm/batches/batch",
mode="read",
)
res3 = exp.run(data)
res3['dcg'] = DcgScore(res3)
res3['dcg'].mean()
# # turicreate
exp = ExternalModelExperiment(
period_length=60 * 60 * 24 * 7 * 4,
in_name_base="../turicreate/batches/batch",
mode="read",
)
res4 = exp.run(data)
res4['dcg'] = DcgScore(res4)
res4['dcg'].mean()
# # libfm
exp = ExternalModelExperiment(
period_length=60 * 60 * 24 * 7 * 4,
in_name_base="../libfm/batches/batch",
mode="read",
)
res5 = exp.run(data)
res5['dcg'] = DcgScore(res5)
res5['dcg'].mean()
# # alpenglow batch factor model
exp = BatchFactorExperiment(
period_length=60 * 60 * 24 * 7 * 4,
negative_rate=5,
learning_rate = 0.05,
clear_model=True
)
res2 = exp.run(data)
res2['dcg'] = DcgScore(res2)
res2['dcg'].mean()
# # plotting
plt.figure(figsize=(15,8))
res.groupby(res.time//(60*60*24)).dcg.mean().plot()
res2.groupby(res2.time//(60*60*24)).dcg.mean().plot()
res3.groupby(res2.time//(60*60*24)).dcg.mean().plot()
res4.groupby(res2.time//(60*60*24)).dcg.mean().plot()
res5.groupby(res2.time//(60*60*24)).dcg.mean().plot()
plt.ylabel('daily average ndcg')
plt.legend(['alpenglow external', 'alpenglow', 'lightfm', 'turicreate', 'libfm'])
| examples/external_models/alpenglow/evaluate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# cd /Users/cfe/Dev/fastapi-nosql/
from app import db, crud, models
session = db.get_session()
data = models.data
data
crud.add_scrape_event(data, fresh=True)
asin = "TESTING123D"
for obj in models.Product.objects().all():
print(obj.asin)
if asin is not None:
print(f"Scrape events for {asin}")
q = models.ProductScrapeEvent.objects.filter(asin=asin)
for i, obj in enumerate(q):
print(i, obj.asin, obj.uuid)
q = models.Product.objects().all()
cquery = str(q._select_query())
print(cquery)
for row in session.execute(cquery):
print(row)
| nbs/Working with Cassandra Models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Truncation
#
# When the prior is very wide and simulation expense is high, it makes sense to focus our simulations on a certain observation $b_o$. We are effectively estimating the likelihood-to-evidence ratio on a small region around the $\theta_o$ which produced $x_o$. We do this marginally, therefore we take the product of marginal estimates and let that be our truncated region on which to estimate the likelihood-to-evidence ratio. This notebook demonstrates that technique.
# %load_ext autoreload
# %autoreload 2
# DON'T FORGET TO ACTIVATE THE GPU when on google colab (Edit > Notebook settings)
from os import environ
GOOGLE_COLAB = True if "COLAB_GPU" in environ else False
if GOOGLE_COLAB:
# !pip install git+https://github.com/undark-lab/swyft.git
# +
import numpy as np
import pylab as plt
import torch
import swyft
# +
device = 'cuda' if swyft.utils.is_cuda_available() else "cpu"
n_training_samples = 3000
n_parameters = 2
marginal_indices_1d, marginal_indices_2d = swyft.utils.get_corner_marginal_indices(n_parameters)
observation_key = "x"
n_posterior_samples_for_truncation = 10_000
n_weighted_samples = 10_000
# +
def model(v, sigma = 0.01):
x = v + np.random.randn(n_parameters)*sigma
return {observation_key: x}
v_o = np.zeros(n_parameters)
observation_o = model(v_o, sigma = 0.)
n_observation_features = observation_o[observation_key].shape[0]
observation_shapes = {key: value.shape for key, value in observation_o.items()}
# +
simulator = swyft.Simulator(
model,
n_parameters,
sim_shapes=observation_shapes,
)
low = -1 * np.ones(n_parameters)
high = 1 * np.ones(n_parameters)
prior = swyft.get_uniform_prior(low, high)
store = swyft.Store.memory_store(simulator)
store.add(n_training_samples, prior)
store.simulate()
# -
# ## creating a `do_round` function
#
# We call the process of training the marginal likelihood-to-evidence ratio estimator and estimating the support of the truncated prior a `round`. The output of a round is a `bound` object which is then used in the next `round`. It makes sense to encapsulate your round in a function which can be called repeatedly.
#
# We will start by truncating in one dimension (with hyperrectangles as bounds)
def do_round_1d(bound, observation_focus):
store.add(n_training_samples, prior, bound=bound)
store.simulate()
dataset = swyft.Dataset(n_training_samples, prior, store, bound = bound)
network_1d = swyft.get_marginal_classifier(
observation_key=observation_key,
marginal_indices=marginal_indices_1d,
observation_shapes=observation_shapes,
n_parameters=n_parameters,
hidden_features=32,
num_blocks=2,
)
mre_1d = swyft.MarginalRatioEstimator(
marginal_indices=marginal_indices_1d,
network=network_1d,
device=device,
)
mre_1d.train(dataset)
posterior_1d = swyft.MarginalPosterior(mre_1d, prior, bound)
new_bound = posterior_1d.truncate(n_posterior_samples_for_truncation, observation_focus)
return posterior_1d, new_bound
# ## Truncating and estimating the marginal posterior over the truncated region
#
# First we train the one-dimensional likelihood-to-evidence ratios, then we train the two-dimensional estimator on the truncated region. This region is defined by the `bound` object.
bound = None
for i in range(3):
posterior_1d, bound = do_round_1d(bound, observation_o)
network_2d = swyft.get_marginal_classifier(
observation_key=observation_key,
marginal_indices=marginal_indices_2d,
observation_shapes=observation_shapes,
n_parameters=n_parameters,
hidden_features=32,
num_blocks=2,
)
mre_2d = swyft.MarginalRatioEstimator(
marginal_indices=marginal_indices_2d,
network=network_2d,
device=device,
)
store.add(n_training_samples, prior, bound=bound)
store.simulate()
dataset = swyft.Dataset(n_training_samples, prior, store, bound = bound)
mre_2d.train(dataset)
# +
weighted_samples_1d = posterior_1d.weighted_sample(n_weighted_samples, observation_o)
posterior_2d = swyft.MarginalPosterior(mre_2d, prior, bound)
weighted_samples_2d = posterior_2d.weighted_sample(n_weighted_samples, observation_o)
# -
_, _ = swyft.plot.corner(
weighted_samples_1d,
weighted_samples_2d,
kde=True,
truth=v_o,
xlim=[-0.15, 0.15],
ylim_lower=[-0.15, 0.15],
bins=200,
)
# ## Repeat but truncate in two dimensions
#
# First we train the two-dimensional likelihood-to-evidence ratio. We use that bound to estimate the one-dimensional likelihood-to-evidence ratios.
def do_round_2d(bound, observation_focus):
store.add(n_training_samples, prior, bound=bound)
store.simulate()
dataset = swyft.Dataset(n_training_samples, prior, store, bound = bound)
network_2d = swyft.get_marginal_classifier(
observation_key=observation_key,
marginal_indices=marginal_indices_2d,
observation_shapes=observation_shapes,
n_parameters=n_parameters,
hidden_features=32,
num_blocks=2,
)
mre_2d = swyft.MarginalRatioEstimator(
marginal_indices=marginal_indices_2d,
network=network_2d,
device=device,
)
mre_2d.train(dataset)
posterior_2d = swyft.MarginalPosterior(mre_2d, prior, bound)
new_bound = posterior_1d.truncate(n_posterior_samples_for_truncation, observation_focus)
return posterior_2d, new_bound
bound = None
for i in range(3):
posterior_2d, bound = do_round_2d(bound, observation_o)
network_1d = swyft.get_marginal_classifier(
observation_key=observation_key,
marginal_indices=marginal_indices_1d,
observation_shapes=observation_shapes,
n_parameters=n_parameters,
hidden_features=32,
num_blocks=2,
)
mre_1d = swyft.MarginalRatioEstimator(
marginal_indices=marginal_indices_1d,
network=network_1d,
device=device,
)
mre_1d.train(dataset)
store.add(n_training_samples + 100, prior, bound=bound)
store.simulate()
dataset = swyft.Dataset(n_training_samples, prior, store, bound = bound)
mre_1d.train(dataset)
# +
posterior_1d = swyft.MarginalPosterior(mre_1d, prior, bound)
weighted_samples_1d = posterior_1d.weighted_sample(n_weighted_samples, observation_o)
weighted_samples_2d = posterior_2d.weighted_sample(n_weighted_samples, observation_o)
# -
_, _ = swyft.plot.corner(
weighted_samples_1d,
weighted_samples_2d,
kde=True,
truth=v_o,
xlim=[-0.15, 0.15],
ylim_lower=[-0.15, 0.15],
bins=200,
)
| notebooks/Examples - 2. Truncation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.9 64-bit
# language: python
# name: python3
# ---
from qiskit.circuit import Parameter, QuantumCircuit
import numpy as np
import matplotlib.pyplot as plt
theta_param = Parameter('θ')
phi_param = Parameter('φ')
# +
# circuit A
circuit_A = QuantumCircuit(1)
circuit_A.h(0)
circuit_A.rz(theta_param, 0)
# %matplotlib inline
circuit_A.draw('mpl')
# +
circuit_B = QuantumCircuit(1)
circuit_B.h(0)
circuit_B.rz(theta_param, 0)
circuit_B.rx(phi_param, 0)
# %matplotlib inline
circuit_B.draw('mpl')
# +
np.random.seed(0)
num_param = 1000
theta = [2*np.pi*np.random.uniform() for i in range(num_param)]
phi = [2*np.pi*np.random.uniform() for i in range(num_param)]
# +
from qiskit.visualization.bloch import Bloch
from qiskit.quantum_info import Statevector
def state_to_bloch(state_vec):
# Converts state vectors to points on the Bloch sphere
phi = np.angle(state_vec.data[1])-np.angle(state_vec.data[0])
theta = 2*np.arccos(np.abs(state_vec.data[0]))
return [np.sin(theta)*np.cos(phi),np.sin(theta)*np.sin(phi),np.cos(theta)]
# +
width, height = plt.figaspect(1/2)
fig = plt.figure(figsize=(width, height))
ax1, ax2 = fig.add_subplot(1, 2, 1, projection='3d'), fig.add_subplot(1, 2, 2, projection='3d')
b1,b2 = Bloch(axes=ax1), Bloch(axes=ax2)
b1.point_color, b2.point_color = ['tab:blue'],['tab:blue']
b1.point_marker, b2.point_marker= ['o'],['o']
b1.point_size, b2.point_size=[2],[2]
for i in range(num_param):
state_1 = Statevector.from_instruction(circuit_A.bind_parameters({theta_param:theta[i]}))
state_2 = Statevector.from_instruction(circuit_B.bind_parameters({theta_param:theta[i], phi_param:phi[i]}))
b1.add_points(state_to_bloch(state_1))
b2.add_points(state_to_bloch(state_2))
b1.show()
b2.show()
# +
# we can also use a circuit defined in qiskit library
from qiskit.circuit.library import PauliFeatureMap
qc_tl = PauliFeatureMap(5, reps = 1)
qc_tl.decompose().draw()
| qml/parametrzed_circuits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Series
import numpy as np
import pandas as pd
np.random.seed(12345)
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
PREVIOUS_MAX_ROWS = pd.options.display.max_rows
pd.options.display.max_rows = 20
np.set_printoptions(precision=4, suppress=True)
# ## Date and Time Data Types and Tools
from datetime import datetime
now = datetime.now()
now
now.year, now.month, now.day
delta = datetime(2011, 1, 7) - datetime(2008, 6, 24, 8, 15)
delta
delta.days
delta.seconds
from datetime import timedelta
start = datetime(2011, 1, 7)
start + timedelta(12)
start - 2 * timedelta(12)
# ### Converting Between String and Datetime
stamp = datetime(2011, 1, 3)
str(stamp)
stamp.strftime('%Y-%m-%d')
value = '2011-01-03'
datetime.strptime(value, '%Y-%m-%d')
datestrs = ['7/6/2011', '8/6/2011']
[datetime.strptime(x, '%m/%d/%Y') for x in datestrs]
from dateutil.parser import parse
parse('2011-01-03')
parse('Jan 31, 1997 10:45 PM')
parse('6/12/2011', dayfirst=True)
datestrs = ['2011-07-06 12:00:00', '2011-08-06 00:00:00']
pd.to_datetime(datestrs)
idx = pd.to_datetime(datestrs + [None])
idx
idx[2]
pd.isnull(idx)
# ## Time Series Basics
from datetime import datetime
dates = [datetime(2011, 1, 2), datetime(2011, 1, 5),
datetime(2011, 1, 7), datetime(2011, 1, 8),
datetime(2011, 1, 10), datetime(2011, 1, 12)]
ts = pd.Series(np.random.randn(6), index=dates)
ts
ts.index
ts + ts[::2]
ts.index.dtype
stamp = ts.index[0]
stamp
# ### Indexing, Selection, Subsetting
stamp = ts.index[2]
ts[stamp]
ts['1/10/2011']
ts['20110110']
longer_ts = pd.Series(np.random.randn(1000),
index=pd.date_range('1/1/2000', periods=1000))
longer_ts
longer_ts['2001']
longer_ts['2001-05']
ts[datetime(2011, 1, 7):]
ts
ts['1/6/2011':'1/11/2011']
ts.truncate(after='1/9/2011')
dates = pd.date_range('1/1/2000', periods=100, freq='W-WED')
long_df = pd.DataFrame(np.random.randn(100, 4),
index=dates,
columns=['Colorado', 'Texas',
'New York', 'Ohio'])
long_df.loc['5-2001']
# ### Time Series with Duplicate Indices
dates = pd.DatetimeIndex(['1/1/2000', '1/2/2000', '1/2/2000',
'1/2/2000', '1/3/2000'])
dup_ts = pd.Series(np.arange(5), index=dates)
dup_ts
dup_ts.index.is_unique
dup_ts['1/3/2000'] # not duplicated
dup_ts['1/2/2000'] # duplicated
grouped = dup_ts.groupby(level=0)
grouped.mean()
grouped.count()
# ## Date Ranges, Frequencies, and Shifting
ts
resampler = ts.resample('D')
# ### Generating Date Ranges
index = pd.date_range('2012-04-01', '2012-06-01')
index
pd.date_range(start='2012-04-01', periods=20)
pd.date_range(end='2012-06-01', periods=20)
pd.date_range('2000-01-01', '2000-12-01', freq='BM')
pd.date_range('2012-05-02 12:56:31', periods=5)
pd.date_range('2012-05-02 12:56:31', periods=5, normalize=True)
# ### Frequencies and Date Offsets
from pandas.tseries.offsets import Hour, Minute
hour = Hour()
hour
four_hours = Hour(4)
four_hours
pd.date_range('2000-01-01', '2000-01-03 23:59', freq='4h')
Hour(2) + Minute(30)
pd.date_range('2000-01-01', periods=10, freq='1h30min')
# #### Week of month dates
rng = pd.date_range('2012-01-01', '2012-09-01', freq='WOM-3FRI')
list(rng)
# ### Shifting (Leading and Lagging) Data
ts = pd.Series(np.random.randn(4),
index=pd.date_range('1/1/2000', periods=4, freq='M'))
ts
ts.shift(2)
ts.shift(-2)
# ts / ts.shift(1) - 1
ts.shift(2, freq='M')
ts.shift(3, freq='D')
ts.shift(1, freq='90T')
# #### Shifting dates with offsets
from pandas.tseries.offsets import Day, MonthEnd
now = datetime(2011, 11, 17)
now + 3 * Day()
now + MonthEnd()
now + MonthEnd(2)
offset = MonthEnd()
offset.rollforward(now)
offset.rollback(now)
ts = pd.Series(np.random.randn(20),
index=pd.date_range('1/15/2000', periods=20, freq='4d'))
ts
ts.groupby(offset.rollforward).mean()
ts.resample('M').mean()
# ## Time Zone Handling
import pytz
pytz.common_timezones[-5:]
tz = pytz.timezone('America/New_York')
tz
# ### Time Zone Localization and Conversion
rng = pd.date_range('3/9/2012 9:30', periods=6, freq='D')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts
print(ts.index.tz)
pd.date_range('3/9/2012 9:30', periods=10, freq='D', tz='UTC')
ts
ts_utc = ts.tz_localize('UTC')
ts_utc
ts_utc.index
ts_utc.tz_convert('America/New_York')
ts_eastern = ts.tz_localize('America/New_York')
ts_eastern.tz_convert('UTC')
ts_eastern.tz_convert('Europe/Berlin')
ts.index.tz_localize('Asia/Shanghai')
# ### Operations with Time Zone−Aware Timestamp Objects
stamp = pd.Timestamp('2011-03-12 04:00')
stamp_utc = stamp.tz_localize('utc')
stamp_utc.tz_convert('America/New_York')
stamp_moscow = pd.Timestamp('2011-03-12 04:00', tz='Europe/Moscow')
stamp_moscow
stamp_utc.value
stamp_utc.tz_convert('America/New_York').value
from pandas.tseries.offsets import Hour
stamp = pd.Timestamp('2012-03-12 01:30', tz='US/Eastern')
stamp
stamp + Hour()
stamp = pd.Timestamp('2012-11-04 00:30', tz='US/Eastern')
stamp
stamp + 2 * Hour()
# ### Operations Between Different Time Zones
rng = pd.date_range('3/7/2012 9:30', periods=10, freq='B')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts
ts1 = ts[:7].tz_localize('Europe/London')
ts2 = ts1[2:].tz_convert('Europe/Moscow')
result = ts1 + ts2
result.index
# ## Periods and Period Arithmetic
p = pd.Period(2007, freq='A-DEC')
p
p + 5
p - 2
pd.Period('2014', freq='A-DEC') - p
rng = pd.period_range('2000-01-01', '2000-06-30', freq='M')
rng
pd.Series(np.random.randn(6), index=rng)
values = ['2001Q3', '2002Q2', '2003Q1']
index = pd.PeriodIndex(values, freq='Q-DEC')
index
# ### Period Frequency Conversion
p = pd.Period('2007', freq='A-DEC')
p
p.asfreq('M', how='start')
p.asfreq('M', how='end')
p = pd.Period('2007', freq='A-JUN')
p
p.asfreq('M', 'start')
p.asfreq('M', 'end')
p = pd.Period('Aug-2007', 'M')
p.asfreq('A-JUN')
rng = pd.period_range('2006', '2009', freq='A-DEC')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts
ts.asfreq('M', how='start')
ts.asfreq('B', how='end')
# ### Quarterly Period Frequencies
p = pd.Period('2012Q4', freq='Q-JAN')
p
p.asfreq('D', 'start')
p.asfreq('D', 'end')
p4pm = (p.asfreq('B', 'e') - 1).asfreq('T', 's') + 16 * 60
p4pm
p4pm.to_timestamp()
rng = pd.period_range('2011Q3', '2012Q4', freq='Q-JAN')
ts = pd.Series(np.arange(len(rng)), index=rng)
ts
new_rng = (rng.asfreq('B', 'e') - 1).asfreq('T', 's') + 16 * 60
ts.index = new_rng.to_timestamp()
ts
# ### Converting Timestamps to Periods (and Back)
rng = pd.date_range('2000-01-01', periods=3, freq='M')
ts = pd.Series(np.random.randn(3), index=rng)
ts
pts = ts.to_period()
pts
rng = pd.date_range('1/29/2000', periods=6, freq='D')
ts2 = pd.Series(np.random.randn(6), index=rng)
ts2
ts2.to_period('M')
pts = ts2.to_period()
pts
pts.to_timestamp(how='end')
# ### Creating a PeriodIndex from Arrays
data = pd.read_csv('examples/macrodata.csv')
data.head(5)
data.year
data.quarter
index = pd.PeriodIndex(year=data.year, quarter=data.quarter,
freq='Q-DEC')
index
data.index = index
data.infl
# ## Resampling and Frequency Conversion
rng = pd.date_range('2000-01-01', periods=100, freq='D')
ts = pd.Series(np.random.randn(len(rng)), index=rng)
ts
ts.resample('M').mean()
ts.resample('M', kind='period').mean()
# ### Downsampling
rng = pd.date_range('2000-01-01 00:03:00', periods=12, freq='T')
ts = pd.Series(np.arange(12), index=rng)
ts
ts.resample('5min').count()
ts.resample('5min', closed='right').count()
ts.resample('5min', closed='right', label='right').sum()
ts.resample('5min', closed='right',
label='right', loffset='-1s').sum()
# #### Open-High-Low-Close (OHLC) resampling
ts.resample('5min').ohlc()
# ### Upsampling and Interpolation
frame = pd.DataFrame(np.random.randn(2, 4),
index=pd.date_range('1/1/2000', periods=2,
freq='W-WED'),
columns=['Colorado', 'Texas', 'New York', 'Ohio'])
frame
df_daily = frame.resample('D').asfreq()
df_daily
frame.resample('D').ffill()
frame.resample('D').ffill(limit=2)
frame.resample('W-THU').asfreq()
# ### Resampling with Periods
frame = pd.DataFrame(np.random.randn(24, 4),
index=pd.period_range('1-2000', '12-2001',
freq='M'),
columns=['Colorado', 'Texas', 'New York', 'Ohio'])
frame[:5]
annual_frame = frame.resample('A-DEC').mean()
annual_frame
# Q-DEC: Quarterly, year ending in December
annual_frame.resample('Q-DEC').ffill()
annual_frame.resample('Q-DEC', convention='end').ffill()
annual_frame.resample('Q-MAR').ffill()
# ## Moving Window Functions
close_px_all = pd.read_csv('examples/stock_px_2.csv',
parse_dates=True, index_col=0)
close_px = close_px_all[['AAPL', 'MSFT', 'XOM']]
close_px = close_px.resample('B').ffill()
close_px.AAPL.plot()
close_px.AAPL.rolling(250).mean().plot()
plt.figure()
appl_std250 = close_px.AAPL.rolling(250, min_periods=10).std()
appl_std250[5:12]
appl_std250.plot()
expanding_mean = appl_std250.expanding().mean()
plt.figure()
close_px.rolling(60).mean().plot(logy=True)
close_px.rolling('20D').mean()
# ### Exponentially Weighted Functions
plt.figure()
aapl_px = close_px.AAPL['2006':'2007']
ma60 = aapl_px.rolling(30, min_periods=20).mean()
ewma60 = aapl_px.ewm(span=30).mean()
ma60.plot(style='k--', label='Simple MA')
ewma60.plot(style='k-', label='EW MA')
plt.legend()
# ### Binary Moving Window Functions
plt.figure()
spx_px = close_px_all['SPX']
spx_rets = spx_px.pct_change()
returns = close_px.pct_change()
corr = returns.AAPL.rolling(125, min_periods=100).corr(spx_rets)
corr.plot()
plt.figure()
corr = returns.rolling(125, min_periods=100).corr(spx_rets)
corr.plot()
# ### User-Defined Moving Window Functions
plt.figure()
from scipy.stats import percentileofscore
score_at_2percent = lambda x: percentileofscore(x, 0.02)
result = returns.AAPL.rolling(250).apply(score_at_2percent)
result.plot()
pd.options.display.max_rows = PREVIOUS_MAX_ROWS
# ## Conclusion
| ch11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1A93lbv8AFP_" colab_type="text"
# ---
# #Notebook #3: Clasificación de imágenes y almacenamiento en base de datos
# ---
#
# Este notebook forma parte del trabajo de fin de grado realizado por:
#
# * <NAME>
#
# Con la colaboración de los tutores:
#
# * <NAME>
# * <NAME>
#
# El resto de este proyecto se puede encontrar en el siguiente repositorio de GitHub: [TFG-Amazon-Scraper](https://github.com/daniarnaizg/TFG-Amazon-Scraper)
# + [markdown] id="sIYyQ7pSOupt" colab_type="text"
# En la parte izquierda de esta ventana se encuentra la tabla de contenido de este notebook.
# + [markdown] id="blc29jDQCqZh" colab_type="text"
# ---
# # Importación de los clasificadores ya entrenados
#
# Para poder usar los clasificadores que hemos entrenado previamente, es necesario descargarlos desde el repositorio y cargarlos con la función load_model de Keras.
# + id="I8set9x1CuRY" colab_type="code" outputId="ccce055c-2003-4249-d380-d30d98c1f578" colab={"base_uri": "https://localhost:8080/", "height": 593}
# Descargamos los clasificadores ya entrenados.
# !wget https://github.com/daniarnaizg/TFG-Amazon-Scraper/raw/master/keras_models/clasificador-modelos.h5
# !wget https://github.com/daniarnaizg/TFG-Amazon-Scraper/raw/master/keras_models/clasificador-caras.h5
# + id="-Blsdf20Cy8Y" colab_type="code" outputId="bbca96d1-54fd-47cd-eb82-dc492f1505ff" colab={"base_uri": "https://localhost:8080/", "height": 35}
# !ls
# + id="XTxcOn3wC1B6" colab_type="code" outputId="4be14322-220d-44de-e15c-a9fb4ed8b526" colab={"base_uri": "https://localhost:8080/", "height": 413}
# Cargamos los modelos
from keras.models import load_model
calsificador_modelos = load_model('clasificador-modelos.h5')
calsificador_caras = load_model('clasificador-caras.h5')
# + [markdown] id="ty1jBHdLDv15" colab_type="text"
# ---
# # Implementación del clasificador
#
# En esta sección se implementan las funciones utilizadas para clasificar las imágenes.
# La función principal, `evaluate` se encarga de cargar la imagen, redimensionarla y convertirla en formato array. Las funciones `model_finder` y `face_finder` son las encargadas de clasificar las imágenes.
# + id="36RIUjQhC536" colab_type="code" colab={}
# Clasifica una imagen en función de si encuentr aun modelo o no con ayuda de calsificador previamente entrenado:
# 0 --> Modelo
# 1 --> Sin modelo
def model_finder(image):
predictions = calsificador_modelos.predict(image)[0]
return int(predictions.argmax())
# + id="Qdud--bzC56G" colab_type="code" colab={}
# Clasifica una imagen en función de si encuentra una cara o no con ayuda de calsificador previamente entrenado:
# 0 --> Cara
# 1 --> Sin cara
def face_finder(image):
predictions = calsificador_caras.predict(image)[0]
return int(predictions.argmax())
# + id="FHBgyIaiC58g" colab_type="code" colab={}
import cv2
import numpy as np
from keras.preprocessing.image import img_to_array
def evaluate(image_dir):
# load the image
image = cv2.imread(image_dir)
original = image.copy()
# pre-process the image for classification
image = cv2.resize(image, (28, 28))
image = image.astype("float") / 255.0
image = img_to_array(image)
image = np.expand_dims(image, axis=0)
prediction = model_finder(image)
final_prediction = ''
if prediction is 1:
final_prediction = "Sin modelo"
else:
if face_finder(image) is 0:
final_prediction = "Modelo con cara"
else:
final_prediction = "Modelo sin cara"
return final_prediction
# + [markdown] id="wU_7txO2FHow" colab_type="text"
# ---
# # Productos a clasificar
# La siguiente celda se encarga de descargar el archivo JSON generado por Scrapy que contiene los productos cuyas imágenes se desean clasificar:
# + id="GVa6ftr1C5-8" colab_type="code" outputId="14d8e7a4-bd4e-4ec5-f648-ae4cc9afa302" colab={"base_uri": "https://localhost:8080/", "height": 215}
# Descargamos el archivos JSON que contiene los productos a clasificar.
# !wget https://raw.githubusercontent.com/daniarnaizg/TFG-Amazon-Scraper/master/outputs/ejemplo_PRODUCTS.json
# + [markdown] id="fPE_F3BHFWrS" colab_type="text"
# ---
#
# # Descarga de imágenes
#
# Una vez tenemos el archivo JSON hay que descargar las imágenes asociadas a cada producto. Esto se hace a través de la librería [Requests](https://2.python-requests.org//es/latest/).
#
# Cada imagen se descarga con el código identificativo del producto al que pertenece como nombre, así podrán ser asociadas a dicho producto a la hpra de almacenarse.
# + id="hLeToge6C6BY" colab_type="code" colab={}
import json, requests
from PIL import Image
from io import BytesIO
# Recorrer el json y por cada asin descargar la primera url y asociarla a su asin (nombre de la imagen descargada).
def requestImg(image_name, url):
r = requests.get(url)
img = Image.open(BytesIO(r.content))
img.save("image_folder/" + image_name)
def downloadImage():
with open('ejemplo_PRODUCTS.json', 'r') as f:
data = json.load(f)
for i in range(len(data)):
try:
url = data[i]['image_urls'][0]
asin = data[i]['asin']
image_name = asin + '.jpg'
requestImg(image_name, url)
except IndexError:
print('Error con el producto ' + str(i + 1) + ', continuando...')
continue
f.close()
# + id="XaKW9Ty2C6Dl" colab_type="code" colab={}
# !mkdir image_folder
# + id="yWjWSZPxC6Fj" colab_type="code" outputId="03478609-742a-4791-fc5d-e4471e324f03" colab={"base_uri": "https://localhost:8080/", "height": 35}
downloadImage()
# + [markdown] id="q-LzroHUGnZW" colab_type="text"
# ---
# # Prueba manual
#
# A continuación se clasifica una imagen cualquiera para demostrar el funcionamiento del clasificador.
# Se puede cambiar la imagen a clasificar eligiendo otro código ASIN.
# + id="mPTOjNPRC6Kb" colab_type="code" outputId="c2560a10-06d6-46f7-f234-65dc0b7b1281" colab={"base_uri": "https://localhost:8080/", "height": 535}
from google.colab.patches import cv2_imshow
image_dir = 'image_folder/B07C81DMXS.jpg'
pred = evaluate(image_dir)
print('Predicción: '+ pred)
image = cv2.imread(image_dir)
peq = cv2.resize(image, (500, 500))
cv2_imshow(peq)
# + [markdown] id="z1YP13JGHAWJ" colab_type="text"
# ---
# # Descarga de base de datos
#
# A continuación se descarga la base de datos SQLite desde el repositorio del proyecto para poder almacenar la clasificación de las imágenes.
# + id="t6aiHQ9yC6M3" colab_type="code" outputId="7ec11ec3-2bc7-49db-b15e-f1f99b241b11" colab={"base_uri": "https://localhost:8080/", "height": 305}
# Descargamos la base de datos para poder trabajar sobre ella.
# !wget https://github.com/daniarnaizg/TFG-Amazon-Scraper/raw/master/outputs/ejemplo_DB.db
# + [markdown] id="8BMAin7bHVol" colab_type="text"
# ---
# #Proceso de clasificación y almacenamiento
#
# Las siguientes funciones serán las encargadas de clasificar, una a una cada imagen, y guardar la predicción en la base de datos.
# Primero se crea la conexión con la base de datos, a continuación se crea la nueva tabla llamada **'Predictions'** y por último se itera entre todas las imágenes que hemos desargado previamente y se clasifican por medio de la función `evaluate` que hemos definido al principio del notebook.
# + id="yvGph9ZAC6Pl" colab_type="code" colab={}
# Creación de la tabla PREDICTIONS en la base de datos actual-
# También sería posible añadir este campo a la tabla MAIN_AMAZON en vez de crear una nueva tabla.
# Esta tabla se crea con vistas al futuro, como por ejemplo clasificar todas las imagenes de un prodcuto en vez de la principal.
import sqlite3
def createTables(curr):
# self.dropAmazonTables()
curr.execute("""CREATE TABLE IF NOT EXISTS PREDICTIONS(
asin TEXT FOREING KEY NOT NULL,
prediction TEXT
)""")
def storePredictionInDb(curr, conn, item):
curr.execute("""INSERT INTO PREDICTIONS VALUES( ?, ?)""",(
item['asin'],
item['prediction']
))
conn.commit()
def closeDB(conn):
conn.close()
# + id="iJZY68VpDK9r" colab_type="code" colab={}
import json, glob
def savePredictions(curr, conn):
for imagePath in glob.glob('image_folder/*.jpg'):
item = {}
item['asin'] = imagePath[13:23]
item['prediction'] = evaluate(imagePath)
storePredictionInDb(curr, conn, item)
# + id="EyGCDPJXDLAQ" colab_type="code" colab={}
# Creamos la conexión con la base de datos
conn = sqlite3.connect('ejemplo_DB.db')
curr = conn.cursor()
# Creamos la tabla
createTables(curr)
# Guardamos la predicción en la tabla asociando cada una al ASIN del producto
savePredictions(curr, conn)
# Cerramos la conexión con la base de datos
closeDB(conn)
# + [markdown] id="oyGAXEchIgT_" colab_type="text"
# ---
# # Exportación de la base de datos ya actualizada
#
# Una vez terminado el proceso, la base de datos ha quedado actualizada con la nueva tabla.
#
# Cada predicción generada por el clasificador a quedado asociada al producto al que pertenece por medio del ASIN (Código identificativo de cada producto).
#
# + id="woOIWWN4DLCy" colab_type="code" colab={}
# Descargamos las base de datos actualizada en caso de que sea necesario
from google.colab import files
files.download('ejemplo_DB.db')
# + [markdown] id="xl80saeFJF-J" colab_type="text"
# ---
# # Generación del documento EXCEL
#
# En las siguiente celdas se va a proceder a la generación del documento excel que contiene todos los campos de todos los productos, así como los comentarios, los enlaces de las imágenes, y las predicciones generadas por los clasificadores.
#
# Esto se consigue a partir de la base de datos y por medio de una librería de Python llamada [Pandas](https://pandas.pydata.org/).
#
# Este proceso comienza con la creación de dataframes, que son las estructuras en forma de tablas de las que hace uso Pandas. Se generan a partir de las tablas de la base de datos con la función `read_sql_query`.
#
# A continuación podemos ver una muestra del contenido de cada dataframe creado:
# + id="8aBdtOn-DLFJ" colab_type="code" colab={}
# A continuación generamos un documento excel con las tablas de la base de datos
# Conectamos con la base de datos de nuevo
conn = sqlite3.connect('ejemplo_DB.db')
# Generamos un dataframe con cada tabla de la abse de datos
import pandas as pd
main_df = pd.read_sql_query("SELECT * FROM PRODUCTOS", conn)
images_df = pd.read_sql_query("SELECT * FROM IMAGENES", conn)
comments_df = pd.read_sql_query("SELECT * FROM COMENTARIOS", conn)
predictions_df = pd.read_sql_query("SELECT * FROM PREDICTIONS", conn)
# + id="p97lu1zhDLLp" colab_type="code" outputId="fda21625-b642-430c-b516-b61decd32d8b" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Dataframe que contiene los campos principales de cada producto. Se ha creado
# a partir de la tabla 'Productos' en la base de datos.
main_df.head()
# + id="yprKbQxODLJk" colab_type="code" outputId="43ea00e7-0079-4ee1-9f4c-798c6a75b807" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Dataframe que contiene los enlaces de las imágenes de cada producto.
# Se ha creado a partir de la tabla 'Imagenes' en la base de datos.
images_df.head()
# + id="UdyvLsBDC6SB" colab_type="code" outputId="1c203665-f785-40a1-9a37-9658f769d211" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Dataframe que contiene los comentarios asociados a cada producto.
# Se ha creado a partir de la tabla 'Comentarios' en la base de datos.
comments_df.head()
# + id="5NNT1wAQC6UU" colab_type="code" outputId="59fb90c4-2773-48a7-aa68-e3bfd5370f4d" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Dataframe que contiene las predicciones de las imágenes de cada producto.
# Se ha creado a partir de la tabla 'Predicciones' en la base de datos.
predictions_df.head()
# + [markdown] id="nbJh3X4SKv7Y" colab_type="text"
# ---
# # Union de los dataframes 'Productos' y 'Predicciones'
#
# A diferencia que en la base de datos, para el documento Excel se va a almacenar el campo 'prediction' junto al resto de campos de cada producto. En la base de datos se ha decidido utilizar una tabla a parte ya que una de las posibles líneas de futuro del proyecto sería la clasificación de otras carácterísticas de un producto, las cuales se almacenarías en esa tabla ya existente.
#
# Esta unión se consigue con la ayuda de la función `merge` implementada en la ibrería de Pandas. Esto es posible gracias a la asociación a cada producto por medio del ASIN.
#
# En este caso también se va a proceder a añadir el campo 'url' a este dataframe para una mejor visualización general una vez generado el documento Excel.
# + id="XIxvZRqhDZBe" colab_type="code" outputId="c220d9e1-7ab3-45a1-b1af-f132d7070ffb" colab={"base_uri": "https://localhost:8080/", "height": 204}
# Hacer un merge del dataframen predictions con el dataframe principal para añadir el campo 'predicción'
final_df = pd.merge(main_df, predictions_df, on='asin')
# Añadir la url del producto en un nuevo campo llamardo url
# Se sacará a partir del asin -> https://www.amazon.com/dp/[ASIN]/
final_df['url'] = final_df.apply(lambda row: 'https://www.amazon.com/dp/' + row.asin + '/', axis=1)
final_df.head()
# + [markdown] id="O8HsOnvWMggT" colab_type="text"
# ---
# #Generación del documento Excel
#
# En esta sección se va a generar el documento Excel a partir de los dataframes anteriores. Se generará una página por cada dataframe, por lo que consisitirá de:
#
# * **Productos**: Contiene los princiaples campos de cada producto, además de la predicción de la imagen generada por los clasficadores.
# * **Imágenes**: Contiene los enlaces de las imágenes asociadas a cada producto.
# * **Comentarios**: Contiene los comentarios que los clientes han dejado de cada producto.
#
# Este proceso se consigue a través de [XlsxWriter](https://xlsxwriter.readthedocs.io/), un módulo de Python que funciona a su vez a través de Pandas.
#
# Al finalizar la generación del docuemento, lo descargamos para su posterior uso.
# + id="TlSoWQ2oTSdw" colab_type="code" outputId="cf052172-b4e7-42df-ac3f-88e57fdf338c" colab={"base_uri": "https://localhost:8080/", "height": 107}
# El siguiente paso será exportar los dataframes a un documento excel.
# Herramienta para escribir en formato xlsx
# !pip install XlsxWriter
# + id="rONdKLi3O1YK" colab_type="code" colab={}
# Create a Pandas Excel writer using XlsxWriter as the engine.
writer = pd.ExcelWriter('ejemplo-EXCEL.xlsx', engine='xlsxwriter')
# + id="NPTdZE_JR1Fc" colab_type="code" colab={}
# Write each dataframe to a different worksheet.
final_df.to_excel(writer, index=False, sheet_name='Productos')
images_df.to_excel(writer, index=False, sheet_name='Imagenes')
comments_df.to_excel(writer, index=False, sheet_name='Comentarios')
# + id="Pfk2zEPWYRqm" colab_type="code" colab={}
# Close the Pandas Excel writer and output the Excel file.
writer.save()
# + id="ZJs45c-wYb46" colab_type="code" outputId="64cf17f8-5145-4f05-8702-229599110469" colab={"base_uri": "https://localhost:8080/", "height": 53}
# !ls
# + id="rx0pjjYqYf7B" colab_type="code" colab={}
# Descargamos el documento creado
files.download('ejemplo-EXCEL.xlsx')
# + [markdown] id="sO1CYTITN9aI" colab_type="text"
# ---
# #Visualización de datos
#
# Por último, en las siguientes celdas se muestra una potente herramienta que hace posible la visualización de forma totalmente interactiva de los datos que hemos extraído y almacenado.
#
# Esta herramienta es [pivottablejs](https://pivottable.js.org/). Funciona con JavaScript para crear un docuemtno HTML que podemos ver a continuación:
# + id="ySUYvVzvWLvv" colab_type="code" outputId="c71904e0-edec-4f48-aec9-cfa28ce7d442" colab={"base_uri": "https://localhost:8080/", "height": 89}
# !pip install pivottablejs
# + id="NIw6ddXPWRlI" colab_type="code" outputId="7f243311-e8df-4ae0-b322-68e58bbd503f" colab={"base_uri": "https://localhost:8080/", "height": 203}
import IPython
from pivottablejs import pivot_ui
pivot_ui(final_df, outfile_path='pivottablejs.html')
IPython.display.HTML(filename='pivottablejs.html')
# + id="OZlxvK2OWZoZ" colab_type="code" colab={}
# + id="N3wElyDnWv-7" colab_type="code" colab={}
# + id="1VqQwoXrW3eO" colab_type="code" colab={}
| colab notebooks/Clasificador_final_+_excel_+_db.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="1LEMD4f9dG3U"
# # Character Recurrent Neural Network
# - mimicing Shakespeare's writing style
# - Naive RNN
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" executionInfo={"elapsed": 4497, "status": "ok", "timestamp": 1559555773673, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="ODxhI-YBeYKz" outputId="ce716e54-acc0-40e1-d04f-33bf670cc3d0"
# !rm -r data
import os
try:
os.mkdir("./data")
except:
pass
# !wget https://raw.githubusercontent.com/GunhoChoi/PyTorch-FastCampus/master/05_RNN/2_Char_RNN/data/linux.txt -P ./data
# + [markdown] colab_type="text" id="X0FyjLatdG3V"
# ## 1. Settings
# ### 1) Import required libraries
# + colab={} colab_type="code" id="VVeUr2FjdG3X"
import torch
import torch.nn as nn
# + colab={} colab_type="code" id="TwWbv_-KdG3a"
import unidecode
import string
import random
import re
import time, math
# + [markdown] colab_type="text" id="62S5HbY2dG3d"
# ## 2) Hyperparameter
# + colab={} colab_type="code" id="oVeZybezdG3e"
num_epochs = 1000
print_every = 100
plot_every = 10
chunk_len = 200
hidden_size = 100
batch_size = 1
num_layers = 1
embedding_size = 70
lr = 0.002
# + [markdown] colab_type="text" id="_eKGWH4BdG3h"
# ## 2. Data
# ### 1) Prepare characters
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 5120, "status": "ok", "timestamp": 1559555774317, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="2Cg5-Wp0dG3i" outputId="096340c6-5d7b-4052-9e13-fa4832ba8cc9"
all_characters = string.printable
n_characters = len(all_characters)
print(all_characters)
print('num_chars = ', n_characters)
# + [markdown] colab_type="text" id="AxeLPVWsdG3m"
# ### 2) Get text data
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 5093, "status": "ok", "timestamp": 1559555774318, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="5LEkE0bxdG3n" outputId="0cc4bdfd-ca9f-4d1e-acba-cda20b40dec3"
file = unidecode.unidecode(open('./data/linux.txt').read())
file_len = len(file)
print('file_len =', file_len)
# + [markdown] colab_type="text" id="xmTf-IBOdG3r"
# ## 3. Functions for text processing
# ### 1) Random Chunk
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" executionInfo={"elapsed": 5081, "status": "ok", "timestamp": 1559555774319, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="bXDt_MmzdG3s" outputId="2d5ecd26-2dc3-4302-fd0c-963cfe376bf6"
def random_chunk():
start_index = random.randint(0, file_len - chunk_len)
end_index = start_index + chunk_len + 1
return file[start_index:end_index]
print(random_chunk())
# + [markdown] colab_type="text" id="DsSgzWIRdG3v"
# ### 2) Character to tensor
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 5071, "status": "ok", "timestamp": 1559555774320, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="fhY5WxwsdG3v" outputId="00c4759e-4334-4a99-8fe6-f404fb2f52e4"
def char_tensor(string):
tensor = torch.zeros(len(string)).long()
for c in range(len(string)):
tensor[c] = all_characters.index(string[c])
return tensor
print(char_tensor('ABCdef'))
# + [markdown] colab_type="text" id="Boc9LQimdG3y"
# ### 3) Chunk into input & label
# + colab={} colab_type="code" id="nDoMmRY2dG3z"
def random_training_set():
chunk = random_chunk()
inp = char_tensor(chunk[:-1])
target = char_tensor(chunk[1:])
return inp, target
# + [markdown] colab_type="text" id="zsHsRxlodG31"
# ## 3. Model & Optimizer
# ### 1) Model
# + colab={} colab_type="code" id="abZpjt5udG33"
class RNN(nn.Module):
def __init__(self, input_size, embedding_size, hidden_size, output_size, num_layers=1):
super(RNN, self).__init__()
self.input_size = input_size
self.embedding_size = embedding_size
self.hidden_size = hidden_size
self.output_size = output_size
self.num_layers = num_layers
self.encoder = nn.Embedding(self.input_size, self.embedding_size)
self.rnn = nn.GRU(self.embedding_size,self.hidden_size,self.num_layers)
self.decoder = nn.Linear(self.hidden_size, self.output_size)
def forward(self, input, hidden):
out = self.encoder(input.view(1,-1))
out,hidden = self.rnn(out,hidden)
out = self.decoder(out.view(batch_size,-1))
return out,hidden
def init_hidden(self):
hidden = torch.zeros(self.num_layers, batch_size, self.hidden_size)
return hidden
model = RNN(n_characters, embedding_size, hidden_size, n_characters, num_layers)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" executionInfo={"elapsed": 5063, "status": "ok", "timestamp": 1559555774325, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="rOVPmyxodG37" outputId="49e30928-095f-4ade-c137-a9525276994e"
inp = char_tensor("A")
print(inp)
hidden = model.init_hidden()
print(hidden.size())
out,hidden = model(inp,hidden)
print(out.size())
# + [markdown] colab_type="text" id="yfpAh13edG3_"
# ### 2) Loss & Optimizer
# + colab={} colab_type="code" id="8wRW02JXdG3_"
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
loss_func = nn.CrossEntropyLoss()
# + [markdown] colab_type="text" id="hoiLqf3_dG4C"
# ### 3) Test function
# + colab={} colab_type="code" id="rVTQky8vdG4D"
def test():
start_str = "b"
inp = char_tensor(start_str)
hidden = model.init_hidden()
x = inp
print(start_str,end="")
for i in range(200):
output,hidden = model(x,hidden)
output_dist = output.data.view(-1).div(0.8).exp()
top_i = torch.multinomial(output_dist, 1)[0]
predicted_char = all_characters[top_i]
print(predicted_char,end="")
x = char_tensor(predicted_char)
# + [markdown] colab_type="text" id="JNaIweaudG4G"
# ## 4. Train
# + colab={"base_uri": "https://localhost:8080/", "height": 3641} colab_type="code" executionInfo={"elapsed": 298689, "status": "ok", "timestamp": 1559556067982, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "04388737836176863066"}, "user_tz": -540} id="6CnV3_andG4H" outputId="9e4681f5-2ecf-4439-9b0b-42c034933c92"
for i in range(num_epochs):
inp,label = random_training_set()
hidden = model.init_hidden()
loss = torch.tensor([0]).type(torch.FloatTensor)
optimizer.zero_grad()
for j in range(chunk_len-1):
x = inp[j]
y_ = label[j].unsqueeze(0).type(torch.LongTensor)
y,hidden = model(x,hidden)
loss += loss_func(y,y_)
loss.backward()
optimizer.step()
if i % 200 == 0:
print("\n",loss/chunk_len,"\n")
test()
print("\n","="*100)
# + colab={} colab_type="code" id="SV7mBpxtjJsy"
| Recurrent Neural Network/Char_RNN_GRU_Linux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Разработать программу, для решения численным методом гравитационной задачи N тел.
# Эволюция системы N материальных точек описывается следующей системой уравнений:
# \begin{equation}
# \begin{cases}
# \frac{dr_i}{dt} = v_i\\
# \frac{dv_i}{dt} = \displaystyle\sum_{i \ne j}^{N} Gm_j\frac{r_j - r_i}{|r_j - r_i|^3}
# \end{cases}
# \end{equation}
import numpy as np
import time
from scipy.constants import g
import matplotlib.pyplot as plt
from matplotlib.animation import ArtistAnimation
from IPython.display import HTML
# #### Функция `getAcc` находит проекции ускорения на оси x и y.
def getAcc(pos, mass, G, softening):
x = pos[:, 0:1]
y = pos[:, 1:2]
dx = x.T - x
dy = y.T - y
inv_r3 = (dx ** 2 + dy ** 2 + softening ** 2) ** (-1.5)
ax = G * (dx * inv_r3) @ mass
ay = G * (dy * inv_r3) @ mass
a = np.hstack((ax, ay))
return a
# ### Задание начальных условий:
t = 0
t_end = 10.0
dt = 0.01
softening = 0.1
field_size = 500
n = int(input("Введите количество частиц: "))
des_1 = int(input("Введите 1, если масса различна, иначе 2: "))
des_2 = int(input("Введите 1, если первоначальная скорость = 0, иначе 2: "))
# +
if des_1 == 1:
mass = np.random.random((n, 1)).astype(float) * 10 ** 4
else:
mass = np.ones((n, 1)) * 2 * 10**4
if des_2 == 1:
vel = np.zeros((n, 2))
else:
vel = np.random.randn(n, 2)
pos = np.random.randn(n, 2) * 100
# -
acc = getAcc(pos, mass, g, softening)
color = []
pos_x = {}
pos_y = {}
for i in range(n):
pos_x[i] = []
pos_y[i] = []
color.append((
np.random.random(),
np.random.random(),
np.random.random()
))
# ### Построение модели
# #### Переменная `step_count` содержит количетсво шагов моделирования.
step_count = int(np.ceil(t_end / dt))
# +
fig, ax = plt.subplots()
ax.set_xlim((-field_size, field_size))
ax.set_ylim((-field_size, field_size))
plt.close()
frames = []
# -
# ### Осуществление вычислений
# Код просчитывает ускорение и скорость для всех заданных точек.
#
# А также сохраняет кадры для анимации.
for i in range(step_count):
vel += acc * dt / 2.0
pos += vel * dt
line = []
for j in range(len(pos)):
pos_x[j].append(pos[j][0])
pos_y[j].append(pos[j][1])
temp, = ax.plot(pos_x[j], pos_y[j], color=color[j], linewidth=1)
line.append(temp,)
frames.append(line)
acc = getAcc(pos, mass, g, softening)
vel += acc * dt / 2.0
t += dt
# Создать анимацию:
anim = ArtistAnimation(
fig,
frames,
interval=60,
blit=True,
repeat=True
)
# Отобразить анимацию.
HTML(anim.to_html5_video())
| individual tasks/N Body simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeClassifier
data = pd.read_csv("../data/house_location/house_location_data.csv")
test = pd.read_csv("../data/house_location/house_location_new_data_with_labels.csv")
# +
cols = ["beds", "bath", "price", "year_built", "sqft", "price_per_sqft", "elevation"]
X = data.loc[:, cols].values
y = data.in_sf.values
X_test = test.loc[:, cols].values
y_test = test.in_sf.values
# -
model = DecisionTreeClassifier(max_depth=10).fit(X, y)
model.score(X, y)
model.score(X_test, y_test)
with open("house_location_model.pkl", "w") as file:
pickle.dump(model, file)
model2 = pickle.load("house_location_model.pkl")
model2.score(X_test, y_test)
model2.predict(X_test)
import joblib
joblib.dump(model, 'house_location_model.pkl', compress=9)
model_clone = joblib.load('house_location_model.pkl')
model_clone
model_clone.score(X_test, y_test)
| 1_5_vorhersagen_treffen/Tutorial - Vorhersagen treffen - preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fh2cpWFgKlwv"
# # **AULA 03**
#
# * Introdução ao Python
#
# * Variáveis
#
# * Manipulação de Strings
#
# * Comando Input
#
# + [markdown] id="PfwIhhSvIf4S"
# # **Comando Print()**
# + id="RsYbievAu_9i" colab={"base_uri": "https://localhost:8080/"} outputId="e0bd83ae-710a-44b2-a6c1-e47827d772a3"
print('Olá mundo!!!')
# + colab={"base_uri": "https://localhost:8080/"} id="nkEC1i5WqnTo" outputId="bb78eec6-2dad-4115-a8c9-1decbefdad84"
print("São Paulo é o melhor time")
# + colab={"base_uri": "https://localhost:8080/"} id="EWOvSSuWvU0_" outputId="3037158d-d85a-4cb8-eadb-9a276c99b2db"
print(7+4)
print (55*43)
print("55*43")
print ("55""43")
# + [markdown] id="s3_POfLKI3Fe"
# Se colocarmos "aspas" nos números o resultado é outro
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="HzopBKyHI314" outputId="453a8b35-ada2-486f-cbeb-b19f91154651"
print('7'+'4')
# + [markdown] id="UBlwfG0MDUE0"
# # **Operadores em Python**
# + [markdown] id="AH9aupTPJ2sN"
# * Operadores Aritméticos
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="Ju8VKML7G_HO" outputId="3a70bb10-b306-4598-b9f0-bb34315d77eb"
a = 2 + 3
b = 2 * 3
c = 2 - 3
d = 2 / 3
e = 8 % 4
print(a)
print(b)
print(c)
print(d)
print(e)
# + id="U2-OamwBIKpS" colab={"base_uri": "https://localhost:8080/"} outputId="42127315-0140-45ed-ad75-2efd68ec4751"
soma = 5 + 5
subtracao = 10 - 5
multiplicacao = 10 * 5
divisao = 10 / 2
print(soma)
print(subtracao)
print(multiplicacao)
print(divisao)
# + [markdown] id="B6EAI1ByJ8aM"
# * Operadores Relacionais
# + colab={"base_uri": "https://localhost:8080/"} id="PQLLXrXmFZCT" outputId="487313ab-df52-4352-8e0c-f04da0b2e804"
2>3
4!=5
4!=4
"gustavo" == "GUSTAVO"
# + [markdown] id="XOs9OQG8KDNi"
# * Operadores Lógicos
# + colab={"base_uri": "https://localhost:8080/"} id="ROnieyJeFrQT" outputId="f476acef-f7a6-4269-e7b3-0833af5c0b2b"
nota = 3
nota <= 6 and nota >= 4
# + colab={"base_uri": "https://localhost:8080/"} id="L07VwY1vu3q1" outputId="4930a3fb-3475-4713-8b8c-c6d469c827bd"
2<3 and 1<8
# + colab={"base_uri": "https://localhost:8080/"} id="fMFmfk66u_AO" outputId="acc5f65e-4edf-4091-a923-1bb0450eaf42"
2<4 and 1>3
# + colab={"base_uri": "https://localhost:8080/"} id="BHrSKEhrvLlu" outputId="b11f26e7-c712-4187-d8bc-5d6979d51120"
2<4 or 1>3
# + colab={"base_uri": "https://localhost:8080/"} id="1Ar54L6-vk7u" outputId="304b40fc-e7e2-4167-c2dc-78364b5bdc95"
not True
# + colab={"base_uri": "https://localhost:8080/"} id="dX5dfGXWvyPD" outputId="80502da6-f31d-43f4-c940-0c89e25c1f07"
not False
# + colab={"base_uri": "https://localhost:8080/"} id="KLU9rvfXv5_D" outputId="020d3496-6e0b-405e-e8a7-29669ee98407"
not not True
# + [markdown] id="7Dk2_jXq7pmV"
# # **`VARIÁVEIS`**
#
#
#
#
# + [markdown] id="Lk7MeOMsnsV-"
# Exemplo 1
# + colab={"base_uri": "https://localhost:8080/"} id="bN8hJ1hwntpM" outputId="c10fdb23-e6bf-4312-da24-d551ee1a23bc"
b = 42
print (b)
print(type(b))
# + [markdown] id="D0F-aT_fn96-"
# Exemplo 2:
# + colab={"base_uri": "https://localhost:8080/"} id="xHirzCAhoaYa" outputId="2a02b57f-9015-4150-df2a-f86a6c4e57b4"
b = 1.8
print(b)
print(type(b))
# + [markdown] id="CYSBwP92odE5"
# Exemplo 3:
# + colab={"base_uri": "https://localhost:8080/"} id="82QMlZeJoe0I" outputId="9df9dbe7-6e0b-4056-b08a-f1ce86a01547"
c = "<NAME>! Eu tenho a força!"
print(c)
print (type(c))
# + [markdown] id="293GNmrjoogI"
# Tipos de variáveis
# + colab={"base_uri": "https://localhost:8080/"} id="DagjUieX7wXr" outputId="c6bbff72-f6be-4220-c426-0baa4ad72557"
print(type(c))
print(type(b))
print(type(a))
# + [markdown] id="nXjKf5FV0XXH"
# ________________________________________________________________________________
# + [markdown] id="VTwn121qpiaB"
# **Sintaxe elegante**
# + colab={"base_uri": "https://localhost:8080/"} id="mFAX4nv5pljv" outputId="df8ff7bc-12a5-48b6-a5bd-8246e2d05afe"
print('Olá Mundo')
# + colab={"base_uri": "https://localhost:8080/"} id="0brqbZv2prE1" outputId="b120205f-47f1-4518-c480-31ed6c946424"
x = 'Blue'
print(x)
# + [markdown] id="4_4DnczJpwyU"
# **Tipagem dinâmica**
# + colab={"base_uri": "https://localhost:8080/"} id="CIjcCQLip1-0" outputId="3c7aa047-1a9d-4e68-af42-746f0d0b5b1e"
b,t = 10,20
print('Resultado: ', b+t)
# + colab={"base_uri": "https://localhost:8080/"} id="-StxZZaa25x3" outputId="7ec0a5b4-e3f6-436c-ae4a-73982f5475c5"
a,b,c,d = 1,2,3,4
print ("Soma:", a+b, "Multiplicação:",c*d)
print ("Soma", a+b+c+d)
# + [markdown] id="MG0OEAlEqBbi"
# **Tipagem forte**
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="NWAtC6OyqEJQ" outputId="b3d00a30-9f1a-4263-aee4-3e24620be27e"
i,j = 10, 'Gustavo'
print(i)
print(j)
print(i+j)
# + [markdown] id="R4lDgs_2qRGi"
# **Estrutura de dados de alto nível**
# + colab={"base_uri": "https://localhost:8080/"} id="U-mkbzUVqVJE" outputId="0f894025-01a3-4443-f08f-22b284f09617"
l = [3,2,1]
m = l
l[1] = 8
print(m)
print()
m = 2
l=m
l=3
m = l
print(l)
print(m)
# + colab={"base_uri": "https://localhost:8080/"} id="Db33l46Uqjd4" outputId="6775be62-47a6-4e46-943c-4eaee10e8f5b"
d = ['23', 'Sexta=feira', 'Março','2021']
print(d)
# + [markdown] id="03Oijd418Aaz"
# # **COMANDO INPUT**
# + colab={"base_uri": "https://localhost:8080/"} id="QzkOLy_f8GRt" outputId="7c1a3fb7-fc57-4d5f-81e4-fa15f6c7aeb1"
nome = input("Digite seu nome completo")
print(nome)
# + colab={"base_uri": "https://localhost:8080/"} id="V4gupj7Y9PLE" outputId="cadcf5d2-0b29-4e68-a272-b716186fd660"
num = int(input("Digite seu número da sorte: "))
print(num)
# + colab={"base_uri": "https://localhost:8080/"} id="_Jx1NRDc9ali" outputId="512eb2e4-0b2c-4eda-9266-333647155a9e"
num = float(input("Digite sua altura: "))
print (num)
# + [markdown] id="0EwoQN303Xj7"
# # **STRINGS**
# + colab={"base_uri": "https://localhost:8080/"} id="7cqs6DGN2BL0" outputId="9461976f-ab1d-4494-cc83-9a68ff535659"
print("Qual é o seu nome?")
# + [markdown] id="Rj_N_O_K3djP"
# **CONCATENAÇÃO DE STRINGS**
# + [markdown] id="VzKPwNfn3kJs"
# **Exemplo 1:**
# + colab={"base_uri": "https://localhost:8080/"} id="bU7756132yRr" outputId="d478d2de-d6dd-42fa-eea2-ba65e46e9ae2"
print("Programação"+"Python")
# + [markdown] id="I4HmjZNK3lum"
# **Exemplo 2:**
# + colab={"base_uri": "https://localhost:8080/"} id="gkJeFUJ63A6l" outputId="07e891b5-4b1e-4450-b149-cfb0e87f41a9"
p = "Programação"
py = "Python"
prog = p + py
print(prog)
# + [markdown] id="Tb1gqCTs-Zvt"
# **MANIPULAÇÃO DE STRINGS**
# + [markdown] id="05Cesvxj-fGp"
# Utilizando *len()*
# + colab={"base_uri": "https://localhost:8080/"} id="pKoXhhOt5WLY" outputId="59afd2d0-d34d-4a0d-bfa5-f5446d8e38d5"
teste = "Programação Python"
len(teste)
# + [markdown] id="9S9nIyvH_S2J"
# Utilizando *capitalize()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="949xGhAu_GAx" outputId="9eeb6369-49a3-4a10-a58b-4bfb675243f6"
a = "python"
a.capitalize()
# + [markdown] id="yYeymIKh_3MT"
# Utilizando *count()*
# + colab={"base_uri": "https://localhost:8080/"} id="FJKsW4cO_77h" outputId="d34213b2-718b-46aa-b716-c5d15f3ba9e0"
b = "Linguagem Python"
b.count("y")
# + [markdown] id="dtubZ_KuBQqX"
# Utilizando *starswith()*
# + colab={"base_uri": "https://localhost:8080/"} id="JHGXavHiBdCz" outputId="013c8287-5cd6-4cd5-e246-7005c83922a4"
d = "Python"
d.startswith("Py")
# + [markdown] id="vVMFV5j0B9-5"
# Utilizando *endswith()*
# + colab={"base_uri": "https://localhost:8080/"} id="KQBrMxwBBs-z" outputId="831456fc-3812-42a9-bc7b-80dc76035ca5"
d = "Python"
d.endswith("Py")
# + [markdown] id="JbCc9VWTWQjx"
# Utilizando *isalpha()*
# + colab={"base_uri": "https://localhost:8080/"} id="CKshmwCIWEAa" outputId="28a99d39-95d2-4bf0-c821-fd4774be212a"
f = "Python"
f.isalpha()
# + [markdown] id="A_C8yMA-WZWi"
# Utilizando *islower()*
# + colab={"base_uri": "https://localhost:8080/"} id="Ut0wFxBrWfWO" outputId="b48ff2c3-6ba2-45a5-b002-daec0d73de59"
g = "Python"
g.islower()
# + [markdown] id="W4875nFiWts2"
# Utilizando issuper()
# + colab={"base_uri": "https://localhost:8080/"} id="0d08gxlTWwCR" outputId="c8cfbe64-ec28-40a3-cb58-b794291bab6c"
h = "# PYTHON 12"
h.isupper()
# + [markdown] id="yjgo_zFCpc6o"
# Utilizando *lower()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="s26M_XQBpfKK" outputId="6e46524c-8474-4a2c-97b1-50c2a44888a1"
i = "#PYTHON 3"
i.lower()
# + [markdown] id="x-cdcVLlp1Zq"
# Utilizando *upper()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="4wUIQvfap4Ni" outputId="47f298a4-c886-452b-b7a9-9785f1a41b74"
j = "Python"
j.upper()
# + [markdown] id="ZOLHPyqLqKfb"
# Utlizando *swapcase()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="SZQtvoNQqOdi" outputId="2f87478b-1b4f-4f06-f89a-f2d6ca32f33b"
k ="Python"
k.swapcase()
# + [markdown] id="x_LjjTsjqgFc"
# Utilizando *title()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="L5-UczBFqa_-" outputId="a9ec06e1-d86a-4581-a9b8-27f75983a3d3"
l = "apostila de python"
l.title()
# + [markdown] id="yr4EFPxtqsNG"
# Utilizando *split()*
# + colab={"base_uri": "https://localhost:8080/"} id="FKojoQBKqu23" outputId="21b3ca03-80eb-4c4c-e3bd-eb87b2a0ae39"
m = "cana de açucar"
m.split()
# + [markdown] id="xak5FtKhq8Fb"
# Utilizando *replace(S1, S2)*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="UYRI73NIq_Ji" outputId="8434c0f2-9b66-424b-94de-9f99d46b877e"
n = "Apostila teste"
n.replace("teste","Python")
# + [markdown] id="PzefHEGBzNvc"
# Utilizando *find()*
# + colab={"base_uri": "https://localhost:8080/"} id="2qcfd7jLzQle" outputId="0a7d8cdd-c969-465e-808c-56f8f3056894"
o = "Python"
o.find("h")
# + [markdown] id="gNfm1HOr0zKz"
# Utilizando *ljust()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="BKLcU0E-0ovt" outputId="2116bd7a-8b39-4e63-ca0a-f58c1a70b2af"
p = "Python"
p.ljust(15)
# + [markdown] id="1BKX5q8A1JNp"
# Utilizando *center()*
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="H-PRROC61C9A" outputId="1cd2a187-3ee6-4fd3-c0db-42c955929be0"
r = "Python"
r.center(10)
# + [markdown] id="eGS4DaDQ118v"
# Utilizando strip()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="nDeQS74s13n2" outputId="01cdc4c1-f1ed-46af-8d97-54e652a8606a"
u = " Python "
u.strip()
# + [markdown] id="8_LI2wJB3MAO"
# **FATIAMENTO DE STRINGS**
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="l85vSNgL3VPe" outputId="c5091dab-2eef-447b-ea66-992101fea305"
p = "Python"
p[1:4]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="NNxwdlmz3qjt" outputId="062635ac-e5c2-4e2c-c42c-faf373a0982c"
p = "Python"
p[2]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="hBwtkUki3tmw" outputId="d4679bc2-de59-4837-b5a9-1973a0e220e5"
p = "Python"
p[:4]
# + [markdown] id="XvYj73ck32ui"
# # **Exercício 1**
#
# Considere a string A = "Os limites só existem se você os deixar existir.(goku)".
#
# Que fatia corresponde a (goku)?
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="w9lyUUj04clK" outputId="e6ae25fd-4c3b-43cd-9234-cdaf59ad8521"
A = "Os limites só existem se você os deixar existir.(goku)"
print(len(A))
A[48:54]
# + [markdown] id="u5xxsKKR6S76"
# # **Exercício 2**
#
# Escreva um programa que solicite uma frase ao usuário e escreva a frase toda em maiúscula e sem espaços em branco.
# + colab={"base_uri": "https://localhost:8080/"} id="uvtVOOLy6qV8" outputId="61a44c5a-3ae4-4cf4-8fd0-95d08a6d06e8"
frase = input("Digite uma frase: ")
frase_sem_espaços = frase.replace(' ','')
frase_maiuscula = frase_sem_espaços.upper()
print(frase_maiuscula)
| Imagens/AULA03_Introducao_Python_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Regresión
#
# Revisaremos los conceptos de regresión vistos en el teórico.
#
# Haremos pruebas con datos de entrada de **una dimensión**.
import numpy as np
import matplotlib.pyplot as plt
import numpy as np
np.set_printoptions(suppress=True) # no usar notacion "e"
# ## Función Verdadera Oculta
#
# Usaremos como función oculta un sinusoide.
def create_sinusoidal_data(spread=0.25, data_size=50):
np.random.seed(0)
x = np.linspace(0, 1, data_size)
y = np.sin(2 * np.pi * x) + np.random.normal(scale=spread, size=x.shape)
return x, y
x, f_x = create_sinusoidal_data(0, 100)
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
# ## Muestra Ruidosa
#
# Tomaremos puntos uniformes en $x$, ruidosos en $y$.
# +
data_size = 20
X, y = create_sinusoidal_data(0.10, data_size)
plt.scatter(X, y, color="blue", label="datos")
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
# -
X
y
# ## División en Entrenamiento y Evaluación
#
# Dividiremos aleatoriamente los datos en una parte para entrenamiento y otra para evaluación.
#
# Usaremos
# [train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) de scikit-learn:
# +
from sklearn.model_selection import train_test_split
train_size = 5
test_size = data_size - train_size
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=train_size, random_state=0)
# -
X_train.shape, X_test.shape
plt.scatter(X_train, y_train, color="blue", label="train")
plt.scatter(X_test, y_test, color="white", edgecolor="k", label="test")
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
# ## Regresión Lineal
#
# Probaremos ajustar los puntos usando una recta.
#
# Vamos a programar a mano el aprendizaje y la predicción.
#
# ### Solución de Cuadrados Mínimos
#
# Datos de entrenamiento:
# - $X \in R^{N \times K}:$ $N$ vectores de entrada, de $K$ dimensiones cada uno.
# - $y \in R^N:$ $N$ valores de salida.
#
# Aprendizaje:
#
# $$w^* = (X^\top X)^{-1} X^\top y$$
# Predicción:
#
# $$f_{w^*}(x) = x^\top w^* = \sum_{k=1}^K x_k w^*_k$$
#
# Con $K=1$ tendríamos:
#
# $$f_{w^*}(x) = x_1 w_1^*$$
#
# Para que sea una recta nos falta un $w_0$ ("bias").
# Esto se puede resolver haciendo $K=2$ y agregando un valor constante 1 a cada dato:
#
# $$f_{w^*}((1, x)) = w_0 + x_1 w_1^*$$
#
X_train
X_train.shape
X_train_bias = np.stack((np.ones(X_train.shape[0]), X_train), axis=1) # add bias
X_train_bias
X_train_bias.shape
# +
def linear_least_squares(X, y):
X_b = np.stack((X, np.ones(X.shape[0])), axis=1) # add bias
return np.linalg.pinv(X_b.T.dot(X_b)).dot(X_b.T.dot(y))
def f(X, w):
X_b = np.stack((X, np.ones(X.shape[0])), axis=1) # add bias: (1, x)
return X_b.dot(w)
# -
# ### Entrenar
X_train, y_train
w = linear_least_squares(X_train, y_train)
w # qué dimensiones tiene w?
# ### Graficar
#
# Graficaremos la función aprendida a partir de los datos de entrenamiento. También graficaremos con los datos de evaluación y la función oculta, para comparar.
plt.scatter(X_train, y_train, color="blue", label="train")
plt.scatter(X_test, y_test, color="white", edgecolor="k", label="test")
plt.plot(x, f(x, w), color="red", label="model")
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
# ### Predecir y Evaluar: Error Cuadrático Medio
#
# Obtendremos los valores predichos para los datos de entrenamiento y de evaluación.
# Calcularemos el error cuadrático medio sobre ambos conjuntos de datos.
y_train_pred = f(X_train, w)
y_test_pred = f(X_test, w)
# Usamos la función [mean_squared_error](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.mean_squared_error.html) de scikit-learn:
# +
from sklearn.metrics import mean_squared_error
train_error = mean_squared_error(y_train, y_train_pred)
test_error = mean_squared_error(y_test, y_test_pred)
print(f'Train error: {train_error:f}')
print(f'Test error: {test_error:f}')
# -
# ## Regresión Polinomial
#
# Ahora haremos regresión polinomial. En este caso usaremos scikit-learn para definir el modelo, entrenar y predecir.
#
# En scikit-learn cada dato de entrada debe ser un vector, no un número. Debemos convertir cada dato en un vector de una dimensión:
X_train.shape, X_test.shape
X_train = X_train.reshape(-1, 1)
X_test = X_test.reshape(-1, 1)
X_train.shape, X_test.shape
# ### Features Polinomiales
#
# En scikit-learn, la regresión polinomial se implementa como un modelo de dos pasos.
#
# El primer paso genera vectores de características polinomiales, y el segundo paso aplica una regresión lineal sobre estos vectores (ver [Polynomial interpolation](https://scikit-learn.org/stable/auto_examples/linear_model/plot_polynomial_interpolation.html)).
#
# Por ejemplo, para generar características polinomiales de grado 5:
# +
from sklearn.preprocessing import PolynomialFeatures
pf = PolynomialFeatures(5) # polinomio de grado 5
pf.fit(X_train) # no necesita y_train!
# -
X_train.shape
X_train2 = pf.transform(X_train)
X_train2.shape # qué forma tiene esto?
X_train[0]
X_train2[0]
# ### Instanciar y Entrenar
#
# Crearemos y entrenaremos un modelo de grado 2.
#
# Como siempre en scikit-learn, para entrenar usamos la función **fit**.
# +
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
degree = 2
pf = PolynomialFeatures(degree)
lr = LinearRegression(fit_intercept=False) # el bias ya esta como feature
model = make_pipeline(pf, lr)
# -
model.fit(X_train, y_train)
# equivalente a:
Z_train = pf.fit(X_train, y_train)
lr.fit(Z_train, y_train)
# ### Inspeccionar Parámetros
#
# Ver [LinearRegression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html).
lr.coef_
lr.intercept_
# ### Graficar Resultado
plt.scatter(X_train, y_train, color="blue", label="train")
plt.scatter(X_test, y_test, color="white", edgecolor="k", label="test")
plt.plot(x, model.predict(x.reshape(-1, 1)), color="red", label="model")
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
# ### Predecir y Evaluar
#
# Para predecir, usamos la función **predict**:
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
train_error = mean_squared_error(y_train, y_train_pred)
test_error = mean_squared_error(y_test, y_test_pred)
print(f'Train error: {train_error:0.2}')
print(f'Test error: {test_error:0.2}')
# ## Sobreajuste vs. Generalización
#
# Probaremos polinomios de varios grados, obteniendo valores de error en entrenamiento y evaluación:
train_errors = []
test_errors = []
degrees = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
for degree in degrees:
# train:
pf = PolynomialFeatures(degree)
lr = LinearRegression(fit_intercept=False)
model = make_pipeline(pf, lr)
model.fit(X_train, y_train)
# predict:
y_train_pred = model.predict(X_train)
y_test_pred = model.predict(X_test)
# evaluate:
train_error = mean_squared_error(y_train, y_train_pred)
test_error = mean_squared_error(y_test, y_test_pred)
train_errors.append(train_error)
test_errors.append(test_error)
train_errors, test_errors
# Graficaremos las curvas de error en términos del grado del polinomio.
plt.plot(degrees, train_errors, color="blue", label="train")
plt.plot(degrees, test_errors, color="red", label="test")
plt.legend()
plt.xlabel("degree")
plt.ylabel("error")
plt.show()
# Se puede ver que el error en entrenamiento siempre baja, pero que en algún punto comienza el sobreajuste, ya que el error en evaluación empieza a subir.
# ### Mejor Modelo
#
# De acuerdo a la gráfica anterior, y como era de esperarse, el modelo que mejor ajusta los datos es el de grado 3:
degree = 3
model = make_pipeline(PolynomialFeatures(degree), LinearRegression())
model.fit(X_train, y_train);
# Graficamos:
plt.scatter(X_train, y_train, color="blue", label="train")
plt.scatter(X_test, y_test, color="white", edgecolor="k", label="test")
plt.plot(x, model.predict(x.reshape(-1, 1)), color="red", label="model")
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
train_errors[3], test_errors[3]
# ### Modelo Sobreajustado
#
# Veamos cómo es la gráfica de uno de los modelos que sufre de sobreajuste:
degree = 8
model = make_pipeline(PolynomialFeatures(degree), LinearRegression())
model.fit(X_train, y_train);
plt.scatter(X_train, y_train, color="blue", label="train")
plt.scatter(X_test, y_test, color="white", edgecolor="k", label="test")
plt.plot(x, model.predict(x.reshape(-1, 1)), color="red", label="model")
plt.plot(x, f_x, color="green", label="$\sin(2\pi x)$")
plt.legend()
plt.show()
# ## Ejercicios
#
# 1. Probar agregando puntos a los datos de entrenamiento para prevenir el sobreajuste en polinomios de grado alto.
# 2. Imprimir los parámetros de los modelos para cada grado.
# 3. Probar usando regularización para prevenir el sobreajuste en polinomios de grado alto
# (ver [Ridge](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.Ridge.html)).
# 4. Hicimos todo con datos de entrada de una dimensión. ¿Cómo serían los features polinomiales en datos de 2 o más dimensiones?
# ## Referencias
#
# Scikit-learn:
#
# - [train_test_split](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html)
# - [mean_squared_error](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.mean_squared_error.html)
# - [PolynomialFeatures](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PolynomialFeatures.html)
# - [LinearRegression](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html)
#
| 01 Regresion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 작업할 데이터 받기
df_ols = pd.read_csv('df_OLS.csv', encoding='utf-8-sig', index_col=0)
df_ols
# # position 데이터 정리 - 반올림
df_ols.position.unique()
df_ols.position = df_ols.position.round()
df_ols.position.unique()
# # Follower 제거
df_ols = df_ols.drop('follower', axis=1)
df_ols
# # 아웃라이어 확인
# +
from statsmodels.graphics import utils
dfX0 = df_ols.drop('value', axis=1)
dfX = sm.add_constant(dfX0)
dfy = df_ols['value']
model_p = sm.OLS(dfy, dfX)
result_p = model_p.fit()
pred = result_p.predict(dfX)
influence_p = result_p.get_influence()
cooks_d2, pvals = influence_p.cooks_distance
K = influence_p.k_vars
fox_cr = 4 / (len(dfy) - K - 1)
idx = np.where(cooks_d2 > fox_cr)[0]
ax = plt.subplot()
plt.scatter(dfy, pred)
plt.scatter(dfy[idx], pred[idx], s=300, c="r", alpha=0.5)
utils.annotate_axes(range(len(idx)), idx, list(zip(dfy[idx], pred[idx])), [(-20, 15)] * len(idx), size="small", ax=ax)
plt.title("아웃라이어")
plt.show()
print(idx)
# -
# # 아웃라이어 제거
idx2 = list(set(range(len(dfX))).difference(idx))
df_out = df_ols.iloc[idx2].reset_index(drop=True)
# # 아웃라이어 먼저 제거 후, OLS
# +
from sklearn.model_selection import train_test_split
dfX = df_out.drop(['value'], axis=1)
dfy = df_out['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean()) ** 2).sum()
rsquared = 1 - rss / tss
rsquared
# -
# # VIF Factor 확인
# +
from statsmodels.stats.outliers_influence import variance_inflation_factor
vif = pd.DataFrame()
vif["VIF Factor"] = [variance_inflation_factor(dfX.values, i) for i in range(dfX.shape[1])]
vif["features"] = dfX.columns
vif.sort_values(by='VIF Factor', ascending=False)
# -
# # OLS - 1
df_out_1 = df_out.drop(['duelsWonTotal'], axis=1)
len(df_out.columns), len(df_out_1.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_1
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # OLS - 2
df_out_2 = df_out_1.drop(['cards_red'], axis=1)
len(df_out_2.columns), len(df_out_1.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_2
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # OLS - 3
df_out_3 = df_out_2.drop(['penalty_missed'], axis=1)
len(df_out_3.columns), len(df_out_2.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_3
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # OLS - 4
df_out_4 = df_out_3.drop(['cards_yellowred'], axis=1)
len(df_out_4.columns), len(df_out_3.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_4
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # OLS - 5
df_out_5 = df_out_4.drop(['rating'], axis=1)
len(df_out_5.columns), len(df_out_4.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_5
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # OLS - 6
df_out_6 = df_out_5.drop(['position'], axis=1)
len(df_out_6.columns), len(df_out_5.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_6
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # OLS - 7
df_out_7 = df_out_6.drop(['cards_yellow'], axis=1)
len(df_out_7.columns), len(df_out_6.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_7
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 8
# -
df_out_8 = df_out_7.drop(['goalsAssist_passesKey'], axis=1)
len(df_out_8.columns), len(df_out_7.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_8
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 9
# -
df_out_9 = df_out_8.drop(['tackles_total'], axis=1)
len(df_out_9.columns), len(df_out_8.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_9
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 10
# -
df_out_10 = df_out_9.drop(['penalty_commited'], axis=1)
len(df_out_10.columns), len(df_out_9.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_10
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 11
# -
df_out_11 = df_out_10.drop(['penalty_won'], axis=1)
len(df_out_11.columns), len(df_out_10.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_11
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 12
# -
df_out_12 = df_out_11.drop(['fouls_drawn'], axis=1)
len(df_out_12.columns), len(df_out_11.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_12
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 13
# -
df_out_13 = df_out_12.drop(['hw'], axis=1)
len(df_out_13.columns), len(df_out_12.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_13
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 14
# -
df_out_14 = df_out_13.drop(['tackles_interceptions'], axis=1)
len(df_out_14.columns), len(df_out_13.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_14
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 15
# -
df_out_15 = df_out_14.drop(['penalty_success'], axis=1)
len(df_out_15.columns), len(df_out_14.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_15
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 16
# -
df_out_16 = df_out_15.drop(['tackles_blocks'], axis=1)
len(df_out_16.columns), len(df_out_15.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_16
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# +
# OLS - 17
# -
df_out_17 = df_out_16.drop(['games_lineups'], axis=1)
len(df_out_17.columns), len(df_out_16.columns)
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_17
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# +
from sklearn.model_selection import train_test_split
df_ols = df_out_17
dfX = df_ols.drop(['value'], axis=1)
dfy = df_ols['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
#feature_names = list(dfX.columns)
#feature_names = ["scale({})".format(name) for name in feature_names]
formula = "value ~ scale(age) + \
scale(passes_total) + \
scale(passes_accuracy) + \
scale(fouls_committed) + \
scale(games_played) + \
scale(shotsOnTotal_goalsTotal) + \
scale(goasConceded_penaltySaved) + \
scale(dribblesAtmptsSuc) + scale(gamesApperance_sub)"
model = sm.OLS.from_formula(formula, data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula(formula, data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # 검증 1차
# +
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_out_20)):
df_train = df_out_20.iloc[idx_train]
df_test = df_out_20.iloc[idx_test]
model = sm.OLS.from_formula("value ~ scale(age) + scale(follower) + scale(passes_accuracy) + \
scale(games_played) + scale(shotsOnTotal_goalsTotal)", data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
# -
# # 검증 2차
# +
from sklearn.base import BaseEstimator, RegressorMixin
import statsmodels.formula.api as smf
import statsmodels.api as sm
class StatsmodelsOLS(BaseEstimator, RegressorMixin):
def __init__(self, formula):
self.formula = formula
self.model = None
self.data = None
self.result = None
def fit(self, dfX, dfy):
self.data = pd.concat([dfX, dfy], axis=1)
self.model = smf.ols(self.formula, data=self.data)
self.result = self.model.fit()
def predict(self, new_data):
return self.result.predict(new_data)
# +
from sklearn.model_selection import cross_val_score
model = StatsmodelsOLS("value ~ scale(age) + scale(follower) + scale(passes_accuracy) + \
scale(games_played) + scale(shotsOnTotal_goalsTotal)")
cv = KFold(10, shuffle=True, random_state=0)
cross_val_score(model, dfX, dfy, scoring="r2", cv=cv).mean()
# -
# +
from sklearn.model_selection import train_test_split
dfX = df_out_20.drop(['value'], axis=1)
dfy = df_out_20['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
formula = "value ~ " + "+".join(feature_names)
model = sm.OLS.from_formula(formula, data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula(formula, data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
# # 아웃라이어 2차 제거
# +
from statsmodels.graphics import utils
dfX0 = df_out_20.drop('value', axis=1)
dfX = sm.add_constant(dfX0)
dfy = df_out_20['value']
model_p = sm.OLS(dfy, dfX)
result_p = model_p.fit()
pred = result_p.predict(dfX)
influence_p = result_p.get_influence()
cooks_d2, pvals = influence_p.cooks_distance
K = influence_p.k_vars
fox_cr = 4 / (len(dfy) - K - 1)
idx = np.where(cooks_d2 > fox_cr)[0]
ax = plt.subplot()
plt.scatter(dfy, pred)
plt.scatter(dfy[idx], pred[idx], s=300, c="r", alpha=0.5)
utils.annotate_axes(range(len(idx)), idx, list(zip(dfy[idx], pred[idx])), [(-20, 15)] * len(idx), size="small", ax=ax)
plt.title("아웃라이어")
plt.show()
print(idx)
# -
idx2 = list(set(range(len(dfX))).difference(idx))
len(idx2)
df_out2 = df_out_20.iloc[idx2].reset_index(drop=True)
df_out2
# +
from sklearn.model_selection import train_test_split
dfX = df_out2.drop(['value'], axis=1)
dfy = df_out2['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
model = sm.OLS.from_formula("value ~ " + "+".join(feature_names), data=df_train)
result = model.fit()
print(result.summary())
# +
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean()) ** 2).sum()
rsquared = 1 - rss / tss
rsquared
# -
# # 검증 1차
# +
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_out2)):
df_train = df_out2.iloc[idx_train]
df_test = df_out2.iloc[idx_test]
model = sm.OLS.from_formula("value ~ scale(age) + scale(follower) + scale(passes_accuracy) + \
scale(games_played) + scale(shotsOnTotal_goalsTotal)", data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
# -
# # 검증 2차
# +
from sklearn.base import BaseEstimator, RegressorMixin
import statsmodels.formula.api as smf
import statsmodels.api as sm
class StatsmodelsOLS(BaseEstimator, RegressorMixin):
def __init__(self, formula):
self.formula = formula
self.model = None
self.data = None
self.result = None
def fit(self, dfX, dfy):
self.data = pd.concat([dfX, dfy], axis=1)
self.model = smf.ols(self.formula, data=self.data)
self.result = self.model.fit()
def predict(self, new_data):
return self.result.predict(new_data)
# +
from sklearn.model_selection import cross_val_score
model = StatsmodelsOLS("value ~ scale(age) + scale(follower) + scale(passes_accuracy) + \
scale(games_played) + scale(shotsOnTotal_goalsTotal)")
cv = KFold(10, shuffle=True, random_state=0)
cross_val_score(model, dfX, dfy, scoring="r2", cv=cv).mean()
# -
# # Data 가중 - 20200622
# +
from sklearn.model_selection import train_test_split
dfX = df_out_15.drop(['value'], axis=1)
dfy = df_out_15['value']
df = pd.concat([dfX, dfy], axis=1)
df_train, df_test = train_test_split(df, test_size=0.3, random_state=0)
feature_names = list(dfX.columns)
feature_names = ["scale({})".format(name) for name in feature_names]
formula = "value ~ scale(age) + scale(I(age**2)) + scale(I(age**3)) + \
scale(follower) + scale(I(follower**2)) +\
scale(passes_total) + scale(I(passes_total**2)) +\
scale(passes_accuracy) + \
scale(fouls_drawn) + \
scale(games_lineups) + \
scale(games_played) + \
scale(shotsOnTotal_goalsTotal) + \
scale(dribblesAtmptsSuc) + \
scale(gamesApperance_sub)"
model = sm.OLS.from_formula(formula, data=df_train)
result = model.fit()
print(result.summary())
##############################################################################
from sklearn.model_selection import KFold
scores = np.zeros(10)
cv = KFold(10, shuffle=True, random_state=0)
for i, (idx_train, idx_test) in enumerate(cv.split(df_ols)):
df_train = df_ols.iloc[idx_train]
df_test = df_ols.iloc[idx_test]
model = sm.OLS.from_formula(formula, data=df_train)
result = model.fit()
pred = result.predict(df_test)
rss = ((df_test.value - pred) ** 2).sum()
tss = ((df_test.value - df_test.value.mean())** 2).sum()
rsquared = 1 - rss / tss
scores[i] = rsquared
# print("학습 R2 = {:.8f}, 검증 R2 = {:.8f}".format(result.rsquared, rsquared))
print("모델 성능 : {}".format(scores.mean()))
# -
| Analysis/4.8_DataAnalysis_AllData-Follower_removeOutlierFirst_20200624.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Virtual Agent for Frequently Asked Questions
# Building Virtual Agent that understands the semantics of user utterances has become simple with transformers based models out there and with the support of large collection of open-source libraries
# <b>Note:</b> Update variables under <b>Variables</b> section if required before running the notebook. To run notebook cell by cell, click on a cell and click <b>Run</b> button below the <b>Menu</b> bar. Or to run all cells, select <b>Cell --> Run All<b> from Menu bar.
# ##### Variables
# The variable <b>DATA_SOURCE_PATH</b> should be set to the path of <b>input</b> data to be used for training.
# MODEL_PATH variable refers to the location to store embeddings of input text.
# The dataset is expected to contain questions under column <b>Q</b> and answers under column <b>A</b>. Please reference <b>faqs.csv</b> from downloaded repository that can be found in the kit_installer file location.
# The execution of <b>last cell</b> helps to try Virtual Agent built.
DATA_SOURCE_PATH = r"faqs.csv"
# Default model location
MODEL_PATH = r"models/model_va.pickle"
# ###### Import libraries for data analysis
import numpy as np
import pandas as pd
# ###### Import libraries for text mining
from lingualytics.preprocessing import remove_lessthan, remove_punctuation, remove_stopwords
from lingualytics.stopwords import en_stopwords
from texthero.preprocessing import remove_digits
# ###### Import libraries for transformers
from sentence_transformers import SentenceTransformer
# ###### Import libraries for computing similarities
from torch.nn import CosineSimilarity
import torch
# ###### Import library for storing into binary file
import pickle
# ###### Load data source into dataframe
df = pd.read_csv(DATA_SOURCE_PATH, encoding_errors="ignore")
pd.set_option('display.max_colwidth', None)
df
# ###### Cleanse data by removing numbers and punctutation
# This process is part of pre-processing that aids in getting rid of unnecessary text, which would otherwise hinder the learning process of the model. Techniques like stemming, lemmatisation can also help here.
#
# As we're using sentence embedding, we wouldn't be doing extensive pre-processing here. The pre-processing complexity decreases with increase in the quality of the dataset
df['procd_Q'] = df['Q'].pipe(remove_digits).pipe(remove_punctuation)#.pipe(remove_lessthan,length=3)\
#.pipe(remove_stopwords,stopwords=en_stopwords.union(hi_stopwords))
df
# ###### Load sentence transformer model of your choice for getting sentence embeddings
# The model can be chosen by considering various aspects and comparing available models from this link.
# https://www.sbert.net/docs/pretrained_models.html
model = SentenceTransformer('paraphrase-MiniLM-L6-v2')
# ###### Find embeddings of sentences and store in a binary file
#
# The binary file storage helps to load and use embeddings later without having the need to computing them again. We use pickle here to store in binary files. You may also use joblib
q_embs = model.encode(df["procd_Q"]) # computes encode for all the questions from the dataset.
#Embeddings can be computed in batches for massive dataset.
with open(MODEL_PATH, "wb") as file:
pickle.dump(q_embs, file)
# ###### Load embeddings from binary file into memory
with open(MODEL_PATH, "rb") as file:
q_embs = pickle.load(file)
# ###### Predict answer to user query
# The user query is cleansed and pre-processed as earlier, and then a matching query from data source is predicted. The predicted query is used to look up to find corresponding answer
def pred_answer(usr_query):
df_query = pd.DataFrame([usr_query], columns=["usr_query"]) # use similar pipeline that was used for computing embeddings from dataset
df_query["clean_usr_q"] = df_query["usr_query"].pipe(remove_digits).pipe(remove_punctuation)
usr_q_emb = model.encode(df_query["clean_usr_q"]) # compute embedding
cosine_similarity = CosineSimilarity()
q_idx = np.argmax(cosine_similarity(torch.from_numpy(usr_q_emb), torch.from_numpy(q_embs))) # compute cosine similarity and find the matched query
return df["A"][q_idx.item()] # look up answer of the matched query from the dataframe of input dataset
usr_query = "tell me about kandi"
pred_answer(usr_query)
# ###### Simulating Virtual Agent
while True:
usr_q = input("Ask a query(or type 'exit' to exit):")
if usr_q == "exit":
break
else:
print("Answer: ", pred_answer(usr_q))
print("-----------------")
| Virtual Agent for FAQ.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (nlp_new)
# language: python
# name: nlp_new
# ---
# ## Source
# 1. https://jovian.ml/aakanksha-ns/shelter-outcome
# 2. https://github.com/fastai/fastai/blob/master/courses/dl1/lesson3-rossman.ipynb
import pandas as pd
import numpy as np
from collections import Counter
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
import torch
from torch.utils.data import Dataset, DataLoader
import torch.optim as torch_optim
import torch.nn as nn
import torch.nn.functional as F
from torchvision import models
from datetime import datetime
| notebooks/8.nba_entity_embeddings__test_fastai.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="npFemIWusYQi" colab_type="code" outputId="4f767764-1062-4649-c072-f5a58941da16" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 73}
import numpy as np
from google.colab import files
import pandas as pd
uploaded = files.upload()
# + id="XdRWg0lfTTa4" colab_type="code" outputId="7128fb7b-4f10-40fd-d177-51ed5a249deb" colab={"base_uri": "https://localhost:8080/", "height": 177}
import io
data = pd.read_csv(io.BytesIO(uploaded['IRIS.csv']))
# + id="c3wNhzzoiB2o" colab_type="code" outputId="b74d0085-2f54-4cee-813a-8af66c809f4f" colab={"base_uri": "https://localhost:8080/", "height": 33}
from keras.utils import np_utils
# + id="AFiRnfvgT1oh" colab_type="code" outputId="5f746465-9232-4ffa-d915-d86d23b5a43b" colab={"base_uri": "https://localhost:8080/", "height": 227}
y = data.drop('variety', axis=1)
x = data.variety.astype('category').cat.codes
y = y.rename(columns=y.iloc[0]).drop(y.index[0])
y = y.to_numpy()
x = x.to_numpy()
x = np.delete(x, 0)
y = y.reshape(149, 1, 4)
x = x.reshape(149, 1, 1)
x = np_utils.to_categorical(x)
# + id="2KSVvSqrXX_s" colab_type="code" colab={}
# + id="oTrTMpTwtLXd" colab_type="code" colab={}
class FCLayer:
def __init__(self, input_size, output_size):
self.input_size = input_size
self.output_size = output_size
self.weights = np.random.randn(input_size, output_size) / np.sqrt(input_size + output_size)
self.bias = np.random.randn(1, output_size) / np.sqrt(input_size + output_size)
def forward(self, input):
self.input = input
return np.dot(input, self.weights) + self.bias
def backward(self, output_error, learning_rate):
input_error = np.dot(output_error, self.weights.T)
weights_error = np.dot(self.input.T, output_error)
# bias_error = output_error
self.weights -= learning_rate * weights_error
self.bias -= learning_rate * output_error
return input_error
# + id="E6nSYAB2sam3" colab_type="code" colab={}
class ActivationLayer:
def __init__(self, activation, activation_prime):
self.activation = activation
self.activation_prime = activation_prime
def forward(self, input):
self.input = input
return self.activation(input)
def backward(self, output_error, learning_rate):
return output_error * self.activation_prime(self.input)
# + id="hl8LxP1lAEiN" colab_type="code" colab={}
# + id="RQeuIfkK3vyl" colab_type="code" colab={}
# bonus
class SoftmaxLayer:
def __init__(self, input_size):
self.input_size = input_size
def forward(self, input):
self.input = input
tmp = np.exp(input)
self.output = tmp / np.sum(tmp)
return self.output
def backward(self, output_error, learning_rate):
input_error = np.zeros(output_error.shape)
out = np.tile(self.output.T, self.input_size)
return self.output * np.dot(output_error, np.identity(self.input_size) - out)
# + id="LuPbn70Wt8Q7" colab_type="code" colab={}
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def sigmoid_prime(x):
return np.exp(-x) / (1 + np.exp(-x))**2
def tanh(x):
return np.tanh(x)
def tanh_prime(x):
return 1 - np.tanh(x)**2
def relu(x):
return np.maximum(x, 0)
def relu_prime(x):
return np.array(x >= 0).astype('int')
# + id="rXY7jkUzuqEk" colab_type="code" colab={}
def mse(y_true, y_pred):
return np.mean(np.power(y_true - y_pred, 2))
def mse_prime(y_true, y_pred):
return 2 * (y_pred - y_true) / y_pred.size
def sse(y_true, y_pred):
return 0.5 * np.sum(np.power(y_true - y_pred, 2))
def sse_prime(y_true, y_pred):
return y_pred - y_true
# + id="-whGNp8Joaur" colab_type="code" outputId="b2974514-a85f-4729-f9ed-e2c6e05a8289" colab={"base_uri": "https://localhost:8080/", "height": 33}
# + id="oHQpwN8LpKiN" colab_type="code" outputId="a2848a74-9d8d-4e7c-8323-c176e81508f2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
network = [
FCLayer(4, 16),
ActivationLayer(sigmoid, sigmoid_prime),
FCLayer(16, 3),
ActivationLayer(sigmoid, sigmoid_prime)
]
epochs = 1000
learning_rate = 0.01
# training
for epoch in range(epochs):
error = 0
for xi, y_true in zip(y, x):
# forward
output = xi
for layer in network:
output = layer.forward(output)
# error (display purpose only)
error += mse(y_true, output)
# backward
output_error = mse_prime(y_true, output)
for layer in reversed(network):
output_error = layer.backward(output_error, learning_rate)
error /= len(y)
print('%d/%d, error=%f' % (epoch + 1, epochs, error))
# + id="mrMLx3eGv3jk" colab_type="code" colab={}
def predict(network, input):
output = input
for layer in network:
output = layer.forward(output)
return output
# + id="5zBBU8ANbd1o" colab_type="code" outputId="e89b4687-78d7-44f1-aa45-9383e718bf45" colab={"base_uri": "https://localhost:8080/", "height": 33}
predict(network, [[4.8,3,1.4,0.1]])
# + id="6fhIV3dibsws" colab_type="code" colab={}
| AI_4_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
import tensorflow as tf
import simplejson
import matplotlib.pyplot as plt
# %matplotlib inline
import threading
import tensorflow.contrib.slim as slim
from utils import data_utils, train_utils
import datetime
import os
import time
import numpy as np
import matplotlib.pyplot as plt
import matplotlib
import cv2
import train
import pandas as pd
from shapely import wkt
import time
import sys
from inference import pred_for_each_quarter, test_input, stitch_mask
# + deletable=true editable=true
hypes = './hypes/hypes.json'
with open(hypes, 'r') as f:
H = simplejson.load(f)
H['batch_size'] = 1
H['pad'] = 100
H['x_width'] = 1920
H['x_height'] = 1920
H['print_iter'] = 100
H['save_iter'] = 500
H['crop_size'] = [1700, 1700]
print_iter = H['print_iter']
num_channel = H['num_channel']
x_width = H['x_width']
x_height = H['x_height']
batch_size = H['batch_size']
class_type = H['class_type']
pad = H['pad']
class_type = H['class_type']
log_dir = H['log_dir']
save_iter = H['save_iter']
# + deletable=true editable=true
img_in = tf.placeholder(dtype=tf.float32, shape=[batch_size, x_width, x_height, 16])
logits, pred = train.build_pred(img_in, H, 'test')
# + deletable=true editable=true
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
saver = tf.train.Saver()
sess = tf.Session(config = config)
saver.restore(sess, save_path='log_dir/ckpt_new/ckpt-12000')
# + deletable=true editable=true
ids_with_instance = train_utils.generate_train_ids(class_type)
print 'IDs of training data with instance of class {} ({}): {}'.format(
class_type, data_utils.CLASSES[class_type + 1], ids_with_instance)
# -
jaccard_indices = {}
# + [markdown] deletable=true editable=true
# # Compare training data with predictions
# + deletable=true editable=true
for img_id in ids_with_instance:
img_data = data_utils.ImageData(img_id)
img_data.load_image()
img_data.create_train_feature()
img_data.create_label()
mask_stack, shape_stack = pred_for_each_quarter(sess, img_in, pred, img_data, H)
mask = stitch_mask(mask_stack, img_data.image_size, shape_stack, H)
polygons = data_utils.mask_to_polygons(mask=mask, img_id=img_id, test=False, epsilon=1)
true_polygons = data_utils.get_polygon_list(
image_id=data_utils.train_IDs_dict[img_id], class_type=class_type + 1)
jaccard_indices[data_utils.train_IDs_dict[img_id]] = \
polygons.intersection(true_polygons).area / polygons.union(true_polygons).area
alpha = 0.4
fig, axs = plt.subplots(2, 2, figsize = [20, 20])
print 'Processing ImageId: {} (No. {}); Class ({}): {}'.format(
img_id, data_utils.train_IDs_dict[img_id], class_type, data_utils.CLASSES[class_type + 1])
fig.suptitle('Image (No. {}) Id {}; Class ({}): {}'.format(
img_id, data_utils.train_IDs_dict[img_id], class_type, data_utils.CLASSES[class_type + 1]),
fontsize = 16)
for i in range(2):
for j in range(2):
axs[0, 0].imshow(img_data.label[:, :, class_type], cmap = plt.cm.gray)
axs[0, 0].set_title('True label for image: {}, class: {}'.format(
data_utils.train_IDs_dict[img_id], data_utils.CLASSES[class_type + 1]))
axs[0, 1].imshow(data_utils.scale_percentile(img_data.three_band_image))
axs[0, 1].imshow(img_data.label[:, :, class_type], cmap = plt.cm.gray, alpha = alpha)
axs[0, 1].set_title('3-band image with true label for image: {}, class: {}'.format(
data_utils.train_IDs_dict[img_id], data_utils.CLASSES[class_type + 1]))
axs[1, 0].imshow(mask, cmap = plt.cm.gray)
axs[1, 0].set_title('Predicted label for image: {}, class: {}'.format(
data_utils.train_IDs_dict[img_id], data_utils.CLASSES[class_type + 1]))
axs[1, 1].imshow(data_utils.scale_percentile(img_data.three_band_image))
axs[1, 1].imshow(mask, cmap = plt.cm.gray, alpha = alpha)
axs[1, 1].set_title('3-band image with predicted label for image: {}, class: {}'.format(
data_utils.train_IDs_dict[img_id], data_utils.CLASSES[class_type + 1]))
fig.tight_layout()
fig.subplots_adjust(top=0.95)
print 'Jaccard indices {}'.format(jaccard_indices)
print 'Mean Jaccard index {}'.format(np.mean(jaccard_indices.values()))
# -
ids_w_o_instance = sorted(list(set(range(25)) - set(ids_with_instance)))
print 'IDs of training data w/o instance of class {} ({}): {}'.format(
class_type, data_utils.CLASSES[class_type + 1], ids_w_o_instance)
# +
for img_id in ids_w_o_instance:
print 'Processing ImageId (No. {}): {}; Class ({}): {}'.format(
img_id, data_utils.train_IDs_dict[img_id], class_type, data_utils.CLASSES[class_type + 1])
img_data = data_utils.ImageData(img_id)
img_data.load_image()
img_data.create_train_feature()
mask_stack, shape_stack = pred_for_each_quarter(sess, img_in, pred, img_data, H)
mask = stitch_mask(mask_stack, img_data.image_size, shape_stack, H)
polygons = data_utils.mask_to_polygons(mask=mask, img_id=img_id, test=False, epsilon=1)
true_polygons = data_utils.get_polygon_list(
image_id=data_utils.train_IDs_dict[img_id], class_type=class_type + 1)
jaccard_indices[data_utils.train_IDs_dict[img_id]] = \
polygons.intersection(true_polygons).area / polygons.union(true_polygons).area \
if polygons.union(true_polygons).area else 1.
print 'Jaccard indices {}'.format(jaccard_indices)
print 'Mean Jaccard index {}'.format(np.mean(jaccard_indices.values()))
# + [markdown] deletable=true editable=true
# # Predictions on the test data
# + deletable=true editable=true
for img_id in range(30, 35):
img_data = data_utils.ImageData(img_id, phase='test')
img_data.load_image()
img_data.create_train_feature()
mask_stack, shape_stack = pred_for_each_quarter(sess, img_in, pred, img_data, H)
mask = stitch_mask(mask_stack, img_data.image_size, shape_stack, H)
alpha = 0.4
fig, axs = plt.subplots(1, 2, figsize = [20, 10])
print 'Processing ImageId: {} (No. {}); Class ({}): {}'.format(
img_id, data_utils.test_IDs_dict[img_id], class_type, data_utils.CLASSES[class_type + 1])
fig.suptitle('Image (No. {}) Id {}; Class ({}): {}'.format(
img_id, data_utils.test_IDs_dict[img_id], class_type, data_utils.CLASSES[class_type + 1]),
fontsize = 16)
for i in range(2):
for j in range(2):
axs[0].imshow(mask, cmap = plt.cm.gray)
axs[0].set_title('Predicted label for image: {}, class: {}'.format(
data_utils.test_IDs_dict[img_id], data_utils.CLASSES[class_type + 1]))
axs[1].imshow(data_utils.scale_percentile(img_data.three_band_image))
axs[1].imshow(mask, cmap = plt.cm.gray, alpha = alpha)
axs[1].set_title('3-band image with predicted label for image: {}, class: {}'.format(
data_utils.test_IDs_dict[img_id], data_utils.CLASSES[class_type + 1]))
fig.tight_layout()
fig.subplots_adjust(top=0.95)
# + deletable=true editable=true
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Our functional calculator … so far
#
# This is the calculator that we implemented in the first section. But it suffers from a few drawbacks:
#
# - No input validation
# - No looping
# +
OPERATORS = '+', '-', '*', '/'
def f_get_number():
return int(input('Enter an integer: '))
def f_get_operator():
return input('Enter an operator (+, -, *, /): ')
def f_calculate(number1, operator, number2):
return number1+number2 if operator == '+' \
else number1-number2 if operator == '-' \
else number1/number2 if operator == '/' \
else number1*number2 if operator == '*' \
else None
def f_main():
return f_calculate(
f_get_number(),
f_get_operator(),
f_get_number(),
)
print('The result is: %s' % f_main())
# -
# ## Let's get to work!
#
# Our toolkit contains:
#
# - Lambda expressions
# - Decorators
# - Higher-order functions
# +
OPERATORS = '+', '-', '*', '/'
def maybe(fnc):
"""Turns Exceptions into return values."""
def inner(*args):
for a in args:
if isinstance(a, Exception):
return a
try:
return fnc(*args)
except Exception as e:
return e
return inner
def repeat(fnc, until):
"""Repeats a function until its return value meets
the stop criterion."""
def inner(*args):
while True:
result = fnc(*args)
if until(result):
return result
return inner
is_int = lambda i: isinstance(i, int)
get_number = lambda: int(input('Enter an integer: '))
safe_get_number = repeat(maybe(get_number), until=is_int)
is_operator = lambda o: o in OPERATORS
get_operator = lambda: input('Enter an operator (+, -, *, /): ')
safe_get_operator = repeat(get_operator, until=is_operator)
calculate = lambda number1, operator, number2: \
number1+number2 if operator == '+' \
else number1-number2 if operator == '-' \
else number1/number2 if operator == '/' \
else number1*number2 if operator == '*' \
else None
main = lambda: calculate(
safe_get_number(),
safe_get_operator(),
safe_get_number(),
)
forever = lambda retval: False
main_loop = repeat(lambda: print(main()), until=forever)
main_loop()
| 7181_05_code_ACC_SB/A fully functional, interactive calculator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# # Intro to Python - Exercises - Part 6
# ## 6. Strings
# Until now, most examples and exercises have been using numbers. In daily life, it is far more commonplace to deal with textual information. So are you ever going to learn how to deal with texts?
#
# The reason that dealing with texts was postponed until this point, is that dealing with numbers is simply easier than dealing with texts. But in the present section, the first steps are taken to learn to manipulate textual information.
#
# Texts, in programming languages, are dealt with in the form of strings. This section is on the details of strings, and on readily-available functions to juggle them.
# ### Multi-line strings
# Strings in Python may span across multiple lines. This can be useful when you have a very long string, or when you want to format the output of the string in a certain way. Multi-line strings can be achieved in two ways:
#
# 1. With single or double quotes, and an indication that the remainder of the string continues on the next line with a backslash.
# 2. With triple single or double quotes.
#
# I first demonstrate how this works when you use the regular string enclosure with one double or single quote at each end of the string:
longString = "I'm fed up with being treated like sheep. \
What's the point of going abroad if you're just another \
tourist carted around in buses surrounded by sweaty \
mindless oafs from Kettering and Coventry in their \
cloth caps and their cardigans and their transistor \
radios and their Sunday Mirrors, complaining about \
the tea - 'Oh they don't make it properly here, do they, \
not like at home' - and stopping at Majorcan bodegas \
selling fish and chips and Watney's Red Barrel and \
calamaris and two veg and sitting in their cotton frocks \
squirting <NAME>'s suncream all over their puffy \
raw swollen purulent flesh 'cos they 'overdid it on the first day."
print(longString)
# As you can see, Python now interprets this example as a single line of text. The backslash (`\`) can actually be included after any Python statement to indicate that it continues on the next line, and it can be quite useful for that, for instance when you write long calculations.
#
# The recommended way to write multi-line strings in Python is, however, to use triple double or single quotes. I indicated earlier that you can use those to write multi-line comments. Such comments are basically large strings in the middle of your Python program, which do nothing as they are not assigned to a variable.
#
# Here is an example of a long string with triple double quotes:
longString = """And being herded into endless Hotel Miramars and Bellevueses
and Continentales with their modern international luxury
roomettes and draught Red Barrel and swimming pools full
of fat German businessmen pretending they're acrobats forming
pyramids and frightening the children and barging into queues
and if you're not at your table spot on seven you miss the
bowl of Campbell's Cream of Mushroom soup, the first item on
the menu of International Cuisine, and every Thursday night
the hotel has a bloody cabaret in the bar, featuring a tiny
emaciated dago with nine-inch hips and some bloated fat tart
with her hair brylcreemed down and a big arse presenting
Flamenco for Foreigners."""
print(longString)
# The interesting difference between these two examples is that in the first example, the string was interpreted as one long, continuous series of characters, while in the second example the different lines are all printed on different lines on the output. The reason that this happens is that there is an invisible character included at the end of each line in the second example that indicates that Python should move to the next line before continuing. This is the so-called "newline" character, and you can actually insert it explicitly into a string, using the code "`\n`". So this code should not be read as two characters, the backslash and the "n", but as a single newline character. By using it, you can ensure that you print the output on multiple lines, even if you use the backslash to indicate the continuation of the string, as was done in the first example. For example:
longstring = "And then some adenoidal typists from Birmingham with flabby\n\
white legs and diarrhoea trying to pick up hairy bandy-legged\n\
wop waiters called Manuel and once a week there's an excursion\n\
to the local Roman Ruins to buy cherryade and melted ice cream\n\
and bleeding Watney's Red Barrel and one evening you visit the\n\
so called typical restaurant with local colour and atmosphere\n\
and you sit next to a party from Rhyl who keep singing\n\
'Torremolinos, torremolinos' and complaining about the food -\n\
'It's so greasy here, isn't it?' - and you get cornered by some\n\
drunken greengrocer from Luton with an Instamatic camera and\n\
Dr. Scholl sandals and last Tuesday's Daily Express and he\n\
drones on and on and on about how Mr. Smith should be running\n\
this country and how many languages Enoch Powell can speak and\n\
then he throws up over the Cuba Libres."
print(longstring)
# This means that if you do not want automatic newline characters inserted into a multi-line string, you have to use the first approach, with the backslash at the end of the line. If you are okay with newline characters in your multi-line string, the second approach is probably the easiest to read.
# ### Escape sequences
# "`\n`" is a so-called "escape sequence". An escape sequence is a string character written as a backslash followed by a code, which can be one or multiple characters. Python interprets escape sequences in a string as a special character; a control character.
# +
word1 = "orange"
word2 = "banana"
def add_newline_between_words(word1,word2):
new_line = word1 + "\n" + word2
return(new_line)
print(add_newline_between_words(word1,word2))
# -
# Besides the newline character there are more special characters "`\'`" and "`\"`", which can be used to place a single respectively double quote in a string, regardless of what characters surround the string. I also mentioned that you can use "`\\`" to insert a "real" backslash in a string.
#
# There are a few more "backslash sequences" which lead to a special character. Most of these are archaic and you do not need to worry about them. The one I want to mention are "`\t`" which represents a single tabulation (also known as the 'tab').
# +
d = "test"
m = "me"
def place_word_between_single_quotes(w1):
new_line = '\'' + word1 + "\'"
return(new_line)
print(place_word_between_single_quotes(m))
def place_word_between_double_quotes(w1):
new_line = '\t' + word1 + '"'
return(new_line)
print(place_word_between_double_quotes(d))
# -
# Extra information for students who want to know more, but not necessary for this course:
#
# There is another character "`\xnn`" whereby `nn` stands for two hexadecimal digits, which represents the character with hexadecimal number `nn`. For example, "`\x20`" is the character expressed by the hexadecimal number `20`, which is the same as the decimal number `32`, which is the space (this will be explained later in this chapter).
#
# In case you never learned about hexadecimal counting: hexadecimals use a numbering scheme that uses 16 different digits. We use ten (`0` to `9`), binary uses two (`0` to `1`), and hexidecimal then uses `0` to `9` and then continues from `A` to `F`. A direct translation from hexadecimals to decimals turns `A` into `10`, `B` into `11`, etcetera. In decimal counting, the value of a multi-digit number is found by multiplying the digits by increasing powers of `10`, from right to left, e.g., the number `1426` is `6 + 2*10 + 4*100 + 1*1000`. For hexadecimal numbers you do the same thing, but multiply by powers of `16`, e.g., the hexadecimal number `4AF2` is `2 + 15*16 + 10*256 + 4*4096`. Programmers tend to like hexadecimal numbers, as computers work with bytes as the smallest unit of memory storage, and a byte can store 256 different values, i.e., any byte value can be expressed by a hexadecimal number of two digits.
# ### Accessing characters of a string
# As I showed several times before, a string is a collection of characters in a specific order. You can access the individual characters of a string using indices.
# ### String indices
# Each symbol in a string has a position, this position can be referred to by the index number of the position. The index numbers start at 0 and then increase to the length of the string. The following table shows the word "orange" in the first row and the indices for each letter in the second and third rows:
#
# __` o r a n g e`__<br>
# ` 0 1 2 3 4 5`<br>
# ` -6 -5 -4 -3 -2 -1`
#
# As you can see, you can use positive indices, which start at the first letter of the string and increase until the end of the string is reached, or negative indices, which start with -1 for the last letter of the string and decrease until the first letter of the string is reached.
#
# As the length of a string `s` is `len(s)`, the last letter of the string has index `len(s)-1`. With negative indices, the first letter of the string has index `-len(s)`.
#
# If a string is stored in a variable, the individual letters of the string can be accessed by the variable name and the index of the requested letter between square brackets (`[]`) next to it.
# +
fruit = "orange"
def print_indices(fruit,n):
print(fruit[n])
print_indices(fruit,1)
print_indices(fruit,2)
print_indices(fruit,4)
print_indices(fruit,-1)
print_indices(fruit,-6)
print_indices(fruit,-3)
# -
# Besides using single indices you can also access a substring (also called a "slice") from a string by using two numbers between the square brackets with a colon (`:`) in between. The first of these numbers is the index where the substring starts, the second where it ends. The substring does *not* include the letter at the second index. By leaving out the left number you indicate that the substring starts at the beginning of the string (i.e., at index 0). By leaving out the right number you indicate that the substring ranges up to and includes the last character of the string.
#
# If you try to access a character using an index that is beyond the reaches of a string, you get a runtime error ("index out of bounds"). For a range of indices to access substrings such limitations do not exist; you can use numbers that are outside the bounds of the string.
fruit = "orange"
print(fruit[:])
print(fruit[0:])
print(fruit[:5])
print(fruit[:100])
print(fruit[:len(fruit)])
print(fruit[1:-1])
print(fruit[2], fruit[1:6])
# ### Traversing strings
# We already saw how you can traverse the characters of a string using a `for` loop:
# +
fruit = 'apple'
def traverse_characters(word):
new_word = ""
for char in word:
new_word+=(char + ' - ')
return new_word
print(traverse_characters(fruit))
# -
# Now you know about indices, you probably realize you can also use those to traverse the characters of a string:
# +
fruit = 'apple'
def traverse_characters2(word):
new_word = ""
for i in range(0, len(word)):
new_word += word[i] + " - "
return new_word
def traverse_characters3(word):
new_word = ""
i = 0
while i < len(word):
new_word += word[i] + " - "
i += 1
return new_word
print(traverse_characters2(fruit)+"\n"+traverse_characters3(fruit))
# -
# If you just want to traverse the individual characters of a string, the first method, using `for <character> in <string>:`, is by far the most elegant and readable. However, occasionally you have to solve problems in which you might prefer one of the other methods.
#
# **Exercise (optional)**: Write code that for a string prints the indices of all of its vowels (`a`, `e`, `i`, `o`, and `u`). This can be done with a `for` loop or a `while` loop.
# +
# Indices of vowels
def index_vowels (text):
index_vowels("apple")
# -
# **Exercise (optional)**: Write code that uses two strings. For each character in the first string that has exactly the same character at the same index in the second string, you print the character and the index. Watch out that you do not get an "index out of bounds" runtime error.
# +
# Your function
s1 = "The Holy Grail"
s2 = "Life of Brian"
def similar_char(text1, text2):
print(similar_char(s1, s2))
# -
# **Exercise (optional)**: Write a function that takes a string as argument, and creates a new string that is a copy of the argument, except that every non-letter is replaced by a space (e.g., "`ph@t l00t`" is changed to "`ph t l t`"). To write such a function, you will start with an empty string, and traverse the characters of the argument one by one. When you encounter a character that is acceptable, you add it to the new string. When it is not acceptable, you add a space to the new string. Note that you can check whether a character is acceptable by simple comparisons, e.g., any lower case letter can be found using the test `if ch >= 'a' and ch <= 'z':`.
# +
# String cleaning function
def clean_string(string):
clean_string("Aph@t 100t")
# -
# ### Extended slices
# Slices (substrings) in python can take a third argument, which is the step size (or "stride") that is taken between indices. It is similar to the third argument for the `range()` function. The format for slices then becomes `<string>[<begin>:<end>:<step>]`. By default the step size is 1.
#
# The most common use for the step size is to use a negative step size in order to create a reversed version of a string.
fruit = "banana"
print(fruit[::2])
print(fruit[1::2])
print(fruit[::-1])
print(fruit[::-2])
# Reversing a string using `[::-1]` is conceptually similar to traversing the string from the last character to the beginning of the string using backward steps of size 1.
fruit = "banana"
for i in range(len(fruit), -1):
print(fruit[i])
# ### Strings are immutable
# A core property of strings is that they are *immutable*. This means that they cannot be changed. For instance, you cannot change a character of a string by assigning a new value to it. As a demonstration, the following code leads to a runtime error if you try to run it:
fruit = "oringe"
fruit[2] = "a"
print(fruit)
# If you want to make a change to a string, you have to create a new string that contains the change; you can then assign the new string to the existing variable if you want. For instance:
fruit = "oringe"
fruit = fruit[:2] + "a" + fruit[3:]
print(fruit)
# The reasons for why strings are immutable are beyond the scope of this course. Just remember that if you want to modify a string you need to overwrite the entire string, and you cannot modify individual indices.
# ### `string` methods
# There is a collection of methods that are designed to operate on strings. All of these methods are applied to a string to perform some operation. Since strings are immutable, they *never change* the string they work on, but they always `return` a changed version of the string.
#
# All these methods are called as `<string>.<method>()`, i.e., you have to write the string that they work on before the method call, with a period in between. You will encounter this more often, and why this is implemented in this way will be explained later in the course, in the chapters about object orientation.
#
# Most of these methods are not part of a specific module, but can be called without importing them. There is a `string` module that contains specific constants and methods that can be used in your programs, but the methods I discuss here can all be used without importing the `string` module.
# ### `strip()`
# `strip()` removes from a string leading and trailing spaces, including leading and trailing newlines and other characters that may be viewed as spaces. There are no parameters. See the following example (the string is bordered by [ and ] to show the effect):
s = " And now for something completely different\n "
print("["+s+"]")
s = s.strip()
print("["+s+"]")
# ### `upper()` and `lower()`
# `upper()` creates a version of a string of which all letters are capitals. `lower()` is equivalent, but uses only lower case letters. Neither method uses parameters.
s = "The Meaning of Life"
print(s)
print(s.upper())
print(s.lower())
# ### `find()`
# `find()` can be used to search in a string for the starting index of a particular substring. As parameters it gets the substring, and optionally a starting index to search from, and an ending index. It returns the lowest index where the substring starts, or `-1` if the substring is not found.
# +
s = "Humpty Dumpty sat on the wall"
print(s.find("sat"))
print(s.find("t"))
print(s.find("t", 12))
print(s.find("q"))
s.find(" ")
# -
# ### `replace()`
# `replace()` replaces all occurrences of a substring with another substring. As parameters it gets the substring to look for, and the substring to replace it with. Optionally, it gets a parameter that indicates the maximum number of replacements to be made.
#
# I must stress again that strings are immutable, so the `replace()` function is not actually changing the string. It returns a new string that is a copy of the string with the replacements made.
# +
s = ' <NAME> sat on the wall '
new_s = s.replace('sat on', 'fell off')
print(new_s)
print(s)
# -
# ### `split()`
# `split()` splits a string up in words, based on a given character or substring which is used as separator. The separator is given as the parameter, and if no separator is given, the white space is used, i.e., you split a string in the actual words (though punctuation attached to words is considered part of the words). If there are multiple occurrences of the separator next to each other, the extra ones are ignored (i.e., with the white space as separator, it does not matter if there is a single white space between two words, or multiple).
#
# The result of this split is a so-called "list" of strings. Lists are discussed in a coming chapter. However, note that if you want to access the separate words, you can use the `for <word> in <list>:` construction.
s = '<NAME> sat on the wall'
wordlist = s.split()
for i in wordlist:
print(i)
print(wordlist)
# A very useful property of splitting is that we can decode some basic file formats. For example, a comma separated value (CSV) file is a very simple format, of which the basic setup is that each line consists of values that are separated by a comma. These values can be split from each other using the `split()` method. (Note: In actuality it will be a bit more convoluted as there might be commas in the fields that are stored in the CSV file, so it depends a bit on the contents of the file whether this simple approach will work. More on CSV files will be said in a later chapter in the course, where file formats are discussed.)
# +
csv = "2016,September,28,Data Processing,Tilburg University,Tilburg"
values = csv.split(',')
for value in values:
print(value)
print("")
print(values)
print (values[1][0])
# -
# ### `join()`
# `join()` is the opposite of `split()`. `join()` joins a list of words together, separated by a specific separator. This sounds like it would be a method of lists, but for historic reasons it is defined as a string method. Since all string methods are called with the format `<string>.<method>()`, there must be a string in front of the call to `join()`. That string is the separator that you want to use, while the parameter of the method is the list that you want to join together. The return value, as always, is the resulting string. In the following example, note the notation of each of these steps:
s = "Humpty;Dumpty;sat;on;the;wall"
print (s)
wordlist = s.split(';')
print (wordlist)
s = " ".join(wordlist)
print(s)
# ### What you learned
# In this chapter, you learned about:
#
# - Strings
# - Multi-line strings
# - Accessing string characters with positive and negative indices
# - Slices
# - Immutability of strings
# - String methods `strip()`, `upper()`, `lower()`, `find()`, `replace()`, `split()`, and `join()`
# - Escape sequences
# # Exercises
# **Exercise 6.1:** The text string in the next cell contains several words which are enclosed by square brackets (`[` and `]`). Scan the string and print out all words which are between square brackets. For example, if the text string would be "`[a]n example[ string]`", you are expected to print out "`a string`".
# +
# Distilling text.
text = """The quick, brown fox jumps over a lazy dog. DJs flock by when MTV ax quiz prog.
Junk MTV quiz graced by fox whelps. [Never gonna ] Bawds jog, flick quartz, vex nymphs.
[give you up\n] Waltz, bad nymph, for quick jigs vex! Fox nymphs grab quick-jived waltz.
Brick quiz whangs jumpy veldt fox. [Never ] Bright vixens jump; [gonna let ] dozy fowl
quack. Quick wafting zephyrs vex bold Jim. Quick zephyrs blow, vexing daft Jim. Charged
[you down\n] fop blew my junk TV quiz. How quickly daft jumping zebras vex. Two driven
jocks help fax my big quiz. Quick, Baz, get my woven flax jodhpurs! "Now fax quiz Jack!"
my brave ghost pled. [Never ] Five quacking zephyrs jolt my wax bed. [gonna ] Flummoxed
by job, kvetching W. zaps Iraq. Cozy sphinx waves quart jug of bad milk. [run around ]
A very bad quack might jinx zippy fowls. Few quips galvanized the mock jury box. Quick
brown dogs jump over the lazy fox. The jay, pig, fox, zebra, and my wolves quack!
[and desert you] Blowzy red vixens fight for a quick jump. <NAME> was gazed
by MTV for luck. A wizard’s job is to vex chumps quickly in fog. Watch "Jeopardy!",
<NAME>'s fun TV quiz game."""
text_split = text.split("[")
for i in range(1, len(text_split)) :
bracket_text = text_split[i].split("]")
print(bracket_text[0])
# -
# **Exercise 6.2:** Print a line of all the capital letters "A" to "Z". Below it, print a line of the letters that are 13 positions in the alphabet away from the letters that are above them. E.g., below the "A" you print an "N", below the "B" you print an "O", etcetera. You have to consider the alphabet to be circular, i.e., after the "Z", it loops back to the "A" again.
# +
# ROTR-13
letters = "ABCDEFGHIJKLMNOPQRSTUVWXYZ"
new_letters = letters[13:] + letters[:13]
print(letters)
print(new_letters)
# -
# **Exercise 6.3:** In the text below, count how often the word "wood" occurs (using program code, of course). Capitals and lower case letters may both be used, and you have to consider that the word "wood" should be a separate word, and not part of another word. Hint: If you did the exercises from this chapter, you already developed a function that "cleans" a text. Combining that function with the `split()` function more or less solves the problem for you.
# +
text = """How much wood would a woodchuck chuck
If a woodchuck could chuck wood?
He would chuck, he would, as much as he could,
And chuck as much as a woodchuck would
If a Mr. Smith could chuck wood\n\r\t."""
# read whole text
# create a counter
# get rid \n
# get rid of ?. special grammar
# lowercase my sentence
"if a woodchuck could chuck wood"
# split the string by some character
["if", "a", "woodchuck"]
#check if wood is in the list
# if yes
# counter = counter +1 <--> counter += 1
# else
#pass
#return
# +
# Counting wood.
def wood_counter(text):
clean_text = text.replace("?", " ").replace("\n"," ").replace("\r", " ").replace("\t", " ").replace(",", " ").replace(".", " ")
lower_text = clean_text.lower()
split_text = lower_text.split()
counter = 0
print(split_text)
for word in split_text:
if word == "wood":
counter += 1
return counter
wood_counter(text)
# -
help(str)
| Chapter 1/3. Python/String and Files/exercises/Into to Python - part 6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="q9PN64nMyo7p"
# #Data Pre-Processing
# -
# !pip install pillow
# + id="0JpvmghRxdnG"
# import the necessary packages
import PIL
import PIL.Image
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.applications import MobileNetV2
from tensorflow.keras.layers import AveragePooling2D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.preprocessing.image import load_img
from tensorflow.keras.utils import to_categorical
from sklearn.preprocessing import LabelBinarizer
from sklearn.model_selection import train_test_split
from sklearn.metrics import classification_report
from imutils import paths
import matplotlib.pyplot as plt
import numpy as np
import os
# + colab={"base_uri": "https://localhost:8080/", "height": 231} id="ckvARvyMyg1B" outputId="5b62afe4-d0e5-458e-b390-896398e84a5c"
data_dir = "dataset/"
outputs = ["with_mask", "without_mask"]
data = []
labels = []
for out in outputs:
path = os.path.join(data_dir, out)
for img in os.listdir(path):
image_path = os.path.join(path, img)
image = load_img(image_path, target_size = (224, 224, 3))
image = img_to_array(image)
image = preprocess_input(image)
data.append(image)
labels.append(out)
# + [markdown] id="NpAfuGiu0whN"
# Binarize Labels
# + colab={"base_uri": "https://localhost:8080/", "height": 374} id="Etykf7xf0q7O" outputId="d7b0f358-6580-41ae-ea17-1e8d445397d0"
lb = LabelBinarizer()
labels = lb.fit_transform(labels)
labels = to_categorical(labels)
data = np.array(data, dtype="float32")
labels = np.array(labels)
# + [markdown] id="W0cFK9Tp1LbC"
# Train_Test_Split
#
# + colab={"base_uri": "https://localhost:8080/", "height": 306} id="9dSOFMeJ1I30" outputId="d24594a8-eccc-4472-8ef8-e2c5fe7b3d04"
# + [markdown] id="nCf_fsOi2rCa"
# Custom keras sequential model
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="HuxYVGEa2ERh" outputId="b2a304cb-c6b8-486a-925a-e24f753d3641"
import numpy
import time
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import Flatten
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.layers import BatchNormalization
seed = 7
numpy.random.seed(seed)
model = Sequential()
model.add(Conv2D(filters=32, kernel_size=(3, 3), input_shape=(224, 224, 3), activation='relu'))
model.add(BatchNormalization(axis=3))
model.add(Conv2D(filters=32, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(BatchNormalization(axis=3))
model.add(Dropout(0.1))
model.add(Conv2D(filters=64, kernel_size=(3, 3), activation='relu'))
model.add(BatchNormalization(axis=3))
model.add(Conv2D(filters=64, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(BatchNormalization(axis=3))
model.add(Dropout(0.1))
model.add(Conv2D(filters=128, kernel_size=(3, 3), activation='relu'))
model.add(BatchNormalization(axis=3))
model.add(Conv2D(filters=128, kernel_size=(3, 3), activation='relu'))
model.add(MaxPooling2D((2, 2)))
model.add(BatchNormalization(axis=3))
model.add(Dropout(0.1))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(128, activation='relu'))
model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(12, activation='softmax'))
model.summary()
# compile model
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="tAMBOVxY2kU3"
INIT_LR = 1e-4
EPOCHS = 20
BS = 32
# + [markdown] id="HYPsvF474eOB"
# Mobile net cnn
#
# + colab={"base_uri": "https://localhost:8080/", "height": 374} id="U-SHjfvd4gH6" outputId="9142330d-8b78-48c0-d20d-0de1bf7e2bc3"
train_x, test_x, train_y, test_y = train_test_split(data, labels, test_size = 0.20, stratify = labels, random_state = 30)
image_aug = ImageDataGenerator(
rotation_range=180, # randomly rotate images in the range
zoom_range = 0.1, # Randomly zoom image
width_shift_range=0.1, # randomly shift images horizontally
height_shift_range=0.1, # randomly shift images vertically
horizontal_flip=True, # randomly flip images horizontally
vertical_flip=True # randomly flip images vertically
)
image_aug.fit(train_x)
# load the MobileNetV2 network, ensuring the head FC layer sets are
# left off
baseModel = MobileNetV2(weights="imagenet", include_top=False,
input_tensor=Input(shape=(224, 224, 3)))
# construct the head of the model that will be placed on top of the
# the base model
headModel = baseModel.output
headModel = AveragePooling2D(pool_size=(7, 7))(headModel)
headModel = Flatten(name="flatten")(headModel)
headModel = Dense(128, activation="relu")(headModel)
headModel = Dropout(0.5)(headModel)
headModel = Dense(2, activation="softmax")(headModel)
# place the head FC model on top of the base model (this will become
# the actual model we will train)
model = Model(inputs=baseModel.input, outputs=headModel)
# loop over all layers in the base model and freeze them so they will
# *not* be updated during the first training process
for layer in baseModel.layers:
layer.trainable = False
# compile our model
print("[INFO] compiling model...")
opt = Adam(lr=INIT_LR, decay=INIT_LR / EPOCHS)
model.compile(loss="binary_crossentropy", optimizer=opt,
metrics=["accuracy"])
# train the head of the network
print("[INFO] training head...")
H = model.fit(
image_aug.flow(train_x, train_y, batch_size=BS),
steps_per_epoch=len(trainX) // BS,
validation_data=(testX, testY),
validation_steps=len(testX) // BS,
epochs=EPOCHS)
# make predictions on the testing set
print("[INFO] evaluating network...")
predIdxs = model.predict(testX, batch_size=BS)
# for each image in the testing set we need to find the index of the
# label with corresponding largest predicted probability
predIdxs = np.argmax(predIdxs, axis=1)
# show a nicely formatted classification report
print(classification_report(testY.argmax(axis=1), predIdxs,
target_names=lb.classes_))
# serialize the model to disk
print("[INFO] saving mask detector model...")
model.save("mask_detector.model", save_format="h5")
# + id="xxoNFJIX5eCl"
# import the necessary packages
from tensorflow.keras.applications.mobilenet_v2 import preprocess_input
from tensorflow.keras.preprocessing.image import img_to_array
from tensorflow.keras.models import load_model
from imutils.video import VideoStream
import numpy as np
import imutils
import time
import cv2
import os
def detect_and_predict_mask(frame, faceNet, maskNet):
# grab the dimensions of the frame and then construct a blob
# from it
(h, w) = frame.shape[:2]
blob = cv2.dnn.blobFromImage(frame, 1.0, (224, 224),
(104.0, 177.0, 123.0))
# pass the blob through the network and obtain the face detections
faceNet.setInput(blob)
detections = faceNet.forward()
print(detections.shape)
# initialize our list of faces, their corresponding locations,
# and the list of predictions from our face mask network
faces = []
locs = []
preds = []
# loop over the detections
for i in range(0, detections.shape[2]):
# extract the confidence (i.e., probability) associated with
# the detection
confidence = detections[0, 0, i, 2]
# filter out weak detections by ensuring the confidence is
# greater than the minimum confidence
if confidence > 0.5:
# compute the (x, y)-coordinates of the bounding box for
# the object
box = detections[0, 0, i, 3:7] * np.array([w, h, w, h])
(startX, startY, endX, endY) = box.astype("int")
# ensure the bounding boxes fall within the dimensions of
# the frame
(startX, startY) = (max(0, startX), max(0, startY))
(endX, endY) = (min(w - 1, endX), min(h - 1, endY))
# extract the face ROI, convert it from BGR to RGB channel
# ordering, resize it to 224x224, and preprocess it
face = frame[startY:endY, startX:endX]
face = cv2.cvtColor(face, cv2.COLOR_BGR2RGB)
face = cv2.resize(face, (224, 224))
face = img_to_array(face)
face = preprocess_input(face)
# add the face and bounding boxes to their respective
# lists
faces.append(face)
locs.append((startX, startY, endX, endY))
# only make a predictions if at least one face was detected
if len(faces) > 0:
# for faster inference we'll make batch predictions on *all*
# faces at the same time rather than one-by-one predictions
# in the above `for` loop
faces = np.array(faces, dtype="float32")
preds = maskNet.predict(faces, batch_size=32)
# return a 2-tuple of the face locations and their corresponding
# locations
return (locs, preds)
# load our serialized face detector model from disk
prototxtPath = r"face_detector/deploy.prototxt"
weightsPath = r"face_detector/res10_300x300_ssd_iter_140000.caffemodel"
faceNet = cv2.dnn.readNet(prototxtPath, weightsPath)
# load the face mask detector model from disk
maskNet = load_model("mask_detector.model")
# initialize the video stream
print("[INFO] starting video stream...")
vs = VideoStream(src=0).start()
# loop over the frames from the video stream
while True:
# grab the frame from the threaded video stream and resize it
# to have a maximum width of 400 pixels
frame = vs.read()
frame = imutils.resize(frame, width=900)
# detect faces in the frame and determine if they are wearing a
# face mask or not
(locs, preds) = detect_and_predict_mask(frame, faceNet, maskNet)
# loop over the detected face locations and their corresponding
# locations
for (box, pred) in zip(locs, preds):
# unpack the bounding box and predictions
(startX, startY, endX, endY) = box
(mask, withoutMask) = pred
# determine the class label and color we'll use to draw
# the bounding box and text
label = "Mask" if mask > withoutMask else "No Mask"
color = (0, 255, 0) if label == "Mask" else (0, 0, 255)
# include the probability in the label
label = "{}: {:.2f}%".format(label, max(mask, withoutMask) * 100)
# display the label and bounding box rectangle on the output
# frame
cv2.putText(frame, label, (startX, startY - 10),
cv2.FONT_HERSHEY_SIMPLEX, 0.70, color, 2)
cv2.rectangle(frame, (startX, startY), (endX, endY), color, 4)
# show the output frame
cv2.imshow("Frame", frame)
key = cv2.waitKey(1) & 0xFF
# if the `q` key was pressed, break from the loop
if key == ord("q"):
break
# do a bit of cleanup
cv2.destroyAllWindows()
vs.stop()
# + [markdown] id="7U5HVH367YGo"
# #### New Section
# -
| Untitled0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pistarlab
# language: python
# name: pistarlab
# ---
# # Load Snapshot
# +
import sys
# sys.path.append("?../")
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# %config Completer.use_jedi = False
# -
from pistarlab import ctx
ctx.initialize()
# !ls /home/brandyn/pistarlab/snapshot_repo/agent/
ctx.connect()
ctx.get_snapshot_index()
ctx.get_snapshot_index()
ctx.create_agent_from_snapshot('agent_stable_baselines_PPO_default_0x7554a16a_0-dev')
source = "https://raw.githubusercontent.com/pistarlab/pistarlab-repo/main/snapshots/"
ctx.load_remote_snapshot_index(source)
| tests/test_remote_snapshot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Note: 1번 DiscreteREINFORCE에서 겹치는 변수나 함수는 설명이 기재되어있지 않습니다.
# #### 따라서 설명이 있는 부분만 보시면, 1번과의 차이점을 확인하실 수 있습니다.
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torchsummary import summary
import gym
import os
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import clear_output
# for using sampling with gradient-tracking when selecting an action
# Discrete action과는 달리, Normal 함수를 활용하는 것을 보실 수 있습니다 (Categorical은 불연속 확률 변수에 대해서 정의되어지는 확률 분포이기 때문).
# 이름에서 짐작하실 수 있듯이, gaussian 확률 분포입니다.
from torch.distributions import Normal
# -
class Policy(nn.Module):
def __init__(self, input_dim, action_dim, hidden):
super(Policy, self).__init__()
self.fc1 = nn.Linear(input_dim, hidden)
self.mean = nn.Linear(hidden, action_dim) # Discrete 과의 차이점. 각 action의 mean, std를 구한다.
self.mean_scale = 1 # 일반적으로 초기 학습의 안정성을 위해서 mean값의 상한과 하한을 정하는데, 예를 들어 tanh를 쓰면 -1~1로 정할 수 있습니다. 이러한 상한, 하한의 scaling을 위한 변수.
self.std = nn.Linear(hidden, action_dim)
def forward(self, state):
out = F.relu(self.fc1(state))
mean = torch.tanh(self.mean(out)) * self.mean_scale # 평균값 상한과 하한을 위해 tanh를 적용하였습니다.
std = torch.exp(self.std(out)) # sutton책 chapter.13의 13.7에 exponential을 쓰는 이유가 나와있습니다 (양수로 만들기 위함 + 좀 더 나은 approximate 성능).
return mean, std
class Agent():
def __init__(self, env, n_epi, max_steps, gamma, plot_freq, input_dim, action_dim, hidden, learning_rate, device, save_mode, model_name, saving_start_epi):
# environment parameter
self.gamma = gamma
self.env = env
self.input_dim = input_dim
self.action_dim = action_dim
# trainig parameter
self.device = device
self.n_epi = n_epi
self.max_steps = max_steps
self.plot_freq = plot_freq
self.frame_cnt = 0
# network paramter
self.lr = learning_rate
self.policy = Policy(self.input_dim, self.action_dim, hidden).to(self.device)
self.optimizer = optim.Adam(self.policy.parameters(), lr=self.lr)
# log parameter
self.avg = []
self.scores = []
self.losses = []
self.save_mode = save_mode
self.model_name = model_name
self.saving_start = saving_start_epi
def select_action(self, state):
"""
Discrete action space일 때 뽑는 것과 달리, policy net에서 구한 mean, std를 이용하여 Gaussian 분포를 생성한 후(Normal 함수),
그 함수에서 action을 sampling 합니다. 그런데 이렇게 sampling하면, 예를 들어 mean값이 -1~1로 정해졌다 할지라도
gaussian 분포에서 뽑는 것이기 때문에 절대값이 매우 큰 양수 or 음수가 action으로 뽑힐 수 있습니다.
그러므로 이렇게 뽑힌 action을 다시 한번 상한과 하한을 정할 수 있는데요,
다만, 이때 주의할 것이 미분가능을 염두하여야 하므로, torch.clamp를 일반적으로 활용합니다.
예) torch.clamp(tensor, min=-1.0, max=1.0)
"""
mean, std = self.policy(torch.FloatTensor(state).view(1, -1).to(self.device))
nor_dist = Normal(mean, std)
action = torch.clamp(nor_dist.sample(), min=-2.0, max=2.0)
# [0]을 넣은 이유: nor_dist.sample()의 차원이 (1, 2)라서 0차원을 없애기 위해서. .unsqueeze()를 활용해도 됩니다.
return action.detach().cpu().numpy()[0], nor_dist.log_prob(action)[0].sum(), mean.detach().cpu().numpy(), std.detach().cpu().numpy()
def train(self):
for i_episode in range(1, self.n_epi+1):
epi_history = []
rewards = []
score = 0
state = self.env.reset()
for step in range(self.max_steps):
action, log_prob, mean, std = self.select_action(state)
next_state, reward, done, _ = env.step(action)
epi_history.append([log_prob, reward])
state = next_state
score += reward
self.frame_cnt += 1
if done: break
self.scores.append(score)
loss = self._update_policy(epi_history)
self.losses.append(loss)
if i_episode%self.plot_freq == 0:
self._plot_status(i_episode, self.losses, self.scores)
if self.save_mode & (i_episode > self.saving_start): self._save_model()
self.avg.append(np.mean(self.scores[-10:]))
def test(self, model_path):
''' 저장된 pt파일을 불러와 test 합니다 '''
self.policy.load_state_dict(torch.load(model_path))
self.policy.eval()
state = self.env.reset()
done = False
score = 0
accum_frames = []
while not done:
accum_frames.append(self.env.render(mode="rgb_array"))
action, log_prob, prob = self.select_action(state)
next_state, reward, done, _ = self.env.step(action)
state = next_state
score += reward
print("score: ", score)
self.env.close()
return accum_frames
def _update_policy(self, epi_history):
accum_R = 0
loss = 0
epi_len = len(epi_history) - 1
for idx, [log_p, r] in enumerate(epi_history[::-1]):
accum_R = r + (self.gamma*accum_R)
gamma_pow_t = self.gamma**(epi_len-idx)
one_step_loss = -log_p * gamma_pow_t * accum_R
loss += one_step_loss
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
return loss.item()
def _save_model(self):
last_mean = np.mean(self.scores[-11:-1])
if max(self.avg) < last_mean:
torch.save(self.policy.state_dict(), self.model_name+f'Score_{round(last_mean, 3)}.pt')
def _plot_status(self, i_episode, losses, scores):
subplot_params = [
(121, f"Scores in episode_{i_episode}", scores),
(122, f"Policy loss in episode_{i_episode}", losses),
]
clear_output(True)
plt.figure(figsize=(10, 5), facecolor='w')
for loc, title, values in subplot_params:
plt.subplot(loc)
plt.title(f'Frame:{self.frame_cnt} '+title)
plt.plot(values)
plt.show()
# +
device = torch.device(
"cuda:1" if torch.cuda.is_available() else "cpu"
)
env_list = ["Pendulum-v0", "MountainCarContinuous-v0", "LunarLanderContinuous-v2"]
env_name = env_list[0]
# 몇 episode 이후부터 저장할 것인지 정하는 변수 입니다.
saving_start_epi = 100
# model을 저장할 폴더를 지정합니다. 기본은 저장하지 않는 mode 입니다.
save_mode = True
model_save_folder = './model_save'
if not os.path.exists(model_save_folder):
os.mkdir(model_save_folder)
model_name = f"./{model_save_folder}/Continuous_REINFORCE_{env_name}_"
env = gym.make(env_name)
state_dim = env.observation_space.shape[0]
action_dim = env.action_space.shape[0]
print("Environment: ", env_name)
print("State Dimension:", state_dim, " Action Dimension:", action_dim)
n_epi = 100000
max_steps = 500
gamma = 0.999
plot_freq = 10
hidden = 64
learning_rate = 0.00001
# -
agent = Agent(
env,
n_epi,
max_steps,
gamma,
plot_freq,
state_dim,
action_dim,
hidden,
learning_rate,
device,
save_mode,
model_name,
saving_start_epi)
# +
### Continous action space의 경우는 learning rate 등의 hyper parameter에 더욱 민감한 것 같습니다. 학습이 잘 안되는 경우가 생각보다 많았습니다.
agent.train()
# +
# 모델을 test하고 게임 플레이한 결과를 배열로 저장합니다.
model_path = f"./{model_save_folder}/Discrete_REINFORCE_CartPole-v0_Score_200.0.pt"
frames = agent.test(model_path)
# 위의 배열로 저장한 것을 test.mp4 파일을 만들고 재생합니다.
# 배열을 재생시키는 방법은 여러가지가 있습니다. 만일,
import imageio
from IPython.display import Video
imageio.mimwrite('./test.mp4', frames, fps=30)
Video('./test.mp4', width=480, height=360)
# -
# LunarLander 환경
#
# n_epi = 100000
# max_steps = 500
# gamma = 0.999
# plot_freq = 10
# hidden = 64
# learning_rate = 0.0008
# 
# 
# LunarLander 환경
#
# n_epi = 100000
# max_steps = 500
# gamma = 0.95
# plot_freq = 10
# hidden = 128
# learning_rate = 0.00075
# 
# LunarLander 환경
#
# n_epi = 100000
# max_steps = 500
# gamma = 0.95
# plot_freq = 10
# hidden = 128
# learning_rate = 0.002
# 
# LunarLander 환경
#
# n_epi = 100000
# max_steps = 500
# gamma = 0.95
# plot_freq = 10
# hidden = 64
# learning_rate = 0.006
# 
| Policy_Based/REINFORCE/2. ContinuousREINFORCE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/felipefreitas93/Colab-XLNet-FineTuning/blob/master/clean_bert_gpu_svm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qBHO2O30RHt3" colab_type="code" colab={}
DATASET_NAME = 'MPQA'
# + id="-xViSbArPQgi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="6e86492b-82e4-47a8-9035-9ccd4ba77fbf"
#downloads
# !git clone https://github.com/felipefreitas93/NLPdatasets.git
# !pip install bert-tensorflow
# + id="3Wmu131YPWhd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 98} outputId="c01f065b-199a-4812-ce69-37799d2fcd02"
from tqdm import tqdm
import pandas as pd
from sklearn.model_selection import train_test_split
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
from datetime import datetime
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
# + id="wjEuoy_3QITG" colab_type="code" colab={}
BERT_MODEL_HUB = "https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1"
DATA_COLUMN = 'x'
LABEL_COLUMN = 'y'
label_list = [0, 1]
OUTPUT_DIR = 'OUTPUT'
MAX_SEQ_LENGTH = 256
NUMBER_EXAMPLES = 250
BATCH_SIZE = 16
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 3.0
WARMUP_PROPORTION = 0.1
SAVE_CHECKPOINTS_STEPS = 5000
SAVE_SUMMARY_STEPS = 1000
# + id="-2q9TrbCPA6h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="8274c32c-ab32-4e1e-a6fd-fda2e4d72dfb"
results_list = []
for RANDOM_STATE in tqdm(range(30)):
#create the output dir file
tf.gfile.MakeDirs(OUTPUT_DIR)
train = pd.read_csv(f'./NLPdatasets/{DATASET_NAME}/train.tsv', sep='\t', names=['y','x']).dropna().sample(n=NUMBER_EXAMPLES, random_state=RANDOM_STATE)
test = pd.read_csv(f'./NLPdatasets/{DATASET_NAME}/dev.tsv', sep='\t', names=['y','x']).dropna()
train_InputExamples = train.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
test_InputExamples = test.apply(lambda x: bert.run_classifier.InputExample(guid=None,
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
def create_tokenizer_from_hub_module():
"""Get the vocab file and casing info from the Hub module."""
with tf.Graph().as_default():
bert_module = hub.Module(BERT_MODEL_HUB)
tokenization_info = bert_module(signature="tokenization_info", as_dict=True)
with tf.Session() as sess:
vocab_file, do_lower_case = sess.run([tokenization_info["vocab_file"],
tokenization_info["do_lower_case"]])
return bert.tokenization.FullTokenizer(
vocab_file=vocab_file, do_lower_case=do_lower_case)
#creating tokenizer
tokenizer = create_tokenizer_from_hub_module()
# Convert our train and test features to InputFeatures that BERT understands.
train_features = bert.run_classifier.convert_examples_to_features(train_InputExamples, label_list, MAX_SEQ_LENGTH, tokenizer);
test_features = bert.run_classifier.convert_examples_to_features(test_InputExamples, label_list, MAX_SEQ_LENGTH, tokenizer);
#create a model that uses hinge loss
def create_model(is_predicting, input_ids, input_mask, segment_ids, labels,
num_labels):
"""Creates a classification model."""
bert_module = hub.Module(
BERT_MODEL_HUB,
trainable=True)
bert_inputs = dict(
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids)
bert_outputs = bert_module(
inputs=bert_inputs,
signature="tokens",
as_dict=True)
# Use "pooled_output" for classification tasks on an entire sentence.
# Use "sequence_outputs" for token-level output.
output_layer = bert_outputs["pooled_output"]
hidden_size = output_layer.shape[-1].value
# Create our own layer to tune for politeness data.
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
# Dropout helps prevent overfitting
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
logits = tf.nn.bias_add(logits, output_bias)
log_probs = tf.nn.log_softmax(logits, axis=-1)
# Convert labels into one-hot encoding
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
# If we're predicting, we want predicted labels and the probabiltiies.
if is_predicting:
return (predicted_labels, log_probs)
# If we're train/eval, compute loss between predicted and actual label
# per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
# loss = tf.reduce_mean(per_example_loss)
#attempt 1 at creating a hinge loss function
loss = tf.losses.hinge_loss(one_hot_labels, logits, scope='loss')
return (loss, predicted_labels, log_probs)
# model_fn_builder actually creates our model function
# using the passed parameters for num_labels, learning_rate, etc.
def model_fn_builder(num_labels, learning_rate, num_train_steps,
num_warmup_steps):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
is_predicting = (mode == tf.estimator.ModeKeys.PREDICT)
# TRAIN and EVAL
if not is_predicting:
(loss, predicted_labels, log_probs) = create_model(
is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
train_op = bert.optimization.create_optimizer(
loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu=False)
# Calculate evaluation metrics.
def metric_fn(label_ids, predicted_labels):
accuracy = tf.metrics.accuracy(label_ids, predicted_labels)
f1_score = tf.contrib.metrics.f1_score(
label_ids,
predicted_labels)
auc = tf.metrics.auc(
label_ids,
predicted_labels)
recall = tf.metrics.recall(
label_ids,
predicted_labels)
precision = tf.metrics.precision(
label_ids,
predicted_labels)
true_pos = tf.metrics.true_positives(
label_ids,
predicted_labels)
true_neg = tf.metrics.true_negatives(
label_ids,
predicted_labels)
false_pos = tf.metrics.false_positives(
label_ids,
predicted_labels)
false_neg = tf.metrics.false_negatives(
label_ids,
predicted_labels)
return {
"eval_accuracy": accuracy,
"f1_score": f1_score,
"auc": auc,
"precision": precision,
"recall": recall,
"true_positives": true_pos,
"true_negatives": true_neg,
"false_positives": false_pos,
"false_negatives": false_neg
}
eval_metrics = metric_fn(label_ids, predicted_labels)
if mode == tf.estimator.ModeKeys.TRAIN:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
train_op=train_op)
else:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
eval_metric_ops=eval_metrics)
else:
(predicted_labels, log_probs) = create_model(
is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
predictions = {
'probabilities': log_probs,
'labels': predicted_labels
}
return tf.estimator.EstimatorSpec(mode, predictions=predictions)
# Return the actual model function in the closure
return model_fn
num_train_steps = int(len(train_features) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
# Specify outpit directory and number of checkpoint steps to save
run_config = tf.estimator.RunConfig(
model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
model_fn = model_fn_builder(
num_labels=len(label_list),
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
train_input_fn = bert.run_classifier.input_fn_builder(
features=train_features,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
print(f'Beginning Training!')
current_time = datetime.now()
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps);
print("Training took time ", datetime.now() - current_time)
test_input_fn = run_classifier.input_fn_builder(
features=test_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
results = estimator.evaluate(input_fn=test_input_fn, steps=None);
#save results to result list
results_list.append(results['eval_accuracy'])
#delete output dir file (cleanup)
tf.gfile.DeleteRecursively(OUTPUT_DIR)
| clean_bert_gpu_svm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="https://upload.wikimedia.org/wikipedia/commons/4/47/Logo_UTFSM.png" width="200" alt="utfsm-logo" align="left"/>
#
# # MAT281
# ### Aplicaciones de la Matemática en la Ingeniería
# + [markdown] slideshow={"slide_type": "slide"}
# ## Módulo 02
# ## Laboratorio Clase 06: Desarrollo de Algoritmos
# -
# ### Instrucciones
#
#
# * Completa tus datos personales (nombre y rol USM) en siguiente celda.
# * La escala es de 0 a 4 considerando solo valores enteros.
# * Debes _pushear_ tus cambios a tu repositorio personal del curso.
# * Como respaldo, debes enviar un archivo .zip con el siguiente formato `mXX_cYY_lab_apellido_nombre.zip` a <EMAIL>.
# * Se evaluará:
# - Soluciones
# - Código
# - Que Binder esté bien configurado.
# - Al presionar `Kernel -> Restart Kernel and Run All Cells` deben ejecutarse todas las celdas sin error.
# * __La entrega es al final de esta clase.__
# __Nombre__: <NAME>
#
# __Rol__: 201510008-K
# ## Ejercicio 1 (2 ptos.):
# Utilizando los datos del Gasto Fiscal Neto de Chile, crea una nueva columna del tipo `datetime` llamada `dt_date` utilizando `anio`, `mes` y el día primero de cada mes.
import os
import numpy as np
import pandas as pd
# Utilizaremos como ejemplo un dataset de gasto fiscal neto en Chile, obtenidos de una [datathon de DataCampfire](https://datacampfire.com/datathon/).
gasto_raw = pd.read_csv(os.path.join("data", "gasto_fiscal.csv"), sep=";")
gasto_raw.head()
# Pasos a seguir:
#
# 1. Renombra la columna `anio` por `year`.
# 2. Crea la columna `month` utilizando el diccionario `es_month_dict` definido abajo. Hint: Usar un mapeo.
# 3. Crea la columna `day` en que todos los registros sean igual a `1`.
# 4. Crea la columna `dt_date` con la función `pd.to_datetime`. Lee la documentación!
# 5. Finalmente, elimina las columnas `year`, `mes`, `month`, `day`.
es_month_dict = {
'enero': 1,
'febrero': 2,
'marzo': 3,
'abril': 4,
'mayo': 5,
'junio': 6,
'julio': 7,
'agosto': 8,
'septiembre': 9,
'octubre': 10,
'noviembre': 11,
'diciembre': 12
}
gasto = (
gasto_raw.rename(columns={"anio":"year"}) ## FIX ME - Renombrar columna ##
.assign(
month=lambda x: x["mes"].str.lower().map(es_month_dict), ## FIX ME - Map ##
day=1, ## FIX ME - Día 1 de cada mes ##
dt_date=lambda x: pd.to_datetime(x.loc[:,["year","month","day"]])
).drop(columns=["year","mes","month","day"])
)
gasto.head()
# ## Ejercicio 2 (1 pto.)
#
# Pivotea el dataframe `gasto_raw` tal que:
#
# - Los índices sean los ministerios (partidas).
# - Las columnas sean los años.
# - Cada celda sea la suma de los montos.
# - Rellenar las celdas vacías con `""`.
#
# ¿Cuáles son las combinaciones de Año-Ministerio que no tienen gasto?
gasto_raw.pivot_table(
index="partida",
columns="anio",
values="monto",
aggfunc="sum",
fill_value="",
)
# __Respuesta__: Ministerio de Energía (2006), Ministerio de la Mujer y la Equidad de Género (2009-2015), Ministerio del Deporte (2006-2013), Ministerio del Medio Ambiente y Servicio Electoral (2006-2016)
# ## Ejercicio 3 (1 pto.)
#
# Realiza los benchmarks del archivo `benchmark_loop.py` que se encuentra en el directorio `fast_pandas`.
#
# ¿Cuál forma dirías que es la más eficiente?
#
# Utiliza el comando mágico `%load` y edita de tal manera que el módulo `Benchmarker` se importe correctamente.
# +
# # %load fast_pandas/benchmark_loop.py
from fast_pandas.Benchmarker import Benchmarker
def iterrows_function(df):
for index, row in df.iterrows():
pass
def itertuples_function(df):
for row in df.itertuples():
pass
def df_values(df):
for row in df.values:
pass
params = {
"df_generator": 'pd.DataFrame(np.random.randint(1, df_size, (df_size, 4)), columns=list("ABCD"))',
"functions_to_evaluate": [df_values, itertuples_function, iterrows_function],
"title": "Benchmark for iterating over all rows",
"user_df_size_powers": [2, 3, 4, 5, 6],
"user_loop_size_powers": [2, 2, 1, 1, 1],
"user_df_size_powers": [2, 3, 4, 5]
}
benchmark = Benchmarker(**params)
benchmark.benchmark_all()
benchmark.print_results()
benchmark.plot_results()
# -
# __Respuesta__: Según la gráficas obtenidas df_values es la más eficiente.
| m02_data_analysis/m02_c06_development/m02_c06_lab.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # assignment
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/assignment.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/assignment.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
Assignment problem in Google CP Solver.
Winston 'Operations Research', Assignment Problems, page 393f
(generalized version with added test column)
Compare with the following models:
* Comet : http://www.hakank.org/comet/assignment.co
* ECLiPSE : http://www.hakank.org/eclipse/assignment.ecl
* Gecode : http://www.hakank.org/gecode/assignment.cpp
* MiniZinc: http://www.hakank.org/minizinc/assignment.mzn
* Tailor/Essence': http://www.hakank.org/tailor/assignment.eprime
* SICStus: http://hakank.org/sicstus/assignment.pl
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver("n-queens")
#
# data
#
# declare variables
total_cost = solver.IntVar(0, 100, "total_cost")
x = []
for i in range(rows):
t = []
for j in range(cols):
t.append(solver.IntVar(0, 1, "x[%i,%i]" % (i, j)))
x.append(t)
x_flat = [x[i][j] for i in range(rows) for j in range(cols)]
#
# constraints
#
# total_cost
solver.Add(total_cost == solver.Sum(
[solver.ScalProd(x_row, cost_row) for (x_row, cost_row) in zip(x, cost)]))
# exacly one assignment per row, all rows must be assigned
[
solver.Add(solver.Sum([x[row][j]
for j in range(cols)]) == 1)
for row in range(rows)
]
# zero or one assignments per column
[
solver.Add(solver.Sum([x[i][col]
for i in range(rows)]) <= 1)
for col in range(cols)
]
objective = solver.Minimize(total_cost, 1)
#
# solution and search
#
solution = solver.Assignment()
solution.Add(x_flat)
solution.Add(total_cost)
# db: DecisionBuilder
db = solver.Phase(x_flat, solver.INT_VAR_SIMPLE, solver.ASSIGN_MIN_VALUE)
solver.NewSearch(db, [objective])
num_solutions = 0
while solver.NextSolution():
print("total_cost:", total_cost.Value())
for i in range(rows):
for j in range(cols):
print(x[i][j].Value(), end=" ")
print()
print()
for i in range(rows):
print("Task:", i, end=" ")
for j in range(cols):
if x[i][j].Value() == 1:
print(" is done by ", j)
print()
num_solutions += 1
solver.EndSearch()
print()
print("num_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
# Problem instance
# hakank: I added the fifth column to make it more
# interestingrows = 4
cols = 5
cost = [[14, 5, 8, 7, 15], [2, 12, 6, 5, 3], [7, 8, 3, 9, 7], [2, 4, 6, 10, 1]]
| examples/notebook/contrib/assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 1 - Problem 1
# ## Introduction
# #### Ex 1: Say "Hello, World!" With Python
if __name__ == '__main__':
print "Hello, World!"
# ### Ex 2: Python If-Else
# +
# #!/bin/python
import math
import os
import random
import re
import sys
n= int(raw_input().strip())
a=range(100)
if n%2!=0:
print('Weird')
elif n%2==0:
if n in a[3:6]:
print('Not Weird')
elif n in a[7:21]:
print('Weird')
elif n>20:
print('Not Weird')
else:
print('Not Weird')
# -
# ### Ex 3: Arithmetic Operators
if __name__ == '__main__':
a=int(raw_input().strip())
b=int(raw_input().strip())
print(a+b)
print(a-b)
print(a*b)
# ### Ex 4: Python: Division
# +
from __future__ import division
if __name__ == '__main__':
a = int(raw_input())
b = int(raw_input())
print(a//b)
print(a/b)
# -
# ### Ex 5: Loops
if __name__ == '__main__':
n = int(raw_input())
for i in range(0,n,1):
print(i*i)
# ### Ex 6: Write a function
def is_leap(year):
leap = year%4==0 and (year%400==0 or year%100!=0)
return leap
# ### Ex 7: Print function
from __future__ import print_function
n = int(input())
for i in range(1,n+1):
print(i,end="")
# ## Data types
# ### Ex 1: List Comprehensions
x,y,z,n = [int(input()) for i in range(4)]
print([[i,j,k] for i in range(x+1) for j in range(y+1) for k in range(z+1) if ((i+j+k) != n)])
# ### Ex 2: Find the Runner-Up Score!
# +
n = int(input())
arr = list(map(int,raw_input().strip().split()))[:n]
m = max(arr)
while max(arr) == m:
arr.remove(max(arr))
print max(arr)
# -
# ### Ex 3: Nested Lists
# +
students = {}
n = input()
for x in range(n):
name = raw_input()
grade = input()
if grade in students:
students[grade].append(name)
else:
students[grade] = [name]
all_students = list(set(students.keys()))
all_students.sort()
names_ordered = students[all_students[1]]
names_ordered.sort()
for name in names_ordered:
print name
# -
# ### Ex 4: Finding the percentage
if __name__ == '__main__':
n = int(raw_input())
student_marks = {}
for _ in range(n):
line = raw_input().split()
name, scores = line[0], line[1:]
scores = map(float, scores)
student_marks[name] = scores
query_name = raw_input()
lq = student_marks[query_name]
print("{0:.2f}".format(sum(lq)/(len(lq))))
# ### Ex 5: Lists
if __name__ == '__main__':
N = int(raw_input())
arr=[]
for i in range(N):
ask = raw_input().split()
if ask[0] == 'insert':
arr.insert(int(ask[1]),int(ask[2]))
elif ask[0] == 'append':
arr.append(int(ask[1]))
elif ask[0] == 'remove':
arr.remove(int(ask[1]))
elif ask[0] == 'sort':
arr.sort()
elif ask[0] == 'pop':
arr.pop()
elif ask[0] == 'reverse':
arr.reverse()
else:
print arr
# ### Ex 6: Tuples
if __name__ == '__main__':
n = int(raw_input())
integer_list = map(int, input().split())
t=tuple(integer_list)
print(hash(t))
# ## Strings
# ### Ex 1: sWAP cASE
def swap_case(s):
word = ""
for i in s:
if i == i.upper():
word += i.lower()
else:
word += i.upper()
return word
# ### Ex 2: What's your name?
# +
def print_full_name(first, last):
print('Hello {} {}! You just delved into python.'.format(first, last))
if __name__ == '__main__':
first_name = input()
last_name = input()
print_full_name(first_name, last_name)
# -
# ### Ex 3: Mutations
# +
def mutate_string(string, position, character):
l = list(string)
l[int(position)] = str(character)
string = ''.join(l)
return string
if __name__ == '__main__':
s = input()
i, c = input().split()
s_new = mutate_string(s, int(i), c)
print(s_new)
# -
# ### Ex 4: String Validators
if __name__ == '__main__':
s = input() #here I initialize the string
print(any(a.isalnum() for a in s)) #looking for alphanumeric characters
print(any(a.isalpha() for a in s)) #looking for alphabetical characters
print(any(a.isdigit() for a in s)) #looking for digits characters
print(any(a.islower() for a in s)) #looking for lowercase characters
print(any(a.isupper() for a in s)) #looking for uppercase characters
# ### Ex 5: Deigner Door Mat
n, m = map(int,input().split())
pattern = [('.|.'*(2*i + 1)).center(m, '-') for i in range(n//2)]
'''The flag is symmetrical, so you have the top and the bottom consequently. You can just work on n // 2. '''
print('\n'.join(pattern + ['WELCOME'.center(m, '-')] + pattern[::-1]))
'''[ : :-1] reverses the pattern'''
# ### Ex 6: Text Alignment
# +
thickness = int(input()) #This must be an odd number
c = 'H'
#Top Cone
for i in range(thickness):
print((c*i).rjust(thickness-1)+c+(c*i).ljust(thickness-1))
#Top Pillars
for i in range(thickness+1):
print((c*thickness).center(thickness*2)+(c*thickness).center(thickness*6))
#Middle Belt
for i in range((thickness+1)//2):
print((c*thickness*5).center(thickness*6))
#Bottom Pillars
for i in range(thickness+1):
print((c*thickness).center(thickness*2)+(c*thickness).center(thickness*6))
#Bottom Cone
for i in range(thickness):
print(((c*(thickness-i-1)).rjust(thickness)+c+(c*(thickness-i-1)).ljust(thickness)).rjust(thickness*6))
# -
# ### Ex 7: String Split and Join
# +
def split_and_join(line):
line = line.split(" ") # line is converted to a list of strings.
line = "-".join(line)
return line
if __name__ == '__main__':
line = input()
result = split_and_join(line)
print(result)
# -
# ### Ex 8: Text Wrap
# +
import textwrap
def wrap(string, max_width):
return "\n".join([string[ind:ind+max_width] for ind in range(0, len(string), max_width)])
#\n is for starting a new line
#join in for concatenate these lines
#string[ind:ind+max_width] is the expression that I added in the line
if __name__ == '__main__':
string, max_width = input(), int(input())
result = wrap(string, max_width)
print(result)
# -
# ### Ex 9: Alphabet Rangoli
# +
def print_rangoli(size):
alpha = "abcdefghijklmnopqrstuvwxyz"
data = [alpha[i] for i in range(n)]
items = list(range(n))
items = items[:-1]+items[::-1]
for i in items:
temp = data[-(i+1):]
row = temp[::-1]+temp[1:]
print("-".join(row).center(n*4-3, "-"))
if __name__ == '__main__':
n = int(input())
print_rangoli(n)
# -
# ### Ex 10: String Formatting
# +
def print_formatted(n):
lst = len(str(bin(n)).replace('0b',''))
for i in range(1, n+1):
b = bin(int(i)).replace('0b','').rjust(lst, ' ')
o = oct(int(i)).replace('0o','',1).rjust(lst, ' ')
h = hex(int(i)).replace('0x','').upper().rjust(lst, ' ')
j = str(i).rjust(lst, ' ')
print(j, o, h, b)
if __name__ == '__main__':
n = int(input())
print_formatted(n)
# -
# ### Ex 11: Capitalize!
def solve(s):
wr = s.split(' ') #for obtaining a list of the words in s I use split
cap_wr = [w.capitalize() for w in wr]
return(" ".join(cap_wr))
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
s = input()
result = solve(s)
fptr.write(result + '\n')
fptr.close()
# ### Ex 12: Find a string
# +
def count_substring(string, sub_string):
conta = 0
for ind in range(0,len(string)-len(sub_string)+3): #I have to add 3 or my loop won't take the last 3 letters
'''I can iterate all over the string just once because
I know that every character could be the first one of the substring,
so I don't need two nested loops, I can use just one loop'''
if string[ind:ind+len(sub_string)]==sub_string:
''' For every iteration I want to see if the substring between these indexes is in the string '''
conta += 1
return(conta)
if __name__ == '__main__':
string = input().strip()
sub_string = input().strip()
count = count_substring(string, sub_string)
print(count)
# -
# ### Ex 13: The Minion Game
# +
def minion_game(string):
ck = 0
cs = 0
vowels = 'AEIOU'
cons = 'BCDFGHLMNPQRSTVZWXY'
for sub in range(len(string)):
if string[sub] in vowels:
ck += (len(string) - sub)
else:
cs += (len(string) - sub)
if cs > ck:
print("Stuart", cs)
elif ck > cs:
print("Kevin", ck)
else:
print("Draw")
if __name__ == '__main__':
s = input()
minion_game(s)
# -
# ### Ex 14: Merge the Tools!
# +
def merge_the_tools(string, k):
for x in zip(*[iter(string)] * k):
d = dict()
print(''.join([d.setdefault(c, c) for c in x if c not in d]))
if __name__ == '__main__':
string, k = input(), int(input())
merge_the_tools(string, k)
# -
# ## Sets
# ### Ex 1: Introduction to Sets
# +
def average(array):
mean = round(sum(set(array))/len(set(array)),3)
return(mean)
if __name__ == '__main__':
n = int(input())
arr = list(map(int, input().split()))
result = average(arr)
print(result)
# -
# ### Ex 2: Symmetic Difference
m,m1=(int(input()),input().split())
n,n1=(int(input()),input().split())
m = set(m1)
n = set(n1)
d_mn=m.difference(n)
d_nm=n.difference(m)
un=d_mn.union(d_nm)
print ('\n'.join(sorted(un, key=int)))
# ### Ex 3: Set. add()
n = int(input())
stdin = set()
for _ in range(n):
stdin.add(input())
print(len(stdin))
# ### Ex 4: Set. discard(), .remove() & .pop()
n = int(input()) #elements to take
s = set(map(int, input().split())) #element to take in the array
oth = int(input()) #number of operations to do
for i in range(oth):
job = input().split()
if job[0] == "remove": #if job wants to remove, remove the "int(job[1])" element
s.remove(int(job[1]))
elif job[0] == "discard":
s.discard(int(job[1]))
else:
s.pop()
print(sum(s))
# ### Ex 5: Set .union() Operation
n = input()
eng_roll = set(map(int, input().split()))
b = input()
fra_roll = set(map(int, input().split()))
print(len(eng_roll.union(fra_roll)))
# ### Ex 6: Set .intersection()
n = input()
eng_roll = set(map(int, input().split()))
b = input()
fra_roll = set(map(int, input().split()))
print(len(eng_roll.intersection(fra_roll)))
# ### Ex 7: Set. difference()
n = input()
eng_roll = set(map(int, input().split()))
b = input()
fra_roll = set(map(int, input().split()))
print(len(eng_roll.difference(fra_roll)))
# ### Ex 8: Set. symmetric_difference()
n = input()
eng_roll = set(map(int, input().split()))
b = input()
fra_roll = set(map(int, input().split()))
print(len(eng_roll.symmetric_difference(fra_roll)))
# ### Ex 9: Set Mutations
# +
_ = int(input()) #the number of elements in set A
el_A = set(map(int, input().split())) #the space separated list of elements in set A
n_ots = int(input()) #the number of other sets
for _ in range(n_ots):
ope, _ = input().split()
el_B = set(map(int,input().split()))
if ope == "intersection_update":
el_A.intersection_update(el_B)
elif ope == "update":
el_A.update(el_B)
elif ope == "difference_update":
el_A.difference_update(el_B)
elif ope == "symmetric_difference_update":
el_A.symmetric_difference_update(el_B)
print(sum(el_A))
# -
# ### Ex 10: Check Subset
nT = int(input()) #number of test cases
for _ in range(nT):
nA = int(input()) #the number of elements in set A
el_A = set(map(int, input().split())) #the space separated list of elements in set A
nB = int(input()) #the number of elements in set B
el_B = set(map(int, input().split())) #the space separated list of elements in set B
if el_A.intersection(el_B) == el_A:
print("True")
else:
print("False")
# ### Ex 11: Check Strict Superset
el_A = set(map(int, input().split())) #the space separated list of elements in set A
n = int(input()) #the number of other sets
el_n = set(map(int, input().split()))
if el_A>set(map(int, input().split())):
print("True")
else:
print("False")
# ### Ex 12: The Capitain's Room
k,el = int(input()),list(map(int, input().split()))
r = set(el)
print(((sum(r)*k)-(sum(el)))//(k-1))
# ### Ex 13: No Idea!
n, m = input().split()
elms = input().split()
A = set(input().split())
B = set(input().split())
print(sum([(i in A) - (i in B) for i in elms]))
# ## Collections
# ### Ex 1: DefaultDict Tutorial
# +
from collections import defaultdict
n, m = map(int, input().split())
l1 = defaultdict(list)
l2 = []
for i in range(0,n):
l1[input()].append(i+1)
for i in range(0,m):
l2 = l2 + [input()]
for i in l2:
if i in l1:
print(" ".join(map(str,l1[i])))
else:
print(-1)
# -
# ### Ex 2: Collections.namedtuple()
from collections import namedtuple
std = namedtuple('std','Id Name Marks Class')
n, Marks, somm = int(input()), list(input().split()), 0 #I launch the variables
for _ in range(n):
somm += int(list(input().split())[Marks.index('MARKS')])
print(somm/n)
# ### Ex 3: Collections.OrderedDict()
from collections import OrderedDict
D = OrderedDict()
for _ in range(int(input())):
item, space, price = input().rpartition(' ')
D[item] = D.get(item, 0) + int(price)
print(*[" ".join([item, str(price)]) for item, price in D.items()], sep="\n")
# ### Ex 4: Collections.deque()
from collections import deque
n = int(input()) #number of operations
d = deque()
for _ in range(n):
Input = input().split()
if Input[0] == 'append':
d.append(Input[1])
elif Input[0] == 'appendleft':
d.appendleft(Input[1])
elif Input[0] == 'pop':
d.pop()
elif Input[0] == 'popleft':
d.popleft()
print(' '.join(d))
# ### Ex 5: Company Logo
# +
# #!/bin/python3
from operator import itemgetter, attrgetter, methodcaller
import math
import os
import random
import re
import sys
if __name__ == '__main__':
s = input()
md = {}
for i in s:
if i in md:
md[i] += 1
else:
md[i] = 1
l = []
for x, y in md.items(): # .items() returns a generator
# a generator is an object that "creates" one object at a time each time next () is called on it
l.append([x,-y])
l2 = sorted(l, key=itemgetter(1,0))
print(l2[0][0], -l2[0][1])
print(l2[1][0], -l2[1][1])
print(l2[2][0], -l2[2][1])
# -
# ### Ex 6: Pilling Up!
# +
from collections import deque
for _ in range(int(input())):
_, queue =input(), deque(map(int, input().split()))
for cube in reversed(sorted(queue)):
if queue[-1] == cube: queue.pop()
elif queue[0] == cube: queue.popleft()
else:
print('No')
break
else:
print('Yes')
# -
# ### Ex 7: collections.Counter()
import collections
X = int(input()) #n of shoes
earned_money = 0
shoe_size = collections.Counter(map(int, input().split()))
# a Counter creates a collection where elements are stored as dictionary keys and their counts are stored as dictionary values
custom = int(input())
for _ in range(custom):
shoe_wanted, price_shoe = map(int, input().split())
if shoe_size[shoe_wanted]:
earned_money += price_shoe #he earns what he sells
shoe_size[shoe_wanted] -= 1 #when he sells the shoes, he has a pair less
print(earned_money)
# ### Ex 8: Word Order
# +
from collections import OrderedDict
words = OrderedDict()
for _ in range(int(input())):
abc = input()
words.setdefault(abc, 0)
words[abc] += 1
print(len(words))
print(*words.values())
# -
# ## Date and Time
# ### Ex 1: Calendar Module
import calendar
y, m, d = map(int, input().split())
print(calendar.day_name[calendar.weekday(d, y, m)].upper())
# ### Ex 2: Time Delta
# +
# #!/bin/python3
import math
import os
import random
import re
import sys
import calendar
import datetime
# Complete the time_delta function below.
def time_delta(t1, t2):
t1 = datetime.datetime.strptime(t1,'%a %d %b %Y %H:%M:%S %z') #
t2 = datetime.datetime.strptime(t2,'%a %d %b %Y %H:%M:%S %z')
return (str(abs(int((t1-t2).total_seconds()))))
if __name__ == '__main__':
fptr = open(os.environ['OUTPUT_PATH'], 'w')
t = int(input())
for t_itr in range(t):
t1 = input()
t2 = input()
delta = time_delta(t1, t2)
fptr.write(delta + '\n')
fptr.close()
# -
# ## Exceptions
# ### Ex 1: Exceptions
for i in range(int(input())):
try:
a, b = map(int, input().split())
print(a//b)
except ZeroDivisionError as e:
print("Error Code:",e)
except ValueError as vr:
print("Error Code:",vr)
# ## Built-ins
# ### Ex 1: Zipped!
n, x = map(int, input().split()) #n stud, x subj
# creating an empty list
lst = []
# iterating till the range
for _ in range(0, x):
lst.append(map(float, input().split()))
for i in zip(*lst):
print(sum(i)/len(i))
# ### Ex 2: Athlete Sort
# +
# #!/bin/python3
import math
import os
import random
import re
import sys
N,M = map(int,input().split())
rows = [list(map(int,input().split())) for i in range(N)]
i = int(input())
rows = sorted(rows, key=lambda x:x[i])
for i in rows:
print(*i)
if __name__ == '__main__':
nm = input().split()
n = int(nm[0])
m = int(nm[1])
arr = []
for _ in range(n):
arr.append(list(map(int, input().rstrip().split())))
k = int(input())
# -
# ### Ex 3: ginortS
import string as s
print(*sorted(input(), key=(s.ascii_letters + '1357902468').index), sep='')
# ## Python Functionals
# ### Ex 1: Map and Lambda Function
# +
cube = lambda x: x ** 3
def fibonacci(n):
l=[0,1]
for i in range(2,n):
l.append(l[i-2] + l[i-1])
return(l[0:n])
if __name__ == '__main__':
n = int(input())
print(list(map(cube, fibonacci(n))))
# -
# ## Regex and Parsing challenges
# ### Ex 1: Detect Floating Point Number
import re
for _ in range(int(input())):
print(bool(re.match(r'^[-+]?[0-9]*\.[0-9]+$', input())))
# ### Ex 2: Re.split()
# +
from __future__ import print_function
import sys, re
def resplit(text):
return [x for x in re.split(r'[,.]+', text) if len(x) > 0]
def main():
for line in resplit(sys.stdin.read().rstrip()):
print(line)
if __name__ == '__main__':
main()
regex_pattern = r"[.,]" # Do not delete 'r'.
import re
print("\n".join(re.split(regex_pattern, input())))
# -
# ### Ex 3: Group(), Groups() & Groupdict()
#Valid chars . a-z 0-9
import re
s = input()
pattern = re.search(r'([a-z0-9])\1+', s) #'1+' indicates one or more
#search() --> Scan through a string, looking for any location where this RE matches.
if pattern:
print(pattern.group(1))
else:
print('-1')
# ### Ex 4: Re.findall() & Re.finditer()
s = input()
import re
vow = 'AEIOUaeiou'
m = re.compile('[^{0}][{0}][{0}]+[^{0}]'.format(vow),re.I)
pattern = m.search(s)
if not pattern:
print('-1')
while pattern:
print(s[pattern.start()+1:pattern.end()-1]) #removing 1st & last consonant
pattern = m.search(s,pattern.start()+1)
# ### Ex 5: Regex Substitution
from sys import stdin
import re
n = input()
print(re.sub( r"(?<= )(&&|\|\|)(?= )", lambda x: "and" if x.group()=="&&" else "or", stdin.read()))
# ### Ex 6: Validating Roman Numerals
# +
regex_pattern = r"" # Do not delete 'r'.
import re
n_rm = input()
if re.search(r"^(I|V|X|L|C|D|M)+$", n_rm) == None:
print("False")
elif re.search(r"(IIII|XXXX|CCCC|MMMM|VV|LL|DD)", n_rm) != None:
print("False")
else:
print("True")
import re
print(str(bool(re.match(regex_pattern, input()))))
# -
# ### Ex 7: Validatin phone numbers
import re
n = int(input())
for i in range(n):
if re.match(r'[789]\d{9}$', input()): #\d{9} means exactly 9 digits
print ('YES')
else:
print ('NO')
# ### Ex 8: Validating and Parsing Email Addresses
#
#
import re
m = r'^([a-z]+ \<[a-z][-.\w]*@[a-z]+\.[a-z]{1,3}\>)'
for _ in range(int(input())):
match = re.match(m, input(), re.I)
if match:
print(match.group(1))
# ### Ex 9: Hex Color Code
#
#
import re
n = int(input())
m = r'(#[0-9A-Fa-f]{3,6}){1,2}[^\n ]'
for _ in range(n):
for c in re.findall(m, input()):
print(c)
# ### Ex 10: HTML Parser - Part 1
import re
import html
from html.parser import HTMLParser
# create a subclass and override the handler methods
class MyHTMLParser(HTMLParser):
def handle_starttag(self, tag, attrs):
print("Start :", tag)
for name,value in attrs:
print("->",name+" >",value)
def handle_endtag(self, tag):
print ("End :", tag)
def handle_startendtag(self, tag, attrs):
print ("Empty :", tag)
for name,value in attrs:
print("->",name+" >",value)
# instantiate the parser and fed it some HTML
parser = MyHTMLParser()
n = int(input())
for _ in range(n):
parser.feed(input())
# ### Ex 11: HTML Parser - Part 2
# +
from html.parser import HTMLParser
class MyHTMLParser(HTMLParser):
def handle_comment(self, comment):
if '\n' in comment:
print('>>> Multi-line Comment')
else:
print('>>> Single-line Comment')
print(comment)
def handle_data(self, data):
if data == '\n': return
print('>>> Data')
print(data)
html = ""
for i in range(int(input())):
html += input().rstrip()
html += '\n'
parser = MyHTMLParser()
parser.feed(html)
parser.close()
# -
# ### Ex 12: Detect HTML Tags, Attributes and Attribute Values
#
#
from html.parser import HTMLParser
class Parser(HTMLParser):
def handle_starttag(self, tag, attrb):
print(tag)
for attr in attrb:
prop,value = attr
print("-> " + prop + " > " + value)
n=int(input())
s=""
for i in range(n):
s+= " " + input()
parser = Parser()
parser.feed(s)
# ### Ex 13: Validating UID
import re
t = int(input())
for i in range(t):
if re.search(r'^(?!.*(.).*\1)(?=(?:.*[A-Z]){2,})(?=(?:.*\d){3,})[a-zA-Z0-9]{10}$', input()):
print ('Valid')
else:
print ('Invalid')
# ### Ex 14: Validating Credit Card Numbers
#
#
import re
m = re.compile(
r"^"
r"(?!.*(\d)(-?\1){3})"
r"[456]"
r"\d{3}"
r"(?:-?\d{4}){3}"
r"$")
for _ in range(int(input().strip())):
if m.search(input().strip()):
print("Valid")
else:
print("Invalid")
# ### Ex 15: Validating Postal Codes
# +
regex_integer_in_range = r"^[1-9][\d]{5}$" # Do not delete 'r'.
regex_alternating_repetitive_digit_pair = r"(\d)(?=\d\1)" # Do not delete 'r'.
import re
P = input()
print (bool(re.match(regex_integer_in_range, P))
and len(re.findall(regex_alternating_repetitive_digit_pair, P)) < 2)
# -
# ### Ex 16: Matrix Script
# +
import sys
import re
N, M = map(int, sys.stdin.readline().split())
rows = [sys.stdin.readline()[:M] for i in range(N)]
cols = [''.join([rows[i][j] for i in range(N)]) for j in range(M)]
decode = ''.join(cols)
print (re.sub('([0-9a-zA-Z])[^0-9a-zA-Z]+([0-9a-zA-Z])', '\g<1> \g<2>', decode))
# -
# ### Ex 17: Re.split()
# +
from __future__ import print_function
import sys, re
def resplit(text):
return [x for x in re.split(r'[,.]+', text) if len(x) > 0]
def main():
for line in resplit(sys.stdin.read().rstrip()):
print(line)
if __name__ == '__main__':
main()
regex_pattern = r"[.,]"
import re
print("\n".join(re.split(regex_pattern, input())))
# -
# ## XML
# ### Ex 1: XML 1 - Find the Score
#
#
# +
import sys
import xml.etree.ElementTree as etree
def get_attr_number(node):
return etree.tostring(node).count(b'=') #per eseguire il cast di oggetti di diverso tipo in stringhe
if __name__ == '__main__':
sys.stdin.readline()
xml = sys.stdin.read()
tree = etree.ElementTree(etree.fromstring(xml))
root = tree.getroot()
print(get_attr_number(root))
# -
# ### Ex 2: XML2 - Find the Maximum Depth
# +
import xml.etree.ElementTree as etree
maxdepth = 0
def depth(elem, level):
global maxdepth
level += 1
if level >= maxdepth:
maxdepth = level
for child in elem:
depth(child, level)
if __name__ == '__main__':
n = int(input())
xml = ""
for i in range(n):
xml = xml + input() + "\n"
tree = etree.ElementTree(etree.fromstring(xml))
depth(tree.getroot(), -1)
print(maxdepth)
# -
# ## Closures and Decorations
# ### Ex 1: Standardize Mobile Number Using Decorators
#
#
# +
def wrapper(f):
def fun(l):
f(["+91 "+c[-10:-5]+" "+c[-5:] for c in l])
# 6 characters counting from the end
# last 5 characters of the phone number
return fun
@wrapper
def sort_phone(l):
print(*sorted(l), sep='\n')
if __name__ == '__main__':
l = [input() for _ in range(int(input()))]
sort_phone(l)
# -
# ### Ex 2: Decorators 2 - Name Directory
# +
import operator
def et(n):
return (int(n[2]))
def person_lister(f):
def inner(people):
return (map(f, sorted(people, key=et)))
return inner
@person_lister
def name_format(person):
return ("Mr. " if person[3] == "M" else "Ms. ") + person[0] + " " + person[1]
if __name__ == '__main__':
people = [input().split() for i in range(int(input()))]
print(*name_format(people), sep='\n')
# -
# ## Numpy
# ### Ex 1: Arrays
# +
import numpy
def arrays(arr):
ris = numpy.array(arr, float)
ris_rev = ris[::-1]
return (ris_rev)
arr = input().strip().split(' ')
result = arrays(arr)
print(result)
# -
# ### Ex 2: Shape and Reshape
# +
import numpy
arr = input().strip().split(' ')
arr = numpy.array(arr, int)
arr.shape = (3, 3)
print(arr)
# -
# ### Ex 3: Transpose and Flatten
# +
import numpy
n, m = map(int, input().split())
array = numpy.array([input().strip().split() for _ in range(n)], int)
print (array.transpose())
print (array.flatten())
# -
# ### Ex 4: Concatenate
# +
import numpy
n, m, p = map(int, input().split())
array_n = numpy.array([input().strip().split() for _ in range(n)], int)
array_m = numpy.array([input().strip().split() for _ in range(m)], int)
print(numpy.concatenate((array_n, array_m), axis = 0))
# -
# ### Ex 5: Zeros and Ones
# +
import numpy
nmr = list(map(int, input().split()))
print(numpy.zeros((nmr), dtype = numpy.int))
print(numpy.ones((nmr), dtype = numpy.int))
# -
# ### Ex 6: Eye and Identity
# +
import numpy
numpy.set_printoptions(legacy='1.13')
n,m = map(int, input().split())
print(numpy.identity(n))
# -
# ### Ex 7: Array Mathematics
# +
import numpy
n, m = map(int, input().split())
a = numpy.array([input().strip().split() for _ in range(n)], int)
b = numpy.array([input().strip().split() for _ in range(n)], int)
print(numpy.add(a, b))
print(numpy.subtract(a, b))
print(numpy.multiply(a, b))
print(a // b )
print(numpy.mod(a, b))
print(numpy.power(a, b))
# -
# ### Ex 8: Floor, Ceil and Rint
# +
import numpy
numpy.set_printoptions(legacy='1.13')
arr = input().strip().split(' ')
my_array = numpy.array(arr, float)
print(numpy.floor(my_array))
print(numpy.ceil(my_array))
print(numpy.rint(my_array))
# -
# ### Ex 9: Sum and Prod
#
#
# +
import numpy
n, m = map(int, input().split())
my_array = numpy.array([input().strip().split() for _ in range(n)], int)
a2 = numpy.sum(my_array, axis = 0)
print(numpy.prod(a2))
# -
# ### Ex 10: Min and Max
# +
import numpy
n, m = map(int, input().split())
my_array = numpy.array([input().strip().split() for _ in range(n)], int)
finder = numpy.min(my_array, axis = 1)
print(numpy.max(finder))
# -
# ### Ex 11: Mean, Var and Std
import numpy
n, m = map(int, input().split())
my_array = numpy.array([input().strip().split() for _ in range(n)], float)
print(numpy.mean(my_array, axis = 1))
print(numpy.var(my_array, axis = 0))
a = numpy.std(my_array)
b = round(a,11)
print(b)
# ### Ex 12: Dot and Cross
# +
import numpy
n = int(input())
A = numpy.array([input().strip().split() for _ in range(n)], int)
B = numpy.array([input().strip().split() for _ in range(n)], int)
print(numpy.dot(A, B))
# -
# ### Ex 13: Inner and Outer
# +
import numpy
A = numpy.array(input().split(), int)
B = numpy.array(input().split(), int)
print(numpy.inner(A,B), numpy.outer(A,B), sep='\n')
# -
# ### Ex 14: Polynomials
# +
import numpy
my_array = numpy.array(input().split(), float)
x = int(input())
print(numpy.polyval(my_array, x))
# -
# ### Ex 15: Linear Algebra
# +
import numpy
n = int(input())
A = numpy.array([input().strip().split() for _ in range(n)], float)
det2 = round(numpy.linalg.det(A),2)
print(det2)
| HW1_LetiziaRusso_Problem1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/siddhantkushwaha/palladium/blob/master/samples/Palladium.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="CTuVyABOWnD8" colab_type="code" colab={}
# ! pip install palladium-python
# + pycharm={"name": "#%%\n"}
from palladium.chrome_custom import ChromeCustom
# + id="Lt3rZrY_Wrh8" colab_type="code" colab={}
chrome = ChromeCustom(headless=True)
# + id="Y2R5Zo0wYqDJ" colab_type="code" colab={}
chrome.get('https://github.com/siddhantkushwaha')
chrome.save_screenshot('/content/test.png')
# + id="zXHENxAPZr3f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6108728d-e7db-46ac-a1ed-094c913df0e3" pycharm={"name": "#%%\n"}
from IPython.display import Image
Image('test.png')
# + pycharm={"name": "#%%"}
| samples/Palladium.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pytorch-env]
# language: python
# name: conda-env-pytorch-env-py
# ---
# # Expander Dictionary Learning
# In this notebook we provide experimental results pertaining to the paper XXX, in which an algorithm to solve a particular dictionary learning problem is proposed. To be clear, the problem is given a matrix $Y$ of dimension $m \times N$, can we recover the dictionary $A$ and latent representation $X$ of the data under the model $Y = AX$? To make this problem tractable we assume the following,
# - $A$ is a binary matrix of dimension $m \times n$ and is the adjacency matrix of a $k, \epsilon , d$ bipartite expander graph.
# - $X$ is a sparse, real matrix whose columns are $k$ sparse (have exactly k non-zeros) and are dissociated, meaning that for any column $supp(x_i) = S$, then for any $T_1, T_2 \subset S$, $T_1 \neq T_2$, it holds that $\sum_{j \in T_1} x_{ji} \neq \sum_{j \in T_2} x_{ji} $.
#
# In the paper linked an algorithm is proposed to solve this dictionary learning problem, up to some permutation of the columns of $A$ / rows of $X$, based on the insight that partial supports of the columns of $A$ can be identified and recovered from the columns of $Y$. This is possible due to the the unique neighbour property which $k, \epsilon , d$ bipartite expander graphs satisfy.
# ## Setup
# ### Functions for generating data
# Firstly we import the libraries we will need and also define the functions we will use to generate the data for our experiments. The functions for generating and checking $A$ and $X$ are as follows,
# - **generate_latent** produces a k sparse vector x with non-zeros drawn uniformly from the set $[-c_2, -c_1] \cup [c_1, c_2]$.
# - **generate_exp_dictionary** returns a binary matrix which, under certain parameter regimes, is the adjacency matrix of an unbalanced fixed degree expander graph with high probability. Each column's support is chosen uniformly from random from the set of all subsets of cardinality $d$ from $[m]$. Note that the algorithms we propose can still work when $A$ is only approximatly an expander.
# - **generate_exp_dictionary2** is similar to generate_exp_dictionary but generates a dictionary heuristically more likely to be the adjacency matrix of an unbalanced fixed degree expander graph. This is created by creating a random permutation of the row indicces and assigning the first d to the first column, the next set of d to the 2nd etc. Once this permutation has been used up repeat until all columns of A are constructed.
# - **check_dictionary** checks that a matrix A satisfies a necessary condition for it to be the adjacency matrix of a fixed left degree $k, \epsilon, d$ bipartite expander graph. This condition ensures that no two columns overlap by at most $2\epsilon d$. Note this condition is not a sufficient condition.
# - ** check_recon_A** checks the final reconstruction of A against the true A to see if there are any errors and or missing entries.
# - ** check_recon_X** checks the final reconstruction of X against the true X to see if there are any errors and or missing entries. This function uses the permutation found when A is checked, so one need to runs ** check_recon_A** first.
# +
# IMPORTS
import numpy as np
import copy
import matplotlib.pyplot as plt
from timeit import default_timer as timer
# DATA GENERATION AND PERFORMANCE CHECKING FUNCTIONS
def generate_latent(n,k,c1,c2):
b = c2-c1
a = c1/b
x = np.zeros(n)
locs = np.random.permutation(n)[:k]
x[locs] = 1
signs = np.sign(np.random.randn(k))
vals = b*(np.random.rand(k)+a)
x[locs] = x[locs]*vals*signs
return x
def generate_exp_dictionary1(m,n,d):
A = np.zeros((m,n))
for i in range(n):
locs = np.random.permutation(m)[:d]
A[locs,i] = 1
return A
def generate_exp_dictionary2(m,n,d):
A = np.zeros((m,n))
alpha = int(np.floor(m/d))
c = 0
while c<n:
perm = np.random.permutation(m)
zeta = min(n-c, alpha)
for j in range(zeta):
locs = perm[j*d:(j+1)*d]
A[locs,c] = 1
c = c+1
return A
def check_dictionary(eps,n,d,A):
t = (1-4*eps)*d
mu_A = np.max(np.transpose(A)@A - d*np.identity(n))
if mu_A >= t:
print("Generation error, A is not expander for this value of epsilon, mu_A = ", str.format('{0:.2f}', mu_A),", tau = ", str.format('{0:.2f}', t), ".")
passed = False
else:
print("A passes coherence test, mu_A = ", str.format('{0:.2f}', mu_A),", tau = ", str.format('{0:.2f}', t), ".")
passed = True
return passed
def check_recon_A(A,A_rec,d,eps,n):
print("Checking reconstruction of A.")
col_sum = sum(A_rec)
print(str.format('{0:.1f}',100*len(col_sum[col_sum==d])/n), "% of columns fully formed.")
print(str.format('{0:.1f}',100*(n - len(col_sum[col_sum==d]) - len(col_sum[col_sum==0]))/n) , "% of columns partially formed.")
print(str.format('{0:.1f}', 100*len(col_sum[col_sum==0])/n), "% of columns have no entries.")
overlaps = np.transpose(A_rec)@A
A_rec_col, A_col = np.where(overlaps>(1-4*eps)*d)
if len(col_sum[col_sum==0])<n:
print(str.format('{0:.1f}', 100*len(A_rec_col)/(n - len(col_sum[col_sum==0]))), "% of reconstructed (fully or partially) columns match with a column in target matrix.")
error = 0
missing = 0
P = np.zeros((n,n))
for i in range(len(A_rec_col)):
diff = A_rec[:, A_rec_col[i]] - A[:,A_col[i]]
error = error + sum(abs(diff[diff==1]))
missing = missing + sum(abs(diff[diff==-1]))
P[A_col[i], A_rec_col[i]] = 1
print(str.format('{0:.0f}', 100*(missing+d*n-len(A_rec_col)*d)/(d*n)), "% of entries missing in total.")
print(str.format('{0:.0f}', missing), "entries missing in matched columns.")
print(str.format('{0:.0f}', error), "entries wrong in matched columns.")
return P
def check_recon_X(X,X_rec,P,N,k):
X_perm = P@X_rec
print("Checking the values of X recovered.")
tol = 10**-6
total_entries = N*k
total_entries_recovered = 0
correct_loc_correct_val = 0
correct_loc_wrong_val = 0
wrong_loc = 0
missing = 0
passed = True
for i in range(N):
for j in range(n):
if abs(X[j,i])>tol and abs(X_perm[j,i])>tol:
error = abs(X[j,i] - X_perm[j,i])
if error < tol:
correct_loc_correct_val +=1
else:
correct_loc_wrong_val += 1
print("Column number = ", i, ", row number = ", j, ", true value = ", X[j,i], "value found = ", X_perm[j,i])
elif abs(X[j,i])>tol and abs(X_perm[j,i])<tol:
missing += 1
elif abs(X[j,i])<tol and abs(X_perm[j,i])>tol:
wrong_loc += 1
print(str.format('{0:.1f}', 100*(correct_loc_correct_val/total_entries)), "% of entries correctly recovered.")
print(str.format('{0:.1f}', 100*(correct_loc_wrong_val/total_entries)), "% of entries in correct location but have wrong value for location.")
print(str.format('{0:.1f}', 100*(missing/total_entries)), "% of entries missing.")
return X_perm
# -
# ### Auxillary functions
# As previously stated, we exploit the unique neighbour property of expander graphs to extract partial supports of the columns of $A$ and then cluster these partial supports to reconstruct each column. The following 'auxillary' encapsulate this idea.
# - **xps** extracts all partial supports of columns of $A$ from $y$. It returns both the partial supports in $W$ as well as the value of the coefficients in $vals$.
# For experiment 1, the demonstration of $A$ can be reconstructed from partial supports extracted from the columns of $Y$ there are the following two auxillary functions.
# - **extract_supports** is used in experiment 1, this function process a data point by first extracting any partial supports, then matches these partial supports to a dictionary element before finally updating said dictionary elements.
# - **update_column** updates the reconstruction of a column with the information gathered from a partial support of that column.
# For experiment 2, recovering $A$ and $X$ from $Y$, we have the following three auxillary functions.
# - **process_column** takes a column of $Y$, extracts partial supports and returns them along with the column index of $A$ which the partial support matches to, the index the column (data id), the value or coefficient of the partial measurement and finally whether or not the partial support matches with an already partially reconstructed column of $A$.
# - **sort_unmatched_columns** takes the partial supports extracted from all columns of $Y$ that do not match with a partially reconstructed column of A and clusters them.
# - **update_reconstruction** takes all the partial supports extracted during an epoch and uses them to update the reconstruction of $A$ and $X$.
# +
# USED IN BOTH EXPERIMENTS 1 and 2
def xps(y,m,eps,d,k):
tol1 = 10**-8
tol2 = 10**-8
y_copy = copy.deepcopy(y)
t = (1-2*eps)*d
z1 = len(y_copy);
z2 = int(z1 - np.ceil(t) +1);
W = np.zeros((m,k))
vals = np.zeros(k)
q = 0
for i in range(z2):
if np.abs(y_copy[i])> tol1:
locs = list()
locs.append(i)
for j in range(i+1, m):
if np.abs(y_copy[j] - y_copy[i]) < tol2:
locs.append(j)
y_copy[j] = 0
if len(locs)>t:
W[locs,q] = 1
vals[q] = y_copy[i]
q = q+1
y_copy[i] = 0
W = W[:,:q]
vals = vals[:q]
return [W, vals, q]
# AUXILLARY FUNCTIONS USED IN EXPERIMENT 1: DEMONSTRATING HOW SUPPORTS OF A CAN BE EXTRACTED FROM Y
def extract_supports(y, A_rec, m, eps, d, k, c, per):
tau = (1-4*eps)*d
[W, vals, q] = xps(y,m,eps,d,k)
if q > 0:
if c == 0:
A_rec[:,:q] = W
c = q
else:
matches = np.transpose(W)@A_rec[:,:c]
matches[matches<=tau] = 0
matches[matches>tau] = 1
num_matches = matches.sum(axis=1)
col_ind = matches@np.arange(c)
for i in range(q):
if num_matches[i] == 0:
if c<n:
old = sum(A_rec[:,c])
A_rec[:,c]=update_column(A_rec[:,c],W[:,i])
per = per + sum(A_rec[:,c]) - old
c=c+1
else:
print("Error in extract_supports: trying to add new column which exceeds total number of columns")
elif num_matches[i] == 1:
z = int(col_ind[i])
old = sum(A_rec[:,z])
A_rec[:,z]=update_column(A_rec[:,z],W[:,i])
per = per + sum(A_rec[:,z]) - old
elif num_matches[i] > 1:
print("Error in extract_supports: partial support matches with ", num_matches," partially reconstructed columns of A.")
# else:
# # print("Error in extract_supports: xps returned no partial supports for this data point.")
return [A_rec, c, per]
def update_column(a,w):
a = a+w
a[a>0] = 1
return a
# AUXILLARY FUNCTION USED IN EXPERIMENT 2: RECOVERING A AND X FROM Y
def process_column(y, A_rec, m, eps, d, k, c, dpid):
exit_flag = True
tau = (1-4*eps)*d
[W, vals, q] = xps(y,m,eps,d,k)
if q > 0:
match_check = np.zeros(q)
match_col_id = -0.5*np.ones(q)
data_id = dpid*np.ones(q)
if c > 0:
matches = np.transpose(W)@A_rec
matches[matches<=tau] = 0
matches[matches>tau] = 1
num_matches = matches.sum(axis=1)
col_ind = matches@np.arange(c)
for i in range(q):
if num_matches[i] == 0:
if c>=n:
print("Error: trying to add new column which exceeds total number of columns")
exit_flag==False
elif num_matches[i] == 1:
match_check[i] = 1
z = int(col_ind[i])
match_col_id[i] = z
elif num_matches[i] > 1:
print("Error: partial support matches with ", num_matches," partially reconstructed columns of A.")
exit_flag = False
else:
exit_flag = False
match_check = []
match_col_id = []
data_id = []
return [W, match_check, match_col_id, data_id, vals, exit_flag]
def sort_unmatched_columns(W, eps, d, c):
q = W.shape[1]
col_id_new = -0.5*np.zeros(q)
tau = (1-4*eps)*d
matches = np.transpose(W)@W
matches[matches<=tau] = 0
matches[matches>tau] = 1
i =0
while i<q:
if sum(matches[:,i]) > 0:
temp = np.where(matches[:,i]==1)[0]
col_id_new[temp] = c
c +=1
for j in range(len(temp)):
matches[temp[j],:] = np.zeros(q)
matches[:,temp[j]] = np.zeros(q)
i +=1
return [col_id_new, c]
def update_reconstruction(W, col_id, data_id, vals, A_rec, X_rec):
for i in range(len(col_id)):
A_rec[:, int(col_id[i])] += W[:,i]
X_rec[int(col_id[i]), int(data_id[i])] = vals[i]
A_rec[A_rec>0] = 1
return [A_rec, X_rec]
# -
# # Experiments
# ### Generating a valid dictionary
# We firstly generate a dictionary to use in the following experiments. Note that the algorithm can succeed in recovering $A$ even if $A$ is not truly the bipartite of an expander graph. However if A is not an expander it is not guaranteed that the algorithm will recover X, due to the fact that the unique neighbour property may not hold. This means that columns of $Y$, or the residue of $Y$, $R = Y - A_{rec} X_{rec}$, may not have any partial supports that can extracted since the respective columns of $A$ are to overlapping. Checking that $A$ is indeed an expander graph is NP hard since one is required to check that all $\binom{n}{k}$ subsets of the columns satisfy the unique neighbour property. Hence we do not know the true value of $\epsilon$!
# +
# Dictionary parameters parameters
#n = 1200;
#m = int(np.ceil(4*n/5));
#d = 9;
#eps = 1/6;
#N = 500
n = 5000;
m = int(np.ceil(4*n/5));
d = 10;
eps = 1/6;
N = 500
# Generate a dictioary and check that passes necessary test.
num_generations = 0
passed = False
while passed==False and num_generations < 100:
A = generate_exp_dictionary1(m,n,d);
passed = check_dictionary(eps,n,d,A)
num_generations += 0
if passed == True:
print("Generated dictionary that passes the coherency test")
else:
print("Failed to generate dictionary that passes the coherency test, consider different parameters")
# -
# ## Dictionary Statistics
row_count = np.sum(A, 1)
print(A.shape)
print(sum(sum(A)))
print(min(row_count))
print(max(row_count))
print(np.mean(row_count))
print(n*d/m)
# +
num_bins = int(max(row_count) - min(row_count))
freq, bins, patches = plt.hist(row_count, bins=num_bins, color='#0504aa',
alpha=0.7, rwidth=0.85)
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Number of non-zeros')
plt.ylabel('Frequency')
plt.title('Row density of A with n=%s'%(n))
maxfreq = freq.max()
# +
# FOR COMPARISON AGAINST IID BERNOULLI ENTRIES
B = np.random.binomial(1, d/m, size=(m,n))
print(B.shape)
print(sum(sum(B)))
row_count_B = np.sum(B, 1)
num_bins_B = int(max(row_count_B) - min(row_count_B))
freq, bins, patches = plt.hist(row_count_B, bins=num_bins_B, color='#0504aa',
alpha=0.7, rwidth=0.85)
plt.grid(axis='y', alpha=0.75)
plt.xlabel('Number of non-zeros')
plt.ylabel('Frequency')
plt.title('Row density of B with n=%s'%(n))
maxfreq = freq.max()
# -
# ## Experiment 1 - Demonstrating extraction of partial supports
# In this experiment we demonstrate how we can extract partial supports to reconstruct the columns of A. In this experiment the algorithm is presented with a stream of columns of $Y$, which one could think of as a stream of data points, the objective being to reconstruct $A$ using the minimum number of data points. Hence for a fixed dictionary $A$, we sequentially generate latent representations of the data (i.e., columns the in X) with a given sparsity, generate a data point (column of $Y$), pass this to the algorithm and record the total number of entries of $A$ that have been recovered.
#
# Note that as already stated the it is NP hard to check that $A$ is an expander, regardless the algorithm can still succeed in recovering $A$ so long as enough submatrices satisfy the unique neighbour property condition. This does mean however that some data points may not return any partial supports, this is increasingly likely for larger $k$.
# +
b1=1;
b2=5;
k = [int(np.ceil(0.02*n)), int(np.ceil(0.04*n)), int(np.ceil(0.06*n))]
limit = N
if passed == True:
per1=np.zeros((len(k), limit));
comp_time1 = np.zeros((len(k), limit))
done = False
for j in range(len(k)):
print("Solving problem with k/n=", str.format('{0:.2f}', 100*k[j]/n),"%.")
A_rec1 = np.zeros((m,n))
c1=0
entry_total1 = 0
total_comp_time1 = 0
N=0
while done == False and N < limit:
x = generate_latent(n,k[j],b1,b2)
y = A@x
if done == False:
start_time = timer()
[A_rec1, c1, entry_total1] = extract_supports(y, A_rec1, m, eps, d, k[j], c1, entry_total1)
end_time = timer()
total_comp_time1 = total_comp_time1 + (end_time - start_time)
comp_time1[j,N] = total_comp_time1
per1[j, N] = entry_total1
if entry_total1 >= n*d:
comp_time1 = comp_time1[:N+1]
per1[j, (N+1):limit] = n*d*np.ones(limit - N - 1)
done = True
print("Finished reconstructing the dictionary.")
N = N + 1
if np.mod(N, limit/10)==0:
print(str.format('{0:.2f}', 100*N/limit), "% of max data points processed.")
if N == limit:
print("Max number of data points reached, checking reconstruction...")
print("Finished attempts to solve reconstruction problem with k/n =",str.format('{0:.2f}', 100*k[j]/n),"%%.")
print("Checking reconstruction of A with k/n=", str.format('{0:.2f}', 100*k[j]/n),"%.")
P = check_recon_A(A,A_rec1,d,eps,n)
# +
# Plots
plot_per1 = 100*per1/(d*n)
rel_comp_time1 = comp_time1
for j in range (len(k)):
rel_comp_time1[j,:] = 100*rel_comp_time1[j,:]/comp_time1[j, N-1]
plt.figure(1)
plt.figure(figsize=(20,10))
plt.plot(range(N), plot_per1[0], 'g', label="k/n=%s %%"%(100*k[0]/n))
plt.plot(range(N), plot_per1[1], 'b', label="k/n=%s %%"%(100*k[1]/n))
plt.plot(range(N), plot_per1[2], 'r', label="k/n=%s %%"%(100*k[2]/n))
plt.grid(True)
plt.title('Recovery of dictionary entries with N fixed as %s'%(N))
plt.xlabel('N - # of data points')
plt.ylabel('Percentage of entries of A recovered')
plt.legend()
plt.savefig('./figures/recovery_curve_N.eps', format='eps', bbox_inches='tight')
plt.figure(2)
plt.figure(figsize=(20,10))
plt.plot(rel_comp_time1[0], plot_per1[0], 'g', label="SECER k/n=%s %%"%(100*k[0]/n))
plt.plot(rel_comp_time1[1], plot_per1[1], 'b', label="SECER k/n=%s %%"%(100*k[1]/n))
plt.plot(rel_comp_time1[2], plot_per1[2], 'r', label="SECER k/n=%s %%"%(100*k[2]/n))
plt.grid(True)
plt.title('Recovery of dictionary entries with N fixed as %s'%(N))
plt.xlabel('Computational time as percentage of total running time (%)')
plt.ylabel('Percentage of entries of A recovered (%)')
plt.legend()
plt.savefig('./figures/recovery_curve_time.eps', format='eps', bbox_inches='tight')
# -
# ### Remarks
# As the ratio of $k/n$ is increased the rate at which the algorithm is able to extract partial supports and hence 'learn' $A$ decreases. This is inline with our intuition since a larger $k$ means that more columns are being added together to form a column of $Y$, thereby incurring a higher risk of overlap and hence occlusion of entries from potential partial supports.
#
# Secondly in terms of computational running time the rate at which entries of $A$ are recovered is logarithmic in the number of data points. This also agrees with our intuition since the rate at which we observe partial supports for new or only partially completed should be akin to the waiting time for a coupon collector problem.
# ## Experiment 2 - recovering $A$ and $X$ from $Y$
# In this experiment the challenge is, given a block of data $Y$, to recover $A$ and $X$ under the assumptions previously stated. We now demonstrate that this can be achieved by iteratively passing through the data sequentially or indeed in a parallelised fashion, each time extracting partial supports, updating the reconstructions of $A$ and $X$ and before updating the residual $R=Y - A_{rec}X_{rec}$ and then repeating. Algorithm proceeds until one of the following three conditions has been met,<br>
# 1) Both $A$ and $X$ have been recovered.<br>
# 2) The algorithm stops making progress (implying that $A$ is not an expander and/or $N$ is too small).<br>
# 3) The maximum number of allowed epochs (computational budget) has been reached.<br>
# GENERATE LATENET REPRESENTATION
#N = 500
b1=1;
b2=5;
#k = [int(np.ceil(0.02*n))]#, int(np.ceil(0.05*n))]
X = np.zeros((len(k), n, N))
for j in range(len(k)):
for i in range(N):
X[j,:,i] = generate_latent(n,k[j],b1,b2)
# +
# LEARN DICTIONARY
max_epoch = 40
tol = 10**-8
X_sparsity = np.zeros(len(k))
total_sparsity = np.zeros(len(k))
Y_frob = np.zeros(len(k))
comp_time = np.zeros(len(k))
number_k_to_process = len(k)
if passed == True:
A_sparsity = np.count_nonzero(A)
A_rec = np.zeros((len(k),m,n))
X_rec = np.zeros((len(k),n,N))
frob_error_series = np.ones((len(k), max_epoch))
A_entries = np.zeros((len(k), max_epoch))
X_entries = np.zeros((len(k), max_epoch))
total_entries = np.zeros((len(k), max_epoch))
for j in range(number_k_to_process):
print("Solving dictionary learning problem with k/n = ", str.format('{0:.1f}', 100*k[j]/n), '%, N = ', str.format('{0:.0f}', N))
X_sparsity[j] = k[j]*N
total_sparsity[j] = A_sparsity + X_sparsity[j]
Y = A@X[j]
R = copy.deepcopy(Y)
Y_frob[j] = np.sqrt(sum(sum(Y**2)))
num_entries = 0
epoch = 0
c=0
exit_program = False
frob_error = Y_frob[j]
start_time = timer()
while epoch < max_epoch and frob_error > tol and exit_program==False:
# Serial case so use a for loop to iterate through the data.
num_matched = 0
num_new = 0
for i in range(N):
[W, match_check, match_col_id, data_id, vals, supports_found]=process_column(R[:,i], A_rec[j,:,:c], m, eps, d, k[j], c, i)
matches = np.where(match_check==1)[0]
new = np.where(match_check==0)[0]
if supports_found == True:
if len(matches)>0:
if num_matched > 0:
W_match = np.concatenate((W_match, W[:,matches]), axis=1)
col_id_match = np.concatenate((col_id_match, match_col_id[matches]), axis=0)
data_id_match = np.concatenate((data_id_match, data_id[matches]), axis=0)
val_match = np.concatenate((val_match, vals[matches]), axis=0)
else:
W_match = W[:,matches]
col_id_match = match_col_id[matches]
data_id_match = data_id[matches]
val_match = vals[matches]
num_matched += W_match.shape[1]
if len(new)>0:
if num_new > 0:
W_new = np.concatenate((W_new, W[:,new]), axis=1)
data_id_new = np.concatenate((data_id_new, data_id[new]), axis=0)
val_new = np.concatenate((val_new, vals[new]), axis=0)
else:
W_new = W[:,new]
data_id_new = data_id[new]
val_new = vals[new]
num_new += W_new.shape[1]
if num_matched > 0 and num_new > 0:
[col_id_new, c] = sort_unmatched_columns(W_new, eps, d, c)
W_all = np.concatenate((W_match, W_new), axis=1)
col_id_all = np.concatenate((col_id_match, col_id_new), axis=0)
data_id_all = np.concatenate((data_id_match, data_id_new), axis=0)
val_all = np.concatenate((val_match, val_new), axis=0)
elif num_matched > 0 and num_new == 0:
W_all = W_match
col_id_all = col_id_match
data_id_all = data_id_match
val_all = val_match
elif num_matched == 0 and num_new > 0:
[col_id_all, c] = sort_unmatched_columns(W_new, eps, d, c)
W_all = W_new
data_id_all = data_id_new
val_all = val_new
elif num_matched == 0 and num_new == 0:
print("No partial supports recovered, terminating algorithm")
frob_error_series[j, epoch:] = frob_error
A_entries[j,epoch:] = A_entries[j,epoch-1]
X_entries[j,epoch:] = X_entries[j,epoch-1]
total_entries[j, epoch:] = total_entries[j, epoch-1]
exit_program = True
if exit_program == False:
[A_rec[j], X_rec[j]] = update_reconstruction(W_all, col_id_all, data_id_all, val_all, A_rec[j], X_rec[j])
R = Y - A_rec[j]@X_rec[j]
frob_error = np.sqrt(sum(sum(R**2)))
frob_error_series[j, epoch] = frob_error
A_entries[j,epoch] = np.count_nonzero(A_rec[j])
X_entries[j,epoch] = np.count_nonzero(X_rec[j])
total_entries[j, epoch] = A_entries[j,epoch] + X_entries[j,epoch]
epoch +=1
print('Epoch ', str.format('{0:.0f}', epoch), " processed, l_2 error = ", str.format('{0:.1f}', 100*frob_error/Y_frob[j]), '%, l_0 error =', str.format('{0:.1f}', 100*(total_sparsity[j] - total_entries[j,epoch-1])/total_sparsity[j]), "%.")
if epoch == max_epoch:
print("Maximum number of epochs reached.")
end_time = timer()
comp_time[j] = end_time - start_time
print("")
P = check_recon_A(A,A_rec[j],d,eps,n)
print("")
check_recon_X(X[j],X_rec[j],P,N,k[j])
print("")
# +
# Plots
plt.figure(1).clear()
plt.figure(2).clear()
plot_A_entries = 100*A_entries/A_sparsity
plot_X_entries = np.zeros((len(k), max_epoch))
plot_total_entries = np.zeros((len(k), max_epoch))
plot_frob = np.zeros((len(k), max_epoch))
for j in range(number_k_to_process):
plot_frob[j,:] = 100*frob_error_series[j,:]/Y_frob[j]
plot_X_entries[j,:] = 100*X_entries[j,:]/X_sparsity[j]
plot_total_entries[j,:] = 100*total_entries[j,:]/total_sparsity[j]
plot_epoch = 6
print("Running times:")
print("k/n=%s %%, %s s"%(100*k[0]/n, comp_time[0]))
print("k/n=%s %%, %s s"%(100*k[1]/n, comp_time[1]))
print("k/n=%s %%, %s s"%(100*k[2]/n, comp_time[2]))
plt.figure(1)
plt.figure(figsize=(20,10))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[0], 'g-', label="A, k/n=%s %%"%(100*k[0]/n))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[0], 'g--',label="X, k/n=%s %%"%(100*k[0]/n))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[1], 'b-', label="A, k/n=%s %%"%(100*k[1]/n))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[1], 'b--',label="X, k/n=%s %%"%(100*k[1]/n))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[2], 'r-', label="A, k/n=%s %%"%(100*k[2]/n))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[2], 'r--',label="X, k/n=%s %%"%(100*k[2]/n))
plt.grid(True)
plt.title('Percentage of entries of A and X Recovered with N = %s'%(N))
plt.xlabel('Epochs')
plt.ylabel('Percentage of entries of A and X recovered')
plt.xlim(1,plot_epoch)
plt.legend()
plt.savefig('./figures/A_X_recovery.eps', format='eps', bbox_inches='tight')
plt.figure(2)
plt.figure(figsize=(20,10))
plt.plot(np.arange(max_epoch)+1, plot_frob[0], 'g-', label="k/n=%s %%"%(100*k[0]/n))
plt.plot(np.arange(max_epoch)+1, plot_frob[1], 'b-', label="k/n=%s %%"%(100*k[1]/n))
plt.plot(np.arange(max_epoch)+1, plot_frob[2], 'r-', label="k/n=%s %%"%(100*k[2]/n))
plt.title('Frobenius error percentage of Y-AX with N fixed as %s'%(N))
plt.grid(True)
plt.xlabel('Epochs')
plt.ylabel('Frobenius error percentage of Y-AX')
plt.xlim(1,plot_epoch)
plt.legend()
plt.savefig('./figures/frob_error.eps', format='eps', bbox_inches='tight')
# -
plt.figure(1)
plt.figure(figsize=(15,20))
plt.subplot(2,2,1)
plt.plot(range(N), plot_per1[0], 'g', label="k/n=%s %%"%(100*k[0]/n))
plt.plot(range(N), plot_per1[1], 'b', label="k/n=%s %%"%(100*k[1]/n))
plt.plot(range(N), plot_per1[2], 'r', label="k/n=%s %%"%(100*k[2]/n))
plt.grid(True)
plt.title('Recovery of dictionary entries with N fixed as %s'%(N))
plt.xlabel('N - # of data points')
plt.ylabel('Percentage of entries of A recovered')
plt.legend()
plt.subplot(2,2,2)
plt.plot(np.arange(max_epoch)+1, plot_A_entries[0], 'g-', label="A, k/n=%s %%"%(100*k[0]/n))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[0], 'g--',label="X, k/n=%s %%"%(100*k[0]/n))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[1], 'b-', label="A, k/n=%s %%"%(100*k[1]/n))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[1], 'b--',label="X, k/n=%s %%"%(100*k[1]/n))
plt.plot(np.arange(max_epoch)+1, plot_A_entries[2], 'r-', label="A, k/n=%s %%"%(100*k[2]/n))
plt.plot(np.arange(max_epoch)+1, plot_X_entries[2], 'r--',label="X, k/n=%s %%"%(100*k[2]/n))
plt.grid(True)
plt.title('Percentage of entries of A and X Recovered with N = %s'%(N))
plt.xlabel('Epochs')
plt.ylabel('Percentage of entries of A and X recovered')
plt.xlim(1,plot_epoch)
plt.legend()
plt.savefig('./figures/both.eps', format='eps', bbox_inches='tight')
# ### Remarks
# 1) Firstly we note that the for loop on line 32 can be trivially parallelized so that the extraction of partial supports from each column can be done simultaneously. This should result in a very significant speedup over the current serial implementation, particularly in the case when $N$ is large.<br>
# 2) Secondly we will often observe that $A$ is learnt far quicker than $X$, indeed we learn an X value as a by-product of extracting or finding a partial support. Once A has been recovered it still requires multiple epochs to pass through and decode the data points.<br>
# 3) Sometimes certain entries in $X$ will not be recovered by the algorithm, i.e. a certain number of entries will be missing from the recovery. This is as a result of the columns of $A$ whose indices are in the support of a given column of $X$ overlapping too much, and not satisfyign the unique neighbour property. Although our check ensures that no two columns overlap by more than $2 \epsilon d$, it may be the case that a set of $l \geq 3$ columns of $A$ overlap enough that partial support can be extracted.<br>
#
| Old/Expander_DL_v04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Classifying Fashion Items using Deep Learning
#
# ### Motivation
#
# Image classification is the problem of assigning an input image one label from a fixed set of categories. Image classification is one of the core problems in computer vision. But despite it's simplicity, image classification has a ton of practical applications.
#
# ### Our Approach
#
# In this example we shall use a basic 2 layer convolutional neural net to classify images from the [Fashion-MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset. We shall use 60,000 images to train our neural net and test our model using 10,000 images. We shall save our model checkpoints so that we can use them later with tensorboard to visualize the learning.
#
# ### Why Fashion-MNIST?
#
# **MNIST is too easy!!**
#
# We have been using MNIST to train and evaluate our neural network models since the 90's. Although, we must agree that MNIST has aged well and still is much respected in the machine learning community but it's too damn easy now-a-days to prepare a model that can achieve more than [99% accuracy on it](http://rodrigob.github.io/are_we_there_yet/build/classification_datasets_results.html#4d4e495354). Fashion-MNIST is more complicated than MNIST and has been prepared to replace the age old dataset.
# +
"""
These are all the modules we'll be using later. Make sure you can import them before proceeding further.
"""
from __future__ import print_function
import os
import sys
import gzip
import numpy as np
from six.moves.urllib.request import urlretrieve
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
tf.logging.set_verbosity(tf.logging.INFO)
# -
"""
Global variables that we need to set.
"""
url = "http://fashion-mnist.s3-website.eu-central-1.amazonaws.com/"
last_percent_reported = None
DATA_DIR = "data"
# +
"""
A hook to report the progress of a download. This is mostly intended for users with
slow internet connections. Reports every 1% change in download progress.
"""
def download_progress_hook(count, blockSize, totalSize):
global last_percent_reported
percent = int(count * blockSize * 100 / totalSize)
if last_percent_reported != percent:
if percent % 5 == 0:
sys.stdout.write("%s%%" % percent)
sys.stdout.flush()
else:
sys.stdout.write(".")
sys.stdout.flush()
last_percent_reported = percent
"""
Download a file if not present, and make sure it's the right size.
"""
def maybe_download(download_path, filename, expected_bytes, force=False):
if not os.path.exists(download_path):
os.makedirs(download_path)
fullfilename = os.path.join(download_path, filename)
if force or not os.path.exists(fullfilename):
print('Attempting to download:', fullfilename)
filename, _ = urlretrieve(url + filename, fullfilename, reporthook=download_progress_hook)
print('\nDownload Complete!')
statinfo = os.stat(fullfilename)
if statinfo.st_size == expected_bytes:
print('Found and verified', fullfilename)
else:
raise Exception('Failed to verify ' + fullfilename + '. Can you get to it with a browser?')
maybe_download(DATA_DIR, 'train-images-idx3-ubyte.gz', 26421880)
maybe_download(DATA_DIR, 'train-labels-idx1-ubyte.gz', 29515)
maybe_download(DATA_DIR, 't10k-images-idx3-ubyte.gz', 4422102)
maybe_download(DATA_DIR, 't10k-labels-idx1-ubyte.gz', 5148)
# -
"""
Load data.
"""
fashion_data = input_data.read_data_sets(DATA_DIR, one_hot=False, validation_size=0)
train_X = fashion_data.train.images # This returns a numpy array
train_y = np.asarray(fashion_data.train.labels, dtype=np.int32)
test_X = fashion_data.test.images
test_y = np.asarray(fashion_data.test.labels, dtype=np.int32)
# +
"""
Optionally we can suffle the data.
"""
from sklearn.utils import shuffle
train_X, train_y = shuffle(train_X, train_y)
test_X, test_y = shuffle(test_X, test_y)
# -
print("train_X: ", train_X.shape)
print("train_y: ", train_y.shape)
print("test_X: ", test_X.shape)
print("test_y: ", test_y.shape)
# +
"""
We can visualize some of our training data using matplotlib. Visualization is optional but highly recommended.
"""
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
# Config the matplotlib backend as plotting inline in IPython
# %matplotlib inline
fig = plt.figure()
gs = gridspec.GridSpec(2, 5)
for i in np.arange(10):
""" Pick a random sample from the loaded set of images and display using matplotlib """
sample_idx = np.random.randint(len(train_X))
sample_img = train_X[sample_idx, :].reshape(28, 28)
fig.add_subplot(gs[i])
plt.imshow(sample_img, cmap='gray')
# -
# ### Our Model
#
# Our model is a 2 layer ConvNet. Each conv layer is followed by a max pooling layer. Finally, we are using one dense layer for classification. To make sure our model does not overfit, we are using a dropout.
#
# Graphically, our model looks something like this:
#
# 
def cnn_model_fn_1(features, labels, mode):
# Input Layer
# Reshape X to 4-D tensor: [batch_size, width, height, channels]
input_layer = tf.reshape(features["x"], [-1, 28, 28, 1])
# Convolutional Layer #1
# Computes 32 features using a 5x5 filter with ReLU activation.
# Padding is added to preserve width and height.
# Input Tensor Shape: [batch_size, 28, 28, 1]
# Output Tensor Shape: [batch_size, 28, 28, 32]
conv1 = tf.layers.conv2d(
inputs=input_layer,
filters=32,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
# Pooling Layer #1
# First max pooling layer with a 2x2 filter and stride of 2
# Input Tensor Shape: [batch_size, 28, 28, 32]
# Output Tensor Shape: [batch_size, 14, 14, 32]
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2, 2], strides=2)
# Convolutional Layer #2
# Computes 64 features using a 5x5 filter.
# Padding is added to preserve width and height.
# Input Tensor Shape: [batch_size, 14, 14, 32]
# Output Tensor Shape: [batch_size, 14, 14, 64]
conv2 = tf.layers.conv2d(
inputs=pool1,
filters=64,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
# Pooling Layer #2
# Second max pooling layer with a 2x2 filter and stride of 2
# Input Tensor Shape: [batch_size, 14, 14, 64]
# Output Tensor Shape: [batch_size, 7, 7, 64]
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2, 2], strides=2)
# Flatten tensor into a batch of vectors
# Input Tensor Shape: [batch_size, 7, 7, 64]
# Output Tensor Shape: [batch_size, 7 * 7 * 64]
pool2_flat = tf.reshape(pool2, [-1, 7 * 7 * 64])
# Dense Layer
# Densely connected layer with 1024 neurons
# Input Tensor Shape: [batch_size, 7 * 7 * 64]
# Output Tensor Shape: [batch_size, 1024]
dense = tf.layers.dense(inputs=pool2_flat, units=1024, activation=tf.nn.relu)
# Add dropout operation; 0.6 probability that element will be kept
dropout = tf.layers.dropout(inputs=dense, rate=0.4, training=mode == tf.estimator.ModeKeys.TRAIN)
# Logits layer
# Input Tensor Shape: [batch_size, 1024]
# Output Tensor Shape: [batch_size, 10]
logits = tf.layers.dense(inputs=dropout, units=10)
predictions = {
# Generate predictions (for PREDICT and EVAL mode)
"classes": tf.argmax(input=logits, axis=1),
# Add `softmax_tensor` to the graph. It is used for PREDICT and by the `logging_hook`.
"probabilities": tf.nn.softmax(logits, name="softmax_tensor")
}
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode=mode, predictions=predictions)
# Calculate Loss (for both TRAIN and EVAL modes)
onehot_labels = tf.one_hot(indices=tf.cast(labels, tf.int32), depth=10)
loss = tf.losses.softmax_cross_entropy(onehot_labels=onehot_labels, logits=logits)
# Configure the Training Op (for TRAIN mode)
if mode == tf.estimator.ModeKeys.TRAIN:
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train_op = optimizer.minimize(loss=loss, global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
# Add evaluation metrics (for EVAL mode)
eval_metric_ops = {"accuracy": tf.metrics.accuracy(labels=labels, predictions=predictions["classes"])}
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, eval_metric_ops=eval_metric_ops)
# Create the Estimator
mnist_classifier = tf.estimator.Estimator(model_fn=cnn_model_fn_1, model_dir="./mnist_convnet_model")
# Train the model
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": train_X},
y=train_y,
batch_size=400,
num_epochs=None,
shuffle=True)
# Evaluate the model and print results
eval_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": test_X},
y=test_y,
num_epochs=1,
shuffle=False)
for j in range(100):
mnist_classifier.train(input_fn=train_input_fn, steps=2000)
eval_results = mnist_classifier.evaluate(input_fn=eval_input_fn)
print(eval_results)
| section_2/00_fashion_image_classification/fashion-mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.8 64-bit
# language: python
# name: python3
# ---
# + [markdown] id="vO1xwQMynOnC"
# # Naive Bayes w/o WE
#
# -
# ## Preprocessing
# + colab={"base_uri": "https://localhost:8080/"} id="bRqNl6G-mPT7" outputId="68b29eb7-ad2e-49f9-af64-ab03ba0add06"
# from google.colab import drive
# drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="H7Swf4P8H5xv" outputId="c6fdfb30-7d8e-4a4f-b28a-927f6cb8e76d"
# # cd drive/MyDrive/NLP_Project
# + id="6135jyy5IQ2B"
# # !pip install pandas matplotlib tqdm seaborn sklearn numpy graphviz
import pandas as pd
import matplotlib.pyplot as plt
from tqdm import tqdm
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.model_selection import GridSearchCV
import numpy as np
import warnings
warnings.filterwarnings('always')
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
import pickle as pk
# + id="wCsgwo7tIvAs"
trainData = np.load('../../../dataFinal/npy_files/fin_noWE_t2_train.npy')
trainLabels = open('../../../dataFinal/finalTrainLabels.labels', 'r').readlines()
testData = np.load('../../../dataFinal/npy_files/fin_noWE_t2_test.npy')
testLabels = open('../../../dataFinal/finalTestLabels.labels', 'r').readlines()
validationData = np.load('../../../dataFinal/npy_files/fin_noWE_t2_trial.npy')
validationLabels = open('../../../dataFinal/finalDevLabels.labels', 'r').readlines()
# + colab={"base_uri": "https://localhost:8080/"} id="1QjcQGoVGDJB" outputId="ff4a6bee-c555-4753-a288-9b5baed2bb0b"
for i in tqdm(range(len(trainLabels))):
trainLabels[i] = int(trainLabels[i])
for i in tqdm(range(len(testLabels))):
testLabels[i] = int(testLabels[i])
for i in tqdm(range(len(validationLabels))):
validationLabels[i] = int(validationLabels[i])
# + id="A6A_MlQgJlte"
trainLabels = np.array(trainLabels)
testLabels = np.array(testLabels)
validationLabels = np.array(validationLabels)
trainLabels = trainLabels.reshape((-1, ))
testLabels = testLabels.reshape((-1, ))
validationLabels = validationLabels.reshape((-1, ))
X_train, X_test, y_train, y_test, X_val, y_val = trainData, testData, trainLabels, testLabels, validationData, validationLabels
# + [markdown] id="13pLC5g7nUJO"
# ## Gaussian - Better
# + id="eSVyYN35d4d_" outputId="7b5add08-2107-4166-ccd6-d2c5237283e8"
from sklearn.naive_bayes import GaussianNB
clf = GaussianNB()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
y_true = testLabels
print("Test Data:")
accuracy = round(accuracy_score(y_pred = y_pred, y_true = y_true) * 100,2)
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
y_pred = clf.predict(X_train)
y_true = trainLabels
print("Train Data:")
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
y_pred = clf.predict(X_val)
y_true = validationLabels
print("Validation Data:")
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
# -
filename = 'finalModelGNB_NoWE'
pk.dump(clf,open(filename,'wb'))
# + [markdown] id="vBGw25QQndDk"
# ## Bernoulli
# + id="0cbQ_Tndd4d_" outputId="d9212cd4-e54c-4891-dae0-f0f508220908"
from sklearn.naive_bayes import BernoulliNB
clf = BernoulliNB()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
y_true = testLabels
print("Test Data:")
accuracy = round(accuracy_score(y_pred = y_pred, y_true = y_true) * 100,2)
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
y_pred = clf.predict(X_train)
y_true = trainLabels
print("Train Data:")
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
y_pred = clf.predict(X_val)
y_true = validationLabels
print("Validation Data:")
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
# -
filename = 'finalModelBNBNoWE'
pk.dump(clf,open(filename,'wb'))
# + [markdown] id="0J9r1GCFnhNu"
# ## Multinomial
# + id="W8uIr5sHd4d_" outputId="e4fc82c2-fbc2-45cf-c4ab-0b9688134b1f"
import numpy as np
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
y_true = testLabels
print("Test Data:")
accuracy = round(accuracy_score(y_pred = y_pred, y_true = y_true) * 100,2)
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
y_pred = clf.predict(X_train)
y_true = trainLabels
print("Train Data:")
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
y_pred = clf.predict(X_val)
y_true = validationLabels
print("Validation Data:")
print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
# + id="W8uIr5sHd4d_" outputId="e4fc82c2-fbc2-45cf-c4ab-0b9688134b1f"
filename = 'finalModelMNBNoWE'
pk.dump(clf,open(filename,'wb'))
# import numpy as np
# from sklearn.naive_bayes import MultinomialNB
# clf = MultinomialNB()
# clf.fit(X_train, y_train)
# y_pred = clf.predict(X_test)
# y_true = testLabels
# print("Test Data:")
# accuracy = round(accuracy_score(y_pred = y_pred, y_true = y_true) * 100,2)
# print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
# print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
# y_pred = clf.predict(X_train)
# y_true = trainLabels
# print("Train Data:")
# print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
# print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
# y_pred = clf.predict(X_val)
# y_true = validationLabels
# print("Validation Data:")
# print("Accuracy % : ",round(accuracy_score(y_pred = y_pred, y_true=y_true) * 100,2))
# print(classification_report(y_true,y_pred,labels = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19]))
| src/finalModels/NaiveBayes/NB_noWE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import h5py, os
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from imp import reload
import pandas as pd
from sklearn.decomposition import PCA
from scipy.stats import spearmanr
from dca import analysis, data_util, style, DynamicalComponentsAnalysis
# -
T_pi = 10
dim = 3
n_init = 5
# +
data = []
X = data_util.load_sabes_data('neural/indy_20160627_01.mat')['M1']
unit = 'Neurons'
name = 'M1'
data.append((X, name, unit))
X = data_util.load_kording_paper_data('neural/example_data_hc.pickle')['neural']
unit = 'Neurons'
name = 'HC'
data.append((X, name, unit))
X = data_util.load_weather_data('weather/temperature.csv')
unit = 'Cities'
name = 'Temperature'
data.append((X, name, unit))
X = data_util.load_accel_data('accel/sub_19.csv')
unit = 'Sensors'
name = 'Accelerometer'
data.append((X, name, unit))
# -
results = []
for X, name, unit in data:
dca = DynamicalComponentsAnalysis(T=T_pi, d=dim, n_init=n_init)
pca = PCA(n_components=3)
Xd = PCA().fit_transform(dca.fit_transform(X))
Xp = pca.fit_transform(X)
dca_ls = np.linalg.norm(dca.coef_, axis=1)**2 / dim
pca_ls = np.linalg.norm(pca.components_, axis=0)**2 / dim
results.append((dca_ls, pca_ls, Xd, Xp))
# +
fig, axes_m = plt.subplots(4, 3, figsize=(12, 16))
for (_, name, unit), (dca_ls, pca_ls, _, _), axes in zip(data, results, axes_m):
c, p = spearmanr(dca_ls, pca_ls)
c = np.round(c, 2)
pl10 = int(np.floor(np.log10(p)))
pr = np.round(p / 10**pl10, 2)
ax = axes[0]
ax.plot(np.sort(dca_ls)[::-1], c='r', label='DCA')
ax.plot(np.sort(pca_ls)[::-1], c='k', label='PCA')
ax.set_xlabel('{} sorted per model'.format(unit))
ax.set_ylabel('{}\nLeverage Scores'.format(name))
ax.text(.9, .5, 'Spearman-r: {}\np-value: {}e{}'.format(c, pr, pl10),
ha='right', transform=ax.transAxes)
ax.legend()
ax = axes[1]
idxs = np.argsort(dca_ls)[::-1]
ax.plot(dca_ls[idxs], c='r', label='DCA')
ax.plot(pca_ls[idxs], c='k', label='PCA')
ax.set_xlabel('{} sorted by DCA'.format(unit))
ax = axes[2]
idxs = np.argsort(pca_ls)[::-1]
ax.plot(dca_ls[idxs], c='r', label='DCA')
ax.plot(pca_ls[idxs], c='k', label='PCA')
ax.set_xlabel('{} sorted by PCA'.format(unit))
fig.tight_layout()
plt.savefig('leverage_scores_3d_old.pdf'.format(name))
# +
te = .01
be = .05
le = .13
re = .02
lg = .05
sm = .01
h = (1 - te - be - 4*sm - 3*lg) / 8
w = 1 - le - re
fig = plt.figure(figsize=(4, 8))
axes = []
names = ['M1', 'HC', 'Temperature', 'Accelerometer']
colors = ['r', 'k', 'gray']
for ii in range(4):
x = le
y = 1 - te - (h * 2 + sm + lg) * ii - h
fig.text(0, y - sm / 2, names[ii], rotation=90, va='center', ha='left')
axes.append(fig.add_axes([le, y, w, h]))
y = y - sm - h
axes.append(fig.add_axes([le, y, w, h]))
slices = [slice(1000, 1250), slice(2000, 2250), slice(0, 600), slice(2000, 7000)]
bin_sizes = [.05, .05, 1, .02]
for ii, ((X, name, unit), (_, _, Xd, Xp), sl, bs) in enumerate(zip(data, results, slices, bin_sizes)):
xd = Xd[sl, :]
xp = Xp[sl, :]
xd -= xd.mean(axis=0, keepdims=True)
xd /= xd.std(axis=0, keepdims=True)
xp -= xp.mean(axis=0, keepdims=True)
xp /= xp.std(axis=0, keepdims=True)
maxval = max(abs(xd).max(), abs(xp).max())
t = np.linspace(0, xd.shape[0] * bs, xd.shape[0])
for jj in np.arange(dim)[::-1]:
label = 'Dim {}'.format(jj+1)
axes[ii*2].plot(t, xd[:, jj], c=colors[jj], label=label, lw=1)
axes[ii*2+1].plot(t, xp[:, jj], c=colors[jj], lw=1)
axes[ii*2].set_ylim(-maxval * 1.1, maxval * 1.1)
axes[ii*2+1].set_ylim(-maxval * 1.1, maxval * 1.1)
axes[ii*2].set_ylabel('DCA', fontsize=style.axis_label_fontsize, labelpad=-5)
axes[ii*2].set_xticks([])
axes[ii*2+1].set_ylabel('PCA', fontsize=style.axis_label_fontsize, labelpad=-5)
print(spearmanr(Xd[:, jj], Xp[:, jj]))
print()
axes[0].legend(ncol=3, fontsize=style.axis_label_fontsize,
loc='lower left', frameon=True, bbox_to_anchor=(0., -.06), borderpad=.15)
axes[1].set_xlabel('Time (s)', fontsize=style.axis_label_fontsize)
axes[3].set_xlabel('Time (s)', fontsize=style.axis_label_fontsize)
axes[5].set_xlabel('Time (days)', fontsize=style.axis_label_fontsize)
axes[7].set_xlabel('Time (s)', fontsize=style.axis_label_fontsize)
for ax in axes:
ax.tick_params(labelsize=style.ticklabel_fontsize)
plt.savefig('inferred_dynamics_3d.pdf'.format(name))
# +
fig, axes_m = plt.subplots(4, 2, figsize=(4.75, 8))
for (_, name, unit), (dca_ls, pca_ls, _, _), axes in zip(data, results, axes_m):
c, p = spearmanr(dca_ls, pca_ls)
c = np.round(c, 2)
pl10 = int(np.floor(np.log10(p)))
pr = np.round(p / 10**pl10, 2)
ax = axes[0]
ax.plot(np.sort(dca_ls)[::-1], c='r', label='DCA')
ax.plot(np.sort(pca_ls)[::-1], c='k', label='PCA')
ax.set_xlabel('{} sorted per model'.format(unit), fontsize=style.axis_label_fontsize)
ax.set_ylabel('{}\nLeverage Scores'.format(name), fontsize=style.axis_label_fontsize)
ax = axes[1]
maxval = max(dca_ls.max(), pca_ls.max())
minval = min(dca_ls.min(), pca_ls.min())
ax.scatter(dca_ls, pca_ls, marker='.', c='k')
ax.set_xlabel('DCA leverage scores', fontsize=style.axis_label_fontsize)
ax.set_ylabel('PCA leverage scores', fontsize=style.axis_label_fontsize)
ax.set_xlim(minval / 2, maxval*2)
ax.set_ylim(minval / 2, maxval*2)
ax.set_xscale('log')
ax.set_yscale('log')
ax.text(.05, .98, 'RC: {}\np: {}e{}'.format(c, pr, pl10),
ha='left', va='top', transform=ax.transAxes, fontsize=style.ticklabel_fontsize)
axes_m[0, 0].legend(fontsize=style.axis_label_fontsize)
for ax in axes_m.ravel():
ax.tick_params(labelsize=style.ticklabel_fontsize)
fig.tight_layout()
plt.savefig('leverage_scores_3d.pdf'.format(name))
# -
| notebooks/real_data_latent_loadings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load the modules
# +
import cudf
from collections import OrderedDict
import numpy as np
import datetime as dt
from dask_cuda import LocalCUDACluster
from dask.distributed import Client
from blazingsql import BlazingContext
import dask_cudf
# -
print(cudf.__version__)
cluster = LocalCUDACluster()
client = Client(cluster)
client.restart()
# # Read the data
# !head -n 10 ../../data/parking_MayJun2019.csv
transactions_path = '../../data/seattle_parking/parking_MayJun2019.parquet/partition_idx={partition}/*'
transactions_parq = [transactions_path.format(partition=p) for p in range(transactions_partitions_cnt)]
# +
parking_transactions = dask_cudf.read_parquet(
transactions_parq
)
parking_locations = dask_cudf.read_parquet('../../data/seattle_parking/parking_locations.parquet/')
# -
# Features
#
# 1. Occupancy at T minus 15mins, 1h, 3h, 1d, 2d, 3d, 7d, 14d
# 2. PaidParkingArea & SubArea
bc = BlazingContext(dask_client=client)
bc.create_table('parking_transactions', parking_transactions)
bc.sql('SELECT * FROM parking_transactions LIMIT 10').compute()
bc.sql('''
SELECT SourceElementKey
, OccupancyDateTime
, dow AS DOW
, HOUR(OccupancyDateTime) - 1 AS hr_minus_1
, HOUR(OccupancyDateTime) - 3 AS hr_minus_3
, DAYOFMONTH(OccupancyDateTime) - 1 AS day_minus_1
FROM parking_transactions
ORDER BY SourceElementKey
, OccupancyDateTime
LIMIT 10
''').compute()
# +
df_locations['ParkingArea_concat'] = (
df_locations['PaidParkingArea'].str.replace(' ', '_')
.str.cat(df_locations['PaidParkingSubArea'], sep='__')
)
df_locations.head()
| codes/parking/rapids_seattleParking_ML.ipynb |