code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Word Embeddings: Hands On
#
# In previous lecture notebooks you saw all the steps needed to train the CBOW model. This notebook will walk you through how to extract the word embedding vectors from a model.
#
# Let's dive into it!
import numpy as np
from utils2 import get_dict
# Before moving on, you will be provided with some variables needed for further procedures, which should be familiar by now. Also a trained CBOW model will be simulated, the corresponding weights and biases are provided:
# +
# Define the tokenized version of the corpus
words = ['i', 'am', 'happy', 'because', 'i', 'am', 'learning']
# Define V. Remember this is the size of the vocabulary
V = 5
# Get 'word2Ind' and 'Ind2word' dictionaries for the tokenized corpus
word2Ind, Ind2word = get_dict(words)
# Define first matrix of weights
W1 = np.array([[ 0.41687358, 0.08854191, -0.23495225, 0.28320538, 0.41800106],
[ 0.32735501, 0.22795148, -0.23951958, 0.4117634 , -0.23924344],
[ 0.26637602, -0.23846886, -0.37770863, -0.11399446, 0.34008124]])
# Define second matrix of weights
W2 = np.array([[-0.22182064, -0.43008631, 0.13310965],
[ 0.08476603, 0.08123194, 0.1772054 ],
[ 0.1871551 , -0.06107263, -0.1790735 ],
[ 0.07055222, -0.02015138, 0.36107434],
[ 0.33480474, -0.39423389, -0.43959196]])
# Define first vector of biases
b1 = np.array([[ 0.09688219],
[ 0.29239497],
[-0.27364426]])
# Define second vector of biases
b2 = np.array([[ 0.0352008 ],
[-0.36393384],
[-0.12775555],
[-0.34802326],
[-0.07017815]])
# -
#
#
# ## Extracting word embedding vectors
#
# Once you have finished training the neural network, you have three options to get word embedding vectors for the words of your vocabulary, based on the weight matrices $\mathbf{W_1}$ and/or $\mathbf{W_2}$.
#
# ### Option 1: extract embedding vectors from $\mathbf{W_1}$
#
# The first option is to take the columns of $\mathbf{W_1}$ as the embedding vectors of the words of the vocabulary, using the same order of the words as for the input and output vectors.
#
# > Note: in this practice notebooks the values of the word embedding vectors are meaningless after a single iteration with just one training example, but here's how you would proceed after the training process is complete.
#
# For example $\mathbf{W_1}$ is this matrix:
# Print W1
W1
# The first column, which is a 3-element vector, is the embedding vector of the first word of your vocabulary. The second column is the word embedding vector for the second word, and so on.
#
# The first, second, etc. words are ordered as follows.
# Print corresponding word for each index within vocabulary's range
for i in range(V):
print(Ind2word[i])
# So the word embedding vectors corresponding to each word are:
# Loop through each word of the vocabulary
for word in word2Ind:
# Extract the column corresponding to the index of the word in the vocabulary
word_embedding_vector = W1[:, word2Ind[word]]
# Print word alongside word embedding vector
print(f'{word}: {word_embedding_vector}')
# ### Option 2: extract embedding vectors from $\mathbf{W_2}$
# The second option is to take $\mathbf{W_2}$ transposed, and take its columns as the word embedding vectors just like you did for $\mathbf{W_1}$.
# Print transposed W2
W2.T
# Loop through each word of the vocabulary
for word in word2Ind:
# Extract the column corresponding to the index of the word in the vocabulary
word_embedding_vector = W2.T[:, word2Ind[word]]
# Print word alongside word embedding vector
print(f'{word}: {word_embedding_vector}')
# ### Option 3: extract embedding vectors from $\mathbf{W_1}$ and $\mathbf{W_2}$
# The third option, which is the one you will use in this week's assignment, uses the average of $\mathbf{W_1}$ and $\mathbf{W_2^\top}$.
# **Calculate the average of $\mathbf{W_1}$ and $\mathbf{W_2^\top}$, and store the result in `W3`.**
# +
# Compute W3 as the average of W1 and W2 transposed
W3 = (W1+W2.T)/2
# Print W3
W3
# -
# Expected output:
#
# array([[ 0.09752647, 0.08665397, -0.02389858, 0.1768788 , 0.3764029 ],
# [-0.05136565, 0.15459171, -0.15029611, 0.19580601, -0.31673866],
# [ 0.19974284, -0.03063173, -0.27839106, 0.12353994, -0.04975536]])
# Extracting the word embedding vectors works just like the two previous options, by taking the columns of the matrix you've just created.
# Loop through each word of the vocabulary
for word in word2Ind:
# Extract the column corresponding to the index of the word in the vocabulary
word_embedding_vector = W3[:, word2Ind[word]]
# Print word alongside word embedding vector
print(f'{word}: {word_embedding_vector}')
# Now you know 3 different options to get the word embedding vectors from a model!
# ### How this practice relates to and differs from the upcoming graded assignment
#
# - After extracting the word embedding vectors, you will use principal component analysis (PCA) to visualize the vectors, which will enable you to perform an intrinsic evaluation of the quality of the vectors, as explained in the lecture.
# **Congratulations on finishing all lecture notebooks for this week!**
#
# You're now ready to take on this week's assignment!
#
# **Keep it up!**
| Part2_Probabilistic_Models/C2_W4_lecture_nb_4_word_embeddings_hands_on.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
filename = './exampleFiles/truncateFile.txt'
changes = False
with open(filename, 'r+') as f:
doc = f.read()
doc = doc.replace('line', '+line')
changes = True
if changes:
f.truncate(0)
f.write(doc)
| notebooks/truncateWrite.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab={} colab_type="code" id="S4k18nivWAPG"
# # Extract User and Item Latent Factors Using Neural Collaborative Filtering
#
# Here we train neural collaborative filtering (NCF) model to predict the ratings given the user_ids and item_ids. The neural collaborative filtering model consists of Generalized Matrix Factorization (GMF) stream and Multi-Layer Perceptron (MLP) stream, which represents the matrix factorization and the non-linear relation of the user embedding and the item embedding. The neural collaborative filtering model fuses GMF stream and MLP stream and is able to predict the ratings given the ids of the user and the item. After training, the user and item embeddings from GMF and MLP stream are treated as user and item latent factors. The latent factors are stored into "user_latent.csv" and "item_latent.csv" in GCP bucket.
#
# ---
# The implementation is related to the following paper:
# - [Neural Collaborative Filtering](https://arxiv.org/abs/1708.05031)
# -
# ## 1. import libraries
# import libraries
import os
import pandas as pd
import tensorflow as tf
from google.cloud import bigquery
# +
# set constants
PROJECT = "hybrid-recsys-gcp"
BUCKET = "hybrid-recsys-gcp-bucket"
REGION = 'us-central1'
DATASET = 'news_recommend_dataset'
MODEL = "neural_collaborate_filter_trained_model"
os.environ["PROJECT"] = PROJECT
os.environ["BUCKET"] = BUCKET
os.environ["REGION"] = REGION
os.environ["DATASET"] = DATASET
os.environ["MODEL"] = MODEL
# -
# ## 2. create neural_collaborate_filter package
#
# This is the package for the neural colaborative filtering model. The trainer/task.py defines the input arguments for training. In trainer/model.py, the "create_dataset" function creates tf.dataset for input user and item ids. The "NeuMF_Model" class defines the tf.keras.Model which takes user and item id to predict rating. And the "train_model_and_save_latent_factors" function defines the custom training loop for training the model.
# The NeuMF_Model uses the following architecture:
#
# <img src="./img/neural_CF.png" width="65%" height="65%" />
# + colab={} colab_type="code" id="_weFIvdVwp5o" language="bash"
# mkdir -p neural_collaborate_filter/trainer
# touch neural_collaborate_filter/trainer/__init__.py
# +
# %%writefile neural_collaborate_filter/trainer/task.py
import argparse
import tensorflow as tf
from trainer import model
if __name__ == "__main__":
parser = argparse.ArgumentParser()
parser.add_argument(
"--job-dir",
help="job dir to store training outputs and other data",
required=True
)
parser.add_argument(
"--train_data_path",
help="path to import train data",
required=True
)
parser.add_argument(
"--test_data_path",
help="path to import test data",
required=True
)
parser.add_argument(
"--output_dir",
help="output dir to export checkpoints or trained model",
required=True
)
parser.add_argument(
"--batch_size",
help="batch size for training",
type=int,
default=2048
)
parser.add_argument(
"--epochs",
help="number of epochs for training",
type=int,
default=1
)
parser.add_argument(
"--latent_num",
help="number of latent factors for gmf and mlp each",
type=int,
default=8
)
parser.add_argument(
"--user_id_path",
help="path to import user_id_list.txt",
required=True
)
parser.add_argument(
"--item_id_path",
help="path to import item_id_list.txt",
required=True
)
parser.add_argument(
"--user_latent_path",
help="output path to save user latent factors",
default="./"
)
parser.add_argument(
"--item_latent_path",
help="output path to save item latent factors",
default="./"
)
parser.add_argument(
"--save_latent_factors",
help="set to save latent factors",
default=False,
action="store_true"
)
args = parser.parse_args()
args = args.__dict__
model.train_model_and_save_latent_factors(args)
# +
# %%writefile neural_collaborate_filter/trainer/model.py
import os
import pandas as pd
import tensorflow as tf
import shutil
import datetime
# create dataset function
def create_dataset(path, column_names, label_name, defaults, batch_size, shuffle):
""" Create tf.dataset from csv file.
Args:
path (str): Path to the csv file.
column_names (list:str): List of string to specify which columns to use in dataset (including label).
label_name (str): Column name for the label.
defaults (list:str): List of string to set default values for columns.
batch_size (str): Batchsize of the dataset.
shuffle (bool): True for shuffling dataset and False otherwise.
Returns:
(tf.dataset): dataset used for training or testing
"""
dataset = tf.data.experimental.make_csv_dataset(
file_pattern=path,
select_columns=column_names,
label_name=label_name,
column_defaults = defaults,
batch_size=batch_size,
num_epochs=1,
shuffle=shuffle
)
return dataset
# model class
class NeuMF_Model(tf.keras.Model):
""" The NeuMF_Model class. Takes user ids and item ids to predict ratings through matrix factorization and
multilayer perceptron. The user and item embeddings are user and item latent factors.
Attributes:
user_index_table (tf.lookup.StaticVocabularyTable): Table to transform user_id to user indices.
item_index_table (tf.lookup.StaticVocabularyTable): Table to transform item_id to item indices.
gmf_u_embed (tf.keras.layers.Embedding): User embedding layer for General Matrix Factorization.
gmf_i_embed (tf.keras.layers.Embedding): Item embedding layer for General Matrix Factorization.
mlp_u_embed (tf.keras.layers.Embedding): User embedding layer for Multilayer Perceptron.
mlp_i_embed (tf.keras.layers.Embedding): Item embedding layer for Multilayer Perceptron.
dense_1 (tf.keras.layers.Dense): First dense layer of multilayer perceptron stream.
dense_2 (tf.keras.layers.Dense): Second dense layer of multilayer perceptron stream.
dense_3 (tf.keras.layers.Dense): Third dense layer of multilayer perceptron stream.
dense_4 (tf.keras.layers.Dense): Fourth dense layer of multilayer perceptron stream.
output_layer (tf.keras.layers.Dense): Output dense layer for concat of mlp and gmf stream.
mlp_concat (tf.keras.layers.Concatenate): Concatenate layer to combine user and item embedding in mlp stream.
stream_concat (tf.keras.layers.Concatenate): Concatenate layer to combine mlp and gmf stream.
"""
def __init__(self, user_file_path, item_file_path, latent_num):
""" init method for NeuMF_Model class
Args:
user_file_path (str): Path to txt file containing user ids.
item_file_path (str): Path to txt file containing item ids.
Returns:
None
"""
super(NeuMF_Model, self).__init__()
self.user_index_table = self.create_lookup_table(user_file_path)
self.item_index_table = self.create_lookup_table(item_file_path)
user_id_size = self.get_size(user_file_path)
item_id_size = self.get_size(item_file_path)
self.gmf_u_embed = tf.keras.layers.Embedding(user_id_size, latent_num, name='gmf_u_embed')
self.gmf_i_embed = tf.keras.layers.Embedding(item_id_size, latent_num, name='gmf_i_embed')
self.mlp_u_embed = tf.keras.layers.Embedding(user_id_size, latent_num, name='mlp_u_embed')
self.mlp_i_embed = tf.keras.layers.Embedding(item_id_size, latent_num, name='mlp_i_embed')
self.dense_1 = tf.keras.layers.Dense(64, activation='relu', name='mlp_dense_1')
self.dense_2 = tf.keras.layers.Dense(32, activation='relu', name='mlp_dense_2')
self.dense_3 = tf.keras.layers.Dense(16, activation='relu', name='mlp_dense_3')
self.dense_4 = tf.keras.layers.Dense(8, activation='relu', name='mlp_dense_4')
self.output_layer = tf.keras.layers.Dense(1, activation='sigmoid', name='output_layer')
self.mlp_concat = tf.keras.layers.Concatenate(axis=1, name='mlp_concat')
self.stream_concat = tf.keras.layers.Concatenate(axis=1, name='stream_concat')
def create_lookup_table(self, file_path):
""" create lookup table to translate ids to indices
Args:
file_path (str): Path to txt file containing ids.
Returns:
(tf.lookup.StaticVocabularyTable): The lookup table.
"""
file_initializer = tf.lookup.TextFileInitializer(file_path, key_dtype=tf.string, key_index=tf.lookup.TextFileIndex.WHOLE_LINE, \
value_dtype=tf.int64, value_index=tf.lookup.TextFileIndex.LINE_NUMBER, delimiter="\n")
lookup_table = tf.lookup.StaticVocabularyTable(file_initializer, num_oov_buckets=1)
return lookup_table
def get_size(self, file_path):
""" Get the total number of lines for a txt file, indicating the size of the column.
Args:
file_path (str): Path to txt file.
Returns:
(int): The total number of lines for the txt file.
"""
id_text = tf.io.read_file(file_path)
id_tensor = tf.strings.split(id_text, '\n')
return id_tensor.shape[0]
@tf.function
def call(self, inputs, training):
"""The call method for NeuMF_Model class.
Args:
inputs (OrderedDict:tf.Tensor): OrderedDict of tensor containing user_id and item_id
Returns:
output (tf.Tensor): The predicted rating for the user and item combination.
"""
user_id = inputs['user_id']
item_id = inputs['item_id']
# convert id to index
user_index = self.user_index_table.lookup(user_id)
item_index = self.item_index_table.lookup(item_id)
# GMF stream
gmf_u_latent = self.gmf_u_embed(user_index)
gmf_i_latent = self.gmf_i_embed(item_index)
# multiply latent factors
gmf_out = gmf_u_latent * gmf_i_latent
# MLP stream
mlp_u_latent = self.mlp_u_embed(user_index)
mlp_i_latent = self.mlp_i_embed(item_index)
# concat latent factors and pass to dense layers
mlp_concat_out = self.mlp_concat([mlp_u_latent, mlp_i_latent])
dense_1_out = self.dense_1(mlp_concat_out)
dense_2_out = self.dense_2(dense_1_out)
dense_3_out = self.dense_3(dense_2_out)
mlp_out = self.dense_4(dense_3_out)
# concat GMF and MLP stream
stream_concat_out = self.stream_concat([gmf_out, mlp_out])
output = self.output_layer(stream_concat_out)
return output
def save_latent_factors_to_bucket(col_name, col_path, tensor_weight, output_path, latent_num):
"""Store tensor weights as user or item latent factors to bucket in csv file.
Args:
col_name (str): Column name for the latent factors (user or item).
col_path (str): Path to user or item ids.
tensor_weight (tf.Tensor): Tensors of the embedding layer used as latent factors.
output_path (str): Path to ouput file in bucket.
latent_num (int): Number of latent factors
Returns:
None
"""
id_tensors = tf.strings.split(tf.io.read_file(col_path), "\n")
id_list = [tf.compat.as_str_any(x) for x in id_tensors.numpy()]
latent_df = pd.DataFrame(tensor_weight)
latent_df[col_name] = id_list
key = range(latent_num * 2)
value = ['{}_latent_'.format(col_name[0]) + str(x) for x in key]
column_dict = dict(zip(key, value))
latent_df = latent_df.rename(columns=column_dict)
latent_df = latent_df[[col_name] + value]
latent_df.to_csv("./latent.csv", index=False)
script = "gsutil mv ./latent.csv {}".format(output_path)
os.system(script)
def train_model_and_save_latent_factors(args):
""" Train the NeuMF_Model and save embeddings as latent_factors to bucket in csv files.
Args:
args (dict): dict of arguments from task.py
Returns:
None
"""
# create dataset
column_name = ['user_id', 'item_id', 'rating']
label_name = 'rating'
defaults = ['unknown', 'unknown', 0.0]
batch_size = args["batch_size"]
train_path = args["train_data_path"]
test_path = args["test_data_path"]
train_dataset = create_dataset(train_path, column_name, label_name, defaults, batch_size, True)
test_dataset = create_dataset(test_path, column_name, label_name, defaults, batch_size, False)
# create model
model = NeuMF_Model(args["user_id_path"], args["item_id_path"], args["latent_num"])
# loss function and optimizers
bc_loss_object = tf.keras.losses.BinaryCrossentropy()
optimizer = tf.keras.optimizers.Adam(learning_rate = 0.001)
# loss metrics
train_bc_loss = tf.keras.metrics.Mean(name='train_bc_loss')
train_mae_loss = tf.keras.metrics.MeanAbsoluteError(name='train_mae_loss')
train_rmse_loss = tf.keras.metrics.RootMeanSquaredError(name='train_rmse_loss')
test_bc_loss = tf.keras.metrics.Mean(name='test_bc_loss')
test_mae_loss = tf.keras.metrics.MeanAbsoluteError(name='test_mae_loss')
test_rmse_loss = tf.keras.metrics.RootMeanSquaredError(name='test_rmse_loss')
@tf.function
def train_step(features, labels):
""" Concrete function for train setp and update train metircs
Args:
features (OrderedDict:tf.Tensor): OrderedDict of tensor containing user_id and item_id as features.
labels (tf.Tensor): labels (rating) of the training examples
Returns:
None
"""
with tf.GradientTape() as tape:
preds = model(features, training=True)
bc_loss = bc_loss_object(labels, preds)
gradients = tape.gradient(bc_loss, model.trainable_variables)
optimizer.apply_gradients(zip(gradients, model.trainable_variables))
train_bc_loss(bc_loss)
train_mae_loss(labels, preds)
train_rmse_loss(labels, preds)
@tf.function
def test_step(features, labels):
""" Concrete function for test setp and update test metircs
Args:
features (OrderedDict:tf.Tensor): OrderedDict of tensor containing user_id and item_id as features.
labels (tf.Tensor): labels (rating) of the training examples
Returns:
None
"""
preds = model(features, training=False)
bc_loss = bc_loss_object(labels, preds)
test_bc_loss(bc_loss)
test_mae_loss(labels, preds)
test_rmse_loss(labels, preds)
# custom train loop
EPOCHS = args["epochs"]
for epoch in range(EPOCHS):
train_bc_loss.reset_states()
train_mae_loss.reset_states()
train_rmse_loss.reset_states()
test_bc_loss.reset_states()
test_mae_loss.reset_states()
test_rmse_loss.reset_states()
for features, labels in train_dataset:
train_step(features, labels)
for features, labels in test_dataset:
test_step(features, labels)
template = "Epoch {:d}, train [bc_loss: {:.5f}, mae_loss: {:.5f}, rmse_loss: {:.5f}], test [bc_loss: {:.5f}, mae_loss: {:.5f}, rmse_loss: {:.5f}]"
print(template.format(epoch + 1, train_bc_loss.result(), train_mae_loss.result(), train_rmse_loss.result(), \
test_bc_loss.result(), test_mae_loss.result(), test_rmse_loss.result()))
# export model
EXPORT_PATH = os.path.join(args["output_dir"], datetime.datetime.now().strftime("%Y%m%d%H%M%S"))
tf.saved_model.save(obj=model, export_dir=EXPORT_PATH)
if args["save_latent_factors"]:
# get embedding weights
user_weight = tf.concat([model.gmf_u_embed.get_weights()[0], model.mlp_u_embed.get_weights()[0]], axis = 1).numpy()
item_weight = tf.concat([model.gmf_i_embed.get_weights()[0], model.mlp_i_embed.get_weights()[0]], axis = 1).numpy()
# store embedding weights to csv
save_latent_factors_to_bucket('user_id', args["user_id_path"], user_weight, args["user_latent_path"], args["latent_num"])
save_latent_factors_to_bucket('item_id', args["item_id_path"], item_weight, args["item_latent_path"], args["latent_num"])
# -
# ## 3. train model locally
#
# Run package as a python module in local environment.
# + language="bash"
#
# JOBDIR=./${MODEL}
# OUTDIR=./${MODEL}
#
# rm -rf ${JOBDIR}
# export PYTHONPATH=${PYTHONPATH}:${PWD}/neural_collaborate_filter
#
# python -m trainer.task \
# --job-dir=${JOBDIR} \
# --train_data_path=gs://${BUCKET}/${DATASET}/preprocess_train.csv \
# --test_data_path=gs://${BUCKET}/${DATASET}/preprocess_test.csv \
# --output_dir=${OUTDIR} \
# --batch_size=2048 \
# --epochs=8 \
# --latent_num=10 \
# --user_id_path=gs://${BUCKET}/${DATASET}/user_id_list.txt \
# --item_id_path=gs://${BUCKET}/${DATASET}/item_id_list.txt
# -
# ## 4. train model on gcloud
#
# Submit a training job in gcloud ai-platform to train the package.
# + language="bash"
#
# JOBDIR=gs://${BUCKET}/${MODEL}
# OUTDIR=gs://${BUCKET}/${MODEL}
# JOBID=neural_collaborate_filter_train_job_$(date -u +%y%m%d_%H%M%S)
#
# gcloud ai-platform jobs submit training ${JOBID} \
# --region=${REGION} \
# --module-name=trainer.task \
# --package-path=$(pwd)/neural_collaborate_filter/trainer \
# --staging-bucket=gs://${BUCKET} \
# --scale-tier=CUSTOM \
# --master-machine-type=n1-highcpu-16 \
# --runtime-version=2.1 \
# --python-version=3.7 \
# -- \
# --job-dir=${JOBDIR} \
# --train_data_path=gs://${BUCKET}/${DATASET}/preprocess_train.csv \
# --test_data_path=gs://${BUCKET}/${DATASET}/preprocess_test.csv \
# --output_dir=${OUTDIR} \
# --batch_size=2048 \
# --epochs=8 \
# --latent_num=10 \
# --user_id_path=gs://${BUCKET}/${DATASET}/user_id_list.txt \
# --item_id_path=gs://${BUCKET}/${DATASET}/item_id_list.txt \
# --user_latent_path=gs://${BUCKET}/${DATASET}/user_latent.csv \
# --item_latent_path=gs://${BUCKET}/${DATASET}/item_latent.csv \
# --save_latent_factors
# -
# The ai-platform training log should look like the following. The final test result is: bc_loss: 0.60900, mae_loss: 0.24119, rmse_loss: 0.30784.
#
# <img src="./img/ncf_train_log.png" width="80%" height="80%" />
# ## 5. save latent factors in bigquery dataset
#
# Save user and item laten factors into "user_latent.csv" and "item_latent.csv" in GCS bucket.
def load_csv_to_bigquery_table(project_id, dataset_id, table_id, schema, soruce_uri):
""" Load content from csv file into bigquery table.
Args:
project_id (str): ID of the project.
dataset_id (str): ID of the dataset.
table_id (str): ID of the table.
schema (list:bigquery.SchemaField): Schema of the csv file.
soruce_uri (str): Path to the csv file.
Returns:
None
"""
client = bigquery.Client(project_id)
dataset_ref = client.dataset(dataset_id)
job_config = bigquery.LoadJobConfig()
job_config.schema = schema
job_config.write_disposition = bigquery.WriteDisposition.WRITE_EMPTY
job_config.skip_leading_rows = 1
job_config.source_format = bigquery.SourceFormat.CSV
load_job = client.load_table_from_uri(soruce_uri, dataset_ref.table(table_id), job_config=job_config)
print("Starting job {}".format(load_job.job_id))
load_job.result() # Waits for table load to complete.
print("Job finished.")
destination_table = client.get_table(dataset_ref.table(table_id))
print("Loaded {} rows.".format(destination_table.num_rows))
# +
latent_num = 10
user_schema = [bigquery.SchemaField("user_id", "STRING", mode="REQUIRED")] + \
[bigquery.SchemaField("user_latent_{}".format(i), "FLOAT", mode="REQUIRED") for i in range(2 * latent_num)]
item_schema = [bigquery.SchemaField("item_id", "STRING", mode="REQUIRED")] + \
[bigquery.SchemaField("item_latent_{}".format(i), "FLOAT", mode="REQUIRED") for i in range(2 * latent_num)]
load_csv_to_bigquery_table(PROJECT, DATASET, "user_latent", user_schema, "gs://{}/{}/user_latent.csv".format(BUCKET, DATASET))
load_csv_to_bigquery_table(PROJECT, DATASET, "item_latent", item_schema, "gs://{}/{}/item_latent.csv".format(BUCKET, DATASET))
# -
# ## 6. view user and item latent factors
# !gsutil cp gs://{BUCKET}/{DATASET}/user_latent.csv ./{DATASET}/user_latent.csv
# !gsutil cp gs://{BUCKET}/{DATASET}/item_latent.csv ./{DATASET}/item_latent.csv
user_df = pd.read_csv("./{}/user_latent.csv".format(DATASET))
item_df = pd.read_csv("./{}/item_latent.csv".format(DATASET))
user_df.head()
item_df.head()
| neural_collaborate_filter_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import urllib3
import json
from bs4 import BeautifulSoup
import requests
import pandas as pd
dataset=pd.DataFrame()
# #### Key: Made a list of keywords for which I need the reddit posts
key=["husband", "wife", "girlfriend", "boyfriend", "bf", "gf"]
# #### Now traverse the Key list and fetch the reddit data. Some key parameters in url.
# #### 1) q: Keyword name
# #### 2) subreddit: Subreddit name (eg. CasualConv or Offmychest)
# #### 3) filter: Required data I need from the JSON fetched for a particular keyword.
# #### 4) size: Maximum number of post fetched.
# #### 5) after & before: Timespan of the posts fecthed.
# #### 6) sort_type: The way in which posts are sorted (eg. sort by score or date of publish)
# #### 7) sort: Sort order i.e. ascending or descending.
#
# #### Fetched the data for all keywords of "Key" list and appended the data all together to generate single CSV file i.e. "dataset".
#
# code below
for i in range(len(key)):
url = 'https://api.pushshift.io/reddit/search/submission/?q='+str(key[i])+'&subreddit=CasualConversation&filter=id,subreddit,num_comments,score,author,title,selftext,full_link,created_utc,over_18,num_comments,allow_live_comments,no_follow,pinned&size=1000&after=1483228800&before=1546300800&sort_type=score&sort=desc'
print(url)
response = requests.get(url)
dict = response.json()
df = pd.DataFrame(dict['data'])
df["Label"]=str(key[i])
dataset = pd.concat([dataset, df], ignore_index=True)
# #### Number of posts fetched
dataset.shape
# #### Count of number of posts for each keyword
dataset.Label.value_counts()
# #### Saving csv
dataset.to_csv('post_dataset.csv', index=False)
# #### Now create a list of ids of all the posts fetched. This will be used to get the comments of the posts fetched above
ids = list(dataset['id'])
len(ids)
# #### Now traverse through the "ids" list to fetch the comment data one by one.
# #### In this case also, all the commented data is for each post is appended together to form a single comment CSV file i.e. "dataset2"
# Code below
#
dataset2=pd.DataFrame()
for i in range(0,10880):
print(i)
url = 'http://api.pushshift.io/reddit/comment/search/?link_id='+str(ids[i])+'&size=50&sort_type=created_utc&sort=asc&filter=author,body,created_utc,score,subreddit,no_follow&score=>-100'
response = requests.get(url)
dict = response.json()
df = pd.DataFrame(dict['data'])
df["id"]=str(ids[i])
dataset2 = pd.concat([dataset2, df], ignore_index=True)
# #### Saving comment data as CSV
dataset2.to_csv('comment_dataset.csv', index=False)
| claff-offmychest-master/scripts/reddit_scarper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simon Fraser University - Mechatronic Systems Engineering
# ## Spring 2021 - MSE491 - Application of Machine Learning in Mechatronic Systems
# ## Instructor: Dr. <NAME>
#
# ## Lab 2 - Classification
# ### @author: <NAME> - <EMAIL>
#
# Load General Libraries
import numpy as np
import pandas as pd
import pickle
import matplotlib.pyplot as plt
# Importing the dataset
df0 = pd.read_csv("EMG\EMG_0.csv", header=None)
df1 = pd.read_csv("EMG\EMG_1.csv", header=None)
df2 = pd.read_csv("EMG\EMG_2.csv", header=None)
df3 = pd.read_csv("EMG\EMG_3.csv", header=None)
dataset = pd.concat([df0,df1,df2,df3], axis = 0)
# +
# Display What Each Label Refers To
import matplotlib.image as mpimg
img0 = mpimg.imread('EMG/0.jpg')
img1 = mpimg.imread('EMG/1.jpg')
img2 = mpimg.imread('EMG/2.jpg')
img3 = mpimg.imread('EMG/3.jpg')
fig = plt.figure()
plt.subplot(2, 2, 1)
plt.imshow(img0)
plt.axis('off')
plt.title('0 - rock')
plt.subplot(2, 2, 2)
plt.imshow(img1)
plt.axis('off')
plt.title('1 - scissors')
plt.subplot(2, 2, 3)
plt.imshow(img2)
plt.axis('off')
plt.title('2 - paper')
plt.subplot(2, 2, 4)
plt.imshow(img3)
plt.axis('off')
plt.title('3 - okay')
plt.show()
# -
# Split features and targets - X: Features, y: Targets
X = dataset.iloc[:, :-1]
y = dataset.iloc[:, -1].values
# +
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
# Save Test Set
np.savetxt("features_emg_test.csv", X_test, delimiter=",")
np.savetxt("targets_emg_test.csv", y_test, delimiter=",")
# -
# Standardize features by removing the mean and scaling to unit variance
from sklearn.preprocessing import StandardScaler
X_train = pd.DataFrame(StandardScaler().fit_transform(X_train))
X_test = pd.DataFrame(StandardScaler().fit_transform(X_test))
# # Classification Models
# +
# Some Functions for Showing the Classifier Performance
from sklearn.metrics import classification_report
from sklearn.metrics import plot_confusion_matrix
def classifier_performance(model,y_pred):
print('Classification Report: \n', classification_report(y_test,y_pred))
# Plot normalized confusion matrix
titles_options = [("Confusion matrix, without normalization", None),
("Normalized confusion matrix", 'true')]
for title, normalize in titles_options:
disp = plot_confusion_matrix(model, X_test, y_test,
display_labels=['rock','scissors','paper','ok'],
cmap=plt.cm.Blues,
normalize=normalize)
disp.ax_.set_title(title)
print(title)
print(disp.confusion_matrix)
plt.show()
return
# -
# ### Decision Trees Classification
# +
from sklearn.tree import DecisionTreeClassifier
MODEL_DT = DecisionTreeClassifier()
# Train the Model
MODEL_DT.fit(X_train,y_train)
# Save the Trained Model
pickle.dump(MODEL_DT, open('MODEL_CLASSIFICATION_DT.pkl', 'wb'))
# +
# Evaluate the Trained Model
# Predict the Trained Model on our Test data
y_pred_DT = MODEL_DT.predict(X_test)
# Print the Classification Report and Confusion Matrix
classifier_performance(MODEL_DT,y_pred_DT)
# -
# ### K Nearest Neighbors Classification
# +
from sklearn.neighbors import KNeighborsClassifier
MODEL_KNN = KNeighborsClassifier(n_neighbors=5)
# Train the Model
MODEL_KNN.fit(X_train,y_train)
# Save the Trained Model
pickle.dump(MODEL_KNN, open('MODEL_CLASSIFICATION_KNN.pkl', 'wb'))
# +
# Evaluate the Trained Model
# Predict the Trained Model on our Test data
y_pred_KNN = MODEL_KNN.predict(X_test)
# Print the Classification Report and Confusion Matrix
classifier_performance(MODEL_KNN,y_pred_KNN)
# -
# ### Gaussian Naive Bayes Classification
# +
from sklearn.naive_bayes import GaussianNB
MODEL_GNB = GaussianNB()
# Train the Model
MODEL_GNB.fit(X_train,y_train)
# Save the Trained Model
pickle.dump(MODEL_GNB, open('MODEL_CLASSIFICATION_GNB.pkl', 'wb'))
# +
# Evaluate the Trained Model
# Predict the Trained Model on our Test data
y_pred_GNB = MODEL_GNB.predict(X_test)
# Print the Classification Report and Confusion Matrix
classifier_performance(MODEL_GNB,y_pred_GNB)
# -
# ### Support Vector Machines
# +
from sklearn import svm
MODEL_SVM = svm.SVC()
# Train the Model
MODEL_SVM.fit(X_train,y_train)
# Save the Trained Model
pickle.dump(MODEL_SVM, open('MODEL_CLASSIFICATION_SVM.pkl', 'wb'))
# +
# Evaluate the Trained Model
# Predict the Trained Model on our Test data
y_pred_SVM = MODEL_SVM.predict(X_test)
# Print the Classification Report and Confusion Matrix
classifier_performance(MODEL_SVM,y_pred_SVM)
| LAB_2/MSE491_Lab2_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import scipy as stats
import seaborn as sns
import pandas_profiling
from sklearn import tree
from sklearn import ensemble
from sklearn import metrics
import matplotlib.pyplot as plt
from sklearn.ensemble import BaggingClassifier
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import cross_val_score
data=pd.read_csv("/home/manikanta/Documents/ML/classification/Ennsembling_learning/creditcard.csv")
data.head()
data.tail()
data.describe()
data.isnull().sum()
sns.heatmap(data.isnull(),cbar=False,cmap='viridis')
# +
#sns.pairplot(data)
# +
#pandas_profiling.ProfileReport(data)
# -
from sklearn.model_selection import train_test_split
x=data.drop('Class',axis=1)
y=data['Class']
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.20)
print(x_train.shape)
print(y_train.shape)
print(x_test.shape)
print(y_test.shape)
#SVM ALgorithm
from sklearn.svm import SVC
clf=SVC()
clf.fit(x,y)
trine_score=clf.score(x_train,y_train)
trine_score
test_score=clf.score(x_test,y_test)
test_score
yhat=clf.predict(x_test)
pd.DataFrame({'Acutual Data':y_test,'New_prediction':yhat})
# +
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.metrics import accuracy_score, classification_report, precision_score, recall_score
from sklearn.metrics import confusion_matrix, precision_recall_curve, roc_auc_score, roc_curve, auc, log_loss
print(confusion_matrix(y_test,yhat))
print(classification_report(y_test,yhat))
# -
#Decision Tree Algorithm
x1=data.drop('Class',axis=1)
y1=data['Class']
x_train1,x_test1,y_train1,y_test1=train_test_split(x1,y1,test_size=0.20)
print(x_train1.shape)
print(y_train1.shape)
print(x_test1.shape)
print(y_test1.shape)
from sklearn import tree
clf1 = tree.DecisionTreeClassifier()
clf1 = clf.fit(x1,y1)
clf1
trine_score=clf.score(x_train1,y_train1)
trine_score
test_score=clf.score(x_test1,y_test1)
test_score
yhat1=clf1.predict(x_test1)
pd.DataFrame({'Acutual Data':y_test1,'New_prediction':yhat1})
print(confusion_matrix(y_test1,yhat1))
print(classification_report(y_test1,yhat1))
tree.plot_tree(clf1)
#AdaBoosting Algorithm
x2=data.drop('Class',axis=1)
y2=data['Class']
x_train2,x_test2,y_train2,y_test2=train_test_split(x2,y2,test_size=0.20)
print(x_train2.shape)
print(y_train2.shape)
print(x_test2.shape)
print(y_test2.shape)
from sklearn import model_selection
from sklearn.ensemble import AdaBoostClassifier
clf2 = AdaBoostClassifier(tree.DecisionTreeClassifier(random_state=0))
clf2.fit(x2,y2)
clf2
trine_score=clf2.score(x_train2,y_train2)
trine_score
test_score=clf2.score(x_test2,y_test2)
test_score
yhat2=clf2.predict(x_test2)
pd.DataFrame({'Acutual Data':y_test2,'New_prediction':yhat2})
print(confusion_matrix(y_test2,yhat2))
print(classification_report(y_test2,yhat2))
#RandomForest Algorithm
x3=data.drop('Class',axis=1)
y3=data['Class']
x_train3,x_test3,y_train3,y_test3=train_test_split(x3,y3,test_size=0.20)
from sklearn.ensemble import RandomForestClassifier
clf3 = RandomForestClassifier()
clf3.fit(x3, y3)
clf3
trine_score=clf3.score(x_train3,y_train3)
trine_score
test_score=clf3.score(x_test3,y_test3)
test_score
yhat3=clf3.predict(x_test3)
pd.DataFrame({'Acutual Data':y_test3,'New_prediction':yhat3})
print(confusion_matrix(y_test3,yhat3))
print(classification_report(y_test3,yhat3))
| Ensemble Learning/DT_AB_RF_Train_Test_Splite_Credicard_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python TensorFlow 1.15
# language: python
# name: tf1.1_py36
# ---
# # Licences / Notes
# + colab={} colab_type="code" id="j0a4mTk9o1Qg"
# Copyright 2019 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# +
#Adapted by <NAME> in November,2019 from this Colab notebook:
#https://colab.research.google.com/github/google-research/bert/blob/master/predicting_movie_reviews_with_bert_on_tf_hub.ipynb.
#Changes includes
# - Reading our stressor data and parsing it properly
# - reconfiguring the last layer to include N neurons corresponding to N categories
# - correcting the probability output so that it follows [0,1] proper pattern
# - better analysis with confusion matrix
# - exporting to pb format for tensorflow serving api
# + [markdown] colab_type="text" id="xiYrZKaHwV81"
# Intro:
#
# If you’ve been following Natural Language Processing over the past year, you’ve probably heard of BERT: Bidirectional Encoder Representations from Transformers. It’s a neural network architecture designed by Google researchers that’s totally transformed what’s state-of-the-art for NLP tasks, like text classification, translation, summarization, and question answering.
#
# Now that BERT's been added to [TF Hub](https://www.tensorflow.org/hub) as a loadable module, it's easy(ish) to add into existing Tensorflow text pipelines. In an existing pipeline, BERT can replace text embedding layers like ELMO and GloVE. Alternatively, [finetuning](http://wiki.fast.ai/index.php/Fine_tuning) BERT can provide both an accuracy boost and faster training time in many cases.
#
# Some code was adapted from [this colab notebook](https://colab.sandbox.google.com/github/tensorflow/tpu/blob/master/tools/colab/bert_finetuning_with_cloud_tpus.ipynb). Let's get started!
# -
# # Loading Libraries
os.environ['LD_LIBRARY_PATH'] = '/usr/local/cuda-10.0/lib64:/usr/local/cuda-10.0/lib'
#os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = "0" #(or "1" or "2")
import sys
print(sys.executable)
# +
#export LD_LIBRARY_PATH=/usr/local/cuda-10.0/lib64:/usr/local/cuda-10.0/lib
#export CUDA_VISIBLE_DEVICES=0
# + colab={} colab_type="code" id="hsZvic2YxnTz"
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedShuffleSplit
import pandas as pd
import tensorflow as tf
import tensorflow_hub as hub
from datetime import datetime
import matplotlib.pyplot as plt
from sklearn.utils.multiclass import unique_labels
from sklearn.metrics import f1_score,confusion_matrix,classification_report,accuracy_score
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_colwidth', 1000)
# -
print(tf.__version__) #needs to be version 1.15.0, version 2.0 doesn't work with this notebook
# +
config = tf.ConfigProto()
#config.graph_options.optimizer_options.global_jit_level = tf.OptimizerOptions.ON_1
#config.gpu_options.visible_device_list="0"
from tensorflow.python.client import device_lib
device_lib.list_local_devices()
# + [markdown] colab_type="text" id="cp5wfXDx5SPH"
# In addition to the standard libraries we imported above, we'll need to install BERT's python package.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="jviywGyWyKsA" outputId="166f3005-d219-404f-b201-2a0b75480360"
# #!pip install bert-tensorflow
# + colab={} colab_type="code" id="hhbGEfwgdEtw"
import bert
from bert import run_classifier_with_tfhub
from bert import optimization
from bert import tokenization
import numpy as np
# + [markdown] colab_type="text" id="KVB3eOcjxxm1"
# Below, we'll set an output directory location to store our model output and checkpoints. This can be a local directory, in which case you'd set OUTPUT_DIR to the name of the directory you'd like to create.
#
#
# Set DO_DELETE to rewrite the OUTPUT_DIR if it exists. Otherwise, Tensorflow will load existing model checkpoints from that directory (if they exist).
# -
# ## Utils functions
# +
def f(x):
n = 2 # index of the second proability to get labeled
index = np.argsort(x.values.flatten().tolist())[-n:][0]
print(f"index is {index}")
label = label_list_text[index]
print(f"label is {label}")
return label
def manual_eval_metrics(df):
df["SD"] = df[label_list_text].std(axis=1)
df["SD-THRESHOLD"] = df[label_list_text].max(axis=1) - 2*df["SD"]
df["Above SD-THRESHOLD"] = df.apply(lambda x: len(np.where(x[label_list_text]>x["SD-THRESHOLD"])[0]),axis=1)
df["Branch"] = df.apply(lambda x: 1 if x["Above SD-THRESHOLD"]>1 else 0,axis=1)
return df
final_columns = ["sOrder","Input.text","is_stressor","is_stressor_conf","top_label","second_label","Branch", "Above SD-THRESHOLD","SD-THRESHOLD","SD","Other","Everyday Decision Making","Work","Social Relationships","Financial Problem","Health, Fatigue, or Physical Pain","Emotional Turmoil","Family Issues","School","avg_severity","median_severity","SD_severity","Votes","Source"]
def get_test_experiment_df(test):
test_predictions = [x[0]['probabilities'] for x in zip(getListPrediction(in_sentences=list(test[DATA_COLUMN])))]
test_live_labels = np.array(test_predictions).argmax(axis=1)
test[LABEL_COLUMN_RAW] = [label_list_text[x] for x in test_live_labels] # appending the labels to the dataframe
probabilities_df_live = pd.DataFrame(test_predictions) # creating a proabilities dataset
probabilities_df_live.columns = [x for x in label_list_text] # naming the columns
probabilities_df_live['second_label'] = probabilities_df_live.apply(lambda x:f(x),axis=1)
#print(test)
#label_df = create_examples_prediction(test)
#label_df.columns = label_list_text
#label_df['label 2'] = label_df.apply(lambda x:f(x),axis=1)
test.reset_index(inplace=True,drop=True) # resetting index
test_removed_columns = list(set(test.columns)-set(probabilities_df_live.columns))
test_temp = test[test_removed_columns]
experiment_df = pd.concat([test_temp,probabilities_df_live],axis=1, ignore_index=False)
missing_cols = list(set(experiment_df.columns)-set(final_columns))
experiment_df[missing_cols] = np.nan#.loc[:, missing_cols] = np.nan
experiment_df = experiment_df.reindex(columns = final_columns)
experiment_df = manual_eval_metrics(experiment_df)
#experiment_df = experiment_df.reindex(sorted(experiment_df.columns), axis=1)
return test,experiment_df
# -
def getListPrediction(in_sentences):
#1
input_examples = [bert.run_classifier.InputExample(guid="", text_a = x, text_b = None, label = 0) for x in in_sentences] # here, "" is just a dummy label
#2
input_features = bert.run_classifier.convert_examples_to_features(input_examples, label_list, MAX_SEQ_LENGTH, tokenizer)
#3
predict_input_fn = bert.run_classifier.input_fn_builder(features=input_features, seq_length=MAX_SEQ_LENGTH, is_training=False, drop_remainder=False)
print(input_features[0].input_ids)
#4
predictions = estimator.predict(input_fn=predict_input_fn,yield_single_examples=True)
return predictions
# +
is_normalize_active=False
def get_confusion_matrix(y_test,predicted,labels):
class_names=labels
# plotting confusion matrix
np.set_printoptions(precision=2)
# Plot non-normalized confusion matrix
plot_confusion_matrix(y_test, predicted, classes=class_names,
title='Confusion matrix, without normalization')
# Plot normalized confusion matrix
plot_confusion_matrix(y_test, predicted, classes=class_names, normalize=True,
title='Normalized confusion matrix')
plt.show()
def plot_confusion_matrix(y_true, y_pred, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if not title:
if normalize:
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
# Compute confusion matrix
cm = confusion_matrix(y_true, y_pred)
# Only use the labels that appear in the data
classes =classes
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
#print("Normalized confusion matrix")
else:
test =1
#print('Confusion matrix, without normalization')
fig, ax = plt.subplots()
im = ax.imshow(cm, interpolation='nearest', cmap=cmap)
#ax.figure.colorbar(im, ax=ax)
# We want to show all ticks...
ax.set(xticks=np.arange(cm.shape[1]),
yticks=np.arange(cm.shape[0]),
# ... and label them with the respective list entries
xticklabels=classes, yticklabels=classes,
title=title,
ylabel='True label',
xlabel='Predicted label')
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i in range(cm.shape[0]):
for j in range(cm.shape[1]):
ax.text(j, i, format(cm[i, j], fmt),
ha="center", va="center",
color="white" if cm[i, j] > thresh else "black")
#fig.tight_layout()
return ax
# -
# # Loading the data
def data_prep_bert(df,test_size):
print("Filling missing values")
df[DATA_COLUMN] = df[DATA_COLUMN].fillna('_NA_')
print("Splitting dataframe with shape {} into training and test datasets".format(df.shape))
X_train, X_test = train_test_split(df, test_size=test_size, random_state=2018,stratify = df[LABEL_COLUMN_RAW])
return X_train, X_test
def open_dataset(NAME,mapping_index,excluded_categories):
df = pd.read_csv(PATH+NAME+'.csv',sep =',')
df.head(10)
df = df[df[LABEL_COLUMN_RAW].notna()]
#df.columns = [LABEL_COLUMN_RAW,'Severity',DATA_COLUMN,'Source']
if excluded_categories is not None:
for category in excluded_categories:
df = df[df[LABEL_COLUMN_RAW] !=category]
label_list=[]
label_list_final =[]
if(mapping_index is None):
df[LABEL_COLUMN_RAW] = df[LABEL_COLUMN_RAW].astype('category')
df[LABEL_COLUMN], mapping_index = pd.Series(df[LABEL_COLUMN_RAW]).factorize() #uses pandas factorize() to convert to numerical index
else:
df[LABEL_COLUMN] = df[LABEL_COLUMN_RAW].apply(lambda x: mapping_index.get_loc(x))
label_list_final = [None] * len(mapping_index.categories)
label_list_number = [None] * len(mapping_index.categories)
for index,ele in enumerate(list(mapping_index.categories)):
lindex = mapping_index.get_loc(ele)
label_list_number[lindex] = lindex
label_list_final[lindex] = ele
frequency_dict = df[LABEL_COLUMN_RAW].value_counts().to_dict()
df["class_freq"] = df[LABEL_COLUMN_RAW].apply(lambda x: frequency_dict[x])
return df,mapping_index,label_list_number,label_list_final
# # Require user changes > Start Here
# ### Experiment Name
PATH = './datasets/'
TODAY_DATE = "31_07_2020/"
EXPERIMENT_NAME = 'ryukas_single_label_big_data'
EXPERIMENTS_PATH = PATH + 'experiments/'+TODAY_DATE+EXPERIMENT_NAME
if not os.path.exists(PATH + 'experiments/'+TODAY_DATE):
os.mkdir(PATH + 'experiments/'+TODAY_DATE)
if not os.path.exists(EXPERIMENTS_PATH):
os.mkdir(EXPERIMENTS_PATH)
# ### Model Hyperparameters
# + colab={} colab_type="code" id="OjwJ4bTeWXD8"
# Compute train and warmup steps from batch size
# These hyperparameters are copied from this colab notebook (https://colab.sandbox.google.com/github/tensorflow/tpu/blob/master/tools/colab/bert_finetuning_with_cloud_tpus.ipynb)
BATCH_SIZE = 16
LEARNING_RATE = 2e-5
NUM_TRAIN_EPOCHS = 3.0
# Warmup is a period of time where hte learning rate
# is small and gradually increases--usually helps training.
WARMUP_PROPORTION = 0.1
# Model configs
SAVE_CHECKPOINTS_STEPS = 1000
SAVE_SUMMARY_STEPS = 100
# We'll set sequences to be at most 32 tokens long.
MAX_SEQ_LENGTH = 32
OUTPUT_DIR = './models/'+EXPERIMENT_NAME+ '/' #_01_04_2020/
# +
DATASET_NAME = 'Filtered_df_everything'
DATA_COLUMN = 'Answer.Stressor'
LABEL_COLUMN_RAW = 'top_label'#'Answer.Label'
LABEL_COLUMN = 'label_numeric'
#dataset,mapping_index,label_list, label_list_text = open_dataset('mturk900balanced',None)
EXCLUDED_CATEGORIES = None #['Other'] #None # # if nothing to exclude put None, THIS ALWAYS MUST BE A LIST
mapping_dict = {'Other': 0, 'Everyday Decision Making': 1, 'Work': 2, 'Social Relationships': 3, 'Financial Problem': 4, 'Emotional Turmoil': 5, 'Health, Fatigue, or Physical Pain': 6, 'School': 7, 'Family Issues': 8}#,'Not Stressful':9}
mapping_index = pd.CategoricalIndex([key for key,value in mapping_dict.items()])
dataset,mapping_index,label_list, label_list_text = open_dataset(DATASET_NAME,mapping_index,EXCLUDED_CATEGORIES)
test_on_mturk_and_popbots_live = True # include live data in training + include mturk in testing
if test_on_mturk_and_popbots_live:
mturk = dataset[dataset['Source']== 'mTurk']
live = dataset[dataset['Source']== 'Popbots']
live = live.sample(frac=1).reset_index(drop=True) # shuffle live
TEST_PERCENTAGE = len(live)/(2*len(mturk)) # given to set the percentage of mturk used as test set to have 50/50
print(f"Test percentage is {TEST_PERCENTAGE}")
train,test = data_prep_bert(mturk,TEST_PERCENTAGE) # test size from mturk
train = train.append(live.loc[0:int(len(live)/2)]) # taking 1/2 of that dataset for training
test = test.append(live.loc[int(len(live)/2):int(len(live))]) # taking 1/2 of live dataset for testing
else:
# or taking live only for testing
train,test = dataset[dataset['Source']== 'mTurk'],dataset[dataset['Source']== 'Popbots']
#train,test = data_prep_bert(dataset,0.2)
#print(f"Dataset has {len(dataset)} training examples")
print(f"Normal label list is {label_list}")
print(f"The labels text is {label_list_text}")
print(f"test size is{len(test)}")
print(f"train size is{len(train)}")
#Export train test to csv
#train.to_csv(PATH+'900_CSV_SPLITTED/train.csv')
#test.to_csv(PATH+'900_CSV_SPLITTED/test.csv')
# +
DATA_COLUMN = 'Input.text'
LABEL_COLUMN_RAW = 'top_label'#'Answer.Label'
LABEL_COLUMN = 'label_numeric'
MTURK_NAME = 'mTurk_synthetic'
LIVE_NAME = 'popbots_live'
INQUIRE_NAME = 'Inquire'
MTURK_COVID_NAME = 'mTurk_synthetic_covid'
LABEL_HOT_VECTOR = 'label_conf'
#train.to_csv(EXPERIMENTS_PATH+'/TRAIN_'+DATASET_NAME+'.csv')
#test.to_csv(EXPERIMENTS_PATH+'/TEST_'+DATASET_NAME+'.csv')
#train = pd.read_csv('./datasets/experiments/29_07_2020/ryukas_evaluation/TRAIN_2020-07-27-MainTurkAggregation-5-Turkers_v0.csv')
test = pd.read_csv('./datasets/experiments/29_07_2020/ryukas_evaluation/TEST_2020-07-27-MainTurkAggregation-5-Turkers_v0.csv')
#test_down.to_csv(EXPERIMENTS_PATH+'/SMALL_TEST_'+DATASET_NAME+'.csv')
test_down = pd.read_csv('./datasets/experiments/29_07_2020/ryukas_evaluation/SMALL_TEST_2020-07-27-MainTurkAggregation-5-Turkers_v0.csv')
train = pd.read_csv('./datasets/experiments/29_07_2020/ryukas_evaluation/SMALL_TRAIN_2020-07-27-MainTurkAggregation-5-Turkers_v0.csv')
# -
train = train.sample(frac=1).reset_index(drop=True) #reshuffle everything
test = test.sample(frac=1).reset_index(drop=True)
# ### Train set and test set analysis
def print_dataset_info(train,test):
print(f"Train size {len(train)} with {len(train[train['Source']== 'Popbots'])} from Popbots and {len(train[train['Source']== 'mTurk'])} from mturk")
print(f"Test size {len(test)} with {len(test[test['Source']== 'Popbots'])} from Popbots and {len(test[test['Source']== 'mTurk'])} from mturk")
print('\nTraining distribution:')
print(pd.pivot_table(train[[LABEL_COLUMN_RAW, 'Source']],index=[LABEL_COLUMN_RAW, 'Source'],columns=None, aggfunc=len)) #.to_clipboard(excel=True)
print('\nTesting distribution:')
print(pd.pivot_table(test[[LABEL_COLUMN_RAW, 'Source']],index=[LABEL_COLUMN_RAW, 'Source'],columns=None, aggfunc=len)) #.to_clipboard(excel=True)
train = train.sample(frac=1).reset_index(drop=True) #reshuffle everything
test = test.sample(frac=1).reset_index(drop=True)
print('\nAll dataset distribution:')
print(pd.pivot_table(dataset[[LABEL_COLUMN_RAW, 'Source']],index=[LABEL_COLUMN_RAW, 'Source'],columns=None, aggfunc=len)) #.to_clipboard(excel=T
print_dataset_info(train,test)
# ### Step to reduce the most dominant categories and balance the dataset
# +
sampling_cutoff = 100 # all the categories which had less than 100 example won't be sampled down
total_training_size = 1501
REVERSE_FREQ = 'Max_reverse_sampling_chance'
train[REVERSE_FREQ] = train['class_freq'].apply(lambda x: (max(train['class_freq'])/x)*(max(train['class_freq'])/x))
sampling_boolean = (train['Source'] != 'Popbots') & (train['class_freq'].astype(float) > sampling_cutoff)
train_to_be_balanced = train[sampling_boolean]
train_not_resampled = train[~sampling_boolean]
train_temp = train_to_be_balanced.sample(n=(total_training_size-len(train_not_resampled)), weights=REVERSE_FREQ, random_state=2020)
train = pd.concat([train_temp,train_not_resampled])
# -
print_dataset_info(train,test)
mapping_index.categories
train.to_csv(EXPERIMENTS_PATH+'/TRAIN_'+DATASET_NAME+'.csv')
test.to_csv(EXPERIMENTS_PATH+'/TEST_'+DATASET_NAME+'.csv')
# # Require user changes > STOP Here
# # Data Preprocessing
# + [markdown] colab_type="text" id="sfRnHSz3iSXz"
# For us, our input data is the 'sentence' column and our label is the 'polarity' column
# + [markdown] colab_type="text" id="V399W0rqNJ-Z"
# #Data Preprocessing
# We'll need to transform our data into a format BERT understands. This involves two steps. First, we create `InputExample`'s using the constructor provided in the BERT library.
#
# - `text_a` is the text we want to classify, which in this case, is the `Request` field in our Dataframe.
# - `text_b` is used if we're training a model to understand the relationship between sentences (i.e. is `text_b` a translation of `text_a`? Is `text_b` an answer to the question asked by `text_a`?). This doesn't apply to our task, so we can leave `text_b` blank.
# - `label` is the label for our example, i.e. True, False
# + colab={} colab_type="code" id="p9gEt5SmM6i6"
# Use the InputExample class from BERT's run_classifier code to create examples from the data
train_InputExamples = train.apply(lambda x: bert.run_classifier.InputExample(guid=None, # Globally unique ID for bookkeeping, unused in this example
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
test_InputExamples = test.apply(lambda x: bert.run_classifier.InputExample(guid=None,
text_a = x[DATA_COLUMN],
text_b = None,
label = x[LABEL_COLUMN]), axis = 1)
# + [markdown] colab_type="text" id="SCZWZtKxObjh"
# Next, we need to preprocess our data so that it matches the data BERT was
#
# 1. List item
# 2. List item
#
# trained on. For this, we'll need to do a couple of things (but don't worry--this is also included in the Python library):
#
#
# 1. Lowercase our text (if we're using a BERT lowercase model)
# 2. Tokenize it (i.e. "sally says hi" -> ["sally", "says", "hi"])
# 3. Break words into WordPieces (i.e. "calling" -> ["call", "##ing"])
# 4. Map our words to indexes using a vocab file that BERT provides
# 5. Add special "CLS" and "SEP" tokens (see the [readme](https://github.com/google-research/bert))
# 6. Append "index" and "segment" tokens to each input (see the [BERT paper](https://arxiv.org/pdf/1810.04805.pdf))
#
# Happily, we don't have to worry about most of these details.
#
#
#
# + [markdown] colab_type="text" id="qMWiDtpyQSoU"
# To start, we'll need to load a vocabulary file and lowercasing information directly from the BERT tf hub module:
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="IhJSe0QHNG7U" outputId="20b28cc7-3cb3-4ce6-bfff-a7847ce3bbaa"
# This is a path to an uncased (all lowercase) version of BERT
BERT_MODEL_HUB = "https://tfhub.dev/google/bert_uncased_L-12_H-768_A-12/1"
def create_tokenizer_from_hub_module():
"""Get the vocab file and casing info from the Hub module."""
with tf.Graph().as_default():
bert_module = hub.Module(BERT_MODEL_HUB)
tokenization_info = bert_module(signature="tokenization_info", as_dict=True)
with tf.Session() as sess:
vocab_file, do_lower_case = sess.run([tokenization_info["vocab_file"],
tokenization_info["do_lower_case"]])
return bert.tokenization.FullTokenizer(
vocab_file=vocab_file, do_lower_case=do_lower_case)
tokenizer = create_tokenizer_from_hub_module()
# + [markdown] colab_type="text" id="z4oFkhpZBDKm"
# Great--we just learned that the BERT model we're using expects lowercase data (that's what stored in tokenization_info["do_lower_case"]) and we also loaded BERT's vocab file. We also created a tokenizer, which breaks words into word pieces:
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="dsBo6RCtQmwx" outputId="9af8c917-90ec-4fe9-897b-79dc89ca88e1"
tokenizer.tokenize("This here's an example of using the BERT tokenizer")
# + [markdown] colab_type="text" id="0OEzfFIt6GIc"
# Using our tokenizer, we'll call `run_classifier.convert_examples_to_features` on our InputExamples to convert them into features BERT understands.
# + colab={"base_uri": "https://localhost:8080/", "height": 1261} colab_type="code" id="LL5W8gEGRTAf" outputId="65001dda-155b-48fc-b5fc-1e4cabc8dfbf"
# Convert our train and test features to InputFeatures that BERT understands.
train_features = bert.run_classifier.convert_examples_to_features(train_InputExamples, label_list, MAX_SEQ_LENGTH, tokenizer)
test_features = bert.run_classifier.convert_examples_to_features(test_InputExamples, label_list, MAX_SEQ_LENGTH, tokenizer)
# -
# # Creating a model
# + [markdown] colab_type="text" id="ccp5trMwRtmr"
# Now that we've prepared our data, let's focus on building a model. `create_model` does just this below. First, it loads the BERT tf hub module again (this time to extract the computation graph). Next, it creates a single new layer that will be trained to adapt BERT to our classification task. This strategy of using a mostly trained model is called [fine-tuning](http://wiki.fast.ai/index.php/Fine_tuning).
# -
# To understand the `pooled ouput` vs `sequence output` refer to https://www.kaggle.com/questions-and-answers/86510
# + colab={} colab_type="code" id="6o2a5ZIvRcJq"
def create_model(is_predicting, input_ids, input_mask, segment_ids, labels,
num_labels):
"""Creates a classification model."""
bert_module = hub.Module(
BERT_MODEL_HUB,
trainable=True) # fined tuning the complete weights of all the model
bert_inputs = dict(
input_ids=input_ids,
input_mask=input_mask,
segment_ids=segment_ids)
bert_outputs = bert_module(
inputs=bert_inputs,
signature="tokens",
as_dict=True)
# Use "pooled_output" for classification tasks on an entire sentence.
# Use "sequence_outputs" for token-level output.
output_layer = bert_outputs["pooled_output"] # 768 dimention vector
hidden_size = output_layer.shape[-1].value
# Create our own layer to tune for politeness data.
output_weights = tf.get_variable(
"output_weights", [num_labels, hidden_size],
initializer=tf.truncated_normal_initializer(stddev=0.02))
output_bias = tf.get_variable(
"output_bias", [num_labels], initializer=tf.zeros_initializer())
with tf.variable_scope("loss"):
# Dropout helps prevent overfitting
output_layer = tf.nn.dropout(output_layer, keep_prob=0.9)
# does the Ax multiplication
logits = tf.matmul(output_layer, output_weights, transpose_b=True)
# add the bias eg: Ax+B
logits = tf.nn.bias_add(logits, output_bias)
########################### HERE ADDITIONAL LAYERS CAN BE ADDED ######################
# compute the log softmax for each neurons/logit
#log_probs = tf.nn.log_softmax(logits, axis=-1)
#compute the normal softmax to get the probabilities
#probs = tf.nn.softmax(logits, axis=-1)
# Convert labels into one-hot encoding
#one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
#classes_weights = tf.constant([1.0,1.0,1.0,1.0,1.0,1.0,0.7], dtype=tf.float32)
#sample_weights = tf.multiply(one_hot_labels, classes_weights)
# probabilities = tf.nn.softmax(logits, axis=-1) ### multiclass case
probabilities = tf.nn.sigmoid(logits)#### multi-label case
#labels = tf.cast(labels, tf.float32)
one_hot_labels = tf.one_hot(labels, depth=num_labels, dtype=tf.float32)
#tf.logging.info("num_labels:{};logits:{};labels:{}".format(num_labels, logits, labels))
per_example_loss = tf.nn.sigmoid_cross_entropy_with_logits(labels=one_hot_labels, logits=logits)
loss = tf.reduce_mean(per_example_loss)
#predicted_labels = tf.squeeze(tf.argmax(log_probs, axis=-1, output_type=tf.int32))
predicted_labels = tf.squeeze(tf.argmax(probabilities, axis=-1, output_type=tf.int32))
# If we're predicting, we want predicted labels and the probabiltiies.
if is_predicting:
return (predicted_labels, probabilities,probabilities)#log_probs,probs)
# If we're train/eval, compute loss between predicted and actual label
#per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
#per_example_loss = -tf.reduce_sum(one_hot_labels * log_probs, axis=-1)
#loss = tf.reduce_mean(per_example_loss)
return (loss, predicted_labels, probabilities)#log_probs)
# + [markdown] colab_type="text" id="qpE0ZIDOCQzE"
# Next we'll wrap our model function in a `model_fn_builder` function that adapts our model to work for training, evaluation, and prediction.
# + colab={} colab_type="code" id="FnH-AnOQ9KKW"
# model_fn_builder actually creates our model function
# using the passed parameters for num_labels, learning_rate, etc.
def model_fn_builder(num_labels, learning_rate, num_train_steps,
num_warmup_steps):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
label_ids = features["label_ids"]
is_predicting = (mode == tf.estimator.ModeKeys.PREDICT)
# TRAIN and EVAL
if not is_predicting:
(loss, predicted_labels, log_probs) = create_model(
is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
train_op = bert.optimization.create_optimizer(
loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu=False)
# Calculate evaluation metrics.
def metric_fn(label_ids, predicted_labels):
accuracy = tf.metrics.accuracy(label_ids, predicted_labels)
"""
f1_score = tf.contrib.metrics.f1_score(
label_ids,
predicted_labels)
auc = tf.metrics.auc(
label_ids,
predicted_labels)"""
recall = tf.metrics.recall(
label_ids,
predicted_labels)
precision = tf.metrics.precision(
label_ids,
predicted_labels)
true_pos = tf.metrics.true_positives(
label_ids,
predicted_labels)
true_neg = tf.metrics.true_negatives(
label_ids,
predicted_labels)
false_pos = tf.metrics.false_positives(
label_ids,
predicted_labels)
false_neg = tf.metrics.false_negatives(
label_ids,
predicted_labels)
return {
"eval_accuracy": accuracy,
#"f1_score": f1_score,
#"auc": auc,
"precision": precision,
"recall": recall,
"true_positives": true_pos,
"true_negatives": true_neg,
"false_positives": false_pos,
"false_negatives": false_neg
}
eval_metrics = metric_fn(label_ids, predicted_labels)
if mode == tf.estimator.ModeKeys.TRAIN:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
train_op=train_op)
else:
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
eval_metric_ops=eval_metrics)
else:
(predicted_labels, log_probs,probs) = create_model(
is_predicting, input_ids, input_mask, segment_ids, label_ids, num_labels)
predictions = {
'probabilities': probs#,
#'labels': predicted_labels
}
return tf.estimator.EstimatorSpec(mode, predictions=predictions)
# Return the actual model function in the closure
return model_fn
# + colab={} colab_type="code" id="emHf9GhfWBZ_"
# Compute # train and warmup steps from batch size
num_train_steps = int(len(train_features) / BATCH_SIZE * NUM_TRAIN_EPOCHS)
num_warmup_steps = int(num_train_steps * WARMUP_PROPORTION)
# + colab={} colab_type="code" id="oEJldMr3WYZa"
# Specify outpit directory and number of checkpoint steps to save
run_config = tf.estimator.RunConfig(
model_dir=OUTPUT_DIR,
save_summary_steps=SAVE_SUMMARY_STEPS,
save_checkpoints_steps=SAVE_CHECKPOINTS_STEPS)
# + colab={"base_uri": "https://localhost:8080/", "height": 156} colab_type="code" id="q_WebpS1X97v" outputId="1648932a-7391-49d3-8af7-52d514e226e8"
model_fn = model_fn_builder(
num_labels=len(label_list),
learning_rate=LEARNING_RATE,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps)
estimator = tf.estimator.Estimator(
model_fn=model_fn,
config=run_config,
params={"batch_size": BATCH_SIZE})
# + [markdown] colab_type="text" id="NOO3RfG1DYLo"
# Next we create an input builder function that takes our training feature set (`train_features`) and produces a generator. This is a pretty standard design pattern for working with Tensorflow [Estimators](https://www.tensorflow.org/guide/estimators).
# + colab={} colab_type="code" id="1Pv2bAlOX_-K"
# Create an input function for training. drop_remainder = True for using TPUs.
train_input_fn = bert.run_classifier.input_fn_builder(
features=train_features,
seq_length=MAX_SEQ_LENGTH,
is_training=True,
drop_remainder=False)
# + [markdown] colab_type="text" id="t6Nukby2EB6-"
# # Training the model
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="nucD4gluYJmK" outputId="5d728e72-4631-42bf-c48d-3f51d4b968ce"
print(f'Beginning Training!')
current_time = datetime.now()
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
print("Training took time ", datetime.now() - current_time)
# + [markdown] colab_type="text" id="CmbLTVniARy3"
# # Evaluating the model on Test Set
# + colab={} colab_type="code" id="JIhejfpyJ8Bx"
test_input_fn = bert.run_classifier.input_fn_builder(
features=test_features,
seq_length=MAX_SEQ_LENGTH,
is_training=False,
drop_remainder=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 445} colab_type="code" id="PPVEXhNjYXC-" outputId="dd5482cd-c558-465f-c854-ec11a0175316"
estimator.evaluate(input_fn=test_input_fn, steps=None)
# +
#fetching all the probabilities for each line of the test set
test_probabilities = [x[0]['probabilities'] for x in zip(estimator.predict(test_input_fn,yield_single_examples=True))]
#taking the argmex for the highest category
test_final_labels = np.array(test_probabilities).argmax(axis=1)
# -
# ### Classification Report
# +
def replace_by_multi_hot(array, threshold):
two_std = threshold*np.std(array)
array = [1 if x > two_std else 0 for x in array]
return array
y_true = pd.DataFrame(test[label_list_text]).apply(lambda x: replace_by_multi_hot(x,1),axis=1).to_list()
y_pred = pd.DataFrame(test_probabilities).apply(lambda x: replace_by_multi_hot(x,1),axis=1).to_list()
from sklearn.metrics import hamming_loss, accuracy_score , f1_score
print("accuracy_score:", accuracy_score(y_true, y_pred)) # exact match
print("f1_score:", f1_score(y_true, y_pred, average='weighted'))
print("Hamming_loss:", hamming_loss(y_true, y_pred))
print("Exact match:",np.all(np.array(y_pred) == np.array(y_true), axis=1).mean())
print("Hamming_score:", (np.array(y_pred) == np.array(y_true)).mean())
# -
for i in np.arange(0.5, 4, 0.1):
y_true = pd.DataFrame(test[label_list_text]).apply(lambda x: replace_by_multi_hot(x,i),axis=1).to_list()
y_pred = pd.DataFrame(test_probabilities).apply(lambda x: replace_by_multi_hot(x,i),axis=1).to_list()
print(f"For {i:.2f} std, hamming lost of {hamming_loss(y_true, y_pred):.2f} weighted f1 of {f1_score(y_true, y_pred, average='weighted'):.2f} accuracy of {accuracy_score(y_true, y_pred):.2f}")
# +
report = pd.DataFrame(classification_report(list(test[LABEL_COLUMN]),list(test_final_labels),zero_division=0, output_dict=True)).T
print(report)
# -
# ### Confusion Matrix
get_confusion_matrix(y_test=test[LABEL_COLUMN],predicted=test_final_labels,labels=label_list_text)
# ### Exporting test set with probabilities
# +
test, experiment_df = get_test_experiment_df(test)
experiment_df.to_csv(EXPERIMENTS_PATH+'/test_with_probabilities.csv')
test_down, experiment_df = get_test_experiment_df(test_down)
experiment_df.to_csv(EXPERIMENTS_PATH+'/testdown_with_probabilities.csv')
# -
experiment_df
# ### RUN ALL CELLS ABOVE ON HERE
experiment_df[experiment_df['Predicted'] != experiment_df['Answer.Label']].head(10) # change head(n) to see more
# # Exporting the model as Pb format
def export_model(dir_path):
MAX_SEQ_LEN = 128
def serving_input_receiver_fn():
"""An input receiver that expects a serialized tf.Example."""
reciever_tensors = {
"input_ids": tf.placeholder(dtype=tf.int32,
shape=[1, MAX_SEQ_LEN])
}
features = {
"label_ids":tf.placeholder(tf.int32, [None], name='label_ids'),
"input_ids": reciever_tensors['input_ids'],
"input_mask": 1 - tf.cast(tf.equal(reciever_tensors['input_ids'], 0), dtype=tf.int32),
"segment_ids": tf.zeros(dtype=tf.int32,
shape=[1, MAX_SEQ_LEN])
}
return tf.estimator.export.ServingInputReceiver(features, reciever_tensors)
estimator._export_to_tpu = False
estimator.export_saved_model(dir_path, serving_input_receiver_fn)
export_model('./tfmode/pbformat/')
# ## Getting analysis for a another dataset
# +
test_all_live = pd.read_csv(PATH+'PopbotsLive_TestSet_213.csv')
test_all_live, experiment_df_live = get_test_experiment_df(test_all_live)
# -
| bert-pipeline/Predicting_Stressor_with_BERT_on_TF_Hub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# 
#
# <center><h2>Scale your pandas workflows by changing one line of code</h2>
#
# # Exercise 5: Setting up cluster environment
#
# **GOAL**: Learn how to set up a cluster for Modin.
#
# **NOTE**: This exercise has extra requirements. Read instructions carefully before attempting.
#
# **This exercise instructs the user on how to start a 700+ core cluster, and it is not shut down until the end of Exercise 5. Read instructions carefully.**
# Often in practice we have a need to exceed the capabilities of a single machine. Modin works and performs well in both local mode and in a cluster environment. The key advantage of Modin is that your notebook does not change between local development and cluster execution. Users are not required to think about how many workers exist or how to distribute and partition their data; Modin handles all of this seamlessly and transparently.
#
# 
#
# **Extra Requirements for this exercise**
#
# Detailed instructions can be found here: https://docs.ray.io/en/latest/cluster/cloud.html
#
# From command line:
# - `pip install boto3`
# - `aws configure`
# - `ray up modin-cluster.yaml`
#
# Included in this directory is a file named [`modin-cluster.yaml`](https://github.com/modin-project/modin/blob/master/examples/tutorial/tutorial_notebooks/cluster/modin-cluster.yaml). We will use this to start the cluster.
# +
# # !pip install boto3
# +
# # !aws configure
# -
# ## Starting and connecting to the cluster
#
# This example starts 1 head node (m5.24xlarge) and 7 workers (m5.24xlarge), 768 total CPUs.
#
# Cost of this cluster can be found here: https://aws.amazon.com/ec2/pricing/on-demand/.
# +
# # !ray up modin-cluster.yaml
# -
# Connect to the cluster with `ray attach`
# +
# # !ray attach modin-cluster.yaml
# +
# DO NOT CHANGE THIS CODE!
# Changing this code risks breaking further exercises
import time
time.sleep(600) # We need to give ray enough time to start up all the workers
import ray
ray.init(address="auto")
from modin.config import NPartitions
assert NPartitions.get() == 768, "Not all Ray nodes are started up yet"
ray.shutdown()
# -
# ### Please move on to Exercise 6
| examples/tutorial/tutorial_notebooks/cluster/exercise_5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analysis
# +
# Import basic libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
pd.pandas.set_option('display.max_columns', None)
# +
# Load dataset
data = pd.read_csv('train.csv')
print(data.shape)
data.head(5)
# -
# data types in the dataset
data.info()
# Following the same order as proposed in the course we will identify:
#
# 1. Missing values
# 2. Numerical variables
# 3. Distribution of the numerical variables
# 4. Outliers
# 5. Categorical variables
# 6. Cardinality of the categorical variables
# 7. Potential relationship between the variables and the target: Survived
# ### Missing Values
# +
# Find the variables that contains missing values
vars_na = [var for var in data.columns if data[var].isnull().sum() > 0]
# Calculate the percentage of missing values
data[vars_na].isnull().mean()
# -
# Let's see what the impact is in our target variable *"Survived"* when considering the variables with missing data
# +
def impact_na_value (df, var):
# Copy the original dataset to avoid changes on it
df = df.copy()
# Create a variable with 1 if the observation is missing and 0 otherwise
df[var] = np.where(df[var].isnull(), 1, 0)
df.groupby(var)['Survived'].mean().plot.bar()
plt.title(var)
plt.show()
# Call the function for each variable
for var in vars_na:
impact_na_value(data, var)
# -
# In Age, and Cabin variables the missing values contribute to lower the chances of surviving in the Titanic. However, in the embarked variable missing values contributes to survival.
# ### Numerical Variables
#
# We can find the numerical variables in the dataset using data.info() but that method is limited when the dataset is low dimensional. The following method scales easily
# +
# Make a list of numberical variabes
num_var = [var for var in data.columns if data[var].dtypes != 'O']
print('Number of numerical variables: ', len(num_var))
# Visualise numerical variables
data[num_var].head()
# -
# We can get rid of *PassengerID* as it seems it is a unique indicator:
print('Number of Unique Ids: ', len(data['PassengerId'].unique()))
print('Number of passengers in Titanic: ', len(data))
# #### Numerical discrete variables
#
# Let's identify those numerical variable that exhibit a discrete behaviour, i.e., show a finite number of values
# +
# List of discrete variables
dis_var = [var for var in data.columns if data[var].dtypes != 'O'
and len(data[var].unique()) < 20 and var != 'Survived']
print('Number of numerical discrete variables: ', len(dis_var))
# Visualise numerical variables
data[dis_var].head()
# -
# Let's see the analysis of these discrete variables
# +
def dis_analysis(df, var):
df = df.copy()
df.groupby(var)['Survived'].mean().plot.bar()
plt.title(var)
plt.ylabel('Mean survival rate')
plt.show()
for var in dis_var:
dis_analysis(data, var)
# -
# The ticket class represented by *Pclass* clearly influences the likelihood of surviving in Titatic. First class more than doubles third class the likelihood of surviving.
#
# *SibSp* denotes the number of siblings/spouses aboard the Titanic. 1 and 2 number of siblings/spouses makes it more likely for the passenger to survive.
#
# *Parch* represents the number of parents/children in the Titanic. Having fewer children (up to 3) helped to survive.
# #### Numerical continious variables
#
# Let's do the same with continious variables this time.
# +
# List of continious variables
cont_var = [var for var in num_var if var not in dis_var and var != 'Survived'
and var != 'PassengerId']
print('Number of numerical continious variables: ', len(cont_var))
# Visualise numerical variables
data[cont_var].head()
# +
def cont_analysis(df, var):
df = df.copy()
df[var].hist(bins=30)
plt.ylabel('Number of passengers')
plt.xlabel(var)
plt.title(var)
plt.show()
for var in cont_var:
cont_analysis(data, var)
# -
# The variables are not normally distributed.
#
# The idea is to build a logistic regression model. This kind of model applies a sigmoid function to a linear combination of variables, therefore to optimize performance we want to work with normally distributed variables.
#
# Let's try a logarithmic transformation.
# +
def trans_cont_var(df, var):
df = df.copy()
# Log is not defined with 0 or negative values, so let's avoid those.
if any(df[var] <= 0):
pass
else:
df[var] = np.log(df[var])
df[var].hist(bins=30)
plt.ylabel('Number of passengers')
plt.title(var)
plt.show()
for var in cont_var:
trans_cont_var(data, var)
# -
# Log transformation can't be applied to *fare* variable as there are some negative values. On top of that, the age looks more similar to a normally distribution but there are extreme values out of the bulk of data.
# Let's see how is the relationship with the transformed *age* variable and the *survived* target variable.
# +
def cont_relationship(df, var):
df = df.copy()
if any(df[var] <=0):
pass
else:
df[var] = np.log(df[var])
plt.scatter(df[var], df['Survived'])
plt.ylabel('Survived')
plt.title(var)
plt.show()
for var in cont_var:
cont_relationship(data, var)
# -
# It is not clear from the scatter plot that age is a relevant factor in the survival of a passgener.
# ### Outliers
#
# Extreme values may affect the performance of a linear model. Therefore, it is necessary to discover if there are outliers in our data set by looking at the numerical continious variables
# +
def outliers(df, var):
df = df.copy()
if any(data[var] <= 0):
pass
else:
df[var] = np.log(df[var])
df.boxplot(column=var)
plt.ylabel(var)
plt.show()
for var in cont_var:
outliers(data, var)
# -
# The age variable seems to have some outliers, once it is transformed into a normal distribution. Let's take this into consideration when doing the feature engineering step.
# ### Categorical variables
#
# Let's analyse now the categorical variables in the dataset.
# +
# List of continious variables
cat_var = [var for var in data.columns if data[var].dtypes == 'O']
print('Number of categorical variables: ', len(cat_var))
# Visualise numerical variables
data[cat_var].head()
# -
# #### Cardinality of categorical variables
#
# Let's assess how many different classes are present in each of the variables
data[cat_var].nunique()
# *Sex* and *Embarked* variables exhibit low cardinality. Name is unique for each passenger so it makes sense that it is the same as the number of passengers. The fact that *Ticket* and *Cabin* are below 891 unique classes maybe due to several passengers sharing a group ticket and the same cabin.
# #### Rare labels
# Let's go ahead and explore now if there are labels present just in a few number of passengers. Although, using the insights so far it is reasonable to expect there won't be rare labels.
# +
def rare_labels(df, var, perc):
df = df.copy()
# compute the % of observations per category
rate = df.groupby(var)['Survived'].count()/len(df)
# return categories that are rare
return rate[rate < perc]
# print those categories that are present less that certain %
for var in cat_var:
print(rare_labels(data, var, 0.001))
print()
# -
# The results depend on the threshold established to consider "rare" categories. For instance, considering that we should consider as "rare" those observation that contain a unique cabin number does not make sense, so we will not change the labels of the categorical variables.
# Let's finish the data analysis with some plots that analyse how the categorical variables influence the target variable. We will use the same function that was applied to the numerical discrete variables.
for var in cat_var:
dis_analysis(data, var)
# *Sex* and *Embarked* provide useful information about their effect on the target variable.
#
# It is clear that female passengers had a much greater survival rate than male passengers.
#
# The port of embarkation seems to affect as well the target variable. The people that embarked in Cherbour (C) had greater chances of surviving than those who embarked in Queenstown (Q) and SouthHampton (S).
#
# Nonetheless, *Cabin*, *Ticket* and *Name* do not provide any useful information as the are many unique values that can't be aggregated into a group. Besides that, the reason of being name John or David does not seem to affect the target variable.
#
# Based on that the best option would be to drop those variable in the feature selection stage.
| Research_environment/ML pipeline research environment/ML_pipeline_research_data_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
pd.Series(['hello', 'world', 15, True])
x = pd.Series(['hello', 'world', 15, True])
y = pd.Series([i for i in range(15) if i%2==1])
x,y
y
dataframe = pd.DataFrame({
'x': x,
'y': y
})
dataframe
dataframe['z'] = pd.Series(['hi', 'hello', 'khana', 'kha', 'ke', 'jana', 'ha'])
dataframe
dataframe = pd.DataFrame(
data = [[0,0,0], [0,1,0], [1,0,0], [1,1,1]]
columns = ['x1', 'x2', 'y']
)
dataframe
dataframe = pd.DataFrame(
data = [[0,0,0], [0,1,0], [1,0,0], [1,1,1]],
columns = ['x1', 'x2', 'y']
)
dataframe
df = pd.DataFrame(
columns = ['x1', 'x2', 'y'],
data = [[0,0,0], [0,1,0], [1,0,0], [1,1,1]]
)
df
# +
df = pd.DataFrame([
{'a': '1', 'b': '1', 'c': 3},
{'a': '0', 'b': '10', 'c': 5}
])
df
# +
df = pd.DataFrame([
{'a': '1', 'b': '1', 'c': 3},
{'d': '0', 'b': '10', 'c': 5}
])
df
# -
| Workshops/agnostech-ml/day1/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import tensorflow as tf
import numpy as np
import gc
import pandas as pd
from datetime import datetime
from sklearn.utils import shuffle
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from tensorflow import keras
df = pd.read_csv("../Dataset/01-03-2018.csv", low_memory = False)
df = df.drop([0,1])
input_label = np.array(df.loc[:, df.columns != "Label"]).astype(np.float)
output_label = np.array(df["Label"])
out = []
for o in output_label:
if(o == "Benign"):out.append(0)
else: out.append(1)
output_label = out
scaler = MinMaxScaler(feature_range=(0,1))
scaler.fit(input_label)
input_label = scaler.transform(input_label)
input_label, output_label = shuffle(input_label, output_label)
# <h2>AutoEncoder</h2>
inp_train,inp_test,out_train,out_test = train_test_split(input_label, input_label, test_size=0.2)
input_model = keras.layers.Input(shape = (78,))
enc = keras.layers.Dense(units = 64, activation = "relu", use_bias = True)(input_model)
enc = keras.layers.Dense(units = 36, activation = "relu", use_bias = True)(enc)
enc = keras.layers.Dense(units = 18, activation = "relu")(enc)
dec = keras.layers.Dense(units = 36, activation = "relu", use_bias = True)(enc)
dec = keras.layers.Dense(units = 64, activation = "relu", use_bias = True)(dec)
dec = keras.layers.Dense(units = 78, activation = "relu", use_bias = True)(dec)
auto_encoder = keras.Model(input_model, dec)
encoder = keras.Model(input_model, enc)
decoder_input = keras.layers.Input(shape = (18,))
decoder_layer = auto_encoder.layers[-3](decoder_input)
decoder_layer = auto_encoder.layers[-2](decoder_layer)
decoder_layer = auto_encoder.layers[-1](decoder_layer)
decoder = keras.Model(decoder_input, decoder_layer)
auto_encoder.compile(optimizer=keras.optimizers.Adam(lr=0.00025), loss = "mean_squared_error", metrics = ['accuracy'])
train = auto_encoder.fit(x = np.array(inp_train), y = np.array(out_train),validation_split= 0.1, epochs = 10, verbose = 2, shuffle = True)
# <h2>cross validation</h2>
input_label = encoder.predict(input_label).reshape(len(input_label), 18, 1)
def createModel():
model = keras.Sequential([
keras.layers.Conv1D(filters = 16, input_shape = (18,1), kernel_size = 3, padding = "same", activation = "relu", use_bias = True),
keras.layers.MaxPool1D(pool_size = 3),
keras.layers.Conv1D(filters = 8, kernel_size = 3, padding = "same", activation = "relu", use_bias = True),
keras.layers.MaxPool1D(pool_size = 3),
keras.layers.Flatten(),
keras.layers.Dense(units = 2, activation = "softmax")
])
model.compile(optimizer= keras.optimizers.Adam(lr= 0.00025), loss="sparse_categorical_crossentropy", metrics=['accuracy'])
return model
skf = StratifiedKFold(n_splits = 10, shuffle = True, random_state=1)
confusion_matrixs = []
roc_curvs = []
for i, (train, test) in enumerate(skf.split(input_label, output_label)):
print("Modelo " + str(i))
inp_train, out_train = np.array(input_label)[train], np.array(output_label)[train]
inp_test, out_test = np.array(input_label)[test], np.array(output_label)[test]
model = createModel()
model.fit(x = inp_train, y = out_train, validation_split= 0.1, epochs = 10, shuffle = True,verbose = 2)
res = np.array([np.argmax(resu) for resu in model.predict(inp_test)])
confusion_matrixs.append(confusion_matrix(out_test, res))
fpr, tpr, _ = roc_curve(out_test, res)
auc = roc_auc_score(out_test, res)
roc_curvs.append([fpr, tpr, auc])
print("\n\n")
# <h2>Roc Curves</h2>
for i in range(10):
print("------------------------------------")
print("Modelo " + str(i))
print(roc_curvs[i])
print(confusion_matrixs[i])
print("------------------------------------")
| Modelos_com_reducao/Servidor/CNN/AutoEncoder/CNNInfiltrationIDS(01-03-2018).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Handling Missing Data
import numpy as np
import pandas as pd
vals1 = np.array([1, None, 3, 4])
vals1
for dtype in ['object', 'int']:
print("dtype =", dtype)
# %timeit np.arange(1E6, dtype=dtype).sum()
print()
vals1.sum()
vals2 = np.array([1, np.nan, 3, 4])
vals2.dtype
1 + np.nan
0 * np.nan
vals2.sum(), vals2.min(), vals2.max()
np.nansum(vals2), np.nanmin(vals2), np.nanmax(vals2)
pd.Series([1, np.nan, 2, None])
x = pd.Series(range(2), dtype=int)
x
x[0] = None
x
data = pd.Series([1, np.nan, 'hello', None])
data.isnull()
data[data.notnull()]
data.dropna()
df = pd.DataFrame([[1, np.nan, 2],
[2, 3, 5],
[np.nan, 4, 6]])
df
df.dropna()
df.dropna(axis='columns')
df[3] = np.nan
df
df.dropna(axis='columns', how='all')
df.dropna(axis='rows', thresh=3)
data = pd.Series([1, np.nan, 2, None, 3], index=list('abcde'))
data
data.fillna(0)
# forward-fill
data.fillna(method='ffill')
# back-fill
data.fillna(method='bfill')
df
df.fillna(method='ffill', axis=1)
| code_listings/03.04-Missing-Values.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
ad_data = pd.read_csv('advertising.csv')
ad_data.head()
ad_data.info()
ad_data.describe()
sns.set_style('whitegrid')
sns.displot(ad_data['Age'],bins=30)
ad_data['Age'].plot.hist(bins=30)
sns.jointplot(x='Age',y='Area Income',data=ad_data)
sns.jointplot(x='Age',y='Daily Time Spent on Site',data=ad_data,kind='kde')
sns.jointplot(x='Daily Time Spent on Site',y='Daily Internet Usage',data=ad_data,color='green')
sns.pairplot(data=ad_data,hue='Clicked on Ad')
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
X = ad_data.drop(['Ad Topic Line','City','Country','Timestamp','Clicked on Ad'], axis=1)
y = ad_data['Clicked on Ad']
ad_data.columns
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.3, random_state=101)
logmodel = LogisticRegression(solver='lbfgs',max_iter=1000)
logmodel.fit(X_train,y_train)
prediction = logmodel.predict(X_test)
from sklearn.metrics import classification_report
print(classification_report(y_test,prediction))
| Logistic Regression/Logistic Regression Project.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # Problema
#
# GoNuts produce diversi succhi realizzati interamente con varie noci esotiche. Il loro mercato principale è la Cina.
# Gestiscono tre stabilimenti, situati in Etiopia, Tanzania e Nigeria.
# Si vuole determinare dove produrre i due più recenti succhi che offrono, Gingko Nut e Kola Nut, considerando che ogni impianto ha una diversa capacità di produzione dei diversi succhi.
# Il costo fisso si applica solo se la fabbrica produce succo.
# Inoltre, ogni succo ha una domanda prevista.
# Quanto di ogni succo dovrebbe essere prodotto in ogni impianto per ridurre al minimo il costo totale, soddisfacendo la domanda e rispettando la capacità dell'impianto?
#
#
# |Costo/Unità |Ginko|Kola| Capacità Mensile Impianto|
# | --- | --- | --- | ---|
# |Ethiopia|21.00 dol| 22.50 dol|       425|
# |Tanzania|22.50 dol| 24.50 dol|       400|
# |Nigeria|23.00 dol | 25.50 dol|       750|
#
#
#
# |Domanda|Unità/Mese
# |---|---
# |Ginko|  550
# |Kola|  450
#
#
#
#
# Inizializziamo le librerie
using JuMP, GLPK, StatsPlots, Plots
# Definiamo il modello
m = Model(with_optimizer(GLPK.Optimizer));
# ### Variabili utili
#
# Le variabili che verranno dichiarate nel modello sono riportate di seguito.
#
# Si usa una notazione matriciale per rendere il codice più snello.
#
# * <b>x[1:3][1:2]</b>: la matrice incognita delle quantità dei succhi, così costruito:
# * <b>x[1][1]</b>: la quantità di Ginko prodotta in Ethiopia
# * <b>x[1][2]</b>: la quantità di Kola prodotta in Ethiopia
# * <b>x[2][1]</b>: la quantità di Ginko prodotta in Tanzania
# * <b>x[2][2]</b>: la quantità di Kola prodotta in Tanzania
# * <b>x[3][1]</b>: la quantità di Ginko prodotta in Nigeria
# * <b>x[3][2]</b>: la quantità di Kola prodotta in Nigeria
#
# * <b>costo</b>: la matrice del costo/unità dei due succhi, rispettivamente Ginko e Kola, prodotti rispettivamente in Ethiopia, Tanzania e Nigeria;
# * <b>capacità</b>: il vettore delle capacità dei tre impianti, rispettivamente Ethiopia, Tanzania e Nigeria;
# * <b>domanda</b>: il vettore della domanda di mercato delle quantità dei due succhi in ordine, Ginko e Kola.
#
# Definiamo la variabile x[1:6].
#
# Richiediamo che le quantità dei succhi prodotti in ogni impianto siano non nulle e numeri interi.
@variable(m, x[1:3 , 1:2] >= 0, Int);
# Definiamo il vettore della capacità e della domanda e la matrice dei costi.
capacita = [425 400 750];
domanda = [550 450];
costo = [21 22.5; 22.5 24.5; 23 25.5];
# Costruiamo la funzione obiettivo.
@objective(m, Min, sum(x .* costo))
# Dichiariamo i vincoli del problema:
# - la quantità di Ginko e Kola prodotta, deve soddisfare la domanda;
# - la quantità complessiva di succo prodotta non deve superare la capacità dei tre impianti.
@constraint(m, constraint1, sum(x, dims=2) .<= capacita')
@constraint(m, constraint2, sum(x, dims=1) .>= domanda)
# Il problema espresso in termini matematici:
# $$
# \text{Minimize } 21 x_{G,E} + 22.5 x_{K,E} + 22.5 x_{G,T}+ 24.5 x_{K,T} + 23 x_{G,N} + 25.5 x_{K,N}
# \text{ s.t } \begin{cases} x_{G,E} + x_{K,E} \leq 425 \\ x_{G,T} + x_{K,T} \leq 400 \\ x_{G,N} + x_{K,N} \leq 750 \\ x_{G,E} + x_{G,T} + x_{G,N} \geq 550 \\ x_{K,E} + x_{K,T} + x_{K,N} \geq 450 \\
# x_{G,E}, x_{K,E}, x_{G,T}, x_{K,T}, x_{G,N}, x_{K,N} \geq 0 \end{cases}
# $$
m
# Avviamo l'algoritmo di ottimizzazione.
optimize!(m)
# Calcoliamo il tempo di esecuzione
solve_time(m)
# Verifichiamo che l'algoritmo abbia fornito risultato con esito positivo.
termination_status(m)
# <p style="border: 3px solid crimson;box-shadow: 5px 6px indianRed;padding:20px;">La soluzione ottimale</p>
#
#
#
# +
println("x_{G,E} = ", value(x[1,1]))
println("x_{K,E} = ", value(x[1,2]))
println("x_{G,T} = ", value(x[2,1]))
println("x_{K,T} = ", value(x[2,2]))
println("x_{G,N} = ", value(x[3,1]))
println("x_{K,N} = ", value(x[3,2]))
println("** Optimal objective function value = ", JuMP.objective_value(m))
# -
# Disegnamo un grafico a barre con le quantità dei succhi ottimali da produrre, al fine di minimizzare i costi.
# +
ctg = repeat(["Ginko", "Kola"], inner = 3)
nam = ["Ethiopia", "Tanzania", "Nigeria","Ethiopia", "Tanzania","Nigeria"]
groupedbar(nam, [ value(x[1,1]) value(x[1,2]) ; value(x[2,1]) value(x[2,2]) ; value(x[3,1]) value(x[3,2]) ], group = ctg, xlabel = "Stabilimenti", ylabel = "Unità di succo",
title = "Quantità di succo prodotta", bar_width = 0.67,
lw = 0, framestyle = :box)
# +
Plots.gr()
totale= sum([value(x[1,1]) value(x[1,2]) value(x[2,1]) value(x[2,2]) value(x[3,1]) value(x[3,2])])
etichette = ["Ginko-Ethiopia","Kola-Ethiopia","Ginko-Tanzania","Kola-Tanzania","Ginko-Nigeria","Kola-Nigeria"]
y = [value(x[1,1]) ; value(x[1,2]) ; value(x[2,1]) ; value(x[2,2]) ; value(x[3,1]) ; value(x[3,2])]./totale
pie(etichette, y, title="Quantità di succo prodotta", l=0.5)
# -
y
| Progetto Ottimizzazione (versione interi).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
data = [115, 140, 175]
max = max(data)
min = min(data)
list(map(lambda x: (x - min)/(max-min), data) )
[float(e-min)/(max-min) for e in data ]
# ### Feature Scaling
from sklearn.preprocessing import MinMaxScaler
import numpy
data = numpy.array([[115.], [140.], [175.]])
scaler = MinMaxScaler()
scaler.fit_transform(data)
| classroom_examples/naive_bayes/Misc.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .java
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Java
// language: java
// name: java
// ---
// # Concise Implementation of Multilayer Perceptron
//
// :label:`sec_mlp_djl`
//
//
// As you might expect, by relying on the DJL library,
// we can implement MLPs even more concisely. <br>
// Let's setup the relevant libraries first.
// +
// %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// %maven ai.djl:api:0.7.0-SNAPSHOT
// %maven ai.djl:model-zoo:0.7.0-SNAPSHOT
// %maven ai.djl:basicdataset:0.7.0-SNAPSHOT
// %maven org.slf4j:slf4j-api:1.7.26
// %maven org.slf4j:slf4j-simple:1.7.26
// %maven ai.djl.mxnet:mxnet-engine:0.7.0-SNAPSHOT
// %maven ai.djl.mxnet:mxnet-native-auto:1.7.0-b
// -
// %%loadFromPOM
<dependency>
<groupId>tech.tablesaw</groupId>
<artifactId>tablesaw-jsplot</artifactId>
<version>0.30.4</version>
</dependency>
// %load ../utils/plot-utils.ipynb
import java.nio.file.*;
import ai.djl.Device;
import ai.djl.*;
import ai.djl.metric.*;
import ai.djl.ndarray.*;
import ai.djl.ndarray.types.*;
import ai.djl.ndarray.index.*;
import ai.djl.nn.*;
import ai.djl.nn.core.*;
import ai.djl.training.*;
import ai.djl.training.initializer.*;
import ai.djl.training.loss.*;
import ai.djl.training.listener.*;
import ai.djl.training.evaluator.*;
import ai.djl.training.optimizer.*;
import ai.djl.training.tracker.*;
import ai.djl.training.dataset.*;
import ai.djl.util.*;
import java.util.Random;
import java.util.stream.LongStream;
import ai.djl.basicdataset.FashionMnist;
import ai.djl.training.dataset.Dataset;
import tech.tablesaw.api.*;
import tech.tablesaw.plotly.api.*;
import tech.tablesaw.plotly.components.*;
import tech.tablesaw.plotly.Plot;
import tech.tablesaw.plotly.components.Figure;
import org.apache.commons.lang3.ArrayUtils;
// ## The Model
//
// As compared to our gluon implementation
// of softmax regression implementation
// (:numref:`sec_softmax_gluon`),
// the only difference is that we add
// *two* `Linear` (fully-connected) layers
// (previously, we added *one*).
// The first is our hidden layer,
// which contains *256* hidden units
// and applies the ReLU activation function.
// The second is our output layer.
// + attributes={"classes": [], "id": "", "n": "5"}
SequentialBlock net = new SequentialBlock();
net.add(Blocks.batchFlattenBlock(784));
net.add(Linear.builder().setUnits(256).build());
net.add(Activation::relu);
net.add(Linear.builder().setUnits(10).build());
net.setInitializer(new NormalInitializer());
// -
// Note that DJL, as usual, automatically
// infers the missing input dimensions to each layer.
//
// The training loop is *exactly* the same
// as when we implemented softmax regression.
// This modularity enables us to separate
// matters concerning the model architecture
// from orthogonal considerations.
// +
int batchSize = 256;
int numEpochs = 10;
double[] trainLoss;
double[] testAccuracy;
double[] epochCount;
double[] trainAccuracy;
trainLoss = new double[numEpochs];
trainAccuracy = new double[numEpochs];
testAccuracy = new double[numEpochs];
epochCount = new double[numEpochs];
FashionMnist trainIter = FashionMnist.builder()
.optUsage(Dataset.Usage.TRAIN)
.setSampling(batchSize, true)
.build();
FashionMnist testIter = FashionMnist.builder()
.optUsage(Dataset.Usage.TEST)
.setSampling(batchSize, true)
.build();
trainIter.prepare();
testIter.prepare();
for(int i = 0; i < epochCount.length; i++) {
epochCount[i] = (i + 1);
}
Map<String, double[]> evaluatorMetrics = new HashMap<>();
// + attributes={"classes": [], "id": "", "n": "6"}
Tracker lrt = Tracker.fixed(0.5f);
Optimizer sgd = Optimizer.sgd().setLearningRateTracker(lrt).build();
Loss loss = Loss.softmaxCrossEntropyLoss();
DefaultTrainingConfig config = new DefaultTrainingConfig(loss)
.optOptimizer(sgd) // Optimizer (loss function)
.addEvaluator(new Accuracy()) // Model Accuracy
.addTrainingListeners(TrainingListener.Defaults.logging()); // Logging
try (Model model = Model.newInstance("mlp")) {
model.setBlock(net);
try (Trainer trainer = model.newTrainer(config)) {
trainer.initialize(new Shape(1, 784));
trainer.setMetrics(new Metrics());
EasyTrain.fit(trainer, numEpochs, trainIter, testIter);
// collect results from evaluators
Metrics metrics = trainer.getMetrics();
trainer.getEvaluators().stream()
.forEach(evaluator -> {
evaluatorMetrics.put("train_epoch_" + evaluator.getName(), metrics.getMetric("train_epoch_" + evaluator.getName()).stream()
.mapToDouble(x -> x.getValue().doubleValue()).toArray());
evaluatorMetrics.put("validate_epoch_" + evaluator.getName(), metrics.getMetric("validate_epoch_" + evaluator.getName()).stream()
.mapToDouble(x -> x.getValue().doubleValue()).toArray());
});
}
}
// +
trainLoss = evaluatorMetrics.get("train_epoch_SoftmaxCrossEntropyLoss");
trainAccuracy = evaluatorMetrics.get("train_epoch_Accuracy");
testAccuracy = evaluatorMetrics.get("validate_epoch_Accuracy");
String[] lossLabel = new String[trainLoss.length + testAccuracy.length + trainAccuracy.length];
Arrays.fill(lossLabel, 0, trainLoss.length, "test acc");
Arrays.fill(lossLabel, trainAccuracy.length, trainLoss.length + trainAccuracy.length, "train acc");
Arrays.fill(lossLabel, trainLoss.length + trainAccuracy.length,
trainLoss.length + testAccuracy.length + trainAccuracy.length, "train loss");
Table data = Table.create("Data").addColumns(
DoubleColumn.create("epochCount", ArrayUtils.addAll(epochCount, ArrayUtils.addAll(epochCount, epochCount))),
DoubleColumn.create("loss", ArrayUtils.addAll(testAccuracy , ArrayUtils.addAll(trainAccuracy, trainLoss))),
StringColumn.create("lossLabel", lossLabel)
);
render(LinePlot.create("", data, "epochCount", "loss", "lossLabel"),"text/html");
// -
// 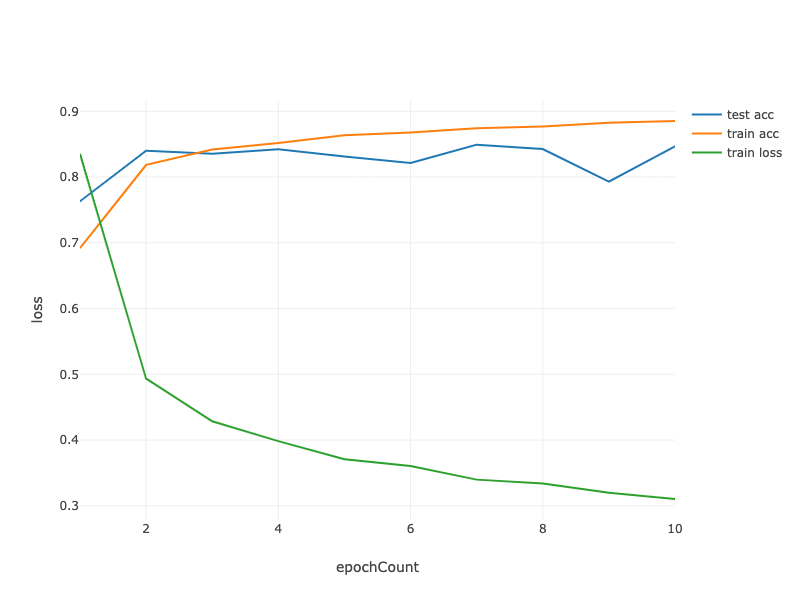
// ## Exercises
//
// 1. Try adding different numbers of hidden layers. What setting (keeping other parameters and hyperparameters constant) works best?
// 1. Try out different activation functions. Which ones work best?
// 1. Try different schemes for initializing the weights. What method works best?
//
//
| jupyter/d2l-java/chapter_multilayer-perceptrons/mlp-djl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import logging
import sklearn.datasets as ds
import pandas as pd
import numpy as np
import math as mt
import time
import datetime
from tensorflow.keras.optimizers import Adadelta
from tensorflow.keras.optimizers import Adagrad
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.optimizers import Adamax
from tensorflow.keras.optimizers import Ftrl
from tensorflow.keras.optimizers import Nadam
from tensorflow.keras.optimizers import Optimizer
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.optimizers import SGD
from itertools import product
from sklearn.metrics import mean_squared_error
from mulearn import FuzzyInductor
from mulearn.optimization import TensorFlowSolver
from mulearn.kernel import GaussianKernel
logger = logging.getLogger()
logger.setLevel(logging.INFO)
# -
def create_dataset(name):
#load dataset, in iris_X values, in iris_y labels 0 1 2
iris_X, iris_y = ds.load_iris(return_X_y=True)
labels = ("Setosa", "Versicolor", "Virginica")
#dataframe with correct labels for respective values
df = pd.DataFrame(iris_X, columns=["Sepal length", "Sepal width",
"Petal length", "Petal width"])
#associating
df['Class'] = iris_y
df['Class'] = df['Class'].map(lambda c: labels[c])
#dataset copy for labels 0 1
selected_iris_dataset = iris_y.copy()
#dataset selected with labels
if(name == "Setosa"):
selected_iris_dataset[selected_iris_dataset != 0] = 2
selected_iris_dataset[selected_iris_dataset == 0] = 1
selected_iris_dataset[selected_iris_dataset == 2] = 0
elif(name == "Versicolor"):
selected_iris_dataset[selected_iris_dataset==2] = 0
elif(name == "Virginica"):
selected_iris_dataset[selected_iris_dataset != 2] = 0
selected_iris_dataset[selected_iris_dataset == 2] = 1
return iris_X, selected_iris_dataset
def create_handler(path):
fhandler = logging.FileHandler(filename = path)
formatter = logging.Formatter('%(asctime)s - %(name)s - %(levelname)s - %(message)s')
fhandler.setFormatter(formatter)
logger.addHandler(fhandler)
return fhandler
def main(nome, c, sigma, penalization, optimizer):
handler = create_handler(f"../../../log/Prove-6/Different-Optimizer/{nome}_c{str(c).replace('.','')}"
f"_sigma{str(sigma).replace('.','')}"
f"_penalization{str(penalization).replace('.','')}"
f"_{str(optimizer)}.log")
#salvo i parametri
logger.info(f"PARAMETRI DI PARTENZA: nome={nome}, c={c}, sigma={sigma}, penalization={penalization},"
f"optimizer={optimizer}")
iris_X, selected_iris_dataset = create_dataset(nome)
n_iter = 100
# Gurobi solver & fit
fi = FuzzyInductor(c=c, k=GaussianKernel(sigma=sigma))
start = time.time()
fi.fit(iris_X, selected_iris_dataset)
end = time.time()
# rmse gurobi
gurobi_chis = fi.chis_
logger.info(f"GUROBI_CHIS: {gurobi_chis}")
logger.info(f"GUROBI_START: {start}")
logger.info(f"GUROBI_END: {end}")
logger.info(f"TEMPO_ESECUZIONE GUROBI(IN EPOCH): {(end-start)}")
# TensorFlow solver
fi = FuzzyInductor(solver=TensorFlowSolver(n_iter=n_iter, optimizer=optimizer,
penalization=penalization),c=c,k=GaussianKernel(sigma=sigma))
try:
start = time.time()
fi.fit(iris_X, selected_iris_dataset)
end = time.time()
except (ModuleNotFoundError, ValueError):
print('Tensorflow not available')
# rmse TensorFlow
tensorflow_chis = fi.chis_
logger.info(f"TENSORFLOW_CHIS: {tensorflow_chis}")
logger.info(f"TENSORFLOW_START: {start}")
logger.info(f"TENSORFLOW_END: {end}")
total_tensorflow = end-start
logger.info(f"TEMPO PER ITERAZIONI (N_ITER,TEMPO) TENSORFLOW: [({n_iter},{total_tensorflow})]")
# calcolo distanza
rmse_distance = abs(mean_squared_error(gurobi_chis, tensorflow_chis, squared=False))
logger.info(f"RMSE_DISTANCE_CHIS: {rmse_distance}")
#coppia n_iter, distance
couple = [(n_iter, rmse_distance)]
logger.info(f"COUPLE(N_ITER,DISTANCE RMSE): {couple}")
#incremento n
n = 200
iter_tot = 5000
# faccio ciclo fino a 5000
while n <= iter_tot:
chi_ = tensorflow_chis
# TensorFlow solver
fi = FuzzyInductor(solver=TensorFlowSolver(initial_values=chi_, n_iter=n_iter,
optimizer=optimizer,
penalization=penalization),c=c,k=GaussianKernel(sigma=sigma))
try:
start = time.time()
fi.fit(iris_X, selected_iris_dataset)
end = time.time()
except (ModuleNotFoundError, ValueError):
print('Tensorflow not available')
total_tensorflow += (end-start)
# rmse TensorFlow
tensorflow_chis = fi.chis_
logger.info(f"TENSORFLOW_CHIS: {tensorflow_chis}")
logger.info(f"TENSORFLOW_START: {start}")
logger.info(f"TENSORFLOW_END: {end}")
logger.info(f"TEMPO PER ITERAZIONI (N_ITER,TEMPO) TENSORFLOW: [({n},{total_tensorflow})]")
# calcolo distanza
rmse_distance = abs(mean_squared_error(gurobi_chis, tensorflow_chis, squared=False))
logger.info(f"RMSE_DISTANCE_CHIS: {rmse_distance}")
#coppia n_iter, distance
couple = [(n, rmse_distance)]
logger.info(f"COUPLE(N_ITER,DISTANCE RMSE): {couple}")
if(n == iter_tot):
logger.info(f"TENSORFLOW_END: {end}")
logger.info(f"TEMPO_ESECUZIONE TENSORFLOW(IN EPOCH): {total_tensorflow}")
#incremento n_iter
n += 100
logger.removeHandler(handler)
main("Setosa",1,0.25,0.1,Adadelta(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,Adagrad(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,Adamax(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,Ftrl(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,Nadam(learning_rate=1e-4))
# +
#main("Setosa",1,0.25,0.1,Optimizer(learning_rate=1e-4))
# -
main("Setosa",1,0.25,0.1,RMSprop(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,SGD(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,Adam(learning_rate=1e-4))
main("Setosa",1,0.25,0.1,Adadelta(learning_rate=1e-3))
# +
#lista = list(product(["Setosa","Versicolor","Virginica"],[1],[0.25],[0.1,10],[1e-4], repeat=1))
#print(lista)
# +
#len(lista)
# -
#for i in lista:
#if i not in [('Setosa', 0.05, 0.1, 0.1),('Versicolor', 0.05, 0.1, 0.1),('Virginica', 0.05, 0.1, 0.1)]:
#main(*i)
| notebook/Prove/Prove-6/Prove-Different-Optimizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Geothermal doublets
#
# In this notebook, we will take a look at geothermal doublets. It will cover the definition of a geothermal doublet installation, explain some _buzzwords_ such as "thermal breakthrough" or "kalina cycle", and show some results of a numerical doublet simulation.
# This simulation is run with SHEMAT-Suite, a numerical code for simulating heat- and mass transfer in a porous medium. The simulations are run on a synthetic model of a graben system over a time of 35 years. That means, we simulate geothermal power production over a lifespan of 35 years.
#
# <img src='https://images.fd.nl/archive/89002_def-tno-geothermie.jpg?fit=crop&crop=faces&auto=format%2Ccompress&q=45&w=1280' style="float: left; width: 27%; margin-right: 3%; margin-bottom: 0.5em;">
# ### Introduction
# A geothermal doublet system usually consists of -at least- two boreholes, connected via a surface application installation (such as a powerplant for producing electricity, or an installation for district heating). One geothermal well produces the hot fluid (or steam) from the subsurface. This well is called the producer. The heat of the produced fluid is used accordingly, and the significant cooler fluid is then re-injected in the geothermal reservoir. In the figure to the left, the producing well is marked as a red line, while the injecting well is marked as a blue line, representing the difference in fluid temperature.
#
# If the heat content of the produced "fluid" is large enough, i.e. dry steam is produced, a turbine can be directly operated in one circuit. If the temperatures of the fluide are lower and do not really suffice to operate a turbine directly, a binary cycle is often installed. In such a cycle, the produced fluid heats up a secondary fluid in a heat-exchanger. This secondary fluid has a significantly lower boiling point, an can thus be used to operate a turbine instead. Such a system is often called a [Kalina installation](https://en.wikipedia.org/wiki/Kalina_cycle), where the secondary fluid is a mixture of water and ammonia.
#
# Operating such a system over a prolonged time will eventually cause a decrease in production temperature, as the cooling front of the re-injected water reaches the producing borehole. The point in time, where production temperature starts to decrease significantly is called a _thermal breakthrough_. In the results of a simplified doublet simulation, which we study in this notebook, we will also look for a thermal breakthrough.
# <br>[Image source](https://images.fd.nl/archive/89002_def-tno-geothermie.jpg?fit=crop&crop=faces&auto=format%2Ccompress&q=45&w=1280)
#
# +
# necessary libraries
import h5py
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# for improving plot aesthetics
import seaborn as sns
sns.set_style('ticks')
sns.set_context('poster')
# -
# ## 3D reservoir model
# For assessing this (rather cold) geothermal system, we simulate the heat- and mass transfer in a synthetic, three dimensional model. The model consists of 4 geological units:
# * Basement
# * Bottom Unit
# * Reservoir Unit
# * Top Unit
#
# The reservoir Unit is a geological body with a high primary porosity and permeability, which enables a natural darcy flow, and also shows advective heat transport by convection. Convection increases the geothermal potential of a system, as hot fluids are flowing upwards due to lower density, cooling down, and descending again. Kind of like [lava lamps](https://lavalamp.com/wp-content/uploads/2016/08/6825_1500x2000.jpg), which were popular back in the days.
# In the plot below, we see a vertical cross-section of the model in x-direction (let's say an East-West cross-section). On the left, you can see the geological units with annotations, on the right the natural temperature field. There, we can already see strongly bent isothermes, which are caused by an interplay of differences in thermal conductivity of the geological units, and advective heat transport.
# For assessing, which of those processes, i.e. conductive vs. advective heat transport, we could perform a peclet number analysis (there will also be a notebook on peclet number analysis in this repository). For the moment, it is sufficient to state, that both processes are present in this model.
# + code_folding=[]
# 3D model file
mod = h5py.File('data/Input_f_final.h5')
x = mod['x'][0,0,:]
y = mod['y'][0,:,0]
z = mod['z'][:,0,0]
t = mod['temp'][:,:,:]
ui = mod['uindex'][:,:,:]
vx = mod['vx'][:,:,:]
vy = mod['vy'][:,:,:]
vz = mod['vz'][:,:,:]
mod.close()
# doublet position
prod = np.array([74, 29, 15])
inj = np.array([41, 29, 15])
# plot routine
cs = 29 # cross -section position
fig, axs = plt.subplots(1, 2, figsize=[15,5], sharey=True, sharex=True)
ax1 = axs[0].contourf(x, z-2000, ui[:,cs,:], 4, cmap= 'viridis') # geological units
ax2 = axs[1].contourf(x, z-2000, t[:,cs,:], cmap='viridis') # temperature
# plot doublet positions
axs[0].plot(prod[0]*20, (prod[2]*50-2000), 'ro')
axs[0].plot(inj[0]*20, (inj[2]*50-2000), 'bo')
axs[1].plot(prod[0]*20, (prod[2]*50-2000), 'ro')
axs[1].plot(inj[0]*20, (inj[2]*50-2000), 'bo')
axs[1].streamplot(x, z-2000, vx[:,cs,:], vz[:,cs,:], density=[1, .5], color='white')
# axes arguments
axs[0].set_ylabel('model height [m]')
axs[0].set_xlabel('y [m]')
axs[1].set_xlabel('y [m]')
# annotations
axs[0].text(200, -1750, 'Basement', color=[.9,.9,.9])
axs[0].text(200, -1335, 'Bottom Unit', color=[.9,.9,.9])
axs[0].text(200, -850, 'Reservoir Unit', color=[.9,.9,.9])
axs[0].text(200, -280, 'Top Unit', color=[.3,.3,.3])
# colormap
fig.subplots_adjust(right=0.8)
cbar_ax = fig.add_axes([1., 0.21, 0.03, 0.7])
fig.colorbar(ax2, cax=cbar_ax, label= 'T °C')
# layout option
plt.tight_layout()
# -
# Temperatures in the model range from 10 °C to 80 °C. These temperatures are not sufficiently high for direct geothermal electricity generation, but potentially appropriate for a binary system or a system for direct-heat usage. One interesting characteristic of this model is the convective heat transport. Next to temperature, you can see streamlines in the right plot above. These lines visualise the flow field (and may be used to visualise every vector field you have). Essentially, with a stable convective system, producing from upwelling areas (where hot water rises due to its relatively lower density) is good, as higher temperatures are encountered at shallower depths.
# Despite the low temperature range we simulated, the following assessment can generally be applied to doublet systems, regardless of the produced temperatures. Speaking of produced temperatures, in the model above, we implemented a doublet system, where the flow rate (i.e. the rate at which the pump operates) is varied.
# Below, we plot the temperature at the producing borehole with time.
# +
# load monitoring files
pr_50L = pd.read_csv('data/doublet_50ls.csv', comment='%')
pr_75L = pd.read_csv('data/doublet_75ls.csv', comment='%')
pr_100L = pd.read_csv('data/doublet_100ls.csv', comment='%')
fig = plt.figure(figsize=[12,6])
p50 = plt.plot(pr_50L['time'], pr_50L['temp'], '--', label='50 L s$^{-1}$')
p75 = plt.plot(pr_75L['time'], pr_75L['temp'], '--', label='75 L s$^{-1}$')
p100 = plt.plot(pr_100L['time'], pr_100L['temp'], '--', label='100 L s$^{-1}$')
plt.legend()
plt.ylabel('Temperature [°C]')
plt.xlabel('time [a]')
fig.tight_layout()
# -
# Over the simulated time of 35 years, you can see the temperatures produced from the production borehole (red circle in the cross sections above). Note how the temperatures rapidly decrease at the beginning of the simulation. As we implement the doublet as a boundary condition (i.e. a source/sink term), the system has to react to the newly applied boundary conditions. However, after around 1 year, the system is re-equilibrated and produced temperatures can be assessed.
# One thing of interest for example is the thermal breakthrough of a system, the point in time, when the temperature signal of the cold, re-injected water reaches the production borehole.
# Subsurface parameters, such as hydraulic head distribution or reservoir permeability affect the timing of a thermal breakthrough. But it is also influenced by operation parameters, e.g. pumping rate and injection temperature. In the plot above, we model the same doublet layout with three different pumping rates, and we see that 100 l s$^{-1}$ correlate to an earlier thermal breakthrough, when compared to the other pumping rates (75 l s$^{-1}$ and 50 l s$^{-1}$).
# While knowledge about the development of the production temperature over time is important, other parameters, such as the transient change in obtained thermal power are equally or even more interesting. Thermal power of a geothermal doublet can be calculated using the following equation:
#
# $$ P_{th} = Q (\rho c_p)_w (T_{pr} - T_{in}) $$
#
# where P$_{th}$ is the thermal power in W, Q the pumping rate in m³ s$^{-1}$ ($\rho c_p)_w$) the thermal capacity of water in W m$^{-3}$ K$^{-1}$ and (T$_{pr}$ - T$_{in}$) the difference between production temperature and injection temperature (in K). Let's have a look at the thermal power produced by the doublet with three different pumping rates.
def therm_pow(Q, rhocp, Tin, Tpr):
"""
A short function for calculating the thermal power
Q - scalar, flow rate in m³/s
rhocp - scalar, thermal capacity of water in W/(m³K)
Tin - scaler, re-injection temperature of the water in °C
Tpr - vector, production temperature of the water in °C
"""
return Q*rhocp*(Tpr-Tin)
# +
# parameter
Q = np.array([0.05, 0.075, 0.1]) # 50 L/s, 75 L/s, 100 L/s
rhocp = 988 * 4180 # density and specific heat capacity of water at a temperature of about 50 °C
Tin = 30 # injection temeprature in °C
# calculate thermal power
thp50 = therm_pow(Q[0], rhocp, Tin, pr_50L['temp'])
thp75 = therm_pow(Q[1], rhocp, Tin, pr_75L['temp'])
thp100 = therm_pow(Q[2], rhocp, Tin, pr_100L['temp'])
# +
# plot
d2MW = 1e6 # use for calculating in MW
fig = plt.figure(figsize=[12,6])
p50 = plt.plot(pr_50L['time'], thp50/d2MW, '--', label='50 L s$^{-1}$')
p75 = plt.plot(pr_75L['time'], thp75/d2MW, '--', label='75 L s$^{-1}$')
p100 = plt.plot(pr_100L['time'], thp100/d2MW, '--', label='100 L s$^{-1}$')
plt.legend()
plt.ylabel('therm. power [MW]')
plt.xlabel('time [a]')
fig.tight_layout()
# -
# Logically, the highest flow rate yields the highest thermal power. The timing of thermal breakthrough and its intensity are however visible, as the produced thermal power with a flow rate of 100 L s$^{-1}$ declines significantly stronger than lower flow rates. In this scenario, this may not cause a problem.
# In the exercise course, we will assess a different reservoir system where the significantly different evolution of production temperatures depending on the chosen pumping rate will visualise the importance of monitoring the reservoir and the production temperatures.
| 09_Geothermal_doublets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating synthetic data
#
# This notebook walks through training a probabilistic, generative RNN model<br>
# on a rental scooter location dataset, and then generating a synthetic<br>
# dataset with greater privacy guarantees.
#
# For both training and generating data, we can use the ``config.py`` module and<br>
# create a ``LocalConfig`` instance that contains all the attributes that we need<br>
# for both activities.
# +
# Google Colab support
# Note: Click "Runtime->Change Runtime Type" set Hardware Accelerator to "GPU"
# Note: Use pip install gretel-synthetics[tf] to install tensorflow if necessary
#
# #!pip install gretel-synthetics --upgrade
# +
from pathlib import Path
from gretel_synthetics.config import LocalConfig
# Create a config that we can use for both training and generating data
# The default values for ``max_lines`` and ``epochs`` are optimized for training on a GPU.
config = LocalConfig(
max_line_len=2048, # the max line length for input training data
vocab_size=20000, # tokenizer model vocabulary size
field_delimiter=",", # specify if the training text is structured, else ``None``
overwrite=True, # overwrite previously trained model checkpoints
checkpoint_dir=(Path.cwd() / 'checkpoints').as_posix(),
input_data_path="https://gretel-public-website.s3-us-west-2.amazonaws.com/datasets/uber_scooter_rides_1day.csv" # filepath or S3
)
# +
# Train a model
# The training function only requires our config as a single arg
from gretel_synthetics.train import train_rnn
train_rnn(config)
# +
# Let's generate some text!
#
# The ``generate_text`` funtion is a generator that will return
# a line of predicted text based on the ``gen_lines`` setting in your
# config.
#
# There is no limit on the line length as with proper training, your model
# should learn where newlines generally occur. However, if you want to
# specify a maximum char len for each line, you may set the ``gen_chars``
# attribute in your config object
from gretel_synthetics.generate import generate_text
# Optionally, when generating text, you can provide a callable that takes the
# generated line as a single arg. If this function raises any errors, the
# line will fail validation and will not be returned. The exception message
# will be provided as a ``explain`` field in the resulting dict that gets
# created by ``generate_text``
def validate_record(line):
rec = line.split(", ")
if len(rec) == 6:
float(rec[5])
float(rec[4])
float(rec[3])
float(rec[2])
int(rec[0])
else:
raise Exception('record not 6 parts')
for line in generate_text(config, line_validator=validate_record, num_lines=1000):
print(line)
| examples/synthetic_records.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Series
import pandas as pd
from IPython.display import Image, YouTubeVideo
YouTubeVideo(id='zmdjNSmRXF4', width=900, height=400)
# In data science, we very often have labels that are associated with the data that we manipulate. And we would like to be able to put labels on the tables we handle. This is exactly what pandas can do.
# Let's look at an example. Here I have an array that represents ages for example, and I would like to be able to associate each entry in this array with a first name.
Image('serie.png')
# Well, in pandas, I can do it by adding labels which are represented by an object called an index. And this one-dimensional object is called in pandas a Series.
Image('serie1.png')
# Now, imagine that I have a two-dimensional array, where the first column represents ages and the second column represents sizes, I would like to be able to add labels on the columns, Age and Height, and I would like to be able to add labels on the lines, which correspond to first names.
Image('df.png')
# Again, this is what pandas does with a data structure called the DataFrame.
Image('df1.png')
# So in summary, there are two major data structures in pandas, **`Series`**, for one-dimensional data, and **` DataFrames`**, for two-dimensional data. Much of the complexity of pandas comes from mastering this notion of **index**. An index is an object which is very powerful and which has two major characteristics. It allows optimized access to the data in the table, and it also allows a notion of automatic alignment during operations.Let's take an example, imagine that you have to add two DataFrames, the addition operation will **only apply** to elements that have exactly the **same label**.
# An index is an immutable object that is on the border between `set` and `list`. An index contains items that are hasable, like a set, and it's sliceable, like a list. In addition, the index will define an order relationship on the items that are stored, and it may contain duplicate items. An index on a Series will provide the Series with an interface that is both a list interface and a dictionary interface. In practice, we very rarely create our own indexes. In fact, the indexes will be created automatically when we import our data. And pandas supports a wide variety of data formats, both import and export. In pandas, you can import data in `csv` format, in `json` format, in `html`, `excel`, `sql` or `pickle` format.
# ## Creating a Series
# Create a series and pass it a list of items, 20, 30, 40, 50, and pass it an index; the index is going to be given names, for example, Eve, Bill, Liz and Bob.
s = pd.Series([20, 30, 40, 50], index=['eve', 'bill', 'liz', 'bob'])
s
# In this series, I can retrieve the values and so it will output me the NumPy array corresponding to the values hosted by this series.
s.values
# I can access the index with the index attribute, and so it's going to return me the index object that is in this series.
s.index
# And then I can access the elements of that object.
s.loc['eve']
# I write `s.loc['eve']` This will allow me to access the value corresponding to the label eve. So you notice here that I used the loc attribute; it is very important to always use this attribute. You will see in some documentation or on the internet that it is possible to access the value stored by eve directly in this way `s['eve']`; but this notation has many side effects.
# So I can access eve but I can also make a slice; I can go for example from eve to liz. Let's look at the result and I get the values ranging from eve to liz.
s.loc['eve': 'liz']
# There is a noticeable difference here with the slice you usually know is that here I make a slice of i colon j but I go from i to j inclusive, so I go from eve to liz included, the right terminal is indeed a terminal that is taken into account and which is included. As we can make slices on the labels, you are probably wondering what is the order relation that I have on my labels. This order relation is determined when I create my Series by the order in which I will specify my labels when I define my index.
# Let's look at an example, I will take the previous case, so I take this series, which I copy below, but now I'm going to swap bill and liz. I am running this series. I recreate the same slice as the one above, s.loc from eve to liz and what is the result I get? Now, I'm fine from eve to liz but with an order relation which is defined by the order of the elements when creating my index.
s = pd.Series([20, 30, 40, 50], index=['eve', 'liz', 'bill', 'bob'])
s.loc['eve':'liz']
# However, there is one case where **slicing** doesn't work. Slicing on labels will not be defined if you have **duplicate** labels and in addition your index has not been **sorted**. So in other words, if you don't have a duplicate label, slicing will still work, and if your index has been sorted, slicing will still work. In practice, we do not control the labels since the labels are given by our dataset. However, you can still sort your index. Therefore, it is recommended that you always do this. When you `sort` the `index`, you are guaranteed that the slicing will always work, and in addition, sorting the index significantly improves the performance of that index. Let's take an example:
pets = ['dog', 'cat', 'cat', 'dog', 'fish']
owner = ['eve', 'bob', 'eve', 'bill', 'liz']
s = pd.Series(pets, index=owner)
s
# So now let's try to do some slicing on this streak. I will make slicing from eve to liz. So now what's going to happen?
s.loc['eve': 'liz']
# I have a KeyError exception: Cannot get left slice bound for non-unique label: "eve". My label eve has been duplicated, I cannot no slicing on it. So how do you solve this problem? It's very simple, you just have to sort the index.
s = s.sort_index()
s.loc['eve':'liz']
# Let's look at an example. I'm going to take my series s, and I'm going to get all the items for which the value is dog. I run and I get the series for which the value is dog.
s.loc[s=='dog']
# I can obviously do more sophisticated indexing, so I take s equals dog or s equals fish, and so I'm going to grab the list, the series that contains dogs and fish
s.loc[(s=='dog') | (s=='fish')]
# And I can end up doing some assignment on this advanced index, and so I can say here that all those who are worth dog or fish are now replaced by other. If I look at my series now, all I have left is a cat, and dog and fish have been replaced by other.
s.loc[(s =='dog') | (s == 'fish')] = 'other'
s
# ## Index alignement
# The notion of index alignment is an extremely important notion when talking about Series in Pandas.
# Let's create two series s1 and s2
s1 = pd.Series([1, 2, 3], index=list('abc'))
s2 = pd.Series([5, 6, 7], index=list('acd'))
s1
s2
# Now let's look at what 'it happens if I do an addition.
s1 + s2
# Pandas will automatically align the labels, i.e. the operation will only be defined for values that have the same label on the left and on the right.So when I miss a value for a given label in one of the two series, the result of the operation will be NaN. You also notice that my two series are series which are int64. And that the result is a float64. Why ? Well because there are operations which are not defined, I have to represent the operation not defined by a NaN, and a limitation of NumPy is that the NaN only exists for floats so everything is converted to float64.
# We can obviously control this behavior with the `add` method, but now i can pass an argument which is called `fill_value` and which will consist of saying: if i am missing an element on the left or on the right, i will replace it with a default value. And so here, the default value that we are going to put is 50.
s1.add(s2, fill_value=50)
| Lecture 30 Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: couchers
# language: python
# name: couchers
# ---
from dotenv import load_dotenv
load_dotenv("readonly.env")
# +
from google.protobuf import message_factory, any_pb2
from sqlalchemy.sql import or_
from couchers.descriptor_pool import get_descriptor_pool
from couchers.db import session_scope
from couchers.models import APICall
from couchers.utils import now
from datetime import timedelta
from trace_pb2 import Trace
# -
TYPE_DOMAIN = "type.couchers.org"
# +
pool = get_descriptor_pool()
factory = message_factory.MessageFactory(pool=pool)
# -
def get_method_descriptor(method):
_, service_name, method_name = method.split("/")
service_descriptor = pool.FindServiceByName(service_name)
method_descriptor = service_descriptor.FindMethodByName(method_name)
return method_descriptor
def get_proto(type_url):
domain, version, type_name = type_url.split("/")
return factory.GetPrototype(pool.FindMessageTypeByName(type_name))
def deserialize_trace(trace: Trace):
request_proto = get_proto(trace.request.type_url)
response_proto = get_proto(trace.response.type_url)
return request_proto.FromString(trace.request.value), response_proto.FromString(trace.response.value) if trace.response.value else None
# +
traces = []
with session_scope() as session:
trace_q = (session.query(APICall)
#.filter(APICall.method.startswith("/org.couchers.api.admin"))
#.filter(APICall.method == "/org.couchers.bugs.Bugs/ReportBug")
.filter(APICall.time > now() - timedelta(hours=24))
#.filter(APICall.time < "2021-06-30T21:08:55.054000+00:00")
#.filter(or_(APICall.user_id == user_id, APICall.user_id == None))
.filter(APICall.status_code != None)
.filter(APICall.traceback != None)
.order_by(APICall.id.desc())
.limit(100))
for trace in trace_q.all():
mdesc = get_method_descriptor(trace.method)
input_url = f"{TYPE_DOMAIN}/{trace.version}/{mdesc.input_type.full_name}"
output_url = f"{TYPE_DOMAIN}/{trace.version}/{mdesc.output_type.full_name}"
trace = Trace(
id=trace.id,
version=trace.version,
time=trace.time.isoformat(),
method=trace.method,
status_code=trace.status_code,
duration=trace.duration,
user_id=trace.user_id,
request=any_pb2.Any(type_url=input_url, value=trace.request),
response=any_pb2.Any(type_url=output_url, value=trace.response),
traceback=trace.traceback,
)
traces.append(trace)
# -
for trace in traces:
print(f"*** Call ***")
print(f"status_code={trace.status_code or ('TB' if trace.traceback else 'OK')}")
print(f"method={trace.method}")
print(f"user_id={trace.user_id or 'NA'}")
print(f"time={trace.time}")
print(f"duration={trace.duration} ms")
req, res = deserialize_trace(trace)
if req:
print(f"Request: ({trace.request.type_url})")
print(req)
if res:
print(f"Response: ({trace.response.type_url})")
print(res)
print()
| ds/Trace tool.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.1
# language: julia
# name: julia-1.3
# ---
# +
using HDF5
using LaTeXStrings
using Plots
pyplot()
# -
# give the directory where the data from all the runs are saved
your_dir = "/home/thanasis/repos/BondiToy/examples/run00"
# +
# uncomment the model you want
# SH model
data_dir = your_dir*"/SH_smooth_B1"
# WH model
#data_dir = your_dir*"/WH_smooth_B0"
Nx = 129#17 # the overal course graining
Nz = 128#16
n=0
# load the x grid
xc = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/x.h5","x")
# load the z grid
zc = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/z.h5","z")
hzc = zc[2]-zc[1]
# load all the timesteps that data is writen
#tc = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0000.h5","time");
# load initial data
#i_final=length(tc)
ψc_0 = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0000.h5","ψ")
ψvc_0 = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0000.h5","ψv")
ϕc_0 = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0000.h5","ϕ")
# load final field configurations
#i_final=length(tc)
# the above is the final timstep of the evolution. You can modify it to your desired step
ψc_f = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0512.h5","ψ");
ψvc_f = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0512.h5","ψv");
ϕc_f = h5read(data_dir*"/data_$((Nx-1)*2^n + 1)_$(Nz*2^n)/data_0512.h5","ϕ");
# -
# field ψ at t=tf=1
surface(xc,zc,transpose(ψc_f),
xlabel="x",xtick=(0.0:0.25:1.0),xtickfont = font(12),
ylabel="z",ytick=(0.0:2:6),ytickfont = font(12),
ztick=(-1.0:0.5:1.0),ztickfont = font(12),
zlim=(-1.5,1.5), # change this it pic does not fit
camera=(-35,55),legend = true)
# field ψv at t=tf=1
surface(xc,zc,transpose(ψvc_f),
xlabel="x",xtick=(0.0:0.25:1.0),xtickfont = font(12),
ylabel="z",ytick=(0.0:2:6),ytickfont = font(12),
#uncomment the following for SH
ztick=(-1.0:0.5:1.0),ztickfont = font(12),
zlim=(-1,1), # change this it pic does not fit
#uncomment the following for WH
#ztick=(-4*1e-11:2*1e-11:4*1e-11),ztickfont = font(12),
#zlim=(-4*1e-11,4*1e-11), # change this it pic does not fit
camera=(-35,55),legend = true)
# field ϕ at t=tf=1
surface(xc,zc,transpose(ϕc_f),
xlabel="x",xtick=(0.0:0.25:1.0),xtickfont = font(12),
ylabel="z",ytick=(0.0:2:6),ytickfont = font(12),
#uncomment the following for SH
ztick=(-1.0:0.5:1.0),ztickfont = font(12),
zlim=(-1,1), # change this it pic does not fit
#uncomment the following for WH
#ztick=(-4*1e-11:2*1e-11:4*1e-11),ztickfont = font(12),
#zlim=(-4*1e-11,4*1e-11), # change this it pic does not fit
camera=(-35,55),legend = true)
| examples/data_analysis/surface_plots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {}}}}
import json
import copy
from ipyleaflet import (
Map,
GeoJSON,
Layer
)
# -
m = Map(center=[53.88, 27.45], zoom=3, name='Base map')
m
# + extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {}, "report_default": {}}}}
# geojson layer with hover handler
with open('./europe_110.geo.json') as f:
data = json.load(f)
for feature in data['features']:
feature['properties']['style'] = {
'color': 'grey',
'weight': 1,
'fillColor': 'grey',
'fillOpacity': 0.2
}
selected_set = set()
selected_layer = None
def convert_selected_set_to_geojson(selected_set):
geojson = {'type': 'FeatureCollection', 'features': []}
geojson['features'] = [feature for feature in data['features'] if feature['properties']['iso_a3'] in selected_set]
for feature in data['features']:
feature['properties']['style'] = {'color': 'yellow', 'weight': 2,'fillColor': 'grey', 'fillOpacity': 0.2}
return geojson
def selected_onclick_handler(event=None, id=None, properties=None, **args):
global selected_layer
if properties is None:
return
cid = properties['iso_a3']
selected_set.remove(cid)
if selected_layer is not None:
m.remove_layer(selected_layer)
selected_layer = GeoJSON(data=convert_selected_set_to_geojson(selected_set),
name='Selected EU Countries',
hover_style={'fillColor': 'yellow'}
)
selected_layer.on_click(selected_onclick_handler)
m.add_layer(selected_layer)
def geojson_onclick_handler(event=None, id=None, properties=None, **args):
global selected_layer
if properties is None:
return
cid = properties['iso_a3']
selected_set.add(cid)
if selected_layer is not None:
m.remove_layer(selected_layer)
selected_layer = GeoJSON(data=convert_selected_set_to_geojson(selected_set),
name='Selected EU Countries',
hover_style={'fillColor': 'yellow'}
)
selected_layer.on_click(selected_onclick_handler)
m.add_layer(selected_layer)
geojson_layer = GeoJSON(data = data, name='EU Countries', hover_style={'fillColor': 'red'})
geojson_layer.on_click(geojson_onclick_handler)
m.add_layer(geojson_layer)
# -
| examples/Select-GeoJson.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/giswqs/geog-312/blob/master/labs/lab_04.ipynb)
# [](https://gishub.org/geog312-pangeo)
#
# **Firstname Lastname**
# +
from datetime import datetime
now = datetime.now()
print(f"Submitted time: {now}")
# -
# ## Question 1
# **Alien Colors # 1:** Imagine an alien was just shot down in a game. Create a variable called `alien_color` and assign it a value of `green, yellow,` or `red`.
#
# * Write an if statement to test whether the alien’s color is green. If it is, print a message that the player just earned 5 points.
# * Write one version of this program that passes the if test and another that fails. (The version that fails will have no output.)
# ## Question 2
# **Alien Colors # 2:** Choose a color for an alien as you did in Question 1, and write an `if-else` chain.
#
# * If the alien’s color is green, print a statement that the player just earned 5 points for shooting the alien.
# * If the alien’s color isn’t green, print a statement that the player just earned 10 points.
# * Write one version of this program that runs the if block and another that runs the else block.
# ## Question 3
# **Alien Colors # 3:** Turn your `if-else` chain from Question 2 into an `if-elif-else` chain.
#
# * If the alien is green, print a message that the player earned 5 points.
# * If the alien is yellow, print a message that the player earned 10 points.
# * If the alien is red, print a message that the player earned 15 points.
# * Write three versions of this program, making sure each message is printed for the appropriate color alien.
# ## Question 4
# **Stages of Life:** Write an `if-elif-else` chain that determines a person’s stage of life. Set a value for the variable `age`, and then:
#
# * If the person is less than 2 years old, print a message that the person is a baby.
# * If the person is at least 2 years old but less than 4, print a message that the person is a toddler.
# * If the person is at least 4 years old but less than 13, print a message that the person is a kid.
# * If the person is at least 13 years old but less than 20, print a message that the person is a teenager.
# * If the person is at least 20 years old but less than 65, print a message that the person is an adult.
# ## Question 5
# **Favorite Fruit:** Make a list of your favorite fruits, and then write a series of independent `if` statements that check for certain fruits in your list.
#
# * Make a list of your three favorite fruits and call it favorite_fruits.
# * Write five if statements. Each should check whether a certain kind of fruit is in your list. If the fruit is in your list, the if block should print a statement, such as You really like bananas!
# ## Question 6
# **Hello Admin:** Make a list of five or more usernames, including the name `admin`. Imagine you are writing code that will print a greeting to each user after they log in to a website. Loop through the list, and print a greeting to each user:
#
# * If the username is 'admin', print a special greeting, such as *Hello admin, would you like to see a status report?*
# * Otherwise, print a generic greeting, such as *Hello Jaden, thank you for logging in again*.
# ## Question 7
# **No Users:** Based on Question 6, add an `if` test to make sure the list of users is not empty.
#
# * If the list is empty, print the message *We need to find some users!*
# * Remove all of the usernames from your list, and make sure the correct message is printed.
# ## Question 8
# **Checking Usernames:** Do the following to create a program that simulates how websites ensure that everyone has a unique username.
#
# * Make a list of five or more usernames called `current_users`.
# * Make another list of five usernames called `new_users`. Make sure one or two of the new usernames are also in the `current_users` list.
# * Loop through the `new_users` list to see if each new username has already been used. If it has, print a message that the person will need to enter a new username. If a username has not been used, print a message saying that the username is available.
# * Make sure your comparison is case insensitive. If 'John' has been used, 'JOHN' should not be accepted. (To do this, you’ll need to make a copy of `current_users` containing the lowercase versions of all existing users.)
# ## Question 9
# **Ordinal Numbers:** Ordinal numbers indicate their position in a list, such as *1st* or *2nd*. Most ordinal numbers end in *th*, except 1, 2, and 3.
#
# * Store the numbers 1 through 9 in a list.
# * Loop through the list.
# * Use an `if-elif-else` chain inside the loop to print the proper ordinal ending for each number. Your output should read "1st 2nd 3rd 4th 5th 6th 7th 8th 9th", and each result should be on a separate line.
| docs/labs/lab_04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IoT Microdemos
#
#
# ## Timeseries
# A common pattern to store and retrieve time series data is to leverage the document model with the so called bucketing schema pattern. Instead of storing each measurement into a single document, multiple measurements are stored into one single document. This provides the benefits of:
# * Reducing Storage space (as less data is stored multiple times, e.g. device id and other metadata, as well as better compression ratios on larger documents)
# * Reduce Index sizes (by bucket size), larger parts of the index will fit into memory and increase performance
# * Reduce IO by less documents (reading time series at scale is usually IO-bound load)
#
# The following examples will guide through the typical patterns:
# * [Ingesting Data](#Ingesting-Data)
# * [Indexing Data](#Indexing-Strategy)
# * [Querying Data](Querying-the-Data)
# ## Ingesting Data
#
# The following statement will search for a document of device 4711 and where the count of measurements is less than 3 entries in the bucket. In reality, this will be a higher number, e.g. 60 or 100. The new measurement is pushed to the array called m.
#
# Because of the upsert option, a new document will be inserted, if no available bucket can be found. Increasing the cnt by one during each insert will automatically create a new document once the exiting bucket is full.
#
# ### Initialize the database and drop the collection:
# + pycharm={"is_executing": false, "name": "#%%\n"}
import pymongo
import os
import datetime
import bson
from bson.json_util import loads, dumps, RELAXED_JSON_OPTIONS
import random
from pprint import pprint
CONNECTIONSTRING = "localhost:27017"
# Establish Database Connection
client = pymongo.MongoClient(CONNECTIONSTRING)
db = client.iot
collection = db.iot_raw
# Drop the collection before we start with the demo
collection.drop()
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Insert the first measurements:
# MongoDB Query Language offer rich operators that we leverage here to automatically bucket the data, i.e. we do not store each individual measurement into one document, but store multiple measurements into an array.
#
# By using upsert, we automatically start a new bucket, i.e. create a new document if no bucket with additional space can be found. Otherwise, we push the new measurement into the bucket.
#
# The following statement will find an open bucket of device 4711, i.e. where the count of measurements is less than 3 entries in the bucket. In reality, this will be a higher number, e.g. 60 or 100. The new measurement is pushed to the array called m, the bucket size is increased by one. For the later query on time ranges, we also store the minimal and maximal timestamp within this bucket.
# + pycharm={"name": "#%%\n"}
# The timestamp of the new measurement
# Note: For better readability, we work with datetime objects.
# For higher precision of timestamps, e.g. nanoseconds,
# it is recommended to work with decimal values representing seconds and nanoseconds
date = datetime.datetime.now()
# Add the new measurement to the bucket
collection.update_one({
"device": 4711,
"cnt": { "$lt": 3 }
},
{
"$push": {
"m": {
"ts": date,
"temperature": random.randint(0,100),
"rpm": random.randint(0,10000),
"status": "operating"
}
},
"$max": { "max_ts": date },
"$min": { "min_ts": date },
"$inc": { "cnt": 1 }
},
upsert=True);
# + [markdown] pycharm={"name": "#%% md\n"}
# The target document looks like the following:
# + pycharm={"name": "#%%\n"}
result = collection.find_one()
pprint(result)
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Add additional measurements
#
# Insert some more data in order to have multiple buckets (again, here we use a bucket size of 3, in reality this number will be much higher). We Iinsert four more measurements, so there will be two documents with 3 and 2 measurements, respectively.
# + pycharm={"name": "#%%\n"}
for i in range(4):
date = datetime.datetime.now()
collection.update_one(
{
"device": 4711,
"cnt": { "$lt": 3 }
},
{
"$push": {
"m": {
"ts": date,
"temperature": random.randint(0,100),
"rpm": random.randint(0,10000),
"status": "operating",
"new_field": { "subfield1": "s1", "subfield2": random.randint(0,100)}
}
},
"$max": { "max_ts": date },
"$min": { "min_ts": date },
"$inc": { "cnt": 1 }
},
upsert=True
)
# + [markdown] pycharm={"name": "#%% md\n"}
# The result will look like the following:
# + pycharm={"name": "#%%\n"}
res = collection.find()
for doc in res:
pprint(doc)
# -
# ## Indexing Strategy
#
# A proper indexing strategy is key for efficient querying of data. The first index is mandatory for efficient time series queries in historical data. The second one is needed for efficient retreival of the current, i.e. open, bucket for each device. If all device types have the same bucket size, it can be created as a partial index - this will only keep the open buckets in the index. For varying bucket sizes, e.g. per device type, the type could be added to the index. The savings can be huge for large implementations.
# +
# Efficient queries per device and timespan
result = collection.create_index([("device",pymongo.ASCENDING),
("min_ts",pymongo.ASCENDING),
("max_ts",pymongo.ASCENDING)])
print("Created Index: " + result)
# Efficient retreival of open buckets per device
result = collection.create_index([("device",pymongo.ASCENDING),
("cnt",pymongo.ASCENDING)],
partialFilterExpression={"cnt": {"$lt":3}})
print("Created Index: " + result)
# -
# These indexes will be used during the ingestion as well as the retreival process. And we will have a closer look at them later on.
# ## Querying the Data
#
# With Aggregation Pipelines it is easy to query, filter, and format the data. This is the query for two timeseries (temperature and rpm). The sort should use the full index prefix in order to be executed on the index and not in memory.
# +
result = collection.aggregate([
{ "$match": { "device": 4711 } },
{ "$sort": { "device": 1, "min_ts": 1 } },
{ "$unwind": "$m" },
{ "$sort": { "m.ts": 1 } },
{ "$project": { "_id": 0, "device": 1, "ts": "$m.ts", "temperature": "$m.temperature", "rpm": "$m.rpm" } }
]);
for doc in result:
print(doc)
# -
# In order to query for a certain timeframe, the following $match stage can be used to search for a certain timeframe (please replace LOWER_BOUND and UPPER_BOUND with appropriate ISODate values).
# +
LOWER_BOUND = datetime.datetime(2020, 4, 20, 13, 26, 43, 18000) # Replace with lower bound (copy & paste from results above)
UPPER_BOUND = datetime.datetime(2020, 4, 20, 13, 30, 26, 130000) # Replace with upper bound (copy & paste from results above)
result = collection.aggregate([
{ "$match": { "device": 4711, "min_ts": { "$lte": UPPER_BOUND }, "max_ts": { "$gte": LOWER_BOUND } } },
{ "$sort": { "device": 1, "min_ts": 1 } },
{ "$unwind": "$m" },
{ "$match": { "$and": [ { "m.ts": { "$lte": UPPER_BOUND } }, { "m.ts": { "$gte": LOWER_BOUND } } ] } },
{ "$project": { "_id": 0, "device": 1, "ts": "$m.ts", "temperature": "$m.temperature", "rpm": "$m.rpm" } }
]);
for doc in result:
print(doc)
# -
# ### How to explain this query pattern
#
# We want to get the data from timestamps 8 to 17 that are spread across 5 buckets:
# ```
# (1) 1 2 3 4 5
# (2) 6 7 8 9 10
# (3) 11 12 13 14 15
# (4) 16 17 18 19 20
# (5) 21 22 23
# ```
#
# We could use a complex condition, but this will end up in expensive index scans:
# ```
# min <= 8 and max >= 8 [ bucket (1) ]
# OR: min >= 8 and max <= 17 [ bucket (3) ]
# OR: min <= 17 and max >= 8 [ bucket (4) ]
# ```
#
# The following statement leads to the same result and allows efficient index traversal and selects exactly the buckets of our interest:
# ```
# max >= 8
# AND: min <= 17
# ```
| mongodb-timeseries/iot_timeseries_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (augur)
# language: python
# name: augur
# ---
# +
import augur
from augur.githubapi import GitHubAPI
from augur.ghtorrent import GHTorrent
# import everything from githubapi.py and ghtorrent.py so we can
# just copy and paste our function later
import json
import re
from dateutil.parser import parse
import pandas as pd
import github
import numpy as np
import sqlalchemy as s
import datetime
import requests
import time
# %matplotlib inline
# create an Augur application so we can test our function
augurApp = augur.Application('../augur.config.json')
github = augurApp.githubapi()
ghtorrent = augurApp.ghtorrent()
# -
#
# ## GitHub API Example Function
# We are writing this function to be run as part of the GitHub class, so
# the "self" in this function will be an instance of an augur.GitHubAPI
def lines_deleted(self, owner, repo=None):
"""
Additions and deletions each week
:param owner: The name of the project owner
:param repo: The name of the repo
:return: DataFrame with each row being am issue
"""
# get the data we need from the GitHub API
# see <project_root>/augur/githubapi.py for examples using the GraphQL API
url = "https://api.github.com/repos/{}/{}/stats/code_frequency".format(owner, repo)
json = requests.get(url, auth=('user', self.GITHUB_API_KEY)).json()
# get our data into a dataframe
df = pd.DataFrame(json, columns=['date', 'additions', 'deletions'])
# all timeseries metrics need a 'date' column
df['date'] = pd.to_datetime(df['date'], unit='s', infer_datetime_format=True)
# normalize our data and create useful aggregates
df['deletions'] = df['deletions'] * -1
df.drop(columns=['additions'])
# return the dataframe
return df
# +
# add our new function to the class
GitHubAPI.lines_deleted = lines_deleted
time.sleep(10)
# test our function on the initialized instance
ld = github.lines_deleted('osshealth', 'augur')
ld['deletions'].plot()
# -
#
# ## GHTorrent Example Function
# +
# all Augur functions that provide data should accept at least these arguments
#
# we are writing this function to be run as part of the GHTorrent class, so
# the "self" in this function will be an instance of an augur.GHTorrent
def issues_closed_by_month(self, owner, repo=None):
# get the repoid we will need in our query
repoid = self.repoid(owner, repo)
# write our query
sql = s.sql.text("""
SELECT DATE_FORMAT(issue_events.created_at, '%Y-%m') as "date", COUNT(*) as issues_closed
FROM issue_events, issues
WHERE issue_events.issue_id = issues.id
AND issue_events.action = "closed"
AND issues.repo_id = :repoid
GROUP BY YEAR(issue_events.created_at), MONTH(issue_events.created_at)
""")
# return a dataframe of the results of the query
return pd.read_sql(sql, self.db, params={"repoid": str(repoid)})
# add our new function to the class
GHTorrent.issues_closed_by_month = issues_closed_by_month
# -
# test our function on the initialized instance
rails_closed_per_month = ghtorrent.issues_closed_by_month('rails', 'rails')
rails_closed_per_month.plot()
| notebooks/example-metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
data = [ [1,2,3], [4,5,6], [7,8,9] ]
data
list('321')
df = pd.DataFrame(data, list('321'), list('ZYX'))
df
df.sort_index(inplace = True)
df
df.sort_index(inplace = True, axis = 1)
df
df.sort_values(by = 'X', inplace = True)
df
df.sort_values(by = '3', axis = 1, inplace = True)
df
df.sort_values(by = ['X', 'Y'], inplace = True)
df
| extras/Organizando DataFrame (Sort).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from pymongo import MongoClient, UpdateOne # https://pymongo.readthedocs.io/en/stable/tutorial.html
import dns
import requests
import requests_cache # https://requests-cache.readthedocs.io/en/stable/
import datetime
from typing import List, Dict, Tuple, Optional
import json
import time
from dotenv import load_dotenv
import os
from bs4 import BeautifulSoup
import random
from scrape.stack import StackOverflowScraper
from urllib.parse import quote_plus
load_dotenv()
None
# +
# Connect to MongoDB
def get_mongo_client() -> MongoClient:
# A MongoDB connection string _must_ be provided
mongo_uri = os.getenv('MONGO_URI')
if not mongo_uri:
raise Exception('MONGO_URI not set')
# URI may either be a fully usable mongo connection string, or a printf-style string
# where the username and password are specified as %s and %s respectively
# e.g. mongodb+srv://%s:%s@mongodb.mydomain.com:27017/someDatabase
# See:
# https://pymongo.readthedocs.io/en/stable/api/pymongo/mongo_client.html#pymongo.mongo_client.MongoClient
# https://docs.mongodb.com/manual/reference/connection-string/
user = os.getenv('MONGO_USERNAME') or os.getenv('MONGO_PUBLIC_KEY')
pw = os.getenv('MONGO_PASSWORD') or os.getenv('MONGO_PRIVATE_KEY')
if user and pw:
mongo_uri = mongo_uri % ( quote_plus(user), quote_plus(pw))
else:
assert not user and not pw, 'MONGO_USERNAME and MONGO_PASSWORD must both be set or both be unset'
client = MongoClient(mongo_uri, tls=True)
return client
client = get_mongo_client()
db = client['stackOverflowDB']
questions = db['questions']
answers = db['answers']
# -
# Set up request caching for StackOverflow API
session = requests_cache.CachedSession('.cache/stack_cache', cache_control=True, stale_if_error=True, backend='filesystem')
# # Getting StackOverflow Questions
#
# Questions are procured from the [StackOverflow REST API](https://api.stackexchange.com/docs), specifically the [/questions endpoint](https://api.stackexchange.com/docs/questions#order=desc&sort=activity&tagged=c%3Bc%2B%2B&filter=default&site=stackoverflow). We'll be limiting our search to C/C++ code snippets for simplicity.
def get_stackoverflow_questions(**kwargs):
"""
"""
pagesize: int = kwargs.get('pagesize', 100) # How many questions to return per page
assert 1 <= pagesize <= 100 # Stack allows [0, 100] but why waste API calls?
page: int = kwargs.get('page', 1) # Starting page index, 1-indexed
assert page >= 1
maxpages: int = kwargs.get('maxpages', 10) # Max number of pages to return
assert maxpages >= 1
api_key: Optional[str] = kwargs.get('api_key')
question_boundary_younger = datetime.datetime(2021, 12, 4) # No questions posted more recently than this will be returned
done = False # Set to True if we hit our request quota or no more question data is available
requests_made = 0
# StackOverflow API query parameters common across all queries
base_query_params: dict = {
'site': 'stackoverflow',
'sort': 'activity',
'order': 'desc',
'tagged': 'c',
'pagesize': pagesize,
'todate': int(question_boundary_younger.timestamp())
}
# Include the API key if one was provided
if api_key:
base_query_params['key'] = api_key
while not done and requests_made < maxpages:
query_params = base_query_params.copy()
query_params['page'] = page
# Returns a Common Wrapper Object
# https://api.stackexchange.com/docs/wrapper
r = session.get('https://api.stackexchange.com/2.3/questions', params=query_params)
if r.status_code > 299:
if r.headers['content-length'] == 0:
r.raise_for_status()
elif 'json' in r.headers['content-type']:
error_json = r.json()
raise requests.HTTPError(f'{r.status_code} {r.reason} API returned error {error_json["error_id"]}: {error_json["error_message"]}')
else:
raise requests.HTTPError(f'{r.status_code} {r.reason}: {r.text}')
assert 'json' in r.headers['content-type'] # We're expecting JSON back
requests_made += 1
page += 1
# Yield each question in the response
body = r.json()
assert 'items' in body
assert isinstance(body['items'], list)
yield body['items']
# Check if we're done
quota_remaining = body['quota_remaining']
quota_max = body['quota_max']
has_more: bool = body['has_more']
done = not body['has_more'] or body['quota_remaining'] <= 0
print('\r', f'Got {pagesize} questions from page #{page} (quota: {quota_remaining}/{quota_max})', end='')
# Check if we need to back off before sending more requests. Only necessary if we're not done.
backoff = body.get('backoff', 0)
if not done and backoff > 0:
print(f'Backoff requested, sleeping for {backoff} seconds')
time.sleep(backoff)
stack = StackOverflowScraper(cache=session)
# +
# This takes a while, is expensive, and is only necessary once. This flag
# lets you skip this step if you've already run it.
should_scrape = False
drop = False
page_size = 100 # Number of questions to return per page
page = 185 # Starting page index, 1-indexed. Useful for continuing where you left off in the event of a crash
if should_scrape:
if drop:
print('Dropping questions collection')
questions.drop()
print('Scraping questions')
# Scrape each page, bulk inserting each one into mongo
for page in stack.get_questions(page=page, maxpages=100, pagesize=page_size):
if type(page) is not list:
assert type(page) is dict
page = [page]
# Remove questions with no answers. Also, questions with low scores are less likely
# to have useful answers, it's probably just someone insulting the poster for
# being a noob.
page = filter(lambda q: q['answer_count'] > 0 and q['score'] >= 0, page)
upserts = [UpdateOne({'_id': q['question_id']}, {'$set': q}, upsert=True) for q in page]
questions.bulk_write(upserts)
time.sleep(0.5 + random.random() / 2) # Sleep for a bit to avoid hitting the API too hard
# -
# # Scraping StackOverflow Answers
def get_questions(**kwargs):
pagesize = kwargs.get('pagesize', 100) # How many questions to return per page
assert 1 <= pagesize
page = kwargs.get('page', 1) # Starting page index, 1-indexed
assert page >= 1
# Calculate number of documents to skip
skips = page_size * (page_num - 1)
# Skip and limit
cursor = questions.find().skip(skips).limit(page_size)
for doc in cursor:
yield doc
# +
def scrape_stackoverflow_page(url: str) -> List[Dict]:
# Load the page into BeautifulSoup
r = session.get(url)
html_doc = r.text
soup = BeautifulSoup(html_doc, 'html.parser')
answers = soup.select('.answer')
answers_parsed = []
for answer in answers:
answer_cell = answer.select_one('.answercell')
answer_id = int(answer['data-answerid'])
# Get all code snippet elements for the answer, skipping if there are none
snippet_elems = answer_cell.select('pre > code')
if not len(snippet_elems):
continue
# Contains the user name and id of the answerer
user_details = answer.select_one('.post-signature .user-details > a')
# Extract the answer author's user id. Anonymous users have no user id
if user_details is None:
user_id = None
user_name = 'anonymous'
else:
_, _, user_id, user_name = user_details['href'].split('/') # takes form /users/:id/:name
user_id = int(user_id) # May be -1 if posted by 'community'
answer_data = {
# 'question_id': question_id,
'snippets': '\n'.join([code_block.text for code_block in snippet_elems]),
'score': int(answer['data-score']),
'answer_id': answer_id,
'page_pos': int(answer['data-position-on-page']),
'is_highest_scored': answer['data-highest-scored'] == '1',
'question_has_highest_accepted_answer': answer['data-question-has-accepted-highest-score'] == '1',
# 'is_accepted': answer.has_class('accepted-answer'),
'is_accepted': 'accepted-answer' in answer['class'],
# 'source': answer.select_one('a.js-share-link').get('href').strip(),
'source': f'https://stackoverflow.com/a/{answer_id}',
'author_id': user_id,
'author_username': user_name,
}
answers_parsed.append(answer_data)
return answers_parsed
# Test that the scraper works
test_data = scrape_stackoverflow_page('https://stackoverflow.com/questions/69729326/endless-sine-generation-in-c')
assert type(test_data) is list
assert len(test_data) > 0
for answer in test_data:
assert type(answer) is dict
# assert answer['question_id'] == 69729326 # This is the question we're scraping
assert 'snippets' in answer
assert 'score' in answer
assert 'answer_id' in answer
assert 'page_pos' in answer
assert 'is_highest_scored' in answer
assert 'question_has_highest_accepted_answer' in answer
assert 'is_accepted' in answer
assert 'source' in answer
assert 'author_id' in answer
assert 'author_username' in answer
print(test_data[0]['snippets'])
# +
drop = False # Set to True to drop the collection before scraping
page_size = 100 # The number of questions to scrape in each page
start_page = 170 # The page to start scraping at, allows for resuming scraping after a crash
should_scrape = True # Set to True to scrape the questions collection
num_pages = int(questions.count_documents({}) / page_size)
if should_scrape:
# Drop the collection if we're dropping it
if drop:
print('Dropping answers collection')
answers.drop()
# Scrape each page of questions, bulk inserting answers into mongo
assert start_page > 0
for page_num in range(start_page, num_pages + 1):
# page_num = i + 1
print(f'Scraping page {page_num}/{num_pages} ', end = '')
for question in get_questions(page=page_num, pagesize=page_size):
# Get the answers for this question, skipping if no relevant answers are available
answers_data = scrape_stackoverflow_page(question['link'])
if not len(answers_data):
print('x', end = '')
continue
# Add the question id to each answer
for answer_data in answers_data:
answer_data['question_id'] = question['question_id']
# Bulk insert the answers
upserts = [UpdateOne({'_id': answer['answer_id']}, {'$set': answer}, upsert=True) for answer in answers_data]
answers.bulk_write(upserts)
print('.', end = '')
time.sleep(0.60 + random.random()) # Don't spam the server, otherwise CloudFlare will complain
print('')
# -
| final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %pylab inline
# %config InlineBackend.figure_format = 'retina'
import seaborn as sns
from scipy.stats import gaussian_kde
import scipy.interpolate as sip
import scipy.integrate as cumtrapz
import h5py
# -
# ### set up mixture model for spin magnitude distribution
HM_spin_samples = loadtxt("/Users/mayafishbach/Desktop/BHspins/spin_for_plots/4HBRmerger_69_2500.txt") #change filepath appropriately
HM_spin_kde = gaussian_kde(HM_spin_samples)
sns.distplot(HM_spin_samples)
chis = linspace(0.,1.,100)
plt.plot(chis,HM_spin_kde(chis))
plt.xlabel(r'$a$')
plt.ylabel(r'$p(a)$')
plt.show()
HM_spin_interp = sip.interp1d(chis,HM_spin_kde(chis))
# +
def pa1a2_mixture(a1,a2,f):
return (f*HM_spin_interp(a1)+(1.-f))*(f*HM_spin_interp(a2)+(1.-f))
def pa1a2_mixture_smallspins(a1,a2,f):
return (f*HM_spin_interp(a1)+(1.-f)*2.*(1.-a1))*(f*HM_spin_interp(a2)+(1.-f)*2.*(1.-a2))
def pa1a2_mixture_primary(a1,a2,f):
return (f*HM_spin_interp(a1)+(1.-f))
# -
# ### hierarchical Bayesian likelihood, assuming no selection effects
def loglikelihood(a1s,a2s,pa1a2):
fs = linspace(0.0,1.0,100)
ll_sum = zeros_like(fs)
def onesysmeanl(a1,a2,f):
return mean(pa1a2(a1,a2,f))
def logl(f):
ll = 0.0
for a1, a2 in zip(a1s, a2s):
res = log(onesysmeanl(a1, a2, f))
ll += res
return ll
for i,f in enumerate(fs):
ll_sum[i] = logl(f)
return fs, ll_sum
# ### Read in samples
import glob
a1_list_SEOB = []
a2_list_SEOB = []
a1_list_IMRP = []
a2_list_IMRP = []
a1_list_comb = []
a2_list_comb = []
evt_names = ['GW150914', 'GW151012', 'GW151226', 'GW170104',
'GW170608', 'GW170729', 'GW170809', 'GW170814', 'GW170818', 'GW170823']
for evt_name in evt_names:
fname = glob.glob("/Users/mayafishbach/Downloads/GWTC-1_sample_release/*"+evt_name+"*.hdf5")[0] #change file path
with h5py.File(fname,'r') as inp:
print fname
a1_list_SEOB.append(array(inp['SEOBNRv3_posterior']['spin1']))
a2_list_SEOB.append(array(inp['SEOBNRv3_posterior']['spin2']))
a1_list_IMRP.append(array(inp['IMRPhenomPv2_posterior']['spin1']))
a2_list_IMRP.append(array(inp['IMRPhenomPv2_posterior']['spin2']))
a1_list_comb.append(array(inp['Overall_posterior']['spin1']))
a2_list_comb.append(array(inp['Overall_posterior']['spin2']))
sns.set_palette('husl',5)
for i in range(5):
sns.kdeplot(a1_list_IMRP[i],label=evt_names[i])
sns.kdeplot(a1_list_SEOB[i],ls='--',c=sns.color_palette()[i])
sns.kdeplot(a1_list_comb[i],ls=':',c=sns.color_palette()[i])
plt.legend(loc='best')
plt.show()
for i in range(5):
sns.kdeplot(a1_list_IMRP[i+5],label=evt_names[i+5])
sns.kdeplot(a1_list_SEOB[i+5],ls='--',c=sns.color_palette()[i])
sns.kdeplot(a1_list_comb[i+5],ls=':',c=sns.color_palette()[i])
plt.legend(loc='best')
plt.show()
# ### calculate posterior on f, the mixing fraction for different models and different sets of PE samples
fs, ll_sum_mixture_IMRP = loglikelihood(a1_list_IMRP,a2_list_IMRP,pa1a2_mixture)
fs, ll_sum_mixture_SEOB = loglikelihood(a1_list_SEOB,a2_list_SEOB,pa1a2_mixture)
fs, ll_sum_mixture_comb = loglikelihood(a1_list_comb,a2_list_comb,pa1a2_mixture)
fs, ll_sum_mixture_low_IMRP = loglikelihood(a1_list_IMRP,a2_list_IMRP,pa1a2_mixture_smallspins)
fs, ll_sum_mixture_low_SEOB = loglikelihood(a1_list_SEOB,a2_list_SEOB,pa1a2_mixture_smallspins)
fs, ll_sum_mixture_low_comb = loglikelihood(a1_list_comb,a2_list_comb,pa1a2_mixture_smallspins)
# +
plt.plot(fs,exp(ll_sum_mixture_IMRP)/trapz(exp(ll_sum_mixture_IMRP),fs),label='IMRP samples')
plt.plot(fs,exp(ll_sum_mixture_SEOB)/trapz(exp(ll_sum_mixture_SEOB),fs),label='SEOB samples')
plt.plot(fs,exp(ll_sum_mixture_comb)/trapz(exp(ll_sum_mixture_comb),fs),label='combined samples')
plt.xlabel('$f$')
plt.ylabel('$p(f)$')
plt.title('fraction formed from hierarchical mergers vs. uniform spin magnitude distribution')
plt.show()
plt.plot(fs,exp(ll_sum_mixture_low_IMRP)/trapz(exp(ll_sum_mixture_low_IMRP),fs),label='IMRP asmples')
plt.plot(fs,exp(ll_sum_mixture_low_SEOB)/trapz(exp(ll_sum_mixture_low_SEOB),fs),label='SEOB samples')
plt.plot(fs,exp(ll_sum_mixture_low_comb)/trapz(exp(ll_sum_mixture_low_comb),fs),label='combined samples')
plt.title('fraction formed from hierarchical mergers vs. low spin magnitude distribution')
plt.xlabel('$f$')
plt.ylabel('$p(f)$')
plt.show()
# -
# ### the ratio p(f=1)/p(f=0) gives odds ratio between the two models
print exp(ll_sum_mixture_IMRP[0]-ll_sum_mixture_IMRP[-1])
print exp(ll_sum_mixture_SEOB[0]-ll_sum_mixture_SEOB[-1])
print exp(ll_sum_mixture_comb[0]-ll_sum_mixture_comb[-1])
print exp(ll_sum_mixture_low_IMRP[0]-ll_sum_mixture_low_IMRP[-1])
print exp(ll_sum_mixture_low_SEOB[0]-ll_sum_mixture_low_SEOB[-1])
print exp(ll_sum_mixture_low_comb[0]-ll_sum_mixture_low_comb[-1])
# ### Reading in data for the BNS
#
# This is the data relevant for the H0 measurement
#
# Use either the distance samples from the analysis with the high spin prior (called simply distance below) or the distance from the low spin prior analysis (distance_lowspin)
with h5py.File('/Users/mayafishbach/Downloads/GWTC-1_sample_release/GW170817_GWTC-1.hdf5','r') as inp: #change file path appropriately
distance = array(inp['IMRPhenomPv2NRT_highSpin_posterior']['luminosity_distance_Mpc'])
cosinc = array(inp['IMRPhenomPv2NRT_highSpin_posterior']['costheta_jn'])
distance_lowspin = array(inp['IMRPhenomPv2NRT_lowSpin_posterior']['luminosity_distance_Mpc'])
sns.kdeplot(distance,cosinc)
sns.distplot(distance)
sns.distplot(distance_lowspin)
| lecture8/HMspins.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # NY Parking Violation Data Analysis
#
# **Mini Project**
#
# IISC | M.Tech (Online) | DSBA
# DA 231-O Data Engineering at Scale
#
# Mentor: **<NAME>** (<EMAIL>)
# ## Team
#
# 1. <NAME> | <EMAIL> | 9092696415
# 2. <NAME> | <EMAIL> | 9986778909
# 3. <NAME> | <EMAIL> | 9886702749
# 4. <NAME> | <EMAIL> | 9035076656
# ## Problem
#
# * Exploratory data analysis of **NYC Parking violation** data for **2017** – **2021** years (5 years)
# * Infer from the data analysis findings
#
# > **Dataset** : https://data.cityofnewyork.us/City-Government/Parking-Violations-Issued-Fiscal-Year-2022/pvqr-7yc4
# ## Environment
#
# * Python 3
# * Spark 3.0 (colab) / 3.1 (local)
# * Spark Data Frames & Spark SQL
# * Local & Google Colab
# ## Dataset Details
# Dataset contains New York city Police Department issued parking violations details
# ### Raw Dataset details
# - Fiscal year data (July 1 - June 30) is available. That means one file contains data for 2 years
# - All years data has same 43 columns
# - Total Number of records: **xxxx** # TODO
# ### Processed Data details
# - Removed 30 columns which are not used in our analysis (Most of the removed columns don't have data for > 50% records)
# - Total Columns: 13
# - Total Number of records: **xxxx** # TODO
# ### Columns Summary
# | S. No. | Source Column Name | Description/Comment |
# |--------|-------------------------|------------------------------------------------------|
# | 1 | SUMMONS NUMBER | UNIQUE IDENTIFIER OF SUMMONS |
# | 2 | PLATE ID | REGISTERED PLATE ID |
# | 3 | REGISTRATION STATE | STATE OF PLATE REGISTRATION |
# | 4 | PLATE TYPE | TYPE OF PLATE |
# | 5 | ISSUE DATE | ISSUE DATE |
# | 6 | VIOLATION CODE | TYPE OF VIOLATION |
# | 7 | VEHICLE BODY TYPE | VEHICLE BODY TYPE WRITTEN ON SUMMONS (SEDAN, ETC.) |
# | 8 | VEHICLE MAKE | MAKE OF CAR WRITTEN ON SUMMONS |
# | 9 | VIOLATION PRECINCT | POLICE STATION OF VIOLATION |
# | 10 | ISSUER PRECINCT | PRECINCT OF ISSUANCE |
# | 11 | VIOLATION TIME | VIOLATION TIME |
# | 12 | VEHICLE COLOR | CAR COLOR WRITTEN ON SUMMONS |
# | 13 | VIOLATION DESCRIPTION | DESCRIPTION OF VIOLATION |
# ### Columns Details
# #### Plate Type
#
# Registration Class Codes for vehicles. 3 letters code
#
# Common Plate types are
# * Passenger Vehicles (PAS): standard issue plates
# * Commercial Vehicles (COM): Full-size vans and most pickups
# * Medallion (OMT): Taxis
# * Personalized Plates (SRF): cars, mini-vans, SUVs and some pick-ups registered as passenger class
# * Special Omnibus Rentals (OMS)
#
# https://dmv.ny.gov/registration/registration-class-codes
#
# #### Violation Code
# Type of violation. Codes are from 1-99. Fines are charged based on this
#
# Some of the common violation codes & their fines
#
# | Violation Code | Description | Fine |
# |----------------|----------------------------------|------|
# | 36 | PHTO SCHOOL ZN SPEED VIOLATION | 50 |
# | 21 | NO PARKING-STREET CLEANING | 45 |
# | 38 | FAIL TO DSPLY MUNI METER RECPT | 35 |
# | 14 | NO STANDING-DAY/TIME LIMITS | 115 |
# | 20 | NO PARKING-DAY/TIME LIMITS | 60 |
#
# https://data.cityofnewyork.us/api/views/pvqr-7yc4/files/7875fa68-3a29-4825-9dfb-63ef30576f9e?download=true&filename=ParkingViolationCodes_January2020.xlsx
#
# #### Vehicle Body Type
#
# Common Vehicle body types are
# * suburban(SUBN): Vehicle that can be used to carry passengers and cargo
# * four-door sedan (4DSD)
# * Van Truck (VAN
# * Delivery Truck (DELV)
# * Pick-up Truck (PICK)
# * two-door sedan (2DSD)
# * Sedan (SEDN)
#
# https://nysdmv.custhelp.com/app/answers/detail/a_id/491/kw/body%20type%20subn
#
# #### Vehicle Make
#
# The DMV code for the make of a vehicle that appears on the registration. The DMV make code is the first 5 letters of the vehicle’s make name. If the vehicle make is more than one word, the make code is the first 2 letters of the first two words with a slash in between
#
# Common Vehicle Makes are
# * Honda (HONDA)
# * Toyota (TOYOT)
# * Ford (FORD)
# * Nissan (NISSA)
# * Chevrolet (CHEVR)
# * mercedes benz (ME/BE)
#
# https://data.ny.gov/Transportation/Vehicle-Makes-and-Body-Types-Most-Popular-in-New-Y/3pxy-wy2i
# https://data.ny.gov/api/assets/83055271-29A6-4ED4-9374-E159F30DB5AE
#
# #### Vehicle Colors
#
# Common colors are
# * Gray (GY)
# * White (WH)
# * Black (BK)
# * Blue (BL)
# * Red (RD)
# ## Config
# +
# Execution environment. Use False for local execution
colab_env = False
drive_path = "/content/drive/MyDrive/Project/data_eng_at_scale" # Project home
# Use Sample file for speedy execution
processed_file = True
# Set to full-sanitized-data.csv for full sanitized file (4.6 GB)
processed_file_path = f'{drive_path}/data/sample-100000.csv' if colab_env else "../data/sample-100000.csv"
# Specify the raw CSV files path
raw_CSV_path = f'{drive_path}/data/Parking_*.csv' if colab_env else "../data/Parking*.csv"
csv_files = processed_file_path if processed_file else raw_CSV_path
# For faster execution. Some statements are skipped based on this check
presenting = False
# Specify the years for which we are reading the data from CSV
years = [2017, 2018, 2019, 2020, 2021]
# Schema Types. Only specify for the non-string type & NULL columns. Others are considered as string
schema_types = {
"Summons Number": {"type": "long", "null": False},
"Issue Date" : {"type": "date" if processed_file else "string", "null": True},
"Violation Code": {"type": "integer", "null": True},
"Violation Precinct": {"type": "integer", "null": True},
"Issuer Precinct": {"type": "integer", "null": True},
}
# Columns which are used in the analysis. Other columns will be removed
used_columns = ["Summons Number", "Plate ID", "Registration State", "Plate Type", "Issue Date", "Violation Code", "Vehicle Body Type", "Vehicle Make", "Violation Precinct", "Issuer Precinct", "Violation Time", "Vehicle Color", "Violation Description"]
# All the columns which are there in the datset (Need to be in CSV file order)
schema_columns = ["Summons Number", "Plate ID", "Registration State", "Plate Type", "Issue Date", "Violation Code", "Vehicle Body Type", "Vehicle Make", "Issuing Agency", "Street Code1", "Street Code2", "Street Code3", "Vehicle Expiration Date", "Violation Location", "Violation Precinct", "Issuer Precinct", "Issuer Code", "Issuer Command", "Issuer Squad", "Violation Time", "Time First Observed", "Violation County", "Violation In Front Of Or Opposite", "House Number", "Street Name", "Intersecting Street", "Date First Observed", "Law Section", "Sub Division", "Violation Legal Code", "Days Parking In Effect ", "From Hours In Effect", "To Hours In Effect", "Vehicle Color", "Unregistered Vehicle?", "Vehicle Year", "Meter Number", "Feet From Curb", "Violation Post Code", "Violation Description", "No Standing or Stopping Violation", "Hydrant Violation", "Double Parking Violation"] if not processed_file else [i.lower().replace(" ", '_') for i in used_columns]
# Generates the sample CSV
if not processed_file:
sample_CSV_generate = False # Generate the sample CSV
sample_CSV_records = 100000 # No. of records to write into the sample CSV file
sample_CSV_path = f"{drive_path}/data/sample-{sample_CSV_records}.csv" if colab_env else f"../data/sample-{sample_CSV_records}.csv" # path to save
sample_seed = sample_CSV_records # Seed value so that we get same random records
# -
# ## Setup
# ### Spark Setup
# + tags=[]
# For google Colab
if colab_env:
# !apt-get install openjdk-8-jdk-headless -qq > /dev/null
# !wget -q https://mirrors.estointernet.in/apache/spark/spark-3.0.3/spark-3.0.3-bin-hadoop2.7.tgz -P /content/
# !tar xf /content/drive/Shareddrives/DA231-2021-Aug-Public/spark-3.0.3-bin-hadoop2.7.tgz
# !pip install -q findspark
import os
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["SPARK_HOME"] = "/content/spark-3.0.3-bin-hadoop2.7"
# -
# ### Imports
# Reload all changed modules every time before executing a new line
# %load_ext autoreload
# %autoreload 2
# +
# PySpark related
import findspark
findspark.init()
findspark.find()
from pyspark.sql import SparkSession
import pyspark.sql.functions as F
from pyspark.sql.functions import col # Frequently using this. hence imported separately
# user defined modules import
import data_preprocess as dp
import basic_analysis as ba
import precinct_based_analysis as pba
import time_based_analysis as tba
import season_based_analysis as sba
import revenue_based_analysis as rba
import mis_analysis as ma
# -
# ### Spark Session
spark = (
SparkSession
.builder
.master("local[*]") # Using all available cores
.appName("NYC Parking Violation Analysis")
.getOrCreate()
)
spark
# ## Data Pre-processing
# ### Reading CSV files into DataFrame
# +
# Better performance than inferSchema=True
NY_schema = dp.get_schema(schema_columns, schema_types)
org_df = spark.read.option("header", True).schema(NY_schema).csv(csv_files)
# -
if not processed_file:
print(f'Shape : {(org_df.count(), len(org_df.columns))}')
org_df.printSchema()
org_df.show(2)
if not processed_file:
presenting and org_df.summary().toPandas().transpose() # More execution time
# ### Pre-processing
# No pre-processing while using Sample file
df = org_df
if not processed_file:
# Need this setting to process the legacy time formats like 07/16/2019 12:00:00 AM # TODO: Pick it in challenges
spark.sql("set spark.sql.legacy.timeParserPolicy=LEGACY")
# Remove 30 unused columns
df = dp.remove_unused_columns(df, used_columns)
# Removes duplicate rows. Also drops the rows which are having null in all the columns
df = dp.drop_duplicates_nulls(df)
# Converting column names to lower case & replacing spaces with _
df = dp.santize_column_names(df)
# Ensure all the values in 'Summons Number' column are unique
dp.assert_uniqueness(df, column_name="summons_number")
# Converting issue date string type to Date type
df = dp.convert_to_date(df, column_name="issue_date", format="MM/dd/yyyy")
# Removing the rows which are outside of 2017-2021
df = dp.remove_outside_years_data(df, years, "issue_date")
# ## Sample file generation
# +
from pathlib import Path
from shutil import rmtree
def write_CSV(df, CSV_path):
# Creates CSV in a folder. But memory efficient
df.coalesce(1).write.mode("overwrite").csv(CSV_path, header=True)
# OOM
# df.toPandas().to_csv(CSV_path, index=False)
# Moving file to data folder
f_path = list(Path(CSV_path).glob('*.csv'))[0]
Path(f_path).rename(CSV_path+'.tmp')
rmtree(CSV_path)
Path(CSV_path+'.tmp').rename(CSV_path)
# -
if (not processed_file and sample_CSV_generate):
total_records = df.count()
fraction = (sample_CSV_records+10000)/total_records # Exact records are not coming. Hence increasing the fraction using 10k
sample_df = df.sample(fraction=fraction, seed=sample_seed).limit(sample_CSV_records) # Getting exact number of records
print(f"Sample records: {sample_df.count()}")
write_CSV(sample_df, sample_CSV_path)
df = sample_df
# ## Final DF / Table for Analysis
df.cache() # Cache the data
total_records = df.count() # Materialize the cache
print(f'Shape : {(total_records, len(df.columns))}')
df.printSchema()
df.show(2)
presenting and df.describe().toPandas().transpose() # More execution time
df.createOrReplaceTempView("NYCPV")
# ## Analysis
# ### 01 Basic Analysis
# #### **How often does each violation code occur? (frequency of violation codes find the top 10) ?**
#
# **Top violation codes**
#
# Speed violations are more than no parking violations
#
# | Violation Code | Description | Fine |
# |----------------|----------------------------------|------|
# | 36 | PHTO SCHOOL ZN SPEED VIOLATION | 50 |
# | 21 | NO PARKING-STREET CLEANING | 45 |
# | 38 | FAIL TO DSPLY MUNI METER RECPT | 35 |
# | 14 | NO STANDING-DAY/TIME LIMITS | 115 |
# | 20 | NO PARKING-DAY/TIME LIMITS | 60 |
# print('Top5 frequency of violation codes')
ba.violation_frequencey(df)
# TODO Verify the revenue & violation code count
# #### **How often does each vehicle body type get a parking ticket? (find the top 10)**
#
# Passenger vehicles have been getting more violations than commercial vehicles
print('Top5 Body type')
ba.violations_by_bodytype(df)
# #### **How often does each vehicle make get a parking ticket? (find the top 10)**
#
# TODO: Toyota, Ford & Honda vehicles are having more violations
# +
print('Top5 Vehicle Make')
ba.violations_by_make(df)
# -
# ### 02 Precinct based Analysis
#
# A **precinct** is a police station that has a certain zone of the city under its command
# #### **Find the (10 highest) frequencies of violating Precincts (this is the precinct of the zone where the violation occurred)**
# print('Top 5 Violation precicts')
pba.violating_precicts(df, True)
# #### **Find the (10 highest) frequencies of Issuing Precincts (this is the precinct that issued the ticket)**
# print('Top 5 Issuer precicts')
pba.issuing_precincts(df, True)
# #### **Find the violation code frequency across 3 precincts which have issued the most number of tickets (Top 5)**
# print('Violation frequencies from top 3 issuer precicts')
pba.violation_code_frequency_top3_precincts(df, True)
# ### 03 Time based Analysis
# #### **Divide 24 hours into 6 equal discrete bins of time. For each of these groups, find the 3 most commonly occurring violations**
# Divide 24 hours into six equal discrete bins of time.
# Bin Time Interval
# 1 12:00 AM to 4:00 AM
# 2 4:00 AM to 8:00 AM
# 3 8:00 AM to 12:00 PM
# 4 12:00 PM to 4:00 PM
# 5 4:00 PM to 8:00 PM
# 6 8:00 PM to 12:00 AM
tba.three_most_common_violations_in_6_time_bins (spark, True)
# #### **For the 5 most commonly occurring violation codes, find the most common times of day (in terms of the bins from the previous part)**
# Divide 24 hours into six equal discrete bins of time.
# Bin Time Interval
# 1 12:00 AM to 4:00 AM
# 2 4:00 AM to 8:00 AM
# 3 8:00 AM to 12:00 PM
# 4 12:00 PM to 4:00 PM
# 5 4:00 PM to 8:00 PM
# 6 8:00 PM to 12:00 AM
tba.five_most_common_Violations_with_times(spark, True)
# ### 04 Year / Season based Analysis
# #### **What is the average reduction in violations for the year 2020 compared to 2019 (due to COVID), and year 2019 compared to 2018**
# print("Reduction in violations due to covid")
sba.reduction_in_violations(df)
# #### **Divide the year into 3 number of seasons, and find frequencies of tickets for each season**
# print("Frequencies of violation in each season")
sba.season_violation_frequencies(df)
# #### **Find the 3 most common violations for each of these seasons**
# print("Frequencies of violation in each season")
sba.common_violations_season(df)
# ### 05 Revenue based Analysis
#
# The fines collected from all the parking violation constitute a revenue source for the NYC police department
# Fine amounts for each violation code are listed here https://www1.nyc.gov/site/finance/vehicles/services-violation-codes.page
# #### **Find the total amount collected year wise**
#
# TODO
# * Violation revenue is keep on increasing year on year
rba.yearly_revenue(df)
# #### **Find the top 5 violation codes which collected highest amount**
#
# TODO
# * Violation revenue is keep on increasing year on year
#
# | Violation Code | Description | Fine |
# |----------------|----------------------------------|------|
# | 36 | PHTO SCHOOL ZN SPEED VIOLATION | 50 |
# | 21 | NO PARKING-STREET CLEANING | 45 |
# | 38 | FAIL TO DSPLY MUNI METER RECPT | 35 |
# | 14 | NO STANDING-DAY/TIME LIMITS | 115 |
# | 20 | NO PARKING-DAY/TIME LIMITS | 60 |
rba.highest_revenue(df)
# ### 06 Miscellaneous Analysis
# #### **Find the top 10 Repeat offenders**
#
# * There is a invalid blank plate ID with more number of violations
# * TODO: **64638MD ** plate has the highest violations - 17
ma.repeat_offenders(spark)
# #### **Compare in state violations vs Out of state violations**
#
# * Violations are registered for **61**(TODO) out of state vehicles. But USA has 50 states. So we might have invalid entires in state column
# * **New Jersey** state registered vehicles got highest violations (TODO)
ma.in_out_state(spark)
# #### **Find number of violations on weekdays & weekends**
#
# TODO
# * Weekends violations are ~50% less compared to week days
ma.weekends(spark)
# #### **Find which days(holidays) has less number of violations**
#
# TODO
# * Christmas & New year holidays registered less number of violations
ma.holidays(spark)
# # Timing evaluation
def execute_all(time_eval_df):
ba.violation_frequencey(time_eval_df, False)
ba.violations_by_bodytype(time_eval_df, False)
ba.violations_by_make(time_eval_df, False)
pba.violating_precicts(time_eval_df, False)
pba.issuing_precincts(time_eval_df, False)
pba.violation_code_frequency_top3_precincts(time_eval_df, False)
tba.three_most_common_voilations_in_6_time_bins (spark, False)
tba.five_most_common_Voilations_with_times(spark, False)
sba.reduction_in_violations(time_eval_df, False)
sba.season_violation_frequencies(time_eval_df, False)
sba.common_violations_season(time_eval_df, False)
rba.yearly_revenue(time_eval_df, False)
rba.highest_revenue(time_eval_df, False)
ma.repeat_offenders(spark, False)
ma.in_out_state(spark, False)
ma.weekends(spark, False)
ma.holidays(spark, False)
# +
from timeit import timeit
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
NY_schema = dp.get_schema(schema_columns, schema_types)
entries = [15000, 100000, 1000000, 10000000]
time_data = []
for entry in entries:
data_file = '../data/time_eval_sample'+ f'{int(entry)}'+'.csv'
time_eval_df = spark.read.option("header", True).schema(NY_schema).csv(data_file)
time_eval_df.createOrReplaceTempView("NYCPV")
execution_time = timeit(lambda : execute_all(time_eval_df), number=1)
time_data.append(execution_time)
fig, ax = plt.subplots(figsize=(5,5))
ax.set_title("Data Entries Vs Time(s)")
ax.set_xlabel("Number of Data Entries")
ax.set_ylabel("Time of execution(seconds)")
x_data = output = [str(x) for x in entries]
ax.bar(x_data, time_data, color=['cyan', 'green', 'yellow', 'blue'])
fig.savefig('../output/entries_execution_time.png')
time_pd = pd.DataFrame(time_data, columns = ['Exectuin Time'], index = entries)
time_pd.index.name = 'Data processed'
time_pd
# +
# spark.stop()
| notebooks/nyc_parking_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Approximate Rates
#
# pynucastro can use rate approximations for $A(\alpha,\gamma)B$ and $A(\alpha,p)X(p,\gamma)B$,
# combining them into a single effective rate by assuming that the protons and nucleus $X$ are in equilibrium.
import pynucastro as pyna
import numpy as np
import matplotlib.pyplot as plt
from scipy.integrate import solve_ivp
# Let's create a simple network that has both an $(\alpha, \gamma)$ and
# $(\alpha, p)(p, \gamma)$ sequence.
reaclib_library = pyna.ReacLibLibrary()
mylib = reaclib_library.linking_nuclei(["mg24", "al27", "si28", "p31", "s32", "he4", "p"])
pynet = pyna.PythonNetwork(libraries=[mylib])
# + tags=[]
pynet.plot(rotated=True, curved_edges=True)
# -
pynet.write_network("full_net.py")
import full_net
# ## Integrating the full network
#
# Now let's integrate this. We'll start with half ${}^{24}\mathrm{Mg}$ and half $\alpha$ by mass.
# +
rho = 1.e7
T = 3e9
X0 = np.zeros(full_net.nnuc)
X0[full_net.jhe4] = 0.5
X0[full_net.jmg24] = 0.5
Y0 = X0 / full_net.A
# -
tmax = 1.e-3
sol = solve_ivp(full_net.rhs, [0, tmax], Y0, method="BDF",
dense_output=True, args=(rho, T), rtol=1.e-6, atol=1.e-10)
# +
fig = plt.figure()
ax = fig.add_subplot(111)
for i in range(full_net.nnuc):
ax.loglog(sol.t, sol.y[i,:] * full_net.A[i], label=f"X({full_net.names[i].capitalize()})")
ax.legend()
ax.set_xlim(1.e-10, 1.e-3)
ax.set_ylim(1.e-12, 1)
fig.set_size_inches((10, 8))
# -
# ## Approximate Version
#
# Now we will approximate the rates, combining $(\alpha, \gamma)$ and
# $(\alpha, p)(p, \gamma)$ into a single effective rate.
#
# The routine `make_ap_pg_approx()` will find all of the rates that make up that sequence and create a
# single `ApproximateRate` that captures the effective rate. The original rates will still be stored in the `ApproximateRate` object and will be evaluated to compute the needed approximation when the effective rate is evaluated.
pynet.make_ap_pg_approx()
pynet
# Since we no longer care about the ${}^{27}\mathrm{Al}$ and ${}^{31}\mathrm{P}$, we can remove them from the network. The `ApproximateRate` object still knows that these are the intermediate nucleus, but now they
# won't explicitly appear as one of the nuclei in the network.
pynet.remove_nuclei(["al27", "p31"])
# Note that since no reactions consume protons after that removal, the protons are all removed from the network, reducing its size from 7 nuclei to 4
print(pynet.network_overview())
# + tags=[]
pynet.plot(rotated=True, curved_edges=True)
# -
# As we see above, the nuclei ${}^{27}\mathrm{Al}$ and ${}^{31}\mathrm{P}$ no longer appear in the network, but the links to them are still understood to the network. This reduces the size of the network, while still preserving those flows.
pynet.write_network("approx_net.py")
import approx_net
# The `PythonNetwork` knows how to write out the code needed to evaluate the rate approximation. For instance, the evolution of ${}^{4}\mathrm{He}$ is determined as:
print(pynet.full_ydot_string(pyna.Nucleus("he4")))
# And the rate approximations are computed as:
r = pynet.get_rate("mg24_he4__si28__approx")
print(pynet.approx_function_string(r))
# where the 4 calls before the rate approximation is made are evaluating the original, unapproximated rates.
# ## Integrating the approximate network
#
# Let's integrate this approximate net and compare to above
# +
rho = 1.e7
T = 3.e9
X0 = np.zeros(approx_net.nnuc)
X0[approx_net.jhe4] = 0.5
X0[approx_net.jmg24] = 0.5
Y0 = X0 / approx_net.A
# -
tmax = 1.e-3
approx_sol = solve_ivp(approx_net.rhs, [0, tmax], Y0, method="BDF",
dense_output=True, args=(rho, T), rtol=1.e-6, atol=1.e-10)
# +
fig = plt.figure()
ax = fig.add_subplot(111)
for i in range(approx_net.nnuc):
ax.loglog(approx_sol.t, approx_sol.y[i,:] * approx_net.A[i], label=f"X({approx_net.names[i].capitalize()})")
ax.legend()
ax.set_xlim(1.e-10, 1.e-3)
ax.set_ylim(1.e-12, 1)
fig.set_size_inches((10, 8))
# -
# ## Comparison
#
# Let's plot both on the same axes to see the comparison.
# +
fig = plt.figure()
ax = fig.add_subplot(111)
for i in range(approx_net.nnuc):
ax.loglog(approx_sol.t, approx_sol.y[i,:] * approx_net.A[i],
linestyle=":", color=f"C{i}")
idx = full_net.names.index(approx_net.names[i])
ax.loglog(sol.t, sol.y[idx,:] * full_net.A[idx],
label=f"X({full_net.names[idx].capitalize()})",
linestyle="-", color=f"C{i}")
ax.legend()
ax.set_xlim(1.e-10, 1.e-3)
ax.set_ylim(1.e-12, 1)
fig.set_size_inches((10, 8))
# -
# Here the dotted line is the approximate network. We see that the results agree well.
# ## No approximation
#
# What if we just create a 4 nuclei network without the $(\alpha,p)(p,\gamma)$ links? How does this compare?
newlib = reaclib_library.linking_nuclei(["he4", "mg24", "si28", "s32"])
newpynet = pyna.PythonNetwork(libraries=[newlib])
newpynet.plot(rotated=True, curved_edges=True)
newpynet.write_network("simple_net.py")
import simple_net
# +
rho = 1.e7
T = 3e9
X0 = np.zeros(simple_net.nnuc)
X0[simple_net.jhe4] = 0.5
X0[simple_net.jmg24] = 0.5
Y0 = X0 / simple_net.A
# -
simple_net.names == approx_net.names
tmax = 1.e-3
simple_sol = solve_ivp(simple_net.rhs, [0, tmax], Y0, method="BDF",
dense_output=True, args=(rho, T), rtol=1.e-6, atol=1.e-10)
# +
fig = plt.figure()
ax = fig.add_subplot(111)
for i in range(approx_net.nnuc):
ax.loglog(approx_sol.t, approx_sol.y[i,:] * approx_net.A[i],
linestyle=":", color=f"C{i}")
idx = full_net.names.index(approx_net.names[i])
ax.loglog(sol.t, sol.y[idx,:] * full_net.A[idx],
label=f"X({full_net.names[idx].capitalize()})",
linestyle="-", color=f"C{i}")
idx = simple_net.names.index(approx_net.names[i])
ax.loglog(simple_sol.t, simple_sol.y[idx,:] * simple_net.A[idx],
linestyle="--", color=f"C{i}")
ax.legend()
ax.set_xlim(1.e-10, 1.e-3)
ax.set_ylim(1.e-12, 1)
fig.set_size_inches((10, 8))
# -
# Here we see all 3 networks. The full network (7 nuclei) is the solid lines. The approximate version of that is the dotted line. We see that they track reasonably well, especially when the abundance is high. The dashed line is the version of the network that has the same 4 nuclei as the approximate network, but with out approximating the $(\alpha, p)(p,\gamma)$ links, so we see that the ${}^{24}\mathrm{Mg}$ takes longer to burn.
| docs/source/approx-rates-examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## You can use firehose to ingest data near real time to your data lake, without writing any code on the consumer side.
# 
# ## 1) Create your stream using the console.
# Use prefix data/analytics/realtime/
# 
# 
# 
# 
# ## 2) Wait until your stream is created, and run the code below:
# ## 3) Create a firehose client
# +
import json
import boto3
session = boto3.Session(profile_name='default')
client = boto3.client('firehose')
# -
# ## 4) Send some records
with open("ratings.json") as json_file:
observations = json.load(json_file)
for observation in observations:
print(observation)
response = client.put_record(
DeliveryStreamName='mystream',
Record={
'Data': json.dumps(observation)
}
)
print(response)
# ## 5) Run your glue crawler again, and it will create a new table. Thenquery your new table using Athena.
| LAB12_Firehose.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="b2cfb934-16d2-462d-870c-379581678d79"
# ## Artificial Neural Network:
# + colab={"base_uri": "https://localhost:8080/"} id="tsWpmvCor_0b" outputId="d27a889e-ed3c-4f22-9ff2-1ca73eb73d12"
from google.colab import drive
drive.mount('/content/drive')
# + id="2588307f-587b-4dee-a0d1-d5f900fc27f7"
# importing packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats as stats
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
# Importing the Keras libraries and packages
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
import warnings
warnings.filterwarnings('ignore')
# + colab={"base_uri": "https://localhost:8080/", "height": 203} id="84316854-3369-4718-b301-12eb74a08052" outputId="333ad927-485e-4915-ddb0-e983b70f6acf"
# reading the dataset
df = pd.read_csv("/content/drive/MyDrive/Deep Learning/Churn_Modelling.csv")
df.head()
# + id="a7a27250-f458-4efc-86c1-1333f995672f" colab={"base_uri": "https://localhost:8080/"} outputId="7fc70a02-632d-4637-8d06-b7ab891ad4ff"
# check if there is any null value
df.isnull().sum().sum()
# + [markdown] id="a26683e6-62e9-4186-b035-0cce16b640e3"
# ### Data preprocessing:
# + id="d992d132-4d47-4f71-a4ba-90726cf94f85" colab={"base_uri": "https://localhost:8080/"} outputId="fde6127d-25d1-4e96-c9a1-b205391e5694"
# checking how many unique value in that columnn for replacing
df.Geography.unique()
# + id="491dd6b7-3a5c-471f-9c1e-d2c6cce8b703"
# replacing categorical value with numerical values
df['Geography'] = df['Geography'].replace(['France', 'Spain', 'Germany'], [0,1,2])
df['Gender'] = df['Gender'].replace(['Male', 'Female'], [0, 1])
# + id="c4d989d9-da7d-44fa-8be7-935051ee8f92"
# drop unwanted columns
df = df.drop(['RowNumber', 'CustomerId', 'Surname'], axis=1)
# + id="736c8c08-fcd5-4021-a5a2-ae404c6d2fa1" colab={"base_uri": "https://localhost:8080/"} outputId="86ca059f-c2b9-4b42-8db5-0dae229e1966"
df.Exited.value_counts()
# + id="05794edc-f7c2-4df4-91a3-588469518eb7" colab={"base_uri": "https://localhost:8080/", "height": 203} outputId="05ac49ab-ad64-4631-bf57-8d8b86ddfb5b"
df.head()
# + id="e22eaa8b-b6e7-4e34-86ec-dfc1c367833a"
X = df.drop(columns='Exited')
y = df['Exited']
# + id="7cdbe688-7fa0-4f2b-8216-51ef9a24ab75"
# split dataset into train and test dataset
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
# + [markdown] id="e89bdd39-404d-4542-a9a6-2137bbe2e7e4"
# ### Feature scaling:
# + id="1bdad7eb-5629-4757-bc5b-ce1ed1f799b2"
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.fit_transform(X_test)
# + [markdown] id="204d227a-b6d4-42bf-b06c-74dd35a33ba2"
# ### Initialising the ANN:
# + id="80a543c5-e82d-472f-9a55-d2a21399a0c5"
# initialising the model
model = Sequential()
# + id="a9e941df-4ca3-41fb-a437-be78751a331c"
# Adding the input layer (first hidden layer)
model.add(Dense(units=10, kernel_initializer='he_uniform', activation='relu', input_dim = 10))
# + id="59a8a5f5-f455-4848-82da-35f3349a7141"
# Adding the another input layer (second hidden layer)
model.add(Dense(units=20, kernel_initializer='he_uniform', activation='relu')) # for 2nd hidden layer dont use input_dim
# + id="824e9a3a-ecb4-4833-ba72-95c8941b1522"
# Adding the another input layer (third hidden layer)
model.add(Dense(units=10, kernel_initializer='he_uniform', activation='relu')) # for 3rd hidden layer dont use input_dim
# + id="68aa5ca5-3f86-42f1-b554-2ba3575ccf75"
# Adding the output layer
model.add(Dense(units = 1, kernel_initializer = 'glorot_uniform', activation = 'sigmoid'))
# + id="cf4c5df0-45c7-469e-a8f3-f54f0354d41e"
# compile the ANN
model.compile(optimizer='adam', loss='binary_crossentropy', metrics=['accuracy'])
# + id="70bc416c-53e3-4b19-9633-484ebfc1cf16" colab={"base_uri": "https://localhost:8080/"} outputId="9f9d5421-d356-4958-fc21-cf6fd7519b88"
# Fitting the ANN to the Training set
Model = model.fit(X_train, y_train, validation_split=0.1, epochs=10, batch_size=100, )
# + id="b36a3e9a-3ecb-4eda-a26e-98b9705ae425" colab={"base_uri": "https://localhost:8080/"} outputId="71c33e30-a2af-4b06-ebe1-fe46c01131bd"
# list all data in history
print(Model.history.keys())
# + id="eb7c600e-0f93-4e28-bd85-18330d69db64" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="156f7514-0b98-4f44-96bf-c0945d1ba156"
# summarize history for accuracy
plt.plot(Model.history['accuracy'])
plt.plot(Model.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower right')
plt.show()
# + id="4f941242-229a-41d5-b8c3-92d822381810" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="3c68154b-5c0b-4896-8f28-0edfa4b36de9"
# summarize history for loss
plt.plot(Model.history['loss'])
plt.plot(Model.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
# + id="dbb2dab3-8fe0-4ba7-9ef2-114c3c2c73ec"
# fitting the test dataset
y_pred = model.predict(X_test)
y_pred = y_pred > 0.5
# + id="497a1162-fb25-4638-8ef9-51799c4080ba" colab={"base_uri": "https://localhost:8080/"} outputId="3acf9c6f-6423-47e6-f1af-514a27b80a72"
# confusion matrix
print(f"Confusion matrix:\n {confusion_matrix(y_pred, y_test)}")
# classification report
print(f"Classification report:\n {classification_report(y_pred, y_test)}")
# + [markdown] id="96d88de8-59ff-47ae-9a0c-b2aa9ca68947"
# ### HyperParameter tunning:
# + id="9f33ac1c-f055-446d-b4be-42caf7bbe742"
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import GridSearchCV
from keras.models import Sequential
from keras.layers import Dense, Activation, Embedding, Flatten, LeakyReLU, BatchNormalization, Dropout
from keras.activations import relu, sigmoid
# + id="8bc1867a-0c0f-4645-98ca-4c7830ad68a4"
def create_model(layer, activation, kernel):
model = Sequential()
for i, nodes in enumerate(layer):
if i == 0:
model.add(Dense(nodes, input_dim=X_train.shape[1]))
model.add(Activation(activation))
else:
model.add(Dense(nodes), kernel_initializer=kernel)
model.add(Activation(activation))
model.add(Dense(units=1, activation='sigmoid', kernel_initializer="glorot_uniform")) # this one is output layer so we use sigmoid af
model.compile(optimizer='adam', loss="binary_crossentropy", metrics=['accuracy'])
return model
# + id="1ca60311-f192-495b-a2eb-2f7ee519ca8e"
# creating an object for previous function
model = KerasClassifier(build_fn=create_model)
# + id="e157d359-283c-412b-8ac0-611ec4bf566a"
layers = [[20], [40, 20], [40, 20, 10]]
activations = ['relu', 'sigmoid']
kernel = [['he_uniform', 'he_normal'], ['glorot_uniform', 'glorot_normal']]
para_grid = dict(layer=layers, activation=activations, kernel=kernel)
grid = GridSearchCV(model, para_grid, cv=5)
# + id="decfe152-2275-45c2-ac72-b0aa3c379500" colab={"base_uri": "https://localhost:8080/"} outputId="c303b790-8c2d-4f2b-d821-ce4969dcff2f"
# fitting dataset with created model
grid.fit(X_train, y_train, validation_split=0.3, validation_data=(X_test, y_test), epochs=10)
# + id="d8f49757-4c7a-4b35-bcba-a827c352eb02" colab={"base_uri": "https://localhost:8080/"} outputId="26737218-b366-422e-f996-12ecb430856b"
print(grid.best_params_, grid.best_score_)
# + [markdown] id="be4ec82b-2a34-48aa-9301-918db617ab1a"
# ### HyperParameter Tunnig using KerasTunner:
# + id="ed49839f-3e3a-4a51-8695-dc751dc2bf72"
# # !pip install keras_tuner
from tensorflow import keras
from keras import Sequential
from keras.layers import Dense, Activation
from keras_tuner import RandomSearch
# + id="5cc6d254-f1d4-4166-bf29-7084520fe0c4"
def create_model(hp):
model = Sequential()
for i in range(hp.Int("num_layer", 2, 20)):
model.add(Dense(units= hp.Int("units_" + str(i), min_value=32, max_value=512, step=32),
activation='ELU', kernel_initializer='he_uniform',
input_dim=X_train.shape[1]))
model.add(Dense(units=1, activation="sigmoid"))
model.compile(optimizer=keras.optimizers.Adam(hp.Choice('learning_rate', [1e-2, 1e-3, 1e-4])),
loss='binary_crossentropy', metrics=['accuracy'])
return model
# + id="5e02656a-4aea-447d-bc43-34b330327e83" colab={"base_uri": "https://localhost:8080/"} outputId="56e7dae0-99f8-4e99-9aeb-197296f55e66"
tuner = RandomSearch(hypermodel=create_model,
objective="val_accuracy",
max_trials=5,
executions_per_trial=3,
directory='Log',
project_name='HyperTuning')
# + id="dca58625-3dff-4869-a05f-2db0698bf521" colab={"base_uri": "https://localhost:8080/"} outputId="e944e254-2eb2-404a-d2ff-d4f11a074a02"
tuner.search(X_train, y_train, epochs=10, validation_split=0.3)
# + id="b0fd60e6-016c-43b7-b8e0-f5af9f50a008"
model = tuner.get_best_models(num_models=1)[0]
# + id="798b35ab-d485-4739-b624-c22bdaa1f63d" colab={"base_uri": "https://localhost:8080/"} outputId="628800c8-67d9-4a05-d9c2-59c6979d51be"
model.summary()
# + id="d23a76cd-32c0-4984-a98f-94e97dea29df" colab={"base_uri": "https://localhost:8080/"} outputId="0c064727-26f6-4aa5-cfe4-588e4ec01d2e"
Model = model.fit(X_train, y_train, epochs=10, validation_split=0.3, validation_data=(X_test, y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="QXYRrf6EchDS" outputId="2c9f5e62-df55-42d0-e26a-b480a6eb46df"
print(Model.history.keys())
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="jfVnjepjcfna" outputId="43cd36eb-1182-43fd-bbba-ab2ee38bbd60"
# summarize history for accuracy
plt.plot(Model.history['accuracy'])
plt.plot(Model.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower right')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="tCRVVg3Ccla7" outputId="3a2218e4-68f0-441f-9d75-25f5d98b94b8"
# summarize history for loss
plt.plot(Model.history['loss'])
plt.plot(Model.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
# + id="6OJHWs3JclGr"
# fitting the test dataset
y_pred = model.predict(X_test)
y_pred = y_pred > 0.5
# + colab={"base_uri": "https://localhost:8080/"} id="ChWWaOgadp8u" outputId="cbf204a3-02da-4b47-e8e2-e8a61a11b24d"
# confusion matrix
print(f"Confusion matrix:\n {confusion_matrix(y_pred, y_test)}")
# classification report
print(f"Classification report:\n {classification_report(y_pred, y_test)}")
# + [markdown] id="370faaa1-c25d-4aad-9e8b-b06363133534"
# ### Sample Code from the documentation
# + id="7cbec061-9c52-483b-98b8-624c37e1d439"
from tensorflow import keras
import numpy as np
(x, y), (x_test, y_test) = keras.datasets.mnist.load_data()
x_train = x[:-10000]
x_val = x[-10000:]
y_train = y[:-10000]
y_val = y[-10000:]
x_train = np.expand_dims(x_train, -1).astype("float32") / 255.0
x_val = np.expand_dims(x_val, -1).astype("float32") / 255.0
x_test = np.expand_dims(x_test, -1).astype("float32") / 255.0
num_classes = 10
y_train = keras.utils.to_categorical(y_train, num_classes)
y_val = keras.utils.to_categorical(y_val, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
# + id="ad879e5a-3a28-41b0-b3f1-66051504b0a7"
from tensorflow.keras import layers
from keras_tuner import RandomSearch
def build_model(hp):
model = keras.Sequential()
model.add(layers.Flatten())
model.add(
layers.Dense(
units=hp.Int("units", min_value=32, max_value=512, step=32),
activation="relu",
)
)
model.add(layers.Dense(10, activation="softmax"))
model.compile(
optimizer=keras.optimizers.Adam(
hp.Choice("learning_rate", values=[1e-2, 1e-3, 1e-4])
),
loss="categorical_crossentropy",
metrics=["accuracy"],
)
return model
# + id="efb8c519-5e1d-435e-a720-83c94797b042"
tuner = RandomSearch(
build_model,
objective="val_accuracy",
max_trials=3,
executions_per_trial=2,
overwrite=True,
directory="my_dir",
project_name="helloworld",
)
# + id="f21159a3-5939-44d3-abb0-b1710eb0e892" colab={"base_uri": "https://localhost:8080/"} outputId="bbc40c4d-a041-4914-ecaf-dc30dbb127e8"
tuner.search_space_summary()
# + id="902ff586-fbe5-4c7f-8c6e-9acfdb13f4b9" colab={"base_uri": "https://localhost:8080/"} outputId="0ffc113e-65ae-4678-a76d-81c5b87413a1"
tuner.search(x_train, y_train, epochs=2, validation_data=(x_val, y_val))
# + id="4521b27e-e1e9-416a-b757-79013e1b4a5f"
model = tuner.get_best_models(num_models=1)[0]
# + colab={"base_uri": "https://localhost:8080/"} id="Iq2ue4Gnd_Sw" outputId="38b6366e-e62f-45d0-9de8-c9870e799551"
Model = model.fit(x_train, y_train, epochs=10, validation_data=(x_val, y_val))
# + colab={"base_uri": "https://localhost:8080/"} id="9yTRstEaeS0e" outputId="17de9f60-88c6-4490-925f-a34ddf3bcc0d"
Model.history.keys()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="MjpKx1yJeWQG" outputId="56f470e5-1ae3-463d-e73b-4d8fa46fd5eb"
# summarize history for accuracy
plt.plot(Model.history['accuracy'])
plt.plot(Model.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='lower right')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="CZY6sSGkhVRi" outputId="539fe29e-221d-4bea-fd8b-3668c9ee5d11"
# summarize history for loss
plt.plot(Model.history['loss'])
plt.plot(Model.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper right')
plt.show()
# + id="RVd7Fj2_l87L"
| ANN/ANN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# + [markdown] id="NdrXRgjU7Zih"
# # Getting Started
#
# In this tutorial, we demonstrate some of the basic features of TorchGeo and show how easy it is to use if you're already familiar with other PyTorch domain libraries like torchvision.
#
# It's recommended to run this notebook on Google Colab if you don't have your own GPU. Click the "Open in Colab" button above to get started.
# + [markdown] id="lCqHTGRYBZcz"
# ## Setup
#
# First, we install TorchGeo.
# -
# %pip install torchgeo
# + [markdown] id="dV0NLHfGBMWl"
# ## Imports
#
# Next, we import TorchGeo and any other libraries we need.
# + id="entire-albania"
import os
import tempfile
from torch.utils.data import DataLoader
from torchgeo.datasets import NAIP, ChesapeakeDE, stack_samples
from torchgeo.datasets.utils import download_url
from torchgeo.samplers import RandomGeoSampler
# + [markdown] id="5rLknZxrBEMz"
# ## Datasets
#
# For this tutorial, we'll be using imagery from the [National Agriculture Imagery Program (NAIP)](https://www.fsa.usda.gov/programs-and-services/aerial-photography/imagery-programs/naip-imagery/) and labels from the [Chesapeake Bay High-Resolution Land Cover Project](https://www.chesapeakeconservancy.org/conservation-innovation-center/high-resolution-data/land-cover-data-project/). First, we manually download a few NAIP tiles and create a PyTorch Dataset.
# + colab={"base_uri": "https://localhost:8080/", "height": 232, "referenced_widgets": ["d00a2177bf4b4b8191bfc8796f0e749f", "17d6b81aec50455989276b595457cc7f", "<KEY>", "<KEY>", "<KEY>", "f7ef78d6f87a4a2685788e395525fa7c", "5b2450e316e64b4ba432c78b63275124", "<KEY>", "<KEY>", "<KEY>", "8dd61c8479d74c95a55de147e04446b3", "<KEY>", "<KEY>", "e680eda3c84c440083e2959f04431bea", "a073e33fd9ae4125822fc17971233770", "<KEY>", "b3d4c9c99bec4e69a199e45920d52ce4", "a215f3310ea543d1a8991f57ec824872", "569f60397fd6440d825e8afb83b4e1ae", "<KEY>", "737fa148dfae49a18cc0eabbe05f2d0f", "0b6613adbcc74165a9d9f74988af366e", "<KEY>", "ef0fc75ff5044171be942a6b3ba0c2da", "<KEY>", "9a689285370646ab800155432ea042a5", "<KEY>", "<KEY>", "8a8538a91a74439b81e3f7c6516763e3", "<KEY>", "99cd2e65fb104380953745f2e0a93fac", "<KEY>", "54f5db9555c44efa9370cbb7ab58e142", "<KEY>", "<KEY>", "bd2e44a8eb1a4c19a32da5a1edd647d1", "0f9feea4b8344a7f8054c9417150825e", "31acb7a1ca8940078e1aacd72e547f47", "0d0ca8d64d3e4c2f88d87342808dd677", "<KEY>", "910b98584fa74bb5ad308fe770f5b40e", "b2dce834ee044d69858389178b493a2b", "237f2e31bcfe476baafae8d922877e07", "43ac7d95481b4ea3866feef6ace2f043"]} id="e3138ac3" outputId="11589c46-eee6-455d-839b-390f2934d834"
data_root = tempfile.gettempdir()
naip_root = os.path.join(data_root, "naip")
naip_url = "https://naipblobs.blob.core.windows.net/naip/v002/de/2018/de_060cm_2018/38075/"
tiles = [
"m_3807511_ne_18_060_20181104.tif",
"m_3807511_se_18_060_20181104.tif",
"m_3807512_nw_18_060_20180815.tif",
"m_3807512_sw_18_060_20180815.tif",
]
for tile in tiles:
download_url(naip_url + tile, naip_root)
naip = NAIP(naip_root)
# + [markdown] id="HQVji2B22Qfu"
# Next, we tell TorchGeo to automatically download the corresponding Chesapeake labels.
# + colab={"base_uri": "https://localhost:8080/"} id="2Ah34KAw2biY" outputId="03b7bdf0-78c1-4a13-ac56-59de740d7f59"
chesapeake_root = os.path.join(data_root, "chesapeake")
chesapeake = ChesapeakeDE(chesapeake_root, crs=naip.crs, res=naip.res, download=True)
# + [markdown] id="OWUhlfpD22IX"
# Finally, we create an IntersectionDataset so that we can automatically sample from both GeoDatasets simultaneously.
# + id="WXxy8F8l2-aC"
dataset = naip & chesapeake
# + [markdown] id="yF_R54Yf3EUd"
# ## Sampler
#
# Unlike typical PyTorch Datasets, TorchGeo GeoDatasets are indexed using lat/long/time bounding boxes. This requires us to use a custom GeoSampler instead of the default sampler/batch_sampler that comes with PyTorch.
# + id="RLczuU293itT"
sampler = RandomGeoSampler(naip, size=1000, length=10)
# + [markdown] id="OWa-mmYd8S6K"
# ## DataLoader
#
# Now that we have a Dataset and Sampler, we can combine these into a single DataLoader.
# + id="jfx-9ZmU8ZTc"
dataloader = DataLoader(dataset, sampler=sampler, collate_fn=stack_samples)
# + [markdown] id="HZIfqqW58oZe"
# ## Training
#
# Other than that, the rest of the training pipeline is the same as it is for torchvision.
# + id="7sGmNvBy8uIg"
for sample in dataloader:
image = sample["image"]
target = sample["mask"]
| docs/tutorials/getting_started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Simple example for performing symbolic regression for a set of points
from nesymres.architectures.model import Model
from nesymres.utils import load_metadata_hdf5
from nesymres.dclasses import FitParams, NNEquation, BFGSParams
from pathlib import Path
from functools import partial
import torch
from sympy import lambdify
import json
# +
## Load equation configuration and architecture configuration
import omegaconf
with open('100M/eq_setting.json', 'r') as json_file:
eq_setting = json.load(json_file)
cfg = omegaconf.OmegaConf.load("100M/config.yaml")
# -
## Set up BFGS load rom the hydra config yaml
bfgs = BFGSParams(
activated= cfg.inference.bfgs.activated,
n_restarts=cfg.inference.bfgs.n_restarts,
add_coefficients_if_not_existing=cfg.inference.bfgs.add_coefficients_if_not_existing,
normalization_o=cfg.inference.bfgs.normalization_o,
idx_remove=cfg.inference.bfgs.idx_remove,
normalization_type=cfg.inference.bfgs.normalization_type,
stop_time=cfg.inference.bfgs.stop_time,
)
params_fit = FitParams(word2id=eq_setting["word2id"],
id2word={int(k): v for k,v in eq_setting["id2word"].items()},
una_ops=eq_setting["una_ops"],
bin_ops=eq_setting["bin_ops"],
total_variables=list(eq_setting["total_variables"]),
total_coefficients=list(eq_setting["total_coefficients"]),
rewrite_functions=list(eq_setting["rewrite_functions"]),
bfgs=bfgs,
beam_size=cfg.inference.beam_size #This parameter is a tradeoff between accuracy and fitting time
)
weights_path = "../weights/100M.ckpt"
## Load architecture, set into eval mode, and pass the config parameters
model = Model.load_from_checkpoint(weights_path, cfg=cfg.architecture)
model.eval()
if torch.cuda.is_available():
model.cuda()
fitfunc = partial(model.fitfunc,cfg_params=params_fit)
# +
# Create points from an equation
number_of_points = 500
n_variables = 1
#To get best results make sure that your support inside the max and mix support
max_supp = cfg.dataset_train.fun_support["max"]
min_supp = cfg.dataset_train.fun_support["min"]
X = torch.rand(number_of_points,len(list(eq_setting["total_variables"])))*(max_supp-min_supp)+min_supp
X[:,n_variables:] = 0
target_eq = "x_1*sin(x_1)" #Use x_1,x_2 and x_3 as independent variables
X_dict = {x:X[:,idx].cpu() for idx, x in enumerate(eq_setting["total_variables"])}
y = lambdify(",".join(eq_setting["total_variables"]), target_eq)(**X_dict)
# -
print("X shape: ", X.shape)
print("y shape: ", y.shape)
output = fitfunc(X,y)
output
| jupyter/fit_func.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="eTdCMVl9YAXw" colab_type="text"
# # Embeddings
#
# In this lesson we will learn how to map tokens to vectors (embeddings) that capture the contextual, semantic and syntactic value of a token in text.
# + [markdown] id="xuabAj4PYj57" colab_type="text"
# <div align="left">
# <a href="https://github.com/madewithml/basics/blob/master/notebooks/14_Embeddings/14_PT_Embeddings.ipynb" role="button"><img class="notebook-badge-image" src="https://img.shields.io/static/v1?label=&message=View%20On%20GitHub&color=586069&logo=github&labelColor=2f363d"></a>
# <a href="https://colab.research.google.com/github/madewithml/basics/blob/master/notebooks/14_Embeddings/14_PT_Embeddings.ipynb"><img class="notebook-badge-image" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"></a>
# </div>
# + [markdown] id="paO18xGs7kGX" colab_type="text"
# # Overview
# + [markdown] id="JqxyljU18hvt" colab_type="text"
# So far, we've also represented our text data in a one-hot encoded form where each token is represented by an n-dimensional array.
#
# ```python
# [[0. 0. 0. ... 0. 0. 0.]
# [0. 0. 1. ... 0. 0. 0.]
# [0. 0. 0. ... 0. 0. 0.]
# ...
# [0. 0. 0. ... 0. 0. 0.]
# [0. 0. 0. ... 0. 0. 0.]
# [0. 0. 0. ... 0. 0. 0.]]
# ```
#
# This allows us to preserve the structural information but there are two major disadvantages here. We used character level representations in the CNN lessons because the number of characters is small. Suppose we wanted to one-hot encode each word instead. Now the vocabulary sizes quickly grows leading to large computes. And though we preserve the structure within the text, the actual representation for each token does not preserve any relationship with respect to other tokens.
#
# In this notebook, we're going to learn about embeddings and how they address all the shortcomings of the representation methods we've seen so far.
#
#
#
#
# + [markdown] id="yN73ZCCnjezh" colab_type="text"
# * **Objective:** Represent tokens in text that capture the intrinsic semantic relationships.
# * **Advantages:**
# * Low-dimensionality while capturing relationships.
# * Interpretable token representations
# * **Disadvantages:** None
# * **Miscellaneous:** There are lot's of pretrained embeddings to choose from but you can also train your own from scratch.
# + [markdown] id="MrDStrYbjsnW" colab_type="text"
# # Data
# + [markdown] id="nH_O4MZ294jk" colab_type="text"
# ## Load data
# + [markdown] id="F47IiPgUupAk" colab_type="text"
# We can learn embeddings by creating our models in TensorFLow but instead, we're going to use a library that specializes in embeddings and topic modeling called [Gensim](https://radimrehurek.com/gensim/).
# + id="NUuFGxRI8xxl" colab_type="code" colab={}
import urllib
# + id="lVfE26vR9O-y" colab_type="code" colab={}
DATA_FILE = 'harrypotter.txt'
# + id="LegtLIr-lxxZ" colab_type="code" colab={}
# Load data from GitHub to this notebook's local drive
url = "https://raw.githubusercontent.com/madewithml/basics/master/data/harrypotter.txt"
response = urllib.request.urlopen(url)
html = response.read()
with open(DATA_FILE, 'wb') as fp:
fp.write(html)
# + [markdown] id="4ZDGEBa2-Ccf" colab_type="text"
# ## Preprocess
# + id="62qsAAZ5gG9M" colab_type="code" outputId="19d9a314-6b13-4ada-e42c-98837d5e9a61" executionInfo={"status": "ok", "timestamp": 1584550699062, "user_tz": 420, "elapsed": 5299, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Use TensorFlow 2.x
# %tensorflow_version 2.x
# + id="_pZljlaCgG6Y" colab_type="code" outputId="4d2314c0-0b20-4015-9a03-9ec242ec9bd8" executionInfo={"status": "ok", "timestamp": 1584550701360, "user_tz": 420, "elapsed": 7579, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
import nltk; nltk.download('punkt')
import tensorflow as tf
from tensorflow.keras.preprocessing.text import text_to_word_sequence
print("GPU Available: ", tf.config.list_physical_devices('GPU'))
# + id="oektJd55gG1p" colab_type="code" colab={}
SEED = 1234
# + id="tqbnugiD-SW0" colab_type="code" colab={}
# Set seed for reproducibility
tf.random.set_seed(SEED)
# + id="fdtG_em9YaLc" colab_type="code" colab={}
FILTERS = "!\"'#$%&()*+,-./:;<=>?@[\\]^_`{|}~"
LOWER = True
# + id="vF5D_nNjlx2d" colab_type="code" outputId="4429d5bf-4710-4d6f-c336-7be26feead7f" executionInfo={"status": "ok", "timestamp": 1584550701362, "user_tz": 420, "elapsed": 7536, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Split text into sentences
tokenizer = nltk.data.load('tokenizers/punkt/english.pickle')
with open(DATA_FILE, encoding='cp1252') as fp:
book = fp.read()
sentences = tokenizer.tokenize(book)
print (f"{len(sentences)} sentences")
# + id="NsZz5jfMlx0d" colab_type="code" outputId="e48716b7-fe2c-455d-ffa4-229b5d55e947" executionInfo={"status": "ok", "timestamp": 1584550701592, "user_tz": 420, "elapsed": 7736, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# Preprocess sentences
print (sentences[11])
sentences = [text_to_word_sequence(
text=sentence, filters=FILTERS,
lower=LOWER, split=' ') for sentence in sentences]
print (sentences[11])
# + [markdown] id="yjIUhBwxkBHc" colab_type="text"
# # Learning embeddings
# + [markdown] id="rozFTf06ji1b" colab_type="text"
# The main idea of embeddings is to have fixed length representations for the tokens in a text regardless of the number of tokens in the vocabulary. So instead of each token representation having the shape [1 X V] where V is vocab size, each token now has the shape [1 X D] where D is the embedding size (usually 50, 100, 200, 300). The numbers in the representation will no longer be 0s and 1s but rather floats that represent that token in a D-dimensional latent space. If the embeddings really did capture the relationship between tokens, then we should be able to inspect this latent space and confirm known relationships (we'll do this soon).
#
# But how do we learn the embeddings the first place? The intuition behind embeddings is that the definition of a token depends on the token itself but on its context. There are several different ways of doing this:
#
# 1. Given the word in the context, predict the target word (CBOW - continuous bag of words).
# 2. Given the target word, predict the context word (skip-gram).
# 3. Given a sequence of words, predict the next word (LM - language modeling).
#
# All of these approaches involve create data to train our model on. Every word in a sentence becomes the target word and the context words are determines by a window. In the image below (skip-gram), the window size is 2 (2 words to the left and right of the target word). We repeat this for every sentence in our corpus and this results in our training data for the unsupervised task. This in an unsupervised learning technique since we don't have official labels for contexts. The idea is that similar target words will appear with similar contexts and we can learn this relationship by repeatedly training our mode with (context, target) pairs.
#
# <div align="left">
# <img src="https://raw.githubusercontent.com/madewithml/images/master/basics/14_Embeddings/skipgram.png" width="600">
# </div>
#
# We can learn embeddings using any of these approaches above and some work better than others. You can inspect the learned embeddings but the best way to choose an approach is to empirically validate the performance on a supervised task.
# + [markdown] id="No6c943C-P7o" colab_type="text"
# ## Word2Vec
# + [markdown] id="VeszvcMOji4u" colab_type="text"
# When we have large vocabularies to learn embeddings for, things can get complex very quickly. Recall that the backpropagation with softmax updates both the correct and incorrect class weights. This becomes a massive computation for every backwas pass we do so a workaround is to use [negative sampling](http://mccormickml.com/2017/01/11/word2vec-tutorial-part-2-negative-sampling/) which only updates the correct class and a few arbitrary incorrect classes (negative_sampling=20). We're able to do this because of the large amount of training data where we'll see the same word as the target class multiple times.
#
#
# + id="TqKCr--k-f9e" colab_type="code" colab={}
import gensim
from gensim.models import KeyedVectors
from gensim.models import Word2Vec
# + id="ufU-9l_W-QKj" colab_type="code" colab={}
EMBEDDING_DIM = 100
WINDOW = 5
MIN_COUNT = 3 # Ignores all words with total frequency lower than this
SKIP_GRAM = 1 # 0 = CBOW
NEGATIVE_SAMPLING = 20
# + id="Ha3I2oSsmhJa" colab_type="code" outputId="fdb857f0-b642-487f-abc9-653630f9e25a" executionInfo={"status": "ok", "timestamp": 1584550713317, "user_tz": 420, "elapsed": 19432, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Super fast because of optimized C code under the hood
w2v = Word2Vec(sentences=sentences, size=EMBEDDING_DIM,
window=WINDOW, min_count=MIN_COUNT,
sg=SKIP_GRAM, negative=NEGATIVE_SAMPLING)
print (w2v)
# + id="Cl6oJv8jmhHE" colab_type="code" outputId="04b46ec1-bb3e-4dc3-ed64-e1d30b94ea93" executionInfo={"status": "ok", "timestamp": 1584550713318, "user_tz": 420, "elapsed": 19413, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 374}
# Vector for each word
w2v.wv.get_vector("potter")
# + id="DyuLX9DTnLvM" colab_type="code" outputId="3a5e590c-f65c-49bd-966e-283bd124253f" executionInfo={"status": "ok", "timestamp": 1584550713318, "user_tz": 420, "elapsed": 19394, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 156}
# Get nearest neighbors (excluding itself)
w2v.wv.most_similar(positive="scar", topn=5)
# + id="YT7B0KRVTFew" colab_type="code" outputId="b64faca7-c609-492e-bb08-a7d0dddbd46a" executionInfo={"status": "ok", "timestamp": 1584550713319, "user_tz": 420, "elapsed": 19371, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 71}
# Saving and loading
w2v.wv.save_word2vec_format('model.bin', binary=True)
w2v = KeyedVectors.load_word2vec_format('model.bin', binary=True)
# + [markdown] id="JZXVP5vfuiD5" colab_type="text"
# ## FastText
# + [markdown] id="uvuoeWYMuqsa" colab_type="text"
# What happen's when a word doesn't exist in our vocabulary? We could assign an UNK token which is used for all OOV (out of vocabulary) words or we could use [FastText](https://radimrehurek.com/gensim/models/fasttext.html), which uses character-level n-grams to embed a word. This helps embed rare words, misspelled words, and also words that don't exist in our corpus but are similar to words in our corpus.
# + id="fVg3PBeD-kAa" colab_type="code" colab={}
from gensim.models import FastText
# + id="eTNW4Mfgrpo0" colab_type="code" outputId="7b6f9472-6ad5-4fd4-d151-80870c18f381" executionInfo={"status": "ok", "timestamp": 1584550726329, "user_tz": 420, "elapsed": 32338, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Super fast because of optimized C code under the hood
ft = FastText(sentences=sentences, size=EMBEDDING_DIM,
window=WINDOW, min_count=MIN_COUNT,
sg=SKIP_GRAM, negative=NEGATIVE_SAMPLING)
print (ft)
# + id="LbA4vU5uxiw3" colab_type="code" colab={}
# This word doesn't exist so the word2vec model will error out
# w2v.wv.most_similar(positive="scarring", topn=5)
# + id="eRG30aE4sMjt" colab_type="code" outputId="fcb1ebf7-8d36-4ccd-fbb2-d28eac7969d4" executionInfo={"status": "ok", "timestamp": 1584550726330, "user_tz": 420, "elapsed": 32317, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 156}
# FastText will use n-grams to embed an OOV word
ft.wv.most_similar(positive="scarring", topn=5)
# + id="7SE5fPMUnLyP" colab_type="code" outputId="3ad0f818-a265-487a-bf2a-15606734da66" executionInfo={"status": "ok", "timestamp": 1584550726331, "user_tz": 420, "elapsed": 32287, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 71}
# Save and loading
ft.wv.save('model.bin')
ft = KeyedVectors.load('model.bin')
# + [markdown] id="67UmjtK0pF9X" colab_type="text"
# # Pretrained embeddings
# + [markdown] id="Xm1GPn4spF6x" colab_type="text"
# We can learn embeddings from scratch using one of the approaches above but we can also leverage pretrained embeddings that have been trained on millions of documents. Popular ones include Word2Vec (skip-gram) or GloVe (global word-word co-occurrence). We can validate that these embeddings captured meaningful semantic relationships by confirming them.
# + id="Hh42Mb4lLbuB" colab_type="code" colab={}
from gensim.scripts.glove2word2vec import glove2word2vec
from io import BytesIO
import matplotlib.pyplot as plt
import numpy as np
from sklearn.decomposition import PCA
from urllib.request import urlopen
from zipfile import ZipFile
# + id="pZIn8oAaBiZv" colab_type="code" colab={}
# Set seed for reproducibility
np.random.seed(SEED)
# + id="m9gxHJA9M8hK" colab_type="code" colab={}
# Arguments
EMBEDDING_DIM = 100
# + id="ANfQHxGrMKTe" colab_type="code" colab={}
def plot_embeddings(words, embeddings, pca_results):
for word in words:
index = embeddings.index2word.index(word)
plt.scatter(pca_results[index, 0], pca_results[index, 1])
plt.annotate(word, xy=(pca_results[index, 0], pca_results[index, 1]))
plt.show()
# + id="ZW9Qtkz3LfdY" colab_type="code" outputId="3d127209-370f-4202-c3d7-62cc160aad31" executionInfo={"status": "ok", "timestamp": 1584551115734, "user_tz": 420, "elapsed": 421660, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
# Unzip the file (may take ~3-5 minutes)
resp = urlopen('http://nlp.stanford.edu/data/glove.6B.zip')
zipfile = ZipFile(BytesIO(resp.read()))
zipfile.namelist()
# + id="bWnVBrOaLjIC" colab_type="code" outputId="77d82016-3b1c-43e3-e24e-7cee672ac5d3" executionInfo={"status": "ok", "timestamp": 1584551118950, "user_tz": 420, "elapsed": 424855, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Write embeddings to file
embeddings_file = 'glove.6B.{0}d.txt'.format(EMBEDDING_DIM)
zipfile.extract(embeddings_file)
# + id="qFLyIqIxrUIs" colab_type="code" outputId="3cc33b26-57e4-4cb2-e86b-26d37160b3f1" executionInfo={"status": "ok", "timestamp": 1584551118950, "user_tz": 420, "elapsed": 424835, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 323}
# Preview of the GloVe embeddings file
with open(embeddings_file, 'r') as fp:
line = next(fp)
values = line.split()
word = values[0]
embedding = np.asarray(values[1:], dtype='float32')
print (f"word: {word}")
print (f"embedding:\n{embedding}")
print (f"embedding dim: {len(embedding)}")
# + id="9eD5doqFLjFY" colab_type="code" outputId="06a8d0ec-a862-4495-8f3c-140e57d9d33b" executionInfo={"status": "ok", "timestamp": 1584551120204, "user_tz": 420, "elapsed": 426068, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 88}
# Save GloVe embeddings to local directory in word2vec format
word2vec_output_file = '{0}.word2vec'.format(embeddings_file)
glove2word2vec(embeddings_file, word2vec_output_file)
# + id="To4sx_1iMCX0" colab_type="code" outputId="0a9afc39-967d-4a9f-9f36-e6c2a1861ba4" executionInfo={"status": "ok", "timestamp": 1584551155794, "user_tz": 420, "elapsed": 461634, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 71}
# Load embeddings (may take a minute)
glove = KeyedVectors.load_word2vec_format(word2vec_output_file, binary=False)
# + id="UEhBhvgHMEH9" colab_type="code" outputId="55aff262-b918-4f7f-f2c5-c28c6694fe3e" executionInfo={"status": "ok", "timestamp": 1584551155796, "user_tz": 420, "elapsed": 461611, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 156}
# (king - man) + woman = ?
glove.most_similar(positive=['woman', 'king'], negative=['man'], topn=5)
# + id="xR94AICkMEFV" colab_type="code" outputId="558efa58-3085-4cf5-b041-3fe62614cc59" executionInfo={"status": "ok", "timestamp": 1584551155796, "user_tz": 420, "elapsed": 461584, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 190}
# Get nearest neighbors (exlcusing itself)
glove.wv.most_similar(positive="goku", topn=5)
# + id="gseqjBmzMECq" colab_type="code" outputId="3ea3efef-c735-4b50-fd37-5c1d70554576" executionInfo={"status": "ok", "timestamp": 1584551158749, "user_tz": 420, "elapsed": 464520, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# Reduce dimensionality for plotting
X = glove[glove.wv.vocab]
pca = PCA(n_components=2)
pca_results = pca.fit_transform(X)
# + id="LFQWGyncMHgK" colab_type="code" outputId="94aeb639-72db-47a8-a08a-7d29a3ec53ca" executionInfo={"status": "ok", "timestamp": 1584551158751, "user_tz": 420, "elapsed": 464505, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 265}
# Visualize
plot_embeddings(words=["king", "queen", "man", "woman"],
embeddings=glove,
pca_results=pca_results)
# + id="MzrZ2_RBMHdn" colab_type="code" outputId="d1827acc-0867-4a8f-a0eb-a96b9f6dd822" executionInfo={"status": "ok", "timestamp": 1584551158751, "user_tz": 420, "elapsed": 464486, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 156}
# Bias in embeddings
glove.most_similar(positive=['woman', 'doctor'], negative=['man'], topn=5)
# + [markdown] id="xF1olr2citGG" colab_type="text"
# # Data
# + [markdown] id="c69z9wpJ56nE" colab_type="text"
# ## Load data
# + [markdown] id="2V_nEp5G58M0" colab_type="text"
# We will download the [AG News dataset](http://www.di.unipi.it/~gulli/AG_corpus_of_news_articles.html), which consists of 120000 text samples from 4 unique classes ('Business', 'Sci/Tech', 'Sports', 'World')
# + id="y3qKSoEe57na" colab_type="code" colab={}
import pandas as pd
import re
import urllib
# + id="N18Rcha5CL_t" colab_type="code" colab={}
DATA_FILE = 'news.csv'
INPUT_FEATURE = 'title'
OUTPUT_FEATURE = 'category'
# + id="cGQo98566GIV" colab_type="code" colab={}
# Upload data from GitHub to notebook's local drive
url = "https://raw.githubusercontent.com/madewithml/basics/master/data/news.csv"
response = urllib.request.urlopen(url)
html = response.read()
with open(DATA_FILE, 'wb') as fp:
fp.write(html)
# + id="dG_Oltib6G-9" colab_type="code" outputId="a7b38751-d973-48c9-f965-20b61ec54152" executionInfo={"status": "ok", "timestamp": 1584551159502, "user_tz": 420, "elapsed": 465207, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
# Load data
df = pd.read_csv(DATA_FILE, header=0)
X = df[INPUT_FEATURE].values
y = df[OUTPUT_FEATURE].values
df.head(5)
# + [markdown] id="hxo6RKCQ71dl" colab_type="text"
# ## Split data
# + id="eS6kCcfY6IHE" colab_type="code" colab={}
import collections
from sklearn.model_selection import train_test_split
# + id="55M6iPpZCRWH" colab_type="code" colab={}
TRAIN_SIZE = 0.7
VAL_SIZE = 0.15
TEST_SIZE = 0.15
SHUFFLE = True
# + id="-ZFVitqVWY4J" colab_type="code" colab={}
def train_val_test_split(X, y, val_size, test_size, shuffle):
"""Split data into train/val/test datasets.
"""
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=test_size, stratify=y, shuffle=shuffle)
X_train, X_val, y_train, y_val = train_test_split(
X_train, y_train, test_size=val_size, stratify=y_train, shuffle=shuffle)
return X_train, X_val, X_test, y_train, y_val, y_test
# + id="kqiQd2j_76gP" colab_type="code" outputId="81586394-a8b3-4718-b3cb-db12e67853b1" executionInfo={"status": "ok", "timestamp": 1584551159683, "user_tz": 420, "elapsed": 465360, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
# Create data splits
X_train, X_val, X_test, y_train, y_val, y_test = train_val_test_split(
X=X, y=y, val_size=VAL_SIZE, test_size=TEST_SIZE, shuffle=SHUFFLE)
class_counts = dict(collections.Counter(y))
print (f"X_train: {X_train.shape}, y_train: {y_train.shape}")
print (f"X_val: {X_val.shape}, y_val: {y_val.shape}")
print (f"X_test: {X_test.shape}, y_test: {y_test.shape}")
print (f"Sample point: {X_train[0]} → {y_train[0]}")
print (f"Classes: {class_counts}")
# + [markdown] id="dIfmW7vJ8Jx1" colab_type="text"
# ## Tokenizer
# + [markdown] id="JP4VCO0LAJUt" colab_type="text"
# Unlike the previous notebook, we will be processing our text at a word-level (as opposed to character-level).
# + id="DHPAxkKR7736" colab_type="code" colab={}
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.utils import to_categorical
# + id="rDOj4s1ECa8q" colab_type="code" colab={}
FILTERS = "!\"'#$%&()*+,-./:;<=>?@[\\]^_`{|}~"
LOWER = True
CHAR_LEVEL = False
# + id="dMg5QhVybVfL" colab_type="code" colab={}
def decode(indices, tokenizer):
"""Decode a list of indices into string."""
return " ".join([tokenizer.index_word[index] for index in indices])
# + id="WcscM_vL8KvP" colab_type="code" colab={}
# Input vectorizer
X_tokenizer = Tokenizer(
filters=FILTERS, lower=LOWER, char_level=CHAR_LEVEL, oov_token='<UNK>')
# + id="xV2JgpOA8PwO" colab_type="code" outputId="067cb0d8-a514-464d-868b-12031175b6b7" executionInfo={"status": "ok", "timestamp": 1584551161792, "user_tz": 420, "elapsed": 467433, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Fit only on train data
X_tokenizer.fit_on_texts(X_train)
vocab_size = len(X_tokenizer.word_index) + 1
print (f"# tokens: {vocab_size}")
# + id="ybb-YZSz8Qno" colab_type="code" outputId="6541a5ad-a45a-4859-b0c5-e4786974ae8f" executionInfo={"status": "ok", "timestamp": 1584551162742, "user_tz": 420, "elapsed": 468360, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 68}
# Convert text to sequence of tokens
original_text = X_train[0]
X_train = np.array(X_tokenizer.texts_to_sequences(X_train))
X_val = np.array(X_tokenizer.texts_to_sequences(X_val))
X_test = np.array(X_tokenizer.texts_to_sequences(X_test))
preprocessed_text = decode(X_train[0], X_tokenizer)
print (f"{original_text} \n\t→ {preprocessed_text} \n\t→ {X_train[0]}")
# + [markdown] id="ORGuhjCf8TKh" colab_type="text"
# ## LabelEncoder
# + id="7aBBgzkW8Rxv" colab_type="code" colab={}
from sklearn.preprocessing import LabelEncoder
# + id="ckM_MnQi8UTH" colab_type="code" colab={}
# Output vectorizer
y_tokenizer = LabelEncoder()
# + id="0-FkxqCT8WUk" colab_type="code" outputId="e95f41ab-0e7a-47b2-d67d-ecfa784b3ff9" executionInfo={"status": "ok", "timestamp": 1584551162744, "user_tz": 420, "elapsed": 468335, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Fit on train data
y_tokenizer = y_tokenizer.fit(y_train)
classes = list(y_tokenizer.classes_)
print (f"classes: {classes}")
# + id="yrLHd1i_8XAJ" colab_type="code" outputId="b2b25efb-0c4b-4b8f-bf21-8a5827a9056f" executionInfo={"status": "ok", "timestamp": 1584551162744, "user_tz": 420, "elapsed": 468315, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Convert labels to tokens
y_train = y_tokenizer.transform(y_train)
y_val = y_tokenizer.transform(y_val)
y_test = y_tokenizer.transform(y_test)
print (f"y_train[0]: {y_train[0]}")
# + id="DY91F44BR15z" colab_type="code" outputId="1d60b27e-0ee0-4c6d-e205-47aee11488c4" executionInfo={"status": "ok", "timestamp": 1584551162745, "user_tz": 420, "elapsed": 468297, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# Class weights
counts = np.bincount(y_train)
class_weights = {i: 1.0/count for i, count in enumerate(counts)}
print (f"class counts: {counts},\nclass weights: {class_weights}")
# + [markdown] id="eoWQk0hO9bK2" colab_type="text"
# ## Datasets
# + id="GVxnbzgW8X1V" colab_type="code" colab={}
import math
from tensorflow.keras.preprocessing.sequence import pad_sequences
import torch
import torch.nn as nn
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
# + id="i-ta3se1Cq-4" colab_type="code" colab={}
BATCH_SIZE = 64
FILTER_SIZES = [2, 3, 4]
# + id="dGMp4t7Wkdy0" colab_type="code" outputId="235c0521-465a-40b4-be17-cf384b8b8e95" executionInfo={"status": "ok", "timestamp": 1584551166506, "user_tz": 420, "elapsed": 472036, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Set seed for reproducibility
torch.manual_seed(SEED)
# + id="YiJHwJPXkfBw" colab_type="code" outputId="7df2d03f-1509-4ffc-e858-4631663d8106" executionInfo={"status": "ok", "timestamp": 1584551166509, "user_tz": 420, "elapsed": 472010, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print (device)
# + id="1w6wVKJe9fxk" colab_type="code" colab={}
class TextDataset(Dataset):
"""Text dataset."""
def __init__(self, X, y, batch_size, max_filter_size):
self.X = X
self.y = y
self.batch_size = batch_size
self.max_filter_size = max_filter_size
def __str__(self):
return f"<Dataset(N={len(self)}, batch_size={self.batch_size}, num_batches={self.get_num_batches()})>"
def __len__(self):
return len(self.y)
def __getitem__(self, index):
X = self.X[index]
y = self.y[index]
return X, y
def get_num_batches(self):
return math.ceil(len(self)/self.batch_size)
def collate_fn(self, batch):
"""Processing on a batch."""
# Get inputs
X = np.array(batch)[:, 0]
y = np.array(batch)[:, 1]
# Pad inputs
max_seq_len = max(self.max_filter_size, max([len(x) for x in X]))
X = pad_sequences(X, padding="post", maxlen=max_seq_len)
# Cast
X = torch.LongTensor(X.astype(np.int32))
y = torch.LongTensor(y.astype(np.int32))
return X, y
def generate_batches(self, shuffle=False, drop_last=False):
dataloader = DataLoader(dataset=self, batch_size=self.batch_size,
collate_fn=self.collate_fn,
shuffle=shuffle, drop_last=drop_last)
for (X, y) in dataloader:
yield X, y
# + id="5T8mVj9d9hNI" colab_type="code" outputId="2ab38c2e-ecd4-473d-febd-86c694fdd582" executionInfo={"status": "ok", "timestamp": 1584551166510, "user_tz": 420, "elapsed": 471987, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# Create datasets
train_set = TextDataset(X=X_train, y=y_train, batch_size=BATCH_SIZE, max_filter_size=max(FILTER_SIZES))
val_set = TextDataset(X=X_val, y=y_val, batch_size=BATCH_SIZE, max_filter_size=max(FILTER_SIZES))
test_set = TextDataset(X=X_test, y=y_test, batch_size=BATCH_SIZE, max_filter_size=max(FILTER_SIZES))
print (train_set)
print (train_set[0])
# + id="drbY5WDX9kcL" colab_type="code" outputId="1b436756-da4c-4be4-a5b2-0e77d4ba5010" executionInfo={"status": "ok", "timestamp": 1584551166510, "user_tz": 420, "elapsed": 471969, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# Generate batch
batch_X, batch_y = next(iter(test_set.generate_batches()))
print (batch_X.shape)
print (batch_y.shape)
# + [markdown] id="581nl9EYFAsS" colab_type="text"
# # Embedding
# + [markdown] id="JbOzzfLNFCtW" colab_type="text"
# We can embed our inputs using the [embedding layer](https://pytorch.org/docs/stable/nn.html#torch.nn.Embedding).
# + id="sqgPAuRLFC63" colab_type="code" colab={}
import torch
import torch.nn as nn
# + id="1tHb3v_KH53e" colab_type="code" outputId="7588b4cc-650b-4c2a-bfeb-75326a59400c" executionInfo={"status": "ok", "timestamp": 1584551166511, "user_tz": 420, "elapsed": 471937, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
# Input
x = torch.randint(high=10, size=(1,5)) # high = vocab size
print (x)
print (x.shape)
# + id="FXUpmH7AFOJh" colab_type="code" outputId="dd4ec4ec-1555-4118-83df-9ac6a4a1d687" executionInfo={"status": "ok", "timestamp": 1584551166511, "user_tz": 420, "elapsed": 471913, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Embedding layer
embeddings = nn.Embedding(embedding_dim=100,
num_embeddings=10, # vocab size
padding_idx=0) # which id is for padding
print (embeddings.weight.shape)
# + id="bVGWIgEGGmHn" colab_type="code" outputId="d070b211-a920-459d-8e06-fff0117252da" executionInfo={"status": "ok", "timestamp": 1584551166512, "user_tz": 420, "elapsed": 471896, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Embed the input
embeddings(x).shape
# + [markdown] id="WbO8HYjaGxZY" colab_type="text"
# Each id (token) in the input has been embedded using the embeddings. In the model below, we'll see how to preset our embeddings with our GloVe embeddings and how to choose whether to freeze (don't change during training) those embeddings or not.
# + [markdown] id="yV0AhZWOjXG0" colab_type="text"
# # Modeling
# + [markdown] id="pfhjWZRD94hK" colab_type="text"
# ## Model
# + [markdown] id="eI5xEaMN-vT9" colab_type="text"
# Let's visualize the model's forward pass.
#
# 1. We'll first tokenize our inputs (`batch_size`, `max_seq_len`).
# 2. Then we'll embed our tokenized inputs (`batch_size`, `max_seq_len`, `embedding_dim`).
# 3. We'll apply convolution via filters (`filter_size`, `vocab_size`, `num_filters`) followed by batch normalization. Our filters act as character level n-gram detecors. We have three different filter sizes (2, 3 and 4) and they will act as bi-gram, tri-gram and 4-gram feature extractors, respectivelyy.
# 4. We'll apply 1D global max pooling which will extract the most relevant information from the feature maps for making the decision.
# 5. We feed the pool outputs to a fully-connected (FC) layer (with dropout).
# 6. We use one more FC layer with softmax to derive class probabilities.
# + [markdown] id="zVmJGm8m-KIz" colab_type="text"
# <div align="left">
# <img src="https://raw.githubusercontent.com/madewithml/images/master/basics/14_Embeddings/forward_pass.png" width="1000">
# </div>
# + [markdown] id="JrVDcLC9kNMq" colab_type="text"
# The `FILTER_SIZES` are [2, 3, 4] which effectively act as bi-gram, tri-gram and 4th-gram feature extractors when applied to our text.
# + id="_I3dmAFtsfy6" colab_type="code" colab={}
import torch.nn.functional as F
# + id="UPP5ROd69mXC" colab_type="code" colab={}
class CNN(nn.Module):
def __init__(self, embedding_dim, vocab_size, num_filters,
filter_sizes, hidden_dim, dropout_p, num_classes,
pretrained_embeddings=None, freeze_embeddings=False,
padding_idx=0):
super(CNN, self).__init__()
# Filter sizes
self.filter_sizes = filter_sizes
# Initialize embeddings
if pretrained_embeddings is None:
self.embeddings = nn.Embedding(embedding_dim=embedding_dim,
num_embeddings=vocab_size,
padding_idx=padding_idx)
else:
pretrained_embeddings = torch.from_numpy(pretrained_embeddings).float()
self.embeddings = nn.Embedding(embedding_dim=embedding_dim,
num_embeddings=vocab_size,
padding_idx=padding_idx,
_weight=pretrained_embeddings)
# Freeze embeddings or not
if freeze_embeddings:
self.embeddings.weight.requires_grad = False
# Conv weights
self.conv = nn.ModuleList(
[nn.Conv1d(in_channels=embedding_dim,
out_channels=num_filters,
kernel_size=f) for f in filter_sizes])
# FC weights
self.dropout = nn.Dropout(dropout_p)
self.fc1 = nn.Linear(num_filters*len(filter_sizes), hidden_dim)
self.fc2 = nn.Linear(hidden_dim, num_classes)
def forward(self, x_in, channel_first=False, apply_softmax=False):
# Embed
x_in = self.embeddings(x_in)
# Rearrange input so num_channels is in dim 1 (N, C, L)
if not channel_first:
x_in = x_in.transpose(1, 2)
# Conv outputs
z = []
max_seq_len = x_in.shape[2]
for i, f in enumerate(self.filter_sizes):
# `SAME` padding
padding_left = int((self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2)
padding_right = int(math.ceil((self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2))
# Conv + pool
_z = self.conv[i](F.pad(x_in, (padding_left, padding_right)))
_z = F.max_pool1d(_z, _z.size(2)).squeeze(2)
z.append(_z)
# Concat conv outputs
z = torch.cat(z, 1)
# FC layers
z = self.fc1(z)
z = self.dropout(z)
y_pred = self.fc2(z)
if apply_softmax:
y_pred = F.softmax(y_pred, dim=1)
return y_pred
# + [markdown] id="QBmYu6wjkgf0" colab_type="text"
# ## GloVe embeddings
# + id="x9uev5AGsuqq" colab_type="code" colab={}
def load_glove_embeddings(embeddings_file):
"""Load embeddings from a file."""
embeddings = {}
with open(embeddings_file, "r") as fp:
for index, line in enumerate(fp):
values = line.split()
word = values[0]
embedding = np.asarray(values[1:], dtype='float32')
embeddings[word] = embedding
return embeddings
# + id="tQHD-ThwWnjD" colab_type="code" colab={}
def make_embeddings_matrix(embeddings, word_index, embedding_dim):
"""Create embeddings matrix to use in Embedding layer."""
embedding_matrix = np.zeros((len(word_index) + 1, embedding_dim))
for word, i in word_index.items():
embedding_vector = embeddings.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
return embedding_matrix
# + id="9WxP2GR3LmrO" colab_type="code" outputId="4758001e-8a27-4c7e-db4e-3323c765f9c8" executionInfo={"status": "ok", "timestamp": 1584551176851, "user_tz": 420, "elapsed": 482193, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Create embeddings
embeddings_file = 'glove.6B.{0}d.txt'.format(EMBEDDING_DIM)
glove_embeddings = load_glove_embeddings(embeddings_file=embeddings_file)
embedding_matrix = make_embeddings_matrix(embeddings=glove_embeddings,
word_index=X_tokenizer.word_index,
embedding_dim=EMBEDDING_DIM)
print (f"<Embeddings(words={embedding_matrix.shape[0]}, dim={embedding_matrix.shape[1]})>")
# + [markdown] id="Sf-yZn5CzIex" colab_type="text"
# ## Training
# + colab_type="code" id="FqYSeju9u0O2" colab={}
class Trainer(object):
def __init__(self, **kwargs):
self.__dict__ = kwargs
def train_loop(self, num_epochs):
"""Training and validation steps."""
# Metrics
self.train_loss = []
self.train_acc = []
self.val_loss = []
self.val_acc = []
best_val_loss = np.inf
# Epochs
for epoch in range(num_epochs):
# Steps
self.train_step(epoch)
self.val_step(epoch)
print (f"Epoch: {epoch} | train_loss: {self.train_loss[-1]:.2f}, train_acc: {self.train_acc[-1]:.1f}, val_loss: {self.val_loss[-1]:.2f}, val_acc: {self.val_acc[-1]:.1f}")
# Early stopping
if self.val_loss[-1] < best_val_loss:
best_val_loss = self.val_loss[-1]
patience = self.patience # reset patience
else:
patience -= 1
if not patience: # 0
print ("Stopping early!")
break
return self.train_loss, self.train_acc, self.val_loss, self.val_acc, best_val_loss
def train_step(self, epoch):
"""Training one epoch."""
# Set model to train mode
self.model.train()
# Reset batch metrics
running_train_loss = 0.0
running_train_acc = 0.0
# Iterate over train batches
for i, (X, y) in enumerate(self.train_set.generate_batches()):
# Set device
X = X.to(self.device)
y = y.to(self.device)
# Forward pass
y_pred = self.model(X)
loss = self.loss_fn(y_pred, y)
# Backward pass + optimize
self.optimizer.zero_grad()
loss.backward()
self.optimizer.step()
# Metrics
predictions = y_pred.max(dim=1)[1] # class
accuracy = self.accuracy_fn(y_pred=predictions, y_true=y)
# Update batch metrics
running_train_loss += (loss - running_train_loss) / (i + 1)
running_train_acc += (accuracy - running_train_acc) / (i + 1)
# Update epoch metrics
self.train_loss.append(running_train_loss)
self.train_acc.append(running_train_acc)
# Write to TensorBoard
self.writer.add_scalar(tag='training loss', scalar_value=running_train_loss, global_step=epoch)
self.writer.add_scalar(tag='training accuracy', scalar_value=running_train_acc, global_step=epoch)
def val_step(self, epoch):
"""Validate one epoch."""
# Set model to eval mode
self.model.eval()
# Reset batch metrics
running_val_loss = 0.0
running_val_acc = 0.0
# Iterate over val batches
for i, (X, y) in enumerate(self.val_set.generate_batches()):
# Set device
X = X.to(self.device)
y = y.to(self.device)
# Forward pass
with torch.no_grad():
y_pred = self.model(X)
loss = self.loss_fn(y_pred, y)
# Metrics
predictions = y_pred.max(dim=1)[1] # class
accuracy = self.accuracy_fn(y_pred=predictions, y_true=y)
# Update batch metrics
running_val_loss += (loss - running_val_loss) / (i + 1)
running_val_acc += (accuracy - running_val_acc) / (i + 1)
# Update epoch metrics
self.val_loss.append(running_val_loss)
self.val_acc.append(running_val_acc)
# Write to TensorBoard
self.writer.add_scalar(tag='validation loss', scalar_value=running_val_loss, global_step=epoch)
self.writer.add_scalar(tag='validation accuracy', scalar_value=running_val_acc, global_step=epoch)
# Adjust learning rate
self.scheduler.step(running_val_loss)
def test_loop(self):
"""Evalution of the test set."""
# Metrics
running_test_loss = 0.0
running_test_acc = 0.0
y_preds = []
y_targets = []
# Iterate over val batches
for i, (X, y) in enumerate(self.test_set.generate_batches()):
# Set device
X = X.to(self.device)
y = y.to(self.device)
# Forward pass
with torch.no_grad():
y_pred = self.model(X)
loss = self.loss_fn(y_pred, y)
# Metrics
predictions = y_pred.max(dim=1)[1] # class
accuracy = self.accuracy_fn(y_pred=predictions, y_true=y)
# Update batch metrics
running_test_loss += (loss - running_test_loss) / (i + 1)
running_test_acc += (accuracy - running_test_acc) / (i + 1)
# Store values
y_preds.extend(predictions.cpu().numpy())
y_targets.extend(y.cpu().numpy())
return running_test_loss, running_test_acc, y_preds, y_targets
# + [markdown] id="C26maF-9Goit" colab_type="text"
# ## Experiments
# + [markdown] id="eTWQcUJ_GrIx" colab_type="text"
# Once you have chosen your embeddings, you can choose to freeze them or continue to train them using the supervised data (this could lead to overfitting). In this example, we will do three experiments:
# * frozen GloVe embeddings
# * fine-tuned (unfrozen) GloVe embeddings
# * randomly initialized embeddings
# + id="geKOPVzVK6S9" colab_type="code" colab={}
import matplotlib.pyplot as plt
from torch.optim import Adam
from torch.optim.lr_scheduler import ReduceLROnPlateau
from torch.utils.tensorboard import SummaryWriter
from torchsummary import summary
# %load_ext tensorboard
# + id="64iPmq2lDv2h" colab_type="code" colab={}
EMBEDDING_DIM = 100
NUM_FILTERS = 50
HIDDEN_DIM = 100
DROPOUT_P = 0.1
LEARNING_RATE = 1e-3
PATIENCE = 3
NUM_EPOCHS = 10
# + [markdown] id="To_CB7ibLesP" colab_type="text"
# ### GloVe embeddings (frozen)
# + id="oT9w__AMkqfG" colab_type="code" colab={}
FREEZE_EMBEDDINGS = True
# + id="yg13AyoUkqcJ" colab_type="code" outputId="9c858ee3-844d-40f1-ac3f-8dd09793586a" executionInfo={"status": "ok", "timestamp": 1584551185672, "user_tz": 420, "elapsed": 490964, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
# Initialize model
glove_frozen_model = CNN(embedding_dim=EMBEDDING_DIM,
vocab_size=vocab_size,
num_filters=NUM_FILTERS,
filter_sizes=FILTER_SIZES,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes),
pretrained_embeddings=embedding_matrix,
freeze_embeddings=FREEZE_EMBEDDINGS).to(device)
model = glove_frozen_model
print (model.named_parameters)
# summary(model, input_size=(10, vocab_size)) # bug: can't make inputs into LongTensor
# + id="n-OQ-PRfJFdR" colab_type="code" colab={}
# Loss
weights = torch.Tensor([class_weights[key] for key in sorted(class_weights.keys())]).to(device)
loss_fn = nn.CrossEntropyLoss(weight=weights)
# + id="gi9DSAYGkuBW" colab_type="code" colab={}
# Accuracy
def accuracy_fn(y_pred, y_true):
n_correct = torch.eq(y_pred, y_true).sum().item()
accuracy = (n_correct / len(y_pred)) * 100
return accuracy
# + id="olAw2yp8t4bu" colab_type="code" colab={}
# Optimizer
optimizer = Adam(model.parameters(), lr=LEARNING_RATE)
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=3)
# + id="-uvFsxH34T7K" colab_type="code" colab={}
# Create writer to store values
log_dir = 'tensorboard/glove_frozen'
# !rm -rf log_dir # remove if it already exists
writer = SummaryWriter(log_dir=log_dir)
# + colab_type="code" id="aoM7tPq_u0AL" outputId="2f124a83-b9c0-4263-88d2-e5b98985ce53" executionInfo={"status": "ok", "timestamp": 1584551214873, "user_tz": 420, "elapsed": 520124, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
# Train
trainer = Trainer(train_set=train_set, val_set=val_set, test_set=test_set,
model=model, optimizer=optimizer, scheduler=scheduler,
loss_fn=loss_fn, accuracy_fn=accuracy_fn, patience=PATIENCE,
writer=writer, device=device)
train_loss, train_acc, val_loss, val_acc, best_val_loss = trainer.train_loop(num_epochs=NUM_EPOCHS)
# + [markdown] id="dUVkeDbNqO7V" colab_type="text"
# ### Fine-tuned GloVe embeddings (unfrozen)
# + id="eubLrHydkt_J" colab_type="code" colab={}
# Arguments
FREEZE_EMBEDDINGS = False
# + id="IGeZwoy9qUpa" colab_type="code" outputId="be5a4a64-5900-4854-d625-213be3b360e6" executionInfo={"status": "ok", "timestamp": 1584551214875, "user_tz": 420, "elapsed": 520100, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
# Initialize model
glove_finetuned_model = CNN(embedding_dim=EMBEDDING_DIM,
vocab_size=vocab_size,
num_filters=NUM_FILTERS,
filter_sizes=FILTER_SIZES,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes),
pretrained_embeddings=embedding_matrix,
freeze_embeddings=FREEZE_EMBEDDINGS).to(device)
model = glove_finetuned_model
print (model.named_parameters)
# + id="oUaEr92PqUml" colab_type="code" colab={}
# Optimizer
optimizer = Adam(model.parameters(), lr=LEARNING_RATE)
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=3)
# + id="3qf5oYZd4RZ3" colab_type="code" colab={}
# Create writer to store values
log_dir = 'tensorboard/glove_finetuned'
# !rm -rf log_dir # remove if it already exists
writer = SummaryWriter(log_dir=log_dir)
# + id="NpyhLUK2qUjb" colab_type="code" outputId="e58bcfbe-ef4c-46f4-baa6-79710abacb0f" executionInfo={"status": "ok", "timestamp": 1584551244557, "user_tz": 420, "elapsed": 549757, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
# Train
trainer = Trainer(train_set=train_set, val_set=val_set, test_set=test_set,
model=model, optimizer=optimizer, scheduler=scheduler,
loss_fn=loss_fn, accuracy_fn=accuracy_fn, patience=PATIENCE,
writer=writer, device=device)
train_loss, train_acc, val_loss, val_acc, best_val_loss = trainer.train_loop(num_epochs=NUM_EPOCHS)
# + [markdown] id="Y8JzMrcv_p8a" colab_type="text"
# ### Randomly initialized embeddings
# + id="TnLSYV0WKo8x" colab_type="code" colab={}
# Arguments
FREEZE_EMBEDDINGS = False
# + id="wD4sRUS5_lwq" colab_type="code" outputId="5109d441-87fb-42eb-e96f-1e8dd49a3912" executionInfo={"status": "ok", "timestamp": 1584551244558, "user_tz": 420, "elapsed": 549724, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
randomly_initialized_model = CNN(embedding_dim=EMBEDDING_DIM,
vocab_size=vocab_size,
num_filters=NUM_FILTERS,
filter_sizes=FILTER_SIZES,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes),
pretrained_embeddings=None,
freeze_embeddings=FREEZE_EMBEDDINGS).to(device)
model = randomly_initialized_model
print (model.named_parameters)
# + id="Ucn3tYq1_sE1" colab_type="code" colab={}
# Optimizer
optimizer = Adam(model.parameters(), lr=LEARNING_RATE)
scheduler = ReduceLROnPlateau(optimizer, mode='min', factor=0.1, patience=3)
# + id="by1-D8cN4W2u" colab_type="code" colab={}
# Create writer to store values
log_dir = 'tensorboard/randomly_initialized'
# !rm -rf log_dir # remove if it already exists
writer = SummaryWriter(log_dir=log_dir)
# + id="F7bTmNdCJA0g" colab_type="code" outputId="d377761a-6207-4747-8570-680c4683a55f" executionInfo={"status": "ok", "timestamp": 1584551282592, "user_tz": 420, "elapsed": 587728, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 119}
# Train
trainer = Trainer(train_set=train_set, val_set=val_set, test_set=test_set,
model=model, optimizer=optimizer, scheduler=scheduler,
loss_fn=loss_fn, accuracy_fn=accuracy_fn, patience=PATIENCE,
writer=writer, device=device)
train_loss, train_acc, val_loss, val_acc, best_val_loss = trainer.train_loop(num_epochs=NUM_EPOCHS)
# + [markdown] id="vskwiiI3V3S6" colab_type="text"
# ## Evaluation
# + [markdown] id="6tO2hX8OLQ5s" colab_type="text"
# Looks like fine-tuned glove embeddings had the best test performance (based on validation accuracy) so let's do proper evaluation and inference with that strategy.
# + id="Itq7lT9qV9Y8" colab_type="code" colab={}
import io
import itertools
import json
import matplotlib.pyplot as plt
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import precision_recall_fscore_support
# + id="NNeyYs3tW3VN" colab_type="code" colab={}
def plot_confusion_matrix(y_true, y_pred, classes, cmap=plt.cm.Blues):
"""Plot a confusion matrix using ground truth and predictions."""
# Confusion matrix
cm = confusion_matrix(y_true, y_pred)
cm_norm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
# Figure
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(cm, cmap=plt.cm.Blues)
fig.colorbar(cax)
# Axis
plt.title("Confusion matrix")
plt.ylabel("True label")
plt.xlabel("Predicted label")
ax.set_xticklabels([''] + classes)
ax.set_yticklabels([''] + classes)
ax.xaxis.set_label_position('bottom')
ax.xaxis.tick_bottom()
# Values
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, f"{cm[i, j]:d} ({cm_norm[i, j]*100:.1f}%)",
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
# Display
plt.show()
# + id="smP8T1bEW3fH" colab_type="code" colab={}
def get_performance(y_true, y_pred, classes):
"""Per-class performance metrics. """
performance = {'overall': {}, 'class': {}}
metrics = precision_recall_fscore_support(y_true, y_pred)
# Overall performance
performance['overall']['precision'] = np.mean(metrics[0])
performance['overall']['recall'] = np.mean(metrics[1])
performance['overall']['f1'] = np.mean(metrics[2])
performance['overall']['num_samples'] = np.float64(np.sum(metrics[3]))
# Per-class performance
for i in range(len(classes)):
performance['class'][classes[i]] = {
"precision": metrics[0][i],
"recall": metrics[1][i],
"f1": metrics[2][i],
"num_samples": np.float64(metrics[3][i])
}
return performance
# + id="avmwpr5syKHY" colab_type="code" outputId="b4486135-c371-49f8-e693-7199e677e630" executionInfo={"status": "ok", "timestamp": 1584551600028, "user_tz": 420, "elapsed": 968, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Test loop
model = glove_finetuned_model
test_loss, test_acc, y_preds, y_targets = trainer.test_loop()
print (f"test_loss: {test_loss:.2f}, test_acc: {test_acc:.1f}")
# + id="qdAj6KyCU88E" colab_type="code" outputId="120b0a2b-696e-4c39-e79d-78f55c16cd02" executionInfo={"status": "ok", "timestamp": 1584551282594, "user_tz": 420, "elapsed": 587669, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 595}
# Class performance
performance = get_performance(y_true=y_targets, y_pred=y_preds, classes=classes)
print (json.dumps(performance, indent=4))
# + id="nRbPfqgZWaof" colab_type="code" outputId="7da8a938-bb5e-4ac5-f8b3-55d147a7fec4" executionInfo={"status": "ok", "timestamp": 1584551282595, "user_tz": 420, "elapsed": 587650, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 598}
# Confusion matrix
plt.rcParams["figure.figsize"] = (7,7)
plot_confusion_matrix(y_targets, y_preds, classes=classes)
print (classification_report(y_targets, y_preds))
# + [markdown] id="yeiD1T_QZpdk" colab_type="text"
# ## Inference
# + id="z7G7vuSTZHkQ" colab_type="code" colab={}
import collections
# + id="362Bl2chXDOA" colab_type="code" colab={}
def get_probability_distribution(y_prob, classes):
results = {}
for i, class_ in enumerate(classes):
results[class_] = np.float64(y_prob[i])
sorted_results = {k: v for k, v in sorted(
results.items(), key=lambda item: item[1], reverse=True)}
return sorted_results
# + id="CLP2Vzp3Zwth" colab_type="code" outputId="a8b26809-bc68-4bf8-86cd-aae4d3adf5a4" executionInfo={"status": "ok", "timestamp": 1584551606237, "user_tz": 420, "elapsed": 505, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 85}
# Inputs
texts = ["The Wimbledon tennis tournament starts next week."]
num_samples = len(texts)
X_infer = np.array(X_tokenizer.texts_to_sequences(texts))
print (f"{texts[0]} \n\t→ {decode(X_infer[0], X_tokenizer)} \n\t→ {X_infer[0]}")
print (f"len(X_infer[0]): {len(X_infer[0])} words")
y_filler = np.array([0]*num_samples)
# + id="q1gFlI5MZ143" colab_type="code" colab={}
# Dataset
infer_set = TextDataset(X=X_infer, y=y_filler, batch_size=BATCH_SIZE, max_filter_size=max(FILTER_SIZES))
# + id="UFE4sp_7aHTq" colab_type="code" colab={}
# Iterate over val batches
probabilities = []
for i, batch in enumerate(infer_set.generate_batches()):
# Cast
X, y = batch
X = X.to(device)
y = y.to(device)
# Forward pass
with torch.no_grad():
y_pred = model(X, apply_softmax=True)
# Save probabilities
probabilities.extend(y_pred.cpu().numpy())
# + id="bGi_NvbBaMap" colab_type="code" outputId="bc853c9d-c477-4c77-d197-d613a9617645" executionInfo={"status": "ok", "timestamp": 1584551609255, "user_tz": 420, "elapsed": 751, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 221}
# Results
results = []
for index in range(num_samples):
results.append({
'raw_input': texts[index],
'preprocessed_input': decode(indices=X_infer[index], tokenizer=X_tokenizer),
'probabilities': get_probability_distribution(probabilities[index], y_tokenizer.classes_)
})
print (json.dumps(results, indent=4))
# + [markdown] id="Y4-WkjN595lO" colab_type="text"
# # Interpretability
# + [markdown] id="Uo0FqqEY98El" colab_type="text"
# Recall that each our unique filter sizes (2, 3 and 4) act as n-gram feature detectors. When these filters convolve on our embedded input (`N`, `max_seq_len`, `embedding_dim`), they produce feature maps which are shape ((`N`, `max_seq_len`, `num_filters`) for each filter size. Since we used `SAME` padding with stride=1, our feature maps have the same length as our inputs ('max_seq_len') which you can think of as what the filters extracted from each n-gram window. When we apply 1d global max-pooling we're effectively extracting the most relevant information from the feature maps. We can inspect the trained model at the pooling step to determine which n-grams were most relevant towards the prediction.
# + id="Zv2uqi6mOe9Z" colab_type="code" colab={}
import seaborn as sns
from statistics import mode
# + [markdown] id="M-aGz2BgCCKq" colab_type="text"
# We're going to copy the same model structure as before but now we'll stop just after convolution since those are the outputs we care about.
# + id="_nzdZ2_tBsfc" colab_type="code" colab={}
class ConvOutputsModels(nn.Module):
def __init__(self, embedding_dim, vocab_size, num_filters,
filter_sizes, hidden_dim, dropout_p, num_classes,
pretrained_embeddings=None, freeze_embeddings=False,
padding_idx=0):
super(ConvOutputsModels, self).__init__()
# Filter sizes
self.filter_sizes = filter_sizes
# Initialize embeddings
if pretrained_embeddings is None:
self.embeddings = nn.Embedding(embedding_dim=embedding_dim,
num_embeddings=vocab_size,
padding_idx=padding_idx)
else:
pretrained_embeddings = torch.from_numpy(pretrained_embeddings).float()
self.embeddings = nn.Embedding(embedding_dim=embedding_dim,
num_embeddings=vocab_size,
padding_idx=padding_idx,
_weight=pretrained_embeddings)
# Freeze embeddings or not
if freeze_embeddings:
self.embeddings.weight.requires_grad = False
# Conv weights
self.conv = nn.ModuleList(
[nn.Conv1d(in_channels=embedding_dim,
out_channels=num_filters,
kernel_size=f) for f in filter_sizes])
# FC weights
self.dropout = nn.Dropout(dropout_p)
self.fc1 = nn.Linear(num_filters*len(filter_sizes), hidden_dim)
self.fc2 = nn.Linear(hidden_dim, num_classes)
def forward(self, x_in, channel_first=False, apply_softmax=False):
# Embed
x_in = self.embeddings(x_in)
# Rearrange input so num_channels is in dim 1 (N, C, L)
if not channel_first:
x_in = x_in.transpose(1, 2)
# Conv outputs
z = []
max_seq_len = x_in.shape[2]
for i, f in enumerate(self.filter_sizes):
# `SAME` padding
padding_left = int((self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2)
padding_right = int(math.ceil((self.conv[i].stride[0]*(max_seq_len-1) - max_seq_len + self.filter_sizes[i])/2))
# Conv
_z = self.conv[i](F.pad(x_in, (padding_left, padding_right)))
z.append(_z.cpu().numpy())
return z
# + id="XfWHwZ7DB2gf" colab_type="code" outputId="5363a5c9-4e7c-484b-db70-03e965212a09" executionInfo={"status": "ok", "timestamp": 1584551616473, "user_tz": 420, "elapsed": 1059, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
# Initialize model
model = ConvOutputsModels(embedding_dim=EMBEDDING_DIM,
vocab_size=vocab_size,
num_filters=NUM_FILTERS,
filter_sizes=FILTER_SIZES,
hidden_dim=HIDDEN_DIM,
dropout_p=DROPOUT_P,
num_classes=len(classes),
pretrained_embeddings=embedding_matrix,
freeze_embeddings=FREEZE_EMBEDDINGS).to(device)
print (model.named_parameters)
# + [markdown] id="4fL_exZ2CMP0" colab_type="text"
# Since we already trained our model, we'll transfer those weights to our new model.
# + id="Q24ZsZofCkNV" colab_type="code" outputId="1ea15888-c1d8-4c09-fc2e-7edeccf9233b" executionInfo={"status": "ok", "timestamp": 1584551618116, "user_tz": 420, "elapsed": 269, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# Transfer weights
model.load_state_dict(glove_finetuned_model.state_dict())
# + id="pZQY75xXC4rZ" colab_type="code" colab={}
# Iterate over val batches
conv_outputs = []
for i, batch in enumerate(infer_set.generate_batches()):
# Cast
X, y = batch
X = X.to(device)
y = y.to(device)
# Forward pass
with torch.no_grad():
z = model(X, apply_softmax=True)
# Save
conv_outputs.extend(z)
# + id="RyC7FJndIFaE" colab_type="code" outputId="8f8c47da-e636-4d79-eaf3-0dcc46b31eb4" executionInfo={"status": "ok", "timestamp": 1584551629600, "user_tz": 420, "elapsed": 822, "user": {"displayName": "<NAME>", "photoUrl": "https://<KEY>", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 494}
# Visualize bi-gram filters
tokens = decode(X_infer[0], X_tokenizer).split(' ')
sns.heatmap(conv_outputs[1][0], xticklabels=tokens)
# + [markdown] id="cTHPBDzWPI8s" colab_type="text"
# 1D global max-pooling would extract the highest value from each of our num_filters for each filter size. We could also follow this same approach to figure out which n-gram is most relevant but notice in the heatmap above that many filters don't have much variance. To mitigate this, this [paper](https://www.aclweb.org/anthology/W18-5408/) uses threshold values to determine which filters to use for interpretability.
#
# But to keep things simple, let's extract which tokens' filter outputs were extracted via max-pooling the most frequenctly.
# + id="P72CZhU0CtGa" colab_type="code" outputId="d6d177de-6aea-4ec3-cc0b-1a7fb8a806df" executionInfo={"status": "ok", "timestamp": 1584551640663, "user_tz": 420, "elapsed": 978, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjMIOf3R_zwS_zZx4ZyPMtQe0lOkGpPOEUEKWpM7g=s64", "userId": "00378334517810298963"}} colab={"base_uri": "https://localhost:8080/", "height": 136}
sample_index = 0
print (f"Preprocessed text:\n{decode(indices=X_infer[sample_index], tokenizer=X_tokenizer)}")
print ("\nMost important n-grams:")
# Process conv outputs for each unique filter size
for i, filter_size in enumerate(FILTER_SIZES):
# Identify most important n-gram (excluding last token)
popular_indices = collections.Counter([np.argmax(conv_output) \
for conv_output in conv_outputs[i][0]])
# Get corresponding text
start = popular_indices.most_common(1)[-1][0]
n_gram = " ".join([token for token in tokens[start:start+filter_size]])
print (f"[{filter_size}-gram]: {n_gram}")
# + [markdown] id="kbZPYQ2TH1Jt" colab_type="text"
# ---
# Share and discover ML projects at <a href="https://madewithml.com/">Made With ML</a>.
#
# <div align="left">
# <a class="ai-header-badge" target="_blank" href="https://github.com/madewithml/basics"><img src="https://img.shields.io/github/stars/madewithml/basics.svg?style=social&label=Star"></a>
# <a class="ai-header-badge" target="_blank" href="https://www.linkedin.com/company/madewithml"><img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
# <a class="ai-header-badge" target="_blank" href="https://twitter.com/madewithml"><img src="https://img.shields.io/twitter/follow/madewithml.svg?label=Follow&style=social"></a>
# </div>
#
| notebooks/14_Embeddings/14_PT_Embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Imports
from bs4 import BeautifulSoup
import pandas as pd
import time
import os
import re
from tqdm import tqdm
# Liste aller HTMLs
lst = os.listdir('pages/')
# Seiten einlesen, hier zuerst mit einer Seite
file = open('pages/' + lst[0], 'r')
text = file.read()
h = BeautifulSoup(text, 'html.parser')
# Alle Elemente, die uns interessieren
h.find_all('tr', {'class':'ng-scope'})
# Alle Elemente
h.find_all('tr', {'class':'ng-scope'})[0].find('div')['title']
h.find_all('tr', {'class':'ng-scope'})[0].find_next('div').find_next('div').find('a')['href']
h.find_all('tr', {'class':'ng-scope'})[0].find_next('div') \
.find_next('div').find_next('div').find_next('div').find_next('div').text
h.find_all('tr', {'class':'ng-scope'})[0].find_next('div') \
.find_next('div').find_next('div').find_next('div') \
.find_next('div').find_next('div').text
# Und jetzt alles in einen For_loop packen
soup_lst = h.find_all('tr', {'class':'ng-scope'})
# +
bäck_lst = []
for bäck in soup_lst:
name = bäck.find('div')['title']
url = bäck.find_next('div').find_next('div').find('a')['href']
gem = bäck.find_next('div') \
.find_next('div').find_next('div').find_next('div').find_next('div').text
kt = bäck.find_next('div') \
.find_next('div').find_next('div').find_next('div') \
.find_next('div').find_next('div').text
minidict = {'Bäckerei': name,
'URL': url,
'Gemeinde': gem,
'Kanton': kt}
bäck_lst.append(minidict)
# -
# Jetzt machen wir daraus eine Funktion
def get_bäcks(html_code):
soup_lst = html_code.find_all('tr', {'class':'ng-scope'})
bäck_lst = []
for bäck in soup_lst:
name = bäck.find('div')['title']
url = bäck.find_next('div').find_next('div').find('a')['href']
gem = bäck.find_next('div') \
.find_next('div').find_next('div').find_next('div').find_next('div').text
kt = bäck.find_next('div') \
.find_next('div').find_next('div').find_next('div') \
.find_next('div').find_next('div').text
minidict = {'Bäckerei': name,
'URL': url,
'Gemeinde': gem,
'Kanton': kt}
bäck_lst.append(minidict)
return bäck_lst
file = open('pages/' + lst[0], 'r')
text = file.read()
h = BeautifulSoup(text, 'html.parser')
get_bäcks(h)
# Und jetzt wenden wir das auf alle html files an
full_bäck_lst = []
for html_file in tqdm(lst):
file = open('pages/' + html_file, 'r')
text = file.read()
h = BeautifulSoup(text, 'html.parser')
full_bäck_lst = full_bäck_lst + get_bäcks(h)
# Einlesen in die Liste und abspeichern
df = pd.DataFrame(full_bäck_lst)
df.to_csv('bäckereien_list.csv')
df['Kanton'].value_counts()
| 10 Selenium/2 Htmls auslesen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Estructuras de Control Iterativas
# A diferencia de las estructuras de control condicionales, las iterativas (también llamadas cíclicas o bucles), nos permiten ejecutar un mismo código, de manera repetida, mientras se cumpla una condición. En Python se dispone de dos estructuras cíclicas:
# - El bucle while
# - El bucle for
#
# Las veremos en detalle a continuación.
# ## Bucle While
# ------------------------------
# Se basa en <b>repetir un bloque a partir de evaluar una condición lógica, siempre que ésta sea True</b>. Queda en las manos del programador decidir el momento en que la condición cambie a False para hacer que el While finalice.
# ### Ejemplo
#
# Mientras que año sea menor o igual a 2012, imprimir la frase “Informes del Año año”
#
# +
# -*- coding: utf-8 -*
anio = 2001
while anio <= 2012:
print("Informes del Año {}".format(anio))
anio += 1
# anio = anio + 2
# anio += 1
# -
# Si miras la última línea:
#
# <code>anio += 1</code>
#
# Podrás notar que en cada iteración, incrementamos el valor de la variable que condiciona el bucle (anio). Si no lo hiciéramos, esta variable siempre sería igual a 2001 y el bucle se ejecutaría de forma infinita, ya que la condición (anio <= 2012) siempre se estaría cumpliendo.
# ### - Instruccion <b>break</b>
# Sirve para "romper" la ejecución del While en cualquier momento. No se ejecutará el Else, ya que éste sólo se llama al finalizar la iteración.:
c = 0
while c <= 5:
c+=1
if c == 4:
print("Rompemos el bucle cuando c vale", c)
break
print("c vale",c)
# ### - Instruccion <b>continue</b>
# Sirve para "saltarse" la iteración actual sin romper el bucle.
c = 0
while c <= 5:
c+=1
#c = c+1
if c==3 or c==4:
# print("Continuamos con la siguiente iteración", c)
continue
print("c vale",c)
# ### Ejemplo Menú Interactivo
### print("Bienvenido al menú interactivo")
while(True):
print("""¿Qué quieres hacer? Escribe una opción
1) Saludar
2) Sumar dos números
3) Salir""")
opcion = input()
if opcion == '1':
print("Hola, espero que te lo estés pasando bien")
elif opcion == '2':
n1 = float(input("Introduce el primer número: "))
n2 = float(input("Introduce el segundo número: "))
print("El resultado de la suma es: ",n1+n2)
elif opcion =='3':
print("¡Hasta luego! Ha sido un placer ayudarte")
break
else:
print("Comando desconocido, vuelve a intentarlo")
# ### EJERCICIOS
# Realiza un programa que lea dos números por teclado y permita elegir entre 3 opciones en un menú:
#
# - Mostrar una suma de los dos números
# - Mostrar una resta de los dos números (el primero menos el segundo)
# - Mostrar una multiplicación de los dos números
#
# En caso de introducir una opción inválida, el programa informará de que no es correcta
# ## Bucle For
# ----------------------------------------------
# El bucle <code>for</code>, en Python, es aquel que nos permitirá iterar sobre una variable compleja, del tipo lista o tupla:
# Iterando sobre listas
mi_lista = ['Juan', 'Antonio', 'Pedro', 'Herminio']
for elemento in mi_lista:
print(elemento)
# Modificando valores sobre listas
mi_lista = ['Juan', 'Antonio', 'Pedro', 'Herminio']
for indice,nombre in enumerate(mi_lista):
if nombre=='Juan':
mi_lista[indice]='Maria'
# valor cambiado
mi_lista
# +
# Iterando sobre diccionarios
dicx = {'val1':1,'val2':2,'val3':3}
for key,val in dicx.items():
print(key,val)
# -
dicx.items
# ### Funcion Range
# Sirve para generar una lista de números que podemos recorrer fácilmente, pero no ocupa memoria porque se interpreta sobre la marcha:
#
#
# <img src='https://cdn.techbeamers.com/wp-content/uploads/2019/05/Python-range-function-explained.png'>
[*range(1,6,2)]
##es una especie de lista
# -*- coding: utf-8 -*-
# Generarndo un rango de valores del 2001 al 2012
for anio in range(2001, 2013):
print("Informes del Año", str(anio))
# por defecto el inicio de la funcion es 0
for i in range(10):
print(i)
text= hola que tal
for
# ## EJERCICIOS
# 1. Realiza un programa que pida al usuario cuantos números quiere introducir. Luego lee todos los números y realiza una media aritmética.
# #### Nota
# Usar listas
#
#
cantidad = int(input('ingrese la cantidad de numeros a introducir'))
numero = []
for n in range(cantidad):
x = int(input('ingrese el numero a ser sumado: '))
numero.append(x)
sum(numero)/cantidad
# 2. Escribir un programa que pida al usuario un número entero y muestre por pantalla un triángulo rectángulo como el de más abajo, de altura el número introducido.
#
#
# Para n = 4
#
# <code>#
# ##
# ###
# ####</code>
q=int(input("ingrese la cantidad de #s: "))
# +
# forma 1
for i in range(q):
print('#'*(i+1))
# -
# prueba
'#'*4
# forma 2
nums=[]
v=1
for elem in range(v,q+1):
nums.append(v*("#"))
v=v+1
for b in (nums):
print (b)
# 3. Escribir un programa que pida al usuario un número entero y muestre por pantalla si es un número primo o no.
#
numero = int(input("ingrese un numero entero:"))
numero
primo = True
for n in range(2, numero, 1):
if numero % n ==0:
primo = False
break
7 % 2?
7 % 3?
7 % 4?
7 % 5?
7 % 6?
if primo:
print(f'el numero: {numero} es primo')
else:
print(f'el numero: {numero} no es primo')
# 4. Dadas dos listas, debes generar una tercera con todos los elementos que se repitan en ellas, pero no debe repetirse ningún elemento en la nueva lista:
#
#
lista_1 = ["h",'o','l','a',' ', 'm','u','n','d','o']
lista_2 = ["h",'o','l','a',' ', 'l','u','n','a']
l1=set(lista_1)
l2=set(lista_2)
l1
l2
l3 = l1.intersection(l2)
list(l3)
| Modulo2/1. Estructuras de Control Iterativas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
excel = pd.read_excel('/Users/alicia/Downloads/2019_05_10 TMNA Master Site Aqueduct Input 2019_08_19 (1).xlsx')
excel.head()
excel.columns
| Aqueduct/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### AOS 575 Application Lab 4 - aos575_applab4_fft_EPICA.ipynb - Dome-C Ice Core (EPICA)
# Coded by <NAME> (University of Colorado) with functions from <NAME> (CSU) <br>
# Peer-reviewed by <NAME>/<NAME> (University of Colorado), last updated Monday October 19, 2020 <br>
# Updated/adapted for AOS 575 by <NAME> (UWisc), 10/25/2020
#
# #### Learning Goals
# 1. Calculate the power spectrum for data that started on a non-uniform grid
# 2. Contrast the Boxcar and Hanning wind approaches. What are the advantages/disadvantages of these two windows? How do they change the spectra?
# 3. Apply a Hanning window of different window lengths. How does that affect the statistical significance and temporal precision?
# 4. Appling a Hanning window with Welch's Method (Windows of Overlapping Segment Analysis, WOSA). How does WOSA change the results?
#
# #### Underlying science and data
# This notebook has code that computes the power spectrum of the temperature record from the Dome-C ice core (Antarctica) taken as (123E,75S) (Jouzel et al. 2007, https://science.sciencemag.org/content/317/5839/793). The data goes back 800,000 years before present and are unevenly spaced in time because the further down you go, the more compressed ice gets (older ice samples a longer segment of time). You can obtain the data online here from the NOAA Paleoclimatology Program and World Data Center for Paleoclimatology:
# ftp://ftp.ncdc.noaa.gov/pub/data/paleo/icecore/antarctica/epica_domec/edc3deuttemp2007.txt More information on the data is available at:
# https://www.ncdc.noaa.gov/paleo-search/study/6080 <br>
#
# Power spectral analysis of this dataset will illuminate the periodic behavior of ice ages due to Milankovitch cycles. Milankovitch cycles are periodic variations in climate due to the Earth's gradual orbital variations in eccentricity, tilt, and precession. If Milankovitch cyles are a new phenomena for you, check out this website after class: https://climate.nasa.gov/news/2948/milankovitch-orbital-cycles-and-their-role-in-earths-climate/
#
# #### How to complete this notebook
# 1. Power spectrum analysis via fast Fourier transform (FFT) requires evenly spaced data. Regrid the data to an evenly spaced dataset. Plot the before and after data to make sure things worked and make sense. <br>
# 2. What is the autocorrelation and e-folding time of your data? Calculate the power spectra using the Numpy method (which uses a boxcar window by default. Graph the power spectrum, a red noise fit to the data, and the 99% confidence interval. What statistically significant spectral peaks do you find and how much power do they have? What do they represent?
# Insert answer here.
# 3. Compare the Boxcar and Hanning window: Calculate and plot the power spectra using the scipy periodogram method which allows quick comparison between different window types. Compare the power spectrum results using the two different windows. What are the stiatistically significant peaks? What are the differences that occur from using the two different windows? Why?
# Insert answer here.
# 4. Compare Hanning windows of different lengths and use multiple chunks of the dataset to increase spectral confidence. Calculate and plot the power spectra using the scipy method but with different window lengths. Try a couple different window lengths to see how the results change. How does window length change the statistically signficant peaks that you obtain? How does decreasing the window length affect the temporal precision of the spectral peaks and the statistical significance? Do you find the tradeoff between (1) high spectral/temporal resolution but low quality statistics AND (2) high quality statistics but low spectral/temporal resolution?
# Insert answer here.
# 5. Apply WOSA: Apply Welch's method with a Hanning window with different window lengths. Use the same window lengths that you tried in 4. How does using overlaps with Welch's method change your results?
# Insert answer here.
# Import modules
import numpy as np #for calculations
import matplotlib.pyplot as plt #for plotting
import pandas as pd #for reading in data
import datetime #for manipulation by dtae
import collections #for additional dictionary functionality
from matplotlib.gridspec import GridSpec #for subplots
import scipy.signal as signal #for signal processing
import scipy.stats as stats
# A couple helpful functions
# +
def create_normalized_redfit(data_length,Te):
freq = np.arange(0,(data_length/2)+1,1)/float(data_length) # to Nyquist
red_fit = (2 * Te)/(1 + ((2*np.pi*freq)**2)*(Te**2)) # After Hartmann 6.64, 6.91
return red_fit/np.sum(red_fit)
def create_f_bounds(alpha,dof,red_fit_n):
f_ratio = stats.f.ppf(alpha,dof,200) # Note: 200 = large degree of freedom for red noise
return f_ratio*red_fit_n
# -
# Read in the ice core data <br>
# Data are from ftp://ftp.ncdc.noaa.gov/pub/data/paleo/icecore/antarctica/epica_domec/edc3deuttemp2007.txt
filename = 'edc3deuttemp2007_nohead.txt'
data_all=pd.read_csv(filename,sep='\s+',names = ['bag', 'ztop','age','Deuterium','TempAnom'])
data_all.head()
# We're using the temperature data (which is derived from different paleoproxies - how that's done is a different class! Go talk to the geologists in the building next door...)<br>
# Read in the temperature data from the ice core:
data_raw=data_all['TempAnom']
time_raw=data_all['age']
var_str = "Temp"
data_units = "C"
# Plot raw temperature data
fig = plt.figure(figsize=(10,4))
plt.plot(time_raw.values,data_raw, lw=.75)
plt.ylabel(data_units,fontsize=9)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.xlabel('Age (years)',fontsize=14)
plt.ylabel('Temperature Anomaly (K)',fontsize=14)
plt.title("Dome-C Ice Core Antarctica",fontsize=14)
plt.grid()
#plt.savefig('XX.pdf', dpi=300, format='png')
# PROBLEM FOR FFT ANALYSIS: Raw data are not evenly spaced in time. <br>
# Need to fix that as FFT requires data on evenly spaced grid <br>
#
# +
## Print statements and calculate 1st derivative to show raw data are not evenly spaced
#print(time.values)
dt=np.diff(time_raw.values)
#print(dt)
#print(round(np.mean(dt),0))
print(min(time_raw.values),0)
print(max(time_raw.values),0)
### Define the grid
time=np.linspace(min(time_raw.values),max(time_raw.values),800)
#time=np.linspace(min(time_raw.values),max(time_raw.values),max(time_raw.values)-min(time_raw.values)+1)
#print(time)
dt=np.diff(time)
print(dt[0])
dt_regrid=dt[0]
## Interpolate raw data to an evenly spaced grid
data=np.interp(time,time_raw,data_raw)
print('regridded spacing in years',dt[1])
# -
# Plot the evenly spaced data:
fig = plt.figure(figsize=(10,4))
plt.plot(time,data, lw=.75)
plt.ylabel(data_units,fontsize=14)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
plt.title("Vostok Ice Core Antarctica - regridded on evenly spaced grid",fontsize=14)
plt.grid()
plt.ylabel('Temperature anomaly (K)')
plt.xlabel('Age (years)')
#plt.savefig('XX.pdf', dpi=300, format='png')
print(len(data))
print(len(time))
## calculate the anomaly note: need to do this to get equivalent results with numpy and scipy method (not sure why?)
data=data-data.mean()
# Calculate the power spectrum of red noise with lag1_r to use for significance testing
# +
alpha = 0.99 ## set statistical significance level
### step 1: calculate lag-1 autocorrelation (lag1_r, rho) and the associated p value (lag1_p)
lag1_r,lag1_p = stats.pearsonr(data[0:len(data)-1],data[1:len(data)])
### step 2: Calculate e-folding time for a red-noise process with this lag-1 autocorrelation
Te = -1./np.log(lag1_r) # After Hartman 6.62 with delta t = 1
print(var_str,'lag-1 autocorrelation =',round(lag1_r,2),'and Te =',round(Te,0))
## calculate the power spectrum of red noise with lag1_r to use for significance testing
red_fit_n = create_normalized_redfit(len(data),Te)
dof_entirewindow=2 ### note dof=2 because using whole record for FFT with no chunking
f_bounds = create_f_bounds(alpha,dof_entirewindow,red_fit_n) ## using f-test for variance, see function
# -
# Calculate the power spectrum. First with the numpy method:
## Method #1: Numpy Method - Calculate Power Spectrum
freq = np.fft.rfftfreq(len(data),d=1.) ###
#print(freq)
A = np.fft.rfft(data) ### r = real
ps = (np.abs(A)**2)/2.
ps_n = (np.abs(A)**2)/np.sum(np.abs(A)**2) ## normalizing - calculate percent of the total
# Plot the results -- replicate a figure from a paper in Science that has 6000+ citations (Petit et al. 1999)
fig = plt.figure(figsize=(15,6))
method_str = "Boxcar Window on Full Dataset"
plt.plot(freq,ps_n, color='C0', lw=1.3, label='Boxcar')
plt.plot(freq,red_fit_n, color='firebrick', lw=1, alpha=0.9, label='Red Noise Fit') # freq plots 0...0.5
plt.plot(freq,f_bounds, color='firebrick', lw=1, ls='--', alpha=0.9, label=str(int(alpha*100))+'% CI')
plt.ylim(0.0,0.3)
plt.xlim(0.0,0.06)
plt.xlabel("Frequency (approximately millennia$^{-1}$)",fontsize=14)
plt.ylabel("Normalized Power",fontsize=14)
plt.title("Normalized Power Spectrum of "+var_str+"\n"+method_str,fontsize=14)
plt.legend(loc='upper right',fontsize=14)
plt.grid()
fig.tight_layout()
# Find the significant spectral peaks:
for i in range(len(ps_n)):
if (ps_n[i] - f_bounds[i]) > 5e-03:
print('##### FOUND IT - spectral peak exceeds red noise ####')
print('exceeds by...',ps_n[i]-f_bounds[i])
print('at frequency....',freq[i])
print('which in years is approximately...',round((dt_regrid/freq[i]))) ## use the regridded regular spacing
print()
# Scipy Method: Compare Boxcar (sqaure) Window with a Hanning (tapered) Window. <br>
# http://docs.scipy.org/doc/scipy/reference/generated/scipy.signal.periodogram.html#scipy.signal.periodogram
# Which would you prefer??? Boxcar or Hanning??
# +
## Use Boxcar Window
f,Pxx = signal.periodogram(data,fs=1.,window='boxcar',nfft=None,return_onesided=True,scaling='spectrum')
Pxx_n = Pxx/np.sum(Pxx)
## Use Hanning Window
f_h,Pxx_h = signal.periodogram(data,fs=1.,window='hanning',nfft=None,return_onesided=True,scaling='spectrum')
Pxx_h_n = Pxx_h/np.sum(Pxx_h)
fig = plt.figure(figsize=(15,6))
plt.plot(f,Pxx_n, color='C0', lw=1.3, label='Boxcar')
plt.plot(f_h,Pxx_h_n, color='C1', lw=1.3, label='Hanning')
plt.plot(freq,red_fit_n, color='firebrick', lw=1, alpha=0.9, label='Red Noise Fit')
plt.plot(freq,f_bounds, color='firebrick', lw=1, ls='--', alpha=0.9, label=str(int(alpha*100))+'% CI')
plt.ylim(0.0,0.3)
plt.xlim(0.0,0.06)
plt.xlabel("Frequency (approximately millennia$^{-1}$)",fontsize=14)
plt.ylabel("Normalized Power",fontsize=11)
plt.title("Normalized Power Spectrum of "+var_str,fontsize=14)
plt.legend(loc='upper right',fontsize=14)
plt.grid()
# -
# Use the Hanning Window AND change the length of the data window. <br>
#
# Tinker, try different window lengths, you should see the peaks broaden as window length shortens. Taking a small window decreases the sample, erodes confidence in the location of the peaks. But the smaller the window the more DOF and the higher the statistical significance.<br>
# Take home: window length is a tradeoff between accuracy and significance. Also beware of what is happening at the lowest frequencies (longest period waves) when you reduce the window_length... remember that your dataset is *only* 800,000 years long :)
# +
window_length=int(len(data)/4) #CHANGE WINDOW LENGTH HERE; divide by bigger/smaller numbers
print(f'window length: {window_length}')
f_h_wl,Pxx_h_wl = signal.periodogram(data,fs=1.,window='hanning',nfft=window_length,return_onesided=True,scaling='spectrum')
Pxx_h_wl_n = Pxx_h_wl/np.sum(Pxx_h_wl)
### Need to update the frequencies to plot, the DOF, the red noise fit and 99% confidence interval
### Note: Check that I am calculating the significance level appropriately for the window.
### Vineel/Elizabeth to double check!!
#EM: looks good to me.
freq_wl = np.arange(0.,(window_length/2)+1.)/window_length
#print(freq_wl) #EM: added these three lines to make sure that the above line is doing what I think it is- and it is
#freq_test=np.fft.rfftfreq(window_length,d=1.) #EM: these can be removed later
#print(freq_test)
red_fit_n_wl = create_normalized_redfit(window_length,Te) ## generate red_fit_n using window_length
print(np.size(red_fit_n_wl))
dof_wl=1.2*len(data)/(window_length/2) ### Barnes Eq. 26
print('DOF with window =',dof_wl)
print('alpha, statistical significance level',alpha)
f_bounds_wl = create_f_bounds(alpha,dof_wl,red_fit_n_wl) ## f-test for variance;
### plot results with data window of length window_length and compare to using the entire window
fig = plt.figure(figsize=(14,8))
gs=GridSpec(2,1)
plt.subplot(gs[0,0])
plt.plot(f_h_wl,Pxx_h_wl_n, color='C0', lw=1.3, label='Hanning,wl='+str(window_length))
plt.plot(freq_wl,red_fit_n_wl, color='firebrick', lw=1, alpha=0.9, label='Red Noise Fit')
plt.plot(freq_wl,f_bounds_wl, color='firebrick', lw=1, ls='--', alpha=0.9, label=str(int(alpha*100))+'% CI')
plt.ylim(0.0,0.5)
plt.xlim(0.0,0.06)
plt.xlabel("Frequency (approximately millennia$^{-1}$)",fontsize=14)
plt.ylabel("Normalized Power",fontsize=14)
plt.title("Normalized Power Spectrum: Hanning window using window length="+str(window_length)+",DOF="+str(dof_wl),fontsize=14)
plt.legend(loc='upper right',fontsize=14)
plt.grid()
fig.tight_layout()
plt.subplot(gs[1,0])
plt.plot(f_h,Pxx_h_n, color='C0', lw=1.3, label='Hanning')
plt.plot(freq,red_fit_n, color='firebrick', lw=1, alpha=0.9, label='Red Noise Fit')
plt.plot(freq,f_bounds, color='firebrick', lw=1, ls='--', alpha=0.9, label=str(int(alpha*100))+'% CI')
plt.ylim(0.0,0.5)
plt.xlim(0.0,0.06)
plt.xlabel("Frequency (approximately millennia$^{-1}$)",fontsize=14)
plt.ylabel("Normalized Power",fontsize=14)
plt.title("Normalized Power Spectrum: Hanning window with full data as window length="+str(len(data))+",DOF=2",fontsize=14)
plt.legend(loc='upper right',fontsize=14)
plt.grid()
fig.tight_layout()
###
# -
# Take-home message for 100,000ish year peak. If you reduce your window length too much -- your data chunks are too short to see the low frequency oscillations!
# Use the Hanning Window AND change the length of the data window AND use WOSA (Welch’s Overlapping Segment Analysis) <br>
# How does adding WOSA change the answer??
# +
window_length=int(len(data)/4) ##Here's where you change the window length if tinkering
T2 = window_length/2
freq_w = np.arange(0.,T2+1.)/window_length
#print(len(freq))
P_welch = signal.welch(data,window='hanning',nperseg=window_length); #Here's where Welch's method is changed
#print(np.size(P_welch))
P_welch_n = P_welch[1]/np.sum(P_welch[1]) #normalize
## Construct the expected rednoise spectrum - functions
## Need to update the 99% confidence interval - we have more DOF
red_fit_n_welch = create_normalized_redfit(window_length,Te) ## generate red_fit_n using window_length
print(np.size(red_fit_n_welch))
dof_welch=1.2*len(data)/(window_length/2) ### Barnes Eq. 26
print('DOF =',dof_welch)
f_bounds_welch = create_f_bounds(alpha,dof_welch,red_fit_n_welch) ## f-test for variance;
## plot
fig = plt.figure(figsize=(14,4))
plt.xlabel('Frequency')
plt.ylabel('Normalized Power')
plt.ylim(0.0,0.5)
plt.xlim(0.0,0.06)
plt.grid()
plt.plot(freq_w,P_welch_n, color='C0', lw=1.3, label='Hanning,wl='+str(window_length));
plt.plot(freq_w,red_fit_n_welch, color='firebrick', lw=1, alpha=0.9, label='Red Noise Fit');
plt.plot(freq_w,f_bounds_welch, color='firebrick', lw=1, ls='--', alpha=0.9, label=str(int(alpha*100))+'% CI');
plt.title("Normalized Power Spectrum: WOSA, Hanning window using window length="+str(window_length)+",DOF="+str(round(dof_welch,0)),fontsize=14);
# -
| aos575_applab4_fft_EPICA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Data Structures
#
# In this notebook, we'll explore different types of data structures that Python can use to store information, namely **lists, tuples, and dictionaries.**
# + [markdown] slideshow={"slide_type": "slide"}
#
# ## At the end of this notebook, you'll be able to:
# * Compare & contrast the types of structures that Python uses to store data points
# * Recognize & create lists, tuples, and dictionaries in Python
# * Index, slice, cast, and mutate lists
# * Understand the implications of mutability and object-oriented programming
#
# <hr>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Lists
# A _list_ is a mutable collection of ordered items, that can be of mixed type.
#
# **Mutable** means that individual items in the object can be changed. Lists are mutable. Tuples and strings are not -- they're **immutable**.
#
# Lists are created using square brackets `[ ]`, and individual elements are separated by commas.
# + slideshow={"slide_type": "fragment"}
# Create a list of fruits
# + [markdown] slideshow={"slide_type": "slide"}
# ### Useful list methods
# - Check the length of your list by using `len(my_list)`
# - Use `my_list.append()` to add elements to a list
# - Remove elements by index using `del my_list[index]`
# - Remove elements by value by using `my_list.remove('value')`
# - Sort by using `my_list.sort()`
# + slideshow={"slide_type": "fragment"}
# Try different list methods here
# + [markdown] slideshow={"slide_type": "slide"}
# ### List indexing & slicing
# **Indexing** refers to selecting an item from within a collection (e.g., lists, tuples, and strings). Indexing is done by placing the **index number** in square brackets, directly after the list variable.
#
# For example, if `my_list = [1,3,5]`, we can get the second value using `my_list[1]`. (Remember that Python starts indexing at zero!)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Reminders
# - Python is zero-based (The first index is '0')
# - Negative indices index backwards through a collection
# + slideshow={"slide_type": "slide"}
# Try indexing our list of fruits here
# + [markdown] slideshow={"slide_type": "slide"}
# ### If we want multiple items, we can **slice** the list.
#
# There are a few ways to slice:
#
# 1. We can **slice** a part of a list using the syntax `[start:stop]`, which extracts characters between index start and stop (-1).
#
# **Notes**
# - `start` is __included__ then every element __until__ `stop` is included.
# - Negative values count backwards through the list.
# + [markdown] slideshow={"slide_type": "slide"}
# 2. If we omit either (or both) of start or stop from `[start:stop]`, the default is the beginning and the end of the string, respectively, e.g. `[:3]`
# 3. We can also define the step size (instead of default 1) using the syntax `[start:stop:step]`
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-success"><b>Task:</b> For our list of fruits, create three different slices, and save them as different variables:
#
# 1. A slice of the first two fruits.
# 2. A slice of the middle three fruits.
# 3. A slice of the last fruit.
#
# </div>
# + slideshow={"slide_type": "fragment"}
# Your code here!
# + [markdown] slideshow={"slide_type": "slide"}
# ### Checking length
# We can use the function `len( )` to check the length of lists.
#
# **Note**: We can also use this to get the number of characters in a string!
# -
# + [markdown] slideshow={"slide_type": "slide"}
# ### Checking membership
# We can use `in` to see if an item exists in a list. The `in` operator checks whether an element is present in a collection, and can be negated with `not`. _(More on operators in the next lecture)_
# + slideshow={"slide_type": "fragment"}
# + [markdown] slideshow={"slide_type": "slide"}
# ### Mutating lists
# After definition, we can update members of our list _because lists are mutable!_ This also impacts aliases of our lists.
# + slideshow={"slide_type": "fragment"}
# + [markdown] slideshow={"slide_type": "slide"}
# ### Creating lists of lists
# Sometimes, it's useful to create lists of lists. Often, if we import big datasets as lists, this is how it will be organized.
#
# 
# + slideshow={"slide_type": "slide"}
gene_1 = ['gene1',0.48,0.55]
gene_2 = ['gene2',0.38,0.85]
gene_3 = ['gene3',0.21,0.81]
all_genes = [gene_1, gene_2, gene_3]
# We can use this syntax to get a specific value
print(all_genes[0])
# + [markdown] slideshow={"slide_type": "slide"}
# ## Tuples
# A _tuple_ is an **immutable** collection of ordered items, that can be of mixed type.
#
# * Tuples are created using parentheses.
# * Indexing works similar to lists.
# + slideshow={"slide_type": "fragment"}
# Define a tuple
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-success"><b>Question</b>: Before running the cell below, try to predict: What will be printed out from running this code?</div>
# + slideshow={"slide_type": "fragment"}
lst = ['a', 'b', 'c']
tup = ('b', 'c', 'd')
if lst[-1] == tup[-1]:
print('EndMatch')
elif tup[1] in lst:
print('Overlap')
elif len(lst) == tup:
print('Length')
else:
print('None')
# + [markdown] slideshow={"slide_type": "slide"}
# ### Casting between variable types
# We can use `list( )` or `tuple( )` to convert variables into different types. This is called **casting**.
#
# This is particularly useful when we use an operator like `range( )` which generates a range, but in the form of an **iterator**.
# + [markdown] slideshow={"slide_type": "slide"}
# **Note**: `range`, like indexing, is defined with `start`,`stop`, and `step`, but commas in between each. Remember that you can always use `?range` or `help(range)` to get details on how a function works.
# + slideshow={"slide_type": "fragment"}
# Test range here
# + [markdown] slideshow={"slide_type": "slide"}
# # Dictionaries
# Dictionaries are also like lists, except that each element is a key-value pair. The syntax for dictionaries is `{key1 : value1, ...}:`
#
# ### When dictionaries are useful
# 1. Flexible & efficient way to associate labels with heterogeneous data
# 2. Use where data items have, or can be given, labels
# 3. Appropriate for collecting data of different kinds (e.g., name, addresses, ages)
#
# > In the cell below, create a dictionary for three countries and capitals using the syntax `{country:capital,...}`. Remember that strings still need parentheses!
#
# **Note**: You can also create an empty dicitionary using `{}` and fill it using `dictionary['key'] = 'value'`.
# + slideshow={"slide_type": "fragment"}
capitals = {'United Kingdom':'London','France':'Paris','Spain':'Madrid'}
capitals
# + [markdown] slideshow={"slide_type": "slide"}
# <div class="alert alert-success"><b>Question:</b> Before running the cell below, predict: What would the following code produce?</div>
# + slideshow={"slide_type": "fragment"}
capitals.update({'United Kingdom':'England'})
capitals
# + [markdown] slideshow={"slide_type": "fragment"}
# <div class="alert alert-success"><b>Task</b>: What happens if we look for a key that doesn't exist? Try this above.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Additional dictionary functionality
# - Use `capitals.update(morecapitals)` to add another dictionary entry
# - Use `del capitals['US']` to delete entries
# - Loop by key or values, or both
# + [markdown] slideshow={"slide_type": "slide"}
# <hr>
#
# ## Additional resources
# <a href="https://swcarpentry.github.io/python-novice-gapminder/11-lists/index.html">Software Carpentries Lists</a>
#
# <a href="https://python101.pythonlibrary.org/chapter3_lists_dicts.html">Python 101: Lists, Tuples, and Dictionaries</a>
#
# <a href="https://github.com/jakevdp/WhirlwindTourOfPython/blob/6f1daf714fe52a8dde6a288674ba46a7feed8816/06-Built-in-Data-Structures.ipynb">Whirlwind Tour of Python: Built-In Data Structures</a>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## About this notebook
# This notebook is largely derived from UCSD COGS18 Materials, created by <NAME> & <NAME>, as well as the <a href="https://github.com/jrjohansson/scientific-python-lectures/blob/master/Lecture-1-Introduction-to-Python-Programming.ipynb">Scientific Python Lecture</a> by <NAME>.
#
# Want to run this notebook as a slideshow? If you have Python (or Anaconda) follow <a href="http://www.blog.pythonlibrary.org/2018/09/25/creating-presentations-with-jupyter-notebook/">these instructions</a> to setup your computer with the RISE plugin.
| 04-DataStructures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
# -
# Load Data
train_data = pd.read_csv("/kaggle/input/robi-datathon-2-pre-assessment/train.csv")
test_data = pd.read_csv("/kaggle/input/robi-datathon-2-pre-assessment/test.csv")
# submission file
submission_data = pd.read_csv("/kaggle/input/robi-datathon-2-pre-assessment/sample_submission.csv")
train_data.head(2)
# <b> Data Cleaning
train_data.isna().sum()
print(f"Train dataset shape: {train_data.shape}")
print(f"Test dataset shape: {test_data.shape}")
# Drop null columns
train_data = train_data.drop(['s56','s57','s59'],axis=1)
train_data.shape
train_data.columns.values
train_data.info()
# +
# for colm in train_data:
# print(train_data[colm].unique())
# -
train_data.describe(include=["object"])
# <b> Visualization
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# %config InlineBackend.figure_format = 'retina'
sns.countplot(x="s16", hue="gender", data=train_data);
# Feature Engineering
numerical_feat = train_data.select_dtypes(include=np.number).columns
print('Numerical columns:',numerical_feat)
# +
categorical_feat = train_data.select_dtypes(include='object').columns
print('Categorical columns:',categorical_feat)
# -
train_data.label.value_counts()
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# train_data.label.value_counts().plot.bar()
# plt.show()
# -
for c in categorical_feat:
if len(train_data[c].value_counts()) < 100:
print(c)
train_data[c].value_counts().plot.bar()
plt.show()
for c in numerical_feat:
print(c)
train_data[c].plot.hist(bins=5)
plt.show()
for c in numerical_feat:
print(c)
#print('min:', train_data[c].min(), 'max:', train_data[c].max())
dropIndexes = train_data[train_data[c] > train_data[c].max()*9/10].index
train_data.drop(dropIndexes , inplace=True)
for c in numerical_feat:
print(c)
print(train_data[c].value_counts(bins=2, sort=False))
plt.show()
for c in numerical_feat:
print(c)
train_data[c].plot.hist(bins=100)
plt.show()
fig, axes = plt.subplots(len(numerical_feat), len(numerical_feat), figsize=(16, 16), sharex=False, sharey=False)
for i in range(0,len(numerical_feat)):
for j in range(0,len(numerical_feat)):
axes[i,j].scatter(x = train_data[numerical_feat[i]], y = train_data[numerical_feat[j]])
fig.tight_layout()
# Feature Selection
target = 'label'
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
numerical_features = ['s13', 's48', 'n1', 'n2', 'n3', 'n4', 'n5', 'n6', 'n7', 'n8', 'n9',
'n10', 'n11', 'n12', 'n13', 'n14', 'n15']
categorical_features = ['id', 'gender', 's11', 's12', 's16', 's17', 's18', 's52', 's53', 's54',
's55', 's58', 's69', 's70', 's71']
model_features = numerical_features + categorical_features
# Get train data to train the pipeline
X_train = train_data[model_features]
y_train = train_data[target]
# Maximum number of decision trees. The effective number of trained trees can be smaller if early stopping is enabled.
NUM_TREES = 250
# Minimum number of examples in a node.
MIN_EXAMPLES = 6
# Maximum depth of the tree. max_depth=1 means that all trees will be roots.
MAX_DEPTH = 5
# Ratio of the dataset (sampling without replacement) used to train individual trees for the random sampling method.
SUBSAMPLE = 0.65
# Control the sampling of the datasets used to train individual trees.
SAMPLING_METHOD = "RANDOM"
# Ratio of the training dataset used to monitor the training. Require to be >0 if early stopping is enabled.
VALIDATION_RATIO = 0.1
import tensorflow as tf
ann = tf.keras.models.Sequential()
ann.add(tf.keras.layers.Dense(units=6, activation='relu'))
ann.add(tf.keras.layers.Dense(units=6, activation='relu'))
ann.add(tf.keras.layers.Dense(units=1, activation='sigmoid'))
ann.compile(optimizer='adam' , loss='binary_crossentropy' , metrics=['accuracy'] )
ann.fit(X_train, y_train, batch_size=32, epochs=100)
| core/ml/pre-datathon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Pipeline
# - 직접 코드를 하나씩 실행하면 혼돈이 생긴다
# - 데이터 전처리와 여러 알고리즘의 반복시행, 하이퍼파라미터 튜닝과정을 번갈아 하다보면 코드의 실행 순서에 혼돈이 생길 수 있다
# - 이런 경우 class로 만들어서 진행하면 좋지만, sklearn 유저는 꼭 그럴 필요 없이 pipeline 사용하면 됨
# 와인데이터
# +
import pandas as pd
red_url = 'https://raw.githubusercontent.com/PinkWink/ML_tutorial/master/dataset/winequality-red.csv'
white_url = 'https://raw.githubusercontent.com/PinkWink/ML_tutorial/master/dataset/winequality-white.csv'
red_wine = pd.read_csv(red_url, sep=';')
white_wine = pd.read_csv(white_url, sep=';')
red_wine['color'] = 1.
white_wine['color']= 0.
wine = pd.concat([red_wine, white_wine])
# -
x = wine.drop(['color'], axis=1)
y = wine['color']
# - **Pipeline 구현**
# +
from sklearn.pipeline import Pipeline
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import StandardScaler
estimators = [('scaler', StandardScaler()),
('clf', DecisionTreeClassifier())]
pipe = Pipeline(estimators)
# -
pipe.steps
pipe.steps[0]
pipe.steps[1]
# - 스텝별로 객체 호출
pipe[0]
pipe['scaler']
# - set_params
# - step이름 + __ (언더바 2개) + 속성이름 = 속성값
#
pipe.set_params(clf__max_depth=2)
pipe.set_params(clf__random_state=13)
# - pipeline을 이용한 분류기 구성
# +
from sklearn.model_selection import train_test_split
x_scaled = pipe['scaler'].fit_transform(x)
x_train, x_test, y_train, y_test = train_test_split(x_scaled, y, test_size=0.2, random_state=13, stratify=y)
pipe.fit(x_train, y_train)
# -
# - 성과
# +
from sklearn.metrics import accuracy_score
y_pred_tr = pipe.predict(x_train)
y_pred_test = pipe.predict(x_test)
print('Train Acc : ', accuracy_score(y_train, y_pred_tr))
print('Test Acc : ', accuracy_score(y_test, y_pred_test))
# -
# - 모델 구조 확인
# +
from graphviz import Source
from sklearn.tree import export_graphviz
Source(export_graphviz(pipe['clf'], feature_names=x.columns, class_names=['W', 'R'], rounded=True, filled=True))
| machine_learning/datascienceschool/0320_pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
"""
Get data of publications (year) for each author on fictionDB.com by selenium.
functions used:
search(author) from fictiondb.py
Added more data on 1/20/2021
Output to pickle 'author_book_df'.
"""
# +
from bs4 import BeautifulSoup
import requests
import time, os
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.common.by import By
from selenium.webdriver.support import expected_conditions as EC
from selenium.webdriver.common.action_chains import ActionChains
import pandas as pd
import numpy as np
import sys
sys.path.append('/Users/katiehuang/Documents/metis/projects/onl_ds5_project_2/py')
from scrape import *
import importlib
from fictiondb import *
# -
chromedriver = "/Applications/chromedriver" # path to the chromedriver executable
os.environ["webdriver.chrome.driver"] = chromedriver
# Open automated Chrome with fictionDB webpage
driver = webdriver.Chrome(chromedriver)
driver.get("https://www.fictiondb.com/search/search.htm")
# +
# Find books survived in the merge and create author list to search on fictionDB
# Load 'movie_data','book_data', 'found_book' and merge/concat
movie_df = pd.read_pickle('../data/movie_data')
book_df = pd.read_pickle('../data/book_data')
found_book_df = pd.read_pickle('../data/found_book')
overlap_df = pd.merge(book_df, movie_df,left_on='book_title',\
right_on='movie_title', how='inner')
overlap_df2 = pd.merge(found_book_df, movie_df,left_on='book_title',\
right_on='movie_title', how='inner')
combined_book_df = pd.concat([overlap_df,overlap_df2])
# -
combined_book_df.head(3)
# Create list of author and remove duplicates by converting to set
author_list = list(combined_book_df.author)
author_list_set = set(author_list)
len(author_list_set)
# Save the list of unique author
import pickle
with open('../dump/author_list_set', 'wb') as f:
pickle.dump(author_list_set, f)
import pandas as pd
author_list_set = pd.read_pickle('../dump/author_list_set')
# # Open automated Chrome with fictionDB webpage
driver = webdriver.Chrome(chromedriver)
driver.get("https://www.fictiondb.com/search/search.htm")
c = 0
author_book_list=[]
for author in author_list_set:
c += 1
print(c,author)
result = search(author)
author_book_list.append(result)
# Save to pickle
import pickle
with open('../dump/author_book_list', 'wb') as f:
pickle.dump(author_book_list, f)
import pandas as pd
author_book_list = pd.read_pickle('../dump/author_book_list')
# +
# Ran into error, search for the rest of the authors
c = len(author_book_list)
done_author=[]
for dic in author_book_list:
for key, value in dic.items():
print(key)
done_author.append(key)
# authors still need to be looked up
left_author_set = author_list_set.difference(set(done_author))
# -
# %run -i '../py/fictiondb.py'
for author in left_author_set:
c += 1
print(c,author)
result = search(author)
author_book_list.append(result)
len(author_book_list)
# Save to pickle
import pickle
with open('../dump/author_book_list', 'wb') as f:
pickle.dump(author_book_list, f)
# Create DataFrame of author and publication years and pickle
author_book_df = pd.DataFrame(author_book_list_test)
author_book_df.to_pickle('../dump/author_book_data')
# ### Increase author_book data (1/20/2021)
import pandas as pd
# 1/20/2021
# Load data from the second scrape of book data
author_book_df1 = pd.read_pickle('../dump/author_book_data')
author_book_df2 = pd.read_pickle('../dump/author_book_data_add')
# Merge two author_book_df2
author_book_df = pd.concat([author_book_df1,author_book_df2])
author_book_df.to_pickle('../data/author_book_data')
author_book_df.head()
| notebooks/06_scrape_fictiondb.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// <a name="top"></a><img src="images/chisel_1024.png" alt="Chisel logo" style="width:480px;" />
// # Module 2.5: Putting it all Together: An FIR Filter
// **Prev: [Sequential Logic](2.4_sequential_logic.ipynb)**<br>
// **Next: [Generators: Parameters](3.1_parameters.ipynb)**
//
// ## Motivation
// Now that you've learned the basics of Chisel, let's use that knowledge to build a FIR (finite impulse response) filter module! FIR filters are very common in digital signal processing applications. Also, the FIR filter will reappear frequently in module 3, so it's important that you don't filter out this module by skipping ahead! If you are unfamiliar with FIR filters, head over to the article on [trusty Wikipedia](https://en.wikipedia.org/wiki/Finite_impulse_response) to learn more.
//
// ## Setup
val path = System.getProperty("user.dir") + "/source/load-ivy.sc"
interp.load.module(ammonite.ops.Path(java.nio.file.FileSystems.getDefault().getPath(path)))
import chisel3._
import chisel3.util._
import chisel3.iotesters.{ChiselFlatSpec, Driver, PeekPokeTester}
// + [markdown] nbpresent={"id": "2e849671-a1e9-48b2-9bba-eb916ac623c6"}
// ---
// # FIR Filter
//
// The FIR filter you will design performs the following operation.
//
// <img src="images/fir.jpg" width="720">
//
// Basically, this does a elementwise multiplication of the element of the filter coefficients with the elements of the input signal and outputs the sum (also called a _convolution_).
//
// Or, a signals definition:
//
// $y[n] = b_0 x[n] + b_1 x[n-1] + b_2 x[n-2] + ...$
// - $y[n]$ is the output signal at time $n$
// - $x[n]$ is the input signal
// - $b_i$ are the filter coefficients or impulse response
// - $n-1$, $n-2$, ... are time $n$ delayed by 1, 2, ... cycles
//
// ## 8-bit Specification
//
// Build a 4-element FIR filter where the four filter coefficients are parameters. A module skeleton and basic tests are provided for you.
// Note that both the input and output are 8-bit unsigned integers. You will need to save necessary state (like delayed signal values) using constructs like shift registers. Use the provided testers to check your implementation.
// Registers with constant inputs can be created using a `ShiftRegister` of shift value 1, or by using the `RegNext` construct.
//
// Note: for the tests to pass, your registers must be initialized to `0.U`.
// + nbpresent={"id": "26e4a686-0397-4306-985c-813909256c95"}
class My4ElementFir(b0: Int, b1: Int, b2: Int, b3: Int) extends Module {
val io = IO(new Bundle {
val in = Input(UInt(8.W))
val out = Output(UInt(8.W))
})
// Don't forget to initialize to 0!
val delayed_1 = RegNext(io.in, 0.U)
val delayed_2 = RegNext(delayed_1, 0.U)
val delayed_3 = RegNext(delayed_2, 0.U)
io.out := (io.in * b0.asUInt(8.W)) + (delayed_1 * b1.asUInt(8.W)) + (delayed_2 * b2.asUInt(8.W)) + (delayed_3 * b3.asUInt(8.W))
}
// + nbpresent={"id": "ddf24b7b-09a2-46f0-b1d8-cb2ca7976b4b"}
// Simple sanity check: a element with all zero coefficients should always produce zero
Driver(() => new My4ElementFir(0, 0, 0, 0)) {
c => new PeekPokeTester(c) {
poke(c.io.in, 0)
expect(c.io.out, 0)
step(1)
poke(c.io.in, 4)
expect(c.io.out, 0)
step(1)
poke(c.io.in, 5)
expect(c.io.out, 0)
step(1)
poke(c.io.in, 2)
expect(c.io.out, 0)
}
}
// -
// Simple 4-point moving average
Driver(() => new My4ElementFir(1, 1, 1, 1)) {
c => new PeekPokeTester(c) {
poke(c.io.in, 1)
expect(c.io.out, 1) // 1, 0, 0, 0
step(1)
poke(c.io.in, 4)
expect(c.io.out, 5) // 4, 1, 0, 0
step(1)
poke(c.io.in, 3)
expect(c.io.out, 8) // 3, 4, 1, 0
step(1)
poke(c.io.in, 2)
expect(c.io.out, 10) // 2, 3, 4, 1
step(1)
poke(c.io.in, 7)
expect(c.io.out, 16) // 7, 2, 3, 4
step(1)
poke(c.io.in, 0)
expect(c.io.out, 12) // 0, 7, 2, 3
}
}
// Nonsymmetric filter
Driver(() => new My4ElementFir(1, 2, 3, 4)) {
c => new PeekPokeTester(c) {
poke(c.io.in, 1)
expect(c.io.out, 1) // 1*1, 0*2, 0*3, 0*4
step(1)
poke(c.io.in, 4)
expect(c.io.out, 6) // 4*1, 1*2, 0*3, 0*4
step(1)
poke(c.io.in, 3)
expect(c.io.out, 14) // 3*1, 4*2, 1*3, 0*4
step(1)
poke(c.io.in, 2)
expect(c.io.out, 24) // 2*1, 3*2, 4*3, 1*4
step(1)
poke(c.io.in, 7)
expect(c.io.out, 36) // 7*1, 2*2, 3*3, 4*4
step(1)
poke(c.io.in, 0)
expect(c.io.out, 32) // 0*1, 7*2, 2*3, 3*4
}
}
// <div id="container"><section id="accordion"><div>
// <input type="checkbox" id="check-1" />
// <label for="check-1"><strong>Solution</strong></label>
// <article>
// <pre style="background-color:#f7f7f7">
// val x_n1 = RegNext(io.in, 0.U)
// val x_n2 = RegNext(x_n1, 0.U)
// val x_n3 = RegNext(x_n2, 0.U)
// io.out := io.in \* b0.U(8.W) +
// x_n1 \* b1.U(8.W) +
// x_n2 \* b2.U(8.W) +
// x_n3 \* b3.U(8.W)
// </pre></article></div></section></div>
// ---
// # IPXact Example
//
// Your Chisel generator produces Verilog, but just handing someone else a piece of Verilog is insufficient to verify and integrate the design. What other information is needed to capture a design’s intent? IPXact solves this by providing an XML description of a design and its metadata, including the interfaces, parameters, address mapping, etc. So for this portion of the lab, we'll standardize the FIR interfaces and write out IPXact to ease implementation and verification. You'll need the results of this generator in later labs.
//
// ## Setup
// We have compiled the necessary depedencies, which are not all published on Maven, into a jar file. Contact the contributors if you want a copy. If you are not at the bootcamp, you can safely skip this section.
// depdencies for ipxact
val path = System.getProperty("user.dir") + "/../rocket-dsp-utils-assembly-1.0.jar"
interp.load.cp(ammonite.ops.Path(java.nio.file.FileSystems.getDefault().getPath(path)))
// ---
// # FIR Filter Generator
//
// For this module, we'll be using a slightly modified example from [Module 3.2: Generators: Collection](3.2_collections.ipynb).
// If you haven't started Module 3.2, don't worry.
// You'll learn about the details of how `MyManyDynamicElementVecFir` works, but the basic idea is that it is a FIR filter generator.
//
// The generator has one parameter: length.
// That parameter dictates how many taps the filter has, and the taps are inputs to the hardware `Module`.
//
// The generator has 3 inputs:
// * in, the input to the filter
// * valid, a boolean that says when the input is valid
// * consts, a vector for all the taps
//
// and 1 output:
// * out, the filtered input
//
// <img src="images/fir.jpg" style="width:450px;"/>
class MyManyDynamicElementVecFir(length: Int) extends Module {
val io = IO(new Bundle {
val in = Input(UInt(8.W))
val valid = Input(Bool())
val out = Output(UInt(8.W))
val consts = Input(Vec(length, UInt(8.W)))
})
// Such concision! You'll learn what all this means later.
val taps = Seq(io.in) ++ Seq.fill(io.consts.length - 1)(RegInit(0.U(8.W)))
taps.zip(taps.tail).foreach { case (a, b) => when (io.valid) { b := a } }
io.out := taps.zip(io.consts).map { case (a, b) => a * b }.reduce(_ + _)
}
// ---
// # DspBlock
//
// Integrating DSP components into a larger system can be challenging and error prone.
// The [rocket-dsp-utils](https://github.com/ucb-art/rocket-dsp-utils) repository consists of useful generators that should help with such tasks.
//
// One of the core abstractions is the notion of a DSPBlock.
// A DSPBlock has:
// * AXI-4 Stream input and output
// * AXI-4 memory-mapped status and control
//
//
// <img src="images/fir_filter.png" style="width:800px;"/>
//
// The following code wraps the FIR filter in AXI4 interfaces.
// +
import dspblocks._
import freechips.rocketchip.amba.axi4._
import freechips.rocketchip.amba.axi4stream._
import freechips.rocketchip.config._
import freechips.rocketchip.diplomacy._
case object NumFilters extends CSRField {
val name = "firQueueDepth"
}
case class FIRTap(filterIdx: Int, tapIdx: Int) extends CSRField {
val name = s"firTap${filterIdx}_$tapIdx"
}
class FIRBlock(nFilters: Int, nTaps: Int)(implicit p: Parameters) extends
AXI4DspBlock with AXI4HasCSR with HasIPXactParameters {
outer =>
addStatus(NumFilters)
for (i <- 0 until nFilters) {
for (j <- 0 until nTaps) {
addControl(FIRTap(i, j))
}
}
makeCSRs()
val streamNode = AXI4StreamIdentityNode()
override def ipxactParameters = Map(
"nFilters" -> nFilters.toString,
"nTaps" -> nTaps.toString
)
def beatBytes: Int = 8
def csrAddress = freechips.rocketchip.diplomacy.AddressSet(0x0, 0xffffL)
def csrBase: Int = 0
def csrSize: Int = 4096
lazy val module = new LazyModuleImp(this) {
val (in, _) = streamNode.in(0)
val (out, _) = streamNode.out(0)
val mem = outer.mem.map { m => m.in(0) }
require(in.params.n >= nFilters,
s"""AXI-4 Stream port must be big enough for all
|the filters (need $nFilters,, only have ${in.params.n})""".stripMargin)
status(NumFilters) := nFilters.U
val outs = (0 until nFilters).map(i => {
val fir = Module(new MyManyDynamicElementVecFir(nTaps))
fir.io.in := in.bits.data((i+1)*8, i*8)
fir.io.valid := in.valid && out.ready
for (j <- 0 until nTaps) {
fir.io.consts(j) := control(FIRTap(i, j))
}
fir.io.out
})
val output = if (outs.length == 1) {
outs(0)
} else {
outs.reduce((x: UInt, y: UInt) => Cat(y, x))
}
out.bits.data := output
in.ready := out.ready
out.valid := in.valid
}
}
// -
// ---
// # Invoking the Generator
//
// The following code invokes the generator.
// It will produce a file called `BlindModule.fir` which contains the firrtl generated by our code, as well as a file called `BlindModule.v` which is the compiled verilog.
// `BlindModule` is a wrapper for our `DspBlock`- it is necessary because of how [diplomacy](https://carrv.github.io/papers/cook-diplomacy-carrv2017.pdf) in rocket works.
// You'll notice that `BlindModule` instantiates an `FIRBlock`.
//
// The line
// ```scala
// AXI4StreamBundleParameters(n = 8)
// ```
// sets the AXI-4 streams' data widths to 8 bytes.
object FIRGenerator extends ipxact.IPXactGeneratorApp with App {
override val verilogFilename: String = "BlindModule.v"
override val ipxactDir: String = "./"
implicit val p: Parameters = Parameters.root((new freechips.rocketchip.coreplex.BaseCoreplexConfig).toInstance)
val blindNodes = DspBlockBlindNodes.apply(
AXI4StreamBundleParameters(n = 8),
() => AXI4MasterNode(Seq(AXI4MasterPortParameters(Seq(AXI4MasterParameters("fir"))))))
val dut = () => {
val lazyMod = LazyModule(DspBlock.blindWrapper(() => new FIRBlock(4, 8), blindNodes))
val m = lazyMod.module
IPXactComponents._ipxactComponents += DspIPXact.makeDspBlockComponent(lazyMod.internal)
m
}
chisel3.Driver.dumpFirrtl(chisel3.Driver.elaborate(dut), None)
chisel3.Driver.compileFirrtlToVerilog("BlindModule", new java.io.File(System.getProperty("user.dir")))
generateIPXact(IPXactComponents.ipxactComponents())
}
FIRGenerator.main(Array[String]())
// ---
// # IPXact
// Look in the file browser for a file called `craft_BlindModule_edu.berkeley.cs_1.0.xml`.
// This is the ipxact file.
// It contains information about:
// * Port mappings
// * Interfaces
// * Memory maps
// * Generator parameters
//
// You'll notice that the parameters from the scala code
// ```scala
// override def ipxactParameters = Map(
// "nFilters" -> nFilters.toString,
// "nTaps" -> nTaps.toString
// )
// ```
// appears in the IPXact output.
// This gives verification generators the information they need to generate appropriate test vectors for the given instance.
// ---
// # You're done!
//
// [Return to the top.](#top)
| 2.5_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
counter_fail = 0
total_attempts = 3
category_selection = []
selection = True
while selection:
choice = input("Do you want to play Hangman?\nYes = Press Y / No = Press N for other games ").lower()
if choice == 'y' or choice == 'yes':
print(" ")
print('GAME RULES: 3 maximum fail attempts AND letters from a to z')
print(" ")
choice_category = input("Select a category:\nfor cities:(Press c) for states:(Press s), for anything(Press a) ").lower()
if choice_category == 'c':
print(" ")
print("You are playing cities, Good Luck")
category_selection = ["delhi","beirut","berlin"]
if choice_category == 's':
print(" ")
print("You are playing States, Good Luck")
category_selection = ["florida","idaho","texas"]
if choice_category == 'a':
print(" ")
print("You are playing anything, Good Luck")
category_selection = ["iphone","computer","netflix"]
secret_word = list(random.choice(category_selection))
len_words_options = int(len(secret_word))
found_letters = ["_"] * len_words_options
found_letters[0] = secret_word[0]
found_letters[-1] = secret_word[-1]
print(" ")
print("Guess the secret word", " ".join(found_letters))
while counter_fail < total_attempts:
if counter_fail== 1:
print("1 fail - 2 Attempts remaining")
print(" + ---+\n | |")
print(" ")
if counter_fail== 2:
print("2 fail - 1 Attempt remaining")
print(" + ---+ \n | | \n O | \n / |\ | \n")
guess_letter = input("insert a letter: ").lower()
if guess_letter not in secret_word:
print('Letter not in secret word')
counter_fail += 1
print(" ".join(found_letters))
print(" ")
if guess_letter in secret_word:
print('Congratulation,you found a letter ')
index_letter = secret_word.index(guess_letter)
found_letters[index_letter] = guess_letter
print(" ".join(found_letters))
print(" ")
if secret_word == found_letters:
print('Congratulation you guessed the secret word ')
break
else:
print('3 fail attempts reached - You did not guess the secrete word')
print(" + ---+\n | |\n O |\n / |\ |\n / \ |\n |\n=========\n")
print('The word was ' + "".join(secret_word).upper())
elif choice == 'n' or choice == 'no':
print("\n")
print("Ask Sky or Denisse, they have more games")
else:
print("Option not recognized")
| module-1/Lambda-Functions/your-code/Leandro's Game.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="OC5nJawdBX97" colab_type="text"
# **Python string methods**
#
#
#
# + [markdown] id="4JrmwkI58SaI" colab_type="text"
# **capitalize()**
# + id="ElwkWucW7Ogj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="7d104639-44f9-4077-e758-f4e99293c6a4"
""" capitalize() method capitalizes the first character of the string """
string1 = "<NAME>"
print(string1.capitalize())
# + [markdown] id="VB3KA6yQ9Kof" colab_type="text"
# **casefold()**
# + id="YowtjkJ18l9M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="b98c7310-9f85-4052-8c07-a9a8d3ee9a9e"
""" casefold() method converts the string into lowercase """
string2 = "<NAME> Adnan"
print(string2.casefold())
# + [markdown] id="WRRZZlfi93jL" colab_type="text"
# **center()**
# + id="iYQTvbA69r8H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="747f10aa-0b56-44e7-e00c-97e084686810"
""" center() method returns a centered string, padded with specific character(optional) """
string3 = "center"
# Without specific character
print(string3.center(15))
# With specific character for padding, here '1' is used for padding
print(string3.center(15,'1'))
# + [markdown] id="K4IKD0ATRQef" colab_type="text"
# **count()**
# + id="JEexWNpZQIjB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="7d57db39-e606-4f15-ec4d-58347dc90452"
"""count() method returns the number of count of the mentioned substring in a string"""
string4 = "so so is not so good"
substring = "so"
print(string4.count(substring))
# We can optionally mention the start and end point
print(string4.count(substring, 0, 8))
# + [markdown] id="45giVdarS01I" colab_type="text"
# **endswith()**
# + id="QdVR9QnrSBsN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="fe84fca1-74fc-47d5-bc16-31288985ff96"
"""endswith() is a binary method which returns True if the specified value is present at the end"""
string5 = "May the force be with you"
substring = "you"
print(string5.endswith(substring))
# We can optionally mention the start and end point
print(string5.endswith(substring, 0, 8))
# + [markdown] id="nkMInVuVVpnq" colab_type="text"
# **expandtabs()**
# + id="KrYCmwPAUZMq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="56e80e2f-2280-4c11-84d6-e262daaf6854"
"""expandtabs() method returns the string with specified tabsize or whitespace"""
# \t denotes tab, by deafult tab size is set to 8
string6 = "A\td\tn\ta\tn"
print(string6.expandtabs())
# Setting the tab size to 12
print(string6.expandtabs(12))
# + [markdown] id="G_BmlOU3Wszt" colab_type="text"
# **find()**
# + id="GTJRXP0uWVBE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="93e1e8f2-527e-4a85-ca2d-d7cc88ccdb0b"
"""find() method searches for a specific value in the string, and returns it's
first occurance index, if no match found it returns -1"""
string7 = "If I have seen further it is by standing on the shoulders of Giants"
print(string7.find("Giants"))
# We can optionally mention the start and end point
print(string7.find("Giants",1,50))
# + [markdown] id="vo7m4obmcqWR" colab_type="text"
# **format()**
# + id="Zh0EV05aYQsY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="82d65acd-b007-4dd0-bc5a-3f5e58091159"
"""The format() method returns a formatted string"""
string8 = "I am {}.I love {}"
print(string8.format("Adnan","reading books"))
# + [markdown] id="kdOLvxK0fhmV" colab_type="text"
# **isalnum()**
# + id="_WkCTXkfeqde" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="eefabe64-af80-4c02-faf1-e219e3967c6c"
""" isalnum() is a boolean method that returns true if the string contains alphanumeric values only """
string9 = "se7en"
print(string9.isalnum())
# + [markdown] id="pXR4o6IIgIX5" colab_type="text"
# **isalpha()**
# + id="RZSX0G1qgHvO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="1ef1e07c-5dd7-4716-baf6-5dcefe719ce9"
""" isalpha() is a boolean method that returns true if the string contains alphabet only """
string10 = "Awesome"
print(string10.isalpha())
# + [markdown] id="SQxtAh2Hg1TF" colab_type="text"
# **isnumeric()**
# + id="-Ey_xRBvgBWF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="705e559e-b50b-4127-b84c-65ccf217aa37"
""" isnumeric() is a boolean method that returns true if the string contains numeric character only """
string11 = "2441139"
print(string11.isnumeric())
# + [markdown] id="EvqwimFWh_J5" colab_type="text"
# **isupper()**
# + id="9YTA5xDZh2to" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="89e5bb72-5623-4259-f682-2080caae29bb"
""" isupper() is a boolean method that returns true if the string contains uppercase characters only """
string12 = "GITHUB"
print(string12.isupper())
# + [markdown] id="jN6NJcvAid5w" colab_type="text"
# **islower()**
# + id="ycjTvOfMia65" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="71011d47-1f8b-4d6d-88c9-161bc1e071ef"
""" islower() is a boolean method that returns true if the string contains lowercase characters only """
string13 = "octocat"
print(string13.islower())
# + [markdown] id="okYdE33kwZ1c" colab_type="text"
# **upper()**
# + id="YZ_9v91gi2xK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="a248dd5d-d266-4fd8-87ba-c14119129ac3"
""" upper() method converts a string into uppercase """
string14 = "The higher you rise,the harder you fall"
print(string14.upper())
# + [markdown] id="C7OXiDYUy5LN" colab_type="text"
# **lower()**
# + id="ZP67QoL5y3Iy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="796a1aa0-3b92-467b-b966-2704678a4469"
""" lower() method converts a string into lowercase """
string15 = "THE HIGHER YOU RISE,THE HARDER YOU FALL"
print(string15.lower())
# + [markdown] id="m72CPkzJz1ef" colab_type="text"
# **replace()**
# + id="A51KbM6xzLIZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="4175f730-aab3-4a09-a1bb-b1b362fc29dc"
""" In a string, replace() method replaces a substring with another one """
string16 = "Let's fly higher and higher"
print(string16.replace("higher","lower"))
# We can also mention the number of occurance we want to replace,
# here we replaced just the first occurance of 'higher'
print(string16.replace('higher', 'lower', 1))
# + [markdown] id="2nj2tu4J3ARx" colab_type="text"
# **split()**
# + id="YxY-z6x20W-O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="31048de6-b097-4f65-8f22-1aaa277d5a96"
""" split() method splits the string with the given parameters,
and retruns a list of strings.By default it splits using space """
string17 = "Divided we fall"
print(string17.split())
string17a = "comma,separated,string"
# Here we use comma as separator
print(string17a.split(','))
# We can also mention the maximum number of time split would occur (optional)
print(string17a.split(',',1))
# + [markdown] id="VIqwdDUc5B9L" colab_type="text"
# **strip()**
# + id="vt8U9I1s38kk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="c01b32b0-6a7a-43b8-bcbb-bfdcbc5c27a3"
""" strip() method returns the trimmed version of the string, by default it trims heading and trailing whitespace """
string18 = " we need space "
print(string18.strip())
string18a = "******Thus a star is born******"
# Here we trim the heading and trailing '*'
print(string18a.strip('*'))
# + [markdown] id="cq5ucPgE7b4k" colab_type="text"
# **swapcase()**
# + id="06fxlEqn5vpv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="5cecc189-8dbd-4cec-d7cc-743f7c197ea5"
""" swapcase() method converts uppercase into lowercase and vice-versa """
string19 = "nEVER wRITE lIKE tHIS"
print(string19.swapcase())
# + [markdown] id="zdTvKEkQ8rbj" colab_type="text"
# **zfill()**
# + id="RHEViQ9Z71Ex" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="c903e757-89f3-4062-e564-4c188fa16de8"
""" zfill() method adds zero at the begining of string, width of the returned
string is passed as parameter """
string20 = "is a number"
# Here the given string is of length 11, we passed 10 as the parameter of zfill,
# so it added one 0 at the left of the string
print(string20.zfill(12))
# If a string starts with + or - operator the zeros are filled after the 1st operator
string20a = "+is needed"
print(string20a.zfill(15))
string20b = "++is needed"
# Here zero is filled after the first '+' operator
print(string20b.zfill(15))
# + id="adCxloKXDTU7" colab_type="code" colab={}
| 4-assets/BOOKS/Jupyter-Notebooks/Overflow/String_methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GDP and life expectancy
#
# Richer countries can afford to invest more on healthcare, on work and road safety, and other measures that reduce mortality. On the other hand, richer countries may have less healthy lifestyles. Is there any relation between the wealth of a country and the life expectancy of its inhabitants?
#
# The following analysis checks whether there is any correlation between the total gross domestic product (GDP) of a country in 2013 and the life expectancy of people born in that country in 2013.
# ## Getting the data
#
# Two datasets of the World Bank are considered. One dataset, available at <http://data.worldbank.org/indicator/NY.GDP.MKTP.CD>, lists the GDP of the world's countries in current US dollars, for various years. The use of a common currency allows us to compare GDP values across countries. The other dataset, available at <http://data.worldbank.org/indicator/SP.DYN.LE00.IN>, lists the life expectancy of the world's countries. The datasets were downloaded as CSV files in March 2016.
# +
import warnings
warnings.simplefilter('ignore', FutureWarning)
import pandas as pd
YEAR = 2013
GDP_INDICATOR = 'NY.GDP.MKTP.CD'
gdpReset = pd.read_csv('WB GDP 2013.csv')
LIFE_INDICATOR = 'SP.DYN.LE00.IN'
lifeReset = pd.read_csv('WB LE 2013.csv')
lifeReset.head()
# -
# ## Cleaning the data
#
# While inspecting the data with `head()` and `tail()` shows that, it was observed that the first 34 rows are aggregated data, that is they are country groups. Individual countries starts in row number 34
# It was observed that GDP and life expectancy values are missing for some countries.
#
# Therefore wwe have to clean the data by removing the first 34 rows and removing rows with unavailable values.
gdpCountries = gdpReset[34:].dropna()
lifeCountries = lifeReset[34:].dropna()
# ## Transforming the data
#
# The World Bank reports GDP in US dollars and cents. To make the data easier to read, the GDP is converted to millions of British pounds (the author's local currency) with the following auxiliary functions, using the average 2013 dollar-to-pound conversion rate provided by <http://www.ukforex.co.uk/forex-tools/historical-rate-tools/yearly-average-rates>.
# +
def roundToMillions (value):
return round(value / 1000000)
def usdToGBP (usd):
return usd / 1.564768
GDP = 'GDP (£m)'
gdpCountries[GDP] = gdpCountries[GDP_INDICATOR].apply(usdToGBP).apply(roundToMillions)
gdpCountries.head()
# -
# The unnecessary columns can be dropped.
COUNTRY = 'country'
headings = [COUNTRY, GDP]
gdpClean = gdpCountries[headings]
gdpClean.head()
# The World Bank reports the life expectancy with several decimal places. After rounding, the original column is discarded.
LIFE = 'Life expectancy (years)'
lifeCountries[LIFE] = lifeCountries[LIFE_INDICATOR].apply(round)
headings = [COUNTRY, LIFE]
lifeClean = lifeCountries[headings]
lifeClean.head()
# ## Combining the data
#
# The tables are combined through an inner join on the common 'country' column.
gdpVsLife = pd.merge(gdpClean, lifeClean, on=COUNTRY, how='inner')
gdpVsLife.head()
# ## Calculating the correlation
#
# To measure if the life expectancy and the GDP grow together, the Spearman rank correlation coefficient is used. It is a number from -1 (perfect inverse rank correlation: if one indicator increases, the other decreases) to 1 (perfect direct rank correlation: if one indicator increases, so does the other), with 0 meaning there is no rank correlation. A perfect correlation doesn't imply any cause-effect relation between the two indicators. A p-value below 0.05 means the correlation is statistically significant.
# +
from scipy.stats import spearmanr
gdpColumn = gdpVsLife[GDP]
lifeColumn = gdpVsLife[LIFE]
(correlation, pValue) = spearmanr(gdpColumn, lifeColumn)
print('The correlation is', correlation)
print('The p-value is', pValue)
if pValue < 0.05:
print('It is statistically significant.')
else:
print('It is not statistically significant.')
# -
# The value shows a direct correlation, i.e. richer countries tend to have longer life expectancy, but it is not very strong.
# ## Showing the data
#
# Measures of correlation can be misleading, so it is best to see the overall picture with a scatterplot. The GDP axis uses a logarithmic scale to better display the vast range of GDP values, from a few million to several billion (million of million) pounds.
# %matplotlib inline
gdpVsLife.plot(x=GDP, y=LIFE, kind='scatter', grid=True, logx=True, figsize=(10, 4))
# The plot shows there is no clear correlation: there are rich countries with low life expectancy, poor countries with high expectancy, and countries with around 10 thousand (10<sup>4</sup>) million pounds GDP have almost the full range of values, from below 50 to over 80 years. Towards the lower and higher end of GDP, the variation diminishes. Above 40 thousand million pounds of GDP (3rd tick mark to the right of 10<sup>4</sup>), most countries have an expectancy of 70 years or more, whilst below that threshold most countries' life expectancy is below 70 years.
#
# Comparing the 10 poorest countries and the 10 countries with the lowest life expectancy shows that total GDP is a rather crude measure. The population size should be taken into account for a more precise definiton of what 'poor' and 'rich' means. Furthermore, looking at the countries below, droughts and internal conflicts may also play a role in life expectancy.
# the 10 countries with lowest GDP
gdpVsLife.sort_values(GDP).head(10)
# the 10 countries with lowest life expectancy
gdpVsLife.sort_values(LIFE).head(10)
# ## Conclusions
#
# To sum up, there is no strong correlation between a country's wealth and the life expectancy of its inhabitants: there is often a wide variation of life expectancy for countries with similar GDP, countries with the lowest life expectancy are not the poorest countries, and countries with the highest expectancy are not the richest countries. Nevertheless there is some relationship, because the vast majority of countries with a life expectancy below 70 years is on the left half of the scatterplot.
#
# Using the [NY.GDP.PCAP.PP.CD](http://data.worldbank.org/indicator/NY.GDP.PCAP.PP.CD) indicator, GDP per capita in current 'international dollars', would make for a better like-for-like comparison between countries, because it would take population and purchasing power into account. Using more specific data, like expediture on health, could also lead to a better analysis.
# ## NOW IS YOUR TURN
#
#
# ## GOOD LUCK!
# Activity
# Extend the project
# Make a copy of the Project Notebook: GDP and Life expectancy and change it to answer one or more of the following questions:
#
# To what extent do the ten countries with the highest GDP coincide with the ten countries with the longest life expectancy?
#
# Which are the two countries in the right half of the plot (higher GDP) with life expectancy below 60 years?
#
# What factors could explain their lower life expectancy compared to countries with similar GDP? Hint: use the filtering techniques you learned in Week 2 to find the two countries.
#
# Redo the analysis using the countries’ GDP per capita (i.e. per inhabitant) instead of their total GDP. If you’ve done the workbook task, you already have a column with the population data.
#
# Hint: write an expression involving the GDP and population columns, as you learned in Calculating over columns in previous modules. Think about the units in which you display GDP per capita.
#
# Redo the analysis using the indicator suggested at the end of the project notebook.
# Create your own project Create a completely new project and choose another two of the hundreds of World Bank indicators and see if there is any correlation between them. If there is a choice of similar indicators, choose one that leads to meaningful comparisons between countries.
#
# Look at the results you obtained and take a few moments to assess how they differ from mine.
# Below is ten countries with the highest GDP
highestGDP = gdpVsLife.sort_values(GDP).tail(10)
highestGDP
# Below is ten countries with the longest life expectancy
longestLife = gdpVsLife.sort_values(LIFE).tail(10)
longestLife
# countries with both highest GDP and longest life expectancy is gotten by merging the countries with highest GDP and countries with longest life expectancy. this is shown below
pd.merge(highestGDP, longestLife, how='inner', on='country')
# From the figure above we can say that the countries with highest GDPand longest life expectancy is Italy, France and Japan
# +
# right half of the graph is above 10**5
# Therefore, to get the countries in the right half of the plot
# (higher GDP) with life expectancy below 60 years we use the comparison operator and bitwise opeartor '&'
gdpVsLife[(gdpVsLife[GDP] > 10**5) & (gdpVsLife[LIFE] < 60)]
# -
# it was observed that Nigeria and South Africa are the two countries in the right half of the plot (higher GDP) and with life expectancy below 60 years
#
#
#
# +
# countries in the right half of the plot (higher GDP) and with high life expectancy of above 75years
gdpVsLife[(gdpVsLife[GDP] > 10**6) & (gdpVsLife[LIFE] >= 75)]
# -
gdpVsLife[(gdpVsLife[GDP] > 175000) & (gdpVsLife[GDP] < 375000)].sort_values(LIFE)
# Most countries with GDP Above 40 thousand million pounds have an expectancy of 70 years or more expect two countries Nigeria and South Africa. FRom the figure above we observe a great difference in life expectancy exists between South Africa and Colombia
#
# The lower life expectancy of South Africa and Nigeria maybe as a result of poor socio-economic status in Africa.
# # Analysis using the countries’ GDP per capita (i.e. per inhabitant)
# +
import pandas as pd
YEAR = 2013
GDP_INDICATOR = 'NY.GDP.MKTP.CD'
gdpReset = pd.read_csv('WB GDP 2013.csv')
LIFE_INDICATOR = 'SP.DYN.LE00.IN'
lifeReset = pd.read_csv('WB LE 2013.csv')
POP_INDICATOR = 'SP.POP.TOTL'
popReset = pd.read_csv("WB POP 2013.csv")
popReset.head()
# -
# # Cleaning data
# While inspecting the data with `head()` and `tail()` shows that, it was observed that the first 34 rows are aggregated data, that is they are country groups. Individual countries starts in row number 34
# It was observed that GDP and life expectancy values are missing for some countries.
#
# Therefore wwe have to clean the data by removing the first 34 rows and removing rows with unavailable values.
# +
gdpCountries = gdpReset[34:].dropna()
lifeCountries = lifeReset[34:].dropna()
popCountries = popReset[34:].dropna()
# -
# # Transform data
# The World Bank reports GDP in US dollars and cents. in order to make the data easy to read we then converted the GDP to millions of British pounds, using the average 2013 dollar-to-pound conversion rate provided by http://www.ukforex.co.uk/forex-tools/historical-rate-tools/yearly-average-rates.
# +
def roundToMillions (value):
return round(value / 1000000)
def usdToGBP (usd):
return usd / 1.564768
GDP = 'GDP (£m)'
gdpCountries[GDP] = gdpCountries[GDP_INDICATOR].apply(usdToGBP)
gdpCountries.head()
# -
# We need to need remove the column we don't need and focus the the main data
COUNTRY = 'country'
headings = [COUNTRY, GDP]
gdpClean = gdpCountries[headings]
gdpClean.head()
POP = ('Population')
popCountries[POP] = popCountries[POP_INDICATOR]
headings = [COUNTRY, POP]
popClean = popCountries[headings]
popClean.head()
gdpVspop = pd.merge(gdpClean, popClean, on=COUNTRY, how='inner')
gdpVspop.head()
GDP_PC = 'GDP per capita (£)'
gdpVspop[GDP_PC] = gdpVspop[GDP]/gdpVspop[POP]
# From the data, it was observed that the life expectancy are in decimals. Inorder to make the data easy to read, we need to round it up nearest whole number.
LIFE = 'Life expectancy (years)'
lifeCountries[LIFE] = lifeCountries[LIFE_INDICATOR].apply(round)
headings = [COUNTRY, LIFE]
lifeClean = lifeCountries[headings]
lifeClean.head()
gdpPC = gdpVspop[[COUNTRY, GDP_PC]]
gdpPC.head(10)
gdpPCVslife = pd.merge(gdpPC, lifeClean, on=COUNTRY, how='inner')
gdpPCVslife.head()
highestGDP_PC = gdpPCVslife.sort_values(GDP_PC).tail(10)
highestGDP_PC
longestlife = gdpPCVslife.sort_values(LIFE).tail(10)
longestlife
# countries with both the highest GDP per capital and longest life expectancy are shown below
pd.merge(highestGDP_PC, longestlife, how='inner', on='country')
# +
from scipy.stats import spearmanr
gdppcColumn = gdpPCVslife[GDP_PC]
lifeColumn = gdpPCVslife[LIFE]
(r, p) = spearmanr(gdppcColumn, lifeColumn)
print('The correlation coefficient is', r)
print('The p-value is', p)
if p < 0.05:
print('This result is statistically significant.')
else:
print('This result is not statistically significant.')
# -
# The value 0.8502554597715132 shows a strong correlation
# +
# %matplotlib inline
gdpPCVslife.plot(x=GDP_PC, y=LIFE, grid=True, kind='scatter', logx=True, figsize=(10,5))
# -
# from the graph above it is observed that that the life expectancy increases as the GDP per capita increases. Therefore, there's a strong correlation between both
| Omobolanle Adeyemi WT 192/Project1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example HDF5 Insights
#
# This Jupyter Notebook should give a brief overview how to programmatically analyze the HDF5 files produced by *mycelyso*. Please note that you can always inspect these files with *mycelyso Inspector* as well, this tutorial should just give you a hint how to open these files if you might want to write your own analyses.
#
# First, it is assumed that an `output.h5` is present in the current directory, with an analysis of the *example dataset* contained.
#
# You can fetch the *example dataset* by running `get-dataseth.sh` or download it manually at https://zenodo.org/record/376281.
#
# Afterwards, analyze it with:
# ```
# > python -m mycelyso S_lividans_TK24_Complex_Medium_nd046_138.ome.tiff -t BoxDetection=1
# ```
#
# Afterwards, you will have an `output.h5` in the residing in the directory.
#
# We will be using [Pandas](https://pandas.pydata.org) to read our data, while the non-tabular data could easily be read with any other HDF5 compatible tool, the tabular data is layed out in a chunked format particular to Pandas, and as such it is easiest to open it with Pandas.
# First, some general setup …
# +
# %matplotlib inline
# %config InlineBackend.figure_formats=['svg']
import pandas
pandas.options.display.max_columns = None
import numpy as np
import networkx as nx
from networkx.readwrite import GraphMLReader
from matplotlib import pyplot, ticker
pyplot.rcParams.update({
'figure.figsize': (10, 6), 'svg.fonttype': 'none',
'font.sans-serif': 'Arial', 'font.family': 'sans-serif',
'image.cmap': 'gray_r', 'image.interpolation': 'none'
})
# -
# ## Opening the HDF5 file
# We will load the `output.h5` using `pandas.HDFStore` …
store = pandas.HDFStore('output.h5', 'r')
store
# Now let's dive a bit into the HDF5 file.
#
# Remember that HDF5 stands for *Hierarchical* Data Format 5 …
# +
root = store.get_node('/')
print("Root:")
print(repr(root))
print()
print("/results:")
print(repr(root.results))
# -
# The key names are dependent on the on-disk path of the analyzed file.
# Assuming there is only one file analyzed with one position in the file, we pick the first …
for image_file in root.results:
print(image_file)
for position in image_file:
print(position)
break
# We can now investigate what data is available for that particular position
#
# There is e.g., (binary) data, there are images, and there are various tabular datasets
# +
print("data")
print(position.data)
for node in position.data:
print(node)
print()
print("nodes")
print(position.images)
for node in position.images:
print(node)
print()
# -
# ## Accessing Graph and Image Data
# Let's for example start with pulling out an image from the file, and displaying it …
# +
binary_images = list(position.images.binary)
skeleton_images = list(position.images.skeleton)
n = 120
total = len(binary_images)
assert 0 <= n < total
print("Total count of images: %d" % (total,))
fig, (ax_l, ax_r) = pyplot.subplots(1, 2, sharey=True)
fig.suptitle('Images of Timepoint #%d:' % (n,))
ax_l.imshow(binary_images[n])
ax_l.set_title('Binary Image')
ax_r.imshow(skeleton_images[n])
ax_r.set_title('Skeleton')
# -
# Let's now take a look at the graph data present for the position, display it and overlay it onto the image data …
# +
# The graph structure is saved in GraphML
draw_parameters = dict(node_size=25, node_color='darkgray', linewidths=0, edge_color='darkgray', with_labels=False)
#graphml_data = list([np.array(graphml).tobytes() for graphml in list(position.data.graphml)])
graphml_data = list(position.data.graphml)
graph, = GraphMLReader()(string=np.array(graphml_data[n]).tobytes())
# the following draw function needs separate positions...
# each node has its position saved as attributes:
example_node_id = list(sorted(graph.node.keys()))[1]
print("Example node: %s: %r" % (example_node_id, graph.node[example_node_id],))
other_node_id = list(sorted(graph.adj[example_node_id].keys(), reverse=True))[0]
print("Some other node: %s" % (other_node_id,))
print("The distance between the two nodes is: %.2f px" % (graph.adj[example_node_id][other_node_id]['weight']))
pyplot.title('Graph Representation of Timepoint #%d:' % (n,))
# first draw the graph,
pos = {n_id: (n['x'], n['y']) for n_id, n in graph.node.items()}
nx.draw_networkx(graph, pos=pos, **draw_parameters)
example_nodes = [graph.node[node_id] for node_id in [example_node_id, other_node_id]]
# mark on top the two choosen sample nodes
pyplot.scatter([p['x'] for p in example_nodes], [p['y'] for p in example_nodes], zorder=2)
# then show the corresponding binarized image
pyplot.imshow(binary_images[n])
# -
# ## Accessing Tabular Data
#
# In the next few cells we'll take a look at the tabular data stored in the HDF5 file.
#
# There is for example the `result_table`, which contains compounded information about the whole position:
result_table = store[position.result_table._v_pathname]
result_table
# Then there is the `result_table_collected`, which contains collected information about every single frame of the time series of one position:
result_table_collected = store[position.result_table_collected._v_pathname]
result_table_collected
# The per-frame informations contain e.g. the graph length (i.e. the mycelium length), which can be plotted over time:
# +
timepoint = result_table_collected.timepoint / (60*60)
length = result_table_collected.graph_edge_length
pyplot.title('Length over Time')
pyplot.xlabel('Time [h]')
pyplot.ylabel('Length [µm]')
pyplot.plot(timepoint, length)
# -
# Last but not least, we will look at mycelium level tracking data in the `track_table`.
# The `track_table` is a level deeper in the HDF5 structure, next to tables with individual tracks.
track_table = store[list(position.tables.track_table)[0]._v_pathname]
track_table
# Let's find the longest track and try to visualize it:
# +
track_table.sort_values(by=['count'], ascending=False, inplace=True)
particular_tracking_table = track_table.aux_table[0] # the first
_mapping_track_table_aux_tables = store[list(position.tables._mapping_track_table_aux_tables)[0]._v_pathname]
index = _mapping_track_table_aux_tables.query('_index == @particular_tracking_table').individual_table
the_longest_track = store[getattr(position.tables._individual_track_table_aux_tables, 'track_table_aux_tables_%09d' % (index,))._v_pathname]
the_longest_track
# +
timepoint = the_longest_track.timepoint / (60*60)
length = the_longest_track.distance
pyplot.title('Length over Time')
pyplot.xlabel('Time [h]')
pyplot.ylabel('Length [µm]')
pyplot.plot(timepoint, length)
# -
# Now all tracked hyphae:
# +
pyplot.title('Length over Time')
pyplot.xlabel('Time [h]')
pyplot.ylabel('Length [µm]')
for idx, row in track_table.iterrows():
particular_tracking_table = int(row.aux_table)
index = _mapping_track_table_aux_tables.query('_index == @particular_tracking_table').individual_table
track = store[getattr(position.tables._individual_track_table_aux_tables, 'track_table_aux_tables_%09d' % (index,))._v_pathname]
timepoint = track.timepoint / (60*60)
length = track.distance - track.distance.min()
pyplot.plot(timepoint, length)
pyplot.xlim(0, None)
# -
| examples/Example_HDF5_Insights.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Output data preparation for dataset Deezer
# #### Plots and figures in separate notebook
# IMPORTS
import matplotlib.pyplot as plt
import numpy as np
import csv
import networkx as nx
from random import sample
import time
import math
import random
import scipy
import pandas as pd
# +
# Define necessary functions
def arccosh_og(x): ##note that x*x-1 might be less than zero :( And then log(t) could be negative (negative distance?!?!?!?!)
t = x + math.sqrt(x * x - 1)
return math.log(t)
def arccosh(x):
t = x + math.sqrt(max(x * x, 1) - 1)
return max(math.log(t), 0.5)
def query(coordinates, source, destination, curvature):
if source == destination:
return 0
sourceCoords = coordinates[source]
destinationCoords = coordinates[destination]
i = 0
ts = 1.0
td = 1.0
tt = 1.0
for i in range(len(sourceCoords)):
ts += math.pow(sourceCoords[i], 2)
td += math.pow(destinationCoords[i], 2)
tt += (sourceCoords[i] * destinationCoords[i])
#print(ts, td, tt)
t = math.sqrt(ts * td) - tt
#print('t:', t)
return arccosh(t) * math.fabs(curvature)
def intersection_similarity(u,v):
return len(set(u).intersection(set(v)))
def weighted_intersection_similarity(u,v, alpha):
similarity = 0
if len(u)==len(v):
n = len(u)
for i in range(n):
if u[i] in v:
j = v.index(u[i])
similarity += (n-abs(i-j))**alpha
else:
print('not equal vector lengths')
similarity = -1
return similarity
# +
# READ REAL NETWORK - Giant Connected Component
dataset = 'HU_edges.csv'
data = pd.read_csv(dataset, header = 0, sep = ',')
data = data[[data.columns[0], data.columns[1]]]
data.head()
graph = nx.from_pandas_edgelist(data, data.columns[0], data.columns[1])
Gcc = sorted(nx.connected_components(graph), key=len, reverse=True)
giant = graph.subgraph(Gcc[0])
# +
# SPECIFY THESE INPUTS
output_file_name = 'deezer/out'
partitions = 1
curvature = -1
number_of_nodes = 47538
#######################
landFile = output_file_name + '.land'
coordFiles = [output_file_name + str(i) + '.coord' for i in range(partitions)]
coordinates = dict()
with open(landFile) as infile:
for line in infile:
linesplit = line.split()
id = int(linesplit[0])
coords = [float(c) for c in linesplit[1:]]
coordinates[id] = coords
for coordFile in coordFiles:
with open(coordFile) as infile:
for line in infile:
linesplit = line.split()
id = int(linesplit[0])
coords = [float(c) for c in linesplit[1:]]
coordinates[id] = coords
#while True:
# query_input = input("Enter ID of 2 nodes: ")
# if query_input == 'exit' or query_input == 'q' or query_input == 'quit':
# break
# querysplit = query_input.split()
# source = int(querysplit[0])
# destination = int(querysplit[1])
# estimate = query(coordinates, source, destination, curvature)
# print('Rigel estimates the distance between %d and %d to be %f.\n' % (source, destination, estimate))
# +
# Relative errors - approximation: select 'select_count = 1000' nodes from where distances (to all nodes) are calculated
### This is necessary due to slow EXACT path calculation
result_avg_path_length_estimated = []
result_avg_path_length_exact = []
result_radius_estimated = []
result_radius_exact = []
result_diameter_estimated = []
result_diameter_exact = []
top_cent_exact = []
top_cent_estimate = []
top_ecc_exact = []
top_ecc_estimate = []
for sed in range(5):
print('START OF SEED', sed, '.')
np.random.seed(sed)
select_count = 1000
selected_nodes = random.sample(range(number_of_nodes), select_count)
relative_errors = dict()
exact_distances = dict()
estimated_distances= dict()
avg_path_length_exact = 0
avg_path_length_estimated = 0
radius_estimated = number_of_nodes
diameter_estimated = 0
radius_exact = number_of_nodes
diameter_exact = 0
eccentricites_estimated = []
eccentricites_exact =[]
centralities_exact = []
centralities_estimated = []
node_names = list(giant.nodes())
iters = 0
for source in selected_nodes:
iters += 1
if iters % int(select_count/10) == 0:
print('Processed ', 10 * iters / int(select_count/10), '% of total calculations...')
eccentricity_curr_est = 0
eccentricity_curr_ex = 0
exact_distances[source] = []
estimated_distances[source] = []
relative_errors[source] = []
for target in selected_nodes:
#print('points:', source, target)
if source != target:
estimate = query(coordinates, source, target, curvature)
exact = nx.shortest_path_length(giant, node_names[source], node_names[target])
avg_path_length_estimated += estimate
avg_path_length_exact += exact
eccentricity_curr_est = max(eccentricity_curr_est, estimate)
diameter_estimated = max(diameter_estimated, estimate)
eccentricity_curr_ex = max(eccentricity_curr_ex,exact)
diameter_exact = max(diameter_exact,exact)
relative_errors[source].append(abs(estimate-exact)/exact)
exact_distances[source].append(exact)
estimated_distances[source].append(estimate)
else:
relative_errors[source].append(0)
exact_distances[source].append(0)
estimated_distances[source].append(0)
radius_estimated = min(eccentricity_curr_est, radius_estimated)
radius_exact = min(eccentricity_curr_ex, radius_exact)
eccentricites_estimated.append(0-eccentricity_curr_est)
eccentricites_exact.append(0-eccentricity_curr_ex)
centralities_exact.append(0-np.mean(list(exact_distances.values())))
centralities_estimated.append(0-np.mean(list(estimated_distances.values())))
avg_path_length_estimated = avg_path_length_estimated / (select_count * (select_count - 1) )
avg_path_length_exact = avg_path_length_exact / (select_count * (select_count - 1) )
result_avg_path_length_estimated.append(avg_path_length_estimated)
result_avg_path_length_exact.append(avg_path_length_exact)
result_radius_estimated.append(radius_estimated)
result_radius_exact.append(radius_exact)
result_diameter_estimated.append(diameter_estimated)
result_diameter_exact.append(diameter_exact)
ind = np.argpartition(centralities_exact, -80)[-80:]
top_cent_exact.append(ind[np.argsort(np.array(centralities_exact)[ind])])
ind = np.argpartition(centralities_estimated, -80)[-80:]
top_cent_estimate.append(ind[np.argsort(np.array(centralities_estimated)[ind])])
ind = np.argpartition(eccentricites_exact, -80)[-80:]
top_ecc_exact.append(ind[np.argsort(np.array(eccentricites_exact)[ind])])
ind = np.argpartition(eccentricites_estimated, -80)[-80:]
top_ecc_estimate.append(ind[np.argsort(np.array(eccentricites_estimated)[ind])])
# -
#ESTIMATIONs:
print(np.std(result_avg_path_length_estimated))
print(np.std(result_avg_path_length_exact))
print(np.std(result_radius_estimated))
print(np.std(result_radius_exact))
print(np.std(result_diameter_estimated))
print(np.std(result_diameter_exact))
# Average similarity calculations
avg_wsims_c = []
avg_wsims_e = []
avg_sims_c = []
avg_sims_e = []
for j in [10,20,30,40,50,60,70,80]:
swc = 0
swe = 0
sc = 0
se = 0
for i in range(5):
swc += weighted_intersection_similarity(list(top_cent_estimate[i][-j:]),list(top_cent_exact[i][-j:]),1)
swe += weighted_intersection_similarity(list(top_ecc_estimate[i][-j:]),list(top_ecc_exact[i][-j:]),1)
sc += intersection_similarity(list(top_cent_estimate[i][-j:]),list(top_cent_exact[i][-j:]))
se += intersection_similarity(list(top_ecc_estimate[i][-j:]),list(top_ecc_exact[i][-j:]))
avg_wsims_c.append(swc/5.0/j)
avg_wsims_e.append(swe/5.0/j)
avg_sims_c.append(sc/5.0/j)
avg_sims_e.append(se/5.0/j)
plt.plot([10,20,30,40,50,60,70,80],avg_sims_c, 'x-', label = 'closeness')
plt.plot([10,20,30,40,50,60,70,80],avg_sims_e, 'x-', label = 'eccentricity')
plt.title('Deezer - Similarity of top nodes', fontsize = 15)
plt.xlabel('top k')
plt.ylabel('similarity')
plt.legend()
plt.show()
# +
# save data for reuse (plotting)
with open('deezer_tops_c.pickle', 'wb') as handle:
pickle.dump(avg_sims_c, handle)
with open('deezer_tops_e.pickle', 'wb') as handle:
pickle.dump(avg_sims_e, handle)
with open('deezer_diam_ex.pickle', 'wb') as handle:
pickle.dump(result_diameter_exact, handle)
# -
# similarity measures - in detail
for i in range(5):
print('Weighted Centrality similarity of top 20: ', weighted_intersection_similarity(list(top_cent_estimate[i]),list(top_cent_exact[i]),1))
print('Weighted Eccentricity similarity of top 20: ', weighted_intersection_similarity(list(top_ecc_estimate[i]),list(top_ecc_exact[i]),1))
print('Centrality similarity of top 20: ', intersection_similarity(list(top_cent_estimate[i]),list(top_cent_exact[i])))
print('Eccentricity similarity of top 20: ', intersection_similarity(list(top_ecc_estimate[i]),list(top_ecc_exact[i])))
# similarity measure - top j nodes
j= 20
for i in range(5):
print('Weighted Centrality similarity of top 20: ', weighted_intersection_similarity(list(top_cent_estimate[i][-j:]),list(top_cent_exact[i][-j:]),1))
print('Weighted Eccentricity similarity of top 20: ', weighted_intersection_similarity(list(top_ecc_estimate[i]),list(top_ecc_exact[i]),1))
print('Centrality similarity of top 20: ', intersection_similarity(list(top_cent_estimate[i][-j:]),list(top_cent_exact[i][-j:])))
print('Eccentricity similarity of top 20: ', intersection_similarity(list(top_ecc_estimate[i]),list(top_ecc_exact[i])))
#Average relative error of last selection
ARE_per_source = [np.mean(relative_errors[node]) for node in relative_errors.keys()]
ARE_total = np.mean(ARE_per_source)
print('Relative error (approximated): ', ARE_total)
# +
# distribution of relative error in total (last selection)
relative_errors_total = []
for source in relative_errors.keys():
relative_errors_total += relative_errors[source]
#print(source, ': ' ,min(relative_errors[source]))
plt.hist(relative_errors_total, bins = 100)
plt.title('RE distribution')
plt.xlabel('RE')
plt.ylabel('#occurance')
plt.show()
plt.hist([relative_errors_total[i] for i in range(len(relative_errors_total)) if (relative_errors_total[i] < 1.0 and relative_errors_total[i] > 0.0)], bins = 100)
plt.title('RE distribution - in [0,1]')
plt.xlabel('RE')
plt.ylabel('#occurance')
plt.show()
# +
# save data for later reuse (figures)
with open('deezer_erdist.pickle', 'wb') as handle:
pickle.dump([relative_errors_total[i] for i in range(len(relative_errors_total)) if (relative_errors_total[i] < 1.0 and relative_errors_total[i] > 0.0)], handle)
with open('deezer_cdf.pickle', 'wb') as handle:
pickle.dump({'bins': bins_count[1:], 'cdf':cdf }, handle)
# +
# Cumulative Distribution Function of the Distribution if Relative Errors
base = [relative_errors_total[i] for i in range(len(relative_errors_total)) if (relative_errors_total[i] < 1.5 and relative_errors_total[i] > 0.0)]
count, bins_count = np.histogram(base, bins=1000)
pdf = count / sum(count)
cdf = np.cumsum(pdf)
plt.plot(bins_count[1:], cdf, label="CDF")
plt.title('CDF of Relative Error')
plt.xlabel('Relative Error')
plt.ylabel('CDF')
plt.show()
# +
# Is there any extremely bad node (last selection)
bad_nodes = sorted(range(len(ARE_per_source)), key=lambda i: ARE_per_source[i])[-10:]
print('Nodes with the highest REs: ', bad_nodes)
print('Highest REs:', [round(ARE_per_source[bad_node], 3) for bad_node in bad_nodes])
# -
| embed/result_report_deezer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: default_3.7.9
# language: python
# name: default_3.7.9
# ---
import pandas as pd
import numpy as np
import xgboost as xgb
import lightgbm as gbm
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.linear_model import LogisticRegression
# +
from sklearn.model_selection import train_test_split, cross_val_score, cross_validate, StratifiedKFold, GridSearchCV
from sklearn.metrics import f1_score, classification_report, confusion_matrix, make_scorer
from functools import partial
# -
train = pd.read_parquet('../data/processed/train.parquet')
test = pd.read_parquet('../data/processed/test.parquet')
metric = partial(f1_score, average='macro')
score = make_scorer(f1_score, average='macro')
# +
X, y = train.loc[:, ~train.columns.isin(['revenue', "id"])], train.revenue
from imblearn.over_sampling import SMOTE
resampler = SMOTE(random_state=0)
X, y = resampler.fit_resample(X, y)
# -
X_train, X_test, y_train, y_test = train_test_split(X,y, stratify=y, test_size=0.2)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, stratify=y_train, test_size=0.2)
def evaluate_model(clf, metric, score, data):
"""Evaluates a model and returns a summary for visualize. [just internal use]
"""
X_train, X_test, y_train, y_test = data
f1_cross = cross_val_score(clf, X_train, y_train, cv=5, scoring=score)
clf.fit(X_train, y_train)
return {
"model": clf.__class__.__name__,
"test": metric(y_test, clf.predict(X_test)),
"train": metric(y_train, clf.predict(X_train)),
"f1_cross_mean": np.mean(f1_cross),
"f1_cross_std": np.std(f1_cross)
}
lgb_params = {
'n_estimators':450,
'boosting_type': 'dart',
'objective': 'binary',
'learning_rate': 0.1,
'max_depth' : 8,
'num_leaves': 64,
# 'is_unbalance':True, # is_unbalance vs scale_pos_weight
## see https://towardsdatascience.com/understanding-lightgbm-parameters-and-how-to-tune-them-6764e20c6e5b
'scale_pos_weight': y_train.sum() / (y_train-1).abs().sum()
# 'max_bin': 512,
# 'subsample_for_bin': 200,
# 'subsample': 1,
# 'subsample_freq': 1,
# 'colsample_bytree': 0.8,
# 'reg_alpha': 5,
# 'reg_lambda': 10,
# 'min_split_gain': 1,
# 'min_child_weight': 1,
# 'min_child_samples': 5,
}
# +
# I've tested out other models but I select LightGBM
models = [
gbm.LGBMClassifier(**lgb_params,),
# xgb.XGBClassifier(
# n_estimators=400
# ),
# RandomForestClassifier(),
# LogisticRegression(),
# AdaBoostClassifier()
]
# -
results = [evaluate_model(model, metric, score, (X_train, X_val, y_train, y_val)) for model in models]
pd.DataFrame.from_dict(results)
models[0].get_params()
# ## Model evaluation
# I planned to use LightGBM.
# +
# Create parameters to search
# these hypermarateres were obtained from 004 notebook with optuna.
# Hence, I let it to have a reproducible workflow.
grid_params = {
'learning_rate': [0.08],
'boosting_type' : ['dart', 'goss'],
'n_estimators': [300, 350, 400],
'lambda_l1': [0.001, 0.02],
'lambda_l2': [1e-3, 0.1, 5],
'max_depth': [8,16,25],
'num_leaves': [23, 75],
# "random_state": [42],
# 'random_state' : [501], # Updated from 'seed'
# 'colsample_bytree' : [0.65, 0.66],
# 'subsample' : [0.60, 0.75],
# 'reg_alpha' : [1,1.2],
# 'reg_lambda' : [1,1.2,1.4],
}
# -
clf = gbm.LGBMClassifier(**lgb_params)
# +
# Create the grid
grid = GridSearchCV(
clf,
grid_params,
cv=StratifiedKFold(2),
n_jobs=-1,
scoring=score,
verbose=1
)
# Run the grid
# I setted fit_params for lightgbm, if needed, remove it (in case of changing the model)
grid.fit(
X_train, y_train,
eval_set=[(X_train, y_train), (X_val, y_val)],
early_stopping_rounds=15,
eval_metric=[lambda y_true, y_pred: ("F1_MACRO", metric(y_true, np.round(y_pred)), True)],
feature_name=X_train.columns.tolist(),
categorical_feature=["Month", "TrafficType", "Browser", "SpecialDay", "OperatingSystems", "Region", "VisitorType", "Weekend"]
)
# Print the best parameters found
print(grid.best_params_)
print(grid.best_score_)
# -
print(classification_report(y_test, grid.best_estimator_.predict(X_test)))
print(confusion_matrix(y_test, grid.best_estimator_.predict(X_test)))
print(metric(y_test, grid.best_estimator_.predict(X_test)))
print(metric(y_train, grid.best_estimator_.predict(X_train)))
# +
### for submition, parameters from optuna + grid-search
### dictionary update: {**dict1, **dict2}, or with dict.update(dict2)
clf = gbm.LGBMClassifier(**{**lgb_params, **grid.best_params_})
# -
clf.fit(
pd.concat([X_train, X_val], axis=0), pd.concat([y_train,y_val], axis=0),
eval_set=[(X_train, y_train), (X_val, y_val)],
early_stopping_rounds=15,
eval_metric=[lambda y_true, y_pred: ("F1_MACRO", metric(y_true, np.round(y_pred)), True)],
feature_name=X_train.columns.tolist(),
categorical_feature=["Month", "TrafficType", "Browser", "SpecialDay", "OperatingSystems", "Region", "VisitorType", "Weekend"]
)
# +
print(classification_report(y_train, clf.predict(X_train)))
print(classification_report(y_test, clf.predict(X_test)))
print(metric(y_train, clf.predict(X_train)))
print(metric(y_test, clf.predict(X_test)))
# -
test_pred = test.loc[:, ~test.columns.isin(['revenue', "id"])]
submit = clf.predict(test_pred)
from datetime import datetime
get_date = lambda : datetime.now().strftime("%Y%m%d_%H%M%S")
# +
date_part = get_date()
# save results
pd.DataFrame({"id":test.id, "revenue":submit}).to_csv(f'../data/results/{date_part}_lgb.csv', index=False)
# just if necessary, persist the parameters
pd.DataFrame([clf.get_params()]).to_csv(f'../data/results/{date_part}_params.csv')
# -
clf._Booster.save_model(f'../data/results/{date_part}_model.txt')
# __________________________
# ## Model Interpretability
# - *TODO*
clf.feature_importances_
gbm.plot_importance(clf, max_num_features=10)
| notebooks/003_Model_evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ee]
# language: python
# name: conda-env-ee-py
# ---
#https://github.com/giswqs/geemap/blob/master/examples/notebooks/11_export_image.ipynb
import ee
import geemap
import numpy as np
import matplotlib.pyplot as plt
# Initialize the Earth Engine module.
ee.Initialize()
#// Create a domain name to attach to your output. Optional.
domain_name = 'WY'
'''
// These are the min and max corners of your domain in Lat, Long
// Western Wyoming
// Input the minimum lat, lower left corner
'''
minLat = 42.363116
#// Input the minimum long, lower left corner
minLong = -111.155208
#// Input the max lat, upper right corner
maxLat = 44.582480
#// Input the max Long, upper right corner
maxLong = -109.477849
#/ These are the min and max corners of your reanalysis in Lat, Long (create a slightly larger box)
#// Input the minimum lat, lower left corner
minLat2 = (minLat - 0.25);
#// print(minLat2);
#// Input the minimum long, lower left corner
minLong2 = (minLong - 0.5);
#// Input the max lat, upper right corner
maxLat2 = (maxLat + 0.25);
#// Input the max Long, upper right corner
maxLong2 = (maxLong + 0.5);
# +
# This resolution for the NLCD and DEM outputs for the SnowModel domain in meters
sm_resolution = 100
'''// Resolution for the PRISM output. This shoud change by Latitude of the domain
// because the PRISM product spatial resolution is 2.5 minutes, which equals 150 arc seconds.
// You can use this arc-second calculator to estimate the correct value for the PRISM resolution by latitude
// https://opendem.info/arc2meters.html
// This is one arc-second in meters for 43 degrees N Latitude'''
one_arcsecond = 22.57
PRISM_resolution = one_arcsecond * 150
'''// Define the final output projection using EPSG codes
// These are the EPSG codes for the final projection of your SnowModel simulation.
// WGS UTM Zone 12 Code for Idaho/Wyoming = 32612
// WGS UTM Zone 11 Code for Nevada = 32611
// WGS UTM Zone 10 Code for West Coast = 32610
// WGS 84 4326
// WGS UTM 10
// WGS Alaska Albers = 3338'''
epsg_code = 'EPSG:32612'
#// Name the DEM output
dem_name = 'DEM'
#// Name the Land Cover output
lc_name = 'NLCD2016'
'''// The Beginning and End Dates you care about //
// This will start on the 'begin' date at 0:00 and the last iteration will be
// on the day before the 'end' date below.'''
begin = '2014-09-01'
end = '2014-09-02'
# -
# # Domain
my_domain = ee.Geometry.Rectangle(**{'coords':[minLong,minLat,maxLong,maxLat],'proj': 'EPSG:4326','geodesic':True,});
my_domain2 = ee.Geometry.Rectangle([minLong2,minLat2,maxLong2,maxLat2])
'''//////////////// Datasets of Interest //////////////////////
//////// Digital Elevation Models and Land Cover /////////
///////////////////////////////////////////////////////////////
// NOTE: several choices below for DEM. Uncomment your preferred option
//////// Import 30m SRTM Data ///////////////////
// NOTE: This only covers through 60 degrees latitude. See visualization layers.
//var SRTM30 = ee.Image('USGS/SRTMGL1_003');
// Find out some info about this image (hint: look in the console)
//var bands30 = SRTM30.bandNames();
//var info30 = SRTM30.getInfo();
//print(bands30,'Band Names');
//print(info30,'Band Info');
//Map.addLayer(SRTM30,visparams,'SRTM30');
//////// Import 100m ASTER data //////////////
// NOTE: this works above 60 deg lat; better for Alaska...
//var ASTER = ee.Image('NASA/ASTER_GED/AG100_003');
// Find out some info about this image (hint: look in the console)
//var bands100 = ASTER.bandNames();
//var info100 = ASTER.getInfo();
//print(bands100,'Band Names');
//print(info100,'Band Info');
//Map.addLayer(ASTER,visparams_aster,'ASTER');
'''
#///////// Import 90m SRTM Data ////////////////////
#// NOTE: This only covers through 60 degrees latitude. See visualization layers.
SRTM90 = ee.Image('CGIAR/SRTM90_V4')
bands90 = SRTM90.bandNames()
info90 = SRTM90.getInfo()
#print(bands90,'Band Names')
#print(info90,'Band Info')
'''//////// Import NLCD Dataset ////////////////////
var NLCD = ee.ImageCollection('USGS/NLCD');
//Note: the NLCD has numerous images for different years.'''
landcover = NLCD.select('landcover');
#// Define the timeframe of NLCD images to select. Currently, its set to the previous 5 years.
landcoverfiltered=landcover.filterDate('2015-01-01','2020-01-01');
'''landcoverVis = {
'min': 0.0,
'max': 95.0,
'palette': [
'466b9f', 'd1def8', 'dec5c5', 'd99282', 'eb0000', 'ab0000', 'b3ac9f',
'68ab5f', '1c5f2c', 'b5c58f', 'af963c', 'ccb879', 'dfdfc2', 'd1d182',
'a3cc51', '82ba9e', 'dcd939', 'ab6c28', 'b8d9eb', '6c9fb8'
],
};'''
#// Create a single image out of the image collection using the most common land cover
#// designation from the previous 5 years.
lcsingle=landcoverfiltered.mode();
#Map.addLayer(lcsingle, landcoverVis, 'Landcover');
# # download ee image
#
# Here I am just downloading the dem tif, but you could also download the lc data
out_dir = os.path.abspath('/nfs/attic/dfh/Aragon2/Notebooks/preprocess_python/Downloads')
filename = os.path.join(out_dir, 'dem.tif')
image = SRTM90.clip(my_domain).unmask()
geemap.ee_export_image(image, filename=filename, scale=sm_resolution, region=my_domain, file_per_band=False, crs = epsg_code)
# # download ee collection
#
# It appears that I am able to access the data, but I have to convert the collection to images before downloading.
'''//////////////// Datasets of Interest //////////////////////
//////// Reanalysis DATA /////////
///////////////////////////////////////////////////////////////'''
cfsv2 = ee.ImageCollection('NOAA/CFSV2/FOR6H') \
.filterBounds(my_domain2) \
.filter(ee.Filter.date(begin,end))
# This method exports daily .tif files of the collection that have all 22 bands.
# https://developers.google.com/earth-engine/datasets/catalog/NOAA_CFSV2_FOR6H#bands
cfsv2 = ee.ImageCollection('NOAA/CFSV2/FOR6H') \
.filterBounds(my_domain) \
.filter(ee.Filter.date(begin,end))
geemap.ee_export_image_collection(cfsv2,out_dir=out_dir,region=my_domain2,crs=epsg_code,scale=22200)
# This method exports daily .tif files of the selected variables needed for SM
# +
cfsv2 = ee.ImageCollection('NOAA/CFSV2/FOR6H') \
.filterBounds(my_domain2) \
.filter(ee.Filter.date(begin,end))
data = cfsv2.select('Temperature_height_above_ground', \
'Geopotential_height_surface', \
'v-component_of_wind_height_above_ground', \
'Pressure_surface', \
'Specific_humidity_height_above_ground', \
'Precipitation_rate_surface_6_Hour_Average', \
'Downward_Long-Wave_Radp_Flux_surface_6_Hour_Average', \
'Downward_Short-Wave_Radiation_Flux_surface_6_Hour_Average')
geemap.ee_export_image_collection(data, out_dir=out_dir,region=my_domain2,scale=22200,crs=epsg_code)
# -
# # GEE data directly to numpy
# I think there should be a way to iterare through these to pull the data directly into a numpy array - but I cannot quite figure it out.
# This is a different package that could maybe be useful: https://github.com/gee-community/gee_tools/issues/27
tair = cfsv2.select('Temperature_height_above_ground').toBands()
elev = cfsv2.select('Geopotential_height_surface').toBands()
uwind = cfsv2.select('u-component_of_wind_height_above_ground').toBands()
vwind = cfsv2.select('v-component_of_wind_height_above_ground').toBands()
surfpres = cfsv2.select('Pressure_surface').toBands()
spechum = cfsv2.select('Specific_humidity_height_above_ground').toBands()
prec = cfsv2.select('Precipitation_rate_surface_6_Hour_Average').toBands()
lwr = cfsv2.select('Downward_Long-Wave_Radp_Flux_surface_6_Hour_Average').toBands()
swr = cfsv2.select('Downward_Short-Wave_Radiation_Flux_surface_6_Hour_Average').toBands()
rgb_img = geemap.ee_to_numpy(swr, region=my_domain)
rgb_img
# +
# from geetools import batch
# import ee
# geom = ee.Geometry.Point([-72,-42]).buffer(1000)
# col = ee.ImageCollection.fromImages([ee.Image.random(1), ee.Image.random(2), ee.Image.random(3)])
# size = col.size().getInfo()
# clist = col.toList(size)
# for i in range(size):
# image = ee.Image(clist.get(i))
# iid = image.id().getInfo()
# name = 'test_'+iid
# print('downloading '+name)
# batch.Image.toLocal(image, name, scale=100, region=geom)
| GEE_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
## data 선언
x_data = [[0.,0.], [0.,1.], [1.,0.],[1.,1.]]
y_data = [[1.], [0.], [0.], [1.]]
# +
## tf.keras를 활용한 perceptron 모델 구현.
model = tf.keras.Sequential() ## 모델 선언
model.add(tf.keras.layers.Dense(1, input_dim=2)) # 선언된 모델에 add를통해 쌓아감.은닉층
model.summary()
# +
# 모델 loss, 학습 방법 결정하기
optimizer=tf.keras.optimizers.SGD(learning_rate=0.5) ### 경사 하강법으로 global min 에 찾아가는 최적화 방법 선언.
loss=tf.keras.losses.binary_crossentropy ## 예측값 과 정답의 오차값 정의. mse는 mean squre error로 (예측값 - 정답)^2 를 의미
metrics=tf.keras.metrics.binary_accuracy ### 학습하면서 평가할 메트릭스 선언언
# 모델 컴파일하기
model.compile(loss=loss, optimizer=optimizer, metrics=[metrics])
# 모델 동작하기
model.fit(x_data, y_data, epochs=3000, batch_size=4)
| tensorflow/day2/answer/A_02_00_XOR_problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.0 64-bit
# metadata:
# interpreter:
# hash: 3ee23a5df1e42c0866af1bc156f33b0e74320a58edd3956f52bd54dba858fc5a
# name: python3
# ---
# +
def reverse(str):
output = ''
index = len(str)
while index:
index -= 1
output += str[index]
return output
reverse('lo')
# +
def reverse(string):
if len(string) == 1 or string == '':
return string
else:
return string[-1] + reverse(string[:-1])
reverse('motor')
# +
def replace(a):
end = len(a)
start = 0
while end > 0:
end -= 1
a[start], a[end] = a[end], a[start]
start += 1
return a
replace([1,2,3,4,5])
# +
def replace(a):
if len(a) <= 1:
return a
else:
return [a[-1]] + replace(a[:-1])
replace([1,2,3,4,5,6])
# +
# Checks whether element occurs in an unsorted array.
def check(num, array):
for elem in array:
if elem == num:
return True
return False
check(3,[2,5,6,8,1])
# +
def occur(num, a):
if a[0] == num:
return True
else:
return occur(a[:-1])
return false
check(2, [2, 8, 5, 3, 1, 9, 6])
# +
# Computes sum of an array of numbers.
def sum(a):
if len(a) == 1:
return a[0]
else:
return a[-1] + sum(a[:-1])
sum([1,2,3,4,5])
# +
# Computes how many times substring appears in the string.
# Iterative
def find(sub, str):
length = len(sub)
count = 0
for char in range(len(str)-1):
if str[char:length] == sub:
count += 1
length += 1
return count
find('he', 'her and herself')
# +
def search(sub,str):
if len(str) == 0 or len(sub) > len(str):
return 0
elif sub == str[:len(sub)]:
return 1 + search(sub,str[1:])
else:
return search(sub,str[1:])
search('he', 'her and herself')
# -
| temp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import json
import pandas as pd
import re
path = os.path.abspath("/group/ag_abi/seiler/IBF/")
pattern = re.compile(r"(\w+)_(\w+)<(\w+), (\w+)<\d+>, (\w+)(?:, Chunks<(\d+)>){0,1}>/(\d+)/(\d+)(?:/(\d+)/(\d+)){0,1}")
# -
# # Insert Benchmark Evaluation
filename = os.path.join(path, "insertCompChunk.json")
with open(filename, "r") as f:
x = json.load(f)
row_list = []
for benchmark in x['benchmarks']:
row_dict = {}
[test, spec, alphabet, strategy, bitvector, chunks, bins, k, ram, h] = re.match(pattern, benchmark['name']).groups()
if ram is None:
ram = int(4**int(k)*int(bins)/1024/1024/8)
else:
ram = int(2**int(ram)/1024/1024/8)
if h is None:
h = 1
time = "{0:,.2f}".format(benchmark['real_time']/10**9/60)
size = "{0:,}".format(int(benchmark['Size']))
row_dict['Function'] = test
row_dict['BD'] = spec
row_dict['Alphabet'] = alphabet
row_dict['Strategy'] = strategy
row_dict['Bitvector'] = bitvector
row_dict['Chunks'] = chunks
row_dict['bins'] = bins
row_dict['k'] = k
row_dict['RAM'] = "{0:,}".format(ram)
row_dict['h'] = h
row_dict['Time'] = time
row_dict['Size'] = size
row_list.append(row_dict)
df = pd.DataFrame(row_list)
df = df[["Function",
"BD",
"Alphabet",
"Strategy",
"Bitvector",
"Chunks",
"bins",
"k",
"h",
"RAM",
"Size",
"Time"]]
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
display(df)
df.to_csv(os.path.join(path, "insertCompChunkBenchmark.tsv"), sep='\t', index=False)
# # Select Benchmark Evaluation
filename = os.path.join(path, "selectCompChunk.json")
with open(filename, "r") as f:
x = json.load(f)
row_list = []
for benchmark in x['benchmarks']:
row_dict = {}
[test, spec, alphabet, strategy, bitvector, chunks, bins, k, ram, h] = re.match(pattern, benchmark['name']).groups()
if ram is None:
ram = int(4**int(k)*int(bins)/1024/1024/8)
else:
ram = int(2**int(ram)/1024/1024/8)
if h is None:
h = 1
#time = round(benchmark['real_time']/10**9,2)
#size = int(benchmark['Size'])
row_dict['Full Time'] = "{0:,.2f}".format(benchmark['fullTime'])
row_dict['load BD'] = "{0:,.2f}".format(benchmark['loadingTime'])
row_dict['load Reads'] = "{0:,.2f}".format(benchmark['ioTime'])
row_dict['sum Select'] = "{0:,.2f}".format(benchmark['selectTime'])
row_dict['avg Select'] = "{0:,.2f}".format(benchmark['selectTime'] / 32)
row_dict['Threads'] = 32
row_dict['TP'] = "{0:,}".format(int(benchmark['TP']))
row_dict['FN'] = "{0:,}".format(int(benchmark['FN']))
row_dict['FP'] = "{0:,}".format(int(benchmark['FP']))
row_dict['P'] = "{0:,}".format(int(benchmark['P']))
row_dict['readNo'] = "{0:,}".format(int(benchmark['readNo']))
row_dict['Absolute Verifications'] = "{0:,}".format(int(benchmark['verifications']))
row_dict['Verifications per read'] = "{0:,.2f}".format(benchmark['Verifications'])
row_dict['Sensitivity'] = benchmark['Sensitivity']
row_dict['Precision'] = benchmark['Precision']
row_dict['FNR'] = "{0:,.2f}".format(benchmark['FNR'])
row_dict['FDR'] = "{0:,.2f}".format(benchmark['FDR'])
row_dict['Function'] = test
row_dict['BD'] = spec
row_dict['Alphabet'] = alphabet
row_dict['Strategy'] = strategy
row_dict['Bitvector'] = bitvector
row_dict['Chunks'] = chunks
row_dict['bins'] = bins
row_dict['k'] = k
row_dict['RAM'] = "{0:,}".format(int(ram))
row_dict['h'] = h
#row_dict['Time'] = time
#row_dict['Size'] = size
row_list.append(row_dict)
df = pd.DataFrame(row_list)
df = df[["Function",
"BD",
"Alphabet",
"Strategy",
"Bitvector",
"Chunks",
"bins",
"k",
"h",
"RAM",
"Full Time",
"load BD",
"load Reads",
"sum Select",
"avg Select",
"TP",
"FN",
"FP",
"P",
"readNo",
"Absolute Verifications",
"Verifications per read",
"Sensitivity",
"Precision",
"FNR",
"FDR",
]]
with pd.option_context('display.max_rows', None, 'display.max_columns', None):
display(df)
df.to_csv(os.path.join(path, "selectCompChunkBenchmark.tsv"), sep='\t', index=False)
| IBF/benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
ews = np.loadtxt('measured_brg_ews.txt',usecols=(1,2,3,4,5))
ews = ews.T
measured_ews = ews[0]
measured_ew_unc = ews[1]
measured_ew_unc_log = 0.434 * (measured_ew_unc / measured_ews)
joel_ews = ews[3]
joel_minus = ews[2]
joel_plus = ews[4]
joel_errs = np.array([joel_ews-joel_minus,joel_plus-joel_ews])
joel_errs_log = 0.434 * (joel_errs / joel_ews)
#plot_bounds = np.array([np.max([measured_ews,joel_plus]),np.max([measured_ews,joel_plus])])
#plt.rcParams.update({'font.size': 15})
fig, ax = plt.subplots()
ax.errorbar(np.log10(joel_ews),np.log10(measured_ews),yerr=measured_ew_unc_log,xerr=joel_errs_log,fmt='None',color='#113166',alpha=0.15)
ax.plot(np.log10(joel_ews),np.log10(measured_ews),'s',color='#006289',ms=7,alpha=0.9)
#ax.plot([0.0001,np.max([measured_ews,joel_plus])],[0.0001,np.max([measured_ews,joel_plus])],color='k',alpha=0.8)
#plt.subplots_adjust(right=0.98,top=0.98)
#ax.set_xscale('log')
#ax.set_yscale('log')
#ax.tick_params(axis='both',which='both',direction='in',top=True,right=True)
ax.set_ylabel('log EW[Br-$\gamma$] (Measured)')
ax.set_xlabel('log EW[Br-$\gamma$] (predicted from photometry)')
ax.set_xlim((-0.3,1.8))
ax.set_ylim((-1.05,2.5))
#ax.set_xticks([-0.5,0.0,0.5,1.0,])
#ax.set_yticks([-0.5,0.0,0.5,1.0,1.5])
ax.plot([-5,5],[-5,5],'k',alpha=0.5)
ax.text(1.6,-0.72,'mean offset = 0.07 dex',horizontalalignment='right')
ax.text(1.6,-0.9,'biweight scatter = 0.27 dex',horizontalalignment='right')
#ax.plot(0.55,-0.48,'v',color='#7c0b0b',alpha=0.8)
plt.show()
#residuals = measured_ews - joel_ews
#std = np.log10(np.std(residuals))
#print(std)
#fig2, ax2 = plt.subplots(figsize=(5,5))
#ax2.errorbar(measured_ews,joel_ews-measured_ews,yerr=joel_errs,xerr=measured_ew_unc,fmt='s',alpha=0.8)
#plt.show()
# +
#p = plt.rcParams.find_all(pattern='labelsize')
#p.update(plt.rcParams.find_all(pattern='font.size'))
p = plt.rcParams.find_all(pattern='size')
p
# -
from astroplots import astroplots
astro = astroplots()
astro.scale_text(3.3333)
astro.scale_text('default')
| saurabh_sample_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Utilisation de base
#
# Importer, hériter `AbstractSessionContainer` et insérer l'objet `session` dans les méthodes l'utilisant avec `attach_session`.
# +
from asynctools import AbstractSessionContainer, attach_session
MATH_API_URL = "http://api.mathjs.org/v4"
class MathRequests(AbstractSessionContainer):
@attach_session
async def get(self, url, session=None, **kwargs):
""" Takes as input the URL and all keywords arguments that aiohttp.get takes:
params - (optional) Dictionary or bytes to be sent in the query
string of the new request
data - (optional) Dictionary, bytes, or file-like object to
send in the body of the request
json - (optional) Any json compatible python object
headers - (optional) Dictionary of HTTP Headers to send with
the request
cookies - (optional) Dict object to send with the request
auth - (optional) BasicAuth named tuple represent HTTP Basic Auth
"""
async with session.get(url, **kwargs) as response:
return await response.text()
async def get_square(self, value):
params = {
"expr" : f"{value}^2"
}
return await self.get(MATH_API_URL, params=params)
# +
import asyncio
values = [1, 2, 3, 4]
async with MathRequests() as maths:
squares = await asyncio.gather(*(maths.get_square(v) for v in values))
print(squares)
# +
from asynctools import AbstractSessionContainer, attach_session
MATH_API_URL = "http://api.mathjs.org/v4"
class SessionContainer(AbstractSessionContainer):
@attach_session
async def get(self, *args, session=None, **kwargs):
return session.get(url, params=params)
# -
| experiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="hrXv0rU9sIma"
# # Autograph: Basic
# In this ungraded lab, you will go through some of the basics of autograph so you can explore what the generated code looks like.
# -
# ## Imports
# + colab={} colab_type="code" id="NiolgWMPgpwI"
import tensorflow as tf
# -
# ## Addition in autograph
# You can use the `@tf.function` decorator to automatically generate the graph-style code as shown below:
# + tags=[]
@tf.function
def add(a, b):
return a + b
a = tf.Variable([[1.,2.],[3.,4.]])
b = tf.Variable([[4.,0.],[1.,5.]])
print(tf.add(a, b))
# See what the generated code looks like
print(tf.autograph.to_code(add.python_function))
# -
# ## if-statements in autograph
# Control flow statements which are very intuitive to write in eager mode can look very complex in graph mode. You can see that in the next examples: first a simple function, then a more complicated one that involves lots of ops and conditionals (fizzbuzz).
# + tags=[]
# simple function that returns the square if the input is greater than zero
@tf.function
def f(x):
if x>0:
x = x * x
return x
print(tf.autograph.to_code(f.python_function))
# -
# ## Fizzbuzz in autograph
#
# You may remember implementing [fizzbuzz](http://wiki.c2.com/?FizzBuzzTest) in preparation for a coding interview.
# - Imagine how much fun it would be if you were asked to impement the graph mode version of that code!
#
# Fortunately, you can just use `@tf.function` and then call `tf.autograph.to_code`!
# + tags=[]
@tf.function
def fizzbuzz(max_num):
counter = 0
for num in range(max_num):
if num % 3 == 0 and num % 5 == 0:
print('FizzBuzz')
elif num % 3 == 0:
print('Fizz')
elif num % 5 == 0:
print('Buzz')
else:
print(num)
counter += 1
return counter
print(tf.autograph.to_code(fizzbuzz.python_function))
# -
fizzbuzz(20)
| C2_custom_training/C2_W3_Lab_1_autograph-basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div>
# <a href="https://bokeh.org/"><img src="images/bokeh-header.png"></a>
# </div>
# Welcome to [Bokeh](https://bokeh.org) in the Jupyter Notebook!
#
# Bokeh is a Python interactive visualization library that targets modern web browsers for presentation. Its goal is to provide elegant, concise construction of novel graphics in the style of D3.js, and to extend this capability with high-performance interactivity over very large or streaming datasets. Bokeh can help anyone who would like to quickly and easily create interactive plots, dashboards, and data applications.
# ## Quickstart
#
# Get started with a [5-min introduction to Bokeh](quickstart/quickstart.ipynb).
# ## Tutorial
#
# Start with the [Introduction and Setup](tutorial/00%20-%20Introduction%20and%20Setup.ipynb) notebook and jump to any of the specific topic sections from there.
# ## More information
#
# Find more details and contact information at https://bokeh.org.
#
# ## Thanks
#
# Bokeh was originally developed with financial support from [Anaconda, Inc.](https://anaconda.com) and the Darpa XDATA initiative, and continues due to support from NumFOCUS and individual community contributions. Many thanks to [all of the Bokeh Github contributors](https://github.com/bokeh/bokeh/graphs/contributors).
# <script>
# (function(i,s,o,g,r,a,m){i['GoogleAnalyticsObject']=r;i[r]=i[r]||function(){
# (i[r].q=i[r].q||[]).push(arguments)},i[r].l=1*new Date();a=s.createElement(o),
# m=s.getElementsByTagName(o)[0];a.async=1;a.src=g;m.parentNode.insertBefore(a,m)
# })(window,document,'script','//www.google-analytics.com/analytics.js','ga');
#
# ga('create', 'UA-27761864-9', 'auto');
# ga('send', 'pageview');
# </script>
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: weldx
# language: python
# name: weldx
# ---
# # 3D Geometries
#
# ## Introduction
#
# In this tutorial we will describe how to create 3 dimensional structures that are based on one or more 2d profiles.
# We assume that you already know how to create 2d profiles using the `weldx` package.
# If this is not the case, please read the corresponding tutorial first.
#
# You will learn:
#
# - about the `Trace` class
# - how to define a 3 dimensional geometry using the `Geometry` class
# - how to specify geometries with varying cross sections using the `VariableProfile` class
#
# Before we start, run the following code cells to include everything we need.
# + jupyter={"outputs_hidden": false} nbsphinx="hidden" pycharm={"name": "#%%\n"}
# if the package is not installed in your python environment, run this to execute the notebook directly from inside the GitHub repository
# %cd -q ..
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
import numpy as np
from mpl_toolkits.mplot3d import Axes3D # noqa: F401 unused import
import matplotlib.pyplot as plt
from weldx.transformations import LocalCoordinateSystem
from weldx.geometry import (Geometry,
LinearHorizontalTraceSegment,
linear_profile_interpolation_sbs,
Profile,
RadialHorizontalTraceSegment,
Shape,
Trace,
VariableProfile
)
import weldx.transformations as tf
from weldx import Q_
# -
# ## Trace
#
# The `Trace` class describes an arbitrary path through the 3 dimensional space.
# It is build from multiple segments that need to be passed to it when we create the `Trace`.
# Currently there are 2 segment types available: the `LinearHorizontalTraceSegment` that represents a straight line and
# the `RadialHorizontalTraceSegment` which describes a circular path.
# Both segment types have in common that the `z`-value remains constant and that they are free from torsion.
#
# Let's create one instance of each segment type.
# All you need to specify when creating a `LinearHorizontalTraceSegment` is its length:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
line_segment = LinearHorizontalTraceSegment(length=Q_(10, "cm"))
# -
# That's it.
# The `RadialHorizontalTraceSegment` needs a little bit more information.
# You need to provide its radius, rotation angle and if the rotation is clockwise or counter-clockwise:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
arc_segment = RadialHorizontalTraceSegment(radius=Q_(15, "cm"),
angle=Q_(90, "degree"),
clockwise=True)
# -
# Now that we have some segments, we can create a `Trace`.
# It expects a single segment or a list of segments as first argument.
# The individual segments get attached to each other in the same order as they are attached to the list.
# Each segment except for the first one gets its start orientation and coordinates from the end of the previous segment.
# The initial orientation and coordinates of the `Trace` can be provided in form of a `LocalCoordinateSystem` as optional
# second argument during the creation of the class instance.
# If you omit the second parameter, the `Trace` starts at the origin and is oriented in positive x-direction.
#
#
# Here is an example:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
trace = Trace([line_segment, arc_segment,line_segment])
# -
# We can plot our `Trace` using the `plot` method:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
trace.plot(raster_width=Q_(2, "cm"))
# -
# Let's provide a initial coordinate system that is rotated by 45 degrees around the y-axis and see how that affects the
# result:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
lcs_rot = LocalCoordinateSystem(tf.rotation_matrix_y(Q_(45, "degree")))
trace_rot = Trace([line_segment, arc_segment,line_segment], lcs_rot)
trace_rot.plot(raster_width=Q_(2, "cm"))
# -
# Linear and radial trace segments already cover a lot of use cases but they are far from being enough to cover all
# possible needs.
# If you need more than those two basic segment types, there are two options.
# The first one is to implement your own segment type.
# This is discussed in detail in another tutorial.
# > TODO: Add link to tutorial once it is written
#
# The second option, that you should prefer is the `TODO: SOME_FANCY_NAME_SEGMENT`.
# It enables you to describe the course of the trace segment with a mathematical function.
# > TODO: Continue once the segment type is implemented
# ## Describing a 3d geometry
#
# The simplest possible way to describe a 3 dimensional geometry using the `weldx.geometry` package is to create a 2d
# `Profile` and extrude it into 3d.
# The class that performs this action is called `Geometry`.
# It expects a `Profile` that defines the cross section as first argument and a `Trace` that describes
# the extrusion path as second one.
# Since we already have created a `Trace` all that is missing is a `Profile`.
# So let's create one:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
shape_right = Shape().add_line_segments(Q_([[10, 8], [5, 8], [1, 2], [1, 0], [10, 0]], "cm"))
shape_left = shape_right.reflect_across_line([0, 0], [0, 1])
profile = Profile([shape_left, shape_right])
profile.plot(raster_width=Q_(1, "cm"))
# -
# Now we can create and plot our `Geometry`:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
geometry = Geometry(profile=profile, trace=trace)
geometry.plot(profile_raster_width=Q_(1, "cm"),
trace_raster_width=Q_(10, "cm"))
trace.plot(raster_width=Q_(1, "cm"), axes=plt.gca(), fmt="r-")
# -
# ## Geometries with varying cross section
#
# The `VariableProfile` class gives us the opportunity to handle 3 dimensional geometries with a varying cross section.
# It can be used instead of a `Profile` during the creation of the `Geometry` class.
# Three pieces of information are necessary to create a `VariableProfile`.
#
# First a list of profiles is required.
# This list represents "snapshots" of known cross sections along the extrusion path.
#
# The second piece of information is a list of numbers that has the same number of elements as the list of profiles.
# Each of those numbers defines a coordinate on a 1 dimensional axis for the profile with the same list index.
# Their magnitude can be chosen arbitrarily with the exception that the first entry has to be 0 and the list items must be
# in ascending order.
# Due to the previously mentioned association between the coordinates/numbers and the profiles, the profiles are ordered
# by ascending coordinates.
# When you pass a `VariableProfile` to a `Geometry`, its first profile is used as the initial profile at the start of
# the `Trace` and the last as the one at the end.
# This relation enables the `Geometry` class to establish a linear coordinate transformation between the coordinates you
# specified in the `VariableProfile` and the corresponding position on the trace.
# For example, if you use three profiles with the coordinates `[0, 25, 100]` and your `Trace` has a length of $40 mm$,
# then the second profile occurs after $10 mm$ on the `Trace`.
#
# The last thing we need to provide in order to create a `VariableProfile` is how the space between two profiles gets
# interpolated.
# We pass this information is form of a list that contains the corresponding interpolation functions.
# For each pair of profiles we need to specify an interpolation where the first list entry is associated with the first
# pair, the second entry with the second pair and so on.
# The number of list items is therefore one less than the number of profiles.
#
# Lets have look at an example after this rather long text section. First, we define two more profiles.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
shape_right_2 = Shape().add_line_segments(Q_([[10, 8], [5, 8], [5, 2], [5, 0], [10, 0]], "cm"))
shape_left_2 = shape_right_2.reflect_across_line([0, 0], [0, 1])
profile_2 = Profile([shape_left_2, shape_right_2])
shape_right_3 = Shape().add_line_segments(Q_([[10, 12], [2, 12], [4, 4], [4, 0], [10, 0]], "cm"))
shape_left_3 = shape_right_3.reflect_across_line([0, 0], [0, 1])
profile_3 = Profile([shape_left_3, shape_right_3])
profile.plot(raster_width=Q_(1, "cm"), label="Profile 1")
profile_2.plot(raster_width=Q_(1, "cm"), label="Profile 2", ax=plt.gca())
profile_3.plot(raster_width=Q_(1, "cm"), label="Profile 3", ax=plt.gca())
plt.gca().legend()
# -
# Now we create a `VariableProfile`, pass it to a new `Geometry` and plot it:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
v_profile = VariableProfile([profile_3, profile_2, profile],
[0, 40, 100],
[linear_profile_interpolation_sbs, linear_profile_interpolation_sbs]
)
geometry_vp = Geometry(profile=v_profile,trace=trace)
geometry_vp.plot(profile_raster_width=Q_(1, "cm"),
trace_raster_width=Q_(8, "cm"))
# -
# ## Custom interpolation schemes
#
# You might notice that we used a interpolation function called `linear_profile_interpolation_sbs`.
# The "sbs" stands for "segment-by-segment".
# It interpolates all of the profile segments linearly in the order as they appear in both involved profiles.
# So you need to assure that the segments are ordered accordingly in both profiles.
# It also requires that the number of segments is identical and it only works with the `LineSegment`.
#
# You might wonder what alternative functions are provided, but currently there are none.
# The reason is that as soon as you start thinking about it, there are so many options that need to be considered that
# don't have a unique solution that fits all use cases.
# To circumvent this issue, we provide the possibility to use custom interpolation functions.
# All that you need to do in order to use your own interpolation scheme is to define a python
# function with the following interface:
#
# ~~~
# def custom_interpolation(Profile, Profile, float) -> Profile
# ~~~
#
#
# The `float` parameter is a weighting factor representing the position on the trace segment between both profiles.
# The `Geometry` class will pass values between `0` and `1` where `0` is the beginning of the segment and associates with
# the first passed `Profile`.
# In conclusion, a value of `1` represents the end of the segment and is associated with the second `Profile`.
# What you do with this information is up to you.
# All that matters is that the function returns a valid `Profile` in the end.
#
# Lets create an example interpolation that returns the first passed profile for weighting factors below `0.4` and the
# second one otherwise:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
def my_interp(p0, p1, weight):
if weight < 0.4:
return p0
return p1
# + [markdown] pycharm={"name": "#%% md\n"}
# Now we just create a `VariableProfile` with our new interpolation, combine it with a linear `Trace` inside of a
# `Geometry` and plot it:
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
v_profile_custom_interp = VariableProfile([profile_3, profile_2], [0, 1], my_interp)
trace_custom_interp = Trace(LinearHorizontalTraceSegment(Q_(20, "cm")))
geometry_custom_interp = Geometry(v_profile_custom_interp, trace_custom_interp)
geometry_custom_interp.plot(profile_raster_width=Q_(1, "cm"),
trace_raster_width=Q_(2, "cm"))
| tutorials/geometry_02_geometry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
base_dir = 'C:\\Users\heine\github\PlateReaderTools'
os.chdir(base_dir)
#import sys
from core import plate_reader_tools
import pandas as pd
import numpy as np
# import math
# import matplotlib
import matplotlib.pyplot as plt
import string
import seaborn as sns
from itertools import chain
# import os
# import sys
# path = '/Users/andrewng/Documents/PlateReaderTools'
# if not path in sys.path:
# sys.path.insert(1, path)
# del path
# from plate_reader_tools import *
# import seaborn as sns
# sns.set_style("white")
# sns.set_style("ticks")
# %load_ext autoreload
# %autoreload 2
# %matplotlib notebook
# #%matplotlib inline
# -
dirname = "C:\\Users\\heine\\Google Drive\\UCSF\\ElSamad_Lab\\CRISPR iGRNA\\experiments\\plate_reader\\"
fname = dirname + "20170504_BMH_24H_OD_mRFP_sfGFP_Venus_20170505_141316.xlsx"
#date = fname[-13:-5]
sheetname = 'Result sheet'
skiprows = 100 #skip until the row that has the label (i.e. OD) for the readout.
rows = 8
columns = 12
datalabels = ['OD','sfGFP','mRFP','Venus']
cycles = 95
horz = 0
fname
[d,time_list] = plate_reader_tools.readplate(fname,sheetname, skiprows, rows, columns, datalabels, cycles, horz)
OD_data = d['OD']
sfGFP_data = d['sfGFP']
# +
OD_data_filtered, OVER_list = over_filter(OD_data)
# OVER_list = [index for (index, value) in enumerate(OD_data.isin(['OVER']).any()) if value]
# if len(OVER_list)==0:
# print('No OVER values in data')
# else:
# print('OVER value found in well(s): \n' + '\n'.join(OVER_list))
# OD_data_filtered = OD_data.replace('OVER',np.nan)
| scripts/Ben/.ipynb_checkpoints/20170504_igrna_CCW12_test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="text-align:center">Undamped Response to Harmonic Direct-Force Inputs</h1>
# <h3 style="text-align:center"> MCHE 485: Mechanical Vibrations</h3>
# <p style="text-align:center">Dr. <NAME> <br>
# <a href="mailto:<EMAIL>"><EMAIL></a><br>
# http://www.ucs.louisiana.edu/~jev9637/ </p>
# <p style="text-align:center">
# <img src="http://shared.crawlab.org/MassSpring_DirectForce_Horiz.png" alt="A Mass-Spring-Damper System" width=50%></a><br>
# <strong> Figure 1: A Mass-Spring System </strong>
# </p>
#
# This notebook examines the frequency response of a simple mass-spring system like the one shown in Figure 1 to a harmonic, direct-force input.
#
# The equation of motion for the system is:
#
# <!-- the \quad commmand just adds a space in the math mode -->
# $ \quad m \ddot{x} + kx = f $
#
# We could also write this equation in terms of the damping ratio, $\zeta$, and natural frequency, $\omega_n$.
#
# $ \quad \ddot{x} + \omega_n^2x = \frac{f}{m}$
#
# For information on how to obtain this equation, you can see the lectures at the [class website](http://www.ucs.louisiana.edu/~jev9637/MCHE485.html).
import numpy as np # Grab all of the NumPy functions with nickname np
# We want our plots to be displayed inline, not in a separate window
# %matplotlib inline
# Import the plotting functions
import matplotlib.pyplot as plt
# Define the System Parameters
m = 1.0 # kg
k = (2.0 * np.pi)**2. # N/m (Selected to give an undamped natrual frequency of 1Hz)
wn = np.sqrt(k / m) # Natural Frequency (rad/s)
# Let's use the closed-form, steady-state solution we developed in lecture:
#
# Assume:
#
# $ \quad f(t) = \bar{f} \sin{\omega t} $
#
# Then, the solution $x(t)$ should have the form:
#
# $ \quad x(t) = \bar{x} \sin{\omega t} $
#
# We can then find the amplitude of the frequency response, $ \bar{x} $, as a function of of the frequency of the input, $ \omega $, and the amplitude of the force, $ \bar{f} $.
#
# $ \quad \bar{x} = \frac{\bar{f}}{m} \left(\frac{1}{\omega_n^2 - \omega^2}\right) $
#
# So,
#
# $ \quad x(t) = \frac{\bar{f}}{m} \left(\frac{1}{\omega_n^2 - \omega^2}\right) \sin{\omega t} $
#
# or
#
# $ \quad x(t) = \frac{1}{m} \left(\frac{1}{\omega_n^2 - \omega^2} \right) f(t) $
#
# So, $ \left(\frac{1/m}{\omega_n^2 - \omega^2} \right) $ gives us the relationship between the input $ f(t) $ and the system response $ x(t) $. Let's plot that for a range of frequencies.
# +
# Set up input parameters
w = np.linspace(1e-6, wn*3, 1000) # Frequency range for freq response plot, 0-3x wn with 1000 points in-between
x_amp = (1/m) / (wn**2 - w**2)
# Let's mask the discontinuity, so it isn't plotted
pos = np.where(np.abs(x_amp) >= 5)
x_amp[pos] = np.nan
w[pos] = np.nan
# +
# Make the figure pretty, then plot the results
# "pretty" parameters selected based on pdf output, not screen output
# Many of these setting could also be made default by the .matplotlibrc file
fig = plt.figure(figsize=(6,4))
ax = plt.gca()
plt.subplots_adjust(bottom=0.2,left=0.15,top=0.96,right=0.96)
plt.setp(ax.get_ymajorticklabels(),family='serif',fontsize=18)
plt.setp(ax.get_xmajorticklabels(),family='serif',fontsize=18)
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.grid(True,linestyle=':',color='0.75')
ax.set_axisbelow(True)
plt.xlabel(r'Input Frequency $\left(\omega\right)$',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylabel(r'$ \frac{1}{m\left(\omega_n^2 - \omega^2\right)} $',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylim(-1.0,1.0)
plt.xticks([1],['$\omega = \omega_n$'])
plt.yticks([0])
plt.plot(w/wn,x_amp,linewidth=2)
# If you want to save the figure, uncomment the commands below.
# The figure will be saved in the same directory as your IPython Notebook.
# Save the figure as a high-res pdf in the current folder
# plt.savefig('MassSpring_ForcedFreqResp_Amplitude.pdf',dpi=300)
fig.set_size_inches(9,6) # Resize the figure for better display in the notebook
# -
# ### Magnitude of the Response
# We can also plot the magnitude of this.
# +
x_mag = np.abs(x_amp)
# Make the figure pretty, then plot the results
# "pretty" parameters selected based on pdf output, not screen output
# Many of these setting could also be made default by the .matplotlibrc file
fig = plt.figure(figsize=(6,4))
ax = plt.gca()
plt.subplots_adjust(bottom=0.2,left=0.15,top=0.96,right=0.96)
plt.setp(ax.get_ymajorticklabels(),family='serif',fontsize=18)
plt.setp(ax.get_xmajorticklabels(),family='serif',fontsize=18)
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.grid(True,linestyle=':',color='0.75')
ax.set_axisbelow(True)
plt.xlabel(r'Input Frequency $\left(\omega\right)$',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylabel(r'$\left| \frac{1}{m\left(\omega_n^2 - \omega^2\right)\right|} $',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylim(0.0,0.25)
plt.xticks([1],[r'$\omega = \omega_n$'])
plt.yticks([0])
plt.plot(w/wn, x_mag, linewidth=2)
# If you want to save the figure, uncomment the commands below.
# The figure will be saved in the same directory as your IPython Notebook.
# Save the figure as a high-res pdf in the current folder
# plt.savefig('MassSpring_ForcedFreqResp_Magnitude.pdf',dpi=300)
fig.set_size_inches(9,6) # Resize the figure for better display in the notebook
# -
# ### Normalization
# Just as we did for seismic inputs, we can also normalize the frequency response by dividing both the numerator and denominator of the expression for $\bar{x}$ by the natural frequency $ \omega_n $. We find that:
#
# $ \quad \bar{x} = \frac{\bar{f}}{m \omega_n^2 \left( 1 - \Omega^2\right)}$
#
# As a final normalization step, we can normalize the amplitude by plotting $\frac{m \omega_n^2}{\bar{f}} \bar{x}$ as a function of $\Omega$.
# +
# Set up input parameters
wnorm = np.linspace(0,4,500) # Frequency range for freq response plot, 0-4 Omega with 500 points in-between
x_amp = 1 / ((wn**2 * m) * (1 - wnorm**2))
xnorm_amp = x_amp * (m * wn**2)
# Let's mask the discontinuity, so it isn't plotted
pos = np.where(np.abs(xnorm_amp) >= 100)
xnorm_amp[pos] = np.nan
wnorm[pos] = np.nan
# +
# Make the figure pretty, then plot the results
# "pretty" parameters selected based on pdf output, not screen output
# Many of these setting could also be made default by the .matplotlibrc file
fig = plt.figure(figsize=(6,4))
ax = plt.gca()
plt.subplots_adjust(bottom=0.2,left=0.15,top=0.96,right=0.96)
plt.setp(ax.get_ymajorticklabels(),family='serif',fontsize=18)
plt.setp(ax.get_xmajorticklabels(),family='serif',fontsize=18)
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.grid(True,linestyle=':',color='0.75')
ax.set_axisbelow(True)
plt.xlabel(r'Normalized Frequency $\left(\Omega\right)$',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylabel(r'$\frac{m \omega_n^2}{\bar{f}} \bar{x}$',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylim(-4.0,4.0)
plt.xticks([0,1],['0','1'])
plt.yticks([0,1])
plt.plot(wnorm,xnorm_amp,linewidth=2)
# If you want to save the figure, uncomment the commands below.
# The figure will be saved in the same directory as your IPython notebook.
# Save the figure as a high-res pdf in the current folder
# plt.savefig('MassSpring_ForcedFreqResp_NormAmp.pdf',dpi=300)
fig.set_size_inches(9,6) # Resize the figure for better display in the notebook
# -
# ### Magnitude of the Response
# We can also plot the magnitude of this.
# +
xnorm_mag = np.abs(xnorm_amp)
# Make the figure pretty, then plot the results
# "pretty" parameters selected based on pdf output, not screen output
# Many of these setting could also be made default by the .matplotlibrc file
fig = plt.figure(figsize=(6,4))
ax = plt.gca()
plt.subplots_adjust(bottom=0.2,left=0.15,top=0.96,right=0.96)
plt.setp(ax.get_ymajorticklabels(),family='serif',fontsize=18)
plt.setp(ax.get_xmajorticklabels(),family='serif',fontsize=18)
ax.spines['right'].set_color('none')
ax.spines['top'].set_color('none')
ax.xaxis.set_ticks_position('bottom')
ax.yaxis.set_ticks_position('left')
ax.grid(True,linestyle=':',color='0.75')
ax.set_axisbelow(True)
plt.xlabel(r'Normalized Frequency $\left(\Omega\right)$',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylabel(r'$\left| \frac{m \omega_n^2}{\bar{f}} \bar{x} \right|$',family='serif',fontsize=22,weight='bold',labelpad=10)
plt.ylim(0.0,5.0)
plt.xticks([0,1],['0','1'])
plt.yticks([0,1])
plt.plot(wnorm, xnorm_mag, linewidth=2)
# If you want to save the figure, uncomment the commands below.
# The figure will be saved in the same directory as your IPython notebook.
# Save the figure as a high-res pdf in the current folder
# savefig('MassSpring_ForcedFreqResp_NormMag.pdf',dpi=300)
fig.set_size_inches(9,6) # Resize the figure for better display in the notebook
# -
# <hr style="border: 0px;
# height: 1px;
# text-align: center;
# background: #333;
# background-image: -webkit-linear-gradient(left, #ccc, #333, #ccc);
# background-image: -moz-linear-gradient(left, #ccc, #333, #ccc);
# background-image: -ms-linear-gradient(left, #ccc, #333, #ccc);
# background-image: -o-linear-gradient(left, #ccc, #333, #ccc);">
# #### Licenses
# Code is licensed under a 3-clause BSD style license. See the licenses/LICENSE.md file.
#
# Other content is provided under a [Creative Commons Attribution-NonCommercial 4.0 International License](http://creativecommons.org/licenses/by-nc/4.0/), CC-BY-NC 4.0.
# This cell will just improve the styling of the notebook
# You can ignore it, if you are okay with the default sytling
from IPython.core.display import HTML
import urllib.request
response = urllib.request.urlopen("https://cl.ly/1B1y452Z1d35")
HTML(response.read().decode("utf-8"))
| Jupyter Notebooks/Undamped Response to Harmonic Direct-Force Inputs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Single Target Regression Example
# This example will show you how to use one of the provided data sets and ```amorf.problemTransformation.SingleTargetMethod``` to perform a basic multi-output Regression.
#
# The Single Target Method trains an estimator for each target variable, predicts each target variable and concatenates the results to yield a multi output regression.
#
# ## Setting Up Training and Test Set
# +
from amorf.datasets import RiverFlow1
from sklearn.model_selection import train_test_split
X, y = RiverFlow1().get_numpy()
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, shuffle=True)
# -
# ## Normalize Data
# +
from sklearn.preprocessing import normalize
X_train = normalize(X_train)
X_test = normalize(X_test)
# -
# ## Initialize all Selectors, Perform Prediction and Calculate Error
selectors = ['linear', 'kneighbors',
'adaboost', 'gradientboost', 'mlp', 'svr', 'xgb']
from amorf.problemTransformation import SingleTargetMethod
from amorf.metrics import average_relative_root_mean_squared_error
for selector in selectors:
regressor = SingleTargetMethod(selector)
regressor.fit(X_train, y_train)
result = regressor.predict(X_test)
print(selector)
print(average_relative_root_mean_squared_error(result, y_test))
# ## Use Custom Regressor
#
# +
from sklearn.linear_model import RidgeCV
ridgeCV = RidgeCV()
regressor = SingleTargetMethod(custom_regressor=ridgeCV)
regressor.fit(X_train,y_train)
result = regressor.predict(X_test)
print(average_relative_root_mean_squared_error(result,y_test))
# -
#
| examples/05_singleTarget_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Iris-Tutorial local
# language: python
# name: Iris-Tutorial_local
# ---
# Use the Azure Machine Learning data collector to log various metrics
from azureml.logging import get_azureml_logger
logger = get_azureml_logger()
# +
# Use Azure Machine Learning history magic to control history collection
# History is off by default, options are "on", "off", or "show"
# # %azureml history on
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
CODE_DIR = 'code'
sys.path.append(f"{os.path.join(os.getcwd(), CODE_DIR)}")
# -
# !pip install -r requirements.txt
# !python code/pretrain_DAMSM.py --cfg code/cfg/DAMSM/bird.yml --data_dir data/birds --gpu 0
# !python code/main.py --cfg code/cfg/eval_bird.yml --data_dir data/birds --gpu 0
# !python code/main.py --cfg code/cfg/bird_attn2.yml --data_dir data/birds --gpu 0
| Playground.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # In Class Notebook, Week 16
# You can always paste the URL of this notebook (https://github.com/UIUC-iSchool-DataViz/is445_AOUAOG_fall2021/blob/master/week16/inClass_week16.ipynb ) into the nbviewer interface for a plain-text rendering:
#
# https://kokes.github.io/nbviewer.js/viewer.html
import nltk
# +
# nltk.download('stopwords')
# nltk.download('punkt')
# nltk.download('words')
# +
from nltk.corpus import stopwords, words
import string
import itertools
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
import matplotlib.pyplot as plt
# -
filename = '/Users/jillnaiman/Downloads/othello.txt'
with open(filename) as f:
text = f.read()
# +
#text.split('\n') # individual lines
# -
myWords = text.split()
myWords
wordsDict = {} # store words and their frequency
for word in myWords: # loop over all text
if word in wordsDict: # if the word is already in dictionary
wordsDict[word] += 1 #... update count
else:# otherwise
wordsDict[word] = 1 # ... add to dictionary
wordsDict
import operator
wordsDict = sorted(wordsDict.items(), # I want to sort each key-value pair in the dictionary
key=operator.itemgetter(1)) # sort by the 2nd of those -- i.e. the value
# this returns a list object
wordsDict[::-1] # list of tuples of key-value pairs, ordered with most frequent on top
# back into a dictionary, that has now been sorted
wordsDict = dict(wordsDict[::-1])
wordsDict
# +
# finally, a plot!
n = 50 # 50 most frequent words in Othello
wordsPlot = {k:wordsDict[k] for k in list(wordsDict)[:n]} # grabbing only the nth most frequent words into a dict.
fig, ax = plt.subplots(figsize=(16,5))
ax.bar(range(len(wordsPlot)), list(wordsPlot.values()), align='center')
plt.show()
# +
# finally, a plot!
n = 50 # 50 most frequent words in Othello
wordsPlot = {k:wordsDict[k] for k in list(wordsDict)[:n]} # grabbing only the nth most frequent words into a dict.
fig, ax = plt.subplots(figsize=(16,5))
ax.bar(range(len(wordsPlot)), list(wordsPlot.values()), align='center')
# add a tickmark for each bar
ax.set_xticks(range(len(wordsPlot)))
plt.show()
# +
# finally, a plot!
n = 50 # 50 most frequent words in Othello
wordsPlot = {k:wordsDict[k] for k in list(wordsDict)[:n]} # grabbing only the nth most frequent words into a dict.
fig, ax = plt.subplots(figsize=(16,5))
ax.bar(range(len(wordsPlot)), list(wordsPlot.values()), align='center')
# add a tickmark for each bar
ax.set_xticks(range(len(wordsPlot)))
# replace tick numbers by words
ax.set_xticklabels(list(wordsPlot.keys()))
plt.show()
# +
# finally, a plot!
n = 50 # 50 most frequent words in Othello
wordsPlot = {k:wordsDict[k] for k in list(wordsDict)[:n]} # grabbing only the nth most frequent words into a dict.
fig, ax = plt.subplots(figsize=(16,3))
ax.bar(range(len(wordsPlot)), list(wordsPlot.values()), align='center')
# add a tickmark for each bar
ax.set_xticks(range(len(wordsPlot)))
# replace tick numbers by words
ax.set_xticklabels(list(wordsPlot.keys()))
plt.xticks(rotation=90)
plt.show()
# -
len(wordsDict)
stop_words=set(stopwords.words('english'))
stop_words
less_words = {}
for w,num in wordsDict.items():
if w.lower() not in stop_words:
less_words[w] = num
len(less_words)
# +
# finally, a plot!
n = 50 # 50 most frequent words in Othello
wordsPlot = {k:less_words[k] for k in list(less_words)[:n]} # grabbing only the nth most frequent words into a dict.
fig, ax = plt.subplots(figsize=(16,3))
ax.bar(range(len(wordsPlot)), list(wordsPlot.values()), align='center')
# add a tickmark for each bar
ax.set_xticks(range(len(wordsPlot)))
# replace tick numbers by words
ax.set_xticklabels(list(wordsPlot.keys()))
plt.xticks(rotation=90)
plt.show()
# -
# we going to be a little fancier in our splitting of text data, using the NLTK package (spaCy is another NLP package)
word_tokens = nltk.word_tokenize(" ".join(myWords))
word_tokens
less_words = [w for w in word_tokens if not w.lower() in stop_words]
# make a new list if that particular word in word_tokens is not also in stop_words
less_words
string.punctuation
no_punc_words = []
for word in less_words:
if word not in string.punctuation:
no_punc_words.append(word)
no_punc_words
# Note: could be removing more things like 's or 'd or --, but moving on for now.
wordsDict = {}
for word in no_punc_words:
if word in wordsDict:
wordsDict[word] += 1
else:
wordsDict[word] = 1
wordsDict
wordsDict = sorted(wordsDict.items(), # I want to sort each key-value pair in the dictionary
key=operator.itemgetter(1)) # sort by the 2nd of those -- i.e. the value
wordsDict = dict(wordsDict[::-1])
wordsDict
# +
# finally, a plot!
n = 50 # 50 most frequent words in Othello
wordsPlot = {k:wordsDict[k] for k in list(wordsDict)[:n]} # grabbing only the nth most frequent words into a dict.
fig, ax = plt.subplots(figsize=(16,3))
ax.bar(range(len(wordsPlot)), list(wordsPlot.values()), align='center')
# add a tickmark for each bar
ax.set_xticks(range(len(wordsPlot)))
# replace tick numbers by words
ax.set_xticklabels(list(wordsPlot.keys()))
plt.xticks(rotation=90)
plt.show()
# -
wordcloud = WordCloud().generate_from_frequencies(wordsDict)
fig, ax = plt.subplots(figsize=(10,5))
ax.imshow(wordcloud)
plt.show()
# # Time for network viz!
import bqplot
import ipywidgets
import pandas as pd
# +
# data
node_data = [
{"label": "<NAME>", "media": "Star Wars", "shape": "rect"},
{"label": "<NAME>", "media": "Star Trek", "shape": "rect"},
{"label": "Doctor Who", "media": "Doctor Who", "shape": "rect"},
{"label": "Pikachu", "media": "Detective Pikachu", "shape": "circle"},
]
#mark that is using bqplot's Graph
graph = bqplot.Graph(node_data=node_data,
colors=["red","red","red","red"])
fig = bqplot.Figure(marks=[graph])
fig
# +
# data
# nodes
node_data = [
{"label": "<NAME>", "media": "Star Wars", "shape": "rect"},
{"label": "<NAME>", "media": "Star Trek", "shape": "rect"},
{"label": "Doctor Who", "media": "Doctor Who", "shape": "rect"},
{"label": "Pikachu", "media": "Detective Pikachu", "shape": "circle"},
]
# links
# link <NAME> (0th node) to <NAME> (1st node) AND Pikachu (3rd node)
link_data = [{'source':0, 'target':1}, {'source':0, 'target':3}]
#mark that is using bqplot's Graph
graph = bqplot.Graph(node_data=node_data, link_data=link_data,
colors=["red","red","red","red"])
# link lines instead of link arcs
graph.link_type = 'line'
graph.directed = False # not directionally linked
graph.charge = -300
fig = bqplot.Figure(marks=[graph])
fig
# -
# ### Facebook data
# +
filename = 'facebook_combined_sm000090_000010.txt' # broad
filename = 'facebook_combined_sm000030_000000.txt' # more central
network = pd.read_csv('/Users/jillnaiman/Downloads/'+filename, sep=' ', names=['ind1','ind2'])
# -
network = network.drop_duplicates()
network
import numpy as np
node_data = []
link_data = []
color_data = []
# how many unique nodes?
u_nodes = np.unique(np.append(network['ind1'].values, network['ind2'].values))
u_nodes
for un in u_nodes:
node_data.append({'label':str(un),"shape":"circle"})
node_data[:5]
for iu,un in enumerate(u_nodes): # for each of these nodes
target_ids = network.loc[network['ind1']==un] # taking all rows with this unique node number
tnodes = np.unique(target_ids['ind2'].values) # all of the unique nodes that are linked with un
for t in tnodes:
tind = np.where(t==u_nodes)[0][0] # index in u_nodes that is this target
link_data.append({'source':iu, 'target':tind})
color_data.append('blue')
# +
graph = bqplot.Graph(node_data=node_data, link_data=link_data, colors=color_data)
graph.link_type='line'
graph.link_distance=50
graph.charge=-200
graph.directed=False
fig = bqplot.Figure(marks=[graph])
fig.layout.min_width='1000px'
fig.layout.min_height='900px'
fig
# -
| week16/inClass_week16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Residual Networks
#
# Welcome to the first assignment of this week! You'll be building a very deep convolutional network, using Residual Networks (ResNets). In theory, very deep networks can represent very complex functions; but in practice, they are hard to train. Residual Networks, introduced by [He et al.](https://arxiv.org/pdf/1512.03385.pdf), allow you to train much deeper networks than were previously feasible.
#
# **By the end of this assignment, you'll be able to:**
#
# - Implement the basic building blocks of ResNets in a deep neural network using Keras
# - Put together these building blocks to implement and train a state-of-the-art neural network for image classification
# - Implement a skip connection in your network
#
# For this assignment, you'll use Keras.
#
# Before jumping into the problem, run the cell below to load the required packages.
# ## Table of Content
#
# - [1 - Packages](#1)
# - [2 - The Problem of Very Deep Neural Networks](#2)
# - [3 - Building a Residual Network](#3)
# - [3.1 - The Identity Block](#3-1)
# - [Exercise 1 - identity_block](#ex-1)
# - [3.2 - The Convolutional Block](#3-2)
# - [Exercise 2 - convolutional_block](#ex-2)
# - [4 - Building Your First ResNet Model (50 layers)](#4)
# - [Exercise 3 - ResNet50](#ex-3)
# - [5 - Test on Your Own Image (Optional/Ungraded)](#5)
# - [6 - Bibliography](#6)
# <a name='1'></a>
# ## 1 - Packages
# +
import tensorflow as tf
import numpy as np
import scipy.misc
from tensorflow.keras.applications.resnet_v2 import ResNet50V2
from tensorflow.keras.preprocessing import image
from tensorflow.keras.applications.resnet_v2 import preprocess_input, decode_predictions
from tensorflow.keras import layers
from tensorflow.keras.layers import Input, Add, Dense, Activation, ZeroPadding2D, BatchNormalization, Flatten, Conv2D, AveragePooling2D, MaxPooling2D, GlobalMaxPooling2D
from tensorflow.keras.models import Model, load_model
from resnets_utils import *
from tensorflow.keras.initializers import random_uniform, glorot_uniform, constant, identity
from tensorflow.python.framework.ops import EagerTensor
from matplotlib.pyplot import imshow
from test_utils import summary, comparator
import public_tests
# %matplotlib inline
# -
# <a name='2'></a>
# ## 2 - The Problem of Very Deep Neural Networks
#
# Last week, you built your first convolutional neural networks: first manually with numpy, then using Tensorflow and Keras.
#
# In recent years, neural networks have become much deeper, with state-of-the-art networks evolving from having just a few layers (e.g., AlexNet) to over a hundred layers.
#
# * The main benefit of a very deep network is that it can represent very complex functions. It can also learn features at many different levels of abstraction, from edges (at the shallower layers, closer to the input) to very complex features (at the deeper layers, closer to the output).
#
# * However, using a deeper network doesn't always help. A huge barrier to training them is vanishing gradients: very deep networks often have a gradient signal that goes to zero quickly, thus making gradient descent prohibitively slow.
#
# * More specifically, during gradient descent, as you backpropagate from the final layer back to the first layer, you are multiplying by the weight matrix on each step, and thus the gradient can decrease exponentially quickly to zero (or, in rare cases, grow exponentially quickly and "explode," from gaining very large values).
#
# * During training, you might therefore see the magnitude (or norm) of the gradient for the shallower layers decrease to zero very rapidly as training proceeds, as shown below:
# <img src="images/vanishing_grad_kiank.png" style="width:450px;height:220px;">
# <caption><center> <u> <font color='purple'> <b>Figure 1</b> </u><font color='purple'> : <b>Vanishing gradient</b> <br> The speed of learning decreases very rapidly for the shallower layers as the network trains </center></caption>
#
# Not to worry! You are now going to solve this problem by building a Residual Network!
# <a name='3'></a>
# ## 3 - Building a Residual Network
#
# In ResNets, a "shortcut" or a "skip connection" allows the model to skip layers:
#
# <img src="images/skip_connection_kiank.png" style="width:650px;height:200px;">
# <caption><center> <u> <font color='purple'> <b>Figure 2</b> </u><font color='purple'> : A ResNet block showing a skip-connection <br> </center></caption>
#
# The image on the left shows the "main path" through the network. The image on the right adds a shortcut to the main path. By stacking these ResNet blocks on top of each other, you can form a very deep network.
#
# The lecture mentioned that having ResNet blocks with the shortcut also makes it very easy for one of the blocks to learn an identity function. This means that you can stack on additional ResNet blocks with little risk of harming training set performance.
#
# On that note, there is also some evidence that the ease of learning an identity function accounts for ResNets' remarkable performance even more than skip connections help with vanishing gradients.
#
# Two main types of blocks are used in a ResNet, depending mainly on whether the input/output dimensions are the same or different. You are going to implement both of them: the "identity block" and the "convolutional block."
# <a name='3-1'></a>
# ### 3.1 - The Identity Block
#
# The identity block is the standard block used in ResNets, and corresponds to the case where the input activation (say $a^{[l]}$) has the same dimension as the output activation (say $a^{[l+2]}$). To flesh out the different steps of what happens in a ResNet's identity block, here is an alternative diagram showing the individual steps:
#
# <img src="images/idblock2_kiank.png" style="width:650px;height:150px;">
# <caption><center> <u> <font color='purple'> <b>Figure 3</b> </u><font color='purple'> : <b>Identity block.</b> Skip connection "skips over" 2 layers. </center></caption>
#
# The upper path is the "shortcut path." The lower path is the "main path." In this diagram, notice the CONV2D and ReLU steps in each layer. To speed up training, a BatchNorm step has been added. Don't worry about this being complicated to implement--you'll see that BatchNorm is just one line of code in Keras!
#
# In this exercise, you'll actually implement a slightly more powerful version of this identity block, in which the skip connection "skips over" 3 hidden layers rather than 2 layers. It looks like this:
#
# <img src="images/idblock3_kiank.png" style="width:650px;height:150px;">
# <caption><center> <u> <font color='purple'> <b>Figure 4</b> </u><font color='purple'> : <b>Identity block.</b> Skip connection "skips over" 3 layers.</center></caption>
# These are the individual steps:
#
# First component of main path:
# - The first CONV2D has $F_1$ filters of shape (1,1) and a stride of (1,1). Its padding is "valid". Use 0 as the seed for the random uniform initialization: `kernel_initializer = initializer(seed=0)`.
# - The first BatchNorm is normalizing the 'channels' axis.
# - Then apply the ReLU activation function. This has no hyperparameters.
#
# Second component of main path:
# - The second CONV2D has $F_2$ filters of shape $(f,f)$ and a stride of (1,1). Its padding is "same". Use 0 as the seed for the random uniform initialization: `kernel_initializer = initializer(seed=0)`.
# - The second BatchNorm is normalizing the 'channels' axis.
# - Then apply the ReLU activation function. This has no hyperparameters.
#
# Third component of main path:
# - The third CONV2D has $F_3$ filters of shape (1,1) and a stride of (1,1). Its padding is "valid". Use 0 as the seed for the random uniform initialization: `kernel_initializer = initializer(seed=0)`.
# - The third BatchNorm is normalizing the 'channels' axis.
# - Note that there is **no** ReLU activation function in this component.
#
# Final step:
# - The `X_shortcut` and the output from the 3rd layer `X` are added together.
# - **Hint**: The syntax will look something like `Add()([var1,var2])`
# - Then apply the ReLU activation function. This has no hyperparameters.
#
# <a name='ex-1'></a>
# ### Exercise 1 - identity_block
#
# Implement the ResNet identity block. The first component of the main path has been implemented for you already! First, you should read these docs carefully to make sure you understand what's happening. Then, implement the rest.
# - To implement the Conv2D step: [Conv2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D)
# - To implement BatchNorm: [BatchNormalization](https://www.tensorflow.org/api_docs/python/tf/keras/layers/BatchNormalization) `BatchNormalization(axis = 3)(X, training = training)`. If training is set to False, its weights are not updated with the new examples. I.e when the model is used in prediction mode.
# - For the activation, use: `Activation('relu')(X)`
# - To add the value passed forward by the shortcut: [Add](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Add)
#
# We have added the initializer argument to our functions. This parameter receives an initializer function like the ones included in the package [tensorflow.keras.initializers](https://www.tensorflow.org/api_docs/python/tf/keras/initializers) or any other custom initializer. By default it will be set to [random_uniform](https://www.tensorflow.org/api_docs/python/tf/keras/initializers/RandomUniform)
#
# Remember that these functions accept a `seed` argument that can be any value you want, but that in this notebook must set to 0 for **grading purposes**.
# Here is where you're actually using the power of the Functional API to create a shortcut path:
# + nbgrader={"grade": false, "grade_id": "cell-0017b68317ffa974", "locked": false, "schema_version": 3, "solution": true, "task": false}
# UNQ_C1
# GRADED FUNCTION: identity_block
def identity_block(X, f, filters, training=True, initializer=random_uniform):
"""
Implementation of the identity block as defined in Figure 4
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
training -- True: Behave in training mode
False: Behave in inference mode
initializer -- to set up the initial weights of a layer. Equals to random uniform initializer
Returns:
X -- output of the identity block, tensor of shape (n_H, n_W, n_C)
"""
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value. You'll need this later to add back to the main path.
X_shortcut = X
# First component of main path
X = Conv2D(filters = F1, kernel_size = 1, strides = (1,1), padding = 'valid', kernel_initializer = initializer(seed=0))(X)
X = BatchNormalization(axis = 3)(X, training = training) # Default axis
X = Activation('relu')(X)
### START CODE HERE
## Second component of main path (≈3 lines)
X = Conv2D(filters = F2, kernel_size = f, strides = (1,1), padding = 'same', kernel_initializer = initializer(seed=0))(X)
X = BatchNormalization(axis = 3)(X, training = training) # Default axis
X = Activation('relu')(X)
## Third component of main path (≈2 lines)
X = Conv2D(filters = F3, kernel_size = 1, strides = (1,1), padding = 'valid', kernel_initializer = initializer(seed=0))(X)
X = BatchNormalization(axis = 3)(X, training = training) # Default axis
## Final step: Add shortcut value to main path, and pass it through a RELU activation (≈2 lines)
X = X + X_shortcut
X = Activation('relu')(X)
### END CODE HERE
return X
# + nbgrader={"grade": true, "grade_id": "cell-e73a8466b807e261", "locked": true, "points": 10, "schema_version": 3, "solution": false, "task": false}
np.random.seed(1)
X1 = np.ones((1, 4, 4, 3)) * -1
X2 = np.ones((1, 4, 4, 3)) * 1
X3 = np.ones((1, 4, 4, 3)) * 3
X = np.concatenate((X1, X2, X3), axis = 0).astype(np.float32)
A3 = identity_block(X, f=2, filters=[4, 4, 3],
initializer=lambda seed=0:constant(value=1),
training=False)
print('\033[1mWith training=False\033[0m\n')
A3np = A3.numpy()
print(np.around(A3.numpy()[:,(0,-1),:,:].mean(axis = 3), 5))
resume = A3np[:,(0,-1),:,:].mean(axis = 3)
print(resume[1, 1, 0])
print('\n\033[1mWith training=True\033[0m\n')
np.random.seed(1)
A4 = identity_block(X, f=2, filters=[3, 3, 3],
initializer=lambda seed=0:constant(value=1),
training=True)
print(np.around(A4.numpy()[:,(0,-1),:,:].mean(axis = 3), 5))
public_tests.identity_block_test(identity_block)
# -
# **Expected value**
#
# ```
# With training=False
#
# [[[ 0. 0. 0. 0. ]
# [ 0. 0. 0. 0. ]]
#
# [[192.71234 192.71234 192.71234 96.85617]
# [ 96.85617 96.85617 96.85617 48.92808]]
#
# [[578.1371 578.1371 578.1371 290.5685 ]
# [290.5685 290.5685 290.5685 146.78426]]]
# 96.85617
#
# With training=True
#
# [[[0. 0. 0. 0. ]
# [0. 0. 0. 0. ]]
#
# [[0.40739 0.40739 0.40739 0.40739]
# [0.40739 0.40739 0.40739 0.40739]]
#
# [[4.99991 4.99991 4.99991 3.25948]
# [3.25948 3.25948 3.25948 2.40739]]]
# ```
# <a name='3-2'></a>
# ### 3.2 - The Convolutional Block
#
# The ResNet "convolutional block" is the second block type. You can use this type of block when the input and output dimensions don't match up. The difference with the identity block is that there is a CONV2D layer in the shortcut path:
#
# <img src="images/convblock_kiank.png" style="width:650px;height:150px;">
# <caption><center> <u> <font color='purple'> <b>Figure 4</b> </u><font color='purple'> : <b>Convolutional block</b> </center></caption>
#
# * The CONV2D layer in the shortcut path is used to resize the input $x$ to a different dimension, so that the dimensions match up in the final addition needed to add the shortcut value back to the main path. (This plays a similar role as the matrix $W_s$ discussed in lecture.)
# * For example, to reduce the activation dimensions's height and width by a factor of 2, you can use a 1x1 convolution with a stride of 2.
# * The CONV2D layer on the shortcut path does not use any non-linear activation function. Its main role is to just apply a (learned) linear function that reduces the dimension of the input, so that the dimensions match up for the later addition step.
# * As for the previous exercise, the additional `initializer` argument is required for grading purposes, and it has been set by default to [glorot_uniform](https://www.tensorflow.org/api_docs/python/tf/keras/initializers/GlorotUniform)
#
# The details of the convolutional block are as follows.
#
# First component of main path:
# - The first CONV2D has $F_1$ filters of shape (1,1) and a stride of (s,s). Its padding is "valid". Use 0 as the `glorot_uniform` seed `kernel_initializer = initializer(seed=0)`.
# - The first BatchNorm is normalizing the 'channels' axis.
# - Then apply the ReLU activation function. This has no hyperparameters.
#
# Second component of main path:
# - The second CONV2D has $F_2$ filters of shape (f,f) and a stride of (1,1). Its padding is "same". Use 0 as the `glorot_uniform` seed `kernel_initializer = initializer(seed=0)`.
# - The second BatchNorm is normalizing the 'channels' axis.
# - Then apply the ReLU activation function. This has no hyperparameters.
#
# Third component of main path:
# - The third CONV2D has $F_3$ filters of shape (1,1) and a stride of (1,1). Its padding is "valid". Use 0 as the `glorot_uniform` seed `kernel_initializer = initializer(seed=0)`.
# - The third BatchNorm is normalizing the 'channels' axis. Note that there is no ReLU activation function in this component.
#
# Shortcut path:
# - The CONV2D has $F_3$ filters of shape (1,1) and a stride of (s,s). Its padding is "valid". Use 0 as the `glorot_uniform` seed `kernel_initializer = initializer(seed=0)`.
# - The BatchNorm is normalizing the 'channels' axis.
#
# Final step:
# - The shortcut and the main path values are added together.
# - Then apply the ReLU activation function. This has no hyperparameters.
#
# <a name='ex-2'></a>
# ### Exercise 2 - convolutional_block
#
# Implement the convolutional block. The first component of the main path is already implemented; then it's your turn to implement the rest! As before, always use 0 as the seed for the random initialization, to ensure consistency with the grader.
# - [Conv2D](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D)
# - [BatchNormalization](https://www.tensorflow.org/api_docs/python/tf/keras/layers/BatchNormalization) (axis: Integer, the axis that should be normalized (typically the features axis)) `BatchNormalization(axis = 3)(X, training = training)`. If training is set to False, its weights are not updated with the new examples. I.e when the model is used in prediction mode.
# - For the activation, use: `Activation('relu')(X)`
# - [Add](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Add)
#
# We have added the initializer argument to our functions. This parameter receives an initializer function like the ones included in the package [tensorflow.keras.initializers](https://www.tensorflow.org/api_docs/python/tf/keras/initializers) or any other custom initializer. By default it will be set to [random_uniform](https://www.tensorflow.org/api_docs/python/tf/keras/initializers/RandomUniform)
#
# Remember that these functions accept a `seed` argument that can be any value you want, but that in this notebook must set to 0 for **grading purposes**.
# + nbgrader={"grade": false, "grade_id": "cell-df47af4847e5335f", "locked": false, "schema_version": 3, "solution": true, "task": false}
# UNQ_C2
# GRADED FUNCTION: convolutional_block
def convolutional_block(X, f, filters, s = 2, training=True, initializer=glorot_uniform):
"""
Implementation of the convolutional block as defined in Figure 4
Arguments:
X -- input tensor of shape (m, n_H_prev, n_W_prev, n_C_prev)
f -- integer, specifying the shape of the middle CONV's window for the main path
filters -- python list of integers, defining the number of filters in the CONV layers of the main path
s -- Integer, specifying the stride to be used
training -- True: Behave in training mode
False: Behave in inference mode
initializer -- to set up the initial weights of a layer. Equals to Glorot uniform initializer,
also called Xavier uniform initializer.
Returns:
X -- output of the convolutional block, tensor of shape (n_H, n_W, n_C)
"""
# Retrieve Filters
F1, F2, F3 = filters
# Save the input value
X_shortcut = X
##### MAIN PATH #####
# First component of main path glorot_uniform(seed=0)
X = Conv2D(filters = F1, kernel_size = 1, strides = (s, s), padding='valid', kernel_initializer = initializer(seed=0))(X)
X = BatchNormalization(axis = 3)(X, training=training)
X = Activation('relu')(X)
### START CODE HERE
## Second component of main path (≈3 lines)
X = Conv2D(filters = F2, kernel_size = f, strides = (1, 1), padding='same', kernel_initializer = initializer(seed=0))(X)
X = BatchNormalization(axis = 3)(X, training=training)
X = Activation('relu')(X)
## Third component of main path (≈2 lines)
X = Conv2D(filters = F3, kernel_size = 1, strides = (1, 1), padding='valid', kernel_initializer = initializer(seed=0))(X)
X = BatchNormalization(axis = 3)(X, training=training)
##### SHORTCUT PATH ##### (≈2 lines)
X_shortcut = Conv2D(filters = F3, kernel_size = 1, strides = (s, s), padding='valid', kernel_initializer = initializer(seed=0))(X_shortcut)
X_shortcut = BatchNormalization(axis = 3)(X_shortcut, training=training)
### END CODE HERE
# Final step: Add shortcut value to main path (Use this order [X, X_shortcut]), and pass it through a RELU activation
X = Add()([X, X_shortcut])
X = Activation('relu')(X)
return X
# + nbgrader={"grade": true, "grade_id": "cell-95c291eb244218fe", "locked": true, "points": 10, "schema_version": 3, "solution": false, "task": false}
from outputs import convolutional_block_output1, convolutional_block_output2
np.random.seed(1)
#X = np.random.randn(3, 4, 4, 6).astype(np.float32)
X1 = np.ones((1, 4, 4, 3)) * -1
X2 = np.ones((1, 4, 4, 3)) * 1
X3 = np.ones((1, 4, 4, 3)) * 3
X = np.concatenate((X1, X2, X3), axis = 0).astype(np.float32)
A = convolutional_block(X, f = 2, filters = [2, 4, 6], training=False)
assert type(A) == EagerTensor, "Use only tensorflow and keras functions"
assert tuple(tf.shape(A).numpy()) == (3, 2, 2, 6), "Wrong shape."
assert np.allclose(A.numpy(), convolutional_block_output1), "Wrong values when training=False."
print(A[0])
B = convolutional_block(X, f = 2, filters = [2, 4, 6], training=True)
assert np.allclose(B.numpy(), convolutional_block_output2), "Wrong values when training=True."
print('\033[92mAll tests passed!')
# -
# **Expected value**
#
# ```
# tf.Tensor(
# [[[0. 0.66683817 0. 0. 0.88853896 0.5274254 ]
# [0. 0.65053666 0. 0. 0.89592844 0.49965227]]
#
# [[0. 0.6312079 0. 0. 0.8636247 0.47643146]
# [0. 0.5688321 0. 0. 0.85534114 0.41709304]]], shape=(2, 2, 6), dtype=float32)
# ```
# <a name='4'></a>
# ## 4 - Building Your First ResNet Model (50 layers)
#
# You now have the necessary blocks to build a very deep ResNet. The following figure describes in detail the architecture of this neural network. "ID BLOCK" in the diagram stands for "Identity block," and "ID BLOCK x3" means you should stack 3 identity blocks together.
#
# <img src="images/resnet_kiank.png" style="width:850px;height:150px;">
# <caption><center> <u> <font color='purple'> <b>Figure 5</b> </u><font color='purple'> : <b>ResNet-50 model</b> </center></caption>
#
# The details of this ResNet-50 model are:
# - Zero-padding pads the input with a pad of (3,3)
# - Stage 1:
# - The 2D Convolution has 64 filters of shape (7,7) and uses a stride of (2,2).
# - BatchNorm is applied to the 'channels' axis of the input.
# - MaxPooling uses a (3,3) window and a (2,2) stride.
# - Stage 2:
# - The convolutional block uses three sets of filters of size [64,64,256], "f" is 3, and "s" is 1.
# - The 2 identity blocks use three sets of filters of size [64,64,256], and "f" is 3.
# - Stage 3:
# - The convolutional block uses three sets of filters of size [128,128,512], "f" is 3 and "s" is 2.
# - The 3 identity blocks use three sets of filters of size [128,128,512] and "f" is 3.
# - Stage 4:
# - The convolutional block uses three sets of filters of size [256, 256, 1024], "f" is 3 and "s" is 2.
# - The 5 identity blocks use three sets of filters of size [256, 256, 1024] and "f" is 3.
# - Stage 5:
# - The convolutional block uses three sets of filters of size [512, 512, 2048], "f" is 3 and "s" is 2.
# - The 2 identity blocks use three sets of filters of size [512, 512, 2048] and "f" is 3.
# - The 2D Average Pooling uses a window of shape (2,2).
# - The 'flatten' layer doesn't have any hyperparameters.
# - The Fully Connected (Dense) layer reduces its input to the number of classes using a softmax activation.
#
#
# <a name='ex-3'></a>
# ### Exercise 3 - ResNet50
#
# Implement the ResNet with 50 layers described in the figure above. We have implemented Stages 1 and 2. Please implement the rest. (The syntax for implementing Stages 3-5 should be quite similar to that of Stage 2) Make sure you follow the naming convention in the text above.
#
# You'll need to use this function:
# - Average pooling [see reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/AveragePooling2D)
#
# Here are some other functions we used in the code below:
# - Conv2D: [See reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Conv2D)
# - BatchNorm: [See reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/BatchNormalization) (axis: Integer, the axis that should be normalized (typically the features axis))
# - Zero padding: [See reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/ZeroPadding2D)
# - Max pooling: [See reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/MaxPool2D)
# - Fully connected layer: [See reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense)
# - Addition: [See reference](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Add)
# + nbgrader={"grade": false, "grade_id": "cell-10dc95a4cf6275b9", "locked": false, "schema_version": 3, "solution": true, "task": false}
# UNQ_C3
# GRADED FUNCTION: ResNet50
def ResNet50(input_shape = (64, 64, 3), classes = 6):
"""
Stage-wise implementation of the architecture of the popular ResNet50:
CONV2D -> BATCHNORM -> RELU -> MAXPOOL -> CONVBLOCK -> IDBLOCK*2 -> CONVBLOCK -> IDBLOCK*3
-> CONVBLOCK -> IDBLOCK*5 -> CONVBLOCK -> IDBLOCK*2 -> AVGPOOL -> FLATTEN -> DENSE
Arguments:
input_shape -- shape of the images of the dataset
classes -- integer, number of classes
Returns:
model -- a Model() instance in Keras
"""
# Define the input as a tensor with shape input_shape
X_input = Input(input_shape)
# Zero-Padding
X = ZeroPadding2D((3, 3))(X_input)
# Stage 1
X = Conv2D(64, (7, 7), strides = (2, 2), kernel_initializer = glorot_uniform(seed=0))(X)
X = BatchNormalization(axis = 3)(X)
X = Activation('relu')(X)
X = MaxPooling2D((3, 3), strides=(2, 2))(X)
# Stage 2
X = convolutional_block(X, f = 3, filters = [64, 64, 256], s = 1)
X = identity_block(X, 3, [64, 64, 256])
X = identity_block(X, 3, [64, 64, 256])
### START CODE HERE
## Stage 3 (≈4 lines)
X = convolutional_block(X, f = 3, filters = [128,128,512], s = 2)
X = identity_block(X, f = 3, filters = [128,128,512])
X = identity_block(X, f = 3, filters = [128,128,512])
X = identity_block(X, f = 3, filters = [128,128,512])
## Stage 4 (≈6 lines)
X = convolutional_block(X, f = 3, filters = [256,256,1024], s = 2)
X = identity_block(X, f = 3, filters = [256,256,1024])
X = identity_block(X, f = 3, filters = [256,256,1024])
X = identity_block(X, f = 3, filters = [256,256,1024])
X = identity_block(X, f = 3, filters = [256,256,1024])
X = identity_block(X, f = 3, filters = [256,256,1024])
## Stage 5 (≈3 lines)
X = convolutional_block(X, f = 3, filters = [512,512,2048], s = 2)
X = identity_block(X, f = 3, filters = [512,512,2048])
X = identity_block(X, f = 3, filters = [512,512,2048])
## AVGPOOL (≈1 line). Use "X = AveragePooling2D(...)(X)"
X = AveragePooling2D(pool_size=(2, 2))(X)
### END CODE HERE
# output layer
X = Flatten()(X)
X = Dense(classes, activation='softmax', kernel_initializer = glorot_uniform(seed=0))(X)
# Create model
model = Model(inputs = X_input, outputs = X)
return model
# -
# Run the following code to build the model's graph. If your implementation is incorrect, you'll know it by checking your accuracy when running `model.fit(...)` below.
model = ResNet50(input_shape = (64, 64, 3), classes = 6)
print(model.summary())
# + nbgrader={"grade": true, "grade_id": "cell-866b891ec47ccb7b", "locked": true, "points": 10, "schema_version": 3, "solution": false, "task": false}
from outputs import ResNet50_summary
model = ResNet50(input_shape = (64, 64, 3), classes = 6)
comparator(summary(model), ResNet50_summary)
# -
# As shown in the Keras Tutorial Notebook, prior to training a model, you need to configure the learning process by compiling the model.
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
# The model is now ready to be trained. The only thing you need now is a dataset!
# Let's load your old friend, the SIGNS dataset.
#
# <img src="images/signs_data_kiank.png" style="width:450px;height:250px;">
# <caption><center> <u> <font color='purple'> <b>Figure 6</b> </u><font color='purple'> : <b>SIGNS dataset</b> </center></caption>
#
# +
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
# Normalize image vectors
X_train = X_train_orig / 255.
X_test = X_test_orig / 255.
# Convert training and test labels to one hot matrices
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
# -
# Run the following cell to train your model on 10 epochs with a batch size of 32. On a GPU, it should take less than 2 minutes.
model.fit(X_train, Y_train, epochs = 10, batch_size = 32)
# **Expected Output**:
#
# ```
# Epoch 1/10
# 34/34 [==============================] - 1s 34ms/step - loss: 1.9241 - accuracy: 0.4620
# Epoch 2/10
# 34/34 [==============================] - 2s 57ms/step - loss: 0.6403 - accuracy: 0.7898
# Epoch 3/10
# 34/34 [==============================] - 1s 24ms/step - loss: 0.3744 - accuracy: 0.8731
# Epoch 4/10
# 34/34 [==============================] - 2s 44ms/step - loss: 0.2220 - accuracy: 0.9231
# Epoch 5/10
# 34/34 [==============================] - 2s 57ms/step - loss: 0.1333 - accuracy: 0.9583
# Epoch 6/10
# 34/34 [==============================] - 2s 52ms/step - loss: 0.2243 - accuracy: 0.9444
# Epoch 7/10
# 34/34 [==============================] - 2s 48ms/step - loss: 0.2913 - accuracy: 0.9102
# Epoch 8/10
# 34/34 [==============================] - 1s 30ms/step - loss: 0.2269 - accuracy: 0.9306
# Epoch 9/10
# 34/34 [==============================] - 2s 46ms/step - loss: 0.1113 - accuracy: 0.9630
# Epoch 10/10
# 34/34 [==============================] - 2s 57ms/step - loss: 0.0709 - accuracy: 0.9778
# ```
#
# The exact values could not match, but don't worry about that. The important thing that you must see is that the loss value decreases, and the accuracy increases for the firsts 5 epochs.
# Let's see how this model (trained on only two epochs) performs on the test set.
preds = model.evaluate(X_test, Y_test)
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
# **Expected Output**:
#
# <table>
# <tr>
# <td>
# <b>Test Accuracy</b>
# </td>
# <td>
# >0.80
# </td>
# </tr>
#
# </table>
# For the purposes of this assignment, you've been asked to train the model for just two epochs. You can see that it performs pretty poorly, but that's ok! The online grader will only run your code for a small number of epochs as well. Please go ahead and submit your assignment as is.
# After you have finished this official (graded) part of this assignment, you can also optionally train the ResNet for more iterations, if you want. It tends to get much better performance when trained for ~20 epochs, but this does take more than an hour when training on a CPU.
#
# Using a GPU, this ResNet50 model's weights were trained on the SIGNS dataset. You can load and run the trained model on the test set in the cells below. It may take ≈1min to load the model. Have fun!
pre_trained_model = tf.keras.models.load_model('resnet50.h5')
preds = pre_trained_model.evaluate(X_test, Y_test)
print ("Loss = " + str(preds[0]))
print ("Test Accuracy = " + str(preds[1]))
# **Congratulations** on finishing this assignment! You've now implemented a state-of-the-art image classification system! Woo hoo!
#
# ResNet50 is a powerful model for image classification when it's trained for an adequate number of iterations. Hopefully, from this point, you can use what you've learned and apply it to your own classification problem to perform state-of-the-art accuracy.
# <font color = 'blue'>
#
# **What you should remember**:
#
# - Very deep "plain" networks don't work in practice because vanishing gradients make them hard to train.
# - Skip connections help address the Vanishing Gradient problem. They also make it easy for a ResNet block to learn an identity function.
# - There are two main types of blocks: The **identity block** and the **convolutional block**.
# - Very deep Residual Networks are built by stacking these blocks together.
# <a name='5'></a>
# ## 5 - Test on Your Own Image (Optional/Ungraded)
# If you wish, you can also take a picture of your own hand and see the output of the model. To do this:
# 1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
# 2. Add your image to this Jupyter Notebook's directory, in the "images" folder
# 3. Write your image's name in the following code
# 4. Run the code and check if the algorithm is right!
img_path = 'images/my_image.jpg'
img = image.load_img(img_path, target_size=(64, 64))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = x/255.0
print('Input image shape:', x.shape)
imshow(img)
prediction = pre_trained_model.predict(x)
print("Class prediction vector [p(0), p(1), p(2), p(3), p(4), p(5)] = ", prediction)
print("Class:", np.argmax(prediction))
# You can also print a summary of your model by running the following code.
pre_trained_model.summary()
# <a name='6'></a>
# ## 6 - Bibliography
#
# This notebook presents the ResNet algorithm from He et al. (2015). The implementation here also took significant inspiration and follows the structure given in the GitHub repository of Francois Chollet:
#
# - <NAME>, <NAME>, <NAME>, <NAME> - [Deep Residual Learning for Image Recognition (2015)](https://arxiv.org/abs/1512.03385)
# - Francois Chollet's GitHub repository: https://github.com/fchollet/deep-learning-models/blob/master/resnet50.py
#
| convolution_neural_networks/week_2/Residual_Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.applications.xception import Xception
from keras.models import load_model
from pickle import load
import numpy as np
from PIL import Image
import matplotlib.pyplot as plt
import argparse
from pathlib import Path
from IPython.display import display
from nltk.translate.bleu_score import corpus_bleu
#ap = argparse.ArgumentParser()
#ap.add_argument('-i', '--image', required=True, help="Image Path")
#args = vars(ap.parse_args())
#img_path = args['image']
def extract_features(filename, model):
try:
image = Image.open(filename)
except:
print("ERROR: Couldn't open image! Make sure the image path and extension is correct")
image = image.resize((299,299))
image = np.array(image)
# for images that has 4 channels, we convert them into 3 channels
if image.shape[2] == 4:
image = image[..., :3]
image = np.expand_dims(image, axis=0)
image = image/127.5
image = image - 1.0
feature = model.predict(image)
return feature
def word_for_id(integer, tokenizer):
for word, index in tokenizer.word_index.items():
if index == integer:
return word
return None
def generate_desc(model, tokenizer, photo, max_length):
in_text = 'start'
for i in range(max_length):
sequence = tokenizer.texts_to_sequences([in_text])[0]
sequence = pad_sequences([sequence], maxlen=max_length)
pred = model.predict([photo,sequence], verbose=0)
pred = np.argmax(pred)
word = word_for_id(pred, tokenizer)
if word is None:
break
in_text += ' ' + word
if word == 'end':
break
return in_text
path = Path("C:\\Users\Harsh\Desktop\Test Images")
image_label = 'images.jpg'
img_path = path / image_label
max_length = 72
tokenizer = load(open("tokenizer.p","rb"))
model = load_model('model_8.h5')
xception_model = Xception(include_top=False, pooling="avg")
photo = extract_features(img_path, xception_model)
img = Image.open(img_path)
description = generate_desc(model, tokenizer, photo, max_length)
print(description)
display(img.resize((299,299)))
| Models/Flickr30k/.ipynb_checkpoints/Flickr30k_testing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# Experimento muy simple solo para ilustrar el funcionamiento del QuantileTransformer cuando se tiene un target de distribución gaussiana.
#
# sklearn.preprocessing.QuantileTransformer:
# 1. https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.QuantileTransformer.html#sklearn.preprocessing.QuantileTransformer.inverse_transform
# 1. https://en.wikipedia.org/wiki/Quantile_normalization
# 1. https://stats.stackexchange.com/questions/325570/quantile-transformation-with-gaussian-distribution-sklearn-implementation
import sys
sys.path.append('../')
# +
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from mismatch.quantile_transform import naive_quantile_normal
from fuzzytools.matplotlib.plots import plot_hist_bins
'''
cdf = cumulative density function
ppf = Percent point function (inverse of cdf — percentiles).
'''
fig, axs = plt.subplots(1, 2, figsize=(15,6))
n = 500
to_plot_points = 30
hist_kwargs = {
'fig':fig,
'ax':axs[0],
'bins':50,
'cmap':matplotlib.colors.ListedColormap(['r','b'], name=''),
'title':'data distribution',
'legend_loc':'upper left',
}
n_quantiles = 100 # or percentiles
x = np.random.RandomState(0).normal(loc=0.8, scale=0.5, size=(n,1)) # generate random data
normx, to_plot = naive_quantile_normal(x, n_quantiles) # get nromalized data
x_percentiles, x_distr_ppf, x_percentiles_i, normx_percentiles_i, norm_ppf = to_plot
### plots
to_plot = {'x':x, 'normx':normx[~(np.abs(normx)==np.infty)]} # for this example, we don't care for infty values
plot_hist_bins(to_plot, **hist_kwargs)
ax = axs[1]
ax.plot(x_percentiles, x_distr_ppf, '-r', label='x-distribution ppf')
ax.plot(x_percentiles_i[:30], x[:30], 'or', label='x-ppf per each data point')
ax.plot(x_percentiles, norm_ppf, 'b-', label='normal-distribution ppf')
ax.plot(normx_percentiles_i[:30], normx[:30], 'ob', label='normx-ppf per each data point')
for i in range(0,30):
ax.plot([x_percentiles_i[i], normx_percentiles_i[i]], [x[i], normx[i]], 'k', label='map' if i==0 else None)
ax.grid(alpha=0.5)
ax.set_title('ppf functions for distributions (ppf = inverse of cdf)\nnew data points in normx share the same percentiles as x')
ax.set_xlabel('percentiles')
ax.set_ylabel('ppf values')
ax.legend()
| experiments/.ipynb_checkpoints/quantile_transformer-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Git versioning system
# **git** is a version control system (VCS) which keeps track of different version of a project and allows people to collaborate. It is most widely used for code development or website development.
#
# Git was created by <NAME> in 2005 for the development of the Linux kernel. It's current maintainer is <NAME>, a Google employee.
# The advantages of git are:
# * you can come back to a previous version
# * you have a backup of the current and all previous checkpoints
# * you can collaborate with others to work on large software projects
# Sources :
# * https://en.wikipedia.org/wiki/Git
# * https://git-scm.com
# ## Documentation
# Excellent documentation is available for free on the web. <NAME> and <NAME> wrote **Pro Git**, a 500-page book which is available for free, and had been translated to multiple languages (fully translated in 8 different languages, in 2018.
#
# * English: https://git-scm.com/book/en/v2
# * French: https://git-scm.com/book/fr/v2
#
# The book is available as
# * printed book version by Apress, available on Amazon
# * HTML verion
# * PDF version
# * eBook version (the most popular e-publishing platform)
#
# The source code for the book is hosted on GitHub
# https://github.com/progit/progit2
#
# ### About running git examples within a Jupyter notebook
# * the command **git** must be preceded by `!` to run it in the Linux command shell
# * Use `alt+enter` to run a code cell and create a new one
# ## Getting help
# Just running `git` without any arguments at the command line gives a short summary of all it's command.
# !git
# For example the first command mentioned allows to find out what version is running
# !git --version
# This is an old version, so let's update git to the latest one.
# !git --version
# !git help
# # Basics
# There are two ways of starting a repository:
# 1. cloning an existing repository from GitHub
# * turning an empty directory into a git repository
# ## Create a local repository
# For this exercice we create a local git repository `git-demo` inside a temporary directory `tmp` inside the user's home directory `~`.
# !mkdir ~/tmp/git-demo
# !cd ~/tmp/git-demo
# In order to test git, we must make sure that we are not already in a git-versioned sub-directory. For example the directory in which this Jupyter notebook resides is under git version control. However the newly create directory is not. And it is also completely empty.
# !git status
# ls -lA
# So let's create a new repository here.
# !git init
# Now the base directory contains a hidden `.git` directory which is the git repository which contains all the versioning information. The information is in the subfolders.
# ls -A
# ls -A .git/
# cat .git/config
# ## Configure git
# The commande `git config` allows to configure git. There are three levels of configuration:
# * **system** - for all users in the system
# * **global** - for one specific user
# * **local** - for one specific repository
# !git config
# To show all configuration key=value pairs use the command `git config --list`
# !git config --list
# The minimum configuration for git is a user name and a user email. Let's check this.
# !git config user.name
# !git config user.email
# ## Create a directory for the new project
# Let's add a new file to the newly created repository.
# %%writefile README.md
# git demo-project
This is the first line
# It is there.
# ls
# Check the git status. The newly created file is not yet under version control and will show up in red.
# !git status
# We have to add it to the index (staging area) using `git add`
# !git add README.md
# To make the changes permanent in the repository database, we need to use the command `commit`, which has the option `-m` of adding a commit message.
# !git commit README.md -m 'initial commit'
# With `git log` we can show all the versioning history. Each commit is identified not by a file name or a time-stamp, but by the SHA1 key of the file commited.
# !git log
# This is how we can calculate the SHA1 sum on the Mac:
# !shasum README.md
# Let's modify the README.md file
# !echo 'add another line' >> README.md
# !git status
# !shasum -h
# !git diff
# !git commit -a -m 'modified README file'
# !git log
# !shasum README.md
# # Cloning an existing directory
# Go to the temporary directory in the user directory
# cd ~/tmp
# ls
# !git clone https://github.com/progit/progit2-fr.git
# ls
# cd progit2-fr/
# +
# # %load README.asc
image:https://travis-ci.org/progit/progit2-fr.svg?branch=master["Build Status", link="https://travis-ci.org/progit/progit2-fr"]
= Pro Git 2ème Édition, Effort de traduction française
Ce dépôt concerne l'effort de traduction en français de la deuxième édition du livre Pro Git.
La dernière version du livre est compilée et proposée en téléchargement sur https://git-scm.com/book/fr aux formats web, epub et PDF.
== Liste de diffusion
Notre liste de diffusion est ici : http://groups.google.com/group/progit-fr
Si vous travaillez ou souhaitez travailler sur la traduction française de Progit 2ème édition, faites-nous le savoir, de sorte à ne pas dupliquer le travail.
== Méthode de contribution
Pour simplifier la gestion et utiliser pleinement les capacités de Git et GitHub, la manière la plus directe de collaborer consiste à faire un fork sur GitHub du dépôt progit/progit2-fr sur votre propre compte pour y générer vos modifications, si possible sur une branche thématique.
Ensuite, il suffit de lancer une requête de tirage pour nous avertir que les modifications peuvent être revues et intégrées.
== Comment générer le livre
Vous pouvez générer les différentes versions du livre manuellement avec Asciidoctor.
Pour cela, vous devez installer les paquets `ruby`, `rubygems`, `rubygem-asciidoctor` et `ruby-devel` s'ils ne sont pas déjà installés.
Vous pouvez générer le livre aux formats PDF, e-pub, mobi et HTML avec les commandes suivantes :
[source,console]
----
$ bundle install
$ bundle exec rake book:build
Converting to HTML...
-- HTML output at progit.html
Converting to EPub...
-- Epub output at progit.epub
Converting to Mobi (kf8)...
-- Mobi output at progit.mobi
Converting to PDF...
-- PDF output at progit.pdf
----
Une alternative à l'appel de la commande `bundle` est d'appeler directement la commande `asciidoctor`.
Utilisez les commandes suivantes :
[source,console]
----
$ asciidoctor progit.asc
$ asciidoctor-pdf progit.asc
$ asciidoctor-epub3 progit.asc
$ asciidoctor-epub3 -a ebook-format=kf8 progit.asc
----
Cela utilise les projets `asciidoctor`, `asciidoctor-pdf` et `asciidoctor-epub`.
Pour plus d'informations, veuillez vous référer à link:generer_livre.asc[la page dédiée].
= Références pour la traduction
== Fichier glossaire
Le fichier fr/glossaire-git.adoc tente de rassembler les traductions choisies pour les termes spécifiques à Git. Si d'autres termes nécessitent une uniformisation, il ne faut pas hésiter à les y ajouter. De même, il est toujours possible de changer les termes déjà renseignés si une meilleure traduction est proposée.
L'uniformisation de la traduction par le respect de ce glossaire est une tâche permanente d'amélioration de la qualité finale du texte.
== Glossaires généraux sur internet
Les glossaires de traduction informatiques disponibles sur Internet, sans être des références dogmatiques donnent des indications sur les termes les plus utilisés dans les documents français sur l'informatique.
* http://glossaire.traduc.org/
* http://www.dglf.culture.gouv.fr/cogeter/16-03-99-internet-listes.html
* http://deschamp.free.fr/exinria/RETIF/
== Typographie française
La version française du livre se veut un document valant un document écrit nativement en français. À ce titre, la traduction doit suivre autant que possible les règles de typographie française en vigueur.
Voici ci-dessous quelques liens :
* http://devernay.free.fr/typo.html
* http://jacques-andre.fr/faqtypo/lessons.pdf
# -
# Now let's install the code for the open-source book **Pro Git, 2nd Edition**, using `git clone url`:
# !pwd
# !git clone https://github.com/progit/progit2.git
# Update Homebrew
# !brew update
# Install Ruby
# !brew install rbenv ruby-build
# !ruby --version
# # Git Branching
# Branching is seperating the working flow into two different strands to develop a separte aspect or a new feature. Git has an extremely light-weight branching mechanism. It's lightning fast and it is encouraged to branch and merge frequently.
# ## Branches in a Nutshell
# To understand the Git branching mechanism, we have to examine how Git stores commits. Git stores changes as a series of snapshots (or commits).
# A commit consists of three elements:
# * commit element : pointer to tree (snapshot), pointer to parent, name, email, message
# * tree element : the snapshot
# * data blobs : files referred to by their SHA1 key
# At this moment there is only one single file in our repository.
# ls
# ls
# There is an untracked file called `.DS_Store` which is a Mac specific hidden file.
# !git status
# We add a new file to our repository.
# !echo 'licence file' > LICENSE.txt
# The two files are present in the listing.
# ls
# `git status` now shows anothre untracked file.
# !git status
# We add the file to the staging area.
# !git add LICENSE.txt
# and commit the
# !git commit -m 'added license'
# ls
# !git status
# !git rm LICENCSE.txt
# !git status
# !git commit -m 'removed file with typo in name'
# !git log
# A commit is just a pointer to a tree. It's an extremely light-weight data structure:
# * pointer to the previous commit
# * info (name, email, message)
# * tree
# * blobs corresponding to the tree entries
# cd .git
# !tree
# cd ..
# !shasum LICENSE.txt
# !shasum README.md
# !cat .git/objects/12/54d0c5234e9c5458bcec150e17c817b21177eb
# !cat .git/objects/19/564902f3e7eb2834ed890d185ad5359e7b9e30
# !git branch testing
# ls .git/objects/
# ls .git/logs/refs/heads/
# cat .git/logs/refs/heads/master
# cat .git/logs/refs/heads/testing
# !git log
# !git log --decorate --oneline
# To switch to the `testing` branch use the `checkout` command
# !git checkout testing
# !echo 'change' >> LICENSE.txt
# !git commit -a -m 'changed license'
# !git checkout master
# !echo 'made another change' > LICENSE.txt
# !git commit -a -m 'changed licenses again (in master)'
# !git log --oneline --decorate --graph --all
# cd .git/
# !tree
# !cat refs/heads/master
# !cat refs/heads/testing
# ls -l refs/heads/
# ## Basic Branching and Merging
# !git checkout -b 'hotfix'
# !echo '<html></html>' > index.html
# !git commit -a -m 'added HTML index file'
# !git add index.html
# !git commit -m 'added HTML file'
# !git log --oneline --decorate --graph --all
# !git checkout master
# !git merge hotfix
# !git branch -d hotfix
# ## Branch Management
# ## Branching Workflows
# ## Remote Branches
# cd git-demo/
# !git branch -vv
# !git fetch --all
# !git branch -v
# ## Rebasing
# ## Summary
# !ls
# !shasum README.md
# !cat README.md
# # Git internals
# ## Plumbing and Porcelain
# cd ~/tmp
# !git init test
# cd test/
# ls -F1 .git
# cd .git
# !tree
# ## Git Objects
pwd
# !find .git/objects
# !echo 'test content' | git hash-object -w --stdin
# !find .git/objects -type f
# !git cat-file -p d670460b4b4aece5915caf5c68d12f560a9fe3e4
# !echo 'version 1' > test.txt
# !git hash-object -w test.txt
# !echo 'version 2' > test.txt
# !git hash-object -w test.txt
# !find .git/objects -type f
# !git cat-file -p 83baae61804e65cc73a7201a7252750c76066a30 > test.txt
# !cat test.txt
# !git cat-file -p 1f7a7a472abf3dd9643fd615f6da379c4acb3e3a > test.txt
# !cat test.txt
# !git cat-file -t 1f7a7a472abf3dd9643fd615f6da379c4acb3e3a
# !echo 'read me' > README.md
# !git status
# !git add README.md test.txt
# !git commit -m 'initial commit'
# !git cat-file -p master^{tree}
# !git update-index --add --cacheinfo 100644 \
# 83baae61804e65cc73a7201a7252750c76066a30 test.txt
# !git write-tree
# !git cat-file -p a14c904cf2467053919c908ecbbfd159bea6bc77
# !git cat-file -t a14c904cf2467053919c908ecbbfd159bea6bc77
# !echo 'new file' > new.txt
# !git update-index --add --cacheinfo 100644 \
# 1f7a7a472abf3dd9643fd615f6da379c4acb3e3a test.txt
# !git update-index --add new.txt
# !git write-tree
# !git cat-file -p 726169a6aca1f3820e89fac001f9ffced90c3d31
# !git read-tree --prefix=bak 726169a6aca1f3820e89fac001f9ffced90c3d31
# !git write-tree
# !git cat-file -p 754d31f5d80d1fa8fd00da2d8e16a7d4a8055055
# !git cat-file -p 726169a6aca1f3820e89fac001f9ffced90c3d31
# !echo 'second commit' | git commit-tree 754d31 -p 85e9c5
# !git cat-file -p e9697aabb8dfa1eebfd85f0168446bae8c5b45c5
# !echo 'second commit' | git commit-tree 726169a6aca1f3820e89fac001f9ffced90c3d31 -p 85e9c50676d2639601b32eee52173ab05f814917
# !git log --stat 85e9c5
# !git log
# !find .git/objects -type f
content = 'what is up, doc?'
content
header = 'blob '+str(len(content))+'\x00'
header
store = header+content
store
bytes(store, 'utf-8')
import hashlib
hashlib.sha1(b'blob 16\x00what is up, doc?').hexdigest()
hashlib.sha1(bytes(store, 'utf-8')).hexdigest()
# !echo -n "what is up, doc?" | git hash-object --stdin
# ## Git References
# !find .git/refs
# !find .git/refs -type f
# !cat .git/refs/heads/master
# ## Packfiles
# ## The Refspec
# ## Transfer Protocols
# ## Maintenance and Data Recovery
# ## Environment Variables
# ## Summary
# ls -l /tmp
# ls /private/tmp/
# # Distributed Git
# !gpg --list-keys
# !git tag v0.1 -m 'my signed 0.1 tag'
# !git describe master
# !git archive master --prefix='project/' | gzip > `git describe master`.tar.gz
# ls
# !git archive master --prefix='project/' --format=zip > `git describe master`.zip
# ls
# !git shortlog --no-merges master
| nb/git/git.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.datasets import fetch_olivetti_faces
import matplotlib.pyplot as plt
np.random.seed(42)
# %matplotlib inline
# -
data = fetch_olivetti_faces()
x = data.data
y = data.target
print(x.shape)
print(y.shape)
plt.imshow(x[0].reshape(64, 64), cmap='gray')
# Looking on a random set of images
fig = plt.figure(figsize=(9, 9))
cols = 4
rows = 5
for ind in range(1, cols*rows+1):
img = x[np.random.randint(x.shape[0])].reshape(64, 64)
fig.add_subplot(rows, cols, ind)
plt.imshow(img, cmap='gray')
plt.axis("off")
plt.show()
x.shape
# +
# Splitting into train and test set and having equal proportions
from sklearn.model_selection import StratifiedShuffleSplit
split_test = StratifiedShuffleSplit(n_splits=1, test_size=0.1, random_state=42)
for train_valid_ind, test_ind in split_test.split(x, y):
x_train_valid, x_test = x[train_valid_ind], x[test_ind]
y_train_valid, y_test = y[train_valid_ind], y[test_ind]
split_valid = StratifiedShuffleSplit(n_splits=1, test_size=0.2, random_state=42)
for train_ind, valid_ind in split_valid.split(x_train_valid, y_train_valid):
x_train, x_valid = x_train_valid[train_ind], x_train_valid[valid_ind]
y_train, y_valid = y_train_valid[train_ind], y_train_valid[valid_ind]
# -
# ### PCA Reduction
# +
from sklearn.decomposition import PCA
pca = PCA(n_components=0.99)
x_train_pca = pca.fit_transform(x_train)
x_valid_pca = pca.transform(x_valid)
# +
from sklearn.mixture import GaussianMixture
gm = GaussianMixture(n_components=40, random_state=42)
y_pred = gm.fit_predict(x_train_pca)
# -
# Generating random samples for further processing
x_gen, y_gen = gm.sample(20)
# Converting samples to full size for displaying
print(x_gen.shape)
x_gen = pca.inverse_transform(x_gen)
print(x_gen.shape)
# ### Visualizing Sampled Images by GMM
# +
def plot_faces(faces, label, n_rows = 4, n_cols = 5):
plt.figure(figsize=(8, 5))
for index, (face, label) in enumerate(zip(faces, label)):
plt.subplot(n_rows, n_cols, index+1)
plt.imshow(face.reshape(64, 64), cmap='gray')
plt.axis("off")
plt.title(label)
plt.show()
plot_faces(x_gen, y_gen)
# -
# ### Modifying Images and detecting anomalies
# +
from scipy import ndimage
# rotated, flipped and darkened the images
# flipping and darkening has been used from solution as turned out to be easier
x_transformed = []
for face in x_train[:20]:
transform = ndimage.rotate(face.reshape(64, 64), angle=np.random.choice([90, 180]),
mode='constant')[:,::-1]
transform[:, 1:-1] *= np.random.choice([1, 0.3])
x_transformed.append(transform)
x_transformed = np.array(x_transformed)
# -
plot_faces(x_transformed, y_train[:20])
# +
# Performing dimensionality reduction and passing through GMM
x_trans_pca = pca.transform(x_transformed.reshape(-1, 4096))
gm.score_samples(x_trans_pca)
# -
# ### GMM scores them very poorly
gm.score_samples(x_train_pca[:20])
# ### In Comparison the original data had much higher scores
| exercise/chap_9/12_olivetti_gaussian.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup
# +
# %matplotlib inline
import numpy as np
import scipy.signal as sig
import scipy.stats as stat
import matplotlib.pyplot as plt
import seaborn as sns
import os
import h5py
import datetime
import pandas as pd
from pandas import DataFrame,Series,read_table
# -
# General info
# +
makePlots = True # whether or not to save plots
saveAsPath = './Fig 01/'
if not os.path.exists(saveAsPath):
os.mkdir(saveAsPath)
saveAsName = 'PSG_example_'
# +
birdPaths = ['../data_copies/01_PreprocessedData/01_BudgieFemale_green1/00_Baseline_night/',
'../data_copies/01_PreprocessedData/02_BudgieMale_yellow1/00_Baseline_night/',
'../data_copies/01_PreprocessedData/03_BudgieFemale_white1/00_Baseline_night/',
'../data_copies/01_PreprocessedData/04_BudgieMale_yellow2/00_Baseline_night/',
'../data_copies/01_PreprocessedData/05_BudgieFemale_green2/00_Baseline_night/']
arfFilePaths = ['EEG 2 scored/',
'EEG 3 scored/',
'EEG 3 scored/',
'EEG 4 scored/',
'EEG 4 scored/']
### load BEST EEG channels - as determined during manual scoring ####
channelsToLoadEEG_best = [['5 LEEGf-LEEGp', '6 LEEGm-LEEGp'],
['5 LEEGf-LEEGm'], #, '4 LEEGf-Fgr'] this channel seems to have too much artifact
['6LEEGm-LEEGp', '9REEGm-REEGp'],
['9REEGf-REEGp', '6LEEGm-LEEGf'],
['4LEEGf-LEEGp','7REEGf-REEGp']]
### load ALL of EEG channels ####
channelsToLoadEEG = [['4 LEEGf-Fgr', '5 LEEGf-LEEGp', '6 LEEGm-LEEGp', '7 LEEGp-Fgr', '8 REEGp-Fgr','9 REEGp-LEEGp'],
['4 LEEGf-Fgr','5 LEEGf-LEEGm', '6 LEEGm-LEEGp', '7 REEGf-Fgr', '8 REEGm-Fgr', '9 REEGf-REEGm'],
['4LEEGf-LEEGp', '5LEEGf-LEEGm', '6LEEGm-LEEGp', '7REEGf-REEGp', '8REEGf-REEGm', '9REEGm-REEGp'],
['4LEEGf-LEEGp', '5LEEGm-LEEGp', '6LEEGm-LEEGf', '7REEGf-Fgr', '8REEGf-REEGm','9REEGf-REEGp',],
['4LEEGf-LEEGp', '5LEEGf-LEEGm', '6LEEGm-LEEGp', '7REEGf-REEGp', '8REEGf-REEGm', '9REEGm-REEGp']]
channelsToLoadEOG = [['1 LEOG-Fgr', '2 REOG-Fgr'],
['2 LEOG-Fgr', '3 REOG-Fgr'],
['2LEOG-Fgr', '3REOG-Fgr'],
['2LEOG-Fgr', '3REOG-Fgr'],
['2LEOG-Fgr', '3REOG-Fgr']]
birds_LL = [1,2,3]
nBirds_LL = len(birds_LL)
birdPaths_LL = ['../data_copies/01_PreprocessedData/02_BudgieMale_yellow1/01_Constant_light/',
'../data_copies/01_PreprocessedData/03_BudgieFemale_white1/01_Constant_light/',
'../data_copies/01_PreprocessedData/04_BudgieMale_yellow2/01_Constant_light/',]
arfFilePaths_LL = ['EEG 2 preprocessed/',
'EEG 2 preprocessed/',
'EEG 2 preprocessed/']
lightsOffSec = np.array([7947, 9675, 9861 + 8*3600, 9873, 13467]) # lights off times in seconds from beginning of file
lightsOnSec = np.array([46449, 48168, 48375+ 8*3600, 48381, 52005]) # Bird 3 gets 8 hours added b/c file starts at 8:00 instead of 16:00
epochLength = 3
sr = 200
scalingFactor = (2**15)*0.195 # scaling/conversion factor from amplitude to uV (when recording arf from jrecord)
stages = ['w','d','u','i','s','r'] # wake, drowsy, unihem sleep, intermediate sleep, SWS, REM
stagesSleep = ['u','i','s','r']
stagesVideo = ['m','q','d','s','u'] # moving wake, quiet wake, drowsy, sleep, unclear
## Path to scores formatted as CSVs
formatted_scores_path = '../formatted_scores/'
# +
colors = sns.color_palette(np.array([[234,103,99],
[218,142,60],
[174,174,62],
[97,188,101],
[140,133,232],
[225,113,190]])
/255)
sns.palplot(colors)
# colorpalette from iWantHue
# -
# Plot-specific info
# +
sns.set_context("notebook", font_scale=1.5)
sns.set_style("white")
# Markers for legends of EEG scoring colors
legendMarkersEEG = []
for stage in range(len(stages)):
legendMarkersEEG.append(plt.Line2D([0],[0], color=colors[stage], marker='o', linestyle='', alpha=0.7))
# -
# Calculate general variables
# +
lightsOffEp = lightsOffSec / epochLength
lightsOnEp = lightsOnSec / epochLength
nBirds = len(birdPaths)
epochLengthPts = epochLength*sr
nStages = len(stagesSleep)
# -
# ## Read in files
# ### Load formatted scores
AllScores = {}
for b in range(nBirds):
bird_name = 'Bird ' + str(b+1)
file = formatted_scores_path + 'All_scores_' + bird_name + '.csv'
data = pd.read_csv(file, index_col=0)
AllScores[bird_name] = data
# ### loadData
def loadData(b):
bird = 'Bird ' + str(b+1)
EEGdataAll = {}
arf_path = birdPaths[b] + arfFilePaths[b]
# Load EEG channels
for channel in channelsToLoadEEG[b]:
all_data_array = np.array([])
for file in np.sort(os.listdir(arf_path)):
if file.endswith('.arf'):
arffile = h5py.File(arf_path+file, 'r')
data_array = arffile['.'][channel][()]
data_array = np.ndarray.flatten(data_array)
all_data_array = np.append(all_data_array,data_array)
# Save in dict under bird number and channel
data_name = 'Bird ' + str(b+1) + ': ' + channel
EEGdataAll[data_name] = scalingFactor * all_data_array
EEGchannels = np.sort(list(EEGdataAll.keys()))
# Create time index for EEG
all_time_array = np.array([], dtype='datetime64')
for file in np.sort(os.listdir(arf_path)):
if file.endswith('.arf'):
arffile = h5py.File(arf_path+file, 'r')
date = file.split('_')[2]
if b == 0:
hours = '17'
minutes = '32'
else:
time = file.split('_')[3]
hours = time.split('-')[0]
minutes = time.split('-')[1]
datetime_start = np.datetime64(date + 'T' + hours + ':' + minutes + ':06') # assume 6-s delay in starting recording
# time index in datetime format
length_s = len(arffile['.'][channel][()])/sr
length_ms = np.timedelta64(int(1000 * length_s), 'ms')
datetime_end = datetime_start + length_ms
time_array = np.arange(datetime_start, datetime_end, np.timedelta64(int(1000/sr),'ms'))
# Add to end of whole-night time index
all_time_array = np.append(all_time_array, time_array)
TimeIndexEEG = {}
data_name = 'Bird ' + str(b+1)
TimeIndexEEG[data_name] = all_time_array
# Read in EOG traces
EOGdataAll = {}
arf_path = birdPaths[b] + arfFilePaths[b]
for channel in channelsToLoadEOG[b]:
all_data_array = np.array([])
for file in np.sort(os.listdir(arf_path)):
if file.endswith('.arf'):
arffile = h5py.File(arf_path+file, 'r')
data_array = arffile['.'][channel][()]
data_array = np.ndarray.flatten(data_array)
all_data_array = np.append(all_data_array,data_array)
# Save in dict under bird number and channel
data_name = 'Bird ' + str(b+1) + ': ' + channel
EOGdataAll[data_name] = scalingFactor * all_data_array
EOGchannels = np.sort(list(EOGdataAll.keys()))
return(bird, EEGdataAll, EEGchannels, TimeIndexEEG, EOGdataAll, EOGchannels)
# ## User-defined: which EEG channels to plot
AllEEGchannel_to_use = {}
# +
# Bird 1
AllEEGchannel_to_use['Bird 1'] = ['Bird 1: 5 LEEGf-LEEGp',
'Bird 1: 6 LEEGm-LEEGp',
'Bird 1: 8 REEGp-Fgr', ]
# +
# Bird 2
AllEEGchannel_to_use['Bird 2'] = ['Bird 2: 4 LEEGf-Fgr',
'Bird 2: 5 LEEGf-LEEGm']
# +
# Bird 3
AllEEGchannel_to_use['Bird 3'] = ['Bird 3: 4LEEGf-LEEGp',
'Bird 3: 6LEEGm-LEEGp',
'Bird 3: 9REEGm-REEGp', ]
# +
# Bird 4
AllEEGchannel_to_use['Bird 4'] = ['Bird 4: 4LEEGf-LEEGp',
'Bird 4: 5LEEGm-LEEGp',
'Bird 4: 6LEEGm-LEEGf',
'Bird 4: 7REEGf-Fgr',
'Bird 4: 9REEGf-REEGp']
# +
# Bird 5
AllEEGchannel_to_use['Bird 5'] = ['Bird 5: 4LEEGf-LEEGp',
'Bird 5: 6LEEGm-LEEGp',
'Bird 5: 7REEGf-REEGp', ]
# -
# # PlotFig1
def PlotFig1(b, startTime, EOGchannels, EEGchannels, dT=20, scoreBarWidth=10, ylimAmtEOG=250, yCalibBarEOG=200, xCalibBarEOG=1,
ylimAmtEEG=150, yCalibBarEEG=100, xCalibBarEEG=1,
sr=sr, colors=colors, stages=stages,
linewidth=1, plotSizeMultiplier=1):
'''Plot Figure 1: sleep score, EEG & EOG
b = bird name
startTime = where to start plotting, in seconds
dT = number of seconds to plot
ylimAmtEOG / EEG = set range of y axis above & below 0
yCalibBarEOG / EEG = how big to make the calibration bar for uV
xCalibBarEOG / EEG = how big to make the calibration bar for sec
sr = sampling rate
colors = list of colors to use for plotting sleep stages
stages = list of sleep/wake stages
EOGchannels = dictionary of EOG channels to use (for all birds)
EEGchannels = dictionary of EEG channels to use (for all birds)
'''
stopTime = startTime + dT
# Bird number from 0-4:
birdID = int(b[5])-1
# Get datetime index
time_index = TimeIndexEEG[b]
start_datetime_rec = time_index[0]
# calc start and stop datetimes
start_timedelta = startTime.astype('timedelta64[s]')
dt_timedelta = np.timedelta64(dT, 's')
start_datetime = start_datetime_rec + start_timedelta
stop_datetime = start_datetime + dt_timedelta
# Calculate start and stop in points
dP = dT*sr
startPts = np.where(time_index >= np.datetime64(start_datetime))[0][0]
stopPts = np.where(time_index <= np.datetime64(stop_datetime))[0][-1]
EOGtoPlot = [EOGchannels[x] for x in range(len(EOGchannels)) if b in EOGchannels[x]]
EEGtoPlot = [EEGchannels[x] for x in range(len(EEGchannels)) if b in EEGchannels[x]]
allChToPlot = np.concatenate((np.array(EOGtoPlot), np.array(EEGtoPlot)))
nChToPlot = len(allChToPlot)
plt.figure(figsize=(plotSizeMultiplier*.325*dT, (5/6)*nChToPlot+1))
row = 1
offset = 1/15 # fraction of plot size to leave blank on either side
###########################################################################
# PLOT SLEEP SCORES AS BAR AT TOP
plt.subplot(nChToPlot+1, 1, row)
# Plotting parameters
width = scoreBarWidth
scoreLoc = 0
# Get scores to plot
bird_scores = AllScores[b]
scoresToPlot = bird_scores[((bird_scores['Time (s)']+epochLength)>startTime)&(bird_scores['Time (s)']<=stopTime)]['Label'].values
firstEpOffset = (startTime%epochLength)*sr # how much of first epoch is cut off at beginning, in pts
nEpochs = len(scoresToPlot)
# replace 'l' or 'g' in "Scores to Plot" with 'u' for unihem
unihem_inds = [x for x in range(nEpochs) if ('l' in scoresToPlot[x])|('g' in scoresToPlot[x])]
scoresToPlot[unihem_inds] = 'u'
# 1. Plot first epoch (which might be cut off at beginning):
# determine color based on sleep stage
scoreNum = [x for x in range(len(stages)) if stages[x] in scoresToPlot[0]][0]
scoreColor = colors[scoreNum]
# determine where to draw the bar
start = 0
stop = epochLengthPts - firstEpOffset
# draw the bar
plt.hlines(scoreLoc, start, stop, color=scoreColor, linewidth=width)
# 2. Plot middle epochs
for ep in np.arange(1,nEpochs-1):
# determine color based on sleep stage
scoreNum = [x for x in range(len(stages)) if stages[x] in scoresToPlot[ep]][0]
scoreColor = colors[scoreNum]
# determine where to draw the bar
start = ep*epochLengthPts - firstEpOffset
stop = ep*epochLengthPts - firstEpOffset + epochLengthPts
# draw the bar
plt.hlines(scoreLoc, start, stop, color=scoreColor, linewidth=width)
# 3. Plot last epoch (which might be cut off at end)
lastEp = nEpochs-1
# determine color based on sleep stage
scoreNum = [x for x in range(len(stages)) if stages[x] in scoresToPlot[lastEp]][0]
scoreColor = colors[scoreNum]
# determine where to draw the bar
start = lastEp*epochLengthPts - firstEpOffset
stop = dP
# draw the bar
plt.hlines(scoreLoc, start, stop, color=scoreColor, linewidth=width)
# Get rid of axes
plt.yticks([])
plt.xticks([])
sns.despine(left=True, bottom=True)
plt.xlim(plt.xlim(-dP*offset, dP + dP*offset))
row = row+1
###########################################################################
# PLOT EOG CHANNELS
# Set plot characteristics
ylimAmt = ylimAmtEOG # set range of y axis above & below 0
yCalibBar = yCalibBarEOG # how big to make the calibration bar for uV
xCalibBar = xCalibBarEOG # how big to make the calibration bar for sec
eog_color = [142/255,134/255,137/255]
for ch in EOGtoPlot:
plt.subplot(nChToPlot+1, 1, row)
# Plot!
plt.plot(EOGdataAll[ch][startPts:stopPts], color=eog_color, lw=linewidth);
# Set axis limits
plt.ylim(-ylimAmt, ylimAmt)
plt.xlim(-dP*offset, dP + dP*offset)
# leave room on left side for labels
# leave room on right side for calibration bars
# Labels
plt.ylabel(ch[len(ch)-8:len(ch)-4], rotation=0, fontsize=14, color=eog_color)
# Get rid of axes
plt.yticks([])
plt.xticks([])
sns.despine(left=True, bottom=True)
row = row+1
# Calibration bars
plt.vlines(dP + dP*(offset/4), -yCalibBar - .75*yCalibBar, 0 - .75*yCalibBar);
#plt.hlines(-yCalibBar - .75*yCalibBar, dP + dP*(offset/4) - xCalibBar*sr, dP + dP*(offset/4) );
plt.text(1.01*(dP + dP*(offset/4)), -1.25*yCalibBar, str(yCalibBar)+' uV', fontsize=10)
#plt.text(dP - .5*xCalibBar*sr, -2.75*yCalibBar, str(xCalibBar)+' s', fontsize=10)
###########################################################################
# PLOT EEG CHANNELS
# Set plot characteristics
ylimAmt = ylimAmtEEG # set range of y axis above & below 0
yCalibBar = yCalibBarEEG # how big to make the calibration bar for uV
xCalibBar = xCalibBarEEG # how big to make the calibration bar for sec
for ch in EEGtoPlot:
plt.subplot(nChToPlot+1, 1, row)
# Plot!
plt.plot(EEGdataAll[ch][startPts:stopPts], color='k', lw=linewidth);
# Set axis limits
plt.ylim(-ylimAmt, ylimAmt)
plt.xlim(-dP*offset, dP + dP*offset)
# leave room on left side for labels
# leave room on right side for calibration bars
# Labels
if 'LEEG' in ch:
label_to_plot = 'LEEG'
elif 'REEG' in ch:
label_to_plot = 'REEG'
plt.ylabel(label_to_plot, rotation=0, fontsize=14)
# Get rid of axes
plt.yticks([])
plt.xticks([])
sns.despine(left=True, bottom=True)
row = row+1
# Calibration bars
plt.vlines(dP + dP*(offset/2), -yCalibBar - .75*yCalibBar, 0 - .75*yCalibBar);
plt.hlines(-yCalibBar - .75*yCalibBar, dP + dP*(offset/2) - xCalibBar*sr, dP + dP*(offset/2) );
plt.text(1.01*(dP + dP*(offset/2)), -0.6*yCalibBar, str(yCalibBar)+' uV', fontsize=10)
plt.text(dP - .75*xCalibBar*sr, -1.7*yCalibBar, str(xCalibBar)+' s', fontsize=10)
# # Bird 1
# +
bird = 'Bird 1'
b_num = int(bird[5]) - 1
bird, EEGdataAll, EEGchannels, TimeIndexEEG, EOGdataAll, EOGchannels = loadData(b_num)
EEGchannels_toUse = AllEEGchannel_to_use[bird]
# -
# ## Drowsy & Wake
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(4371,'s')).astype('int')
# hours minutes seconds
PlotFig1(bird, startTime, dT=20, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels)
if makePlots:
name = "Fig01b_drowsy_vsWake_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# ## Unihemispheric sleep
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(1647,'s')).astype('int')
# hours minutes seconds
PlotFig1(bird, startTime, dT=20, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels)
if makePlots:
name = "Fig01c_unihem_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# ## REM
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(23160,'s')).astype('int')
# hours minutes seconds
PlotFig1(bird, startTime, dT=20, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels)
if makePlots:
name = "Fig01d_REM_classic_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# ## IS
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(23421,'s')).astype('int')
# hours minutes seconds
PlotFig1(bird, startTime, dT=20, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels)
if makePlots:
name = "Fig01e_IS_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# ## SWS
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(18648,'s')).astype('int')
# hours minutes seconds
PlotFig1(bird, startTime, dT=20, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels)
if makePlots:
name = "Fig01f_SWS_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# ## REM theta
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(16800,'s')).astype('int')
# hours minutes seconds
# Double figure size to make this plot
PlotFig1(bird, startTime, dT=9, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels,
plotSizeMultiplier=2)
if makePlots:
name = "Fig01g_REM_theta_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# # Bird 4
bird, EEGdataAll, EEGchannels, TimeIndexEEG, EOGdataAll, EOGchannels = loadData(3)
bird
# +
# Bird 4
EEGchannels_toUse = ['Bird 4: 4LEEGf-LEEGp', 'Bird 4: 5LEEGm-LEEGp',
'Bird 4: 6LEEGm-LEEGf', 'Bird 4: 7REEGf-Fgr', 'Bird 4: 9REEGf-REEGp']
# +
bird = 'Bird 4'
b_num = int(bird[5]) - 1
bird, EEGdataAll, EEGchannels, TimeIndexEEG, EOGdataAll, EOGchannels = loadData(b_num)
EEGchannels_toUse = AllEEGchannel_to_use[bird]
# -
# ## K complexes
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(32154,'s')).astype('int')
# hours minutes seconds
# Double figure size to make this plot
PlotFig1(bird, startTime, dT=9, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels,
plotSizeMultiplier=2)
if makePlots:
name = "Fig01h_IS_Kcomplexes_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
# ## SWS vs IS
# +
startTime = (np.timedelta64(0,'h') + np.timedelta64(0,'m') + np.timedelta64(22431,'s')).astype('int')
# hours minutes seconds
PlotFig1(bird, startTime, dT=20, ylimAmtEEG=200, yCalibBarEEG=100,
ylimAmtEOG=200, yCalibBarEOG=100,
EEGchannels=EEGchannels_toUse, EOGchannels=EOGchannels)
if makePlots:
name = "Fig01i_SWS_toIS_quickly_" + bird + "_" + str(startTime) + "s"
plt.savefig(saveAsPath + name + ".pdf")
plt.savefig(saveAsPath + name + ".tiff")
# -
| Fig01 - PSG examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: keras
# language: python
# name: keras
# ---
# Load MNIST data.
# +
import utils
import numpy as np
import matplotlib.pyplot as plt
(train_x, train_y), (test_x, test_y) = utils.processed_data()
# -
# A fuzzy classifier is trained on the original feature space.
# +
import keras
import keras.layers as layers
import keras.models as models
import keras.regularizers as regularizers
import keras.backend as K
from model import LogGaussMF
def train_baseline():
model = keras.Sequential([
layers.Reshape((784,), input_shape=(28,28)),
LogGaussMF(10),
layers.Lambda(lambda x: K.sum(x, axis=-1)),
layers.Activation("softmax")
])
model.compile(
optimizer=keras.optimizers.adam(lr=0.0005),
loss="binary_crossentropy",
metrics=[keras.metrics.categorical_accuracy])
# train the model
history = model.fit(
x=train_x,
y=train_y,
batch_size=64,
epochs=20,
validation_data=(test_x, test_y),
verbose=1,
shuffle=True)
# plot the loss and accuracy
fig, axes = plt.subplots(1, 2, figsize=(8, 4), squeeze=True)
axes[0].set_title("Loss")
axes[0].plot(history.history["loss"], c="b")
axes[0].plot(history.history["val_loss"], c="r")
axes[1].set_title("Accuracy")
axes[1].plot(history.history["categorical_accuracy"], c="b")
axes[1].plot(history.history["val_categorical_accuracy"], c="r")
return model
baseline_model = train_baseline()
# -
# When the classifier is trained in the original feature space, mu and beta can be plotted as images in the original feature space. This will not work when convolutional layers are being used. Additionally, the images of mu and beta don't provide a lot of information.
# +
mu, beta = baseline_model.layers[-3].get_weights()
fig, axes = plt.subplots(2, 10, figsize=(10, 2))
for i in range(10):
axes[0][i].imshow(mu[i,:].reshape(28,28))
axes[1][i].imshow(beta[i,:].reshape(28,28))
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# One convoltutional layer is added to the model to improve is classification power.
# +
def train_conv1():
model = keras.Sequential([
layers.Reshape((28,28,1), input_shape=(28,28)),
layers.Conv2D(16, 3, padding="same"),
layers.Activation("relu"),
layers.Flatten(),
LogGaussMF(10),
layers.Lambda(lambda x: K.sum(x, axis=-1)),
layers.Activation("softmax")
])
model.compile(
optimizer=keras.optimizers.adam(lr=0.0001),
loss="binary_crossentropy",
metrics=[keras.metrics.categorical_accuracy])
# train the model
history = model.fit(
x=train_x,
y=train_y,
batch_size=128,
epochs=20,
validation_data=(test_x, test_y),
verbose=1,
shuffle=True)
# plot the loss and accuracy
fig, axes = plt.subplots(1, 2, figsize=(8, 4), squeeze=True)
axes[0].set_title("Loss")
axes[0].plot(history.history["loss"], c="b")
axes[0].plot(history.history["val_loss"], c="r")
axes[1].set_title("Accuracy")
axes[1].plot(history.history["categorical_accuracy"], c="b")
axes[1].plot(history.history["val_categorical_accuracy"], c="r")
return model
conv1_model = train_conv1()
# -
# The most important pixels for each class of image are determined using the following process. Guided backprop is run on each training sample of the same class, with the target neuron being that images class. The resulting images are then rectified and a single representative image is created by taking the mean of all the images. This is repeated for all 10 classes.
#
# The resulting images are scaled so that the values range from 0 to 1 and fed back into the classifier to ensure that it gives the correct prediction. Sometimes the mispredicts the image, but this is dealt with in the next cell.
# +
import innvestigate
from keras.models import Model
labels = np.argmax(train_y, axis=1)
model_wo_softmax = Model(conv1_model.inputs, [conv1_model.layers[-2].output])
analyzer = innvestigate.create_analyzer(
"guided_backprop",
model_wo_softmax,
neuron_selection_mode="index",
allow_lambda_layers=True)
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
images = train_x[labels == i]
analysis = analyzer.analyze(images, i)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
axes[i].imshow(analysis)
print(np.argmax(conv1_model.predict(analysis.reshape(1,28,28))))
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# A threshold between 0 and 1 is added, and all pixels below the threshold are set to zero. This seems to solved the problem of the model incorrectly classifying the representative images.
# +
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
images = train_x[labels == i]
analysis = analyzer.analyze(images, i)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.4)
axes[i].imshow(analysis)
print(np.argmax(conv1_model.predict(analysis.reshape(1,28,28))))
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# Changing the index of the neuron analyzed does not seem to affect the image at all, nor effect the classification result.
# +
fig, axes = plt.subplots(10, 10, figsize=(10,10))
for i in range(10):
for j in range(10):
images = train_x[labels == i]
analysis = analyzer.analyze(images, j)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.4)
axes[i][j].imshow(analysis)
print(np.argmax(conv1_model.predict(analysis.reshape(1,28,28))), end=", ")
print()
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# A larger convolutional model is trained.
# +
def train_conv2():
model = keras.Sequential([
layers.Reshape((28,28,1), input_shape=(28,28)),
layers.Conv2D(16, 3, padding="same"),
layers.Activation("relu"),
layers.Conv2D(32, 3, strides=2, padding="same"),
layers.Activation("relu"),
layers.Conv2D(64, 3, strides=2, padding="same"),
layers.Activation("relu"),
layers.Flatten(),
LogGaussMF(10),
layers.Lambda(lambda x: K.sum(x, axis=-1)),
layers.Activation("softmax")
])
model.compile(
optimizer=keras.optimizers.adam(lr=0.0001),
loss="binary_crossentropy",
metrics=[keras.metrics.categorical_accuracy])
# train the model
history = model.fit(
x=train_x,
y=train_y,
batch_size=128,
epochs=20,
validation_data=(test_x, test_y),
verbose=1,
shuffle=True)
# plot the loss and accuracy
fig, axes = plt.subplots(1, 2, figsize=(8, 4), squeeze=True)
axes[0].set_title("Loss")
axes[0].plot(history.history["loss"], c="b")
axes[0].plot(history.history["val_loss"], c="r")
axes[1].set_title("Accuracy")
axes[1].plot(history.history["categorical_accuracy"], c="b")
axes[1].plot(history.history["val_categorical_accuracy"], c="r")
return model
conv2_model = train_conv2()
# -
# The images are not as clear as the model with the single convolutional layer, but still can be clearly made out. The threshold value had to be changed to 0.3 for the network to correctly classify the representatives.
# +
model_wo_softmax = Model(conv2_model.inputs, [conv2_model.layers[-2].output])
analyzer = innvestigate.create_analyzer(
"guided_backprop",
model_wo_softmax,
neuron_selection_mode="index",
allow_lambda_layers=True)
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
images = train_x[labels == i]
analysis = analyzer.analyze(images, i)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.3)
axes[i].imshow(analysis)
print(np.argmax(conv1_model.predict(analysis.reshape(1,28,28))))
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# Again, changing out the neuron index has little to no effect on the image generated. I am not sure why this is case and it something that should be investigated.
# +
fig, axes = plt.subplots(10, 10, figsize=(10,10))
for i in range(10):
for j in range(10):
images = train_x[labels == i]
analysis = analyzer.analyze(images, j)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.4)
axes[i][j].imshow(analysis)
print(np.argmax(conv1_model.predict(analysis.reshape(1,28,28))), end=", ")
print()
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# The resnet20 model is loaded for analysis.
#
# **The cell after this cell will fail the first time the notebook is run. This cell will have to run again for the next cell to work correctly.**
# +
from keras.models import load_model
# this has to be run twice to avoid an error with layer names
resnet20 = load_model(
"saved_models/mnist_ResNet20v2_model.114.h5",
custom_objects={
"LogGaussMF": lambda **x: LogGaussMF(rules=10, **x)})
model_wo_softmax = Model(resnet20.inputs, [resnet20.layers[-2].output])
analyzer = innvestigate.create_analyzer(
"guided_backprop",
model_wo_softmax,
neuron_selection_mode="index",
allow_lambda_layers=True)
# -
# The images are fuzzier yet and the threshold value had to be changed to 0.38 in order for the model to make the correct predictions about the representatives. The results here imply that deeper, more complicated models cannot not be reversed as successfully as shallower models, and there is a trade-off between performance and interpretibility.
#
# Additionally only 1000 samples from each class are used to create the representatives, in order to reduce memory usage. This may have effected their appearance.
# +
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
images = train_x[labels == i][:1000].reshape(-1, 28, 28, 1)
analysis = analyzer.analyze(images, i)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.38)
axes[i].imshow(analysis.squeeze())
print(np.argmax(conv1_model.predict(analysis.reshape(1,28,28))))
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# The second convolutional model will be used for the analysis in the next part of this notebook for the sake of convenience. The representative images are generated and stored for later used.
# +
model_wo_softmax = Model(conv2_model.inputs, [conv2_model.layers[-2].output])
analyzer = innvestigate.create_analyzer(
"guided_backprop",
model_wo_softmax,
neuron_selection_mode="index",
allow_lambda_layers=True)
representatives = np.empty((10,28,28))
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
images = train_x[labels == i]
analysis = analyzer.analyze(images, i)
# rectify guided backprop results and get mean image
analysis = np.maximum(analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.3)
axes[i].imshow(analysis)
representatives[i] = analysis
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# The 10x10 grid below shows each of the representative images multiplied with another one of the 10 representatives. This is a heuristic way to determine how similar any two digits representatives are to each other.
#
# Some interesting things can be seen, like that the digit eight shared important pixels with nearly every other digit.
# +
fig, axes = plt.subplots(10, 10, figsize=(10,10))
for i in range(10):
for j in range(10):
image = representatives[i] * representatives[j]
print("{:4.1f}".format(np.sum(image)), end=", ")
axes[i][j].imshow(image)
print()
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# This grid shows the squared error between any two digits, another way to heuristically determine similarity between images.
# +
fig, axes = plt.subplots(10, 10, figsize=(10,10))
for i in range(10):
for j in range(10):
image = np.square(representatives[i] - representatives[j])
print("{:4.1f}".format(np.sum(image)), end=", ")
axes[i][j].imshow(image)
print()
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# The index of all incorrectly classified images is determined.
# +
test_labels = np.argmax(test_y, axis=1)
predictions = np.argmax(conv2_model.predict(test_x), axis=1)
incorrect_indexes = np.nonzero(predictions != test_labels)[0]
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
image = test_x[incorrect_indexes[i]]
axes[i].imshow(image)
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# This function compared the investigates the incorrectly labeled images looks at the similarities between the image and the digit representatives using multiplication (bottom row) and squared error (top row) heuristic methods.
#
# This examples is not explained well using these methods, but that is to be expected.
# +
def compare_image(idx):
print("Label: {}".format(test_labels[idx]))
print("Predictions: {}".format(predictions[idx]))
print(" MSE Overlap")
plt.figure(figsize=(1,1))
plt.imshow(test_x[idx])
fig, axes = plt.subplots(2, 10, figsize=(10,2))
for i in range(10):
print(i, end=": ")
image = test_x[idx]
difference = np.square(image - representatives[i])
overlap = image * representatives[i]
print("{:5.1f}".format(np.sum(difference)), end=", ")
axes[0][i].imshow(difference)
print("{:5.1f}".format(np.sum(overlap)))
axes[1][i].imshow(overlap)
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
compare_image(incorrect_indexes[0])
# -
# In this example, the 4 is misclassified as a 9. This can be explained as the image has the biggest overlap with the representative of 9, rather than 4. Additionally the sum squared error is lower for 4 than 9. Some of the next few examples display the same characteristics, while some do not (like the first example).
compare_image(incorrect_indexes[1])
compare_image(incorrect_indexes[2])
compare_image(incorrect_indexes[3])
compare_image(incorrect_indexes[4])
# There is a lot of overlap between these representatives, meaning showing only what pixels should be positive doesn't give us the full picture. The code below uses a technique similar to guided backprop to find what pixels need to not have data for an image to be classified correctly. Positive values are shown in red, and negative values are shown in blue.
# +
import tensorflow as tf
def guided_backprop(model, input_image, neuron_index=0):
grad = tf.gradients(
ys=model.layers[-2].output[:,neuron_index],
xs=model.layers[-2].input,
grad_ys=model.layers[-2].output[:,neuron_index],
stop_gradients=model.layers[-2].input)
for layer in model.layers[-3:0:-1]:
grad = tf.gradients(
ys=layer.output,
xs=layer.input,
grad_ys=grad[0],
stop_gradients=layer.input)
if getattr(layer, 'activation', None) == keras.activations.get("relu"):
pass
#grad = [tf.maximum(grad[0], 0.)]
sess = K.get_session()
grad_val = sess.run(
grad,
feed_dict={model.input: input_image})
return grad_val[0]
heatmaps = np.empty((10,28,28))
fig, axes = plt.subplots(1, 10, figsize=(10,1))
for i in range(10):
images = train_x[labels == i]
analysis = guided_backprop(conv2_model, images, i).squeeze()
# rectify guided backprop results and get mean image
analysis = np.maximum(-analysis, 0.)
analysis = np.mean(analysis, axis=0)
# normalize between 0 and 1
analysis -= np.min(analysis)
analysis /= np.max(analysis)
# remove any values below abritrary threshold
# improves the likelihood that image is classified correctly
analysis *= (analysis > 0.3)
axes[i].imshow(representatives[i] - analysis, cmap="seismic")
heatmaps[i] = representatives[i] - analysis
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
# -
# This function shows what makes similar comparisons, only now it uses the updated representatives.
# +
def compare_image_v2(idx):
print("Label: {}".format(test_labels[idx]))
print("Predictions: {}".format(predictions[idx]))
print(" MSE Overlap")
plt.figure(figsize=(1,1))
plt.imshow(test_x[idx])
fig, axes = plt.subplots(2, 10, figsize=(10,2))
for i in range(10):
print(i, end=": ")
image = test_x[idx]
difference = np.square(image - heatmaps[i])
overlap = image * heatmaps[i]
print("{:5.1f}".format(np.sum(difference)), end=", ")
axes[0][i].imshow(difference, vmin=0., vmax=1.)
print("{:5.1f}".format(np.sum(overlap)))
axes[1][i].imshow(overlap, cmap="seismic", vmin=-1., vmax=1.)
plt.setp(axes, xticks=[], yticks=[], frame_on=False)
plt.tight_layout(h_pad=0, w_pad=0)
compare_image_v2(incorrect_indexes[0])
# -
compare_image_v2(incorrect_indexes[1])
compare_image_v2(incorrect_indexes[2])
# This sample can now be explained, where as before it was misclassified despite having the more overlap with the representative for the digit 7 rather than 2. Now we can see it has more positive overlap with the representative for 2 as opposed to 7. This can be used to explain why its misclassified.
compare_image_v2(incorrect_indexes[3])
compare_image_v2(incorrect_indexes[4])
| mnist/rule-propagation-centroids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '2'
import bert
from bert import run_classifier
from bert import optimization
from bert import tokenization
from bert import modeling
import numpy as np
import tensorflow as tf
import pandas as pd
from tqdm import tqdm
# +
from prepro_utils import preprocess_text, encode_ids, encode_pieces
import sentencepiece as spm
sp_model = spm.SentencePieceProcessor()
sp_model.Load('bert-base/sp10m.cased.v4.model')
with open('bert-base/sp10m.cased.v4.vocab') as fopen:
v = fopen.read().split('\n')[:-1]
v = [i.split('\t') for i in v]
v = {i[0]: i[1] for i in v}
class Tokenizer:
def __init__(self, v):
self.vocab = v
pass
def tokenize(self, string):
return encode_pieces(sp_model, string, return_unicode=False, sample=False)
def convert_tokens_to_ids(self, tokens):
return [sp_model.PieceToId(piece) for piece in tokens]
def convert_ids_to_tokens(self, ids):
return [sp_model.IdToPiece(i) for i in ids]
tokenizer = Tokenizer(v)
# +
import json
import glob
left, right, label = [], [], []
for file in glob.glob('../Malaya-Dataset/text-similarity/quora/*.json'):
with open(file) as fopen:
x = json.load(fopen)
for i in x:
splitted = i[0].split(' <> ')
if len(splitted) != 2:
continue
left.append(splitted[0])
right.append(splitted[1])
label.append(i[1])
# -
BERT_INIT_CHKPNT = 'bert-base/model.ckpt'
BERT_CONFIG = 'bert-base/bert_config.json'
left[0], right[0], label[0]
np.unique(label, return_counts = True)
# +
from tqdm import tqdm
MAX_SEQ_LENGTH = 100
def _truncate_seq_pair(tokens_a, tokens_b, max_length):
while True:
total_length = len(tokens_a) + len(tokens_b)
if total_length <= max_length:
break
if len(tokens_a) > len(tokens_b):
tokens_a.pop()
else:
tokens_b.pop()
def get_inputs(left, right):
input_ids, input_masks, segment_ids = [], [], []
for i in tqdm(range(len(left))):
tokens_a = tokenizer.tokenize(left[i])
tokens_b = tokenizer.tokenize(right[i])
_truncate_seq_pair(tokens_a, tokens_b, MAX_SEQ_LENGTH - 3)
tokens = []
segment_id = []
tokens.append("<cls>")
segment_id.append(0)
for token in tokens_a:
tokens.append(token)
segment_id.append(0)
tokens.append("<sep>")
segment_id.append(0)
for token in tokens_b:
tokens.append(token)
segment_id.append(1)
tokens.append("<sep>")
segment_id.append(1)
input_id = tokenizer.convert_tokens_to_ids(tokens)
input_mask = [1] * len(input_id)
while len(input_id) < MAX_SEQ_LENGTH:
input_id.append(0)
input_mask.append(0)
segment_id.append(0)
input_ids.append(input_id)
input_masks.append(input_mask)
segment_ids.append(segment_id)
return input_ids, input_masks, segment_ids
# -
input_ids, input_masks, segment_ids = get_inputs(left, right)
bert_config = modeling.BertConfig.from_json_file(BERT_CONFIG)
epoch = 20
batch_size = 60
warmup_proportion = 0.1
num_train_steps = int(len(left) / batch_size * epoch)
num_warmup_steps = int(num_train_steps * warmup_proportion)
class Model:
def __init__(
self,
dimension_output,
learning_rate = 2e-5,
training = True
):
self.X = tf.placeholder(tf.int32, [None, None])
self.segment_ids = tf.placeholder(tf.int32, [None, None])
self.input_masks = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None])
model = modeling.BertModel(
config=bert_config,
is_training=training,
input_ids=self.X,
input_mask=self.input_masks,
token_type_ids=self.segment_ids,
use_one_hot_embeddings=False)
output_layer = model.get_pooled_output()
self.logits = tf.layers.dense(output_layer, dimension_output)
self.logits = tf.identity(self.logits, name = 'logits')
self.cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits = self.logits, labels = self.Y
)
)
self.optimizer = optimization.create_optimizer(self.cost, learning_rate,
num_train_steps, num_warmup_steps, False)
correct_pred = tf.equal(
tf.argmax(self.logits, 1, output_type = tf.int32), self.Y
)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# +
dimension_output = 2
learning_rate = 2e-5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
dimension_output,
learning_rate
)
sess.run(tf.global_variables_initializer())
var_lists = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope = 'bert')
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, BERT_INIT_CHKPNT)
# +
from sklearn.model_selection import train_test_split
train_input_ids, test_input_ids, train_input_masks, test_input_masks, train_segment_ids, test_segment_ids, train_Y, test_Y = train_test_split(
input_ids, input_masks, segment_ids, label, test_size = 0.2)
# +
from tqdm import tqdm
import time
EARLY_STOPPING, CURRENT_CHECKPOINT, CURRENT_ACC, EPOCH = 2, 0, 0, 0
while True:
lasttime = time.time()
if CURRENT_CHECKPOINT == EARLY_STOPPING:
print('break epoch:%d\n' % (EPOCH))
break
train_acc, train_loss, test_acc, test_loss = [], [], [], []
pbar = tqdm(
range(0, len(train_input_ids), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_input_ids))
batch_x = train_input_ids[i: index]
batch_masks = train_input_masks[i: index]
batch_segment = train_segment_ids[i: index]
batch_y = train_Y[i: index]
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.segment_ids: batch_segment,
model.input_masks: batch_masks
},
)
assert not np.isnan(cost)
train_loss.append(cost)
train_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_input_ids), batch_size), desc = 'test minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_masks = test_input_masks[i: index]
batch_segment = test_segment_ids[i: index]
batch_y = test_Y[i: index]
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.segment_ids: batch_segment,
model.input_masks: batch_masks
},
)
test_loss.append(cost)
test_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss = np.mean(train_loss)
train_acc = np.mean(train_acc)
test_loss = np.mean(test_loss)
test_acc = np.mean(test_acc)
if test_acc > CURRENT_ACC:
print(
'epoch: %d, pass acc: %f, current acc: %f'
% (EPOCH, CURRENT_ACC, test_acc)
)
CURRENT_ACC = test_acc
CURRENT_CHECKPOINT = 0
else:
CURRENT_CHECKPOINT += 1
print('time taken:', time.time() - lasttime)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
EPOCH += 1
# -
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'bert-base-similarity/model.ckpt')
# +
dimension_output = 2
learning_rate = 2e-5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
dimension_output,
learning_rate,
training = False
)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.trainable_variables())
saver.restore(sess, 'bert-base-similarity/model.ckpt')
# -
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
]
)
strings.split(',')
# +
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_input_ids), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_masks = test_input_masks[i: index]
batch_segment = test_segment_ids[i: index]
batch_y = test_Y[i: index]
predict_Y += np.argmax(sess.run(model.logits,
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
model.segment_ids: batch_segment,
model.input_masks: batch_masks
},
), 1, ).tolist()
real_Y += batch_y
# +
from sklearn import metrics
print(
metrics.classification_report(
real_Y, predict_Y, target_names = ['not similar', 'similar'],
digits = 5
)
)
# -
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('bert-base-similarity', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
g = load_graph('bert-base-similarity/frozen_model.pb')
x = g.get_tensor_by_name('import/Placeholder:0')
segment_ids = g.get_tensor_by_name('import/Placeholder_1:0')
input_masks = g.get_tensor_by_name('import/Placeholder_2:0')
logits = g.get_tensor_by_name('import/logits:0')
test_sess = tf.InteractiveSession(graph = g)
result = test_sess.run(tf.nn.softmax(logits), feed_dict = {x: batch_x,
segment_ids: batch_segment,
input_masks: batch_masks})
result
# +
import boto3
bucketName = 'huseinhouse-storage'
Key = 'bert-base-similarity/frozen_model.pb'
outPutname = "v30/similarity/bert-base-similarity.pb"
s3 = boto3.client('s3',
aws_access_key_id='',
aws_secret_access_key='')
s3.upload_file(Key,bucketName,outPutname)
# -
| session/similarity/bert-base.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 ('data_science_py395')
# language: python
# name: python3
# ---
# Programación para *Data Science*
# ============================
#
# Intro101 - 07a Conceptos avanzados de Python
# --------------------------------------
#
# En este Notebook encontraréis dos conjuntos de ejercicios de Python.
# ### Ejercicio 1
#
# Un número primo es aquél que solo es divisible por él mismo y por 1.
#
# a) Escribe un código que compruebe si un número `x = 15` es solo divisible por 1 o por el mismo. Escribe este código usando un iterador (un `for` o un `while`) que barra todos los valores desde `2` a `x-1`. Crea una variable `divisible` que tenga por defecto valor `False` y asigne el valor `True` si a lo largo de la iteración encuentra un número natural divisible. Puedes usar el operador modulo `a % b` para saber si un numero `b` es divisible por `a`.
# +
# Respuesta
# -
# b) Convierte tu código anterior en una función que compruebe si el número del argumento es primo o no, devolviendo True is es primo y False si no es primo. Comprueba tu función con los valores 492366587, 492366585, 48947 y 2,
#
# +
# Respuesta
# -
# c) En el cálculo de la función anterior, una vez se ha encontrado un número que es divisible dentro del rango ya no tiene sentido comprobar el resto de números del rango. Por ejemplo si 10 ya es divisble entre 2, ya no hace falta probar de 3 en adelante pues ya sabemos que el número no es primo.
#
# Modifica la función anterior de la siguiente forma:
# - Una vez se encuentra el divisor, la iteración se interrumpe para no probar el resto de enteros.
# - La función devuelve
# - **Si es primo**: True
# - **Si no es primo**, el primer divisor mayor que 1.
#
# Puedes hacer uso del comando *break* dentro de un bucle para interrumpir este, puedes consultar más información sobre break en la documentación de python [aquí](https://docs.python.org/2/tutorial/controlflow.html).
#
# Comprueba tu función con los valores 492366585, 492366587, 48947 y 2,
# +
# Respuesta
# -
# ### Ejercicio 2
#
# La Covid-19 es una enfermedad producida por la infección del virus SARS-CoV-2. La infección es transmisible de persona a persona y su contagiosidad depende de la cantidad del virus en las vías respiratorias. Si cada persona contagiada transmite la enfermedad a $\beta$ contactos en promedio por periodo de tiempo $t$, es posible estimar la evolución del contagio con un modelo matemático sencillo.
#
# Para $t=1$día, las transmisiones en España se han estimado a partir de su histórico de las semanas de Febrero y Marzo del 2020 una $\beta = 0.35$ transmissiones por día por infectado.
#
# Durante un periodo de tiempo (por ejempo un día $d$) la tasa de nuevos contagios se puede estimar como una proporción al número de contagiados del periodo anterior $N$:
#
# $ \Delta N = N_{1} - N = \beta \cdot $ (1)
#
# Por tanto, podemos proyectar el número futuro de afectados como
#
# $ N_{1} = N + \beta \cdot N = (1+\beta) \cdot $ (2)
#
# En dos días:
#
# $ N_{2} = (1+\beta) \cdot N_{1} = (1+\beta)^2 \cdot $ (3)
#
# Y en general en D días tendremos
#
# $ N_{D} = (1+\beta)^D \cdot N$ (4)
#
# Asumiendo este sencillo modelo:
#
# a) Implementa una función de dos parámetros (N: población infectada inicial, D: número de días), que devuelva el cálculo de afectados para D días siguiendo la ecuación (4). Suponiendo una población afectada de 4250 (población afectada en españa a día 13 de Marzo de 2020), usa la función para calcular la población estimada en 1, 2, 7 y 30 días.
# Respuesta
# b) Sabiendo que los Servicios de Medicina Intensiva (SMI) disponen de 3363 camas para enfermos graves, y suponiendo que un 10% de los afectados por el covid-19 requerirán de SMI y una supervivencia del 2,5% (Exitus), escribe un código que calcule:
# - El día en curso (Día)
# - El total de afectados por el virus para cada día d (Afectados)
# - El total de ingresados en SMI por el virus para cada día d (Críticos)
# - El total de Exitus por el virus para cada día d (Exitus)
# - Si los servicios de SMI no pueden aceptar los ingresados para cada día $d$ (Estado: indicando Saturación/No Saturación)
#
# Imprime en pantalla la información de cada día durante una simulación de tres semanas, suponiendo que no hay recuperaciones, con una población afectada inicial 4250 y una $\beta = 0.35$ constante.
# Respuesta
# c) Convierte el código anterior en una función que genere un archivo de texto con nombre `output.txt`, siguiendo este formato:
# ```
# Dia, Afectados, Críticos, Exitus, Estado
# 0, 4250, 425, 106, No Saturación
# 1, 5737, 573, 143, No Saturación
# 2, 7745, 774, 193, No Saturación
# ...
# ```
# Con los parámetros de entrada $N$, $D$, $\beta$, camas SMI.
# Respuesta
# ### Ejercicio 3
#
# Dado el siguiente diccionario:
d = {"Alex":344334443, "Eva":5533443, "Cristina":443355, "Jonas":33223324}
# Escribid una función que pregunte al usuario que introduzca el nombre de una persona y muestre por pantalla el nombre de la persona y su teléfono.
#
# Tened en cuenta que:
#
# - La función debe controlar que el valor introducido por el usuario es un nombre que existe en el diccionario. En caso contrario, mostrará un mensaje de error ("El nombre introducido no corresponde a ninguna persona") y devolverá el valor False.
# - Debéis tener en cuenta que el nombre de las personas que nos pasan por parámetro puede ser en minúsculas, mayúsculas o una combinación de ambas, y que debemos encontrar el número de teléfono aunque la capitalización de la cadena entrada por el usuario no sea exactamente la misma que hemos almacenado en el diccionario.
# - Suponed que no hay acentos en los nombres.
#
# Nota 1: Para realizar la actividad, tendréis que capturar un texto que entrará el usuario. Consultad la [documentación oficial de la función input](https://docs.python.org/3/library/functions.html#input) para ver cómo hacerlo.
#
# Nota 2: También tendréis que pensar cómo tratar el hecho de que el usuario pueda utilizar mayúsculas y minúsculas en la escritura del nombre en el diccionario. ¡Os animamos a usar un buscador para intentar encontrar alguna alternativa para resolver este subproblema! ¡Recordad citar las referencias que hayáis usado para resolverlo!
#
# +
d = {"Alex":344334443, "Eva":5533443, "Cristina":443355, "Jonas":33223324}
# Respuesta
# -
# ### Ejercicio 4
#
# Python dispone de un **idiom** muy útil conocido como `list comprehension`. Utilizando este **idiom**, proporcionad una expresión que devuelva las listas siguientes.
#
# Nota: Para realizar esta actividad necesitaréis investigar qué son las `list comprehension` y qué sintaxis utilizan. Para ello, se recomienda en primer lugar que utilicéis un buscador para encontrar información genérica sobre esta construcción. Después, os recomendamos que consultéis stackoverflow para ver algunos ejemplos de problemas que se pueden resolver con esta construcción.
#
#
# [stackoverflow](https://stackoverflow.com/) es un sitio de preguntas-y-respuestas muy popular entre programadores. Veréis que para la gran mayoría de las dudas que tengáis, habrá alguien que ya les habrá tenido (y consultado) anteriormente! Así pues, más allá de preguntar vosotros mismos las dudas allí (nosotros ya tenemos el foro del aula para ello!), consultar esta web os permitirá ver qué soluciones proponen otros programadores a estas dudas. A menudo habrá más de una solución a un mismo problema, y podréis valorar cuál es la más adecuada para vuestro problema.
#
# Para ver ejemplos de problemas que son adecuados para resolver con **list comprehensions**, os recomendamos leer las siguientes páginas:
# * https://stackoverflow.com/questions/12555443/squaring-all-elements-in-a-list
# * https://stackoverflow.com/questions/18551458/how-to-frame-two-for-loops-in-list-comprehension-python
# * https://stackoverflow.com/questions/24442091/list-comprehension-with-condition
# * https://stackoverflow.com/questions/41676212/i-want-to-return-only-the-odd-numbers-in-a-list
# * https://stackoverflow.com/questions/4260280/if-else-in-a-list-comprehension
#
# a) Una lista con los valores $4 x^2$ donde $x$ es cada uno de los números de la lista `list_1`:
# +
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9]
# Respuesta
# -
# b) Una lista con los valores $x/(x+1)$ donde $x$ es cada uno de los números de la lista `list_1`:
# +
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9]
# Respuesta
# -
# c) Una lista con los valores $4x^2/(4x^2-1)$ donde $x$ es cada uno de los números de la lista `list_1`:
# +
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9]
# Respuesta
# -
# ### Ejercicio 5
#
# El siguiente ejercicio consiste en pasar un número en base 16 (hexadecimal, 0-9/A-F) a base 10 (decimal). Para hacerlo, debéis crear una **función** que dado un _string_ que representa un número en hexadecimal, por ejemplo, `AE3F`, devuelva el número
# natural correspondiente. En este caso, el resultado sería `44607`.
# +
# Respuesta
# -
# ### Ejercicio 6
#
# Las excepciones son errores detectados en tiempo de ejecución. Pueden y deben ser manejadas por el programador para minimizar el riesgo de que un determinado programa falle de forma no controlada. Escribid, en lenguaje Python, cómo generar y capturar la siguiente excepción: **ZeroDivisionError**.
# +
# Respuesta
# -
# ### Ejercicio 7
#
# Completad el código necesario para calcular el número de vocales y de consonantes respectivamente de un texto.
# +
def contar_vocales_y_consonantes(texto):
# Cuenta las vocales contenidas en el string texto y también las consonantes.
num_vocales = 0
num_consonantes = 0
# Código que hay que completar.
return num_vocales, num_consonantes
texto = "Orbiting Earth in the spaceship, I saw how beautiful our planet is. \
People, let us preserve and increase this beauty, not destroy it!"
num_vocales, num_consonantes = contar_vocales_y_consonantes(texto)
print("El número de vocales es %d." % num_vocales)
print("El número de consonantes es %d." % num_consonantes)
# -
# ### Ejercicio 8
#
# Escribid una función que dada una lista de planetas del sistema solar, pregunte al usuario que introduzca una posición y muestre el plante correspondiente a dicha posición. Por ejemplo, si tenemos la siguiente lista: `['Mercurio', 'Venus', 'Tierra', 'Marte']` y el usuario nos ha introducido la posición `3`, hemos de mostrar como resultado por pantalla: `Tierra`.
#
# Consideraciones:
#
# - La posición que introduzca el usuario tiene que ser un número entero estrictamente positivo.
# - La función debe controlar el acceso a una posición fuera de la lista mediante una **excepción**. Por ejemplo, en el caso anterior debemos mostrar una mensaje de error si el usuario pide acceder a la posición 10.
# +
# Respuesta
# -
# ### Ejercicio 9
#
# Dada una lista de planetas del sistema solar, determinad cuales de estos planetas tienen una masa superior a la de la Tierra. Por ejemplo, si la lista inicial es `['Venus', 'Marte', 'Saturno']`, el resultado que mostraríamos por pantalla sería `['Saturno']` ya que el planeta Saturno tiene una masa `95.2` veces superior a la Tierra.
#
# Consideraciones:
#
# - Debéis tener en cuenta que el nombre de los planetas que nos pasan por parámetro puede estar en minúsculas, mayúsculas o una combinación de ambas.
# - Podéis asumir que no habrá acentos en el nombre de los planetas.
# - Debéis determinar aquellos planetas que tiene una massa estrictamente superior a la de la Tierra.
# - No habrá planetas repetidos en la lista que nos pasan por parámetro.
masas = {'Mercurio': 0.06, 'Venus': 0.82, 'Tierra': 1, 'Marte': 0.11, 'Jupiter': 317.8,
'Saturno': 95.2, 'Urano': 14.6, 'Neptuno': 17.2, 'Pluto': 0.0022}
# +
def planetas_mas_grandes_que_Tierra(planetas):
"""
Planetas con una masa superior a la de la Tierra
"""
planetas_masa_superior = []
# Código a completar
return planetas_masa_superior
# Ejemplos de uso de la función anterior
print(planetas_mas_grandes_que_Tierra(['Venus', 'Mercurio', 'Marte']))
print(planetas_mas_grandes_que_Tierra(['Jupiter', 'Saturno', 'Pluto']))
print(planetas_mas_grandes_que_Tierra(['urano', 'tierra', 'neptuno', 'marte', 'Venus']))
print(planetas_mas_grandes_que_Tierra(['Tierra', 'MeRcUrIo', 'PLUTO', 'SATURNO']))
# Podéis añadir más ejemplos si lo consideráis oportuno
# -
# ### Ejercicio 10
#
# Dada una cadena de caracteres, `s`, de longitud `n` y un número entero positivo `k`, siendo `k` un divisor de `n`, podemos dividir la cadena `s` en `n / k` sub-cadenas de la misma longitud.
#
# Escribid una función que, dada una cadena `s` y un número entero `k`, devuelva las `n/k` sub-cadenas teniendo en cuenta las siguientes consideraciones:
#
# - El orden de los caracteres en las sub-cadenas debe ser el mismo que en la cadena original.
# - Todos los caracteres de las sub-cadenas deben aparecer una única vez. Es decir, si un caracter se repite dentro de una sub-cadena, sólo hemos de mostrar la primera ocurrencia.
#
# Por ejemplo, si tenemmos
# <code>
# s = AABCCAADA
# k = 3
# </code>
#
# el resultado a mostrar por pantalla sería:
# <code>
# AB
# CA
# AD
# </code>
#
# Tenemos que la longitud de la cadena es 9 y por lo tanto, podemos formar 3 sub-cadenas:
#
# `AAB -> AB` (el caracter A se repite dos veces)
#
# `CCA -> CA` (el caracter C se repite dos veces)
#
# `ADA -> AD` (el caracter A se repite dos veces)
# +
# Respuesta
| 02_PYTHON/week06/pra/07a_python_avanzando.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
# !pip install nbdev -q
#hide
from nbdev.showdoc import *
# # Icevision
#
# > a computer vision framework for end-to-end training of curated models
# IceVision is the first agnostic computer vision framework to offer a curated collection with hundreds of high-quality pre-trained models from [torchvision](https://pytorch.org/vision/stable/index.html), [MMLab](https://openmmlab.com/), Ultralytics' [yolov5](https://github.com/ultralytics/yolov5) and <NAME>ightman's [EfficientDet](https://github.com/rwightman/efficientdet-pytorch). It orchestrates the end-to-end deep learning workflow allowing to train networks with easy-to-use robust high-performance libraries such as [Pytorch-Lightning](https://www.pytorchlightning.ai/) and [Fastai](https://docs.fast.ai/).
# ##Features of Icevision
# * Data curation/cleaning with auto-fix
# * Access to an exploratory data analysis dashboard
# * Pluggable transforms for better model generalization
# * Access to hundreds of neural net models
# * Access to multiple training loop libraries
# * Multi-task training to efficiently combine object detection, segmentation, and classification models
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# # [Join our Forum](https://discord.gg/JDBeZYK)
# ##Quick Example: How to train the Fridge Objects Dataset
# ```python
# from icevision.all import *
#
# #Model
# model_type = models.mmdet.retinanet
#
# #Backbone form MMDetection Example
# backbone = model_type.backbones.resnet50_fpn_1x
#
# #Loading Data
# data_dir = icedata.fridge.load_data()
#
# #Parsing
# parser = icedata.fridge.parser(data_dir)
# train_records, valid_records = parser.parse()
#
# #Transforms
# train_tfms = tfms.A.Adapter([*tfms.A.aug_tfms(size=image_size, presize=512), tfms.A.Normalize()])
# valid_tfms = tfms.A.Adapter([*tfms.A.resize_and_pad(image_size), tfms.A.Normalize()])
#
# # Datasets
# train_ds = Dataset(train_records, train_tfms)
# valid_ds = Dataset(valid_records, valid_tfms)
#
# #Create model object
# model = model_type.model(backbone=backbone(pretrained=True), num_classes=len(parser.class_map))
#
# #Learner
# metrics = [COCOMetric(metric_type=COCOMetricType.bbox)]
# learn = model_type.fastai.learner(dls=[train_dl, valid_dl], model=model, metrics=metrics)
#
# #LR Finder
# learn.lr_find()
#
# #Training
# learn.fine_tune(20, 1e-4, freeze_epochs=1)
# ```
# #Installing Icevision
# > Important: We currently only support Linux/MacOS installations
# > Note: Please do not forget to install the other optional dependencies if you would like to use them: MMCV+MMDetection, and/or YOLOv5
# ##Installation on colab
# ```python
# #check cuda version
# import torch
# cuda_version_major = int(torch.version.cuda.split('.')[0])
# cuda_version_major
# ```
# ```python
# #install packages based on your cuda version
# # !wget https://raw.githubusercontent.com/airctic/icevision/master/install_colab.sh
# # !bash install_colab.sh {cuda_version_major}
# ```
# ```python
# # Restart kernel
# import IPython
# IPython.Application.instance().kernel.do_shutdown(True)
# ```
# ##Installation using pip
# Option 1: Installing from pypi repository [Stable Version]
#
# `pip install icevision[all]`
# Option 2: Installing an editable package locally [For Developers]
# > Note: This method is used by developers who are usually either: actively contributing to icevision project by adding new features or fixing bugs, or
# creating their own extensions, and making sure that their source code stay in sync with the icevision latest version.
#
# ```bash
# $ got clone --depth=1
# https://github.com/airctic/icevision.git
#
# $ cd icevision
# $ pip install -e .[all,dev]
# $ pre-commit istall
# ```
# Option 3: Installing a non-editable package from GitHub:
# To install the icevision package from its GitHub repo, run the command here below. This option can be used in Google Colab, for example, where you might install the icevision latest version (from the master branch)
# ```pip install git+git://github.com/airctic/icevision.git#egg=icevision[all] --upgrade```
# ##Installation using conda
# Creating a conda environment is considered as a best practice because it avoids polluting the default (base) environment, and reduces dependencies conflicts. Use the following command in order to create a conda environment called icevision.
# ```
# $ conda create -n icevision python=3.8 anaconda
# $ conda activate icevision
# $ pip install icevision[all]
# ```
# #Installing optional dependencies
# ##MMDetection Installation
# We need to provide the appropriate version of the mmcv-full package as well as the cuda and the torch versions. Here are some examples for both the CUDA and the CPU versions.
# > Important: For the torch version use `torch.__version__` and replace the last number with 0. For the cuda version use: `torch.version.cuda.
# Example: TORCH_VERSION = torch1.8.0; CUDA_VERSION = cu101`
# For CUDA version,
# ```python
# $ pip install mmcv-full=="1.3.3" -f https://download.openmmlab.com/mmcv/dist/CUDA_VERSION/TORCH_VERSION/index.html --upgrade
# $ pip install mmdet
# ```
# For CPU version,
# ```python
# $ pip install mmcv-full=="1.3.3+torch.1.8.0+cpu" -f https://download.openmmlab.com/mmcv/dist/index.html --upgrade
# $ pip install mmdet
# ```
# ##YOLOv5 Installation
# You can install yolov5 by:
#
# `pip install yolov5-icevision --upgrade`
# ##Troubleshooting
# ###MMCV is not installing with cuda support
#
# If you are installing MMCV from the wheel like described above and still are having problems with CUDA you will probably have to compile it locally. Do that by running:
#
# `pip install mmcv-full`
#
# If you encounter the following error it means you will have to install CUDA manually (the one that comes with conda installation will not do).
#
# `OSError: CUDA_HOME environment variable is not set. Please set it to your CUDA install root.`
#
# Try installing it with:
#
# `sudo apt install nvidia-cuda-toolkit`
#
# Check the installation by running:
#
# `nvcc --version`
#
# Error: Failed building wheel for pycocotools
# If you encounter the following error, when installation process is building wheel for pycocotools:
#
# ```python
# unable to execute 'gcc': No such file or directory
# error: command 'gcc' failed with exit status 1
# ```
#
# Try installing gcc with:
#
# `sudo apt install gcc`
#
# Check the installation by running:
#
# `gcc --version`
#
# It should return something similar:
#
# ```
# gcc (Ubuntu 9.3.0-17ubuntu1~20.04) 9.3.0
# Copyright (C) 2019 Free Software Foundation, Inc.
# This is free software; see the source for copying conditions. There is NO
# warranty; not even for MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE.
# ```
#
# After that try installing icevision again.
#
# If you need any assistance, feel free to:
#
# # [Join our Forum](https://discord.gg/JDBeZYK)
| index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import face_recognition
import cv2
import pandas as pd
import numpy as np
# This is a demo of running face recognition on live video from your webcam. It's a little more complicated than the
# other example, but it includes some basic performance tweaks to make things run a lot faster:
# 1. Process each video frame at 1/4 resolution (though still display it at full resolution)
# 2. Only detect faces in every other frame of video.
# PLEASE NOTE: This example requires OpenCV (the `cv2` library) to be installed only to read from your webcam.
# OpenCV is *not* required to use the face_recognition library. It's only required if you want to run this
# specific demo. If you have trouble installing it, try any of the other demos that don't require it instead.
# Get a reference to webcam #0 (the default one)
# Create arrays of known face encodings and their names
known_face=[]
known_face_names=[]
known_face=pd.read_csv("names", header=None).values
#print((known_face[0][0]))
for i in range(len(known_face)):
known_face_names=np.append(known_face_names,known_face[i][0])
"""
known_face_names = [
"<NAME>",
"<NAME>"
]
"""
# Load a sample picture and learn how to recognize it.
#obama_image = face_recognition.load_image_file("known/Chetan Madan.jpg")
# -
known_face_names
len(df)
# +
known_face_encodings=[[]]
known_face_encoding=[[]]
face_encoding=[]
i=0
for name in known_face_names:
i+=1
print(name,i)
file_name="known/"+name+".jpg"
img_file=face_recognition.load_image_file(str(file_name))
df=face_recognition.face_encodings(img_file)[0]
known_face_encoding=np.append(known_face_encoding,df)
known_face_encodings=known_face_encoding.reshape(-1,128)
#known_face_encodings=np.append(known_face_encodings,known_face_encoding)
#obama_face_encoding = face_recognition.face_encodings(face_recognition.load_image_file("known/Chetan Madan.jpg"))[0]
i=0
# -
# for name in known_face_names:
# known_face_encodings=np.append(known_face_encodings, known_face_encoding[i])
#
#
#
# # Load a second sample picture and learn how to recognize it.
# biden_image = face_recognition.load_image_file("known/Saksham Madan.jpg")
# biden_face_encoding = face_recognition.face_encodings(biden_image)[0]
#
# known_face_encodings=known_face_encoding
# known_face_encodings = [
#
# biden_face_encoding
# ]
len(known_face_encodings)
#len(biden_face_encoding)
known_face_encodings
#len(biden_face_encoding)
# for name in known_face_names:
# known_face_encodings=np.append(known_face_encodings, known_face_encoding[i])
#
#
#
# # Load a second sample picture and learn how to recognize it.
# biden_image = face_recognition.load_image_file("known/Saksham Madan.jpg")
# biden_face_encoding = face_recognition.face_encodings(biden_image)[0]
# """
#
# known_face_encodings = [
#
# biden_face_encoding
# ]
# """
len(known_face_encodings)
# +
process_this_frame = True
video_capture = cv2.VideoCapture(0)
while True:
# Grab a single frame of video
ret, frame = video_capture.read()
# Resize frame of video to 1/4 size for faster face recognition processing
small_frame = cv2.resize(frame, (0, 0), fx=0.25, fy=0.25)
# Convert the image from BGR color (which OpenCV uses) to RGB color (which face_recognition uses)
rgb_small_frame = small_frame[:, :, ::-1]
# Only process every other frame of video to save time
if process_this_frame:
# Find all the faces and face encodings in the current frame of video
face_locations = face_recognition.face_locations(rgb_small_frame)
face_encodings = face_recognition.face_encodings(rgb_small_frame, face_locations)
face_names = []
for face_encoding in face_encodings:
# See if the face is a match for the known face(s)
matches = face_recognition.compare_faces(known_face_encodings, face_encoding)
name = "Unknown"
# If a match was found in known_face_encodings, just use the first one.
if True in matches:
first_match_index = matches.index(True)
name = known_face_names[first_match_index]
face_names.append(name)
process_this_frame = not process_this_frame
# Display the results
for (top, right, bottom, left), name in zip(face_locations, face_names):
# Scale back up face locations since the frame we detected in was scaled to 1/4 size
top *= 4
right *= 4
bottom *= 4
left *= 4
# Draw a box around the face
cv2.rectangle(frame, (left, top), (right, bottom), (0, 0, 255), 2)
# Draw a label with a name below the face
cv2.rectangle(frame, (left, bottom - 35), (right, bottom), (0, 0, 255), cv2.FILLED)
font = cv2.FONT_HERSHEY_DUPLEX
cv2.putText(frame, name, (left + 6, bottom - 6), font, 1.0, (255, 255, 255), 1)
# Display the resulting image
cv2.imshow('Video', frame)
# Hit 'q' on the keyboard to quit!
if cv2.waitKey(1) & 0xFF == ord('q'):
break
# Release handle to the webcam
video_capture.release()
cv2.destroyAllWindows()
# -
biden_face_encoding
len(biden_face_encoding)
len(known_face_encodings)
known_face_encodings
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Start by importing the functions
# +
from pathlib import Path
from xnemogcm import open_domain_cfg, open_nemo, open_namelist, open_nemo_and_domain_cfg
from xnemogcm import __version__ as xnemogcm_version
# -
xnemogcm_version
# # First open the domain_cfg and nemo files into 2 datasets
# ## domain
# +
# open_domain_cfg?
# -
# ---
#
# You can provide the file names / folder using 3 similar methods:
# 1. Give the path to the files and xnemogcm opens the domain_cfg_out and mesh_mesk files
# 2. Give the path to the data folder + the name of the files
# 3. Give the name of the files that already contain the tree (e.g. ['/path/to/file1', '/path/to/file2']
#
# These 3 methods are equivalent, however if your domain files don't have the strandard names you need to provide them by hand.
#
# We use one of the test folder:
datadir = Path('../xnemogcm/test/data/open_and_merge/')
# !ls ../xnemogcm/test/data/open_and_merge/
domcfg = open_domain_cfg(datadir=datadir)
# or
domcfg = open_domain_cfg(datadir=datadir, files=['domain_cfg_out.nc'])
# or
domcfg = open_domain_cfg(files=datadir.glob('*domain*.nc'))
domcfg
# ## Nemo
# +
# open_nemo?
# -
# ---
# We can provide the files folder / name following the same convention as for the `open_domain_cfg` function. We also need to provide the `domcfg` dataset so xnemogcm knows how to set the variables on the proper grid position. We can also provide extra kwargs to the underlying call to `xarray.open_mfdataset` function.
nemo = open_nemo(domcfg=domcfg, datadir=datadir)
# or
nemo = open_nemo(domcfg=domcfg, files=datadir.glob('*grid*.nc'))
nemo
# # Open both at once
#
# It is possible to open the domain and nemo output at once in one unique dataset. What happens is that 2 datasets are created and then merged. Thus all option possible for the `open_nemo` and `open_domain_cfg` functions are still possible.
# +
# open_nemo_and_domain_cfg?
# -
# ---
# Again, mutiple equivalent arguments are possible to open the data
# the simplest for simple cases, provide the path
ds = open_nemo_and_domain_cfg(nemo_files=datadir, domcfg_files=datadir)
# or provide the files
ds = open_nemo_and_domain_cfg(nemo_files=datadir.glob('*grid*.nc'), domcfg_files=datadir.glob('*domain*.nc'))
# or use the nemo_kwargs and domcfg_kwargs dictionnaries
ds = open_nemo_and_domain_cfg(nemo_kwargs=dict(datadir=datadir), domcfg_kwargs={'files':datadir.glob('*domain*.nc')})
ds
# ## Remark
#
# All opening are lazy using dask, which makes files quick to open, until you actually load the data you need
# # Namelist
#
# It can be convenient to open the namelist used for the run (e.g. to compare different runs with different parameters). This is possible using the `f90nml` package (it needs to be installed, this is an optional dependency).
# +
# open_namelist?
# -
# ---
# Here you provide the folder path containing the reference and configuration namelists, or the filenames (as for nemo and domcfg). You can choose to load both, or only one of them. The configration namelist will overwrite the default one.
#
# For this we need to use another folder of the test data (with simplified namelists for the exemple):
datadir = Path('../xnemogcm/test/data/namelist/')
# !ls ../xnemogcm/test/data/namelist/
nam = open_namelist(datadir)
nam
nam.nn_it000
| exemple/open_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# # Programming language interoperability
# ## Python
using PyCall
np = pyimport("numpy")
np.linalg.eigvals(np.random.rand(5,5))
M = rand(5,5)
np.linalg.eigvals(M)
py"""
import numpy as np
def sinpi(x):
return np.sin(np.pi * x)
"""
py_sinpi(x) = py"sinpi"(x)
py_sinpi(10)
using BenchmarkTools
@btime py_sinpi(10);
# ## C
c_code = """
#include <stddef.h>
double c_sum(size_t n, double *X) {
double s = 0.0;
for (size_t i = 0; i < n; ++i) {
s += X[i];
}
return s;
}
""";
# Compile to a shared library by piping `c_code` to gcc:
# +
using Libdl
const Clib = tempname()
open(`gcc -fPIC -O3 -msse3 -xc -shared -o $(Clib * "." * Libdl.dlext) -`, "w") do f
print(f, c_code)
end
# -
# Binding the function from the shared library:
c_sum(X::Array{Float64}) = @ccall Clib.c_sum(length(X)::Csize_t, X::Ptr{Float64})::Float64
c_sum(rand(10))
x = rand(10)
@btime c_sum($x);
# ## Mixing Julia, Python, and C
# Julia (`real`), Python/numpy (`py_sinpi`), C (`c_sum`)
x = rand(10);
py_sinpi(real(c_sum(x)))
@btime py_sinpi(real(c_sum($x)));
# See [JuliaInterop](https://github.com/JuliaInterop) for more, such as [RCall.jl](https://github.com/JuliaInterop/RCall.jl), [JavaCall.jl](https://github.com/JuliaInterop/JavaCall.jl), and [MATLAB.jl](https://github.com/JuliaInterop/MATLAB.jl).
| Day2/1_interop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [serenata_de_amor]
# language: python
# name: Python [serenata_de_amor]
# ---
# ### TSE data example
# This is just a small example of how to use tse data.
# In this script we filter the candidates in order to obtain a dataframe containing only politicians that were elected at least once.
import pandas as pd
import numpy as np
import os
DATASET_PATH= os.path.join(os.pardir,'data','2017-05-10-tse-candidates.xz')
# Loading csv
cand_df=pd.read_csv(DATASET_PATH,encoding='utf-8',dtype='category',)# setting dtype to category instead of str cuts by more than a half RAM usage
cand_df.columns
### Here, we quickly check the data using value_counts() to get the frequency on each column
for name, col in cand_df.iteritems():
print ('\t',name,'\n')
print (col.value_counts())
# ### Only elected politicians
# Now, we process candidacies data to obtain a list of elected politicians
# We use 'result' to figure out who has been elected. It is better to use the description column then the code column, since the codes dont seem to be consistent along the years.
#
ind_elected= (cand_df.result=='elected') | (cand_df.result=='elected_by_party_quota')
# ind_elected|=cand_df.result=='alternate'# should we consider it?
ind_elected= cand_df.index[ind_elected]
politicians_df=cand_df.loc[ind_elected,['cpf','name','post','location','state','year']]
politicians_df.sort_values('name')
# It is quite curious that alphabetically ordered in this way, the last politician is called...
# If we want to keep only the list of politicians we keep only cpf and name and remove the duplicates
politicians_df[['cpf','name']].drop_duplicates().sort_values('name')
| research/develop/2017-05-10-rafonseca-use-tse-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exmple
# .ipynb
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import datasets
from sklearn.preprocessing import scale
# -
# ## load test data
# +
ds = datasets.load_diabetes()
data_X = ds.data
data_y = ds.target
num_samples, num_features = data_X.shape
print(data_X.shape, data_y.shape)
## standardize data
X, y = scale(data_X, axis=0), scale(data_y)
## resahpe data into 1D
X_resh = X.reshape((num_samples*num_features,))
# -
# ### A 'standard' reference with scikit-learn
from sklearn.linear_model import ElasticNet
enet = ElasticNet(alpha=1.0, l1_ratio=0.5)
enet.fit(X, y)
enet.coef_
from td3a_cpp.linreg.all_methods import regularizedLinearRegression
# ### naive python method
demo_naive_py = regularizedLinearRegression(alpha=1.0, L1_ratio=0.5, method='nav')
demo_naive_py.fit(X, y)
# ## C++ wrapped method
demo_cpp_py = regularizedLinearRegression(alpha=1.0, L1_ratio=0.5, method='cpp')
demo_cpp_py.fit(X, y)
# ## multi-processing method
# which is not fast due to the core of the coordinate descent
demo_mp_py = regularizedLinearRegression(alpha=1.0, L1_ratio=0.5, method='mp')
demo_mp_py.fit(X, y)
| .ipynb_checkpoints/report-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# +
# configure
means = [1, 4, 1, 4]
N = 30
f1_name = 'pobudzenie'
f2_name = 'walencja'
dv_name = '<NAME>'
labels_factor1 = ['niski', 'wysoki']
labels_factor2 = ['pozytywny', 'negatywny']
fname = 'dane4.csv'
res = [np.random.normal(m, 1, N) for m in means]
dv = np.concatenate(res)
f1 = np.repeat(labels_factor1, 2*N)
f2 = np.tile(np.repeat(labels_factor2, N), 2)
df = pd.DataFrame({
f1_name: f1,
f2_name: f2,
dv_name: dv
})
df.to_csv(fname)
# -
len(dv)
| plany_zlozone/make_datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="xtaLPWSdhSQX"
#
# Audio Classifier Tutorial (Fast.ai)
# =========================
# **Author**: `<NAME> <https://github.com/limazix>`
#
# This is the recreation of the PyTorch tutorial for audio classification using the Fastai framework.
# - [torchaudio](https://github.com/pytorch/audio)
# - [fastai](https://docs.fast.ai/)
#
#
# -
# First, let’s import the common packages ``pandas`` and ``numpy``.
# + colab={} colab_type="code" id="z8KGhRRshSQR"
# %matplotlib inline
import os
import pandas as pd
import numpy as np
# + [markdown] colab_type="text" id="nF73jgL2hSQk"
# Importing the Dataset
# ---------------------
#
# We will use the UrbanSound8K dataset to train our network. It is
# available for free `here <https://urbansounddataset.weebly.com/>`_ and contains
# 10 audio classes with over 8000 audio samples! Once you have downloaded
# the compressed dataset, extract it to your current working directory.
# First, we will look at the csv file that provides information about the
# individual sound files. ``pandas`` allows us to open the csv file and
# use ``.iloc()`` to access the data within it.
#
#
#
# -
# After the download at the data folder, let's map the data paths into global variables.
DATA_ROOT_DIR=os.path.normpath(os.path.join(os.getcwd(), 'data/UrbanSound8k'))
DATA_META_FILE=os.path.join(DATA_ROOT_DIR, 'metadata/UrbanSound8k.csv')
DATA_AUDIO_DIR=os.path.join(DATA_ROOT_DIR, 'audio')
# + colab={} colab_type="code" id="8btdLhPdhSQl"
data_meta = pd.read_csv(DATA_META_FILE)
print(data_meta.iloc[0, :])
# + [markdown] colab_type="text" id="7miODyWlhSQp"
# The 10 audio classes in the UrbanSound8K dataset are: `air_conditioner`, `car_horn`, `children_playing`, `dog_bark`, `drilling`, `enginge_idling`, `gun_shot`, `jackhammer`, `siren`, and `street_music`.
#
# Each class has its own folder and id. For instance, the last print shows that the audio file has:
# - **class** - dog_bark
# - **id** - 3
# - **folder** - 5
#
# We will need the classID and the full file path only.
# +
def build_file_path(item):
item['path'] = 'fold{}/{}'.format(item['fold'], item['slice_file_name'])
item['label'] = item['class']
return item[['path', 'label']]
data = data_meta.apply(build_file_path, axis=1)
data.info()
# -
# ---
# ## Fast.ai Data Structure
#
# In order to run DNN with Fast.ai, it's necessary to create a DataBunch, the basic framework data structure.
#
# The Fast.ai has a few built-in DataBanch, such as ImageDataBunch for image processing and TextDataBunch for NLP, but none for audio processing yet. Therefore, we'll have to create a custom one.
#
# The simplest way to do that is using the ItemList and ItemBase classes. The first one creates a structure that allows operations to manipulate the dataset, while the second one defines the operation bahavior for one single item.
# +
import torchaudio
from fastai.data_block import ItemList, ItemBase
class AudioItem(ItemBase):
def apply_tfms(self, tfms):
if not tfms: return self
for t in tfms:
self.data = t(self.data)
return self
class AudioItemList(ItemList):
def get(self, i):
fn = super().get(i)
file_path = os.path.join(self.path, fn)
return self.open(file_path)
def open(self, fn):
audio, sample_rate = torchaudio.load(fn)
return AudioItem(audio[0, :].view(1, -1))
# -
# To keep it simple, we create two methods for the custom ItemList:
# - **get** - Method to retrieve one single item from the dataset given its position;
# - **open** - Method used to load the audio file and build the AudioItem object.
#
# And one for the custom ItemBase:
# - **apply_ftms** - Method used to apply transformations in each data from the dataset.
# ### Data Transformation
#
# Like any machine learning system, it's necessary to perform a few preprocessing steps on the data in order to run the intent algorithm. The original tutorial perform three transformations:
# - Resample - It perform a downsample from 44.1KHz to 8KHz
# - MaxClips - It uses a fixed sample size of 160000 clips by completting with zeros or remove when necessary
# - ClipFrequency - It group every 5th clip to the final sample and ignore de rest
#
# For the first transformation, the `torchaudio` library already have an implementation. But not for the last two, so we need to create then.
# +
import torch
class MaxClips(object):
def __init__(self, max_clips):
self.max_clips = max_clips
def __call__(self, sound_base):
sound_base = sound_base.long()
#tempData accounts for audio clips that are too short
sound = torch.zeros([self.max_clips, 1])
if sound_base.numel() < self.max_clips:
sound[:sound_base.numel()] = sound_base.view(-1, 1)[:]
else:
sound[:] = sound_base.view(-1, 1)[:self.max_clips]
return sound
class ClipFrequency(object):
def __init__(self, size, frequency):
self.frequency = frequency
self.size = size
def __call__(self, sound_base):
sound = torch.zeros([self.size, 1])
#take every fifth sample of soundData
sound[:self.size] = sound_base[::self.frequency]
sound = sound.permute(1, 0)
return sound
# -
# Now, we can create the transformation pipelie.
# +
from torchaudio.transforms import Resample
transforms = [
Resample(orig_freq=44100, new_freq=8000),
MaxClips(max_clips=160000),
ClipFrequency(size=32000, frequency=5)
]
print(transforms)
# -
# ### DataBunch Creation
#
# We already have the minimun necessary to create the DataBunch. Here, we perform five operations:
# - **from_df** - It transform the DataFrame with two columns into an AudioItemList instance;
# - **split_subsets** - It splits the dataset in two, train and validation dataset, based on the given proportions;
# - **label_from_df** - It informs the ItemList to use the column `label` from the dataset as the item class;
# - **transform** - It applys the transformation pipeline to bouth datasets, train and validation;
# - **databunch** - It creates the databunch with a batch size of 128 inputs per batch
data_bunch = (AudioItemList.from_df(data, path=DATA_AUDIO_DIR)
.split_subsets(train_size=0.8, valid_size=0.2)
.label_from_df(cols='label')
.transform((transforms, transforms))
.databunch(bs=128))
# ---
# ## Model
#
# The tutorial built de model class after the M5 CNN architecture for audio processing and classification as presented by [<NAME> Et al.](https://arxiv.org/pdf/1610.00087.pdf)
# +
from torch import nn
from torch.functional import F
class M5Model(nn.Module):
def __init__(self, n_classes):
super(M5Model, self).__init__()
self.conv1 = nn.Conv1d(1, 128, 80, 4)
self.bn1 = nn.BatchNorm1d(128)
self.pool1 = nn.MaxPool1d(4)
self.conv2 = nn.Conv1d(128, 128, 3)
self.bn2 = nn.BatchNorm1d(128)
self.pool2 = nn.MaxPool1d(4)
self.conv3 = nn.Conv1d(128, 256, 3)
self.bn3 = nn.BatchNorm1d(256)
self.pool3 = nn.MaxPool1d(4)
self.conv4 = nn.Conv1d(256, 512, 3)
self.bn4 = nn.BatchNorm1d(512)
self.pool4 = nn.MaxPool1d(4)
#input should be 512x30 so this outputs a 512x1
self.avgPool = nn.AvgPool1d(30)
self.fc1 = nn.Linear(512, n_classes)
self.softmax = nn.LogSoftmax(dim=2)
def forward(self, x):
x = self.conv1(x)
x = F.relu(self.bn1(x))
x = self.pool1(x)
x = self.conv2(x)
x = F.relu(self.bn2(x))
x = self.pool2(x)
x = self.conv3(x)
x = F.relu(self.bn3(x))
x = self.pool3(x)
x = self.conv4(x)
x = F.relu(self.bn4(x))
x = self.pool4(x)
x = self.avgPool(x)
#change the 512x1 to 1x512
x = x.permute(0, 2, 1)
x = self.fc1(x)
return self.softmax(x)
# -
# ## Wrap-up
#
# After defined the `databunch` and the `model`, it is time to create the main responsible by the larning operations, the `Learner`.
#
# Usually, with `pytorch`, that's the moment to define the `optimizer`, the `learning rate`(lr), the `loss function` and methods for training and testing for each epoch. All those things are define manually.
#
# With time, it becomes clear that not everything has to be custom made for each experiment scenario. The [Learner](https://docs.fast.ai/basic_train.html#Learner) class provide all those setups already built-in and it uses the the concept of partial to allow a few enhancements.
#
# Tho original tutorial uses the `Adan` optimizer with lr `0.1`. The `Learner` use the same optimizer by default but, if experiment request a different one, it can be changed by using the `opt_func` parament from the constructor. Likewise, the `loss function` with the parameter `loss_func`.
# +
from fastai.callbacks import EarlyStoppingCallback, ReduceLROnPlateauCallback
from fastai.metrics import accuracy, partial, error_rate
from fastai.basic_data import DataBunch
from fastai.basic_train import Learner
model = M5Model(n_classes=data_bunch.c)
learn = Learner(data_bunch, model, metrics=[error_rate, accuracy],
callback_fns=[
partial(EarlyStoppingCallback, monitor='accuracy', patience=10, min_delta=5e-4),
partial(ReduceLROnPlateauCallback, monitor='accuracy', patience=5, factor=0.2, min_delta=0)
])
# -
# It's important to notice that the above setup is using `error_rate` and `accuracy` as metrics to evaluate the performace of the model while it performs the training and validation phases.
#
# In adition, there are two partials:
# - **EarlyStoppingCallback** - It stops the trainig if a given metric, in this case `accuracy`, does not increases `5e-4` in `10` epochs
# - **ReduceLROnPlateauCallback** - It changes the lr by `0.2` times if the `accuracy` does not increases in `5` epochs
#
# Those two partials avoid overtrainig and helps to scape from a possible plateau.
#
# To run the training and the validation phases, the `Learner` has a method called `fit`. In the follow example, it receives two parameters: number of epochs and lr.
learn.fit(40, 0.01)
# ## Final Considerations
#
# It shows how simplier and cleaner is to wright using fast.ai even when no previous module exists, such as `vision` or `text`.
#
# Now, it is possible to dig a little deeper on audio processing universe using fast.ai. For the next article, it'll try to improve the accuracy of the model used here by playing with the input data in the preprocessing phase and with the parameters in the training phase.
#
# The notebook used in this tutorial is available [here](https://github.com/limazix/audio-processing).
| audio_classifier_fastai.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a name="oov-words"></a>
# # Out of vocabulary words (OOV)
# <a name="vocabulary"></a>
# ### Vocabulary
# In the video about the out of vocabulary words, you saw that the first step in dealing with the unknown words is to decide which words belong to the vocabulary.
#
# In the code assignment, you will try the method based on minimum frequency - all words appearing in the training set with frequency >= minimum frequency are added to the vocabulary.
#
# Here is a code for the other method, where the target size of the vocabulary is known in advance and the vocabulary is filled with words based on their frequency in the training set.
# +
# build the vocabulary from M most frequent words
# use Counter object from the collections library to find M most common words
from collections import Counter
# the target size of the vocabulary
M = 3
# pre-calculated word counts
# Counter could be used to build this dictionary from the source corpus
word_counts = {'happy': 5, 'because': 3, 'i': 2, 'am': 2, 'learning': 3, '.': 1}
vocabulary = Counter(word_counts).most_common(M)
# remove the frequencies and leave just the words
vocabulary = [w[0] for w in vocabulary]
print(f"the new vocabulary containing {M} most frequent words: {vocabulary}\n")
# -
# Now that the vocabulary is ready, you can use it to replace the OOV words with $<UNK>$ as you saw in the lecture.
# +
# test if words in the input sentences are in the vocabulary, if OOV, print <UNK>
sentence = ['am', 'i', 'learning']
output_sentence = []
print(f"input sentence: {sentence}")
for w in sentence:
# test if word w is in vocabulary
if w in vocabulary:
output_sentence.append(w)
else:
output_sentence.append('<UNK>')
print(f"output sentence: {output_sentence}")
# -
# When building the vocabulary in the code assignment, you will need to know how to iterate through the word counts dictionary.
#
# Here is an example of a similar task showing how to go through all the word counts and print out only the words with the frequency equal to f.
# +
# iterate through all word counts and print words with given frequency f
f = 3
word_counts = {'happy': 5, 'because': 3, 'i': 2, 'am': 2, 'learning':3, '.': 1}
for word, freq in word_counts.items():
if freq == f:
print(word)
# -
# As mentioned in the videos, if there are many $<UNK>$ replacements in your train and test set, you may get a very low perplexity even though the model itself wouldn't be very helpful.
#
# Here is a sample code showing this unwanted effect.
# +
# many <unk> low perplexity
training_set = ['i', 'am', 'happy', 'because','i', 'am', 'learning', '.']
training_set_unk = ['i', 'am', '<UNK>', '<UNK>','i', 'am', '<UNK>', '<UNK>']
test_set = ['i', 'am', 'learning']
test_set_unk = ['i', 'am', '<UNK>']
M = len(test_set)
probability = 1
probability_unk = 1
# pre-calculated probabilities
bigram_probabilities = {('i', 'am'): 1.0, ('am', 'happy'): 0.5, ('happy', 'because'): 1.0, ('because', 'i'): 1.0, ('am', 'learning'): 0.5, ('learning', '.'): 1.0}
bigram_probabilities_unk = {('i', 'am'): 1.0, ('am', '<UNK>'): 1.0, ('<UNK>', '<UNK>'): 0.5, ('<UNK>', 'i'): 0.25}
# got through the test set and calculate its bigram probability
for i in range(len(test_set) - 2 + 1):
bigram = tuple(test_set[i: i + 2])
probability = probability * bigram_probabilities[bigram]
bigram_unk = tuple(test_set_unk[i: i + 2])
probability_unk = probability_unk * bigram_probabilities_unk[bigram_unk]
# calculate perplexity for both original test set and test set with <UNK>
perplexity = probability ** (-1 / M)
perplexity_unk = probability_unk ** (-1 / M)
print(f"perplexity for the training set: {perplexity}")
print(f"perplexity for the training set with <UNK>: {perplexity_unk}")
# -
# <a name="smoothing"></a>
# ### Smoothing
# Add-k smoothing was described as a method for smoothing of the probabilities for previously unseen n-grams.
#
# Here is an example code that shows how to implement add-k smoothing but also highlights a disadvantage of this method. The downside is that n-grams not previously seen in the training dataset get too high probability.
#
# In the code output bellow you'll see that a phrase that is in the training set gets the same probability as an unknown phrase.
# +
def add_k_smooting_probability(k, vocabulary_size, n_gram_count, n_gram_prefix_count):
numerator = n_gram_count + k
denominator = n_gram_prefix_count + k * vocabulary_size
return numerator / denominator
trigram_probabilities = {('i', 'am', 'happy') : 2}
bigram_probabilities = {( 'am', 'happy') : 10}
vocabulary_size = 5
k = 1
probability_known_trigram = add_k_smooting_probability(k, vocabulary_size, trigram_probabilities[('i', 'am', 'happy')],
bigram_probabilities[( 'am', 'happy')])
probability_unknown_trigram = add_k_smooting_probability(k, vocabulary_size, 0, 0)
print(f"probability_known_trigram: {probability_known_trigram}")
print(f"probability_unknown_trigram: {probability_unknown_trigram}")
# -
# <a name="backoff"></a>
# ### Back-off
# Back-off is a model generalization method that leverages information from lower order n-grams in case information about the high order n-grams is missing. For example, if the probability of an trigram is missing, use bigram information and so on.
#
# Here you can see an example of a simple back-off technique.
# +
# pre-calculated probabilities of all types of n-grams
trigram_probabilities = {('i', 'am', 'happy'): 0}
bigram_probabilities = {( 'am', 'happy'): 0.3}
unigram_probabilities = {'happy': 0.4}
# this is the input trigram we need to estimate
trigram = ('are', 'you', 'happy')
# find the last bigram and unigram of the input
bigram = trigram[1: 3]
unigram = trigram[2]
print(f"besides the trigram {trigram} we also use bigram {bigram} and unigram ({unigram})\n")
# 0.4 is used as an example, experimentally found for web-scale corpuses when using the "stupid" back-off
lambda_factor = 0.4
probability_hat_trigram = 0
# search for first non-zero probability starting with trigram
# to generalize this for any order of n-gram hierarchy,
# you could loop through the probability dictionaries instead of if/else cascade
if trigram not in trigram_probabilities or trigram_probabilities[trigram] == 0:
print(f"probability for trigram {trigram} not found")
if bigram not in bigram_probabilities or bigram_probabilities[bigram] == 0:
print(f"probability for bigram {bigram} not found")
if unigram in unigram_probabilities:
print(f"probability for unigram {unigram} found\n")
probability_hat_trigram = lambda_factor * lambda_factor * unigram_probabilities[unigram]
else:
probability_hat_trigram = 0
else:
probability_hat_trigram = lambda_factor * bigram_probabilities[bigram]
else:
probability_hat_trigram = trigram_probabilities[trigram]
print(f"probability for trigram {trigram} estimated as {probability_hat_trigram}")
# -
# <a name="interpolation"></a>
# ### Interpolation
# The other method for using probabilities of lower order n-grams is the interpolation. In this case, you use weighted probabilities of n-grams of all orders every time, not just when high order information is missing.
#
# For example, you always combine trigram, bigram and unigram probability. You can see how this in the following code snippet.
# +
# pre-calculated probabilities of all types of n-grams
trigram_probabilities = {('i', 'am', 'happy'): 0.15}
bigram_probabilities = {( 'am', 'happy'): 0.3}
unigram_probabilities = {'happy': 0.4}
# the weights come from optimization on a validation set
lambda_1 = 0.8
lambda_2 = 0.15
lambda_3 = 0.05
# this is the input trigram we need to estimate
trigram = ('i', 'am', 'happy')
# find the last bigram and unigram of the input
bigram = trigram[1: 3]
unigram = trigram[2]
print(f"besides the trigram {trigram} we also use bigram {bigram} and unigram ({unigram})\n")
# in the production code, you would need to check if the probability n-gram dictionary contains the n-gram
probability_hat_trigram = lambda_1 * trigram_probabilities[trigram]
+ lambda_2 * bigram_probabilities[bigram]
+ lambda_3 * unigram_probabilities[unigram]
print(f"estimated probability of the input trigram {trigram} is {probability_hat_trigram}")
# -
# That's it for week 3, you should be ready now for the code assignment.
| Natural Language Processing/Course 2 - Natural Language Processing with Probabilistic Models/Labs/Week 3/Out of vocabulary words (OOV).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# Zeroing out gradients in PyTorch
# ================================
# It is beneficial to zero out gradients when building a neural network.
# This is because by default, gradients are accumulated in buffers (i.e,
# not overwritten) whenever ``.backward()`` is called.
#
# Introduction
# ------------
# When training your neural network, models are able to increase their
# accuracy through gradient decent. In short, gradient descent is the
# process of minimizing our loss (or error) by tweaking the weights and
# biases in our model.
#
# ``torch.Tensor`` is the central class of PyTorch. When you create a
# tensor, if you set its attribute ``.requires_grad`` as ``True``, the
# package tracks all operations on it. This happens on subsequent backward
# passes. The gradient for this tensor will be accumulated into ``.grad``
# attribute. The accumulation (or sum) of all the gradients is calculated
# when .backward() is called on the loss tensor.
#
# There are cases where it may be necessary to zero-out the gradients of a
# tensor. For example: when you start your training loop, you should zero
# out the gradients so that you can perform this tracking correctly.
# In this recipe, we will learn how to zero out gradients using the
# PyTorch library. We will demonstrate how to do this by training a neural
# network on the ``CIFAR10`` dataset built into PyTorch.
#
# Setup
# -----
# Since we will be training data in this recipe, if you are in a runable
# notebook, it is best to switch the runtime to GPU or TPU.
# Before we begin, we need to install ``torch`` and ``torchvision`` if
# they aren’t already available.
#
# ::
#
# pip install torchvision
#
#
#
#
# Steps
# -----
#
# Steps 1 through 4 set up our data and neural network for training. The
# process of zeroing out the gradients happens in step 5. If you already
# have your data and neural network built, skip to 5.
#
# 1. Import all necessary libraries for loading our data
# 2. Load and normalize the dataset
# 3. Build the neural network
# 4. Define the loss function
# 5. Zero the gradients while training the network
#
# 1. Import necessary libraries for loading our data
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# For this recipe, we will just be using ``torch`` and ``torchvision`` to
# access the dataset.
#
#
#
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torchvision
import torchvision.transforms as transforms
# -
# 2. Load and normalize the dataset
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# PyTorch features various built-in datasets (see the Loading Data recipe
# for more information).
#
#
#
# +
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=4,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=4,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# -
# 3. Build the neural network
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# We will use a convolutional neural network. To learn more see the
# Defining a Neural Network recipe.
#
#
#
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
# 4. Define a Loss function and optimizer
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# Let’s use a Classification Cross-Entropy loss and SGD with momentum.
#
#
#
net = Net()
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
# 5. Zero the gradients while training the network
# ~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# This is when things start to get interesting. We simply have to loop
# over our data iterator, and feed the inputs to the network and optimize.
#
# Notice that for each entity of data, we zero out the gradients. This is
# to ensure that we aren’t tracking any unnecessary information when we
# train our neural network.
#
#
#
# +
for epoch in range(2): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
# -
# You can also use ``model.zero_grad()``. This is the same as using
# ``optimizer.zero_grad()`` as long as all your model parameters are in
# that optimizer. Use your best judgement to decide which one to use.
#
# Congratulations! You have successfully zeroed out gradients PyTorch.
#
# Learn More
# ----------
#
# Take a look at these other recipes to continue your learning:
#
# - `Loading data in PyTorch <https://pytorch.org/tutorials/recipes/recipes/loading_data_recipe.html>`__
# - `Saving and loading models across devices in PyTorch <https://pytorch.org/tutorials/recipes/recipes/save_load_across_devices.html>`__
#
#
| PyTorch Recipes/Part 2/zeroing_out_gradients.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bravoeight98/FFN_498R/blob/main/498r_7k.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="o1yt6LTmyvK6"
# # Imporing Packages and Libraries
# + [markdown] id="rjrHm517yz7H"
# Importing all necessary packages
# + id="Yi3hM-qb6LR-" colab={"base_uri": "https://localhost:8080/"} outputId="f881d713-7db5-413f-e6c2-ca842f31ffc9"
#import package
# !pip install ipython-autotime
# %load_ext autotime
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.stem.porter import PorterStemmer
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer, HashingVectorizer
import re
import string
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn import metrics
from sklearn.feature_extraction.text import TfidfVectorizer
# + [markdown] id="wEsIoybTlpY3"
# # Getting and preparing Datasets
# + [markdown] id="ZZRQJMvLiMAe"
# Mount from Google Drive
# + colab={"base_uri": "https://localhost:8080/"} id="NOaBzjJC8Typ" outputId="25f34a4a-afc5-4c4a-8d16-5a41043d81cd"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="eJFaCWHG9lHJ" outputId="94a0ca64-977a-4a5e-b8c8-fac795926005"
# Reading from file
fake = pd.read_csv('/content/drive/MyDrive/BanFakeNewsDataset/Fake-1K.csv')
true = pd.read_csv('/content/drive/MyDrive/BanFakeNewsDataset/LabeledAuthentic-7K.csv')
# + [markdown] id="MBop7t4flK_3"
# Show Info
# + colab={"base_uri": "https://localhost:8080/", "height": 667} id="j80ejHDH9oQN" outputId="a5812dcb-d4af-4661-b8f6-82cb4d9ef441"
print(true.shape)
print(true.info())
true.head()
# + [markdown] id="rL_AuHkzlvdk"
# # Preprocessing Data
# + colab={"base_uri": "https://localhost:8080/"} id="QSorWdYI9vH2" outputId="4ad210e6-2e01-4617-bac0-2d1a3786262d"
fake['Label'] = 1
true['Label'] = 0
# + colab={"base_uri": "https://localhost:8080/", "height": 372} id="FeFS5Jmc-Das" outputId="f15af86f-5e52-475d-b77b-84d176b0728f"
data = pd.concat([true,fake],axis=0,ignore_index=True)
print(data.shape)
data.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 318} id="KOT5JhMX-Fhk" outputId="327d2450-e07c-417d-92d0-13890bc6c2f3"
data.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="6YcJG8J1-Jva" outputId="86bd8c69-3cf5-424a-fac1-aec3c3458935"
data['content']=data['headline']+data['content']
data=data.drop(['headline'], axis=1)
# + [markdown] id="6B2DFdjYs_n-"
# Chek if dataset is Balanced
# + colab={"base_uri": "https://localhost:8080/", "height": 368} id="oZJT7Bcp-Kaq" outputId="7c2c2b32-7382-4028-e72a-66c18259703c"
sns.countplot(data.Label)
# + [markdown] id="8ceK6nIOtE2G"
# Conclusion: Imbalanced Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="jfR7-5pf-QJx" outputId="08435bac-578a-4e3c-9ac4-2cd7ed1c2d40"
data.category.value_counts()
# + [markdown] id="fambCLALooHz"
# # Data Cleaning
# + [markdown] id="zTHknxdNoqu5"
# Removing stop words in order to root words that are retaining in the data and fed into the models
# + id="OPjR0yuHb_xk" colab={"base_uri": "https://localhost:8080/"} outputId="e6214d72-ae56-43ab-c22f-c6e4de2237d9"
#stop = stopwords.words('bangla')
stop = frozenset(["অবশ্য", "অনেক", "অনেকে", "অনেকেই", "অন্তত", "অথবা", "অথচ",
"অর্থাত",
"অন্য",
"আজ",
"আছে",
"আপনার",
"আপনি",
"আবার",
"আমরা",
"আমাকে",
"আমাদের",
"আমার",
"আমি",
"আরও",
"আর",
"আগে",
"আগেই",
"আই",
"অতএব",
"আগামী",
"অবধি",
"অনুযায়ী",
"আদ্যভাগে",
"এই",
"একই",
"একে",
"একটি",
"এখন",
"এখনও",
"এখানে",
"এখানেই",
"এটি",
"এটা",
"এটাই",
"এতটাই",
"এবং",
"একবার",
"এবার",
"এদের",
"এঁদের",
"এমন",
"এমনকী",
"এল",
"এর",
"এরা",
"এঁরা",
"এস",
"এত",
"এতে",
"এসে",
"একে",
"এ",
"ঐ",
"ই",
"ইহা",
"ইত্যাদি",
"উনি",
"উপর",
"উপরে",
"উচিত",
"ও",
"ওই",
"ওর",
"ওরা",
"ওঁর",
"ওঁরা",
"ওকে",
"ওদের",
"ওঁদের"
"ওখানে",
"কত",
"কবে",
"করতে",
"কয়েক",
"কয়েকটি",
"করবে",
"করলেন",
"করার",
"কারও",
"করা",
"করি",
"করিয়ে",
"করার",
"করাই",
"করলে",
"করলেন",
"করিতে",
"করিয়া",
"করেছিলেন",
"করছে",
"করছেন",
"করেছেন",
"করেছে",
"করেন",
"করবেন",
"করায়",
"করে",
"করেই",
"কাছ",
"কাছে",
"কাজে",
"কারণ",
"কিছু",
"কিছুই",
"কিন্তু",
"কিংবা",
"কি",
"কী",
"কেউ",
"কেউই",
"কাউকে",
"কেন",
"কে",
"কোনও",
"কোনো",
"কোন",
"কখনও",
"ক্ষেত্রে",
"খুব",
"গুলি",
"গিয়ে",
"গিয়েছে",
"গেছে",
"গেল",
"গেলে",
"গোটা",
"চলে",
"ছাড়া",
"ছাড়াও",
"ছিলেন",
"ছিল",
"জন্য",
"জানা",
"ঠিক",
"তিনি",
"তিনঐ",
"তিনিও",
"তখন",
"তবে",
"তবু",
"তাঁদের",
"তাঁহারা",
"তাঁরা",
"তাঁর",
"তাঁকে",
"তাই",
"তেমন",
"তাকে",
"তাহা",
"তাহাতে",
"তাহার",
"তাদের",
"তারপর",
"তারা",
"তারৈ",
"তার",
"তাহলে",
"তিনি",
"তা",
"তাও",
"তাতে",
"তো",
"তত",
"তুমি",
"তোমার",
"তথা",
"থাকে",
"থাকা",
"থাকায়",
"থেকে",
"থেকেও",
"থাকবে",
"থাকেন",
"থাকবেন",
"থেকেই",
"দিকে",
"দিতে",
"দিয়ে",
"দিয়েছে",
"দিয়েছেন",
"দিলেন",
"দু",
"দুটি",
"দুটো",
"দেয়",
"দেওয়া",
"দেওয়ার",
"দেখা",
"দেখে",
"দেখতে",
"দ্বারা",
"ধরে",
"ধরা",
"নয়",
"নানা",
"না",
"নাকি",
"নাগাদ",
"নিতে",
"নিজে",
"নিজেই",
"নিজের",
"নিজেদের",
"নিয়ে",
"নেওয়া",
"নেওয়ার",
"নেই",
"নাই",
"পক্ষে",
"পর্যন্ত",
"পাওয়া",
"পারেন",
"পারি",
"পারে",
"পরে",
"পরেই",
"পরেও",
"পর",
"পেয়ে",
"প্রতি",
"প্রভৃতি",
"প্রায়",
"ফের",
"ফলে",
"ফিরে",
"ব্যবহার",
"বলতে",
"বললেন",
"বলেছেন",
"বলল",
"বলা",
"বলেন",
"বলে",
"বহু",
"বসে",
"বার",
"বা",
"বিনা",
"বরং",
"বদলে",
"বাদে",
"বার",
"বিশেষ",
"বিভিন্ন বিষয়টি",
"ব্যবহার",
"ব্যাপারে",
"ভাবে",
"ভাবেই",
"মধ্যে",
"মধ্যেই",
"মধ্যেও",
"মধ্যভাগে",
"মাধ্যমে",
"মাত্র",
"মতো",
"মতোই",
"মোটেই",
"যখন",
"যদি",
"যদিও",
"যাবে",
"যায়",
"যাকে",
"যাওয়া",
"যাওয়ার",
"যত",
"যতটা",
"যা",
"যার",
"যারা",
"যাঁর",
"যাঁরা",
"যাদের",
"যান",
"যাচ্ছে",
"যেতে",
"যাতে",
"যেন",
"যেমন",
"যেখানে",
"যিনি",
"যে",
"রেখে",
"রাখা",
"রয়েছে",
"রকম",
"শুধু",
"সঙ্গে",
"সঙ্গেও",
"সমস্ত",
"সব",
"সবার",
"সহ",
"সুতরাং",
"সহিত",
"সেই",
"সেটা",
"সেটি",
"সেটাই",
"সেটাও",
"সম্প্রতি",
"সেখান",
"সেখানে",
"সে",
"স্পষ্ট",
"স্বয়ং",
"হইতে",
"হইবে",
"হৈলে",
"হইয়া",
"হচ্ছে",
"হত",
"হতে",
"হতেই",
"হবে",
"হবেন",
"হয়েছিল",
"হয়েছে",
"হয়েছেন",
"হয়ে",
"হয়নি",
"হয়",
"হয়েই",
"হয়তো",
"হল",
"হলে",
"হলেই",
"হলেও",
"হলো",
"হিসাবে",
"হওয়া",
"হওয়ার",
"হওয়ায়",
"হন",
"হোক",
"জন",
"জনকে",
"জনের",
"জানতে",
"জানায়",
"জানিয়ে",
"জানানো",
"জানিয়েছে",
"জন্য", "জন্যওজে", "জে", "বেশ", "দেন", "তুলে", "ছিলেন", "চান", "চায়", "চেয়ে", "মোট", "যথেষ্ট", "টি"])
data['content'] = data['content'].apply(lambda x: ' '.join([word for word in x.split() if word not in (stop)]))
# + [markdown] id="ezZDHJ-0fbc1"
# #Bulding Model
# + [markdown] id="06ReTE9Z6pLv"
# Spliting datas into training set and test set
# + id="yU7d4N1gff4b" colab={"base_uri": "https://localhost:8080/"} outputId="f087d5c7-15c8-45aa-836c-e9cd51eeb49a"
y = data['Label']
X_train, X_test, y_train, y_test = train_test_split(data['content'], y,test_size=0.33,random_state=53)
# + id="s-iIarNYfm7B" colab={"base_uri": "https://localhost:8080/"} outputId="1dc2bda6-23a6-41c5-9876-00a191fd6fc2"
count_vectorizer = CountVectorizer(stop_words = frozenset(["অবশ্য", "অনেক", "অনেকে", "অনেকেই", "অন্তত", "অথবা", "অথচ",
"অর্থাত",
"অন্য",
"আজ",
"আছে",
"আপনার",
"আপনি",
"আবার",
"আমরা",
"আমাকে",
"আমাদের",
"আমার",
"আমি",
"আরও",
"আর",
"আগে",
"আগেই",
"আই",
"অতএব",
"আগামী",
"অবধি",
"অনুযায়ী",
"আদ্যভাগে",
"এই",
"একই",
"একে",
"একটি",
"এখন",
"এখনও",
"এখানে",
"এখানেই",
"এটি",
"এটা",
"এটাই",
"এতটাই",
"এবং",
"একবার",
"এবার",
"এদের",
"এঁদের",
"এমন",
"এমনকী",
"এল",
"এর",
"এরা",
"এঁরা",
"এস",
"এত",
"এতে",
"এসে",
"একে",
"এ",
"ঐ",
"ই",
"ইহা",
"ইত্যাদি",
"উনি",
"উপর",
"উপরে",
"উচিত",
"ও",
"ওই",
"ওর",
"ওরা",
"ওঁর",
"ওঁরা",
"ওকে",
"ওদের",
"ওঁদের"
"ওখানে",
"কত",
"কবে",
"করতে",
"কয়েক",
"কয়েকটি",
"করবে",
"করলেন",
"করার",
"কারও",
"করা",
"করি",
"করিয়ে",
"করার",
"করাই",
"করলে",
"করলেন",
"করিতে",
"করিয়া",
"করেছিলেন",
"করছে",
"করছেন",
"করেছেন",
"করেছে",
"করেন",
"করবেন",
"করায়",
"করে",
"করেই",
"কাছ",
"কাছে",
"কাজে",
"কারণ",
"কিছু",
"কিছুই",
"কিন্তু",
"কিংবা",
"কি",
"কী",
"কেউ",
"কেউই",
"কাউকে",
"কেন",
"কে",
"কোনও",
"কোনো",
"কোন",
"কখনও",
"ক্ষেত্রে",
"খুব",
"গুলি",
"গিয়ে",
"গিয়েছে",
"গেছে",
"গেল",
"গেলে",
"গোটা",
"চলে",
"ছাড়া",
"ছাড়াও",
"ছিলেন",
"ছিল",
"জন্য",
"জানা",
"ঠিক",
"তিনি",
"তিনঐ",
"তিনিও",
"তখন",
"তবে",
"তবু",
"তাঁদের",
"তাঁহারা",
"তাঁরা",
"তাঁর",
"তাঁকে",
"তাই",
"তেমন",
"তাকে",
"তাহা",
"তাহাতে",
"তাহার",
"তাদের",
"তারপর",
"তারা",
"তারৈ",
"তার",
"তাহলে",
"তিনি",
"তা",
"তাও",
"তাতে",
"তো",
"তত",
"তুমি",
"তোমার",
"তথা",
"থাকে",
"থাকা",
"থাকায়",
"থেকে",
"থেকেও",
"থাকবে",
"থাকেন",
"থাকবেন",
"থেকেই",
"দিকে",
"দিতে",
"দিয়ে",
"দিয়েছে",
"দিয়েছেন",
"দিলেন",
"দু",
"দুটি",
"দুটো",
"দেয়",
"দেওয়া",
"দেওয়ার",
"দেখা",
"দেখে",
"দেখতে",
"দ্বারা",
"ধরে",
"ধরা",
"নয়",
"নানা",
"না",
"নাকি",
"নাগাদ",
"নিতে",
"নিজে",
"নিজেই",
"নিজের",
"নিজেদের",
"নিয়ে",
"নেওয়া",
"নেওয়ার",
"নেই",
"নাই",
"পক্ষে",
"পর্যন্ত",
"পাওয়া",
"পারেন",
"পারি",
"পারে",
"পরে",
"পরেই",
"পরেও",
"পর",
"পেয়ে",
"প্রতি",
"প্রভৃতি",
"প্রায়",
"ফের",
"ফলে",
"ফিরে",
"ব্যবহার",
"বলতে",
"বললেন",
"বলেছেন",
"বলল",
"বলা",
"বলেন",
"বলে",
"বহু",
"বসে",
"বার",
"বা",
"বিনা",
"বরং",
"বদলে",
"বাদে",
"বার",
"বিশেষ",
"বিভিন্ন বিষয়টি",
"ব্যবহার",
"ব্যাপারে",
"ভাবে",
"ভাবেই",
"মধ্যে",
"মধ্যেই",
"মধ্যেও",
"মধ্যভাগে",
"মাধ্যমে",
"মাত্র",
"মতো",
"মতোই",
"মোটেই",
"যখন",
"যদি",
"যদিও",
"যাবে",
"যায়",
"যাকে",
"যাওয়া",
"যাওয়ার",
"যত",
"যতটা",
"যা",
"যার",
"যারা",
"যাঁর",
"যাঁরা",
"যাদের",
"যান",
"যাচ্ছে",
"যেতে",
"যাতে",
"যেন",
"যেমন",
"যেখানে",
"যিনি",
"যে",
"রেখে",
"রাখা",
"রয়েছে",
"রকম",
"শুধু",
"সঙ্গে",
"সঙ্গেও",
"সমস্ত",
"সব",
"সবার",
"সহ",
"সুতরাং",
"সহিত",
"সেই",
"সেটা",
"সেটি",
"সেটাই",
"সেটাও",
"সম্প্রতি",
"সেখান",
"সেখানে",
"সে",
"স্পষ্ট",
"স্বয়ং",
"হইতে",
"হইবে",
"হৈলে",
"হইয়া",
"হচ্ছে",
"হত",
"হতে",
"হতেই",
"হবে",
"হবেন",
"হয়েছিল",
"হয়েছে",
"হয়েছেন",
"হয়ে",
"হয়নি",
"হয়",
"হয়েই",
"হয়তো",
"হল",
"হলে",
"হলেই",
"হলেও",
"হলো",
"হিসাবে",
"হওয়া",
"হওয়ার",
"হওয়ায়",
"হন",
"হোক",
"জন",
"জনকে",
"জনের",
"জানতে",
"জানায়",
"জানিয়ে",
"জানানো",
"জানিয়েছে",
"জন্য", "জন্যওজে", "জে", "বেশ", "দেন", "তুলে", "ছিলেন", "চান", "চায়", "চেয়ে", "মোট", "যথেষ্ট", "টি"]))
count_train = count_vectorizer.fit_transform(X_train.values)
count_test = count_vectorizer.transform(X_test.values)
print(count_train.shape)
# + [markdown] id="citg6dki-5fI"
# #Trainig Model
# + [markdown] id="-zFUEVye--Dy"
# ##RNN
# + colab={"base_uri": "https://localhost:8080/"} id="mA7Wo_bgC35l" outputId="b382b855-2ef9-48d9-caa8-a30d7f49350f"
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import re
from tensorflow.keras.preprocessing.text import Tokenizer
import tensorflow as tf
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix, precision_score, recall_score
import seaborn as sns
plt.style.use('ggplot')
# + colab={"base_uri": "https://localhost:8080/"} id="nUBxBE35-9nL" outputId="f6ebd9e4-4dea-4ee4-9aa5-f7a072a6b169"
max_vocab = 10000
# + colab={"base_uri": "https://localhost:8080/"} id="TiMq02N9BkiZ" outputId="07c6859a-fd02-464d-a48b-8d303af7f1b2"
X_train = tf.keras.preprocessing.sequence.pad_sequences(X_train, padding='post', maxlen=256)
X_test = tf.keras.preprocessing.sequence.pad_sequences(X_test, padding='post', maxlen=256)
# + colab={"base_uri": "https://localhost:8080/"} id="fTYPp7FQBm-M" outputId="341d51b6-47d9-414d-9d82-3f5d08b1245c"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(max_vocab, 32),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64, return_sequences=True)),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(16)),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dropout(0.5),
tf.keras.layers.Dense(1)
])
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="5aLoBzW0HA-a" outputId="654037a5-daf3-4a6c-eee1-741044ff3aae"
from sklearn.metrics import classification_report
# + colab={"base_uri": "https://localhost:8080/"} id="eRH7YXIIBskT" outputId="2bf69389-cee9-4381-ddc7-315fa8f8fd91"
early_stop = tf.keras.callbacks.EarlyStopping(monitor='val_loss', patience=2, restore_best_weights=True)
model.compile(loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
optimizer=tf.keras.optimizers.Adam(1e-4),
metrics=['accuracy'])
history = model.fit(X_train, y_train, epochs=10,validation_split=0.1, batch_size=30, shuffle=True, callbacks=[early_stop])
pred = model.predict(X_test)
binary_predictions = []
for i in pred:
if i >= 0.5:
binary_predictions.append(1)
else:
binary_predictions.append(0)
print(classification_report(y_test, binary_predictions, target_names = ['Fake','True']))
# + colab={"base_uri": "https://localhost:8080/", "height": 319} id="c5_mrWkiBwaK" outputId="e4eb9da9-5a66-4185-9885-4937dbd97694"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, binary_predictions)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (RNN)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="bfoCFBj8CTB6" outputId="e9d652c9-1195-4ad2-d61f-0aff75670c05"
# compute ROC AUC
from sklearn.metrics import roc_auc_score
ROC_AUC = roc_auc_score(y_test, binary_predictions)
print('ROC AUC : {:.4f}'.format(ROC_AUC))
# + colab={"base_uri": "https://localhost:8080/"} id="K7htgPPUCb3b" outputId="e53e606c-3823-407b-8a47-1837db568db8"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, binary_predictions)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + colab={"base_uri": "https://localhost:8080/", "height": 303} id="Tp6GrQV7Cuwp" outputId="52d946ad-a991-4993-f890-06168b1f7575"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
# + [markdown] id="ZQoyPWfaTwNr"
# ##Naive Bayes
# + colab={"base_uri": "https://localhost:8080/"} id="FD7Gx372SvEW" outputId="4ba67922-7d61-4b3d-ea8a-8b0251c17735"
from sklearn.naive_bayes import MultinomialNB
from sklearn import metrics
classifier=MultinomialNB()
classifier.fit(X_train, y_train)
pred = classifier.predict(X_test)
score = metrics.accuracy_score(y_test, pred)
print(classification_report(y_test, pred, target_names = ['Fake','True']))
# + colab={"base_uri": "https://localhost:8080/", "height": 323} id="oyJ59n43UPad" outputId="a9aa0f74-a609-4493-f7c2-15f1cb16cf35"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, pred)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (RNN)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="Dy2QcLV_UZwx" outputId="a039e1df-4362-4ace-d487-439730a1f29a"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, pred)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + colab={"base_uri": "https://localhost:8080/", "height": 303} id="n6bsQQVTUtDO" outputId="67acca86-9f42-45e6-cfe9-bdd26a7cf7cf"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
# + [markdown] id="54wjDMUDfxAz"
# ##SVM (Linear)
# + id="-JdlSUEl8ACA" colab={"base_uri": "https://localhost:8080/"} outputId="398e384e-e014-4160-9049-ead6706640c1"
from sklearn.svm import SVC
from sklearn.metrics import classification_report
# + colab={"base_uri": "https://localhost:8080/"} id="bBPR_JGff1Ql" outputId="9c514a9e-80d0-4952-d735-15fa77cead7a"
svc_model1 = SVC(C=1, kernel='linear', gamma= 1)
svc_model1.fit(count_train, y_train)
prediction1 = svc_model1.predict(count_test)
print(classification_report(y_test, prediction1, target_names = ['Fake','True']))
# + id="VYvOnx5CprS_" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="019ed002-9e51-49f8-c3b3-b403518c12b4"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, prediction1)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (SVM Linear Kernel)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + id="Z2-HKvYGzCT1" colab={"base_uri": "https://localhost:8080/"} outputId="2fbcf002-6c3c-430d-85be-d0e47a9b0e6f"
# compute ROC AUC
from sklearn.metrics import roc_auc_score
ROC_AUC = roc_auc_score(y_test, prediction1)
print('ROC AUC : {:.4f}'.format(ROC_AUC))
# + id="7I_s-3Buq_3v" colab={"base_uri": "https://localhost:8080/"} outputId="df494b6c-dcfe-4451-f5dc-9914cdc5160a"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, prediction1)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + id="SQ52ep5JrBUa" colab={"base_uri": "https://localhost:8080/", "height": 304} outputId="ecbb6ae7-ac31-43ad-8019-b68cd17e5013"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
# + [markdown] id="5LTMd8HIVNW5"
# ##SVM (Polynomial)
# + id="6h_kadEH74jo" colab={"base_uri": "https://localhost:8080/"} outputId="8976d5a9-3dec-4b16-f99b-dfab5ba3b42e"
svc_model1 = SVC(C=1, kernel='poly', gamma= 1)
svc_model1.fit(count_train, y_train)
prediction2 = svc_model1.predict(count_test)
print(classification_report(y_test, prediction2, target_names = ['Fake','True']))
# + id="uaImSO8EzUVl" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="76e6ff75-1369-4200-9ff5-e6215118696c"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, prediction2)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (SVM Polynomial Kernel)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + id="B8f6rvVqza7b" colab={"base_uri": "https://localhost:8080/"} outputId="084270e4-e513-4621-e269-481b0afcb0e3"
# compute ROC AUC
from sklearn.metrics import roc_auc_score
ROC_AUC = roc_auc_score(y_test, prediction2)
print('ROC AUC : {:.4f}'.format(ROC_AUC))
# + id="9uI2SbKMzfzG" colab={"base_uri": "https://localhost:8080/"} outputId="4a9256c2-3fcb-4b34-d019-67a9ab3de657"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, prediction2)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + id="TLca3uDazmJM" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="aaffef43-1d7a-4586-e60f-eea0593bc144"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
# + [markdown] id="GIg9SzwhVhSr"
# ##SVM (RBF)
# + id="tQVoR5Lb3biv" colab={"base_uri": "https://localhost:8080/"} outputId="36ebc5b6-d49e-4423-d5ae-63cd8e5376b7"
svc_model1 = SVC(C=1, kernel='rbf', gamma= 1)
svc_model1.fit(count_train, y_train)
prediction3 = svc_model1.predict(count_test)
print(classification_report(y_test, prediction3, target_names = ['Fake','True']))
# + id="SIEDYjyx2zdi" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="6a4e1573-1f56-4381-ca21-fef9da5be4ac"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, prediction3)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (SVM RBF Kernel)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + id="Dd6PogmN2000" colab={"base_uri": "https://localhost:8080/"} outputId="84dc91a3-edec-4140-a449-625d5cdd3c2b"
# compute ROC AUC
from sklearn.metrics import roc_auc_score
ROC_AUC = roc_auc_score(y_test, prediction3)
print('ROC AUC : {:.4f}'.format(ROC_AUC))
# + id="TyClOgfwKW8f" colab={"base_uri": "https://localhost:8080/"} outputId="58a516e3-f775-47c1-fcfe-8ffac117cf95"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, prediction3)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + id="5BaNsV1I29OX" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="389ab323-0b9f-4428-a294-9fa7660f7c33"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
# + [markdown] id="Nl-_nd7QV1Ua"
# ##SVM (Sigmoid)
# + id="_thb5WsE3ca0" colab={"base_uri": "https://localhost:8080/"} outputId="7bc9478c-fa5c-4dc7-e594-41a1fcc065f6"
svc_model1 = SVC(C=1, kernel='sigmoid', gamma= 1)
svc_model1.fit(count_train, y_train)
prediction4 = svc_model1.predict(count_test)
print(classification_report(y_test, prediction4, target_names = ['Fake','True']))
# + id="GlYEV3lYJuRK" colab={"base_uri": "https://localhost:8080/", "height": 323} outputId="09a28147-f14b-4ba0-d8f4-7eb3e3372a77"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, prediction4)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (SVM Sigmoid Kernel)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + id="SR2tGEQtJwZA" colab={"base_uri": "https://localhost:8080/"} outputId="f1a57eb1-052e-4bd5-cf4e-85397f2c58bb"
# compute ROC AUC
from sklearn.metrics import roc_auc_score
ROC_AUC = roc_auc_score(y_test, prediction4)
print('ROC AUC : {:.4f}'.format(ROC_AUC))
# + id="1IxY_HcRKshR" colab={"base_uri": "https://localhost:8080/"} outputId="26d3e764-d667-4468-b27a-de0bcc990f91"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, prediction4)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + id="UIwkzNqSJy8T" colab={"base_uri": "https://localhost:8080/", "height": 303} outputId="3e1189e9-b596-4e8e-99ab-40e27af7e890"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
# + [markdown] id="jMmK3aYjV7ct"
# ##LSTM
# + colab={"base_uri": "https://localhost:8080/"} id="4wXFvhnWWH51" outputId="72a6450e-96b5-417c-a9e9-d766c7a5c9db"
# importing neural network libraries
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Embedding, GRU, LSTM, RNN, SpatialDropout1D
# + colab={"base_uri": "https://localhost:8080/"} id="K2LpizcKYQFa" outputId="46253702-7c6c-48d9-d39e-40961c29408b"
max_features = 100000
# + colab={"base_uri": "https://localhost:8080/"} id="ALR4ZPjVWgQ8" outputId="36abd4bd-0365-4c51-c371-a4262d9d9630"
# LSTM Neural Network
lstm_model = Sequential(name = 'lstm_nn_model')
lstm_model.add(layer = Embedding(input_dim = max_features, output_dim = 120, name = '1st_layer'))
lstm_model.add(layer = LSTM(units = 120, dropout = 0.2, recurrent_dropout = 0.2, name = '2nd_layer'))
lstm_model.add(layer = Dropout(rate = 0.5, name = '3rd_layer'))
lstm_model.add(layer = Dense(units = 120, activation = 'relu', name = '4th_layer'))
lstm_model.add(layer = Dropout(rate = 0.5, name = '5th_layer'))
lstm_model.add(layer = Dense(units = len(set(y)), activation = 'sigmoid', name = 'output_layer'))
# compiling the model
lstm_model.compile(optimizer = 'adam', loss = 'sparse_categorical_crossentropy', metrics = ['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="GygZjCZSYja6" outputId="d4fb9b55-c15d-4716-e4b1-4909260c4a8f"
lstm_model_fit = lstm_model.fit(X_train, y_train, epochs = 1)
# + colab={"base_uri": "https://localhost:8080/"} id="qkHV-mgPaUxb" outputId="e81162b3-2b6e-4d49-c806-a2d9bb71aee3"
#pred = lstm_model.predict(X_test)
#pred = np.round(pred).astype(int)
predict_x=lstm_model.predict(X_test)
classes_x=np.argmax(predict_x,axis=1)
binary_predictions = []
for i in classes_x:
if i >= 0.5:
binary_predictions.append(1)
else:
binary_predictions.append(0)
print(classification_report(y_test, binary_predictions, target_names = ['Fake','True']))
# + colab={"base_uri": "https://localhost:8080/", "height": 323} id="bRFLCRHBadvX" outputId="777089d7-3989-4e30-8a31-7a397ba47f9e"
# plot ROC Curve
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, classes_x)
plt.figure(figsize=(6,4))
plt.plot(fpr, tpr, linewidth=2)
plt.plot([0,1], [0,1], 'k--' )
plt.rcParams['font.size'] = 12
plt.title('ROC curve for Predicting Bangla Fake News (LSTM)')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="2iR_2gyxfPdX" outputId="49877640-3815-4ec8-8a17-cc49c16d7f04"
# Print the Confusion Matrix and slice it into four pieces
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, classes_x)
print('Confusion matrix\n\n', cm)
print('\nTrue Positives(TP) = ', cm[0,0])
print('\nTrue Negatives(TN) = ', cm[1,1])
print('\nFalse Positives(FP) = ', cm[0,1])
print('\nFalse Negatives(FN) = ', cm[1,0])
# + colab={"base_uri": "https://localhost:8080/", "height": 303} id="cyHFqH_WfpaJ" outputId="9282ee2a-a393-4e94-db54-04145b920786"
# visualize confusion matrix with seaborn heatmap
cm_matrix = pd.DataFrame(data=cm, columns=['Actual Positive:1', 'Actual Negative:0'],
index=['Predict Positive:1', 'Predict Negative:0'])
sns.heatmap(cm_matrix, annot=True, fmt='d', cmap='YlGnBu')
| 498r_7k.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data and Imports
# +
import pandas as pd
import numpy as np
import jsonlines
import seaborn as sns
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import torch.nn as nn
import torch
import torch.nn.functional as F
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch_optimizer as optim
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from importlib import reload
pd.set_option('display.max_rows', 500)
pd.set_option('display.float_format', '{:0.3f}'.format)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
pd.options.display.width = 0
import warnings
import torchvision
warnings.filterwarnings('ignore')
from facebook_hateful_memes_detector.utils.globals import set_global, get_global
set_global("cache_dir", "/home/ahemf/cache/cache")
set_global("dataloader_workers", 4)
set_global("use_autocast", True)
set_global("models_dir", "/home/ahemf/cache/")
from facebook_hateful_memes_detector.utils import read_json_lines_into_df, in_notebook, set_device
get_global("cache_dir")
from facebook_hateful_memes_detector.models import Fasttext1DCNNModel, MultiImageMultiTextAttentionEarlyFusionModel, LangFeaturesModel, AlbertClassifer
from facebook_hateful_memes_detector.preprocessing import TextImageDataset, my_collate, get_datasets, get_image2torchvision_transforms, TextAugment
from facebook_hateful_memes_detector.preprocessing import DefinedRotation, QuadrantCut, ImageAugment
from facebook_hateful_memes_detector.training import *
import facebook_hateful_memes_detector
reload(facebook_hateful_memes_detector)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
set_device(device)
# -
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
set_device(device)
device
aug_speeds = {"keyboard": 117, "char_substitute": 109, "char_insert": 109, "char_swap": 114,
"ocr": 114, "char_delete": 108,
"word_insert": 0.0, "word_substitute": 0.0, "text_rotate": 32,
"stopword_insert": 34, "word_join": 32, "word_cutout": 36,
"w2v_insert": 0.0, "w2v_substitute": 0.0,
"fasttext": 137, "glove_twitter": 88, "glove_wiki": 82, "word2vec": 137,
"synonym": 522, "split": 110, "sentence_shuffle": 67, "one_third_cut": 0.0, "half_cut":0.0}
# - What Augs are useful
# - What Text models perform best
# +
choice_probas = {"keyboard": 0.1, "char_substitute": 0.0, "char_insert": 0.1, "char_swap": 0.1, "ocr": 0.0, "char_delete": 0.1,
"fasttext": 0.0, "glove_twitter": 0.0, "glove_wiki": 0.0, "word2vec": 0.0, "split": 0.1,
"stopword_insert": 0.3, "word_join": 0.1, "word_cutout": 0.8,
"text_rotate": 0.5, "sentence_shuffle": 0.5, "one_third_cut": 0.3, "half_cut":0.1}
preprocess_text = TextAugment([0.05, 0.05, 0.05, 0.35, 0.3, 0.2], choice_probas, fasttext_file="wiki-news-300d-1M-subword.bin")
data = get_datasets(data_dir="../data/", train_text_transform=preprocess_text, train_image_transform=None,
test_text_transform=None, test_image_transform=None,
cache_images = True, use_images = False, dev=False, test_dev=True,
keep_original_text=False, keep_original_image=False,
keep_processed_image=True, keep_torchvision_image=False,)
# images = list(data["train"].img) + list(data["test"].img)
# pd.DataFrame({"img":images}).to_csv("image.csv", header=None, index=None)
# ImageAugment([0.2, 0.5, 0.3])
# +
# data["train"].label.value_counts()
# train = data["train"]
# ones = train[train["label"] == 1]
# zeros = train[train["label"] == 0].sample(n=len(ones), replace=False)
# data["train"] = pd.concat((ones, zeros)).sample(frac=1.0)
# data["train"].label.value_counts()
# len(set(data["train"]["id"])) == data["train"].shape[0]
# -
# https://discuss.pytorch.org/t/how-to-implement-torch-optim-lr-scheduler-cosineannealinglr/28797/11
# # Params
# +
sgd = torch.optim.SGD
sgd_params = dict(lr=2e-2, momentum=0.9, dampening=0, weight_decay=0, nesterov=False)
rangerQH = optim.RangerQH
rangerQHparams = dict(lr=1e-3, betas=(0.9, 0.999), nus=(.7, 1.0),
weight_decay=0.0,
k=6,
alpha=.5,
decouple_weight_decay=True,
eps=1e-8,)
adam = torch.optim.Adam
adam_params = params=dict(lr=1e-3, weight_decay=1e-7)
adamw = torch.optim.AdamW
adamw_params = dict(lr=0.001, betas=(0.9, 0.999), eps=1e-08, weight_decay=1e-2)
novograd = optim.NovoGrad
novograd_params = dict(lr= 1e-3,
betas=(0.9, 0.999),
eps=1e-8,
weight_decay=0,
grad_averaging=False,
amsgrad=False,)
qhadam = optim.QHAdam
qhadam_params = dict(lr= 1e-3,
betas=(0.9, 0.999),
nus=(1.0, 1.0),
weight_decay=0,
decouple_weight_decay=False,
eps=1e-8,)
radam = optim.RAdam
radam_params = dict(lr= 1e-3,
betas=(0.9, 0.999),
eps=1e-8,
weight_decay=0,)
yogi = optim.Yogi
yogi_params = dict(lr= 1e-2,
betas=(0.9, 0.999),
eps=1e-3,
initial_accumulator=1e-6,
weight_decay=0)
# +
batch_size=256
epochs = 10
optimizer = adam
optimizer_params = adam_params
scheduler_init_fn = get_multistep_lr([11, 13], gamma=0.25) # get_cosine_schedule_with_warmup # get_cosine_with_hard_restarts_schedule_with_warmup
scheduler_init_fn = get_cosine_schedule_with_warmup()
reg_sched = get_regularizer_scheduler()
# + [markdown] heading_collapsed=true
# # Fasttext 1D CNN
# + hidden=true
epochs = 15
model_fn = model_builder(Fasttext1DCNNModel,
dict(
classifier_dims=256,
num_classes=2,
n_layers=2,
final_layer_builder=fb_1d_loss_builder,
gaussian_noise=0.15,
dropout=0.2,
embedding_dims=256,
internal_dims=512,
fasttext_file="crawl-300d-2M-subword.bin",
featurizer="transformer",
loss="focal",
dice_loss_coef=0.0,
auc_loss_coef=0.0,
),
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
model_call_back=reg_sched,
scheduler_init_fn=scheduler_init_fn,
show_model_stats=False,
sampling_policy="without_replacement",
validation_epochs=[4, 7, 9, 11, 14, 17, 19])
r1, p1 = results, prfs
results
prfs
# 0.738 0.734
# 0.730 0.715
# 0.730 0.715
# 0.734 0.731
# 0.746 0.712
# + [markdown] heading_collapsed=true
# # Lang Features Model
# + hidden=true
epochs = 15
batch_size = 256
# fasttext_crawl 1.9s
# spacy 1.8s
# full_view 1.4s 0.659 0.651
# gensim 7.5s # 0.718 0.737
# nltk 3s 0.609 0.585
# "spacy", "key_phrases" 4.2s 0.688 0.670
# "fasttext_crawl", "spacy", "key_phrases", "gensim" 20s 0.763 0.729 2h 49m
# "fasttext_crawl", "gensim" 11s 0.749 0.733 1h 47m
# gensim 8s 0.751 0.733 1h 20m
all_caps = [
"fasttext_crawl", "spacy", "full_view", "key_phrases", "nltk", "gensim"
] # "snlp", "ibm_max", "tmoji", "key_phrases", "full_view", "spacy", "nltk", "fasttext_crawl"
all_caps = [
"full_view",
]
all_caps = ["fasttext_crawl", "spacy", "key_phrases", "nltk"]
all_caps = ["fasttext_crawl", "gensim"]
model_fn = model_builder(LangFeaturesModel,
dict(classifier_dims=256,
num_classes=2,
gaussian_noise=0.2,
dropout=0.2,
embedding_dims=256,
internal_dims=512,
capabilities=all_caps,
featurizer="transformer",
loss="focal",
dice_loss_coef=0.0,
auc_loss_coef=0.0,
n_layers=2,
final_layer_builder=fb_1d_loss_builder),
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched,
show_model_stats=False,
sampling_policy="without_replacement",
validation_epochs=[1, 4, 7, 9, 11, 14, 17, 19])
r1, p1 = results, prfs
results
prfs
# + hidden=true
# -
# # BERT Models
# + [markdown] heading_collapsed=true
# ## Actibus/Bert_REview
# + hidden=true
epochs = 24
batch_size = 256
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"9": {
"lr": optimizer_params["lr"] / 1e3,
"finetune": False
},
"10": {
"lr": optimizer_params["lr"] / 1e2,
"finetune": True
},
"11": {
"lr": optimizer_params["lr"] / 1e2,
"finetune": True
}
}
},
}
}
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
embedding_dims=768,
gaussian_noise=0.0,
dropout=0.1,
word_masking_proba=0.15,
internal_dims=512,
final_layer_builder=fb_1d_loss_builder,
n_layers=2,
n_encoders=2,
n_decoders=2,
n_tokens_in=96,
n_tokens_out=16,
featurizer="transformer",
model="activebus/BERT_Review",
loss="focal",
dice_loss_coef=0.0,
auc_loss_coef=0.0,
),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched,
validation_epochs=[4, 7, 9, 11, 14, 17, 19, 23, 27, 31, 34, 37, 41, 44, 47, 51, 54],
show_model_stats=False,
sampling_policy="without_replacement")
r2, p2 = results, prfs
results
prfs
# 0.761 0.749 (0.703 0.691)
# -
# ## NSP Style Finetuned
# ### Non DETR
# +
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"2": {
"lr": optimizer_params["lr"] / 1e3,
"finetune": False
},
"3": {
"lr": optimizer_params["lr"] / 1e3,
"finetune": False
},
"4": {
"lr": optimizer_params["lr"] / 1e2,
"finetune": True
},
"5": {
"lr": optimizer_params["lr"] / 1e2,
"finetune": True
}
}
},
}
}
epochs = 24
batch_size = 256
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
embedding_dims=768,
gaussian_noise=0.75,
dropout=0.1,
word_masking_proba=0.15,
internal_dims=512,
final_layer_builder=fb_1d_loss_builder,
n_layers=2,
n_encoders=3,
n_decoders=3,
n_tokens_in=96,
n_tokens_out=16,
featurizer="transformer",
model='./distilbert-nsp',
loss="focal",
classification_head="decoder_ensemble",
dice_loss_coef=0.0,
auc_loss_coef=0.0, # fasttext_vector_config=dict(n_decoders=2, gru_layers=2)
finetune=False),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched,
validation_epochs=[4, 7, 9, 11, 14, 17, 19, 23, 27, 31, 34, 37, 41, 44, 47, 51, 54],
show_model_stats=False,
sampling_policy="without_replacement")
r2, p2 = results, prfs
results
prfs
# Try with_replacement
# 0.810 0.661 (0.724 0.600) (gaussian_noise=0.75, dropout=0.1, word_masking_proba=0.2,)
# +
adamw = torch.optim.AdamW
adamw_params = dict(lr=1e-4, betas=(0.9, 0.98), eps=1e-08, weight_decay=1e-2)
optimizer = adamw
optimizer_params = adamw_params
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"2": {
"lr": optimizer_params["lr"] / 1e3,
"finetune": False
},
"3": {
"lr": optimizer_params["lr"],
"finetune": True
},
"4": {
"lr": optimizer_params["lr"],
"finetune": True
},
"5": {
"lr": optimizer_params["lr"],
"finetune": True
}
}
},
}
}
epochs = 24
batch_size = 256
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
gaussian_noise=0.2,
dropout=0.2,
word_masking_proba=0.25,
internal_dims=512,
final_layer_builder=fb_1d_loss_builder,
n_layers=2,
n_encoders=2,
n_decoders=2,
n_tokens_in=96,
n_tokens_out=16,
featurizer="transformer",
model='./distilbert-nsp',
loss="focal",
classification_head="decoder_ensemble", # head_ensemble
dice_loss_coef=0.0,
auc_loss_coef=0.5,
attention_drop_proba=0.2,
finetune=False),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched, # reg_sched
validation_epochs=[4, 7, 9, 11, 14, 17, 19, 23, 27, 31, 34, 37, 41, 44, 47, 51, 54],
show_model_stats=False,
sampling_policy="without_replacement")
r2, p2 = results, prfs
results
prfs
# auc_loss_coef=0.5
# 0.853 0.661 (0.757 0.596) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# 0.852 0.658 (0.748 0.604) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.2,
# 0.857 0.661 (0.761 0.590) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.845 0.657 (0.757 0.592) gaussian_noise=0.1, dropout=0.25, word_masking_proba=0.25,
# 0.841 0.644 (0.753 0.578) gaussian_noise=0.1, dropout=0.25, word_masking_proba=0.2,
# 0.861 0.647 (0.759 0.594) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.857 0.652 (0.756 0.576) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.848 0.661 (0.751 0.592) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.853 0.657 (0.755 0.588) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# 0.853 0.661 (0.750 0.602) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# 0.852 0.649 (0.757 0.578) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# No reg_sched testing
# 0.848 0.652 (0.754 0.590) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# Cnn1D head
# +
adamw = torch.optim.AdamW
adamw_params = dict(lr=1e-4, betas=(0.9, 0.98), eps=1e-08, weight_decay=1e-3)
optimizer = adamw
optimizer_params = adamw_params
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"2": {
"lr": optimizer_params["lr"] / 1e3,
"finetune": False
},
"3": {
"lr": optimizer_params["lr"],
"finetune": True
},
"4": {
"lr": optimizer_params["lr"],
"finetune": True
},
"5": {
"lr": optimizer_params["lr"],
"finetune": True
}
}
},
}
}
epochs = 24
batch_size = 256
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
gaussian_noise=0.1,
dropout=0.25,
word_masking_proba=0.25,
internal_dims=512,
final_layer_builder=fb_1d_loss_builder,
n_layers=2,
n_encoders=2,
n_decoders=2,
n_tokens_in=96,
n_tokens_out=16,
featurizer="transformer",
model='distilbert-cor',
loss="focal",
classification_head="decoder_ensemble", # decoder_ensemble
dice_loss_coef=0.0,
auc_loss_coef=0.5,
attention_drop_proba=0.2,
finetune=False),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched, # reg_sched
validation_epochs=[4, 7, 9, 11, 14, 17, 19, 23, 27, 31, 34, 37, 41, 44, 47, 51, 54],
show_model_stats=False,
sampling_policy="without_replacement")
r3, p3 = results, prfs
results
prfs
# auc_loss_coef=0.5
# 0.853 0.661 (0.757 0.596) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# 0.852 0.658 (0.748 0.604) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.2,
# 0.857 0.661 (0.761 0.590) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.845 0.657 (0.757 0.592) gaussian_noise=0.1, dropout=0.25, word_masking_proba=0.25,
# 0.841 0.644 (0.753 0.578) gaussian_noise=0.1, dropout=0.25, word_masking_proba=0.2,
# 0.861 0.647 (0.759 0.594) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.857 0.652 (0.756 0.576) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.848 0.661 (0.751 0.592) gaussian_noise=0.1, dropout=0.15, word_masking_proba=0.15,
# 0.853 0.657 (0.755 0.588) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# 0.853 0.661 (0.750 0.602) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# 0.852 0.649 (0.757 0.578) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# No reg_sched testing
# 0.848 0.652 (0.754 0.590) gaussian_noise=0.1, dropout=0.2, word_masking_proba=0.2,
# Cnn1D head
# -
# ### DETR Style
# +
from facebook_hateful_memes_detector.utils import in_notebook, CNNHead, MultiLayerTransformerDecoderHead, AveragedLinearHead, OneTokenPositionLinearHead, MultiTaskForward, CNN2DHead
def fb_detr_loss_builder(n_dims, n_tokens, n_out, dropout, **kwargs):
loss = kwargs.pop("loss", "classification")
cnn = MultiLayerTransformerDecoderHead(n_dims, n_tokens, n_out, dropout=0.4,
gaussian_noise=0.75, n_layers=3, loss=loss)
return cnn
# +
epochs = 24
batch_size = 256
adamw = torch.optim.AdamW
adamw_params = dict(lr=5e-5, betas=(0.9, 0.98), eps=1e-08, weight_decay=1e-2)
optimizer = adamw
optimizer_params = adamw_params
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"3": {
"lr": optimizer_params["lr"],
"finetune": True
},
"4": {
"lr": optimizer_params["lr"],
"finetune": True
},
"5": {
"lr": optimizer_params["lr"],
"finetune": True
}
}
},
}
}
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
gaussian_noise=0.15,
dropout=0.25,
word_masking_proba=0.25,
internal_dims=512,
final_layer_builder=fb_detr_loss_builder,
n_layers=2,
n_encoders=2,
n_decoders=0,
n_tokens_in=96,
n_tokens_out=96,
featurizer="transformer",
model='./distilbert-nsp',
classification_head="decoder_ensemble",
loss="focal",
dice_loss_coef=0.0,
auc_loss_coef=0.5,
attention_drop_proba=0.2,
finetune=False),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
kfold = False
results, prfs = train_validate_ntimes(
model_fn,
data,
batch_size,
epochs,
kfold=kfold,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched,
validation_epochs=[4, 7, 9, 11, 14, 17, 19, 23, 27, 31, 34, 37, 41, 44, 47, 51, 54],
show_model_stats=False,
sampling_policy="without_replacement")
r2, p2 = results, prfs
results
prfs
# 0.824 0.649 (0.748 0.576) (gaussian_noise=0.75, dropout=0.25, word_masking_proba=0.25,)
# 0.829 0.654 (0.744 0.584) (gaussian_noise=0.5, dropout=0.25, word_masking_proba=0.25,)
# 0.811 0.635 (0.740 0.578) (gaussian_noise=0.5, dropout=0.25, word_masking_proba=0.25,)
# 0.834 0.630 (0.755 0.566) (gaussian_noise=0.5, dropout=0.2, word_masking_proba=0.25,)
#
# + [markdown] heading_collapsed=true
# # Predict
# + [markdown] hidden=true
# ## Normal Head
# + hidden=true
adamw = torch.optim.AdamW
adamw_params = dict(lr=1e-4, betas=(0.9, 0.98), eps=1e-08, weight_decay=1e-2)
optimizer = adamw
optimizer_params = adamw_params
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"2": {
"lr": optimizer_params["lr"] / 1e3,
"finetune": False
},
"3": {
"lr": optimizer_params["lr"] / 1e1,
"finetune": True
},
"4": {
"lr": optimizer_params["lr"],
"finetune": True
},
"5": {
"lr": optimizer_params["lr"],
"finetune": True
}
}
},
}
}
epochs = 24
batch_size = 256
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
gaussian_noise=0.1,
dropout=0.2,
word_masking_proba=0.2,
internal_dims=512,
final_layer_builder=fb_1d_loss_builder,
n_layers=2,
n_encoders=3,
n_decoders=3,
n_tokens_in=96,
n_tokens_out=32,
featurizer="transformer",
model='./distilbert-nsp',
loss="focal",
classification_head="decoder_ensemble", # decoder_ensemble
dice_loss_coef=0.0,
auc_loss_coef=0.5,
finetune=False),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
# + hidden=true
submission, text_model = train_and_predict(model_fn, data, batch_size, epochs,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched,
sampling_policy="without_replacement",
validation_epochs=[15, 31, 34, 42],
)
submission.to_csv("submission.csv",index=False)
submission.sample(5)
# + [markdown] heading_collapsed=true hidden=true
# ## DETR Style head
# + hidden=true
lr_strategy = {
"model": {
"lr": optimizer_params["lr"] / 1000,
"finetune": False,
"encoder": {
"layer": {
"3": {
"lr": optimizer_params["lr"] / 1e2,
"finetune": True
},
"4": {
"lr": optimizer_params["lr"] / 1e2,
"finetune": True
},
"5": {
"lr": optimizer_params["lr"] / 1e1,
"finetune": True
}
}
},
}
}
model_fn = model_builder(AlbertClassifer,
dict(classifier_dims=256,
num_classes=2,
embedding_dims=768,
gaussian_noise=0.75,
dropout=0.25,
word_masking_proba=0.25,
internal_dims=512,
final_layer_builder=fb_detr_loss_builder,
n_layers=2,
n_encoders=2,
n_decoders=0,
n_tokens_in=96,
n_tokens_out=16,
featurizer="transformer",
model='./distilbert-nsp',
loss="focal",
dice_loss_coef=0.0,
auc_loss_coef=0.0,
finetune=False),
per_param_opts_fn=lr_strategy,
optimiser_class=optimizer,
optimiser_params=optimizer_params)
epochs = 24
batch_size = 256
submission, text_model = train_and_predict(model_fn, data, batch_size, epochs,
scheduler_init_fn=scheduler_init_fn,
model_call_back=reg_sched, validation_epochs=[2, 5, 7],
sampling_policy="without_replacement")
submission.to_csv("submission.csv",index=False)
submission.sample(10)
# 0.6723
# + hidden=true
# What train-auc does we achieve if all examples have same score.
# What train-auc does we achieve if all examples have random score.
# + hidden=true
submission.sample(10)
# + hidden=true
| notebooks/Fasttext1DCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Writing Photon-HDF5 files in python
#
# > This notebook shows how to create a [Photon-HDF5](http://photon-hdf5.org) file
# > from scratch using dummy data and [phconvert](http://photon-hdf5.github.io/phconvert/).
# >
# > For more info see [Writing Photon-HDF5 files](http://photon-hdf5.readthedocs.org/en/latest/writing.html).
import phconvert as phc
phc.__version__
# # 1. Create some dummy data
import numpy as np
timestamps = np.random.randint(low=0, high=2e8, size=10000).astype('int64')
timestamps.sort()
timestamps_unit = 10e-9 # 10 ns, units are always S.I.
detectors = np.random.randint(low=0, high=1, size=10000).astype('uint8')
# # 2. Create some metadata
# +
description = 'This is a fake dataset which mimics smFRET data.'
author = '<NAME>'
author_affiliation = 'Name of Research Institution'
sample_name = 'describe the sample here'
buffer_name = 'describe the buffer here'
dye_names = 'Cy3B, ATTO647N' # Comma separates names of fluorophores
# -
# # 3. Create Photon-HDF5 data structure
#
# In this section we create all the mandatory and non mandatory groups.
# Not all of the are required to save a valid Photon-HDF5 file
# (see example in section 4).
#
# ## 3.1 `photon_data` group
#
# Contains arrays of photon-data: timestamps, detectors, nanotimes, etc...
#
# *See [photon_data group reference](http://photon-hdf5.readthedocs.org/en/latest/phdata.html#photon-data-group)*
photon_data = dict(
timestamps=timestamps,
detectors=detectors,
timestamps_specs={'timestamps_unit': timestamps_unit})
# ## 3.2 `setup` group
#
# The `/setup` group contains information about the measurement setup.
#
# *See [setup group reference](http://photon-hdf5.readthedocs.org/en/latest/phdata.html#setup-group).*
setup = dict(
## Mandatory fields
num_pixels = 2, # using 2 detectors
num_spots = 1, # a single confoca excitation
num_spectral_ch = 2, # donor and acceptor detection
num_polarization_ch = 1, # no polarization selection
num_split_ch = 1, # no beam splitter
modulated_excitation = False, # CW excitation, no modulation
excitation_alternated = [False], # CW excitation, no modulation
lifetime = False, # no TCSPC in detection
## Optional fields
excitation_wavelengths = [532e-9], # List of excitation wavelenghts
excitation_cw = [True], # List of booleans, True if wavelength is CW
detection_wavelengths = [580e-9, 640e-9], # Nominal center wavelength
# each for detection ch
)
# ## 3.3 `provenance` group
#
# Non-mandatory group containing info about the original file
# prior to Photon-HDF5 conversion. If some information is not
# available the relative field may be omitted.
#
# *See [provenance group documentation](http://photon-hdf5.readthedocs.org/en/latest/phdata.html#provenance-group).*
provenance = dict(
filename='original_data_file.dat',
software='Acquisition Software Name')
# ## 3.4 `identity` group
#
# Non-mandatory group containing info about information
# this specific Photon-HDF5 file.
#
# *See [identity group documentation](http://photon-hdf5.readthedocs.org/en/latest/phdata.html#identity-group).*
identity = dict(
author=author,
author_affiliation=author_affiliation)
# ## 3.5 `measurement_specs` group
#
# The optional /photon_data/measurement_specs group contains
# additional information allowing unambiguous interpretation
# of the data for each specific type of measurement.
#
# *See [measurement_specs group documentation](http://photon-hdf5.readthedocs.org/en/latest/phdata.html#measurement-specs).*
measurement_specs = dict(
measurement_type = 'smFRET',
detectors_specs = {'spectral_ch1': [0], # list of donor's detector IDs
'spectral_ch2': [1]} # list of acceptor's detector IDs
)
# # 4. Save Photon-HDF5 files
#
# To save a file we need to join together the root fields and group
# in a single dictionary. Here we provide a few examples.
#
# ## 4.1 Minimal file
#
# Create a bare-bone Photon-HDF5 file with only mandatory fields.
data = dict(
description=description,
photon_data = photon_data,
setup=setup,
)
phc.hdf5.save_photon_hdf5(data, h5_fname='dummy_dataset_barebone.h5', overwrite=True)
# > **NOTE:** a user of this file can read the data but does not know
# > what kind of measurement it is (e.g. smFRET with single laser
# > excitation and 2-colors detection).
# ## 4.2 Expanded Photon-HDF5
#
# Create a Photon-HDF5 with non-mandatory fields (including `measurement_specs`):
# +
photon_data['measurement_specs'] = measurement_specs
data = dict(
description=description,
photon_data = photon_data,
setup=setup,
identity=identity,
provenance=provenance
)
# -
phc.hdf5.save_photon_hdf5(data, h5_fname='dummy_dataset_complete.h5', overwrite=True)
# > **NOTE:** a user of this file can correctly interpret the data
# > reading that the measurement type is 'smFRET' (meaning smFRET with single laser
# > excitation and 2-colors detection) and the IDs of donor and acceptor detectors
# > (from `detectors_specs/spectral_ch1` and `spectral_ch2` respectively).
| notebooks/Writing Photon-HDF5 files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Legend Place
#
# Place legend at `bottom`, `top`, `left` or `right`.
#
# See [theme()](https://jetbrains.github.io/lets-plot-docs/pages/api/lets_plot.theme.html#lets_plot.theme).
# +
import pandas as pd
from lets_plot import *
LetsPlot.setup_html()
# -
df = pd.read_csv('https://raw.githubusercontent.com/JetBrains/lets-plot-docs/master/data/mpg.csv')
ggplot(df, aes('class', 'hwy')) + \
geom_jitter(aes(color='fl'), width=.3, height=0) + \
theme(legend_position='top')
| source/examples/basics/gog/legend_place.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''venv'': venv)'
# name: python382jvsc74a57bd0d46d14bfc67a8d69f82f77af2e60e5b02fcabf25a759a56cf61072a2d590541d
# ---
import pandas as pd
log = pd.read_csv("/Users/alexigna/projects/skillfactory/unit1/00_log.csv", header = None)
users = pd.read_csv("/Users/alexigna/projects/skillfactory/unit1/00_users.csv", encoding="koi8-r", sep="\t")
sample = pd.read_csv("/Users/alexigna/projects/skillfactory/unit1/00_sample.csv")
sample.columns = map(str.lower, sample.columns)
sample.columns
log.columns = ["user_id","time","bet","win"]
log.head()
users
users.columns = ["user_id", "email", "geo"]
users
sample["name"].unique()
sample.info()
log.user_id.unique()
sample2 = sample[sample.age < 30]
sample2
log_win = log[log.win > 0]
win_count = log_win.win.count()
win_count
sample
sample[(sample.age<30) & (sample.profession=="Рабочий")]
sample.query("age>20")
sample.query('city in ["Рига", "Сочи","Чебоксары", "Сургут"] & 21<age<50 & profession!="Менеджер"')
log2 = log.query("bet<2000 & win>0")
log2
sample.name.str.match("К", na=False)
sample[sample.name.str.match("К", na=False)]
sample[~sample.name.str.match("К", na=True)]
sample3 = sample[sample.city.str.contains("о", na=False)]
sample3
sample4 = sample[~sample.city.str.contains("о", na=True)]
sample4
new_log = log[~log.user_id.str.contains("error", na=False)]
new_log.user_id.unique()
sample2 = sample.copy()
sample2["Age"] = sample2["Age"].apply(lambda x:x+1)
sample2
sample3 = sample.copy()
sample3["City"] = sample3["City"].apply(lambda s: str(s).lower())
sample3
# +
def age_category(age):
if age > 35:
return "зрелый"
elif age < 23:
return "молодой"
else:
return "средний"
sample['Age_category'] = sample.Age.apply(age_category)
# -
sample
# +
def user_format(username):
if username.startswith("Запись"):
return username.split(" ")[-1]
else:
return ""
log = pd.read_csv('00_log.csv', header=None)
log.columns = ['user_id','time','bet','win']
log["user_id"] = log["user_id"].apply(user_format)
log
# +
log = pd.read_csv("00_log.csv",header=None)
log.columns = ['user_id','time','bet','win']
def user_format(username):
if username.startswith("Запись"):
return username.split(" ")[-1]
else:
return ""
log = log[~log.user_id.str.contains("error", na=False)]
log["user_id"] = log["user_id"].apply(lambda s: s.split(" ")[-1] if s.startswith("Запись") else "")
log["time"] = log["time"].apply(lambda s: s[1:] if type(s) == str else s)
log
# -
lambda s: s[1:] if type(s) == str else s
| unit_1/python-8.1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
# ## For colab:
# +
# import tensorflow as tf
# tf.test.gpu_device_name()
# +
# from tensorflow.python.client import device_lib
# device_lib.list_local_devices()
# +
# # !nvidia-smi
# +
# from google.colab import drive
# drive.mount('/content/drive')
# +
# os.chdir(r'/content/drive/MyDrive/higgs boson/')
# -
# ## Display all rows:
# To display all columns:
pd.pandas.set_option('display.max_columns', None)
# ## Loding Data:
# +
# For colab:
# df1 = pd.read_csv('train_MICE_median_ind_QuantileTransformer.csv')
# df2 = pd.read_csv('train_MICE_med_ind_QT_dropOutlier.csv')
# For local
df1 = pd.read_csv('../data/interim/train_MICE_median_ind_QuantileTransformer.csv')
df2 = pd.read_csv('../data/interim/train_MICE_med_ind_QT_dropOutlier.csv')
# -
X, y = df1.drop(columns=['Weight', 'Label']), df1['Weight']
X2, y2 = df2.drop(columns=['Weight', 'Label']), df2['Weight']
print(df1.shape, df2.shape, sep='\n')
# ## Train-test split:
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=30000) # about 12%
X_train2, X_test2, y_train2, y_test2 = train_test_split(X2, y2, test_size=30000) # about 12%
# -
# # Feature Selection:
# ## Using Correlation:
plt.figure(figsize=(25, 20))
sns.heatmap(df1.corr(), annot=True, linewidth=0.2, cmap='coolwarm')
df1.corr()['Weight'].abs().sort_values(ascending=False)
# ## Univariate Selection:
# +
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import chi2
#apply SelectKBest class to extract top 10 best features
bestfeatures = SelectKBest(score_func=chi2, k=25)
# fit = bestfeatures.fit(X,y) # ValueError: Input X must be non-negative.
# -
# ## ExtraTreeRegressor:
# +
from sklearn.ensemble import ExtraTreesRegressor
model = ExtraTreesRegressor(n_estimators=100, max_depth=11, min_samples_split=50, n_jobs=-1)
model.fit(X_train, y_train)
# -
model.score(X_test, y_test) # Returns R2 value
from sklearn.metrics import mean_squared_error
pred = model.predict(X_test)
mean_squared_error(pred, y_test)
etr_imp = model.feature_importances_
fi_etr = pd.Series(etr_imp, index=X.columns)
fi_etr.sort_values(ascending=False)
# +
#Dropping the last 5 columns and checking:
drop_cols = ['PRI_jet_leading_phi',
'PRI_jet_subleading_phi',
'PRI_tau_phi',
'PRI_met_phi',
'PRI_lep_phi']
X_train_new = X_train.drop(columns=drop_cols)
X_test_new = X_test.drop(columns=drop_cols)
# -
model_new = ExtraTreesRegressor(n_estimators=151, max_depth=11, min_samples_split=50, n_jobs=-1)
model_new.fit(X_train_new, y_train)
model_new.score(X_test_new, y_test) # Returns R2 value
from sklearn.metrics import mean_squared_error
pred = model_new.predict(X_test_new)
mean_squared_error(pred, y_test)
#
| notebooks/05-Applying_Feature_Selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: kgtk-env-ckg07
# language: python
# name: kgtk-env-ckg07
# ---
import pandas as pd
import os
import sys
from kgtk.configure_kgtk_notebooks import ConfigureKGTK
from kgtk.functions import kgtk, kypher
from kgtk.io.kgtkreader import KgtkReader
from kgtk.io.kgtkwriter import KgtkWriter
from pathlib import Path
# + tags=["parameters"]
# Parameters
input_path = "/data/amandeep/wikidata-20220505-dwd-v4"
output_path = "/data/amandeep/wikidata-20220505-dwd-v4"
kgtk_path = "/data/amandeep/Github/kgtk"
graph_cache_path = None
project_name = "browser-claims-file"
files = 'claims'
debug=True
# -
files = files.split(',')
ck = ConfigureKGTK(files, kgtk_path=kgtk_path)
ck.configure_kgtk(input_graph_path=input_path,
output_path=output_path,
project_name=project_name,
graph_cache_path=graph_cache_path,
debug=debug)
ck.print_env_variables()
def separate_edges_and_qualifiers(input_file, output_edge_file, output_qualifier_file):
kr: KgtkReader = KgtkReader.open(Path(input_file),
error_file=sys.stderr)
ids = set()
for row in kr:
ids.add(row[kr.id_column_idx])
kr.close()
kr: KgtkReader = KgtkReader.open(Path(input_file),
error_file=sys.stderr)
kw_edge: KgtkWriter = KgtkWriter.open(file_path=Path(output_edge_file),
error_file=sys.stderr,
column_names=kr.column_names,
mode=KgtkWriter.Mode.EDGE,
no_header=False)
kw_qualifier: KgtkWriter = KgtkWriter.open(file_path=Path(output_qualifier_file),
error_file=sys.stderr,
column_names=kr.column_names,
mode=KgtkWriter.Mode.EDGE,
no_header=False)
for row in kr:
id = row[kr.id_column_idx]
node1 = row[kr.node1_column_idx]
if node1 in ids:
kw_qualifier.write(row)
else:
kw_edge.write(row)
kr.close()
kw_edge.close()
kw_qualifier.close()
# !curl https://raw.githubusercontent.com/usc-isi-i2/kgtk/dev/kgtk-properties/kgtk.properties.tsv -o $TEMP/kgtk.properties.tsv
kgtk("""filter -i $TEMP/kgtk.properties.tsv
-p ';label;'
-o $TEMP/kgtk.properties.labels.tsv
""")
kgtk("""filter -i $TEMP/kgtk.properties.tsv
-p ';alias;'
-o $TEMP/kgtk.properties.aliases.tsv
""")
kgtk("""filter -i $TEMP/kgtk.properties.tsv
-p ';description;'
-o $TEMP/kgtk.properties.descriptions.tsv
""")
kgtk("""filter -i $TEMP/kgtk.properties.tsv
-p ';datatype;'
-o $TEMP/kgtk.properties.datatypes.tsv
""")
kgtk("""filter -i $TEMP/kgtk.properties.tsv
-p ';label,alias,description,datatype;'
--invert
-o $TEMP/kgtk.properties.claims.tsv
""")
kgtk("""cat
-i $GRAPH/statistics.Pinstance_count.star.tsv.gz
-i $GRAPH/statistics.Pinstance_count.tsv.gz
-i $GRAPH/statistics.Psubclass_count.star.tsv.gz
-i $GRAPH/derived.class.P1963computed.count.tsv.gz
-i $GRAPH/derived.P1963computed.subclass.count.star.tsv.gz
-i $GRAPH/derived.Pproperty_domain.tsv.gz
-i $TEMP/kgtk.properties.claims.tsv
-i $GRAPH/derived.P1963computed.count.star.tsv.gz
-i $GRAPH/metadata.pagerank.undirected.tsv.gz
-o $TEMP/derived.claims.augmented.tsv.gz
""")
separate_edges_and_qualifiers(f"{os.environ['TEMP']}/derived.claims.augmented.tsv.gz",
f"{os.environ['TEMP']}/derived.claims.edges.tsv.gz",
f"{os.environ['TEMP']}/derived.claims.qualifiers.tsv.gz")
kgtk(f"""cat
-i $GRAPH/labels.en.tsv.gz
-i $TEMP/kgtk.properties.labels.tsv
/ sort
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/labels.en.tsv.gz
""")
kgtk(f"""cat
-i $GRAPH/aliases.en.tsv.gz
-i $TEMP/kgtk.properties.aliases.tsv
/ sort
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/aliases.en.tsv.gz
""")
kgtk(f"""cat
-i $GRAPH/descriptions.en.tsv.gz
-i $TEMP/kgtk.properties.descriptions.tsv
/ sort
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/descriptions.en.tsv.gz
""")
kgtk(f"""cat
-i $GRAPH/metadata.property.datatypes.tsv.gz
-i $TEMP/kgtk.properties.datatypes.tsv
/ sort
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/metadata.property.datatypes.tsv.gz
""")
kgtk(f"""cat
-i $TEMP/derived.claims.qualifiers.tsv.gz
-i $GRAPH/qualifiers.tsv.gz
/ sort
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/qualifiers.tsv.gz
""")
kgtk("""filter -i $GRAPH/sitelinks.tsv.gz
-o $TEMP/sitelinks.filtered.tsv.gz
-p ';wikipedia_sitelink;'
""")
kgtk(f"""cat
-i $TEMP/derived.claims.edges.tsv.gz
-i $GRAPH/claims.tsv.gz
-i $GRAPH/downloaded.wikipedia.short_abstracts.tsv.gz
-i $TEMP/sitelinks.filtered.tsv.gz
/ sort -c node1 label node2
--extra '--parallel 24 --buffer-size 30% --temporary-directory {os.environ['TEMP']}'
-o $OUT/claims.tsv.gz
""")
| use-cases/create_wikidata/Create-claims-augmented-for-browser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="8LDziy681qsA" outputId="2f1d5921-7a1a-4a16-e241-72997436efeb"
# !pip install opendatasets
import opendatasets as od
od.download("https://www.kaggle.com/eemanmajumder/the-anime-dataset")
# + colab={"base_uri": "https://localhost:8080/"} id="WihkJ9X9rrhW" outputId="3f283d75-766d-4697-a63f-a8bac02eccfc"
import pandas as pd
import random
import re
from tqdm import tqdm
import numpy as np
from spacy.tokenizer import Tokenizer
from spacy.lang.en import English
import torch
import spacy
nlp = English()
import torch.nn as nn
import nltk
pd.options.display.max_columns = 500
import warnings
warnings.filterwarnings(action='ignore')
nltk.download('punkt')
# + colab={"base_uri": "https://localhost:8080/"} id="jfnPAtuesCRf" outputId="10efc464-734b-4902-b02e-be0739e0d69e"
data = pd.read_csv('eda-data.csv',index_col=0)
synopsis = data.synopsis
print('Number of Anime synopsis we have: ',len(synopsis))
# + colab={"base_uri": "https://localhost:8080/"} id="DEiPGR7wsGfk" outputId="52299374-d8a5-4539-d318-d515b6cbc06a"
i = random.randint(0,len(synopsis))
print('Synopsis example\n\nAnime:{} \nSynopsis:{}\n'.format(data['anime_name'].values[i],synopsis.values[i]))
# + id="JhXJ0DRVsNzc"
def remove_source(text):
cln_text = text
if '(Source' in cln_text:
cln_text,_,_ = cln_text.partition('(Source')
elif '[Written ' in cln_text:
cln_text,_,_ = cln_text.partition('[Written')
return cln_text
# + colab={"base_uri": "https://localhost:8080/"} id="z9wZvvNpsR2Z" outputId="f3148222-538c-4473-ab9f-bb47f27d1944"
def clean_synopsis(data):
# removing hentai and kids tags
data = data[(data.Hentai != 1) & (data.Kids != 1)]
synopsis = data.synopsis
# removing very small synopsis
synopsis = synopsis.apply(lambda x: x if ((len(str(x).strip().split())<=300) and len(str(x).strip().split())>30 ) else -1)
synopsis = synopsis[synopsis!=-1]
# removing source text
synopsis = synopsis.apply(lambda x: remove_source(x))
# removing japanese characters
synopsis = synopsis.apply(lambda x: re.sub("([^\x00-\x7F])+"," ",x))
# remove symbols
rx = re.compile('^[&#/@`)(;<=\'"$%>]')
synopsis = synopsis.apply(lambda x: rx.sub('',x))
synopsis = synopsis.apply(lambda x: x.replace('>',""))
synopsis = synopsis.apply(lambda x: x.replace('`',""))
synopsis = synopsis.apply(lambda x: x.replace(')',""))
synopsis = synopsis.apply(lambda x: x.replace('(',""))
# removing adaptation animes (some relevant might get deleted but there aren`t a lot so we wont be affected as much)
synopsis = synopsis[synopsis.apply(lambda x: 'adaptation' not in str(x).lower())]
synopsis = synopsis[synopsis.apply(lambda x: 'music video' not in str(x).lower())]
synopsis = synopsis[synopsis.apply(lambda x: 'based on' not in str(x).lower())]
synopsis = synopsis[synopsis.apply(lambda x: 'spin-off' not in str(x).lower())]
return synopsis.reset_index(drop=True)
cleaned_synopsis = clean_synopsis(data)
print('Size: ',len(cleaned_synopsis))
# + id="Nq_0QFAusSqQ"
class config:
tokenizer = nltk.word_tokenize
#data = AnimeDataset(cleaned_synopsis)
batch_size = 32
#vocab_size = data.vocab_size
seq_len = 30
emb_dim = 100
epochs = 15
hidden_dim = 512
model_path = 'lm_lrdecay_drop.bin'
# + id="0E6IZEyosVlS"
def create_dataset(synopsis,batch_size,seq_len):
np.random.seed(0)
synopsis = synopsis.apply(lambda x: str(x).lower()).values
synopsis_text = ' '.join(synopsis)
tokens = config.tokenizer(synopsis_text)
global num_batches
num_batches = int(len(tokens)/(seq_len*batch_size))
tokens = tokens[:num_batches*batch_size*seq_len]
words = sorted(set(tokens))
w2i = {w:i for i,w in enumerate(words)}
i2w = {i:w for i,w in enumerate(words)}
tokens = [w2i[tok] for tok in tokens]
target = np.zeros_like((tokens))
target[:-1] = tokens[1:]
target[-1] = tokens[0]
input_tok = np.reshape(tokens,(batch_size,-1))
target_tok = np.reshape(target,(batch_size,-1))
print(input_tok.shape)
print(target_tok.shape)
vocab_size = len(i2w)
return input_tok,target_tok,vocab_size,w2i,i2w
def create_batches(input_tok,target_tok,batch_size,seq_len):
num_batches = np.prod(input_tok.shape)//(batch_size*seq_len)
for i in range(0,num_batches*seq_len,seq_len):
yield input_tok[:,i:i+seq_len], target_tok[:,i:i+seq_len]
# + id="sJAl1xn1saEw"
class LSTMModel(nn.Module):
def __init__(self,hid_dim,emb_dim,vocab_size,num_layers=1):
super(LSTMModel,self).__init__()
self.hid_dim = hid_dim
self.emb_dim = emb_dim
self.num_layers = num_layers
self.vocab_size = vocab_size+1
self.embedding = nn.Embedding(self.vocab_size,self.emb_dim)
self.lstm = nn.LSTM(self.emb_dim,self.hid_dim,batch_first = True,num_layers = self.num_layers)
self.drop = nn.Dropout(0.3)
self.linear = nn.Linear(self.hid_dim,vocab_size) # from here we will randomly sample a word
def forward(self,x,prev_hid):
x = self.embedding(x)
x,hid = self.lstm(x,prev_hid)
x = self.drop(x)
x = self.linear(x)
return x,hid
def zero_state(self,batch_size):
return (torch.zeros(self.num_layers,batch_size,self.hid_dim),torch.zeros(self.num_layers,batch_size,self.hid_dim))
# + id="-LWoTuK3sgoO"
class AverageMeter:
"""Computes and stores the average and current value"""
def __init__(self):
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
# + id="Orz9PpNysli9"
def loss_fn(predicted,target):
loss = nn.CrossEntropyLoss()
return loss(predicted,target)
# + id="nNzkJpfusp6-"
def train_fn(model,device,dataloader,optimizer):
model.train()
tk0 = tqdm(dataloader,position=0,leave=True,total = num_batches)
train_loss = AverageMeter()
hid_state,cell_state = model.zero_state(config.batch_size)
hid_state = hid_state.to(device)
cell_state = cell_state.to(device)
losses = []
for inp,target in tk0:
inp = torch.tensor(inp,dtype=torch.long).to(device)
target = torch.tensor(target,dtype=torch.long).to(device)
optimizer.zero_grad()
pred,(hid_state,cell_state) = model(inp,(hid_state,cell_state))
#print(pred.transpose(1,2).shape)
loss = loss_fn(pred.transpose(1,2),target)
hid_state = hid_state.detach()
cell_state = cell_state.detach()
loss.backward()
_ = torch.nn.utils.clip_grad_norm_(model.parameters(),max_norm=2) # to avoid gradient explosion
optimizer.step()
train_loss.update(loss.detach().item())
tk0.set_postfix(loss = train_loss.avg)
losses.append(loss.detach().item())
return np.mean(losses)
# + colab={"base_uri": "https://localhost:8080/"} id="J0n7rG71sqmK" outputId="49b9656c-61a5-4242-fa20-a23c9373d57c"
input_tok,target_tok,vocab_size,w2i,i2w = create_dataset(cleaned_synopsis,batch_size=config.batch_size,seq_len=config.seq_len)
# + id="OUT5t7Mgst9q"
def run():
device = 'cuda'
model = LSTMModel(vocab_size=vocab_size,emb_dim=config.emb_dim,hid_dim=config.hidden_dim,num_layers=3).to(device)
optimizer = torch.optim.Adam(model.parameters(),lr=0.001)
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(optimizer=optimizer, mode = 'min', patience=2, verbose=True, factor=0.5)
epochs = config.epochs
best_loss = 999
for i in range(1,epochs+1):
train_dataloader = create_batches(batch_size=config.batch_size,input_tok=input_tok,seq_len=config.seq_len,target_tok=target_tok)
print('Epoch..',i)
loss = train_fn(model,device,train_dataloader,optimizer)
if loss<best_loss:
best_loss = loss
torch.save(model.state_dict(),config.model_path)
scheduler.step(loss)
torch.cuda.empty_cache()
return model
# + colab={"base_uri": "https://localhost:8080/"} id="l-F3YAB-s18f" outputId="a42215b2-1b7c-46ca-909a-a637742e2e71"
model = run()
# + id="9w7kcU9As5az"
def inference(model,input_text,device,top_k=5,length = 100):
output = ''
model.eval()
tokens = config.tokenizer(input_text)
h,c = model.zero_state(1)
h = h.to(device)
c = c.to(device)
for t in tokens:
output = output+t+' '
pred,(h,c) = model(torch.tensor(w2i[t.lower()]).view(1,-1).to(device),(h,c))
#print(pred.shape)
for i in range(length):
_,top_ix = torch.topk(pred[0],k = top_k)
choices = top_ix[0].tolist()
choice = np.random.choice(choices)
out = i2w[choice]
output = output + out + ' '
pred,(h,c) = model(torch.tensor(choice,dtype=torch.long).view(1,-1).to(device),(h,c))
return output
# + colab={"base_uri": "https://localhost:8080/", "height": 175} id="gywEtnIIs6KK" outputId="7e06b063-87f1-45ac-cb20-a326a3c2a2d8"
device = 'cpu'
mod = LSTMModel(emb_dim=config.emb_dim,hid_dim=config.hidden_dim,vocab_size=vocab_size,num_layers=3).to(device)
mod.load_state_dict(torch.load(config.model_path))
print('AI generated Anime synopsis:')
inference(model = mod, input_text = 'Bob the Architect ', top_k = 30, length = 1000, device = device)
| Anime_Generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="vE9BuidfHmGC"
import numpy as np
import pandas as pd
import os
# 1.1 Call sklearn libraries
# 1.1.1 Split data into train and test data
from sklearn.model_selection import train_test_split
# 1.1.2 PReprocessong:
from sklearn.preprocessing import StandardScaler
# 1.1.3 Import class DecisionTreeClassifier
from sklearn.tree import DecisionTreeClassifier as dtree
# 1.1.4 To draw decision tree
from sklearn.tree import export_graphviz
# 1.1.5 Import iris
from sklearn.datasets import load_iris
# 1.1.6
# Install as:
# conda install -c conda-forge python-graphviz
# conda install -c anaconda pydot
import graphviz
# 1.1.7
from sklearn.tree import plot_tree
import matplotlib.pyplot as plt
# + id="95Ke5bQR3dDl"
import seaborn as sns
# + colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 74} id="1rQvimlEHnw1" outputId="f632e82b-ebf5-4767-f5c7-10d5e477fbaa"
from google.colab import files
uploaded = files.upload()
# + id="sjYDSn_LsZy-"
df = pd.read_csv("Real_fake_news.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="Oo49Kn5fzAWy" outputId="f21b0824-a3ab-4386-d823-eeb3f0256966"
df.head()
# + id="zJgjYr-W8waY"
# removing redundant spaces from column names
df.columns = list(map(lambda a: a.lstrip(), df.columns))
# + colab={"base_uri": "https://localhost:8080/"} id="UbuBPlo3zCIg" outputId="9cc986fb-087e-43b8-d03b-074531429c17"
df.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="Xdbr8VP9zMC5" outputId="a1dca28a-3db7-4b35-bb86-d4c31e3a4872"
df.shape
# + colab={"base_uri": "https://localhost:8080/"} id="n9SRAoSQ2Zcw" outputId="21c6b7da-6a19-4de2-9bdb-8a458326cd36"
df['author'].value_counts()
# + id="uiOa63a_Qbx0"
df = df.drop(columns=['Unnamed: 0','Unnamed: 0.1','target','BinaryTarget'])
# + colab={"base_uri": "https://localhost:8080/", "height": 403} id="Lt2kBjMxR-tP" outputId="55c35cb0-f84c-481f-e453-fb78d87a3268"
df_final = pd.get_dummies(df)
df_final.head()
# + id="ZCpaDeiCSMrg"
X = df_final.drop(columns=['BinaryNumTarget'])
y = df_final['BinaryNumTarget']
# + id="78Muxya-SWHr"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.30, random_state=42)
# + id="FzeOGjQTBMp2"
ct = dtree(
criterion="entropy", # Alternative 'entropy'
max_depth=None # Alternative, specify an integer
# 'None' means full tree till single leaf
)
# 5.1
_=ct.fit(X_train,y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="8LuvOP41Blzx" outputId="61372da6-96d5-48de-8a04-afbea1708af9"
y_te = ct.predict(X_test)
np.sum((y_test == y_te))/y_test.size
# + colab={"base_uri": "https://localhost:8080/"} id="qmjH5jpkBy-h" outputId="6fecd550-1111-43a4-b78e-34df2f1a0f1a"
fi = ct.feature_importances_
fi
# + id="2TZFxOZdCRhU"
from sklearn.ensemble import RandomForestClassifier # using random forest
# + id="LalcEYxACdbF"
#Create a Gaussian Classifier
clf=RandomForestClassifier(n_estimators=100)
#Train the model using the training sets y_pred=clf.predict(X_test)
clf.fit(X_train,y_train)
y_pred=clf.predict(X_test)
# + colab={"base_uri": "https://localhost:8080/"} id="SgZLATPvJGwR" outputId="d8ecd2e5-6321-4730-f213-d74cead46c07"
#Import scikit-learn metrics module for accuracy calculation
from sklearn import metrics
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# + [markdown] id="8iQIu8X7JKd8"
# When using decision tree , the accuracy was 88% , while using the Random Forest Classifier accuracy has increased to 90%
# + id="B2FWjzwC2EBp"
| Fake_News_or_Not_Using_random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
print(sys.version)
from astropy import units as u
from astropy.io import fits
import numpy as np
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
# -
rawdir = os.path.expanduser('~/SuPrimeCam/SuPrimeCam_S17A-UH16A/o16308')
# # Assembling an MEF
#
# Each file has one chip with 4 channels. Therefore we will generate a 40 extension MEF, one extension per channel. This allows for different properties in each channel (i.e. gain) to be accounted for.
def assemble_MEF(file0, write=False):
path, filename0 = os.path.split(file0)
files = ['{}{:d}.fits'.format(filename0[0:11], n) for n in range(0,10)]
rawhduls = [fits.open(os.path.join(rawdir, file), 'readonly') for file in files]
expid0 = rawhduls[0][0].header.get('EXP-ID')
# Confirm that EXP-IDs match indicating that these files are from the same exposure
for hdul in rawhduls:
expid = hdul[0].header.get('EXP-ID')
if not expid == expid0:
print(f'WARNING: EXP-ID mismatch!! {expid0}, {expid}')
# Split each file out in to channels and assemble a multi extension fits (MEF)
# file from the individual amplifiers (4 per chip)
MEF = fits.HDUList([])
extver = 0
for i,hdul in enumerate(rawhduls):
detid = hdul[0].header.get('DET-ID')
detx = hdul[0].header.get('DET-P101')
dety = hdul[0].header.get('DET-P201')
chxpos = []
chypos = []
minyos = {}
minyef = {}
minxos = {}
minxef = {}
maxyos = {}
maxyef = {}
maxxos = {}
maxxef = {}
for ch in [1,2,3,4]:
minxos[ch] = hdul[0].header.get(f'S_OSMN{ch}1')
maxxos[ch] = hdul[0].header.get(f'S_OSMX{ch}1')
minyos[ch] = hdul[0].header.get(f'S_OSMN{ch}2')
maxyos[ch] = hdul[0].header.get(f'S_OSMX{ch}2')
minxef[ch] = hdul[0].header.get(f'S_EFMN{ch}1')
maxxef[ch] = hdul[0].header.get(f'S_EFMX{ch}1')
minyef[ch] = hdul[0].header.get(f'S_EFMN{ch}2')
maxyef[ch] = hdul[0].header.get(f'S_EFMX{ch}2')
chxpos.append(minxef[ch])
chypos.append(minyef[ch])
minxpix = min(chxpos)
minypix = min(chypos)
tb = {True: 2, False: 1}[dety > -30]
DETSECy2 = tb * (maxyef[1] - minypix)
DETSECy1 = DETSECy2 - (maxyef[1] - minyef[1]) + 1
for ch in [1,2,3,4]:
extver += 1
miny = min([minyos[ch], minyef[ch]])
minx = min([minxos[ch], minxef[ch]])
maxy = max([maxyos[ch], maxyef[ch]])
maxx = max([maxxos[ch], maxxef[ch]])
chdata = hdul[0].data[miny:maxy,minx:maxx]
if extver == 1:
phdu = fits.PrimaryHDU(None, hdul[0].header)
MEF.append(phdu)
chhdu = fits.ImageHDU(chdata, hdul[0].header, name=f'd{detid}c{ch}')
chhdu.header.set('EXTNAME', f'im{extver}')
chhdu.header.set('EXTVER', extver)
chhdu.header.set('IMAGEID', f'd{detid}c{ch}')
chhdu.header.set('CCDNAME', hdul[0].header.get('DETECTOR'))
binx = int(hdul[0].header.get('BIN-FCT1'))
biny = int(hdul[0].header.get('BIN-FCT2'))
chhdu.header.set('CCDSUM', f'{binx:d} {biny:d}')
chhdu.header.set('FILTER', hdul[0].header.get('FILTER01'))
# OBSTYPE: "zero", "dark", flat", and "object"
obstype_trans = {'DOMEFLAT': 'flat', 'SKYFLAT': 'twiflat',
'BIAS': 'zero', 'OBJECT': 'object', 'DARK': 'dark'}
obstype = obstype_trans[hdul[0].header.get('DATA-TYP').strip()]
chhdu.header.set('OBSTYPE', obstype)
BIASSECx1 = minxos[ch] - minx + 1
BIASSECx2 = maxxos[ch] - minx
BIASSECy1 = minyef[ch] - miny + 1
BIASSECy2 = maxyef[ch] - miny
biassec = f'[{BIASSECx1:d}:{BIASSECx2:d},{BIASSECy1:d}:{BIASSECy2:d}]'
chhdu.header.set('BIASSEC', biassec)
DATASECx1 = minxef[ch] - minx + 1
DATASECx2 = maxxef[ch] - minx
DATASECy1 = minyef[ch] - miny + 1
DATASECy2 = maxyef[ch] - miny
datasec = f'[{DATASECx1:d}:{DATASECx2:d},{DATASECy1:d}:{DATASECy2:d}]'
chhdu.header.set('DATASEC', datasec)
print(biassec, datasec)
detsecy = {True: 2, False: 1}[dety > 0]
ampwidth = maxxef[ch] - minxef[ch]
chipwidth = 4 * ampwidth
if detx > 30:
DETSECx1 = 4 * chipwidth + (maxxef[ch] - ampwidth - minxef[ch] % ampwidth) + 1
elif detx > 0:
DETSECx1 = 3 * chipwidth + (maxxef[ch] - ampwidth - minxef[ch] % ampwidth) + 1
elif detx > -30:
DETSECx1 = 2 * chipwidth + (maxxef[ch] - ampwidth - minxef[ch] % ampwidth) + 1
elif detx > -60:
DETSECx1 = 1 * chipwidth + (maxxef[ch] - ampwidth - minxef[ch] % ampwidth) + 1
else:
DETSECx1 = 0 * chipwidth + (maxxef[ch] - ampwidth - minxef[ch] % ampwidth) + 1
DETSEXx2 = DETSECx1 + ampwidth -1
chhdu.header.set('DETSEC', f'[{DETSECx1:d}:{DETSEXx2:d},{DETSECy1:d}:{DETSECy2:d}]')
MEF.append(chhdu)
return MEF
filename = 'SUPA01564270.fits'
MEF = assemble_MEF(os.path.join(rawdir, filename))
MEF.writeto(filename.replace('SUPA', 'MEF_'), overwrite=True)
# +
plt.figure(figsize=(16,14))
for i,hdu in enumerate(MEF[1:5]):
plt.subplot(1,4,i+1)
vmin = np.percentile(hdu.data, 0.75)
vmax = np.percentile(hdu.data, 98.5)
plt.imshow(hdu.data, vmin=vmin-100, vmax=vmax)
plt.show()
# -
| notebooks/Generate MEF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from gs_quant.session import Environment, GsSession
from gs_quant.instrument import IRSwaption
from gs_quant.markets.portfolio import Portfolio
from gs_quant.markets import PricingContext
from gs_quant.risk import CarryScenario
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# external users should substitute their client id and secret; please skip this step if using internal jupyterhub
GsSession.use(Environment.PROD, client_id=None, client_secret=None, scopes=('run_analytics',))
# +
# basic usage of carry scenario
eur1y10y = IRSwaption('Pay', '10y', 'EUR', expiration_date='1y')
# care needs to be taken when creating relative trades like the one above.
# If you don't resolve the trade, the resolution of the trade parameters will be done with
# reference to the active pricing context. Under the carry scenario this means that
# if you don't resolve the trade will be a different trade when priced under the carry scenario.
eur1y10y.resolve()
# Shift forward 22 business days (1 month)
carry_scenario = CarryScenario(time_shift=22)
with carry_scenario:
fwd_price = eur1y10y.price()
print('Base price: {:,.2f}'.format(eur1y10y.price()))
print('Scenario price: {:,.2f}'.format(fwd_price))
# +
# show how the option value will roll down moving forward 66 business days assuming either fwds
# or spot rates are realised.
short_swaption = IRSwaption('Pay', '5y', 'USD', expirationDate='6m', notionalAmount=1e8)
short_swaption.resolve()
prices = []
roll_spot_prices = []
with PricingContext():
for time_shift in range(66):
with CarryScenario(time_shift, roll_to_fwds=True):
prices.append(short_swaption.price())
with CarryScenario(time_shift, roll_to_fwds=False):
roll_spot_prices.append(short_swaption.price())
pd.Series([p.result() for p in prices], dtype=np.dtype(float)).plot(figsize=(10, 6),
title="Swaption Price Forward in Time",
label='roll to fwd')
pd.Series([rp.result() for rp in roll_spot_prices], dtype=np.dtype(float)).plot(figsize=(10, 6),
label='roll to spot')
plt.xlabel('TimeShift')
plt.ylabel('PV')
# -
# create a grid of expiry by tenor swaptions showing the pv under the carry scenario minus the base pv.
def calc_risk_matrix(ccy, strike, pay_rec, time_shift, roll_to_fwds, expiries, tenors):
portfolio = Portfolio([IRSwaption(pay_rec, tenor, ccy,
expiration_date=expiry, strike=strike, name='{}_{}'.format(expiry, tenor))
for expiry in expiries for tenor in tenors])
portfolio.resolve()
with CarryScenario(time_shift, roll_to_fwds):
carry_results = portfolio.price()
base_results = portfolio.price()
carry_records = [(carry_results[t]-base_results[t], t.name.split('_')[0], t.name.split('_')[1]) for t in portfolio]
carry_df = pd.DataFrame(carry_records, columns=['Value', 'Expiry', 'Tenor'])
pivot_df = carry_df.pivot_table(values='Value', index='Expiry', columns='Tenor')
return pivot_df[tenors].reindex(expiries)
# +
ccy = 'EUR'
strike = 'ATM'
pay_rec = 'Pay' # or 'Receive' or 'Straddle'
time_shift = 22 # in business days for carry scenario
roll_to_fwds = True
expiries = ['1m', '2m', '3m', '6m', '9m', '1y', '18m', '2y', '3y', '5y', '7y', '10y']
tenors = ['1m', '3m', '6m', '1y', '2y', '3y', '5y', '7y', '10y', '15y', '20y', '25y', '30y']
calc_risk_matrix(ccy, strike, pay_rec, time_shift, roll_to_fwds, expiries, tenors)
| gs_quant/examples/04_scenario/01_carry_shock/040101_basic_use_of_carry_scenario.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/xhang24/xiaotong/blob/master/src/hw9.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="jllLxl8tjIvN" colab_type="code" colab={}
import numpy as np
# + id="h2SQn0gQjL6P" colab_type="code" colab={}
def init_value(N):
h=1/N
v=np.zeros([N+1,N+1])
for i in range(N+1):
for j in range(N+1):
if i==0 or i==N or j==0 or j==N:
v[i,j]=((i/N-0.5)**2+(j/N-0.5)**2)
return v
# + id="jS7upf_sjOHW" colab_type="code" colab={}
def F(u,N):
h=1/N
v=np.zeros([N+1,N+1])
for i in range(N+1):
for j in range(N+1):
if i==0 or i==N or j==0 or j==N:
v[i,j]=u[i,j]
else:
v[i,j]=(2/(2+h**2))*((h**2/2)*((i*h)**2+(j*h)**2-(i*h)-(j*h)-(3/2))+(1/4)*(u[i+1,j]+u[i,j+1]+u[i,j-1]+u[i-1,j]))
return v
# + id="EAftAH7MjQTi" colab_type="code" colab={}
def VI(N, tolerance):
v=init_value(N)
error=1
step=0
while error>tolerance:
step+=1
u=v
v=F(u,N)
error=np.max(np.abs(u - v))
return [error, step, v]
# + id="8cczpt91jSWr" colab_type="code" colab={}
def exact_value(N):
v=np.zeros([N+1,N+1])
for i in range(0,N+1):
for j in range(0,N+1):
v[i,j]=(i/N-0.5)**2+(j/N-0.5)**2
return v
# + id="1SvNgpVCjUot" colab_type="code" outputId="65935267-462c-4682-a004-a6014cff3362" colab={"base_uri": "https://localhost:8080/", "height": 1000}
for i in range(4):
N=2**(i+2)
h=1/N
soln=VI(N,0.0001)[2]
error=np.max(np.abs(VI(N,0.0001)[2]-exact_value(N)))
print('>>>>>>For h=' + str(h) +',the CFD solution is \n ' , soln)
print('>>>>>>The maxnorm of error is ' , error)
| src/hw9.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import statistics
import collections
import itertools
import functools
import operator
import multiprocessing
import numpy as np
import scipy as sp
import scipy.stats
import pandas as pd
from matplotlib import pyplot as plt
from IPython.display import clear_output
# %load_ext snakeviz
# +
train = pd.read_csv("../data/raw/train.csv", index_col="PassengerId")
test = pd.read_csv("../data/raw/test.csv", index_col="PassengerId")
def enrich(ds):
ds["Desk"] = ds["Cabin"].dropna().apply(lambda x: x[0])
enrich(train)
enrich(test)
train
# -
def hist(cats, vals, catc):
histo, bins = np.histogram(vals, bins="auto")
s = np.sum(histo)
plt.xticks(bins)
for cat, col in catc.items():
h, _ = np.histogram(vals[cats == cat], bins=bins)
# dist = sp.stats.rv_histogram((h, bins))
plt.bar(bins[:-1], h / s, width=(bins[-1] - bins[0]) / (len(bins) - 1), fc=col + (0.5,))
plt.show()
survived = train["Survived"]
color = {
0: (1, 0, 0),
1: (0, 1, 0)
}
hist(survived, train["Age"].dropna().astype(int), color)
# +
def normal_dist_generator(sigma):
k = 1 / (sigma * math.sqrt(2 * math.pi))
s = (2 * sigma ** 2)
def normal_dist_func(z, rel):
return k * math.exp(-(z - rel)**2 / s)
return normal_dist_func
def discr(peak, miss):
return lambda z, rel: (peak if z == rel else miss)
class Model:
def __init__(self):
self._df = None
self._cols = None
self._wf = None
def relearn(self, df):
self._df = df
return self
def reset_features(self, **kwargs):
self._cols = list(kwargs.keys())
self._wf = {}
for feature, config in kwargs.items():
if config["type"] == "finite":
peak = config["peak"]
miss = (1 - peak) / (len(config["values"]) - 1)
self._wf[feature] = np.vectorize(discr(peak, miss))
elif config["type"] == "gauss":
sigma = config["sigma"]
self._wf[feature] = np.vectorize(normal_dist_generator(sigma))
else:
raise ValueError("Unknown type: {}".format(config["type"]))
return self
def get_weight(self, z):
col_weights = []
for col, zv in z.iteritems():
if zv != zv:
continue
binop = self._wf[col]
col_weights.append(binop(zv, self._df[col].values))
cw = np.column_stack(col_weights)
# cw = pd.DataFrame(cw)
return np.nansum(np.nanprod(cw, axis=1))
def prob(self, z):
z = z[set(z.columns) & set(self._cols)]
total_count = np.size(self._df.index)
weight = z.apply(self.get_weight, axis=1)
return weight / total_count
def cond_prob(self, z, prop, vals):
assert prop not in z.columns, "Conditional probability of known value"
assert prop in self._cols, "Conditional probability on ignored column"
p = self.prob(z)
c_props = []
for val in vals:
z_with_val = z.assign(**{prop: val})
p_with_val = self.prob(z_with_val)
c_prop = p_with_val / p
c_props.append(c_prop)
return pd.concat(c_props, axis=1)
def mean_succ_prob(m, sample, target, classes, splits=10):
sz = len(sample.index) // splits
res = []
for split_id in range(splits):
tra = pd.concat([sample[:split_id * sz], sample[(split_id + 1)* sz:]])
tes = sample[split_id * sz:(split_id + 1)* sz]
m.relearn(tra)
pred = m.cond_prob(tes.drop(columns=[target]), target, classes)
pred["best_prediction"] = pred.apply(lambda r: r.argmax(), axis=1)
pred["actual"] = tes[target]
pred["correct"] = pred["best_prediction"] == pred["actual"]
res.append(np.mean(pred["correct"]))
return np.mean(res)
# +
exps = pd.DataFrame()
m = Model()
for sibsp in [.9, .8, .5]:
for parch in [.9, .8, .5]:
for pclass in [.9, .8, .5]:
for sex in [.9, .8, .5]:
for desk in [.9, .8, .5]:
for age in [.1, 1.0]:
m.reset_features(**{
"Sex": {"type": "finite", "peak": sex, "values": ["male", "female"]},
"Age": {"type": "gauss", "sigma": age},
"Pclass": {"type": "finite", "peak": pclass, "values": [1, 2, 3]},
"Desk": {"type": "finite", "peak": desk, "values": ["A", "B", "C", "D", "E", "F", "G"]},
"Survived": {"type": "finite", "peak": 1.0, "values": [0, 1]},
"SibSp": {"type": "gauss", "sigma": sibsp},
"Parch": {"type": "gauss", "sigma": parch},
})
p = mean_succ_prob(m, train, "Survived", [0, 1])
exps = exps.append(
pd.Series(
dict(
sibsp=sibsp,
parch=parch,
pclass=pclass,
sex=sex,
desk=desk,
age=age,
p=p
)
),
ignore_index=True
).sort_values(by="p", ascending=False)
clear_output(wait=True)
display(exps)
# -
non_parametric_v01 = Model().reset_features(**{
"Sex": {"type": "finite", "peak": .9, "values": ["male", "female"]},
"Age": {"type": "gauss", "sigma": .1},
"Pclass": {"type": "finite", "peak": .5, "values": [1, 2, 3]},
"Desk": {"type": "finite", "peak": .8, "values": ["A", "B", "C", "D", "E", "F", "G"]},
"Survived": {"type": "finite", "peak": 1.0, "values": [0, 1]},
"SibSp": {"type": "gauss", "sigma": .8},
"Parch": {"type": "gauss", "sigma": .9},
})
mean_succ_prob(non_parametric_v01, train, "Survived", [0, 1])
non_parametric_v01.relearn(train)
(
(non_parametric_v01.cond_prob(test, "Survived", [0, 1])[1] > 0.5)
.rename("Survived")
.astype(int)
.to_csv("../models/non_parametric_v01/non_parametric_v01.csv")
)
| drafts/kaggle/titanic/notebooks/non_parametric.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ZfpTLD7GajFb"
# # Find duplicate questions on StackOverflow by their embeddings
#
# In this assignment you will learn how to calculate a similarity for pieces of text. Using this approach you will know how to find duplicate questions from [StackOverflow](https://stackoverflow.com).
# + [markdown] id="MJ91mGTTajFb"
# ### Libraries
#
# In this task you will you will need the following libraries:
# - [StarSpace](https://github.com/facebookresearch/StarSpace) — a general-purpose model for efficient learning of entity embeddings from Facebook
# - [Gensim](https://radimrehurek.com/gensim/) — a tool for solving various NLP-related tasks (topic modeling, text representation, ...)
# - [Numpy](http://www.numpy.org) — a package for scientific computing.
# - [scikit-learn](http://scikit-learn.org/stable/index.html) — a tool for data mining and data analysis.
# - [Nltk](http://www.nltk.org) — a platform to work with human language data.
# + [markdown] id="I-Qiql4ZajFb"
# ### Data
#
# The following cell will download all data required for this assignment into the folder `week3/data`.
# + colab={"base_uri": "https://localhost:8080/", "height": 538, "referenced_widgets": ["f595857c272f4d42873e39cccdaa06da", "33110456b0f448409bcbbb28ad29303e", "ce4a42704dda45798cfd20bb6f2ba712", "365334c7de1149e9ac58716e8fae3003", "d17e253365fb48188247e330a5a7e1ac", "7af4c8da0c154beca10d27586ca928c6", "35bde971a25f4047a1c485feeed02d71", "7fecbe50d029497a9a17b6dae0a641d0", "098c7a6be01d4933bef052bf68e61f27", "<KEY>", "6ea1117104f34335a903c73d544ea112", "<KEY>", "375c9a8278f4424bad3bebafec579405", "7c25de210ad74d27a4996335e403d52d", "<KEY>", "0cf0ed1319a1484b9edca61a444e6fea", "d931cdfeb07d405c9e85b9eec06aa338", "<KEY>", "<KEY>", "660c8adc0aae437e9ed0e34a31f3bdeb", "e0ca71ff3697411dbb1ab1dde0b4a8b4", "<KEY>", "9ed941b47695495ca315893a970d5fdb", "d55e4afa8d3a42b1a719e678aaede953", "<KEY>", "<KEY>", "3abe7492e94c47ae8bc0a279c3a28f1e", "e4b659297ee4423e9440ca53b2eceaba", "<KEY>", "3b038500d2c2406eb5266a7df0d5d1f0", "6f21e6ac202f48ca9c8f8c9fdfe10e5d", "b1b24dee4e4a490e95eb1e5d43e40591", "025b6383dfde498d8dc12b59e907716f", "<KEY>", "7f3bb13d1e8d4f9e9d4a03710c15fc02", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "40d4be5a731040c4a62b932842322514", "7f88af7fba0a4b7d98f32c05e27e75e1", "<KEY>", "9101ab0b96b145e28ed3806b74cb83e5", "<KEY>", "0e36e220bdac4894877fac4e5df46af4", "e22d9706722c4068ae299f6295767549", "<KEY>", "<KEY>", "<KEY>", "d0b0da7479f543d1861f7f235fad169d", "7f1680c596a74abe842c56831cd77f61", "50cacc91ad2149f8b0d11a3bab19443d", "fb18e5ab48304c54b38a505fe06ba9db", "e37fe0656abe4977a839c1a844d12263", "7a309e98f9974946943c3ca33050aa10"]} id="OYK8gdN7ajFb" outputId="32971b87-769d-4c91-942b-ff8e0314c943"
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
# ! wget https://raw.githubusercontent.com/hse-aml/natural-language-processing/master/setup_google_colab.py -O setup_google_colab.py
import setup_google_colab
setup_google_colab.setup_week3()
import sys
sys.path.append("..")
from common.download_utils import download_week3_resources
download_week3_resources()
# + colab={"base_uri": "https://localhost:8080/"} id="AsoFsHIClvAl" outputId="c899bfb0-ab6c-4cfb-af77-c87d7a2b1502"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="uJsMQifsl1ww" outputId="7b68f8ed-d27c-4d2d-ffce-7d927cd95db7"
# !ls Starspace
# + colab={"base_uri": "https://localhost:8080/"} id="VFGWaF0kmCbl" outputId="80e2f6ec-4746-411a-cbf8-1a4c6d2a1210"
# ! ls data
# + [markdown] id="AEfGIGxfajFd"
# ### Grading
# We will create a grader instace below and use it to collect your answers. Note that these outputs will be stored locally inside grader and will be uploaded to platform only after running submiting function in the last part of this assignment. If you want to make partial submission, you can run that cell any time you want.
# + id="KUUVybzwajFd"
from grader import Grader
# + id="x1YXeXV9ajFd"
grader = Grader()
# + [markdown] id="64LMGOwKajFd"
# ## Word embedding
#
# To solve the problem, you will use two different models of embeddings:
#
# - [Pre-trained word vectors](https://code.google.com/archive/p/word2vec/) from Google which were trained on a part of Google News dataset (about 100 billion words). The model contains 300-dimensional vectors for 3 million words and phrases. `GoogleNews-vectors-negative300.bin.gz` will be downloaded in `download_week3_resources()`.
# - Representations using StarSpace on StackOverflow data sample. You will need to train them from scratch.
# + [markdown] id="EemqZ8XlajFd"
# It's always easier to start with pre-trained embeddings. Unpack the pre-trained Goggle's vectors and upload them using the function [KeyedVectors.load_word2vec_format](https://radimrehurek.com/gensim/models/keyedvectors.html) from gensim library with the parameter *binary=True*. If the size of the embeddings is larger than the avaliable memory, you could load only a part of the embeddings by defining the parameter *limit* (recommended: 500000).
# + id="0HywiRsrajFd"
import gensim
# + id="itAKbZ8fajFd"
from gensim.models import KeyedVectors
wv_embeddings = KeyedVectors.load_word2vec_format('GoogleNews-vectors-negative300.bin.gz', binary=True, limit=500000)
# + [markdown] id="AaRv5GbJajFd"
# ### How to work with Google's word2vec embeddings?
#
# Once you have loaded the representations, make sure you can access them. First, you can check if the loaded embeddings contain a word:
#
# 'word' in wv_embeddings
#
# Second, to get the corresponding embedding you can use the square brackets:
#
# wv_embeddings['word']
#
# ### Checking that the embeddings are correct
#
# To prevent any errors during the first stage, we can check that the loaded embeddings are correct. You can call the function *check_embeddings*, implemented below, which runs 3 tests:
# 1. Find the most similar word for provided "positive" and "negative" words.
# 2. Find which word from the given list doesn’t go with the others.
# 3. Find the most similar word for the provided one.
#
# In the right case the function will return the string *These embeddings look good*. Othervise, you need to validate the previous steps.
# + id="wbZBlo5rajFd"
def check_embeddings(embeddings):
error_text = "Something wrong with your embeddings ('%s test isn't correct)."
most_similar = embeddings.most_similar(positive=['woman', 'king'], negative=['man'])
if len(most_similar) < 1 or most_similar[0][0] != 'queen':
return error_text % "Most similar"
doesnt_match = embeddings.doesnt_match(['breakfast', 'cereal', 'dinner', 'lunch'])
if doesnt_match != 'cereal':
return error_text % "Doesn't match"
most_similar_to_given = embeddings.most_similar_to_given('music', ['water', 'sound', 'backpack', 'mouse'])
if most_similar_to_given != 'sound':
return error_text % "Most similar to given"
return "These embeddings look good."
# + colab={"base_uri": "https://localhost:8080/"} id="i-X_w6-EajFd" outputId="fa706eba-90b7-4a99-ae9f-58f5d2e7f97a"
print(check_embeddings(wv_embeddings))
# + [markdown] id="l4rBXWfxajFd"
# ## From word to text embeddings
#
# **Task 1 (Question2Vec).** Usually, we have word-based embeddings, but for the task we need to create a representation for the whole question. It could be done in different ways. In our case we will use a **mean** of all word vectors in the question. Now you need to implement the function *question_to_vec*, which calculates the question representation described above. This function should work with the input text as is without any preprocessing.
#
# Note that there could be words without the corresponding embeddings. In this case, you can just skip these words and don't take them into account during calculating the result. If the question doesn't contain any known word with embedding, the function should return a zero vector.
# + id="kPWm-WW9ajFd"
import numpy as np
# + id="kYqTrdtHajFd"
def question_to_vec(question, embeddings, dim=300):
"""
question: a string
embeddings: dict where the key is a word and a value is its' embedding
dim: size of the representation
result: vector representation for the question
"""
######################################
######### YOUR CODE HERE #############
######################################
result = np.zeros(dim)
count = 0
for word in question.split():
if word in embeddings:
result += embeddings[word]
count += 1
return result / count if count != 0 else result
# + [markdown] id="z3huS2ZmajFd"
# To check the basic correctness of your implementation, run the function *question_to_vec_tests*.
# + id="pAM_dmnmajFd"
def question_to_vec_tests():
if (np.zeros(300) != question_to_vec('', wv_embeddings)).any():
return "You need to return zero vector for empty question."
if (np.zeros(300) != question_to_vec('thereisnosuchword', wv_embeddings)).any():
return "You need to return zero vector for the question, which consists only unknown words."
if (wv_embeddings['word'] != question_to_vec('word', wv_embeddings)).any():
return "You need to check the corectness of your function."
if ((wv_embeddings['I'] + wv_embeddings['am']) / 2 != question_to_vec('I am', wv_embeddings)).any():
return "Your function should calculate a mean of word vectors."
if (wv_embeddings['word'] != question_to_vec('thereisnosuchword word', wv_embeddings)).any():
return "You should not consider words which embeddings are unknown."
return "Basic tests are passed."
# + colab={"base_uri": "https://localhost:8080/"} id="21gmQbcuajFd" outputId="c22c8fb0-f186-4ded-a4b9-5e85193f4973"
print(question_to_vec_tests())
# + [markdown] id="do97-Rp6ajFd"
# You can submit embeddings for the questions from the file *test_embeddings.tsv* to earn the points. In this task you don't need to transform the text of a question somehow.
# + colab={"base_uri": "https://localhost:8080/"} id="4nKukh6sajFd" outputId="dbe48ef4-d7bd-4117-e950-1291c1be0827"
import nltk
nltk.download('stopwords')
from util import array_to_string
# + colab={"base_uri": "https://localhost:8080/"} id="UYeXRm9uajFd" outputId="9568879a-c02d-4b42-be6f-2fdcca785d4c"
question2vec_result = []
for question in open('data/test_embeddings.tsv'):
question = question.strip()
answer = question_to_vec(question, wv_embeddings)
question2vec_result = np.append(question2vec_result, answer)
grader.submit_tag('Question2Vec', array_to_string(question2vec_result))
# + [markdown] id="ACvR3KMDajFd"
# Now we have a method to create a representation of any sentence and we are ready for the first evaluation. So, let's check how well our solution (Google's vectors + *question_to_vec*) will work.
#
# ## Evaluation of text similarity
#
# We can imagine that if we use good embeddings, the cosine similarity between the duplicate sentences should be less than for the random ones. Overall, for each pair of duplicate sentences we can generate *R* random negative examples and find out the position of the correct duplicate.
#
# For example, we have the question *"Exceptions What really happens"* and we are sure that another question *"How does the catch keyword determine the type of exception that was thrown"* is a duplicate. But our model doesn't know it and tries to find out the best option also among questions like *"How Can I Make These Links Rotate in PHP"*, *"NSLog array description not memory address"* and *"PECL_HTTP not recognised php ubuntu"*. The goal of the model is to rank all these 4 questions (1 *positive* and *R* = 3 *negative*) in the way that the correct one is in the first place.
#
# However, it is unnatural to count on that the best candidate will be always in the first place. So let us consider the place of the best candidate in the sorted list of candidates and formulate a metric based on it. We can fix some *K* — a reasonalble number of top-ranked elements and *N* — a number of queries (size of the sample).
#
# ### Hits@K
#
# The first simple metric will be a number of correct hits for some *K*:
# $$ \text{Hits@K} = \frac{1}{N}\sum_{i=1}^N \, [dup_i \in topK(q_i)]$$
#
# where $q_i$ is the i-th query, $dup_i$ is its duplicate, $topK(q_i)$ is the top K elements of the ranked sentences provided by our model and the operation $[dup_i \in topK(q_i)]$ equals 1 if the condition is true and 0 otherwise (more details about this operation could be found [here](https://en.wikipedia.org/wiki/Iverson_bracket)).
#
#
# ### DCG@K
# The second one is a simplified [DCG metric](https://en.wikipedia.org/wiki/Discounted_cumulative_gain):
#
# $$ \text{DCG@K} = \frac{1}{N} \sum_{i=1}^N\frac{1}{\log_2(1+rank_{dup_i})}\cdot[rank_{dup_i} \le K] $$
#
# where $rank_{dup_i}$ is a position of the duplicate in the sorted list of the nearest sentences for the query $q_i$. According to this metric, the model gets a higher reward for a higher position of the correct answer. If the answer does not appear in topK at all, the reward is zero.
# + [markdown] id="01tFbc2CajFd"
# ### Evaluation examples
#
# Let's calculate the described metrics for the toy example introduced above. In this case $N$ = 1 and the correct candidate for $q_1$ is *"How does the catch keyword determine the type of exception that was thrown"*. Consider the following ranking of the candidates:
# 1. *"How Can I Make These Links Rotate in PHP"*
# 2. *"How does the catch keyword determine the type of exception that was thrown"*
# 3. *"NSLog array description not memory address"*
# 4. *"PECL_HTTP not recognised php ubuntu"*
#
# Using the ranking above, calculate *Hits@K* metric for *K = 1, 2, 4*:
#
# - [K = 1] $\text{Hits@1} = \frac{1}{1}\sum_{i=1}^1 \, [dup_i \in top1(q_i)] = [dup_1 \in top1(q_1)] = 0$ because the correct answer doesn't appear in the *top1* list.
# - [K = 2] $\text{Hits@2} = \frac{1}{1}\sum_{i=1}^1 \, [dup_i \in top2(q_i)] = [dup_1 \in top2(q_1)] = 1$ because $rank_{dup_1} = 2$.
# - [K = 4] $\text{Hits@4} = \frac{1}{1}\sum_{i=1}^1 \, [dup_i \in top4(q_i)] = [dup_1 \in top4(q_1)] = 1$
#
# Using the ranking above, calculate *DCG@K* metric for *K = 1, 2, 4*:
#
# - [K = 1] $\text{DCG@1} = \frac{1}{1} \sum_{i=1}^1\frac{1}{\log_2(1+rank_{dup_i})}\cdot[rank_{dup_i} \le 1] = \frac{1}{\log_2(1+rank_{dup_i})}\cdot[rank_{dup_i} \le 1] = 0$ because the correct answer doesn't appear in the top1 list.
# - [K = 2] $\text{DCG@2} = \frac{1}{1} \sum_{i=1}^1\frac{1}{\log_2(1+rank_{dup_i})}\cdot[rank_{dup_i} \le 2] = \frac{1}{\log_2{3}}$, because $rank_{dup_1} = 2$.
# - [K = 4] $\text{DCG@4} = \frac{1}{1} \sum_{i=1}^1\frac{1}{\log_2(1+rank_{dup_i})}\cdot[rank_{dup_i} \le 4] = \frac{1}{\log_2{3}}$.
#
# + [markdown] id="FFko08SFajFd"
# **Tasks 2 and 3 (HitsCount and DCGScore).** Implement the functions *hits_count* and *dcg_score* as described above. Each function has two arguments: *dup_ranks* and *k*. *dup_ranks* is a list which contains *values of ranks* of duplicates. For example, *dup_ranks* is *[2]* for the example provided above.
# + id="OJbF3v8hajFd"
def hits_count(dup_ranks, k):
"""
dup_ranks: list of duplicates' ranks; one rank per question;
length is a number of questions which we are looking for duplicates;
rank is a number from 1 to len(candidates of the question);
e.g. [2, 3] means that the first duplicate has the rank 2, the second one — 3.
k: number of top-ranked elements (k in Hits@k metric)
result: return Hits@k value for current ranking
"""
######################################
######### YOUR CODE HERE #############
######################################
return sum(rank <= k for rank in dup_ranks) / len(dup_ranks)
# + [markdown] id="_geDdfFvajFd"
# Test your code on the tiny examples:
# + id="Hkk8HhuyajFd"
def test_hits():
# *Evaluation example*
# answers — dup_i
answers = ["How does the catch keyword determine the type of exception that was thrown"]
# candidates_ranking — the ranked sentences provided by our model
candidates_ranking = [["How Can I Make These Links Rotate in PHP",
"How does the catch keyword determine the type of exception that was thrown",
"NSLog array description not memory address",
"PECL_HTTP not recognised php ubuntu"]]
# dup_ranks — position of the dup_i in the list of ranks +1
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
# correct_answers — the expected values of the result for each k from 1 to 4
correct_answers = [0, 1, 1, 1]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(hits_count(dup_ranks, k), correct):
return "Check the function."
# Other tests
answers = ["How does the catch keyword determine the type of exception that was thrown",
"Convert Google results object (pure js) to Python object"]
# The first test: both duplicates on the first position in ranked list
candidates_ranking = [["How does the catch keyword determine the type of exception that was thrown",
"How Can I Make These Links Rotate in PHP"],
["Convert Google results object (pure js) to Python object",
"WPF- How to update the changes in list item of a list"]]
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
correct_answers = [1, 1]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(hits_count(dup_ranks, k), correct):
return "Check the function (test: both duplicates on the first position in ranked list)."
# The second test: one candidate on the first position, another — on the second
candidates_ranking = [["How Can I Make These Links Rotate in PHP",
"How does the catch keyword determine the type of exception that was thrown"],
["Convert Google results object (pure js) to Python object",
"WPF- How to update the changes in list item of a list"]]
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
correct_answers = [0.5, 1]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(hits_count(dup_ranks, k), correct):
return "Check the function (test: one candidate on the first position, another — on the second)."
# The third test: both candidates on the second position
candidates_ranking = [["How Can I Make These Links Rotate in PHP",
"How does the catch keyword determine the type of exception that was thrown"],
["WPF- How to update the changes in list item of a list",
"Convert Google results object (pure js) to Python object"]]
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
correct_answers = [0, 1]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(hits_count(dup_ranks, k), correct):
return "Check the function (test: both candidates on the second position)."
return "Basic test are passed."
# + colab={"base_uri": "https://localhost:8080/"} id="3QbuZB0fajFd" outputId="2bc7320e-833b-4de7-822e-515280931fa6"
print(test_hits())
# + id="hlodhjPmajFd"
def dcg_score(dup_ranks, k):
"""
dup_ranks: list of duplicates' ranks; one rank per question;
length is a number of questions which we are looking for duplicates;
rank is a number from 1 to len(candidates of the question);
e.g. [2, 3] means that the first duplicate has the rank 2, the second one — 3.
k: number of top-ranked elements (k in DCG@k metric)
result: return DCG@k value for current ranking
"""
######################################
######### YOUR CODE HERE #############
######################################
return sum(1 / (np.log2(1 + rank)) for rank in dup_ranks if rank <= k) / len(dup_ranks)
# + id="15siQ6rCajFd"
def test_dcg():
# *Evaluation example*
# answers — dup_i
answers = ["How does the catch keyword determine the type of exception that was thrown"]
# candidates_ranking — the ranked sentences provided by our model
candidates_ranking = [["How Can I Make These Links Rotate in PHP",
"How does the catch keyword determine the type of exception that was thrown",
"NSLog array description not memory address",
"PECL_HTTP not recognised php ubuntu"]]
# dup_ranks — position of the dup_i in the list of ranks +1
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
# correct_answers — the expected values of the result for each k from 1 to 4
correct_answers = [0, 1 / (np.log2(3)), 1 / (np.log2(3)), 1 / (np.log2(3))]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(dcg_score(dup_ranks, k), correct):
return "Check the function."
# Other tests
answers = ["How does the catch keyword determine the type of exception that was thrown",
"Convert Google results object (pure js) to Python object"]
# The first test: both duplicates on the first position in ranked list
candidates_ranking = [["How does the catch keyword determine the type of exception that was thrown",
"How Can I Make These Links Rotate in PHP"],
["Convert Google results object (pure js) to Python object",
"WPF- How to update the changes in list item of a list"]]
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
correct_answers = [1, 1]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(dcg_score(dup_ranks, k), correct):
return "Check the function (test: both duplicates on the first position in ranked list)."
# The second test: one candidate on the first position, another — on the second
candidates_ranking = [["How Can I Make These Links Rotate in PHP",
"How does the catch keyword determine the type of exception that was thrown"],
["Convert Google results object (pure js) to Python object",
"WPF- How to update the changes in list item of a list"]]
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
correct_answers = [0.5, (1 + (1 / (np.log2(3)))) / 2]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(dcg_score(dup_ranks, k), correct):
return "Check the function (test: one candidate on the first position, another — on the second)."
# The third test: both candidates on the second position
candidates_ranking = [["How Can I Make These Links Rotate in PHP",
"How does the catch keyword determine the type of exception that was thrown"],
["WPF- How to update the changes in list item of a list",
"Convert Google results object (pure js) to Python object"]]
dup_ranks = [candidates_ranking[i].index(answers[i]) + 1 for i in range(len(answers))]
correct_answers = [0, 1 / (np.log2(3))]
for k, correct in enumerate(correct_answers, 1):
if not np.isclose(dcg_score(dup_ranks, k), correct):
return "Check the function (test: both candidates on the second position)."
return "Basic test are passed."
# + colab={"base_uri": "https://localhost:8080/"} id="_-J4rqszajFe" outputId="e5409e91-8e6a-4841-ea86-e5edf74cf92d"
print(test_dcg())
# + [markdown] id="ivce5MPiajFe"
# Submit results of the functions *hits_count* and *dcg_score* for the following examples to earn the points.
# + id="CTv1xPqXajFe"
test_examples = [
[1],
[1, 2],
[2, 1],
[1, 2, 3],
[1, 2, 3, 4, 5, 6, 7, 8, 9, 10],
[9, 5, 4, 2, 8, 10, 7, 6, 1, 3],
[4, 3, 5, 1, 9, 10, 7, 8, 2, 6],
[5, 1, 7, 6, 2, 3, 8, 9, 10, 4],
[6, 3, 1, 4, 7, 2, 9, 8, 10, 5],
[10, 9, 8, 7, 6, 5, 4, 3, 2, 1],
]
# + colab={"base_uri": "https://localhost:8080/"} id="0eo5HiNfajFe" outputId="21cfa2d8-1b89-4986-bc74-57ca8d49a911"
hits_results = []
for example in test_examples:
for k in range(len(example)):
hits_results.append(hits_count(example, k + 1))
grader.submit_tag('HitsCount', array_to_string(hits_results))
# + colab={"base_uri": "https://localhost:8080/"} id="EnqeczDpajFf" outputId="e07aadc9-2b94-4f44-c761-c57b26604cf5"
dcg_results = []
for example in test_examples:
for k in range(len(example)):
dcg_results.append(dcg_score(example, k + 1))
grader.submit_tag('DCGScore', array_to_string(dcg_results))
# + [markdown] id="6mHLh7YwajFf"
# ## First solution: pre-trained embeddings
# + [markdown] id="UtscqfzwajFf"
# We will work with predefined train, validation and test corpora. All the files are tab-separated, but have a different format:
# - *train* corpus contains similar sentences at the same row.
# - *validation* corpus contains the following columns: *question*, *similar question*, *negative example 1*, *negative example 2*, ...
# - *test* corpus contains the following columns: *question*, *example 1*, *example 2*, ...
#
# Validation corpus will be used for the intermediate validation of models. The test data will be necessary for submitting the quality of your model in the system.
# + [markdown] id="p5KGkuZ8ajFf"
# Now you should read *validation* corpus, located at `data/validation.tsv`. You will use it later to evaluate current solution.
# + id="MI2O3H-HajFf"
def read_corpus(filename):
data = []
for line in open(filename, encoding='utf-8'):
data.append(line.strip().split('\t'))
return data
# + id="MJU8n5NrajFf"
validation = read_corpus('data/validation.tsv')
# + id="XEzUBPbVajFf"
from sklearn.metrics.pairwise import cosine_similarity
# + [markdown] id="_wod_6V1ajFf"
# We will use cosine distance to rank candidate questions which you need to implement in the function *rank_candidates*. The function should return a sorted list of pairs *(initial position in candidates list, candidate)*. Index of some pair corresponds to its rank (the first is the best). For example, if the list of candidates was *[a, b, c]* and the most similar is *c*, then *a* and *b*, the function should return a list *[(2, c), (0, a), (1, b)]*.
#
# Pay attention, if you use the function *cosine_similarity* from *sklearn.metrics.pairwise* to calculate similarity because it works in a different way: most similar objects has greatest similarity. It's preferable to use a vectorized version of *cosine_similarity* function. Try to compute similarity at once and not use list comprehension. It should speed up your computations significantly.
# + id="r03wQUhFajFf"
def rank_candidates(question, candidates, embeddings, dim=300):
"""
question: a string
candidates: a list of strings (candidates) which we want to rank
embeddings: some embeddings
dim: dimension of the current embeddings
result: a list of pairs (initial position in the list, question)
"""
######################################
######### YOUR CODE HERE #############
######################################
qv = question_to_vec(question, embeddings, dim)[np.newaxis, :]
cvs = np.array([question_to_vec(candidate, embeddings, dim) for candidate in candidates])
sims = cosine_similarity(qv, cvs)[0]
idxs = np.argsort(sims)[::-1]
return [(i, candidates[i]) for i in idxs]
# + [markdown] id="rTSX-pELajFf"
# Test your code on the tiny examples:
# + id="bnHMmb4ZajFf"
def test_rank_candidates():
questions = ['converting string to list', 'Sending array via Ajax fails']
candidates = [['Convert Google results object (pure js) to Python object',
'C# create cookie from string and send it',
'How to use jQuery AJAX for an outside domain?'],
['Getting all list items of an unordered list in PHP',
'WPF- How to update the changes in list item of a list',
'select2 not displaying search results']]
results = [[(1, 'C# create cookie from string and send it'),
(0, 'Convert Google results object (pure js) to Python object'),
(2, 'How to use jQuery AJAX for an outside domain?')],
[(0, 'Getting all list items of an unordered list in PHP'),
(2, 'select2 not displaying search results'),
(1, 'WPF- How to update the changes in list item of a list')]]
for question, q_candidates, result in zip(questions, candidates, results):
ranks = rank_candidates(question, q_candidates, wv_embeddings, 300)
if not np.all(ranks == result):
return "Check the function."
return "Basic tests are passed."
# + colab={"base_uri": "https://localhost:8080/"} id="LgpbIRKuajFf" outputId="bcc47a04-7327-4639-ffb3-4d2c2824fbc2"
print(test_rank_candidates())
# + [markdown] id="CMzd821majFf"
# Now we can test the quality of the current approach. Run the next two cells to get the results. Pay attention that calculation of similarity between vectors takes time and this calculation is computed approximately in 10 minutes.
# + id="o9g6HJNlajFf"
wv_ranking = []
for line in validation:
q, *ex = line
ranks = rank_candidates(q, ex, wv_embeddings)
wv_ranking.append([r[0] for r in ranks].index(0) + 1)
# + colab={"base_uri": "https://localhost:8080/"} id="6Q4c9Y_oajFf" outputId="f4c793d5-4376-44f2-8cb1-3713fa7bfb01"
for k in [1, 5, 10, 100, 500, 1000]:
print("DCG@%4d: %.3f | Hits@%4d: %.3f" % (k, dcg_score(wv_ranking, k), k, hits_count(wv_ranking, k)))
# + [markdown] id="NQBXY4eRajFf"
# If you did all the steps correctly, you should be frustrated by the received results. Let's try to understand why the quality is so low. First of all, when you work with some data it is necessary to have an idea how the data looks like. Print several questions from the data:
# + colab={"base_uri": "https://localhost:8080/"} id="pLtaJ3JlajFf" outputId="276af284-0e85-4577-ffda-9b8d74e463f0"
for line in validation[:3]:
q, *examples = line
print(q, *examples[:3])
# + [markdown] id="5SDYIh_3ajFf"
# As you can see, we deal with the raw data. It means that we have many punctuation marks, special characters and unlowercased letters. In our case, it could lead to the situation where we can't find some embeddings, e.g. for the word "grid?".
#
# To solve this problem you should use the functions *text_prepare* from the previous assignments to prepare the data.
# + id="CaeRbRsJajFf"
from util import text_prepare
# + [markdown] id="hyPAidl6ajFf"
# Now transform all the questions from the validation set:
# + id="tj36EAsSajFf"
prepared_validation = []
for line in validation:
######### YOUR CODE HERE #############
prepared_validation.append([text_prepare(text) for text in line])
# + [markdown] id="rsfOhDVcajFf"
# Let's evaluate the approach again after the preparation:
# + id="RmRwDSsxajFf"
wv_prepared_ranking = []
for line in prepared_validation:
q, *ex = line
ranks = rank_candidates(q, ex, wv_embeddings)
wv_prepared_ranking.append([r[0] for r in ranks].index(0) + 1)
# + colab={"base_uri": "https://localhost:8080/"} id="w2b58_G4ajFf" outputId="9f2cbb4e-d7d7-4b58-c6dd-78a51a34dbe0"
for k in [1, 5, 10, 100, 500, 1000]:
print("DCG@%4d: %.3f | Hits@%4d: %.3f" % (k, dcg_score(wv_prepared_ranking, k),
k, hits_count(wv_prepared_ranking, k)))
# + [markdown] id="MrlgovOpajFf"
# Now, prepare also train and test data, because you will need it in the future:
# + id="dSRlRsPIajFf"
def prepare_file(in_, out_):
out = open(out_, 'w')
for line in open(in_, encoding='utf8'):
line = line.strip().split('\t')
new_line = [text_prepare(q) for q in line]
print(*new_line, sep='\t', file=out)
out.close()
# + id="joDPDHr4ajFf"
######################################
######### YOUR CODE HERE #############
######################################
prepare_file('data/train.tsv', 'data/train_prepared.tsv')
prepare_file('data/validation.tsv', 'data/validation_prepared.tsv')
prepare_file('data/test.tsv', 'data/test_prepared.tsv')
# + [markdown] id="wS-zapgwajFf"
# **Task 4 (W2VTokenizedRanks).** For each question from prepared *test.tsv* submit the ranks of the candidates to earn the points. The calculations should take about 3-5 minutes. Pay attention that the function *rank_candidates* returns a ranking, while in this case you should find a position in this ranking. Ranks should start with 1.
# + id="FvUcgSz5ajFf"
from util import matrix_to_string
# + colab={"base_uri": "https://localhost:8080/"} id="xEjONDjYajFf" outputId="8f08fdc0-c3a2-4846-8b28-6398a5926295"
w2v_ranks_results = []
prepared_test_data = 'data/test_prepared.tsv'
for line in open(prepared_test_data):
q, *ex = line.strip().split('\t')
ranks = rank_candidates(q, ex, wv_embeddings, 300)
ranked_candidates = [r[0] for r in ranks]
w2v_ranks_results.append([ranked_candidates.index(i) + 1 for i in range(len(ranked_candidates))])
grader.submit_tag('W2VTokenizedRanks', matrix_to_string(w2v_ranks_results))
# + [markdown] id="yufRh7wuajFf"
# ## Advanced solution: StarSpace embeddings
#
# Now you are ready to train your own word embeddings! In particular, you need to train embeddings specially for our task of duplicates detection. Unfortunately, StarSpace cannot be run on Windows and we recommend to use provided
# [docker container](https://github.com/hse-aml/natural-language-processing/blob/master/Docker-tutorial.md) or other alternatives. Don't delete results of this task because you will need it in the final project.
#
# ### How it works and what's the main difference with word2vec?
# The main point in this section is that StarSpace can be trained specifically for some tasks. In contrast to word2vec model, which tries to train similar embeddings for words in similar contexts, StarSpace uses embeddings for the whole sentence (just as a sum of embeddings of words and phrases). Despite the fact that in both cases we get word embeddings as a result of the training, StarSpace embeddings are trained using some supervised data, e.g. a set of similar sentence pairs, and thus they can better suit the task.
#
# In our case, StarSpace should use two types of sentence pairs for training: "positive" and "negative". "Positive" examples are extracted from the train sample (duplicates, high similarity) and the "negative" examples are generated randomly (low similarity assumed).
#
# ### How to choose the best params for the model?
# Normally, you would start with some default choice and then run extensive experiments to compare different strategies. However, we have some recommendations ready for you to save your time:
# - Be careful with choosing the suitable training mode. In this task we want to explore texts similarity which corresponds to *trainMode = 3*.
# - Use adagrad optimization (parameter *adagrad = true*).
# - Set the length of phrase equal to 1 (parameter *ngrams*), because we need embeddings only for words.
# - Don't use a large number of *epochs* (we think that 5 should be enough).
# - Try dimension *dim* equal to 100.
# - To compare embeddings usually *cosine* *similarity* is used.
# - Set *minCount* greater than 1 (for example, 2) if you don't want to get embeddings for extremely rare words.
# - Parameter *verbose = true* could show you the progress of the training process.
# - Set parameter *fileFormat* equals *labelDoc*.
# - Parameter *negSearchLimit* is responsible for a number of negative examples which is used during the training. We think that 10 will be enought for this task.
# - To increase a speed of training we recommend to set *learning rate* to 0.05.
# + [markdown] id="VyuKVOhIajFf"
# Train StarSpace embeddings for unigrams on the train dataset. You don't need to change the format of the input data. Just don't forget to use prepared version of the training data.
#
# If you follow the instruction, the training process will take about 1 hour. The size of the embeddings' dictionary should be approximately 100 000 (number of lines in the result file). If you got significantly more than this number, try to check all the instructions above.
# + colab={"base_uri": "https://localhost:8080/"} id="ZRXbPMyemY5P" outputId="bf7da470-8825-40a1-fd2a-a395e1f6d8e9" language="bash"
# ./Starspace/starspace train \
# -trainFile "data/train_prepared.tsv" \
# -model MODEL_NAME \
# -trainMode 3 \
# + colab={"base_uri": "https://localhost:8080/"} id="Ouek0HAFm2wy" outputId="bb74d85d-428d-4543-9f73-72a43c44f354"
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="42KPz-U8ajFf" outputId="527742a1-7946-410d-aa5a-c3165a99023d"
######### TRAINING HAPPENING HERE #############
# %%bash
./Starspace/starspace train \
-trainFile "data/train_prepared.tsv" \
-model StarSpace_embeddings \
-trainMode 3 \
-adagrad true \
-ngrams 1 \
-epoch 5 \
-dim 100 \
-similarity cosine \
-minCount 2 \
-verbose true \
-fileFormat labelDoc \
-negSearchLimit 10 \
-lr 0.05
# + [markdown] id="VIAGJoDiajFf"
# And now we can compare the new embeddings with the previous ones. You can find trained word vectors in the file *[model_file_name].tsv*. Upload the embeddings from StarSpace into a dict.
# + id="EZk0wu_NajFf"
starspace_embeddings = {}
for line in open('StarSpace_embeddings.tsv', encoding='utf-8'):
word, *vec = line.strip().split('\t')
starspace_embeddings[word] = np.array(vec, dtype=np.float)
# + id="JDjnYUJAajFf"
ss_prepared_ranking = []
for line in prepared_validation:
q, *ex = line
ranks = rank_candidates(q, ex, starspace_embeddings, 100)
ss_prepared_ranking.append([r[0] for r in ranks].index(0) + 1)
# + colab={"base_uri": "https://localhost:8080/"} id="yyJV123RajFf" outputId="1342b3a2-0c1f-485f-fc67-5e91896d1c93"
for k in [1, 5, 10, 100, 500, 1000]:
print("DCG@%4d: %.3f | Hits@%4d: %.3f" % (k, dcg_score(ss_prepared_ranking, k),
k, hits_count(ss_prepared_ranking, k)))
# + [markdown] id="UM2uCcR9ajFf"
# Due to training for the particular task with the supervised data, you should expect to obtain a higher quality than for the previous approach. In additiion, despite the fact that StarSpace's trained vectors have a smaller dimension than word2vec's, it provides better results in this task.
# + [markdown] id="1Foxym-BajFf"
# **Task 5 (StarSpaceRanks).** For each question from prepared *test.tsv* submit the ranks of the candidates for trained representation.
# + colab={"base_uri": "https://localhost:8080/"} id="3dgOJncpajFf" outputId="b074c96b-cfc7-4a81-fd21-f26ce2e112f2"
starspace_ranks_results = []
prepared_test_data = 'data/test_prepared.tsv'
for line in open(prepared_test_data):
q, *ex = line.strip().split('\t')
ranks = rank_candidates(q, ex, starspace_embeddings, 100)
ranked_candidates = [r[0] for r in ranks]
starspace_ranks_results.append([ranked_candidates.index(i) + 1 for i in range(len(ranked_candidates))])
grader.submit_tag('StarSpaceRanks', matrix_to_string(starspace_ranks_results))
# + [markdown] id="sbaqw2qmajFf"
# Please, **don't remove** the file with these embeddings because you will need them in the final project.
# + [markdown] id="UxjmhER1ajFf"
# ### Authorization & Submission
# To submit assignment parts to Cousera platform, please, enter your e-mail and token into variables below. You can generate token on this programming assignment page. <b>Note:</b> Token expires 30 minutes after generation.
# + colab={"base_uri": "https://localhost:8080/"} id="Csj01BPVajFg" outputId="62612cfe-3f51-4f56-eb0d-ff61e64fc564"
STUDENT_EMAIL = '<EMAIL>'
STUDENT_TOKEN = '<PASSWORD>'
grader.status()
# + [markdown] id="NPxwcrq_ajFg"
# If you want to submit these answers, run cell below
# + colab={"base_uri": "https://localhost:8080/"} id="m_skMckOajFg" outputId="b348e6e1-94b2-485e-a082-b0afffce3acd"
grader.submit(STUDENT_EMAIL, STUDENT_TOKEN)
# + id="TcZSmqFAajFg"
| Natural Language Processing/Week3/z-week3-Embeddings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyBaMM development (env)
# language: python
# name: pybamm-dev
# ---
# # Expression trees in PyBaMM
#
# The basic data structure that PyBaMM uses to express models is an expression tree. This data structure encodes a tree representation of a given equation. The expression tree is used to encode the equations of both the original symbolic model, and the discretised equations of that model. Once discretised, the model equations are then passed to the solver, which must then evaluate the discretised expression trees in order to perform the time-stepping.
#
# The expression tree must therefore satisfy three requirements:
# 1. To encode the model equations, it must be able to encode an arbitrary equation, including unary and binary operators such as `*`, `-`, spatial gradients or divergence, symbolic parameters, scalar, matrices and vectors.
# 2. To perform the time-stepping, it must be able to be evaluated, given the current state vector $\mathbf{y}$ and the current time $t$
# 3. For solvers that require it, its gradient with respect to a given variable must be able to be evaluated (once again given $\mathbf{y}$ and $t$)
#
# As an initial example, the code below shows how to construct an expression tree of the equation $2y(1 - y) + t$. We use the `pybamm.StateVector` to represent $\mathbf{y}$, which in this case will be a vector of size 1. The time variable $t$ is already provided by PyBaMM and is of class `pybamm.Time`.
# +
# %pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import numpy as np
y = pybamm.StateVector(slice(0,1))
t = pybamm.t
equation = 2*y * (1 - y) + t
equation.visualise('expression_tree1.png')
# -
# 
#
# Once the equation is constructed, we can evaluate it at a given $t=1$ and $\mathbf{y}=\begin{pmatrix} 2 \end{pmatrix}$.
equation.evaluate(1, np.array([2]))
# We can also calculate the expression tree representing the gradient of the equation with respect to $t$,
diff_wrt_equation = equation.diff(t).simplify()
diff_wrt_equation.visualise('expression_tree2.png')
# 
#
#
# ...and evaluate this expression,
diff_wrt_equation.evaluate(t=1, y=np.array([2]), y_dot=np.array([2]))
# ## The PyBaMM Pipeline
#
# Proposing, parameter setting and discretising a model in PyBaMM is a pipeline process, consisting of the following steps:
#
# 1. The model is proposed, consisting of equations representing the right-hand-side of an ordinary differential equation (ODE), and/or algebraic equations for a differential algebraic equation (DAE), and also associated boundary condition equations
# 2. The parameters present in the model are replaced by actual scalar values from a parameter file, using the [`pybamm.ParamterValues`](https://pybamm.readthedocs.io/en/latest/source/parameters/parameter_values.html) class
# 3. The equations in the model are discretised onto a mesh, any spatial gradients are replaced with linear algebra expressions and the variables of the model are replaced with state vector slices. This is done using the [`pybamm.Discretisation`](https://pybamm.readthedocs.io/en/latest/source/discretisations/discretisation.html) class.
#
# ## Stage 1 - Symbolic Expression Trees
#
# At each stage, the expression tree consists of certain types of nodes. In the first stage, the model is first proposed using [`pybamm.Parameter`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/parameter.html), [`pybamm.Variable`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/variable.html), and other [unary](https://pybamm.readthedocs.io/en/latest/source/expression_tree/unary_operator.html) and [binary](https://pybamm.readthedocs.io/en/latest/source/expression_tree/binary_operator.html) operators (which also includes spatial operators such as [`pybamm.Gradient`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/unary_operator.html#pybamm.Gradient) and [`pybamm.Divergence`](https://pybamm.readthedocs.io/en/latest/source/expression_tree/unary_operator.html#pybamm.Divergence)). For example, the right hand side of the equation
#
# $$\frac{d c}{dt} = D \nabla \cdot \nabla c$$
#
# can be constructed as an expression tree like so:
# +
D = pybamm.Parameter('D')
c = pybamm.Variable('c', domain=['negative electrode'])
dcdt = D * pybamm.div(pybamm.grad(c))
dcdt.visualise('expression_tree3.png')
# -
# 
#
# ## Stage 2 - Setting parameters
#
# In the second stage, the `pybamm.ParameterValues` class is used to replace all the parameter nodes with scalar values, according to an input parameter file. For example, we'll use a this class to set $D = 2$
parameter_values = pybamm.ParameterValues({'D': 2})
dcdt = parameter_values.process_symbol(dcdt)
dcdt.visualise('expression_tree4.png')
# 
#
# ## Stage 3 - Linear Algebra Expression Trees
#
# The third and final stage uses the `pybamm.Discretisation` class to discretise the spatial gradients and variables over a given mesh. After this stage the expression tree will encode a linear algebra expression that can be evaluated given the state vector $\mathbf{y}$ and $t$.
#
# **Note:** for demonstration purposes, we use a dummy discretisation below. For a more complete description of the `pybamm.Discretisation` class, see the example notebook [here](https://github.com/pybamm-team/PyBaMM/blob/master/examples/notebooks/discretisations/finite-volumes.ipynb).
from tests import get_discretisation_for_testing
disc = get_discretisation_for_testing()
disc.y_slices = {c.id: [slice(0, 40)]}
dcdt = disc.process_symbol(dcdt)
dcdt.visualise('expression_tree5.png')
# 
#
# After the third stage, our expression tree is now able to be evaluated by one of the solver classes. Note that we have used a single equation above to illustrate the different types of expression trees in PyBaMM, but any given models will consist of many RHS or algebraic equations, along with boundary conditions. See [here](https://github.com/pybamm-team/PyBaMM/blob/master/examples/notebooks/add-model.ipynb) for more details of PyBaMM models.
# ## References
#
# The relevant papers for this notebook are:
pybamm.print_citations()
| examples/notebooks/expression_tree/expression-tree.ipynb |