code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="o8rERttgz2WZ" endofcell="--"
# 
#
# [](https://colab.research.google.com/github/JohnSnowLabs/nlu/blob/master/examples/webinars_conferences_etc/graph_ai_summit/Healthcare_Graph_NLU_COVID_Tigergraph.ipynb)
#
#
# <div>
# <img src="http://ckl-it.de/wp-content/uploads/2021/04/WhatsApp-Image-2021-04-19-at-7.35.04-AM.jpeg" width="400" height="250" >
# </div>
#
#
#
# # Graph NLU 20 Minutes Crashcourse - State of the Art Text Mining for Graphs
# This notebook is used to feed the `Tiger Graph` engine with features derived by `John Snow Labs` python `NLU` library
# This short notebook will teach you a lot of things!
# - Sentiment classification, binary, multi class and regressive
# - Extract Parts of Speech (POS)
# - Extract Named Entities (NER)
# - Extract Keywords (YAKE!)
# - Answer Open and Closed book questions with T5
# - Summarize text and more with Multi task T5
# - Translate text with Microsofts Marian Model
# - Train a Multi Lingual Classifier for 100+ languages from a dataset with just one language
# - Extract Medical Named Entities (Medical NER)
# - Resolve Medical Entities to codes
# - Classify and extrat relations between entities
# -
#
#
# ## More ressources
# - [Join our Slack](https://join.slack.com/t/spark-nlp/shared_invite/zt-<KEY>)
# - [NLU Website](https://nlu.johnsnowlabs.com/)
# - [NLU Github](https://github.com/JohnSnowLabs/nlu)
# - [Many more NLU example tutorials](https://github.com/JohnSnowLabs/nlu/tree/master/examples)
# - [Overview of every powerful nlu 1-liner](https://nlu.johnsnowlabs.com/docs/en/examples)
# - [Checkout the Modelshub for an overview of all models](https://nlp.johnsnowlabs.com/models)
# - [Checkout the NLU Namespace where you can find every model as a tabel](https://nlu.johnsnowlabs.com/docs/en/spellbook)
# - [Intro to NLU article](https://medium.com/spark-nlp/1-line-of-code-350-nlp-models-with-john-snow-labs-nlu-in-python-2f1c55bba619)
# - [Indepth and easy Sentence Similarity Tutorial, with StackOverflow Questions using BERTology embeddings](https://medium.com/spark-nlp/easy-sentence-similarity-with-bert-sentence-embeddings-using-john-snow-labs-nlu-ea078deb6ebf)
# - [1 line of Python code for BERT, ALBERT, ELMO, ELECTRA, XLNET, GLOVE, Part of Speech with NLU and t-SNE](https://medium.com/spark-nlp/1-line-of-code-for-bert-albert-elmo-electra-xlnet-glove-part-of-speech-with-nlu-and-t-sne-9ebcd5379cd)
# --
# + [markdown] id="GAVkEjc2l_jv"
# # Install NLU and authorize licensed enviroment
# - Run the install script
# - Upload your`spark_nlp_for_healthcare.json`
# - Have fun
#
#
# #### Instructions for non Google Colab enviroment :
# - [See the installation guide](https://nlu.johnsnowlabs.com/docs/en/install) and the [Autorization guide](TODO) for detailed instructions
# - If you need help or run into troubles, [ping us on slack :)](https://join.slack.com/t/spark-nlp/shared_invite/zt-<KEY>)
# + colab={"base_uri": "https://localhost:8080/"} id="fjAIW3p2Lhx4" outputId="3aca4e0e-8f4e-4bf1-e7c4-c3946c1fc2bb" executionInfo={"status": "ok", "timestamp": 1650023785732, "user_tz": -300, "elapsed": 117175, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# !wget https://setup.johnsnowlabs.com/nlu/colab.sh -O - | bash
import nlu
# + [markdown] id="jNw8mtj2jJyS"
# # Simple NLU basics on Strings
# + [markdown] id="cr3HrpqX0Lju"
# ## Context based spell Checking in 1 line
#
# 
# + colab={"base_uri": "https://localhost:8080/", "height": 342} id="yiU5oCWGz31e" outputId="28e94045-eaa6-4b6f-f3b9-4e1df10e3cd7" executionInfo={"status": "ok", "timestamp": 1650023889409, "user_tz": -300, "elapsed": 103697, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
import nlu
nlu.load('spell').predict('I also liek to live dangertus')
# + [markdown] id="EbRSJNNE0XH7"
# ## Binary Sentiment classification in 1 Line
# 
#
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="Kr7JAcnr0Khb" outputId="aa55eafa-08a7-4be9-d939-d1996c8ff3af" executionInfo={"status": "ok", "timestamp": 1650023907540, "user_tz": -300, "elapsed": 18148, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
nlu.load('sentiment').predict('I love NLU and rainy days!')
# + [markdown] id="p0z4S7kF0aeT"
# ## Part of Speech (POS) in 1 line
# 
#
# |Tag |Description | Example|
# |------|------------|------|
# |CC| Coordinating conjunction | This batch of mushroom stew is savory **and** delicious |
# |CD| Cardinal number | Here are **five** coins |
# |DT| Determiner | **The** bunny went home |
# |EX| Existential there | **There** is a storm coming |
# |FW| Foreign word | I'm having a **déjà vu** |
# |IN| Preposition or subordinating conjunction | He is cleverer **than** I am |
# |JJ| Adjective | She wore a **beautiful** dress |
# |JJR| Adjective, comparative | My house is **bigger** than yours |
# |JJS| Adjective, superlative | I am the **shortest** person in my family |
# |LS| List item marker | A number of things need to be considered before starting a business **,** such as premises **,** finance **,** product demand **,** staffing and access to customers |
# |MD| Modal | You **must** stop when the traffic lights turn red |
# |NN| Noun, singular or mass | The **dog** likes to run |
# |NNS| Noun, plural | The **cars** are fast |
# |NNP| Proper noun, singular | I ordered the chair from **Amazon** |
# |NNPS| Proper noun, plural | We visted the **Kennedys** |
# |PDT| Predeterminer | **Both** the children had a toy |
# |POS| Possessive ending | I built the dog'**s** house |
# |PRP| Personal pronoun | **You** need to stop |
# |PRP$| Possessive pronoun | Remember not to judge a book by **its** cover |
# |RB| Adverb | The dog barks **loudly** |
# |RBR| Adverb, comparative | Could you sing more **quietly** please? |
# |RBS| Adverb, superlative | Everyone in the race ran fast, but John ran **the fastest** of all |
# |RP| Particle | He ate **up** all his dinner |
# |SYM| Symbol | What are you doing **?** |
# |TO| to | Please send it back **to** me |
# |UH| Interjection | **Wow!** You look gorgeous |
# |VB| Verb, base form | We **play** soccer |
# |VBD| Verb, past tense | I **worked** at a restaurant |
# |VBG| Verb, gerund or present participle | **Smoking** kills people |
# |VBN| Verb, past participle | She has **done** her homework |
# |VBP| Verb, non-3rd person singular present | You **flit** from place to place |
# |VBZ| Verb, 3rd person singular present | He never **calls** me |
# |WDT| Wh-determiner | The store honored the complaints, **which** were less than 25 days old |
# |WP| Wh-pronoun | **Who** can help me? |
# |WP\$| Possessive wh-pronoun | **Whose** fault is it? |
# |WRB| Wh-adverb | **Where** are you going? |
# + colab={"base_uri": "https://localhost:8080/", "height": 467} id="49-eCSZQ0cQa" outputId="f1611137-f757-4089-92a8-f6b1183e0879" executionInfo={"status": "ok", "timestamp": 1650023926450, "user_tz": -300, "elapsed": 18936, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
nlu.load('pos').predict('POS assigns each token in a sentence a grammatical label')
# + [markdown] id="2-ih6TXc0d6x"
# ## Named Entity Recognition (NER) in 1 line
#
# 
#
# |Type | Description |
# |------|--------------|
# | PERSON | People, including fictional like **<NAME>** |
# | NORP | Nationalities or religious or political groups like the **Germans** |
# | FAC | Buildings, airports, highways, bridges, etc. like **New York Airport** |
# | ORG | Companies, agencies, institutions, etc. like **Microsoft** |
# | GPE | Countries, cities, states. like **Germany** |
# | LOC | Non-GPE locations, mountain ranges, bodies of water. Like the **Sahara desert**|
# | PRODUCT | Objects, vehicles, foods, etc. (Not services.) like **playstation** |
# | EVENT | Named hurricanes, battles, wars, sports events, etc. like **hurricane Katrina**|
# | WORK_OF_ART | Titles of books, songs, etc. Like **Mona Lisa** |
# | LAW | Named documents made into laws. Like : **Declaration of Independence** |
# | LANGUAGE | Any named language. Like **Turkish**|
# | DATE | Absolute or relative dates or periods. Like every second **friday**|
# | TIME | Times smaller than a day. Like **every minute**|
# | PERCENT | Percentage, including ”%“. Like **55%** of workers enjoy their work |
# | MONEY | Monetary values, including unit. Like **50$** for those pants |
# | QUANTITY | Measurements, as of weight or distance. Like this person weights **50kg** |
# | ORDINAL | “first”, “second”, etc. Like David placed **first** in the tournament |
# | CARDINAL | Numerals that do not fall under another type. Like **hundreds** of models are avaiable in NLU |
#
# + colab={"base_uri": "https://localhost:8080/", "height": 340} id="zoDjsSxx0eJY" outputId="08e23666-8ac8-4fa3-a2d5-b3f8b4df1f8c" executionInfo={"status": "ok", "timestamp": 1650023945682, "user_tz": -300, "elapsed": 19254, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
nlu.load('ner').predict("John Snow Labs congratulates the Amarican John Biden to winning the American election!", output_level='chunk')
# + [markdown] id="RcvxNNeXt83u"
# # Let's apply NLU to a COVID dataset!
#
#
# <div>
# <img src="http://ckl-it.de/wp-content/uploads/2021/04/WhatsApp-Image-2021-04-19-at-7.35.04-AM-1.jpeg" width="600" height="450" >
# </div>
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="lKbaoFLMsIRA" outputId="6d85b5ad-0c6c-43ab-814f-2350277d6441" executionInfo={"status": "ok", "timestamp": 1650023949803, "user_tz": -300, "elapsed": 4137, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# ! wget http://ckl-it.de/wp-content/uploads/2021/04/covid19_tweets.csv
import pandas as pd
df = pd.read_csv('covid19_tweets.csv')
df
# + colab={"base_uri": "https://localhost:8080/", "height": 758} id="YWjXqWlFLGOB" outputId="61c12df4-b6c9-4a07-fd99-77f27401a62f" executionInfo={"status": "ok", "timestamp": 1650024077314, "user_tz": -300, "elapsed": 2420, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
df.hashtags = df.hashtags.astype(str)
df["hashtags"][(df["hashtags"] != "nan")].apply(eval).explode().value_counts()[:50].plot.barh(figsize=(20,14), title='Top 50 Hashtag Distribution of COVID dataset')
# + [markdown] id="AMFwC0jX_dCT"
# ## General NER on a COVID News dataset
# ### The **NER** model which you can load via `nlu.load('ner')` recognizes 18 different classes in your dataset.
# We set output level to chunk, so that we get 1 row per NER class.
#
#
# #### Predicted entities:
#
#
# NER is avaiable in many languages, which you can [find in the John Snow Labs Modelshub](https://nlp.johnsnowlabs.com/models)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="aUGAomeusNVv" outputId="f2566bb8-6f86-4aa4-a63e-32120bb113c0" executionInfo={"status": "error", "timestamp": 1650025387012, "user_tz": -300, "elapsed": 1300315, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
ner_df = nlu.load('ner').predict(df, output_level = 'chunk')
ner_df
# + [markdown] id="6gC7S2vqBpT1"
# ### Top 50 Named Entities
# + id="aKSSgTC-sVq8" executionInfo={"status": "aborted", "timestamp": 1650025386505, "user_tz": -300, "elapsed": 64, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
ner_df['entities<EMAIL>'].value_counts()[:100].plot.barh(figsize = (16,20))
# + [markdown] id="B8oAF_MCBxyN"
# ### Top 50 Named Entities which are Countries/Cities/States
# + id="IEiIMsFj9v2n" executionInfo={"status": "aborted", "timestamp": 1650025386511, "user_tz": -300, "elapsed": 68, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
ner_df[ner_df['meta_<EMAIL>'] == 'GPE']['entities@<EMAIL>'].value_counts()[:50].plot.barh(figsize=(18,20), title ='Top 50 Occuring Countries/Cities/States in the dataset')
# + [markdown] id="JiHofegqB0sM"
# ### Top 50 Named Entities which are PRODUCTS
# + id="coeW-Kgs92fH" executionInfo={"status": "aborted", "timestamp": 1650025386513, "user_tz": -300, "elapsed": 70, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
ner_df[ner_df['meta_entities@ner_entity'] == 'PRODUCT']['entities@ner_results'].value_counts()[:50].plot.barh(figsize=(18,20), title ='Top 50 Occuring products in the dataset')
# + id="4ng-TNYu--09" executionInfo={"status": "aborted", "timestamp": 1650025386515, "user_tz": -300, "elapsed": 71, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
ner_df[ner_df['meta_entities@<EMAIL>'] == 'ORG']['entities@ner_results'].value_counts()[:50].plot.barh(figsize=(18,20), title ='Top 50 products occuring in the dataset')
# + [markdown] id="p73T7MyeU2kS"
# ## YAKE on COVID Tweet dataset
# ### The **YAKE!** model (Yet Another Keyword Extractor) is a **unsupervised** keyword extraction algorithm.
# You can load it via which you can load via `nlu.load('yake')`. It has no weights and is very fast.
# It has various parameters that can be configured to influence which keywords are beeing extracted, [here for an more indepth YAKE guide](https://github.com/JohnSnowLabs/nlu/blob/master/examples/webinars_conferences_etc/multi_lingual_webinar/1_NLU_base_features_on_dataset_with_YAKE_Lemma_Stemm_classifiers_NER_.ipynb)
# + id="Zu8-yar9VLqO" executionInfo={"status": "aborted", "timestamp": 1650025386517, "user_tz": -300, "elapsed": 72, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
yake_df = nlu.load('yake').predict(df.text)
yake_df
# + [markdown] id="VJxjIrgdWZ0M"
# ### Top 50 extracted Keywords with YAKE!
# + id="CMXkeiCLVo4u" executionInfo={"status": "aborted", "timestamp": 1650025386520, "user_tz": -300, "elapsed": 75, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
yake_df.explode('keywords_results').keywords_results.value_counts()[:50].plot.barh(title='Keyword Distribution in COVID twitter dataset', figsize = (16,20) )
# + [markdown] id="gXoYzE_RBaaj"
# ## Binary Sentimental Analysis and Distribution on a dataset
# + id="yjHx4TR3_SMe" executionInfo={"status": "aborted", "timestamp": 1650025386523, "user_tz": -300, "elapsed": 78, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
df = pd.read_csv('covid19_tweets.csv').fillna('na').iloc[:5000]
sent_df = nlu.load('sentiment').predict(df)
sent_df
# + id="3LHQb5biErAv" executionInfo={"status": "aborted", "timestamp": 1650025386524, "user_tz": -300, "elapsed": 77, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
sent_df.sentiment_results.value_counts().plot.bar(title='Sentiment distribution')
# + [markdown] id="3a3xxhUSCDhJ"
# ## Emotional Analysis and Distribution of Headlines
# + id="rrYi4f1PEpV3" executionInfo={"status": "aborted", "timestamp": 1650025386524, "user_tz": -300, "elapsed": 77, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
emo_df = nlu.load('emotion').predict(df)
emo_df
# + id="anyAVNjFCG9H" executionInfo={"status": "aborted", "timestamp": 1650025386537, "user_tz": -300, "elapsed": 89, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
emo_df.category_results.value_counts().plot.bar(title='Emotion Distribution')
# + [markdown] id="MHfPEwbA4FrM"
# **Make sure to restart your notebook again** before starting the next section
#
# + id="yZQDoM1p4FS9" executionInfo={"status": "aborted", "timestamp": 1650025386538, "user_tz": -300, "elapsed": 90, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
print("Please restart kernel if you are in google colab and run next cell after the restart to configure java 8 back")
1+'wait'
# + [markdown] id="_kThBMV3LBhg"
# # Let's apply some medical models to the COVID dataset!
#
# + [markdown] id="NcOss-wxREfR"
# ## Medical Named Entitiy recognition
# The medical named entity recognizers are pretrained to extract various `medical` named entities.
# Here are the medical classes it predicts :
#
# ` Kidney_Disease, HDL, Diet, Test, Imaging_Technique, Triglycerides, Obesity, Duration, Weight, Social_History_Header, ImagingTest, Labour_Delivery, Disease_Syndrome_Disorder, Communicable_Disease, Overweight, Units, Smoking, Score, Substance_Quantity, Form, Race_Ethnicity, Modifier, Hyperlipidemia, ImagingFindings, Psychological_Condition, OtherFindings, Cerebrovascular_Disease, Date, Test_Result, VS_Finding, Employment, Death_Entity, Gender, Oncological, Heart_Disease, Medical_Device, Total_Cholesterol, ManualFix, Time, Route, Pulse, Admission_Discharge, RelativeDate, O2_Saturation, Frequency, RelativeTime, Hypertension, Alcohol, Allergen, Fetus_NewBorn, Birth_Entity, Age, Respiration, Medical_History_Header, Oxygen_Therapy, Section_Header, LDL, Treatment, Vital_Signs_Header, Direction, BMI, Pregnancy, Sexually_Active_or_Sexual_Orientation, Symptom, Clinical_Dept, Measurements, Height, Family_History_Header, Substance, Strength, Injury_or_Poisoning, Relationship_Status, Blood_Pressure, Drug, Temperature, EKG_Findings, Diabetes, BodyPart, Vaccine, Procedure, Dosage`
# + id="8nug0OtXLBHf" executionInfo={"status": "aborted", "timestamp": 1650025386540, "user_tz": -300, "elapsed": 92, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
import pandas as pd
import nlu
df = pd.read_csv('covid19_tweets.csv').fillna('na')
vac_names = ['vaccine', 'Pfizer', 'BioNTech','Sinopharm','Sinovac','Moderna','Oxford', 'AstraZeneca','Covaxin','SputnikV']#,
r = "|".join(vac_names)
f_d = df.fillna('na')[df.fillna('na').hashtags.str.contains(r ,flags=re.IGNORECASE, regex=True)]
disease_df = nlu.load('med_ner.jsl.wip.clinical').predict(f_d.text.iloc[:2000], output_level='chunk',drop_irrelevant_cols=False)
disease_df
# + id="3wQZmE9TlwjX" executionInfo={"status": "aborted", "timestamp": 1650025386544, "user_tz": -300, "elapsed": 96, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
disease_df['meta_entities@clinical_entity'].value_counts().plot.barh(figsize=(20,14),title='Distribution of predicted medical entity labels ')
# + [markdown] id="7ILw5508FR6h"
# ### Top 50 **named Entities of type vacine** found in the dataset
#
# The Medical named entity recognizer extracted various entities and classified them as vaccine
# + id="7MLJumSlFQTo" executionInfo={"status": "aborted", "timestamp": 1650025386550, "user_tz": -300, "elapsed": 101, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
disease_df[disease_df['meta_<EMAIL>'] == 'Vaccine']['entities@<EMAIL>'].value_counts()[:50].plot.barh(figsize=(18,20), title ='Top 50 Occuring products in the dataset')
# + [markdown] id="4xEMGtwctN4i"
# ### Top 50 **named entities of type symptom** found in the dataset
#
# The Medical named entity recognizer extracted various entities and classified them as vaccine
# + id="dsWiyY99bzuK" executionInfo={"status": "aborted", "timestamp": 1650025386552, "user_tz": -300, "elapsed": 103, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
disease_df[disease_df['meta_<EMAIL>'] == 'Symptom']['entities@<EMAIL>'].value_counts().plot.barh(figsize=(18,20), title ='Top 50 Occuring smpyoms in the dataset')
# + [markdown] id="n5j2Dw_CvdyG"
# ## Medical Entitiy resolution for `medcical procedures`
#
# Let's mine the text data and extract relevant `procedures` and their `procedure icd 10 pcs codes` to understand what the twitter world is thinking about `treatment procedures` of COVID
#
# For this we are filtering for tweets that conain `vaccine`
# + id="tBdsgVUydCSv" executionInfo={"status": "aborted", "timestamp": 1650025386553, "user_tz": -300, "elapsed": 104, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
vac_names = ['vaccine','Pfizer', 'BioNTech','Sinopharm','Sinovac','Moderna','Oxford', 'AstraZeneca','Covaxin','SputnikV']#,
symptoms = ['sick','cough','sore','fever', 'tired','diarrhoea','taste','smell','loss','rash','skin','breath','short','difficult','ventilator' ]
r = "|".join(symptoms + vac_names)
f_d = df[df.text.str.contains(r ,flags=re.IGNORECASE, regex=True)]
# Extract COVID procedures
resolve_df = nlu.load('med_ner.jsl.wip.clinical resolve.icd10pcs').predict(f_d, output_level='sentence')
resolve_df
# + id="LS9q0j78R_NY" executionInfo={"status": "aborted", "timestamp": 1650025386554, "user_tz": -300, "elapsed": 105, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
resolve_df.sentence_resolution_results.value_counts()[:50]
# + [markdown] id="ZVusxTa3OJ6h"
# ### Top 50 medical proceduress resolved
#
#
#
# + id="MA8AHgZxJyD-" executionInfo={"status": "aborted", "timestamp": 1650025386555, "user_tz": -300, "elapsed": 105, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
resolve_df.sentence_resolution_results.value_counts()[:50].plot.barh(title='Top 50 resolved procedures', figsize=(16,20))
# + [markdown] id="SAz42g2DJd9S"
# ## Resolve Medical Symptoms
# + id="loserzTXQkw1" executionInfo={"status": "aborted", "timestamp": 1650025386556, "user_tz": -300, "elapsed": 106, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
import pandas as pd
# java reset
# ! echo 2 | update-alternatives --config java
# ! java -version
import nlu
import re
df = pd.read_csv('covid19_tweets.csv').fillna('na')
vac_names = ['vaccine','sick','cough','sore','fever','Pfizer', 'BioNTech','Sinopharm','Sinovac','Moderna','Oxford', 'AstraZeneca','Covaxin','SputnikV']
symptoms = ['sick','cough','sore','fever', 'tired','diarrhoea','taste','smell','loss','rash','skin','breath','short','difficult','ventilator' ]
r = "|".join(symptoms+vac_names)
# f_d = df.fillna('na')[df.fillna('na').hashtags.str.contains(r ,flags=re.IGNORECASE, regex=True)]
f_d = df[df.text.str.contains(r ,flags=re.IGNORECASE, regex=True)]
f_d
resolve_df = nlu.load('med_ner.jsl.wip.clinical resolve.icd10cm').predict(f_d[:2000], output_level='sentence')
resolve_df
# + [markdown] id="39sHQNpFBD2v"
# ### Top 50 Medical symptoms
# + id="2StNqCUcAdz8" executionInfo={"status": "aborted", "timestamp": 1650025386558, "user_tz": -300, "elapsed": 108, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
resolve_df.sentence_resolution_results.value_counts()[:50].plot.barh(title='Top 50 resolved procedures', figsize=(16,20))
# + [markdown] id="JJNLjhrjYAFV"
# ## Assert status of medical entities
# + id="t6jKxxEkX_bt" executionInfo={"status": "aborted", "timestamp": 1650025386561, "user_tz": -300, "elapsed": 111, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
df = pd.read_csv('covid19_tweets.csv').fillna('na')
vac_names = ['vaccine','sick','cough','sore','fever','Pfizer', 'BioNTech','Sinopharm','Sinovac','Moderna','Oxford', 'AstraZeneca','Covaxin','SputnikV']
symptoms = ['sick','cough','sore','fever', 'tired','diarrhoea','taste','smell','loss','rash','skin','breath','short','difficult','ventilator' ]
r = "|".join(symptoms+vac_names)
f_d = df[df.text.str.contains(r ,flags=re.IGNORECASE, regex=True)]
f_d
assert_df = nlu.load('med_ner.jsl.wip.clinical assert').predict(f_d[:2000], output_level='chunk')
assert_df
# + [markdown] id="BIMUo8XOXw8i"
# ## Extract relation between entities
# + id="0JmDnL2fYVFr" executionInfo={"status": "aborted", "timestamp": 1650025386563, "user_tz": -300, "elapsed": 112, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
import pandas as pd
import nlu
import re
df = pd.read_csv('covid19_tweets.csv').fillna('na')
vac_names = ['vaccine','sick','cough','sore','fever','Pfizer', 'BioNTech','Sinopharm','Sinovac','Moderna','Oxford', 'AstraZeneca','Covaxin','SputnikV']
symptoms = ['sick','cough','sore','fever', 'tired','diarrhoea','taste','smell','loss','rash','skin','breath','short','difficult','ventilator' ]
r = "|".join(symptoms+vac_names)
f_d = df[df.text.str.contains(r ,flags=re.IGNORECASE, regex=True)]
f_d
relation_df = nlu.load('med_ner.jsl.wip.clinical en.relation.bodypart.problem ').predict(f_d[:2000], output_level='relation')
relation_df
# + [markdown] id="lRA-zGz2D6N_"
# **Make sure to restart your notebook again** before starting the next section
#
# + id="ajINWfeC0jT2" executionInfo={"status": "aborted", "timestamp": 1650025386565, "user_tz": -300, "elapsed": 114, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
print("Please restart kernel if you are in google colab and run next cell after the restart to configure java 8 back")
1+'wait'
# + [markdown] id="rN_H9hKmApll"
# # Answer **Closed Book** and Open **Book Questions** with Google's T5!
#
# <!-- [T5]() -->
# 
#
# You can load the **question answering** model with `nlu.load('en.t5')`
# + id="sKmud8AHN9yo" executionInfo={"status": "aborted", "timestamp": 1650025386566, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Load question answering T5 model
t5_closed_question = nlu.load('en.t5')
# + [markdown] id="-F9rrWbfNyPZ"
# ## Answer **Closed Book Questions**
# Closed book means that no additional context is given and the model must answer the question with the knowledge stored in it's weights
# + id="QsvnphOwfzVQ" executionInfo={"status": "aborted", "timestamp": 1650025386566, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
t5_closed_question.predict("Who is president of Nigeria?")
# + id="DcTbqAGmM6YY" executionInfo={"status": "aborted", "timestamp": 1650025386567, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
t5_closed_question.predict("What is the most common language in India?")
# + id="2Rb4EhK_NAb3" executionInfo={"status": "aborted", "timestamp": 1650025386567, "user_tz": -300, "elapsed": 114, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
t5_closed_question.predict("What is the capital of Germany?")
# + [markdown] id="ogxJNa5MOOQj"
# ## Answer **Open Book Questions**
# These are questions where we give the model some additional context, that is used to answer the question
# + id="e9cwqQGtaTa5" executionInfo={"status": "aborted", "timestamp": 1650025386568, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
t5_open_book = nlu.load('answer_question')
# + id="OB5GOHxPYUYM" executionInfo={"status": "aborted", "timestamp": 1650025386569, "user_tz": -300, "elapsed": 114, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
context = 'Peters last week was terrible! He had an accident and broke his leg while skiing!'
question1 = 'Why was peters week so bad?'
question2 = 'How did peter broke his leg?'
t5_open_book.predict([question1+context, question2 + context])
# + id="kZb_BdGm1-yc" executionInfo={"status": "aborted", "timestamp": 1650025386570, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Ask T5 questions in the context of a News Article
question1 = 'Who is <NAME>?'
question2 = 'Who is founder of Alibaba Group?'
question3 = 'When did <NAME> re-appear?'
question4 = 'How did Alibaba stocks react?'
question5 = 'Whom did <NAME> meet?'
question6 = 'Who did <NAME> hide from?'
# from https://www.bbc.com/news/business-55728338
news_article_context = """ context:
Alibaba Group founder <NAME> has made his first appearance since Chinese regulators cracked down on his business empire.
His absence had fuelled speculation over his whereabouts amid increasing official scrutiny of his businesses.
The billionaire met 100 rural teachers in China via a video meeting on Wednesday, according to local government media.
Alibaba shares surged 5% on Hong Kong's stock exchange on the news.
"""
questions = [
question1+ news_article_context,
question2+ news_article_context,
question3+ news_article_context,
question4+ news_article_context,
question5+ news_article_context,
question6+ news_article_context,]
# + id="e0kTj4ZN4kJi" executionInfo={"status": "aborted", "timestamp": 1650025386571, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
t5_open_book.predict(questions)
# + [markdown] id="xJIuT3ZhOhoc"
# # Multi Problem T5 model for Summarization and more
# The main T5 model was trained for over 20 tasks from the SQUAD/GLUE/SUPERGLUE datasets. See [this notebook](https://github.com/JohnSnowLabs/nlu/blob/master/examples/webinars_conferences_etc/multi_lingual_webinar/7_T5_SQUAD_GLUE_SUPER_GLUE_TASKS.ipynb) for a demo of all tasks
#
#
# # Overview of every task available with T5
# [The T5 model](https://arxiv.org/pdf/1910.10683.pdf) is trained on various datasets for 17 different tasks which fall into 8 categories.
#
#
#
# 1. Text summarization
# 2. Question answering
# 3. Translation
# 4. Sentiment analysis
# 5. Natural Language inference
# 6. Coreference resolution
# 7. Sentence Completion
# 8. Word sense disambiguation
#
# ### Every T5 Task with explanation:
# |Task Name | Explanation |
# |----------|--------------|
# |[1.CoLA](https://nyu-mll.github.io/CoLA/) | Classify if a sentence is gramaticaly correct|
# |[2.RTE](https://dl.acm.org/doi/10.1007/11736790_9) | Classify whether if a statement can be deducted from a sentence|
# |[3.MNLI](https://arxiv.org/abs/1704.05426) | Classify for a hypothesis and premise whether they contradict or contradict each other or neither of both (3 class).|
# |[4.MRPC](https://www.aclweb.org/anthology/I05-5002.pdf) | Classify whether a pair of sentences is a re-phrasing of each other (semantically equivalent)|
# |[5.QNLI](https://arxiv.org/pdf/1804.07461.pdf) | Classify whether the answer to a question can be deducted from an answer candidate.|
# |[6.QQP](https://www.quora.com/q/quoradata/First-Quora-Dataset-Release-Question-Pairs) | Classify whether a pair of questions is a re-phrasing of each other (semantically equivalent)|
# |[7.SST2](https://www.aclweb.org/anthology/D13-1170.pdf) | Classify the sentiment of a sentence as positive or negative|
# |[8.STSB](https://www.aclweb.org/anthology/S17-2001/) | Classify the sentiment of a sentence on a scale from 1 to 5 (21 Sentiment classes)|
# |[9.CB](https://ojs.ub.uni-konstanz.de/sub/index.php/sub/article/view/601) | Classify for a premise and a hypothesis whether they contradict each other or not (binary).|
# |[10.COPA](https://www.aaai.org/ocs/index.php/SSS/SSS11/paper/view/2418/0) | Classify for a question, premise, and 2 choices which choice the correct choice is (binary).|
# |[11.MultiRc](https://www.aclweb.org/anthology/N18-1023.pdf) | Classify for a question, a paragraph of text, and an answer candidate, if the answer is correct (binary),|
# |[12.WiC](https://arxiv.org/abs/1808.09121) | Classify for a pair of sentences and a disambigous word if the word has the same meaning in both sentences.|
# |[13.WSC/DPR](https://www.aaai.org/ocs/index.php/KR/KR12/paper/view/4492/0) | Predict for an ambiguous pronoun in a sentence what it is referring to. |
# |[14.Summarization](https://arxiv.org/abs/1506.03340) | Summarize text into a shorter representation.|
# |[15.SQuAD](https://arxiv.org/abs/1606.05250) | Answer a question for a given context.|
# |[16.WMT1.](https://arxiv.org/abs/1706.03762) | Translate English to German|
# |[17.WMT2.](https://arxiv.org/abs/1706.03762) | Translate English to French|
# |[18.WMT3.](https://arxiv.org/abs/1706.03762) | Translate English to Romanian|
#
#
# + id="XJw187r91QKN" executionInfo={"status": "aborted", "timestamp": 1650025386572, "user_tz": -300, "elapsed": 115, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
# Load the Multi Task Model T5
t5_multi = nlu.load('en.t5.base')
# + id="_F6jE7IN1U-G" executionInfo={"status": "aborted", "timestamp": 1650025386573, "user_tz": -300, "elapsed": 116, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
# https://www.reuters.com/article/instant-article/idCAKBN2AA2WF
text = """(Reuters) - Mastercard Inc said on Wednesday it was planning to offer support for some cryptocurrencies on its network this year, joining a string of big-ticket firms that have pledged similar support.
The credit-card giant’s announcement comes days after <NAME>’s Tesla Inc revealed it had purchased $1.5 billion of bitcoin and would soon accept it as a form of payment.
Asset manager BlackRock Inc and payments companies Square and PayPal have also recently backed cryptocurrencies.
Mastercard already offers customers cards that allow people to transact using their cryptocurrencies, although without going through its network.
"Doing this work will create a lot more possibilities for shoppers and merchants, allowing them to transact in an entirely new form of payment. This change may open merchants up to new customers who are already flocking to digital assets," Mastercard said. (mstr.cd/3tLaPZM)
Mastercard specified that not all cryptocurrencies will be supported on its network, adding that many of the hundreds of digital assets in circulation still need to tighten their compliance measures.
Many cryptocurrencies have struggled to win the trust of mainstream investors and the general public due to their speculative nature and potential for money laundering.
"""
t5_multi['t5'].setTask('summarize ')
short = t5_multi.predict(text)
short
# + id="1MtQlr_8PucN" executionInfo={"status": "aborted", "timestamp": 1650025386573, "user_tz": -300, "elapsed": 115, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
print(f"Original Length {len(short.document.iloc[0])} Summarized Length : {len(short.T5.iloc[0])} \n summarized text :{short.T5.iloc[0]} ")
# + id="ZqOJSkrWQQA9" executionInfo={"status": "aborted", "timestamp": 1650025386574, "user_tz": -300, "elapsed": 116, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
short.T5.iloc[0]
# + [markdown] id="Sd_4hzC9hz8K"
# **Make sure to restart your notebook again** before starting the next section
# + id="RQizVR2WhzTY" executionInfo={"status": "aborted", "timestamp": 1650025386576, "user_tz": -300, "elapsed": 117, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
print("Please restart kernel if you are in google colab and run next cell after the restart to configure java 8 back")
1+'wait'
# + id="41bGg_s0ioKK" executionInfo={"status": "aborted", "timestamp": 1650025386578, "user_tz": -300, "elapsed": 118, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# This configures colab to use Java 8 again.
# You need to run this in Google colab, because after restart it likes to set Java 11 as default, which will cause issues
# ! echo 2 | update-alternatives --config java
# + [markdown] id="PDmjkRoHhrqn"
# # Translate between more than 200 Languages with [ Microsofts Marian Models](https://marian-nmt.github.io/publications/)
#
# Marian is an efficient, free Neural Machine Translation framework mainly being developed by the Microsoft Translator team (646+ pretrained models & pipelines in 192+ languages)
# You need to specify the language your data is in as `start_language` and the language you want to translate to as `target_language`.
# The language references must be [ISO language codes](https://en.wikipedia.org/wiki/List_of_ISO_639-1_codes)
#
# `nlu.load('<start_language>.translate_to.<target_language>')`
#
# **Translate Turkish to English:**
# `nlu.load('tr.translate_to.en')`
#
# **Translate English to French:**
# `nlu.load('en.translate_to.fr')`
#
#
# **Translate French to Hebrew:**
# `nlu.load('fr.translate_to.he')`
#
#
#
#
#
# 
# + id="AjiWgkvQwxBy" executionInfo={"status": "aborted", "timestamp": 1650025386579, "user_tz": -300, "elapsed": 118, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
import nlu
import pandas as pd
# !wget http://ckl-it.de/wp-content/uploads/2020/12/small_btc.csv
df = pd.read_csv('/content/small_btc.csv').iloc[0:20].title
# + [markdown] id="Q_dx5jDkeaGO"
# ## Translate to German
# + id="_DrnIRUlXpM6" executionInfo={"status": "aborted", "timestamp": 1650025386579, "user_tz": -300, "elapsed": 118, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
translate_pipe = nlu.load('en.translate_to.de')
translate_pipe.predict(df)
# + [markdown] id="9zyhBUxFeP6u"
# ## Translate to Chinese
# + id="B0Z3Ilt0eR3c" executionInfo={"status": "aborted", "timestamp": 1650025386580, "user_tz": -300, "elapsed": 119, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
translate_pipe = nlu.load('en.translate_to.zh')
translate_pipe.predict(df)
# + [markdown] id="SbE1KJQgeTiB"
# ## Translate to Hindi
# + id="5U2Xy6JAeXcj" executionInfo={"status": "aborted", "timestamp": 1650025386581, "user_tz": -300, "elapsed": 119, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
translate_pipe = nlu.load('en.translate_to.hi')
translate_pipe.predict(df)
# + [markdown] id="50Ap0BIujDWr"
# # Train a Multi Lingual Classifier for 100+ languages from a dataset with just one language
#
# [Leverage Language-agnostic BERT Sentence Embedding (LABSE) and acheive state of the art!](https://arxiv.org/abs/2007.01852)
#
# Training a classifier with LABSE embeddings enables the knowledge to be transferred to 109 languages!
# With the [SentimentDL model](https://nlp.johnsnowlabs.com/docs/en/annotators#sentimentdl-multi-class-sentiment-analysis-annotator) from Spark NLP you can achieve State Of the Art results on any binary class text classification problem.
#
# ### Languages suppoted by LABSE
# 
#
#
# + id="y4xSRWIhwT28" executionInfo={"status": "aborted", "timestamp": 1650025386582, "user_tz": -300, "elapsed": 120, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Download French twitter Sentiment dataset https://www.kaggle.com/hbaflast/french-twitter-sentiment-analysis
# ! wget http://ckl-it.de/wp-content/uploads/2021/02/french_tweets.csv
import pandas as pd
train_path = '/content/french_tweets.csv'
train_df = pd.read_csv(train_path)
# the text data to use for classification should be in a column named 'text'
columns=['text','y']
train_df = train_df[columns]
train_df = train_df.sample(frac=1).reset_index(drop=True)
train_df
# + [markdown] id="0296Om2C5anY"
# ## Train Deep Learning Classifier using `nlu.load('train.sentiment')`
#
# Al you need is a Pandas Dataframe with a label column named `y` and the column with text data should be named `text`
#
# We are training on a french dataset and can then predict classes correct **in 100+ langauges**
# + id="mptfvHx-MMMX" executionInfo={"status": "aborted", "timestamp": 1650025386583, "user_tz": -300, "elapsed": 121, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Train longer!
trainable_pipe = nlu.load('xx.embed_sentence.labse train.sentiment')
trainable_pipe['sentiment_dl'].setMaxEpochs(60)
trainable_pipe['sentiment_dl'].setLr(0.005)
fitted_pipe = trainable_pipe.fit(train_df.iloc[:2000])
# predict with the trainable pipeline on dataset and get predictions
preds = fitted_pipe.predict(train_df.iloc[:2000],output_level='document')
#sentence detector that is part of the pipe generates sone NaNs. lets drop them first
preds.dropna(inplace=True)
print(classification_report(preds['y'], preds['sentiment']))
preds
# + [markdown] id="lVyOE2wV0fw_"
# ### Test the fitted pipe on new example
# + [markdown] id="RjtuNUcvuJTT"
# #### The Model understands Englsih
# 
# + id="o0vu7PaWkcI7" executionInfo={"status": "aborted", "timestamp": 1650025386584, "user_tz": -300, "elapsed": 122, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
fitted_pipe.predict("This was awful!")
# + id="1ykjRQhCtQ4w" executionInfo={"status": "aborted", "timestamp": 1650025386586, "user_tz": -300, "elapsed": 123, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
fitted_pipe.predict("This was great!")
# + [markdown] id="vohym-XbuNHn"
# #### The Model understands German
# 
# + id="dzaaZrI4tVWc" executionInfo={"status": "aborted", "timestamp": 1650025386586, "user_tz": -300, "elapsed": 123, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# German for:' this movie was great!'
fitted_pipe.predict("Der Film war echt klasse!")
# + id="BbhgTSBGtTtJ" executionInfo={"status": "aborted", "timestamp": 1650025386587, "user_tz": -300, "elapsed": 123, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# German for: 'This movie was really boring'
fitted_pipe.predict("Der Film war echt langweilig!")
# + [markdown] id="a1JbtmWquQwj"
# #### The Model understands Chinese
# 
# + id="kYSYqtoRtc-P" executionInfo={"status": "aborted", "timestamp": 1650025386587, "user_tz": -300, "elapsed": 123, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Chinese for: "This model was awful!"
fitted_pipe.predict("这部电影太糟糕了!")
# + id="06v9SD-QtlBU" executionInfo={"status": "aborted", "timestamp": 1650025386587, "user_tz": -300, "elapsed": 122, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Chine for : "This move was great!"
fitted_pipe.predict("此举很棒!")
# + [markdown] id="9h7CvN4uu9Pb"
# #### Model understanda Afrikaans
#
# 
#
#
# + id="VMPhbgw9twtf" executionInfo={"status": "aborted", "timestamp": 1650025386588, "user_tz": -300, "elapsed": 123, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Afrikaans for 'This movie was amazing!'
fitted_pipe.predict("Hierdie film was ongelooflik!")
# + id="zWgNTIdkumhX" executionInfo={"status": "aborted", "timestamp": 1650025386588, "user_tz": -300, "elapsed": 122, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
# Afrikaans for :'The movie made me fall asleep, it's awful!'
fitted_pipe.predict('Die film het my aan die slaap laat raak, dit is verskriklik!')
# + [markdown] id="rSEPkC-Bwnpg"
# #### The model understands Vietnamese
# 
# + id="wCcTS5gIu511" executionInfo={"status": "aborted", "timestamp": 1650025386589, "user_tz": -300, "elapsed": 123, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
# Vietnamese for : 'The movie was painful to watch'
fitted_pipe.predict('Phim đau điếng người xem')
# + id="M6giDPK-wm2G" executionInfo={"status": "aborted", "timestamp": 1650025386589, "user_tz": -300, "elapsed": 122, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Vietnamese for : 'This was the best movie ever'
fitted_pipe.predict('Đây là bộ phim hay nhất từ trước đến nay')
# + [markdown] id="IlkmAaMoxTuy"
# #### The model understands Japanese
# 
#
# + id="1IfJu3q8wwUt" executionInfo={"status": "aborted", "timestamp": 1650025386590, "user_tz": -300, "elapsed": 122, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Japanese for : 'This is now my favorite movie!'
fitted_pipe.predict('これが私のお気に入りの映画です!')
# + id="h3k7_PFhxOve" executionInfo={"status": "aborted", "timestamp": 1650025386590, "user_tz": -300, "elapsed": 122, "user": {"displayName": "<NAME>", "userId": "02458088882398909889"}}
# Japanese for : 'I would rather kill myself than watch that movie again'
fitted_pipe.predict('その映画をもう一度見るよりも自殺したい')
# + [markdown] id="VXu21c0iQRSC"
# # There are many more models you can put to use in 1 line of code!
# ## Checkout [the Modelshub](https://nlp.johnsnowlabs.com/models) and the [NLU Namespace](https://nlu.johnsnowlabs.com/docs/en/spellbook) for more models
#
#
# ### More ressources
# - [Join our Slack](https://join.slack.com/t/spark-nlp/shared_invite/<KEY>)
# - [NLU Website](https://nlu.johnsnowlabs.com/)
# - [NLU Github](https://github.com/JohnSnowLabs/nlu)
# - [Many more NLU example tutorials](https://github.com/JohnSnowLabs/nlu/tree/master/examples)
# - [Overview of every powerful nlu 1-liner](https://nlu.johnsnowlabs.com/docs/en/examples)
# - [Checkout the Modelshub for an overview of all models](https://nlp.johnsnowlabs.com/models)
# - [Checkout the NLU Namespace where you can find every model as a tabel](https://nlu.johnsnowlabs.com/docs/en/spellbook)
# - [Intro to NLU article](https://medium.com/spark-nlp/1-line-of-code-350-nlp-models-with-john-snow-labs-nlu-in-python-2f1c55bba619)
# - [Indepth and easy Sentence Similarity Tutorial, with StackOverflow Questions using BERTology embeddings](https://medium.com/spark-nlp/easy-sentence-similarity-with-bert-sentence-embeddings-using-john-snow-labs-nlu-ea078deb6ebf)
# - [1 line of Python code for BERT, ALBERT, ELMO, ELECTRA, XLNET, GLOVE, Part of Speech with NLU and t-SNE](https://medium.com/spark-nlp/1-line-of-code-for-bert-albert-elmo-electra-xlnet-glove-part-of-speech-with-nlu-and-t-sne-9ebcd5379cd)
# + id="R8hBZm_zo5EI" executionInfo={"status": "aborted", "timestamp": 1650025386591, "user_tz": -300, "elapsed": 122, "user": {"displayName": "ahmed lone", "userId": "02458088882398909889"}}
while 1 : 1
| nlu/webinars_conferences_etc/graph_ai_summit/Healthcare_Graph_NLU_COVID_Tigergraph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # part 1: chunked datasets of 300
# + pycharm={"name": "#%%\n"}
import json
# + pycharm={"name": "#%%\n"}
import csv
# + pycharm={"name": "#%%\n"}
from pandas import *
# + pycharm={"name": "#%%\n"}
data = read_csv("master_list_of_comments.csv")
# + pycharm={"name": "#%%\n"}
all_comments = data['comment'].to_dict()
# + pycharm={"name": "#%%\n"}
count = 0
for idx in all_comments:
try:
all_comments[idx] = all_comments[idx].strip()
except:
print(f"{idx}: {all_comments[idx]}")
if count <= 100:
print(f"{idx}: {all_comments[idx]}")
count += 1
# + pycharm={"name": "#%%\n"}
all_comments = {value: key for key, value in all_comments.items()}
# + pycharm={"name": "#%%\n"}
all_comments
# + pycharm={"name": "#%%\n"}
f = open('labeled_dataset_-1.json')
labeled_comments = json.load(f)
# + pycharm={"name": "#%%\n"}
for comment in labeled_comments:
comment_text_stripped = comment['text'].strip()
if not comment_text_stripped in all_comments:
print(comment['text'], "not in dataset")
raise Exception("Comment not found")
for comment in labeled_comments:
comment_text_stripped = comment['text'].strip()
if comment_text_stripped in all_comments:
del all_comments[comment_text_stripped]
# + pycharm={"name": "#%%\n"}
remaining_comments = []
for comment in all_comments:
remaining_comments.append({'idx': all_comments[comment], 'text': comment})
# + pycharm={"name": "#%%\n"}
len(remaining_comments)
# + pycharm={"name": "#%%\n"}
def chunks(lst, n):
"""Yield successive n-sized chunks from lst."""
for i in range(0, len(lst), n):
yield lst[i:i + n]
# + pycharm={"name": "#%%\n"}
chunked_datasets = list(chunks(remaining_comments, 300))
# + pycharm={"name": "#%%\n"}
for i, dataset in enumerate(chunked_datasets):
keys = dataset[0].keys()
a_file = open(f"chunked_datasets/dataset_{i}.csv", "w")
dict_writer = csv.DictWriter(a_file, keys)
dict_writer.writeheader()
dict_writer.writerows(dataset)
a_file.close()
# -
# # part 2: randomized chunked datasets of 25
# + pycharm={"name": "#%%\n"}
for comment in remaining_comments:
comment['query'] = data['query'].iloc[comment['idx']]
# + pycharm={"name": "#%%\n"}
import random
# + pycharm={"name": "#%%\n"}
random.shuffle(remaining_comments)
# + pycharm={"name": "#%%\n"}
desired_order_list = ["idx", "query", "text"]
new_remaining_comments = []
for comment in remaining_comments:
new_remaining_comments.append({k: comment[k] for k in desired_order_list})
# + pycharm={"name": "#%%\n"}
remaining_comments = new_remaining_comments
# + pycharm={"name": "#%%\n"}
chunked_datasets = list(chunks(remaining_comments, 25))
# + pycharm={"name": "#%%\n"}
for i, dataset in enumerate(chunked_datasets):
keys = dataset[0].keys()
a_file = open(f"randomized_chunked_datasets_of_25/dataset_{i}.csv", "w")
dict_writer = csv.DictWriter(a_file, keys)
dict_writer.writeheader()
dict_writer.writerows(dataset)
a_file.close()
# + pycharm={"name": "#%%\n"}
raw_chunked_datasets = [[] for _ in range(len(chunked_datasets))]
for i, dataset in enumerate(chunked_datasets):
for comment in dataset:
raw_chunked_datasets[i].append(comment['text'])
# + pycharm={"name": "#%%\n"}
raw_chunked_datasets[0:5]
# + pycharm={"name": "#%%\n"}
for i, dataset in enumerate(raw_chunked_datasets):
with open(f'randomized_chunked_datasets_of_25/dataset_{i}_raw.csv', 'w') as f:
writer = csv.writer(f)
for val in dataset:
writer.writerow([val])
| MateoStuff/Generate300Dataset/remove_labeled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Purpose
#
# This figure justifies our choice of critic features. It should show that using both counts
# is better than using either or none.
#
# ### Method
#
# I did 3 very large runs. Pretraining for 200 for each, and then training against
# critic with one, other, and both critic-counts missing.
#
# We'll show the grahs over epochs. We'll also show the training critic-loss, to see how well it can fit that curve.
#
# ### Data structure
#
# Each of the files is a JSON object. Its keys are the 3 training phases: ACTOR, CRITIC, and AC.
# Underneath this, it has subfields for what it was recording. For example, NDCG, or TRAINING_CRITIC_ERROR.
#
# ### Conclusions
#
# It seems as though BOTH_COUNTS is BEST, NO_COUNTS is WORST, WITHOUT_SEEN is pretty good, and WITHOUT_UNSEEN is pretty bad.
#
# +
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import json
import seaborn as sns
sns.set()
DATA = {}
DATA['without_either'] = {}
DATA['without_seen'] = {}
DATA['without_unseen'] = {}
DATA['tentative_with_both'] = {}
print("Now, loading data")
with open("./data/without_either/actor_training.json", "r") as f:
DATA['without_either']['actor'] = json.loads(f.read())
with open("./data/without_either/critic_training.json", "r") as f:
DATA['without_either']['critic'] = json.loads(f.read())
with open("./data/without_seen/actor_training.json", "r") as f:
DATA['without_seen']['actor'] = json.loads(f.read())
with open("./data/without_seen/critic_training.json", "r") as f:
DATA['without_seen']['critic'] = json.loads(f.read())
with open("./data/without_unseen/actor_training.json", "r") as f:
DATA['without_unseen']['actor'] = json.loads(f.read())
with open("./data/without_unseen/critic_training.json", "r") as f:
DATA['without_unseen']['critic'] = json.loads(f.read())
with open("./data/tentative_with_both/actor_training.json", "r") as f:
DATA['tentative_with_both']['actor'] = json.loads(f.read())
with open("./data/tentative_with_both/critic_training.json", "r") as f:
DATA['tentative_with_both']['critic'] = json.loads(f.read())
print("Data loaded")
print("I'm calling it tentatitive_with_both, because ")
# +
print("First, we'll plot just the actors")
# https://stackoverflow.com/questions/20618804/how-to-smooth-a-curve-in-the-right-way
plt.clf()
plt.plot(range(200), DATA['without_either']['actor']['ACTOR']['ndcg'], color="red")
plt.plot(range(200), DATA['without_seen']['actor']['ACTOR']['ndcg'], color="blue")
plt.plot(range(200), DATA['without_unseen']['actor']['ACTOR']['ndcg'], color="green")
plt.plot(range(200), DATA['tentative_with_both']['actor']['ACTOR']['ndcg'], color="orange")
plt.show()
plt.ylim(0.42, 0.44)
# +
actor_parts = np.asarray([
DATA[part]['actor']['ACTOR']['ndcg']
for part in
['without_either', 'without_seen', 'without_unseen', 'tentative_with_both']
])
average_actor_parts = np.mean(actor_parts, axis=0).tolist()
# +
# Now,I want to plot with everything...
all_without_either = DATA['without_either']['actor']['ACTOR']['ndcg'] + DATA['without_either']['critic']['AC']['ndcg']
all_without_seen = DATA['without_seen']['actor']['ACTOR']['ndcg'] + DATA['without_seen']['critic']['AC']['ndcg']
all_without_unseen = DATA['without_unseen']['actor']['ACTOR']['ndcg'] + DATA['without_unseen']['critic']['AC']['ndcg']
all_tentative_with_both = DATA['tentative_with_both']['actor']['ACTOR']['ndcg'] + DATA['tentative_with_both']['critic']['AC']['ndcg']
# I ran this with 50 more training steps than the others.
plt.clf()
# plt.plot(range(350), all_without_either, color="red", label="No Counts")
# plt.plot(range(350), all_without_seen, color="blue", label="Only Seen Count")
# plt.plot(range(350), all_without_unseen, color="green", label="Only Unseen Count")
# plt.plot(range(300), all_tentative_with_both, color='orange', label="Both Counts")
plt.plot(range(300), all_without_either[0:300], color="red", label="No Counts")
plt.plot(range(300), all_without_seen[0:300], color="blue", label="Without Seen Count")
plt.plot(range(300), all_without_unseen[0:300], color="green", label="Without Unseen Count")
plt.plot(range(300), all_tentative_with_both[0:300], color='orange', label="Both Counts")
leg = plt.legend(fontsize=12, shadow=True, loc=(0.05, 0.60))
plt.ylim(0.42, 0.45)
plt.show()
print(max(all_tentative_with_both))
print(max(all_without_seen))
print(max(all_without_unseen))
print("Good prevails! without_seen is pretty good, but not as good as with both.")
print("NOTE: I'm only showing 100 epochs of AC-training, because we want ")
# +
plt.clf()
plt.plot(range(200), average_actor_parts, color="black", label="shared_actor")
plt.plot(range(199, 300), [average_actor_parts[-1]] + DATA['without_either']['critic']['AC']['ndcg'][:100], color="red", label="No Counts")
plt.plot(range(199, 300), [average_actor_parts[-1]] + DATA['without_seen']['critic']['AC']['ndcg'][:100], color="blue", label="Without Seen Count")
plt.plot(range(199, 300), [average_actor_parts[-1]] + DATA['without_unseen']['critic']['AC']['ndcg'][:100], color="green", label="Without Unseen Count")
plt.plot(range(199, 300), [average_actor_parts[-1]] + DATA['tentative_with_both']['critic']['AC']['ndcg'], color='orange', label="Both Counts")
leg = plt.legend(fontsize=12, shadow=True, loc=(0.05, 0.60))
plt.ylim(0.42, 0.45)
print(max(all_tentative_with_both))
print(max(all_without_seen))
print(max(all_without_unseen))
print("Good prevails! without_seen is pretty good, but not as good as with both.")
print("NOTE: I'm only showing 100 epochs of AC-training, because we want ")
# +
fig=plt.figure(figsize=(5.4,3.5))
plt.axvline(x=149, linewidth=1, color='k', linestyle="--")
plt.plot(range(200), average_actor_parts, linewidth=1.5, linestyle="-", color="black", label="VAE")
plt.plot(range(149, 250), [average_actor_parts[150]] + DATA['tentative_with_both']['critic']['AC']['ndcg'],linewidth=2.5, color='red', label="$[\mathcal{L}_{E} , | \mathcal{H}_0 |, |\mathcal{H}_1 |]$")
plt.plot(range(149, 250), [average_actor_parts[150]] + DATA['without_seen']['critic']['AC']['ndcg'][:100], linewidth=2.5, color="deepskyblue", label="$[\mathcal{L}_{E} , | \mathcal{H}_0 |]$")
plt.plot(range(149, 250), [average_actor_parts[150]] + DATA['without_unseen']['critic']['AC']['ndcg'][:100], linewidth=2.5, color="green", label="$[\mathcal{L}_{E} , |\mathcal{H}_1 |]$")
plt.plot(range(149, 250), [average_actor_parts[150]] + DATA['without_either']['critic']['AC']['ndcg'][:100], linewidth=2.5, color="orange", label="$[\mathcal{L}_{E}]$")
leg = plt.legend(fontsize=13, shadow=True, loc=(0.002, 0.45))
plt.grid('on')
plt.xlabel('# Epoch of actor', fontsize=13)
plt.ylabel('Validation NDCG@100', fontsize=13)
plt.xlim(100, 200)
plt.ylim(0.428, 0.44)
plt.show()
fig.savefig('plot_feature_ablation.pdf', bbox_inches='tight')
print(max(all_tentative_with_both))
print(max(all_without_seen))
print(max(all_without_unseen))
print("Good prevails! without_seen is pretty good, but not as good as with both.")
print("NOTE: I'm only showing 100 epochs of AC-training, because we want ")
# -
| paper_plots/critic_term_ablation_study/plot_which_feature.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#all_slow
# -
# # Tutorial - Text Generation
# > Using the text generation API with AdaptNLP
# ## What is Text Generation?
#
# Text generation is the NLP task of generating a coherent sequence of words, usually from a language model. The current leading methods, most notably OpenAI’s GPT-2 and GPT-3, rely on feeding tokens (words or characters) into a pre-trained language model which then uses this seed data to construct a sequence of text. AdaptNLP provides simple methods to easily fine-tune these state-of-the-art models and generate text for any use case.
#
# Below, we'll walk through how we can use AdaptNLP's `EasyTextGenerator` module to generate text to complete a given String.
# ## Getting Started with `TextGeneration`
#
# We'll get started by importing the `EasyTextGenerator` class from AdaptNLP.
from adaptnlp import EasyTextGenerator
# Then we'll write some sample text to use:
text = "China and the U.S. will begin to"
# And finally instantiating our `EasyTextGenerator`:
generator = EasyTextGenerator()
# ## Generating Text
# Now that we have the summarizer instantiated, we are ready to load in a model and compress the text
# with the built-in `generate()` method.
#
# Here is one example using the gpt2 model:
# +
generated_text = generator.generate(text, model_name_or_path="gpt2", mini_batch_size=2, num_tokens_to_produce=50)
print(generated_text)
# -
# ## Finding Models with the `HFModelHub`
#
# Rather than searching through HuggingFace for models to use, we can use Adapt's `HFModelHub` to search for valid text generation models.
#
# First, let's import it:
from adaptnlp import HFModelHub
# And then search for some models by task:
hub = HFModelHub()
models = hub.search_model_by_task('text-generation'); models
# We'll use our `gpt2` model again:
model = models[4]; model
# And pass it into our `generator`:
# +
generated_text = generator.generate(text, model_name_or_path=model, mini_batch_size=2, num_tokens_to_produce=50)
print(generated_text)
| nbs/09a_tutorial.easy_text_generator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Select protfolio-id for trades analysis
portfolio_id = "31cb73e1-5c30-4cff-9f23-2a51cce7ca6f"
end_date = '2021-09-14'
# #### imports
# +
import asyncio
import json
import math
import sys
import datetime
from datetime import date, datetime, timedelta
import iso8601
import matplotlib.pyplot as plt
import nest_asyncio
import numpy as np
import pandas as pd
import pytz
import requests
from dateutil import parser
from IPython.display import HTML, display, Markdown
from liualgotrader.analytics.analysis import (
calc_batch_revenue,
count_trades,
load_trades_by_portfolio,
trades_analysis,
symbol_trade_analytics,
)
from liualgotrader.common import config
from liualgotrader.common.types import DataConnectorType
from pandas import DataFrame as df
from pytz import timezone
from talib import BBANDS, MACD, RSI
from liualgotrader.common.data_loader import DataLoader
# %matplotlib inline
nest_asyncio.apply()
# -
config.data_connector = DataConnectorType.alpaca
end_date = datetime.strptime(end_date, "%Y-%m-%d")
local = pytz.timezone("UTC")
end_date = local.localize(end_date, is_dst=None)
# #### Load batch data
trades = load_trades_by_portfolio(portfolio_id)
trades
if trades.empty:
assert False, "Empty batch. halting execution."
# ## Display trades in details
minute_history = DataLoader(connector=DataConnectorType.alpaca)
nyc = timezone("America/New_York")
for symbol in trades.symbol.unique():
symbol_df = trades.loc[trades["symbol"] == symbol]
start_date = symbol_df["client_time"].min()
start_date = start_date.replace(hour=9, minute=30, second=0, tzinfo=nyc)
end_date = end_date.replace(hour=16, minute=0, second=0, tzinfo=nyc)
cool_down_date = start_date + timedelta(minutes=5)
try:
symbol_data = minute_history[symbol][start_date:end_date]
except Exception:
print(f"failed loading {symbol} from {start_date} to {end_date} -> skipping!")
continue
minute_history_index = symbol_data.close.index.get_loc(
start_date, method="nearest"
)
end_index = symbol_data.close.index.get_loc(
end_date, method="nearest"
)
cool_minute_history_index = symbol_data["close"].index.get_loc(
cool_down_date, method="nearest"
)
open_price = symbol_data.close[cool_minute_history_index]
plt.plot(
symbol_data.close[minute_history_index:end_index].between_time(
"9:30", "16:00"
),
label=symbol,
)
plt.xticks(rotation=45)
d, profit = symbol_trade_analytics(symbol_df, open_price, plt)
print(f"{symbol} analysis with profit {round(profit, 2)}")
display(HTML(pd.DataFrame(data=d).to_html()))
plt.legend()
plt.show()
| analysis/notebooks/portfolio_trades.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from __future__ import print_function
# %matplotlib inline
# %config InlineBackend.figure_format = 'png'
# -
# %%time
df = pd.read_csv("../data/train.csv")
# 4min
df.columns
# %%time
df.info()
# %%time
# histogram of features
skip_cols = ['date_time','orig_destination_distance', 'user_id']
for col in df.columns:
if col in skip_cols:
continue
print('='*50)
print('col : ', col)
f, ax = plt.subplots(figsize=(20, 15))
sns.countplot(x=col, data=df, alpha=0.5)
plt.show()
# +
# site_name : 2가 과도하게 많음 -> 나머지 3,4같은 친구들의 빈도수가 어떻게 될까
# posa_contient : 3이 2.5이상 많음
# user_location_country : 특정 위치가 많음(관광을 많이 가는 나라인가..) / 지금 여기엔 작게 보이는 것들도 아예 없는거에 비해 많은것
# user_location_region : 마찬가지로 특정 지역에서 expedia 검색을 많이 함
# user_location_city : 뾰족한 뾰족한 느낌이 온다
# is_mobile : 0이 압도적으로 많음
# is_package : 0이 압도적으로 많음
# 채널 9 >0 > 1 > 2 > 5 > 3 > 4 ....
# srch_ci : 일정 주기마다 데이터의 양이 반복됨. 이 시기에 대한 무언가가 있지 않을까
# srch_co : check out - check in 몇일 여행을 하는지 일수 feature 추가
# adult : 거의 2명 아니면 혼자 -> 예약하는 인원 수에 따른 무언가가 있지 않을까
# 자녀는 없는 경우가 많고 많으면 1,2
# rm_cnt : 방은 거의 1개 -> 이건 역으로 호텔의 분류를 내가 파악해볼 수 있을 듯
# destination id - 뾰족한 친구가 있음. 음
# destination_type : 1과 6..?
# is_booking : 0> 1
# cnt : 5번 안에모든 것이 결정된다고 보면 될 것 같은데.. 정확한 비율을 찾아보자
# 호텔이 있는 대륙은 2가 많네
# 호텔 마켓..
# 호텔의 클러스터는 0부터 99까지 100가지가 있음 => 예약하냐 안하냐에 과연 중요한 변수일까
# user_id 사람들의 평균 예약 빈도수를 파악해서 평균 이상인 사람과 평균 이하인 사람으로 구분
# +
# 내가 여행을 간다면 일단 expedia에서 검색을 해봄 -> 인원 수, 장소, 시간 체크 후 검색
# -> 검색 결과를 훑어봄 (아마 평점이 높은 친구부터, 그리고 expedia가 추천한 것부터 위에서 아래로 봄)
# -> 친구 혹은 같이 가는 사람들에게 링크 전달
# -> 의논 후 호텔 결정
# -> 항공권과 함께하면 할인되므로 또 고민
# +
# y : 2015년 booking 0 / 1
# data를 나누어 모델을 만든다면
# 1) 최근 1년(14년) 데이터 + 빈도가 낮은 호텔 제외
# 2) 여행을 자주 가는 시즌이 있을테니 이런 경우 평준화
# 3) 여행 때문이 아닌 출장을 하는 경우에도 expedia를 사용할 수 있음 -> 별도로 나누어서 빼기
#
# expeida에서 설정한 호텔 cluster
# 데이터 전처리
# 일단 datetime -> 일과 월로 나누어서 표기(월별/일별 주말, 평일, click 시간이 오전이냐 오후냐 새벽이냐)
# null값이 있는 distance 체크
# 타 feature에 na값이 있는지 확인하기
# +
# %%time
# date_time => datetime 형식으로 변경
df["date_time"] = pd.to_datetime(df["date_time"], coerce=True)
# -
# %%time
df_2013 = df[df["date_time"].dt.year == 2013]
# %%time
df_2013.to_csv("train_2013.csv")
# %%time
df_2014 = df[df["date_time"].dt.year == 2014]
df_2014.to_csv("train_2014.csv")
df["date_time"].head().dt.year == 2014
pd.set_option("max_columns", 40)
print(pd.get_option("display.max_columns"))
| notebook/01. EDA(all data).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import tensorflow as tf
from datetime import datetime
import matplotlib.pyplot as plt
import seaborn as sns
features = pd.read_csv('../Data/training_set_features.csv')
features.describe()
((features.isnull().sum() / len(features)) * 100).sort_values()
# +
categorical_columns = [
'sex',
'hhs_geo_region',
'census_msa',
'race',
'age_group',
'behavioral_face_mask',
'behavioral_wash_hands',
'behavioral_antiviral_meds',
'behavioral_outside_home',
'behavioral_large_gatherings',
'behavioral_touch_face',
'behavioral_avoidance',
'health_worker',
'child_under_6_months',
'chronic_med_condition',
'education',
'marital_status',
'employment_status',
'rent_or_own',
'doctor_recc_h1n1',
'doctor_recc_seasonal',
'income_poverty'
]
numerical_columns = [
'household_children',
'household_adults',
'h1n1_concern',
'h1n1_knowledge',
'opinion_h1n1_risk',
'opinion_h1n1_vacc_effective',
'opinion_h1n1_sick_from_vacc',
'opinion_seas_vacc_effective',
'opinion_seas_risk',
'opinion_seas_sick_from_vacc',
]
# -
features[categorical_columns]
# +
n_bins = 20
fig, axs = plt.subplots(1,2, sharey=True, tight_layout=True)
axs[0].hist('race', bins=n_bins)
# -
fig, axes = plt.subplots(2,3, sharey=True)
sub_num = 0
for num_column in numerical_columns:
ax = sns.distplot(features[num_column], kde=True, bins=4, ax=axes[sub_num])
sub_num += 1
for x in numerical_columns:
plt.figure()
plt.hist(features[x])
plt.xlabel(x)
| Notebooks/.ipynb_checkpoints/H1N1 Exploration Notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nova Tarefa - Implantação
#
# Preencha aqui com detalhes sobre a tarefa.<br>
# ### **Em caso de dúvidas, consulte os [tutoriais da PlatIAgro](https://platiagro.github.io/tutorials/).**
# ## Declaração de Classe para Predições em Tempo Real
#
# A tarefa de implantação cria um serviço REST para predições em tempo-real.<br>
# Para isso você deve criar uma classe `Model` que implementa o método `predict`.
# +
# %%writefile Model.py
# adicione imports aqui...
class Model:
def __init__(self):
# adicione seu código aqui...
pass
def predict(self, X, feature_names, meta=None):
# adicione seu código aqui...
return X
| projects/config/Deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="599c85b07f91bac2ad212af1ea3b4edfc66d2bff"
import pandas as pd
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
from xgboost import XGBRegressor
from learntools.core import *
iowa_file_path = '../input/train.csv'
iowa_test_file_path = '../input/test.csv'
train_data = pd.read_csv(iowa_file_path)
test_data = pd.read_csv(iowa_test_file_path)
y = train_data.SalePrice
train_features = train_data.drop(['SalePrice'], axis = 1)
# + [markdown] _uuid="1cf1ffebcb4a95f15112e2e7750f526baa39fa30"
# # Missing and categorical values
# + _uuid="9760b349094ff1d874615efe613d8cd652be18f0"
# fill in missing numeric values
from sklearn.impute import SimpleImputer
# impute
train_data_num = train_features.select_dtypes(exclude=['object'])
test_data_num = test_data.select_dtypes(exclude=['object'])
imputer = SimpleImputer()
train_num_cleaned = imputer.fit_transform(train_data_num)
test_num_cleaned = imputer.transform(test_data_num)
# columns rename after imputing
train_num_cleaned = pd.DataFrame(train_num_cleaned)
test_num_cleaned = pd.DataFrame(test_num_cleaned)
train_num_cleaned.columns = train_data_num.columns
test_num_cleaned.columns = test_data_num.columns
# + _uuid="4898e1945a5208161a6fcf0e23571bcd06d811b0"
# string columns: transform to dummies
train_data_str = train_data.select_dtypes(include=['object'])
test_data_str = test_data.select_dtypes(include=['object'])
train_str_dummy = pd.get_dummies(train_data_str)
test_str_dummy = pd.get_dummies(test_data_str)
train_dummy, test_dummy = train_str_dummy.align(test_str_dummy,
join = 'left',
axis = 1)
# + _uuid="5af3d1c3cdfe4206bef38e7b1c8ce497dfae9e52"
# convert numpy dummy tables to pandas DataFrame
train_num_cleaned = pd.DataFrame(train_num_cleaned)
test_num_cleaned = pd.DataFrame(test_num_cleaned)
# + _uuid="f1e90fe32706f43ac1d5ea7af242b999703945c2"
# joining numeric (after imputing) and string (converted to dummy) data
train_all_clean = pd.concat([train_num_cleaned, train_dummy], axis = 1)
test_all_clean = pd.concat([test_num_cleaned, test_dummy], axis = 1)
# + _uuid="e9d746b010eb0fdc602de346056144768bb83705"
# detect NaN in already cleaned test data
# (there could be completely empty columns in test data)
cols_with_missing = [col for col in test_all_clean.columns
if test_all_clean[col].isnull().any()]
for col in cols_with_missing:
print(col, test_all_clean[col].isnull().any())
# + _uuid="0a271653aa668a52844a4c56f3195b681fa76d1d"
# since there are empty columns in test we need to drop them in train and test data
train_all_clean_no_nan = train_all_clean.drop(cols_with_missing, axis = 1)
test_all_clean_no_nan = test_all_clean.drop(cols_with_missing, axis = 1)
# + [markdown] _uuid="c3221390f01274b7f875e87abbac46dda8f0c04c"
# # Different models training and validation
# + _uuid="015f66e0c1655124079a8d88d1e00a5399f5e543"
train_X, val_X, train_y, val_y = train_test_split(train_all_clean_no_nan, y, random_state=1)
# default XGBoost
xgb_model = XGBRegressor(random_state = 1)
xgb_model.fit(train_X, train_y, verbose = False)
xgb_predictions = xgb_model.predict(val_X)
xgb_mae = mean_absolute_error(val_y, xgb_predictions)
print("XGBoost MAE default: {:,.0f}".format(xgb_mae))
# fine tuned XGBoost
xgb_model = XGBRegressor(n_estimators = 1000, learning_rate=0.05, random_state = 1)
xgb_model.fit(train_X, train_y, early_stopping_rounds = 25, eval_set = [(val_X, val_y)], verbose = False)
# with verbose = True we have the best iteration on step 757 => n_estimators = 757
# + _uuid="cf6b8944b3c6092e3a544087b2c7caf500ee3b3c"
xgb_predictions = xgb_model.predict(val_X)
xgb_mae = mean_absolute_error(val_y, xgb_predictions)
print("XGBoost MAE default: {:,.0f}".format(xgb_mae))
# + [markdown] _uuid="f7c47011756dd4125ac584cbac751ae5e0d8509f"
# # XGBoost Model (on all training data)
# + _uuid="c7b21de27dd3cc2d79ba9c361789d30a0a8f64ee"
# To improve accuracy, create a new Random Forest model which you will train on all training data
xgb_model_on_full_data = xgb_model = XGBRegressor(n_estimators = 757, learning_rate=0.05)
xgb_model_on_full_data.fit(train_all_clean_no_nan, y)
# + [markdown] _uuid="294a788b8c675e0e42a16d60b41a9846d99f7ae8"
# # Make Predictions
# + _uuid="46cd68a096beb5b969ef7e663f89f4bb22a7b7ad"
test_X = test_all_clean_no_nan
test_preds = xgb_model_on_full_data.predict(test_X)
# + _uuid="e3d6c83e2b612cd6a085c37e504f7e8125eb1fbc"
output = pd.DataFrame({'Id': test_data.Id,
'SalePrice': test_preds})
output.to_csv('submission.csv', index=False)
| machine learning/housing-xgboost-tuned-on-cleaned-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="A_5C1IvdfxkF"
# #pyhpc-benchmarks @ Google Colab
#
# To run all benchmarks, you need to switch the runtime type to match the corresponding section (CPU, TPU, GPU).
# + id="TTViNK-9OfRJ"
# !rm -rf pyhpc-benchmarks; git clone https://github.com/dionhaefner/pyhpc-benchmarks.git
# + id="Eyc45XkjQB1X"
# %cd pyhpc-benchmarks
# + id="RbM7XH04MwFA"
# check CPU model
# !lscpu |grep 'Model name'
# + [markdown] id="cK3jm6V_P4pB"
# ## CPU
# + id="exG5HvsIQtyE"
# !pip install -U -q numba aesara
# + id="tD19gJ_-QAiZ"
# !taskset -c 0 python run.py benchmarks/equation_of_state/
# + id="NYykl19BWfQI"
# !taskset -c 0 python run.py benchmarks/isoneutral_mixing/
# + id="zf2RaRlPXpM6"
# !taskset -c 0 python run.py benchmarks/turbulent_kinetic_energy/
# + [markdown] id="oOIzGKsPP0ui"
# ## TPU
#
# Make sure to set accelerator to "TPU" before executing this.
# + id="JHOxWXecn3kx"
import jax.tools.colab_tpu
jax.tools.colab_tpu.setup_tpu()
# + id="9-tQlcfZOzm0"
# !python run.py benchmarks/equation_of_state -b jax -b numpy --device tpu
# + id="rKcTVbiVPXFu"
# !python run.py benchmarks/isoneutral_mixing -b jax -b numpy --device tpu
# + id="gfIlTgZol9OA"
# !python run.py benchmarks/turbulent_kinetic_energy -b jax -b numpy --device tpu
# + [markdown] id="RDoapE1YPrpN"
# ## GPU
#
# Make sure to set accelerator to "GPU" before executing this.
# + id="b4CQKseuMnzE"
# get GPU model
# !nvidia-smi -L
# + id="Azo78zrdo88Y"
# !for backend in jax tensorflow pytorch cupy; do python run.py benchmarks/equation_of_state/ --device gpu -b $backend -b numpy; done
# + id="Ps8zEacsPWQW"
# !for backend in jax pytorch cupy; do python run.py benchmarks/isoneutral_mixing/ --device gpu -b $backend -b numpy; done
# + id="ogXoFxFAd0KI"
# !for backend in jax pytorch; do python run.py benchmarks/turbulent_kinetic_energy/ --device gpu -b $backend -b numpy; done
| results/pyhpc_benchmarks_colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from collections import defaultdict
from pathlib import Path
import re
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from tensorboard.backend.event_processing.event_accumulator import EventAccumulator
import toml
import tqdm
# +
def logdir2df(logdir):
"""convert tf.events files in a logs directory into a pandas DataFrame
tf.events files are created by SummaryWriter from PyTorch or Tensorflow
Parameters
----------
logdir : str, Path
path to directory containing tfevents file(s) saved by a SummaryWriter
Returns
-------
df : pandas.Dataframe
with columns 'step', 'wall_time', and all Scalars from the tfevents file
"""
if issubclass(type(logdir), Path):
logdir = str(logdir)
ea = EventAccumulator(path=logdir)
ea.Reload() # load all data written so far
scalar_tags = ea.Tags()['scalars'] # list of tags for values written to scalar
dfs = {}
for scalar_tag in scalar_tags:
dfs[scalar_tag] = pd.DataFrame(ea.Scalars(scalar_tag),
columns=["wall_time",
"step",
scalar_tag.replace('val/', '')])
dfs[scalar_tag] = dfs[scalar_tag].set_index("step")
dfs[scalar_tag].drop("wall_time", axis=1, inplace=True)
return pd.concat([v for k, v in dfs.items()], axis=1)
def logdir2csv(logdir):
"""convert tf.events files in a logs directory into a pandas DataFrame
Parameters
----------
logdir
Returns
-------
"""
df = logdir2df(logdir)
name = list(logdir.glob('*tfevents*'))[0].name
csv_fname = name + '.csv'
df.to_csv(logdir.joinpath(csv_fname))
# +
re_int = re.compile(r'[0-9]+')
def int_from_dir_path(dir_path):
name = dir_path.name
return int(re_int.search(name)[0])
# -
BR_RESULTS_ROOT = Path('~/Documents/repos/coding/birdsong/tweetynet/results/BirdsongRecognition').expanduser().resolve()
BIRD_RESULTS_ROOT = BR_RESULTS_ROOT.joinpath('Bird1')
sorted(BIRD_RESULTS_ROOT.iterdir())
RESULTS_ROOT = BIRD_RESULTS_ROOT.joinpath('results_200503_015746')
train_dur_dirs = sorted(RESULTS_ROOT.glob('train_dur_*'), key=int_from_dir_path)
# +
train_history_dfs = {}
for train_dur_dir in train_dur_dirs:
train_dur = int_from_dir_path(train_dur_dir)
print(f'getting tf.events files for training duration: {train_dur}')
train_history_dfs[train_dur] = {}
replicate_dirs = sorted(train_dur_dir.glob('replicate_*'), key=int_from_dir_path)
for replicate_dir in replicate_dirs:
replicate_num = int_from_dir_path(replicate_dir)
print(f'\treplicate: {replicate_num}')
events_file = sorted(replicate_dir.glob('**/events*'))
assert len(events_file) == 1
events_file = events_file[0]
logdir = events_file.parent
log_df = logdir2df(logdir)
train_history_dfs[train_dur][replicate_num] = log_df
# -
for train_dur, replicate_df_dict in train_history_dfs.items():
for replicate, df in replicate_df_dict.items():
df['avg_error/val'] = 1 - df['avg_acc/val']
# +
n_train_durs = len(train_history_dfs)
a_train_dur = list(train_history_dfs)[0]
n_replicates = len(train_history_dfs[a_train_dur])
fig, ax = plt.subplots(n_train_durs, 5, figsize=(25, 20))
train_durs = sorted(train_history_dfs.keys())
for row_ind, train_dur in enumerate(train_durs):
replicate_df_dict = train_history_dfs[train_dur]
replicate_nums = sorted(replicate_df_dict.keys())
for replicate_num, df in replicate_df_dict.items():
sns.lineplot(x=df.index, y='loss/train', data=df, ax=ax[row_ind, 0], alpha=0.5)
sns.lineplot(x=df.index, y='avg_loss/val', data=df, ax=ax[row_ind, 1], alpha=0.5)
ax[row_ind, 1].set_ylim([0.0, 0.4])
sns.lineplot(x=df.index, y='avg_error/val', data=df, ax=ax[row_ind, 2], alpha=0.5)
ax[row_ind, 2].set_ylim([0.0, 0.2])
sns.lineplot(x=df.index, y='avg_levenshtein/val', data=df, ax=ax[row_ind, 3], alpha=0.5)
ax[row_ind, 3].set_ylim([100, 250])
sns.lineplot(x=df.index, y='avg_segment_error_rate/val', data=df, ax=ax[row_ind, 4], alpha=0.5)
ax[row_ind, 4].set_ylim([0.0, 0.1])
# -
learncurve_df = pd.read_csv(RESULTS_ROOT.joinpath('learning_curve.csv'))
learncurve_df['avg_error'] = 1- learncurve_df['avg_acc']
# +
fig, ax = plt.subplots(1, 4, figsize=(20, 4))
ax = ax.ravel()
sns.stripplot(x='train_set_dur', y='avg_loss', data=learncurve_df, ax=ax[0])
sns.boxplot(x='train_set_dur', y='avg_loss', data=learncurve_df, ax=ax[0])
sns.pointplot(x='train_set_dur', y='avg_loss', data=learncurve_df, ax=ax[0]);
sns.stripplot(x='train_set_dur', y='avg_error', data=learncurve_df, ax=ax[1])
sns.boxplot(x='train_set_dur', y='avg_error', data=learncurve_df, ax=ax[1])
sns.pointplot(x='train_set_dur', y='avg_error', data=learncurve_df, ax=ax[1]);
sns.stripplot(x='train_set_dur', y='avg_levenshtein', data=learncurve_df, ax=ax[2])
sns.boxplot(x='train_set_dur', y='avg_levenshtein', data=learncurve_df, ax=ax[2])
sns.pointplot(x='train_set_dur', y='avg_levenshtein', data=learncurve_df, ax=ax[2]);
sns.stripplot(x='train_set_dur', y='avg_segment_error_rate', data=learncurve_df, ax=ax[3])
sns.boxplot(x='train_set_dur', y='avg_segment_error_rate', data=learncurve_df, ax=ax[3])
sns.pointplot(x='train_set_dur', y='avg_segment_error_rate', data=learncurve_df, ax=ax[3]);
# -
| article/src/scripts/BirdsongRecognition/initial_submission/Bird1.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// <a name="top"></a><img src="images/chisel_1024.png" alt="Chisel logo" style="width:480px;" />
// # Module 3.4: Functional Programming
// **Prev: [Higher-Order Functions](3.3_higher-order_functions.ipynb)**<br>
// **Next: [Object Oriented Programming](3.5_object_oriented_programming.ipynb)**
//
// ## Motivation
// You saw functions in many previous modules, but now it's time to make our own and use them effectively.
//
// ## Setup
val path = System.getProperty("user.dir") + "/source/load-ivy.sc"
interp.load.module(ammonite.ops.Path(java.nio.file.FileSystems.getDefault().getPath(path)))
// This module uses the Chisel `FixedPoint` type, which currently resides in the experimental package.
import chisel3._
import chisel3.util._
import chisel3.tester._
import chisel3.tester.RawTester.test
import chisel3.experimental._
import chisel3.internal.firrtl.KnownBinaryPoint
// ---
// # Functional Programming in Scala
// Scala functions were introduced in Module 1, and you saw them being used a lot in the previous module. Here's a refresher on functions. Functions take any number of inputs and produce one output. Inputs are often called arguments to a function. To produce no output, return the `Unit` type.
//
// <span style="color:blue">**Example: Custom Functions**</span><br>
// Below are some examples of functions in Scala.
// +
// No inputs or outputs (two versions).
def hello1(): Unit = print("Hello!")
def hello2 = print("Hello again!")
// Math operation: one input and one output.
def times2(x: Int): Int = 2 * x
// Inputs can have default values, and explicitly specifying the return type is optional.
// Note that we recommend specifying the return types to avoid surprises/bugs.
def timesN(x: Int, n: Int = 2) = n * x
// Call the functions listed above.
hello1()
hello2
times2(4)
timesN(4) // no need to specify n to use the default value
timesN(4, 3) // argument order is the same as the order where the function was defined
timesN(n=7, x=2) // arguments may be reordered and assigned to explicitly
// -
// ## Functions as Objects
// Functions in Scala are first-class objects. That means we can assign a function to a `val` and pass it to classes, objects, or other functions as an argument.
//
// <span style="color:blue">**Example: Function Objects**</span><br>
// Below are the same functions implemented as functions and as objects.
// +
// These are normal functions.
def plus1funct(x: Int): Int = x + 1
def times2funct(x: Int): Int = x * 2
// These are functions as vals.
// The first one explicitly specifies the return type.
val plus1val: Int => Int = x => x + 1
val times2val = (x: Int) => x * 2
// Calling both looks the same.
plus1funct(4)
plus1val(4)
plus1funct(x=4)
//plus1val(x=4) // this doesn't work
// -
// Why would you want to create a `val` instead of a `def`? With a `val`, you can now pass the function around to other functions, as shown below. You can even create your own functions that accept other functions as arguments. Formally, functions that take or produce functions are called *higher-order functions*. You saw them used in the last module, but now you'll make your own!
//
// <span style="color:blue">**Example: Higher-Order Functions**</span><br>
// Here we show `map` again, and we also create a new function, `opN`, that accepts a function, `op`, as an argument.
// +
// create our function
val plus1 = (x: Int) => x + 1
val times2 = (x: Int) => x * 2
// pass it to map, a list function
val myList = List(1, 2, 5, 9)
val myListPlus = myList.map(plus1)
val myListTimes = myList.map(times2)
// create a custom function, which performs an operation on X N times using recursion
def opN(x: Int, n: Int, op: Int => Int): Int = {
if (n <= 0) { x }
else { opN(op(x), n-1, op) }
}
opN(7, 3, plus1)
opN(7, 3, times2)
// -
// <span style="color:blue">**Example: Functions vs. Objects**</span><br>
// A possibly confusing situation arises when using functions without arguments. Functions are evaluated every time they are called, while `val`s are evaluated at instantiation.
// +
import scala.util.Random
// both x and y call the nextInt function, but x is evaluated immediately and y is a function
val x = Random.nextInt
def y = Random.nextInt
// x was previously evaluated, so it is a constant
println(s"x = $x")
println(s"x = $x")
// y is a function and gets reevaluated at each call, thus these produce different results
println(s"y = $y")
println(s"y = $y")
// -
// ## Anonymous Functions
// As the name implies, anonymous functions are nameless. There's no need to create a `val` for a function if we'll only use it once.
//
// <span style="color:blue">**Example: Anonymous Functions**</span><br>
// The following example demonstrates this. They are often scoped (put in curly braces instead of parentheses).
// +
val myList = List(5, 6, 7, 8)
// add one to every item in the list using an anonymous function
// arguments get passed to the underscore variable
// these all do the same thing
myList.map( (x:Int) => x + 1 )
myList.map(_ + 1)
// a common situation is to use case statements within an anonymous function
val myAnyList = List(1, 2, "3", 4L, myList)
myAnyList.map {
case (_:Int|_:Long) => "Number"
case _:String => "String"
case _ => "error"
}
// -
// <span style="color:red">**Exercise: Sequence Manipulation**</span><br>
// A common set of higher-order functions you'll use are `scanLeft`/`scanRight`, `reduceLeft`/`reduceRight`, and `foldLeft`/`foldRight`. It's important to understand how each one works and when to use them. The default directions for `scan`, `reduce`, and `fold` are left, though this is not guaranteed for all cases.
// +
val exList = List(1, 5, 7, 100)
// write a custom function to add two numbers, then use reduce to find the sum of all values in exList
def add(a: Int, b: Int): Int = ???
val sum = ???
// find the sum of exList using an anonymous function (hint: you've seen this before!)
val anon_sum = ???
// find the moving average of exList from right to left using scan; make the result (ma2) a list of doubles
def avg(a: Int, b: Double): Double = ???
val ma2 = ???
// +
assert(add(88, 88) == 176)
assert(sum == 113)
assert(anon_sum == 113)
assert(avg(100, 100.0) == 100.0)
assert(ma2 == List(8.875, 16.75, 28.5, 50.0, 0.0))
// -
// <div id="container"><section id="accordion"><div>
// <input type="checkbox" id="check-1" />
// <label for="check-1"><strong>Solution</strong></label>
// <article>
// <pre style="background-color:#f7f7f7">
// def add(a: Int, b: Int): Int = a + b
// val sum = exList.reduce(add)
//
// val anon\_sum = exList.reduce(\_ + \_)
//
// def avg(a: Int, b: Double): Double = (a + b)/2.0
// val ma2 = exList.scanRight(0.0)(avg)
// </pre></article></div></section></div>
// ---
// # Functional Programming in Chisel
// Let's look at some examples of how to use functional programming when creating hardware generators in Chisel.
// <span style="color:blue">**Example: FIR Filter**</span><br>
// First, we'll revisit the FIR filter from previous examples. Instead of passing in the coefficients as parameters to the class or making them programmable, we'll pass a function to the FIR that defines how the window coefficients are calculated. This function will take the window length and bitwidth to produce a scaled list of coefficients. Here are two example windows. To avoid fractions, we'll scale the coefficients to be between the max and min integer values. For more on these windows, check out the [this Wikipedia page](https://en.wikipedia.org/wiki/Window_function).
// +
// get some math functions
import scala.math.{abs, round, cos, Pi, pow}
// simple triangular window
val TriangularWindow: (Int, Int) => Seq[Int] = (length, bitwidth) => {
val raw_coeffs = (0 until length).map( (x:Int) => 1-abs((x.toDouble-(length-1)/2.0)/((length-1)/2.0)) )
val scaled_coeffs = raw_coeffs.map( (x: Double) => round(x * pow(2, bitwidth)).toInt)
scaled_coeffs
}
// Hamming window
val HammingWindow: (Int, Int) => Seq[Int] = (length, bitwidth) => {
val raw_coeffs = (0 until length).map( (x: Int) => 0.54 - 0.46*cos(2*Pi*x/(length-1)))
val scaled_coeffs = raw_coeffs.map( (x: Double) => round(x * pow(2, bitwidth)).toInt)
scaled_coeffs
}
// check it out! first argument is the window length, and second argument is the bitwidth
TriangularWindow(10, 16)
HammingWindow(10, 16)
// -
// Now we'll create a FIR filter that accepts a window function as the argument. This allows us to define new windows later on and retain the same FIR generator. It also allows us to independently size the FIR, knowing the window will be recalculated for different lengths or bitwidths. Since we are choosing the window at compile time, these coefficients are fixed.
// +
// our FIR has parameterized window length, IO bitwidth, and windowing function
class MyFir(length: Int, bitwidth: Int, window: (Int, Int) => Seq[Int]) extends Module {
val io = IO(new Bundle {
val in = Input(UInt(bitwidth.W))
val out = Output(UInt((bitwidth*2+length-1).W)) // expect bit growth, conservative but lazy
})
// calculate the coefficients using the provided window function, mapping to UInts
val coeffs = window(length, bitwidth).map(_.U)
// create an array holding the output of the delays
// note: we avoid using a Vec here since we don't need dynamic indexing
val delays = Seq.fill(length)(Wire(UInt(bitwidth.W))).scan(io.in)( (prev: UInt, next: UInt) => {
next := RegNext(prev)
next
})
// multiply, putting result in "mults"
val mults = delays.zip(coeffs).map{ case(delay: UInt, coeff: UInt) => delay * coeff }
// add up multiplier outputs with bit growth
val result = mults.reduce(_+&_)
// connect output
io.out := result
}
visualize(() => new MyFir(7, 12, TriangularWindow))
// -
// Those last three lines could be easily combined into one. Also notice how we've handled bitwidth growth conservatively to avoid loss.
//
// <span style="color:blue">**Example: FIR Filter Tester**</span><br>
// Let's test our FIR! Previously, we provided a custom golden model. This time we'll use Breeze, a Scala library of useful linear algebra and signal processing functions, as a golden model for our FIR filter. The code below compares the Chisel output with the golden model output, and any errors cause the tester to fail.
//
// Try uncommenting the print statment at the end just after the expect call. Also try changing the window from triangular to Hamming.
// +
// math imports
import scala.math.{pow, sin, Pi}
import breeze.signal.{filter, OptOverhang}
import breeze.signal.support.{CanFilter, FIRKernel1D}
import breeze.linalg.DenseVector
// test parameters
val length = 7
val bitwidth = 12 // must be less than 15, otherwise Int can't represent the data, need BigInt
val window = TriangularWindow
// test our FIR
test(new MyFir(length, bitwidth, window)) { c =>
// test data
val n = 100 // input length
val sine_freq = 10
val samp_freq = 100
// sample data, scale to between 0 and 2^bitwidth
val max_value = pow(2, bitwidth)-1
val sine = (0 until n).map(i => (max_value/2 + max_value/2*sin(2*Pi*sine_freq/samp_freq*i)).toInt)
//println(s"input = ${sine.toArray.deep.mkString(", ")}")
// coefficients
val coeffs = window(length, bitwidth)
//println(s"coeffs = ${coeffs.toArray.deep.mkString(", ")}")
// use breeze filter as golden model; need to reverse coefficients
val expected = filter(
DenseVector(sine.toArray),
FIRKernel1D(DenseVector(coeffs.reverse.toArray), 1.0, ""),
OptOverhang.None
)
expected.toArray // seems to be necessary
//println(s"exp_out = ${expected.toArray.deep.mkString(", ")}") // this seems to be necessary
// push data through our FIR and check the result
c.reset.poke(true.B)
c.clock.step(5)
c.reset.poke(false.B)
for (i <- 0 until n) {
c.io.in.poke(sine(i).U)
if (i >= length-1) { // wait for all registers to be initialized since we didn't zero-pad the data
val expectValue = expected(i-length+1)
//println(s"expected value is $expectValue")
c.io.out.expect(expected(i-length+1).U)
//println(s"cycle $i, got ${c.io.out.peek()}, expect ${expected(i-length+1)}")
}
c.clock.step(1)
}
}
// -
// ---
// # Chisel Exercises
// Complete the following exercises to practice writing functions, using them as arguments to hardware generators, and avoiding mutable data.
//
// <span style="color:red">**Exercise: Neural Network Neuron**</span><br>
// Our first example will have you build a neuron, the building block of fully-connected layers in artificial neural networks. Neurons take inputs and a set of weights, one per input, and produce one output. The weights and inputs are multiplied and added, and the result is fed through an activation function. In this exercise, you will implement different activation functions and pass them as an argument to your neuron generator.
//
// 
//
// First, complete the following code to create a neuron generator. The parameter `inputs` gives the number of inputs. The parameter `act` is a function that implements the logic of the activation function. We'll make the inputs and outputs 16-bit fixed point values with 8 fractional bits.
class Neuron(inputs: Int, act: FixedPoint => FixedPoint) extends Module {
val io = IO(new Bundle {
val in = Input(Vec(inputs, FixedPoint(16.W, 8.BP)))
val weights = Input(Vec(inputs, FixedPoint(16.W, 8.BP)))
val out = Output(FixedPoint(16.W, 8.BP))
})
???
}
// <div id="container"><section id="accordion"><div>
// <input type="checkbox" id="check-2" />
// <label for="check-2"><strong>Solution</strong></label>
// <article>
// <pre style="background-color:#f7f7f7">
// val mac = io.in.zip(io.weights).map{ case(a:FixedPoint, b:FixedPoint) => a*b}.reduce(_+_)
// io.out := act(mac)
// </pre></article></div></section></div>
// Now let's create some activation functions! We'll use a threshold of zero. Typical activation functions are the sigmoid function and the rectified linear unit (ReLU).
//
// The sigmoid we'll use is called the [logistic function](https://en.wikipedia.org/wiki/Logistic_function), given by
//
// $logistic(x) = \cfrac{1}{1+e^{-\beta x}}$
//
// where $\beta$ is the slope factor. However, computing the exponential function in hardware is quite challenging and expensive. We'll approximate this as the step function.
//
// $step(x) = \begin{cases}
// 0 & \text{if } x \le 0 \\
// 1 & \text{if } x \gt 0
// \end{cases}$
//
// The second function, the ReLU, is given by a similar formula.
//
// $relu(x) = \begin{cases}
// 0 & \text{if } x \le 0 \\
// x & \text{if } x \gt 0
// \end{cases}$
//
// Implement these two functions below. You can specify a fixed-point literal like `-3.14.F(8.BP)`.
val Step: FixedPoint => FixedPoint = ???
val ReLU: FixedPoint => FixedPoint = ???
// <div id="container"><section id="accordion"><div>
// <input type="checkbox" id="check-3" />
// <label for="check-3"><strong>Solution</strong></label>
// <article>
// <pre style="background-color:#f7f7f7">
// val Step: FixedPoint => FixedPoint = x => Mux(x <= 0.F(8.BP), 0.F(8.BP), 1.F(8.BP))
// val ReLU: FixedPoint => FixedPoint = x => Mux(x <= 0.F(8.BP), 0.F(8.BP), x)
// </pre></article></div></section></div>
// Finally, let's create a tester that checks the correctness of our Neuron. With the step activation function, neurons may be used as logic gate approximators. Proper selection of weights and bias can perform binary functions. We'll test our neuron using AND logic. Complete the following tester to check our neuron with the step function.
//
// Note that since the circuit is purely combinational, the `reset(5)` and `step(1)` calls are not necessary.
// +
// test our Neuron
test(new Neuron(2, Step)) { c =>
val inputs = Seq(Seq(-1, -1), Seq(-1, 1), Seq(1, -1), Seq(1, 1))
// make this a sequence of two values
val weights = ???
// push data through our Neuron and check the result (AND gate)
c.reset.poke(true.B)
c.clock.step(5)
c.reset.poke(false.B)
for (i <- inputs) {
c.io.in(0).poke(i(0).F(8.BP))
c.io.in(1).poke(i(1).F(8.BP))
c.io.weights(0).poke(weights(0).F(16.W, 8.BP))
c.io.weights(1).poke(weights(1).F(16.W, 8.BP))
c.io.out.expect((if (i(0) + i(1) > 0) 1 else 0).F(16.W, 8.BP))
c.clock.step(1)
}
}
// -
// <div id="container"><section id="accordion"><div>
// <input type="checkbox" id="check-4" />
// <label for="check-4"><strong>Solution</strong></label>
// <article>
// <pre style="background-color:#f7f7f7">
// val weights = Seq(1.0, 1.0)
// </pre></article></div></section></div>
// ---
// # You're done!
//
// [Return to the top.](#top)
| 3.4_functional_programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The clash of Neighborhoods in Toronto City
#
#
#
# In the first part we will build a code to scrape the following Wikipedia page, https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M, in order to transform the data in the table of postal codes into a pandas dataframe and then clean the data following this main steps:
#
# * Only process the cells that have an assigned borough. Ignore cells with a borough that is Not assigned.
# * More than one neighborhood can exist in one postal code area. For example, in the table on the Wikipedia page, you will notice that - M5A is listed twice and has two neighborhoods: Harbourfront and Regent Park. These two rows will be combined into one row with the - neighborhoods separated with a comma as shown in row 11 in the above table.
# * If a cell has a borough but a Not assigned neighborhood, then the neighborhood will be the same as the borough. So for the 9th cell in the table on the Wikipedia page, the value of the Borough and the Neighborhood columns will be Queen's Park.
# * Use the .shape method to print the number of rows of your dataframe.
import requests
import pandas as pd
from bs4 import BeautifulSoup
List_url = "https://en.wikipedia.org/wiki/List_of_postal_codes_of_Canada:_M"
source = requests.get(List_url).text
soup = BeautifulSoup(source, 'xml')
table = soup.find('table')
#dataframe will consist of three columns: PostalCode, Borough, and Neighborhood
column_names = ['Postalcode','Borough','Neighborhood']
df = pd.DataFrame(columns = column_names)
# +
# Search all the data
for tr_cell in table.find_all('tr'):
row_data=[]
for td_cell in tr_cell.find_all('td'):
row_data.append(td_cell.text.strip())
if len(row_data)==3:
df.loc[len(df)] = row_data
df.head()
# -
# ## Data cleaning
# Get index where Borough = Not assigned and Drop the row
df.drop(labels = df[df['Borough']=='Not assigned'].index, axis = 0, inplace = True)
df.head()
# Make sure that if a cell has a borough but a 'Not assigned' neighborhood, then the neighborhood will be the same as the borough
df.loc[(df.Neighborhood == 'Not assigned'),'Neighborhood']=df['Borough']
df.head()
# Make sure than no more than one neighborhood can exist in one postal code area by groupind them and separate them by a coma
df = df.groupby('Postalcode').agg({'Borough' : 'first', 'Neighborhood' : ','.join}).reset_index()
df.head()
# Verify that all Postalcode data are now unique
boolean = df['Postalcode'].duplicated().any()
print(boolean)
print("The final shape of the dataframe is :", df.shape)
# ## Getting latitude and longitude coordinates
# In the second part of this notebook we will creat a new dataframe with the latitude and the longitude coordinates of each neighborhood.
# +
# Create a loop to get the coordinates
def get_geocode(postal_code):
# initialize your variable to None
lat_lng_coords = None
while(lat_lng_coords is None):
g = geocoder.google('{}, Toronto, Ontario'.format(postal_code))
lat_lng_coords = g.latlng
latitude = lat_lng_coords[0]
longitude = lat_lng_coords[1]
return latitude,longitude
# -
# Get the dataset that includes latitude and longitude coordinates for each neighborhood
df_geo=pd.read_csv('http://cocl.us/Geospatial_data')
df_geo.head()
# Merge the dataframe with latitude and longitude coordinates and the previous dataframe
df_geo.rename(columns={'Postal Code':'Postalcode'},inplace=True)
df_merge = pd.merge(df_geo, df, on='Postalcode')
df_tot = df_merge[['Postalcode','Borough','Neighborhood','Latitude','Longitude']]
df_tot.head()
# ## Segmenting and Clustering Neighborhoods
# In the third part of this notebook we will generate a map to visualize our neighborhoods and how they are clustered together. We will then explore each neighborhood and create a dataframe than contains all the venues depicted in each of them.
# +
import pandas as pd # library for data analsysis
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
import json # library to handle JSON files
#conda config --append channels conda-forge
import requests # library to handle requests
from pandas.io.json import json_normalize # tranform JSON file into a pandas dataframe
# Matplotlib and associated plotting modules
import matplotlib.cm as cm
import matplotlib.colors as colors
# import k-means from clustering stage
from sklearn.cluster import KMeans
# #!conda install -c conda-forge folium=0.5.0 --yes # uncomment this line if you haven't completed the Foursquare API lab
# !pip install folium
import folium # map rendering library
import sys
# !{sys.executable} -m pip install reverse_geocoder
# !pip install geopy
from geopy.geocoders import Nominatim # convert an address into latitude and longitude values
print('Libraries imported.')
# -
print('The dataframe has {} boroughs and {} neighborhoods.'.format(
len(df_tot['Borough'].unique()),
df_tot.shape[0]
)
)
# ### Create a map of Toronto with neighborhoods superimposed on top
# +
# Get latitude and longitude coordinates of Toronto City
address = 'Toronto, Canada'
geolocator = Nominatim(user_agent="can_explorer")
location = geolocator.geocode(address)
latitude = location.latitude
longitude = location.longitude
print('The geograpical coordinate of Toronto are {}, {}.'.format(latitude, longitude))
# +
# create map of Toronto using latitude and longitude values
map_toronto = folium.Map(location=[latitude, longitude], zoom_start=10)
# add markers to map
for lat, lng, borough, neighborhood in zip(df_tot['Latitude'], df_tot['Longitude'], df_tot['Borough'], df_tot['Neighborhood']):
label = '{}, {}'.format(neighborhood, borough)
label = folium.Popup(label, parse_html=True)
folium.CircleMarker(
[lat, lng],
radius=2,
popup=label,
color='blue',
fill=True,
fill_color='#3186cc',
fill_opacity=0.7,
parse_html=False).add_to(map_toronto)
map_toronto
# +
# Define Foursquare Credentials and Version
LIMIT = 100
CLIENT_ID = 'PXNGKJNS1KJ23E5I4MF1H1TW0K4FPJ50M1JZHXPJCN0QNN1Q' # your Foursquare ID
CLIENT_SECRET = '<KEY>' # your Foursquare Secret
ACCESS_TOKEN = '<KEY>' # your FourSquare Access Token
VERSION = '20180604'
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET:' + CLIENT_SECRET)
# -
# ### Explore all neighborhoods and analyzing the data
#
# We will now explore neighborhood by neighborhood in the city of Toronto and retreive neighborhood's name, latitude and longitude values and all venues in the given neighborhood.
#create a function loop over all the neighborhoods in Toronto
def getNearbyVenues(names, latitudes, longitudes, radius=500):
venues_list=[]
for name, lat, lng in zip(names, latitudes, longitudes):
print(name)
# create the API request URL
url = 'https://api.foursquare.com/v2/venues/explore?&client_id={}&client_secret={}&v={}&ll={},{}&radius={}&limit={}'.format(
CLIENT_ID,
CLIENT_SECRET,
VERSION,
lat,
lng,
radius,
LIMIT)
# make the GET request
results = requests.get(url).json()["response"]['groups'][0]['items']
# return only relevant information for each nearby venue
venues_list.append([(
name,
lat,
lng,
v['venue']['name'],
v['venue']['location']['lat'],
v['venue']['location']['lng'],
v['venue']['categories'][0]['name']) for v in results])
nearby_venues = pd.DataFrame([item for venue_list in venues_list for item in venue_list])
nearby_venues.columns = ['Neighborhood',
'Neighborhood Latitude',
'Neighborhood Longitude',
'Venue',
'Venue Latitude',
'Venue Longitude',
'Venue Category']
return(nearby_venues)
# run the above function on each neighborhood and create a new dataframe called toronto_venues
toronto_venues = pd.DataFrame(getNearbyVenues(names=df_tot['Neighborhood'],
latitudes=df_tot['Latitude'],
longitudes=df_tot['Longitude'],
))
# We will print the size of the new dataframe toronto_venues and see more about this dataframe
print(toronto_venues.shape)
toronto_venues.head()
# General informations
print('There are {} venues in Toronto.'.format(toronto_venues['Venue Category'].count()))
print('There are {} uniques categories of venue.'.format(len(toronto_venues['Venue Category'].unique())))
print('There are {} uniques Neighborhood.'.format(len(toronto_venues['Neighborhood'].unique())))
# Count the number of total venue in each Neighborhood
venues_tot = toronto_venues.groupby('Neighborhood').count()
venues_tot.sort_values('Venue',ascending=False,inp lace=True)
venues_tot.drop(labels=['Neighborhood Latitude', 'Neighborhood Longitude', 'Venue Latitude', 'Venue Longitude','Venue Category'], axis=1, inplace=True)
venues_tot.head()
# +
# Take the top 20 of Neighborhood in Toronto city that count the higher number of venues
venues_top20 = venues_tot.head(20)
venues_temp = toronto_venues.groupby('Neighborhood').agg({'Venue Category': 'first', 'Venue Category' : ','.join}).reset_index()
venues_temp.head(20)
#Let's explore the first neighborhood in terms of higher number of venue
Calgary_venues_NE = venues_temp[venues_temp.Neighborhood=='Toronto Dominion Centre, Design Exchange']
Calgary_venues_NE
# +
import matplotlib.pyplot as plt
#plot a bar graph to see which neighborhoods have the most restaurants
bar_graph_data=pd.DataFrame(venues_top20['Venue']).reset_index()
x=bar_graph_data.iloc[:,0]
y=bar_graph_data.iloc[:,1]
plt.figure(figsize=(20,4))
plt.bar(x, y)
#plot labels and titles
plt.xlabel('Neighborhood')
plt.xticks(rotation=70, ha='right')
plt.ylabel('Number of Venue')
plt.show()
# -
# ## Analizing and Clustering of Neighborhoods
# +
# one hot encoding
toronto_onehot = pd.get_dummies(toronto_venues[['Venue Category']], prefix="", prefix_sep="")
# add neighborhood column back to dataframe
toronto_onehot['Neighborhood'] = toronto_venues['Neighborhood']
# move neighborhood column to the first column
fixed_columns = [toronto_onehot.columns[-1]] + list(toronto_onehot.columns[:-1])
toronto_onehot = toronto_onehot[fixed_columns]
toronto_onehot.head()
# -
# Examine the new data frame
toronto_onehot.shape
# +
# Here rows are grouped by neighborhood by taking the mean of the frequency of occurrence of each category
toronto_grouped = toronto_onehot.groupby('Neighborhood').mean().reset_index()
toronto_grouped
# print each neighborhood along with the top 5 most common venues
num_top_venues = 5
for hood in toronto_grouped['Neighborhood']:
print("----"+hood+"----")
temp = toronto_grouped[toronto_grouped['Neighborhood'] == hood].T.reset_index()
temp.columns = ['venue','freq']
temp = temp.iloc[1:]
temp['freq'] = temp['freq'].astype(float)
temp = temp.round({'freq': 2})
print(temp.sort_values('freq', ascending=False).reset_index(drop=True).head(num_top_venues))
print('\n')
# +
# We will write a function to sort the venues and then create a new dataframe and display the top 10 venues for each neighborhood.
import numpy as np
def return_most_common_venues(row, num_top_venues):
row_categories = row.iloc[1:]
row_categories_sorted = row_categories.sort_values(ascending=False)
return row_categories_sorted.index.values[0:num_top_venues]
num_top_venues = 10
indicators = ['st', 'nd', 'rd']
# create columns according to number of top venues
columns = ['Neighborhood']
for ind in np.arange(num_top_venues):
try:
columns.append('{}{} Most Common Venue'.format(ind+1, indicators[ind]))
except:
columns.append('{}th Most Common Venue'.format(ind+1))
# create a new dataframe
neighborhoods_venues_sorted = pd.DataFrame(columns=columns)
neighborhoods_venues_sorted['Neighborhood'] = toronto_grouped['Neighborhood']
for ind in np.arange(toronto_grouped.shape[0]):
neighborhoods_venues_sorted.iloc[ind, 1:] = return_most_common_venues(toronto_grouped.iloc[ind, :], num_top_venues)
neighborhoods_venues_sorted.head()
# -
# We will now group run a k-means algortithm to cluster the neighborhoods into 5 separate clusters.
# +
# set number of clusters
kclusters = 7
toronto_grouped_clustering = toronto_grouped.drop('Neighborhood', 1)
# run k-means clustering
kmeans = KMeans(n_clusters=kclusters, random_state=0).fit(toronto_grouped_clustering)
# check cluster labels generated for each row in the dataframe
kmeans.labels_[0:10]
# +
# add clustering labels
neighborhoods_venues_sorted.insert(0, 'Cluster Labels', kmeans.labels_)
toronto_merged = df_tot
# merge manhattan_grouped with manhattan_data to add latitude/longitude for each neighborhood
toronto_merged = toronto_merged.join(neighborhoods_venues_sorted.set_index('Neighborhood'), on='Neighborhood')
toronto_merged.head() # check the last columns!
# +
# Map all clusters
map_clusters = folium.Map(location=[latitude, longitude], zoom_start=11)
# set color scheme for the clusters
x = np.arange(kclusters)
ys = [i + x + (i*x)**2 for i in range(kclusters)]
colors_array = cm.rainbow(np.linspace(0, 1, len(ys)))
rainbow = [colors.rgb2hex(i) for i in colors_array]
# add markers to the map
markers_colors = []
for lat, lon, poi, cluster in zip(toronto_merged['Latitude'], toronto_merged['Longitude'], toronto_merged['Neighborhood'], toronto_merged['Cluster Labels']):
label = folium.Popup(str(poi) + ' Cluster ' + str(cluster), parse_html=True)
folium.CircleMarker(
[lat, lon],
radius=5,
popup=label,
fill=True,
fill_opacity=0.7).add_to(map_clusters)
map_clusters
# -
# ## Examine Clusters 1 to 5
# ### Cluster 1
toronto_merged.loc[toronto_merged['Cluster Labels'] == 0, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
# ### Cluster 2
toronto_merged.loc[toronto_merged['Cluster Labels'] == 1, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
# ### Cluster 3
toronto_merged.loc[toronto_merged['Cluster Labels'] == 2, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
# ### Cluster 4
toronto_merged.loc[toronto_merged['Cluster Labels'] == 3, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
# ### Cluster 5
toronto_merged.loc[toronto_merged['Cluster Labels'] == 4, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
# ### Cluster 6
toronto_merged.loc[toronto_merged['Cluster Labels'] == 5, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
# ### Cluster 7
toronto_merged.loc[toronto_merged['Cluster Labels'] == 6, toronto_merged.columns[[1] + list(range(5, toronto_merged.shape[1]))]]
| CapsoneProjectWeek5/TheClashofNeighboor-CapsoneProject-week5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
m = '110101'
n = '0011011'
b = [['' for _ in range(len(n)+1)] for _ in range(len(m)+1)]
c = [[0 for _ in range(len(n)+1)] for _ in range(len(m)+1)]
for i in range(1, len(m)+1):
for j in range(1, len(n)+1):
if m[i-1] == n[j-1]: c[i][j], b[i][j] = c[i-1][j-1]+1, 'LeftUp'
elif c[i-1][j] >= c[i][j-1]: c[i][j], b[i][j] = c[i-1][j], 'Up'
else: c[i][j], b[i][j] = c[i][j-1], 'Left'
print('\n'.join(['\t'.join(i) for i in b]))
print()
print('\n'.join(['\t'.join(list(map(str, i))) for i in c]))
# +
a = [1, 3, 8, 3, 9, 6, 7, 2]
c = [[0 for _ in range(len(a)+1)] for _ in range(len(a)+1)]
for i in range(1, len(a)+1):
c[0][i] = 1
for j in range(i+1, len(a)+1):
if a[i-1] <= a[j-1]: c[i][j] = max(c[i-1][i]+1, c[i-1][j])
else: c[i][j] = c[i-1][j]
print('\t'.join(list(map(str, [0, 0] + a))))
print('\n'.join(['\t'.join(list(map(str, [j] + i))) for i, j in zip(c, a)]))
# +
W = 55
value = [2, 7, 30, 31, 9, 5, 5, 1]
weight = [4, 10, 44, 43, 12, 9, 8, 2]
c = [0 for i in range(W+1)]
def knapsack(n, W):
for i in range(1, n):
for j in range(W, 1, -1):
if j < weight[i]: break
c[j] = max(c[j], c[j - weight[i]] + value[i])
return c[W]
knapsack(len(weight), W)
# +
class Item:
def __init__(self, letter, number):
self.letter = letter
self.number = number
def __str__(self):
return '({letter}, {number})'.format(letter=self.letter, number=str(self.number))
class Node:
def __init__(self, left, right):
self.left = left
self.right = right
self.number = left.number + right.number
def __str__(self):
return '[{left} |{number}| {right}]'.format(left=self.left.__str__(), number=self.number, right=self.right.__str__())
letters = ['A', 'B', 'C', 'D', 'E', 'F', 'G']
numbers = [8, 2, 5, 6, 3, 7, 9]
c = [Item(i, j) for i, j in zip(letters, numbers)]
def huffman(c):
for i in range(1, len(c)):
c = sorted(c, key=lambda x: x.number)
c.append(Node(c[0], c[1]))
c = c[2:]
return c[0]
huffman(c).__str__()
| Algorithms HW3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fast GP implementations
# + tags=["hidden"]
# %matplotlib inline
# + tags=["hidden"]
# %config InlineBackend.figure_format = 'retina'
# + tags=["hidden"]
from matplotlib import rcParams
rcParams["savefig.dpi"] = 100
rcParams["figure.dpi"] = 100
rcParams["figure.figsize"] = 12, 4
rcParams["font.size"] = 16
rcParams["text.usetex"] = False
rcParams["font.family"] = ["sans-serif"]
rcParams["font.sans-serif"] = ["cmss10"]
rcParams["axes.unicode_minus"] = False
# + tags=["hidden"]
# https://github.com/matplotlib/matplotlib/issues/12039
try:
old_get_unicode_index
except NameError:
print('Patching matplotlib.mathtext.get_unicode_index')
import matplotlib.mathtext as mathtext
old_get_unicode_index = mathtext.get_unicode_index
mathtext.get_unicode_index = lambda symbol, math=True:\
ord('-') if symbol == '-' else old_get_unicode_index(symbol, math)
# -
# ## Benchmarking our implementation
# Let's do some timing tests and compare them to what we get with two handy GP packages: ``george`` and ``celerite``. We'll learn how to use both along the way.
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 1a</h1>
# </div>
#
# Let's time how long our custom implementation of a GP takes for a rather long dataset. Create a time array of ``10,000`` points between 0 and 10 and time how long it takes to sample the prior of the GP for the default kernel parameters (unit amplitude and timescale). Add a bit of noise to the sample and then time how long it takes to evaluate the log likelihood for the dataset. Make sure to store the value of the log likelihood for later.
# + tags=["hidden"]
import numpy as np
from scipy.linalg import cho_factor
def ExpSquaredKernel(t1, t2=None, A=1.0, l=1.0):
"""
Return the ``N x M`` exponential squared
covariance matrix.
"""
if t2 is None:
t2 = t1
T2, T1 = np.meshgrid(t2, t1)
return A ** 2 * np.exp(-0.5 * (T1 - T2) ** 2 / l ** 2)
def ln_gp_likelihood(t, y, sigma=0, **kwargs):
"""
"""
# The covariance and its determinant
npts = len(t)
kernel = ExpSquaredKernel
K = kernel(t, **kwargs) + sigma ** 2 * np.eye(npts)
# The marginal log likelihood
log_like = -0.5 * np.dot(y.T, np.linalg.solve(K, y))
log_like -= 0.5 * np.linalg.slogdet(K)[1]
log_like -= 0.5 * npts * np.log(2 * np.pi)
return log_like
def draw_from_gaussian(mu, S, ndraws=1, eps=1e-12):
"""
Generate samples from a multivariate gaussian
specified by covariance ``S`` and mean ``mu``.
"""
npts = S.shape[0]
L, _ = cho_factor(S + eps * np.eye(npts), lower=True)
L = np.tril(L)
u = np.random.randn(npts, ndraws)
x = np.dot(L, u) + mu[:, None]
return x.T
def compute_gp(t_train, y_train, t_test, sigma=0, **kwargs):
"""
"""
# Compute the required matrices
kernel = ExpSquaredKernel
Stt = kernel(t_train, **kwargs)
Stt += sigma ** 2 * np.eye(Stt.shape[0])
Spp = kernel(t_test, **kwargs)
Spt = kernel(t_test, t_train, **kwargs)
# Compute the mean and covariance of the GP
mu = np.dot(Spt, np.linalg.solve(Stt, y_train))
S = Spp - np.dot(Spt, np.linalg.solve(Stt, Spt.T))
return mu, S
# + tags=["hidden"]
# %%time
np.random.seed(3)
t = np.linspace(0, 10, 10000)
sigma = np.ones_like(t) * 0.05
gp_mu, gp_S = compute_gp([], [], t, A=1.0, l=1.0)
y = draw_from_gaussian(gp_mu, gp_S)[0] + sigma * np.random.randn(len(t))
# + tags=["hidden"]
# %%time
ln_gp_likelihood(t, y, sigma)
# -
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 1b</h1>
# </div>
#
# Let's time how long it takes to do the same operations using the ``george`` package (``pip install george``).
#
# The kernel we'll use is
#
# ```python
# kernel = amp ** 2 * george.kernels.ExpSquaredKernel(tau ** 2)
# ```
#
# where ``amp = 1`` and ``tau = 1`` in this case.
#
# To instantiate a GP using ``george``, simply run
#
# ```python
# gp = george.GP(kernel)
# ```
#
# The ``george`` package pre-computes a lot of matrices that are re-used in different operations, so before anything else, ask it to compute the GP model for your timeseries:
#
# ```python
# gp.compute(t, sigma)
# ```
#
# Note that we've only given it the time array and the uncertainties, so as long as those remain the same, you don't have to re-compute anything. This will save you a lot of time in the long run!
#
# Finally, the log likelihood is given by ``gp.log_likelihood(y)`` and a sample can be drawn by calling ``gp.sample()``.
#
# How do the speeds compare? Did you get the same value of the likelihood (assuming you computed it for the same sample in both cases)?
# + tags=["hidden"]
import george
# + tags=["hidden"]
# %%time
kernel = george.kernels.ExpSquaredKernel(1.0)
gp = george.GP(kernel)
gp.compute(t, sigma)
# + tags=["hidden"]
# %%time
print(gp.log_likelihood(y))
# + tags=["hidden"]
# %%time
gp.sample()
# -
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 1c</h1>
# </div>
#
# ``george`` offers a fancy GP solver called the HODLR solver, which makes some approximations that dramatically speed up the matrix algebra. Instantiate the GP object again by passing the keyword ``solver=george.HODLRSolver`` and re-compute the log likelihood. How long did that take?
#
# (I wasn't able to draw samples using the HODLR solver; unfortunately this may not be implemented.)
# + tags=["hidden"]
# %%time
gp = george.GP(kernel, solver=george.HODLRSolver)
gp.compute(t, sigma)
# + tags=["hidden"]
# %%time
gp.log_likelihood(y)
# -
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 2</h1>
# </div>
#
# The ``george`` package is super useful for GP modeling, and I recommend you read over the [docs and examples](https://george.readthedocs.io/en/latest/). It implements several different [kernels](https://george.readthedocs.io/en/latest/user/kernels/) that come in handy in different situations, and it has support for multi-dimensional GPs. But if all you care about are GPs in one dimension (in this case, we're only doing GPs in the time domain, so we're good), then ``celerite`` is what it's all about:
#
# ```bash
# pip install celerite
# ```
#
# Check out the [docs](https://celerite.readthedocs.io/en/stable/) here, as well as several tutorials. There is also a [paper](https://arxiv.org/abs/1703.09710) that discusses the math behind ``celerite``. The basic idea is that for certain families of kernels, there exist **extremely efficient** methods of factorizing the covariance matrices. Whereas GP fitting typically scales with the number of datapoints $N$ as $N^3$, ``celerite`` is able to do everything in order $N$ (!!!) This is a **huge** advantage, especially for datasets with tens or hundreds of thousands of data points. Using ``george`` or any homebuilt GP model for datasets larger than about ``10,000`` points is simply intractable, but with ``celerite`` you can do it in a breeze.
#
# Repeat the timing tests, but this time using ``celerite``. Note that the Exponential Squared Kernel is not available in ``celerite``, because it doesn't have the special form needed to make its factorization fast. Instead, use the ``Matern 3/2`` kernel, which is qualitatively similar, and which can be approximated quite well in terms of the ``celerite`` basis functions:
#
# ```python
# kernel = celerite.terms.Matern32Term(np.log(1), np.log(1))
# ```
#
# Note that ``celerite`` accepts the **log** of the amplitude and the **log** of the timescale. Other than this, you should be able to compute the likelihood and draw a sample with the same syntax as ``george``.
#
# How much faster did it run?
# + tags=["hidden"]
import celerite
from celerite import terms
# + tags=["hidden"]
# %%time
kernel = terms.Matern32Term(np.log(1), np.log(1))
gp = celerite.GP(kernel)
gp.compute(t, sigma)
# + tags=["hidden"]
# %%time
gp.log_likelihood(y)
# + tags=["hidden"]
# %%time
gp.sample()
# -
# <div style="background-color: #D6EAF8; border-left: 15px solid #2E86C1;">
# <h1 style="line-height:2.5em; margin-left:1em;">Exercise 3</h1>
# </div>
#
# Let's use ``celerite`` for a real application: fitting an exoplanet transit model in the presence of correlated noise.
#
# Below is a (fictitious) light curve for a star with a transiting planet. There is a transit visible to the eye at $t = 0$, which (say) is when you'd expect the planet to transit if its orbit were perfectly periodic. However, a recent paper claims that the planet shows transit timing variations, which are indicative of a second, perturbing planet in the system, and that a transit at $t = 0$ can be ruled out at 3 $\sigma$. Your task is to verify this claim.
#
# Assume you have no prior information on the planet other than the transit occurs in the observation window, the depth of the transit is somewhere in the range $(0, 1)$, and the transit duration is somewhere between $0.1$ and $1$ day. You don't know the exact process generating the noise, but you are certain that there's correlated noise in the dataset, so you'll have to pick a reasonable kernel and estimate its hyperparameters.
#
#
# Fit the transit with a simple inverted Gaussian with three free parameters:
#
# ```python
# def transit_shape(depth, t0, dur):
# return -depth * np.exp(-0.5 * (t - t0) ** 2 / (0.2 * dur) ** 2)
# ```
#
# Read the celerite docs to figure out how to solve this problem efficiently.
#
# *HINT: I borrowed heavily from [this tutorial](https://celerite.readthedocs.io/en/stable/tutorials/modeling/), so you might want to take a look at it...*
# + tags=["hidden"]
import matplotlib.pyplot as plt
from celerite.modeling import Model
import os
# Define the model
class MeanModel(Model):
parameter_names = ("depth", "t0", "dur")
def get_value(self, t):
return -self.depth * np.exp(-0.5 * (t - self.t0) ** 2 / (0.2 * self.dur) ** 2)
mean_model = MeanModel(depth=0.5, t0=0.05, dur=0.7)
mean_model.parameter_bounds = [(0, 1.0), (-0.1, 0.4), (0.1, 1.0)]
true_params = mean_model.get_parameter_vector()
# Simuate the data
np.random.seed(71)
x = np.sort(np.random.uniform(-1, 1, 70))
yerr = np.random.uniform(0.075, 0.1, len(x))
K = 0.2 * np.exp(-0.5 * (x[:, None] - x[None, :]) ** 2 / 10.5)
K[np.diag_indices(len(x))] += yerr ** 2
y = np.random.multivariate_normal(mean_model.get_value(x), K)
y -= np.nanmedian(y)
# Plot the data
plt.errorbar(x, y, yerr=yerr, fmt=".k", capsize=0)
t = np.linspace(-1, 1, 1000)
plt.plot(t, mean_model.get_value(t))
plt.ylabel(r"$y$")
plt.xlabel(r"$t$")
plt.xlim(-1, 1)
plt.gca().yaxis.set_major_locator(plt.MaxNLocator(5))
plt.title("simulated data");
# Save it
X = np.hstack((x.reshape(-1, 1), y.reshape(-1, 1), yerr.reshape(-1, 1)))
if not (os.path.exists("data")):
os.mkdir("data")
np.savetxt("data/sample_transit.txt", X)
# -
import matplotlib.pyplot as plt
t, y, yerr = np.loadtxt("data/sample_transit.txt", unpack=True)
plt.errorbar(x, y, yerr=yerr, fmt=".k", capsize=0)
plt.xlabel("time")
plt.ylabel("relative flux");
# + tags=["hidden"]
from scipy.optimize import minimize
# Set up the GP model
kernel = terms.RealTerm(log_a=np.log(np.var(y)), log_c=0)
gp = celerite.GP(kernel, mean=mean_model, fit_mean=True)
gp.compute(x, yerr)
print("Initial log-likelihood: {0}".format(gp.log_likelihood(y)))
# Define a cost function
def neg_log_like(params, y, gp):
gp.set_parameter_vector(params)
return -gp.log_likelihood(y)
def grad_neg_log_like(params, y, gp):
gp.set_parameter_vector(params)
return -gp.grad_log_likelihood(y)[1]
# Fit for the maximum likelihood parameters
initial_params = gp.get_parameter_vector()
bounds = gp.get_parameter_bounds()
soln = minimize(neg_log_like, initial_params,
method="L-BFGS-B", bounds=bounds, args=(y, gp))
gp.set_parameter_vector(soln.x)
print("Final log-likelihood: {0}".format(-soln.fun))
# Make the maximum likelihood prediction
t = np.linspace(-1, 1, 500)
mu, var = gp.predict(y, t, return_var=True)
std = np.sqrt(var)
# Plot the data
color = "#ff7f0e"
plt.errorbar(x, y, yerr=yerr, fmt=".k", capsize=0)
plt.plot(t, mu, color=color)
plt.fill_between(t, mu+std, mu-std, color=color, alpha=0.3, edgecolor="none")
plt.ylabel(r"$y$")
plt.xlabel(r"$t$")
plt.xlim(-1, 1)
plt.gca().yaxis.set_major_locator(plt.MaxNLocator(5))
plt.title("maximum likelihood prediction");
# + tags=["hidden"]
def log_probability(params):
gp.set_parameter_vector(params)
lp = gp.log_prior()
if not np.isfinite(lp):
return -np.inf
try:
return gp.log_likelihood(y) + lp
except celerite.solver.LinAlgError:
return -np.inf
# + tags=["hidden"]
import emcee
initial = np.array(soln.x)
ndim, nwalkers = len(initial), 32
sampler = emcee.EnsembleSampler(nwalkers, ndim, log_probability)
print("Running burn-in...")
p0 = initial + 1e-8 * np.random.randn(nwalkers, ndim)
p0, lp, _ = sampler.run_mcmc(p0, 1000)
print("Running production...")
sampler.reset()
sampler.run_mcmc(p0, 2000);
# + tags=["hidden"]
# Plot the data.
plt.errorbar(x, y, yerr=yerr, fmt=".k", capsize=0)
# Plot 24 posterior samples.
samples = sampler.flatchain
for s in samples[np.random.randint(len(samples), size=24)]:
gp.set_parameter_vector(s)
mu = gp.predict(y, t, return_cov=False)
plt.plot(t, mu, color=color, alpha=0.3)
plt.ylabel(r"$y$")
plt.xlabel(r"$t$")
plt.xlim(-1, 1)
plt.gca().yaxis.set_major_locator(plt.MaxNLocator(5))
plt.title("posterior predictions");
# + tags=["hidden"]
import corner
names = gp.get_parameter_names()
cols = mean_model.get_parameter_names()
inds = np.array([names.index("mean:"+k) for k in cols])
corner.corner(sampler.flatchain[:, inds], truths=true_params,
labels=[r"depth", r"$t_0$", r"dur"]);
# + [markdown] tags=["hidden"]
# The transit time is inconsistent with 0 at about 3 sigma, so the claim that TTVs are present is *probably* true. But remember we're making strong assumptions about the nature of the correlated noise (we assumed a specific kernel), so in reality our uncertainty should be a bit higher.
| Session9/Day2/gps/answers/03-FastGPs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#本章需导入的模块
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import warnings
warnings.filterwarnings(action = 'ignore')
# %matplotlib inline
plt.rcParams['font.sans-serif']=['SimHei'] #解决中文显示乱码问题
plt.rcParams['axes.unicode_minus']=False
from sklearn.datasets import make_blobs
from sklearn.feature_selection import f_classif
from sklearn import decomposition
from sklearn.cluster import KMeans,AgglomerativeClustering,MeanShift, estimate_bandwidth
from sklearn.metrics import silhouette_score,calinski_harabasz_score
import scipy.cluster.hierarchy as sch
from itertools import cycle # python自带的迭代器模块
from matplotlib.patches import Ellipse
from sklearn.mixture import GaussianMixture
from scipy.stats.kde import gaussian_kde # ,multivariate_normal
# +
fig=plt.figure(figsize=(10,10))
N=400
K=4
X, y = make_blobs(n_samples=N, centers=K,cluster_std=0.60, random_state=0)
colors = cycle('bgrcmyk')
ax=plt.subplot(221)
for k, col in zip(range(K),colors):
plt.scatter(X[y == k, 0], X[y == k,1], c=col)
ax.set_title('数据的真实小类')
ax.set_xlabel("X1")
ax.set_ylabel("X2")
KM= KMeans(n_clusters=K, n_jobs = 4, max_iter = 500)
KM.fit(X)
lables = KM.labels_
ax=plt.subplot(222)
for k, col in zip(range(K),colors):
ax.scatter(X[lables == k, 0], X[lables == k,1], c=col)
ax.set_title('圆形小类下的K-均值聚类')
ax.set_xlabel("X1")
ax.set_ylabel("X2")
rng = np.random.RandomState(12)
X_stretched = np.dot(X, rng.randn(2, 2))
ax=plt.subplot(223)
for k, col in zip(range(K),colors):
plt.scatter(X_stretched[y == k, 0], X_stretched[y == k,1], c=col)
ax.set_title('数据的真实小类')
ax.set_xlabel("X1")
ax.set_ylabel("X2")
KM= KMeans(n_clusters=K, n_jobs = 4 , max_iter = 500)
KM.fit(X_stretched)
lables = KM.labels_
ax=plt.subplot(224)
for k, col in zip(range(K),colors):
plt.scatter(X_stretched[lables == k, 0], X_stretched[lables == k,1], c=col)
ax.set_title('非圆形小类下的K-均值聚类')
ax.set_xlabel("X1")
ax.set_ylabel("X2")
# -
# 代码说明:
# (1)第2至4行:生成聚类模拟数据,其中包含4个小类,样本量N=400。
# (2)第7至11行:以不同颜色展示数据中真实小类的构成情况。
# (3)第13至15行:采用K-均值聚类,将数据聚成4类。并获得聚类解。
# (4)第17至21行:可视化聚类解
# (5)第23行:定义一个伪随机数生成器对象,后续将用于生成随机数。
# (6)第24行:利用伪随机数生成器对象,生成2个来自二元标准高斯分布的随机数
# (7)第26至30行:以不同颜色展示数据中真实小类的构成情况。
# (8)第32至34行:采用K-均值聚类,将数据聚成4类。并获得聚类解。
# (9)第36至40行:可视化聚类解
# +
fig=plt.figure(figsize=(15,6))
np.random.seed(123)
N1,N2=500,1000
mu1,cov1,y1=[0,0],[[10,3],[3,10]],np.array([0]*N1)
set1= np.random.multivariate_normal(mu1,cov1,N1) #set1 = multivariate_normal(mean=mu1, cov=cov1,size=N1)
mu2,cov2,y2=[15,15],[[10,3],[3,10]],np.array([1]*N2)
set2=np.random.multivariate_normal(mu2,cov2,N2) #set2 = multivariate_normal(mean=mu2, cov=cov2,size=N2)
X=np.vstack([set1,set2])
y=np.vstack([y1.reshape(N1,1),y2.reshape(N2,1)])
ax=plt.subplot(121)
ax.scatter(X[:,0],X[:,1],s=40)
ax.set_title("%d个样本观测点的分布"%(N1+N2))
ax.set_xlabel("X1")
ax.set_ylabel("X2")
X1,X2= np.meshgrid(np.linspace(X[:,0].min(),X[:,0].max(),100), np.linspace(X[:,1].min(),X[:,1].max(),100))
X0=np.hstack((X1.reshape(len(X1)*len(X2),1),X2.reshape(len(X1)*len(X2),1)))
kernel = gaussian_kde(X.T) #要求为p*N形状 高斯核密度估计
Z = np.reshape(kernel(X0.T).T, X1.shape) #得到指定点的密度值
ax = fig.add_subplot(122, projection='3d')
ax.plot_wireframe(X1,X2,Z.reshape(len(X1),len(X2)),linewidth=0.5)
ax.plot_surface(X1,X2,Z.reshape(len(X1),len(X2)),alpha=0.3,rstride =50, cstride = 50)
ax.set_xlabel("X1")
ax.set_ylabel("X2")
ax.set_zlabel("密度")
ax.set_title("混合高斯分布")
# -
def draw_ellipse(position, covariance, ax=None, **kwargs):
"""用给定的位置和协方差画一个椭圆"""
ax = ax or plt.gca()
if covariance.shape == (2, 2):
U, s, Vt = np.linalg.svd(covariance)
angle = np.degrees(np.arctan2(U[1, 0], U[0, 0]))
width, height = 2 * np.sqrt(s)
else:
angle = 0
width, height = 2 * np.sqrt(covariance)
#画出椭圆
for nsig in range(1, 4):
ax.add_patch(Ellipse(position, nsig * width, nsig * height,angle, **kwargs))
# 代码说明:
# (1)第1行:定义用户自定义函数的函数名(draw_ellipse)和形式参数。主要包括椭圆的位置,以及由协方差矩阵决定的椭圆的长短轴和方向。
# (2)第4至7行:指定协方差矩阵对椭圆的对应关系。
# 如果协方差矩阵为2×2的矩阵(一般情况,除非指定聚类变量X_1,X_2的协方差等于0),对协方差矩阵进行奇异值分解。基于U成分计算反正切函数值并转换为度数,作为椭圆方向的角度。基于特征值计算椭圆的长短轴,分别为两个特征值的平方根。
# (3)第8至10行:如果协方差矩阵不是2×2的矩阵,即聚类变量X_1,X_2的协方差等于0,椭圆为水平方向放置,椭圆的长短轴分别为两个聚类变量的标准差。
# (4)第12,13行:利用for循环和函数Ellipse()依次绘制并叠加三个小椭圆,最终得到各小类的椭圆形状
#
gmm = GaussianMixture(n_components=2,covariance_type='full').fit(X)
labels = gmm.predict(X)
fig=plt.figure(figsize=(8,6))
plt.scatter(X[:,0], X[:,1], c=labels, s=40)
w_factor = 0.2 / gmm.weights_.max()
for i in np.unique(labels):
covar=gmm.covariances_[i]
pos=gmm.means_[i]
w=gmm.weights_[i]
draw_ellipse(pos, covar, alpha=w * w_factor)
plt.xlabel("X1")
plt.ylabel("X2")
plt.title("各小类的高斯分布")
probs = gmm.predict_proba(X)
print("前五个样本观测分属于两类的概率:\n{0}".format(probs[:5].round(3)))
# +
N=400
K=4
X, y = make_blobs(n_samples=N, centers=K,cluster_std=0.60, random_state=0)
gmm = GaussianMixture(n_components=K,covariance_type='full').fit(X)
labels = gmm.predict(X)
fig=plt.figure(figsize=(12,6))
ax=plt.subplot(121)
ax.scatter(X[:, 0], X[:, 1], c=labels, s=40)
w_factor = 0.2 / gmm.weights_.max()
for pos, covar, w in zip(gmm.means_, gmm.covariances_, gmm.weights_):
draw_ellipse(pos, covar, alpha=w * w_factor)
ax.set_title("各小类的高斯分布")
ax.set_xlabel("X1")
ax.set_ylabel("X2")
var='tied'
gmm = GaussianMixture(n_components=K,covariance_type=var).fit(X_stretched)
labels = gmm.predict(X_stretched)
ax=plt.subplot(122)
ax.scatter(X_stretched[:, 0], X_stretched[:, 1], c=labels, s=40)
w_factor = 0.2 / gmm.weights_.max()
if var=='tied': #四个分布共享合并的协方差阵
print(gmm.covariances_)
gmm.covariances_=[gmm.covariances_]*K
for pos, covar, w in zip(gmm.means_, gmm.covariances_, gmm.weights_):
draw_ellipse(pos, covar, alpha=w * w_factor)
ax.set_title("各小类的高斯分布")
ax.set_xlabel("X1")
ax.set_ylabel("X2")
# -
# 代码说明:
# (1)第4至14行:对前述的第一组模拟数据(图11.13左上图数据)进行基于高斯分布的EM聚类,并绘制各小类的椭圆。这里指定4个小类的聚类变量X_1,X_2的协方差矩阵不等。
# (2)第16,17行:对前述的第二组模拟数据(图11.13左下数据)进行基于高斯分布的EM聚类,这里指定4个小类的共享协方差矩阵。
# (3)第22,23行:为便于统一处理,将共享的协方差矩阵复制K份
#
| chapter11-4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Global
# +
b = 9
def foo():
print("inside foo", b)
foo()
print("main", b)
# -
# ### Local
# +
def bar():
y = 7
print("inside bar", y)
bar()
print("main", y) # expect error
# -
# ### Pay attention!
# +
b = 9
def magic():
y = 7
print("inside magic", y, b)
b = 6
print("main", b)
magic()
# +
b = 9
def magic_local():
y = 7
b = 6
print("inside magic_local", y, b)
magic_local()
print("main", b)
# -
# ### `global` keyword
# +
b = 9
def magic_global():
global b # <----
y = 7
print("inside magic_local", y, b)
b = 6
magic_global()
print("main", b)
| lectures/lectures/python/02_variables_and_functions/03_global_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: my-first-appyter-2
# language: python
# name: my-first-appyter-2
# ---
# #%%appyter init
from appyter import magic
magic.init(lambda _=globals: _())
# +
# %%appyter markdown
<center>
<h1 id = "top-of-app">
<div style="font-size:3rem;font-weight:500"> <img src="{{ url_for('static', filename='logo.png') }}" style="height:45px;padding:0 5px;display:inline"/> Gene Set Library Synopsis Appyter</div>
</h1>
<br>
<div style="font-size:2rem;font-weight:500">An appyter for the analysis and visualization of gene set libraries</div>
</center>
# +
# %%appyter markdown
# Gene Set Library Synopsis
This Appyter processes, analyzes, and visualizes a collection of gene sets, also known as a gene set library.
First it will generate summary statistics describing the size of the library and its component gene sets, as well as how well studied the genes and gene sets are.
Then the Appyter will use text vectorization (TF-IDF) and dimensionality reduction (UMAP) to visualize the library as a scatterplot.
To assess gene set similarity, pairwise Jaccard Indexes will be calculated, and this statistic will serve as the basis for a heatmap. The Appyter will also produce a set of figures focusing on the gene sets with the highest overlap.
Finally, the Appyter will present additional plots describing the composition of your library, including bar graphs of most frequent and most studied genes, a scatterplot of gene sets arranged by size and publication count, and a scatterplot of the library among all Enrichr libraries.
# +
#%% Imports
import appyter
import pandas as pd
import numpy as np
import base64
import math
import seaborn as sns
import fastcluster
import matplotlib.pyplot as plt; plt.rcdefaults()
import matplotlib.colors as colors
# %matplotlib inline
import IPython
from IPython.display import HTML, display, FileLink, Markdown, IFrame
import urllib
import itertools
from itertools import chain
from scipy.spatial.distance import squareform, pdist, jaccard
from scipy.cluster.hierarchy import linkage
from bokeh.io import output_notebook, export_svg
from bokeh.io.export import get_screenshot_as_png
from bokeh.plotting import figure, show
from bokeh.models import HoverTool, CustomJS, ColumnDataSource, Span, ranges, LabelSet, BasicTicker, ColorBar, LinearColorMapper, PrintfTickFormatter
from bokeh.layouts import layout, row, column, gridplot
from bokeh.palettes import all_palettes, linear_palette, Turbo256, Spectral6
from bokeh.transform import factor_cmap, transform
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.cluster import KMeans
from sklearn.metrics import silhouette_samples, silhouette_score
import umap.umap_ as umap
from sklearn.decomposition import NMF
output_notebook()
# +
# Notebook display functions
def figure_legend(label,title="",content=""):
if len(title)>0:
display(HTML(f"<div><b>{label}</b>: <i>{title}</i>. {content} </div>"))
else:
display(HTML(f"<div><b>{label}</b>: {content} </div>"))
# Figure and table counters
fig_count = 1
table_count = 1
# Output a table of genes and counts.
def create_download_link(df, title, filename):
csv = df.to_csv(index = False)
b64 = base64.b64encode(csv.encode())
payload = b64.decode()
html = '<a download="{filename}" href="data:text/csv;base64,{payload}" target="_blank">{title}</a>'
html = html.format(payload=payload, title=title, filename=filename)
return HTML(html)
# -
# %%appyter hide_code
{% do SectionField(
name='geneEntrySection',
title='1. Submit a Gene Set Library',
subtitle='Upload a GMT file containing a gene set library (gene names must be in gene symbol format) or select an existing Enrichr library. You may also choose to use the default gene library provided.',
img='data-upload-icon.png'
) %}
{% do SectionField(
name='barGraphSection',
title='2. Bar Graphs and Histograms',
subtitle='Bar graphs and histograms describing your library will be generated. Choose parameters to customize these visualizations',
img='bar-icon.png'
) %}
{% do SectionField(
name='simSection',
title='3. Gene Set Similarity',
subtitle='Similarity between gene sets will be assessed using the Jaccard Index. You may choose how this information is displayed.',
img='set-similarity.png'
) %}
# +
# %%appyter code_exec
# Inputting libraries and settings
{% set library_kind = TabField(
name = 'library_kind',
label = 'Library',
default = 'Upload a library',
description = '',
choices = {
'Upload a library': [
FileField(
name = 'library_filename',
label = 'Gene Library File (.gmt or .txt)',
default = 'CellMarker_Augmented_2021.gmt',
examples = {'CellMarker_Augmented_2021.gmt': url_for('static', filename = 'CellMarker_Augmented_2021.gmt'),
'CellMarker_Augmented_2021.txt': url_for('static', filename = 'CellMarker_Augmented_2021.txt')},
description = 'GMT is a tab-delimited file format that describes sets. Visit https://bit.ly/35crtXQ for more information and http://www.molmine.com/magma/fileformats.html to create your own.',
section = 'geneEntrySection'),
MultiCheckboxField(
name='species',
label='Species',
description='Which species are represented by your gene set library?',
default=['human'],
section='geneEntrySection',
choices=[
'human',
'mouse',
'other',
],
)
],
'Select a library from Enrichr': [
ChoiceField(
name = 'enrichr_library',
description = 'Select one Enrichr library whose genes will be counted',
label = 'Enrichr Library',
default = 'CellMarker_Augmented_2021',
section = 'geneEntrySection',
choices = [
'ARCHS4_Cell-lines',
'ARCHS4_IDG_Coexp',
'ARCHS4_Kinases_Coexp',
'ARCHS4_TFs_Coexp',
'ARCHS4_Tissues',
'Achilles_fitness_decrease',
'Achilles_fitness_increase',
'Aging_Perturbations_from_GEO_down',
'Aging_Perturbations_from_GEO_up',
'Allen_Brain_Atlas_10x_scRNA_2021',
'Allen_Brain_Atlas_down',
'Allen_Brain_Atlas_up',
'Azimuth_Cell_Types_2021',
'BioCarta_2013',
'BioCarta_2015',
'BioCarta_2016',
'BioPlanet_2019',
'BioPlex_2017',
'CCLE_Proteomics_2020',
'CORUM',
'COVID-19_Related_Gene_Sets',
'Cancer_Cell_Line_Encyclopedia',
'CellMarker_Augmented_2021',
'ChEA_2013',
'ChEA_2015',
'ChEA_2016',
'Chromosome_Location',
'Chromosome_Location_hg19',
'ClinVar_2019',
'dbGaP',
'DSigDB',
'Data_Acquisition_Method_Most_Popular_Genes',
'DepMap_WG_CRISPR_Screens_Broad_CellLines_2019',
'DepMap_WG_CRISPR_Screens_Sanger_CellLines_2019',
'Descartes_Cell_Types_and_Tissue_2021',
'DisGeNET',
'Disease_Perturbations_from_GEO_down',
'Disease_Perturbations_from_GEO_up',
'Disease_Signatures_from_GEO_down_2014',
'Disease_Signatures_from_GEO_up_2014',
'DrugMatrix',
'Drug_Perturbations_from_GEO_2014',
'Drug_Perturbations_from_GEO_down',
'Drug_Perturbations_from_GEO_up',
'ENCODE_Histone_Modifications_2013',
'ENCODE_Histone_Modifications_2015',
'ENCODE_TF_ChIP-seq_2014',
'ENCODE_TF_ChIP-seq_2015',
'ENCODE_and_ChEA_Consensus_TFs_from_ChIP-X',
'ESCAPE',
'Elsevier_Pathway_Collection',
'Enrichr_Libraries_Most_Popular_Genes',
'Enrichr_Submissions_TF-Gene_Coocurrence',
'Enrichr_Users_Contributed_Lists_2020',
'Epigenomics_Roadmap_HM_ChIP-seq',
'GO_Biological_Process_2013',
'GO_Biological_Process_2015',
'GO_Biological_Process_2017',
'GO_Biological_Process_2017b',
'GO_Biological_Process_2018',
'GO_Cellular_Component_2013',
'GO_Cellular_Component_2015',
'GO_Cellular_Component_2017',
'GO_Cellular_Component_2017b',
'GO_Cellular_Component_2018',
'GO_Molecular_Function_2013',
'GO_Molecular_Function_2015',
'GO_Molecular_Function_2017',
'GO_Molecular_Function_2017b',
'GO_Molecular_Function_2018',
'GTEx_Tissue_Sample_Gene_Expression_Profiles_down',
'GTEx_Tissue_Sample_Gene_Expression_Profiles_up',
'GWAS_Catalog_2019',
'GeneSigDB',
'Gene_Perturbations_from_GEO_down',
'Gene_Perturbations_from_GEO_up',
'Genes_Associated_with_NIH_Grants',
'Genome_Browser_PWMs',
'HMDB_Metabolites',
'HMS_LINCS_KinomeScan',
'HomoloGene',
'HuBMAP_ASCT_plus_B_augmented_w_RNAseq_Coexpression',
'HumanCyc_2015',
'HumanCyc_2016',
'Human_Gene_Atlas',
'Human_Phenotype_Ontology',
'huMAP',
'InterPro_Domains_2019',
'Jensen_COMPARTMENTS',
'Jensen_DISEASES',
'Jensen_TISSUES',
'KEA_2013',
'KEA_2015',
'KEGG_2013',
'KEGG_2015',
'KEGG_2016',
'KEGG_2019_Human',
'KEGG_2019_Mouse',
'Kinase_Perturbations_from_GEO_down',
'Kinase_Perturbations_from_GEO_up',
'L1000_Kinase_and_GPCR_Perturbations_down',
'L1000_Kinase_and_GPCR_Perturbations_up',
'LINCS_L1000_Chem_Pert_down',
'LINCS_L1000_Chem_Pert_up',
'LINCS_L1000_Ligand_Perturbations_down',
'LINCS_L1000_Ligand_Perturbations_up',
'Ligand_Perturbations_from_GEO_down',
'Ligand_Perturbations_from_GEO_up',
'lncHUB_lncRNA_Co-Expression',
'MCF7_Perturbations_from_GEO_down',
'MCF7_Perturbations_from_GEO_up',
'MGI_Mammalian_Phenotype_2013',
'MGI_Mammalian_Phenotype_2017',
'MGI_Mammalian_Phenotype_Level_3',
'MGI_Mammalian_Phenotype_Level_4',
'MGI_Mammalian_Phenotype_Level_4_2019',
'MSigDB_Computational',
'MSigDB_Hallmark_2020',
'MSigDB_Oncogenic_Signatures',
'Microbe_Perturbations_from_GEO_down',
'Microbe_Perturbations_from_GEO_up',
'miRTarBase_2017',
'Mouse_Gene_Atlas',
'NCI-60_Cancer_Cell_Lines',
'NCI-Nature_2015',
'NCI-Nature_2016',
'NIH_Funded_PIs_2017_AutoRIF_ARCHS4_Predictions',
'NIH_Funded_PIs_2017_GeneRIF_ARCHS4_Predictions',
'NIH_Funded_PIs_2017_Human_AutoRIF',
'NIH_Funded_PIs_2017_Human_GeneRIF',
'NURSA_Human_Endogenous_Complexome',
'OMIM_Disease',
'OMIM_Expanded',
'Old_CMAP_down',
'Old_CMAP_up',
'PanglaoDB_Augmented_2021',
'PPI_Hub_Proteins',
'Panther_2015',
'Panther_2016',
'Pfam_Domains_2019',
'Pfam_InterPro_Domains',
'PheWeb_2019',
'Phosphatase_Substrates_from_DEPOD',
'ProteomicsDB_2020',
'RNAseq_Automatic_GEO_Signatures_Human_Down',
'RNAseq_Automatic_GEO_Signatures_Human_Up',
'RNAseq_Automatic_GEO_Signatures_Mouse_Down',
'RNAseq_Automatic_GEO_Signatures_Mouse_Up',
'RNA-Seq_Disease_Gene_and_Drug_Signatures_from_GEO',
'Rare_Diseases_AutoRIF_ARCHS4_Predictions',
'Rare_Diseases_AutoRIF_Gene_Lists',
'Rare_Diseases_GeneRIF_ARCHS4_Predictions',
'Rare_Diseases_GeneRIF_Gene_Lists',
'Reactome_2013',
'Reactome_2015',
'Reactome_2016',
'SILAC_Phosphoproteomics',
'SubCell_BarCode',
'SysMyo_Muscle_Gene_Sets',
'TF-LOF_Expression_from_GEO',
'TF_Perturbations_Followed_by_Expression',
'TG_GATES_2020',
'TRANSFAC_and_JASPAR_PWMs',
'TRRUST_Transcription_Factors_2019',
'Table_Mining_of_CRISPR_Studies',
'TargetScan_microRNA',
'TargetScan_microRNA_2017',
'Tissue_Protein_Expression_from_Human_Proteome_Map',
'Tissue_Protein_Expression_from_ProteomicsDB',
'Transcription_Factor_PPIs',
'UK_Biobank_GWAS_v1',
'Virus-Host_PPI_P-HIPSTer_2020',
'VirusMINT',
'Virus_Perturbations_from_GEO_down',
'Virus_Perturbations_from_GEO_up',
'WikiPathways_2013',
'WikiPathways_2015',
'WikiPathways_2016',
'WikiPathways_2019_Human',
'WikiPathways_2019_Mouse'
]
)
],
},
section = 'geneEntrySection',
) %}
# Choose the orientation of the graph: horizontal or vertical bars
orient_bar = "{{ ChoiceField(name = 'orient_bar', label = 'Orientation', choices = ['Horizontal', 'Vertical'], default = 'Horizontal', description = 'Choose whether your bar graphs will be displayed horizontally or vertically', section = 'barGraphSection') }}"
# Choose color of bars
color_bar = "{{ ChoiceField(name = 'color_bar', label = 'Bar Color', choices = ['Black', 'Blue', 'Red', 'Green', 'Grey', 'Orange', 'Purple', 'Yellow', 'Pink'], default = 'Black', section = 'barGraphSection') }}"
# Choose whether gene counts are displayed on bar graph
counts_bar = {{ BoolField(name = 'counts_bar', label = 'Show Counts?', default = 'true', description = 'Choose \'Yes\' to label the bars with their lengths.', section = 'barGraphSection') }}
# Choose number of genes in bar graph
num_bar_genes = {{ IntField(
name='num_bar_genes',
label='Top Genes',
min=2,
max=20,
default=20,
description='The number of genes that will be included in figures describing top genes (ex: most frequent, most published)',
section='barGraphSection'
)}}
# Choose number of genes in bar graph
jac_cutoff = {{ FloatField(
name='jac_cutoff',
label='Jaccard Index High Threshold',
min=0.10,
max=0.99,
default=0.40,
description='The Jaccard Index will be calculated to measure similarity between sets in your library (0 = no shared genes, 1 = identical). Choose a threshold for what is considered a high Jaccard Index.',
section='simSection'
)}}
# Choose which visualizations are generated for most similar sets
jac_interactive = {{ BoolField(name = 'jac_interactive', label = 'Interactive Heatmap of Most Similar Sets?', default = 'true', description = 'Choose \'Yes\' to generate an interactive heatmap of the gene sets with similarity greater than the threshold you set above.', section = 'simSection') }}
# +
# Color for Bar plot
color_conversion = {
'Black': 'black',
'Blue': 'lightskyblue',
'Red': 'tomato',
'Green': 'mediumspringgreen',
'Grey': 'lightgrey',
'Orange': 'orange',
'Purple': 'plum',
'Yellow': 'yellow',
'Pink': 'lightpink'
}
bar_color = color_conversion[color_bar]
# +
# %%appyter code_exec
{%- if library_kind.raw_value == 'Upload a library' %}
library_kind = "Upload a library"
library_filename = {{ library_kind.value[0] }}
library_name = library_filename.replace("_", " ").replace(".txt", "").replace(".gmt", "")
species = {{library_kind.value[1]}}
{%- else %}
library_kind = "Select a library from Enrichr"
library_filename = "{{ library_kind.value[0] }}"
library_name = "{{ library_kind.value[0] }}"
species = ['human']
if 'MOUSE' in library_name.upper().split("_"):
species = ['mouse']
{%- endif %}
# -
# Download library from the Enrichr site using its file name
def download_library(library_filename):
with open(f"{library_filename}", "w") as fw:
with urllib.request.urlopen(f'https://maayanlab.cloud/Enrichr/geneSetLibrary?mode=text&libraryName={library_filename}') as f:
for line in f.readlines():
fw.write(line.decode('utf-8'))
fw.flush()
# +
# Load library
def remove_comma(gene):
try:
comma = gene.index(',')
return gene[0:comma]
except:
return gene
def load(library_filename, hasNA):
if library_kind == "Select a library from Enrichr":
download_library(library_filename)
library_data, library_genes, hasNA = load_library(library_filename, hasNA)
# to upper case
return library_data, library_genes, hasNA
# Returns a dictionary (library_data) where the values are all the elements
def load_library(library_filename, hasNA):
library_data = dict()
with open(library_filename, "r") as f:
lines = f.readlines()
library_genes = [''] * len(lines)
i = 0
for line in lines:
splited = line.strip().split("\t")
elements = pd.Series(splited[2:]).dropna()
if len(elements) > 0:
# to upper case
allxs = [x.upper() for x in elements]
allxs = pd.Series(allxs).apply(lambda x: remove_comma(x)).to_list()
if 'NA' in allxs:
allxs.remove('NA')
hasNA = True
library_data[splited[0]] = allxs
library_genes[i] = (' ').join(allxs)
i = i + 1
return library_data, library_genes, hasNA
# -
# Method for gene novelty
def gene_novelty_label(pub_count):
if pub_count <= 7:
return 'highly understudied'
if pub_count <= 25:
return 'understudied'
if pub_count <= 87:
return 'studied'
return 'well studied'
# +
# Create geneRIF dictionary and novelty mapping dictionaries
generif_df = pd.read_csv("https://appyters.maayanlab.cloud/storage/Gene_Set_Library_Synopsis/generif.tsv", delimiter="\t", header=None)
generif_df = generif_df.rename(columns={0:'Species',1:'Number',2:'Gene',3:'PMID',4:'Date'})
generif_genes = generif_df['Gene']
generif_s_genes = generif_genes.squeeze().str.upper()
generif_counts = generif_s_genes.value_counts()
generif_dict = generif_counts.to_dict()
novel_map_dict = {"highly understudied": 3, "understudied": 2, "studied": 1, "well studied": 0}
novel_map_dict_rev = {3: "highly understudied", 2: "understudied", 1: "studied", 0: "well studied"}
# +
# %%appyter code_exec
# Load library, create genes list, set list, gene size list
hasNA = False
library_data, library_genes, hasNA = load(library_filename, hasNA)
if library_kind == "Select a library from Enrichr":
library_name = library_name.replace("_", " ")
vals = list(library_data.values())
keys = list(library_data.keys())
all_genes = list(chain(*vals))
all_genes_unique = list(np.unique(np.array(all_genes)))
all_genes_unique = np.array(all_genes_unique)
all_sets = list(library_data.keys())
gs_sizes = [0]*len(vals)
for i in range(0, len(vals)):
gs_sizes[i] = len(vals[i])
# +
# Make dataframes of gene sets and their genes in 1) list form, 2) string form
library_data_onemap = dict()
library_data_onemap_str = dict()
for i in range(len(vals)):
library_data_onemap[keys[i]] = [vals[i]]
library_data_onemap_str[keys[i]] = (" ").join(vals[i])
library_data_onemap = pd.DataFrame(data=library_data_onemap).transpose()
library_data_onemap = library_data_onemap.rename(columns= {0:'Genes'})
library_data_onemap_str = pd.DataFrame(data={0:library_data_onemap_str})
library_data_onemap_str = library_data_onemap_str.rename(columns= {0:'Genes'})
# -
# %%appyter markdown
# 1. Unmapped Gene Names
This Appyter checks whether your gene set library contains unmapped gene names in _-DEC, _-MAR, and _-SEP formats. These conversions frequently occur when gene names are loaded into Excel. For example, either MARC1 or MARCH1 will automatically become '1-MAR'. Read this article for more information: https://genomebiology.biomedcentral.com/articles/10.1186/s13059-016-1044-7. This section also checks for genes labeled NA, which means Not Available.
# +
def month_sorter(month):
return month[-3]
def date_sorter(month):
dash = month.index('-')
return int(month[:dash])
# +
# Check for unmapped genes and display them
month_genes = all_genes_unique.copy()
month_genes.sort()
first = -1
last = -1
for i in range(len(month_genes)):
if len(month_genes[i]) > 4:
substr = month_genes[i][-4:]
if(substr == '-DEC' or substr == '-MAR' or substr == '-SEP'):
if first == -1:
first = i
last = i+1
else:
if first != -1:
break
month_genes = month_genes[first:last]
month_genes = sorted(month_genes,key=lambda x: (month_sorter(x), date_sorter(x)))
if hasNA:
month_genes.append('NA')
month_genes = pd.DataFrame(data=month_genes, columns=['Gene Name'])
# Display if unmapped genes
if len(month_genes) > 0:
month_genes_filename = 'unmapped_gene_names_' + library_name
found_genes_text = '' + str(len(month_genes)) + ' unmapped gene names found.'
display(Markdown(found_genes_text))
display(HTML(month_genes.to_html(index=False)))
figure_legend(f"Table {table_count}", content=f"Unmapped gene names in {library_name}")
display(create_download_link(month_genes, "Download this table as a CSV", month_genes_filename))
table_count = table_count + 1
else:
print("No unmapped gene names found")
# -
# %%appyter markdown
# 2. Descriptive Statistics
The Appyter will present descriptive statistics for your library such as: total genes, total gene sets, average genes per set, and frequency of each gene. Results will be displayed in downloadable tables.
# +
# Count the number of each gene
count_frame = pd.Series(all_genes).value_counts().sort_index().reset_index().reset_index(drop=True)
count_frame.columns = ['Gene', 'Count']
count_frame.dropna()
count_frame.sort_values(by=['Count'], inplace=True, ascending=False)
count_frame = count_frame.reset_index(drop=True)
# Drop skipped rows
mask = count_frame['Gene'].str.len() > 0
count_frame = count_frame.loc[mask]
count_frame = count_frame[~count_frame['Gene'].isin(['NA'])]
count_frame['Publications'] = count_frame['Gene'].map(generif_dict).replace(np.nan, 0)
count_frame['Publications'] = count_frame['Publications'].astype(int)
count_frame['Novelty'] = count_frame['Publications'].apply(lambda x: gene_novelty_label(x))
pubhist_dat = list(count_frame['Publications'].replace(0,np.nan).dropna())
# Make a copy to sort by publications
count_frame2 = count_frame.copy(deep=True)
count_frame2.sort_values(by=['Publications'], inplace=True, ascending=False)
top_genes = count_frame.iloc[0:num_bar_genes]
top_pub_genes = count_frame2.iloc[0:num_bar_genes]
# +
# Calculate novelty statistic for the library as a whole
count_frame2['Novelty Num'] = count_frame2['Novelty'].map(novel_map_dict)
novelty_weighted_sum = count_frame2['Count'] * count_frame2['Novelty Num']
novelty_weighted_sum = novelty_weighted_sum.sum()
count_sum = count_frame2['Count'].sum()
library_nov_exact = round(novelty_weighted_sum/count_sum, 1)
library_nov = math.floor(round(novelty_weighted_sum/count_sum, 1))
library_nov_term = novel_map_dict_rev[library_nov]
# +
# Make table describing gene sets
geneset_df = library_data_onemap.copy(deep=True)
geneset_df = geneset_df.reset_index()
geneset_df = geneset_df.rename(columns={'index':'Gene Set'})
geneset_df['Size'] = [0] * len(all_sets)
geneset_df['Mean Publications/Gene'] = [0] * len(all_sets)
geneset_df['Novelty'] = [''] * len(all_sets)
for i in range(0,len(geneset_df['Gene Set'])):
genes = geneset_df.iloc[i,1]
temp = count_frame2[count_frame2['Gene'].isin(genes)]
tot_pubs = sum(temp['Publications'])
av_pubs = np.mean(temp['Publications'])
tot_genes = len(genes)
novelty_num = sum(temp['Novelty Num']/tot_genes)
novelty = library_nov_term = novel_map_dict_rev[math.floor(novelty_num)]
geneset_df['Size'][i] = tot_genes
geneset_df['Mean Publications/Gene'][i] = av_pubs
geneset_df['Novelty'][i] = novelty
geneset_df = geneset_df.sort_values(by='Size', ascending=False).drop(columns=['Genes']).reset_index(drop=True)
# +
# Descriptive statistics summary table
# Totals
unique_genes = len(count_frame['Gene'])
unique_sets = len(library_data.keys())
avg_genes = round(np.mean(gs_sizes), 2)
median_genes = round(np.median(gs_sizes), 2)
median_publications = round(np.median(count_frame['Publications']), 2)
avg_publications = round(np.mean(count_frame['Publications']), 2)
# Novelty counts and percentages
novelty_counts = count_frame['Novelty'].value_counts()
novelty_counts_gs = geneset_df['Novelty'].value_counts()
tot_pub = np.sum(count_frame['Publications'])
genes_nov_dict = {'highly understudied': 0, 'understudied': 0, 'studied': 0, 'well studied':0}
gs_nov_dict = {'highly understudied': 0, 'understudied': 0, 'studied': 0, 'well studied':0}
genes_nov_pcnt_dict = {'highly understudied': '', 'understudied': '', 'studied': '', 'well studied':''}
gs_nov_pcnt_dict = {'highly understudied': '', 'understudied': '', 'studied': '', 'well studied':''}
# Reassign counts of highly understudied, understudied, etc. only if at least one gene of that type exists
# Calculate percentages of each novelty
for key in genes_nov_dict.keys():
if key in novelty_counts:
genes_nov_dict[key] = novelty_counts[key]
if key in novelty_counts_gs:
gs_nov_dict[key] = novelty_counts_gs[key]
genes_nov_pcnt_dict[key] = str(round(genes_nov_dict[key]/unique_genes * 100, 2)) + "%"
gs_nov_pcnt_dict[key] = str(round(gs_nov_dict[key]/unique_sets * 100, 2)) + "%"
## Load stats for all human and mouse genes
all_human_genes = pd.read_csv("https://appyters.maayanlab.cloud/storage/Gene_Set_Library_Synopsis/all_human_genes_df.csv", header=0)
all_mouse_genes = pd.read_csv("https://appyters.maayanlab.cloud/storage/Gene_Set_Library_Synopsis/all_mouse_genes_df.csv", header=0)
# Change table display based on library composition
other = False
specname = 'Human'
all_species_genes=all_human_genes
if 'human' not in species:
if 'mouse' in species:
all_species_genes = all_mouse_genes
specname = 'Mouse'
if species==['other']:
other = True
spec_col_name = 'All ' + specname + ' Genes'
## Make and display tables
lib_vs_spec_title_col = ["Total Genes", "Total Publications", "Highly Understudied Genes", "Understudied Genes", "Studied Genes", "Well Studied Genes"]
lib_col = [unique_genes, tot_pub, genes_nov_dict['highly understudied'], genes_nov_dict['understudied'], genes_nov_dict['studied'], genes_nov_dict['well studied']]
lib_col_pcnt = ['', '', genes_nov_pcnt_dict['highly understudied'], genes_nov_pcnt_dict['understudied'], genes_nov_pcnt_dict['studied'], genes_nov_pcnt_dict['well studied']]
lib_vs_spec_df = pd.DataFrame(data = {'': lib_vs_spec_title_col, f"{library_name}": lib_col, ' ': lib_col_pcnt})
lib_col = [avg_publications, median_publications]
lib_vs_spec_mm_df = pd.DataFrame(data = {'': ["Mean Publications/Gene", "Median Publications/Gene"], f"{library_name}": lib_col})
if not other:
spec_col = [all_species_genes['Genes'][0], all_species_genes['Publications'][0], all_species_genes['Highly Understudied'][0], all_species_genes['Understudied'][0], all_species_genes['Studied'][0], all_species_genes['Well Studied'][0]]
spec_col_pcnt = ['', '', str(all_species_genes['Highly Understudied'][1]) + "%", str(all_species_genes['Understudied'][1]) + "%", str(all_species_genes['Studied'][1]) + "%", str(all_species_genes['Well Studied'][1]) + "%"]
lib_col = [unique_genes, tot_pub, genes_nov_dict['highly understudied'], genes_nov_dict['understudied'], genes_nov_dict['studied'], genes_nov_dict['well studied']]
lib_col_pcnt = ['', '', genes_nov_pcnt_dict['highly understudied'], genes_nov_pcnt_dict['understudied'], genes_nov_pcnt_dict['studied'], genes_nov_pcnt_dict['well studied']]
lib_vs_spec_df = pd.DataFrame(data = {'': lib_vs_spec_title_col, f"{library_name}": lib_col, ' ': lib_col_pcnt, spec_col_name: spec_col, ' ': spec_col_pcnt})
lib_vs_spec_df[spec_col_name] = lib_vs_spec_df[spec_col_name].astype(int)
lib_col = [avg_publications, median_publications]
spec_col = [all_species_genes['Mean Publications'][0], all_species_genes['Median Publications'][0]]
lib_vs_spec_mm_df = pd.DataFrame(data = {'': ["Mean Publications/Gene", "Median Publications/Gene"], f"{library_name}": lib_col, spec_col_name: spec_col})
genestat_title_col = ["Gene Sets", "Highly Understudied Sets", "Understudied Sets", "Studied Sets", "Well Studied Sets"]
genestat_col = [unique_sets, gs_nov_dict['highly understudied'], gs_nov_dict['understudied'], gs_nov_dict['studied'], gs_nov_dict['well studied']]
genestat_col_pcnt = ['', gs_nov_pcnt_dict['highly understudied'], gs_nov_pcnt_dict['understudied'], gs_nov_pcnt_dict['studied'], gs_nov_pcnt_dict['well studied']]
genestat_df = pd.DataFrame(data = {'': genestat_title_col, 'Total': genestat_col, ' ':genestat_col_pcnt})
genestat_title_col = ["Mean", "Median"]
genestat_col = [avg_genes, median_genes]
genestat_mm_df = pd.DataFrame(data = {'': genestat_title_col, 'Genes / Set': genestat_col})
display(HTML(lib_vs_spec_df.to_html(index=False)))
display(HTML(lib_vs_spec_mm_df.to_html(index=False)))
if other:
figure_legend(f"Tables {table_count}A, {table_count}B", title=f"{library_name} Summary Statistics", content="Descriptive statistics for your library. Novelty ratings are based on Geneshot. Highly understudied genes are associated with 0-7 PubMed IDs (PMIDs); understudied genes with 8-25 PMIDs; studied genes with 26-87 PMIDs; and well studied genes with 88+ PMIDs.")
else:
figure_legend(f"Tables {table_count}A, {table_count}B", title=f"{library_name} vs. All {specname} Genes", content=f"Descriptive statistics comparing your gene set library to the set of all {specname} genes in GeneRIF. Novelty ratings are based on Geneshot. Highly understudied genes are associated with 0-7 PubMed IDs (PMIDs); understudied genes with 8-25 PMIDs; studied genes with 26-87 PMIDs; and well studied genes with 88+ PMIDs.")
table_count = table_count + 1
display(HTML(genestat_df.to_html(index=False)))
display(HTML(genestat_mm_df.to_html(index=False)))
figure_legend(f"Tables {table_count}A, {table_count}B", title=f"Summary Statistics for Gene Sets in {library_name} Library", content="Statistics describing the composition of gene sets within your library. Novelty ratings were calculated by giving each gene in each set a numerical novelty score (based on its rating of highly understudied, understudied, studied, or well studied), taking a weighted average of those scores, and translating the result back into a term that describes the entire gene set.")
table_count = table_count + 1
# +
def make_bok_hist(title, hist, edges, dat, xaxis_lab, yaxis_lab, tooltips, fill_color,xtype='auto', ytype='auto', xtail=5, ytail=5):
yrange = 0
if ytype=='log':
hist, edges = np.histogram(dat, density=False, bins=10)
ycap = math.ceil(math.log10(max(pubhist_dat)))
yrange=(10**0, 10**ycap)
ordered_hist = sorted(hist)
ordered_edges = sorted(edges)
if ytype=='auto':
yrange=[ordered_hist[0], ordered_hist[-1]+ytail]
p = figure(title=title, tooltips=tooltips, background_fill_color="#fafafa", toolbar_location="below",x_axis_type = xtype,y_axis_type = ytype,
x_range=[ordered_edges[0], ordered_edges[-1]+xtail], y_range=yrange)
p.quad(top=hist, bottom=0, left=edges[:-1], right=edges[1:],
fill_color=fill_color, line_color="white", hover_alpha=0.7)
p.y_range.start = 0
p.xaxis.axis_label = xaxis_lab
p.yaxis.axis_label = yaxis_lab
p.grid.grid_line_color="white"
p.title.align = 'center'
return p
def get_bin_params(dat_max, dat_min = 0, bins=40):
rng = dat_max - dat_min
needed = bins - rng%bins
new_rng = rng + needed
maxval = dat_min + new_rng
if needed==bins:
maxval = dat_max
new_rng = rng
interval = new_rng/bins
return maxval, interval
# +
# %%appyter code_exec
# Print out a table of the count_frame dataframe
counts_filename = library_name.replace(" ", "_") + "_gene_counts.csv"
display(HTML(count_frame[0:num_bar_genes].to_html(index=False)))
figure_legend(f"Table {table_count}", title=f"Gene count results for {library_name}", content="This table displays the counts (number of appearances throughout the entire library), number of publication associations (PMIDs), and novelty ratings of each gene. The full chart is also available for download. A weighted average statistic has been used to assign a novelty rating to your library as a whole, representing how well-studied the library is.")
table_count = table_count + 1
display(create_download_link(count_frame, "Download this table as a CSV", counts_filename))
print(f"Based on the novelty and within-library frequencies of each gene, your gene set library is \033[1m{library_nov_term}.")
# -
# Display table of gene set statistics
geneset_df_filename = library_name.replace(" ", "_") + "_gene_set_statistics.csv"
display(HTML(geneset_df.iloc[0:10,:].to_html(index=False)))
figure_legend(f"Table {table_count}", f"Gene Set Statistics in {library_name} Library", "Size (number of genes), average publications per gene, and novelty (calculated using a weighted average across all genes in the set) for each gene set in your library.")
display(create_download_link(geneset_df, "Download this table as a CSV", geneset_df_filename))
table_count = table_count + 1
# %%appyter markdown
# 3. Scatterplot Visualization
In this section, the gene sets in your library will be converted into numerical vectors using TF-IDF, transformed into two dimensions using UMAP, and visualized as a scatterplot.
# +
df = library_data_onemap_str.reset_index().rename(columns={'index':'Name'})
gene_list = df['Genes']
try:
tfidf_vectorizer = TfidfVectorizer(
min_df = 3,
max_df = 0.005,
max_features = 100000,
ngram_range=(1, 1)
)
tfidf = tfidf_vectorizer.fit_transform(gene_list)
except:
factor = 0.005
while factor*unique_sets < 3:
factor = factor + .005
tfidf_vectorizer = TfidfVectorizer(
min_df = 3,
max_df = factor*unique_sets,
max_features = 100000,
ngram_range=(1, 1)
)
tfidf = tfidf_vectorizer.fit_transform(gene_list)
reducer = umap.UMAP()
reducer.fit(tfidf)
embedding = pd.DataFrame(reducer.transform(tfidf), columns=['x','y'])
embedding = pd.concat([embedding, df], axis=1)
# -
# Prepare dimensionality-reduced matrix for clutstering
mapped_df = embedding.copy(deep=True)
mapped_df = mapped_df.set_index('Name')
mapped_df = mapped_df.drop(columns=['Genes'])
mapped_df = mapped_df.rename_axis("Gene Set").reset_index()
# +
# Plot clustered gene sets
xlabel = 'UMAP Dimension 1'
ylabel = 'UMAP Dimension 2'
source2 = ColumnDataSource(
data=dict(
x = mapped_df.x,
y = mapped_df.y,
alpha = [0.7] * mapped_df.shape[0],
size = [7] * mapped_df.shape[0],
gene_set = mapped_df['Gene Set'],
)
)
hover_emb = HoverTool(names=["df"], tooltips="""
<div style="margin: 10">
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Gene Set:</span>
<span style="font-size: 12px">@gene_set</span>
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Coordinates:</span>
<span style="font-size: 12px">(@x,@y)</span>
</div>
</div>
""")
tools_emb = [hover_emb, 'pan', 'wheel_zoom', 'reset', 'save']
title_emb = 'Gene Sets in ' + library_name + ' Library'
plot_emb = figure(plot_width=1000, plot_height=700, tools=tools_emb, title=title_emb, x_axis_label=xlabel, y_axis_label=ylabel)
plot_emb.circle('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source2, name="df",
fill_color='grey')
plot_emb.xaxis.axis_label_text_font_style = 'normal'
plot_emb.xaxis.axis_label_text_font_size = '18px'
plot_emb.yaxis.axis_label_text_font_size = '18px'
plot_emb.yaxis.axis_label_text_font_style = 'normal'
plot_emb.title.align = 'center'
plot_emb.title.text_font_size = '18px'
show(plot_emb)
figure_legend(f"Fig. {fig_count}", f"Scatterplot of Gene Sets in {library_name} Library", "Gene sets plotted by their UMAP dimensions.")
fig_count = fig_count + 1
# -
# %%appyter markdown
# 4. Set Similarity
In this section, the Appyter will compute the pairwise Jaccard index for every pair of gene sets in your library as a measure of set similarity. An index closer to 1 means the gene sets have most genes in common. An index closer to 0 indicates the gene sets share very few terms. The Jaccard indices will serve as the basis for a heatmap. Additional visualizations will be generated for the most similar sets (those with Jaccard indices above the user-specified threshold).
# +
# Put all sets and genes into dataframe where each thing is a list
def jaccard_list(u,v):
setu = set(chain(*u))
setv = set(chain(*v))
return len(setu.intersection(setv)) / len(setu.union(setv))
res = pdist(library_data_onemap[['Genes']], jaccard_list)
distance = pd.DataFrame(squareform(res), index=library_data_onemap.index, columns= library_data_onemap.index)
# Check whether any sets have Jaccard Index > 0; if not, skip all Jaccard plots
jac_zero_tester = pd.Series(res)
jac_zero_tester = jac_zero_tester.replace(0, np.nan)
jac_zero_tester = jac_zero_tester.dropna()
jac_zero = len(jac_zero_tester)
# +
# Filter "distance" to put NA where col# <= row#
mask = np.zeros_like(distance, dtype=bool)
mask[np.triu_indices_from(mask)] = True
masked_dist = distance.mask(mask, '').transpose()
masked_dist_filename = library_name.replace(" ", "_") + "_jaccard_matrix.csv"
display(HTML(masked_dist.iloc[0:10,0:10].to_html(index=True)))
figure_legend(f"Table {table_count}", f"Jaccard Index pairings for {library_name}", "Upper triangle of a pairwise Jaccard matrix comparing each set in the library to each other set.")
display(create_download_link(masked_dist.reset_index(), "Download this table as a CSV", masked_dist_filename))
table_count = table_count + 1
# -
if jac_zero > 0:
maxval, interval = get_bin_params(max(res)*1000)
maxval = maxval/1000
interval = interval/1000
jac_hist_bin_arr = np.arange(0, maxval, interval)
#jac_hist_bin_arr = np.arange(0, max(res)+.015, .01)
hist, edges = np.histogram(res, density=False, bins=jac_hist_bin_arr)
title = f"Jaccard Indices for {library_name}"
tooltips = [
("range", "@left{0.00}" + "-" + "@right{0.00}"),
("pairs", "@top")
]
xaxis_lab = 'Jaccard Index'
yaxis_lab = 'Gene Set Pairs'
jac_hist = make_bok_hist(title, hist, edges, res, xaxis_lab, yaxis_lab, tooltips, bar_color, xtail=max(edges)/15, ytail=max(hist)/15)
else:
print("All pairwise Jaccard Indices in your library are equal to 0. This means that no gene set in your library shares a single term (gene) with another gene set. The remaining tables and visualizations in this section of the analysis will not be generated.")
def make_bok_heatmap(df, title):
colors = ['#ffffb2', '#fed976', '#feb24c', '#fd8d3c', '#f03b20', '#bd0026']
mapper = LinearColorMapper(palette=colors, low=df.jaccard.min(), high=round(df.jaccard.max(), 1), nan_color="white")
rng = pd.unique(df['set1'])
pwidth = 1000* int(math.ceil((len(rng)/(jac_cutoff*100))))
pheight = 700* int(math.ceil((len(rng)/(jac_cutoff*100))))
if pwidth == 0:
pwidth = 1000
if pheight == 0:
pheight = 500
source = ColumnDataSource(df)
p = figure(title=title,
x_range=rng, y_range=list(reversed(rng)),
x_axis_location="above", plot_width=pwidth, plot_height=pheight,
toolbar_location='below',
tooltips=[
('Set 1', '@set1'),
('Set 2', '@set2'),
('Jaccard Index', '@jaccard')
])
p.grid.grid_line_color = None
p.axis.axis_line_color = None
p.axis.major_tick_line_color = None
p.axis.major_label_text_font_size = "7px"
if len(df['set1']) < 15:
p.axis.major_label_text_font_size = "10px"
p.axis.major_label_standoff = 0
p.xaxis.major_label_orientation = np.pi/2
p.title.align = 'center'
p.rect(x = 'set2', y = 'set1', width=1, height=1,
source=source,
fill_color=transform('jaccard', mapper),
line_color=None)
color_bar = ColorBar(color_mapper=mapper, major_label_text_font_size="12px",
ticker=BasicTicker(desired_num_ticks=len(colors)),
formatter=PrintfTickFormatter(format="%.2f"),
label_standoff=6, border_line_color=None)
p.add_layout(color_bar, 'right')
return(p)
if jac_zero > 0:
res_cut = res[res>jac_cutoff]
if len(res_cut) > 0:
datmin = math.floor(jac_cutoff * 100)*10
datmax = math.ceil(max(res_cut)*100)*10
maxval, interval = get_bin_params(datmax, dat_min=datmin)
datmin = datmin/1000
maxval = maxval/1000
interval = interval/1000
#jac_hist_cut_bin_arr = np.arange(math.floor(jac_cutoff * 100)/100.0, max(res_cut)+.01, .005)
jac_hist_cut_bin_arr = np.arange(datmin, maxval, interval)
hist_cut, edges_cut = np.histogram(res_cut, density=False, bins=jac_hist_cut_bin_arr)
title_cut = f"Jaccard Indices > {jac_cutoff} for {library_name}"
tooltips_cut = [
("range", "@left{0.000}" + "-" + "@right{0.000}"),
("pairs", "@top")
]
jac_hist_cut = make_bok_hist(title_cut, hist_cut, edges_cut, res_cut, xaxis_lab, yaxis_lab, tooltips_cut, bar_color, xtail=max(edges_cut)/15, ytail=max(hist_cut)/15)
else:
print(f"There are no gene set pairs with a Jaccard Index greater than {jac_cutoff} in your library. The corresponding tables and visualizations will not be generated.")
# If user wants interactive Jaccard heatmap for high indices, create one
if jac_interactive and jac_zero > 0:
dist_cut_indexes = np.where(distance > jac_cutoff)
dist_cut_rows_cols = np.unique(np.array(list(chain(*list(dist_cut_indexes)))))
dist_cut = distance.iloc[dist_cut_rows_cols, dist_cut_rows_cols]
mask2 = np.zeros_like(dist_cut, dtype=bool)
mask2[np.triu_indices_from(mask2)] = True
dist_cut_masked = dist_cut.mask(mask2, np.nan).transpose()
dist_set1 = np.array(np.repeat(dist_cut_masked.index, len(dist_cut_masked.index)))
sep = ",,"
dist_set2 = (sep.join(list(dist_cut_masked.index))+",,")*len(list(dist_cut_masked.index))
dist_set2 = (dist_set2.split(",,"))[0:-1]
dist_vals = list(chain(*list(dist_cut_masked.values)))
dist_heat_df = pd.DataFrame(data={'set1': dist_set1, 'set2': dist_set2, 'jaccard': dist_vals})
dist_heat_title = f"Jaccard Indices for {library_name}"
# Make smaller heatmap
jac_heat_cut = make_bok_heatmap(dist_heat_df, dist_heat_title)
# Show Jaccard histograms
if jac_zero > 0:
if len(res_cut) > 0:
show(row(jac_hist, jac_hist_cut))
figure_legend(f"Fig. {fig_count} and {fig_count +1}","Jaccard Indices Histograms", content=f"The histogram on the left displays the full range of Jaccard Indices for your library. The histogram on the right displays only those indices greater than {jac_cutoff}.")
fig_count = fig_count + 2
else:
show(jac_hist)
figure_legend(f"Fig. {fig_count}","Jaccard Indices Histogram", content=f"This histogram displays the full range of Jaccard Indices for your library. There were no indices greater than {jac_cutoff}.")
fig_count = fig_count + 1
# Table of highest Jaccard Indices
if jac_zero > 0:
dist_heat_df_disp = dist_heat_df[dist_heat_df['jaccard'] > jac_cutoff].reset_index(drop=True)
dist_heat_df_disp = dist_heat_df_disp.rename(columns={"set1": "Gene Set 1", "set2": "Gene Set 2", "jaccard": "Jaccard Index"})
display(HTML(dist_heat_df_disp.sort_values(by="Jaccard Index").iloc[0:10,:].to_html(index=False)))
figure_legend(f"Table {table_count}",f"Jaccard Indices > {jac_cutoff}", content=f"This table displays all gene set pairings with a Jaccard Index greater than {jac_cutoff}.")
high_jac_filename = library_name.replace(" ", "_") + "_high_jaccard_indices.csv"
display(create_download_link(dist_heat_df_disp.sort_values(by="Jaccard Index"), "Download this table as a CSV", high_jac_filename))
table_count = table_count + 1
# Get row indices in order and offer in table
def sns_heatmap_to_df(g):
rows = g.dendrogram_row.reordered_ind
cmap_df = distance.reset_index().rename(columns={'index': 'gene set'}).reset_index().rename(columns={'index': 'original index'})
cmap_df_dict = dict(zip(cmap_df['original index'], cmap_df['gene set']))
cmap_df = pd.DataFrame(data={'original index': rows})
cmap_df['gene set'] = cmap_df['original index'].map(cmap_df_dict)
cmap_df_ret = pd.DataFrame(data={'gene set': cmap_df['gene set'],
'original index':cmap_df['original index'],
'new index':cmap_df.index})
return cmap_df_ret
# +
# Clustered heatmap of full library- only if selected, or if dendrogram is not possible
sns_clust = None
full_heat_possible = True
if jac_zero > 0 and unique_sets > 0:
try:
sns_clust = sns.clustermap(distance, cmap="Reds", figsize=(13,13))
except:
print("Unable to generate heatmap. Try a smaller library.")
full_heat_possible = False
if full_heat_possible:
sns_clust.ax_row_dendrogram.set_visible(False)
sns_clust.ax_col_dendrogram.set_visible(False)
sns_clust.ax_cbar.set_position((0, 0, .03, .4))
figure_legend(f"Fig. {fig_count}", "Heatmap", f"This heatmap displays the Jaccard Indices of all gene sets in your library.")
fig_count = fig_count + 1
sns_clust_filename = library_name.replace(" ", "_") + "_jaccard_heatmap.png"
plt.savefig(sns_clust_filename, bbox_inches = 'tight')
display(FileLink(sns_clust_filename, result_html_prefix = str('Download png' + ': ')))
# -
if full_heat_possible:
cmap_df = sns_heatmap_to_df(sns_clust)
cmap_filename = library_name.replace(" ", "_") + "_jaccard_heatmap_reordered_gene_sets.csv"
display(HTML(cmap_df.head().to_html(index=False)))
figure_legend(f"Table {table_count}", f"Reordered gene sets in heatmap of {library_name}", "This table lists your gene sets in the order in which they appear in the heatmap. The full table is available for download.")
display(create_download_link(cmap_df, "Download this table as a CSV", cmap_filename))
table_count = table_count + 1
# +
# If interactive heatmap is possible, display it. Otherwise, create a static heatmap and report the new indices.
jac_static_heat=True
if jac_interactive and jac_zero > 0 and len(res_cut) > 0:
if len(dist_cut_rows_cols) < 300:
show(jac_heat_cut)
figure_legend(f"Fig. {fig_count}", "High Jaccard Indices Heatmap", f"This heatmap includes all gene sets with at least one Jaccard Index greater than {jac_cutoff} in comparison with another set.")
fig_count = fig_count + 1
jac_static_heat=False
else:
print("There are too many sets to generate an interactive figure. A static heatmap will be generated instead. To see an interactive heatmap of the highest Jaccard Indices, try selecting a higher threshold value.")
if jac_static_heat:
cmap_cut = sns.clustermap(dist_cut, cmap="Reds")
cmap_cut.ax_row_dendrogram.set_visible(False)
cmap_cut.ax_col_dendrogram.set_visible(False)
cmap_cut.ax_cbar.set_position((0.8, 0, .03, .4))
figure_legend(f"Fig. {fig_count}", "High Jaccard Indices Heatmap", f"This heatmap includes all gene sets with at least one Jaccard Index greater than {jac_cutoff} in comparison with another set.")
fig_count = fig_count + 1
# -
# %%appyter markdown
# 5. Visualization of Novelty and Size Distributions
# Read table of all Enrichr library novelties and append the user's library
all_enrichr_novs = pd.read_csv("https://appyters.maayanlab.cloud/storage/Gene_Set_Library_Synopsis/enrichr_library_novelties.csv", header=0)
all_enrichr_novs = all_enrichr_novs.sort_values(by='Stat', ascending=False)
my_lib_nov = pd.DataFrame(data={'Library':[library_name], 'Novelty':[library_nov_term], 'Stat':[library_nov_exact], 'Genes': [len(all_genes_unique)]})
mouse_libs = ['KEGG 2019 Mouse',
'Mouse Gene Atlas',
'RNAseq Automatic GEO Signatures Mouse Down',
'RNAseq Automatic GEO Signatures Mouse Up',
'WikiPathways 2019 Mouse']
all_enrichr_novs_mouse = all_enrichr_novs[all_enrichr_novs['Library'].isin(mouse_libs)]
all_enrichr_novs_mix = all_enrichr_novs[~all_enrichr_novs['Library'].isin(mouse_libs)]
# +
# Display gene set size and novelty distribution as a scatterplot
novelties = ['well studied', 'studied', 'understudied', 'highly understudied']
xlabel = 'Set Size (Genes)'
ylabel = 'Mean Publications Per Gene'
my_gs_stat_df = pd.DataFrame(data={'Gene Set':['Average of all gene sets'], 'Novelty':[library_nov_term], 'Size':[avg_genes], 'Publications/Gene': [int(round(np.mean(geneset_df['Mean Publications/Gene'],0)))]})
geneset_df['Novelty Num'] = geneset_df['Novelty'].map(novel_map_dict)
geneset_df = geneset_df.sort_values(by='Novelty Num', ascending=False).reset_index(drop=True).drop(columns=['Novelty Num'])
source1 = ColumnDataSource(
data=dict(
x = geneset_df.Size,
y = geneset_df['Mean Publications/Gene'],
alpha = [0.9] * geneset_df.shape[0],
size = [7] * geneset_df.shape[0],
novelty = geneset_df.Novelty,
geneset = geneset_df['Gene Set'],
)
)
source2 = ColumnDataSource(
data=dict(
x = my_gs_stat_df.Size,
y = my_gs_stat_df['Publications/Gene'],
alpha = [0.7] * my_gs_stat_df.shape[0],
size = [10] * my_gs_stat_df.shape[0],
novelty = my_gs_stat_df.Novelty,
geneset = my_gs_stat_df['Gene Set']
)
)
#print(embedding.shape[0])
hover_gs_nov = HoverTool(names=["df"], tooltips="""
<div style="margin: 10">
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Gene Set:</span>
<span style="font-size: 12px">@geneset</span>
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Novelty:</span>
<span style="font-size: 12px">@novelty</span>
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Set Size (Genes):</span>
<span style="font-size: 12px">@x</span>
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Mean Publications Per Gene:</span>
<span style="font-size: 12px">@y</span>
</div>
</div>
""")
tools_gs_nov = [hover_gs_nov, 'pan', 'wheel_zoom', 'reset', 'save']
title_gs_nov = f"Novelties and Sizes of Gene Sets Within {library_name} Library"
plot_gs_nov = figure(plot_width=700, plot_height=700, tools=tools_gs_nov, title=title_gs_nov, x_axis_label=xlabel, y_axis_label=ylabel, x_axis_type='log', y_axis_type='log')
plot_gs_nov.circle('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source1, name="df",
fill_color=factor_cmap('novelty', palette=Spectral6, factors=novelties),
legend_field='novelty')
plot_gs_nov.circle('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source2, name="df",
fill_color='red')
plot_gs_nov.xaxis.axis_label_text_font_style = 'normal'
plot_gs_nov.yaxis.axis_label_text_font_style = 'normal'
plot_gs_nov.title.align = 'center'
plot_gs_nov.legend.location = "bottom_right"
plot_gs_nov.xaxis.axis_label_text_font_size = '18px'
plot_gs_nov.yaxis.axis_label_text_font_size = '18px'
plot_gs_nov.title.text_font_size = '16px'
show(plot_gs_nov)
figure_legend(f"Fig. {fig_count}", title=f"Novelties and Sizes of Gene Sets Within {library_name} Library", content=f"Scatterplot showing the size, publication count, and novelty rating of each gene set within your library.")
fig_count = fig_count + 1
# +
# Make a scatterplot of library size by novelty for all Enrichr libraries
xlabel = 'Library Size (Genes)'
ylabel = 'Novelty Statistic'
source1 = ColumnDataSource(
data=dict(
x = all_enrichr_novs_mix.Genes,
y = all_enrichr_novs_mix.Stat,
alpha = [0.9] * all_enrichr_novs_mix.shape[0],
size = [7] * all_enrichr_novs_mix.shape[0],
novelty = all_enrichr_novs_mix.Novelty,
lib = all_enrichr_novs_mix.Library,
)
)
source2 = ColumnDataSource(
data=dict(
x = my_lib_nov.Genes,
y = my_lib_nov.Stat,
alpha = [0.7] * my_lib_nov.shape[0],
size = [10] * my_lib_nov.shape[0],
novelty = my_lib_nov.Novelty,
lib = my_lib_nov.Library,
)
)
source3 = ColumnDataSource(
data=dict(
x = all_enrichr_novs_mouse.Genes,
y = all_enrichr_novs_mouse.Stat,
alpha = [0.9] * all_enrichr_novs_mouse.shape[0],
size = [7] * all_enrichr_novs_mouse.shape[0],
novelty = all_enrichr_novs_mouse.Novelty,
lib = all_enrichr_novs_mouse.Library,
)
)
#print(embedding.shape[0])
hover_nov = HoverTool(names=["df"], tooltips="""
<div style="margin: 10">
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Gene Set Library:</span>
<span style="font-size: 12px">@lib</span>
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Novelty:</span>
<span style="font-size: 12px">@novelty</span>
<div style="margin: 0 auto; width:300px;">
<span style="font-size: 12px; font-weight: bold;">Total Genes:</span>
<span style="font-size: 12px">@x</span>
</div>
</div>
""")
tools_nov = [hover_nov, 'pan', 'wheel_zoom', 'reset','save']
title_nov = f"Novelty and Size of {library_name} Library Among All Enrichr Libraries"
plot_nov = figure(plot_width=700, plot_height=700, tools=tools_nov, title=title_nov, x_axis_label=xlabel, y_axis_label=ylabel, x_axis_type='log')
plot_nov.circle('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source1, name="df",
fill_color=factor_cmap('novelty', palette=Spectral6, factors=novelties),
legend_field='novelty')
plot_nov.square('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source3, name="df",
fill_color=factor_cmap('novelty', palette=Spectral6, factors=novelties))
if other:
plot_nov.triangle('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source2, name="df",
fill_color='red')
if specname=='Mouse':
plot_nov.square('x', 'y', size='size',
alpha='alpha', line_alpha=0, line_width=0.01, source=source2, name="df",
fill_color='red')
if specname=='Human':
plot_nov.circle('x', 'y', size=10,
alpha='alpha', line_alpha=0, line_width=0.01, source=source2, name="df",
fill_color='red')
plot_nov.xaxis.axis_label_text_font_style = 'normal'
plot_nov.yaxis.axis_label_text_font_style = 'normal'
plot_nov.title.align = 'center'
plot_nov.legend.location = (170,520)
plot_nov.xaxis.axis_label_text_font_size = '18px'
plot_nov.yaxis.axis_label_text_font_size = '18px'
plot_nov.title.text_font_size = '16px'
from bokeh.models import Legend, LegendItem
markers = ['circle','square','triangle']
r = plot_nov.scatter(x=0, y=0, color="grey", size=6, marker=markers)
r.visible = False
shape_legend = Legend(items=[
LegendItem(label="human harmonized", renderers=[r], index=0),
LegendItem(label="mouse harmonized", renderers=[r], index=1),
LegendItem(label="other", renderers=[r], index=2),],
location=(10,520)
)
plot_nov.add_layout(shape_legend)
show(plot_nov)
figure_legend(f"Fig. {fig_count}", title=f"Novelty and Size of {library_name} Library Among All Enrichr Libraries", content=f"Scatterplot showing the size and novelty of your library compared with 174 Enrichr libraries. The Library Novelty Statistic ranges from 0 (well-studied) to 3 (highly understudied). The {library_name} library is shown in red.")
fig_count = fig_count + 1
# -
# Bokeh barplot
def make_bok_barplot(dat, col1name, col2name, title, lab1, lab2, tooltips_vert, tooltips_hor):
barsource_v = ColumnDataSource(
dict(
x = dat[col1name],
y = dat[col2name],
novelty = dat['Novelty'][::-1],
)
)
barsource_h = ColumnDataSource(
dict(
x = dat[col2name][::-1],
y = dat[col1name][::-1],
novelty = dat['Novelty'][::-1],
)
)
bar_title = title
if orient_bar == 'Vertical':
bokbar = figure(x_range=dat[col1name], plot_height=350, title=bar_title, toolbar_location='below', tooltips=tooltips_vert, x_axis_label=lab1, y_axis_label=lab2)
bokbar.vbar(x=dat[col1name], top=dat[col2name], width=.5, color=bar_color, hover_alpha=.7)
bokbar.xaxis.major_label_orientation = math.pi/5
bokbar.xgrid.grid_line_color = None
bokbar.y_range.start = 0
if counts_bar:
labels = LabelSet(x='x', y='y', text='y', level='annotation',
x_offset=-7, y_offset=0, source=barsource_v, render_mode='canvas', text_font_size = '11px')
bokbar.add_layout(labels)
if orient_bar == 'Horizontal':
bokbar = figure(y_range = dat[col1name][::-1], plot_height=400, title=bar_title, toolbar_location='below', tooltips=tooltips_hor, x_axis_label=lab2, y_axis_label=lab1)
bokbar.hbar(y='y',right='x', height=.5, color=bar_color, hover_alpha=.7, source=barsource_h)
bokbar.xgrid.grid_line_color = None
if counts_bar:
labels = LabelSet(x='x', y='y', text='x', level='annotation',
x_offset=2, y_offset=-6, source=barsource_h, render_mode='canvas', text_font_size = '11px')
bokbar.add_layout(labels)
bokbar.xaxis.axis_label_text_font_style = 'normal'
bokbar.yaxis.axis_label_text_font_style = 'normal'
bokbar.title.align = 'center'
return bokbar
# +
bokbar_counts_title = '' + str(num_bar_genes) + ' Most Frequent Genes in ' + library_name
bokbar_pubs_title = '' + str(num_bar_genes) + ' Most Studied Genes in ' + library_name
tooltips_vert = [
("count", "@top")
]
tooltips_hor = [
("count", "@x")
]
bokbar_counts = make_bok_barplot(top_genes, 'Gene', 'Count', bokbar_counts_title, 'Genes', 'Counts', tooltips_vert, tooltips_hor)
show(bokbar_counts)
figure_legend(f"Fig. {fig_count}", title=f"Most Frequent Genes in {library_name}")
fig_count = fig_count + 1
if len(pubhist_dat) > 0:
tooltips_vert = [
("publications", "@top"),
("novelty", "@novelty")
]
tooltips_hor = [
("publications", "@x"),
("novelty", "@novelty")
]
bokbar_pubs = make_bok_barplot(top_pub_genes, 'Gene', 'Publications', bokbar_pubs_title, 'Genes', 'Publications', tooltips_vert, tooltips_hor)
show(bokbar_pubs)
figure_legend(f"Fig. {fig_count}", title=f"Most Studied Genes in {library_name}")
fig_count = fig_count + 1
| appyters/Gene_Set_Library_Synopsis/Gene_Set_Library_Synopsis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Preparation
# ## Append to path and import packages
# In case gumpy is not installed as package, you may have to specify the path to the gumpy directory
# %reset
# %matplotlib inline
import matplotlib.pyplot as plt
import sys, os, os.path
sys.path.append('/.../gumpy')
# ## Import gumpy
# This may take a while, as gumpy as several dependencies that will be loaded automatically
import numpy as np
import gumpy
# ## Select workflow
# Select the actions you want to perform
# # Import data
# To import data, you have to specify the directory in which your data is stored in. For the example given here, the data is in the subfolder ``../EEG-Data/Graz_data/data``.
# Then, one of the classes that subclass from ``dataset`` can be used to load the data. In the example, we will use the GrazB dataset, for which ``gumpy`` already includes a corresponding class. If you have different data, simply subclass from ``gumpy.dataset.Dataset``.
# +
# First specify the location of the data and some
# identifier that is exposed by the dataset (e.g. subject)
data_base_dir = '/.../.../Data'
grazb_base_dir = os.path.join(data_base_dir, 'Graz')
subject = 'B01'
# The next line first initializes the data structure.
# Note that this does not yet load the data! In custom implementations
# of a dataset, this should be used to prepare file transfers,
# for instance check if all files are available, etc.
grazb_data = gumpy.data.GrazB(grazb_base_dir, subject)
# Finally, load the dataset
grazb_data.load()
# -
# The abstract class allows to print some information about the contained data. This is a commodity function that allows quick inspection of the data as long as all necessary fields are provided in the subclassed variant.
grazb_data.print_stats()
# labels = grazb_data.labels
# # Postprocess data
# Usually it is necessary to postprocess the raw data before you can properly use it. ``gumpy`` provides several methods to easily do so, or provides implementations that can be adapted to your needs.
#
# Most methods internally use other Python toolkits, for instance ``sklearn``, which is heavily used throughout ``gumpy``. Thereby, it is easy to extend ``gumpy`` with custom filters. In addition, we expect users to have to manipulate the raw data directly as shown in the following example.
# ## Common average re-referencing the data to Cz
# Some data is required to be re-referenced to a certain electrode. Because this may depend on your dataset, there is no common function provided by ``gumpy`` to do so. However and if sub-classed according to the documentation, you can access the raw-data directly as in the following example.
if False:
grazb_data.raw_data[:, 0] -= 2 * grazb_data.raw_data[:, 1]
grazb_data.raw_data[:, 2] -= 2 * grazb_data.raw_data[:, 2]
# ## Notch and Band-Pass Filters
# ``gumpy`` ships with several filters already implemented. They accept either raw data to be filtered, or a subclass of ``Dataset``. In the latter case, ``gumpy`` will automatically convert all channels using parameters extracted from the dataset.
# +
# this returns a butter-bandpass filtered version of the entire dataset
btr_data = gumpy.signal.butter_bandpass(grazb_data, lo=1, hi=35)
# it is also possible to use filters on individual electrodes using
# the .raw_data field of a dataset. The example here will remove a certain
# from a single electrode using a Notch filter. This example also demonstrates
# that parameters will be forwarded to the internal call to the filter, in this
# case the scipy implementation iirnotch (Note that iirnotch is only available
# in recent versions of scipy, and thus disabled in this example by default)
# frequency to be removed from the signal
if False:
f0 = 50
# quality factor
Q = 50
# get the cutoff frequency
w0 = f0/(grazb_data.sampling_freq/2)
# apply the notch filter
btr_data = gumpy.signal.notch(btr_data[:, 0], w0, Q)
# -
# ## Normalization
# Many datasets require normalization. ``gumpy`` provides functions to normalize either using a mean computation or via min/max computation. As with the filters, this function accepts either an instance of ``Dataset``, or raw_data. In fact, it can be used for postprocessing any row-wise data in a numpy matrix.
if False:
# normalize the data first
norm_data = gumpy.signal.normalize(grazb_data, 'mean_std')
# let's see some statistics
print("""Normalized Data:
Mean = {:.3f}
Min = {:.3f}
Max = {:.3f}
Std.Dev = {:.3f}""".format(
np.nanmean(norm_data),np.nanmin(norm_data),np.nanmax(norm_data),np.nanstd(norm_data)
))
# # Plotting and Feature Extraction
#
# Certainly you wish to plot results. ``gumpy`` provides several functions that show how to implement visualizations. For this purpose it heavily relies on ``matplotlib``, ``pandas``, and ``seaborn``. The following examples will show several of the implemented signal processing methods as well as their corresponding plotting functions. Moreover, the examples will show you how to extract features
#
# That said, let's start with a simple visualization where we access the filtered data from above to show you how to access the data and plot it.
# +
Plot after filtering with a butter bandpass (ignore normalization)
plt.figure()
plt.clf()
plt.plot(btr_data[grazb_data.trials[0]: grazb_data.trials[1], 0], label='C3')
plt.plot(btr_data[grazb_data.trials[0]: grazb_data.trials[1], 1], alpha=0.7, label='C4')
plt.plot(btr_data[grazb_data.trials[0]: grazb_data.trials[1], 2], alpha=0.7, label='Cz')
plt.legend()
plt.title(" Filtered Data")
# -
# ## EEG band visualization
# Using ``gumpy``'s filters and the provided method, it is easy to filter and subsequently plot the EEG bands of a trial.
# +
# determine the trial that we wish to plot
n_trial = 120
# now specify the alpha and beta cutoff frequencies
lo_a, lo_b = 7, 16
hi_a, hi_b = 13, 24
# first step is to filter the data
flt_a = gumpy.signal.butter_bandpass(grazb_data, lo=lo_a, hi=hi_a)
flt_b = gumpy.signal.butter_bandpass(grazb_data, lo=lo_b, hi=hi_b)
# finally we can visualize the data
gumpy.plot.EEG_bandwave_visualizer(grazb_data, flt_a, n_trial, lo_a, hi_a)
gumpy.plot.EEG_bandwave_visualizer(grazb_data, flt_b, n_trial, lo_a, hi_a)
# -
# ## Extract trials
# Now we wish to extract the trials from the data. This operation may heavily depend on your dataset, and thus we cannot guarantee that the function works for your specific dataset. However, the used function ``gumpy.utils.extract_trials`` can be used as a guideline how to extract the trials you wish to examine.
# +
# retrieve the trials from the filtered data. This requires that the function
# knows the number of trials, labels, etc. when only passed a (filtered) data matrix
trial_marks = grazb_data.trials
labels = grazb_data.labels
sampling_freq = grazb_data.sampling_freq
epochs = gumpy.utils.extract_trials(grazb_data, trials=trial_marks, labels=labels, sampling_freq=sampling_freq)
# it is also possible to pass an instance of Dataset and filtered data.
# gumpy will then infer all necessary details from the dataset
data_class_b = gumpy.utils.extract_trials(grazb_data, flt_b)
# similar to other functions, this one allows to pass an entire instance of Dataset
# to operate on the raw data
data_class1 = gumpy.utils.extract_trials(grazb_data)
# -
# ## Visualize the classes
# Given the extracted trials from above, we can proceed to visualize the average power of a class. Again, this depends on the specific data and thus you may have to adapt the function accordingly.
# +
# specify some cutoff values for the visualization
lowcut_a, highcut_a = 14, 30
# and also an interval to display
interval_a = [0, 8]
# visualize logarithmic power?
logarithmic_power = False
# visualize the extracted trial from above
gumpy.plot.average_power(epochs, lowcut_a, highcut_a, interval_a, grazb_data.sampling_freq, logarithmic_power)
# -
# ## Wavelet transform
# ``gumpy`` relies on ``pywt`` to compute wavelet transforms. Furthermore, it contains convenience functions to visualize the results of the discrete wavelet transform as shown in the example below for the Graz dataset and the classes extracted above.
# +
# As with most functions, you can pass arguments to a
# gumpy function that will be forwarded to the backend.
# In this example the decomposition levels are mandatory, and the
# mother wavelet that should be passed is optional
level = 6
wavelet = 'db4'
# now we can retrieve the dwt for the different channels
mean_coeff_ch0_c1 = gumpy.signal.dwt(data_class1[0], level=level, wavelet=wavelet)
mean_coeff_ch1_c1 = gumpy.signal.dwt(data_class1[1], level=level, wavelet=wavelet)
mean_coeff_ch0_c2 = gumpy.signal.dwt(data_class1[3], level=level, wavelet=wavelet)
mean_coeff_ch1_c2 = gumpy.signal.dwt(data_class1[4], level=level, wavelet=wavelet)
# gumpy's signal.dwt function returns the approximation of the
# coefficients as first result, and all the coefficient details as list
# as second return value (this is contrast to the backend, which returns
# the entire set of coefficients as a single list)
approximation_C3 = mean_coeff_ch0_c2[0]
approximation_C4 = mean_coeff_ch1_c2[0]
# as mentioned in the comment above, the list of details are in the second
# return value of gumpy.signal.dwt. Here we save them to additional variables
# to improve clarity
details_c3_c1 = mean_coeff_ch0_c1[1]
details_c4_c1 = mean_coeff_ch1_c1[1]
details_c3_c2 = mean_coeff_ch0_c2[1]
details_c4_c2 = mean_coeff_ch1_c2[1]
# gumpy exhibits a function to plot the dwt results. You must pass three lists,
# i.e. the labels of the data, the approximations, as well as the detailed coeffs,
# so that gumpy can automatically generate appropriate titles and labels.
# you can pass an additional class string that will be incorporated into the title.
# the function returns a matplotlib axis object in case you want to further
# customize the plot.
gumpy.plot.dwt(
[approximation_C3, approximation_C4],
[details_c3_c1, details_c4_c1],
['C3, c1', 'C4, c1'], level, grazb_data.sampling_freq, 'Class: Left')
# -
# ## DWT reconstruction and visualization
# Often a user wantes to reconstruct the power spectrum of a dwt and visualize the results. The functions will return a list of all the reconstructed signals as well as a handle to the figure.
# +
gumpy.plot.reconstruct_without_approx(
[details_c3_c2[4], details_c4_c2[4]],
['C3-c1', 'C4-c1'], level=6)
gumpy.plot.reconstruct_without_approx(
[details_c3_c1[5], details_c4_c1[5]],
['C3-c1', 'C4-c1'], level=6)
# -
gumpy.plot.reconstruct_with_approx(
[details_c3_c1[5], details_c4_c1[5]],
['C3', 'C4'], wavelet=wavelet)
# ## Welch's Power Spectral Density estimate
# Estimating the power spectral density according to Welch's method is =imilar to the power reconstruction shown above
# +
# the function gumpy.plot.welch_psd returns the power densities as
# well as a handle to the figure. You can also pass a figure in if you
# wish to modify the plot
fig = plt.figure()
plt.title('Customized plot')
ps, fig = gumpy.plot.welch_psd(
[details_c3_c1[4], details_c3_c2[4]],
['C3 - c1', 'C4 - c1'],
grazb_data.sampling_freq, fig=fig)
ps, fig = gumpy.plot.welch_psd(
[details_c4_c1[4], details_c4_c2[4]],
['C3 - c1', 'C4 - c1'],
grazb_data.sampling_freq)
# -
# ## Alpha and Beta sub-bands
# Using gumpys functions you can quickly define feature extractors. The following examples will demonstrate how you can use the predefined filters
# +
def alpha_subBP_features(data):
# filter data in sub-bands by specification of low- and high-cut frequencies
alpha1 = gumpy.signal.butter_bandpass(data, 8.5, 11.5, order=6)
alpha2 = gumpy.signal.butter_bandpass(data, 9.0, 12.5, order=6)
alpha3 = gumpy.signal.butter_bandpass(data, 9.5, 11.5, order=6)
alpha4 = gumpy.signal.butter_bandpass(data, 8.0, 10.5, order=6)
# return a list of sub-bands
return [alpha1, alpha2, alpha3, alpha4]
alpha_bands = np.array(alpha_subBP_features(btr_data))
# +
def beta_subBP_features(data):
beta1 = gumpy.signal.butter_bandpass(data, 14.0, 30.0, order=6)
beta2 = gumpy.signal.butter_bandpass(data, 16.0, 17.0, order=6)
beta3 = gumpy.signal.butter_bandpass(data, 17.0, 18.0, order=6)
beta4 = gumpy.signal.butter_bandpass(data, 18.0, 19.0, order=6)
return [beta1, beta2, beta3, beta4]
beta_bands = np.array(beta_subBP_features(btr_data))
# -
# ## Extract features without considering class information
# The following examples show how the sub-bands can be used to extract features. This also shows how the fields of the dataset can be accessed, and how to write methods specific to your data using a mix of gumpy's and numpy's functions.
# ### Method 1: logarithmic sub-band power
# +
def powermean(data, trial, fs, w):
return np.power(data[trial+fs*4+w[0]: trial+fs*4+w[1],0],2).mean(), \
np.power(data[trial+fs*4+w[0]: trial+fs*4+w[1],1],2).mean(), \
np.power(data[trial+fs*4+w[0]: trial+fs*4+w[1],2],2).mean()
def log_subBP_feature_extraction(alpha, beta, trials, fs, w):
# number of features combined for all trials
n_features = 15
# initialize the feature matrix
X = np.zeros((len(trials), n_features))
# Extract features
for t, trial in enumerate(trials):
power_c31, power_c41, power_cz1 = powermean(alpha[0], trial, fs, w)
power_c32, power_c42, power_cz2 = powermean(alpha[1], trial, fs, w)
power_c33, power_c43, power_cz3 = powermean(alpha[2], trial, fs, w)
power_c34, power_c44, power_cz4 = powermean(alpha[3], trial, fs, w)
power_c31_b, power_c41_b, power_cz1_b = powermean(beta[0], trial, fs, w)
X[t, :] = np.array(
[np.log(power_c31), np.log(power_c41), np.log(power_cz1),
np.log(power_c32), np.log(power_c42), np.log(power_cz2),
np.log(power_c33), np.log(power_c43), np.log(power_cz3),
np.log(power_c34), np.log(power_c44), np.log(power_cz4),
np.log(power_c31_b), np.log(power_c41_b), np.log(power_cz1_b)])
return X
# -
if False:
w1 = [0,125]
w2 = [125,256]
w3 = [256,512]
w4 = [512,512+256]
# extract the features
features1 = log_subBP_feature_extraction(
alpha_bands, beta_bands,
grazb_data.trials, grazb_data.sampling_freq,
w1)
features2 = log_subBP_feature_extraction(
alpha_bands, beta_bands,
grazb_data.trials, grazb_data.sampling_freq,
w2)
features3 = log_subBP_feature_extraction(
alpha_bands, beta_bands,
grazb_data.trials, grazb_data.sampling_freq,
w3)
features4 = log_subBP_feature_extraction(
alpha_bands, beta_bands,
grazb_data.trials, grazb_data.sampling_freq,
w4)
print(features4.shape)
# concatenate and normalize the features
features = np.concatenate((features1, features2, features3, features4), axis=1)
features -= np.mean(features)
features = gumpy.signal.normalize(features, 'mean_std')
features = gumpy.signal.normalize(features, 'min_max')
# print shape to quickly check if everything is as expected
print(features.shape)
# ### Method 2: Discrete Wavelet Transform (DWT)
def dwt_features(data, trials, level, sampling_freq, w, n, wavelet):
import pywt
# number of features per trial
n_features = 9
# allocate memory to store the features
X = np.zeros((len(trials), n_features))
# Extract Features
for t, trial in enumerate(trials):
signals = data[trial + fs*4 + (w[0]) : trial + fs*4 + (w[1])]
coeffs_c3 = pywt.wavedec(data = signals[:,0], wavelet=wavelet, level=level)
coeffs_c4 = pywt.wavedec(data = signals[:,1], wavelet=wavelet, level=level)
coeffs_cz = pywt.wavedec(data = signals[:,2], wavelet=wavelet, level=level)
X[t, :] = np.array([
np.std(coeffs_c3[n]), np.mean(coeffs_c3[n]**2),
np.std(coeffs_c4[n]), np.mean(coeffs_c4[n]**2),
np.std(coeffs_cz[n]), np.mean(coeffs_cz[n]**2),
np.mean(coeffs_c3[n]),
np.mean(coeffs_c4[n]),
np.mean(coeffs_cz[n])])
return X
# +
# We'll work with the data that was postprocessed using a butter bandpass
# filter further above
# to see it work, enable here. We'll use the log-power features further
# below, though
if False:
w = [0, 256]
# extract the features
trials = grazb_data.trials
fs = grazb_data.sampling_freq
features1= np.array(dwt_features(btr_data, trials, 5, fs, w, 3, "db4"))
features2= np.array(dwt_features(btr_data, trials, 5, fs, w, 4, "db4"))
# concatenate and normalize the features
features = np.concatenate((features1, features2), axis=1)
features -= np.mean(features)
features = gumpy.signal.normalize(features, 'min_max')
print(features.shape)
# -
# ### Split the data
# Now that we extracted features (and reduced the dimensionality), we can split the data for
# test and training purposes.
# +
# gumpy exposes several methods to split a dataset, as shown in the examples:
if 1:
split_features = np.array(gumpy.split.normal(features, labels,test_size=0.2))
X_train = split_features[0]
X_test = split_features[1]
Y_train = split_features[2]
Y_test = split_features[3]
X_train.shape
if 0:
n_splits=5
split_features = np.array(gumpy.split.time_series_split(features, labels, n_splits))
if 0:
split_features = np.array(gumpy.split.normal(PCA, labels, test_size=0.2))
#ShuffleSplit: Random permutation cross-validator
if 0:
split_features = gumpy.split.shuffle_Split(features, labels, n_splits=10,test_size=0.2,random_state=0)
# #Stratified K-Folds cross-validator
# #Stratification is the process of rearranging the data as to ensure each fold is a good representative of the whole
if 0:
split_features = gumpy.split.stratified_KFold(features, labels, n_splits=3)
#Stratified ShuffleSplit cross-validator
#Repeated random sub-sampling validation
if 0:
split_features = gumpy.split.stratified_shuffle_Split(features, labels, n_splits=10,test_size=0.3,random_state=0)
# # the functions return a list with the data according to the following example
# X_train = split_features[0]
# X_test = split_features[1]
# Y_train = split_features[2]
# Y_test = split_features[3]
# X_train.shape
# -
# ## Extract features considering class information
# ### Method 3: Common Spatial Patterns (CSP)
# #### I - Common prerequesites
# +
def extract_features(epoch, spatial_filter):
feature_matrix = np.dot(spatial_filter, epoch)
variance = np.var(feature_matrix, axis=1)
if np.all(variance == 0):
return np.zeros(spatial_filter.shape[0])
features = np.log(variance/np.sum(variance))
return features
def select_filters(filters, num_components = None, verbose = False):
if verbose:
print("Incomming filters:", filters.shape, "\nNumber of components:", num_components if num_components else " all")
if num_components == None:
return filters
assert num_components <= filters.shape[0]/2, "The requested number of components is too high"
selection = list(range(0, num_components)) + list(range(filters.shape[0] - num_components, filters.shape[0]))
reduced_filters = filters[selection,:]
return reduced_filters
# Select the number of used spatial components
n_components = None; # assign None for all components
# Rearrange epochs to (trials x channels x timesteps)
epochs1 = np.swapaxes(np.array([epochs[0],epochs[1],epochs[2]]),1,0)
epochs2 = np.swapaxes(np.array([epochs[3],epochs[4],epochs[5]]),1,0)
print("Number of trials per class:",epochs1.shape[0],"|",epochs2.shape[0])
#print(epochs1.shape)
#print(epochs2.shape)
# Remove invalid epochs
invalid_entries_1 = np.where([np.all(epoch == 0) for epoch in epochs1])
invalid_entries_2 = np.where([np.all(epoch == 0) for epoch in epochs2])
print("Number of invalid trials per class:",len(invalid_entries_1[0]),"|",len(invalid_entries_2[0]))
#print(invalid_entries_1)
#print(invalid_entries_2)
epochs1 = np.delete(epochs1, invalid_entries_1, 0)
epochs2 = np.delete(epochs2, invalid_entries_2, 0)
print("Number of trials per class after cleanup:",epochs1.shape[0],"|",epochs2.shape[0])
# Concatenate the trials
epochs_re = np.concatenate((epochs1, epochs2), axis=0)
print("Dataset:", epochs_re.shape)
# Update the label vector
labels = np.ones(epochs_re.shape[0])
labels[:epochs1.shape[0]] = 0
# Split data
epochs_train, epochs_test, y_train, y_test = gumpy.split.stratified_shuffle_Split(epochs_re, labels, n_splits=10,test_size=0.2,random_state=0)
print("Training data:", epochs_train.shape, "Testing data:", epochs_test.shape)
# -
# #### II A - Standard implementation
if False:
# Generate the spacial filterns for the training data
temp_filters = np.asarray(gumpy.features.CSP(epochs_train[np.where(y_train==0)], epochs_train[np.where(y_train==1)]))
#print(temp_filters.shape)
spatial_filter = select_filters(temp_filters[0], n_components)
#print(spatial_filter.shape)
# Extract the CSPs
features_train = np.array([extract_features(epoch, spatial_filter) for epoch in epochs_train])
#print(features_train.shape)
features_test = np.array([extract_features(epoch, spatial_filter) for epoch in epochs_test])
# #### II B - Filter Bank CSP
if True:
# Apply the filter bank
freqs = [[1,4],[4,8],[8,13],[13,22],[22,30]] # delta, theta, alpha, low beta, high beta
#freqs = [[1,4],[4,8],[8,13],[13,22],[22,30],[1,30]]
#freqs = [[1,4],[4,8],[8,12],[12,16],[16,20],[20,24],[24,28],[28,32]]
#freqs = [[1,3],[3,5],[5,7],[7,9],[9,11],[11,13],[13,15],[15,17],[17,19],[19,21],[21,23],[23,25],[25,27],[27,29],[29,31],[31,33],[33,35]]
x_train_fb = [gumpy.signal.butter_bandpass(epochs_train, lo=f[0], hi=f[1]) for f in freqs]
x_test_fb = [gumpy.signal.butter_bandpass(epochs_test, lo=f[0], hi=f[1]) for f in freqs]
# Generate the spatial filters for the training data
temp_filters = [np.asarray(gumpy.features.CSP(x_train_fb[f][np.where(y_train==0)], x_train_fb[f][np.where(y_train==1)])) for f in range(len(freqs))]
if n_components is not None:
spatial_filters = [select_filters(temp_filters[f][0], n_components) for f in range(len(freqs))]
else:
spatial_filters = [temp_filters[f][0] for f in range(len(freqs))]
# Extract the CSPs
features_train_tmp = [np.array([extract_features(epoch, spatial_filters[f]) for epoch in x_train_fb[f]]) for f in range(len(freqs))]
features_train = np.concatenate(features_train_tmp, axis=1)
features_test_tmp = [np.array([extract_features(epoch, spatial_filters[f]) for epoch in x_test_fb[f]]) for f in range(len(freqs))]
features_test = np.concatenate(features_test_tmp, axis=1)
print(features_train.shape)
print(features_test.shape)
# #### III - Common postprocessing
# +
if True:
# Feature normalization
features_train = gumpy.signal.normalize(features_train, 'mean_std')
features_test = gumpy.signal.normalize(features_test, 'mean_std')
X_train = gumpy.signal.normalize(features_train, 'min_max')
X_test = gumpy.signal.normalize(features_test, 'min_max')
Y_train = y_train
Y_test = y_test
# Debugging output
#usable_epochs = np.concatenate((epochs1, epochs2), axis=0)
#print(usable_epochs.shape)
#print(features.shape)
# -
# # Select features
# ## Sequential Feature Selection Algorithm
# ``gumpy`` provides a generic function with which you can select features. For a list of the implemented selectors please have a look at the function documentation.
#
if False:
Results = []
classifiers = []
Accuracy=[]
Final_results = {}
for model in gumpy.classification.available_classifiers:
print (model)
feature_idx, cv_scores, algorithm,sfs, clf = gumpy.features.sequential_feature_selector(X_train, Y_train, model,(6, 10), 5, 'SFFS')
classifiers.append(model)
Accuracy.append (cv_scores*100)
Final_results[model]= cv_scores*100
print (Final_results)
# ## PCA
# ``gumpy`` provides a wrapper around sklearn to reduce the dimensionality via PCA in a straightfoward manner.
PCA = gumpy.features.PCA_dim_red(features, 0.95)
# ## Plotting features
# ``gumpy`` wraps 3D plotting of features into a single line
gumpy.plot.PCA("3D", features, split_features[0], split_features[2])
# # Validation and test accuracies
# +
#SVM, RF, KNN, NB, LR, QLDA, LDA
from sklearn.cross_validation import cross_val_score
feature_idx, cv_scores, algorithm,sfs, clf = gumpy.features.sequential_feature_selector(X_train, Y_train, 'RandomForest',(6, 10), 10, 'SFFS')
feature=X_train[:,feature_idx]
scores = cross_val_score(clf, feature, Y_train, cv=10)
print("Validation Accuracy: %0.2f (+/- %0.2f)" % (scores.mean(), scores.std()))
clf.fit(feature, Y_train)
feature1=X_test[:,feature_idx]
clf.predict(feature1)
f=clf.score(feature1, Y_test)
print("Test Accuracy:",f )
# -
# # Voting classifiers
# Because `gumpy.classification.vote` uses `sklearn.ensemble.VotingClassifier` as backend, it is possible to specify different methods for the voting such as 'soft'. In addition, the method can be told to first extract features via `mlxtend.feature_selection.SequentialFeatureSelector` before classification.
if True:
result, _ = gumpy.classification.vote(X_train, Y_train, X_test, Y_test, 'soft', False, (6,12))
print("Classification result for hard voting classifier")
print(result)
print("Accuracy: ", result.accuracy)
if True:
result, _ = gumpy.classification.vote(X_train, Y_train, X_test, Y_test, 'hard', False, (6,12))
print("Classification result for hard voting classifier")
print(result)
print("Accuracy: ", result.accuracy)
# ## Voting Classifier with feature selection
# `gumpy` allows to automatically use all classifiers that are known in `gumpy.classification.available_classifiers` in a voting classifier (for more details see `sklearn.ensemble.VotingClassifier`). In case you developed a custom classifier and registered it using the `@register_classifier` decorator, it will be automatically used as well.
result, _ = gumpy.classification.vote(X_train, Y_train, X_test, Y_test, 'soft', True, (6,12))
print("Classification result for soft voting classifier")
print(result)
print("Accuracy: ", result.accuracy)
# ## Classification without feature selection
if False:
Results = []
classifiers = []
Accuracy=[]
Final_results = {}
for model in gumpy.classification.available_classifiers:
results, clf = gumpy.classify(model, X_train, Y_train, X_test, Y_test)
print (model)
print (results)
classifiers.append(model)
Accuracy.append (results.accuracy)
Final_results[model]= results.accuracy
print (Final_results)
# ## Confusion Matrix
# One of the ideas behind ``gumpy`` is to provide users the means to quickly examine their data. Therefore, gumpy provides mostly wraps existing libraries. This allows to show data with ease, and still be able to modify the plots in any way the underlying libraries allow:
# +
#Method 1
gumpy.plot.confusion_matrix(Y_test, results.pred)
#Method 2
gumpy.plot. plot_confusion_matrix(path='...', cm= , normalize = False, target_names = ['...' ], title = "...")
| examples/notebooks/EEG-motor-imagery-CSP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="rNdWfPXCjTjY"
# # LAB 03: Basic Feature Engineering in Keras
#
# **Learning Objectives**
#
#
# 1. Create an input pipeline using tf.data
# 2. Engineer features to create categorical, crossed, and numerical feature columns
#
#
# ## Introduction
# In this lab, we utilize feature engineering to improve the prediction of housing prices using a Keras Sequential Model.
#
# Each learning objective will correspond to a __#TODO__ in the [student lab notebook](courses/machine_learning/deepdive2/feature_engineering/labs/3_Basic_Feature_Engineering_in_Keras-lab.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
# + [markdown] colab_type="text" id="VxyBFc_kKazA"
# Start by importing the necessary libraries for this lab.
# -
# Install Sklearn
# !python3 -m pip install --user sklearn
# Ensure the right version of Tensorflow is installed.
# !pip freeze | grep tensorflow==2.1 || pip install tensorflow==2.1
# **Note:** After executing the above cell you will see the output
# `tensorflow==2.1.0` that is the installed version of tensorflow.
# + colab={} colab_type="code" id="9dEreb4QKizj"
import os
import tensorflow.keras
import matplotlib.pyplot as plt
import pandas as pd
import tensorflow as tf
from tensorflow import feature_column as fc
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
from keras.utils import plot_model
print("TensorFlow version: ",tf.version.VERSION)
# -
# Many of the Google Machine Learning Courses Programming Exercises use the [California Housing Dataset](https://developers.google.com/machine-learning/crash-course/california-housing-data-description
# ), which contains data drawn from the 1990 U.S. Census. Our lab dataset has been pre-processed so that there are no missing values.
#
# First, let's download the raw .csv data by copying the data from a cloud storage bucket.
#
if not os.path.isdir("../data"):
os.makedirs("../data")
# !gsutil cp gs://cloud-training-demos/feat_eng/housing/housing_pre-proc.csv ../data
# !ls -l ../data/
# + [markdown] colab_type="text" id="lM6-n6xntv3t"
# Now, let's read in the dataset just copied from the cloud storage bucket and create a Pandas dataframe.
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="REZ57BXCLdfG" outputId="a6ef2eda-c7eb-4e2d-92e4-e7fcaa20b0af"
housing_df = pd.read_csv('../data/housing_pre-proc.csv', error_bad_lines=False)
housing_df.head()
# -
# We can use .describe() to see some summary statistics for the numeric fields in our dataframe. Note, for example, the count row and corresponding columns. The count shows 20433.000000 for all feature columns. Thus, there are no missing values.
housing_df.describe()
# + [markdown] colab_type="text" id="u0zhLtQqMPem"
# #### Split the dataset for ML
#
# The dataset we loaded was a single CSV file. We will split this into train, validation, and test sets.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="code" id="YEOpw7LhMYsI" outputId="6161a660-7133-465a-d754-d7acae2b68c8"
train, test = train_test_split(housing_df, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
# + [markdown] colab_type="text" id="dz9kfjOMBX9U"
# Now, we need to output the split files. We will specifically need the test.csv later for testing. You should see the files appear in the home directory.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="ADX23QUu_Wiu" outputId="e97fa59e-4ed4-48a3-8fba-c95f293944ee"
train.to_csv('../data/housing-train.csv', encoding='utf-8', index=False)
# -
val.to_csv('../data/housing-val.csv', encoding='utf-8', index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="CU1FgmKEAmWh" outputId="2cce91e1-2c4a-4fe8-a6c3-3da52cb9458f"
test.to_csv('../data/housing-test.csv', encoding='utf-8', index=False)
# -
# !head ../data/housing*.csv
# + [markdown] colab_type="text" id="Aj35eYy_lutI"
# ## Create an input pipeline using tf.data
# + [markdown] colab_type="text" id="84ef46LXMfvu"
# Next, we will wrap the dataframes with [tf.data](https://www.tensorflow.org/guide/datasets). This will enable us to use feature columns as a bridge to map from the columns in the Pandas dataframe to features used to train the model.
#
# Here, we create an input pipeline using tf.data. This function is missing two lines. Correct and run the cell.
# -
# A utility method to create a tf.data dataset from a Pandas Dataframe
# TODO 1
def df_to_dataset(dataframe, shuffle=True, batch_size=32):
dataframe = dataframe.copy()
labels = dataframe.pop('median_house_value')
ds = tf.data.Dataset.from_tensor_slices((dict(dataframe), labels))
if shuffle:
ds = ds.shuffle(buffer_size=len(dataframe))
ds = ds.batch(batch_size)
return ds
# Next we initialize the training and validation datasets.
batch_size = 32
train_ds = df_to_dataset(train)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
# + [markdown] colab_type="text" id="qRLGSMDzM-dl"
# Now that we have created the input pipeline, let's call it to see the format of the data it returns. We have used a small batch size to keep the output readable.
# + colab={"base_uri": "https://localhost:8080/", "height": 331} colab_type="code" id="CSBo3dUVNFc9" outputId="d1be2646-b1e5-4110-dbba-5bc49d9b30f6"
# TODO 2
for feature_batch, label_batch in train_ds.take(1):
print('Every feature:', list(feature_batch.keys()))
print('A batch of households:', feature_batch['households'])
print('A batch of ocean_proximity:', feature_batch['ocean_proximity'])
print('A batch of targets:', label_batch)
# + [markdown] colab_type="text" id="OT5N6Se-NQsC"
# We can see that the dataset returns a dictionary of column names (from the dataframe) that map to column values from rows in the dataframe.
# + [markdown] colab_type="text" id="YEGEAqaziwfC"
# #### Numeric columns
# The output of a feature column becomes the input to the model. A numeric is the simplest type of column. It is used to represent real valued features. When using this column, your model will receive the column value from the dataframe unchanged.
#
# In the California housing prices dataset, most columns from the dataframe are numeric. Let' create a variable called **numeric_cols** to hold only the numerical feature columns.
# -
# TODO 3
numeric_cols = ['longitude', 'latitude', 'housing_median_age', 'total_rooms',
'total_bedrooms', 'population', 'households', 'median_income']
# + [markdown] colab_type="text" id="EwMEcH_52JT8"
# #### Scaler function
# It is very important for numerical variables to get scaled before they are "fed" into the neural network. Here we use min-max scaling. Here we are creating a function named 'get_scal' which takes a list of numerical features and returns a 'minmax' function, which will be used in tf.feature_column.numeric_column() as normalizer_fn in parameters. 'Minmax' function itself takes a 'numerical' number from a particular feature and return scaled value of that number.
# + [markdown] colab_type="text" id="ig1k5ovWBnN8"
# Next, we scale the numerical feature columns that we assigned to the variable "numeric cols".
# -
# Scalar def get_scal(feature):
# TODO 4
def get_scal(feature):
def minmax(x):
mini = train[feature].min()
maxi = train[feature].max()
return (x - mini)/(maxi-mini)
return(minmax)
# + colab={} colab_type="code" id="Y8IUfcuVaS_g"
# TODO 5
feature_columns = []
for header in numeric_cols:
scal_input_fn = get_scal(header)
feature_columns.append(fc.numeric_column(header,
normalizer_fn=scal_input_fn))
# + [markdown] colab_type="text" id="8v9XoD7WCKRM"
# Next, we should validate the total number of feature columns. Compare this number to the number of numeric features you input earlier.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="4jgPFThi50sS" outputId="23ede6f5-a62a-4767-b3a6-fe8a3b89a212"
print('Total number of feature coLumns: ', len(feature_columns))
# + [markdown] colab_type="text" id="9Ug3hB8Sl0jO"
# ### Using the Keras Sequential Model
#
# Next, we will run this cell to compile and fit the Keras Sequential model.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="_YJPPb3xTPeZ" outputId="2d445722-1d43-4a27-a6c0-c6ce813ab450"
# Model create
feature_layer = tf.keras.layers.DenseFeatures(feature_columns, dtype='float64')
model = tf.keras.Sequential([
feature_layer,
layers.Dense(12, input_dim=8, activation='relu'),
layers.Dense(8, activation='relu'),
layers.Dense(1, activation='linear', name='median_house_value')
])
# Model compile
model.compile(optimizer='adam',
loss='mse',
metrics=['mse'])
# Model Fit
history = model.fit(train_ds,
validation_data=val_ds,
epochs=32)
# -
# Next we show loss as Mean Square Error (MSE). Remember that MSE is the most commonly used regression loss function. MSE is the sum of squared distances between our target variable (e.g. housing median age) and predicted values.
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="vo7hhkPqm6Jx" outputId="938907f6-b6c8-497c-a8f6-0f1cdbf336c9"
loss, mse = model.evaluate(train_ds)
print("Mean Squared Error", mse)
# + [markdown] colab_type="text" id="252EPxGp7-FJ"
# #### Visualize the model loss curve
#
# Next, we will use matplotlib to draw the model's loss curves for training and validation. A line plot is also created showing the mean squared error loss over the training epochs for both the train (blue) and test (orange) sets.
# -
def plot_curves(history, metrics):
nrows = 1
ncols = 2
fig = plt.figure(figsize=(10, 5))
for idx, key in enumerate(metrics):
ax = fig.add_subplot(nrows, ncols, idx+1)
plt.plot(history.history[key])
plt.plot(history.history['val_{}'.format(key)])
plt.title('model {}'.format(key))
plt.ylabel(key)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left');
plot_curves(history, ['loss', 'mse'])
# + [markdown] colab_type="text" id="wqkozY268xi7"
# ### Load test data
# + [markdown] colab_type="text" id="uf4TyVJ_Dzxe"
# Next, we read in the test.csv file and validate that there are no null values.
# -
# Again, we can use .describe() to see some summary statistics for the numeric fields in our dataframe. The count shows 4087.000000 for all feature columns. Thus, there are no missing values.
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="b4C4BmhV8ch9" outputId="82bcc9d3-4432-4068-ab82-6a6abbe4a024"
test_data = pd.read_csv('../data/housing-test.csv')
test_data.describe()
# + [markdown] colab_type="text" id="nY2Yrt8fC7RW"
# Now that we have created an input pipeline using tf.data and compiled a Keras Sequential Model, we now create the input function for the test data and to initialize the test_predict variable.
# + colab={} colab_type="code" id="8rMdDeGDCwpT"
# TODO 6
def test_input_fn(features, batch_size=256):
"""An input function for prediction."""
# Convert the inputs to a Dataset without labels.
return tf.data.Dataset.from_tensor_slices(dict(features)).batch(batch_size)
# -
test_predict = test_input_fn(dict(test_data))
# + [markdown] colab_type="text" id="H5SkINtbDIdr"
# #### Prediction: Linear Regression
#
# Before we begin to feature engineer our feature columns, we should predict the median house value. By predicting the median house value now, we can then compare it with the median house value after feature engineeing.
#
# To predict with Keras, you simply call [model.predict()](https://keras.io/models/model/#predict) and pass in the housing features you want to predict the median_house_value for. Note: We are predicting the model locally.
# + colab={} colab_type="code" id="uNc6TSoJDL7-"
predicted_median_house_value = model.predict(test_predict)
# + [markdown] colab_type="text" id="HFXK1SKPDYgD"
# Next, we run two predictions in separate cells - one where ocean_proximity=INLAND and one where ocean_proximity= NEAR OCEAN.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="xepss0vhoHge" outputId="46842a26-eacd-4801-857b-18c6a8f2005c"
# Ocean_proximity is INLAND
model.predict({
'longitude': tf.convert_to_tensor([-121.86]),
'latitude': tf.convert_to_tensor([39.78]),
'housing_median_age': tf.convert_to_tensor([12.0]),
'total_rooms': tf.convert_to_tensor([7653.0]),
'total_bedrooms': tf.convert_to_tensor([1578.0]),
'population': tf.convert_to_tensor([3628.0]),
'households': tf.convert_to_tensor([1494.0]),
'median_income': tf.convert_to_tensor([3.0905]),
'ocean_proximity': tf.convert_to_tensor(['INLAND'])
}, steps=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="qPssm8p4EZHh" outputId="2a55d427-7857-401c-f60d-edbb36be19ec"
# Ocean_proximity is NEAR OCEAN
model.predict({
'longitude': tf.convert_to_tensor([-122.43]),
'latitude': tf.convert_to_tensor([37.63]),
'housing_median_age': tf.convert_to_tensor([34.0]),
'total_rooms': tf.convert_to_tensor([4135.0]),
'total_bedrooms': tf.convert_to_tensor([687.0]),
'population': tf.convert_to_tensor([2154.0]),
'households': tf.convert_to_tensor([742.0]),
'median_income': tf.convert_to_tensor([4.9732]),
'ocean_proximity': tf.convert_to_tensor(['NEAR OCEAN'])
}, steps=1)
# + [markdown] colab_type="text" id="Txl-MRuLFE_8"
# The arrays returns a predicted value. What do these numbers mean? Let's compare this value to the test set.
#
# Go to the test.csv you read in a few cells up. Locate the first line and find the median_house_value - which should be 249,000 dollars near the ocean. What value did your model predicted for the median_house_value? Was it a solid model performance? Let's see if we can improve this a bit with feature engineering!
#
# -
# ## Engineer features to create categorical and numerical features
# + [markdown] colab_type="text" id="78F1XH1Qwvbt"
# Now we create a cell that indicates which features will be used in the model.
# Note: Be sure to bucketize 'housing_median_age' and ensure that 'ocean_proximity' is one-hot encoded. And, don't forget your numeric values!
# + colab={} colab_type="code" id="ZxSatLUxUmvI"
# TODO 7
numeric_cols = ['longitude', 'latitude', 'housing_median_age', 'total_rooms',
'total_bedrooms', 'population', 'households', 'median_income']
bucketized_cols = ['housing_median_age']
# indicator columns,Categorical features
categorical_cols = ['ocean_proximity']
# + [markdown] colab_type="text" id="5HbypkYHxxwt"
# Next, we scale the numerical, bucktized, and categorical feature columns that we assigned to the variables in the precding cell.
# + colab={} colab_type="code" id="ExX5Akz0UnE-"
# Scalar def get_scal(feature):
def get_scal(feature):
def minmax(x):
mini = train[feature].min()
maxi = train[feature].max()
return (x - mini)/(maxi-mini)
return(minmax)
# + colab={} colab_type="code" id="wzqcddUQUnKn"
# All numerical features - scaling
feature_columns = []
for header in numeric_cols:
scal_input_fn = get_scal(header)
feature_columns.append(fc.numeric_column(header,
normalizer_fn=scal_input_fn))
# + [markdown] colab_type="text" id="yYUpUZvgwrPe"
# ### Categorical Feature
# In this dataset, 'ocean_proximity' is represented as a string. We cannot feed strings directly to a model. Instead, we must first map them to numeric values. The categorical vocabulary columns provide a way to represent strings as a one-hot vector.
# + [markdown] colab_type="text" id="sZnlnFZkyEbe"
# Next, we create a categorical feature using 'ocean_proximity'.
# + colab={} colab_type="code" id="3Cf6SoFTUnc6"
# TODO 8
for feature_name in categorical_cols:
vocabulary = housing_df[feature_name].unique()
categorical_c = fc.categorical_column_with_vocabulary_list(feature_name, vocabulary)
one_hot = fc.indicator_column(categorical_c)
feature_columns.append(one_hot)
# + [markdown] colab_type="text" id="qnGyWaijzShj"
# ### Bucketized Feature
#
# Often, you don't want to feed a number directly into the model, but instead split its value into different categories based on numerical ranges. Consider our raw data that represents a homes' age. Instead of representing the house age as a numeric column, we could split the home age into several buckets using a [bucketized column](https://www.tensorflow.org/api_docs/python/tf/feature_column/bucketized_column). Notice the one-hot values below describe which age range each row matches.
# + [markdown] colab_type="text" id="7ZRlFyP7fOw-"
# Next we create a bucketized column using 'housing_median_age'
#
# + colab={} colab_type="code" id="xB-yiVLmUnXp"
# TODO 9
age = fc.numeric_column("housing_median_age")
# Bucketized cols
age_buckets = fc.bucketized_column(age, boundaries=[10, 20, 30, 40, 50, 60, 80, 100])
feature_columns.append(age_buckets)
# + [markdown] colab_type="text" id="Ri4_wssOg943"
# ### Feature Cross
#
# Combining features into a single feature, better known as [feature crosses](https://developers.google.com/machine-learning/glossary/#feature_cross), enables a model to learn separate weights for each combination of features.
# + [markdown] colab_type="text" id="a6HHJl3J0j0T"
# Next, we create a feature cross of 'housing_median_age' and 'ocean_proximity'.
# + colab={} colab_type="code" id="JVLnG0WbUnkl"
# TODO 10
vocabulary = housing_df['ocean_proximity'].unique()
ocean_proximity = fc.categorical_column_with_vocabulary_list('ocean_proximity',
vocabulary)
crossed_feature = fc.crossed_column([age_buckets, ocean_proximity],
hash_bucket_size=1000)
crossed_feature = fc.indicator_column(crossed_feature)
feature_columns.append(crossed_feature)
# + [markdown] colab_type="text" id="hiz6HCWg1CXO"
# Next, we should validate the total number of feature columns. Compare this number to the number of numeric features you input earlier.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="6P3Ewc3_Unsv" outputId="42c1c4a6-89f8-4685-b2d0-e76a90cdf9ee"
print('Total number of feature coumns: ', len(feature_columns))
# + [markdown] colab_type="text" id="lNr00mP41sJp"
# Next, we will run this cell to compile and fit the Keras Sequential model. This is the same model we ran earlier.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="4Dwal3oxUoCe" outputId="1ae08747-7dbe-47a5-b3e7-87581e460b1b"
# Model create
feature_layer = tf.keras.layers.DenseFeatures(feature_columns,
dtype='float64')
model = tf.keras.Sequential([
feature_layer,
layers.Dense(12, input_dim=8, activation='relu'),
layers.Dense(8, activation='relu'),
layers.Dense(1, activation='linear', name='median_house_value')
])
# Model compile
model.compile(optimizer='adam',
loss='mse',
metrics=['mse'])
# Model Fit
history = model.fit(train_ds,
validation_data=val_ds,
epochs=32)
# + [markdown] colab_type="text" id="3LdUQszM16Oj"
# Next, we show loss and mean squared error then plot the model.
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="ZtFSpkd9UoAW" outputId="bac4836e-c4f1-4b29-876d-91fe1b51a5a7"
loss, mse = model.evaluate(train_ds)
print("Mean Squared Error", mse)
# + colab={"base_uri": "https://localhost:8080/", "height": 350} colab_type="code" id="O8kWMa6xUn-M" outputId="05ed9323-1102-4245-a40b-88543f11b0f3"
plot_curves(history, ['loss', 'mse'])
# + [markdown] colab_type="text" id="C4tWwOQt2e-P"
# Next we create a prediction model. Note: You may use the same values from the previous prediciton.
# -
# TODO 11
# Median_house_value is $249,000, prediction is $234,000 NEAR OCEAN
model.predict({
'longitude': tf.convert_to_tensor([-122.43]),
'latitude': tf.convert_to_tensor([37.63]),
'housing_median_age': tf.convert_to_tensor([34.0]),
'total_rooms': tf.convert_to_tensor([4135.0]),
'total_bedrooms': tf.convert_to_tensor([687.0]),
'population': tf.convert_to_tensor([2154.0]),
'households': tf.convert_to_tensor([742.0]),
'median_income': tf.convert_to_tensor([4.9732]),
'ocean_proximity': tf.convert_to_tensor(['NEAR OCEAN'])
}, steps=1)
# + [markdown] colab_type="text" id="rcbdA3arXkej"
# ### Analysis
#
# The array returns a predicted value. Compare this value to the test set you ran earlier. Your predicted value may be a bit better.
#
# Now that you have your "feature engineering template" setup, you can experiment by creating additional features. For example, you can create derived features, such as households per population, and see how they impact the model. You can also experiment with replacing the features you used to create the feature cross.
#
# -
# Copyright 2020 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| courses/machine_learning/deepdive2/feature_engineering/solutions/3_keras_basic_feat_eng.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ###### Formulation:
# * Stages: $i = 1,2,3,\cdots$.
# * State Space: $\mathcal{X} = \{a,b,c\}$.
# * Control Space: $\mathcal{U} = \{-1,1\}$
# * Feasible control mapping $\mathcal{X}\rightarrow\mathcal{U}: \mathcal{U}(x)=\{-1,1\}$
# * Transition mapping, given $x_i, u_i$, $\mathcal{X}\rightarrow [0,1]$, for $i\in \{1,2,\cdots\}$.
# $P(j\mid i, u=-1)$
#
# | i\\j | a | b | c
# | --- | --- | --- | ---
# | a | 1.0 | 0.0 | 0.0
# | b | 0.8| 0.0 | 0.2
# | c | 0.0 | 1.0 | 0.0
#
# $P(j\mid i, u=1)$
#
# | i\\j | a | b | c
# | --- | --- | --- | ---
# | a | 0.0 | 1.0 | 0.0
# | b | 0.6| 0.0 | 0.4
# | c | 0.0 | 0.0 | 1.0
#
#
# * Reward function $\mathcal{X}\times\mathcal{U}\rightarrow\mathbb{R}$:
# $$r(a, -1)=-1; \quad r(a, 1)=1;\quad r(c, -1)=10;\quad r(c,1)=-10 $$
# For other state and action combinations $(x, u)\in (\mathcal{S}, \mathcal{A})$, $r(x, u)=0$.
# * Let $v^*(x)$ be the optimal value function for state $x$, so it satisfies the following equation:
#
# \begin{align}
# v^*(x_i) = &\max_{u\in \mathcal{U}(x)}\{r(x_i, u)+\alpha\mathbb{E}[v^*(x_{i+1})]\}\\
# \end{align}
#
# And it can be solved by value iteration:
#
# \begin{align}
# v^{k+1}(x_i) = \mathcal{D}(v)(x_i) = &\max_{u\in \mathcal{U}(x)}\{r(x_i, u)+\alpha\mathbb{E}[v^k(x_{i+1})]\}\\
# \end{align}
# with arbitrary $v^0$, since $\mathcal{D}$ is a contraction mapping.
from itertools import product
import numpy as np
import pandas as pd
from copy import deepcopy
import random
import matplotlib.pyplot as plt
import seaborn as sns
pd.DataFrame({0: [1., 0.8, 0.], 1:[0.,0.,1.], 2:[0.,0.2, 0.]})
# +
state_space = [0,1,2]
control_space = [-1,1]
alpha = 0.95
tun1 = pd.DataFrame({0: [1., 1., 0.], 1:[0.,0.,1.], 2:[0.,0., 0.]})
tu1 = pd.DataFrame({0: [0.,0., 0.], 1:[1.,0.,0.], 2:[0.,1., 1.]})
def trans_prob(next_x, x, u):
if u == 1:
t = tu1
elif u == -1:
t = tun1
else:
raise ValueError
return t.loc[x, next_x]
def reward(x, u):
if x == 0 and u == 1:
return 1.
elif x == 2 and u == -1:
return 10.
elif x == 2 and u == 1:
return -10.
elif x == 0 and u == -1:
return -1
else:
return 0.
# -
tu1
tun1
v = pd.Series([0.0 for _ in state_space], index = state_space)
pi = pd.Series([0 for _ in state_space], index = state_space)
def value_iteration(state_space, control_space, trans_prob, reward, alpha, v, pi, MAX_ITER = 1000000, threshold = 1e-6):
for _ in range(MAX_ITER):
next_v = deepcopy(v)
for x in state_space:
tmp = [reward(x, u)+alpha*
np.sum([v[next_x]*trans_prob(next_x, x, u) for next_x in state_space])
for u in control_space]
next_v[x]= np.max(tmp)
pi[x]= control_space[np.argmax(tmp)]
if abs(next_v-v).values.max()<1e-6:
break
else:
v = next_v
return v, pi
v, pi = value_iteration(state_space, control_space, trans_prob, reward, 0.95, v, pi)
v
pi
q_true = pd.DataFrame({x:[0.0]*len(control_space) for x in state_space}, index = control_space)
for x in state_space:
for u in control_space:
q_true.loc[u, x] = reward(x, u)+alpha*np.sum([v[next_x]*trans_prob(next_x, x, u) for next_x in state_space])
print(q_true)
list(product(state_space, control_space))
# +
# Augmented MDP
state_space_d = list(product(state_space, control_space))
control_space_d = [-1,1]
alpha = 0.95
# tun1 = pd.DataFrame({0: [1., 0.8, 0.], 1:[0.,0.,1.], 2:[0.,0.2, 0.]})
# tu1 = pd.DataFrame({0: [0., 0.6, 0.], 1:[1.,0.,0.], 2:[0.,0.4, 1.]})
def trans_prob_d(next_i, i, u):
if next_i[1]==u:
return trans_prob(next_i[0], i[0], i[1])
else:
return 0.0
def reward_d(x, u):
return reward(x[0], x[1])
# -
v_d = pd.Series([0.0 for _ in state_space_d], index = state_space_d)
pi_d = pd.Series([0 for _ in state_space_d], index = state_space_d)
v_d
v_AMDP, pi_AMDP = value_iteration(state_space_d, control_space_d, trans_prob_d, reward_d, 0.95, v_d, pi_d)
v_AMDP
q_true.loc[-1, 0][-1]
next_x = x
v_AMDP[next_x]*trans_prob_d(next_x, x, u)
q_true = pd.DataFrame({x:[0.0]*len(control_space_d) for x in state_space_d}, index = control_space_d)
for x in state_space_d:
for u in control_space_d:
q_true.loc[u, x[0]][x[1]] = reward_d(x, u)+alpha*np.sum([v_AMDP[next_x]*trans_prob_d(next_x, x, u) for next_x in state_space_d])
print(q_true)
q_true.loc[-1, 1][1]
reward
pi_AMDP
# Q_learning env
class Env:
def __init__(self):
# self.gamma = gamma
self.obs = 0
self.control_space = (-1, 1)
self.state_space = tuple([0,1,2])
self.state_dim = 1
self.action_dim = 1
def step(self, action, current_obs = None):
obs = self.obs if current_obs is None else current_obs
self.obs = self._next_state(obs, action)
r = reward(obs, action)
# r = reward((q,p), action)
return self.obs, r, False, {}
def reset(self, random=False):
self.obs = 0 if not random else np.random.choice([0,1,2])
return self.obs
@staticmethod
def _next_state(obs, action):
p = tun1.loc[obs, :] if action == -1 else tu1.loc[obs, :]
return np.random.choice([0,1,2], p = p)
# +
class Q_table:
def __init__(self, q_data=None, random = False):
self.control_space = (-1,1)
self.state_space = tuple([0,1,2])
if q_data is None:
if not random:
self.data = pd.DataFrame({x:[0.0]*len(self.control_space) for x in self.state_space},
index = self.control_space)
else:
self.data = pd.DataFrame({x:np.random.randn(len(self.control_space)) for x in self.state_space},
index = self.control_space)
else:
# assert isinstance(q, Q_table)
self.data = q_data
def __getitem__(self, key):
x, u = key
return self.data.loc[u,x]
def __setitem__(self, key, item):
x, u = key
self.data.loc[u,x] = item
def max_value_act(self, x):
tmp = [self.data.loc[k,x] for k in self.control_space]
return np.max(tmp), self.control_space[np.argmax(tmp)]
def difference(self, q):
assert isinstance(q, Q_table)
return Q_table(self.data-q.data)
def L_infty(self):
return np.max(np.abs(self.data).values)
def L_1(self):
return np.mean(np.abs(self.data).values)
def yield_v_pi(self):
v = pd.Series([0.0 for _ in self.state_space], index = self.state_space)
pi = pd.Series([0 for _ in self.state_space], index = self.state_space)
for i in self.state_space:
v[i], pi[i] = self.max_value_act(i)
return v, pi
def __repr__(self):
return self.data.__repr__()
class Pool:
def __init__(self, length):
self.ls = [None]*length
self.len = length
self.idx = 0
self.isFull = False
def append(self, item):
self.ls[self.idx] = item
self.idx = (self.idx + 1) % self.len
if not self.isFull:
if self.idx + 1 == self.len:
self.isFull = True
def shuffle(self):
cp = deepcopy(self.ls)
random.shuffle(cp)
return cp
def __getitem__(self, key):
idx = (self.idx+key) % self.len
return self.ls[idx]
def __iter__(self):
return iter(self.ls[self.idx:]+self.ls[:self.idx])
def __repr__(self):
return str(self.ls[self.idx:]+self.ls[:self.idx])
# +
# Q-learning with pool
class Q_learning:
def __init__(self, q, env, epsilon = 0.1, MAX_ITER = 10000, episode_length = 100, threshold = 1e-3):
self.q = q
self.env = env
self.epsilon = epsilon
self.MAX_ITER = MAX_ITER
self.episode_length = episode_length
self.threshold = threshold
self.pool = Pool(40)
self.qs = []
def train(self, lr = lambda *args: 0.1):
q = self.q
env = self.env
epsilon = self.epsilon
MAX_ITER = self.MAX_ITER
episode_length = self.episode_length
threshold = self.threshold
for i in range(MAX_ITER):
print(i)
obs = env.reset(random=True)
r_ls = []
for _ in range(10):
action, is_random = (np.random.choice(control_space), True) \
if np.random.uniform() < epsilon else \
(q.max_value_act(obs)[1], False)
next_obs, reward, _, _ = env.step(action)
self.pool.append((obs, next_obs, action, reward, is_random))
r_ls.append(reward)
obs = next_obs
if not self.pool.isFull:
continue
print(f'EP {i} simulation finished, start learning')
q_new = self.update_q(q, self.pool ,i, 0.95, lr)
self.q = q_new
print(np.mean(r_ls))
self.qs.append(q_new)
print(f'EP {i} learning finished: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
if q_new.difference(q).L_1() < threshold and False:
break
else:
q = q_new
return q_new
def update_q(self, q, pool, t ,alpha, lr):
pool = pool.shuffle()
for _ in range(1):
q = deepcopy(q)
#
for s, next_s, a, r, is_random in pool:
# if is_random: continue
q[s, a] = q[s,a] + lr(t, s, a)*(r + alpha*q.max_value_act(next_s)[0]-q[s,a])
# print(f'Learning ep: {i}: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
# if q_new.difference(q).L_1() < self.threshold:
# break
# else:
# q = q_new
return q
# -
# Q-learning with epsilon-greedy policy, learning rate = 1/t
q = Q_table(random = False)
env = Env()
q_one_over_t = Q_learning(q, env, MAX_ITER = 200)
lr_fn = lambda i, s, a: 1/(i+1)
lr_fn = lambda i, s, a: 0.1
class Learning_Rate:
def __init__(self, state_space, control_space):
self.counter = dict(
zip(list(product(state_space, control_space)), [0]*len(state_space)*len(control_space))
)
def __call__(self, i, s, a):
self.counter[(s, a)] += 1
return 1/self.counter[(s, a)]
lr_fn = Learning_Rate(state_space, control_space)
learned_q = q_one_over_t.train(lr_fn)
q_one_over_t.q
v_l, pi_l= q_one_over_t.q.yield_v_pi()
v_l
pi_l
# +
# 1 action delay, 0 obs. delay
class delayed_wrapper:
def __init__(self, env, delay_steps):
self.env = env
self.num = delay_steps
self.state_dim = env.state_dim
self.action_dim = env.action_dim
self.action_buffer = [-1]*delay_steps
def step(self, action):
app_action = self.action_buffer.pop(0)
self.action_buffer.append(action)
return self.env.step(app_action)
def reset(self, random = False):
self.action_buffer = [0]*self.num
return self.env.reset(random)
# +
# Q-learning using model, async reward collection
class Q_learning:
def __init__(self, q, env, delay_step, model, epsilon = 0.1, MAX_ITER = 10000, episode_length = 100, threshold = 1e-3):
self.q = q
self.env = env
self.delay_step = delay_step
self.epsilon = epsilon
self.MAX_ITER = MAX_ITER
self.episode_length = episode_length
self.threshold = threshold
self.pool = Pool(40)
self.qs = []
self.action_memory = [-1]*delay_step
self.model = model
def train(self, lr = lambda *args: 0.1):
q = self.q
env = self.env
epsilon = self.epsilon
MAX_ITER = self.MAX_ITER
episode_length = self.episode_length
threshold = self.threshold
model = self.model
for i in range(MAX_ITER):
print(i)
obs = env.reset(random = True)
r_ls = []
for _ in range(10):
est_obs = model(obs, self.action_memory)
last_action = self.action_memory.pop(0)
action, is_random = (np.random.choice(control_space), True) \
if np.random.uniform() < epsilon else \
(q.max_value_act(est_obs)[1], False)
self.action_memory.append(action)
next_obs, reward, _, _ = env.step(action)
self.pool.append((obs, next_obs, last_action, reward, is_random))
r_ls.append(reward)
obs = next_obs
if not self.pool.isFull:
continue
print(f'EP {i} simulation finished, start learning')
q_new = self.update_q(q, self.pool ,i, 0.95, lr)
self.q = q_new
print(np.mean(r_ls))
self.qs.append(q_new)
print(f'EP {i} learning finished: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
if q_new.difference(q).L_1() < threshold and False:
break
else:
q = q_new
return q_new
def update_q(self, q, pool, t ,alpha, lr):
pool = pool.shuffle()
for _ in range(1):
q = deepcopy(q)
#
for s, next_s, a, r, is_random in pool:
# if is_random: continue
q[s, a] = q[s,a] + lr(t, s, a)*(r + alpha*q.max_value_act(next_s)[0]-q[s,a])
# print(f'Learning ep: {i}: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
# if q_new.difference(q).L_1() < self.threshold:
# break
# else:
# q = q_new
return q
# -
mu1 = np.argmax(tu1.values,axis=1)
mun1 = np.argmax(tun1.values,axis=1)
# +
def model_model(x, u):
assert type(u)==list
u = u[0]
m = mu1
if u == 1:
m = mu1
elif u == -1:
m = mun1
else:
raise ValueError
return m[x]
# -
class Learning_Rate:
def __init__(self, state_space, control_space):
self.counter = dict(
zip(list(product(state_space, control_space)), [0]*len(state_space)*len(control_space))
)
def __call__(self, i, s, a):
self.counter[(s, a)] += 1
return 1/self.counter[(s, a)]
lr_fn = Learning_Rate(state_space, control_space)
# lr_fn = lambda i,s,a: 1/(i+1)
# lr_fn = lambda i,s,a: 0.1
env = delayed_wrapper(Env(), 1)
q = Q_table(random = False)
q_model = Q_learning(q, env, 1, model_model, MAX_ITER = 1000)
learned_q = q_model.train(lr_fn)
v_model, pi_model= q_model.q.yield_v_pi()
v_model
pi_model
# +
for s in state_space:
for a in control_space:
print(f'state: {s}, action_buffer: {[a]}, action to take: {q_model.q.max_value_act(model_model(s, [a]))[1]}')
def pi_model(i):
s, a = i
return q_model.q.max_value_act(model_model(s, [a]))[1]
# +
def pi_wrapper(pi):
def ret(x):
return pi[x]
return ret
def eval_pi_model(env, pi, model, discount_factor, MAX_ITER = 1000000):
path_len = int(np.log(1e-3)/np.log(discount_factor))
path_num = int(MAX_ITER/path_len)
obs = 0
disc_arr = np.array([discount_factor**n for n in range(path_len)])
r_ls = np.array([[np.nan]*path_len for _ in range(path_num)])
ret = []
delay_step = env.num
action_buffer = [-1]*delay_step
for k in range(path_num):
obs = env.reset(random= True)
for i in range(path_len):
action = pi(model(obs, action_buffer))
action_buffer.pop(0)
action_buffer.append(action)
next_obs, reward, _, _ = env.step(action)
# discounted_cum_r = reward + discount_factor*discounted_cum_r
r_ls[k, i] = reward
obs = next_obs
return np.mean(r_ls), r_ls.mean(axis = 1)
def cummean(arr):
return np.cumsum(arr)/np.arange(1, len(arr)+1)
# -
val, v_samples_lin = eval_pi_model(env, pi_wrapper(pi_l), model_model, 0.95)
val
exp2_model = v_samples_lin
sns.distplot(v_samples_lin, label = 'model')
plt.legend()
val, v_samples_lin = eval_pi_model(env, lambda x: np.random.choice([-1,1]), model_model, 0.95)
val
exp2_random = v_samples_lin
sns.distplot(v_samples_lin, label = 'random benchmark')
plt.legend()
# +
# Q-learning using model, async reward collection and optimal action selection
class Q_learning:
def __init__(self, q, env, delay_step, model, epsilon = 0.1, MAX_ITER = 10000, episode_length = 100, threshold = 1e-3):
self.q = q
self.env = env
self.delay_step = delay_step
self.epsilon = epsilon
self.MAX_ITER = MAX_ITER
self.episode_length = episode_length
self.threshold = threshold
self.pool = Pool(400)
self.qs = []
self.action_memory = [-1]*delay_step
self.model = model
def train(self, lr = lambda *args: 0.1):
q = self.q
env = self.env
epsilon = self.epsilon
MAX_ITER = self.MAX_ITER
episode_length = self.episode_length
threshold = self.threshold
model = self.model
for i in range(MAX_ITER):
print(i)
obs = env.reset(random = True)
r_ls = []
for _ in range(100):
action, is_random = (np.random.choice(control_space), True) \
if np.random.uniform() < epsilon else \
(get_opt_act(q, model, obs, self.action_memory), False)
last_action = self.action_memory.pop(0)
self.action_memory.append(action)
next_obs, reward, _, _ = env.step(action)
self.pool.append((obs, next_obs, last_action, reward, is_random))
r_ls.append(reward)
obs = next_obs
if not self.pool.isFull:
continue
print(f'EP {i} simulation finished, start learning')
q_new = self.update_q(q, self.pool ,i, 0.95, lr)
self.q = q_new
print(np.mean(r_ls))
self.qs.append(q_new)
print(f'EP {i} learning finished: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
if q_new.difference(q).L_1() < threshold and False:
break
else:
q = q_new
return q_new
def update_q(self, q, pool, t ,alpha, lr):
pool = pool.shuffle()
for _ in range(1):
q = deepcopy(q)
#
for s, next_s, a, r, is_random in pool:
# if is_random: continue
q[s, a] = q[s,a] + lr(t, s, a)*(r + alpha*q.max_value_act(next_s)[0]-q[s,a])
# print(f'Learning ep: {i}: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
# if q_new.difference(q).L_1() < self.threshold:
# break
# else:
# q = q_new
return q
def get_opt_act(q, model, x, acts):
act_idx = np.argmax([np.sum([q[s, u]*model(s, x, acts) for s in q.state_space]) for u in q.control_space])
return q.control_space[act_idx]
# -
# P(s|x, a_1, a_2, ...)
def model(s, x, acts):
return trans_prob(s, x, acts[0])
class Learning_Rate:
def __init__(self, state_space, control_space):
self.counter = dict(
zip(list(product(state_space, control_space)), [0]*len(state_space)*len(control_space))
)
def __call__(self, i, s, a):
self.counter[(s, a)] += 1
return 1/self.counter[(s, a)]
lr_fn = Learning_Rate(state_space, control_space)
# lr_fn = lambda i,s,a: 1/(i+1)
# lr_fn = lambda i,s,a: 0.1
env = delayed_wrapper(Env(), 1)
q = Q_table(random = False)
q_opt = Q_learning(q, env, 1, model, MAX_ITER = 1000)
learned_q = q_opt.train(lr_fn)
v_opt, pi_opt= q_opt.q.yield_v_pi()
v_opt
pi_opt
# +
for s in state_space:
for a in control_space:
print(f'state: {s}, action_buffer: {[a]}, action to take: {get_opt_act(q_opt.q, model, s, [a])}')
pi_opt = lambda x: get_opt_act(q_opt.q, model, x[0], [x[1]])
# -
q_opt.q
def eval_pi_model(env, q, model, discount_factor, MAX_ITER = 1000000):
path_len = int(np.log(1e-3)/np.log(discount_factor))
path_num = int(MAX_ITER/path_len)
obs = 0
disc_arr = np.array([discount_factor**n for n in range(path_len)])
r_ls = np.array([[np.nan]*path_len for _ in range(path_num)])
ret = []
delay_step = env.num
action_buffer = [-1]*delay_step
for k in range(path_num):
obs = env.reset(random= True)
for i in range(path_len):
action = get_opt_act(q, model, obs, action_buffer)
action_buffer.pop(0)
action_buffer.append(action)
next_obs, reward, _, _ = env.step(action)
# discounted_cum_r = reward + discount_factor*discounted_cum_r
r_ls[k, i] = reward
obs = next_obs
ret.append(np.cumsum(disc_arr*r_ls[k, :]))
# return np.mean(ret) ,ret
return np.mean(r_ls), r_ls.mean(axis = 1)
val, v_samples_lin = eval_pi_model(env, q_opt.q, model, 0.95)
val
exp2_opt = v_samples_lin
# +
from matplotlib.pyplot import figure
figure(figsize=(8, 6), dpi=100)
sns.distplot(exp2_opt, label = 'Opt. Action Selection')
sns.distplot(exp2_model, label = 'model')
sns.distplot(exp2_random, label = 'random')
sns.distplot(exp1_opt, label = 'Augmented MDP')
plt.xlabel("average reward")
plt.ylabel("freq.")
plt.legend()
# plt.set_size_inches(18.5, 10.5)
plt.savefig('exp2.png', dpi=100)
# -
plt.savefig('exp1.png')
# +
# For a given policy function, we solve the system A^{\pi}x=b^{\pi} to get the optimal value function
def linear_system(pi, trans_prob, reward, state_space, gamma):
n = len(state_space)
A = np.zeros(shape = (n,n))
b = np.zeros(n)
for i, s in enumerate(state_space):
for j, next_s in enumerate(state_space):
A[i,j] = trans_prob(next_s, s, pi(s))
b[i] = -reward(s, pi(s))
print(f'policy({s}): {pi(s)}')
A = gamma*A-np.eye(n)
return A, b
# -
from numpy.linalg import solve
# policy of opt action selection
A, b = linear_system(pi_opt, trans_prob_d, reward_d, state_space_d, alpha)
A, b
solve(A, b)
pi_AMDP_func = lambda x:pi_AMDP[x]
# +
# policy of AMDP
A, b = linear_system(pi_AMDP_func, trans_prob_d, reward_d, state_space_d, alpha)
A, b
# -
solve(A, b)
# +
# policy of planning based mathod
# -
A, b = linear_system(pi_model, trans_prob_d, reward_d, state_space_d, alpha)
A, b
solve(A, b)
pi_d
def eval_pi_model(env, pi, discount_factor, MAX_ITER = 1000000):
path_len = int(np.log(1e-3)/np.log(discount_factor))
path_num = int(MAX_ITER/path_len)
obs = 0
disc_arr = np.array([discount_factor**n for n in range(path_len)])
r_ls = np.array([[np.nan]*path_len for _ in range(path_num)])
ret = []
delay_step = env.num
action_buffer = [-1]*delay_step
for k in range(path_num):
obs = env.reset(random= True)
for i in range(path_len):
action = pi[(obs, action_buffer[0])]
action_buffer.pop(0)
action_buffer.append(action)
next_obs, reward, _, _ = env.step(action)
# discounted_cum_r = reward + discount_factor*discounted_cum_r
r_ls[k, i] = reward
obs = next_obs
ret.append(np.cumsum(disc_arr*r_ls[k, :]))
# return np.mean(ret) ,ret
return np.mean(r_ls), r_ls.mean(axis = 1)
val, v_samples_lin = eval_pi_model(env, pi_d, 0.95)
val
# +
# Q-learning using LEARNING-BASED model, async reward collection and optimal action selection
class Q_learning:
def __init__(self, q, env, delay_step, model, model_wrapper, epsilon = 0.1, MAX_ITER = 10000, episode_length = 100, threshold = 1e-3):
self.q = q
self.env = env
self.delay_step = delay_step
self.epsilon = epsilon
self.MAX_ITER = MAX_ITER
self.episode_length = episode_length
self.threshold = threshold
self.pool = Pool(400)
self.qs = []
self.action_memory = [-1]*delay_step
self.model = model
self.model_wrapper = model_wrapper
def train(self, lr = lambda *args: 0.1):
q = self.q
env = self.env
epsilon = self.epsilon
MAX_ITER = self.MAX_ITER
episode_length = self.episode_length
threshold = self.threshold
for i in range(MAX_ITER):
print(i)
obs = env.reset(random = True)
r_ls = []
model = self.model_wrapper(self.model)
for _ in range(100):
action, is_random = (np.random.choice(control_space), True) \
if np.random.uniform() < epsilon else \
(get_opt_act(q, model, obs, self.action_memory), False)
last_action = self.action_memory.pop(0)
self.action_memory.append(action)
next_obs, reward, _, _ = env.step(action)
self.pool.append((obs, next_obs, last_action, reward, is_random))
r_ls.append(reward)
obs = next_obs
if not self.pool.isFull:
continue
print(f'EP {i} simulation finished, start learning')
q_new = self.update_q(q, self.pool ,i, 0.95, lr)
self.update_model(self.pool)
self.q = q_new
print(np.mean(r_ls))
self.qs.append(q_new)
print(f'EP {i} learning finished: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
if q_new.difference(q).L_1() < threshold and False:
break
else:
q = q_new
return q_new
def update_model(self, pool):
tmp = []
for ele in pool:
tmp.append(ele)
x_train = np.array(tmp)[:, [0,2]]
y_train = np.array(tmp)[:, 1]
self.model.fit(x_train, y_train, batch_size=64, epochs=3, validation_split=0.2)
def update_q(self, q, pool, t ,alpha, lr):
pool = pool.shuffle()
for _ in range(1):
q = deepcopy(q)
#
for s, next_s, a, r, is_random in pool:
# if is_random: continue
q[s, a] = q[s,a] + lr(t, s, a)*(r + alpha*q.max_value_act(next_s)[0]-q[s,a])
# print(f'Learning ep: {i}: {q_new.difference(q).L_infty()}, {q_new.difference(q).L_1()}')
# if q_new.difference(q).L_1() < self.threshold:
# break
# else:
# q = q_new
return q
def get_opt_act(q,model, x, acts):
act_idx = np.argmax([np.sum([q[s, u]*model(s, x, acts) for s in q.state_space]) for u in q.control_space])
return q.control_space[act_idx]
# -
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
# +
inputs = keras.Input(shape=(2,))
layer1 = layers.Dense(64, activation="relu", name="layer1")(inputs)
layer1 = layers.Dropout(0.5)(layer1)
layer2 = layers.Dense(64, activation="relu", name="layer2")(layer1)
layer2 = layers.Dropout(0.5)(layer2)
outputs = layers.Dense(3, activation="softmax", name="output_layer")(layer2)
model = keras.Model(inputs = inputs, outputs = outputs)
model.compile(
loss=keras.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer=keras.optimizers.RMSprop(),
metrics=["accuracy"],
)
model.summary()
# +
tmp = []
for ele in q_learn.pool:
tmp.append(ele)
x_train = np.array(tmp)[:, [0,2]]
y_train = np.array(tmp)[:, 1]
def one_hot_encode(y):
y = y.astype(int)
elements = set(y)
n = len(elements)
ret = np.zeros((len(y), n))
ret[np.arange(y.size),y] = 1
return ret
# y_train = one_hot_encode(y_train)
print(x_train.shape, y_train.shape)
# -
model.predict([[1, 1]])[0,0]
history = model.fit(x_train, y_train, batch_size=64, epochs=10, validation_split=0.2)
class Learning_Rate:
def __init__(self, state_space, control_space):
self.counter = dict(
zip(list(product(state_space, control_space)), [0]*len(state_space)*len(control_space))
)
def __call__(self, i, s, a):
self.counter[(s, a)] += 1
return 1/self.counter[(s, a)]
# lr_fn = Learning_Rate(state_space, control_space)
# lr_fn = lambda i,s,a: 1/(i+1)
def model_wrapper(model):
def m(s, x, acts):
inputs = np.array([[x, acts[0]]])
return model.predict(inputs)[0, int(s)]
return m
lr_fn = lambda i,s,a: 0.1
env = delayed_wrapper(Env(), 1)
q = Q_table(random = False)
q_learn = Q_learning(q, env, 1, model, model_wrapper, MAX_ITER = 1000)
learned_q = q_learn.train(lr_fn)
m = model_wrapper(model)
m(0, 1, [1])
v_l, pi_l= q_learn.q.yield_v_pi()
v_l
pi_l
def eval_pi_model(env, q, model, model_wrapper, discount_factor, MAX_ITER = 1000000):
path_len = int(np.log(1e-3)/np.log(discount_factor))
path_num = int(MAX_ITER/path_len)
obs = 0
disc_arr = np.array([discount_factor**n for n in range(path_len)])
r_ls = np.array([[np.nan]*path_len for _ in range(path_num)])
ret = []
delay_step = env.num
action_buffer = [-1]*delay_step
model = model_wrapper(model)
for k in range(path_num):
obs = env.reset(random= True)
for i in range(path_len):
action = get_opt_act(q, model, obs, action_buffer)
action_buffer.pop(0)
action_buffer.append(action)
next_obs, reward, _, _ = env.step(action)
# discounted_cum_r = reward + discount_factor*discounted_cum_r
r_ls[k, i] = reward
obs = next_obs
ret.append(np.cumsum(disc_arr*r_ls[k, :]))
return np.mean(ret) ,ret
val, v_samples_lin = eval_pi_model(env, q_learn.q, model, model_wrapper, 0.95)
val
sns.distplot(v_samples_lin, label = 'model')
plt.legend()
df = np.arange(0, 100).reshape(25, -1)
df
# +
# T = 3
T = 3
[N, D] = df.shape
dataX = np.zeros((N - T + 1, T, D))
for i in range(T, N + 1):
dataX[i - T] = df[i - T:i, :]
dataX
# unfolded LSTM
# -
dataX.shape
[np.random.choice(range(24), replace = True) for _ in range(10)]
dataX.reshape(dataX.shape + (1,)).shape
time = range(1,6)
disc = np.array([np.exp(-0.05*t) for t in time])
disc
# +
def A(lamb):
return 0.016*(np.array([np.exp(-n*lamb) for n in time])*disc).sum()
def B(lamb):
return 0.5*0.016*C(lamb)
def C(lamb):
return (disc*np.array([np.exp(-(n-1)*lamb)-np.exp(-n*lamb) for n in time])).sum()
# -
lamb = 0.0154
A(lamb)+B(lamb), C(lamb)
| simplyabcTask-Copy2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gandalf1819/Data-Science-portfolio/blob/master/CS6053_Homework4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-LK0_tE_6xyE" colab_type="text"
# # CS6053 Foundations of Data Science
# ## Homework 4
# + [markdown] id="fuoDg7R56xyJ" colab_type="text"
# Student Name: <NAME>
#
# Student Netid: cnw282
# ***
# + [markdown] id="DkSQEtB36xyL" colab_type="text"
# In this assignment we will be looking at data generated by particle physicists to test whether machine learning can help classify whether certain particle decay experiments identify the presence of a Higgs Boson. One does not need to know anything about particle physics to do well here, but if you are curious, full feature and data descriptions can be found here:
#
# - https://www.kaggle.com/c/higgs-boson/data
# - http://higgsml.lal.in2p3.fr/files/2014/04/documentation_v1.8.pdf
#
# The goal of this assignment is to learn to use cross-validation for model selection as well as bootstrapping for error estimation. We’ll also use learning curve analysis to understand how well different algorithms make use of limited data. For more documentation on cross-validation with Python, you can consult the following:
#
# - http://scikit-learn.org/stable/modules/cross_validation.html#cross-validation
#
# + [markdown] id="6M2rYlpB6xyN" colab_type="text"
# ### Part 1: Data preparation (5 points)
# Create a data preparation and cleaning function that does the following:
# - Has a single input that is a file name string
# - Reads data (the data is comma separated, has a row header and the first column `EventID` is the index) into a pandas `dataframe`
# - Cleans the data
# - Convert the feature `Label` to numeric (choose the minority class to be equal to 1)
# - Create a feature `Y` with numeric label
# - Drop the feature `Label`
# - If a feature has missing values (i.e., `-999`):
# - Create a dummy variable for the missing value
# - Call the variable `orig_var_name` + `_mv` where `orig_var_name` is the name of the actual var with a missing value
# - Give this new variable a 1 if the original variable is missing
# - Replace the missing value with the average of the feature (make sure to compute the mean on records where the value isn't missing). You may find pandas' `.replace()` function useful.
# - After the above is done, rescales the data so that each feature has zero mean and unit variance (hint: look up sklearn.preprocessing)
# - Returns the cleaned and rescaled dataset
#
# Hint: as a guide, this function can easily be done in less than 15 lines.
# + id="ECMpCNFm6xyQ" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn import preprocessing
from collections import Counter
from sklearn.preprocessing import scale
# + id="mPoJCf-T6xyW" colab_type="code" outputId="c3d69763-076f-4400-914f-9f2627cf8cb6" colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="bU_0F5oR-be9" colab_type="code" colab={}
def cleanBosonData(infile_name):
raw=pd.read_csv(infile_name,header=0,index_col=0,na_values=-999)
raw=raw.replace({'Label':{Counter(raw['Label']).most_common()[0][0]:0,Counter(raw['Label']).most_common()[1][0]:1}}).rename(columns={'Label':'Y'})
for col in raw.columns.values[:-1]:
if raw[col].isnull().any().any():
raw[(col+'_mv')]=np.where(raw[col].isnull(),1,0)
raw[col]=raw[col].fillna(raw[col].mean())
raw[col]=scale(raw[col]) # Use StandardScaler from pre-processing library
return raw
# + [markdown] id="zpD-hJs76xya" colab_type="text"
# ### Part 2: Basic evaluations (5 points)
# In this part you will build an out-of-the box logistic regression (LR) model and support vector machine (SVM). You will then plot ROC for the LR and SVM model.
# + [markdown] id="wB3segBL6xyb" colab_type="text"
# 1\. Read and clean the two data files for this assignment (`boson_training_cut_2000.csv` and `boson_testing_cut.csv`) and use them as training and testing data sets. You can find them inside the data folder.
# + id="uR138aI66xyd" colab_type="code" colab={}
data_train = cleanBosonData("/content/gdrive/My Drive/CS6053_HW4/boson_training_cut_2000.csv")
data_test = cleanBosonData("/content/gdrive/My Drive/CS6053_HW4/boson_testing_cut.csv")
# + id="6Z7f7qUkMk-8" colab_type="code" outputId="bf6b5d09-9538-4d15-a7a4-7d87ab38f4c5" colab={"base_uri": "https://localhost:8080/", "height": 258}
data_train.head()
# + id="Q0uKv-_tO5SU" colab_type="code" outputId="f9e819aa-04d7-4805-e10f-d0f3ed898c1b" colab={"base_uri": "https://localhost:8080/", "height": 258}
data_train.tail()
# + [markdown] id="hd04XNou6xyh" colab_type="text"
# 2\. On the training set, build the following models:
#
# - A logistic regression using sklearn's `linear_model.LogisticRegression()`. For this model, use `C=1e30`.
# - An SVM using sklearn's `svm.svc()`. For this model, specify that `kernel="linear"`.
#
# For each model above, plot the ROC curve of both models on the same plot. Make sure to use the test set for computing and plotting. In the legend, also print out the Area Under the ROC (AUC) for reference.
# + id="IsuHMGNt6xyj" colab_type="code" outputId="a795422c-1e79-4a38-e287-d787b9f0f1d1" colab={"base_uri": "https://localhost:8080/", "height": 295}
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import linear_model,svm
from sklearn import metrics
from sklearn.metrics import roc_auc_score, roc_curve
from sklearn import linear_model
from sklearn.svm import SVC
import warnings
warnings.filterwarnings('ignore')
target_var = "Y"
actual = data_test["Y"].copy()
clf_LR = linear_model.LogisticRegression(C=1e30)
clf_LR.fit(data_train.drop(target_var,1),data_train[target_var])
predictions = clf_LR.predict_proba(data_test.drop(target_var,1))[:,1]
clf_svm = svm.SVC(kernel='linear')
clf_svm.fit(data_train.drop(target_var,1),data_train[target_var])
predictions_svm = clf_svm.decision_function(data_test.drop(target_var,1))
# Creating a model using Logistic regression
logreg = linear_model.LogisticRegression(C = 1e30).fit(data_train.drop('Y', 1), data_train['Y'])
# Creating a model using Support Vector Machines
svm=SVC(kernel="linear").fit(data_train.drop('Y',1),data_train['Y'])
pred_logreg=logreg.predict(data_test.drop('Y',1))
pred_svm=svm.predict(data_test.drop('Y',1))
roc_log=roc_curve(y_true=data_test['Y'],y_score=pred_logreg)
roc_svm=roc_curve(y_true=data_test['Y'],y_score=pred_svm)
# pred_logreg=logreg.predict(data_test.drop('Y',1))
# pred_svm=svm.predict(data_test.drop('Y',1))
def plotAUC(truth, pred, lab):
fpr, tpr, thresholds = metrics.roc_curve(truth, pred)
roc_auc = metrics.auc(fpr, tpr)
c = (np.random.rand(), np.random.rand(), np.random.rand())
plt.plot(fpr, tpr, color=c, label= lab+' (AUC = %0.2f)' % roc_auc)
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.xlabel('FPR')
plt.ylabel('TPR')
plt.title('ROC')
plt.legend(loc="lower right")
plotAUC(actual,predictions, 'LR')
plotAUC(actual,predictions_svm, 'SVM')
plt.show()
# + [markdown] id="Jp38dyn06xyo" colab_type="text"
# 3\. Which of the two models is generally better at ranking the test set? Are there any classification thresholds where the model identified above as "better" would underperform the other in a classification metric (such as TPR)?
# + id="WsEQFverWVav" colab_type="code" outputId="6c5650f3-d070-41ca-be37-3637b15ce49c" colab={"base_uri": "https://localhost:8080/", "height": 242}
from sklearn.metrics import confusion_matrix
from IPython.display import display, HTML
import seaborn as sn
import pandas as pd
import matplotlib.pyplot as plt
# Create confusion matrix to measure performance
conf_logreg = pd.DataFrame(confusion_matrix(y_true=data_test['Y'], y_pred=predictions.round()))
conf_svm = pd.DataFrame(confusion_matrix(y_true=data_test['Y'], y_pred=predictions_svm.round()))
conf_log_reg=pd.DataFrame(confusion_matrix(y_true=data_test['Y'],y_pred=pred_logreg))
conf_support_vm=pd.DataFrame(confusion_matrix(y_true=data_test['Y'],y_pred=pred_svm))
display('Logistic')
display(conf_log_reg)
display('SVM')
display(conf_support_vm)
# + [markdown] id="qocpw3aV6xyq" colab_type="text"
# The Logistic model performs better than the SVM model on an average. We judge them by using the AUC metric which is base rate invariant. There are specific thresholds for which one model outperforms the other and the final selection must be made on the basis of the utility of the model.
#
# In Logistic:
#
# We observe that the TPR (predicted 0 and actual 0) is higher than SvM. Logistic correctly classifies actual true values for 6597 cases as compared to SVM's 3251 values.
# The False Negative Error rate is also less. Logistic has 10566 while SVM has 13912 false negatives.
#
# In SVM:
#
# It is better at classifying True Positives than Logistic. It correctly identifies positives for 31586 cases as compared to logistic's 29399 cases.
# The False Positive rate (1251) is also lower than logistic (3438)
#
#
# The logistic regression model looks better at ranking the test set. It has a higher AUC. We can see on this plot that the ROC for the Logistic Regression is higher in every place. Thus, there are no thresholds where Support Vector Machines would have a higher TPR than the Logisitic Regression.
#
#
# + [markdown] id="bUL0fQrF6xys" colab_type="text"
# ### Part 3: Model selection with cross-validation (10 points)
# We think we might be able to improve the performance of the SVM if we perform a grid search on the hyper-parameter $C$. Because we only have 1000 instances, we will have to use cross-validation to find the optimal $C$.
# + [markdown] id="lTApVVc66xyt" colab_type="text"
# 1\. Write a cross-validation function that does the following:
# - Takes as inputs a dataset, a label name, # of splits/folds (`k`), a sequence of values for $C$ (`cs`)
# - Performs two loops
# - Outer Loop: `for each f in range(k)`:
# - Splits the data into `data_train` & `data_validate` according to cross-validation logic
# - Inner Loop: `for each c in cs`:
# - Trains an SVM on training split with `C=c, kernel="linear"`
# - Computes AUC_c_k on validation data
# - Stores AUC_c_k in a dictionary of values
# - Returns a dictionary, where each key-value pair is: `c:[auc-c1,auc-c2,..auc-ck]`
# + id="ikXRF_8W6xyu" colab_type="code" colab={}
from sklearn.model_selection import KFold
from sklearn.svm import SVC
def xValSVM(dataset, label_name, k, cs):
aucs = {}
n_samples = dataset.shape[0]
ksplits = KFold(n_splits = k) # KFold to split dataset into k-groups
for train_index, test_index in ksplits.split(dataset): # Iterate over indices of training and test sets from kfold function
train_k = dataset.iloc[train_index]
test_k = dataset.iloc[test_index]
for c in cs:
svm_clf = SVC(kernel = 'linear', C = c)
svm_clf.fit(train_k.drop(label_name, 1), train_k[label_name])
met = metrics.roc_auc_score(test_k[label_name], svm_clf.decision_function(test_k.drop(label_name,1)))
if c in aucs:
aucs[c].append(met) # Append the AUC scores for each iteration
else:
aucs[c] = [met]
return aucs
# + [markdown] id="G-2ZgSLG6xyz" colab_type="text"
# 2\. Using the function written above, do the following:
# - Generate a sequence of 10 $C$ values in the interval `[10^(-8), ..., 10^1]` (i.e., do all powers of 10 from -8 to 1).
# 2. Call aucs = xValSVM(train, ‘Y’, 10, cs)
# 3. For each c in cs, get mean(AUC) and StdErr(AUC)
# 4. Compute the value for max(meanAUC-StdErr(AUC)) across all values of c.
# 5. Generate a plot with the following:
# a. Log10(c) on the x-axis
# b. 1 series with mean(AUC) for each c
# c. 1 series with mean(AUC)-stderr(AUC) for each c (use ‘k+’ as color pattern)
# d. 1 series with mean(AUC)+stderr(AUC) for each c (use ‘k--‘ as color pattern)
# e. a reference line for max(AUC-StdErr(AUC)) (use ‘r’ as color pattern)
#
# Then answer the question: Did the model parameters selected beat the out-of-the-box model for SVM?
# + id="heZK6G8CWhEE" colab_type="code" outputId="25bccc30-631c-4e07-ae47-f336a923dc5e" colab={"base_uri": "https://localhost:8080/", "height": 354}
from scipy.stats import sem
import matplotlib.pyplot as plt
import pprint
c_val = np.power(10,np.arange(-8.0,2.0))
aucs=xValSVM(data_train,"Y",10,c_val)
# pprint.pprint(aucs)
alldiff=list()
allmeans=list()
allstderr=list()
for i in aucs.keys():
mean=np.mean(aucs[i])
stderr=sem(aucs[i])
# print("Mean for",i,":",mean," Std. Err: ",stderr)
allmeans.append(mean)
allstderr.append(stderr)
alldiff.append(mean-stderr)
print("Maximum difference:", max(alldiff)," for C value",list(aucs.keys())[(alldiff.index(max(alldiff)))])
plt.figure(figsize=(10,5))
plt.plot(np.log10(c_val),allmeans,label='Mean AUC')
plt.plot(np.log10(c_val),np.array(allmeans)-np.array(allstderr),'k+:',label='Mean-Standard error')
plt.plot(np.log10(c_val),np.array(allmeans)+np.array(allstderr),'k--',label='Mean+Standard error')
plt.axhline(y=max(alldiff),color='r',label='Reference Line')
plt.legend(loc='lower right')
plt.xlabel('Log10(c)')
plt.ylabel('roc_auc_score')
plt.show()
# + [markdown] id="ecjDKPsVY3jP" colab_type="text"
# Yes, our optimised model beats the out-of-box SVM model. However, we observe a only marginal increase in the AUC value. Out-of-box SVM with C=1.0 gives us an AUC of 0.5757 while our optimised SVM with C=10.0 gives us an AUC of 0.5768.
# + [markdown] id="_In0S8To6xy1" colab_type="text"
# ### Part 4: Learning Curve with Bootstrapping
# In this HW we are trying to find the best linear model to predict if a record represents the Higgs Boson. One of the drivers of the performance of a model is the sample size of the training set. As a data scientist, sometimes you have to decide if you have enough data or if you should invest in more. We can use learning curve analysis to determine if we have reached a performance plateau. This will inform us on whether or not we should invest in more data (in this case it would be by running more experiments).
#
# Given a training set of size $N$, we test the performance of a model trained on a subsample of size $N_i$, where $N_i<=N$. We can plot how performance grows as we move $N_i$ from $0$ to $N$.
#
# Because of the inherent randomness of subsamples of size $N_i$, we should expect that any single sample of size $N_i$ might not be representative of an algorithm’s performance at a given training set size. To quantify this variance and get a better generalization, we will also use bootstrap analysis. In bootstrap analysis, we pull multiple samples of size $N_i$, build a model, evaluate on a test set, and then take an average and standard error of the results.
#
#
#
# + [markdown] id="_Oc635n_6xy3" colab_type="text"
# 1\. Create a bootstrap function that can do the following:
#
# def modBootstrapper(train, test, nruns, sampsize, lr, c):
#
# - Takes as input:
# - A master training file (train)
# - A master testing file (test)
# - Number of bootstrap iterations (nruns)
# - Size of a bootstrap sample (sampsize)
# - An indicator variable to specific LR or SVM (lr=1)
# - A c option (only applicable to SVM)
#
# - Runs a loop with (nruns) iterations, and within each loop:
# - Sample (sampsize) instances from train, with replacement
# - Fit either an SVM or LR (depending on options specified). For SVM, use the value of C identified using the 1 standard error method from part 3.
# - Computes AUC on test data using predictions from model in above step
# - Stores the AUC in a list
#
# - Returns the mean(AUC) and Standard Error(mean(AUC)) across all bootstrap samples
#
# + id="-amt9Y6s6xy4" colab_type="code" colab={}
def modBootstrapper(train, test, nruns, sampsize, lr, c):
target = 'Y'
aucs_boot = []
for i in range(nruns):
train_samp = train.iloc[np.random.randint(0, len(train), size = sampsize)]
if (lr == 1):
lr_i = linear_model.LogisticRegression(C = 1e30)
lr_i.fit(train_samp.drop(target,1), train_samp[target])
p = lr_i.predict_proba(test.drop(target,1))[:,1]
else:
svm_i = SVC(kernel='linear', C = 0.1)
svm_i.fit(train_samp.drop(target,1), train_samp[target])
p = svm_i.decision_function(test.drop(target,1))
aucs_boot.append(metrics.roc_auc_score(test[target], p))
return [np.mean(aucs_boot), sem(aucs_boot)]
# + [markdown] id="_Jull6re6xy-" colab_type="text"
# 2\. For both LR and SVM, run 20 bootstrap samples for each samplesize in the following list: samplesizes = [50, 100, 200, 500, 1000, 1500, 2000]. (Note, this might take 10-15 mins … feel free to go grab a drink or watch Youtube while this runs).
#
# Generate a plot with the following:
# - Log2(samplesize) on the x-axis
# - 2 sets of results lines, one for LR and one for SVM, the set should include
# - 1 series with mean(AUC) for each sampsize (use the color options ‘g’ for svm, ‘r’ for lr)
# - 1 series with mean(AUC)-stderr(AUC) for each c (use ‘+’ as color pattern, ‘g’,’r’ for SVM, LR respectively)
# - 1 series with mean(AUC)+stderr(AUC) for each c (use ‘--‘ as color pattern ‘g’,’r’ for SVM, LR respectively)
#
# + id="7nkWzUE26xy-" colab_type="code" outputId="cbccf283-6a19-4810-acfd-80f1bb7609b1" colab={"base_uri": "https://localhost:8080/", "height": 446}
import warnings
warnings.filterwarnings('ignore')
SampleSizes = [50, 100, 200, 500, 1000, 1500, 2000]
LR_means = []
Lr_stderr = []
svm_means = []
svm_stderr = []
for n in SampleSizes:
mean, err = modBootstrapper(data_train, data_test, 20, n, 1, 0.1)# collecting means and stderrs for LR model
LR_means.append(mean)
Lr_stderr.append(err)
mean2, err2 = modBootstrapper(data_train, data_test, 20, n, 0, 0.1)# collecting means and stderrs for SVM model
svm_means.append(mean2)
svm_stderr.append(err2)
plt.figure(figsize=(12,7))
plt.plot(np.log2(SampleSizes), LR_means, 'r', label = 'LR means')
plt.plot(np.log2(SampleSizes), LR_means - np.array(Lr_stderr), 'r+-' , label = 'LR means + stderr')
plt.plot(np.log2(SampleSizes), LR_means + np.array(Lr_stderr), 'r--', label = 'LR means + stderr')
plt.plot(np.log2(SampleSizes), svm_means, 'g', label = 'SVM means')
plt.plot(np.log2(SampleSizes), svm_means - np.array(svm_stderr), 'g+-', label = 'SVM means + stderr')
plt.plot(np.log2(SampleSizes), svm_means + np.array(svm_stderr), 'g--', label = 'SVM means - stderr')
plt.legend(loc = 'lower right')
plt.xlabel('Log2(Sample Sizes)')
plt.ylabel('roc_auc_score')
plt.show()
# + [markdown] id="-7o8ypJS6xzB" colab_type="text"
# 3\. Which of the two algorithms are more suitable for smaller sample sizes, given the set of features? If it costs twice the investment to run enough experiments to double the data, do you think it is a worthy investment?
#
# + [markdown] id="yF94NTap6xzD" colab_type="text"
# For smaller sample sizes, we prefer SVM over Logistic regression. By using SVM, we gain a chance to obtain a marginally better result than Logistic. The intervals that bind the average case of SVM cover a higher value of AUC but are also lower than the lower bound of Logistic.
#
# If the cost is twice the investment to run experiments to double the data, we can surely choose Logistic regression as the accuracy gain by SVM is only marginal (0.01% approx). For cases where accuracy is precious, such as in healthcare industry, we can adopt SVM as we need all the extra precision that we can get to prevent wrong predictions.
# + [markdown] id="Uv_len-a6xzE" colab_type="text"
# 4\. Is there a reason why cross-validation might be biased? If so, in what direction is it biased?
#
#
# + [markdown] id="bAZ5hXe36xzF" colab_type="text"
#
#
# Cross validation is used to remove bias from the dataset but it results in an increase in variance. To remove or reduce bias, we take the value of K to be generally three or greater. In this case, we took the cross-validation as K = 10. This results in the development of a model that exhibits low bias due to repeated sampling and averaging of results. Due to this reason, we can safely say that cross-validation is not biased.
| CS6053_Homework4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import rcParams
# Figure S11
def grad(j, j2, Z, eta):
return 1-j/Z-1/(1+np.power(((2*j*j-2*j*Z+Z*Z-Z)/(2*j*Z-2*j*j)), eta))
fig, axs = plt.subplots(1, 2, figsize=(16, 6))
ax = axs[0]
Z1 = 1000 #populationSize
i=0
for eta in [-0.5, 0.0, 1.0, 5.0]:
xx = np.arange(1, Z1-1, 1)
yy = [grad(x, x, Z1, eta) for x in xx]
ax.plot(xx, yy, alpha=1.0, label='\u03B7='+str(eta))
i = i+1
ax.set_ylabel('Gradient, g(j)')
ax.set_xlabel('# intracommunity links, j')
ticksx = np.round(np.arange(0,Z1+1,200),1)
ax.set_xticks(ticksx)
ax.set_xticklabels(ticksx)
ticksy = [-1.0, 0.0, 1.0]
ax.set_yticks(ticksy)
ax.set_yticklabels(ticksy)
ax.grid(True)
ax.legend()
ax = axs[1]
xx = np.arange(0.8, 2.0, 0.01)
yy = [grad(Z1-1, Z1-1, Z1, eta) for eta in xx]
ax.plot(xx, yy, label='\u03B7='+str(eta))
ax.set_ylabel('g(Z-1, \u03B7)')
ax.set_xlabel('Rewire similarity strength, \u03B7')
ticksx = [0.8, 1.0, 1.2, 1.4, 1.6, 1.8, 2.0]
ax.set_xticks(ticksx)
ax.set_xticklabels(ticksx)
ticksy = [0.0]
ax.set_yticks(ticksy)
ax.set_yticklabels(ticksy)
ax.grid(True)
plt.show()
# +
# Figure S12
def gradOpinions(i1, j, N, i2, Z, alpha):
return (N-i1) / N * ((i1*j+i2*(Z-j))/(N * Z))**alpha - i1/N * ((N*Z-i1*j-i2*(Z-j))/(N*Z))**alpha
Ztest = 250
Ntest = 100
xvalues = np.arange(1,Ntest)
yvalues = np.arange(1,Ztest)
xx, yy = np.meshgrid(xvalues, yvalues)
fig, axs = plt.subplots(1, 2, figsize=(16, 6))
ax = axs[0]
ax.streamplot(xx, yy, gradOpinions(xx, yy, Ntest, Ntest, Ztest, 2), grad(yy, yy, Ztest, 1.0))
ax = axs[1]
ax.streamplot(xx, yy, gradOpinions(xx, yy, Ntest, Ntest, Ztest, 2), grad(yy, yy, Ztest, 5.0))
plt.show()
# -
| Analytic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="H9n05_nQvJNL" colab_type="text"
# # **Fashion Class Classification**
# + [markdown] id="3I5hFWgphGRp" colab_type="text"
# ### **Importing the Libraries**
# + id="Wb-oS9LXhGRq" colab_type="code" colab={}
# import libraries
import pandas as pd
import os
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import random
import keras
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Dense, Flatten, Dropout
from keras.optimizers import Adam
from keras.callbacks import TensorBoard
# + [markdown] id="xvQ-Me7KhgvS" colab_type="text"
# ### **Loading the dataset**
# + id="sf2Ogu_BhGRz" colab_type="code" colab={}
fashion_train_df = pd.read_csv('fashion-mnist_train.csv',sep=',')
fashion_test_df = pd.read_csv('fashion-mnist_test.csv', sep = ',')
# + [markdown] id="FpPxD1LxhGR7" colab_type="text"
# ### **Data Exploration**
# + id="xA8nQyWghGR9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 253} outputId="c4786f0e-840b-4c97-9a58-eb39ae52d573"
#head of the training dataset
fashion_train_df.head()
# + id="UhQ6HZ1khGSF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 253} outputId="5eb3d53c-469a-4749-cc32-5c67d7d9191c"
#last elements in the training dataset
fashion_train_df.tail()
# + id="Xj8OpM5XhGSK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 253} outputId="81dc0c48-9684-4039-9d9e-c116f4728559"
#head of the testing dataset
fashion_test_df.head()
# + id="5_YrbbxQhGSN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 253} outputId="4e738461-efda-4043-9dff-85f3093e5936"
#last elements in the testing dataset
fashion_test_df.tail()
# + id="OGkaetgIhGSS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c6be62d1-0b9c-44f1-fa25-86d8e2555acf"
fashion_train_df.shape
# + id="sppeSAnPhGSW" colab_type="code" colab={}
# Create training and testing arrays
training = np.array(fashion_train_df, dtype = 'float32')
testing = np.array(fashion_test_df, dtype='float32')
# + [markdown] id="6VEu2WRqh7zW" colab_type="text"
# ### **Visualisation**
# + id="ckfEEHEdhGSn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="159f834b-b34c-4738-c64c-9c71c0419de0"
#visualising any one image randomly
i=random.randint(1,60000)
plt.imshow(training[i,1:].reshape(28,28))
label=training[i,0]
label
# + id="m8g_TIf4hGSq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aafc6f54-c8a6-4981-c25a-677bf0e510cb"
label = training[i,0]
label
# + id="554-nIOjhGSt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 943} outputId="bb71f3a9-f958-4997-91b1-0d8f8e701565"
#visualising more images in the grid
W = 15
L = 15
fig, axes = plt.subplots(L, W, figsize = (17,17))
axes = axes.ravel()
n = len(training)
for i in np.arange(0, W * L):
index = np.random.randint(0, n)
axes[i].imshow( training[index,1:].reshape((28,28)) )
plt.subplots_adjust(hspace=0.4)
# + [markdown] id="q6Ya0VwUhGSx" colab_type="text"
# ### **Training the model**
# + id="h0KHcWiRhGSx" colab_type="code" colab={}
# Prepare the training and testing dataset
X_train = training[:,1:]/255
y_train = training[:,0]
X_test = testing[:,1:]/255
y_test = testing[:,0]
# + id="SkMOBvi7hGS0" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
X_train, X_validate, y_train, y_validate = train_test_split(X_train, y_train, test_size = 0.2, random_state = 12345)
# + id="g35qFJ7dhGTA" colab_type="code" colab={}
# reshape the data
X_train = X_train.reshape(X_train.shape[0], *(28, 28, 1))
X_test = X_test.reshape(X_test.shape[0], *(28, 28, 1))
X_validate = X_validate.reshape(X_validate.shape[0], *(28, 28, 1))
# + id="RdWvt-W9hGTN" colab_type="code" colab={}
#Encoding the labels
# y_train = keras.utils.to_categorical(y_train, 10)
# y_test = keras.utils.to_categorical(y_test, 10)
# + id="itPkWSoQhGTV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 527} outputId="bcc50c8b-a86b-4915-8556-3b235d24543d"
#CNN architechture
input_img = tf.keras.layers.Input(shape=(28, 28, 1))
x = tf.keras.layers.Conv2D(64,(2,2),activation='relu',padding='same')(input_img)
x = tf.keras.layers.MaxPool2D((2,2))(x)
x = tf.keras.layers.Dropout(0.25)(x)
x = tf.keras.layers.Conv2D(32,(2,2),activation='relu',padding='same')(x)
x = tf.keras.layers.MaxPool2D((2,2))(x)
x = tf.keras.layers.Dropout(0.25)(x)
x = tf.keras.layers.Flatten()(x)
x = tf.keras.layers.Dense(256, activation='relu')(x)
x = tf.keras.layers.Dropout(0.3)(x)
x = tf.keras.layers.Dense(10,activation='sigmoid')(x)
cnn_model=tf.keras.models.Model(input_img,x)
cnn_model.summary()
# + id="kobRDt60hGTY" colab_type="code" colab={}
cnn_model.compile(loss ='sparse_categorical_crossentropy', optimizer='adam',metrics =['accuracy'])
# + id="WpcWLKswhGTa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0bc7b836-d5a1-4577-b0a1-4ea84f76059c"
cnn_model.fit(X_train, y_train, batch_size = 512,epochs = 100,verbose = 1,
validation_data = (X_validate, y_validate))
# + [markdown] id="jrX5nlFKhGTd" colab_type="text"
# ### **Model Evaluation**
# + id="qag7LZ8phGTd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="1ac57752-0125-48bc-b500-8b2451967336"
# Evaluate the model on test set
score = cnn_model.evaluate(X_test, y_test, verbose=0)
# Print test accuracy
print('\n', 'Test accuracy:', score[1])
# + id="KylYkKlShGTg" colab_type="code" colab={}
#predictions for the test data
y_pred = cnn_model.predict(X_test)
# + [markdown] id="2c99-EFLviH8" colab_type="text"
# ### **Visualise the predictions**
# + id="xjgdkUSBhGTj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 697} outputId="f3508929-bff8-45df-d148-2f590f465e9b"
L = 5
W = 5
fig, axes = plt.subplots(L, W, figsize = (12,12))
axes = axes.ravel() #
for i in np.arange(0, L * W):
axes[i].imshow(X_test[i].reshape(28,28))
predict_index = np.argmax(y_pred[i])
true_index = np.argmax(y_test[i])
axes[i].set_title("Prediction Class = {} \n True Class = ({})".format(predict_index, y_test[i]))
axes[i].axis('off')
plt.subplots_adjust(wspace=0.5)
| Fashion_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Week 05, Part 2
#
# ### Topics
# 1. Review: Plotting Percentiles
# 1. Example: comparing SAT & ACT scores
# 1. BACK TO SLIDES FOR PERCENTILE -> ZSCORE
# resize plots
require(repr)
options(repr.plot.width=4, repr.plot.height=4, repr.plot.res=300)
# ## 1. Review: Plotting Percentiles
#
# (Sometimes we do this in this week instead of the last one so its included here just in case).
#
# Let's use some tools in R to visualize percentiles. To do this, we'll take a little detour into the drawing of shapes in R in general.
#
# Let's begin with our old friend, the sequence:
x = seq(-3,3, length=10)
# Now, let's try to understand a polygon function. We'll be using a pre-compiled version but its worth drawing a few by hand to get a feel.
#
# Let's make a triangle. And let's say the triangle goes from -3 to +3 in x & 0-1 in y. We can use our sequence to plot this.
# +
plot(NULL,xlim=c(-3,3),ylim=c(0,1)) # sets up axes, plots nothing.
# Triangle will be defined by 3 verticies:
# (-3, 0), (0, 1), (3, 0)
xvertices = c(-3, 0, 3) # where x "hits" for each vertex
yvertices = c(0, 1, 0) # where y "hits" for each vertex
polygon(xvertices, yvertices,col="red") # plots on top of previous plot
# Lets try overplotting a little rectangle with bottom vertex at
# x = (-1,1), y = (0.4,0.6)
xvertices = c(-1, -1, 1, 1)
yvertices = c(0.4, 0.6, 0.6, 0.4)
polygon(xvertices, yvertices, col="blue")
# -
# What have we learned?
#
# Essentially, `polygon` just fills in between a list of verticies we give it. We can use this to plot underneath our normal distributions. This will help us get a "feel" for how much of the graph is related to our measurement of interest.
# ONTO Plotting percentiles!
#
# Let's source the `plot_polygons` script so that we can use the function in there to plot polygons for us. This was stored in last week, so we will source from there:
source('../week04/plot_polygons.R')
# Let's recall the sum under the curve of our normal probability distribution has to == 1. Given the laws of probability, we can associate every $Z_{score}$ with a percentile.
#
# Let's go back to our toy example of a "normal" normal distribution centered a 0, with SD=1:
x = seq(-3,3,length=200)
y = dnorm(x,mean=0,sd=1)
plot(x,y,type='l')
# What is the $Z_{score}$ for x=0?
Zscore = (0-0)/1 # = 0
# Plot too:
x = seq(-3,3,length=200)
y = dnorm(x,mean=0,sd=1)
plot(x,y,type='l')
abline(v=Zscore, col='red')
# What is the percentile? i.e. what percent under the curve corresponds to a Zscore of 0 here?
#
# First, let's plot with a polygon.
#
# Let's make sure we are only giving vertices up to $Z_{score}$ = 0.
# +
x = seq(-3,3,length=200)
y = dnorm(x,mean=0,sd=1)
plot(x,y,type='l')
abline(v=Zscore, col='red')
# overplot polygon
x2 = seq(-3, Zscore,length=100) # sequence "up to" our Zscore of interest
y2 = dnorm(x2, mean=0, sd=1) # y-values from dnorm
draw_polygon(x2,y2)
# -
# The above call to polygon might seem a little confusing and weird at first. This is just what we did with our simple drawings of triangles and rectanges but now our shape is just a bit more complicated. You can use this code as a reference and plug in the values for your particular problem.
#
# Given the area under the whole curve is 1.0, what do you think the area of the shaded region is?
#
# **TAKE A MOMENT TO THINK ABOUT IT**
#
# By eye we can see that its probably 1/2 or a percentile of 50%. But we can also get an exact number using "pnorm".
print(pnorm(Zscore,mean=0,sd=1))
# Note the similar calling sequence to "dnorm". This will be a constant source of confusion, so you might want to try to remember that "d" stands for probability density and "p" for percentile.
#
# In fact, we can do this percentage-from-Zscore for any Zscore.
#
# Let's try a few:
#
# Make a plot of the percentile associated with negative "X"
plot(x,y,type='l')
Zscore_of_interest = -2
x2 = seq(-3, Zscore_of_interest,length=100)
y2 = dnorm(x2, mean=0, sd=1)
draw_polygon(x2,y2)
# Before calling pnorm, what percentage do you estimate this to be? **TAKE A MOMENT**
#
# Now, print:
print(pnorm(Zscore_of_interest,mean=0,sd=1))
# This is now showing results for the area < some $Z_{score}$. What about if we want to know the area for > some Zscore?
#
# Example with `lower.tail=FALSE`:
plot(x,y,type='l')
x2 = seq(Zscore_of_interest,3,length=100)
y2 = dnorm(x2,mean=0,sd=1)
# Before we where drawing "up to" our Z-score, now we'll draw "starting from" our Z-score:
draw_polygon(x2,y2)
# There are 2 ways to calculate this red area:
#
# i. since we know the whole area, whole probility distribution, = 1 we can use our P(E) = 1-P(not E) type calculation:
print(1-pnorm(Zscore_of_interest,mean=0,sd=1))
# ii. We can also explicity tell pnorm to use the "upper tail" of the distribution:
#
print(pnorm(Zscore_of_interest,mean=0,sd=1,lower.tail=FALSE))
# I can also ask for the percentage of the curve between 2 observations.
#
# Let's start with our original plot:
x = seq(-3,3,length=200)
y = dnorm(x,mean=0,sd=1)
plot(x,y,type='l')
# What percentage of the curve is between -1 and 1? First, lets plot this with our polygon function.
# +
x = seq(-3,3,length=200)
y = dnorm(x,mean=0,sd=1)
plot(x,y,type='l')
# add in area between curves
x2 = seq(-1,1,length=100)
y2 = dnorm(x2,mean=0,sd=1)
draw_polygon(x2,y2)
# -
# Note here we are plotting "between" two Z-scores -1 to 1.
#
# For the calculation of percentage, we have to subtract one from the other like so:
print(pnorm(1.0,mean=0,sd=1)-pnorm(-1.0,mean=0,sd=1))
# Hey look, we get our "68%" part of the "68-95-99" rule. This makes sense since we are measuring 1 SD from the mean in each direction.
# ## 2. Example: comparing SAT & ACT scores
#
# Let's try with an example.
#
# The mean for SAT scores is 1100 & the SD is 200 points. The mean for ACT scores is 21 and the SD is 6. A student gets 1200 on the SATs and 31 on the ACT.
#
# Q1: Use plots to justify which the student did better on.
#
# Q2: Calculate each $Z_{score}$ $\rightarrow$ was your reasoning justified?
#
# Q3: Make plots showing the percentiles with the polygon function, which looks like the larger percentile?
#
# Q4: Calculate each percentile $\rightarrow$ justified Q3?
# **ANS 1:** Plot the both scores:
# +
options(repr.plot.width=8, repr.plot.height=4)
par(mfrow=c(1,2)) # 2 plots
# Plot SATs -- ~3SD out from mean
x=seq(600,1600,length=200)
plot(x,dnorm(x,mean=1100,sd=200),type='l', ylab='SAT')
abline(v=1200,col="blue")
# Plot ACTs -- ~2SD out from mean
x=seq(6,36,length=200)
plot(x,dnorm(x,mean=21,sd=6),type='l', ylab='ACT')
abline(v=31,col="red")
# -
# By eye ACT $Z_{score}$ > SAT $Z_{score}$.
# **ANS 2:**
#
# NOTE: here I'm assuming the max scores for each are effectively infinite, an ok approximation here.
Z_sat = (1200-1100)/200
Z_act = (31-21)/6
print('SAT ACT')
print(c(Z_sat,Z_act))
# 0.5 < 1.666 so this indeed agrees with our intuition.
#
# **ANS 3:**
# +
par(mfrow=c(1,2)) # 2 plots
# SAT
x=seq(600,1600,length=200)
plot(x,dnorm(x,mean=1100,sd=200),type='l', ylab='SAT')
# SAT Percentile for this test score
x2 = seq(600,1200,length=200)
y2 = dnorm(x2,mean=1100,sd=200)
draw_polygon(x2,y2)
# ACT
x=seq(6,36,length=200)
plot(x,dnorm(x,mean=21,sd=6),type='l', ylab='ACT')
# ACT Percentile for this test score
x2 = seq(6,31,length=200)
y2 = dnorm(x2,mean=21,sd=6)
draw_polygon(x2,y2)
# -
# **ANS 4:**
print('SAT, ACT - percentiles')
print(c(pnorm(1200,mean=1100,sd=200),pnorm(31,mean=21,sd=6)))
# So: much higher percentile for ACT
#
# ## BACK TO SLIDES FOR PERCENTILE -> ZSCORE
| week05/.ipynb_checkpoints/prep_notes_part2_week05_drawingPercentiles-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pymedphys.mudensity import single_mlc_pair
import numpy as np
# +
>>> import matplotlib.pyplot as plt
>>> from pymedphys.mudensity import single_mlc_pair
>>> mlc_left = (-2.3, 3.1) # (start position, end position)
>>> mlc_right = (0, 7.7)
>>> x, mu_density = single_mlc_pair(mlc_left, mlc_right)
>>> fig = plt.plot(x, mu_density, '-o')
>>> x
>>> np.round(mu_density, 3)
# +
# Need to make a min_step_per_pixel, have it default to 10.
# -
x, mu_density = single_mlc_pair(
(-400, 400), (400, 400)
)
fig = plt.plot(x[0:-1], mu_density[0:-1], '-o')
# +
fig = plt.plot(x[0:-1], mu_density[0:-1])
linear = (x[0:-1] + 400) / 800
plt.plot(x[0:-1], linear)
# -
x
x[-2]
np.allclose(linear, mu_density[0:-1], atol=0.001)
mu_density[-2]
x, mu_density = single_mlc_pair(
(-400, 400), (-200, 400)
)
mu_density
min_step_per_pixel = 10
start = np.array([-400, -300])
end = np.array([400, 400])
grid_resolution = 0.1
maximum_travel = np.max(np.abs(end - start))
number_of_pixels = np.ceil(maximum_travel / grid_resolution)
time_steps = number_of_pixels * min_step_per_pixel
time_steps
>>> leaf_pair_widths = (2, 2)
>>> mlc = np.array([
... [
... [-400, 400],
... [-400, 300],
... ],
... [
... [400, 400],
... [400, 400],
... ]
... ])
>>> jaw = np.array([
... [-400, -400],
... [400, 400]
... ])
positions = {
'mlc': {
1: (-mlc[0, :, 0], -mlc[1, :, 0]), # left
-1: (mlc[0, :, 1], mlc[1, :, 1]) # right
},
'jaw': {
1: (-jaw[0::-1, 0], -jaw[1::, 0]), # bot
-1: (jaw[0::-1, 1], jaw[1::, 1]) # top
}
}
# +
def calc_time_steps(positions):
maximum_travel = []
for _, value in positions.items():
for _, (start, end) in value.items():
maximum_travel.append(np.max(np.abs(end - start)))
maximum_travel = np.max(maximum_travel)
number_of_pixels = np.ceil(maximum_travel / grid_resolution)
time_steps = number_of_pixels * min_step_per_pixel
return time_steps
calc_time_steps(positions)
| examples/archive/mudensity/09_need_to_change_time_step_method.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### If, Elif, Else Statements
if 1 < 2:
print('Yep!')
if 1 < 2:
print('yep!')
if 1 < 2:
print('first')
else:
print('last')
if 1 > 2:
print('first')
else:
print('last')
if 2 == 2:
print('first')
elif 3 == 3:
print('middle')
else:
print('Last')
| If-Else.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="WpzpsMsncWz2"
# Strings
# ======
#
# String (ou **str** em Python) é um tipo especializado que trata justamente do armazenamento de textos. O tipo **str** também é imutável, o que significa que, uma vez declarada uma string, ela não poderá mudar senão pela criação de uma nova string.
#
#
#
# + [markdown] colab_type="text" id="RTqKSk6uDi1l"
# Inicialização de Strings
# ---------------------------
#
# A inicialização de strings pode ser feita de diversas maneiras, porém, a mais comum é por aspas **_simples_** ou **_duplas_**.
# + colab_type="code" id="qZzIq_Nie3xW" colab={}
# aspas simples
name = 'Fernando'
# + [markdown] colab_type="text" id="-iOIzPvSmSPs"
# ou
# + colab_type="code" id="s3ptPu4KmUMU" colab={}
# aspas duplas
name = "Fernando"
# + [markdown] colab_type="text" id="nyIR9cm2hxMq"
# Operações com Strings
# ---------------------------
#
# É possível realizar as seguintes operações com strings:
#
# - Concatenação
# - Interpolação
# - String como sequência
# - String como objeto
# + [markdown] colab_type="text" id="JAYua-oSEZ0u"
# ### Concatenação
#
# A concatenação, por exemplo, gera uma nova string a partir de outras já existentes. Utiliza-se, para isso, o operador de concatenação "**+**".
# + colab_type="code" id="7rh4TDbs4Zsa" outputId="4e57cf93-d27c-4ee0-8e91-64d6a61f5ac6" colab={"base_uri": "https://localhost:8080/", "height": 34}
# concatena 3 strings formando uma nova string e imprime
print("Uma" + "Nova" + "String")
# + colab_type="code" id="-Far7h_UAc7d" outputId="ec4c24f0-f36a-463f-d540-537280a952b6" colab={"base_uri": "https://localhost:8080/", "height": 34}
# inicializa uma string name com o valor "Fernando"
name = "Fernando"
# concatena a string "Hello, " com a string name
print("Hello, " + name)
# + [markdown] colab_type="text" id="XxabTIDX4sfT"
# ### Interpolação
#
# A interpolação trata-se da substituição de símbolos em posições fixas por variáveis de diferentes tipos. Para isso usa-se a letra f antes do inicio da string. Caso seja preciso formatar os valores passados usamos os seguintes símbolos interpoladores:
#
# - s - string
# - d - inteiro
# - o - octal
# - x - hexadecimal
# - f - real
# - e - real exponencial
# + colab_type="code" id="uUBt8YzME1Vv" outputId="1547eff5-53e1-41bd-c5ff-7a1547c47138" colab={"base_uri": "https://localhost:8080/", "height": 34}
# coding: utf-8
name = "Jennifer"
# troca o símbolo de interpolação %s pela string name e imprime
print(f"O nome dela é {name}")
# + colab_type="code" id="NhJE-HN9I2Bv" outputId="b8a73f5f-00da-402b-b1b1-4f062e710ef5" colab={"base_uri": "https://localhost:8080/", "height": 34}
name = "Jennifer"
age = 20
# troca o simbolo {name} pela string name e o {age} pela variável inteira age e imprime
print(f"{name} tem {age} anos")
# + colab_type="code" id="2hxijpHntgT7" outputId="a997ed3b-e34e-4cfc-89c2-00abe2632d2d" colab={"base_uri": "https://localhost:8080/", "height": 51}
cash = 189.99
# troca o símbolo {cash:.1f} pelo valor 189.99 com arredondamento e uma casa decimal
print(f"Khan tem {cash:.1f} dólares no League of Legends")
# para mostrar o exato valor
print(f"Khan tem {cash:.2f} dólares no League of Legends")
# + [markdown] colab_type="text" id="0flMzG2rv8eB"
# ### String como sequência
#
# As strings, assim como as listas, podem ser tratadas como sequências. Sendo assim, as strings também podem ser:
#
# - Iteradas (ou percorridas)
# - Acessadas por índices
# - Fatiadas
#
# + colab_type="code" id="aYR2Jqot5T3o" outputId="2d7fbbfd-c63b-4d61-b054-0854ec320e33" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Exemplo com Iteração
name = "Nasus"
# Lê-se: Para cada caractere na variável name, imprima o caractere atual
for char in name:
print(char)
# + colab_type="code" id="Xdts_40HCWNr" outputId="6d8bc066-5cac-4e49-db35-694f983c4745" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Exemplo com acesso por índice
name = "<NAME>"
# Imprime o caractere de índice 0 e o caractere de índice 4
print(f"{name[0]}. {name[4]}.")
# + colab_type="code" id="3JiVmf92DbZX" outputId="19801e51-d7a7-40d2-89ba-63af35d27c2d" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Exemplo com fatiamento (slicing)
#
# O fatiamento funciona com ATÉ 3 dados assim como a função range
#
# string[começo:fim+1:intervalo]
#
# Se o começo ou o intervalo não são definidos, o padrão é 0 e 1, respectivamente.
# Se o fim+1 não for definido, será considerado o tamanho do objeto.
name = "<NAME>"
# Imprime da primeira posição até a sétima posição da string name
print(name[0:8])
# Imprime da nona posição até a décima quinta posição da string name
print(name[9:16])
# Por padrão, será impresso a string name por completo
print(name[:])
# Imprime a string name com intervalos de duas letras
print(name[::2])
# Imprime a string name invertida
print(name[::-1])
# + [markdown] colab_type="text" id="rnTW3V1QKBi_"
# ### String como Objeto
#
# As strings também são objetos no Python, o que significa que pode-se utilizar de alguns métodos nativos da linguagem.
#
# Os métodos são acessados por um ponto (.) logo após a string.
#
# Exemplo:
# *string*__.metodo()__
# + colab_type="code" id="bes7RxvrQGnV" outputId="6dd0d64c-000f-4d91-c53f-56c33b9f5a57" colab={"base_uri": "https://localhost:8080/", "height": 34}
name = "<NAME>"
# O método upper() muda todas as letras da string para MAIÚSCULAS.
print(name.upper())
# + colab_type="code" id="puJgIOIhRih6" outputId="dbf082ac-1ab6-4add-a06c-db6cb2e1c412" colab={"base_uri": "https://localhost:8080/", "height": 34}
# coding:utf-8
name = "<NAME>"
# O método capitalize() muda apenas a primeira letra da string para MAIÚSCULA
# e todas as demais para MINÚSCULAS.
print(name.capitalize())
# + [markdown] colab_type="text" id="EE1Jn288SRWe"
# ### Seção Bônus
#
# Nesta seção bônus, serão mostrados os seguintes tópicos:
#
# - Multiplicação de Strings
# - Método format()
# + colab_type="code" id="Yd5036PrT1G4" outputId="3057ed0d-fdbb-4ef2-866c-35bf56c6e109" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Exemplo de Multiplicação de Strings
# Imprime a string "Hello" 3x
print("Hello" * 3)
# É o equivalente ao seguinte código
print("Hello" + "Hello" + "Hello")
# + colab_type="code" id="S0L7j0juUfTJ" outputId="bd6ec8fc-faa1-4ee4-dcab-80b978543a75" colab={"base_uri": "https://localhost:8080/", "height": 68}
# Exemplos de alguns casos de uso do método format()
best_game = "Fortnite"
company = "Epic Games"
# Neste exemplo os parametros vão ser trocados em ordem como na interpolação
print("{} é o melhor jogo da {}.".format(best_game, company))
# Neste exemplo os parametros vão ser trocados pelas palavras-chave
print("{best_game} é o melhor jogo da {company}.".format(best_game="League of Legends", company="Riot Games"))
# Neste exemplo as palavras-chave hora e minuto são formatadas com 2 inteiros
# se não tiver, como é o caso da hora que é 7, adiciona-se um 0 a esquerda
# para completar
print("{saudacao}, são {hora:02d}:{minuto:02d}.".format(saudacao="Bom dia", hora=7, minuto=30))
| prog1/implementacoes/tutoriais/formatacaoDeStrings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mixture Density Networks
#
# Mixture density networks (MDN) (Bishop, 1994) are a class
# of models obtained by combining a conventional neural network with a
# mixture density model.
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import inferpy as inf
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import tensorflow as tf
import tensorflow_probability as tfp
from scipy import stats
from sklearn.model_selection import train_test_split
# +
def plot_normal_mix(pis, mus, sigmas, ax, label='', comp=True):
"""Plots the mixture of Normal models to axis=ax comp=True plots all
components of mixture model
"""
x = np.linspace(-10.5, 10.5, 250)
final = np.zeros_like(x)
for i, (weight_mix, mu_mix, sigma_mix) in enumerate(zip(pis, mus, sigmas)):
temp = stats.norm.pdf(x, mu_mix, sigma_mix) * weight_mix
final = final + temp
if comp:
ax.plot(x, temp, label='Normal ' + str(i))
ax.plot(x, final, label='Mixture of Normals ' + label)
ax.legend(fontsize=13)
def sample_from_mixture(x, pred_weights, pred_means, pred_std, amount):
"""Draws samples from mixture model.
Returns 2 d array with input X and sample from prediction of mixture model.
"""
samples = np.zeros((amount, 2))
n_mix = len(pred_weights[0])
to_choose_from = np.arange(n_mix)
for j, (weights, means, std_devs) in enumerate(
zip(pred_weights, pred_means, pred_std)):
index = np.random.choice(to_choose_from, p=weights)
samples[j, 1] = np.random.normal(means[index], std_devs[index], size=1)
samples[j, 0] = x[j]
if j == amount - 1:
break
return samples
# -
# ## Data
#
# We use the same toy data from
# [<NAME>'s blog post](http://blog.otoro.net/2015/11/24/mixture-density-networks-with-tensorflow/), where he explains MDNs. It is an inverse problem where
# for every input $x_n$ there are multiple outputs $y_n$.
# +
def build_toy_dataset(N):
y_data = np.random.uniform(-10.5, 10.5, N).astype(np.float32)
r_data = np.random.normal(size=N).astype(np.float32) # random noise
x_data = np.sin(0.75 * y_data) * 7.0 + y_data * 0.5 + r_data * 1.0
x_data = x_data.reshape((N, 1))
return x_data, y_data
import random
tf.random.set_random_seed(42)
np.random.seed(42)
random.seed(42)
#inf.setseed(42)
N = 5000 # number of data points
D = 1 # number of features
K = 20 # number of mixture components
x_train, y_train = build_toy_dataset(N)
print("Size of features in training data: {}".format(x_train.shape))
print("Size of output in training data: {}".format(y_train.shape))
sns.regplot(x_train, y_train, fit_reg=False)
plt.show()
# -
# ## Fitting a Neural Network
#
# We could try to fit a neural network over this data set. However, for each x value in this dataset there are multiple y values. So, it poses problems on the use of standard neural networks.
# Let's first define the neural network. We use `tf.keras.layers` to construct neural networks. We specify a three-layer network with 15 hidden units for each hidden layer.
nnetwork = tf.keras.Sequential([
tf.keras.layers.Dense(15, activation=tf.nn.relu),
tf.keras.layers.Dense(15, activation=tf.nn.relu),
tf.keras.layers.Dense(1, activation=None),
])
# The following code fits the neural network to the data
lossfunc = lambda y_out, y: tf.nn.l2_loss(y_out-y)
nnetwork.compile(tf.train.AdamOptimizer(0.1), lossfunc)
nnetwork.fit(x=x_train, y=y_train, epochs=3000)
# +
sess = tf.keras.backend.get_session()
x_test, _ = build_toy_dataset(200)
y_test = sess.run(nnetwork(x_test))
plt.figure(figsize=(8, 8))
plt.plot(x_train,y_train,'ro',x_test,y_test,'bo',alpha=0.3)
plt.show()
# -
# It can be seen, the neural network is not able to fit this data.
# ## Mixture Density Network (MDN)
#
# We use a MDN with a mixture of 20 normal distributions parameterized by a
# feedforward network. That is, the membership probabilities and
# per-component means and standard deviations are given by the output of a
# feedforward network.
#
#
# We define our probabilistic model using InferPy constructs. Specifically, we use the `MixtureGaussian` distribution, where the the parameters of this network are provided by the feedforwrad network.
# +
def neural_network(X):
"""loc, scale, logits = NN(x; theta)"""
# 2 hidden layers with 15 hidden units
net = tf.keras.layers.Dense(15, activation=tf.nn.relu)(X)
net = tf.keras.layers.Dense(15, activation=tf.nn.relu)(net)
locs = tf.keras.layers.Dense(K, activation=None)(net)
scales = tf.keras.layers.Dense(K, activation=tf.exp)(net)
logits = tf.keras.layers.Dense(K, activation=None)(net)
return locs, scales, logits
@inf.probmodel
def mdn():
with inf.datamodel():
x = inf.Normal(loc = tf.ones([D]), scale = 1.0, name="x")
locs, scales, logits = neural_network(x)
y = inf.MixtureGaussian(locs, scales, logits=logits, name="y")
m = mdn()
# -
# Note that we use the `MixtureGaussian` random variable. It collapses
# out the membership assignments for each data point and makes the model
# differentiable with respect to all its parameters. It takes a
# list as input—denoting the probability or logits for each
# cluster assignment—as well as `components`, which are lists of loc and scale values.
#
# For more background on MDNs, take a look at
# [<NAME>'s blog post](http://cbonnett.github.io/MDN.html) or at Bishop (1994).
# ## Inference
#
# Next we train the MDN model. For details, see the documentation about
# [Inference in Inferpy](https://inferpy.readthedocs.io/projects/develop/en/develop/notes/guideinference.html)
# +
@inf.probmodel
def qmodel():
return;
VI = inf.inference.VI(qmodel(), epochs=4000)
m.fit({"y": y_train, "x":x_train}, VI)
# -
# After training, we can now see how the same network embbeded in a mixture model is able to perfectly capture the training data.
# +
X_test, y_test = build_toy_dataset(N)
y_pred = m.posterior_predictive(["y"], data = {"x": X_test}).sample()
plt.figure(figsize=(8, 8))
sns.regplot(X_test, y_test, fit_reg=False)
sns.regplot(X_test, y_pred, fit_reg=False)
plt.show()
# -
# ## Acknowledgments
#
# This tutorial is inspired by [<NAME>'s blog post](http://blog.otoro.net/2015/11/24/mixture-density-networks-with-tensorflow/) and [Edward's tutorial](http://edwardlib.org/tutorials/mixture-density-network).
| notebooks/mixture_density_networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
using PPLM
tokenizer, model = PPLM.get_gpt2();
text, label = PPLM.load_data_from_csv("./../Datasets/train.csv";text_col="comment_text", label_col="toxic", delim=","); # replace with appropriate Dataset path, text_col and label_col
discrim = PPLM.get_discriminator(model; class_size=2);
ind_true = findall(label.==true);
ind_false = findall(label.==false);
using StatsBase
using Random
rng = MersenneTwister(1234);
true_sample = shuffle(rng, ind_true)[1:15000];
false_sample = shuffle(rng, ind_false)[1:15000];
all_samples = cat(true_sample, false_sample, dims=1);
text_reduced = text[all_samples];
label_reduced = label[all_samples];
using CUDA
CUDA.allowscalar(false)
args = PPLM.HyperParams(lr=5e-6,classification_type="MultiClass", epochs=50)
# # Method 1
PPLM.train_discriminator(text_reduced, label_reduced, 8, "Multiclass", 2; lr=5e-6, discrim=discrim, tokenizer=tokenizer, args=args, train_size=0.85, epochs=50);
# # Method 2
# +
(train_x, train_y), (test_x, test_y) = PPLM.splitobs((text_reduced, label_reduced); at=0.8)
train_loader = PPLM.load_cached_data(discrim, train_x, train_y, tokenizer; truncate=true, classification_type="Multiclass");
test_loader = PPLM.load_cached_data(discrim, test_x, test_y, tokenizer; truncate=true, classification_type="Multiclass");
# -
PPLM.train!(discrim, train_loader; args=args)
PPLM.test!(discrim, test_loader; args=args)
# # Save Discriminator
PPLM.save_discriminator(discrim, "toxicity"; file_name="toxicity_classifier_head_768.bson")
# # Load the saved Discriminator
discrim = PPLM.get_discriminator(model; load_from_pretrained=true, discrim="toxicity", class_size=2);
| examples/.ipynb_checkpoints/Julia_Discriminator_Training-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Chapter 8: Mappings & Sets
# ## Content Review
# Read [Chapter 8](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/08_mappings_00_lecture.ipynb) of the book. Then, work through the questions below.
# ### Essay Questions
# Answer the following questions *briefly*!
# **Q1**: `dict` objects are well-suited **to model** discrete mathematical **functions** and to approximate continuous ones. What property of dictionaries is the basis for that claim, and how does it relate to functions in the mathematical sense?
#
# **Q2**: Explain why **hash tables** are a **trade-off** between **computational speed** and **memory** usage!
#
# **Q3:** The `dict` type is an **iterable** that **contains** a **finite** number of key-value pairs. Despite that, why is it *not* considered a **sequence**?
#
# **Q4**: Whereas *key* **look-ups** in a `dict` object run in so-called **[constant time](https://en.wikipedia.org/wiki/Time_complexity#Constant_time)** (i.e., *extremely* fast), that does not hold for *reverse* look-ups. Why is that?
#
# **Q5**: Why is it conceptually correct that the Python core developers do not implement **slicing** with the `[]` operator for `dict` objects?
#
# **Q6**: **Memoization** is an essential concept to know to solve problems in the real world. Together with the idea of **recursion**, it enables us to solve problems in a "backwards" fashion *effectively*.
#
#
# Compare the **recursive** formulation of `fibonacci()` in [Chapter 8](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/08_mappings_00_lecture.ipynb#"Easy-at-third-Glance"-Example:-Fibonacci-Numbers-%28revisited%29), the "*Easy at third Glance*" example, with the **iterative** version in [Chapter 4](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/master/04_iteration_00_lecture.ipynb#"Hard-at-first-Glance"-Example:-Fibonacci-Numbers-%28revisited%29), the "*Hard at first Glance*" example!
#
# How are they similar and how do they differ?
#
# **Q7**: How are the `set` and the `dict` type related? How could we use the latter to mimic the former?
#
# ### True / False Questions
# Motivate your answer with *one short* sentence!
# **Q8**: We may *not* put `dict` objects inside other `dict` objects because they are **mutable**.
#
# **Q9**: **Mutable** objects (e.g., `list`) may generally *not* be used as keys in a `dict` object. However, if we collect, for example, `list` objects in a `tuple` object, the composite object becomes **hashable**.
#
# **Q10**: **Mutability** of a `dict` object works until the underlying hash table becomes too crowded. Then, we cannot insert any items any more making the `dict` object effectively **immutable**. Luckily, that almost never happens in practice.
#
# **Q11**: A `dict` object's [update()](https://docs.python.org/3/library/stdtypes.html#dict.update) method only inserts key-value pairs whose key is *not* yet in the `dict` object. So, it does *not* overwrite anything.
#
# **Q12**: The `set` type is both a mapping and a sequence.
#
| 08_mappings_01_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="kJIRBmDqBiKH" colab_type="text"
# ## Transformer 구현 과정 (1/2)
# 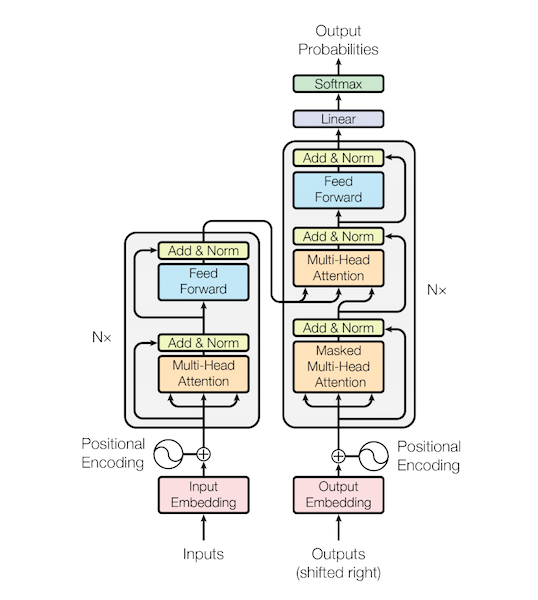
#
# Transformer 모델 구현에 대한 설명 입니다.
#
# 이 내용을 확인하기 전 아래 내용을 확인하시기 바랍니다.
# - [Sentencepiece를 활용해 Vocab 만들기](https://paul-hyun.github.io/vocab-with-sentencepiece/)
# - [Naver 영화리뷰 감정분석 데이터 전처리 하기](https://paul-hyun.github.io/preprocess-nsmc/)
#
# [Colab](https://colab.research.google.com/)에서 실행 했습니다.
# + [markdown] id="ADHWimszCTQB" colab_type="text"
# #### 0. Pip Install
# 필요한 패키지를 pip를 이용해서 설치합니다.
# + id="GvyQ7eeZCV62" colab_type="code" outputId="892adc2b-e3d2-495d-b96f-0ce3669558d5" colab={"base_uri": "https://localhost:8080/", "height": 102}
# !pip install sentencepiece
# + [markdown] id="FWT-P5l0CVde" colab_type="text"
# #### 1. Google Drive Mount
# Colab에서는 컴퓨터에 자원에 접근이 불가능 하므로 Google Drive에 파일을 올려 놓은 후 Google Drive를 mount 에서 로컬 디스크처럼 사용 합니다.
# 1. 아래 블럭을 실행하면 나타나는 링크를 클릭하세요.
# 2. Google 계정을 선택 하시고 허용을 누르면 나타나는 코드를 복사하여 아래 박스에 입력한 후 Enter 키를 입력하면 됩니다.
#
# 학습관련 [데이터 및 결과 파일](https://drive.google.com/open?id=15XGr-L-W6DSoR5TbniPMJASPsA0IDTiN)을 참고 하세요.
# + id="88wMq4r-CdoG" colab_type="code" outputId="3b5835e5-20d4-4fe1-ed10-acb7c192d726" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# data를 저장할 폴더 입니다. 환경에 맞게 수정 하세요.
data_dir = "/content/drive/My Drive/Data/transformer-evolution"
# + [markdown] id="Mw-viWEe_-gb" colab_type="text"
# #### 2. Imports
# + id="AALbOYMx_-Fj" colab_type="code" colab={}
import os
import numpy as np
import math
import matplotlib.pyplot as plt
import sentencepiece as spm
import torch
import torch.nn as nn
import torch.nn.functional as F
# + [markdown] id="kOnh3BmoCzgh" colab_type="text"
# #### 3. 폴더의 목록을 확인
# Google Drive mount가 잘 되었는지 확인하기 위해 data_dir 목록을 확인 합니다.
# + id="AtSMyMiqC1Zz" colab_type="code" outputId="29803586-aeb3-44d0-e5a7-20b7e1047625" colab={"base_uri": "https://localhost:8080/", "height": 136}
for f in os.listdir(data_dir):
print(f)
# + [markdown] id="8Ly5K7I8DMy0" colab_type="text"
# #### 4. Vocab 및 입력
# [Sentencepiece를 활용해 Vocab 만들기](https://paul-hyun.github.io/vocab-with-sentencepiece/)를 통해 만들어 놓은 vocab을 로딩 합니다.
#
# 로딩된 vocab을 이용해 input을 만듭니다.
# + id="S-ubyeJFDbot" colab_type="code" outputId="140e36f4-838c-4eb2-c490-328821139c63" colab={"base_uri": "https://localhost:8080/", "height": 102}
# vocab loading
vocab_file = f"{data_dir}/kowiki.model"
vocab = spm.SentencePieceProcessor()
vocab.load(vocab_file)
# 입력 texts
lines = [
"겨울은 추워요.",
"감기 조심하세요."
]
# text를 tensor로 변환
inputs = []
for line in lines:
pieces = vocab.encode_as_pieces(line)
ids = vocab.encode_as_ids(line)
inputs.append(torch.tensor(ids))
print(pieces)
# 입력 길이가 다르므로 입력 최대 길이에 맟춰 padding(0)을 추가 해 줌
inputs = torch.nn.utils.rnn.pad_sequence(inputs, batch_first=True, padding_value=0)
# shape
print(inputs.size())
# 값
print(inputs)
# + [markdown] id="wOqt_-1p_Z2L" colab_type="text"
# #### 5. Embedding
# + [markdown] id="aAz1I-mG_dSQ" colab_type="text"
# ###### - Input Embedding
# + id="7bjcpE8Z-208" colab_type="code" outputId="b237843e-2451-4ce6-edc6-40ba85aab05a" colab={"base_uri": "https://localhost:8080/", "height": 34}
n_vocab = len(vocab) # vocab count
d_hidn = 128 # hidden size
nn_emb = nn.Embedding(n_vocab, d_hidn) # embedding 객체
input_embs = nn_emb(inputs) # input embedding
print(input_embs.size())
# + [markdown] id="EBriAmpB_cHQ" colab_type="text"
# ###### - Position Embedding
# + id="NlCmyZrkPuOX" colab_type="code" colab={}
""" sinusoid position embedding """
def get_sinusoid_encoding_table(n_seq, d_hidn):
def cal_angle(position, i_hidn):
return position / np.power(10000, 2 * (i_hidn // 2) / d_hidn)
def get_posi_angle_vec(position):
return [cal_angle(position, i_hidn) for i_hidn in range(d_hidn)]
sinusoid_table = np.array([get_posi_angle_vec(i_seq) for i_seq in range(n_seq)])
sinusoid_table[:, 0::2] = np.sin(sinusoid_table[:, 0::2]) # dim 2i
sinusoid_table[:, 1::2] = np.cos(sinusoid_table[:, 1::2]) # dim 2i+1
return sinusoid_table
# + id="x_MrxrxqVyra" colab_type="code" outputId="4559b7b0-5e95-4aef-9587-087eb036bb89" colab={"base_uri": "https://localhost:8080/", "height": 300}
n_seq = 64
pos_encoding = get_sinusoid_encoding_table(n_seq, d_hidn)
print (pos_encoding.shape) # 크기 출력
plt.pcolormesh(pos_encoding, cmap='RdBu')
plt.xlabel('Depth')
plt.xlim((0, d_hidn))
plt.ylabel('Position')
plt.colorbar()
plt.show()
# + id="GmeDRFKqikMy" colab_type="code" outputId="f2ccb64e-8c2f-4468-e70e-58becf9a9965" colab={"base_uri": "https://localhost:8080/", "height": 102}
pos_encoding = torch.FloatTensor(pos_encoding)
nn_pos = nn.Embedding.from_pretrained(pos_encoding, freeze=True)
positions = torch.arange(inputs.size(1), device=inputs.device, dtype=inputs.dtype).expand(inputs.size(0), inputs.size(1)).contiguous() + 1
pos_mask = inputs.eq(0)
positions.masked_fill_(pos_mask, 0)
pos_embs = nn_pos(positions) # position embedding
print(inputs)
print(positions)
print(pos_embs.size())
# + id="ZX63wiFfruY7" colab_type="code" colab={}
input_sums = input_embs + pos_embs
# + [markdown] id="rOYl2XUyt2xc" colab_type="text"
# #### 6. Scale Dot Product Attention
#
# 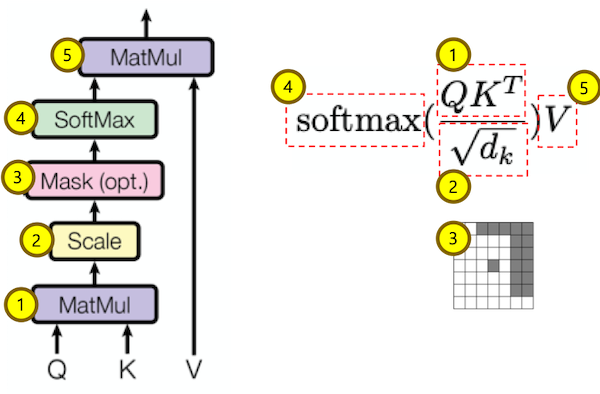
# + [markdown] id="Exfnhn9vA1fl" colab_type="text"
# ###### Input
# + id="aaUTzxeSunt9" colab_type="code" outputId="674683f7-1161-485a-f437-551381d8cdf6" colab={"base_uri": "https://localhost:8080/", "height": 170}
Q = input_sums
K = input_sums
V = input_sums
attn_mask = inputs.eq(0).unsqueeze(1).expand(Q.size(0), Q.size(1), K.size(1))
print(attn_mask.size())
print(attn_mask[0])
# + [markdown] id="st1N5098A7e1" colab_type="text"
# ##### Q * K-transpose
# + id="DQmtb8hcxK1k" colab_type="code" outputId="8f446a21-98c6-46c0-9713-a7af67446c22" colab={"base_uri": "https://localhost:8080/", "height": 306}
scores = torch.matmul(Q, K.transpose(-1, -2))
print(scores.size())
print(scores[0])
# + [markdown] id="2e7jrCXpBCyS" colab_type="text"
# ###### Scale
# + id="tS_LXRbY5hcK" colab_type="code" outputId="67123751-e82b-4a84-d861-5df6a037b7e3" colab={"base_uri": "https://localhost:8080/", "height": 187}
d_head = 64
scores = scores.mul_(1/d_head**0.5)
print(scores.size())
print(scores[0])
# + [markdown] id="lnQ7phqTBGJG" colab_type="text"
# ###### Mask (Opt.)
# + id="BENnRJdKxits" colab_type="code" outputId="ee834637-15cf-4ec7-8f66-26716e903c43" colab={"base_uri": "https://localhost:8080/", "height": 306}
scores.masked_fill_(attn_mask, -1e9)
print(scores.size())
print(scores[0])
# + [markdown] id="GaAzVRmhBJ3B" colab_type="text"
# ###### Softmax
# + id="HKl9riPRxqkj" colab_type="code" outputId="85f1d9ab-ccfe-4f2a-c07e-ad32fb84713e" colab={"base_uri": "https://localhost:8080/", "height": 306}
attn_prob = nn.Softmax(dim=-1)(scores)
print(attn_prob.size())
print(attn_prob[0])
# + [markdown] id="h-MwaJ1oBNOe" colab_type="text"
# ###### atten_prov * V
# + id="P2aRJH-Rxyq8" colab_type="code" outputId="10567189-7867-41ed-fde5-a45e24a8152c" colab={"base_uri": "https://localhost:8080/", "height": 34}
context = torch.matmul(attn_prob, V)
print(context.size())
# + [markdown] id="WR0Wh1ORBRJK" colab_type="text"
# ###### Implementation Class
# + id="EkXGHazDt1rb" colab_type="code" colab={}
""" scale dot product attention """
class ScaledDotProductAttention(nn.Module):
def __init__(self, d_head):
super().__init__()
self.scale = 1 / (d_head ** 0.5)
def forward(self, Q, K, V, attn_mask):
# (bs, n_head, n_q_seq, n_k_seq)
scores = torch.matmul(Q, K.transpose(-1, -2)).mul_(self.scale)
scores.masked_fill_(attn_mask, -1e9)
# (bs, n_head, n_q_seq, n_k_seq)
attn_prob = nn.Softmax(dim=-1)(scores)
# (bs, n_head, n_q_seq, d_v)
context = torch.matmul(attn_prob, V)
# (bs, n_head, n_q_seq, d_v), (bs, n_head, n_q_seq, n_v_seq)
return context, attn_prob
# + [markdown] id="yK38SMqGsiXL" colab_type="text"
# #### 7. Multi-Head Attention
#
# 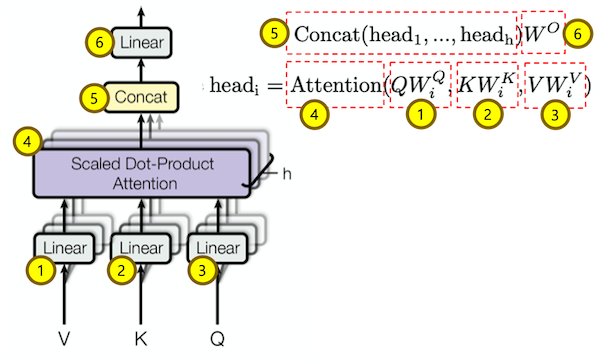
# + [markdown] id="i7F4mEgmBjuw" colab_type="text"
# ###### Input
# + id="rZyMvxiathB-" colab_type="code" colab={}
Q = input_sums
K = input_sums
V = input_sums
attn_mask = inputs.eq(0).unsqueeze(1).expand(Q.size(0), Q.size(1), K.size(1))
batch_size = Q.size(0)
n_head = 2
# + [markdown] id="Mly3n1YcBnR7" colab_type="text"
# ###### Multi Head Q, K, V
# + id="Pes5Jait2EXR" colab_type="code" outputId="c527de1b-b28a-4885-ee72-3d980632223b" colab={"base_uri": "https://localhost:8080/", "height": 68}
W_Q = nn.Linear(d_hidn, n_head * d_head)
W_K = nn.Linear(d_hidn, n_head * d_head)
W_V = nn.Linear(d_hidn, n_head * d_head)
# (bs, n_seq, n_head * d_head)
q_s = W_Q(Q)
print(q_s.size())
# (bs, n_seq, n_head, d_head)
q_s = q_s.view(batch_size, -1, n_head, d_head)
print(q_s.size())
# (bs, n_head, n_seq, d_head)
q_s = q_s.transpose(1,2)
print(q_s.size())
# + id="oW9s0Oam3ZeG" colab_type="code" outputId="ff64f15d-0c2f-487e-e45b-fbcb793b87f8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# (bs, n_head, n_seq, d_head)
q_s = W_Q(Q).view(batch_size, -1, n_head, d_head).transpose(1,2)
# (bs, n_head, n_seq, d_head)
k_s = W_K(K).view(batch_size, -1, n_head, d_head).transpose(1,2)
# (bs, n_head, n_seq, d_head)
v_s = W_V(V).view(batch_size, -1, n_head, d_head).transpose(1,2)
print(q_s.size(), k_s.size(), v_s.size())
# + [markdown] id="u73qrnhfBsYR" colab_type="text"
# ###### Multi Head Attention Mask
# + id="n9ItmgzN32fr" colab_type="code" outputId="1f32a3d3-a145-4671-c15d-a10635608777" colab={"base_uri": "https://localhost:8080/", "height": 51}
print(attn_mask.size())
attn_mask = attn_mask.unsqueeze(1).repeat(1, n_head, 1, 1)
print(attn_mask.size())
# + [markdown] id="ubh168SyBxnw" colab_type="text"
# ###### Attention
# + id="ufWp4KXo6yyi" colab_type="code" outputId="52021897-5830-42e7-d407-fd5b2dff8876" colab={"base_uri": "https://localhost:8080/", "height": 51}
scaled_dot_attn = ScaledDotProductAttention(d_head)
context, attn_prob = scaled_dot_attn(q_s, k_s, v_s, attn_mask)
print(context.size())
print(attn_prob.size())
# + [markdown] id="0zRmgcEfB1lM" colab_type="text"
# ###### Concat
# + id="EjQQPFsn7U8q" colab_type="code" outputId="a3ac5624-652d-46bc-b5e3-b98f37b949e8" colab={"base_uri": "https://localhost:8080/", "height": 34}
# (bs, n_seq, n_head * d_head)
context = context.transpose(1, 2).contiguous().view(batch_size, -1, n_head * d_head)
print(context.size())
# + [markdown] id="6bxvZK6uB4WS" colab_type="text"
# ###### Linear
# + id="kz3C9ABv7st1" colab_type="code" outputId="d0ad361d-e594-409e-e3d6-29ecf84a110b" colab={"base_uri": "https://localhost:8080/", "height": 34}
linear = nn.Linear(n_head * d_head, d_hidn)
# (bs, n_seq, d_hidn)
output = linear(context)
print(output.size())
# + [markdown] id="wwcq-wgvB8Hq" colab_type="text"
# ###### Implementation Class
# + id="pVQZieaCB7xp" colab_type="code" colab={}
""" multi head attention """
class MultiHeadAttention(nn.Module):
def __init__(self, d_hidn, n_head, d_head):
super().__init__()
self.d_hidn = d_hidn
self.n_head = n_head
self.d_head = d_head
self.W_Q = nn.Linear(d_hidn, n_head * d_head)
self.W_K = nn.Linear(d_hidn, n_head * d_head)
self.W_V = nn.Linear(d_hidn, n_head * d_head)
self.scaled_dot_attn = ScaledDotProductAttention(d_head)
self.linear = nn.Linear(n_head * d_head, d_hidn)
def forward(self, Q, K, V, attn_mask):
batch_size = Q.size(0)
# (bs, n_head, n_q_seq, d_head)
q_s = self.W_Q(Q).view(batch_size, -1, self.n_head, self.d_head).transpose(1,2)
# (bs, n_head, n_k_seq, d_head)
k_s = self.W_K(K).view(batch_size, -1, self.n_head, self.d_head).transpose(1,2)
# (bs, n_head, n_v_seq, d_head)
v_s = self.W_V(V).view(batch_size, -1, self.n_head, self.d_head).transpose(1,2)
# (bs, n_head, n_q_seq, n_k_seq)
attn_mask = attn_mask.unsqueeze(1).repeat(1, self.n_head, 1, 1)
# (bs, n_head, n_q_seq, d_head), (bs, n_head, n_q_seq, n_k_seq)
context, attn_prob = self.scaled_dot_attn(q_s, k_s, v_s, attn_mask)
# (bs, n_head, n_q_seq, h_head * d_head)
context = context.transpose(1, 2).contiguous().view(batch_size, -1, self.n_head * self.d_head)
# (bs, n_head, n_q_seq, e_embd)
output = self.linear(context)
# (bs, n_q_seq, d_hidn), (bs, n_head, n_q_seq, n_k_seq)
return output, attn_prob
# + [markdown] id="4O7PH-v-SA9B" colab_type="text"
# #### 8. Masked Multi Head Attention
# + id="sg1uvsXJSAJE" colab_type="code" outputId="d4bcd74c-7bb5-439b-bf71-2594cd4d850a" colab={"base_uri": "https://localhost:8080/", "height": 425}
""" attention decoder mask """
def get_attn_decoder_mask(seq):
subsequent_mask = torch.ones_like(seq).unsqueeze(-1).expand(seq.size(0), seq.size(1), seq.size(1))
subsequent_mask = subsequent_mask.triu(diagonal=1) # upper triangular part of a matrix(2-D)
return subsequent_mask
Q = input_sums
K = input_sums
V = input_sums
attn_pad_mask = inputs.eq(0).unsqueeze(1).expand(Q.size(0), Q.size(1), K.size(1))
print(attn_pad_mask[0])
attn_dec_mask = get_attn_decoder_mask(inputs)
print(attn_dec_mask[0])
attn_mask = torch.gt((attn_pad_mask + attn_dec_mask), 0)
print(attn_mask[0])
batch_size = Q.size(0)
n_head = 2
# + id="-YdCqAETUSzv" colab_type="code" outputId="c927ad28-8967-4fc8-c614-6324467fa053" colab={"base_uri": "https://localhost:8080/", "height": 34}
attention = MultiHeadAttention(d_hidn, n_head, d_head)
output, attn_prob = attention(Q, K, V, attn_mask)
print(output.size(), attn_prob.size())
# + [markdown] id="xPiW76dzGy8D" colab_type="text"
# #### 9. Feed Forward
#
#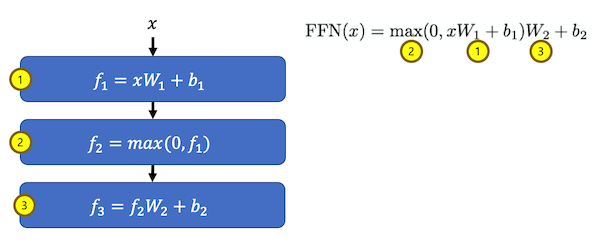 
# + [markdown] id="nLWqtn7UHGz-" colab_type="text"
# ###### f1 (Liear)
# + id="U0iAPIfCHGaa" colab_type="code" outputId="484ee583-3dc9-4401-9007-6f4c79c8cb7f" colab={"base_uri": "https://localhost:8080/", "height": 34}
conv1 = nn.Conv1d(in_channels=d_hidn, out_channels=d_hidn * 4, kernel_size=1)
# (bs, d_hidn * 4, n_seq)
ff_1 = conv1(output.transpose(1, 2))
print(ff_1.size())
# + [markdown] id="ezEDjlVtIqf0" colab_type="text"
# ###### Activation (relu or gelu)
#
# 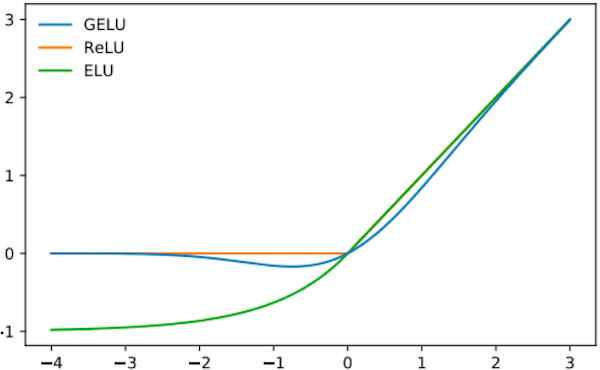
# + id="x7mTDJZiIOrL" colab_type="code" colab={}
# active = F.relu
active = F.gelu
ff_2 = active(ff_1)
# + [markdown] id="fAcrKhAtIvn5" colab_type="text"
# ###### f3 (Liear)
# + id="xGRsfIOcOgLR" colab_type="code" outputId="427c31fb-2691-4524-f382-c0294143286c" colab={"base_uri": "https://localhost:8080/", "height": 34}
conv2 = nn.Conv1d(in_channels=d_hidn * 4, out_channels=d_hidn, kernel_size=1)
ff_3 = conv2(ff_2).transpose(1, 2)
print(ff_3.size())
# + [markdown] id="sE6-A3OhIzLo" colab_type="text"
# ###### Implementation Class
# + id="TaRkgC4CRs60" colab_type="code" colab={}
""" feed forward """
class PoswiseFeedForwardNet(nn.Module):
def __init__(self, d_hidn):
super().__init__()
self.conv1 = nn.Conv1d(in_channels=self.config.d_hidn, out_channels=self.config.d_hidn * 4, kernel_size=1)
self.conv2 = nn.Conv1d(in_channels=self.config.d_hidn * 4, out_channels=self.config.d_hidn, kernel_size=1)
self.active = F.gelu
def forward(self, inputs):
# (bs, d_ff, n_seq)
output = self.active(self.conv1(inputs.transpose(1, 2)))
# (bs, n_seq, d_hidn)
output = self.conv2(output).transpose(1, 2)
# (bs, n_seq, d_hidn)
return output
| tutorial/transformer-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Profiling code in IPython
# We will see how we can use IPython to profile our code.
#
# Here is a python function that does some calculations:
def sum_of_lists(N):
total = 0
for i in range(5):
L = [j ^ (j >> i) for j in range(N)]
total += sum(L)
return total
# ## Profiling execution time
# %prun sum_of_lists(1000000)
# The output looks like this:
# ```
# 14 function calls in 0.512 seconds
#
# Ordered by: internal time
#
# ncalls tottime percall cumtime percall filename:lineno(function)
# 5 0.452 0.090 0.452 0.090 <ipython-input-10-f105717832a2>:4(<listcomp>)
# 1 0.029 0.029 0.503 0.503 <ipython-input-10-f105717832a2>:1(sum_of_lists)
# 5 0.022 0.004 0.022 0.004 {built-in method builtins.sum}
# 1 0.009 0.009 0.512 0.512 <string>:1(<module>)
# 1 0.000 0.000 0.512 0.512 {built-in method builtins.exec}
# 1 0.000 0.000 0.000 0.000 {method 'disable' of '_lsprof.Profiler' objects}
# ```
# ## Profiling code line by line
# !pip -q install line_profiler
# %load_ext line_profiler
# %lprun -f sum_of_lists sum_of_lists(1000000)
# The output looks like this:
# ```
# Timer unit: 1e-06 s
#
# Total time: 0.86063 s
# File: <ipython-input-10-f105717832a2>
# Function: sum_of_lists at line 1
#
# Line # Hits Time Per Hit % Time Line Contents
# ==============================================================
# 1 def sum_of_lists(N):
# 2 1 2 2.0 0.0 total = 0
# 3 6 11 1.8 0.0 for i in range(5):
# 4 5 833118 166623.6 96.8 L = [j ^ (j >> i) for j in range(N)]
# 5 5 27498 5499.6 3.2 total += sum(L)
# 6 1 1 1.0 0.0 return total
# ```
#
| machine_learning/jupyter_101/My first jupyter notebook.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (F#)
// language: F#
// name: .net-fsharp
// ---
#r "nuget:TaskBuilder.fs,2.1.0"
#r "TakensTheorem.Core.dll"
open TakensTheorem.Core
open KaggleClient
//open System
open System.Net.Http
open FSharpKernelHelpers
[<RequireQualifiedAccess>]
module ProgressBar =
let Create className statusText =
sprintf "<div class='%s'>
<div class='progress'>
<div class='progress-bar' role='progressbar'
style='width: 1%%;' aria-valuenow='1' aria-valuemin='0'
aria-valuemax='100'>1%%</div>
</div>
<div><code class='progress-status'>%s</code></div>
</div>" className statusText
|> HtmlString
let CreateWithPocketView (className:string) (statusText: string) =
div.["class",className].innerHTML [
div.["class","progress"].innerHTML (
div.["class", "progress-bar"].["role", "progressbar"]
.["style", "width: 1%"].["aria-valuenow", 1]
.["aria-valuemin", 0 ].["aria-valuemax", 100] )
div.innerHTML (code.["class","progress-status"].innerHTML statusText) ]
let Update className status value =
let str = StringBuilder()
sprintf "$('.%s .progress-status').text('%s');" className status
|> str.AppendLine
sprintf "$('.%s .progress-bar')
.css('width','%.02f%%')
.prop('aria-valuenow',%.02f)
.text('%.02f%%');" className value value value
|> str.AppendLine
str.ToString() |> Javascript
let Report() =
let guid = Guid.NewGuid()
let pbarClass = sprintf "bar-%O" guid
let markerClass = sprintf "code-%O" guid
ProgressBar.CreateWithPocketView pbarClass ""
|> display
|> ignore
sprintf "<div class='%s'></div>" markerClass
|> HtmlString
|> display
|> ignore
fun (file:string, bytesRead : int64, totalBytes: int64) ->
let status =
sprintf "Downloading file [%s] -- %dKB of %.02fMB received."
(file.Replace("\\","/")) bytesRead (float totalBytes/1024.0/1024.0)
let percentage = float bytesRead / float totalBytes * 100.0
ProgressBar.Update pbarClass status percentage
sprintf "$('.%s') \
.closest('.output') \
.find('.output_area') \
.each((i,el)=>{ if (i > 1) $(el).remove(); });" markerClass
|> Javascript
// +
let kaggleJsonPath = "kaggle.json"
let client = new HttpClient()
{ DatasetInfo =
{ Owner = "selfishgene"
Dataset = "historical-hourly-weather-data"
Request = CompleteDatasetZipped }
AuthorizedClient =
kaggleJsonPath
|> Credentials.LoadFrom
|> Credentials.AuthorizeClient client
DestinationFolder = "../Data"
CancellationToken = None
ReportingCallback = Some (Report()) }
|> DownloadDatasetAsync
|> Async.AwaitTask
|> Async.RunSynchronously
// -
| Notebooks/Download Weather Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
# %run main --device-ids 0 1 2 3 --batch-size 1996 --study-num-layer 6 --study-num-ts 16 --study-num-neurons 32
| checklist_transformers/.ipynb_checkpoints/Negation_example_on_subset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 ('tackle')
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# +
np.random.seed(0)
values = np.random.randn(100) # array of normally distributed random numbers
s = pd.Series(values) # generate a pandas series
s.plot(kind='hist', title='Normally distributed random values') # hist computes distribution
plt.show()
| code/hello_world/hello_world.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Pizza-Oven: Cooking with a GPU at home.
#
# > I built a machine with a consumer GPU that keeps me cooking with gas at home. It was a fun project and although I will spin up a Colab instance or other "hosted" solutions from time-to-time this is my everyday _oven_ for cooking up data into something tasty.
#
#
# ## Outline
#
# 1. What I wanted to do, and why.
# 2. What my constraints: \\$<1000
# 3. next?? At some point I will add some security and configure my ATT router to allow my to cook in my pizza oven from wherever I may be.
#
#
# Basically I set up the machine to run linux and act mostly as a jupyter notebook server which i connect thorugh standard SSH tunnels (though sometimes I use it via VSCode instead of a notebook). I can use my laptop wherever I may be in the house, or use my desktop machine in my office (the Pizza-Oven lives in the freezing cold garage).
#
#
# ## Description
#
# Use these commands to run a jupyter server on a remote server and access it locally in a browser. You must have SSH access to the browser. You can use any port you want, but Ill use "8888" for the "server port" and 8889 for the port on the machine I'm connecting from. E.g. on my laptop I type "localhost:8888" in my browser. (Firefox please)
#
# [cite ]
# **Do not forget to change** `username@server` **to the correct value!**
#
# ## TLDR
#
# | where | using | command |
# | ------ | -------- | ------------------------------------------------ |
# | server | terminal | `jupyter notebook --no-browser --port=8888` |
# | local | terminal | `ssh -v -NL 8888:localhost:8888 username@server` |
# | local | browser | `localhost:8888` |
#
# ## Server
#
# ``` bash
# screen -R JUPYT
# conda activate fastai
# jupyter notebook --no-browser --port=8888
#
# ```
#
# This starts the jupyter notebook. Keep the terminal open or run it in the background using screen, tmux, nohup, etc.
#
# ```bash
# jupyter notebook --no-browser --port=8888
# ```
#
# For example, you can start it in the background with screen.
#
# ```bash
# screen -d -m -S JUPYTER jupyter notebook --no-browser --port=8888
# ```
#
# You can reattach to the screen session if need be. Use `Ctrl+a+d` to detach.
#
# ```bash
# screen -r JUPYTER
# ```
#
# You can stop the screen session altogether, killing the jupyter server.
#
# ```bash
# screen -ls
# screen -X -S JUPYTER quit
# ```
#
# ## Local
#
# I typically use a zsh (on a mac) as my terminal. So I created a little function in my .zshrc
#
# This creates an SSH tunnel. It makes localhost:8888 point to the server's 8888 port. Fill in the correct `username@server`!
#
#
#
# ```bash
#
# function jpt(){
# # Fires-up a Jupyter notebook by supplying a specific port
# jupyter notebook --no-browser --port=$1
# } # this runs on the GPU machine
# # you would type jpt 8890 on pizzapie (GPU)
#
#
# function remote_jpt(){
# ssh -N -f -L localhost:$2":localhost:"$1 $desktop_ip
# # ssh -N -L local-address:local-port:remote-address:remote-port username:remote-user@remote-host
# } # type jptt 8890 8888 on pizzadashit (x1e)
#
# ```
# "desktop_ip" is the name of the remote
#
# "-N" is saying don't execute any commands after the "tunnel" is established. That's a little extra security.
#
# "-L" is indicating that we want a Local port forward to the remote machine (-R goes the other way).
#
# "local-address" is optional, but good to use, since it restricts what system can access the port.
# "local-port" is the forwarded port
#
# ```bash
#
# function jpt_tunnel(){
# # "-N" is saying don't execute any commands after the "tunnel" is established. That's a little extra security.
# # "-L" is indicating that we want a Local port forward to the remote machine (-R goes the other way).
# # "local-address" is optional, but good to use, since it restricts what system can access the port.
# # "local-port" is the forwarded port
# screen -d -m -S TUNNEL ssh -N -f -L localhost:$2":localhost:"$1 $desktop_ip
# # ssh -N -L local-address:local-port:remote-address:remote-port username:remote-user@remote-host
# } # type jptt 8890 8888 on pizzadashit (x1e)
#
#
# ```bash
# # USE THIS !!
# function find_remote_jpt(){
# # find the tunnel process on port ARG
# ps aux | grep localhost:$1
# }
# ```
#
#
#
# this is a copy of what runs on the server
# ```bash
#
# # PROBAABLY NEVER USE THIS
# function jpt(){
# # Fires-up a Jupyter notebook by supplying a specific port
# jupyter notebook --no-browser --port=$1
# } # this runs on the GPU machine
# # you would type jpt 8890 on pizzapie (GPU)
# ```
#
#
# this is the crucial one
# ```bash
# # USE THIS !!
# remote_ip=<EMAIL>
# function remote_jpt(){
# # "-N" is saying don't execute any commands after the "tunnel" is established. That's a little extra security.
# # "-L" is indicating that we want a Local port forward to the remote machine (-R goes the other way).
# # "local-address" is optional, but good to use, since it restricts what system can access the port.
# # "local-port" is the forwarded port
# ssh -N -f -L localhost:$2":localhost:"$1 $remote_ip
# # ssh -N -L local-address:local-port:remote-address:remote-port username:remote-user@remote-host
# } # type jptt 8890 8888 on pizzadashit (x1e)
# ```
#
# Not using this...
#
# ```bash
# function jpt_tunnel(){
# # this will only work if we don't need a password... need to use keys...
# screen -d -m -S TUNNEL ssh -N -f -L localhost:$2":localhost:"$1 $remote_ip
# # ssh -N -L local-address:local-port:remote-address:remote-port username:remote-user@remote-host
# } # type jptt 8890 8888 on pizzadashit (x1e)
#
#
# ```
#
#
# ```bash
# ssh -v -NL 8888:localhost:8888 username@server
# ```
#
# Again, you can start it in the background with screen if you don't want to leave the terminal window open.
#
# ```bash
# screen -d -m -S TUNNEL ssh -v -NL 8888:localhost:8888 username@server
# ```
#
# ## Browser
#
# This is the web address you have to open in a browser on the local machine.
#
# ```
# localhost:8888
# ```
#
# ## Screen
#
# Here are some useful screen commands. Use screen to run things in the background (like the jupyter notebook).
#
# ```
# screen -ls # see whats running
# screen -d -m -S NAME command # start a screen
# screen -X -S NAME quit # stop a screen
# screen -r NAME # attach to a screen
# ```
# ## References
#
# * http://www.justinkiggins.com/blog/zero-configuration-remote-jupyter-server/
# ## SSH server settings
#
#
# ### security
#
# - turn off passwords. pass public keys
# - dont use the default SSH port (22). use something else (e.g. 2243)
# -
#
#
# > `vim /etc/ssh/sshd_config`
# ``` bash
# # we will have sudo for proper logins, so turn off root login
# PermitRootLogin no
#
# # don't allow plaintext passwords
# PasswordAuthentication no
#
# ```
#
#
# Save and close the file. To implement your changes, restart the SSH daemon.
#
# >`sudo service ssh restart`
#
#
# > `vim .ssh/config`
# ``` bash
# # aliases to make loging in on the right port easy
#
# Host home_gpu_server
# HostName home_gpu_server.local
# Port 2243
# ```
#
# references: https://www.digitalocean.com/community/tutorials/ssh-essentials-working-with-ssh-servers-clients-and-keys
#
#
# ### router settings
#
#
# ATT Arris BWG210 modem/router. ATT won't let me set it up to just use the modem and pass onto my own router (Google Nest with 2x sattelite mesh) so I need to set it up for "bridge mode". I need to forward the right ports and ips so the Nest router can operate correctly.
#
# NOTE: its set up to use a DHCP range from the Arris which from which the Nest router dynammically allocates IPs to devices on my network.
#
# I need to pass my actuall IP through to my machine with the GPU which i use as a jupyter server. And also expose the nescessare ports for SSH tunnels into the machine from the outside world.
#
# Note that once you do this the router can be found at: http://192.168.48.1/ I also left a low-powered 2.4GHz wireless signal running directly from the Arris (wiht an ssid / password I have handy) so I can easily log into change things withough pluggin in with ethernet. (Because frankly, modern laptops lack an ethernet port, and who wants to dig for a dongle when your network is on the fritz?)
#
# This was the definitive guide I followed, as no "official" guide exists. (this one?)
# https://www.reddit.com/r/Ubiquiti/comments/b1x5l6/how_to_properly_configure_the_arris_bgw210_for/
#
# ## The Build:
#
#
# - PowermacG5
# - Aorus Pro Wifi $169
# - Intel 9900k ~$450
# - GTX 2070 $400
# - 64GB memory
# - coolers/pwr supply
#
#
#
# 1. Gutted the machine
# 2. Cut out the peripherals
# 3. epoxied the motherboard attachments in place
# 4. install pop!_os (great built in nvidia drivers)
# 5. installed cuda / updated drivers...
# 6. COOK!
#
#
#
# side-blog - pics, inspiration, parts list
| _notebooks/2021-03-04-GPU_at_home.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Standard python helper libraries.
import os, sys, re, json, time, wget, csv, string, time, random
import itertools, collections
from importlib import reload
from IPython.display import display
# NumPy and SciPy for matrix ops
import numpy as np
import scipy.sparse
# NLTK for NLP utils
import nltk
nltk.download('punkt')
nltk.download('stopwords')
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem.porter import PorterStemmer
# Helper libraries
from w266_common import utils, vocabulary, tf_embed_viz
utils.require_package("wget") # for fetching dataset
from keras.models import Sequential
from keras.layers import GaussianNoise, LSTM, Bidirectional, Dropout, Dense, Embedding, MaxPool1D, GlobalMaxPool1D, Conv1D
from keras.optimizers import Adam
from pymagnitude import *
# -
# # INTRODUCTION
# ## Phenotype Classification of Electronic Health Records
#
# Electronic Health Record (EHR) data is a rapidly growing source of unstructured biomedical data. This data is extremely rich, often capturing a patient’s phenotype. In a clinical context, phenotype refers to the specific medical condition or disease of a patient. These records captures this data in higher detail compared to structured encodings such as the International Classification of Diseases (ICD) or National Drug Codes (NDC). Traditional methods for extracting phenotypes from this data typically relies on manual review or processing the data through rule-based expert systems. Both approaches are time intensive, rely heavily on human expertise, and scale poorly with increased volume. This project proposes an automated approach to identifying phenotypes in EHR data through word vector clustering and machine learning. An automated approach would greatly reduce time and operation costs, with the potential of even outperforming industry standards.
#
# The data for this project is provided by nlplab, who have induced a biomedical corpus using word2vec. This corpus contains over 5 billion words pulled from biomedical scientific literature and Wikipedia.
#
# # DATA EXPLORATION
# ## Word Embedding
#
# The foundation of this project is based on word embedding models, an approach that converts words into number vectors based on co-occurence. These vectors help capture word meanings and context in a format suitable for machine learning.
#
# Typically these vectors are trained on extremely large corpora, which can take a lot of time and resources. Thankfully, the word embedding space is quite mature and there exists pre-trained models, ready to use out of the box. One such model is Standford's GloVe vectors, which is trained on a corpus of 6B tokens from Wikipedia and Gigaword. These vectors are available at https://nlp.stanford.edu/projects/glove/. We will go through some exercises to explore word vectors.
#
import glove_helper; reload(glove_helper)
hands = glove_helper.Hands(ndim=100)
# +
def find_nn_cos(v, Wv, k=10):
"""Find nearest neighbors of a given word, by cosine similarity.
Returns two parallel lists: indices of nearest neighbors, and
their cosine similarities. Both lists are in descending order,
and inclusive: so nns[0] should be the index of the input word,
nns[1] should be the index of the first nearest neighbor, and so on.
Args:
v: (d-dimensional vector) word vector of interest
Wv: (V x d matrix) word embeddings
k: (int) number of neighbors to return
Returns (nns, ds), where:
nns: (k-dimensional vector of int), row indices of nearest neighbors,
which may include the given word.
similarities: (k-dimensional vector of float), cosine similarity of each
neighbor in nns.
"""
v_norm = np.linalg.norm(v)
Wv_norm = np.linalg.norm(Wv, axis=1)
dot = np.dot(v, Wv.T)
cos_sim = dot / (v_norm * Wv_norm)
nns = np.flipud(np.argsort(cos_sim)[-k:])
ds = np.flipud(np.sort(cos_sim)[-k:])
return [nns, ds]
def show_nns(hands, word, k=10):
"""Helper function to print neighbors of a given word."""
word = word.lower()
print("Nearest neighbors for '{:s}'".format(word))
v = hands.get_vector(word)
for i, sim in zip(*find_nn_cos(v, hands.W, k)):
target_word = hands.vocab.id_to_word[i]
print("{:.03f} : '{:s}'".format(sim, target_word))
print("")
# -
show_nns(hands, "diabetes")
show_nns(hands, "cancer")
show_nns(hands, "depression")
# The results we see make sense and showcase the capability of word embeddings. However, we do run into a few issues. For one,
# loading the file into our workspace requires careful memory management. This can become a problem when dealing with larger models or when we want to tweak our models and reload the data. Another issue is that we have to build our own help functions for performing calculations on the word vectors. Not inherently an issue, but these calculations are fairly standard and it is always a good idea to work smarter, not harder.
#
# As an alternative, we can look at third-party packages that offer fast and simple support for word vector operations. The package we will use for this project is Magnitude (https://github.com/plasticityai/magnitude). This package offers "lazy-loading for faster cold starts in development, LRU memory caching for performance in production, multiple key queries, direct featurization to the inputs for a neural network, performant similiarity calculations, and other nice to have features for edge cases like handling out-of-vocabulary keys or misspelled keys and concatenating multiple vector models together." These are all great features that we can leverage for this project.
# ## Working with Word Vectors - Magnitude
#
# Going through a few simple comparisons and exercises, we can see the difference between working with the raw text file versus working with the magnitude file:
# - The zip file is ~4 times larger than the magnitude file. This is even more impressive consdering the text file still needs to be unpackaged.
# - Load times are extremely quick for the magnitude file, far outperforming the standard file.
# - Querying from the standard file outperforms the magnitude file, but querying from the magnitude file is simpler and offers additional functionality.
#
# While the increased query times is not ideal, especially when it comes to training, the portability and the increased functionality just makes life so much easier.
# +
print('Standard Text File:')
print('\tFile Size: ', os.stat('data/glove/glove.6B.zip').st_size)
start = time.time()
glove_vectors_txt = glove_helper.Hands(ndim=100, quiet=True)
end = time.time()
print('\tFile Load Time: ', end - start)
start = time.time()
glove_vectors_txt.get_vector('diabetes')
glove_vectors_txt.get_vector('cancer')
glove_vectors_txt.get_vector('hypertension')
end = time.time()
print('\tQuery Time: ', end - start)
print('\tHandling out-of-vocabulary words:')
try:
print('\t\t', glove_vectors_txt.get_vector('wordnotfoundinvocab'))
except AssertionError:
print('\t\tWord not found in vocabulary')
print('\nMagnitude File:')
print('\tFile Size: ', os.stat('data/glove-lemmatized.6B.100d.magnitude').st_size)
start = time.time()
glove_vectors_mag = Magnitude("data/glove-lemmatized.6B.100d.magnitude")
end = time.time()
print('\tFile Load Time: ', end - start)
start = time.time()
glove_vectors_mag.query("diabetes")
glove_vectors_mag.query("cancer")
glove_vectors_mag.query("hypertension")
end = time.time()
print('\tQuery Time: ', end - start)
print('\tHandling out-of-vocabulary words:')
try:
print('\t\t', glove_vectors_mag.query('wordnotfoundinvocab'))
except AssertionError:
print('\t\tWord not found in vocabulary')
# -
# ## Corpus Selection - Biomedical Text
#
# -- Talk about importance of base corpora
# -- Reference paper that compares medical coprora to general corpora
# -- Show case actual examples by showing NN of GloVe vs medical
#
# With a framework that allows more freedom in corpus selection, we can move into much more larger word embedding models. The GloVe model we have been previously working with is actually on the smaller side. Of course, a larger corpus offers more data to train on, thus better capturing word contexts and meanings. However, another determininig factor in corpus selection is the source of the text. In general, these pre-trained models are based on general topic sources such as Wikipedia and Gigaword. However, since we know the domain we are working in, it may make sense to pull from relevant text sources.
#
# A Comparison of Word Embeddings for the Biomedical Natural Language Processing (https://arxiv.org/pdf/1802.00400.pdf) explores this idea. The paper concluded that "word embeddings trained on EHR and MedLit can capture the semantics of medical terms better and find semantically relevant medical terms closer to human experts’ judgments than those trained on GloVe and Google News."
#
# We can test these results ourselves by comparing GloVe against a biomedical based word embedding that was trained on text from PubMed and PubMed Central.
# +
print('GloVe length: ', len(glove_vectors_mag))
print('GloVe dimensions: ', glove_vectors_mag.dim)
print('\nNearest Neighbor examples:')
print('10 NN for diabetes:\n', glove_vectors_mag.most_similar("diabetes", topn = 10))
print('10 NN for cancer:\n', glove_vectors_mag.most_similar("cancer", topn = 10))
print('10 NN for hyperlipidemia:\n', glove_vectors_mag.most_similar("hyperlipidemia", topn = 10))
print('10 NN for e119:\n', glove_vectors_mag.most_similar("e119", topn = 10))
# +
med_vectors = Magnitude("data/wikipedia-pubmed-and-PMC-w2v.magnitude", pad_to_length=30)
print('Medical length: ', len(med_vectors))
print('Medical dimensions: ', med_vectors.dim)
# print('\nNearest Neighbor examples:')
# print('10 NN for diabetes:\n', med_vectors.most_similar("diabetes", topn = 10))
# print('10 NN for cancer:\n', med_vectors.most_similar("cancer", topn = 10))
# print('10 NN for hyperlipidemia:\n', med_vectors.most_similar("hyperlipidemia", topn = 10))
# print('10 NN for e119:\n', med_vectors.most_similar("e119", topn = 10))
# -
# ## Training Data - Labeled Electronic Health Record Text
#
# -- Refer back to goal of project
# -- Talk about difficulty of getting medical data (HIPPA)
# -- Reference MTsamples as source of data
# -- Show raw data unprocessed
# -- Briefly talk about transformations
#
# The goal of this project is to classify Eletronic Health Record (EHR) text. This of course means that we need to get our hands on some EHR data. This can be particularly difficult due to the strict rules and guidelines around healthcare data. The Health Insurance Portability and Accountability Act of 1996, or HIPAA, outlines a set of rules that help protect the privacy of our health information. These rules are vital for building a healthcare system where we can trust our healthcare providers and caregivers, so it is important that we adhere to the standards set by HIPAA.
#
# For this project, we will be using a dataset provided by MTSamples.com. They provide ~5,000 transcribed medical reports covering 40 specialty types. All of the notes have been de-identified of protected health information, making them HIPAA compliant. Below we will explore a few rows of the raw data.
# +
ehr_notes = []
with open('data/ehr_samples.csv', newline='') as csvfile:
reader = csv.DictReader(csvfile)
for row in reader:
ehr_notes.append([row['Specialty'], row['Note']])
print('EHR Sentence Example:\n')
print(ehr_notes[0])
print(ehr_notes[1])
# -
# ## Text Processing - Pre-Processing the EHR Notes
#
# -- Talk about how we need to manage our scope. Mention how ML on larger text scales
# -- For simplicity, going to limit ourselves to sentences. Possibly moving on to more text if we see promising results
# -- Talk about the standard set of NLTK functions
# -- Show a new sentence
#
# With the EHR data now loaded, we could technically start applying Machine Learning operations as is. However, as with a lot of text-based data, there are a few characteristics that are less than ideal for this project. The first obstacle is managing our text length. As our input text grows, so does the number of variables and the number of operations. Depending on our algorithm, these values can scale exponentially, causing runtime and resource usage to explode out of hand. To help manage the scope of our input text, we will be breaking up our notes into sentences. This should give us enough context to learn the more complex relationships between our words while minimizing runtime. Of course, if we find that runtime performance is not an issue, we can try further expanding our input text.
#
# Another pre-processing step we can take is to apply basic natural language cleanup techniques that standardize the text and remove non-essential information. Thankfully, python has a package called the Natural Language Toolkit (NLTK) that provides a lot of these transformations as built-in functions. The operations we will use for this project are converting all text to lowercase, removing punctation, filtering out stop words, and removing blanks.
#
# After all of the pre-processing, we can take a look at what the EHR notes now look like.
# +
ehr_sentences = []
for record in ehr_notes:
sent_text = nltk.sent_tokenize(record[1])
for sent in sent_text:
tokens = word_tokenize(sent)
# convert to lower case
tokens = [w.lower() for w in tokens]
# remove punctuation from each word
table = str.maketrans('', '', string.punctuation)
tokens = [w.translate(table) for w in tokens]
# filter out stop words
stop_words = set(stopwords.words('english'))
tokens = [w for w in tokens if not w in stop_words]
# # stem words
# porter = PorterStemmer()
# tokens = [porter.stem(word) for word in tokens]
# remove blanks
tokens = [w for w in tokens if w != '']
ehr_sentences.append([record[0], ' '.join(tokens)])
random.Random(4).shuffle(ehr_sentences)
# +
print(ehr_sentences[:10])
specialties = ['Allergy', 'Autopsy', 'Bariatrics', 'Cardiovascular', 'Chart', 'Chiropractic', 'Consult'
, 'Cosmetic', 'Dentistry', 'Dermatology', 'Diet', 'Discharge', 'Emergency', 'Endocrinology'
, 'Gastroenterology', 'General', 'Gynecology', 'Hospice', 'IME', 'Letters', 'Nephrology', 'Neurology'
, 'Neurosurgery', 'Office Notes', 'Oncology', 'Ophthalmology', 'Orthopedic', 'Otolaryngology'
, 'Pain Management', 'Pathology', 'Pediatrics', 'Podiatry', 'Psychiatry', 'Radiology', 'Rehab'
, 'Rheumatology', 'Sleep', 'Speech', 'Surgery', 'Urology']
# -
# # METHODS AND APPROACHES
# ## Naive Nearest Neighbor
#
# -- Talk about distance vs similarity
# -- Talk about fundametal co-occurence principle of word to vector
# -- How those vectors repsenet context or meaning
# -- If a sentence is more similar to our category, we can simply label it as such
# -- SHow some good examples but emphasize the bad examples
#
#
# The first method we will explore will be to just leverage the word embedding space with no Machine Learning at all. We mentioned earlier that the word vectors capture context and meaning. Additionally position of these vectors in relation to eachother also convey word relationships. At the core of it, vectors clustered together are more similar in context and meaning. Using this principle, we can use our categories as anchors in our word embedding, calculate a similarity score for a sentence, and identify which category is the nearest neighbor to our sentence.
#
# This is a very naive approach but it will be a good exercise and can at least set a baseline for performance.
# +
print('Similarity between diabetes and mellitus: ', med_vectors.similarity("diabetes", "mellitus"))
print('Similarity between diabetes and breast: ', med_vectors.similarity("diabetes", "breast"))
print('\nSimilarity between cancer and mellitus: ', med_vectors.similarity("cancer", "mellitus"))
print('Similarity between cancer and breast: ', med_vectors.similarity("cancer", "breast"))
# +
nn_results = []
for i, ehr_sent in enumerate(ehr_sentences[0:2000]):
# print(ehr_sent)
most_similar_specialty = []
for specialty in specialties:
spec_similarity_sum = 0
for token in ehr_sent[1].split(' '):
# print('\t', token, med_vectors.similarity(specialty, token))
spec_similarity_sum += med_vectors.similarity(specialty, token)
spec_similarity = spec_similarity_sum / len(ehr_sent[1].split(' '))
# print(specialty, spec_similarity)
if not most_similar_specialty:
most_similar_specialty = [i, ehr_sent[0], specialty, spec_similarity]
elif spec_similarity > most_similar_specialty[3]:
most_similar_specialty = [i, ehr_sent[0], specialty, spec_similarity]
nn_results.append(most_similar_specialty)
correct_results = [result for result in nn_results if result[1] == result[2]]
print('# of Correct Classifications: ', len(correct_results))
print('Accuracy: ', len(correct_results) / len(nn_results))
# +
print('Example of correct classification:')
correct_example = correct_results[0]
example_sentence = ehr_sentences[correct_example[0]]
print('\tSentence: ', example_sentence)
print('\n\tTrue category:', correct_example[1])
print('\tPredicted category:', correct_example[2])
print('\n\tTrue/Predicted Similarities:')
for token in example_sentence[1].split(' '):
print('\t\t', token, med_vectors.similarity(correct_example[1], token))
spec_similarity_sum += med_vectors.similarity(correct_example[1], token)
spec_similarity = spec_similarity_sum / len(example_sentence.split(' '))
print('\t\tAverage similarity: ', spec_similarity)
# +
print('Example of incorrect classification:')
incorrect_example = nn_results[0]
example_sentence = ehr_sentences[incorrect_example[0]]
print('\tSentence: ', example_sentence)
print('\n\tTrue category:', incorrect_example[1])
print('\tPredicted category:', incorrect_example[2])
print('\n\tTrue Similarities:')
for token in example_sentence[1].split(' '):
print('\t\t', token, med_vectors.similarity(incorrect_example[1], token))
spec_similarity_sum += med_vectors.similarity(incorrect_example[1], token)
spec_similarity = spec_similarity_sum / len(example_sentence[1].split(' '))
print('\t\tAverage similarity: ', spec_similarity)
print('\n\tPredicted Similarities:')
for token in example_sentence[1].split(' '):
print('\t\t', token, med_vectors.similarity(incorrect_example[2], token))
spec_similarity_sum += med_vectors.similarity(incorrect_example[2], token)
spec_similarity = spec_similarity_sum / len(example_sentence[1].split(' '))
print('\t\tAverage similarity: ', spec_similarity)
# -
# So as we can see, the results are pretty terrible with an accuracy of 5%. Looking at an example the classifier got right, it relied on words that are exclusively and very distinctly related. However, these strong signals are not always present in our sentences. Looking at an incorrect example, we see how the signals are being drowned out or offset by the other words. This emphasizes the need for some type of model that can learn and weigh the words that provide strong signals for particular categories.
# # METHODS AND APPROACHES
# ## Neural Networks
#
# A neural network will allow us to build a model that can take in the word vectors as inputs and learn the complex relationships between those vectors to better classify the target sentence. This is a more holistic approach that tries to capture meaning from the entire sentence rather than token by token.
# ## Defining our Training and Test Data
#
# Before we can start building our neural networks, we first have to define our datasets. Specifically, we have to break up our EHR data so that we have records that we can train on and records that are exclusively used to test on. Maintaining a separate set for testing ensures we avoid overfitting our data.
#
# We will use some built-in functions provided by Magnitude that helps encode our classes/categories. We then partition our data into our train and test sets. For each set we have both data and labels. Initially, we will be making these partitions small to make iterating through model development much quicker. However, once the models are developed, we will expand our datasets to include all of our data. To ensure we defined our data correctly, we can print a few lines from the two sets.
# +
add_intent, intent_to_int, int_to_intent = MagnitudeUtils.class_encoding()
x_train = [ehr_sent[1].split(' ') for ehr_sent in ehr_sentences[:130000]]
x_test = [ehr_sent[1].split(' ') for ehr_sent in ehr_sentences[130001:]]
y_train = [add_intent(ehr_sent[0]) for ehr_sent in ehr_sentences[:130000]]
y_test = [add_intent(ehr_sent[0]) for ehr_sent in ehr_sentences[130001:]]
y_train = list(np.array(y_train).reshape(len(y_train)))
y_test = list(np.array(y_test).reshape(len(y_test)))
num_training = len(x_train)
num_test = len(x_test)
num_outputs = int(max(max(y_train), max(y_test))) + 1
print(int_to_intent(0))
print("First line of train/test data:")
print("\t", x_train[0])
print("\t", y_train[0], int_to_intent(y_train[0]))
print("\t", x_test[0])
print("\t", y_test[0], int_to_intent(y_test[0]))
print("Second line of train/test data:")
print("\t", x_train[1])
print("\t", y_train[1], int_to_intent(y_train[1]))
print("\t", x_test[1])
print("\t", y_test[1], int_to_intent(y_test[1]))
# -
# ## Convultional Neural Network
# -- Explain conv layers, focusing on 1d
# -- how it learns the best filters
# -- talk about exact model structure
# +
MAX_WORDS = 30 # The maximum number of words the sequence model will consider
STD_DEV = 0.01 # Deviation of noise for Gaussian Noise applied to the embeddings
DROPOUT_RATIO = .5 # The ratio to dropout
BATCH_SIZE = 100 # The number of examples per train/validation step
EPOCHS = 100 # The number of times to repeat through all of the training data
LEARNING_RATE = .01 # The learning rate for the optimizer
NUM_FILTERS = 128
model = Sequential()
model.add(GaussianNoise(STD_DEV, input_shape=(MAX_WORDS, med_vectors.dim)))
model.add(Conv1D(NUM_FILTERS, 7, activation='relu', padding='same'))
model.add(MaxPool1D(2))
model.add(Conv1D(NUM_FILTERS, 7, activation='relu', padding='same'))
model.add(GlobalMaxPool1D())
model.add(Dropout(DROPOUT_RATIO))
model.add(Dense(64, activation='relu'))
model.add(Dense(num_outputs, activation='softmax'))
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['categorical_accuracy'])
model.summary()
# +
training_batches = MagnitudeUtils.batchify(x_train, y_train, BATCH_SIZE) # Split the training data into batches
num_batches_per_epoch_train = int(np.ceil(num_training/float(BATCH_SIZE)))
test_batches = MagnitudeUtils.batchify(x_test, y_test, BATCH_SIZE) # Split the test data into batches
num_batches_per_epoch_test = int(np.ceil(num_test/float(BATCH_SIZE)))
# Generates batches of the transformed training data
train_batch_generator = (
(
med_vectors.query(x_train_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_train_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_train_batch, y_train_batch in training_batches
)
# Generates batches of the transformed test data
test_batch_generator = (
(
med_vectors.query(x_test_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_test_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_test_batch, y_test_batch in test_batches
)
# Start training
from keras.utils import np_utils
model.fit_generator(
generator = train_batch_generator,
steps_per_epoch = num_batches_per_epoch_train,
validation_data = test_batch_generator,
validation_steps = num_batches_per_epoch_test,
epochs = EPOCHS,
)
# +
print("Results after training for %d epochs:" % (EPOCHS,))
train_metrics = model.evaluate_generator(
generator = train_batch_generator,
steps = num_batches_per_epoch_train,
)
print("loss: %.4f - categorical_accuracy: %.4f" % tuple(train_metrics))
val_metrics = model.evaluate_generator(
generator = test_batch_generator,
steps = num_batches_per_epoch_test,
)
print("val_loss: %.4f - val_categorical_accuracy: %.4f" % tuple(val_metrics))
# -
len(ehr_sentences)
# ## LSTM Neural Network
# -- talk about LSTM vs conv
# -- advantages
# -- talk about exact model
# +
MAX_WORDS = 30 # The maximum number of words the sequence model will consider
STD_DEV = 0.01 # Deviation of noise for Gaussian Noise applied to the embeddings
HIDDEN_UNITS = 100 # The number of hidden units from the LSTM
DROPOUT_RATIO = .8 # The ratio to dropout
BATCH_SIZE = 100 # The number of examples per train/validation step
EPOCHS = 100 # The number of times to repeat through all of the training data
LEARNING_RATE = .01 # The learning rate for the optimizer
model = Sequential()
model.add(GaussianNoise(STD_DEV, input_shape=(MAX_WORDS, med_vectors.dim)))
model.add(Bidirectional(LSTM(HIDDEN_UNITS, activation='tanh'), merge_mode='concat'))
model.add(Dropout(DROPOUT_RATIO))
model.add(Dense(num_outputs, activation='softmax'))
model.compile(
loss='categorical_crossentropy',
optimizer=Adam(lr=LEARNING_RATE),
metrics=['categorical_accuracy'])
model.summary()
# +
training_batches = MagnitudeUtils.batchify(x_train, y_train, BATCH_SIZE) # Split the training data into batches
num_batches_per_epoch_train = int(np.ceil(num_training/float(BATCH_SIZE)))
test_batches = MagnitudeUtils.batchify(x_test, y_test, BATCH_SIZE) # Split the test data into batches
num_batches_per_epoch_test = int(np.ceil(num_test/float(BATCH_SIZE)))
# Generates batches of the transformed training data
train_batch_generator = (
(
med_vectors.query(x_train_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_train_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_train_batch, y_train_batch in training_batches
)
# Generates batches of the transformed test data
test_batch_generator = (
(
med_vectors.query(x_test_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_test_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_test_batch, y_test_batch in test_batches
)
# Start training
from keras.utils import np_utils
model.fit_generator(
generator = train_batch_generator,
steps_per_epoch = num_batches_per_epoch_train,
validation_data = test_batch_generator,
validation_steps = num_batches_per_epoch_test,
epochs = EPOCHS,
)
# +
print("Results after training for %d epochs:" % (EPOCHS,))
train_metrics = model.evaluate_generator(
generator = train_batch_generator,
steps = num_batches_per_epoch_train,
)
print("loss: %.4f - categorical_accuracy: %.4f" % tuple(train_metrics))
val_metrics = model.evaluate_generator(
generator = test_batch_generator,
steps = num_batches_per_epoch_test,
)
print("val_loss: %.4f - val_categorical_accuracy: %.4f" % tuple(val_metrics))
# +
MAX_WORDS = 30 # The maximum number of words the sequence model will consider
STD_DEV = 0.01 # Deviation of noise for Gaussian Noise applied to the embeddings
HIDDEN_UNITS = 50 # The number of hidden units from the LSTM
DROPOUT_RATIO = .8 # The ratio to dropout
BATCH_SIZE = 100 # The number of examples per train/validation step
EPOCHS = 200 # The number of times to repeat through all of the training data
LEARNING_RATE = .001 # The learning rate for the optimizer
model = Sequential()
model.add(GaussianNoise(STD_DEV, input_shape=(MAX_WORDS, med_vectors.dim)))
model.add(Bidirectional(LSTM(HIDDEN_UNITS, activation='tanh'), merge_mode='concat'))
model.add(Dropout(DROPOUT_RATIO))
model.add(Dense(num_outputs, activation='softmax'))
model.compile(
loss='categorical_crossentropy',
optimizer=Adam(lr=LEARNING_RATE),
metrics=['categorical_accuracy'])
# +
training_batches = MagnitudeUtils.batchify(x_train, y_train, BATCH_SIZE) # Split the training data into batches
num_batches_per_epoch_train = int(np.ceil(num_training/float(BATCH_SIZE)))
test_batches = MagnitudeUtils.batchify(x_test, y_test, BATCH_SIZE) # Split the test data into batches
num_batches_per_epoch_test = int(np.ceil(num_test/float(BATCH_SIZE)))
# Generates batches of the transformed training data
train_batch_generator = (
(
med_vectors.query(x_train_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_train_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_train_batch, y_train_batch in training_batches
)
# Generates batches of the transformed test data
test_batch_generator = (
(
med_vectors.query(x_test_batch), # Magnitude will handle converting the 2D array of text into the 3D word vector representations!
MagnitudeUtils.to_categorical(y_test_batch, num_outputs) # Magnitude will handle converting the class labels into one-hot encodings!
) for x_test_batch, y_test_batch in test_batches
)
# Start training
from keras.utils import np_utils
model.fit_generator(
generator = train_batch_generator,
steps_per_epoch = num_batches_per_epoch_train,
validation_data = test_batch_generator,
validation_steps = num_batches_per_epoch_test,
epochs = EPOCHS,
)
# +
print("Results after training for %d epochs:" % (EPOCHS,))
train_metrics = model.evaluate_generator(
generator = train_batch_generator,
steps = num_batches_per_epoch_train,
)
print("loss: %.4f - categorical_accuracy: %.4f" % tuple(train_metrics))
val_metrics = model.evaluate_generator(
generator = test_batch_generator,
steps = num_batches_per_epoch_test,
)
print("val_loss: %.4f - val_categorical_accuracy: %.4f" % tuple(val_metrics))
# -
print(int_to_intent(MagnitudeUtils.from_categorical(model.predict(med_vectors.query(["past medical history difficulty climbing stairs difficulty airline seats tying shoes used public seating lifting objects floor".split(" ")])))[0]))
with open('data/ehr_sentences.csv', 'w') as outfile:
writer = csv.writer(outfile, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['Specialty', 'Note'])
for sent in ehr_sentences:
writer.writerow(sent)
# +
ehr_labels = []
ehr_vectors = []
for sentence in ehr_sentences:
ehr_labels.append(sentence[0])
sentence_split = sentence[1].split(' ')
ehr_vectors.append(med_vectors.query(sentence_split))
# +
with open('data/ehr_labels.csv', 'w') as outfile:
writer = csv.writer(outfile, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['SpecialtyID'])
for lbl in ehr_labels:
writer.writerow(lbl)
with open('data/ehr_vectors.csv', 'w') as outfile:
writer = csv.writer(outfile, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
writer.writerow(['NoteVector'])
for vctr in ehr_vectors:
writer.writerow(vctr)
| archive/EmbeddingsExploration-CategoricalConv-Completed-4.10.19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### We use function spect_loader extracted from
# (https://github.com/adiyoss/GCommandsPytorch/blob/master/gcommand_loader.py)
# #### to calculate the spectrogram of the audio file
#
# +
import numpy as np
from keras import backend as K
from keras.preprocessing.image import Iterator
from keras.preprocessing.image import img_to_array
import librosa
import os
import multiprocessing.pool
from functools import partial
from random import getrandbits
train_path='C:/Users/Brooks/Desktop/part1/train'
val_path ='C:/Users/Brooks/Desktop/part1/val'
#to count how many files inside each class
classnames=os.listdir(train_path)
train_count_dict = {}
for d in classnames:
train_count_dict[d] = len(os.listdir(os.path.join(train_path, d)))
print('train freq')
for k, v in train_count_dict.items():
print ( '%7s %i' % (k, v))
val_count_dict = {}
for d in classnames:
val_count_dict[d] = len(os.listdir(os.path.join(val_path, d)))
print('\nval freq')
for k, v in val_count_dict.items():
print ( '%7s %i' % (k, v))
print ('')
#print ('test files', len(os.listdir(test_path+'/audio')))
# +
#Convert the sound to melspectrogram#
#We use function spect_loader extracted from
#(https://github.com/adiyoss/GCommandsPytorch/blob/master/gcommand_loader.py)
#to calculate the spectrogram of the audio file
#However we decide to use melspectrogram instead of spectrogram since some research
#found that while using MFCC, the performance will better;
#So we use a similar chart to MFCC which is mel-spectrogram
def spect_loader(path, window_size, window_stride, window, normalize, max_len=101,
augment=False, allow_speedandpitch=False, allow_pitch=False,
allow_speed=False, allow_dyn=False, allow_noise=False,
allow_timeshift=False ):
y, sr = librosa.load(path, sr=None)
#OriginalSampelRate=16Khz;
# n_fft = 4096
n_fft = int(sr * window_size)
win_length = n_fft
hop_length = int(sr * window_stride)
# Let's make and display a mel-scaled power (energy-squared) spectrogram
S = librosa.feature.melspectrogram(y, sr=sr, n_fft = 4096,hop_length=hop_length,n_mels=128)
# Convert to log scale (dB). We'll use the peak power as reference.
log_S = librosa.core.amplitude_to_db(S, ref=np.max)
#made a melspectrogram
spect=log_S
# make all spects with the same dimentions
#In case of the mel-spectrogram size different since the lenght of the sound variate
# TODO: change that in the future
if spect.shape[1] < max_len:
pad = np.zeros((spect.shape[0], max_len - spect.shape[1]))
spect = np.hstack((spect, pad))
elif spect.shape[1] > max_len:
spect = spect[:max_len, ]
spect = np.resize(spect, (1, spect.shape[0], spect.shape[1]))
#spect = torch.FloatTensor(spect)
# z-score normalization
#calculate the mean and standard of the melspect
if normalize:
mean = np.mean(np.ravel(spect))
std = np.std(np.ravel(spect))
if std != 0:
spect = spect -mean
spect = spect / std
return spect
def _count_valid_files_in_directory(directory, white_list_formats, follow_links):
"""Count files with extension in `white_list_formats` contained in a directory.
# Arguments
directory: absolute path to the directory containing files to be counted
white_list_formats: set of strings containing allowed extensions for
the files to be counted.
# Returns
the count of files with extension in `white_list_formats` contained in
the directory.
"""
def _recursive_list(subpath):
return sorted(os.walk(subpath, followlinks=follow_links), key=lambda tpl: tpl[0])
samples = 0
for root, _, files in _recursive_list(directory):
for fname in files:
is_valid = False
for extension in white_list_formats:
if fname.lower().endswith('.' + extension):
is_valid = True
break
if is_valid:
samples += 1
return samples
def _list_valid_filenames_in_directory(directory, white_list_formats,
class_indices, follow_links):
"""List paths of files in `subdir` relative from `directory` whose extensions are in `white_list_formats`.
# Arguments
directory: absolute path to a directory containing the files to list.
The directory name is used as class label and must be a key of `class_indices`.
white_list_formats: set of strings containing allowed extensions for
the files to be counted.
class_indices: dictionary mapping a class name to its index.
# Returns
classes: a list of class indices
filenames: the path of valid files in `directory`, relative from
`directory`'s parent (e.g., if `directory` is "dataset/class1",
the filenames will be ["class1/file1.jpg", "class1/file2.jpg", ...]).
"""
def _recursive_list(subpath):
return sorted(os.walk(subpath, followlinks=follow_links), key=lambda tpl: tpl[0])
classes = []
filenames = []
subdir = os.path.basename(directory)
basedir = os.path.dirname(directory)
for root, _, files in _recursive_list(directory):
for fname in sorted(files):
is_valid = False
for extension in white_list_formats:
if fname.lower().endswith('.' + extension):
is_valid = True
break
if is_valid:
classes.append(class_indices[subdir])
# add filename relative to directory
absolute_path = os.path.join(root, fname)
filenames.append(os.path.relpath(absolute_path, basedir))
return classes, filenames
#window_size=.02
#window_stride=.01
#window_type='hamming'
#normalize=True
#max_len=101
#batch_size = 64
class SpeechDirectoryIterator(Iterator):
"""Iterator capable of reading images from a directory on disk.
# Arguments
"""
def __init__(self, directory, window_size, window_stride,
window_type, normalize, max_len=101,
target_size=(256, 256), color_mode='grayscale',
classes=None, class_mode='categorical',
batch_size=32, shuffle=True, seed=None,
data_format=None, save_to_dir=None,
save_prefix='', save_format='png',
follow_links=False, interpolation='nearest', augment=False,
allow_speedandpitch = False, allow_pitch = False,
allow_speed = False, allow_dyn = False, allow_noise = False, allow_timeshift=False ):
if data_format is None:
data_format = K.image_data_format()
self.window_size = window_size
self.window_stride = window_stride
self.window_type = window_type
self.normalize = normalize
self.max_len = max_len
self.directory = directory
self.allow_speedandpitch = allow_speedandpitch
self.allow_pitch = allow_pitch
self.allow_speed = allow_speed
self.allow_dyn = allow_dyn
self.allow_noise = allow_noise
self.allow_timeshift = allow_timeshift
self.augment = augment
# self.image_data_generator = image_data_generator
self.target_size = tuple(target_size)
if color_mode not in {'rgb', 'grayscale'}:
raise ValueError('Invalid color mode:', color_mode,
'; expected "rgb" or "grayscale".')
self.color_mode = color_mode
self.data_format = data_format
if self.color_mode == 'rgb':
#becasue it is rgb, so the hight of the data is 3
if self.data_format == 'channels_last':
self.image_shape = self.target_size + (3,)
else:
self.image_shape = (3,) + self.target_size
else:
#if it's gray scale, the picture's hight is 1
if self.data_format == 'channels_last':
self.image_shape = self.target_size + (1,)
else:
self.image_shape = (1,) + self.target_size
self.classes = classes
if class_mode not in {'categorical', 'binary', 'sparse',
'input', None}:
raise ValueError('Invalid class_mode:', class_mode,
'; expected one of "categorical", '
'"binary", "sparse", "input"'
' or None.')
self.class_mode = class_mode
self.save_to_dir = save_to_dir
self.save_prefix = save_prefix
self.save_format = save_format
self.interpolation = interpolation
white_list_formats = {'png', 'jpg', 'jpeg', 'bmp', 'ppm', 'wav'}
# first, count the number of samples and classes
self.samples = 0
if not classes:
classes = []
for subdir in sorted(os.listdir(directory)):
if os.path.isdir(os.path.join(directory, subdir)):
classes.append(subdir)
self.num_classes = len(classes)
self.class_indices = dict(zip(classes, range(len(classes))))
pool = multiprocessing.pool.ThreadPool()
function_partial = partial(_count_valid_files_in_directory,
white_list_formats=white_list_formats,
follow_links=follow_links)
self.samples = sum(pool.map(function_partial,
(os.path.join(directory, subdir)
for subdir in classes)))
print('Found %d images belonging to %d classes.' % (self.samples, self.num_classes))
# second, build an index of the images in the different class subfolders
results = []
self.filenames = []
self.classes = np.zeros((self.samples,), dtype='int32')
i = 0
for dirpath in (os.path.join(directory, subdir) for subdir in classes):
results.append(pool.apply_async(_list_valid_filenames_in_directory,
(dirpath, white_list_formats,
self.class_indices, follow_links)))
for res in results:
classes, filenames = res.get()
self.classes[i:i + len(classes)] = classes
self.filenames += filenames
if i==0:
#Use the specloader to trans the .wav file to image
img = spect_loader(os.path.join(self.directory, filenames[0]),
self.window_size,
self.window_stride,
self.window_type,
self.normalize,
self.max_len,
self.augment,
self.allow_speedandpitch,
self.allow_pitch,
self.allow_speed,
self.allow_dyn,
self.allow_noise,
self.allow_timeshift )
img=np.swapaxes(img, 0, 2)
self.target_size = tuple((img.shape[0], img.shape[1]))
print(self.target_size)
if self.color_mode == 'rgb':
if self.data_format == 'channels_last':
self.image_shape = self.target_size + (3,)
else:
self.image_shape = (3,) + self.target_size
else:
if self.data_format == 'channels_last':
self.image_shape = self.target_size + (1,)
else:
self.image_shape = (1,) + self.target_size
i += len(classes)
pool.close()
pool.join()
super(SpeechDirectoryIterator, self).__init__(self.samples, batch_size, shuffle, seed)
def _get_batches_of_transformed_samples(self, index_array):
batch_x = np.zeros((len(index_array),) + self.image_shape, dtype=K.floatx())
batch_f = []
grayscale = self.color_mode == 'grayscale'
# build batch of image data
#print(index_array)
for i, j in enumerate(index_array):
#print(i, j, self.filenames[j])
fname = self.filenames[j]
#img = load_img(os.path.join(self.directory, fname),
# grayscale=grayscale,
# target_size=self.target_size,
# interpolation=self.interpolation)
img = spect_loader(os.path.join(self.directory, fname),
self.window_size,
self.window_stride,
self.window_type,
self.normalize,
self.max_len,
)
img=np.swapaxes(img, 0, 2)
x = img_to_array(img, data_format=self.data_format)
#x = self.image_data_generator.random_transform(x)
#x = self.image_data_generator.standardize(x)
batch_x[i] = x
batch_f.append(fname)
# optionally save augmented images to disk for debugging purposes
if self.save_to_dir:
for i, j in enumerate(index_array):
img = array_to_img(batch_x[i], self.data_format, scale=True)
fname = '{prefix}_{index}_{hash}.{format}'.format(prefix=self.save_prefix,
index=j,
hash=np.random.randint(1e7),
format=self.save_format)
img.save(os.path.join(self.save_to_dir, fname))
# build batch of labels
if self.class_mode == 'input':
batch_y = batch_x.copy()
elif self.class_mode == 'sparse':
batch_y = self.classes[index_array]
elif self.class_mode == 'binary':
batch_y = self.classes[index_array].astype(K.floatx())
elif self.class_mode == 'categorical':
batch_y = np.zeros((len(batch_x), self.num_classes), dtype=K.floatx())
for i, label in enumerate(self.classes[index_array]):
batch_y[i, label] = 1.
else:
return batch_x
return batch_x, batch_y
def next(self):
with self.lock:
index_array = next(self.index_generator)[0]
# The transformation of images is not under thread lock
# so it can be done in parallel
return self._get_batches_of_transformed_samples(index_array)
# -
# window_size: this quantity times the sampling rate (sr) returns the fft window size
#
# window_stride: Each frame of audio is windowed by window_stride * sr
#
# window_type: The type of window function to be applied (hamming, hanning, etc)
#
# normalize: True / Fale
#
# max_len: Keep only max_len frequency components (first dimension of the output numpy array)
window_size=.02
window_stride=.01
window_type='hamming'
normalize=True
max_len=101
batch_size = 64
#put the train file to train_iterator
train_iterator = SpeechDirectoryIterator(directory=train_path,
batch_size=batch_size,
window_size=window_size,
window_stride=window_stride,
window_type=window_type,
normalize=normalize,
max_len=max_len)
#put the validation file to val_iterator
val_iterator = SpeechDirectoryIterator(directory=val_path,
batch_size=batch_size,
window_size=window_size,
window_stride=window_stride,
window_type=window_type,
normalize=normalize,
max_len=max_len)
# +
from keras.models import Sequential
from keras.layers import Conv2D, MaxPooling2D, Flatten, Dense, Dropout
#build the training model by keras
#Use model structure similar to the MNIST recognition
#And we have made some optimize of the parameters include the costfunc and optimizer ect.
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=train_iterator.image_shape))
# add a 2d Convolution with 3*3 kernel and 64 output.
model.add(Conv2D(64, (3, 3), activation='relu'))
#Add a maxpooling with 2*2
model.add(MaxPooling2D(pool_size=(2, 2)))
#with 0.25 margin to Dropout
model.add(Dropout(0.5))
#flat all the pixel
model.add(Flatten())
#use fully connected and the activation is relu
model.add(Dense(128, activation='relu'))
#with 0.5 margin drop out
model.add(Dropout(0.5))
#use the softmax as the activation function
model.add(Dense(len(classnames), activation = 'softmax')) #Last layer with one output per class
#Use crossEntropy as the loss function
#And set the optimizer as Adadelta
model.compile(loss='categorical_crossentropy', optimizer='Adadelta',
metrics=['accuracy'])
model.summary()
# +
from keras.callbacks import EarlyStopping, ReduceLROnPlateau
#Use this func to early stop when the val_loss do not lower
#Use this ReduceLROnPlateau to auto adapt the LearningRate
early = EarlyStopping(monitor='val_loss', min_delta=0, patience=5, verbose=1, mode='auto')
reduce = ReduceLROnPlateau(monitor='val_loss', factor=0.5, patience=3, verbose=1, mode='auto', min_lr=0.00001)
model.fit_generator(train_iterator,
steps_per_epoch=int(np.ceil(train_iterator.n / batch_size)),
epochs=20,
validation_data=val_iterator,
validation_steps=int(np.ceil(val_iterator.n / batch_size)),
verbose=1, callbacks=[early, reduce])
# -
# What will we do in the future:
# draw the acc-loss chart to show the progress of the training;
# continue to optimize the parameter of the network;
| Final_Project/TrainingModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Boosting a decision stump
#
# The goal of this notebook is to implement your own boosting module.
#
# **Brace yourselves**! This is going to be a fun and challenging assignment.
#
#
# * Use SFrames to do some feature engineering.
# * Modify the decision trees to incorporate weights.
# * Implement Adaboost ensembling.
# * Use your implementation of Adaboost to train a boosted decision stump ensemble.
# * Evaluate the effect of boosting (adding more decision stumps) on performance of the model.
# * Explore the robustness of Adaboost to overfitting.
#
# Let's get started!
# ## Fire up Turi Create
# Make sure you have the latest version of Turi Create
import turicreate as tc
import matplotlib.pyplot as plt
# %matplotlib inline
# # Getting the data ready
# We will be using the same [LendingClub](https://www.lendingclub.com/) dataset as in the previous assignment.
loans = tc.SFrame('lending-club-data.sframe/')
# ### Extracting the target and the feature columns
#
# We will now repeat some of the feature processing steps that we saw in the previous assignment:
#
# First, we re-assign the target to have +1 as a safe (good) loan, and -1 as a risky (bad) loan.
#
# Next, we select four categorical features:
# 1. grade of the loan
# 2. the length of the loan term
# 3. the home ownership status: own, mortgage, rent
# 4. number of years of employment.
features = ['grade', # grade of the loan
'term', # the term of the loan
'home_ownership', # home ownership status: own, mortgage or rent
'emp_length', # number of years of employment
]
loans['safe_loans'] = loans['bad_loans'].apply(lambda x : +1 if x==0 else -1)
loans.remove_column('bad_loans')
target = 'safe_loans'
loans = loans[features + [target]]
# ### Subsample dataset to make sure classes are balanced
# Just as we did in the previous assignment, we will undersample the larger class (safe loans) in order to balance out our dataset. This means we are throwing away many data points. We use `seed=1` so everyone gets the same results.
# +
safe_loans_raw = loans[loans[target] == 1]
risky_loans_raw = loans[loans[target] == -1]
# Undersample the safe loans.
percentage = len(risky_loans_raw)/float(len(safe_loans_raw))
risky_loans = risky_loans_raw
safe_loans = safe_loans_raw.sample(percentage, seed=1)
loans_data = risky_loans_raw.append(safe_loans)
print("Percentage of safe loans :", len(safe_loans) / float(len(loans_data)))
print("Percentage of risky loans :", len(risky_loans) / float(len(loans_data)))
print("Total number of loans in our new dataset :", len(loans_data))
# -
# **Note:** There are many approaches for dealing with imbalanced data, including some where we modify the learning algorithm. These approaches are beyond the scope of this course, but some of them are reviewed in this [paper](http://ieeexplore.ieee.org/xpl/login.jsp?tp=&arnumber=5128907&url=http%3A%2F%2Fieeexplore.ieee.org%2Fiel5%2F69%2F5173046%2F05128907.pdf%3Farnumber%3D5128907 ). For this assignment, we use the simplest possible approach, where we subsample the overly represented class to get a more balanced dataset. In general, and especially when the data is highly imbalanced, we recommend using more advanced methods.
# ### Transform categorical data into binary features
# In this assignment, we will work with **binary decision trees**. Since all of our features are currently categorical features, we want to turn them into binary features using 1-hot encoding.
#
# We can do so with the following code block (see the first assignments for more details):
loans_data = risky_loans.append(safe_loans)
for feature in features:
loans_data_one_hot_encoded = loans_data[feature].apply(lambda x: {x: 1})
loans_data_unpacked = loans_data_one_hot_encoded.unpack(column_name_prefix=feature)
# Change None's to 0's
for column in loans_data_unpacked.column_names():
loans_data_unpacked[column] = loans_data_unpacked[column].fillna(0)
loans_data = loans_data.remove_column(feature)
loans_data = loans_data.add_columns(loans_data_unpacked)
# Let's see what the feature columns look like now:
features = loans_data.column_names()
features.remove('safe_loans') # Remove the response variable
features
# ### Train-test split
#
# We split the data into training and test sets with 80% of the data in the training set and 20% of the data in the test set. We use `seed=1` so that everyone gets the same result.
train_data, test_data = loans_data.random_split(0.8, seed=1)
# # Weighted decision trees
# Let's modify our decision tree code from Module 5 to support weighting of individual data points.
# ### Weighted error definition
#
# Consider a model with $N$ data points with:
# * Predictions $\hat{y}_1 ... \hat{y}_n$
# * Target $y_1 ... y_n$
# * Data point weights $\alpha_1 ... \alpha_n$.
#
# Then the **weighted error** is defined by:
# $$
# \mathrm{E}(\mathbf{\alpha}, \mathbf{\hat{y}}) = \frac{\sum_{i=1}^{n} \alpha_i \times 1[y_i \neq \hat{y_i}]}{\sum_{i=1}^{n} \alpha_i}
# $$
# where $1[y_i \neq \hat{y_i}]$ is an indicator function that is set to $1$ if $y_i \neq \hat{y_i}$.
#
#
# ### Write a function to compute weight of mistakes
#
# Write a function that calculates the weight of mistakes for making the "weighted-majority" predictions for a dataset. The function accepts two inputs:
# * `labels_in_node`: Targets $y_1 ... y_n$
# * `data_weights`: Data point weights $\alpha_1 ... \alpha_n$
#
# We are interested in computing the (total) weight of mistakes, i.e.
# $$
# \mathrm{WM}(\mathbf{\alpha}, \mathbf{\hat{y}}) = \sum_{i=1}^{n} \alpha_i \times 1[y_i \neq \hat{y_i}].
# $$
# This quantity is analogous to the number of mistakes, except that each mistake now carries different weight. It is related to the weighted error in the following way:
# $$
# \mathrm{E}(\mathbf{\alpha}, \mathbf{\hat{y}}) = \frac{\mathrm{WM}(\mathbf{\alpha}, \mathbf{\hat{y}})}{\sum_{i=1}^{n} \alpha_i}
# $$
#
# The function **intermediate_node_weighted_mistakes** should first compute two weights:
# * $\mathrm{WM}_{-1}$: weight of mistakes when all predictions are $\hat{y}_i = -1$ i.e $\mathrm{WM}(\mathbf{\alpha}, \mathbf{-1}$)
# * $\mathrm{WM}_{+1}$: weight of mistakes when all predictions are $\hat{y}_i = +1$ i.e $\mbox{WM}(\mathbf{\alpha}, \mathbf{+1}$)
#
# where $\mathbf{-1}$ and $\mathbf{+1}$ are vectors where all values are -1 and +1 respectively.
#
# After computing $\mathrm{WM}_{-1}$ and $\mathrm{WM}_{+1}$, the function **intermediate_node_weighted_mistakes** should return the lower of the two weights of mistakes, along with the class associated with that weight. We have provided a skeleton for you with `YOUR CODE HERE` to be filled in several places.
def intermediate_node_weighted_mistakes(labels_in_node, data_weights):
# Sum the weights of all entries with label +1
total_weight_positive = sum(data_weights[labels_in_node == 1])
# Weight of mistakes for predicting all -1's is equal to the sum above
### YOUR CODE HERE
weighted_mistakes_all_negative = total_weight_positive
# Sum the weights of all entries with label -1
### YOUR CODE HERE
total_weight_negative = sum(data_weights[labels_in_node == -1])
# Weight of mistakes for predicting all +1's is equal to the sum above
### YOUR CODE HERE
weighted_mistakes_all_positive = total_weight_negative
# Return the tuple (weight, class_label) representing the lower of the two weights
# class_label should be an integer of value +1 or -1.
# If the two weights are identical, return (weighted_mistakes_all_positive,+1)
### YOUR CODE HERE
if weighted_mistakes_all_negative < weighted_mistakes_all_positive:
return (weighted_mistakes_all_negative, -1)
else:
return (weighted_mistakes_all_positive, 1)
# **Checkpoint:** Test your **intermediate_node_weighted_mistakes** function, run the following cell:
example_labels = tc.SArray([-1, -1, 1, 1, 1])
example_data_weights = tc.SArray([1., 2., .5, 1., 1.])
if intermediate_node_weighted_mistakes(example_labels, example_data_weights) == (2.5, -1):
print('Test passed!')
else:
print('Test failed... try again!')
# Recall that the **classification error** is defined as follows:
# $$
# \mbox{classification error} = \frac{\mbox{# mistakes}}{\mbox{# all data points}}
# $$
#
# **Quiz Question:** If we set the weights $\mathbf{\alpha} = 1$ for all data points, how is the weight of mistakes $\mbox{WM}(\mathbf{\alpha}, \mathbf{\hat{y}})$ related to the `classification error`?
# ### Function to pick best feature to split on
# We continue modifying our decision tree code from the earlier assignment to incorporate weighting of individual data points. The next step is to pick the best feature to split on.
#
# The **best_splitting_feature** function is similar to the one from the earlier assignment with two minor modifications:
# 1. The function **best_splitting_feature** should now accept an extra parameter `data_weights` to take account of weights of data points.
# 2. Instead of computing the number of mistakes in the left and right side of the split, we compute the weight of mistakes for both sides, add up the two weights, and divide it by the total weight of the data.
#
# Complete the following function. Comments starting with `DIFFERENT HERE` mark the sections where the weighted version differs from the original implementation.
def best_splitting_feature(data, features, target, data_weights):
# These variables will keep track of the best feature and the corresponding error
best_feature = None
best_error = float('+inf')
num_points = float(len(data))
# Loop through each feature to consider splitting on that feature
for feature in features:
# The left split will have all data points where the feature value is 0
# The right split will have all data points where the feature value is 1
left_split = data[data[feature] == 0]
right_split = data[data[feature] == 1]
# Apply the same filtering to data_weights to create left_data_weights, right_data_weights
## YOUR CODE HERE
left_data_weights = data_weights[data[feature] == 0]
right_data_weights = data_weights[data[feature] == 1]
# DIFFERENT HERE
# Calculate the weight of mistakes for left and right sides
## YOUR CODE HERE
left_weighted_mistakes, left_class = intermediate_node_weighted_mistakes(left_split[target], left_data_weights)
right_weighted_mistakes, right_class = intermediate_node_weighted_mistakes(right_split[target], right_data_weights)
# DIFFERENT HERE
# Compute weighted error by computing
# ( [weight of mistakes (left)] + [weight of mistakes (right)] ) / [total weight of all data points]
## YOUR CODE HERE
error = (left_weighted_mistakes + right_weighted_mistakes) / sum(data_weights)
# If this is the best error we have found so far, store the feature and the error
if error < best_error:
best_feature = feature
best_error = error
# Return the best feature we found
return best_feature
# **Checkpoint:** Now, we have another checkpoint to make sure you are on the right track.
example_data_weights = tc.SArray(len(train_data)* [1.5])
if best_splitting_feature(train_data, features, target, example_data_weights) == 'term. 36 months':
print('Test passed!')
else:
print('Test failed... try again!')
# **Note**. If you get an exception in the line of "the logical filter has different size than the array", try upgradting your Turi Create installation to 1.8.3 or newer.
# **Very Optional**. Relationship between weighted error and weight of mistakes
#
# By definition, the weighted error is the weight of mistakes divided by the weight of all data points, so
# $$
# \mathrm{E}(\mathbf{\alpha}, \mathbf{\hat{y}}) = \frac{\sum_{i=1}^{n} \alpha_i \times 1[y_i \neq \hat{y_i}]}{\sum_{i=1}^{n} \alpha_i} = \frac{\mathrm{WM}(\mathbf{\alpha}, \mathbf{\hat{y}})}{\sum_{i=1}^{n} \alpha_i}.
# $$
#
# In the code above, we obtain $\mathrm{E}(\mathbf{\alpha}, \mathbf{\hat{y}})$ from the two weights of mistakes from both sides, $\mathrm{WM}(\mathbf{\alpha}_{\mathrm{left}}, \mathbf{\hat{y}}_{\mathrm{left}})$ and $\mathrm{WM}(\mathbf{\alpha}_{\mathrm{right}}, \mathbf{\hat{y}}_{\mathrm{right}})$. First, notice that the overall weight of mistakes $\mathrm{WM}(\mathbf{\alpha}, \mathbf{\hat{y}})$ can be broken into two weights of mistakes over either side of the split:
# $$
# \mathrm{WM}(\mathbf{\alpha}, \mathbf{\hat{y}})
# = \sum_{i=1}^{n} \alpha_i \times 1[y_i \neq \hat{y_i}]
# = \sum_{\mathrm{left}} \alpha_i \times 1[y_i \neq \hat{y_i}]
# + \sum_{\mathrm{right}} \alpha_i \times 1[y_i \neq \hat{y_i}]\\
# = \mathrm{WM}(\mathbf{\alpha}_{\mathrm{left}}, \mathbf{\hat{y}}_{\mathrm{left}}) + \mathrm{WM}(\mathbf{\alpha}_{\mathrm{right}}, \mathbf{\hat{y}}_{\mathrm{right}})
# $$
# We then divide through by the total weight of all data points to obtain $\mathrm{E}(\mathbf{\alpha}, \mathbf{\hat{y}})$:
# $$
# \mathrm{E}(\mathbf{\alpha}, \mathbf{\hat{y}})
# = \frac{\mathrm{WM}(\mathbf{\alpha}_{\mathrm{left}}, \mathbf{\hat{y}}_{\mathrm{left}}) + \mathrm{WM}(\mathbf{\alpha}_{\mathrm{right}}, \mathbf{\hat{y}}_{\mathrm{right}})}{\sum_{i=1}^{n} \alpha_i}
# $$
# ### Building the tree
#
# With the above functions implemented correctly, we are now ready to build our decision tree. Recall from the previous assignments that each node in the decision tree is represented as a dictionary which contains the following keys:
#
# {
# 'is_leaf' : True/False.
# 'prediction' : Prediction at the leaf node.
# 'left' : (dictionary corresponding to the left tree).
# 'right' : (dictionary corresponding to the right tree).
# 'features_remaining' : List of features that are posible splits.
# }
#
# Let us start with a function that creates a leaf node given a set of target values:
def create_leaf(target_values, data_weights):
# Create a leaf node
leaf = {'splitting_feature' : None,
'is_leaf': True}
# Computed weight of mistakes.
weighted_error, best_class = intermediate_node_weighted_mistakes(target_values, data_weights)
# Store the predicted class (1 or -1) in leaf['prediction']
leaf['prediction'] = best_class
return leaf
# We provide a function that learns a weighted decision tree recursively and implements 3 stopping conditions:
# 1. All data points in a node are from the same class.
# 2. No more features to split on.
# 3. Stop growing the tree when the tree depth reaches **max_depth**.
def weighted_decision_tree_create(data, features, target, data_weights, current_depth = 1, max_depth = 10):
remaining_features = features[:] # Make a copy of the features.
target_values = data[target]
print("--------------------------------------------------------------------")
print("Subtree, depth = %s (%s data points)." % (current_depth, len(target_values)))
# Stopping condition 1. Error is 0.
if intermediate_node_weighted_mistakes(target_values, data_weights)[0] <= 1e-15:
print("Stopping condition 1 reached.")
return create_leaf(target_values, data_weights)
# Stopping condition 2. No more features.
if remaining_features == []:
print("Stopping condition 2 reached.")
return create_leaf(target_values, data_weights)
# Additional stopping condition (limit tree depth)
if current_depth > max_depth:
print("Reached maximum depth. Stopping for now.")
return create_leaf(target_values, data_weights)
splitting_feature = best_splitting_feature(data, features, target, data_weights)
remaining_features.remove(splitting_feature)
left_split = data[data[splitting_feature] == 0]
right_split = data[data[splitting_feature] == 1]
left_data_weights = data_weights[data[splitting_feature] == 0]
right_data_weights = data_weights[data[splitting_feature] == 1]
print("Split on feature %s. (%s, %s)" % (\
splitting_feature, len(left_split), len(right_split)))
# Create a leaf node if the split is "perfect"
if len(left_split) == len(data):
print("Creating leaf node.")
return create_leaf(left_split[target], data_weights)
if len(right_split) == len(data):
print("Creating leaf node.")
return create_leaf(right_split[target], data_weights)
# Repeat (recurse) on left and right subtrees
left_tree = weighted_decision_tree_create(
left_split, remaining_features, target, left_data_weights, current_depth + 1, max_depth)
right_tree = weighted_decision_tree_create(
right_split, remaining_features, target, right_data_weights, current_depth + 1, max_depth)
return {'is_leaf' : False,
'prediction' : None,
'splitting_feature': splitting_feature,
'left' : left_tree,
'right' : right_tree}
# Here is a recursive function to count the nodes in your tree:
def count_nodes(tree):
if tree['is_leaf']:
return 1
return 1 + count_nodes(tree['left']) + count_nodes(tree['right'])
# Run the following test code to check your implementation. Make sure you get **'Test passed'** before proceeding.
example_data_weights = tc.SArray([1.0 for i in range(len(train_data))])
small_data_decision_tree = weighted_decision_tree_create(train_data, features, target,
example_data_weights, max_depth=2)
if count_nodes(small_data_decision_tree) == 7:
print('Test passed!')
else:
print('Test failed... try again!')
print('Number of nodes found:', count_nodes(small_data_decision_tree))
print('Number of nodes that should be there: 7' )
# Let us take a quick look at what the trained tree is like. You should get something that looks like the following
#
# ```
# {'is_leaf': False,
# 'left': {'is_leaf': False,
# 'left': {'is_leaf': True, 'prediction': -1, 'splitting_feature': None},
# 'prediction': None,
# 'right': {'is_leaf': True, 'prediction': 1, 'splitting_feature': None},
# 'splitting_feature': 'grade.A'
# },
# 'prediction': None,
# 'right': {'is_leaf': False,
# 'left': {'is_leaf': True, 'prediction': 1, 'splitting_feature': None},
# 'prediction': None,
# 'right': {'is_leaf': True, 'prediction': -1, 'splitting_feature': None},
# 'splitting_feature': 'grade.D'
# },
# 'splitting_feature': 'term. 36 months'
# }```
small_data_decision_tree
# ### Making predictions with a weighted decision tree
# We give you a function that classifies one data point. It can also return the probability if you want to play around with that as well.
def classify(tree, x, annotate = False):
# If the node is a leaf node.
if tree['is_leaf']:
if annotate:
print("At leaf, predicting %s" % tree['prediction'])
return tree['prediction']
else:
# Split on feature.
split_feature_value = x[tree['splitting_feature']]
if annotate:
print("Split on %s = %s" % (tree['splitting_feature'], split_feature_value))
if split_feature_value == 0:
return classify(tree['left'], x, annotate)
else:
return classify(tree['right'], x, annotate)
# ### Evaluating the tree
#
# Now, we will write a function to evaluate a decision tree by computing the classification error of the tree on the given dataset.
#
# Again, recall that the **classification error** is defined as follows:
# $$
# \mbox{classification error} = \frac{\mbox{# mistakes}}{\mbox{# all data points}}
# $$
#
# The function called **evaluate_classification_error** takes in as input:
# 1. `tree` (as described above)
# 2. `data` (an SFrame)
#
# The function does not change because of adding data point weights.
def evaluate_classification_error(tree, data):
# Apply the classify(tree, x) to each row in your data
prediction = data.apply(lambda x: classify(tree, x))
# Once you've made the predictions, calculate the classification error
return (prediction != data[target]).sum() / float(len(data))
evaluate_classification_error(small_data_decision_tree, test_data)
# ### Example: Training a weighted decision tree
#
# To build intuition on how weighted data points affect the tree being built, consider the following:
#
# Suppose we only care about making good predictions for the **first 10 and last 10 items** in `train_data`, we assign weights:
# * 1 to the last 10 items
# * 1 to the first 10 items
# * and 0 to the rest.
#
# Let us fit a weighted decision tree with `max_depth = 2`.
# +
# Assign weights
example_data_weights = tc.SArray([1.] * 10 + [0.]*(len(train_data) - 20) + [1.] * 10)
# Train a weighted decision tree model.
small_data_decision_tree_subset_20 = weighted_decision_tree_create(train_data, features, target,
example_data_weights, max_depth=2)
# -
# Now, we will compute the classification error on the `subset_20`, i.e. the subset of data points whose weight is 1 (namely the first and last 10 data points).
subset_20 = train_data.head(10).append(train_data.tail(10))
evaluate_classification_error(small_data_decision_tree_subset_20, subset_20)
# Now, let us compare the classification error of the model `small_data_decision_tree_subset_20` on the entire test set `train_data`:
evaluate_classification_error(small_data_decision_tree_subset_20, train_data)
# The model `small_data_decision_tree_subset_20` performs **a lot** better on `subset_20` than on `train_data`.
#
# So, what does this mean?
# * The points with higher weights are the ones that are more important during the training process of the weighted decision tree.
# * The points with zero weights are basically ignored during training.
#
# **Quiz Question**: Will you get the same model as `small_data_decision_tree_subset_20` if you trained a decision tree with only the 20 data points with non-zero weights from the set of points in `subset_20`?
# # Implementing your own Adaboost (on decision stumps)
# Now that we have a weighted decision tree working, it takes only a bit of work to implement Adaboost. For the sake of simplicity, let us stick with **decision tree stumps** by training trees with **`max_depth=1`**.
# Recall from the lecture the procedure for Adaboost:
#
# 1\. Start with unweighted data with $\alpha_j = 1$
#
# 2\. For t = 1,...T:
# * Learn $f_t(x)$ with data weights $\alpha_j$
# * Compute coefficient $\hat{w}_t$:
# $$\hat{w}_t = \frac{1}{2}\ln{\left(\frac{1- \mbox{E}(\mathbf{\alpha}, \mathbf{\hat{y}})}{\mbox{E}(\mathbf{\alpha}, \mathbf{\hat{y}})}\right)}$$
# * Re-compute weights $\alpha_j$:
# $$\alpha_j \gets \begin{cases}
# \alpha_j \exp{(-\hat{w}_t)} & \text{ if }f_t(x_j) = y_j\\
# \alpha_j \exp{(\hat{w}_t)} & \text{ if }f_t(x_j) \neq y_j
# \end{cases}$$
# * Normalize weights $\alpha_j$:
# $$\alpha_j \gets \frac{\alpha_j}{\sum_{i=1}^{N}{\alpha_i}} $$
#
# Complete the skeleton for the following code to implement **adaboost_with_tree_stumps**. Fill in the places with `YOUR CODE HERE`.
# +
from math import log
from math import exp
def adaboost_with_tree_stumps(data, features, target, num_tree_stumps):
# start with unweighted data
alpha = tc.SArray([1.]*len(data))
weights = []
tree_stumps = []
target_values = data[target]
for t in range(num_tree_stumps):
print('=====================================================')
print('Adaboost Iteration %d' % t)
print('=====================================================')
# Learn a weighted decision tree stump. Use max_depth=1
tree_stump = weighted_decision_tree_create(data, features, target, data_weights=alpha, max_depth=1)
tree_stumps.append(tree_stump)
# Make predictions
predictions = data.apply(lambda x: classify(tree_stump, x))
# Produce a Boolean array indicating whether
# each data point was correctly classified
is_correct = predictions == target_values
is_wrong = predictions != target_values
# Compute weighted error
# YOUR CODE HERE
weighted_error = sum(is_wrong * alpha) / sum(alpha)
# Compute model coefficient using weighted error
# YOUR CODE HERE
weight = 0.5 * log((1 - weighted_error) / weighted_error)
weights.append(weight)
# Adjust weights on data point
adjustment = is_correct.apply(lambda is_correct : exp(-weight) if is_correct else exp(weight))
# Scale alpha by multiplying by adjustment
# Then normalize data points weights
## YOUR CODE HERE
alpha = alpha * adjustment
alpha = alpha / sum(alpha)
return weights, tree_stumps
# -
# ### Checking your Adaboost code
#
# Train an ensemble of **two** tree stumps and see which features those stumps split on. We will run the algorithm with the following parameters:
# * `train_data`
# * `features`
# * `target`
# * `num_tree_stumps = 2`
stump_weights, tree_stumps = adaboost_with_tree_stumps(train_data, features, target, num_tree_stumps=2)
def print_stump(tree):
split_name = tree['splitting_feature'] # split_name is something like 'term. 36 months'
if split_name is None:
print("(leaf, label: %s)" % tree['prediction'])
return None
split_feature, split_value = split_name.split('.')
print(' root')
print(' |---------------|----------------|')
print(' | |')
print(' | |')
print(' | |')
print(' [{0} == 0]{1}[{0} == 1] '.format(split_name, ' '*(27-len(split_name))))
print(' | |')
print(' | |')
print(' | |')
print(' (%s) (%s)' \
% (('leaf, label: ' + str(tree['left']['prediction']) if tree['left']['is_leaf'] else 'subtree'),
('leaf, label: ' + str(tree['right']['prediction']) if tree['right']['is_leaf'] else 'subtree')))
# Here is what the first stump looks like:
print_stump(tree_stumps[0])
# Here is what the next stump looks like:
print_stump(tree_stumps[1])
print(stump_weights)
# If your Adaboost is correctly implemented, the following things should be true:
#
# * `tree_stumps[0]` should split on **term. 36 months** with the prediction -1 on the left and +1 on the right.
# * `tree_stumps[1]` should split on **grade.A** with the prediction -1 on the left and +1 on the right.
# * Weights should be approximately `[0.158, 0.177]`
#
# **Reminders**
# - Stump weights ($\mathbf{\hat{w}}$) and data point weights ($\mathbf{\alpha}$) are two different concepts.
# - Stump weights ($\mathbf{\hat{w}}$) tell you how important each stump is while making predictions with the entire boosted ensemble.
# - Data point weights ($\mathbf{\alpha}$) tell you how important each data point is while training a decision stump.
# ### Training a boosted ensemble of 10 stumps
# Let us train an ensemble of 10 decision tree stumps with Adaboost. We run the **adaboost_with_tree_stumps** function with the following parameters:
# * `train_data`
# * `features`
# * `target`
# * `num_tree_stumps = 10`
stump_weights, tree_stumps = adaboost_with_tree_stumps(train_data, features, target, num_tree_stumps=10)
# ## Making predictions
#
# Recall from the lecture that in order to make predictions, we use the following formula:
# $$
# \hat{y} = sign\left(\sum_{t=1}^T \hat{w}_t f_t(x)\right)
# $$
#
# We need to do the following things:
# - Compute the predictions $f_t(x)$ using the $t$-th decision tree
# - Compute $\hat{w}_t f_t(x)$ by multiplying the `stump_weights` with the predictions $f_t(x)$ from the decision trees
# - Sum the weighted predictions over each stump in the ensemble.
#
# Complete the following skeleton for making predictions:
def predict_adaboost(stump_weights, tree_stumps, data):
scores = tc.SArray([0.]*len(data))
for i, tree_stump in enumerate(tree_stumps):
predictions = data.apply(lambda x: classify(tree_stump, x))
# Accumulate predictions on scores array
# YOUR CODE HERE
scores += stump_weights[i] * predictions
return scores.apply(lambda score : +1 if score > 0 else -1)
predictions = predict_adaboost(stump_weights, tree_stumps, test_data)
accuracy = tc.evaluation.accuracy(test_data[target], predictions)
print('Accuracy of 10-component ensemble = %s' % accuracy )
# Now, let us take a quick look what the `stump_weights` look like at the end of each iteration of the 10-stump ensemble:
stump_weights
# **Quiz Question:** Are the weights monotonically decreasing, monotonically increasing, or neither?
#
# **Reminder**: Stump weights ($\mathbf{\hat{w}}$) tell you how important each stump is while making predictions with the entire boosted ensemble.
# # Performance plots
#
# In this section, we will try to reproduce some of the performance plots dicussed in the lecture.
#
# ### How does accuracy change with adding stumps to the ensemble?
#
# We will now train an ensemble with:
# * `train_data`
# * `features`
# * `target`
# * `num_tree_stumps = 30`
#
# Once we are done with this, we will then do the following:
# * Compute the classification error at the end of each iteration.
# * Plot a curve of classification error vs iteration.
#
# First, lets train the model.
# this may take a while...
stump_weights, tree_stumps = adaboost_with_tree_stumps(train_data, features, target, num_tree_stumps=30)
# ### Computing training error at the end of each iteration
#
# Now, we will compute the classification error on the **train_data** and see how it is reduced as trees are added.
error_all = []
for n in range(1, 31):
predictions = predict_adaboost(stump_weights[:n], tree_stumps[:n], train_data)
error = 1.0 - tc.evaluation.accuracy(train_data[target], predictions)
error_all.append(error)
print("Iteration %s, training error = %s" % (n, error_all[n-1]))
# ### Visualizing training error vs number of iterations
#
# We have provided you with a simple code snippet that plots classification error with the number of iterations.
# +
plt.rcParams['figure.figsize'] = 7, 5
plt.plot(range(1,31), error_all, '-', linewidth=4.0, label='Training error')
plt.title('Performance of Adaboost ensemble')
plt.xlabel('# of iterations')
plt.ylabel('Classification error')
plt.legend(loc='best', prop={'size':15})
plt.rcParams.update({'font.size': 16})
# -
# **Quiz Question**: Which of the following best describes a **general trend in accuracy** as we add more and more components? Answer based on the 30 components learned so far.
#
# 1. Training error goes down monotonically, i.e. the training error reduces with each iteration but never increases.
# 2. Training error goes down in general, with some ups and downs in the middle.
# 3. Training error goes up in general, with some ups and downs in the middle.
# 4. Training error goes down in the beginning, achieves the best error, and then goes up sharply.
# 5. None of the above
#
#
# ### Evaluation on the test data
#
# Performing well on the training data is cheating, so lets make sure it works on the `test_data` as well. Here, we will compute the classification error on the `test_data` at the end of each iteration.
test_error_all = []
for n in range(1, 31):
predictions = predict_adaboost(stump_weights[:n], tree_stumps[:n], test_data)
error = 1.0 - tc.evaluation.accuracy(test_data[target], predictions)
test_error_all.append(error)
print("Iteration %s, test error = %s" % (n, test_error_all[n-1]))
# ### Visualize both the training and test errors
#
# Now, let us plot the training & test error with the number of iterations.
# +
plt.rcParams['figure.figsize'] = 7, 5
plt.plot(range(1,31), error_all, '-', linewidth=4.0, label='Training error')
plt.plot(range(1,31), test_error_all, '-', linewidth=4.0, label='Test error')
plt.title('Performance of Adaboost ensemble')
plt.xlabel('# of iterations')
plt.ylabel('Classification error')
plt.rcParams.update({'font.size': 16})
plt.legend(loc='best', prop={'size':15})
plt.tight_layout()
# -
# **Quiz Question:** From this plot (with 30 trees), is there massive overfitting as the # of iterations increases?
| 03_machine_learning_classification/week_5/quiz_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import Libraries
import cv2
import numpy as np
# ### Create Classifier
car_classifier=cv2.CascadeClassifier('Haarcascades/haarcascade_car.xml')
# ### Capture Video
cap=cv2.VideoCapture('Video/Vehicles.mp4')
# ### It's a Kind of Magic
while cap.isOpened():
ret, frame =cap.read()
cars = car_classifier.detectMultiScale(frame,1.4,2)
for (x,y,w,h) in cars:
cv2.rectangle(frame,(x,y),(x+w,y+h),(0,255,225),2)
cv2.imshow('Cars',frame)
if cv2.waitKey(1) & 0xFF == 27:
break
cap.release()
cv2.destroyAllWindows()
| Projects/05_Car_Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:eu-west-1:470317259841:image/datascience-1.0
# ---
# # Targeting Direct Marketing with Amazon SageMaker XGBoost
# _**Supervised Learning with Gradient Boosted Trees: A Binary Prediction Problem With Unbalanced Classes**_
#
# ---
#
# ---
#
# ## Contents
#
# 1. [Background](#Background)
# 1. [Prepration](#Preparation)
# 1. [Data](#Data)
# 1. [Exploration](#Exploration)
# 1. [Transformation](#Transformation)
# 1. [Training](#Training)
# 1. [Hosting](#Hosting)
# 1. [Evaluation](#Evaluation)
# 1. [Exentsions](#Extensions)
#
# ---
#
# ## Background
# Direct marketing, either through mail, email, phone, etc., is a common tactic to acquire customers. Because resources and a customer's attention is limited, the goal is to only target the subset of prospects who are likely to engage with a specific offer. Predicting those potential customers based on readily available information like demographics, past interactions, and environmental factors is a common machine learning problem.
#
# This notebook presents an example problem to predict if a customer will enroll for a term deposit at a bank, after one or more phone calls. The steps include:
#
# * Preparing your Amazon SageMaker notebook
# * Downloading data from the internet into Amazon SageMaker
# * Investigating and transforming the data so that it can be fed to Amazon SageMaker algorithms
# * Estimating a model using the Gradient Boosting algorithm
# * Evaluating the effectiveness of the model
# * Setting the model up to make on-going predictions
#
# ---
#
# ## Preparation
#
# _This notebook was created and tested on an ml.m4.xlarge notebook instance._
#
# Let's start by specifying:
#
# - The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# - The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
conda update pandas
# + isConfigCell=true
import sagemaker
bucket=sagemaker.Session().default_bucket()
prefix = 'sagemaker/DEMO-xgboost-dm'
# Define IAM role
import boto3
import re
from sagemaker import get_execution_role
role = get_execution_role()
# -
# Now let's bring in the Python libraries that we'll use throughout the analysis
import numpy as np # For matrix operations and numerical processing
import pandas as pd # For munging tabular data
import matplotlib.pyplot as plt # For charts and visualizations
from IPython.display import Image # For displaying images in the notebook
from IPython.display import display # For displaying outputs in the notebook
from time import gmtime, strftime # For labeling SageMaker models, endpoints, etc.
import sys # For writing outputs to notebook
import math # For ceiling function
import json # For parsing hosting outputs
import os # For manipulating filepath names
import sagemaker
import zipfile # Amazon SageMaker's Python SDK provides many helper functions
pd.__version__
# Make sure pandas version is set to 1.2.4 or later. If it is not the case, restart the kernel before going further
# ---
#
# ## Data
# Let's start by downloading the [direct marketing dataset](https://sagemaker-sample-data-us-west-2.s3-us-west-2.amazonaws.com/autopilot/direct_marketing/bank-additional.zip) from the sample data s3 bucket.
#
# \[Moro et al., 2014\] <NAME>, <NAME> and <NAME>. A Data-Driven Approach to Predict the Success of Bank Telemarketing. Decision Support Systems, Elsevier, 62:22-31, June 2014
#
# +
# !wget https://sagemaker-sample-data-us-west-2.s3-us-west-2.amazonaws.com/autopilot/direct_marketing/bank-additional.zip
with zipfile.ZipFile('bank-additional.zip', 'r') as zip_ref:
zip_ref.extractall('.')
# -
# Now lets read this into a Pandas data frame and take a look.
data = pd.read_csv('./bank-additional/bank-additional-full.csv')
pd.set_option('display.max_columns', 500) # Make sure we can see all of the columns
pd.set_option('display.max_rows', 20) # Keep the output on one page
data
# Let's talk about the data. At a high level, we can see:
#
# * We have a little over 40K customer records, and 20 features for each customer
# * The features are mixed; some numeric, some categorical
# * The data appears to be sorted, at least by `time` and `contact`, maybe more
#
# _**Specifics on each of the features:**_
#
# *Demographics:*
# * `age`: Customer's age (numeric)
# * `job`: Type of job (categorical: 'admin.', 'services', ...)
# * `marital`: Marital status (categorical: 'married', 'single', ...)
# * `education`: Level of education (categorical: 'basic.4y', 'high.school', ...)
#
# *Past customer events:*
# * `default`: Has credit in default? (categorical: 'no', 'unknown', ...)
# * `housing`: Has housing loan? (categorical: 'no', 'yes', ...)
# * `loan`: Has personal loan? (categorical: 'no', 'yes', ...)
#
# *Past direct marketing contacts:*
# * `contact`: Contact communication type (categorical: 'cellular', 'telephone', ...)
# * `month`: Last contact month of year (categorical: 'may', 'nov', ...)
# * `day_of_week`: Last contact day of the week (categorical: 'mon', 'fri', ...)
# * `duration`: Last contact duration, in seconds (numeric). Important note: If duration = 0 then `y` = 'no'.
#
# *Campaign information:*
# * `campaign`: Number of contacts performed during this campaign and for this client (numeric, includes last contact)
# * `pdays`: Number of days that passed by after the client was last contacted from a previous campaign (numeric)
# * `previous`: Number of contacts performed before this campaign and for this client (numeric)
# * `poutcome`: Outcome of the previous marketing campaign (categorical: 'nonexistent','success', ...)
#
# *External environment factors:*
# * `emp.var.rate`: Employment variation rate - quarterly indicator (numeric)
# * `cons.price.idx`: Consumer price index - monthly indicator (numeric)
# * `cons.conf.idx`: Consumer confidence index - monthly indicator (numeric)
# * `euribor3m`: Euribor 3 month rate - daily indicator (numeric)
# * `nr.employed`: Number of employees - quarterly indicator (numeric)
#
# *Target variable:*
# * `y`: Has the client subscribed a term deposit? (binary: 'yes','no')
# ### Exploration
# Let's start exploring the data. First, let's understand how the features are distributed.
# +
# Frequency tables for each categorical feature
for column in data.select_dtypes(include=['object']).columns:
display(pd.crosstab(index=data[column], columns='% observations', normalize='columns'))
# Histograms for each numeric features
display(data.describe())
# %matplotlib inline
hist = data.hist(bins=30, sharey=True, figsize=(10, 10))
# -
# Notice that:
#
# * Almost 90% of the values for our target variable `y` are "no", so most customers did not subscribe to a term deposit.
# * Many of the predictive features take on values of "unknown". Some are more common than others. We should think carefully as to what causes a value of "unknown" (are these customers non-representative in some way?) and how we that should be handled.
# * Even if "unknown" is included as it's own distinct category, what does it mean given that, in reality, those observations likely fall within one of the other categories of that feature?
# * Many of the predictive features have categories with very few observations in them. If we find a small category to be highly predictive of our target outcome, do we have enough evidence to make a generalization about that?
# * Contact timing is particularly skewed. Almost a third in May and less than 1% in December. What does this mean for predicting our target variable next December?
# * There are no missing values in our numeric features. Or missing values have already been imputed.
# * `pdays` takes a value near 1000 for almost all customers. Likely a placeholder value signifying no previous contact.
# * Several numeric features have a very long tail. Do we need to handle these few observations with extremely large values differently?
# * Several numeric features (particularly the macroeconomic ones) occur in distinct buckets. Should these be treated as categorical?
#
# Next, let's look at how our features relate to the target that we are attempting to predict.
# +
for column in data.select_dtypes(include=['object']).columns:
if column != 'y':
display(pd.crosstab(index=data[column], columns=data['y'], normalize='columns'))
for column in data.select_dtypes(exclude=['object']).columns:
print(column)
hist = data[[column, 'y']].hist(by='y', bins=30)
plt.show()
# -
# Notice that:
#
# * Customers who are-- "blue-collar", "married", "unknown" default status, contacted by "telephone", and/or in "may" are a substantially lower portion of "yes" than "no" for subscribing.
# * Distributions for numeric variables are different across "yes" and "no" subscribing groups, but the relationships may not be straightforward or obvious.
#
# Now let's look at how our features relate to one another.
display(data.corr())
pd.plotting.scatter_matrix(data, figsize=(12, 12))
plt.show()
# Notice that:
# * Features vary widely in their relationship with one another. Some with highly negative correlation, others with highly positive correlation.
# * Relationships between features is non-linear and discrete in many cases.
# ### Transformation
#
# Cleaning up data is part of nearly every machine learning project. It arguably presents the biggest risk if done incorrectly and is one of the more subjective aspects in the process. Several common techniques include:
#
# * Handling missing values: Some machine learning algorithms are capable of handling missing values, but most would rather not. Options include:
# * Removing observations with missing values: This works well if only a very small fraction of observations have incomplete information.
# * Removing features with missing values: This works well if there are a small number of features which have a large number of missing values.
# * Imputing missing values: Entire [books](https://www.amazon.com/Flexible-Imputation-Missing-Interdisciplinary-Statistics/dp/1439868247) have been written on this topic, but common choices are replacing the missing value with the mode or mean of that column's non-missing values.
# * Converting categorical to numeric: The most common method is one hot encoding, which for each feature maps every distinct value of that column to its own feature which takes a value of 1 when the categorical feature is equal to that value, and 0 otherwise.
# * Oddly distributed data: Although for non-linear models like Gradient Boosted Trees, this has very limited implications, parametric models like regression can produce wildly inaccurate estimates when fed highly skewed data. In some cases, simply taking the natural log of the features is sufficient to produce more normally distributed data. In others, bucketing values into discrete ranges is helpful. These buckets can then be treated as categorical variables and included in the model when one hot encoded.
# * Handling more complicated data types: Mainpulating images, text, or data at varying grains is left for other notebook templates.
#
# Luckily, some of these aspects have already been handled for us, and the algorithm we are showcasing tends to do well at handling sparse or oddly distributed data. Therefore, let's keep pre-processing simple.
data['no_previous_contact'] = np.where(data['pdays'] == 999, 1, 0) # Indicator variable to capture when pdays takes a value of 999
data['not_working'] = np.where(np.in1d(data['job'], ['student', 'retired', 'unemployed']), 1, 0) # Indicator for individuals not actively employed
model_data = pd.get_dummies(data) # Convert categorical variables to sets of indicators
# Another question to ask yourself before building a model is whether certain features will add value in your final use case. For example, if your goal is to deliver the best prediction, then will you have access to that data at the moment of prediction? Knowing it's raining is highly predictive for umbrella sales, but forecasting weather far enough out to plan inventory on umbrellas is probably just as difficult as forecasting umbrella sales without knowledge of the weather. So, including this in your model may give you a false sense of precision.
#
# Following this logic, let's remove the economic features and `duration` from our data as they would need to be forecasted with high precision to use as inputs in future predictions.
#
# Even if we were to use values of the economic indicators from the previous quarter, this value is likely not as relevant for prospects contacted early in the next quarter as those contacted later on.
model_data = model_data.drop(['duration', 'emp.var.rate', 'cons.price.idx', 'cons.conf.idx', 'euribor3m', 'nr.employed'], axis=1)
# When building a model whose primary goal is to predict a target value on new data, it is important to understand overfitting. Supervised learning models are designed to minimize error between their predictions of the target value and actuals, in the data they are given. This last part is key, as frequently in their quest for greater accuracy, machine learning models bias themselves toward picking up on minor idiosyncrasies within the data they are shown. These idiosyncrasies then don't repeat themselves in subsequent data, meaning those predictions can actually be made less accurate, at the expense of more accurate predictions in the training phase.
#
# The most common way of preventing this is to build models with the concept that a model shouldn't only be judged on its fit to the data it was trained on, but also on "new" data. There are several different ways of operationalizing this, holdout validation, cross-validation, leave-one-out validation, etc. For our purposes, we'll simply randomly split the data into 3 uneven groups. The model will be trained on 70% of data, it will then be evaluated on 20% of data to give us an estimate of the accuracy we hope to have on "new" data, and 10% will be held back as a final testing dataset which will be used later on.
train_data, validation_data, test_data = np.split(model_data.sample(frac=1, random_state=1729), [int(0.7 * len(model_data)), int(0.9 * len(model_data))]) # Randomly sort the data then split out first 70%, second 20%, and last 10%
# Amazon SageMaker's XGBoost container expects data in the libSVM or CSV data format. For this example, we'll stick to CSV. Note that the first column must be the target variable and the CSV should not include headers. Also, notice that although repetitive it's easiest to do this after the train|validation|test split rather than before. This avoids any misalignment issues due to random reordering.
pd.concat([train_data['y_yes'], train_data.drop(['y_no', 'y_yes'], axis=1)], axis=1).to_csv('train.csv', index=False, header=False)
pd.concat([validation_data['y_yes'], validation_data.drop(['y_no', 'y_yes'], axis=1)], axis=1).to_csv('validation.csv', index=False, header=False)
# Now we'll copy the file to S3 for Amazon SageMaker's managed training to pickup.
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train/train.csv')).upload_file('train.csv')
boto3.Session().resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'validation/validation.csv')).upload_file('validation.csv')
# ---
#
# ## End of Lab 1
#
# ---
#
# ## Training
# Now we know that most of our features have skewed distributions, some are highly correlated with one another, and some appear to have non-linear relationships with our target variable. Also, for targeting future prospects, good predictive accuracy is preferred to being able to explain why that prospect was targeted. Taken together, these aspects make gradient boosted trees a good candidate algorithm.
#
# There are several intricacies to understanding the algorithm, but at a high level, gradient boosted trees works by combining predictions from many simple models, each of which tries to address the weaknesses of the previous models. By doing this the collection of simple models can actually outperform large, complex models. Other Amazon SageMaker notebooks elaborate on gradient boosting trees further and how they differ from similar algorithms.
#
# `xgboost` is an extremely popular, open-source package for gradient boosted trees. It is computationally powerful, fully featured, and has been successfully used in many machine learning competitions. Let's start with a simple `xgboost` model, trained using Amazon SageMaker's managed, distributed training framework.
#
# First we'll need to specify the ECR container location for Amazon SageMaker's implementation of XGBoost.
container = sagemaker.image_uris.retrieve(region=boto3.Session().region_name, framework='xgboost', version='latest')
# Then, because we're training with the CSV file format, we'll create `s3_input`s that our training function can use as a pointer to the files in S3, which also specify that the content type is CSV.
s3_input_train = sagemaker.inputs.TrainingInput(s3_data='s3://{}/{}/train'.format(bucket, prefix), content_type='csv')
s3_input_validation = sagemaker.inputs.TrainingInput(s3_data='s3://{}/{}/validation/'.format(bucket, prefix), content_type='csv')
# First we'll need to specify training parameters to the estimator. This includes:
# 1. The `xgboost` algorithm container
# 1. The IAM role to use
# 1. Training instance type and count
# 1. S3 location for output data
# 1. Algorithm hyperparameters
#
# And then a `.fit()` function which specifies:
# 1. S3 location for output data. In this case we have both a training and validation set which are passed in.
# +
sess = sagemaker.Session()
xgb = sagemaker.estimator.Estimator(container,
role,
instance_count=1,
instance_type='ml.m4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
xgb.set_hyperparameters(max_depth=5,
eta=0.2,
gamma=4,
min_child_weight=6,
subsample=0.8,
silent=0,
objective='binary:logistic',
num_round=100)
xgb.fit({'train': s3_input_train, 'validation': s3_input_validation})
# -
# ---
#
# ## Hosting
# Now that we've trained the `xgboost` algorithm on our data, let's deploy a model that's hosted behind a real-time endpoint.
xgb_predictor = xgb.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
# ---
#
# ## Evaluation
# There are many ways to compare the performance of a machine learning model, but let's start by simply comparing actual to predicted values. In this case, we're simply predicting whether the customer subscribed to a term deposit (`1`) or not (`0`), which produces a simple confusion matrix.
#
# First we'll need to determine how we pass data into and receive data from our endpoint. Our data is currently stored as NumPy arrays in memory of our notebook instance. To send it in an HTTP POST request, we'll serialize it as a CSV string and then decode the resulting CSV.
#
# *Note: For inference with CSV format, SageMaker XGBoost requires that the data does NOT include the target variable.*
xgb_predictor.serializer = sagemaker.serializers.CSVSerializer()
# Now, we'll use a simple function to:
# 1. Loop over our test dataset
# 1. Split it into mini-batches of rows
# 1. Convert those mini-batches to CSV string payloads (notice, we drop the target variable from our dataset first)
# 1. Retrieve mini-batch predictions by invoking the XGBoost endpoint
# 1. Collect predictions and convert from the CSV output our model provides into a NumPy array
# +
def predict(data, predictor, rows=500 ):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = ''
for array in split_array:
predictions = ','.join([predictions, predictor.predict(array).decode('utf-8')])
return np.fromstring(predictions[1:], sep=',')
predictions = predict(test_data.drop(['y_no', 'y_yes'], axis=1).to_numpy(), xgb_predictor)
# -
# Now we'll check our confusion matrix to see how well we predicted versus actuals.
pd.crosstab(index=test_data['y_yes'], columns=np.round(predictions), rownames=['actuals'], colnames=['predictions'])
# So, of the ~4000 potential customers, we predicted 136 would subscribe and 94 of them actually did. We also had 389 subscribers who subscribed that we did not predict would. This is less than desirable, but the model can (and should) be tuned to improve this. Most importantly, note that with minimal effort, our model produced accuracies similar to those published [here](http://media.salford-systems.com/video/tutorial/2015/targeted_marketing.pdf).
#
# _Note that because there is some element of randomness in the algorithm's subsample, your results may differ slightly from the text written above._
# ## Automatic model Tuning (optional)
# Amazon SageMaker automatic model tuning, also known as hyperparameter tuning, finds the best version of a model by running many training jobs on your dataset using the algorithm and ranges of hyperparameters that you specify. It then chooses the hyperparameter values that result in a model that performs the best, as measured by a metric that you choose.
# For example, suppose that you want to solve a binary classification problem on this marketing dataset. Your goal is to maximize the area under the curve (auc) metric of the algorithm by training an XGBoost Algorithm model. You don't know which values of the eta, alpha, min_child_weight, and max_depth hyperparameters to use to train the best model. To find the best values for these hyperparameters, you can specify ranges of values that Amazon SageMaker hyperparameter tuning searches to find the combination of values that results in the training job that performs the best as measured by the objective metric that you chose. Hyperparameter tuning launches training jobs that use hyperparameter values in the ranges that you specified, and returns the training job with highest auc.
#
from sagemaker.tuner import IntegerParameter, CategoricalParameter, ContinuousParameter, HyperparameterTuner
hyperparameter_ranges = {'eta': ContinuousParameter(0, 1),
'min_child_weight': ContinuousParameter(1, 10),
'alpha': ContinuousParameter(0, 2),
'max_depth': IntegerParameter(1, 10)}
objective_metric_name = 'validation:auc'
tuner = HyperparameterTuner(xgb,
objective_metric_name,
hyperparameter_ranges,
max_jobs=20,
max_parallel_jobs=3)
tuner.fit({'train': s3_input_train, 'validation': s3_input_validation})
boto3.client('sagemaker').describe_hyper_parameter_tuning_job(
HyperParameterTuningJobName=tuner.latest_tuning_job.job_name)['HyperParameterTuningJobStatus']
# return the best training job name
tuner.best_training_job()
# Deploy the best trained or user specified model to an Amazon SageMaker endpoint
tuner_predictor = tuner.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
# Create a serializer
tuner_predictor.serializer = sagemaker.serializers.CSVSerializer()
# Predict
predictions = predict(test_data.drop(['y_no', 'y_yes'], axis=1).to_numpy(),tuner_predictor)
# Collect predictions and convert from the CSV output our model provides into a NumPy array
pd.crosstab(index=test_data['y_yes'], columns=np.round(predictions), rownames=['actuals'], colnames=['predictions'])
# ---
#
# ## Extensions
#
# This example analyzed a relatively small dataset, but utilized Amazon SageMaker features such as distributed, managed training and real-time model hosting, which could easily be applied to much larger problems. In order to improve predictive accuracy further, we could tweak value we threshold our predictions at to alter the mix of false-positives and false-negatives, or we could explore techniques like hyperparameter tuning. In a real-world scenario, we would also spend more time engineering features by hand and would likely look for additional datasets to include which contain customer information not available in our initial dataset.
# ### (Optional) Clean-up
#
# If you are done with this notebook, please run the cell below. This will remove the hosted endpoint you created and avoid any charges from a stray instance being left on.
xgb_predictor.delete_endpoint(delete_endpoint_config=True)
tuner_predictor.delete_endpoint(delete_endpoint_config=True)
| xgboost_direct_marketing_sagemaker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2
# The objective of this assignment is to get you familiarizewith the problems of `classification` and `verification`with a popular problem space of `face`
#
# This jupyter notebook is meant to be used in conjunction with the full questions in the assignment pdf.
#
# ## Instructions
# - Write your code and analyses in the indicated cells.
# - Ensure that this notebook runs without errors when the cells are run in sequence.
# - Do not attempt to change the contents of the other cells.
#
# ## Allowed Libraries
# - All libraries are allowed
#
# ## Datasets
# - 3 datasets are provided. Load the data from the drive [link](!https://drive.google.com/file/d/1ujsKv9W5eidb4TXt1pnsqwDKVDFtzZTh/view?usp=sharing).
# - Unzip the downloaded file and store the files in a folder called `datasets`. Keep the `datasets` folder in the same directory as of the jupyter notebook
#
# ## Submission
# - Ensure that this notebook runs without errors when the cells are run in sequence.
# - Rename the notebook to `<roll_number>.ipynb` and submit ONLY the notebook file on moodle.
# - Upload the notebook, report and classification results as a zip file to moodle. Name the zip file as `<rollnumber>_assignment2.zip`
# Installing Libraries
# !pip install scikit-learn matplotlib Pillow
# +
# Basic Imports
import os
import sys
import warnings
import numpy as np
import pandas as pd
from scipy import linalg
# Loading and plotting data
from PIL import Image
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# Features
from sklearn.decomposition import PCA
from sklearn.decomposition import KernelPCA
from sklearn.discriminant_analysis import _class_means,_class_cov
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
from sklearn.manifold import TSNE
plt.ion()
# %matplotlib inline
# -
# # Parameters
# - Image size: Bigger images create better representation but would require more computation. Choose the correct image size based on your Laptop configuration.
# - is_grayscale: Should you take grayscale images? Or rgb images? Choose whichever gives better representation for classification.
opt = {
'image_size': 32,
'is_grayscale': False,
'val_split': 0.75
}
# ### Load Dataset
# + editable=false
cfw_dict = {'Amitabhbachan': 0,
'AamirKhan': 1,
'DwayneJohnson': 2,
'AishwaryaRai': 3,
'BarackObama': 4,
'NarendraModi': 5,
'ManmohanSingh': 6,
'VladimirPutin': 7}
imfdb_dict = {'MadhuriDixit': 0,
'Kajol': 1,
'SharukhKhan': 2,
'ShilpaShetty': 3,
'AmitabhBachan': 4,
'KatrinaKaif': 5,
'AkshayKumar': 6,
'Amir': 7}
# Load Image using PIL for dataset
def load_image(path):
im = Image.open(path).convert('L' if opt['is_grayscale'] else 'RGB')
im = im.resize((opt['image_size'],opt['image_size']))
im = np.array(im)
im = im/256
return im
# Load the full data from directory
def load_data(dir_path):
image_list = []
y_list = []
if "CFW" in dir_path:
label_dict = cfw_dict
elif "yale" in dir_path.lower():
label_dict = {}
for i in range(15):
label_dict[str(i+1)] = i
elif "IMFDB" in dir_path:
label_dict = imfdb_dict
else:
raise KeyError("Dataset not found.")
for filename in sorted(os.listdir(dir_path)):
if filename.endswith(".png"):
im = load_image(os.path.join(dir_path,filename))
y = filename.split('_')[0]
y = label_dict[y]
image_list.append(im)
y_list.append(y)
else:
continue
image_list = np.array(image_list)
y_list = np.array(y_list)
print("Dataset shape:",image_list.shape)
return image_list,y_list
# Display N Images in a nice format
def disply_images(imgs,classes,row=1,col=2,w=64,h=64):
fig=plt.figure(figsize=(8, 8))
for i in range(1, col*row +1):
img = imgs[i-1]
fig.add_subplot(row, col, i)
if opt['is_grayscale']:
plt.imshow(img , cmap='gray')
else:
plt.imshow(img)
plt.title("Class:{}".format(classes[i-1]))
plt.axis('off')
plt.show()
# -
# Loading the dataset
# eg.
# dirpath = './dataset/IMFDB/'
X,y = load_data(dirpath)
N,H,W = X.shape[0:3]
C = 1 if opt['is_grayscale'] else X.shape[3]
# + editable=false
# Show sample images
ind = np.random.randint(0,y.shape[0],6)
disply_images(X[ind,...],y[ind], row=2,col=3)
# -
# # Features
# You are provided 6 Features. These features are:
# - Eigen Faces / PCA
# - Kernel PCA
# - Fisher Face / LDA
# - Kernel Fisher Face
# - VGG Features
# - Resnet Features
#
# **VGG and Resnet features are last layer features learned by training a model for image classification**
#
# ---
#
# + editable=false
# Flatten to apply PCA/LDA
X = X.reshape((N,H*W*C))
# -
# ### 1. Eigen Face:
# Use principal component analysis to get the eigen faces.
# Go through the [documentation](!http://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html) on how to use it
def get_pca(X,k):
"""
Get PCA of K dimension using the top eigen vectors
"""
pca = PCA(n_components=k)
X_k = pca.fit_transform(X)
return X_k
# ### 2. Kernel Face:
# Use Kernel principal component analysis to get the eigen faces.
#
# There are different kernels that can be used. Eg. Poly, rbf, sigmoid. Choose the whichever gives the best result or representation. See [link](!https://data-flair.training/blogs/svm-kernel-functions/) for better understanding of these kernels
#
# Go through the [documentation](!https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.KernelPCA.html#sklearn.decomposition.KernelPCA) on how to use it different kernels in Sklearn.
def get_kernel_pca(X, k,kernel='rbf', degree=3):
"""
Get PCA of K dimension using the top eigen vectors
@param: X => Your data flattened to D dimension
@param: k => Number of components
@param: kernel => which kernel to use (“linear” | “poly” | “rbf” | “sigmoid” | “cosine” )
@param: d => Degree for poly kernels. Ignored by other kernels
"""
kpca = KernelPCA(n_components=k,kernel=kernel,degree=degree)
X_k = kpca.fit_transform(X)
return X_k
# ### 3. Fisher Face
# Another method similar to the eigenface technique is `fisherfaces` which uses linear discriminant analysis.
# This method for facial recognition is less sensitive to variation in lighting and pose of the face than using eigenfaces. Fisherface uses labelled data to retain more of the class-specific information during the dimension reduction stage.
#
# Go through the [documentation](!https://scikit-learn.org/stable/modules/generated/sklearn.discriminant_analysis.LinearDiscriminantAnalysis.html) on how to use it different kernels in Sklearn.
def get_lda(X,y, k):
"""
Get LDA of K dimension
@param: X => Your data flattened to D dimension
@param: k => Number of components
"""
lda = LDA(n_components=k)
X_k = lda.fit_transform(X,y)
return X_k
# ### 4. Kernel Fisher Face
# Use LDA using different kernels similiar to KernelPCA. Here the input is directly transformed instead of using the kernel trick.
def get_kernel_lda(X,y,k,kernel='rbf',degree=3):
"""
Get LDA of K dimension
@param: X => Your data flattened to D dimension
@param: k => Number of components
@param: kernel => which kernel to use ( “poly” | “rbf” | “sigmoid”)
"""
# Transform input
if kernel == "poly":
X_transformed = X**degree
elif kernel == "rbf":
var = np.var(X)
X_transformed= np.exp(-X/(2*var))
elif kernel == "sigmoid":
X_transformed = np.tanh(X)
else:
raise NotImplementedError("Kernel {} Not defined".format(kernel))
klda = LDA(n_components=k)
X_k = klda.fit_transform(X,y)
return X_k
# ### 5. VGG Features
# VGG Neural Networks a 19 layer CNN architecture introduced by <NAME>([Link](!https://arxiv.org/pdf/1409.1556.pdf) to paper). We are providing you with the last fully connected layer of this model.
#
# The model was trained for face classification on each dataset and each feature the dimension of 4096.
def get_vgg_features(dirpath):
features = np.load(os.path.join(dirpath,"VGG19_features.npy"))
return features
# ### 6. Resnet Features
#
# [Residual neural networks](!https://arxiv.org/pdf/1512.03385.pdf) are CNN with large depth, to effectively train these netwrorks they utilize skip connections, or short-cuts to jump over some layers. This helps solving [vanishing gradient problem](!https://en.wikipedia.org/wiki/Vanishing_gradient_problem)
#
# A 50 layer resnet model was trained for face classification on each dataset. Each feature the dimension of 2048
def get_resnet_features(dirpath):
features = np.load(os.path.join(dirpath,"resnet50_features.npy"))
return features
# # Questions
#
# 1(a). What are eigen faces?
#
# ___________________________
#
# Your answers here (double click to edit)
#
# 1(b). How many eigen vec-tors/faces are required to “satisfactorily” reconstruct a person in these three datasets? (Don’t forget to make your argument based on eigen value spectrum) Show appropriate graphs, qualitative examples andmake a convincing argument.
# +
# Compute your features
# eg.
# X_3D = get_kernel_lda(X,y,3)
# +
# Create a scatter plot
# eg.
# fig = plt.figure(figsize=(8,8))
# ax = fig.add_subplot(111, projection='3d')
# ax.scatter(X_3D[:,0],X_3D[:,1],X_3D[:,2],c=y)
# +
# Plot the eigen value spectrum
# -
# 1(c). Reconstruct the image back for each case
#
def reconstruct_images(<input_parameters>,*args,**kwargs):
"""
Reconstruct the images back by just using the selected principal components.
You have to write the code in this code block.
You can change the functions provided above (eg, get_pca, get_lda) for your use case.
@params:
Input parameters
@return reconstructed_X => reconstructed image
"""
pass
reconstruct_X = None
return reconstruct_X
# +
# Display results
# X_reconstruced = reconstruct_images()
# Display random images
# ind = np.random.randint(0,y.shape[0],6)
# disply_images(X_reconstruced_3D[ind,...],y[ind],row=2,col=3)
# Show the reconstruction error
print(np.sqrt(np.mean((X - X_reconstructed)**2)))
# -
# 1(d). Which person/identity is difficult to represent com-pactly with fewer eigen vectors? Why is that? Explain with your empirical observations and intuitive answers
# +
# code goes here
# -
# 2(a). Use any classifier(MLP, Logistic regression, SVM, Decision Trees) and find the classification accuracy.
#
# 2(b)Which method works well? Do a comparitivestudy.
#
#
# You already know the paper [Face Recognition Us-ing Kernel Methods](!http://face-rec.org/algorithms/Kernel/nips01.pdf) .See this as an example for empirical analysis of different features/classification.
# +
# Define your classifier here. You can use libraries like sklearn to create your classifier
class Classifier():
def __init__():
super.__init__()
# Define your parameters eg, W,b, max_iterations etc.
def classify(self,X):
"""
Given an input X classify it into appropriate class.
"""
return prediction
def confusion_matrix(self,pred,y):
"""
A confusion matrix is a table that is often used to describe the performance of a classification
model (or “classifier”) on a set of test data for which the true values are known.
@return confusion_matrix => num_classesxnum_classes martix
where confusion_matrix[i,j] = number of prediction which are i and number of ground truth value equal j
"""
def train(self,X_train,y_train):
"""
Given your training data, learn the parameters of your classifier
@param X_train => NxD tensor. Where N is the number of samples and D is the dimension.
it is the data on which your classifier will be trained.
It can be any combination of features provided above.
@param y_train => N vector. Ground truth label
@return Nothing
"""
def validate(self,X_validate,y_validate):
"""
How good is the classifier on unseen data? Use the function below to calculate different metrics.
Based on these matrix change the hyperparmeters and judge the classification
@param X_validate => NxD tensor. Where N is the number of samples and D is the dimension.
it is the data on which your classifier validated.
It can be any combination of features provided above.
@param y_validate => N vector. Ground truth label
"""
# Create a confusion matrix
# Calculate Validation accuracy
# Calculate precision and recall
# Calculate F1-score
return
# +
# Create a train and validation split to train your classifier
# +
# Create 3 tables simiar to page-6 of the paper. One table per dataset
# Each table will have 5 columns.
# Feature/combination of feature used, reduced dimension space, classification error, accuracy, f1-score
# Print the table. (You can use Pandas)
# +
# For each dataset print the confusion matrix for the best model
# -
# 3. Similiar to 1(b) use t-SNE based visilization of faces? Does it makesense? Do you see similar people coming together?or something else? Can you do visualization datasetwise and combined? Here you will use a popular implementation.(Worth reading and understanding t-SNE. We will not discuss it in the class and out of scope for thiscourse/exams.
# +
# Compute TSNE for different features and create a scatter plot
X = # feature
k = 3 # Number of components in TSNE
# Compute
X_TSNE = TSNE(n_components=k).fit_transform(X)
# Plot the representation in 2d/3d
# -
# 4.`face` is used for verification.
#
# 4(a) How do we formulate the problem using KNN
#
# 4(b) How do we analyze the performance ? suggest the metrics (like accuracy) that is appropriate for this task.
#
# _______________________________________________________________________
#
# 4(c)Show empirical re-sults with all the representations
class FaceVerification():
def __init__():
super.__init__()
# Define your parameters eg, W,b, max_iterations etc.
def verify(self,X,class_id):
"""
Given an input X find if the class id is correct or not.
@return verfication_results => N vector containing True or False.
If the class-id matches with your prediction then true else false.
"""
return verfication_results
def train(self,X_train,y_train):
"""
Given your training data, learn the parameters of your classifier
@param X_train => NxD tensor. Where N is the number of samples and D is the dimension.
it is the data on which your verification system will be trained.
It can be any combination of features provided above.
@param y_train => N vector. Ground truth label
@return Nothing
"""
def validate(self,X_validate,y_validate):
"""
How good is your system on unseen data? Use the function below to calculate different metrics.
Based on these matrix change the hyperparmeters
@param X_validate => NxD tensor. Where N is the number of samples and D is the dimension.
It can be any combination of features provided above.
@param y_validate => N vector. Ground truth label
"""
return
# +
# Create a train and validation split and show your results
# +
# Create 3 tables simiar to page-6 of the paper. One table per dataset
# Each table will have 5 columns.
# Feature/combination of feature used, reduced dimension space, verification error, accuracy, precision
# Print the table. (You can use Pandas)
# -
# ### Extenstion / Application
# Create a system for any one of the following problems:
#
# - Politicians vs Filmstars in a public data set. (eg.LFW)
# You already have seen IIIT-CFW dataset. Use it for classification.
# - Age prediction
# Given different actors/actress in IMFDB create new labels based on their age.
# - Gender prediction
# Given different actors/actress in IMFDB+IIIT-CFW create new labels based on their gender.
# - Emotion classification
# Both the yale dataset and IMFDB contain an `emotion.txt` file. Using that you can create a emotion predicter
# - cartoon vs real images
# Use a combination of IIIT-CFW and other dataset.
#
#
#
# You are free to use a new dataset that is publicly avail-able or even create one by crawling from internet.
# +
# Load data
# +
# Define your features
# +
# Create your classifier
# Validate your classifier
# +
# Show qualitative results such as accuracy, k-fold validation, TSNE/PCA/Isomap plots, etc.
# +
# Show quantitative results such as examples of correct prediction and wrong prediction
| Assignment2/201XXXX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import xml.etree.ElementTree as ET
# from lxml import etree
import re
from tqdm import tqdm
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# -
# load the supp2021
df = pd.read_csv('supp_names_ui_refer_ui.csv')
df
supp_list = df['supp_names'].tolist()
supp_list
# +
pattern = '[A-Z][A-Z]'
acro = []
for name in supp_list:
if re.findall(pattern, name):
acro.append(name)
acro
# -
len(acro)
# +
# Make them as new training data set in this format
# {"id": 9, "document_id": "1253656", "ner_tags": ["O", "O", "O", "O", "O", "O", "O", "O", "B-Chemical", "O", "O", "O", "O", "O", "O"], "tokens": ["The", "sperm", "motion", "parameter", "most", "strongly", "associated", "with", "1N", "was", "straight", "-", "line", "velocity", "."], "spans": [[1397, 1400], [1401, 1406], [1407, 1413], [1414, 1423], [1424, 1428], [1429, 1437], [1438, 1448], [1449, 1453], [1454, 1456], [1457, 1460], [1461, 1469], [1469, 1470], [1470, 1474], [1475, 1483], [1483, 1484]]}
# id starts at 18910
# document_id start at 6032168
# span starts at 36022
import json
import nltk
# nltk.download('punkt')
from nltk.tokenize import word_tokenize
i = 18910
document_id = 6032168
span_0 = 36022
def tokenize(name):
if '-' in name:
t = name.split('-')
tokens = ['-'] * (len(t) * 2 - 1)
tokens[0::2] = t
res = []
for token in tokens:
res += word_tokenize(token)
tokens = res
else:
tokens = word_tokenize(name)
return tokens
def get_spans(tokens, span_0):
spans = []
for t in tokens:
if t not in [',', '-']:
spans += [[span_0, span_0 + len(t)]]
span_0 = span_0 + len(t) + 1
else:
spans += [[span_0 - 1, span_0]]
span_0 += 1
return spans
acro_list = []
for name in acro:
dictionary = {}
dictionary["id"] = i
dictionary['document_id'] = document_id
dictionary['ner_tags'] = ["B-Chemical" if i == 0 else "I-Chemical" for i, t in enumerate(tokenize(name))]
dictionary['tokens'] = tokenize(name)
dictionary['spans'] = get_spans(dictionary['tokens'], span_0)
i += 1
document_id += 1
span_0 = dictionary['spans'][-1][-1] + 1
acro_list.append(dictionary)
# acro_list
# -
filler = {"id": 79998, "document_id": 6093256, "ner_tags": ["O", "O", "O"], "tokens": ["Materials", "and", "Methods"], "spans": [[1463359, 1463368], [1463369, 1463372], [1463373, 1463380]]}
acro_list.append(filler)
len(acro_list)
with open('add_train_1013.json', 'w') as f:
# f.write(json.dumps(i) for i in acro_list[:10] + '\n')
for d in acro_list:
f.write(json.dumps(d))
f.write('\n')
# +
import nltk
nltk.download('punkt')
from nltk.tokenize import word_tokenize
word_tokenize(acro[4])
# -
import spacy
import en_core_web_sm
# +
nlp = en_core_web_sm.load()
nlp(acro[4])
# +
acro[:100]
# do not split '.' as there is not period in this set of data.
for name in acro[:100]:
if '-' in name:
t = name.split('-')
tokens = ['-'] * (len(t) * 2 - 1)
tokens[0::2] = t
res = []
for token in tokens:
res += word_tokenize(token)
tokens = res
else:
tokens = word_tokenize(name)
tokens
# -
tokens
# +
spans = []
span_0 = 36022
for t in tokens:
if t not in [',', '-']:
spans += [[span_0, span_0 + len(t)]]
span_0 = span_0 + len(t) + 1
else:
spans += [[span_0 - 1, span_0]]
span_0 += 1
spans
# -
filler = {"id": 79998, "document_id": 6093256, "ner_tags": ["O", "O", "O"], "tokens": ["Materials", "and", "Methods"], "spans": [[1463359, 1463368], [1463369, 1463372], [1463373, 1463380]]}
# ### Reduce data to drugs with consecutive 4 uppercase letters
# +
pattern_4 = '[A-Z][A-Z][A-Z][A-Z]'
acro_4 = []
for name in supp_list:
if re.findall(pattern_4, name):
acro_4.append(name)
acro_4
# -
len(acro_4)
# +
# Make them as new training data set in this format
# {"id": 9, "document_id": "1253656", "ner_tags": ["O", "O", "O", "O", "O", "O", "O", "O", "B-Chemical", "O", "O", "O", "O", "O", "O"], "tokens": ["The", "sperm", "motion", "parameter", "most", "strongly", "associated", "with", "1N", "was", "straight", "-", "line", "velocity", "."], "spans": [[1397, 1400], [1401, 1406], [1407, 1413], [1414, 1423], [1424, 1428], [1429, 1437], [1438, 1448], [1449, 1453], [1454, 1456], [1457, 1460], [1461, 1469], [1469, 1470], [1470, 1474], [1475, 1483], [1483, 1484]]}
# id starts at 18910
# document_id start at 6032168
# span starts at 36022
import json
import nltk
# nltk.download('punkt')
from nltk.tokenize import word_tokenize
i = 18910
document_id = 6032168
span_0 = 36022
def tokenize(name):
if '-' in name:
t = name.split('-')
tokens = ['-'] * (len(t) * 2 - 1)
tokens[0::2] = t
res = []
for token in tokens:
res += word_tokenize(token)
tokens = res
else:
tokens = word_tokenize(name)
return tokens
def get_spans(tokens, span_0):
spans = []
for t in tokens:
if t not in [',', '-']:
spans += [[span_0, span_0 + len(t)]]
span_0 = span_0 + len(t) + 1
else:
spans += [[span_0 - 1, span_0]]
span_0 += 1
return spans
acro_list = []
for name in acro_4:
dictionary = {}
dictionary["id"] = i
dictionary['document_id'] = document_id
dictionary['ner_tags'] = ["B-Chemical" if i == 0 else "I-Chemical" for i, t in enumerate(tokenize(name))]
dictionary['tokens'] = tokenize(name)
dictionary['spans'] = get_spans(dictionary['tokens'], span_0)
i += 1
document_id += 1
span_0 = dictionary['spans'][-1][-1] + 1
acro_list.append(dictionary)
# acro_list
# -
acro_list[-1]
acro_list.append({"id": 36781, "document_id": 6050038, "ner_tags": ["O", "O", "O"], "tokens": ["Materials", "and", "Methods"], "spans": [[449369, 449378], [449379, 449382], [449383, 449390]]})
len(acro_list)
with open('add_train_1015.json', 'w') as f:
# f.write(json.dumps(i) for i in acro_list[:10] + '\n')
for d in acro_list:
f.write(json.dumps(d))
f.write('\n')
| NER/Acronyms-from-supp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Linear classifiers demo: `fit`
#
# CPSC 340: Machine Learning and Data Mining
#
# The University of British Columbia
#
# 2017 Winter Term 2
#
# <NAME>
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from plot_classifier import plot_loss_diagram, plot_classifier
from sklearn.svm import SVC
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
# %autosave 0
# -
# ### Loss function diagrams and `fit`
# - Below is the 0-1 loss, which, when added over examples, give the number of classification errors.
# - This is intuitive but hard to minimize.
# - Note that we're being clever with the fact that $y_i\in\{-1,+1\}$ when we write $y_iw^Tx_i$.
plot_loss_diagram()
grid = np.linspace(-2,2,1000)
plt.plot(grid, grid<0, color='black', linewidth=3, label="0-1")
plt.legend(loc="best", fontsize=13);
# Below is the squared error loss:
plot_loss_diagram()
grid = np.linspace(-2,2,1000)
plt.plot(grid, (grid-1)**2, color='brown', linewidth=3, label="squared")
plt.legend(loc="best", fontsize=13);
# - Below are two convex approximations to the 0-1 loss which are used a lot in practice.
# - Logistic loss: $\log(1+\exp(-y_iw^Tx_i))$
# - Hinge loss: $\max(0, 1-y_iw^Tx_i)$
# - The subtle difference between them has important implications.
plot_loss_diagram()
plt.plot(grid, grid<0, color='black', linewidth=3, label="0-1")
plt.plot(grid, np.log(1+np.exp(-grid)), color='green', linewidth=3, label="logistic")
plt.plot(grid, np.maximum(0,1-grid), color='orange', linewidth=3, label="hinge")
plt.legend(loc="best", fontsize=13);
# ### Support vectors
# - Support vectors occur when $y_iw^Tx_i < 1$.
# - If something is not a support vector, then its loss is zero and it "doesn't contribute".
n = 20
X = np.random.randn(n,2)
y = np.random.choice((-1,+1),size=n)
X[y>0,0] -= 2
X[y>0,1] += 2
# +
svm = SVC(kernel="linear", C=1e6)
svm.fit(X,y)
plt.figure()
plot_classifier(X,y,svm, ax=plt.gca())
plt.scatter(*svm.support_vectors_.T, marker="o", edgecolor="yellow", facecolor="none", s=120);
# -
# - The support vectors are shown in yellow.
# - These are the example that "support" the boundary. Let's try removing one.
sv = svm.support_
not_sv = list(set(range(n)) - set(sv))
# +
# remove all non-support vectors
X3 = np.delete(X,not_sv,0)
y3 = np.delete(y,not_sv,0)
svm3 = SVC(kernel="linear", C=1e6)
svm3.fit(X3,y3)
plot_classifier(X3,y3,svm3, ax=plt.gca())
# plt.scatter(*svm3.support_vectors_.T, marker="o", edgecolor="yellow", facecolor="none", s=120);
print(svm.coef_)
print(svm3.coef_)
# +
# remove a support vector
X2 = np.delete(X,sv[0],0)
y2 = np.delete(y,sv[0],0)
svm2 = SVC(kernel="linear", C=1e6)
svm2.fit(X2,y2)
plt.figure()
plot_classifier(X2,y2,svm2, ax=plt.gca())
plt.scatter(*svm2.support_vectors_.T, marker="o", edgecolor="yellow", facecolor="none", s=120);
plt.scatter(svm.support_vectors_[0,0], svm.support_vectors_[0,1], marker="x", c="yellow")
print(svm.coef_)
print(svm2.coef_)
# -
# --------
#
# The rest is for fun, if we have time...
#
# --------
# ### Max margin
# this data set should be linearly separable
X = np.random.randn(n,2)
y = np.random.choice((-1,+1),size=n)
X[y>0] += 5
lr = LogisticRegression()
lr.fit(X,y)
plot_classifier(X,y,lr);
svm = SVC(kernel="linear", C=1)
svm.fit(X,y)
plot_classifier(X, y, svm);
plt.scatter(*svm.support_vectors_.T, marker="o", edgecolor="yellow", facecolor="none", s=120);
# - Above: it looks like SVM maximizes the margin whereas logistic regression doesn't.
# - Not sure how meaningful this
# - Also, logistic regression will stop making errors for sufficiently large `C`.
# - (If you want, you could also think of KNN with $k=1$ as maximizing its margin, since the boundary will always be half way between examples of different labels)
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X,y)
plot_classifier(X, y, knn);
# ### Probabilities
#
# - It is not particularly natural to get probabilities out of SVMs, although one could devise ways of doing so.
# ### Regularization
#
# - Larger $\lambda$ makes the coefficients smaller as usual
# - I don't have an elegent intuitive iterpretation
# - But let's revisit this next week when we get to kernels
# ### Preview of upcoming lectures
#
# - Friday: multi-class classification with linear classifiers
# - Monday: kernel methods
# +
svm = SVC() # RBF kernel by default
svm.fit(X,y)
plot_classifier(X,y,svm);
# -
| lectures/L19demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/HadenMoore/DS-Unit-1-Sprint-2-Data-Wrangling-and-Storytelling/blob/master/DS_Unit_1_Sprint_Challenge_2_Data_Wrangling_and_Storytelling_(2).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="4yMHi_PX9hEz"
# # Data Science Unit 1 Sprint Challenge 2
#
# ## Data Wrangling and Storytelling
#
# Taming data from its raw form into informative insights and stories.
# + [markdown] id="9wIvtOss9H_i" colab_type="text"
# ## Data Wrangling
#
# In this Sprint Challenge you will first "wrangle" some data from [Gapminder](https://www.gapminder.org/about-gapminder/), a Swedish non-profit co-founded by <NAME>. "Gapminder produces free teaching resources making the world understandable based on reliable statistics."
# - [Cell phones (total), by country and year](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--cell_phones_total--by--geo--time.csv)
# - [Population (total), by country and year](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv)
# - [Geo country codes](https://github.com/open-numbers/ddf--gapminder--systema_globalis/blob/master/ddf--entities--geo--country.csv)
#
# These two links have everything you need to successfully complete the first part of this sprint challenge.
# - [Pandas documentation: Working with Text Data](https://pandas.pydata.org/pandas-docs/stable/text.html) (one question)
# - [Pandas Cheat Sheet](https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf) (everything else)
# + [markdown] colab_type="text" id="wWEU2GemX68A"
# ### Part 0. Load data
#
# You don't need to add or change anything here. Just run this cell and it loads the data for you, into three dataframes.
# + colab_type="code" id="bxKtSi5sRQOl" colab={}
import pandas as pd
cell_phones = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--cell_phones_total--by--geo--time.csv')
population = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv')
geo_country_codes = (pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv')
.rename(columns={'country': 'geo', 'name': 'country'}))
# + [markdown] colab_type="text" id="AZmVTeCsX9RC"
# ### Part 1. Join data
# + [markdown] colab_type="text" id="GLzX58u4SfEy"
# First, join the `cell_phones` and `population` dataframes (with an inner join on `geo` and `time`).
#
# The resulting dataframe's shape should be: (8590, 4)
# + colab_type="code" id="GVV7Hnj4SXBa" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="4f700b8d-ab19-4854-b553-c0f4565a6152"
# Looking at its head
cell_phones.head()
# + id="ABJMoMvjxPf0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b7c09400-3161-4ca3-fff6-d9966f88da1c"
# Looking at the shape of the dataframe
cell_phones.shape
# + id="XaMv-SVcx4D_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="a44b0080-1414-4dbc-aa96-99fa9c4ee6ee"
#Checking for NaNs
cell_phones.isnull().sum()
# + id="YbZNV2TUxGR5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="1c842b75-43e7-46ad-c7e5-ae7efd851fd7"
# Looking at its head
population.head()
# + id="38coxPTxxgDg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b0b6cd35-05a6-4a44-df32-8d4850722eb5"
# Looking at the shape of the dataframe
population.shape
# + id="JXA3O5_cyGkO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="385135b9-42b3-4bab-f65c-42cce7b9f993"
#Checking for NaNs
population.isnull().sum()
# + id="1fj5wIAszseT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="7ade4940-f8a8-4eb8-ed44-9e56d23bec5c"
#inner joining on Geo and Time
merge1 = pd.merge(cell_phones, population, how='inner', on=['geo','time'])
print(merge1)
# + id="ZurXqs221VaA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="72178927-078f-49f0-c0bf-3e8d3bb4c983"
# I'm just saying, I freaking did this. I'm so proud of myself
# This is to check the final shape after the inner merge
merge1.shape
# + [markdown] colab_type="text" id="xsXpDbwwW241"
# Then, select the `geo` and `country` columns from the `geo_country_codes` dataframe, and join with your population and cell phone data.
#
# The resulting dataframe's shape should be: (8590, 5)
# + id="JPYMjqil8p4d" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 258} outputId="bd3f8a73-f0ed-4880-cb59-31fba14e53c3"
#Looking at columns
geo_country_codes.head()
# + colab_type="code" id="Q2LaZta_W2CE" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="830643db-acca-4aca-cee2-35af2d585de3"
# Checking Shape of DataFrame
geo_country_codes.shape
# + id="MSSI5g5q9tBr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="cd4eb7c6-8d0f-41bb-8bef-e9f4a00c490b"
#Checking Value Counts of Country Column
geo_country_codes['country'].value_counts()
# + id="2ujdPuN2_FAO" colab_type="code" colab={}
#Creating variable to hold country and geo columns
column = ['geo', 'country']
# + id="apZ3IVPTAOi8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="23e89dfa-2675-4ce3-9729-6a975d5ffb9e"
final = pd.merge(merge1, geo_country_codes[column], how='inner', on='geo')
print(final)
# + id="S0WqIE4kBB1w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ba6eb903-3763-4c22-9566-40437b2a3299"
#Answer
final.shape
# + [markdown] colab_type="text" id="oK96Uj7vYjFX"
# ### Part 2. Make features
# + [markdown] colab_type="text" id="AD2fBNrOYzCG"
# Calculate the number of cell phones per person, and add this column onto your dataframe.
#
# (You've calculated correctly if you get 1.220 cell phones per person in the United States in 2017.)
# + id="YJ25cw9wIYEt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ab41caca-bb87-41fa-c36f-e2a0ebf98efc"
# creating function to calculate phones per person
phones_per_person =final['cell_phones_total'] / final['population_total']
print(phones_per_person)
# + id="-_842i-1Kxpi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="13ab663d-8ad3-43ff-9e06-2791133486ef"
final['phones_per_person']= phones_per_person
print(final)
# + colab_type="code" id="wXI9nQthYnFK" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="433f7629-490e-4dd2-a05c-93cdd06f2e9d"
usa = final[final.country=='United States']
usa.head()
# + id="DtHrL-agMM6H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="62784e37-4ffa-4dcd-c597-12c4239b1ee7"
answer = usa[usa.time.isin([2017])]
print(answer)
# + [markdown] colab_type="text" id="S3QFdsnRZMH6"
# Modify the `geo` column to make the geo codes uppercase instead of lowercase.
# + colab_type="code" id="93ADij8_YkOq" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="67a1c9d3-dad6-429f-b98f-ff2c0c48e30a"
final['geo'] = final['geo'].str.upper()
final.head()
# + [markdown] colab_type="text" id="hlPDAFCfaF6C"
# ### Part 3. Process data
# + [markdown] colab_type="text" id="k-pudNWve2SQ"
# Use the describe function, to describe your dataframe's numeric columns, and then its non-numeric columns.
#
# (You'll see the time period ranges from 1960 to 2017, and there are 195 unique countries represented.)
# + colab_type="code" id="g26yemKre2Cu" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="ab3b7c59-175b-4b97-8e6f-3780b3ca9147"
import numpy as np
final.describe(include='all')
# + [markdown] colab_type="text" id="zALg-RrYaLcI"
# In 2017, what were the top 5 countries with the most cell phones total?
#
# Your list of countries should have these totals:
#
# | country | cell phones total |
# |:-------:|:-----------------:|
# | ? | 1,474,097,000 |
# | ? | 1,168,902,277 |
# | ? | 458,923,202 |
# | ? | 395,881,000 |
# | ? | 236,488,548 |
#
#
# + colab_type="code" id="JdlWvezHaZxD" colab={}
# This optional code formats float numbers with comma separators
pd.options.display.float_format = '{:,}'.format
# + id="8XgSphwrpB6e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="05b769cf-6a0d-441d-eea7-6de945c1008b"
final[final.time ==2017].sort_values('cell_phones_total', ascending=False).head(5)
# + [markdown] colab_type="text" id="03V3Wln_h0dj"
# 2017 was the first year that China had more cell phones than people.
#
# What was the first year that the USA had more cell phones than people?
# + colab_type="code" id="KONQkQZ3haNC" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ac064bea-5131-4d5d-da55-18bb27c81d8b"
usa = final[final.country=='United States']
usa[['time','cell_phones_total','population_total']]
# + id="t_EvXJs_dhPf" colab_type="code" colab={}
#Answer is 2014
# + [markdown] colab_type="text" id="6J7iwMnTg8KZ"
# ### Part 4. Reshape data
# + [markdown] colab_type="text" id="LP9InazRkUxG"
# *This part is not needed to pass the sprint challenge, only to get a 3! Only work on this after completing the other sections.*
#
# Create a pivot table:
# - Columns: Years 2007—2017
# - Rows: China, India, United States, Indonesia, Brazil (order doesn't matter)
# - Values: Cell Phones Total
#
# The table's shape should be: (5, 11)
# + id="Dlat5NW65pFO" colab_type="code" colab={}
import seaborn as sns
import matplotlib.pyplot as plt
# + id="7FVsKEnUxDWY" colab_type="code" colab={}
countries = ('China','India','United States','Indonesia','Brazil')
years = ['2007', '2008', '2009', '2010', '2011', '2012', '2013', '2014', '2015','2016','2017']
# + id="RU9H-bI639dP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 307} outputId="ff04a053-1c19-4f95-c978-6327a6a1815f"
final.pivot_table(columns='years', index=countries, values='cell_phones_total').plot
# + [markdown] colab_type="text" id="CNKTu2DCnAo6"
# Sort these 5 countries, by biggest increase in cell phones from 2007 to 2017.
#
# Which country had 935,282,277 more cell phones in 2017 versus 2007?
# + colab_type="code" id="O4Aecv1fmQlj" colab={}
# + [markdown] colab_type="text" id="7iHkMsa3Rorh"
# If you have the time and curiosity, what other questions can you ask and answer with this data?
# + [markdown] id="vtcAJOAV9k3X" colab_type="text"
# ## Data Storytelling
#
# In this part of the sprint challenge you'll work with a dataset from **FiveThirtyEight's article, [Every Guest Jon Stewart Ever Had On ‘The Daily Show’](https://fivethirtyeight.com/features/every-guest-jon-stewart-ever-had-on-the-daily-show/)**!
# + [markdown] id="UtjoIqvm9yFg" colab_type="text"
# ### Part 0 — Run this starter code
#
# You don't need to add or change anything here. Just run this cell and it loads the data for you, into a dataframe named `df`.
#
# (You can explore the data if you want, but it's not required to pass the Sprint Challenge.)
# + id="tYujbhIz9zKU" colab_type="code" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
url = 'https://raw.githubusercontent.com/fivethirtyeight/data/master/daily-show-guests/daily_show_guests.csv'
df = pd.read_csv(url).rename(columns={'YEAR': 'Year', 'Raw_Guest_List': 'Guest'})
def get_occupation(group):
if group in ['Acting', 'Comedy', 'Musician']:
return 'Acting, Comedy & Music'
elif group in ['Media', 'media']:
return 'Media'
elif group in ['Government', 'Politician', 'Political Aide']:
return 'Government and Politics'
else:
return 'Other'
df['Occupation'] = df['Group'].apply(get_occupation)
# + [markdown] id="5hjnMK3j90Rp" colab_type="text"
# ### Part 1 — What's the breakdown of guests’ occupations per year?
#
# For example, in 1999, what percentage of guests were actors, comedians, or musicians? What percentage were in the media? What percentage were in politics? What percentage were from another occupation?
#
# Then, what about in 2000? In 2001? And so on, up through 2015.
#
# So, **for each year of _The Daily Show_, calculate the percentage of guests from each occupation:**
# - Acting, Comedy & Music
# - Government and Politics
# - Media
# - Other
#
# #### Hints:
# You can make a crosstab. (See pandas documentation for examples, explanation, and parameters.)
#
# You'll know you've calculated correctly when the percentage of "Acting, Comedy & Music" guests is 90.36% in 1999, and 45% in 2015.
# + id="EbobyiHv916F" colab_type="code" colab={}
df.head()
# + id="S40AwYTOACWC" colab_type="code" colab={}
# Creating a crosstab to view the data better
pd.crosstab(df['Occupation'], df['Year'])
# + id="-KPglf7WDdQs" colab_type="code" colab={}
pd.crosstab(df['Year'], df['Occupation'], normalize='index')
# + [markdown] id="Kiq56dZb92LY" colab_type="text"
# ### Part 2 — Recreate this explanatory visualization:
# + id="HKLDMWwP98vz" colab_type="code" colab={}
from IPython.display import display, Image
png = 'https://fivethirtyeight.com/wp-content/uploads/2015/08/hickey-datalab-dailyshow.png'
example = Image(png, width=500)
display(example)
# + [markdown] id="TK5fDIag9-F6" colab_type="text"
# **Hints:**
# - You can choose any Python visualization library you want. I've verified the plot can be reproduced with matplotlib, pandas plot, or seaborn. I assume other libraries like altair or plotly would work too.
# - If you choose to use seaborn, you may want to upgrade the version to 0.9.0.
#
# **Expectations:** Your plot should include:
# - 3 lines visualizing "occupation of guests, by year." The shapes of the lines should look roughly identical to 538's example. Each line should be a different color. (But you don't need to use the _same_ colors as 538.)
# - Legend or labels for the lines. (But you don't need each label positioned next to its line or colored like 538.)
# - Title in the upper left: _"Who Got To Be On 'The Daily Show'?"_ with more visual emphasis than the subtitle. (Bolder and/or larger font.)
# - Subtitle underneath the title: _"Occupation of guests, by year"_
#
# **Optional Bonus Challenge:**
# - Give your plot polished aesthetics, with improved resemblance to the 538 example.
# - Any visual element not specifically mentioned in the expectations is an optional bonus.
# + id="yyxGDcYyIllT" colab_type="code" colab={}
ct = pd.crosstab(df['Year'], df['Occupation'], normalize='index')
ct
# + id="ERWK5mbBJzPh" colab_type="code" colab={}
ct.plot(kind='bar', legend=True);
| DS_Unit_1_Sprint_Challenge_2_Data_Wrangling_and_Storytelling_(2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2 散点图Scatterplot
# 散点图能够显示2个维度上2组数据的值。每个点代表一个观察点。X(水平)和Y(垂直)轴上的位置表示变量的值。研究这两个变量之间的关系是非常有用的。在seaborn中通过regplot和lmplot制作散点图,regplot和lmplot核心功能相近,regplot相对简单点,如果要定制图像更深层次功能,需要使用lmplot。此外也用Pairplot制作多变量图。该章节主要内容有:
# 1. 基础散点图绘制 Basic scatterplot
# 2. 更改标记参数 Control marker features
# 3. 自定义线性回归拟合 Custom linear regression fit
# 4. 使用分类变量为散点图着色 Use categorical variable to color scatterplot
# 5. 坐标轴范围设置 Control axis limits of plot
# 6. 在散点图上添加文本注释 Add text annotation on scatterplot
# 7. 自定义相关图 Custom correlogram
#调用seaborn
import seaborn as sns
#调用seaborn自带数据集
df = sns.load_dataset('iris')
#显示数据集
df.head()
# ## 1.基础散点图绘制 Basic scatterplot
# 使用regplot()函数制作散点图。您必须提供至少2个列表:X轴和Y轴上的点的位置。
# 默认情况下绘制线性回归拟合直线,可以使用fit_reg = False将其删除
# use the function regplot to make a scatterplot 有回归曲线
# scipy<1.2会有warning
sns.regplot(x=df["sepal_length"], y=df["sepal_width"]);
# Without regression fit 无回归曲线
sns.regplot(x=df["sepal_length"], y=df["sepal_width"], fit_reg=False);
# ## 2. 更改标记参数 Control marker features
# 可以散点图自定义颜色,透明度,形状和大小
# Change shape of marker控制散点的形状
sns.regplot(x=df["sepal_length"], y=df["sepal_width"], marker="+", fit_reg=False);
# List of available shapes 可用的形状查看
import matplotlib
all_shapes=matplotlib.markers.MarkerStyle.markers.keys()
all_shapes
# More marker customization,更具scatter_kws参数控制颜色,透明度,点的大小
sns.regplot(x=df["sepal_length"], y=df["sepal_width"], fit_reg=False, scatter_kws={"color":"darkred","alpha":0.3,"s":20});
# ## 3. 自定义线性回归拟合 Custom linear regression fit
# 您可以自定义seaborn提出的回归拟合的外观。在此示例中,颜色,透明度和宽度通过line_kws = {}选项进行控制。
sns.regplot(x=df["sepal_length"], y=df["sepal_width"], line_kws={"color":"r","alpha":0.7,"lw":5});
# ## 4. 使用分类变量为散点图着色 Use categorical variable to color scatterplot
# + 每组映射一种颜色 Map a color per group
# + 每组映射一个标记 Map a marker per group
# + 使用其他调色板 Use another palette
# + 控制每组的颜色 Control color of each group
# 每组映射一种颜色 Map a color per group
# Use the 'hue' argument to provide a factor variable hue设置species不同种类的的颜色
sns.lmplot( x="sepal_length", y="sepal_width", data=df, fit_reg=False, hue='species', legend=False);
# Move the legend to an empty part of the plot 需要通过matplotlib更改legend的位置
import matplotlib.pyplot as plt
plt.legend(loc='best');
# 每组映射一个标记 Map a marker per group
# give a list to the marker argument 通过hue设定颜色,markes设定各点的形状
sns.lmplot( x="sepal_length", y="sepal_width", data=df, fit_reg=False, hue='species', legend=False, markers=["o", "x", "1"])
# Move the legend to an empty part of the plot
plt.legend(loc='lower right');
# 使用其他调色板 Use another palette
# Use the 'palette' argument 不同种类设定不同的颜色,颜色类型使用palette设定调色板颜色
sns.lmplot( x="sepal_length", y="sepal_width", data=df, fit_reg=False, hue='species', legend=False, palette="Set2")
# Move the legend to an empty part of the plot
plt.legend(loc='lower right');
# 控制每组的颜色 Control color of each group
# Provide a dictionary to the palette argument 调色盘使用自定义颜色
dict_color=dict(setosa="#9b59b6", virginica="#3498db", versicolor="#95a5a6")
sns.lmplot( x="sepal_length", y="sepal_width", data=df, fit_reg=False, hue='species', legend=False, palette=dict_color)
# Move the legend to an empty part of the plot
plt.legend(loc='lower right');
# ## 5. 坐标轴范围设置 Control axis limits of plot
# basic scatterplot
sns.lmplot( x="sepal_length", y="sepal_width", data=df, fit_reg=False)
# control x and y limits 设置轴的范围,不过需要调用matplotlib.pyplot 模块,通常都是matplotlib和seaborn一起用
plt.ylim(0, 20)
plt.xlim(0, None)
# ## 6. 在散点图上添加文本注释 Add text annotation on scatterplot
# + 添加一个注释 Add one annotation
# + 添加多个注释 Use a loop to annotate each marker
# 添加一个注释 Add one annotation
import pandas as pd
# 制作数据集
df_test = pd.DataFrame({
'x': [1, 1.5, 3, 4, 5],
'y': [5, 15, 5, 10, 2],
'group': ['A','other group','B','C','D']})
# 画散点图
p1=sns.regplot(data=df_test, x="x", y="y", fit_reg=False, marker="o", color="skyblue", scatter_kws={'s':400});
# 添加注释
p1.text(3+0.2, 4.5, "An annotation", horizontalalignment='left', size='medium', color='black', weight='semibold')
# 添加多个注释 Use a loop to annotate each marker
# basic plot
p1=sns.regplot(data=df_test, x="x", y="y", fit_reg=False, marker="o", color="skyblue", scatter_kws={'s':400})
# add annotations one by one with a loop
for line in range(0,df_test.shape[0]):
p1.text(df_test.x[line]+0.2, df_test.y[line], df_test.group[line], horizontalalignment='left', size='medium', color='black', weight='semibold')
# ## 7. 自定义相关图 Custom correlogram
# + 有回归方程的散点相关图 correlogram with regression
# + 无回归方程的散点相关图 correlogram without regression
# + 在相关图上表示组 Represent groups on correlogram
# + 相关图子图设置 Kind of plot for the diagonal subplots
# + 子图参数设置 parameters adjustment of subplots
# +
# 有回归方程的散点相关图 correlogram with regression
# library & dataset
import matplotlib.pyplot as plt
import seaborn as sns
df = sns.load_dataset('iris')
# with regression 有回归方程的散点相关图
# 正对角线上的图表示数据频次的直方图,其他表示散点图
sns.pairplot(df, kind="reg");
# -
# 无回归方程的散点相关图 correlogram without regression
sns.pairplot(df, kind="scatter");
# 在相关图上表示组 Represent groups on correlogram
# 通过hue设定种类,markers不同种类的点的表示方式
# 对角线为核密度图
sns.pairplot(df, kind="scatter", hue="species", markers=["o", "s", "D"], palette="Set2")
# 在相关图上表示组 Represent groups on correlogram
# you can give other arguments with plot_kws plot_kws更改散点图的参数
sns.pairplot(df, kind="scatter", hue="species",plot_kws=dict(s=80, edgecolor="white", linewidth=3));
# 相关图子图设置 Kind of plot for the diagonal subplots
# diag_kind有auto,hist,kde选项,hist为直方图,kde为散点图
sns.pairplot(df, diag_kind="hist");
# 子图参数设置 parameters adjustment of subplots
# You can custom it as a density plot or histogram so see the related sections 通过diag_kws调整子图参数
sns.pairplot(df, diag_kind="kde", diag_kws=dict(shade=True, bw=.05, vertical=False));
| Seaborn Study/sources/2 散点图SCATTERPLOT.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="hjM_sV_AepYf"
# # 🔪 JAX - The Sharp Bits 🔪
#
# [](https://colab.research.google.com/github/google/jax/blob/master/docs/notebooks/Common_Gotchas_in_JAX.ipynb)
# + [markdown] id="4k5PVzEo2uJO"
# *levskaya@ mattjj@*
#
# When walking about the countryside of [Italy](https://iaml.it/blog/jax-intro), the people will not hesitate to tell you that __JAX__ has _"una anima di pura programmazione funzionale"_.
#
# __JAX__ is a language for __expressing__ and __composing__ __transformations__ of numerical programs. __JAX__ is also able to __compile__ numerical programs for CPU or accelerators (GPU/TPU).
# JAX works great for many numerical and scientific programs, but __only if they are written with certain constraints__ that we describe below.
# + id="GoK_PCxPeYcy"
import numpy as np
from jax import grad, jit
from jax import lax
from jax import random
import jax
import jax.numpy as jnp
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib import rcParams
rcParams['image.interpolation'] = 'nearest'
rcParams['image.cmap'] = 'viridis'
rcParams['axes.grid'] = False
# + [markdown] id="gX8CZU1g2agP"
# ## 🔪 Pure functions
# + [markdown] id="2oHigBkW2dPT"
# JAX transformation and compilation are designed to work only on Python functions that are functionally pure: all the input data is passed through the function parameters, all the results are output through the function results. A pure function will always return the same result if invoked with the same inputs.
#
# Here are some examples of functions that are not functially pure for which JAX behaves differently than the Python interpreter. Note that these behaviors are not guaranteed by the JAX system; the proper way to use JAX is to use it only on functionally pure Python functions.
# + id="A6R-pdcm4u3v" outputId="389605df-a4d5-4d4b-8d74-64e9d5d39456"
def impure_print_side_effect(x):
print("Executing function") # This is a side-effect
return x
# The side-effects appear during the first run
print ("First call: ", jit(impure_print_side_effect)(4.))
# Subsequent runs with parameters of same type and shape may not show the side-effect
# This is because JAX now invokes a cached compilation of the function
print ("Second call: ", jit(impure_print_side_effect)(5.))
# JAX re-runs the Python function when the type or shape of the argument changes
print ("Third call, different type: ", jit(impure_print_side_effect)(jnp.array([5.])))
# + id="-N8GhitI2bhD" outputId="f16ce914-1387-43b4-9b8a-1d6e3b97b11d"
g = 0.
def impure_uses_globals(x):
return x + g
# JAX captures the value of the global during the first run
print ("First call: ", jit(impure_uses_globals)(4.))
g = 10. # Update the global
# Subsequent runs may silently use the cached value of the globals
print ("Second call: ", jit(impure_uses_globals)(5.))
# JAX re-runs the Python function when the type or shape of the argument changes
# This will end up reading the latest value of the global
print ("Third call, different type: ", jit(impure_uses_globals)(jnp.array([4.])))
# + id="RTB6iFgu4DL6" outputId="e93d2a70-1c18-477a-d69d-d09ed556305a"
g = 0.
def impure_saves_global(x):
global g
g = x
return x
# JAX runs once the transformed function with special Traced values for arguments
print ("First call: ", jit(impure_saves_global)(4.))
print ("Saved global: ", g) # Saved global has an internal JAX value
# + [markdown] id="Mlc2pQlp6v-9"
# A Python function can be functionally pure even if it actually uses stateful objects internally, as long as it does not read or write external state:
# + id="TP-Mqf_862C0" outputId="78df2d95-2c6f-41c9-84a9-feda6329e75e"
def pure_uses_internal_state(x):
state = dict(even=0, odd=0)
for i in range(10):
state['even' if i % 2 == 0 else 'odd'] += x
return state['even'] + state['odd']
print(jit(pure_uses_internal_state)(5.))
# -
# It is not recommended to use iterators in any JAX function you want to `jit` or in any control-flow primitive. The reason is that an iterator is a python object which introduces state to retrieve the next element. Therefore, it is incompatible with JAX functional programming model. In the code below, there are some examples of incorrect attempts to use iterators with JAX. Most of them return an error, but some give unexpected results.
# +
import jax.numpy as jnp
import jax.lax as lax
from jax import make_jaxpr
# lax.fori_loop
array = jnp.arange(10)
print(lax.fori_loop(0, 10, lambda i,x: x+array[i], 0)) # expected result 45
iterator = iter(range(10))
print(lax.fori_loop(0, 10, lambda i,x: x+next(iterator), 0)) # unexpected result 0
# lax.scan
def func11(arr, extra):
ones = jnp.ones(arr.shape)
def body(carry, aelems):
ae1, ae2 = aelems
return (carry + ae1 * ae2 + extra, carry)
return lax.scan(body, 0., (arr, ones))
make_jaxpr(func11)(jnp.arange(16), 5.)
# make_jaxpr(func11)(iter(range(16)), 5.) # throws error
# lax.cond
array_operand = jnp.array([0.])
lax.cond(True, lambda x: x+1, lambda x: x-1, array_operand)
iter_operand = iter(range(10))
# lax.cond(True, lambda x: next(x)+1, lambda x: next(x)-1, iter_operand) # throws error
# + [markdown] id="oBdKtkVW8Lha"
# ## 🔪 In-Place Updates
# + [markdown] id="JffAqnEW4JEb"
# In Numpy you're used to doing this:
# + id="om4xV7_84N9j" outputId="733f901e-d433-4dc8-b5bb-0c23bf2b1306"
numpy_array = np.zeros((3,3), dtype=np.float32)
print("original array:")
print(numpy_array)
# In place, mutating update
numpy_array[1, :] = 1.0
print("updated array:")
print(numpy_array)
# + [markdown] id="go3L4x3w4-9p"
# If we try to update a JAX device array in-place, however, we get an __error__! (☉_☉)
# + id="2AxeCufq4wAp" outputId="d5d873db-cee0-49dc-981d-ec852347f7ca" tags=["raises-exception"]
jax_array = jnp.zeros((3,3), dtype=jnp.float32)
# In place update of JAX's array will yield an error!
try:
jax_array[1, :] = 1.0
except Exception as e:
print("Exception {}".format(e))
# + [markdown] id="7mo76sS25Wco"
# __What gives?!__
#
# Allowing mutation of variables in-place makes program analysis and transformation very difficult. JAX requires a pure functional expression of a numerical program.
#
# Instead, JAX offers the _functional_ update functions: [__index_update__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_update.html#jax.ops.index_update), [__index_add__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_add.html#jax.ops.index_add), [__index_min__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_min.html#jax.ops.index_min), [__index_max__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index_max.html#jax.ops.index_max), and the [__index__](https://jax.readthedocs.io/en/latest/_autosummary/jax.ops.index.html#jax.ops.index) helper.
#
# ️⚠️ inside `jit`'d code and `lax.while_loop` or `lax.fori_loop` the __size__ of slices can't be functions of argument _values_ but only functions of argument _shapes_ -- the slice start indices have no such restriction. See the below __Control Flow__ Section for more information on this limitation.
# + id="m5lg1RYq5D9p"
from jax.ops import index, index_add, index_update
# + [markdown] id="X2Xjjvd-l8NL"
# ### index_update
# + [markdown] id="eM6MyndXL2NY"
# If the __input values__ of __index_update__ aren't reused, __jit__-compiled code will perform these operations _in-place_.
# + id="ygUJT49b7BBk" outputId="1a3511c4-a480-472f-cccb-5e01620cbe99"
jax_array = jnp.zeros((3, 3))
print("original array:")
print(jax_array)
new_jax_array = index_update(jax_array, index[1, :], 1.)
print("old array unchanged:")
print(jax_array)
print("new array:")
print(new_jax_array)
# + [markdown] id="7to-sF8EmC_y"
# ### index_add
# + [markdown] id="iI5cLY1xMBLs"
# If the __input values__ of __index_update__ aren't reused, __jit__-compiled code will perform these operations _in-place_.
# + id="tsw2svao8FUp" outputId="874acd15-a493-4d63-efe4-9f440d5d2a12"
print("original array:")
jax_array = jnp.ones((5, 6))
print(jax_array)
new_jax_array = index_add(jax_array, index[::2, 3:], 7.)
print("new array post-addition:")
print(new_jax_array)
# + [markdown] id="oZ_jE2WAypdL"
# ## 🔪 Out-of-Bounds Indexing
# + [markdown] id="btRFwEVzypdN"
# In Numpy, you are used to errors being thrown when you index an array outside of its bounds, like this:
# + id="5_ZM-BJUypdO" outputId="461f38cd-9452-4bcc-a44f-a07ddfa12f42" tags=["raises-exception"]
try:
np.arange(10)[11]
except Exception as e:
print("Exception {}".format(e))
# + [markdown] id="eoXrGARWypdR"
# However, raising an error on other accelerators can be more difficult. Therefore, JAX does not raise an error, instead the index is clamped to the bounds of the array, meaning that for this example the last value of the array will be returned.
# + id="cusaAD0NypdR" outputId="48428ad6-6cde-43ad-c12d-2eb9b9fe59cf"
jnp.arange(10)[11]
# -
# Note that due to this behavior jnp.nanargmin and jnp.nanargmax return -1 for slices consisting of NaNs whereas Numpy would throw an error.
# + [markdown] id="MUycRNh6e50W"
# ## 🔪 Random Numbers
# + [markdown] id="O8vvaVt3MRG2"
# > _If all scientific papers whose results are in doubt because of bad
# > `rand()`s were to disappear from library shelves, there would be a
# > gap on each shelf about as big as your fist._ - Numerical Recipes
# + [markdown] id="Qikt9pPW9L5K"
# ### RNGs and State
# You're used to _stateful_ pseudorandom number generators (PRNGs) from numpy and other libraries, which helpfully hide a lot of details under the hood to give you a ready fountain of pseudorandomness:
# + id="rr9FeP41fynt" outputId="849d84cf-04ad-4e8b-9505-a92f6c0d7a39"
print(np.random.random())
print(np.random.random())
print(np.random.random())
# + [markdown] id="ORMVVGZJgSVi"
# Underneath the hood, numpy uses the [Mersenne Twister](https://en.wikipedia.org/wiki/Mersenne_Twister) PRNG to power its pseudorandom functions. The PRNG has a period of $2^{19937}-1$ and at any point can be described by __624 32bit unsigned ints__ and a __position__ indicating how much of this "entropy" has been used up.
# + id="7Pyp2ajzfPO2"
np.random.seed(0)
rng_state = np.random.get_state()
#print(rng_state)
# --> ('MT19937', array([0, 1, 1812433255, 1900727105, 1208447044,
# 2481403966, 4042607538, 337614300, ... 614 more numbers...,
# 3048484911, 1796872496], dtype=uint32), 624, 0, 0.0)
# + [markdown] id="aJIxHVXCiM6m"
# This pseudorandom state vector is automagically updated behind the scenes every time a random number is needed, "consuming" 2 of the uint32s in the Mersenne twister state vector:
# + id="GAHaDCYafpAF"
_ = np.random.uniform()
rng_state = np.random.get_state()
#print(rng_state)
# --> ('MT19937', array([2443250962, 1093594115, 1878467924,
# ..., 2648828502, 1678096082], dtype=uint32), 2, 0, 0.0)
# Let's exhaust the entropy in this PRNG statevector
for i in range(311):
_ = np.random.uniform()
rng_state = np.random.get_state()
#print(rng_state)
# --> ('MT19937', array([2443250962, 1093594115, 1878467924,
# ..., 2648828502, 1678096082], dtype=uint32), 624, 0, 0.0)
# Next call iterates the RNG state for a new batch of fake "entropy".
_ = np.random.uniform()
rng_state = np.random.get_state()
# print(rng_state)
# --> ('MT19937', array([1499117434, 2949980591, 2242547484,
# 4162027047, 3277342478], dtype=uint32), 2, 0, 0.0)
# + [markdown] id="N_mWnleNogps"
# The problem with magic PRNG state is that it's hard to reason about how it's being used and updated across different threads, processes, and devices, and it's _very easy_ to screw up when the details of entropy production and consumption are hidden from the end user.
#
# The Mersenne Twister PRNG is also known to have a [number](https://cs.stackexchange.com/a/53475) of problems, it has a large 2.5Kb state size, which leads to problematic [initialization issues](https://dl.acm.org/citation.cfm?id=1276928). It [fails](http://www.pcg-random.org/pdf/toms-oneill-pcg-family-v1.02.pdf) modern BigCrush tests, and is generally slow.
# + [markdown] id="Uvq7nV-j4vKK"
# ### JAX PRNG
# + [markdown] id="COjzGBpO4tzL"
# JAX instead implements an _explicit_ PRNG where entropy production and consumption are handled by explicitly passing and iterating PRNG state. JAX uses a modern [Threefry counter-based PRNG](https://github.com/google/jax/blob/master/design_notes/prng.md) that's __splittable__. That is, its design allows us to __fork__ the PRNG state into new PRNGs for use with parallel stochastic generation.
#
# The random state is described by two unsigned-int32s that we call a __key__:
# + id="yPHE7KTWgAWs" outputId="329e7757-2461-434c-a08c-fde80a2d10c9"
from jax import random
key = random.PRNGKey(0)
key
# + [markdown] id="XjYyWYNfq0hW"
# JAX's random functions produce pseudorandom numbers from the PRNG state, but __do not__ change the state!
#
# Reusing the same state will cause __sadness__ and __monotony__, depriving the enduser of __lifegiving chaos__:
# + id="7zUdQMynoE5e" outputId="50617324-b887-42f2-a7ff-2a10f92d876a"
print(random.normal(key, shape=(1,)))
print(key)
# No no no!
print(random.normal(key, shape=(1,)))
print(key)
# + [markdown] id="hQN9van8rJgd"
# Instead, we __split__ the PRNG to get usable __subkeys__ every time we need a new pseudorandom number:
# + id="ASj0_rSzqgGh" outputId="bcc2ed60-2e41-4ef8-e84f-c724654aa198"
print("old key", key)
key, subkey = random.split(key)
normal_pseudorandom = random.normal(subkey, shape=(1,))
print(" \---SPLIT --> new key ", key)
print(" \--> new subkey", subkey, "--> normal", normal_pseudorandom)
# + [markdown] id="tqtFVE4MthO3"
# We propagate the __key__ and make new __subkeys__ whenever we need a new random number:
# + id="jbC34XLor2Ek" outputId="6834a812-7160-4646-ee19-a246f683905a"
print("old key", key)
key, subkey = random.split(key)
normal_pseudorandom = random.normal(subkey, shape=(1,))
print(" \---SPLIT --> new key ", key)
print(" \--> new subkey", subkey, "--> normal", normal_pseudorandom)
# + [markdown] id="0KLYUluz3lN3"
# We can generate more than one __subkey__ at a time:
# + id="lEi08PJ4tfkX" outputId="3bb513de-8d14-4d37-ae57-51d6f5eaa762"
key, *subkeys = random.split(key, 4)
for subkey in subkeys:
print(random.normal(subkey, shape=(1,)))
# + [markdown] id="rg4CpMZ8c3ri"
# ## 🔪 Control Flow
# + [markdown] id="izLTvT24dAq0"
# ### ✔ python control_flow + autodiff ✔
#
# If you just want to apply `grad` to your python functions, you can use regular python control-flow constructs with no problems, as if you were using [Autograd](https://github.com/hips/autograd) (or Pytorch or TF Eager).
# + id="aAx0T3F8lLtu" outputId="808cfa77-d924-4586-af19-35a8fd7d2238"
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
print(grad(f)(2.)) # ok!
print(grad(f)(4.)) # ok!
# + [markdown] id="hIfPT7WMmZ2H"
# ### python control flow + JIT
#
# Using control flow with `jit` is more complicated, and by default it has more constraints.
#
# This works:
# + id="OZ_BJX0CplNC" outputId="48ce004c-536a-44f5-b020-9267825e7e4d"
@jit
def f(x):
for i in range(3):
x = 2 * x
return x
print(f(3))
# + [markdown] id="22RzeJ4QqAuX"
# So does this:
# + id="pinVnmRWp6w6" outputId="e3e6f2f7-ba59-4a98-cdfc-905c91b38ed1"
@jit
def g(x):
y = 0.
for i in range(x.shape[0]):
y = y + x[i]
return y
print(g(jnp.array([1., 2., 3.])))
# + [markdown] id="TStltU2dqf8A"
# But this doesn't, at least by default:
# + id="9z38AIKclRNM" outputId="466730dd-df8b-4b80-ac5e-e55b5ea85ec7"
@jit
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
# This will fail!
try:
f(2)
except Exception as e:
print("Exception {}".format(e))
# + [markdown] id="pIbr4TVPqtDN"
# __What gives!?__
#
# When we `jit`-compile a function, we usually want to compile a version of the function that works for many different argument values, so that we can cache and reuse the compiled code. That way we don't have to re-compile on each function evaluation.
#
# For example, if we evaluate an `@jit` function on the array `jnp.array([1., 2., 3.], jnp.float32)`, we might want to compile code that we can reuse to evaluate the function on `jnp.array([4., 5., 6.], jnp.float32)` to save on compile time.
#
# To get a view of your Python code that is valid for many different argument values, JAX traces it on _abstract values_ that represent sets of possible inputs. There are [multiple different levels of abstraction](https://github.com/google/jax/blob/master/jax/abstract_arrays.py), and different transformations use different abstraction levels.
#
# By default, `jit` traces your code on the `ShapedArray` abstraction level, where each abstract value represents the set of all array values with a fixed shape and dtype. For example, if we trace using the abstract value `ShapedArray((3,), jnp.float32)`, we get a view of the function that can be reused for any concrete value in the corresponding set of arrays. That means we can save on compile time.
#
# But there's a tradeoff here: if we trace a Python function on a `ShapedArray((), jnp.float32)` that isn't committed to a specific concrete value, when we hit a line like `if x < 3`, the expression `x < 3` evaluates to an abstract `ShapedArray((), jnp.bool_)` that represents the set `{True, False}`. When Python attempts to coerce that to a concrete `True` or `False`, we get an error: we don't know which branch to take, and can't continue tracing! The tradeoff is that with higher levels of abstraction we gain a more general view of the Python code (and thus save on re-compilations), but we require more constraints on the Python code to complete the trace.
#
# The good news is that you can control this tradeoff yourself. By having `jit` trace on more refined abstract values, you can relax the traceability constraints. For example, using the `static_argnums` argument to `jit`, we can specify to trace on concrete values of some arguments. Here's that example function again:
# + id="-Tzp0H7Bt1Sn" outputId="aba57a88-d8eb-40b0-ff22-7c266d892b13"
def f(x):
if x < 3:
return 3. * x ** 2
else:
return -4 * x
f = jit(f, static_argnums=(0,))
print(f(2.))
# + [markdown] id="MHm1hIQAvBVs"
# Here's another example, this time involving a loop:
# + id="iwY86_JKvD6b" outputId="1ec847ea-df2b-438d-c0a1-fabf7b93b73d"
def f(x, n):
y = 0.
for i in range(n):
y = y + x[i]
return y
f = jit(f, static_argnums=(1,))
f(jnp.array([2., 3., 4.]), 2)
# + [markdown] id="nSPTOX8DvOeO"
# In effect, the loop gets statically unrolled. JAX can also trace at _higher_ levels of abstraction, like `Unshaped`, but that's not currently the default for any transformation
# + [markdown] id="wWdg8LTYwCW3"
# ️⚠️ **functions with argument-__value__ dependent shapes**
#
# These control-flow issues also come up in a more subtle way: numerical functions we want to __jit__ can't specialize the shapes of internal arrays on argument _values_ (specializing on argument __shapes__ is ok). As a trivial example, let's make a function whose output happens to depend on the input variable `length`.
# + id="Tqe9uLmUI_Gv" outputId="fe319758-9959-434c-ab9d-0926e599dbc0"
def example_fun(length, val):
return jnp.ones((length,)) * val
# un-jit'd works fine
print(example_fun(5, 4))
bad_example_jit = jit(example_fun)
# this will fail:
try:
print(bad_example_jit(10, 4))
except Exception as e:
print("Exception {}".format(e))
# static_argnums tells JAX to recompile on changes at these argument positions:
good_example_jit = jit(example_fun, static_argnums=(0,))
# first compile
print(good_example_jit(10, 4))
# recompiles
print(good_example_jit(5, 4))
# + [markdown] id="MStx_r2oKxpp"
# `static_argnums` can be handy if `length` in our example rarely changes, but it would be disastrous if it changed a lot!
#
# Lastly, if your function has global side-effects, JAX's tracer can cause weird things to happen. A common gotcha is trying to print arrays inside __jit__'d functions:
# + id="m2ABpRd8K094" outputId="64da37a0-aa06-46a3-e975-88c676c5b9fa"
@jit
def f(x):
print(x)
y = 2 * x
print(y)
return y
f(2)
# + [markdown] id="uCDcWG4MnVn-"
# ### Structured control flow primitives
#
# There are more options for control flow in JAX. Say you want to avoid re-compilations but still want to use control flow that's traceable, and that avoids un-rolling large loops. Then you can use these 4 structured control flow primitives:
#
# - `lax.cond` _differentiable_
# - `lax.while_loop` __fwd-mode-differentiable__
# - `lax.fori_loop` __fwd-mode-differentiable__
# - `lax.scan` _differentiable_
# + [markdown] id="Sd9xrLMXeK3A"
# #### cond
# python equivalent:
#
# ```python
# def cond(pred, true_fun, false_fun, operand):
# if pred:
# return true_fun(operand)
# else:
# return false_fun(operand)
# ```
# + id="SGxz9JOWeiyH" outputId="b29da06c-037f-4b05-dbd8-ba52ac35a8cf"
from jax import lax
operand = jnp.array([0.])
lax.cond(True, lambda x: x+1, lambda x: x-1, operand)
# --> array([1.], dtype=float32)
lax.cond(False, lambda x: x+1, lambda x: x-1, operand)
# --> array([-1.], dtype=float32)
# + [markdown] id="xkOFAw24eOMg"
# #### while_loop
#
# python equivalent:
# ```
# def while_loop(cond_fun, body_fun, init_val):
# val = init_val
# while cond_fun(val):
# val = body_fun(val)
# return val
# ```
# + id="jM-D39a-c436" outputId="b9c97167-fecf-4559-9ca7-1cb0235d8ad2"
init_val = 0
cond_fun = lambda x: x<10
body_fun = lambda x: x+1
lax.while_loop(cond_fun, body_fun, init_val)
# --> array(10, dtype=int32)
# + [markdown] id="apo3n3HAeQY_"
# #### fori_loop
# python equivalent:
# ```
# def fori_loop(start, stop, body_fun, init_val):
# val = init_val
# for i in range(start, stop):
# val = body_fun(i, val)
# return val
# ```
# + id="dt3tUpOmeR8u" outputId="864f2959-2429-4666-b364-4baf90a57482"
init_val = 0
start = 0
stop = 10
body_fun = lambda i,x: x+i
lax.fori_loop(start, stop, body_fun, init_val)
# --> array(45, dtype=int32)
# + [markdown] id="SipXS5qiqk8e"
# #### Summary
#
# $$
# \begin{array} {r|rr}
# \hline \
# \textrm{construct}
# & \textrm{jit}
# & \textrm{grad} \\
# \hline \
# \textrm{if} & ❌ & ✔ \\
# \textrm{for} & ✔* & ✔\\
# \textrm{while} & ✔* & ✔\\
# \textrm{lax.cond} & ✔ & ✔\\
# \textrm{lax.while_loop} & ✔ & \textrm{fwd}\\
# \textrm{lax.fori_loop} & ✔ & \textrm{fwd}\\
# \textrm{lax.scan} & ✔ & ✔\\
# \hline
# \end{array}
# $$
# <center>$\ast$ = argument-__value__-independent loop condition - unrolls the loop </center>
# + [markdown] id="DKTMw6tRZyK2"
# ## 🔪 NaNs
# + [markdown] id="ncS0NI4jZrwy"
# ### Debugging NaNs
#
# If you want to trace where NaNs are occurring in your functions or gradients, you can turn on the NaN-checker by:
#
# * setting the `JAX_DEBUG_NANS=True` environment variable;
#
# * adding `from jax.config import config` and `config.update("jax_debug_nans", True)` near the top of your main file;
#
# * adding `from jax.config import config` and `config.parse_flags_with_absl()` to your main file, then set the option using a command-line flag like `--jax_debug_nans=True`;
#
# This will cause computations to error-out immediately on production of a NaN. Switching this option on adds a nan check to every floating point type value produced by XLA. That means values are pulled back to the host and checked as ndarrays for every primitive operation not under an `@jit`. For code under an `@jit`, the output of every `@jit` function is checked and if a nan is present it will re-run the function in de-optimized op-by-op mode, effectively removing one level of `@jit` at a time.
#
# There could be tricky situations that arise, like nans that only occur under a `@jit` but don't get produced in de-optimized mode. In that case you'll see a warning message print out but your code will continue to execute.
#
# If the nans are being produced in the backward pass of a gradient evaluation, when an exception is raised several frames up in the stack trace you will be in the backward_pass function, which is essentially a simple jaxpr interpreter that walks the sequence of primitive operations in reverse. In the example below, we started an ipython repl with the command line `env JAX_DEBUG_NANS=True ipython`, then ran this:
# -
# ```
# In [1]: import jax.numpy as jnp
#
# In [2]: jnp.divide(0., 0.)
# ---------------------------------------------------------------------------
# FloatingPointError Traceback (most recent call last)
# <ipython-input-2-f2e2c413b437> in <module>()
# ----> 1 jnp.divide(0., 0.)
#
# .../jax/jax/numpy/lax_numpy.pyc in divide(x1, x2)
# 343 return floor_divide(x1, x2)
# 344 else:
# --> 345 return true_divide(x1, x2)
# 346
# 347
#
# .../jax/jax/numpy/lax_numpy.pyc in true_divide(x1, x2)
# 332 x1, x2 = _promote_shapes(x1, x2)
# 333 return lax.div(lax.convert_element_type(x1, result_dtype),
# --> 334 lax.convert_element_type(x2, result_dtype))
# 335
# 336
#
# .../jax/jax/lax.pyc in div(x, y)
# 244 def div(x, y):
# 245 r"""Elementwise division: :math:`x \over y`."""
# --> 246 return div_p.bind(x, y)
# 247
# 248 def rem(x, y):
#
# ... stack trace ...
#
# .../jax/jax/interpreters/xla.pyc in handle_result(device_buffer)
# 103 py_val = device_buffer.to_py()
# 104 if np.any(np.isnan(py_val)):
# --> 105 raise FloatingPointError("invalid value")
# 106 else:
# 107 return DeviceArray(device_buffer, *result_shape)
#
# FloatingPointError: invalid value
# ```
# The nan generated was caught. By running `%debug`, we can get a post-mortem debugger. This also works with functions under `@jit`, as the example below shows.
# ```
# In [4]: from jax import jit
#
# In [5]: @jit
# ...: def f(x, y):
# ...: a = x * y
# ...: b = (x + y) / (x - y)
# ...: c = a + 2
# ...: return a + b * c
# ...:
#
# In [6]: x = jnp.array([2., 0.])
#
# In [7]: y = jnp.array([3., 0.])
#
# In [8]: f(x, y)
# Invalid value encountered in the output of a jit function. Calling the de-optimized version.
# ---------------------------------------------------------------------------
# FloatingPointError Traceback (most recent call last)
# <ipython-input-8-811b7ddb3300> in <module>()
# ----> 1 f(x, y)
#
# ... stack trace ...
#
# <ipython-input-5-619b39acbaac> in f(x, y)
# 2 def f(x, y):
# 3 a = x * y
# ----> 4 b = (x + y) / (x - y)
# 5 c = a + 2
# 6 return a + b * c
#
# .../jax/jax/numpy/lax_numpy.pyc in divide(x1, x2)
# 343 return floor_divide(x1, x2)
# 344 else:
# --> 345 return true_divide(x1, x2)
# 346
# 347
#
# .../jax/jax/numpy/lax_numpy.pyc in true_divide(x1, x2)
# 332 x1, x2 = _promote_shapes(x1, x2)
# 333 return lax.div(lax.convert_element_type(x1, result_dtype),
# --> 334 lax.convert_element_type(x2, result_dtype))
# 335
# 336
#
# .../jax/jax/lax.pyc in div(x, y)
# 244 def div(x, y):
# 245 r"""Elementwise division: :math:`x \over y`."""
# --> 246 return div_p.bind(x, y)
# 247
# 248 def rem(x, y):
#
# ... stack trace ...
# ```
# When this code sees a nan in the output of an `@jit` function, it calls into the de-optimized code, so we still get a clear stack trace. And we can run a post-mortem debugger with `%debug` to inspect all the values to figure out the error.
#
# ⚠️ You shouldn't have the NaN-checker on if you're not debugging, as it can introduce lots of device-host round-trips and performance regressions!
# + [markdown] id="YTktlwTTMgFl"
# ## Double (64bit) precision
#
# At the moment, JAX by default enforces single-precision numbers to mitigate the Numpy API's tendency to aggressively promote operands to `double`. This is the desired behavior for many machine-learning applications, but it may catch you by surprise!
# + id="CNNGtzM3NDkO" outputId="d1384021-d9bf-450f-a9ae-82024fa5fc1a"
x = random.uniform(random.PRNGKey(0), (1000,), dtype=jnp.float64)
x.dtype
# + [markdown] id="VcvqzobxNPbd"
# To use double-precision numbers, you need to set the `jax_enable_x64` configuration variable __at startup__.
#
# There are a few ways to do this:
#
# 1. You can enable 64bit mode by setting the environment variable `JAX_ENABLE_X64=True`.
#
# 2. You can manually set the `jax_enable_x64` configuration flag at startup:
#
# ```python
# # again, this only works on startup!
# from jax.config import config
# config.update("jax_enable_x64", True)
# ```
#
# 3. You can parse command-line flags with `absl.app.run(main)`
#
# ```python
# from jax.config import config
# config.config_with_absl()
# ```
#
# 4. If you want JAX to run absl parsing for you, i.e. you don't want to do `absl.app.run(main)`, you can instead use
#
# ```python
# from jax.config import config
# if __name__ == '__main__':
# # calls config.config_with_absl() *and* runs absl parsing
# config.parse_flags_with_absl()
# ```
#
# Note that #2-#4 work for _any_ of JAX's configuration options.
#
# We can then confirm that `x64` mode is enabled:
# + id="HqGbBa9Rr-2g" outputId="cd241d63-3d00-4fd7-f9c0-afc6af01ecf4"
import jax.numpy as jnp
from jax import random
x = random.uniform(random.PRNGKey(0), (1000,), dtype=jnp.float64)
x.dtype # --> dtype('float64')
# + [markdown] id="6Cks2_gKsXaW"
# ### Caveats
# ⚠️ XLA doesn't support 64-bit convolutions on all backends!
# + [markdown] id="WAHjmL0E2XwO"
# ## Fin.
#
# If something's not covered here that has caused you weeping and gnashing of teeth, please let us know and we'll extend these introductory _advisos_!
| docs/notebooks/Common_Gotchas_in_JAX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simple completeness stats
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
detected = np.load('../outputs/detected.npy')
not_detected = np.load('../outputs/not_detected.npy')
# +
alpha = 1
lw = 2
props = dict(lw=lw, alpha=alpha, histtype='step')
bins = 50
plt.figure(figsize=(4, 3))
plt.hist(detected[:, 0] * 24 * 60, bins=bins, label='Detected', **props)
plt.hist(not_detected[:, 0] * 24 * 60, bins=bins, label='Not Detected',**props)
plt.legend(loc='lower right')
plt.ylabel('Frequency')
ax = plt.gca()
ax2 = ax.twiny()
ax.set_xlabel('Period [min]')
ax2.set_xlabel('Period [hours]')
for axis in [ax, ax2]:
axis.set_xlim([180, 12*60])
ax2.set_xticklabels(["{0:.1f}".format(float(i)/60)
for i in ax2.get_xticks()])
plt.savefig('plots/period.pdf', bbox_inches='tight')
plt.show()
plt.figure(figsize=(4, 3))
plt.hist(detected[:, 1], bins=bins, label='Detected', **props)
plt.hist(not_detected[:, 1], bins=bins, label='Not Detected', **props)
plt.xlabel('Radius [km]')
plt.ylabel('Frequency')
plt.xlim([500, 2500])
plt.legend(loc='center right')
plt.savefig('plots/radius.pdf', bbox_inches='tight')
plt.show()
plt.hist(detected[:, 2], log=True)
plt.hist(not_detected[:, 2], log=True)
plt.xlabel('S/N')
plt.show()
# -
26/6
| notebooks/analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# +
x = np.linspace(1, 15, 100)
y = np.sin(x / 5) * np.exp(x / 10) + 5 * np.exp(-x / 2)
plt.figure(figsize=(15, 10))
plt.grid(which='both')
plt.plot(x, y)
plt.show()
# -
def f(x):
return np.sin(x / 5) * np.exp(x / 10) + 5 * np.exp(-x / 2)
def poly(x):
return (np.linalg.solve(np.vander(x), f(x)))
# +
p1 = poly(np.array([1, 15]))
p2 = poly(np.array([1, 8, 15]))
p3 = poly(np.array([1, 4, 10, 15]))
print(p1, p2, p3)
# -
plt.figure(figsize=(15, 10))
plt.plot(x, y, lw=3, label="exact")
plt.plot(x, np.polyval(p1, x), label="poly1")
plt.plot(x, np.polyval(p2, x), label="poly2")
plt.plot(x, np.polyval(p3, x), label="poly3")
plt.grid(which='both')
plt.legend()
plt.show()
x = np.array([1, 0, 1, 1, 0, 1, 1])
y = np.array([0, 2, 0, 2, 0, 2, 0])
print(np.count_nonzero(x*y))
print(np.count_nonzero(abs(x) + abs(y)))
x = np.array([1, 2, 0])
N = 4
mat = x + np.zeros((N, 1))
print(mat.T.reshape(N*len(x)))
# +
x = np.array([0, 11, 0, 0, -7, 2, 0, 4, 0])
cond = np.roll(x == 0, 1)
cond[0] = 0
arr = x[cond == 1]
print(max(arr))
# +
x = np.array([8, 0, 0, 2, 1, 0, 0, -17.5, 0])
while len(x[x == 0]) > 0:
a = np.roll((x == 0).astype(np.int), -1) * x
x += np.roll(a, 1)
print(x)
# -
| labs/lab16/lab16_inf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="pzNkLTBDZPsX" colab_type="code" colab={}
# + [markdown] id="tih6cQJWZQQc" colab_type="text"
# **Mean Absolute Error (MAE)**
#
# It is simply the average of the squared difference between the target value and the value predicted by the regression model. As it squares the differences, it penalizes even a small error which leads to over-estimation of how bad the model is.
#
# **Root Mean Squared Error:**
#
# it is the square root of the averaged squared difference between the target value and the value predicted by the model.
#
# RMSE is highly affected by outlier values. As compared to mean absolute error, RMSE gives higher weightage and punishes large errors.
# + [markdown] id="Xr7Hnzs_Z-VF" colab_type="text"
#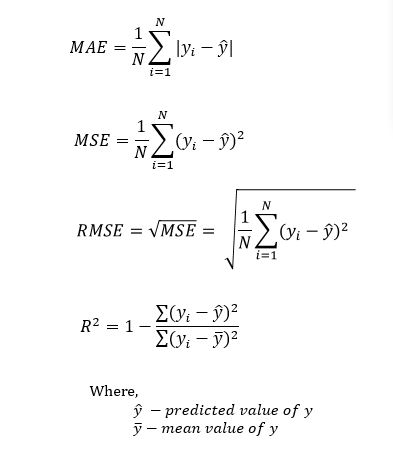 
# + [markdown] id="FPZKa9S6iqBy" colab_type="text"
# **3. Root Mean Squared Logarithmic Error**
#
# 
#
#
# If both predicted and actual values are small: RMSE and RMSLE is same.
#
# If either predicted or the actual value is big: RMSE > RMSLE
#
# If both predicted and actual values are big: RMSE > RMSLE (RMSLE becomes almost negligible)
# + [markdown] id="4RmFp9SRiqLg" colab_type="text"
# Example:
# Y = 60 ,80, 90
#
# X = 67, 78 ,91
#
# On calculating, their RMSE and RMSLE comes out to be: 4.242 and 0.6466 respectively
#
# Now let us introduce an outlier in the data
#
# Y = 60, 80 ,90 ,750
#
# X = 67, 78 ,91, 102
#
# Now, in this case, the RMSE and RMSLE comes out to be: 374.724 and 1.160 respectively.
#
# **RMSE explodes in magnitude as soon as it encounters an outlier. In contrast, even on the introduction of the outlier, the RMSLE error is not affected much.**
#
# **Ex2.**
#
# case1
#
# Y = 100
#
# X = 90
#
# Calculated RMLSE: 0.1053
# Calculated RMSE: 10
#
# Case 2:
#
# Y = 10000
#
# X = 9000
#
# Calculated RMSLE: 0.1053
# Calculated RMSE : 1000
#
# **RMSLE metric only considers the relative error between and the Predicted and the actual value and the scale of the error is not significant. On the other hand, RMSE value Increases in magnitude if the scale of error increases.**
# + [markdown] id="__esuDPtknQ-" colab_type="text"
# RMSLE incurs a larger penalty for the underestimation of the Actual variable than the Overestimation
# + [markdown] id="XX8hRLjclJgm" colab_type="text"
# **R² Error:**
#
# Coefficient of Determination or R² is another metric used for evaluating the performance of a regression model. the R² will always be less than or equal to 1.Nearer the value to 1 means model is good. R-Squared does not penalize for adding features that add no value to the model.
#
# 
#
#
#
# if model has too many terms and too many high-order polynomials you can run into the problem of over-fitting the data. When you over-fit data, a misleadingly high R2 value can lead to misleading projections.
# + [markdown] id="kU4HxZvYlJdE" colab_type="text"
# **Adjusted R-Squared**
#
# A model performing equal to baseline would give R-Squared as 0. Better the model, higher the r2 value. The best model with all correct predictions would give R-Squared as 1. However, on adding new features to the model, the R-Squared value either increases or remains the same. R-Squared does not penalize for adding features that add no value to the model
#
# 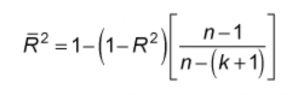
#
# k: number of features
#
# n: number of samples
#
#
# this metric takes the number of features into account. When we add more features, the term in the denominator n-(k +1) decreases, so the whole expression increases.
#
# If R-Squared does not increase, that means the feature added isn’t valuable for our model. So overall we subtract a greater value from 1 and adjusted r2, in turn, would decrease.
#
# R2 shows how well terms (data points) fit a curve or line. Adjusted R2 also indicates how well terms fit a curve or line, but adjusts for the number of terms in a model. If you add more and more useless variables to a model, adjusted r-squared will decrease. If you add more useful variables, adjusted r-squared will increase. Adjusted R2 will always be less than or equal to R2.
# + [markdown] id="uuL90zW7lJbB" colab_type="text"
#
| Regression_metrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LogisticRegressionCV
from sklearn.linear_model import LassoCV
from sklearn.model_selection import cross_val_score
from sklearn.metrics import accuracy_score
from sklearn.model_selection import KFold
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import accuracy_score
from sklearn import metrics
#from prettytable import PrettyTable
import seaborn as sns
#import geopandas
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.ticker import PercentFormatter
# %matplotlib inline
from datetime import datetime
import seaborn as sns
sns.set()
# -
#data from India folder
india_statewise = pd.read_csv('data/India/india_statewise.csv')
covid_19_india = pd.read_csv('data/India/covid_19_india.csv')
india_statewise.head(100)
india_statewise.info()
india_statewise.isnull().sum()
2548/5966
india_statewise.State.nunique()
india_statewise.describe()
india_statewise.drop(columns = ['Negative'], inplace = True)
new_india = india_statewise.ffill(axis = 0)
new_india.isnull().sum()
# +
#new_india = india_statewise.dropna()
# -
new_india
States = set(new_india.State.values)
States = list(States)
new_india[new_india['State'] == States[4]]
sum_by_dates = new_india.groupby(['Date']).count().reset_index()
sum_by_dates
sum_by_dates['Date'] = pd.to_datetime(sum_by_dates['Date'])
sum_by_dates['Date'][0] - sum_by_dates['Date'][189]
# **IGNORE CODE ABOVE THIS FOR NOW**
covid_19_india.head(100)
covid_19_india['Date'] = pd.to_datetime(covid_19_india['Date'], format = '%d/%m/%y')
#covid_19_india[covid_19_india['State/UnionTerritory']=='Telengana' & covid_19_india['Date'] == '12/06/20']
covid_19_india['Date']
#covid_19_india['Date'].sort_values()
covid_19_india[covid_19_india['Date'] == '2020-06-12']
# +
srt_dates = {}
end_dates = {}
states = list(set(covid_19_india[covid_19_india.columns[3]].values))
for i in states:
srt_dates[i] = (covid_19_india[covid_19_india['State/UnionTerritory']==i]['Date'].iloc[0])
end_dates[i] = covid_19_india[covid_19_india['State/UnionTerritory']==i]['Date'].iloc[-1]
#print(srt_dates['Kerala'])
srt_dates = {k: v for k, v in sorted(srt_dates.items(), key=lambda item: item[1])}
print("Starting Dates:",srt_dates)
print("\n\n")
end_dates = {k: v for k, v in sorted(end_dates.items(), key=lambda item: item[1])}
print("Ending Dates:",end_dates)
# -
covid_19_india[covid_19_india['State/UnionTerritory'] == 'Daman & Diu']
cv_by_dates = covid_19_india.groupby('Date').sum()
copy = cv_by_dates['Confirmed'][:]
copy = list(copy)
copy.sort()
copy==list(cv_by_dates['Confirmed'])
cv_by_dates
copy = cv_by_dates['Deaths'][:]
copy = list(copy)
copy.sort()
copy==list(cv_by_dates['Deaths'])
copy = cv_by_dates['Cured'][:]
copy = list(copy)
copy.sort()
copy==list(cv_by_dates['Cured'])
cv_by_dates['Daily Cases'] = cv_by_dates['Confirmed']
cases = list(cv_by_dates['Confirmed'])
cases.insert(0,0)
cases = cases[:-1]
daily = list(cv_by_dates['Daily Cases'])
print(len(daily))
print(len(cases))
daily_cases = []
zip_object = zip(daily, cases)
for list1_i, list2_i in zip_object:
daily_cases.append(list1_i-list2_i)
daily_cases
cv_by_dates['Daily Cases'] = daily_cases
cv_by_dates['Daily Deaths'] = cv_by_dates['Deaths']
cases = list(cv_by_dates['Deaths'])
cases.insert(0,0)
cases = cases[:-1]
daily = list(cv_by_dates['Daily Deaths'])
print(len(daily))
print(len(cases))
daily_cases = []
zip_object = zip(daily, cases)
for list1_i, list2_i in zip_object:
daily_cases.append(list1_i-list2_i)
daily_cases
cv_by_dates['Daily Deaths'] = daily_cases
cv_by_dates['Daily Cured'] = cv_by_dates['Cured']
cases = list(cv_by_dates['Cured'])
cases.insert(0,0)
cases = cases[:-1]
daily = list(cv_by_dates['Daily Cured'])
print(len(daily))
print(len(cases))
daily_cases = []
zip_object = zip(daily, cases)
for list1_i, list2_i in zip_object:
daily_cases.append(list1_i-list2_i)
daily_cases
cv_by_dates['Daily Cured'] = daily_cases
#tcks = np.linspace(cv_by_dates.index[0],cv_by_dates.index[-1], 4)
#cv_by_dates.index[0]
mid = cv_by_dates.index[len(cv_by_dates.index)//2]
#print(tcks)
plt.plot(cv_by_dates.index, cv_by_dates['Daily Cases'])
plt.xlabel('Dates')
plt.ylabel('Daily Number of Cases')
plt.xticks(ticks = [cv_by_dates.index[0], mid, cv_by_dates.index[-1]], rotation = 45)
plt.title('INDIA')
plt.plot(cv_by_dates.index, cv_by_dates['Daily Deaths'])
plt.xlabel('Dates')
plt.ylabel('Daily Number of Deaths')
plt.xticks(ticks = [cv_by_dates.index[0], mid, cv_by_dates.index[-1]], rotation = 45)
plt.title('INDIA')
plt.plot(cv_by_dates.index, cv_by_dates['Daily Cured'])
plt.xlabel('Dates')
plt.ylabel('Daily Number of Recoveries')
plt.xticks(ticks = [cv_by_dates.index[0],mid, cv_by_dates.index[-1]], rotation = 45)
plt.title('INDIA')
states = list(set(covid_19_india['State/UnionTerritory'].values))
df_state = {}
for i in states:
temp = covid_19_india[covid_19_india['State/UnionTerritory'] == i]
df_state[i] = temp.groupby('Date').sum()
df_state
df_state.keys()
#fig, axs = plt.subplots(2)
#j = 0
for i in df_state.keys():
df_state[i]['Daily Cases'] = df_state[i]['Confirmed']
cases = list(df_state[i]['Confirmed'])
cases.insert(0,0)
cases = cases[:-1]
daily = list(df_state[i]['Daily Cases'])
#print(len(daily))
#print(len(cases))
daily_cases = []
zip_object = zip(daily, cases)
for list1_i, list2_i in zip_object:
daily_cases.append(list1_i-list2_i)
daily_cases
df_state[i]['Daily Cases'] = daily_cases
mid = df_state[i].index[len(df_state[i].index)//2]
plt.plot(df_state[i].index, df_state[i]['Daily Cases'])
plt.xlabel('Dates')
plt.ylabel('Daily Number of Cases')
plt.xticks(ticks = [df_state[i].index[0], mid, df_state[i].index[-1]])
plt.title(f'State: {i}')
if(i == 'Telangana***'):
i = 'Telangana_star3'
elif(i == 'Telengana***'):
i = 'Telengana_star3'
plt.savefig(f'State_{i}_plt.png')
plt.clf()
df_state
mid = df_state[i].index[len(df_state[i].index)//2]
plt.plot(df_state[i].index, df_state[i]['Daily Cases'])
plt.xlabel('Dates')
plt.ylabel('Daily Number of Cases')
plt.xticks(ticks = [df_state[i].index[0], mid, df_state[i].index[-1]], rotation = 45)
plt.title(f'State: {i}')
# +
a = True
b = False
c = False
if not a or b:
print ("CORRECT ANSWER")
elif not a or not b and c:
print ("MAYBE CORRECT ANSWER")
elif not a or b or not b and a:
print ("MAYBE WRONG ANSWER")
else:
print ("WRONG ANSWER")
# +
class Sample:
def __init__(self):
self.x = 5
def change(self):
self.x = 10
class Derived_sample(Sample):
def change(self):
self.x=self.x+1
return self.x
def main():
obj = Derived_sample()
print(obj.change())
main()
# +
class A:
def one(self):
return self.two()
def two(self):
return 'A'
class B(A):
def two(self):
return 'B'
obj1=A()
obj2=B()
print(obj1.one(),obj2.one())
# -
import pandas as pd
df = pd.DataFrame({'A':[2,3,5], 'B': [4,6,10]})
print(df.set_index('A').columns)
df
df = pd.DataFrame({'A':[2,3,5], 'B': [4,6,10]})
df.columns
np.random.uniform(0,1,5)
np.random.normal(0,1,5)
| India_Final_Project_20Oct.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Delete duplicates
#
# ```sql
# with r as (SELECT array_agg(cartodb_id), count(the_geom), the_geom, dates, hour, type FROM potential_landslide_areas group by the_geom, dates, hour, type order by 2 desc),
# s as (select unnest(array_remove(array_agg, array_agg[count])) from r where count > 1)
# DELETE FROM potential_landslide_areas where cartodb_id in (select * from s)
# ```
| ResourceWatch/data_management/Useful Queries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:ap-southeast-1:492261229750:image/datascience-1.0
# ---
# **Post-Processing Amazon Textract with Location-Aware Transformers**
#
# # Part 3: Implementing Human Review
#
# > *This notebook works well with the `Python 3 (Data Science)` kernel on SageMaker Studio*
#
# In this final notebook, we'll set up the human review component of the OCR pipeline using [Amazon Augmented AI (A2I)](https://aws.amazon.com/augmented-ai/): Completing the demo pipeline.
#
# The A2I service shares a lot in common with SageMaker Ground Truth, with the main difference that A2I is designed for **near-real-time, single-example** annotation/review to support a live business process, while SMGT is oriented towards **offline, batch** annotation for building datasets.
#
# The two services both use the Liquid HTML templating language, and you might reasonably wonder: "*Are we going to use the same custom boxes-plus-review template from earlier?*"
#
# In fact, no we won't - for reasons we'll get to in a moment.
#
# First though, let's load the required libraries and configuration for the notebook as before:
# +
# %load_ext autoreload
# %autoreload 2
# Python Built-Ins:
import json
from logging import getLogger
import os
# External Dependencies:
import boto3 # AWS SDK for Python
import sagemaker # High-level SDK for SageMaker
# Local Dependencies:
import util
logger = getLogger()
role = sagemaker.get_execution_role()
s3 = boto3.resource("s3")
smclient = boto3.client("sagemaker")
# Manual configuration (check this matches notebook 1):
bucket_name = sagemaker.Session().default_bucket()
bucket_prefix = "textract-transformers/"
print(f"Working in bucket s3://{bucket_name}/{bucket_prefix}")
config = util.project.init("ocr-transformers-demo")
print(config)
# Field configuration saved from first notebook:
with open("data/field-config.json", "r") as f:
fields = [
util.postproc.config.FieldConfiguration.from_dict(cfg)
for cfg in json.loads(f.read())
]
entity_classes = [f.name for f in fields]
# S3 URIs per first notebook:
raw_s3uri = f"s3://{bucket_name}/{bucket_prefix}data/raw"
# -
# ## The rationale for a separate review template
#
# For many ML-powered processes, intercepting low-confidence predictions for human review is important for delivering efficient, accurate service.
#
# To deliver high-performing ML models sustainably, continuous collection of feedback for re-training is also important.
#
# In this section we'll detail some reasons **why**; although joining the two processes together might be ideal; this example will demonstrate a **separate prediction review workflow** from the training data collection.
# ### Tension between process execution and model improvement
#
# As we saw when setting up the pipeline in the last notebook, there's a **post-processing step** after the ML model - whose purpose is:
#
# - To consolidate consecutive `WORD`s of the same class into a single "entity" detection via a simple heuristic
# - To apply (configurable) business rules to consolidate entity detections into "fields" on the document (e.g. selecting a single value from multiple possible matches, etc).
#
# Both of these processes are implemented in a simple Python Lambda function, and so would be technically straightforward to port into the ML model endpoint itself (in [src/inference.py](src/inference.py)). However, it's the **second one** that's important.
#
# For any use case where there's a non-trivial **gap** between what the model is trained to estimate and what the business process consumes, there's a **tension** in the review process:
#
# 1. Reviewing business process fields is efficient, but does not collect model training data (although it may help us understand overall accuracy)
# 2. Reviewing the model inputs & outputs directly collects training data, but:
# - Does not directly review the accuracy of the end-to-end business process, so requires complete trust in the post-processing rules
# - May be inefficient, as the reviewer needs to collect more information than the business process absolutely requires (e.g. having to highlight every instance of `Provider Name` in the doc, when the business process just needs to know what the name is)
# 3. Splitting the review into multiple stages collects training/accuracy data for both components (ML model and business rules), but requires even more time - especially if the hand-off between the review stages might be asynchronous
#
# In many cases the efficiency of the live process is most important for customer experience and cost management, and so approach 1 is taken (as we will in this example): With collection of additional model training data handled as an additional background task.
#
# In some cases it may be possible to fully close the gap to resolve the tension and make a single offline-annotation/online-review UI work for everybody... E.g. for the credit cards use case, we might be able to:
#
# - (Add effort) Move from word classification to a **sequence-to-sequence model**, to support more complex output processing (like OCR error correction, field format standardization, grouping words into matches, etc)... *OR*
# - (Reduce scope) **Focus only on use-cases** where:
# - Each entity class only appears once in the document, *OR* most/every detection of multiple entities is equally important to the business process (may often be the case! E.g. on forms or other well-structured use cases) *AND*
# - Errors in OCR transcription or the heuristic rules to group matched words together are rare enough *or unpredictable enough* that there's no value in a confidence-score-based review (E.g. if "The OCR/business rules aren't making mistakes very often, and even when they do the confidence is still high - so our online review isn't helping these issues")
# ### A small techical challenge
#
# So what if your use case for this model is:
#
# - Seeing **high enough OCR accuracy rates** from Textract, and
# - Enjoying good success with the heuristic for **joining classified words together** into multi-word entities based on the order Textract returned them, and
# - Either having only **one match per entity type** per document; or where it's important to **always return multiple matches** if they're present?
#
# Then maybe it would make sense roll your online review and training data collection into one process! By simply trusting the post-processing logic / OCR quality, and having reviewers use the bounding box tool.
#
# **However,** there's one final hurdle: At the time of writing, the Ground Truth/A2I bounding box annotator works only for individual images, not PDFs. This means you'd also need to either:
#
# - Restrict the pipeline to processing single-page documents/images, or
# - Implement a custom box task UI capable of working with PDFs also, or
# - Orchestrate around the problem by splitting and dispatching each document to multiple single-page A2I reviews.
# ### In summary
#
# For some use cases of technology like this, directly using the training data annotation UI for online review could be the most efficient option.
#
# But to avoid ignoring the (potentially large) set of viable use cases where it's not practical; and to avoid introducing complexity or workarounds for handling multi-page documents; this sample presents a separate tailored online review UI.
#
# ## Develop the review task template
#
# Just as with SageMaker Ground Truth, a custom task UI template has been provided and we can preview it via the SageMaker SDK.
#
# In this case no extra parameter substitutions are required, so we can jump straight to rendering.
#
# One difference is that the input to this stage of the pipeline is a little more complex than a simple image + Amazon Textract result URI as in initial data labelling. We'll use an example JSON file and substitute the Textract URI to match your bucket & prefix (so the displayed file will not match the extracted field content).
# +
# Load the sample input from file:
with open("review/task-input.example.json", "r") as f:
sample_obj = json.loads(f.read())
# Find any `a_pdf_s3uri`, so long as it exists in your account:
textract_s3key_root = f"{bucket_prefix}data/raw"
try:
a_pdf_s3obj = next(filter(
lambda o: o.key.endswith(".pdf"),
s3.Bucket(bucket_name).objects.filter(Prefix=textract_s3key_root)
))
a_pdf_s3uri = f"s3://{a_pdf_s3obj.bucket_name}/{a_pdf_s3obj.key}"
except StopIteration as e:
raise ValueError(
f"Couldn't find any .pdf files in s3://{bucket_name}/{textract_s3key_root}"
) from e
# Substitute the PDF URI in the sample input object:
sample_obj["TaskObject"] = a_pdf_s3uri
# -
# Now render the template:
# +
ui_render_file = "review/render.tmp.html"
with open("review/fields-validation.liquid.html", "r") as fui:
with open(ui_render_file, "w") as frender:
ui_render_resp = smclient.render_ui_template(
UiTemplate={ "Content": fui.read() },
Task={ "Input": json.dumps(sample_obj) },
RoleArn=role,
)
frender.write(ui_render_resp["RenderedContent"])
if "Errors" in ui_render_resp:
if (ui_render_resp["Errors"] and len(ui_render_resp["Errors"])):
print(ui_render_resp["Errors"])
raise ValueError("Template render returned errors")
print(f"▶️ Open {ui_render_file} and click 'Trust HTML' to see the UI in action!")
# -
# ## Set up the human review workflow
#
# Similarly to a SageMaker Ground Truth labelling job, we have 3 main concerns for setting up an A2I review workflow:
#
# - **Who's** doing the labelling
# - **What** the task will look like
# - **Where** the output reviews will be stored to once the review completes (i.e. location on Amazon S3)
#
# Our **workteam** from notebook 1 should already be set up.
#
# ▶️ **Check** the workteam name below matches your setup, and run the cell to store the ARN:
# +
workteam_name = "just-me" # TODO: Update this to match yours, if different
workteam_arn = util.smgt.workteam_arn_from_name(workteam_name)
# -
# Our **template** has been tested as above, so just needs to be registered with A2I.
#
# You can use the below code to register your template and store its ARN, but can also refer to the [A2I Console worker task templates list](https://console.aws.amazon.com/a2i/home?#/worker-task-templates)
# +
with open("review/fields-validation.liquid.html", "r") as f:
create_template_resp = smclient.create_human_task_ui(
HumanTaskUiName="fields-validation-1", # (Can change this name as you like)
UiTemplate={ "Content": f.read() },
)
task_template_arn = create_template_resp["HumanTaskUiArn"]
print(f"Created A2I task template:\n{task_template_arn}")
# -
# To finish setting up the "workflow" itself, we need 2 more pieces of information:
#
# - The **location in S3** where review outputs should be stored
# - An appropriate **execution role** which will give the A2I workflow to read input documents and write review results.
#
# These are determined by the **OCR pipeline solution stack**, because the reviews bucket is created by the pipeline with event triggers to resume the next stage when reviews are uploaded.
#
# The code below should be able to look up these parameters for you automatically:
reviews_bucket_name = config.pipeline_reviews_bucket_name
print(reviews_bucket_name)
reviews_role_arn = config.a2i_execution_role_arn
print(reviews_role_arn)
# Alternatively, you may **find** your pipeline solution stack from the [AWS CloudFormation Console](https://console.aws.amazon.com/cloudformation/home?#/stacks) and click through to the stack detail page. From the **Outputs** tab, you should see the `A2IHumanReviewBucketName` and `A2IHumanReviewExecutionRoleArn` values as shown below.
#
# (You may also note the `A2IHumanReviewFlowParamName`, which we'll use in the next section)
#
# 
# Once these values are populated, you're ready to create your review workflow by running the code below.
#
# Note that you can also manage flows via the [A2I Human Review Workflows Console](https://console.aws.amazon.com/a2i/home?#/human-review-workflows/).
# +
create_flow_resp = smclient.create_flow_definition(
FlowDefinitionName="ocr-fields-validation-1", # (Can change this name as you like)
HumanLoopConfig={
"WorkteamArn": workteam_arn,
"HumanTaskUiArn": task_template_arn,
"TaskTitle": "Review OCR Field Extractions",
"TaskDescription": "Review and correct credit card agreement field extractions",
"TaskCount": 1, # One reviewer per item
"TaskAvailabilityLifetimeInSeconds": 60 * 60, # Availability timeout
"TaskTimeLimitInSeconds": 60 * 60, # Working timeout
},
OutputConfig={
"S3OutputPath": f"s3://{reviews_bucket_name}/reviews",
},
RoleArn=reviews_role_arn,
)
print(f"Created review workflow:\n{create_flow_resp['FlowDefinitionArn']}")
# -
# ## Integrate with the OCR pipeline
#
# Once the human review workflow is created, the final integration step is to point the pipeline at it - just as we did for our SageMaker endpoint earlier.
#
# In code, this can be done as follows:
# +
print(f"Configuring pipeline with review workflow: {create_flow_resp['FlowDefinitionArn']}")
ssm = boto3.client("ssm")
ssm.put_parameter(
Name=config.a2i_review_flow_arn_param,
Overwrite=True,
Value=create_flow_resp["FlowDefinitionArn"],
)
# -
# Alternatively through the console, you would follow these steps:
#
# ▶️ **Check** the `A2IHumanReviewFlowParamName` output of your OCR pipeline stack in [CloudFormation](https://console.aws.amazon.com/cloudformation/home?#/stacks) (as we did above)
#
# ▶️ **Open** the [AWS Systems Manager Parameter Store console](https://console.aws.amazon.com/systems-manager/parameters/?tab=Table) and **find the review flow parameter in the list**.
#
# ▶️ **Click** on the name of the parameter to open its detail page, and then on the **Edit** button in the top right corner. Set the value to the **workflow ARN** (see previous code cell in this notebook) and save the changes.
#
# 
# ## Final testing
#
# Your OCR pipeline should now be fully functional! Let's try it out:
#
# ▶️ **Log in** to the labelling portal (URL available from the [SageMaker Ground Truth Workforces Console](https://console.aws.amazon.com/sagemaker/groundtruth?#/labeling-workforces) for your correct AWS Region)
#
# 
#
# ▶️ **Upload** one of the sample documents to your pipeline's input bucket in Amazon S3, either using the code snippets below or drag and drop in the [Amazon S3 Console](https://console.aws.amazon.com/s3/)
pdfpaths = []
for currpath, dirs, files in os.walk("data/raw"):
if "/." in currpath or "__" in currpath:
continue
pdfpaths += [
os.path.join(currpath, f) for f in files
if f.lower().endswith(".pdf")
]
pdfpaths = sorted(pdfpaths)
# +
test_filepath = pdfpaths[14]
test_s3uri = f"s3://{config.pipeline_input_bucket_name}/{test_filepath}"
# !aws s3 cp '{test_filepath}' '{test_s3uri}'
# -
# ▶️ **Open up** your "Processing Pipeline" state machine in the [AWS Step Functions Console](https://console.aws.amazon.com/states/home?#/statemachines)
#
# After a few seconds you should find that a Step Function execution is automatically triggered and (since we enabled so many fields that at least one is always missing) the example is eventually forwarded for human review in A2I.
#
# As you'll see from the `ModelResult` field in your final *Step Output*, this pipeline produces a rich but usefully-structured output - with good opportunities for onward integration into further Step Functions steps or external systems. You can find more information and sample solutions for integrating AWS Step Functions in the [Step Functions Developer Guide](https://docs.aws.amazon.com/step-functions/latest/dg/welcome.html).
#
# 
# ## Conclusion
#
# In this worked example we showed how advanced, open-source language processing models specifically tailored for document understanding can be integrated with [Amazon Textract](https://aws.amazon.com/textract/): providing a trainable, ML-driven framework for tackling more niche or complex requirements where Textract's [built-in structure extraction tools](https://aws.amazon.com/textract/features/) may not fully solve the challenges out-of-the-box.
#
# The underlying principle of the model - augmenting multi-task neural text processing architectures with positional data - is highly extensible, with potential to tackle a wide range of use cases where joint understanding of the content and presentation of text can deliver better results than considering text alone.
#
# We demonstrated how an end-to-end process automation pipeline applying this technology might look: Developing and deploying the model with [Amazon SageMaker](https://aws.amazon.com/sagemaker/), building a serverless workflow with [AWS Step Functions](https://aws.amazon.com/step-functions/) and [AWS Lambda](https://aws.amazon.com/lambda/), and driving quality with human review of low-confidence documents through [Amazon Augmented AI](https://aws.amazon.com/augmented-ai/).
#
# Thanks for following along, and for more information, don't forget to check out:
#
# - The other published [Amazon Textract Examples](https://docs.aws.amazon.com/textract/latest/dg/other-examples.html) listed in the [Textract Developer Guide](https://docs.aws.amazon.com/textract/latest/dg/what-is.html)
# - The extensive repository of [Amazon SageMaker Examples](https://github.com/aws/amazon-sagemaker-examples) and usage documentation in the [SageMaker Python SDK User Guide](https://sagemaker.readthedocs.io/en/stable/) - as well as the [SageMaker Developer Guide](https://docs.aws.amazon.com/sagemaker/index.html)
# - The wide range of other open algorithms and models published by [HuggingFace Transformers](https://huggingface.co/transformers/), and their specific documentation on [using the library with SageMaker](https://huggingface.co/transformers/sagemaker.html)
# - The conversational AI and NLP area (and others) of Amazon's own [Amazon.Science](https://www.amazon.science/conversational-ai-natural-language-processing) blog
#
# Happy building!
| notebooks/3. Human Review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/camilorey/material_clases/blob/main/visualizadorPruebasSaber11_2019_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="KxLAVIFS1cHJ"
import pandas as Pandas
import matplotlib.pyplot as PyPlot
# %matplotlib inline
# + [markdown] id="Ot7sZw_013-Q"
# Vamos a agregar el archivo de las pruebas Saber 11 2019-2 desde el API de SODA (Socrates) usado por el portal de Datos Abiertos y almacenarlo en un DataFrame
# + id="Mj7zkNqq2C0B" colab={"base_uri": "https://localhost:8080/", "height": 357} outputId="f20f6730-adea-4368-b5d9-16cf2b6627a3"
#read data from Datos Abiertos
documentAddress = 'https://www.datos.gov.co/resource/tkn6-e4ic.json'
saber11DataFrame = Pandas.read_json(documentAddress)
saber11DataFrame.head()
# + id="vyp2IZBR2NEc"
Vamos a dividir las variables en varios tipos. Dependiendo si son variables binarias o personales ordinales (para efectos de una clasificación en árbol más adelantes)
# + id="KrTNQnbZ2QQ2"
#variables en archivo
variablesPersonalesBinarias = ['estu_genero','estu_tieneetnia','fami_tieneinternet', 'fami_tieneserviciotv',\
'fami_tienecomputador','fami_tienelavadora', 'fami_tienehornomicroogas',\
'fami_tieneautomovil','fami_tienemotocicleta', 'fami_tieneconsolavideojuegos','estu_privado_libertad']
variablesPersonalesOrdinales = ['fami_estratovivienda','fami_personashogar','fami_cuartoshogar', \
'fami_educacionpadre', 'fami_educacionmadre','fami_trabajolaborpadre',\
'fami_trabajolabormadre','fami_numlibros', 'fami_comelechederivados',\
'fami_comecarnepescadohuevo', 'fami_comecerealfrutoslegumbre',\
'fami_situacioneconomica', 'estu_dedicacionlecturadiaria',\
'estu_dedicacioninternet', 'estu_horassemanatrabaja','estu_tiporemuneracion',\
'estu_nse_individual']
#variables del examen
variablesExamen = ['percentil_lectura_critica', 'desemp_lectura_critica',\
'percentil_matematicas', 'desemp_matematicas',\
'percentil_c_naturales', 'desemp_c_naturales',\
'percentil_sociales_ciudadanas',\
'desemp_sociales_ciudadanas', 'percentil_ingles',\
'percentil_global','estu_inse_individual']
#variables de puntajes
puntajesExamen = ['punt_matematicas','punt_c_naturales','punt_sociales_ciudadanas','punt_ingles','punt_lectura_critica']
#variables del colegio
variablesColegio = ['cole_naturaleza','cole_bilingue','cole_caracter','cole_genero','cole_calendario','cole_jornada']
#variables objetivo aqui
variableObjetivo = ['desemp_ingles','punt_global']
# + [markdown] id="O-PL44nT3jwu"
# Vamos a hacer una función para extraer las categorías de una variable no numérica (binaria u ordinal) que nos servirá después para ver cómo subcategorizar diferentes puntajes
# + id="tEv21wyw3wNa"
def categoriasDeVariable(nomVariable):
varInfo = saber11DataFrame[nomVariable].value_counts()
categorias = []
for pareja in varInfo.iteritems():
categorias.append(pareja[0])
return categorias
# + [markdown] id="7JnRzSe935JB"
# Vamos a probar este método visualizando las diferencias que hay entre hombres y mujeres en materia de puntajes obtenidos
# + id="AW4fFaKs4K2T" colab={"base_uri": "https://localhost:8080/", "height": 291} outputId="19bfe0e4-99ae-45a9-fdfc-5f5ca40e3928"
#figuras para ver distribución de puntajes entre hombres y mujeres
Figure, axes = PyPlot.subplots(1,len(puntajesExamen),figsize=(12,2.5),dpi=100,sharex=True,sharey=True)
generos = categoriasDeVariable('estu_genero')
for i, (var,ax) in enumerate(zip(puntajesExamen,axes.flatten())):
for genero in generos:
infoGenero = saber11DataFrame[var].loc[saber11DataFrame['estu_genero'] == genero]
ax.hist(infoGenero,bins=50,label=genero,alpha=0.5)
ax.set_title(var)
ax.legend(prop={'size':8})
PyPlot.suptitle('Distribución de Puntajes',fontsize=15,y=1.05)
PyPlot.show()
# + [markdown] id="jDoeRCm45KFt"
# Ahora queremos ver cómo la variable de puntaje de ingles se distribuye con respecto a las variables socio-demográficas antes definidas
# + id="sp8ehYm35P3W" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="9dcbd314-598c-44d1-b1e3-b0eb0310dd71"
variableDeInteres = 'punt_ingles'
variablesSocioDemograficas = variablesPersonalesBinarias+variablesPersonalesOrdinales
numVariables = len(variablesSocioDemograficas)
Figure, axes = PyPlot.subplots(numVariables,1,figsize=(15,5*numVariables),dpi=100,sharex=True,sharey=True)
index = 0
for var in variablesSocioDemograficas:
categorias = categoriasDeVariable(var)
for cat in categorias:
data = saber11DataFrame[variableDeInteres].loc[saber11DataFrame[var] == cat]
axes[index].hist(data,bins=100,label=cat,alpha=0.8)
axes[index].set_xlabel('puntajes')
axes[index].set_ylabel('cantidad')
if len(categorias) <12:
axes[index].legend(loc='best',ncol=10,fontsize=8)
axes[index].set_title(var)
index +=1
PyPlot.show()
| visualizadorPruebasSaber11_2019_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import seaborn as sns
import json
from pandas.io.json import json_normalize
from wordcloud import WordCloud, STOPWORDS
| kaggle/TED.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collision Avoidance - Train Model (with live graph)
#
# Welcome to this host side Jupyter Notebook! This should look familiar if you ran through the notebooks that run on the robot. In this notebook we'll train our image classifier to detect two classes
# ``free`` and ``blocked``, which we'll use for avoiding collisions. For this, we'll use a popular deep learning library *PyTorch*
import torch
import torch.optim as optim
import torch.nn.functional as F
import torchvision
import torchvision.datasets as datasets
import torchvision.models as models
import torchvision.transforms as transforms
# ### Upload and extract dataset
#
# Before you start, you should upload the ``dataset.zip`` file that you created in the ``data_collection.ipynb`` notebook on the robot.
#
# You should then extract this dataset by calling the command below
# !unzip -q dataset.zip
# You should see a folder named ``dataset`` appear in the file browser.
# ### Create dataset instance
# Now we use the ``ImageFolder`` dataset class available with the ``torchvision.datasets`` package. We attach transforms from the ``torchvision.transforms`` package to prepare the data for training.
dataset = datasets.ImageFolder(
'dataset',
transforms.Compose([
transforms.ColorJitter(0.1, 0.1, 0.1, 0.1),
transforms.Resize((224, 224)),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
])
)
# ### Split dataset into train and test sets
# Next, we split the dataset into *training* and *test* sets. The test set will be used to verify the accuracy of the model we train.
train_dataset, test_dataset = torch.utils.data.random_split(dataset, [len(dataset) - 50, 50])
# ### Create data loaders to load data in batches
# We'll create two ``DataLoader`` instances, which provide utilities for shuffling data, producing *batches* of images, and loading the samples in parallel with multiple workers.
# +
train_loader = torch.utils.data.DataLoader(
train_dataset,
batch_size=16,
shuffle=True,
num_workers=4
)
test_loader = torch.utils.data.DataLoader(
test_dataset,
batch_size=16,
shuffle=True,
num_workers=4
)
# -
# ### Define the neural network
#
# Now, we define the neural network we'll be training. The *torchvision* package provides a collection of pre-trained models that we can use.
#
# In a process called *transfer learning*, we can repurpose a pre-trained model (trained on millions of images) for a new task that has possibly much less data available.
#
# Important features that were learned in the original training of the pre-trained model are re-usable for the new task. We'll use the ``alexnet`` model.
model = models.alexnet(pretrained=True)
# The ``alexnet`` model was originally trained for a dataset that had 1000 class labels, but our dataset only has two class labels! We'll replace
# the final layer with a new, untrained layer that has only two outputs.
model.classifier[6] = torch.nn.Linear(model.classifier[6].in_features, 2)
# Finally, we transfer our model for execution on the GPU
device = torch.device('cuda')
model = model.to(device)
# ### Visualization utilities
#
# Exececute the cell below to enable live plotting.
#
# > You need to install bokeh (https://docs.bokeh.org/en/latest/docs/installation.html)
#
# ```bash
# sudo pip3 install bokeh
# sudo jupyter labextension install @jupyter-widgets/jupyterlab-manager
# sudo jupyter labextension install @bokeh/jupyter_bokeh
# ```
# +
from bokeh.io import push_notebook, show, output_notebook
from bokeh.layouts import row
from bokeh.plotting import figure
from bokeh.models import ColumnDataSource
from bokeh.models.tickers import SingleIntervalTicker
output_notebook()
colors = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728']
p1 = figure(title="Loss", x_axis_label="Epoch", plot_height=300, plot_width=360)
p2 = figure(title="Accuracy", x_axis_label="Epoch", plot_height=300, plot_width=360)
source1 = ColumnDataSource(data={'epochs': [], 'trainlosses': [], 'testlosses': [] })
source2 = ColumnDataSource(data={'epochs': [], 'train_accuracies': [], 'test_accuracies': []})
#r = p1.multi_line(ys=['trainlosses', 'testlosses'], xs='epochs', color=colors, alpha=0.8, legend_label=['Training','Test'], source=source)
r1 = p1.line(x='epochs', y='trainlosses', line_width=2, color=colors[0], alpha=0.8, legend_label="Train", source=source1)
r2 = p1.line(x='epochs', y='testlosses', line_width=2, color=colors[1], alpha=0.8, legend_label="Test", source=source1)
r3 = p2.line(x='epochs', y='train_accuracies', line_width=2, color=colors[0], alpha=0.8, legend_label="Train", source=source2)
r4 = p2.line(x='epochs', y='test_accuracies', line_width=2, color=colors[1], alpha=0.8, legend_label="Test", source=source2)
p1.legend.location = "top_right"
p1.legend.click_policy="hide"
p2.legend.location = "bottom_right"
p2.legend.click_policy="hide"
# -
# ### Train the neural network
#
# Using the code below we will train the neural network for 30 epochs, saving the best performing model after each epoch.
#
# > An epoch is a full run through our data.
# +
NUM_EPOCHS = 30
BEST_MODEL_PATH = 'best_model.pth'
best_accuracy = 0.0
optimizer = optim.SGD(model.parameters(), lr=0.001, momentum=0.9)
handle = show(row(p1, p2), notebook_handle=True)
for epoch in range(NUM_EPOCHS):
train_loss = 0.0
train_error_count = 0.0
for images, labels in iter(train_loader):
images = images.to(device)
labels = labels.to(device)
optimizer.zero_grad()
outputs = model(images)
loss = F.cross_entropy(outputs, labels)
train_loss += loss
train_error_count += float(torch.sum(torch.abs(labels - outputs.argmax(1))))
loss.backward()
optimizer.step()
train_loss /= len(train_loader)
test_loss = 0.0
test_error_count = 0.0
for images, labels in iter(test_loader):
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
loss = F.cross_entropy(outputs, labels)
test_loss += loss
test_error_count += float(torch.sum(torch.abs(labels - outputs.argmax(1))))
test_loss /= len(test_loader)
train_accuracy = 1.0 - float(train_error_count) / float(len(train_dataset))
test_accuracy = 1.0 - float(test_error_count) / float(len(test_dataset))
print('%d: %f, %f, %f, %f' % (epoch+1, train_loss, test_loss, train_accuracy, test_accuracy))
new_data1 = {'epochs': [epoch+1],
'trainlosses': [float(train_loss)],
'testlosses': [float(test_loss)] }
source1.stream(new_data1)
new_data2 = {'epochs': [epoch+1],
'train_accuracies': [float(train_accuracy)],
'test_accuracies': [float(test_accuracy)] }
source2.stream(new_data2)
push_notebook(handle=handle)
if test_accuracy > best_accuracy:
torch.save(model.state_dict(), BEST_MODEL_PATH)
best_accuracy = test_accuracy
# -
# Once that is finished, you should see a file ``best_model.pth`` in the Jupyter Lab file browser. Select ``Right click`` -> ``Download`` to download the model to your workstation
| notebooks/collision_avoidance/train_model_plot.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="jtLiiWFaZlUb"
#
# # **Paper Information**
#
#
# **TransGAN: Two Transformers Can Make One Strong GAN, and That Can Scale Up, CVPR 2021**, <NAME>, <NAME>, <NAME>
#
# * Paper Link: https://arxiv.org/pdf/2102.07074v2.pdf
# * Official Implementation: https://github.com/VITA-Group/TransGAN
# * Paper Presentation by <NAME> : https://www.youtube.com/watch?v=xwrUkHiDoiY
#
#
# **Project Group Members:**
#
#
# * <NAME>, <EMAIL>
# * <NAME>, <EMAIL>
# + [markdown] id="GUv7BXE1ZrWc"
# ## **Paper Summary**
# ### **Introduction**
# TransGAN is a transformer-based GAN model which can be considered as a pilot study as being completely free of convolutions. The architecture of TransGAN mainly consists of a memory-friendly transformer-based generator that progressively increases feature resolution, and correspondingly a patch-level discriminator that is also transformer-based. In training of the model, a series of techniques are combined in the original paper such as data augmentation, modified normalization, and relative position encoding to overcome the general training instability issues of the GAN. We implemented data augmentation [(Dosovitskiy et al., 2020)](https://arxiv.org/pdf/2010.11929.pdf), and relative position encoding in our work. In the original paper, performance of the model tested on different datasets such as STL-10, CIFAR-10, CelebA datasets and achieved competitive results compared to current state-of-the-art GANs using convolutions. In our project, we only tested our implementation on CIFAR10 dataset as we stated in our experimental result goals.
#
# ### **TransGAN Architecture**
# The architecture pipeline of TransGAN is shown below in the figure taken from the original paper.
#
# <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/images/transgan.jpg">
#
# Figure 1: The pipeline of the pure transform-based generator and discriminator of TransGAN.
#
# ### **Transformer Encoder as Basic Block**
# We used the transformer encoder [(Vaswani et al., 2017)](https://arxiv.org/pdf/1706.03762.pdf) as our basic block as in the original paper. An encoder is a composition of two parts. The first part is constructed by a multi-head self-attention module and the second part is a feed-forward MLP with GELU non-linearity. We apply layer normalization [(Ba et al., 2016)](https://arxiv.org/pdf/1607.06450.pdf) before both of the two parts.Both parts employ residual connection.
#
# $$Attention(Q, K, V ) = softmax(QK^T√d_k)V$$
# <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/images/vit.gif">
# Credits for illustration of ViT: [@lucidrains](https://github.com/lucidrains)
#
#
# ### **Memory-friendly Generator**
# In building the memory-friendly generator, TransGAN utilizes a common design philosophy in CNN-based GANs which iteratively upscale the resolution at multiple stages. Figure 1 (left) illustrates the memory-friendly generator which consists of multiple stages with several transformer blocks. At each stage, feature map resolution is gradually increased until it meets the target resolution *H × W*. The generator takes the random noise input and passes it through a multiple-layer perceptron (MLP). The output vector reshaped into a $H_0 × W_0$ resolution feature map (by default $H_0$ = $W_0$ = 8), each point a C-dimensional embedding. This “feature map" is next treated as a length-64 sequence of C-dimensional tokens, combined with the learnable positional encoding.
# Then, transformer encoders take embedding tokens as inputs and calculate the correspondence between each token recursively. To synthesize higher resolution images, we insert an upsampling module after each stage, consisting of a pixelshuffle [(Shi et al., 2016)](https://arxiv.org/pdf/1609.05158.pdf) module.
#
# ### **Tokenized-input for Discriminator**
# The authors design the discriminator as shown in Figure 1 (right) that it takes the patches of an image as inputs. Then, they split the input images $Y$ ∈ $R^{H×W×3}$ into 8x8 patches where each patch can be regarded as a "word". The patches are then converted to the 1D sequence of token embeddings through a linear flatten layer. After that, learnable position encoding is added, and tokens pass through the transformer encoder. Finally, tokens are taken by the classification head to output the real/fake prediction.
# + [markdown] id="hBtZpS3T-cl7"
# ### **Training the Model**
# In this section, we show our training code and training score for CIFAR-10 Dataset with the best performance hyperparameters that we found. We trained the largest model TransGAN-XL with data augmentation using different hyperparameters, and record the results in cifar/experiments folder.
# + [markdown] id="718NzrFLOUmr"
# ## Importing Libraries
# + id="0GEJLtggOUms" outputId="6b9e1d8e-f221-4914-ef7e-1971eddc823e"
from __future__ import division
from __future__ import print_function
import time
import argparse
import numpy as np
import torch
import torch.nn.functional as F
import torch.optim as optim
from torchvision.utils import make_grid, save_image
from tensorboardX import SummaryWriter
from tqdm import tqdm
from copy import deepcopy
from utils import *
from models import *
from fid_score import *
from inception_score import *
# !mkdir checkpoint
# !mkdir generated_imgs
# !pip install tensorboardX
# !mkdir fid_stat
# %cd fid_stat
# !wget bioinf.jku.at/research/ttur/ttur_stats/fid_stats_cifar10_train.npz
# %cd ..
# + [markdown] id="BNBeDCAXL6WI"
# ## Hyperparameters for CIFAR-10 Dataset
# Since Google Colab provides limited computational power, we decreased the generated_batch_size from 64 to 32, and we also run it for 10 epochs to show our pre-computed training scores. On our local GPU machine, we train the model with generated_batch_size is 64 and run for 200 epochs.
# + id="Xq4beSAoQSDA"
# training hyperparameters given by code author
lr_gen = 0.0001 #Learning rate for generator
lr_dis = 0.0001 #Learning rate for discriminator
latent_dim = 1024 #Latent dimension
gener_batch_size = 32 #Batch size for generator
dis_batch_size = 32 #Batch size for discriminator
epoch = 10 #Number of epoch
weight_decay = 1e-3 #Weight decay
drop_rate = 0.5 #dropout
n_critic = 5 #
max_iter = 500000
img_name = "img_name"
lr_decay = True
# architecture details by authors
image_size = 32 #H,W size of image for discriminator
initial_size = 8 #Initial size for generator
patch_size = 4 #Patch size for generated image
num_classes = 1 #Number of classes for discriminator
output_dir = 'checkpoint' #saved model path
dim = 384 #Embedding dimension
optimizer = 'Adam' #Optimizer
loss = "wgangp_eps" #Loss function
phi = 1 #
beta1 = 0 #
beta2 = 0.99 #
diff_aug = "translation,cutout,color" #data augmentation
# + [markdown] id="I7iL1KYpMyx5"
# ## Training & Saving Model for CIFAR-10
# As we mentioned above we run the training for 10 epochs due to limitation of Google Colab and showed the decrease in FID score from 253 to 138 in 10 epochs.
# + colab={"base_uri": "https://localhost:8080/"} id="cS5Oxb7WcAts" outputId="90caa77c-f93d-48c1-cc46-1e24c5357d09"
if torch.cuda.is_available():
dev = "cuda:0"
else:
dev = "cpu"
device = torch.device(dev)
generator= Generator(depth1=5, depth2=4, depth3=2, initial_size=8, dim=384, heads=4, mlp_ratio=4, drop_rate=0.5)#,device = device)
generator.to(device)
discriminator = Discriminator(diff_aug = diff_aug, image_size=32, patch_size=4, input_channel=3, num_classes=1,
dim=384, depth=7, heads=4, mlp_ratio=4,
drop_rate=0.5)
discriminator.to(device)
generator.apply(inits_weight)
discriminator.apply(inits_weight)
# + colab={"base_uri": "https://localhost:8080/"} id="-FrwqrkneKdO" outputId="00d3377d-b405-4bec-b98e-659b03dc1c4a"
if optimizer == 'Adam':
optim_gen = optim.Adam(filter(lambda p: p.requires_grad, generator.parameters()), lr=lr_gen, betas=(beta1, beta2))
optim_dis = optim.Adam(filter(lambda p: p.requires_grad, discriminator.parameters()),lr=lr_dis, betas=(beta1, beta2))
elif optimizer == 'SGD':
optim_gen = optim.SGD(filter(lambda p: p.requires_grad, generator.parameters()),
lr=lr_gen, momentum=0.9)
optim_dis = optim.SGD(filter(lambda p: p.requires_grad, discriminator.parameters()),
lr=lr_dis, momentum=0.9)
elif optimizer == 'RMSprop':
optim_gen = optim.RMSprop(filter(lambda p: p.requires_grad, discriminator.parameters()),
lr=lr_dis, eps=1e-08, weight_decay=weight_decay, momentum=0, centered=False)
optim_dis = optim.RMSprop(filter(lambda p: p.requires_grad, discriminator.parameters()), lr=lr_dis, eps=1e-08, weight_decay=weight_decay, momentum=0, centered=False)
gen_scheduler = LinearLrDecay(optim_gen, lr_gen, 0.0, 0, max_iter * n_critic)
dis_scheduler = LinearLrDecay(optim_dis, lr_dis, 0.0, 0, max_iter * n_critic)
#RMSprop(params, lr=0.01, alpha=0.99, eps=1e-08, weight_decay=0, momentum=0, centered=False)
print("optimizer:",optimizer)
fid_stat = 'fid_stat/fid_stats_cifar10_train.npz'
writer=SummaryWriter()
writer_dict = {'writer':writer}
writer_dict["train_global_steps"]=0
writer_dict["valid_global_steps"]=0
# + id="oqrgmomNf01n"
def compute_gradient_penalty(D, real_samples, fake_samples, phi):
"""Calculates the gradient penalty loss for WGAN GP"""
# Random weight term for interpolation between real and fake samples
alpha = torch.Tensor(np.random.random((real_samples.size(0), 1, 1, 1))).to(real_samples.get_device())
# Get random interpolation between real and fake samples
interpolates = (alpha * real_samples + ((1 - alpha) * fake_samples)).requires_grad_(True)
d_interpolates = D(interpolates)
fake = torch.ones([real_samples.shape[0], 1], requires_grad=False).to(real_samples.get_device())
# Get gradient w.r.t. interpolates
gradients = torch.autograd.grad(
outputs=d_interpolates,
inputs=interpolates,
grad_outputs=fake,
create_graph=True,
retain_graph=True,
only_inputs=True,
)[0]
gradients = gradients.contiguous().view(gradients.size(0), -1)
gradient_penalty = ((gradients.norm(2, dim=1) - phi) ** 2).mean()
return gradient_penalty
def train(noise,generator, discriminator, optim_gen, optim_dis,
epoch, writer, schedulers, img_size=32, latent_dim = latent_dim,
n_critic = n_critic,
gener_batch_size=gener_batch_size, device="cuda:0"):
writer = writer_dict['writer']
gen_step = 0
generator = generator.train()
discriminator = discriminator.train()
transform = transforms.Compose([transforms.Resize(size=(img_size, img_size)),transforms.RandomHorizontalFlip(),transforms.ToTensor(),transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
train_set = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
train_loader = torch.utils.data.DataLoader(dataset=train_set, batch_size=30, shuffle=True)
for index, (img, _) in enumerate(train_loader):
global_steps = writer_dict['train_global_steps']
real_imgs = img.type(torch.cuda.FloatTensor)
noise = torch.cuda.FloatTensor(np.random.normal(0, 1, (img.shape[0], latent_dim)))#noise(img, latent_dim)#= args.latent_dim)
optim_dis.zero_grad()
real_valid=discriminator(real_imgs)
fake_imgs = generator(noise).detach()
#assert fake_imgs.size() == real_imgs.size(), f"fake_imgs.size(): {fake_imgs.size()} real_imgs.size(): {real_imgs.size()}"
fake_valid = discriminator(fake_imgs)
if loss == 'hinge':
loss_dis = torch.mean(nn.ReLU(inplace=True)(1.0 - real_valid)).to(device) + torch.mean(nn.ReLU(inplace=True)(1 + fake_valid)).to(device)
elif loss == 'wgangp_eps':
gradient_penalty = compute_gradient_penalty(discriminator, real_imgs, fake_imgs.detach(), phi)
loss_dis = -torch.mean(real_valid) + torch.mean(fake_valid) + gradient_penalty * 10 / (phi ** 2)
loss_dis.backward()
optim_dis.step()
writer.add_scalar("loss_dis", loss_dis.item(), global_steps)
if global_steps % n_critic == 0:
optim_gen.zero_grad()
if schedulers:
gen_scheduler, dis_scheduler = schedulers
g_lr = gen_scheduler.step(global_steps)
d_lr = dis_scheduler.step(global_steps)
writer.add_scalar('LR/g_lr', g_lr, global_steps)
writer.add_scalar('LR/d_lr', d_lr, global_steps)
gener_noise = torch.cuda.FloatTensor(np.random.normal(0, 1, (gener_batch_size, latent_dim)))
generated_imgs= generator(gener_noise)
fake_valid = discriminator(generated_imgs)
gener_loss = -torch.mean(fake_valid).to(device)
gener_loss.backward()
optim_gen.step()
writer.add_scalar("gener_loss", gener_loss.item(), global_steps)
gen_step += 1
#writer_dict['train_global_steps'] = global_steps + 1
if gen_step and index % 100 == 0:
sample_imgs = generated_imgs[:25]
img_grid = make_grid(sample_imgs, nrow=5, normalize=True, scale_each=True)
save_image(sample_imgs, f'generated_images/generated_img_{epoch}_{index % len(train_loader)}.jpg', nrow=5, normalize=True, scale_each=True)
tqdm.write("[Epoch %d] [Batch %d/%d] [D loss: %f] [G loss: %f]" %
(epoch+1, index % len(train_loader), len(train_loader), loss_dis.item(), gener_loss.item()))
# + id="wA0coJ7kgE5X"
def validate(generator, writer_dict, fid_stat):
writer = writer_dict['writer']
global_steps = writer_dict['valid_global_steps']
generator = generator.eval()
fid_score = get_fid(fid_stat, epoch, generator, num_img=5000, val_batch_size=60*2, latent_dim=1024, writer_dict=None, cls_idx=None)
print(f"FID score: {fid_score}")
writer.add_scalar('FID_score', fid_score, global_steps)
writer_dict['valid_global_steps'] = global_steps + 1
return fid_score
# + colab={"base_uri": "https://localhost:8080/"} id="IvwXcTahgPAV" outputId="b92e59f3-23d2-42f1-cf40-c7cb1f2e42e4"
best = 1e4
for epoch in range(epoch):
lr_schedulers = (gen_scheduler, dis_scheduler) if lr_decay else None
train(noise, generator, discriminator, optim_gen, optim_dis,
epoch, writer, lr_schedulers,img_size=32, latent_dim = latent_dim,
n_critic = n_critic,
gener_batch_size=gener_batch_size)
checkpoint = {'epoch':epoch, 'best_fid':best}
checkpoint['generator_state_dict'] = generator.state_dict()
checkpoint['discriminator_state_dict'] = discriminator.state_dict()
score = validate(generator, writer_dict, fid_stat)
print(f'FID score: {score} - best ID score: {best} || @ epoch {epoch+1}.')
if epoch == 0 or epoch > 30:
if score < best:
save_checkpoint(checkpoint, is_best=(score<best), output_dir=output_dir)
print("Saved Latest Model!")
best = score
checkpoint = {'epoch':epoch, 'best_fid':best}
checkpoint['generator_state_dict'] = generator.state_dict()
checkpoint['discriminator_state_dict'] = discriminator.state_dict()
score = validate(generator, writer_dict, fid_stat) ####CHECK AGAIN
save_checkpoint(checkpoint,is_best=(score<best), output_dir=output_dir)
# + [markdown] id="fmPPrYITnQqf"
# ### **Experimental Result Goals vs. Achieved Results**
# In this project, we aimed to reproduce qualitative results(generating image samples by CIFAR-10 Dataset) and quantitative results in Table 2 and Table 4 of the original paper that shown below.
# <table>
# <tr>
# <td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/table2.png" style="width: 400px;"/> </td>
# <td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/table4.png" style="width: 400px;"/> </td>
# </tr></table>
#
# Since we have limited computational resource and time for the training all size of TransGAN model on CIFAR-10 Dataset, we only trained the largest model with data augmentation, TransGAN-XL, for Table 4 results.
# + [markdown] id="WpMhUguJaL4o"
# ## Test Model and Results
# In this section, we loaded pre-trained model and got the following qualitative and quantitative results.
#
# ### Qualitative Results
# The following pictures show our generated images at different epoch numbers.
# <table>
# <tr>
# <td style="text-align: center">0 Epoch</td>
# <td style="text-align: center">40 Epoch</td>
# <td style="text-align: center">100 Epoch</td>
# <td style="text-align: center">200 Epoch</td>
# </tr>
# <trt>
# <p align="center"><img width="30%" src="https://raw.githubusercontent.com/asarigun/TransGAN/main/images/atransgan_cifar.gif"></p>
# </tr>
# <tr>
# <td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/0.jpg" style="width: 400px;"/> </td>
# <td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/40.jpg" style="width: 400px;"/> </td>
# <td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/100.jpg" style="width: 400px;"/> </td>
# <td> <img src="https://raw.githubusercontent.com/asarigun/TransGAN/main/results/200.jpg" style="width: 400px;"/> </td>
# </tr>
# </table>
#
# ### Quantitative Results
# As we mentioned above, due to the lack of computational resource, we did our experiments only with the largest model TransGAN-XL and get the following results. We had decided not to implement 'Co-Training with Self-Supervised Auxiliary Task' and 'Locality-Aware Initialization for Self-Attention' since they made only small differences as shown in the paper. The difference between our result and original paper result can be originated in using some different hyperparameters and abovementioned implementation differences. You can see our quantitative result, FID score 26.82, [here](https://github.com/asarigun/TransGAN/blob/main/results/wgangp_eps_optim_Adam_lr_gen_0_0001_lr_dis_0_0001_epoch_200.txt).
#
#
# + [markdown] id="A7FBDk0Xbc4k"
# ## Challenges and Discussions
#
# Since the authors did not give detailed hyperparameters for each Transformers Block and Multi-Head Attention Mechanism on version 1, we needed to find the best hyperparameters. Also, in the training part, they did not give detailed hyperparameters such as droprate, weight decay, or batch normalization in version 1. But in the last version of the original paper, authors gave more detailed hyperparameters for training, therefore we got more reasonable results.
#
# During the implementation, first we used Hinge loss and faced convergence problem in training. When we tried another loss function, WGAN-GP, that is mentioned in the last version of original paper, we achieved to overcome convergence problem and got better results.
#
# As authors didn't share detailed training process in their previous version, we struggled to converge FID score during training. But in the latest version of the original paper, authors provided more details for training so that we achieved to converge FID score in the training.
#
# Due to lack of computational resource, we only trained the largest model, TransGAN-XL in our project. We implemented data augmentation in our model as it is considered crucial for TransGAN in the original paper. We didn't implement 'Co-Training with Self-Supervised Auxiliary Task' and 'Locality-Aware Initialization for Self-Attention' since they make only small differences as shown in the paper.
# + [markdown] id="uC3MAynNh51x"
# ## Citation
# ```
# @article{jiang2021transgan,
# title={TransGAN: Two Transformers Can Make One Strong GAN},
# author={<NAME> and <NAME> <NAME>},
# journal={arXiv preprint arXiv:2102.07074},
# year={2021}
# }
# ```
# ```
# @article{dosovitskiy2020,
# title={An Image is Worth 16x16 Words: Transformers for Image Recognition at Scale},
# author={Dosovitskiy, Alexey and Beyer, Lucas and Kolesnikov, Alexander and Weissenborn, <NAME>, Xiaohua and Unterthiner, Thomas and Dehghani, Mostafa and Minderer, Matthias and <NAME> and Gelly, Sylvain and Uszkoreit, Jakob and <NAME>},
# journal={arXiv preprint arXiv:2010.11929},
# year={2020}
# }
# ```
# ```
# @inproceedings{zhao2020diffaugment,
# title={Differentiable Augmentation for Data-Efficient GAN Training},
# author={<NAME> <NAME>},
# booktitle={Conference on Neural Information Processing Systems (NeurIPS)},
# year={2020}
# }
# ```
#
| main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py37]
# language: python
# name: conda-env-py37-py
# ---
import pandas as pd
import string
url = "/Users/Artem/Documents/CS 205/NHAHES_VAR_LIST/NHANES Demographics Variable List.html"
df_raw = pd.read_html(url, attrs = {'id' : 'GridView1'})[0]
df_raw.head()
# +
#exclude restricted vars
df_raw = df_raw[df_raw["Use Constraints"] == 'None']
#drop extra columns
drop_columns = ['Variable Description', 'Data File Description', 'Begin Year', 'EndYear', 'Use Constraints']
df_useful = df_raw.drop(drop_columns, axis = 1)
df_useful.columns = ["feature", "file", 'folder']
df_useful.head()
# -
#remove year-specific file names
df_useful.file = df_useful.file.apply(func = lambda x: x[:-2] if x[-2] == '_' else x)
df_useful.drop_duplicates(inplace = True)
df_useful.size
#rewrite everythig we did above as a function
def parse_file(url):
df_raw = pd.read_html(url, attrs = {'id' : 'GridView1'})[0]
df_raw = df_raw[df_raw["Use Constraints"] == 'None']
drop_columns = ['Variable Description', 'Data File Description', 'Begin Year', 'EndYear', 'Use Constraints']
df_useful = df_raw.drop(drop_columns, axis = 1)
df_useful.columns = ["feature", "file", 'folder']
df_useful.file = df_useful.file.apply(func = lambda x: x[:-2] if x[-2] == '_' else x)
df_useful.drop_duplicates(inplace = True)
return df_useful
# +
all_features = pd.DataFrame(columns = ('feature', 'file', 'folder'))
for fname in ["Demographics", "Dietary", "Examination", "Laboratory", "Questionnaire"]:
url = "/Users/Artem/Documents/CS 205/NHAHES_VAR_LIST/NHANES {} Variable List.html".format(fname)
print("Parsing", url)
temp_df = parse_file(url)
all_features = all_features.merge(temp_df, how = 'outer')
print(all_features.shape)
all_features.head()
# -
all_features = all_features[all_features.feature != 'SEQN']
print(all_features.shape)
all_features.to_csv("/Users/Artem/Documents/GitHub/CancerPrediction/all_features.csv", index = False)
original_df.to_pickle("./dummy.pkl")
| NHANES_list_parser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python 3
# ## Version
import sys
print(sys.version)
# ## Spark
import os
from pyspark import SparkContext
from pyspark import SparkConf
from pyspark.sql import SparkSession
from pyspark.sql.functions import *
from pyspark.sql.types import *
spark_home = os.environ.get('SPARK_HOME', None)
print(spark_home)
spark = SparkSession.builder.master("local[*]").appName("spark")
spark = spark.config("spark.driver.memory", "8g")
spark = spark.config("spark.executor.memory", "8g")
spark = spark.config("spark.python.worker.memory", "8g")
spark = spark.getOrCreate()
# !cat /usr/local/spark/examples/src/main/resources/people.json
df = spark.read.json("{}/examples/src/main/resources/people.json".format(spark_home))
df.show()
spark.stop()
| volume/example/python3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sympy import init_session
init_session()
# ## Quadratic
# We want to construct a quadratic polynomal through that points $x_{i-1}$, $x_i$, $x_{i+1}$ that gives the correct averages,
# $f_{i-1}$, $f_i$, and $f_{i-1}$ when integrated over the volume, e.g.
#
# $$\frac{1}{\Delta x} \int_{x_{i-1/2}}^{x_{i+1/2}} f(x) dx = f_i$$
#
# There are 3 unknowns in the quadratic and three constraints, so this is a linear system we can solve.
# Define the quadratic polynomial
a, b, c = symbols("a b c")
x0 = symbols("x0")
f = a*(x-x0)**2 + b*(x-x0) + c
f
dx = symbols("\Delta")
# ### constraints
# Define the 3 constraint equations---here we set them up construct $A$, $B$, and $C$ as the integrals over the 3 control volumes
fm, f0, fp = symbols("f_{i-1} f_i f_{i+1}")
#xm32, xm12, xp12, xp32 = symbols("x_{i-3/2} x_{i-1/2} x_{i+1/2} x_{i+3/2}")
xm32 = x0 - Rational(3,2)*dx
xm12 = x0 - Rational(1,2)*dx
xp12 = x0 + Rational(1,2)*dx
xp32 = x0 + Rational(3,2)*dx
# interfaces
xm32, xm12, xp12, xp32
A = simplify(integrate(f/dx, (x, xm32, xm12)))
B = simplify(integrate(f/dx, (x, xm12, xp12)))
C = simplify(integrate(f/dx, (x, xp12, xp32)))
# The analytic forms of the integrals
A, B, C
# Our linear system is now:
#
# $$A = f_{i-1}$$
# $$B = f_i$$
# $$C = f_{i+1}$$
# Now find the coefficients of the polynomial
coeffs = solve([A-fm, B-f0, C-fp], [a,b,c])
coeffs
# And in pretty form, here's the polynomial
f.subs(a,coeffs[a]).subs(b,coeffs[b]).subs(c,coeffs[c])
# ### Cubic
# We want to construct a cubic polynomal through that points $x_{i-2}$, $x_{i-1}$, $x_i$, $x_{i+1}$ that gives the correct averages,
# $f_{i-2}$, $f_{i-1}$, $f_i$, and $f_{i-1}$ when integrated over the volume of each zone
a, b, c, d = symbols("a b c d")
f = a*(x-x0)**3 + b*(x-x0)**2 + c*(x-x0) + d
f
# Now perform the integals of $f(x)$ over each zone
fm2, fm, f0, fp = symbols("f_{i-2} f_{i-1} f_i f_{i+1}")
xm52 = x0 - Rational(5,2)*dx
xm32 = x0 - Rational(3,2)*dx
xm12 = x0 - Rational(1,2)*dx
xp12 = x0 + Rational(1,2)*dx
xp32 = x0 + Rational(3,2)*dx
# interfaces
xm52, xm32, xm12, xp12, xp32
A = simplify(integrate(f/dx, (x, xm52, xm32)))
B = simplify(integrate(f/dx, (x, xm32, xm12)))
C = simplify(integrate(f/dx, (x, xm12, xp12)))
D = simplify(integrate(f/dx, (x, xp12, xp32)))
A, B, C, D
coeffs = solve([A-fm2, B-fm, C-f0, D-fp], [a,b,c,d], check=False)
coeffs
# and the pretty form of the polynomial
fc = f.subs(a,coeffs[a]).subs(b,coeffs[b]).subs(c,coeffs[c]).subs(d,coeffs[d])
fc
# this interpolant is symmetric about the $i-1/2$ interface---let's see the value there
fc.subs(x,x0-Rational(1,2)*dx)
# Note that this is the interpolating polynomial used to find the interface states in PPM (Colella & Woodward 1984)
# ## Quartic
# Now we define a quartic polynomial that gives the correct averages over 5 zones, $x_{i-2}$, $x_{i-1}$, $x_i$, $x_{i+1}$, $x_{i+2}$,
# with zone averages $f_{i-2}$, $f_{i-1}$, $f_i$, $f_{i+1}$, $f_{i+2}$
a, b, c, d, e = symbols("a b c d e")
x0 = symbols("x0")
f = a*(x-x0)**4 + b*(x-x0)**3 + c*(x-x0)**2 + d*(x-x0) + e
f
# Now we perform the integrals of $f(x)$ over each zone
fm2, fm, f0, fp, fp2 = symbols("f_{i-2} f_{i-1} f_i f_{i+1} f_{i+2}")
#xm32, xm12, xp12, xp32 = symbols("x_{i-3/2} x_{i-1/2} x_{i+1/2} x_{i+3/2}")
xm52 = x0 - Rational(5,2)*dx
xm32 = x0 - Rational(3,2)*dx
xm12 = x0 - Rational(1,2)*dx
xp12 = x0 + Rational(1,2)*dx
xp32 = x0 + Rational(3,2)*dx
xp52 = x0 + Rational(5,2)*dx
# interfaces
xm52, xm32, xm12, xp12, xp32, xp52
A = simplify(integrate(f/dx, (x, xm52, xm32)))
B = simplify(integrate(f/dx, (x, xm32, xm12)))
C = simplify(integrate(f/dx, (x, xm12, xp12)))
D = simplify(integrate(f/dx, (x, xp12, xp32)))
E = simplify(integrate(f/dx, (x, xp32, xp52)))
# The analytic form of the constraints
A, B, C, D, E
# Now find the coefficients
coeffs = solve([A-fm2, B-fm, C-f0, D-fp, E-fp2], [a,b,c,d,e], check=False)
coeffs
# and the pretty form of the polynomial
f.subs(a,coeffs[a]).subs(b,coeffs[b]).subs(c,coeffs[c]).subs(d,coeffs[d]).subs(e,coeffs[e])
| finite-volume/conservative-interpolation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import jax.numpy as np
from jax import value_and_grad, jit
import numpy as onp
import matplotlib.pyplot as plt
# %matplotlib inline
# +
###Original softmax:
# @jit
# def softmax(x):
# x = x + np.diag(np.ones(30)*np.inf)
# """Compute softmax values for each sets of scores in x."""
# e_x = np.exp(x - np.max(x))
# return e_x / e_x.sum()
###Softmax that sets distances to self to -np.inf, meaning the probability goes to zero.
@jit
def softmax(x):
"""Compute softmax values for each sets of scores in x."""
e_x = np.exp((x - np.diag(np.ones(x.shape[0])*np.inf)) - np.max(x))
return e_x / e_x.sum()
#E_fn is just a pairwise distance function,
#stole from https://github.com/google/jax/issues/787 .
@jit
def E_fn(conf):
ri = np.expand_dims(conf, 0)
rj = np.expand_dims(conf, 1)
dxdydz = np.power(ri - rj, 2)
#return squared euclidean:
dij = np.sum(dxdydz, axis=-1)
return dij
@jit
def loss(A, X, y_mask, zero_mask):
embedding = X.dot(A.T)
distances = E_fn(embedding)
p_ij = softmax(-distances)
p_ij_mask = p_ij * y_mask
p_i = p_ij_mask.sum(1)
loss = -p_i.sum()
logprobs = np.log(p_i[zero_mask])
clf_loss = -np.sum(logprobs)
diff_class_distances = distances * ~y_mask
hinge_loss = (np.clip(1- diff_class_distances, 0, np.inf)**2).sum(1).mean()
return clf_loss + hinge_loss
# -
# +
def make_circle(r, num_samples):
t = onp.linspace(0, 2*np.pi, num_samples)
xc, yc = 0, 0 # circle center coordinates
x = r*np.cos(t) + 0.2*onp.random.randn(num_samples) + xc
y = r*np.sin(t) + 0.2*onp.random.randn(num_samples) + yc
return x, y
def gen_data(num_samples, num_classes, mean, std):
"""Generates the data.
"""
num_samples_per = num_samples // num_classes
X = []
y = []
for i, r in enumerate(range(num_classes)):
# first two dimensions are that of a circle
x1, x2 = make_circle(r+1.5, num_samples_per)
# third dimension is Gaussian noise
x3 = std*onp.random.randn(num_samples_per) + mean
X.append(onp.stack([x1, x2, x3]))
y.append(onp.repeat(i, num_samples_per))
X = np.concatenate(X, axis=1)
y = np.concatenate(y)
indices = list(range(X.shape[1]))
onp.random.shuffle(indices)
X = X[:, indices]
y = y[indices]
X = X.T # make it (N, D)
return X, y
a, b = gen_data(1000, 4, 0, 1)
# -
# # nesterov momentum:
# +
D = 3
d = 2
A = onp.random.randn(d,D)*0.1
V = np.zeros_like(A)
mass = 0.1
ayze = list()
velocities = list()
lr = 0.0001 #step size
values = list()
for _ in range(300):
idx = onp.random.choice(range(a.shape[0]), 100, replace=False)
batch = a[idx]
labels = b[idx]
y_mask = (labels[:, None] == labels[None, :])
zero_mask = onp.array(y_mask.sum(1)==1)
pos_idx = (~zero_mask).nonzero()[0]
value, g = value_and_grad(loss)(A,batch,y_mask, pos_idx)
values.append(value)
V = mass * V + g
velocities.append(V)
ayze.append(A)
A = A - lr*(mass*V+g)
#print(value)
# -
plt.plot(values)
# # adadelta
# +
D = 3
d = 2
A = onp.random.randn(d,D)*0.1
g_sq = np.zeros_like(A)
m = np.zeros_like(A)
mass = 1
ayze = list()
g_squares = list()
lr = 0.1 #step size
momentum=0.9
values = list()
for _ in range(300):
idx = onp.random.choice(range(a.shape[0]), 100, replace=False)
batch = a[idx]
labels = b[idx]
y_mask = (labels[:, None] == labels[None, :])
zero_mask = onp.array(y_mask.sum(1)==1)
pos_idx = (~zero_mask).nonzero()[0]
value, g = value_and_grad(loss)(A,batch,y_mask, pos_idx)
values.append(value)
g_sq += g**2
g_sq_inv_sqrt = np.where(g_sq > 0, 1. / np.sqrt(g_sq), 0.0)
m = (1. - momentum) * (g * g_sq_inv_sqrt) + momentum * m
A = A - lr * m
ayze.append(A)
# -
plt.plot(values)
# +
D = 3
d = 2
A = onp.random.randn(d,D)*0.1
avg_sq_grad = np.zeros_like(A)
gamma=0.9
eps=1e-8
mass = 1
ayze = list()
g_squares = list()
lr = 0.001 #step size
values = list()
for _ in range(1000):
idx = onp.random.choice(range(a.shape[0]), 100, replace=False)
batch = a[idx]
labels = b[idx]
y_mask = (labels[:, None] == labels[None, :])
zero_mask = onp.array(y_mask.sum(1)==1)
pos_idx = (~zero_mask).nonzero()[0]
value, g = value_and_grad(loss)(A,batch,y_mask, pos_idx)
values.append(value)
avg_sq_grad = avg_sq_grad * gamma + g**2 * (1. - gamma)
A = A - lr * g / np.sqrt(avg_sq_grad + eps)
ayze.append(A)
#print(value)
# -
plt.plot(values)
# # plot:
# +
fig, ax = plt.subplots(1,2)
fig.set_figwidth(10)
emb = a.dot(ayze[0].T)
ax[0].scatter(emb[:,0], emb[:,1], c=b, cmap='Spectral')
emb = a.dot(ayze[200].T)
ax[1].scatter(emb[:,0], emb[:,1], c=b, cmap='Spectral')
# -
# # save as gif:
# +
def update_plot(i, data, scplot):
A = data[i]
emb = a.dot(A.T)
scplot.set_offsets(emb)
ax.set_xlim(emb[:,0].min()+0.1*emb[:,0].min(), emb[:,0].max()+0.1*emb[:,0].max())
ax.set_ylim(emb[:,1].min()+0.1*emb[:,1].min(), emb[:,1].max()+0.1*emb[:,1].max())
return (scplot,)
emb = a.dot(ayze[0].T)
xp = emb[:,0]
yp = emb[:,1]
fig, ax = plt.subplots()
ax.set_xticklabels([])
ax.set_yticklabels([])
scplot = ax.scatter(xp, yp, c=b, cmap='Spectral')
ani = animation.FuncAnimation(fig, update_plot, frames=1000,
fargs=(ayze, scplot), blit=True)
ani.save('./animation.gif', writer='imagemagick', fps=20)
| NCA in jax example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set_style("whitegrid")
plt.rcParams["patch.force_edgecolor"]=True
from plotly.offline import plot
import plotly.graph_objs as go
import plotly.plotly as py
pd.set_option('display.max_columns', 500)
pd.set_option('display.max_rows', 500)
df= pd.read_csv('globaltd.csv',encoding='ISO-8859-1',low_memory=False)
df.head(3)
df.rename(columns={'iyear': 'Year','imonth':'Month','iday':'Day','country_txt':'Country','region_txt':'Region','city':'City',
'attacktype1_txt':'Type of Attack','targtype1_txt':'Target','target1':'Target Person','natlty1_txt':'Target Nationality',
'weaptype1_txt':'Weapon','gname':'Name of Group','nkill':'Killed','nwound':'Wounded','suicide':'Suicide','summary':'Summary'},inplace=True)
terror= df[['Year','Month','Day','Country','Region','City','Type of Attack','Target','Target Person','Target Nationality',
'Weapon','Name of Group','Killed','Wounded','Suicide','latitude','longitude','Summary','motive']]
terror.isnull().sum()
# +
plt.figure(figsize=(20,10))
x= terror.isnull().sum()
y= terror.columns
sns.barplot(x,y)
plt.xlabel('Count', fontsize=18)
plt.ylabel('Attributes', fontsize=16)
plt.tick_params(axis='both',labelsize=15)
plt.title('Missing Values by Columns',fontdict={'fontsize':40})
# -
plt.figure(figsize=(10,7))
cityterror=terror['City'].value_counts().head(11)
sns.barplot(y=cityterror.index,x=cityterror.values)
plt.xlabel('Count', fontsize=18)
plt.ylabel('City', fontsize=16)
plt.tick_params(axis='both',labelsize=15)
plt.title('Top Terrorist Attacks by City',fontdict={'fontsize':40})
khi = terror[terror['City']=='Karachi']
khi.isnull().sum()
# +
## Data for karachi seems relatively adequate to perform analysis
# -
khi.head(3)
# +
kwg= khi.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=kwg.index,
y=kwg.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=kwg.index,
y=kwg.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People Killed/Wounded in Karachi Attacks')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# -
plt.figure(figsize=(20,10))
sns.countplot(x='Year',hue='Suicide',data=khi)
plt.title('Attacks in Karachi',fontdict={'fontsize':40})
plt.tick_params(axis='y',labelsize=15)
plt.ylabel('Count', fontsize=10)
plt.legend(title='Suicide = 1',prop={'size':30})
# +
## Most attacks were committed in 1995
## 90s and 2010s saw the highest number of attacks
## Suicide attacks only after 2000s
# -
khis=khi[khi['Suicide']==1]
plt.figure(figsize=(20,10))
sns.countplot(x='Year',hue='Suicide',data=khis,color='#eeefff')
plt.title('Suicide Attacks in Karachi',fontdict={'fontsize':40})
plt.tick_params(axis='both',labelsize=15)
plt.ylabel('Count', fontsize=10)
# +
## Records in data base for suicide attacks only have data till 2015
## most recorded in 2014
## trend shows increasing attacks since 2002.
# -
sns.countplot(x=khi['Suicide'])
# +
## >95% attacks in karachi are classified as non suicidal attacks
## All suicide attacks were 2002 onwards
# -
foreign= (khi[khi['Target Nationality'] != 'Pakistan']['Target Nationality']).value_counts()
foreign1= khi[khi['Target Nationality'] != 'Pakistan']
# +
ff1= foreign1.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=ff1.index,
y=ff1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=ff1.index,
y=ff1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People(Foreign/Local) hurt in Karachi Attacks- (Foreign Nationals Targeted)')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# -
plt.figure(figsize=(10,7))
sns.barplot(y=foreign.index,x=foreign.values)
plt.xlabel('Count', fontsize=18)
plt.ylabel('Country', fontsize=16)
plt.tick_params(axis='both',labelsize=15)
plt.title('Nationalities Targeted in Karachi',fontdict={'fontsize':40})
sui= khi[(khi['Target Nationality'] != 'Pakistan')&(khi['Suicide']==1)]
sui[['Year','Target','Target Person','Target Nationality','Name of Group','Killed','Wounded']]
# +
## 3 suicide attacks in which foreigners were targeted. 2 in 2002 and 1 in 2006.
# -
ustarget= khi[khi['Target Nationality']== 'United States']
ustarget.head(3)
# +
uss1= ustarget.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=uss1.index,
y=uss1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=uss1.index,
y=uss1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People(Foreign/Local) hurt in Karachi Attacks- (US Nationals Targeted)')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# +
## Above is a dataframe for all those attacks in which US nationals were targetted
# -
ustype= ustarget['Type of Attack'].value_counts()
trace = go.Pie(labels=ustype.index,values=ustype.values)
layout = go.Layout(title='Type of Attacks on US nationals')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart1')
# +
## Types of attacks on US targets
# -
uskill= ustarget.groupby('Year')['Killed'].sum()
plt.figure(figsize=(10,7))
sns.barplot(y=uskill.values,x=uskill.index)
plt.xlabel('Count', fontsize=18)
plt.ylabel('City', fontsize=16)
plt.tick_params(axis='both',labelsize=15)
plt.title('People killed by the targeting US nationals',fontdict={'fontsize':40})
uswound= ustarget.groupby('Year')['Wounded'].sum()
plt.figure(figsize=(10,7))
sns.barplot(y=uswound.values,x=uswound.index)
plt.xlabel('Count', fontsize=18)
plt.ylabel('City', fontsize=16)
plt.tick_params(axis='both',labelsize=15)
plt.title('People Wounded by attacks targetting US nationals',fontdict={'fontsize':40})
# +
ustype1=ustarget['Target'].value_counts()
trace = go.Pie(labels=ustype1.index,values=ustype1.values)
layout = go.Layout(title='Target of Attacks on US nationals')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart21')
# +
## Targets of attacks on US nationals in karachi
## The only 2 suicide attacks on US nationals were done on Government Diplomatic buildings
# -
khi['Name of Group'].value_counts().head(10)
khigroup= (khi[khi['Name of Group'] != 'Unknown']['Name of Group']).value_counts().head(10)
plt.figure(figsize=(10,7))
sns.barplot(y=khigroup.index,x=khigroup.values)
plt.xlabel('Count', fontsize=18)
plt.ylabel('Name of Group', fontsize=16)
plt.tick_params(axis='both',labelsize=15)
plt.title('Top Groups involved in attacks',fontdict={'fontsize':40})
# +
## Interesting to see MQM at number 2
## MQM claims to be a national political party that has often placed itself in top 5 parties of the country
## Need to see what type of attacks this political party was involved in its headquarters Karachi
# -
mqm= khi[khi['Name of Group'] == 'Muttahida Qami Movement (MQM)']
mqm.head(3)
trace = go.Pie(labels=mqm['Target'])
layout = go.Layout(title='Target of Attacks by MQM')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart1')
trace = go.Pie(labels=mqm['Type of Attack'])
layout = go.Layout(title='Types of Attacks by MQM')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart2')
trace = go.Pie(labels=mqm['Weapon'])
layout = go.Layout(title='Types of Weapons used by MQM')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart2')
mqm['Killed'].value_counts().sum()
# +
## 169 recorded fatilities by MQM..
## This political party appears to be more violent than the biggest terrorist organization in the country
# -
mqm['Suicide'].value_counts()
# +
## No suicide attacks by MQM
# +
## Moving on to TTP- Tehrik e Taliban Pakista, the most notorious and dangerous terrorist outfit in the country.
## TTP does not take part in elections like MQM does.
# -
ttp = khi[khi['Name of Group'] == 'Tehrik-i-Taliban Pakistan (TTP)']
ttp['Suicide'].value_counts()
# +
## 15 suicide attacks recorded for TTP in Karachi.( data only records suicide attacks up till 2015)
# -
trace = go.Pie(labels=ttp['Target'])
layout = go.Layout(title='Target of Attacks by TTP')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart5')
# +
## TTP attacks almost everyone in the city
## Law enforcement agencies- Police and Army make up half of their targets
# -
trace = go.Pie(labels=ttp['Type of Attack'])
layout = go.Layout(title='Types of Attacks by TTP')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart25')
# +
## TTPs signature style of attacks appears to be bombings/explosions.
# -
trace = go.Pie(labels=ttp['Weapon'])
layout = go.Layout(title='Types of Weapons used by TTP')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart2')
# +
## TTP only likes to use explosives and firarms
# -
ttp.head(3)
ttp['Killed'].sum()
ttp['Wounded'].sum()
# +
## 446 fatalities and over 1000 were wounded by TTP- the most violent and dangerous terrorist organization in the country
# +
ttp1= ttp.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=ttp1.index,
y=ttp1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=ttp1.index,
y=ttp1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People hurt in Karachi Attacks by Tehreek-e-Taliban Pakistan(TTP)')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# +
mqm1= mqm.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=mqm1.index,
y=mqm1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=mqm1.index,
y=mqm1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People hurt in Karachi Attacks by MQM')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# +
ukn = khi[khi['Name of Group'] == 'Unknown']
ukn1= ukn.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=ukn1.index,
y=ukn1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=ukn1.index,
y=ukn1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People hurt in Karachi Attacks by Unknown Groups')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# +
yo= (khi.groupby('Name of Group')['Killed'].sum()).sort_values(ascending=False).head(15)
# -
trace = go.Pie(labels=yo.index,values=yo.values)
layout = go.Layout(title='Groups responsible for fatalities above 10')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart2')
# +
## TTP tops the list for recorded fatalities
# -
we=khi['Target'].value_counts()
trace = go.Pie(labels=we.index,values=we.values)
layout = go.Layout(title='Targets in Karachi')
data=[trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic_pie_chart2')
# +
## Private Citizens and Property are the most susceptible,vulnerable and most frequent targets in the city
# +
pvt= khi[khi['Target'] == 'Private Citizens & Property']
pvt1= pvt.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=pvt1.index,
y=pvt1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=pvt1.index,
y=pvt1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People hurt by attacks on Private Citizens and Property')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# +
ass= khi[khi['Type of Attack'] == 'Assassination']
ass1= ass.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=ass1.index,
y=ass1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=ass1.index,
y=ass1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People hurt by Assassinations')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# +
exp= khi[khi['Type of Attack']=='Bombing/Explosion']
exp1= exp.groupby('Year')[['Killed','Wounded']].sum()
trace1 = go.Bar(
x=exp1.index,
y=exp1.Killed,
name='People Killed'
)
trace2 = go.Bar(
x=exp1.index,
y=exp1.Wounded,
name='People Wounded'
)
data = [trace1, trace2]
layout = go.Layout(
barmode='group',
title= 'People hurt by Improvised Explosive Devices')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='grouped-bar')
# -
| Global Terrorism Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Text Classification using TensorFlow/Keras on Cloud ML Engine </h1>
#
# This notebook illustrates:
# <ol>
# <li> Creating datasets for Machine Learning using BigQuery
# <li> Creating a text classification model using the Estimator API with a Keras model
# <li> Training on Cloud ML Engine
# <li> Deploying the model
# <li> Predicting with model
# <li> Rerun with pre-trained embedding
# </ol>
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '1.8'
import tensorflow as tf
print(tf.__version__)
# We will look at the titles of articles and figure out whether the article came from the New York Times, TechCrunch or GitHub.
#
# We will use [hacker news](https://news.ycombinator.com/) as our data source. It is an aggregator that displays tech related headlines from various sources.
# ### Creating Dataset from BigQuery
#
# Hacker news headlines are available as a BigQuery public dataset. The [dataset](https://bigquery.cloud.google.com/table/bigquery-public-data:hacker_news.stories?tab=details) contains all headlines from the sites inception in October 2006 until October 2015.
#
# Here is a sample of the dataset:
import google.datalab.bigquery as bq
query="""
SELECT
url, title, score
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
LENGTH(title) > 10
AND score > 10
LIMIT 10
"""
df = bq.Query(query).execute().result().to_dataframe()
df
# Let's do some regular expression parsing in BigQuery to get the source of the newspaper article from the URL. For example, if the url is http://mobile.nytimes.com/...., I want to be left with <i>nytimes</i>
query="""
SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
COUNT(title) AS num_articles
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
GROUP BY
source
ORDER BY num_articles DESC
LIMIT 10
"""
df = bq.Query(query).execute().result().to_dataframe()
df
# Now that we have good parsing of the URL to get the source, let's put together a dataset of source and titles. This will be our labeled dataset for machine learning.
# +
query="""
SELECT source, LOWER(REGEXP_REPLACE(title, '[^a-zA-Z0-9 $.-]', ' ')) AS title FROM
(SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
title
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
)
WHERE (source = 'github' OR source = 'nytimes' OR source = 'techcrunch')
"""
df = bq.Query(query + " LIMIT 10").execute().result().to_dataframe()
df.head()
# -
# For ML training, we will need to split our dataset into training and evaluation datasets (and perhaps an independent test dataset if we are going to do model or feature selection based on the evaluation dataset).
#
# A simple, repeatable way to do this is to use the hash of a well-distributed column in our data (See https://www.oreilly.com/learning/repeatable-sampling-of-data-sets-in-bigquery-for-machine-learning).
traindf = bq.Query(query + " AND MOD(ABS(FARM_FINGERPRINT(title)),4) > 0").execute().result().to_dataframe()
evaldf = bq.Query(query + " AND MOD(ABS(FARM_FINGERPRINT(title)),4) = 0").execute().result().to_dataframe()
# Below we can see that roughly 75% of the data is used for training, and 25% for evaluation.
#
# We can also see that within each dataset, the classes are roughly balanced.
traindf['source'].value_counts()
evaldf['source'].value_counts()
# Finally we will save our data, which is currently in-memory, to disk.
import os, shutil
DATADIR='data/txtcls'
shutil.rmtree(DATADIR, ignore_errors=True)
os.makedirs(DATADIR)
traindf.to_csv( os.path.join(DATADIR,'train.tsv'), header=False, index=False, encoding='utf-8', sep='\t')
evaldf.to_csv( os.path.join(DATADIR,'eval.tsv'), header=False, index=False, encoding='utf-8', sep='\t')
# !head -3 data/txtcls/train.tsv
# !wc -l data/txtcls/*.tsv
# ### TensorFlow/Keras Code
#
# Please explore the code in this <a href="txtclsmodel/trainer">directory</a>: `model.py` contains the TensorFlow model and `task.py` parses command line arguments and launches off the training job.
#
# There are some TODOs in the `model.py`, **make sure to complete the TODOs before proceeding!**
# ### Run Locally
# Let's make sure the code compiles by running locally for a fraction of an epoch
# + language="bash"
# ## Make sure we have the latest version of Google Cloud Storage package
# pip install --upgrade google-cloud-storage
# rm -rf txtcls_trained
# gcloud ml-engine local train \
# --module-name=trainer.task \
# --package-path=${PWD}/txtclsmodel/trainer \
# -- \
# --output_dir=${PWD}/txtcls_trained \
# --train_data_path=${PWD}/data/txtcls/train.tsv \
# --eval_data_path=${PWD}/data/txtcls/eval.tsv \
# --num_epochs=0.1
# -
# ### Train on the Cloud
#
# Let's first copy our training data to the cloud:
# + language="bash"
# gsutil cp data/txtcls/*.tsv gs://${BUCKET}/txtcls/
# + language="bash"
# OUTDIR=gs://${BUCKET}/txtcls/trained_fromscratch
# JOBNAME=txtcls_$(date -u +%y%m%d_%H%M%S)
# gsutil -m rm -rf $OUTDIR
# gcloud ml-engine jobs submit training $JOBNAME \
# --region=$REGION \
# --module-name=trainer.task \
# --package-path=${PWD}/txtclsmodel/trainer \
# --job-dir=$OUTDIR \
# --scale-tier=BASIC_GPU \
# --runtime-version=$TFVERSION \
# -- \
# --output_dir=$OUTDIR \
# --train_data_path=gs://${BUCKET}/txtcls/train.tsv \
# --eval_data_path=gs://${BUCKET}/txtcls/eval.tsv \
# --num_epochs=5
# -
# ### Monitor training with TensorBoard
# If tensorboard appears blank try refreshing after 10 minutes
from google.datalab.ml import TensorBoard
TensorBoard().start('gs://{}/txtcls/trained_fromscratch'.format(BUCKET))
for pid in TensorBoard.list()['pid']:
TensorBoard().stop(pid)
print 'Stopped TensorBoard with pid {}'.format(pid)
# ### Results
# What accuracy did you get?
# ### Deploy trained model
#
# Once your training completes you will see your exported models in the output directory specified in Google Cloud Storage.
#
# You should see one model for each training checkpoint (default is every 1000 steps).
# + language="bash"
# gsutil ls gs://${BUCKET}/txtcls/trained_fromscratch/export/exporter/
# -
# We will take the last export and deploy it as a REST API using Google Cloud Machine Learning Engine
# + language="bash"
# MODEL_NAME="txtcls"
# MODEL_VERSION="v1_fromscratch"
# MODEL_LOCATION=$(gsutil ls gs://${BUCKET}/txtcls/trained_fromscratch/export/exporter/ | tail -1)
# #gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME} --quiet
# #gcloud ml-engine models delete ${MODEL_NAME}
# gcloud ml-engine models create ${MODEL_NAME} --regions $REGION
# gcloud ml-engine versions create ${MODEL_VERSION} --model ${MODEL_NAME} --origin ${MODEL_LOCATION} --runtime-version=$TFVERSION
# -
# ### Get Predictions
#
# Here are some actual hacker news headlines gathered from July 2018. These titles were not part of the training or evaluation datasets.
techcrunch=[
'Uber shuts down self-driving trucks unit',
'Grover raises €37M Series A to offer latest tech products as a subscription',
'Tech companies can now bid on the Pentagon’s $10B cloud contract'
]
nytimes=[
'‘Lopping,’ ‘Tips’ and the ‘Z-List’: Bias Lawsuit Explores Harvard’s Admissions',
'A $3B Plan to Turn Hoover Dam into a Giant Battery',
'A MeToo Reckoning in China’s Workplace Amid Wave of Accusations'
]
github=[
'Show HN: Moon – 3kb JavaScript UI compiler',
'Show HN: Hello, a CLI tool for managing social media',
'Firefox Nightly added support for time-travel debugging'
]
# Our serving input function expects the already tokenized representations of the headlines, so we do that pre-processing in the code before calling the REST API.
#
# Note: Ideally we would do these transformation in the tensorflow graph directly instead of relying on separate client pre-processing code (see: [training-serving skew](https://developers.google.com/machine-learning/guides/rules-of-ml/#training_serving_skew)), howevever the pre-processing functions we're using are python functions so cannot be embedded in a tensorflow graph.
#
# See the <a href="../text_classification_native.ipynb">text_classification_native</a> notebook for a solution to this.
# +
import pickle
from tensorflow.python.keras.preprocessing import sequence
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
requests = techcrunch+nytimes+github
# Tokenize and pad sentences using same mapping used in the deployed model
tokenizer = pickle.load( open( "txtclsmodel/tokenizer.pickled", "rb" ) )
requests_tokenized = tokenizer.texts_to_sequences(requests)
requests_tokenized = sequence.pad_sequences(requests_tokenized,maxlen=50)
# JSON format the requests
request_data = {'instances':requests_tokenized.tolist()}
# Authenticate and call CMLE prediction API
credentials = GoogleCredentials.get_application_default()
api = discovery.build('ml', 'v1', credentials=credentials,
discoveryServiceUrl='https://storage.googleapis.com/cloud-ml/discovery/ml_v1_discovery.json')
parent = 'projects/%s/models/%s' % (PROJECT, 'txtcls') #version is not specified so uses default
response = api.projects().predict(body=request_data, name=parent).execute()
# Format and print response
for i in range(len(requests)):
print('\n{}'.format(requests[i]))
print(' github : {}'.format(response['predictions'][i]['dense_1'][0]))
print(' nytimes : {}'.format(response['predictions'][i]['dense_1'][1]))
print(' techcrunch: {}'.format(response['predictions'][i]['dense_1'][2]))
# -
# How many of your predictions were correct?
# ### Rerun with Pre-trained Embedding
#
# In the previous model we trained our word embedding from scratch. Often times we get better performance and/or converge faster by leveraging a pre-trained embedding. This is a similar concept to transfer learning during image classification.
#
# We will use the popular GloVe embedding which is trained on Wikipedia as well as various news sources like the New York Times.
#
# You can read more about Glove at the project homepage: https://nlp.stanford.edu/projects/glove/
#
# You can download the embedding files directly from the stanford.edu site, but we've rehosted it in a GCS bucket for faster download speed.
# !gsutil cp gs://cloud-training-demos/courses/machine_learning/deepdive/09_sequence/text_classification/glove.6B.200d.txt gs://$BUCKET/txtcls/
# Once the embedding is downloaded re-run your cloud training job with the added command line argument:
#
# ` --embedding_path=gs://${BUCKET}/txtcls/glove.6B.200d.txt`
#
# Be sure to change your OUTDIR so it doesn't overwrite the previous model.
#
# While the final accuracy may not change significantly, you should notice the model is able to converge to it much more quickly because it no longer has to learn an embedding from scratch.
# #### References
# - This implementation is based on code from: https://github.com/google/eng-edu/tree/master/ml/guides/text_classification.
# - See the full text classification tutorial at: https://developers.google.com/machine-learning/guides/text-classification/
# Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/09_sequence/labs/text_classification.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ # Handling multiple sequences (PyTorch)
/ Install the Transformers and Datasets libraries to run this notebook.
! pip install datasets transformers[sentencepiece]
/ +
import torch
from transformers import AutoTokenizer, AutoModelForSequenceClassification
checkpoint = "distilbert-base-uncased-finetuned-sst-2-english"
tokenizer = AutoTokenizer.from_pretrained(checkpoint)
model = AutoModelForSequenceClassification.from_pretrained(checkpoint)
sequence = "I've been waiting for a HuggingFace course my whole life."
tokens = tokenizer.tokenize(sequence)
ids = tokenizer.convert_tokens_to_ids(tokens)
input_ids = torch.tensor(ids)
model(input_ids)
/ -
tokenized_inputs = tokenizer(sequence, return_tensors="pt")
print(tokenized_inputs["input_ids"])
/ +
import torch
from transformers import AutoTokenizer, AutoModelForSequenceClassification
checkpoint = "distilbert-base-uncased-finetuned-sst-2-english"
tokenizer = AutoTokenizer.from_pretrained(checkpoint)
model = AutoModelForSequenceClassification.from_pretrained(checkpoint)
sequence = "I've been waiting for a HuggingFace course my whole life."
tokens = tokenizer.tokenize(sequence)
ids = tokenizer.convert_tokens_to_ids(tokens)
input_ids = torch.tensor([ids])
print("Input IDs:", input_ids)
output = model(input_ids)
print("Logits:", output.logits)
/ -
batched_ids = [
[200, 200, 200],
[200, 200]
]
/ +
padding_id = 100
batched_ids = [
[200, 200, 200],
[200, 200, padding_id]
]
/ +
model = AutoModelForSequenceClassification.from_pretrained(checkpoint)
sequence1_ids = [[200, 200, 200]]
sequence2_ids = [[200, 200]]
batched_ids = [[200, 200, 200], [200, 200, tokenizer.pad_token_id]]
print(model(torch.tensor(sequence1_ids)).logits)
print(model(torch.tensor(sequence2_ids)).logits)
print(model(torch.tensor(batched_ids)).logits)
/ +
batched_ids = [
[200, 200, 200],
[200, 200, tokenizer.pad_token_id]
]
attention_mask = [
[1, 1, 1],
[1, 1, 0]
]
outputs = model(torch.tensor(batched_ids), attention_mask=torch.tensor(attention_mask))
print(outputs.logits)
/ -
sequence = sequence[:max_sequence_length]
| course/chapter2/section5_pt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vincev16/Numerical-Methods-58011/blob/main/Villasor_Finals_Exam_Numerical_Methods.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LqPeqfDy2SFM"
# ####<NAME>
# 58011
# + [markdown] id="a7Ufu6ALyC9i"
# Problem 3. Create a Python program that integrates the function f(x) = ex from x1=-1 to x2 = 1 as the interval. Save your program to your repository and send your GitHub link here. (30 points)
# + colab={"base_uri": "https://localhost:8080/"} id="K0zC-nRHyKWe" outputId="2429275a-2212-40c4-b3e1-78de33c6b623"
from math import e
def f(x): return e**x #define the function
a = -1
b = 1
n = 10
h = (b-a)/n #width of the trapezoid
S = h* (f(a)+f(b)) #beginning value of summation
for i in range(1,n):
S+=f(a+i*h)
Integral = h*S
print('Integral = %.4f' %Integral)
# + [markdown] id="qekbzMfh3B7I"
# ##another answer with different value of n
# + colab={"base_uri": "https://localhost:8080/"} id="NDfmsexn2-n0" outputId="22513e8c-21fd-46d4-ae42-8c64799093b7"
from math import e
def f(x): return e**x #define the function
a = -1
b = 1
n = 1 #according to the discussion, the value of n should be equal or between the interval
h = (b-a)/n #width of the trapezoid
S = h* (f(a)+f(b)) #beginning value of summation
for i in range(1,n):
S+=f(a+i*h)
Integral = h*S
print('Integral = %.4f' %Integral)
| Villasor_Finals_Exam_Numerical_Methods.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Resources Used
# - wget.download('https://tensorflow-object-detection-api-tutorial.readthedocs.io/en/latest/_downloads/da4babe668a8afb093cc7776d7e630f3/generate_tfrecord.py')
# - Setup https://tensorflow-object-detection-api-tutorial.readthedocs.io/en/latest/install.html
# # 0. Setup Paths
WORKSPACE_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\workspace"
SCRIPTS_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\scripts"
APIMODEL_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\models"
ANNOTATION_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\workspace\annotations"
IMAGE_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\workspace\images"
MODEL_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\workspace\models"
PRETRAINED_MODEL_PATH = r"C:\Users\dell\anaconda3\envs\6th_sense2\RealTimeObjectDetection-main\Tensorflow\workspace\pre-trained-models"
CONFIG_PATH = MODEL_PATH+'\my_ssd_mobnet\pipeline.config'
CHECKPOINT_PATH = MODEL_PATH+'\my_ssd_mobnet'
# # 1. Create Label Map
# +
labels = [
{'name':'hello', 'id':1},
{'name':'thanks', 'id':2},
{'name':'iloveyou', 'id':3},
{'name':'yes', 'id':4},
{'name':'no', 'id':5},
]
with open(ANNOTATION_PATH + '\label_map.pbtxt', 'w') as f:
for label in labels:
f.write('item { \n')
f.write('\tname:\'{}\'\n'.format(label['name']))
f.write('\tid:{}\n'.format(label['id']))
f.write('}\n')
# -
import pandas
# # 2. Create TF records
# !python { r"C:\Users\dell\anaconda3\envs\6th_sense\RealTimeObjectDetection-main\Tensorflow\scripts\generate_tfrecord.py"} -x {r"C:\Users\dell\anaconda3\envs\6th_sense\RealTimeObjectDetection-main\Tensorflow\workspace\images\train"} -l {r"C:\Users\dell\anaconda3\envs\6th_sense\RealTimeObjectDetection-main\Tensorflow\workspace\annotations\label_map.pbtxt"} -o {r"C:\Users\dell\anaconda3\envs\6th_sense\RealTimeObjectDetection-main\Tensorflow\workspace\annotations\train.record"}
# !python {SCRIPTS_PATH + '/generate_tfrecord.py'} -x{IMAGE_PATH + '/test'} -l {ANNOTATION_PATH + '/label_map.pbtxt'} -o {ANNOTATION_PATH + '/test.record'}
# # 3. Download TF Models Pretrained Models from Tensorflow Model Zoo
# !cd Tensorflow && git clone https://github.com/tensorflow/models
# +
#wget.download('http://download.tensorflow.org/models/object_detection/tf2/20200711/ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8.tar.gz')
# #!mv ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8.tar.gz {PRETRAINED_MODEL_PATH}
# #!cd {PRETRAINED_MODEL_PATH} && tar -zxvf ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8.tar.gz
# -
# # 4. Copy Model Config to Training Folder
CUSTOM_MODEL_NAME = 'my_ssd_mobnet'
# !mkdir {'Tensorflow\workspace\models\\'+CUSTOM_MODEL_NAME}
# !copy {PRETRAINED_MODEL_PATH+'/ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8/pipeline.config'} {MODEL_PATH+'/'+CUSTOM_MODEL_NAME}
# # 5. Update Config For Transfer Learning
import tensorflow as tf
from object_detection.utils import config_util
from object_detection.protos import pipeline_pb2
from google.protobuf import text_format
CONFIG_PATH = MODEL_PATH+'/'+CUSTOM_MODEL_NAME+'/pipeline.config'
config = config_util.get_configs_from_pipeline_file(CONFIG_PATH)
config
pipeline_config = pipeline_pb2.TrainEvalPipelineConfig()
with tf.io.gfile.GFile(CONFIG_PATH, "r") as f:
proto_str = f.read()
text_format.Merge(proto_str, pipeline_config)
len(labels)
pipeline_config.model.ssd.num_classes = len(labels)
pipeline_config.train_config.batch_size = 4
pipeline_config.train_config.fine_tune_checkpoint = PRETRAINED_MODEL_PATH+'/ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8/checkpoint/ckpt-0'
pipeline_config.train_config.fine_tune_checkpoint_type = "detection"
pipeline_config.train_input_reader.label_map_path= ANNOTATION_PATH + '/label_map.pbtxt'
pipeline_config.train_input_reader.tf_record_input_reader.input_path[:] = [ANNOTATION_PATH + '/train.record']
pipeline_config.eval_input_reader[0].label_map_path = ANNOTATION_PATH + '/label_map.pbtxt'
pipeline_config.eval_input_reader[0].tf_record_input_reader.input_path[:] = [ANNOTATION_PATH + '/test.record']
config_text = text_format.MessageToString(pipeline_config)
with tf.io.gfile.GFile(CONFIG_PATH, "wb") as f:
f.write(config_text)
# # 6. Train the model
print("""python {}/research/object_detection/model_main_tf2.py --model_dir={}/{} --pipeline_config_path={}/{}/pipeline.config --num_train_steps=5000""".format(APIMODEL_PATH, MODEL_PATH,CUSTOM_MODEL_NAME,MODEL_PATH,CUSTOM_MODEL_NAME))
# # 7. Load Train Model From Checkpoint
import os
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as viz_utils
from object_detection.builders import model_builder
# +
# Load pipeline config and build a detection model
configs = config_util.get_configs_from_pipeline_file(CONFIG_PATH)
detection_model = model_builder.build(model_config=configs['model'], is_training=False)
# Restore checkpoint
ckpt = tf.compat.v2.train.Checkpoint(model=detection_model)
ckpt.restore(os.path.join(CHECKPOINT_PATH, 'ckpt-2')).expect_partial()
@tf.function
def detect_fn(image):
image, shapes = detection_model.preprocess(image)
prediction_dict = detection_model.predict(image, shapes)
detections = detection_model.postprocess(prediction_dict, shapes)
return detections
# -
# # 8. Detect in Real-Time
import cv2
import numpy as np
category_index = label_map_util.create_category_index_from_labelmap(ANNOTATION_PATH+'/label_map.pbtxt')
# Setup capture
cap = cv2.VideoCapture(0)
width = int(cap.get(cv2.CAP_PROP_FRAME_WIDTH))
height = int(cap.get(cv2.CAP_PROP_FRAME_HEIGHT))
while True:
ret, frame = cap.read()
image_np = np.array(frame)
input_tensor = tf.convert_to_tensor(np.expand_dims(image_np, 0), dtype=tf.float32)
detections = detect_fn(input_tensor)
num_detections = int(detections.pop('num_detections'))
detections = {key: value[0, :num_detections].numpy()
for key, value in detections.items()}
detections['num_detections'] = num_detections
# detection_classes should be ints.
detections['detection_classes'] = detections['detection_classes'].astype(np.int64)
label_id_offset = 1
image_np_with_detections = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_detections,
detections['detection_boxes'],
detections['detection_classes']+label_id_offset,
detections['detection_scores'],
category_index,
use_normalized_coordinates=True,
max_boxes_to_draw=5,
min_score_thresh=.5,
agnostic_mode=False)
cv2.imshow('object detection', cv2.resize(image_np_with_detections, (800, 600)))
if cv2.waitKey(1) & 0xFF == ord('q'):
cap.release()
break
detections = detect_fn(input_tensor)
from matplotlib import pyplot as plt
from scipy.optimize import minpack2
import cv2
import numpy as np
# +
import numpy as np
import cv2
cap = cv2.VideoCapture(0)
while True:
#capture frame by frame
ret, img = cap.read()
# gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
cv2.imshow('test', img)
cv2.waitKey(0) & 0xFF
cv2.destroyAllWindows()
cap.release()
# -
| SignLanguage_Live/Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''venv'': venv)'
# language: python
# name: python38364bitvenvvenvf0654de04bec4787b00a87ccaf592e6a
# ---
import pandas as pd
import numpy as np
# +
OScolnames = {'object_number': 'category',
'obs_type': 'category',
'measure_type': 'category',
'?': 'category',
'year': np.uint16,
'month': np.uint8,
'day': np.float32,
'date_accuracy': np.float32,
'RA_HH': np.uint8,
'RA_mm': np.uint8,
'RA_ss': np.float32,
'RA_accuracy': np.float32,
'RA_RMS': np.float32,
#'RA_F': 'category',
'RA_bias': np.float32,
'RA_delta': np.float32,
'DEC_HH': np.uint8,
'DEC_mm': np.uint8,
'DEC_ss': np.float32,
'DEC_accuracy': np.float32,
'DEC_RMS': np.float32,
#'DEC_F': 'category',
'DEC_bias': np.float32,
'DEC_delta': np.float32,
'MAG': np.float32,
'MAG_B': 'category',
'MAG_RMS': np.float32,
'MAG_resid': np.float32,
'catalog': 'category',
'obs_code': 'category',
'xhi': np.float32,
'acceptance': 'bool',
'mag_acceptance': 'bool'}
c=[(0, 11), (11, 13), (13, 15), (15, 17), (17, 21), (22, 25), (25, 40), (40, 50), (50, 53), (53, 56), (56, 64), (64, 77), (77, 83), (87, 94), (94, 103), (103, 107), (107, 110), (110, 117), (117, 130), (128, 136), (141, 148), (149, 156), (156, 161), (161, 164), (164, 170), (170, 178), (178, 180), (180, 186), (189, 194), (194, 196), (196, 197)]
OSdata = pd.read_fwf(r'Data\NEODYS_cleaned.txt', header=None, names=list(OScolnames.keys()), dtype={'obs_code': 'category', 'measure_type': 'category'}, colspecs=c, low_memory=True, index_col=False, usecols=['year', 'obs_code', 'acceptance', 'measure_type', 'RA_delta', 'DEC_delta', 'xhi', 'object_number'], error_bad_lines=False)
OSdata['delta_vect'] = np.sqrt(np.power(OSdata.RA_delta,2) + np.power(OSdata.DEC_delta,2))
# -
# ---
# # Statistics per observation type
types_dict = {'C':'CCD',
'S':'Space observation',
'A':'Observations from B1950.0 converted to J2000.',
'c':'Corrected without republication CCD observation',
'P': 'Photographic',
'T':'Meridian or transit circle ',
'X':'Discovery observation ',
'x':'Discovery observation ',
'M':'Micrometer ',
'H':'Hipparcos geocentric observation ',
'R':'Radar Observation',
'E':'Occultations derived observation ',
'V':'Roving observer observation',
'n':'Video frames',
'e':'Encoder ' }
def get_w_accuracy(typ):
df = OSdata[(OSdata['acceptance']==True) & (OSdata['measure_type'] == typ)]
return np.sqrt(np.power(df.xhi, 2).sum() / len(df))
def get_accuracy(typ):
df = OSdata[(OSdata['acceptance']==True) & (OSdata['measure_type'] == typ)]
return np.sqrt(np.power(df.delta_vect, 2).sum() / len(df))
types = list(OSdata['measure_type'].value_counts().index)
table = pd.DataFrame(index=types, columns=['description', 'count', 'percentage', 'acceptance (%)', 'weighted accuracy (arcsec)', 'accuracy (arcsec)', 'timespan'])
for t in types:
description = types_dict[t]
count = len(OSdata[(OSdata['measure_type'] == t)].index)
percentage = count / len(OSdata.index) * 100
acceptance = (len(OSdata[(OSdata['measure_type'] == t) & (OSdata['acceptance']==True) & (OSdata['xhi'] < 3) & (OSdata['RA_delta'] < 100)]) / count * 100) if count > 0 else np.nan
weighted_accuracy = get_w_accuracy(t)
accuracy = get_accuracy(t)
timespan = str(OSdata[OSdata['measure_type'] == t].year.min()) + '-' + str(OSdata[OSdata['measure_type'] == t].year.max())
table.loc[t] = pd.Series({'description':description, 'count':count, 'percentage':percentage, 'acceptance (%)':acceptance, 'weighted accuracy (arcsec)':weighted_accuracy, 'accuracy (arcsec)':accuracy, 'timespan':timespan})
# + tags=[]
table
# -
# ---
# # Statistics per space telescope
def get_space_waccuracy(code):
df = OSdata[(OSdata['acceptance']==True) & (OSdata['measure_type'] == 'S') & (OSdata['obs_code']==code) & (OSdata['xhi'] < 3) & (OSdata['RA_delta'] < 100) ]
return np.sqrt(np.power(df.xhi, 2).sum() / len(df))
def get_space_accuracy(code):
df = OSdata[(OSdata['acceptance']==True) & (OSdata['measure_type'] == 'S') & (OSdata['obs_code']==code) & (OSdata['xhi'] < 3) & (OSdata['RA_delta'] < 100) ]
return np.sqrt(np.power(df.delta_vect, 2).sum() / len(df))
# + tags=[]
space_telescopes = {'Spitzer Space Telescope':'245',
'Hubble Space Telescope':'250',
'Gaia':'258',
'WISE':'C51',
'NEOSSat':'C53',
'Kepler':'C55',
'TESS':'C57'}
ST_df = pd.DataFrame(index=space_telescopes.keys(), columns=['count', 'acceptance (%)', 'weighted accuracy (arcsec)', 'accuracy (arcsec)'])
for name, obs in space_telescopes.items():
count = len(OSdata[(OSdata['obs_code']==obs) & (OSdata['measure_type']=='S')])
acceptance = (len(OSdata[(OSdata['obs_code']==obs) & (OSdata['acceptance']==True) & (OSdata['measure_type']=='S')]) / count * 100) if count > 0 else np.nan
weighted_accuracy = get_space_waccuracy(obs)
accuracy = get_space_accuracy(obs)
ST_df.loc[name] = pd.Series({'count':count, 'acceptance (%)':acceptance, 'weighted accuracy (arcsec)':weighted_accuracy, 'accuracy (arcsec)':accuracy})
ST_df
# +
#table.to_csv(r'outData\obs_type_stats.csv')
#ST_df.to_csv(r'outData\space_telescope_stats.csv')
# -
table.to_clipboard()
ST_df.to_clipboard()
| observation_types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import qgrid
import seaborn as sns
files = {"all": "../../explanations-for-ner-train-finnish-20190114-total.txt",
"only_target_entities": "../../explanations-for-ner-train-finnish-20190115-total-only_target_entities.txt",
"finnish_model_100_size": "explanations-for-ner-train-finnish_model_100_size.txt",
"turkish_model_100_size": "explanations-for-ner-train-turkish_model_100_size.txt"}
lines = []
records = []
with open(files["finnish_model_100_size"], "r") as f:
lines = f.readlines()
for line in lines:
tokens = line.strip().split("\t")
record = [int(tokens[0]), tokens[1], tuple([int(x) for x in tokens[2].split(" ")])]
record.append({k: float(v) for k, v in [tuple(x.split(" ")) for x in tokens[3:]]})
records.append(record)
records[0]
list(record[3].values())
# +
def log_sum_exp(input_x):
max_value = np.max(input_x)
return np.log(np.sum([np.exp(x-max_value) for x in input_x])) + max_value
log_sum_exp([1, 2])
# -
np.logaddexp(*[1, 2])
group_by_entity_type = {}
for record in records:
entity_type = record[1]
if entity_type not in group_by_entity_type:
group_by_entity_type[entity_type] = {}
if entity_type in group_by_entity_type:
# sum_weights = log_sum_exp(list(record[3].values()))
# min_value = np.min(list(record[3].values()))
# max_value = np.max(list(record[3].values()))
for morpho_tag, weight in record[3].items():
# value = np.exp(weight - sum_weights)
# value = (weight-min_value)/float(max_value-min_value)
value = weight
if morpho_tag in group_by_entity_type[entity_type]:
group_by_entity_type[entity_type][morpho_tag].append(value)
else:
group_by_entity_type[entity_type][morpho_tag] = [value]
group_by_entity_type.keys()
group_by_entity_type['ORG'].keys()
stats_by_entity_type = {key: dict() for key in group_by_entity_type.keys()}
for entity_type in stats_by_entity_type.keys():
for morpho_tag in group_by_entity_type[entity_type]:
l = group_by_entity_type[entity_type][morpho_tag]
stats_by_entity_type[entity_type][morpho_tag] = (np.mean(l), len(l))
for entity_type in stats_by_entity_type.keys():
sorted_l = sorted(stats_by_entity_type[entity_type].items(), key=lambda x: np.abs(x[1][0]), reverse=True)
print(entity_type, sorted_l[:10])
all_morpho_tags = set()
for record in records:
all_morpho_tags.update(set(record[3].keys()))
all_morpho_tags
morpho_tag_to_id = {m: idx for idx, m in enumerate(all_morpho_tags)}
morpho_tag_to_id
record
records_for_panda = []
for record in records:
record_pre_panda = [record[0], record[1], record[2][0], record[2][1]]
morpho_tags = [None] * len(morpho_tag_to_id)
for morpho_tag, idx in morpho_tag_to_id.items():
if morpho_tag in record[3]:
morpho_tags[idx] = record[3][morpho_tag]
record_pre_panda += morpho_tags
records_for_panda.append(record_pre_panda)
# print(record_pre_panda)
id_to_morpho_tag = {idx: morpho_tag for morpho_tag, idx in morpho_tag_to_id.items()}
column_names = ['sentence_idx', 'entity_type', 'entity_start', 'entity_end']
column_names += [id_to_morpho_tag[x] for x in range(len(morpho_tag_to_id))]
explanations = pd.DataFrame(records_for_panda, columns=column_names)
explanations
df_by_entity_type = explanations.groupby('entity_type')
explanations.drop(['sentence_idx', 'entity_start', 'entity_end'], axis=1).groupby('entity_type').mean()
means_over_entity_type = explanations.drop(['sentence_idx', 'entity_start', 'entity_end'], axis=1).groupby('entity_type').mean()
# %matplotlib inline
means_over_entity_type.index
means_over_entity_type.corr()
means_over_entity_type['Ins^DB'].mean()
means_over_entity_type[means_over_entity_type.columns[0]].mean()
explanations_grid = qgrid.show_grid(means_over_entity_type.corr().iloc[:, 0:2], show_toolbar=True)
explanations_grid
df_by_entity_type = explanations.drop(['sentence_idx', 'entity_start', 'entity_end'], axis=1).groupby('entity_type')
explanations[explanations['entity_type'] == "LOC"]
# # LOC type entities - analysis
loc_group_explanations = explanations[explanations['entity_type'] == "LOC"].drop(["sentence_idx", "entity_type", "entity_start", "entity_end"], axis=1)
loc_group_explanations.columns
loc_group_explanations['Loc'].clip(lower=-1.0, upper=1, inplace=False)
len(morpho_tag_to_id)
loc_group_explanations.size
for idx, morpho_tag in enumerate(list(morpho_tag_to_id.keys())):
if idx % 9 == 0:
fig = plt.figure(int(idx/9))
rem = idx % 9
plt.subplot(3, 3, rem+1)
print(morpho_tag)
# sns.violinplot(data=list(loc_group_explanations[morpho_tag].clip(lower=-0.5, upper=0.5)))
data = loc_group_explanations[morpho_tag].dropna().clip(lower=-0.5, upper=0.5)
print(data)
if data.size > 0:
sns.distplot(data)
plt.show()
loc_group_explanations
mean_loc_group_explanations = loc_group_explanations.mean()
mean_loc_group_explanations.sort_values(ascending=False)
loc_group_explanations['Loc'].sort_values()[:10]
loc_group_explanations['Loc'].sort_values(ascending=False)[:10]
loc_group_explanations.hist(['Loc'], range=[-1, 1], bins=100)
loc_group_explanations.hist(['Loc'], range=[-0.015, 0.015], bins=100)
loc_group_explanations['Loc'].value_counts().sort_values(ascending=False)
[(loc_group_explanations['Loc'][loc_group_explanations['Loc'] < 0]).mean(),
(loc_group_explanations['Loc'][loc_group_explanations['Loc'] >= 0]).mean()]
loc_group_explanations.hist(['Loc^DB'], range=[-1, 1])
loc_group_explanations.hist(['Loc'])
loc_group_explanations.hist(['Loc^DB'])
loc_group_explanations.hist(['Loc'], range=[-5000, -10], bins=100)
loc_group_explanations.hist(['Loc'], range=[1, 1000], bins=100)
loc_group_explanations['Loc'][loc_group_explanations['Loc'] < 0].count()
loc_group_explanations['Loc'][loc_group_explanations['Loc'] >= 0].count()
for morpho_tag in ['Loc', 'Loc^DB']:
below_zero = loc_group_explanations[morpho_tag][loc_group_explanations[morpho_tag] < 0].count()
above_zero = loc_group_explanations[morpho_tag][loc_group_explanations[morpho_tag] >= 0].count()
print(morpho_tag, below_zero, above_zero)
# # ORG type entities - analysis
org_group_explanations = explanations[explanations['entity_type'] == "ORG"].drop(["sentence_idx", "entity_type", "entity_start", "entity_end"], axis=1)
org_group_explanations.mean().sort_values(ascending=False)
# # PER type entities - analysis
per_group_explanations = explanations[explanations['entity_type'] == "PER"].drop(["sentence_idx", "entity_type", "entity_start", "entity_end"], axis=1)
per_group_explanations.mean().sort_values(ascending=False)
# !pwd
# !ls ../../explanations-for-ner-train-finnish-201901*
| toolkit/xnlp/Explanations Analysis Run 02 - Finnish.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="center" style="max-width: 1000px" src="banner.png">
# + [markdown] id="dojtwAh1Ww1B"
# <img align="right" style="max-width: 200px; height: auto" src="hsg_logo.png">
#
# ## Lab 04 - "Artificial Neural Networks (ANNs)"
#
# EMBA 58/59 - W8/3 - "AI Coding for Executives", University of St. Gallen
# -
# The lab environment of the "AI Coding for Executives" EMBA course at the University of St. Gallen (HSG) is based on Jupyter Notebooks (https://jupyter.org), which allow to perform a variety of statistical evaluations and data analyses.
# + [markdown] id="aR4Ywe2HWw1M"
# In this lab, we will learn how to implement, train, and apply our first **Artificial Neural Network (ANN)** using a Python library named `PyTorch`. The `PyTorch` library is an open-source machine learning library for Python, used for a variety of applications such as image classification and natural language processing. We will use the implemented neural network to learn to again classify images of fashion articles from the **Fashion-MNIST** dataset.
#
# The figure below illustrates a high-level view of the machine learning process we aim to establish in this lab:
# + [markdown] id="wgQ_ksmaWw1N"
# <img align="center" style="max-width: 700px" src="classification.png">
# + [markdown] id="Z-9LY1AjWw1O"
# As always, pls. don't hesitate to ask all your questions either during the lab, post them in our CANVAS (StudyNet) forum (https://learning.unisg.ch), or send us an email (using the course email).
# + [markdown] id="aut1dJXmWw1O"
# ## 1. Lab Objectives:
# + [markdown] id="7tb0svb4Ww1O"
# After today's lab, you should be able to:
#
# > 1. Understand the basic concepts, intuitions and major building blocks of **Artificial Neural Networks (ANNs)**.
# > 2. Know how to use Python's **PyTorch library** to train and evaluate neural network based models.
# > 3. Understand how to apply neural networks to **classify images** of handwritten digits.
# > 4. Know how to **interpret the detection results** of the network as well as its **reconstruction loss**.
# + [markdown] id="Ks081EJEWw1P"
# ## 2. Setup of the Jupyter Notebook Environment
# + [markdown] id="mdmhjYHFWw1P"
# Similar to the previous labs, we need to import a couple of Python libraries that allow for data analysis and data visualization. We will mostly use the PyTorch, Numpy, Sklearn, Matplotlib, Seaborn and a few utility libraries throughout this lab:
# + id="5rDvIKj-Ww1P"
# import standard python libraries
import os, urllib, io
from datetime import datetime
import numpy as np
# + [markdown] id="heUZY7dgWw1Q"
# Import the Python machine / deep learning libraries:
# + id="-RTb4Mc1Ww1Q"
# import the PyTorch deep learning libary
import torch, torchvision
import torch.nn.functional as F
from torch import nn, optim
# + [markdown] id="FRpZfSImWw1R"
# Import the sklearn classification metrics:
# + id="VJ180J4sWw1R"
# import sklearn classification evaluation library
from sklearn import metrics
from sklearn.metrics import classification_report, confusion_matrix
# + [markdown] id="5tatoV81Ww1R"
# Import Python plotting libraries:
# + id="fTSNWwejWw1R"
# import matplotlib, seaborn, and PIL data visualization libary
import matplotlib.pyplot as plt
import seaborn as sns
from PIL import Image
# + [markdown] id="g83bVkFrWw1S"
# Enable notebook matplotlib inline plotting:
# + id="i0MKI98CWw1S"
# %matplotlib inline
# + [markdown] id="DM3LiXwWWw1S"
# Import Google's GDrive connector and mount your GDrive directories:
# + id="ptQU8CeEWw1S"
# import the Google Colab GDrive connector
from google.colab import drive
# mount GDrive inside the Colab notebook
drive.mount('/content/drive')
# + [markdown] id="I81E5iJOwR6x"
# Create a structure of Colab Notebook sub-directories inside of GDrive to store (1) the data as well as (2) the trained neural network models:
# + id="Oq5F-BV1wSXL"
# create Colab Notebooks directory
notebook_directory = '/content/drive/MyDrive/Colab Notebooks'
if not os.path.exists(notebook_directory): os.makedirs(notebook_directory)
# create data sub-directory inside the Colab Notebooks directory
data_directory = '/content/drive/MyDrive/Colab Notebooks/data'
if not os.path.exists(data_directory): os.makedirs(data_directory)
# create models sub-directory inside the Colab Notebooks directory
models_directory = '/content/drive/MyDrive/Colab Notebooks/models'
if not os.path.exists(models_directory): os.makedirs(models_directory)
# + [markdown] id="32BCRqyA5cD5"
# Set a random `seed` value to obtain reproducable results:
# + id="D3cPTu1R5cmj"
# init deterministic seed
seed_value = 1234
np.random.seed(seed_value) # set numpy seed
torch.manual_seed(seed_value) # set pytorch seed CPU
# + [markdown] id="BHMs9Gf0wakv"
# Google Colab provides the use of free GPUs for running notebooks. However, if you just execute this notebook as is, it will use your device's CPU. To run the lab on a GPU, got to `Runtime` > `Change runtime type` and set the Runtime type to `GPU` in the drop-down. Running this lab on a CPU is fine, but you will find that GPU computing is faster. *CUDA* indicates that the lab is being run on GPU.
#
# Enable GPU computing by setting the device flag and init a CUDA seed:
# + id="Fl2UHzshwdyk"
# set cpu or gpu enabled device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu').type
# init deterministic GPU seed
torch.cuda.manual_seed(seed_value)
# log type of device enabled
print('[LOG] notebook with {} computation enabled'.format(str(device)))
# + [markdown] id="47HnxJHswf05"
# Let's determine if we have access to a GPU provided by e.g. Google's COLab environment:
# + id="907R1nhVwhXb"
# !nvidia-smi
# + [markdown] id="vyqnqndjWw1S"
# ## 3. Dataset Download and Data Assessment
# + [markdown] id="wgyKo34eWw1T"
# The **Fashion-MNIST database** is a large database of Zalando articles that is commonly used for training various image processing systems. The database is widely used for training and testing in the field of machine learning. Let's have a brief look into a couple of sample images contained in the dataset:
# + [markdown] id="-q9TexBXWw1T"
# <img align="center" style="max-width: 700px; height: 300px" src="FashionMNIST.png">
#
# Source: https://www.kaggle.com/c/insar-fashion-mnist-challenge
# + [markdown] id="4YEOHPO5Ww1T"
# Further details on the dataset can be obtained via Zalando research's [github page](https://github.com/zalandoresearch/fashion-mnist).
# + [markdown] id="_B6cw9iEWw1T"
# The **Fashion-MNIST database** is a dataset of Zalando's article images, consisting of a training set of 60,000 examples and a test set of 10,000 examples. Each example is a 28x28 grayscale image, associated with a label from 10 classes. Zalando created this dataset with the intention of providing a replacement for the popular **MNIST** handwritten digits dataset. It is a useful addition as it is a bit more complex, but still very easy to use. It shares the same image size and train/test split structure as MNIST, and can therefore be used as a drop-in replacement. It requires minimal efforts on preprocessing and formatting the distinct images.
# + [markdown] id="igSTsQMKWw1U"
# Let's download, transform and inspect the training images of the dataset. Therefore, let's first define the directory in which we aim to store the training data:
# + id="wfCn1e8MWw1V"
train_path = data_directory + '/train_fashion_mnist'
# + [markdown] id="k4fUsNeLWw1V"
# Now, let's download the training data accordingly:
# + id="X-GZL31YWw1W"
# define pytorch transformation into tensor format
transf = torchvision.transforms.Compose([torchvision.transforms.ToTensor()])
# download and transform training images
fashion_mnist_train_data = torchvision.datasets.FashionMNIST(root=train_path, train=True, transform=transf, download=True)
# + [markdown] id="HaLuUXc4Ww1W"
# Verify the number of training images downloaded:
# + id="OmRdyfxFWw1W"
# determine the number of training data images
len(fashion_mnist_train_data)
# + [markdown] id="wtbIaCpLWw1W"
# Furthermore, let's inspect a couple of the downloaded training images:
# + id="gAtYOeUPWw1X"
# select and set a (random) image id
image_id = 3000
# retrieve image exhibiting the image id
fashion_mnist_train_data[image_id]
# + [markdown] id="nfUMS402Ww1X"
# Ok, that doesn't seem right :). Let's now seperate the image from its label information:
# + id="HDzFFyZfWw1X"
fashion_mnist_train_image, fashion_mnist_train_label = fashion_mnist_train_data[image_id]
# + [markdown] id="3SSorDiP_uFc"
# We can verify the label that our selected image has:
# + id="weOc_Ceb_5dU"
fashion_mnist_train_label
# + [markdown] id="ctD4Dl0C_7RE"
# Ok, we know that the numerical label is 6. Each image is associated with a label from 0 to 9, and this number represents one of the fashion items. So what does 6 mean? Is 6 a bag? A pullover? The order of the classes can be found on Zalando research's [github page](https://github.com/zalandoresearch/fashion-mnist). We need to map each numerical label to its fashion item, which will be useful throughout the lab:
# + id="FmzSMz1FASrm"
fashion_classes = {0: 'T-shirt/top',
1: 'Trouser',
2: 'Pullover',
3: 'Dress',
4: 'Coat',
5: 'Sandal',
6: 'Shirt',
7: 'Sneaker',
8: 'Bag',
9: 'Ankle boot'}
# + [markdown] id="sDvW1PUdKn3a"
# So, we can determine the fashion item that the label represents:
# + id="N8KrM8rfKnHh"
fashion_classes[fashion_mnist_train_label]
# + [markdown] id="JozKTGsCWw1X"
# Great, let's now visually inspect our sample image:
# + id="D80b3WkWWw1Y"
# define tensor to image transformation
trans = torchvision.transforms.ToPILImage()
# set image plot title
plt.title('Example: {}, Label: {}'.format(str(image_id), fashion_classes[fashion_mnist_train_label]))
# plot mnist handwritten digit sample
plt.imshow(trans(fashion_mnist_train_image), cmap='gray')
# + [markdown] id="w5fmhiuzWw1Y"
# Fantastic, right? Let's now define the directory in which we aim to store the evaluation data:
# + id="8G3eQFy3Ww1b"
eval_path = data_directory + '/eval_fashion_mnist'
# + [markdown] id="dNNTmyI7Ww1b"
# And download the evaluation data accordingly:
# + id="PvIBdhlPWw1b"
# define pytorch transformation into tensor format
transf = torchvision.transforms.Compose([torchvision.transforms.ToTensor()])
# download and transform training images
fashion_mnist_eval_data = torchvision.datasets.FashionMNIST(root=eval_path, train=False, transform=transf, download=True)
# + [markdown] id="4skfCFEfWw1c"
# Let's also verify the number of evaluation images downloaded:
# + id="Oq3rW2wKWw1c"
# determine the number of evaluation data images
len(fashion_mnist_eval_data)
# + [markdown] id="ucTxc7GGWw1c"
# ## 4. Neural Network Implementation
# + [markdown] id="xTQ_VZWaWw1d"
# In this section we, will implement the architecture of the **neural network** we aim to utilize to learn a model that is capable to classify the 28x28 pixel FashionMNIST images of fashion items. However, before we start the implementation let's briefly revisit the process to be established. The following cartoon provides a birds-eye view:
# + [markdown] id="9i5LlBmiWw1d"
# <img align="center" style="max-width: 1000px" src="process.png">
# + [markdown] id="OyofP39KWw1d"
# ### 4.1 Implementation of the Neural Network Architecture
# + [markdown] id="loUEinm1Ww1e"
# The neural network, which we name **'FashionMNISTNet'** consists of three **fully-connected layers** (including an “input layer” and two hidden layers). Furthermore, the **FashionMNISTNet** should encompass the following number of neurons per layer: 100 (layer 1), 50 (layer 2) and 10 (layer 3). Meaning the first layer consists of 100 neurons, the second layer of 50 neurons and third layer of 10 neurons (the number of digit classes we aim to classify.
# + [markdown] id="FGxSr-77Ww1e"
# We will now start implementing the network architecture as a separate Python class. Implementing the network architectures as a **separate class** in Python is good practice in deep learning projects. It will allow us to create and train several instances of the same neural network architecture. This provides us, for example, the opportunity to evaluate different initializations of the network parameters or train models using distinct datasets.
# + id="VLrELu2EWw1f"
# implement the MNISTNet network architecture
class FashionMNISTNet(nn.Module):
# define the class constructor
def __init__(self):
# call super class constructor
super(FashionMNISTNet, self).__init__()
# specify fully-connected (fc) layer 1 - in 28*28, out 100
self.linear1 = nn.Linear(28*28, 100, bias=True) # the linearity W*x+b
self.relu1 = nn.ReLU(inplace=True) # the non-linearity
# specify fc layer 2 - in 100, out 50
self.linear2 = nn.Linear(100, 50, bias=True) # the linearity W*x+b
self.relu2 = nn.ReLU(inplace=True) # the non-linarity
# specify fc layer 3 - in 50, out 10
self.linear3 = nn.Linear(50, 10) # the linearity W*x+b
# add a softmax to the last layer
self.logsoftmax = nn.LogSoftmax(dim=1) # the softmax
# define network forward pass
def forward(self, images):
# reshape image pixels
x = images.view(-1, 28*28)
# define fc layer 1 forward pass
x = self.relu1(self.linear1(x))
# define fc layer 2 forward pass
x = self.relu2(self.linear2(x))
# define layer 3 forward pass
x = self.logsoftmax(self.linear3(x))
# return forward pass result
return x
# + [markdown] id="Xy63DaNzWw1f"
# You may have noticed, when reviewing the implementation above, that we applied an additional operator, referred to as **'Softmax'** to the third layer of our neural network.
#
# The **softmax function**, also known as the normalized exponential function, is a function that takes as input a vector of K real numbers, and normalizes it into a probability distribution consisting of K probabilities.
#
# That is, prior to applying softmax, some vector components could be negative, or greater than one; and might not sum to 1; but after application of the softmax, each component will be in the interval $(0,1)$, and the components will add up to 1, so that they can be interpreted as probabilities. In general, the softmax function $\sigma :\mathbb {R} ^{K}\to \mathbb {R} ^{K}$ is defined by the formula:
# + [markdown] id="s5s_1Q7NWw1f"
# <center> $\sigma (\mathbf {z} )_{i}=\ln ({e^{z_{i}} / \sum _{j=1}^{K}e^{z_{j}}})$ </center>
# + [markdown] id="E9JtxNu4Ww1f"
# for $i = 1, …, K$ and ${\mathbf {z}}=(z_{1},\ldots ,z_{K})\in \mathbb {R} ^{K}$ (Source: https://en.wikipedia.org/wiki/Softmax_function ).
#
# Let's have a look at the simplified three-class example below. The scores of the distinct predicted classes $c_i$ are computed from the forward propagation of the network. We then take the softmax and obtain the probabilities as shown:
# + [markdown] id="1ULWQ3RmWw1f"
# <img align="center" style="max-width: 800px" src="softmax.png">
# + [markdown] id="zcrCZgZqWw1g"
# The output of the softmax describes the probability (or if you may, the confidence) of the neural network that a particular sample belongs to a certain class. Thus, for the first example above, the neural network assigns a confidence of 0.49 that it is a 'three', 0.49 that it is a 'four', and 0.03 that it is an 'eight'. The same goes for each of the samples above.
#
# Now, that we have implemented our first neural network we are ready to instantiate a network model to be trained:
# + id="zfvFFCCHWw1g"
model = FashionMNISTNet()
# + [markdown] id="efX9IOPSw8DZ"
# Let's push the initialized `FashionMNISTNet` model to the computing `device` that is enabled:
# + id="W93HbADVw8qe"
model = model.to(device)
# + [markdown] id="9j5xBQ2yxAMw"
# Let's double check if our model was deployed to the GPU if available:
# + id="ZFGVBXGgxAo5"
# !nvidia-smi
# + [markdown] id="2yt4PtPLWw1g"
# Once the model is initialized, we can visualize the model structure and review the implemented network architecture by execution of the following cell:
# + id="SF90-Nk1Ww1g"
# print the initialized architectures
print('[LOG] FashionMNISTNet architecture:\n\n{}\n'.format(model))
# + [markdown] id="PFKJMIjKWw1g"
# Looks like intended? Brilliant! Finally, let's have a look into the number of model parameters that we aim to train in the next steps of the notebook:
# + id="wFOWHzJ3Ww1h"
# init the number of model parameters
num_params = 0
# iterate over the distinct parameters
for param in model.parameters():
# collect number of parameters
num_params += param.numel()
# print the number of model paramters
print('[LOG] Number of to be trained FashionMNISTNet model parameters: {}.'.format(num_params))
# + [markdown] id="vY9rZQ7JWw1h"
# Ok, our "simple" FashionMNISTNet model already encompasses an impressive number 84'060 model parameters to be trained.
# + [markdown] id="pytnqULPWw1h"
# ### 4.2 Specification of the Neural Network Loss Function
# + [markdown] id="bHpHWQ7JWw1h"
# Now that we have implemented the **FashionMNISTNet** we are ready to train the network. However, prior to starting the training, we need to define an appropriate loss function. Remember, we aim to train our model to learn a set of model parameters $\theta$ that minimize the classification error of the true class $c^{i}$ of a given handwritten digit image $x^{i}$ and its predicted class $\hat{c}^{i} = f_\theta(x^{i})$ as faithfully as possible.
#
# Thereby, the training objective is to learn a set of optimal model parameters $\theta^*$ that optimize $\arg\min_{\theta} \|C - f_\theta(X)\|$ over all training images in the FashionMNIST dataset. To achieve this optimization objective, one typically minimizes a loss function $\mathcal{L_{\theta}}$ as part of the network training. In this lab we use the **'Negative Log Likelihood (NLL)'** loss, defined by:
#
# <center> $\mathcal{L}^{NLL}_{\theta} (c_i, \hat c_i) = - \frac{1}{N} \sum_{i=1}^N \log (\hat{c}_i) $, </center>
# + [markdown] id="ID9HqJJqWw1h"
# for a set of $n$-FashionMNIST images $x^{i}$, $i=1,...,n$ and their respective predicted class labels $\hat{c}^{i}$. This is summed for all the correct classes.
#
# Let's have a look at a brief example:
# + [markdown] id="8xRWCsbrWw1i"
# <img align="center" style="max-width: 800px" src="loss.png">
# + [markdown] id="HcObuMZfWw1i"
# During training the **NLL** loss will penalize models that result in a high classification error between the predicted class labels $\hat{c}^{i}$ and their respective true class label $c^{i}$. Luckily, an implementation of the NLL loss is already available in PyTorch! It can be instantiated "off-the-shelf" via the execution of the following PyTorch command:
# + id="qbgFFIDjWw1i"
# define the optimization criterion / loss function
nll_loss = nn.NLLLoss()
# + [markdown] id="YbspDM2dxGrO"
# Let's also push the initialized `nll_loss` computation to the computing `device` that is enabled:
# + id="RgKeoEcaxIdB"
nll_loss = nll_loss.to(device)
# + [markdown] id="hJhKTaHnWw1i"
# ## 5. Neural Network Model Training
# + [markdown] id="uRwd3R8XWw1i"
# In this section, we will train our neural network model (as implemented in the section above) using the transformed images of fashion items. More specifically, we will have a detailed look into the distinct training steps as well as how to monitor the training progress.
# + [markdown] id="OKbH8GgrWw1i"
# ### 5.1. Preparing the Network Training
# + [markdown] id="1K23pmmEWw1j"
# So far, we have pre-processed the dataset, implemented the ANN and defined the classification error. Let's now start to train a corresponding model for **20 epochs** and a **mini-batch size of 128** FashionMNIST images per batch. This implies that the whole dataset will be fed to the ANN 20 times in chunks of 128 images yielding to **469 mini-batches** (60.000 images / 128 images per mini-batch) per epoch.
# + id="4bMEnxc1Ww1j"
# specify the training parameters
num_epochs = 20 # number of training epochs
mini_batch_size = 128 # size of the mini-batches
# + [markdown] id="jX99_DY7Ww1j"
# Based on the loss magnitude of a certain mini-batch PyTorch automatically computes the gradients. But even better, based on the gradient, the library also helps us in the optimization and update of the network parameters $\theta$.
#
# We will use the **Stochastic Gradient Descent (SGD) optimization** and set the learning-rate $l = 0.001$. Each mini-batch step the optimizer will update the model parameters $\theta$ values according to the degree of classification error (the MSE loss).
# + id="84Oq2woEWw1j"
# define learning rate and optimization strategy
learning_rate = 0.001
optimizer = optim.SGD(params=model.parameters(), lr=learning_rate)
# + [markdown] id="HX8MBSDxWw1k"
# Now that we have successfully implemented and defined the three ANN building blocks let's take some time to review the `FashionMNISTNet` model definition as well as the `loss`. Please, read the above code and comments carefully and don't hesitate to let us know any questions you might have.
# + [markdown] id="XO1P2wb3Ww1k"
# Furthermore, lets specify and instantiate a corresponding PyTorch data loader that feeds the image tensors to our neural network:
# + id="vyLwFEMXWw1l"
fashion_mnist_train_dataloader = torch.utils.data.DataLoader(fashion_mnist_train_data, batch_size=mini_batch_size, shuffle=True)
# + [markdown] id="Tp6hmrg7Ww1l"
# ### 5.2. Running the Network Training
# + [markdown] id="Ov5Z6NLvWw1m"
# Finally, we start training the model. The detailed training procedure for each mini-batch is performed as follows:
#
# >1. do a forward pass through the FashionMNISTNet network,
# >2. compute the negative log likelihood classification error $\mathcal{L}^{NLL}_{\theta}(c^{i};\hat{c}^{i})$,
# >3. do a backward pass through the FashionMNISTNet network, and
# >4. update the parameters of the network $f_\theta(\cdot)$.
#
# To ensure learning while training our ANN model, we will monitor whether the loss decreases with progressing training. Therefore, we obtain and evaluate the classification performance of the entire training dataset after each training epoch. Based on this evaluation, we can conclude on the training progress and whether the loss is converging (indicating that the model might not improve any further).
#
# The following elements of the network training code below should be given particular attention:
#
# >- `loss.backward()` computes the gradients based on the magnitude of the reconstruction loss,
# >- `optimizer.step()` updates the network parameters based on the gradient.
# + id="70W8AZWlWw1m"
# init collection of training epoch losses
train_epoch_losses = []
# set the model in training mode
model.train()
# train the MNISTNet model
for epoch in range(num_epochs):
# init collection of mini-batch losses
train_mini_batch_losses = []
# iterate over all-mini batches
for i, (images, labels) in enumerate(fashion_mnist_train_dataloader):
# push mini-batch data to computation device
images = images.to(device)
labels = labels.to(device)
# run forward pass through the network
output = model(images)
# reset graph gradients
model.zero_grad()
# determine classification loss
loss = nll_loss(output, labels)
# run backward pass
loss.backward()
# update network paramaters
optimizer.step()
# collect mini-batch reconstruction loss
train_mini_batch_losses.append(loss.data.item())
# determine mean min-batch loss of epoch
train_epoch_loss = np.mean(train_mini_batch_losses)
# print epoch loss
now = datetime.utcnow().strftime("%Y%m%d-%H:%M:%S")
print('[LOG {}] epoch: {} train-loss: {}'.format(str(now), str(epoch), str(train_epoch_loss)))
# set filename of actual model
model_name = 'fashion_mnist_model_epoch_{}.pth'.format(str(epoch))
# save current model to GDrive models directory
torch.save(model.state_dict(), os.path.join(models_directory, model_name))
# determine mean min-batch loss of epoch
train_epoch_losses.append(train_epoch_loss)
# + [markdown] id="8dqoGo_7Ww1m"
# Upon successfull training let's visualize and inspect the training loss per epoch:
# + id="lLI0Y53VWw1m"
# prepare plot
fig = plt.figure()
ax = fig.add_subplot(111)
# add grid
ax.grid(linestyle='dotted')
# plot the training epochs vs. the epochs' classification error
ax.plot(np.array(range(1, len(train_epoch_losses)+1)), train_epoch_losses, label='epoch loss (blue)')
# add axis legends
ax.set_xlabel("[training epoch $e_i$]", fontsize=10)
ax.set_ylabel("[Classification Error $\mathcal{L}^{NLL}$]", fontsize=10)
# set plot legend
plt.legend(loc="upper right", numpoints=1, fancybox=True)
# add plot title
plt.title('Training Epochs $e_i$ vs. Classification Error $L^{NLL}$', fontsize=10);
# + [markdown] id="QmkQMB6YWw1n"
# Ok, fantastic. The training error is nicely going down. We could train the network a couple more epochs until the error converges. But let's stay with the 20 training epochs for now and continue with evaluating our trained model.
# + [markdown] id="8nyWq1X-Ww1n"
# ## 6. Neural Network Model Evaluation
# + [markdown] id="S7-GBxuwWw1n"
# Before evaluating our model let's load the best performing model. Remember, that we stored a snapshot of the model after each training epoch to our local model directory. We will now load the last snapshot saved.
# + id="-w1B1k_NWw1n"
# restore pre-trained model snapshot
best_model_name = 'https://raw.githubusercontent.com/HSG-AIML/LabAI-Coding/master/resources/lab_04/models/fashion_mnist_model_epoch_19.pth'
# read stored model from the remote location
model_bytes = urllib.request.urlopen(best_model_name)
# load model tensor from io.BytesIO object
model_buffer = io.BytesIO(model_bytes.read())
# init pre-trained model class
best_model = FashionMNISTNet()
# load pre-trained models
best_model.load_state_dict(torch.load(model_buffer, map_location=torch.device('cpu')))
# + [markdown] id="7oqOeiajWw1n"
# Let's inspect if the model was loaded successfully:
# + id="KarjZ3ldWw1n"
# set model in evaluation mode
best_model.eval()
# + [markdown] id="GaFA9LyRWw1o"
# To evaluate our trained model, we need to feed the FashionMNIST images reserved for evaluation (the images that we didn't use as part of the training process) through the model. Therefore, let's again define a corresponding PyTorch data loader that feeds the image tensors to our neural network:
# + id="pQB-RFR8Ww1o"
fashion_mnist_eval_dataloader = torch.utils.data.DataLoader(fashion_mnist_eval_data, batch_size=10000, shuffle=True)
# + [markdown] id="OPP7OowBWw1o"
# We will now evaluate the trained model using the same mini-batch approach as we did throughout the network training and derive the mean negative log-likelihood loss of the mini-batches:
# + id="xpttWy_AWw1o"
# init collection of mini-batch losses
eval_mini_batch_losses = []
# iterate over all-mini batches
for i, (images, labels) in enumerate(fashion_mnist_eval_dataloader):
# run forward pass through the network
output = best_model(images)
# determine classification loss
loss = nll_loss(output, labels)
# collect mini-batch reconstruction loss
eval_mini_batch_losses.append(loss.data.item())
# determine mean min-batch loss of epoch
eval_loss = np.mean(eval_mini_batch_losses)
# print epoch loss
now = datetime.utcnow().strftime("%Y%m%d-%H:%M:%S")
print('[LOG {}] eval-loss: {}'.format(str(now), str(eval_loss)))
# + [markdown] id="oNwLBDGPWw1p"
# Ok, great. The evaluation loss looks in-line with our training loss. Let's now inspect a few sample predictions to get an impression of the model quality. Therefore, we will again pick a random image of our evaluation dataset and retrieve its PyTorch tensor as well as the corresponding label:
# + id="JTM5mTHaWw1p"
# set (random) image id
image_id = 2000
# retrieve image exhibiting the image id
fashion_mnist_eval_image, fashion_mnist_eval_label = fashion_mnist_eval_data[image_id]
# + [markdown] id="wkZowxdUWw1p"
# Let's now inspect the true class of the image we selected:
# + id="IIq9uSnfWw1q"
fashion_classes[fashion_mnist_eval_label]
# + [markdown] id="Hjz4GOpyWw1q"
# Ok, the randomly selected image should contain a one (1). Let's inspect the image accordingly:
# + id="m1hZMQ6oWw1r"
# define tensor to image transformation
trans = torchvision.transforms.ToPILImage()
# set image plot title
plt.title('Example: {}, Label: {}'.format(str(image_id), fashion_classes[fashion_mnist_eval_label]))
# plot mnist handwritten digit sample
plt.imshow(trans(fashion_mnist_eval_image), cmap='gray')
# + [markdown] id="yFW-PYEnWw1r"
# Ok, let's compare the true label with the prediction of our model:
# + id="hfaV80IpWw1s"
best_model(fashion_mnist_eval_image)
# + [markdown] id="mNFAM_deWw1s"
# We can even determine the likelihood of the most probable class:
# + id="A2knLiUqWw1t"
most_probable = torch.argmax(best_model(fashion_mnist_eval_image), dim=1).item()
print('Most probable class: {}'.format(most_probable))
print('This class represents the following fashion article: {}'.format(fashion_classes[most_probable]))
# + [markdown] id="jwOIf2adWw1t"
# Let's now obtain the predictions for all the fashion item images of the evaluation data:
# + id="YRmjfYDZWw1t"
predictions = torch.argmax(best_model(fashion_mnist_eval_data.data.float()), dim=1)
# + [markdown] id="eFKiMKw-Ww1t"
# Furthermore, let's obtain the overall classifcation accuracy:
# + id="BvV-HLcsWw1t"
metrics.accuracy_score(fashion_mnist_eval_data.targets, predictions.detach())
# + [markdown] id="6B-KdgKNWw1u"
# Let's also inspect the confusion matrix to determine major sources of misclassification:
# + id="w5LkSBwpWw1u"
# determine classification matrix of the predicted and target classes
mat = confusion_matrix(fashion_mnist_eval_data.targets, predictions.detach())
# initialize the plot and define size
plt.figure(figsize=(8, 8))
# plot corresponding confusion matrix
sns.heatmap(mat.T, square=True, annot=True, fmt='d', cbar=False, cmap='YlOrRd_r', xticklabels=fashion_classes.values(), yticklabels=fashion_classes.values())
plt.tick_params(axis='both', which='major', labelsize=8, labelbottom = False, bottom=False, top = False, left = False, labeltop=True)
# set plot title
plt.title('FashionMNIST classification matrix')
# set axis labels
plt.xlabel('[true label]')
plt.ylabel('[predicted label]');
# + [markdown] id="Ob6vR-e3Ww1u"
# Ok, we can easily see that our current model confuses sandals with either sneakers or ankle boots. However, the inverse does not really hold. The model sometimes confuses sneakers with ankle boots, and only very rarely with sandals. The same holds ankle boots. Our model also has issues distinguishing shirts from coats (and, to a lesser degree, from T-shirts and pullovers).
#
# These mistakes are not very surprising, as these items exhibit a high similarity.
# + [markdown] id="e1l8HbUzWw1v"
# ## 7. Lab Summary:
# + [markdown] id="S3P5e48aWw1w"
# In this lab, a step by step introduction into the **design, implementation, training and evaluation** of neural networks to classify images of fashion items is presented. The code and exercises presented in this lab may serves as a starting point for developing more complex, more deep and tailored **neural networks**.
| resources/lab_04/colab_04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DI779AZVZbVs"
# # "Python Review for Deep Learning"
# > "Comprehensive python review"
#
# - toc:true
# - branch: master
# - badges: true
# - comments: true
# - author: omsace
# - categories: [python, deep learning, jupyter]
# + [markdown] id="Lb6SZUTtiNit"
# Colab 이용하여 파이썬에 대해 배워보자
# 괜찮아....
# + colab={"base_uri": "https://localhost:8080/"} id="RpBb2edDiQ9J" outputId="6d6b6965-0ba7-4045-86ac-a4902a624514"
2 + 3
# + colab={"base_uri": "https://localhost:8080/"} id="qXRYTsMiiXK7" outputId="51604425-3a32-4e77-ba19-60eaee9a118c"
print("hello world!")
# + colab={"base_uri": "https://localhost:8080/"} id="tGnzi4g5iZPr" outputId="f7cc1602-fd27-4785-868c-cd89b5186b41"
# !python --version
# + [markdown] id="0-lVi0SPitkD"
# ## **변수와 자료형에 대하여**
# + [markdown] id="9S-VfGzHi2zE"
# 변수 : 자료를 저장하는 이시 저장소
# 숫자 또는 문자를 저장
# + [markdown] id="gJ8Cat_injFt"
# **변수(Variables)**
# + [markdown] id="VXgVpxC3k_nS"
# 1. 변수의 선언
# 2. 변수의 초기화
# 3. 변수 값 할당
# 의 순서로 진행한다..
# + id="wpuFDUDFi6Iv"
score = 0 # 변수의 초기화
# + colab={"base_uri": "https://localhost:8080/"} id="DDvzfQS2loJ0" outputId="8dd70a38-24f3-451d-8f68-987fc7943bac"
print(score)
# + id="7gDVO3AGloHy"
score = 10 # 변수값 할당
# + colab={"base_uri": "https://localhost:8080/"} id="cAu8v5HmloFT" outputId="a133fcb9-4483-4c94-a8ac-3dbb9807aea9"
print(score)
# + id="lfhY0_rBloC7"
level = 1
# + [markdown] id="6vzwazVqnqi-"
# Data type
# + id="i7EUYpBDlns1"
sound = True
# + colab={"base_uri": "https://localhost:8080/"} id="H8sgyMLun2dY" outputId="7b5f2547-3f69-4a4a-946d-025ac0c13861"
type(sound)
# + id="dRqM88wFn7bV"
| _notebooks/2022_03_29_theFirstPython.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from path import Path
import pandas as pd
import bjorn_support as bs
import subprocess
import shlex
import mutations as bm
date = '13-01-2021'
out_dir = Path(f'/home/al/analysis/alab_mutations_{date}')
# Path.mkdir(out_dir)
ref_fp = Path('/home/al/data/hcov19/NC045512.fasta')
# bs.align_fasta_reference(fa_fp, num_cpus=25, ref_fp)
in_dir = Path('/home/al/code/HCoV-19-Genomics/consensus_sequences/')
# fa_fp = bs.concat_fasta(in_dir, out_dir/'seqs')
msa_fp = '/home/al/analysis/alab_mutations_01-01-2020/seqs_aligned.fa'
meta_fp = '/home/al/code/HCoV-19-Genomics/metadata.csv'
# +
# seqsdf = bm.identify_replacements_per_sample(msa_fp, meta_fp, bm.GENE2POS)
# seqsdf
# +
# subs = bm.identify_replacements(msa_fp, meta_fp, location='Jordan')
# +
# subs.sort_values('num_samples', ascending=False).to_csv(out_dir/'jordan_substitutions_01-01-2020.csv', index=False)
# +
# dels = bm.identify_deletions(msa_fp, meta_fp, location='Jordan')
# +
# dels.sort_values('num_samples', ascending=False).to_csv(out_dir/'jordan_deletions_01-01-2020.csv', index=False)
# -
gisaid_meta = Path('/home/al/analysis/gisaid/metadata_2021-01-13_09-12.tsv.gz')
gisaid_msa = Path('/home/al/analysis/gisaid/sequences_2021-01-13_08-22_aligned.fasta')
gisaid_subs = bm.identify_replacements(gisaid_msa, gisaid_meta, data_src='gisaid')
gisaid_subs.to_csv(out_dir/f'gisaid_substitutions_agg_{date}.csv', index=False, compression='gzip')
# +
# date = '01-01-2021'
# gisaid_subs_long.to_csv(out_dir/f'gisaid_substitutions_{date}.csv', index=False)
# -
gisaid_subs = pd.read_csv(out_dir/f'gisaid_substitutions_agg_{date}.csv', compression='gzip')
gisaid_subs = (gisaid_subs.rename(columns={'num_samples': 'gisaid_num_samples', 'first_detected': 'gisaid_1st_detected', 'last_detected': 'gisaid_last_detected',
'num_locations': 'gisaid_num_locations', 'locations': 'gisaid_locations', 'location_counts': 'gisaid_location_counts',
'num_divisions': 'gisaid_num_states','divisions': 'gisaid_states', 'division_counts': 'gisaid_state_counts',
'num_countries': 'gisaid_num_countries', 'countries': 'gisaid_countries', 'country_counts': 'gisaid_country_counts'})
.drop(columns=['ref_aa', 'pos']))
old_subs = pd.read_csv('/home/al/code/HCoV-19-Genomics/variants/substitutions_01-01-2021.csv')
old_subs.head()
subs = pd.read_csv(out_dir/f'alab_substitutions_wide_{date}.csv')
subs.sort_values('num_samples', ascending=False)
# subs = pd.read_csv(out_dir/'substitutions_aggregated.csv')
all_subs = pd.merge(subs, gisaid_subs, on=['gene', 'codon_num', 'alt_aa'], how='left').drop_duplicates(subset=['gene', 'codon_num', 'alt_aa'])
cols = ['gene', 'ref_codon', 'alt_codon', 'pos', 'ref_aa', 'codon_num',
'alt_aa', 'num_samples', 'first_detected', 'last_detected', 'gisaid_num_samples',
'locations', 'location_counts', 'samples',
'gisaid_1st_detected', 'gisaid_last_detected', 'gisaid_num_locations',
'gisaid_locations', 'gisaid_location_counts', 'gisaid_num_states',
'gisaid_states', 'gisaid_state_counts',
'gisaid_num_countries', 'gisaid_countries', 'gisaid_country_counts']
# all_subs[cols].sort_values('num_samples', ascending=False).to_csv(out_dir/f'jordan_substitutions_{date}.csv', index=False)
all_subs[cols].sort_values('num_samples', ascending=False).to_csv(out_dir/f'alab_substitutions_{date}.csv', index=False)
all_subs[cols]
all_subs[all_subs['codon_num']==614]
gisaid_dels = bm.identify_deletions(gisaid_msa, gisaid_meta, min_del_len=1, data_src='gisaid')
# +
# gisaid_dels.sort_values('num_samples', ascending=False)
# -
gisaid_dels.to_csv(out_dir/f'gisaid_deletions_agg_{date}.csv', index=False, compression='gzip')
gisaid_dels = pd.read_csv(out_dir/f'gisaid_deletions_agg_{date}.csv', compression='gzip')
gisaid_dels = (gisaid_dels.rename(columns={'num_samples': 'gisaid_num_samples', 'first_detected': 'gisaid_1st_detected', 'last_detected': 'gisaid_last_detected',
'locations': 'gisaid_locations', 'location_counts': 'gisaid_location_counts',
'divisions': 'gisaid_states', 'division_counts': 'gisaid_state_counts',
'countries': 'gisaid_countries', 'country_counts': 'gisaid_country_counts'})
.drop(columns=['ref_aa', 'pos', 'type', 'ref_codon', 'prev_5nts', 'next_5nts', 'relative_coords', 'del_len']))
dels = pd.read_csv(out_dir/f'alab_deletions_wide_{date}.csv')
# dels = pd.read_csv(out_dir/'deletions_aggregated.csv')
all_dels = pd.merge(dels, gisaid_dels, on=['gene', 'codon_num', 'absolute_coords'], how='left').drop_duplicates(subset=['gene', 'codon_num', 'absolute_coords'])
cols = ['type', 'gene', 'absolute_coords', 'del_len', 'pos',
'ref_aa', 'codon_num', 'num_samples',
'first_detected', 'last_detected', 'locations',
'location_counts', 'gisaid_num_samples',
'gisaid_1st_detected', 'gisaid_last_detected', 'gisaid_countries', 'gisaid_country_counts',
'gisaid_states', 'gisaid_state_counts', 'gisaid_locations', 'gisaid_location_counts', 'samples',
'ref_codon', 'prev_5nts', 'next_5nts'
]
all_dels[cols].sort_values('num_samples', ascending=False).to_csv(out_dir/f'alab_deletions_{date}.csv', index=False)
all_dels[cols].sort_values('num_samples', ascending=False).to_excel(out_dir/f'jordan_deletions_{date}.xlsx', index=False)
df = pd.read_csv('/home/al/code/HCoV-19-Genomics/variants/deletions_13-01-2021.csv')
df.head()['samples']
df = pd.read_csv(gisaid_meta, sep='\t')
df.columns
df['virus'].unique()
df.drop_duplicates(subset=['strain'], inplace=True)
df = df[df['host']=='Human']
df['submitting_lab'].value_counts()[:15]
| notebooks/alab_mutations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: LSST
# language: python
# name: lsst
# ---
# # TAP_verify_DP0.1-general
#
# <br>Owner: **<NAME>** ([@douglasleetucker](https://github.com/LSSTScienceCollaborations/StackClub/issues/new?body=@douglasleetucker))
# <br>Updated for DC2 by: <NAME> following in part work for DESC by <NAME> (@yymao) and <NAME> (@johannct)
# <br>Last Verified to Run: **2021-08-17**
# <br>Verified Stack Release: **w_2021_25**
#
# ### Objectives
#
# This notebook is meant to run tests of the basic content of DP0.1 TAPserver tables in the `dp01_dc2_catalogs` schema on the IDF.
#
# ### Logistics
#
# This notebook is intended to be runnable on `data.lsst.cloud` from a local git clone of https://github.com/LSSTScienceCollaborations/StackClub.
#
# +
# https://jira.lsstcorp.org/browse/PREOPS-473
#
# create a separate notebook "for general consumption" (with more notes).
# More important now than comparing against parquet files.
# "Here's a description of what the DB looks like. -- a good product to hand off."
# Maybe do some of this within the SQL query: can SQL test for NaN's?"
# Point towards the data, not so much the DB mechanics.
# Cols: max, min, 1%, 99%
# qserv has a data return limit of 5GB; so beware!
# -
# ### Set Up
# Import general python packages
import numpy as np
import pandas as pd
import os
from datetime import datetime
import gc
# Ignore warnings
import warnings
warnings.filterwarnings('ignore')
# +
# Import the Rubin TAP service utilities
from rubin_jupyter_utils.lab.notebook import get_tap_service, retrieve_query, get_catalog
# Get an instance of the TAP service
service = get_tap_service()
assert service is not None
assert service.baseurl == "https://data.lsst.cloud/api/tap"
# -
# ### User Input
# +
# Which table are we interested in for this notebook?
schema_name = 'dp01_dc2_catalogs'
# Output directory and output file basename info
outputDir = '/home/douglasleetucker/WORK'
baseName = 'TAP_verify_DP01'
# Debug option for quick results for testing purposes:
# True: only runs over a few tracts of data
# False: runs over all tracts available
debug = True
# Do final cleanup?
do_final_cleanup = False
# Level of output to screen (0, 1, 2, ...)
verbose = 2
# -
# ### Basic Tests
# Since some of the following queries take time, let's measure how long it takes to complete each query cell. We will use `%%time` to estimate processor and wall clock time, and `datetime.now()` to estimate the total time to send the query and download its results.
# +
# %%time
now0=datetime.now()
query = """SELECT table_name FROM TAP_SCHEMA.tables WHERE schema_name=%s""" % ("\'"+schema_name+"\'")
print(query)
results = service.search(query)
df = results.to_table().to_pandas()
table_full_name_list = df['table_name'].tolist()
print(table_full_name_list)
# +
# %%time
now0=datetime.now()
for table_full_name in table_full_name_list:
query = """SELECT COUNT(*) FROM %s""" % (table_full_name)
results = service.search(query)
df = results.to_table().to_pandas()
ntot = df['COUNT'].loc[0]
outputLine = """%40s: %d""" % (table_full_name, ntot)
print(outputLine)
now1=datetime.now()
print("Total time:", now1-now0)
| notebooks/TAP_verify_DP0.1-general.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# hide
import numpy as np
import pandas as pd
from forma.judge import FormatJudge
from forma.detector import FormatDetector
from forma.utils import PatternGenerator
# -
# # Forma
#
# > Automatic format error detection on tabular data.
# Forma is an open-source library, written in python, that enables automatic and domain-agnostic format error detection on tabular data. The library is a by-product of the research project [BigDataStack](https://bigdatastack.eu/).
# ## Install
# Run `pip install forma` to install the library in your environment.
# ## How to use
# We will work with the the popular [movielens](https://grouplens.org/datasets/movielens/) dataset.
# local
# load the data
col_names = ['user_id', 'movie_id', 'rating', 'timestamp']
ratings_df = pd.read_csv('../data/ratings.dat', delimiter='::', names=col_names, engine='python')
# local
ratings_df.head()
# Let us introduce some random mistakes.
# +
# local
dirty_df = ratings_df.astype('str').copy()
dirty_df.iloc[3]['timestamp'] = '9783000275'
dirty_df.iloc[2]['movie_id'] = '914.'
dirty_df.iloc[4]['rating'] = '10'
# -
# Initialize the detector, fit and detect. The returned result is a pandas DataFrame with an extra column `p`, which records the probability of a format error being present in the row. We see that the probability for the tuples where we introduced random artificial mistakes is increased.
# +
# local
# initialize detector
detector = FormatDetector()
# fit detector
detector.fit(dirty_df, generator= PatternGenerator(), n=3)
# detect error probability
assessed_df = detector.detect(reduction=np.mean)
# visualize results
assessed_df.head()
| nbs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from text_selector.widget import Widget
from ipywidgets import HBox, Button, Label, VBox
# +
def callback(obj):
# print(obj)
with open('res.txt', 'w') as res:
print(obj['new'], file=res)
txt = "The human TCF-1 gene encodes a nuclear DNA-binding protein uniquely expressed in normal and neoplastic T-lineage lymphocytes ."
tags = [
'protein',
'DNA',
'cell_type',
'cell_line',
'RNA'
]
w1 = Widget(tags=tags, txt=txt, colors=[], callback=callback)
w1.res = [{'start': 3, 'end': 9, 'tag': 'protein'}, {'start': 15, 'end': 20, 'tag': 'DNA'}, {'start': 30, 'end': 38, 'tag': 'cell_type'}, {'start': 42, 'end': 50, 'tag': 'cell_line'}, {'start': 58, 'end': 67, 'tag': 'RNA'}]
w2 = Widget(tags=tags, txt=txt, colors=[], callback=callback, emojify=True)
w2.res = [{'start': 15, 'end': 20, 'tag': 'DNA'}]
# -
VBox([w1, w2])
w2.emojify = True
w2
print(w1.res)
print(w2.res)
| workspace/example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Welcome!
#
# # Practical 1 - An introduction to data types, operations and loops
#
# Welcome to the first practical in our Python programming course. Over the course of this module we are going to take an applied approach to learning Python, covering basic numerical computation to data input and analysis. We cant teach everything Python has to offer in this course, but we can given you a set of tools to begin your journey into the wonderful world of programming and data analysis.
#
# In today's practical we are going to cover some core operations most Python programs will have. This gives you chance to learn the syntax that the Python interpreter expects. Remember, learning to program is similar to learning any other language, including a general appreciation of what the hardware underneath the software is doing.
#
# <div class="alert alert-block alert-success">
# <b>Objectives:</b> The objectives of todays practical are:
#
# - 1) [Understand how data types are represented and common arithmetic operators](#Part1)
# * [Exercise 1: Set the value of two new variables and print to screen](#Exercise1)
# * [Exercise 2: Implement equations as combinations of arithmetic operators](#Exercise2)
# - 2) [Arrays, lists and tuples](#Part2)
# * [Exercise 3: Add two more entries to a list and change an existing entry](#Exercise3)
# - 3) [Loops and conditional statements](#Part3)
# * [Exercise 4: Cycling through arrays and modifying values](#Exercise4)
#
# Please note that you should not feel pressured to complete every exercise in class. These practicals are designed for you to take outside of class and continue working on them. Proposed solutions to all exercises can be found in the 'solutions' folder.
# </div>
# <div class="alert alert-block alert-warning">
# <b>Please note:</b> After reading the instructions and aims of any exercise, search the code snippets for a note that reads ------'INSERT CODE HERE'------ to identify where you need to write your code
# </div>
# ## Understand how data types are represented in Python <a name="Part1">
#
# In the following we will see how both numeric and string variables are handled and manipulated. For this we will first look at the type of variables and then how they are stored and processed.
#
# ### Numbers (integers, floats and complex numbers):
#
# As per the official Python documentation, there are three distinct numeric types:
#
# - integers
# - floating point numbers
# - complex numbers.
#
# We talk briefly about this in class, and will revisit this in more detail as we move into our course. Integers have unlimited precision in that you could use an integer with a growing number of digits that would be limited by the memory on your machine. Floating point numbers are usually implemented using double precision. Complex numbers have a real and imaginary part, which are each a floating point number. Lets see what this means using our first bit of code. In the following 'code box' Im going to use three variables **x,y and z** and assign them either an integer, floating point or complex value.
#
# | Type | Format | Description |
# | --- | --- | --- |
# | int [Integer] | a = 10 | Signed Integer |
# | float [Floating point] | a = 45.67 | Floating point real values |
# | complex | a = 3.14J | (J) Contains integer in the range 0 to 255. |
#
# +
# Whenever you see a hash symbol [#] in Python code, this represents a comment which is ignored by the interpreter.
# This is where a code developer will add some personal comments or add key notes that enable others to understand
# and/or use the code she/he developed.
# So let's assign some values to some variables. By the way, the variable names are not important, but are more
# of an issue regarding style and documentation.
# Also note that Im not defining what each variable is expecting in the first place.
# In other words, I dont have to pre-define to the interpreter what data type to expect in, say, x:
x = 2.3 # float
y = 4 # integer
z = 5 + 6j # complex [we are not going to use complex numbers in our course so this is the first and last instance]
# -
# If you click 'Run' on the above cell, you should notice nothing happens. Not very interesting, but lets print our variables to screen using the 'print' function in Python 3 as per below. Select the cell below and click 'Run' from the top menu:
# Just adding comment here for the sake of it...
print("x =", x)
print("y =", y)
print("z =", z)
# Looks correct! We can also ask Python to check the numeric type using the 'type' function in Python. Thus:
print("The data type of x is",type(x))
print("The data type of y is",type(y))
print("The data type of z is",type(z))
# Let's look at some more specifics for type _float_ from the official Python documentation. We have a number of formatting permutations we could use:
#
# | Property | Syntax |
# |------|------|
# | It can have a sign | "+ -" |
# | We can express infinity as | "Infinity" "inf" |
# | If we dont have a number, referred to as 'Not a Number' [NaN] | "nan" |
#
# For example, look at the following examples where Im using the command 'float' to print out the floating point number of a string in the brackets ().
#
# ```python
# >>> float('+1.23')
# 1.23
# >>> float(' -12345')
# -12345.0
# >>> float('1e-003')
# 0.001
# >>> float('+1E6')
# 1000000.0
# >>> float('-Infinity')
# -inf
# >>> float('NaN')
# nan
# ```
#
# We can use a positive or negative assignment. Notice also the use of lower case 'e' and upper case 'E'. This is used as floats may also be in scientific notation, with E or e indicating the power of 10 (2.5e2 = $2.5 x 10^{2} = 250$).
# <div class="alert alert-block alert-success">
# <b> Exercise 1. Set the value of two new variables and print to screen <a name="Exercise1"> </b>
#
# Set the value of two new variables 'temperature' and 'mass' equal to 298 Kelvin [as an integer] and 3.78 Kilograms [as a float]. Then print their data type to the screen.
#
# </div>
# +
#------'INSERT CODE HERE'------
temperature = 298 # Kelvin
mass = 3.78 # Kg
#------------------------------
print("The data type of temperature is",type(temperature))
print("The data type of mass is",type(mass))
# -
# ### Arithmetic Operators (add, subtract, multiply, divide and raise to a power)
#
# | Operation | Result |
# | --- | --- |
# | x + y | sum of x and y |
# | x - y | difference of x and y |
# | x * y | product of x and y |
# | x / y | quotient of x and y |
# | abs(x) | absolute value or magnitude of x |
# | int(x) | x converted to integer |
# | float(x) | x converted to floating point |
# | pow(x, y) | x to the power y |
# | x ** y | x to the power y |
#
#
# In the following code we demonstrate how to perform basic arithmetic on either defined variables or pure numbers using appropriate Python syntax, and add comments using the # symbol.
# +
# Addition examples [+]
# 1) Define variables and add
x = 2.4
y = 7.5
z = x + y
print("z = ", z)
# Subtraction examples [-]
var1 = -3.4
var2 = 4.7
var3 = var1 + var2
print("var3 = ", var3)
# Multiplication examples [*]
starting_temperature = 273.14 # Kelvin
d_temp_d_time = 3.4 # Kelvin per second
duration = 12.5 # seconds
end_temperature = starting_temperature + d_temp_d_time*duration
print("end_temperature = ", end_temperature)
# Division examples
Price = 12 # £ sterling
weight = 2.3 # kg
price_per_kg = Price / weight # £ per kg
print("price_per_kg = ", price_per_kg)
# Exponetation examples
base = 2.3
power = 4.5
result = base ** power
print("result = ", result)
# -
# In the division example, did you notice we actually asked Python to divide an integer by a float? This is ok in the sense that Python will, by default, change the type of the variable with the lowest precision to the same as the highest.
#
# <div class="alert alert-block alert-success">
# <b> Exercise 2. Implement equations as combinations of arithmetic operators <a name="Exercise2"> </b> Implement the following equations as combinations of arithmetic operators given above:
#
#
# | Description | Form |
# | --- | --- |
# |<img width=200/>|<img width=500/>|
# | Einstein's relativistic mass-energy relation | $E = mc^2$ |
# | De Broglie Wavelength | $\lambda = \frac{h}{{m\upsilon }}$ |
# | Ideal gas equation | $PV = nRT$ |
# | Temperature from Degrees Celsius to Kelvin | $K = ^\circ C + 273$ |
#
# Where we provide values for the following constants referred to in each:
#
# - Planck's constant, h =
# - Speed of light in a vaccuum, c = $3 x 10^{8} m s^{-1}$
# - Ideal Gas constant, R = 8.205736×$10^{-5}$ $\frac{m3⋅atm}{K⋅mol}$
#
# </div>
#
#
#
# +
# Constants used in equations
c = 3.0e8 # Speed of light, [m/s]
h = 6.62607004e-34 # Plank's constant [J s]
m = 9.11e-31 # Mass of Electron in kg
T = 298.15 # Assume temperature is 298.15 K
R = 8.205736e-5 # Ideal Gas Constant [m3⋅atm/ K⋅mol]
V = 1.0 # Assumed volume in ideal gas law
n = 2.15 # Number of moles of gas used in the ideal gas law
vel = 5.31e6 # Assumed velocity of eletron in De Broglies rule [m/sec]
#------'INSERT CODE HERE'------
# Einsteins relativistic mass-energy relation
E = m*c**2.0 # Energy of electron [J]
# De Broglie's Wavelength
Lambda = h/(m*vel) # Wavelength [m]
# Ideal gas equation
P = (n*R*T)/(V) # Pressure [atm]
# Covert above equation to degrees Celsius
T_celsius = 28.5+273.0
#------------------------------
print("The relativistic energy of an electron = ", E)
print("The De Broglie wavelength of an eletronc = ",Lambda)
print("The Pressure of our ideal gas = ",P)
print("The Temperature in Celsius = ",T_celsius)
# -
# ## Arrays: Lists, tuples and dictionaries <a name="Part2">
#
# In most programming applications we want to store and update values in an array, be it 1D, 2D, 3D or even more. Python comes with 3 ways to store this information known as:
#
# - lists
# - tuples
# - dictionaries.
#
# For applications focused on numerical computations, the package known as Numpy is a package used for manipulating and creating/modifying numerical values in arrays. The reason is that Numpy has been optimised for use in this way and can be significantly faster than non-Numpy approaches we will use in this class. We will be using the Numpy package throughout this course, but we will also come across a 'list'.
#
# So, why are there multiple methods?
#
# Lists and tuples are perhaps the most common used data structures in Python. They have a number of similarities:
#
# - They both store a collection of items sequentially
# - They can store items of any data type. So this include numeric and non-numeric.
# - Any item is accessible via its index. We will see this below
#
#
# <div class="alert alert-block alert-danger">
# <b>Indexing </b> Please note, Python indexes start at 0
# </div>
#
# However they have a key difference which is:
#
# >> Lists are mutable while tuples are immutable. This means we can change/modify the values of a list but we cannot change/modify the values of a tuple.
#
# They also have a different syntax. Let's run through some examples below. Notice again Im using whatever variable name I want. To re-run the code in a box, such as that provided below, click anywhere in the box and then click on the 'Run' symbol in the notebook toolbar. For example, once you have read through the below, try accessing a different element from the list.
# +
# First let's create an identical list and tuple containing names of places
# Lists use square brackets, whereas tuples use parentheses
list_capitals = ['London','Paris','Rome','Ankara']
tuple_capitals = ('London','Paris','Rome','Ankara')
# If we want to access a particular element from a list of a tuple, we use the same bracket notation but also
# remember that python indexing starts at 0. Thus, for the first element in the list we would write:
list_capitals[0]
# Now click run on this portion of code and you should see the word 'London' below, if accessing element [0].
# +
# Now lets access the 4th element of both the list and tuple. Note im using the print command here so we can see
# both printed to the screen:
print("4th entry in the list is ",list_capitals[3])
print("4th entry in the tuple is ",tuple_capitals[3])
# -
# The key different between lists and tuples are the ability to change values. This impacts on efficiency, which we wont cover in detail during this course; Lets try to change and then print the 2nd entry in both the list and tuple. Let's try to change this to 'Beijing'. See the code below. When you click 'Run', does it produce an error?
# +
# Change the 2nd entry to 'Beijing'
list_capitals[1] = 'Beijing' # note the index value and the [] bracket notation
# print full array to the screen
print("full list array = ",list_capitals)
tuple_capitals[1] = 'Beijing' # note the index value and the [] bracket notation
# print full array to the screen
print("full list array = ",tuple_capitals)
# -
# You will see an error as we cannot change the entry in the tuple. Following on from this, we can change the size of a list, but we cannot change the size of a tuple. Imagine we want to add a 5th entry to our list. To do this we can append to the list by using the
#
# ```python
# >>[Name of list].append()
# ```
#
# operation. Thus, look at the following code:
# Append another city to our list as follows, and then print the new array
list_capitals.append('Moscow')
# print full array to the screen
print("full list array = ",list_capitals)
# <div class="alert alert-block alert-success">
# <b> Exercise 3. Add two more entries to a list and change an existing entry <a name="Exercise3"> </b>
#
# Append to your existing list the following cities, in sequential order:
# - Manchester
# - Edinburgh
#
# Then change the existing value of the 5th element, which is currently 'Moscow', to 'Hanoi'. When youy have finished this, and run the code, you should see an output resembling:
#
# ```python
# full list array = ['London', 'Beijing', 'Rome', 'Ankara', 'Hanoi', 'Manchester', 'Edinburgh']
# ```
# </div>
# +
#------'INSERT CODE HERE'------
list_capitals.append('Manchester')
list_capitals.append('Edinburgh')
list_capitals[4] = 'Hanoi' # note the index value and the [] bracket notation
#------------------------------
# print full array to screen
print("full list array = ",list_capitals)
# -
# ## Loops and conditional statements <a name="Part3">
#
# Once we have information stored in an array, or wish to generate information iteratively, we start to use a combination of **loops** and **conditional statements**. Let us focus on loops during this practical.
#
#
# ### Loops
# Take the previous list, or array, of city names. We could, as we have done, print each entry individually like so:
# +
print("first entry is",list_capitals[0])
print("second entry is",list_capitals[1])
print("third entry is",list_capitals[2])
#...and so on
# -
# However we often work with very large arrays and this simply wouldnt be feasible or just too time consuming. We might want to generate values to be stored in a very large array, based on a specific algorithm, which would be better automated. Loops allow us to do this. Take, again, our previously constructed list. The following code demonstrates using a loop to print each value in turn. Take note of the way the loop is contructed, the syntax and the need for introducing a space within the loop. We will discuss this shortly. In fact, obeying the white space rule is a fundamental requirement in Python
#
# <div class="alert alert-block alert-danger">
# <b>White space rule </b> Whenever a series of commands, or command, is terminated by a colon :, the following line should be indented. Examples typically include loops and function definitions.
# </div>
#
# Example loop to print entries in our list of capital cities
for step in range(7): # I have introduced a variable called 'step'. It dosnt matter what name we use
# but see how the same variable, once defined by the loop, is used to access entries in our array
print(list_capitals[step])
# Let us break this down a little. The loop counts through a fixed number of integer values using the 'range' function. We will start looking at functions in our third practical. Briefly, this Python function creates a sequence of numbers that define each iteration of the loop. By default, this sequence is based on integer values but you can modify this by passing optional arguments which we will not cover in this practical. For now, this generates values from 0 to 6. Recall, Python indexing starts at 0. In fact, lets add another line within our loop such that we can access the value of our 'step' variable within each loop iteration as follows:
# +
# The following loop counts through a fixed number of integer values using the range function. We have defined the
# limit of values to be iterated through by knowledge of how big our array is. If you ask Python to print
# any entries beyond the boundaries of our array, you would get an error message. Try it!
for step in range(7): # I have introduced a variable called 'step'. It dosnt matter what name we use
# but see how the same variable, once defined by the loop, is used to access entries in our array
print(list_capitals[step])
print("the value of step is ", step)
# -
# We have only looked at a list containing strings, or words. Lets try a numerical example. Assume we want to create a two new lists, x and y. Both have 100 elements. The x array contains integer values from 1 to 100, and the y array is defined by the basic equation y=x*3.4. Check out the code below. This uses a combination of creating a new list, defining a loop and applying a numerical operation:
# +
# Create two empty lists. We are going to fill these in the loop
x = []
y = []
# Create a loop that cycles through 100 values, then implement the equation defined above
for step in range(100):
# First lets define a value in our x array. We know x goes from 1-100 so, knowing Python
# indexing starts at 0, do you understand the following operation
x.append(step+1)
# So for our first iteration, step is 0 and thus we place the value '1' into x
# We might aswell also define our y values.
# Take your time to understand whats going on below.
y.append(x[step]*3.4)
# We can break it down into 3 phases
# 1) We want to append a value to our y array using .append()
# 2) We want to take the associated value from the x array using our index given by the 'step' variable
# 3) We want to implement the function given by equation xx.
# Below the indented space above, the loop has finished. Lets print the first 4 values from both
# our x and y values
print("The 1st element of x is ",x[0])
print("The 1st element of y is ",y[0])
print("The 2nd element of x is ",x[1])
print("The 2nd element of y is ",y[1])
print("The 3rd element of x is ",x[2])
print("The 3rd element of y is ",y[2])
print("The 4th element of x is ",x[3])
print("The 4th element of y is ",y[3])
# What about the 81st element of x and y?
print("The 81st element of x is ",x[80])
print("The 81st element of y is ",y[80])
# -
# <div class="alert alert-block alert-success">
# <b> Exercise 4. Cycling through arrays and modifying values <a name="Exercise4"> </b> Create a loop that implements the function:
#
# \begin{eqnarray}
# Y = X^{2.8}
# \end{eqnarray}
#
# Where x is an array of 50 values from 1 to 50.
# </div>
#
# +
# Initiliase an empty list for both 'x' and 'y'
x = []
y = []
# Now loop through 50 values and append each list accordingly.
# One list contained values for 'x', the other 'y'.
# Please note the operator ** is needed to raise one number to another [e.g 2**3]
#------'INSERT CODE HERE'------
for step in range(50):
# Append a value to our x array
x.append(step+1)
# Append a value to our y array
y.append(x[step]**2.8)
#------'INSERT CODE HERE'------
# Print the last four values from both our x and y arrays
print("The 47th element of x is ",x[46])
print("The 47th element of y is ",y[46])
print("The 48th element of x is ",x[47])
print("The 48th element of y is ",y[47])
print("The 49th element of x is ",x[48])
print("The 49th element of y is ",y[48])
print("The 50th element of x is ",x[49])
print("The 50th element of y is ",y[49])
# -
| solutions/.ipynb_checkpoints/Practical 1. Introduction to data types and numerical operations-checkpoint.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; 変換$T_{pq}$を行列で表すと、
;; $$
;; T_{pq} =
;; \begin{pmatrix}
;; (p+q) && q \\
;; q &&p
;; \end{pmatrix}
;; $$
;; これを用いて、
;; $$
;; \begin{pmatrix}
;; a \\
;; b
;; \end{pmatrix}
;; \leftarrow
;; T_{pq}
;; \begin{pmatrix}
;; a \\
;; b
;; \end{pmatrix}
;; $$
;; $T_{pq}$を二回繰り返すと、
;; $$
;; T_{p'q'}=T_{pq}^2=
;; \begin{pmatrix}
;; (p^2+q^2)+(2pq+q^2) && (2pq+q^2) \\
;; (2pq+q^2) && (p^2+q^2)
;; \end{pmatrix}
;; $$
;; となる。
;; したがって、
;; $$
;; \begin{cases}
;; p'=(p^2+q^2)\\
;; q'=(2pq+q^2)
;; \end{cases}
;; $$
;; + vscode={"languageId": "python"}
(define (square x) (* x x))
(define (fib n)
(fib-iter 1 0 0 1 n))
(define (fib-iter a b p q count)
(cond ((= count 0) b)
((even? count)
(fib-iter a
b
(+ (square p) (square q)) ; compute p'
(+ (* 2 p q) (square q)) ; compute q'
(/ count 2)))
(else (fib-iter (+ (* b q) (* a q) (* a p))
(+ (* b p) (* a q))
p
q
(- count 1)))))
;; + vscode={"languageId": "python"}
;確認
(display (format "~a\n" (fib 0)))
(display (format "~a\n" (fib 1)))
(display (format "~a\n" (fib 2)))
(display (format "~a\n" (fib 3)))
(display (format "~a\n" (fib 4)))
(display (format "~a\n" (fib 5)))
(display (format "~a\n" (fib 6)))
(display (format "~a\n" (fib 7)))
(display (format "~a\n" (fib 8)))
(display (format "~a\n" (fib 9)))
(display (format "~a\n" (fib 10)))
(display (format "~a\n" (fib 11)))
(display (format "~a\n" (fib 12)))
;; -
;; $$
;; T =
;; \begin{pmatrix}
;; p && q \\
;; r &&s
;; \end{pmatrix}
;; $$
;; として、
;; $$
;; \begin{pmatrix}
;; a \\
;; b
;; \end{pmatrix}
;; \leftarrow T
;; \begin{pmatrix}
;; a \\
;; b
;; \end{pmatrix}\\
;; $$
;; $$
;; T^2 =
;; \begin{pmatrix}
;; p^2+qr && pq+qs \\
;; rp+sr && rq+s^2
;; \end{pmatrix} \\
;; $$
;; を利用する。
;; 問題文の方式では変数が2個だが、こちらは4個なので多少遅い。
;; とは言え、行列の冪の問題に行き着くので、整数のべきと同じ考えが適用できるのでわかりやすいし、
;; 思いつくのはこちらのほうが容易いと考えられる。
;; + vscode={"languageId": "python"}
;行列バージョン((p,q),(r,s))の積で考える
(define (fib n)
(fib-iter 1 0 1 1 1 0 n))
(define (fib-iter a b p q r s count)
(cond ((= count 0) b)
((even? count)
(fib-iter a
b
(+ (* p p) (* q r)) ; compute p
(+ (* p q) (* q s)) ; compute q
(+ (* r p) (* s r)) ; compute r
(+ (* r q) (* s s)) ; compute s
(/ count 2)))
(else (fib-iter (+ (* p a) (* q b))
(+ (* r a) (* s b))
p
q
r
s
(- count 1)))))
;; + vscode={"languageId": "python"}
;確認
(display (format "~a\n" (fib 0)))
(display (format "~a\n" (fib 1)))
(display (format "~a\n" (fib 2)))
(display (format "~a\n" (fib 3)))
(display (format "~a\n" (fib 4)))
(display (format "~a\n" (fib 5)))
(display (format "~a\n" (fib 6)))
(display (format "~a\n" (fib 7)))
(display (format "~a\n" (fib 8)))
(display (format "~a\n" (fib 9)))
(display (format "~a\n" (fib 10)))
(display (format "~a\n" (fib 11)))
(display (format "~a\n" (fib 12)))
;; + vscode={"languageId": "python"}
| ch1/1.2/ex.1.19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="2rY0kAPCKo--" colab_type="code" colab={}
# #!wget http://msvocds.blob.core.windows.net/coco2014/train2014.zip
# #!wget http://msvocds.blob.core.windows.net/annotations-1-0-3/instances_train-val2014.zip
# #!wget http://msvocds.blob.core.windows.net/annotations-1-0-3/captions_train-val2014.zip
# #!wget http://msvocds.blob.core.windows.net/annotations-1-0-4/image_info_test2014.zip
# #!wget http://msvocds.blob.core.windows.net/coco2014/val2014.zip
# + id="--Tt-B4PFn4_" colab_type="code" colab={}
# #!ls
# + id="f4bUwUVvGryz" colab_type="code" colab={}
# #!unzip image_info_test2014.zip
# #!unzip train2014.zip
# #!unzip val2014.zip
# #!unzip instances_train-val2014.zip
# #!unzip captions_train-val2014.zip
# + id="jL-uBvEELrTc" colab_type="code" colab={}
# #!ls
# + id="QTNul2V6S5vX" colab_type="code" colab={}
# #!pip install tqdm
import glob
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# from vgg16 import VGG16
import pickle
from tqdm import tqdm
import pandas as pd
from keras.preprocessing import sequence
from keras.models import Sequential
from keras.layers import LSTM, Embedding, TimeDistributed, Dense, RepeatVector, Merge, Activation, Flatten
from keras.optimizers import Adam, RMSprop
from keras.layers.wrappers import Bidirectional
import json
from keras.applications.inception_v3 import InceptionV3
from keras.preprocessing import image
# + id="eYPb-MBXS-ym" colab_type="code" colab={}
images = 'train2014/'
# Contains all the images
img = glob.glob(images+'*.jpg')
img[:5]
# + id="W-XSdkDHTFGf" colab_type="code" colab={}
len(img)
# + id="bQ4pNI7HUj5z" colab_type="code" colab={}
def preprocess_input(x):
x /= 255.
x -= 0.5
x *= 2.
return x
# + id="b6SFHARYUn1o" colab_type="code" colab={}
def preprocess(image_path):
img = image.load_img(image_path, target_size=(299, 299))
x = image.img_to_array(img)
x = np.expand_dims(x, axis=0)
x = preprocess_input(x)
return x
# + id="JrJftWG0Useg" colab_type="code" colab={}
plt.imshow(np.squeeze(preprocess(img[0])))
# + id="aZB92sR4UvEs" colab_type="code" colab={}
model = InceptionV3(weights='imagenet')
# + id="YlpKGc42Uz_U" colab_type="code" colab={}
from keras.models import Model
new_input = model.input
hidden_layer = model.layers[-2].output
model_new = Model(new_input, hidden_layer)
model_new.summary()
# + id="jE1yGQOaU32I" colab_type="code" colab={}
tryi = model_new.predict(preprocess(img[0]))
# + id="1EwcXijgU7Hn" colab_type="code" colab={}
tryi.shape
# + id="jeNg1Ob7U8eB" colab_type="code" colab={}
def encode(image):
image = preprocess(image)
temp_enc = model_new.predict(image)
temp_enc = np.reshape(temp_enc, temp_enc.shape[1])
return temp_enc
# + id="QyTYOecLU-JE" colab_type="code" colab={}
encoding_train = {}
for img in tqdm(img):
encoding_train[img[len(images):]] = encode(img)
# + id="rBBMc3kYVCsc" colab_type="code" colab={}
with open("encoded_images_train_inceptionV3.p", "wb") as encoded_pickle:
pickle.dump(encoding_train, encoded_pickle)
# + id="80YhbeozVR0r" colab_type="code" colab={}
encoding_train = pickle.load(open('encoded_images_train_inceptionV3.p', 'rb'))
# + id="ZVocplTu1McV" colab_type="code" colab={}
images = 'val2014/'
# Contains all the images
img = glob.glob(images+'*.jpg')
img[:5]
# + id="F-CcVzVCnYms" colab_type="code" colab={}
encoding_val = {}
for img in tqdm(img):
encoding_val[img[len(images):]] = encode(img)
# + id="aVjyql9j1A19" colab_type="code" colab={}
with open("encoded_images_val_inceptionV3.p", "wb") as encoded_pickle:
pickle.dump(encoding_val, encoded_pickle)
# + id="P7YjnNSZ1YeK" colab_type="code" colab={}
#encoding_val = pickle.load(open('encoded_images_val_inceptionV3.p', 'rb'))
# + id="1OqGwOCzVZCF" colab_type="code" colab={}
#from google.colab import files
#files.download('encoded_images_train_inceptionV3.p')
#files.download('encoded_images_val_inceptionV3.p')
# + id="-GjQGO4WnEnN" colab_type="code" colab={}
# #!ls
token = 'annotations/captions_train2014.json'
# + id="1HYsnthwtiXL" colab_type="code" colab={}
json_data = open(token)
# + id="aq7D2K-itiap" colab_type="code" colab={}
captions = json.load(json_data)
for i, val in enumerate(captions['annotations']):
if i<10:
print (val)
# + id="x-dBDGWBtieW" colab_type="code" colab={}
for i, val in enumerate(captions['images']):
if i<2:
print (val)
# + [markdown] id="YLpYB17_uD4K" colab_type="text"
# Creating a dictionary containing all the captions of the images
# + id="qP8qbp6Gt_gD" colab_type="code" colab={}
images = 'train2014/'
img = glob.glob(images+'*.jpg')
img[:5]
# + id="cznt-KqhuWQl" colab_type="code" colab={}
image2id = {}
for i, val in enumerate(captions['images']):
image2id[val['id']] = val['file_name']
# + id="Tt-Yz2ykuXhC" colab_type="code" colab={}
id2captions = {}
for i, val in enumerate(captions['annotations']):
id2captions[val['image_id']] = val['caption']
# + id="LgwauVuAuXkA" colab_type="code" colab={}
image2caption = {}
for key, val in id2captions.items():
image2caption[image2id[key]] = val
# + id="SAxg5PDeuXns" colab_type="code" colab={}
len(image2caption)
# + id="nAwj9ro8uXru" colab_type="code" colab={}
image2caption['COCO_train2014_000000522971.jpg']
# + id="LaBcJSIVt_ch" colab_type="code" colab={}
Image.open(img[0])
# + id="vTvGgOHkufQx" colab_type="code" colab={}
caps = []
for key, val in image2caption.items():
caps.append(' '.join(list(val.split())))
caps[:5]
# + id="RWUSZt9Qu44t" colab_type="code" colab={}
words = [i.split() for i in caps]
unique = []
for i in words:
unique.extend(i)
unique.extend("<start>".split())
unique.extend("<end>".split())
unique.extend("<unk>".split())
# + id="JjJ0lv6hufT_" colab_type="code" colab={}
unique = list(set(unique))
# with open("unique.p", "wb") as pickle_d:
# pickle.dump(unique, pickle_d)
len(unique)
# + id="AspcMEsdufax" colab_type="code" colab={}
word2idx = {val:index for index, val in enumerate(unique)}
word2idx['<start>']
# + id="hw-PXMcLzywl" colab_type="code" colab={}
idx2word = {index:val for index, val in enumerate(unique)}
idx2word[5303]
# + id="xvLXUXu3xIAN" colab_type="code" colab={}
max_len = 0
for c in caps:
c = c.split()
if len(c) > max_len:
max_len = len(c)
max_len
# + id="pXwypaDNxILW" colab_type="code" colab={}
vocab_size = len(unique)
# + id="acCKbXBUxIO5" colab_type="code" colab={}
f = open('mscoco_training_dataset.txt', 'w')
f.write("image_id\tcaptions\n")
# + id="bBgyuRvQxOv4" colab_type="code" colab={}
for key, val in image2caption.items():
f.write(key + "\t" + "<start> " + val +" <end>" + "\n")
f.close()
# + id="I7RhGllmxOzh" colab_type="code" colab={}
df = pd.read_csv('mscoco_training_dataset.txt', delimiter='\t')
# + id="mPBZHArUxO3d" colab_type="code" colab={}
df = df.dropna()
df.head()
len(df)
# + id="ITMaFco3zIVw" colab_type="code" colab={}
#files.download('mscoco_training_dataset.txt')
# + id="_lIAYx3A0A15" colab_type="code" colab={}
c = [i for i in df['captions']]
# + id="4z6ABBwp0JyK" colab_type="code" colab={}
imgs = [i for i in df['image_id']]
# + id="GmQH6xRL0LEN" colab_type="code" colab={}
a = c[0]
a, imgs[0]
# + id="mzOr3uOu0U2R" colab_type="code" colab={}
for i in a.split():
print (i, "=>", word2idx[i])
# + id="K8O-UVlS0XDB" colab_type="code" colab={}
samples_per_epoch = 0
for ca in c:
samples_per_epoch = samples_per_epoch + len(ca.split())-1
# + id="BMjeq1Zy0ZQH" colab_type="code" colab={}
samples_per_epoch
# + id="2GpmXmGS0atG" colab_type="code" colab={}
def data_generator(df, batch_size=32):
# Shuffling the dataframe before creating batches
# df = df.sample(frac=1)
# c = [i for i in df['captions']]
# imgs = [i for i in df['image_id']]
partial_caps = []
images = []
next_words = []
count = 0
for i, text in enumerate(c[:1000]):
current_image = encoding_train[imgs[i]]
for j in range(len(text.split())-1):
count+=1
if text.split()[j] in word2idx:
partial = [word2idx[text.split()[j]]]
else:
partial = [word2idx['<unk>']]
partial_caps.append(partial)
# Initializing with zeros to create a one-hot encoding matrix
# This is what we have to predict
# Hence initializing it with vocab_size length
next = np.zeros(vocab_size)
# Setting the next word to 1
if text.split()[j+1] in word2idx:
next[word2idx[text.split()[j+1]]] = 1
else:
next[word2idx['<unk>']] = 1
next_words.append(next)
images.append(current_image)
if count >= batch_size:
images = np.array(images)
next_words = np.array(next_words)
partial_caps = sequence.pad_sequences(partial_caps, maxlen=max_len, padding='post')
yield [[images, partial_caps], next_words]
partial_caps = []
images = []
next_words = []
count = 0
partial_caps = sequence.pad_sequences(partial_caps, maxlen=max_len, padding='post')
return np.array(partial_caps)
# + id="MHf5Ql8w0dOF" colab_type="code" colab={}
for i, val in enumerate(data_generator(df)):
if i==0:
print (val)
partial_caps = []
next_words = []
images = []
partial_caps = data_generator([partial_caps, next_words, images],32)
# + id="h1F1FOY10fX8" colab_type="code" colab={}
#partial_caps.shape
# + id="9PTOnr3E0hXj" colab_type="code" colab={}
embedding_size = 300
# + id="J310odLE1gI4" colab_type="code" colab={}
image_model = Sequential([
Dense(embedding_size, input_shape=(2048,), activation='relu'),
RepeatVector(1)
])
# + id="dpglHJCR1gMN" colab_type="code" colab={}
caption_model = Sequential([
Embedding(vocab_size, embedding_size, input_length=max_len),
LSTM(256, return_sequences=True),
TimeDistributed(Dense(300))
])
# + id="C4jHIiC81juk" colab_type="code" colab={}
final_model = Sequential([
Merge([image_model, caption_model], mode='concat', concat_axis=1),
Bidirectional(LSTM(256, return_sequences=False)),
Dense(vocab_size),
Activation('softmax')
])
# + id="1Cujr_fj1lPS" colab_type="code" colab={}
final_model.compile(loss='categorical_crossentropy', optimizer=RMSprop(), metrics=['accuracy'])
# + [markdown] id="L0jjEI-WX4gZ" colab_type="text"
#
# + id="7D9CIG-N1oau" colab_type="code" colab={}
final_model.fit_generator(data_generator(df, batch_size=128), nb_epoch=1, steps_per_epoch=948248,verbose=2)
# + id="3JZpTuKC1qy3" colab_type="code" colab={}
# + id="PQ9zkyDk11hX" colab_type="code" colab={}
| mscoco_feature_extraction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Losses
"""
# pylint: disable=C0301,C0103,R0902,R0915,W0221,W0622
##
# LIBRARIES
import torch
##
def l1_loss(input, target):
""" L1 Loss without reduce flag.
Args:
input (FloatTensor): Input tensor
target (FloatTensor): Output tensor
Returns:
[FloatTensor]: L1 distance between input and output
"""
return torch.mean(torch.abs(input - target))
##
def l2_loss(input, target, size_average=True):
""" L2 Loss without reduce flag.
Args:
input (FloatTensor): Input tensor
target (FloatTensor): Output tensor
Returns:
[FloatTensor]: L2 distance between input and output
"""
if size_average:
return torch.mean(torch.pow((input-target), 2))
else:
return torch.pow((input-target), 2)
| loss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (AnomalyCableDetection)
# language: python
# name: anomalycabledetection
# ---
# # Cross-correlation
# +
from pathlib import Path
import os
os.chdir(Path(os.getcwd()).parent)
# -
from AnomalyCableDetection.load import Loader, Preprocessor
from AnomalyCableDetection.stl import CableSTL, CrossCorrelation, AdjacencyType, STLType
from AnomalyCableDetection.plot import *
# +
from os.path import join
from pathlib import Path
import matplotlib.dates as mdates
import matplotlib.pylab as plt
import pandas as pd
import numpy as np
import glob
import os
import re
# -
# ### Load STL results
cc = CrossCorrelation('raw_period_3600')
stl_dict = cc.stl_dict
date_list = cc.date_list
cable_list = cc.cable_list
# ### Get cross-correlation
# +
cc_dict = dict()
for cable in cable_list:
cc_df = cc.get_temporal_cross_correlation(cable, STLType.RESIDUAL)
cc_dict.update({cable: cc_df})
# -
| example/STL/4. get cross-correlation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlp]
# language: python
# name: conda-env-nlp-py
# ---
# # Example of loading the classifier model and generating images
from numpy import expand_dims
from keras.models import load_model
from keras.datasets.mnist import load_data
# load the model
model = load_model('c_model_4200.h5')
# load the dataset
(trainX, trainy), (testX, testy) = load_data()
# expand to 3d, e.g. add channels
trainX = expand_dims(trainX, axis=-1)
testX = expand_dims(testX, axis=-1)
# convert from ints to floats
trainX = trainX.astype('float32')
testX = testX.astype('float32')
# scale from [0,255] to [-1,1]
trainX = (trainX - 127.5) / 127.5
testX = (testX - 127.5) / 127.5
# evaluate the model
_, train_acc = model.evaluate(trainX, trainy, verbose=0)
print('Train Accuracy: %.3f%%' % (train_acc * 100))
_, test_acc = model.evaluate(testX, testy, verbose=0)
print('Test Accuracy: %.3f%%' % (test_acc * 100))
| use_model.ipynb |
# + slideshow={"slide_type": "skip"}
############## PLEASE RUN THIS CELL FIRST! ###################
# import everything and define a test runner function
from importlib import reload
from helper import run
import psbt
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 1
#
# #### Create a PSBT from the p2wpkh transaction you've been sent
#
# +
# Exercise 1
from io import BytesIO
from psbt import PSBT
from tx import Tx
hex_tx = '01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060000000000ffffffff01583e0f000000000016001427459b7e4317d1c9e1d0f8320d557c6bb08731ef00000000'
# convert the hex transaction to a tx object
tx_obj = Tx.parse(BytesIO(bytes.fromhex(hex_tx)))
# use the create method to create a PSBT object
psbt_obj = PSBT.create(tx_obj)
# serialize, turn to hex and print it to see what it looks like
print(psbt_obj.serialize().hex())
# + slideshow={"slide_type": "slide"}
# example for updating the PSBT
from helper import read_varstr
from io import BytesIO
from psbt import PSBT, PSBTIn, PSBTOut, NamedHDPublicKey
from tx import Tx
hex_named_hd = '4f01043587cf034d513c1580000000fb406c9fec09b6957a3449d2102318717b0c0d230b657d0ebc6698abd52145eb02eaf3397fea02c5dac747888a9e535eaf3c7e7cb9d5f2da77ddbdd943592a14af10fbfef36f2c0000800100008000000080'
stream = BytesIO(bytes.fromhex(hex_named_hd))
key = read_varstr(stream)
named_hd = NamedHDPublicKey.parse(key, stream)
hex_psbt = '70736274ff0100770100000001192f88eeabc44ac213604adbb5b699678815d24b718b5940f5b1b1853f0887480100000000ffffffff0220a10700000000001976a91426d5d464d148454c76f7095fdf03afc8bc8d82c388ac2c9f0700000000001976a9144df14c8c8873451290c53e95ebd1ee8fe488f0ed88ac0000000000000000'
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
tx_lookup = psbt_obj.tx_obj.get_input_tx_lookup()
pubkey_lookup = named_hd.bip44_lookup()
psbt_obj.update(tx_lookup, pubkey_lookup)
print(psbt_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 2
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_update_p2wpkh`
# + slideshow={"slide_type": "slide"}
# Exercise 2
reload(psbt)
run(psbt.PSBTTest('test_update_p2wpkh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 3
#
# #### Update the PSBT that you got.
#
#
# +
# Exercise 3
from helper import read_varstr
from hd import HDPrivateKey
from io import BytesIO
from psbt import PSBT, PSBTIn, PSBTOut, NamedHDPublicKey
from tx import Tx
mnemonic = 'abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon about'
passphrase = b'<EMAIL> <PASSWORD>'
path = "m/44'/1'/0'"
hd_priv = HDPrivateKey.from_mnemonic(mnemonic, passphrase, testnet=True)
hex_psbt = '70736274ff01005201000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060000000000ffffffff01583e0f000000000016001427459b7e4317d1c9e1d0f8320d557c6bb08731ef00000000000000'
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
# create the NamedHDPublicKey using the HDPrivateKey and path
named_hd = NamedHDPublicKey.from_hd_priv(hd_priv, path)
# get the tx lookup using the psbt_obj's tx_object's get_input_tx_lookup method
tx_lookup = psbt_obj.tx_obj.get_input_tx_lookup()
# get the pubkey lookup using the bip44_lookup method
pubkey_lookup = named_hd.bip44_lookup()
# update the psbt object with the transaction lookup and the pubkey lookup
psbt_obj.update(tx_lookup, pubkey_lookup)
# print the serialized hex to see what it looks like
print(psbt_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 4
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_sign_p2wpkh`
# + slideshow={"slide_type": "slide"}
# Exercise 4
reload(psbt)
run(psbt.PSBTTest('test_sign_p2wpkh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 5
#
# #### Sign the PSBT that you got.
#
#
# +
# Exercise 5
from helper import read_varstr
from hd import HDPrivateKey
from io import BytesIO
from psbt import PSBT, PSBTIn, PSBTOut, NamedHDPublicKey
from tx import Tx
mnemonic = 'abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon about'
passphrase = b'jim<EMAIL> <PASSWORD>'
path = "m/44'/1'/0'"
hd_priv = HDPrivateKey.from_mnemonic(mnemonic, passphrase, testnet=True)
hex_psbt = '70736274ff01005201000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060000000000ffffffff01583e0f000000000016001427459b7e4317d1c9e1d0f8320d557c6bb08731ef000000000001011f40420f0000000000160014f0cd79383f13584bdeca184cecd16135b8a79fc222060247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f218797dcdac2c00008001000080000000800000000000000000002202026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d18797dcdac2c0000800100008000000080010000000000000000'
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
# use the HDPrivateKey to sign the PSBT
psbt_obj.sign(hd_priv)
# print the serialized hex to see what it looks like
print(psbt_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 6
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_finalize_p2wpkh`
# + slideshow={"slide_type": "slide"}
# Exercise 6
reload(psbt)
run(psbt.PSBTTest('test_finalize_p2wpkh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 7
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_final_tx_p2wpkh`
# + slideshow={"slide_type": "slide"}
# Exercise 7
reload(psbt)
run(psbt.PSBTTest('test_final_tx_p2wpkh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 8
#
# #### Finalize, Extract and Broadcast the PSBT that you got.
#
#
# +
# Exercise 8
from psbt import PSBT
hex_psbt = '70736274ff01005201000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060000000000ffffffff01583e0f000000000016001427459b7e4317d1c9e1d0f8320d557c6bb08731ef000000000001011f40420f0000000000160014f0cd79383f13584bdeca184cecd16135b8a79fc222020247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f24730440220575870ef714252a26bc4e61a6ee31db0f3896606a4792d11a42ef7d30c9f1b33022007cd28fb8618b704cbcf1cc6292d9be901bf3c99d967b0cace7307532619811e0122060247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f218797dcdac2c00008001000080000000800000000000000000002202026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d18797dcdac2c0000800100008000000080010000000000000000'
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
# finalize the PSBT
psbt_obj.finalize()
# extract the transaction using final_tx
tx_obj = psbt_obj.final_tx()
# breadcast the transaction
print(tx_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 9
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:NamedHDPublicKeyTest:test_redeem_script_lookup`
# + slideshow={"slide_type": "slide"}
# Exercise 9
reload(psbt)
run(psbt.NamedHDPublicKeyTest('test_redeem_script_lookup'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 10
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_p2sh_p2wpkh`
# + slideshow={"slide_type": "slide"}
# Exercise 10
reload(psbt)
run(psbt.PSBTTest('test_p2sh_p2wpkh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 11
#
# #### You have been sent an empty p2sh-p2wpkh transaction. Update, sign, finalize, extract and broadcast the signed transaction.
#
#
# +
# Exercise 11
from helper import read_varstr
from hd import HDPrivateKey
from io import BytesIO
from psbt import PSBT, PSBTIn, PSBTOut, NamedHDPublicKey
from tx import Tx
mnemonic = 'abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon about'
passphrase = b'j<PASSWORD> <EMAIL>.<PASSWORD> <PASSWORD>'
path = "m/44'/1'/0'"
hd_priv = HDPrivateKey.from_mnemonic(mnemonic, passphrase, testnet=True)
hex_tx = '01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060100000000ffffffff01583e0f00000000001600146e13971913b9aa89659a9f53d327baa8826f2d7500000000'
# convert the hex transaction to a tx object
tx_obj = Tx.parse(BytesIO(bytes.fromhex(hex_tx)))
# use the create method to create a PSBT object
psbt_obj = PSBT.create(tx_obj)
psbt_obj.tx_obj.testnet = True
# create the NamedHDPublicKey using the HDPrivateKey and path
named_hd = NamedHDPublicKey.from_hd_priv(hd_priv, path)
# get the tx lookup using the psbt_obj's tx_object's get_input_tx_lookup method
tx_lookup = psbt_obj.tx_obj.get_input_tx_lookup()
# get the pubkey lookup using the bip44_lookup method
pubkey_lookup = named_hd.bip44_lookup()
# get the RedeemScript lookup using the redeem_script_lookup method
redeem_script_lookup = named_hd.redeem_script_lookup()
# update the psbt object with the transaction lookup and the pubkey lookup
psbt_obj.update(tx_lookup, pubkey_lookup, redeem_script_lookup)
# use the HDPrivateKey to sign the PSBT
psbt_obj.sign(hd_priv)
# finalize the PSBT
psbt_obj.finalize()
# extract the transaction using final_tx
tx_obj = psbt_obj.final_tx()
# breadcast the transaction
print(tx_obj.serialize().hex())
# + slideshow={"slide_type": "slide"}
# example of updating
from helper import serialize_binary_path, encode_varstr
from io import BytesIO
from psbt import PSBT
from script import WitnessScript
hex_psbt = '70736274ff01005e01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060200000000ffffffff01583e0f0000000000220020878ce58b26789632a24ec6b62542e5d4e844dee56a7ddce7db41618049c3928c000000004f01043587cf034d513c1580000000fb406c9fec09b6957a3449d2102318717b0c0d230b657d0ebc6698abd52145eb02eaf3397fea02c5dac747888a9e535eaf3c7e7cb9d5f2da77ddbdd943592a14af10fbfef36f2c0000800100008000000080000000'
hex_witness_scripts = ['47522102c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c210247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f252ae', '47522102db8b701c3210e1bf6f2a8a9a657acad18be1e8bff3f7435d48f973de8408f29021026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d52ae']
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
tx_lookup = psbt_obj.tx_obj.get_input_tx_lookup()
key_1 = bytes.fromhex('<KEY>')
key_2 = bytes.fromhex('<KEY>')
path = "m/44'/1'/0'"
stream_1 = BytesIO(encode_varstr(bytes.fromhex('fbfef36f') + serialize_binary_path(path)))
stream_2 = BytesIO(encode_varstr(bytes.fromhex('797dcdac') + serialize_binary_path(path)))
hd_1 = NamedHDPublicKey.parse(key_1, stream_1)
hd_2 = NamedHDPublicKey.parse(key_2, stream_2)
pubkey_lookup = {**hd_1.bip44_lookup(), **hd_2.bip44_lookup()}
witness_lookup = {}
for hex_witness_script in hex_witness_scripts:
witness_script = WitnessScript.parse(BytesIO(bytes.fromhex(hex_witness_script)))
witness_lookup[witness_script.sha256()] = witness_script
psbt_obj.update(tx_lookup, pubkey_lookup, witness_lookup=witness_lookup)
print(psbt_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 12
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_update_p2wsh`
# + slideshow={"slide_type": "slide"}
# Exercise 12
reload(psbt)
run(psbt.PSBTTest('test_update_p2wsh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 13
#
# #### Update the transaction that's been given to you
#
#
# +
# Exercise 13
from helper import serialize_binary_path, encode_varstr
from io import BytesIO
from psbt import NamedHDPublicKey, PSBT
from script import WitnessScript
hex_psbt = '70736274ff01005e01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060200000000ffffffff01583e0f0000000000220020878ce58b26789632a24ec6b62542e5d4e844dee56a7ddce7db41618049c3928c000000004f01043587cf034d513c1580000000fb406c9fec09b6957a3449d2102318717b0c0d230b657d0ebc6698abd52145eb02eaf3397fea02c5dac747888a9e535eaf3c7e7cb9d5f2da77ddbdd943592a14af10fbfef36f2c0000800100008000000080000000'
hex_witness_scripts = ['47522102c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c210247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f252ae', '<KEY>']
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
mnemonic = 'abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon about'
passphrase = b'<EMAIL> <PASSWORD>'
path = "m/44'/1'/0'"
hd_priv = HDPrivateKey.from_mnemonic(mnemonic, passphrase, testnet=True)
# get the tx lookup using get_input_tx_lookup
tx_lookup = psbt_obj.tx_obj.get_input_tx_lookup()
hd_1 = list(psbt_obj.hd_pubs.values())[0]
hd_2 = NamedHDPublicKey.from_hd_priv(hd_priv, path)
pubkey_lookup = {**hd_1.bip44_lookup(), **hd_2.bip44_lookup()}
witness_lookup = {}
for hex_witness_script in hex_witness_scripts:
witness_script = WitnessScript.parse(BytesIO(bytes.fromhex(hex_witness_script)))
witness_lookup[witness_script.sha256()] = witness_script
psbt_obj.update(tx_lookup, pubkey_lookup, witness_lookup=witness_lookup)
print(psbt_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 14
#
# #### Sign the transaction with your HD private key
#
#
# +
# Exercise 14
from helper import serialize_binary_path, encode_varstr
from io import BytesIO
from psbt import NamedHDPublicKey, PSBT
hex_psbt = '70736274ff01005e01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060200000000ffffffff01583e0f0000000000220020878ce58b26789632a24ec6b62542e5d4e844dee56a7ddce7db41618049c3928c000000004f01043587cf034d513c1580000000fb406c9fec09b6957a3449d2102318717b0c0d230b657d0ebc6698abd52145eb02eaf3397fea02c5dac747888a9e535eaf3c7e7cb9d5f2da77ddbdd943592a14af10fbfef36f2c00008001000080000000800001012b40420f0000000000220020c1b4fff485af1ac26714340af2e13d2e89ad70389332a0756d91a123c7fe7f5d010547522102c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c210247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f252ae22060247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f218797dcdac2c00008001000080000000800000000000000000220602c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c18fbfef36f2c0000800100008000000080000000000000000000010147522102db8b701c3210e1bf6f2a8a9a657acad18be1e8bff3f7435d48f973de8408f29021026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d52ae2202026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d18797dcdac2c00008001000080000000800100000000000000220202db8b701c3210e1bf6f2a8a9a657acad18be1e8bff3f7435d48f973de8408f29018fbfef36f2c0000800100008000000080010000000000000000'
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
mnemonic = 'abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon about'
passphrase = b'<EMAIL> <PASSWORD>'
path = "m/44'/1'/0'"
# get the private key using the mnemonic, passphrase and testnet=True
hd_priv = HDPrivateKey.from_mnemonic(mnemonic, passphrase, testnet=True)
# sign the psbt
print(psbt_obj.sign(hd_priv))
# print the serialized hex of the PSBT to see what it looks like
print(psbt_obj.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 15
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_finalize_p2wsh`
# + slideshow={"slide_type": "slide"}
# Exercise 15
reload(psbt)
run(psbt.PSBTTest('test_finalize_p2wsh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 16
#
# #### Finalize, extract and broadcast the PSBT
#
#
# +
# Exercise 16
from io import BytesIO
from psbt import NamedHDPublicKey, PSBT
hex_psbt = '70736274ff01005e01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060200000000ffffffff01583e0f0000000000220020878ce58b26789632a24ec6b62542e5d4e844dee56a7ddce7db41618049c3928c000000004f01043587cf034d513c1580000000fb406c9fec09b6957a3449d2102318717b0c0d230b657d0ebc6698abd52145eb02eaf3397fea02c5dac747888a9e535eaf3c7e7cb9d5f2da77ddbdd943592a14af10fbfef36f2c00008001000080000000800001012b40420f0000000000220020c1b4fff485af1ac26714340af2e13d2e89ad70389332a0756d91a123c7fe7f5d220202c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c47304402203f26a975aae04a7ae12c964cdcea318c850351a3072aebbab7902e89957008ea022019f895271f70d1515f9da776d6ac17c21bcbca769d87c1beb4ebbf4c7a56fbc20122020247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f247304402204fd654c27002d4c9e53bb001229e3d7587e5be245a81b6f7ead3bf136643af40022060ebf1193a6b3e82615a564f0043e5ae88e661bfdb7fd254c9a30bae8160583901010547522102c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c210247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f252ae22060247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f218797dcdac2c00008001000080000000800000000000000000220602c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c18fbfef36f2c0000800100008000000080000000000000000000010147522102db8b701c3210e1bf6f2a8a9a657acad18be1e8bff3f7435d48f973de8408f29021026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d52ae2202026421c7673552fdad57193e102df96134be00649195b213fec9d07c6d918f418d18797dcdac2c00008001000080000000800100000000000000220202db8b701c3210e1bf6f2a8a9a657acad18be1e8bff3f7435d48f973de8408f29018fbfef36f2c0000800100008000000080010000000000000000'
psbt_obj = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt)))
psbt_obj.tx_obj.testnet = True
# finalize
psbt_obj.finalize()
# get the final Tx
final_tx = psbt_obj.final_tx()
# print the tx serialized hex to see what it looks like
print(final_tx.serialize().hex())
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 17
#
#
#
#
# #### Make [this test](/edit/session5/psbt.py) pass: `psbt.py:PSBTTest:test_p2sh_p2wsh`
# + slideshow={"slide_type": "slide"}
# Exercise 17
reload(psbt)
run(psbt.PSBTTest('test_p2sh_p2wsh'))
# + [markdown] slideshow={"slide_type": "slide"}
# ### Exercise 18
#
# #### Combine, update, sign, finalize, extract and finalize the PSBTs sent to you
#
#
# +
# Exercise 18
from hd import HDPrivateKey
from io import BytesIO
from psbt import NamedHDPublicKey, PSBT
hex_psbt_1 = '70736274ff01005e01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060300000000ffffffff01583e0f0000000000220020878ce58b26789632a24ec6b62542e5d4e844dee56a7ddce7db41618049c3928c000000000001012040420f000000000017a91423358e259fbcf478331138ceb9619d9a8c835073872202031b31547c895b5e301206740ea9890a0d6d127baeebb7fffb07356527323c915b483045022100e7dc3213aff7676be5bc087fe1698b1b04e53555f93bb11478bd22c0b6a6ffe502205c86c17bcb4d9bf7bd7ae82f18af5f7f387f72c82af27ecd4b9f6b68f2f2821b0101042200207fcc2ca7381db4bdfd02e1f2b5eb3d72435b8e09bdbd8bfe3d748bf19d78ef38010569532102c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c21031b31547c895b5e301206740ea9890a0d6d127baeebb7fffb07356527323c915b210247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f253ae220602c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c18fbfef36f2c000080010000800000008000000000000000002206031b31547c895b5e301206740ea9890a0d6d127baeebb7fffb07356527323c915b18fbfef36f2c000080010000800000008000000000010000000000'
hex_psbt_2 = '70736274ff01005e01000000015c89191dc2abf62339e0f114cb4c3bf8fb399d522d112c9afa2dc7a43759f9060300000000ffffffff01583e0f0000000000220020878ce58b26789632a24ec6b62542e5d4e844dee56a7ddce7db41618049c3928c000000000001012040420f000000000017a91423358e259fbcf478331138ceb9619d9a8c83507387220202c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c4830450221008c892608dcbbc5b40fc40a82bb4bdeeb79f4a81b4a8d26d0915d2ba2c3d84a28022076d5507bf6ad60893e9baaf7690823d5c85a8720abab7bb48a64449c1b5c9ff50101042200207fcc2ca7381db4bdfd02e1f2b5eb3d72435b8e09bdbd8bfe3d748bf19d78ef38010569532102c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c21031b31547c895b5e301206740ea9890a0d6d127baeebb7fffb07356527323c915b210247aed77c3def4b8ce74a8db08d7f5fd315f8d96b6cd801729a910c3045d750f253ae220602c1b6ac6e6a625fee295dc2d580f80aae08b7e76eca54ae88a854e956095af77c18fbfef36f2c000080010000800000008000000000000000002206031b31547c895b5e301206740ea9890a0d6d127baeebb7fffb07356527323c915b18fbfef36f2c000080010000800000008000000000010000000000'
psbt_obj_1 = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt_1)))
psbt_obj_1.tx_obj.testnet = True
psbt_obj_2 = PSBT.parse(BytesIO(bytes.fromhex(hex_psbt_2)))
psbt_obj_2.tx_obj.testnet = True
mnemonic = 'abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon abandon about'
passphrase = b'<EMAIL> <PASSWORD>'
path = "m/44'/1'/0'"
# get the private key using the mnemonic, passphrase and testnet=True
hd_priv = HDPrivateKey.from_mnemonic(mnemonic, passphrase, testnet=True)
# create the NamedHDPublicKey using the HDPrivateKey and path
named_hd = NamedHDPublicKey.from_hd_priv(hd_priv, path)
# combine the two objects
psbt_obj_1.combine(psbt_obj_2)
# grab the pubkey lookup using bip44_lookup
pubkey_lookup = named_hd.bip44_lookup()
# update the PSBT
psbt_obj_1.update({}, pubkey_lookup)
# sign the psbt
print(psbt_obj_1.sign(hd_priv))
# finalize the transaction
psbt_obj_1.finalize()
# get the final Tx
final_tx = psbt_obj_1.final_tx()
# print the tx serialized hex to see what it looks like
print(final_tx.serialize().hex())
| session5/complete/session5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Uf6dEaWpP3lo"
# 作業目標:<br>
# 1. 運用索引找出需要資料<br>
# 2. 合併資料的分法應用
# + [markdown] id="3rP6NOVVQZuB"
# 作業重點:<br>
# 1. 分辨索引loc、iloc差別
# 2. 分辨合併資料方法的不同,因應不同情況使用Merge, join, concatenate
# + [markdown] id="02Ld_89FP-4N"
# 題目:<br>
# 讀取STOCK_DAY_0050_202009.csv串聯STOCK_DAY_0050_202010.csv,並且找出open小於close的資料
#
# + id="ysjY4kR5P4rA"
import pandas as pd
print( 'Pandas 版本: ', pd.__version__ )
# + id="hsEDbEogG4DG"
#讀取STOCK_DAY_0050_202009.csv、STOCK_DAY_0050_202010.csv
Data_09 = pd.read_csv('STOCK_DAY_0050_202009.csv')
Data_10 = pd.read_csv('STOCK_DAY_0050_202010.csv')
# -
Data_09
Data_10
# + id="cw3OMgQgG4DG"
#串聯
# 方法一 :
AllData_1 = Data_09.append( Data_10, sort=False, ignore_index=True )
AllData_1
# -
# 方法二 :
AllData_2 = pd.concat( [Data_09,Data_10] ).reset_index( drop=True )
AllData_2
# + executionInfo={"elapsed": 957, "status": "ok", "timestamp": 1610348842287, "user": {"displayName": "\u732e\u7ae4\u9ec3", "photoUrl": "", "userId": "07529243043474362942"}, "user_tz": -480} id="LBBLV7TaG4DH"
#找出open小於close的資料
SelectedData_1 = AllData_1[ AllData_1.open < AllData_1.close ]
print( type(SelectedData_1) )
SelectedData_1
# + id="tZaFA68VG4DH"
SelectedData_2 = AllData_2.loc[ AllData_2.open < AllData_2.close ]
print( type(SelectedData_2) )
SelectedData_2
| Homework/Day_10_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.7.0
# language: julia
# name: julia-0.7
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Solving Double Integrator with SVP
#
# This notebook shows how to use the proposed structured value-based planning (SVP) approach to generate the state-action $Q$-value function for the classic double integrator problem. The correctness of the solution is verified by trajectory simulations.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Problem Definition
#
# The problem is defined as a unit mass brick moving along the $x$-axis on a frictionless surface, with a control input which provides a horizontal force.
# The brick starts from the position-velocity pair $\left(x, \dot{x}\right)$ and follows the dynamics
#
# \begin{align*}
# x &:= x + \dot{x}\cdot\tau, \\
# \dot{x} &:= \dot{x} + u\cdot\tau,
# \end{align*}
#
# where $u \in \left[ -1, 1 \right]$ is the acceleration input, $\tau$ is the time interval between decisions. The brick can take on the state values $\left(x, \dot{x}\right) \in \left[ -3., 3. \right] \times \left[ -3., 3. \right]$. To incentivize getting to the original point of the $x$-axis with minimum control input, the reward function is defined in a quadratic form as
#
# \begin{equation*}
# r(x,\dot{x}) = - \frac{1}{2} \left( x^2 + \dot{x}^2 \right).
# \end{equation*}
# + [markdown] slideshow={"slide_type": "slide"}
# ## Structured Value-based Planning (SVP)
#
# The proposed structured value-based planning (SVP) approach is based on the $Q$-value iteration. At the $t$-th iteration, instead of a full pass over all state-action pairs:
# - SVP first randomly selects a subset $\Omega$ of the state-action pairs. In particular, each state-action pair in $\mathcal{S}\times\mathcal{A}$ is observed (i.e., included in $\Omega$) independently with probability $p$.
# - For each selected $(s,a)$, the intermediate $\hat{Q}(s,a)$ is computed based on the $Q$-value iteration:
# \begin{equation*}\hat{Q}(s,a) \leftarrow \sum_{s'} P(s'|s,a) \left( r(s,a) + \gamma \max_{a'} Q^{(t)}(s',a') \right),\quad\forall\:(s,a)\in\Omega.
# \end{equation*}
# - The current iteration then ends by reconstructing the full $Q$ matrix with matrix estimation, from the set of observations in $\Omega$. That is, $Q^{(t+1)}=\textrm{ME}\big(\{\hat{Q}(s,a)\}_{(s,a)\in\Omega}\big).$
#
# Overall, each iteration reduces the computation cost by roughly $1-p$.
# + [markdown] slideshow={"slide_type": "subslide"}
# Through SVP, we can compute the final state-action $Q$-value function.
# To obtain the optimal policy for state $s$, we compute
#
# \begin{align*}
# \pi^{\star} \left(s\right) = \mbox{argmax}_{a \in \mathcal{A}} Q^{\star}\left(s, a\right).
# \end{align*}
# + [markdown] slideshow={"slide_type": "slide"}
# ### Generate state-action value function with SVP
# + slideshow={"slide_type": "fragment"}
push!(LOAD_PATH, ".")
using MDPs, DoubleIntegrator
mdp = MDP(state_space(), action_space(), transition, reward)
__init__()
policy = value_iteration(mdp, true, "../data/qdi_otf_0.2.csv", true)
print("") # suppress output
# + [markdown] slideshow={"slide_type": "slide"}
# ### Visualize policy as a heat map
# + slideshow={"slide_type": "fragment"}
viz_policy(mdp, policy, "SVP policy (20%)", true, "di/policy_di_0.2.tex")
# + [markdown] slideshow={"slide_type": "slide"}
# ### Verify correctness
# Simulate and visualize trajectory from initial state `[position, speed]`.
# + slideshow={"slide_type": "fragment"}
ss, as = simulate(mdp, policy, [-0.5, 0.0])
viz_trajectory(ss, as, "SVP trajectory (20%)", "SVP input (20%)", true, "di/traj_di_0.2.tex")
# -
nsim = 1000
times = 0
for sim = 1:nsim
state = [rand(XMIN:0.001 * (XMAX - XMIN):XMAX),
rand(VMIN:0.001 * (VMAX - VMIN):VMAX)]
_, as = simulate(mdp, policy, copy(state))
times += length(as)
end # for sim
times /= nsim
@printf("average times: %.3f", times)
| svp/DoubleIntegrator.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Goolgle Play Store Analysis Report
# ### <NAME> [18030142032]
# <b>Google Play Store:</b> Google Play (previously Android Market) is a digital distribution service operated and developed by Google LLC. It serves as the official app store for the Android operating system, allowing users to browse and download applications developed with the Android software development kit (SDK) and published through Google. Google Play also serves as a digital media store, offering music, books, movies, and television programs.<br>
# Applications are available through Google Play either free of charge or at a cost. They can be downloaded directly on an Android device through the Play Store mobile app or by deploying the application to a device from the Google Play website. The Google Play store had over 82 billion app downloads in 2016 and has reached over 3.5 million apps published in 2017.
# <br><br>
# <b><u>Table of Contents</u></b>
# - Dependencies
# - DataSet Description
# - Data Exploration
# - Data Wrangling- Preprocessing, Cleaning & Filtering
# - Visualization and Observation
# ## <br><br><br><br> Import Dependencies
# +
# Data Analysis
import numpy as np
print('Numpy:',np.__version__)
import pandas as pd
print('Pandas:',pd.__version__)
# Data Visualization
import matplotlib
import seaborn as sns
print('Seaborn:', sns.__version__)
import matplotlib.pyplot as plt
print('Matplotlib:', matplotlib.__version__)
# %matplotlib inline
from datetime import datetime,date
# -
# ## <br><br><br><br>DataSet Description
# The dataset has been scraped from Google Play Store on 20th December, 2018 with the help of <a href="https://pypi.org/project/play-scraper/">Play Scraper</a> library available in PyPI. It tooks approx. 4-5 hours to scrape data with the help of this library. Code to scrape data is given below:
#
# <strong>
# <code style="background-color: #eee; border: 1px solid #999; display: block;padding: 20px;">
# import csv
# import play_scraper as ps
# app_category = list(ps.categories()) # Total App Categories: 58
# collections = list(ps.lists.COLLECTIONS) # Total Collections: 6
# with open('play_app.csv','a+') as csvfile:
# writer = csv.writer(csvfile)
# for coll in collections:
# for cat in app_category:
# try:
# c = (ps.collection(collection=coll,category=cat))
# for app in range(0,len(c)):
# writer.writerow(ps.details(app_id = c[app].get('app_id')).values())
# except Exception:
# pass
# </code>
# </strong>
#
# <br><br><br><br>
# There are 31 <b>Headers</b> for the given file :<br>
#
# + active=""
# Name Description Name Description
# ---- ----------- ---- -----------
# title app name icon app icon url
# screenshots app image url video app detail video url
# category app category score app overall rating
# histogram app ratings reviews app comments in number
# description app details description_html app details in HTML format
# recent_changes app update details editors_choice highest quality app
# price app price details free app availability type
# iap in-app purchase developer_id app developer id
# updated app last update date size app size in Mb
# installs app downloads current_version app version
# url app official URL content_rating app content categories
# iap_range in-app purchase range interactive_elements app major interacts
# developer app developer name developer_email developer email
# developer_url app developer website developer_address developer official address
# app_id app unique id
# required_android_version android version details for the app
# -
# ### Data Loading from CSV file.
app_df = pd.read_csv('./play_app.csv',parse_dates=['updated'])
print("DataSet Size on RAM:",app_df.memory_usage(index=True,deep=True).sum(),'bytes')
# ## <br><br><br><br> Data Exploration
## DataSet Description
app_df.info()
# Data Cleaning and Filtering
print('Percentage of NULL in each column')
app_df.isnull().sum()/app_df.shape[0] * 100
pd.set_option('display.max_columns', 31) # Set option to disploy hidden columns
app_df.head(5)
# ## <br><br><br><br> Data Wrangling: Preprocessing, Cleaning and Filtering
# #### [Size]
# +
# Size - Converting App Size to Numeric Values in MB
app_df['size'] = app_df['size'].str.replace('k','e+0') # k represents KB
app_df['size'] = app_df['size'].str.replace('M','e+3') # M represents MB
app_df['size'] = app_df['size'].str.replace(',','') # M represents MB
# Function to check whether the given value can be coverted to float or not
def isConvertable(val):
try:
float(val)
return True
except ValueError:
return False
app_df['size'] = app_df['size'].replace('Varies with device','0') # Replacing the String values to NaN
app_df['size'].apply(lambda x: isConvertable(x)) # converting the convertable values.
app_df['size'] = pd.to_numeric(app_df['size']) # converting to numeric values
# -
# #### [installs]
# Installs - Coverting to Numeric data type
app_df['installs'] = app_df['installs'].str.replace('[+,]','') # removing the special symbols
app_df['installs'] = pd.to_numeric(app_df['installs'])
# #### [price]
# Price - Coverting to Numeric data type after removing the $ symbol
app_df['price'] = app_df['price'].str.replace('$','')
app_df['price'] = pd.to_numeric(app_df['price'])
# #### [score]
# Score - Coverting to Numeric data type after replacing the NaN values
app_df['score'] = app_df['score'].fillna(0) # Filling NULL Values with Zero( cannot update it with mean value )
# app_df['score'].apply(lambda x: isConvertable(x)) # converting the convertable values.
app_df['score'] = pd.to_numeric(app_df['score'])
# #### [categoties]
# New column has been created on the basis of its contents. The categories column contain more than one value in it.
# +
app_df['category'] = app_df['category'].str.replace('[','')
app_df['category'] = app_df['category'].str.replace(']','')
app_df['category'] = app_df['category'].str.replace('[\']','')
def app_categories(word):
if word.startswith('GAME'):
return word.split('_')
else:
return word.split(',')
app_df['flit_cat'] = list(map(lambda x:app_categories(x)[0], app_df.category)) #
# -
# #### [content_ratings]
# New contents has been created on the basis of its contents. This column contains more than one value.
# +
app_df['content_rating'] = app_df['content_rating'].str.replace('[','')
app_df['content_rating'] = app_df['content_rating'].str.replace(']','')
app_df['content_rating'] = app_df['content_rating'].str.replace('[\']','')
app_df['flit_content'] = list(map(lambda x:x[0], app_df.content_rating.str.split(',')))
# -
# #### [required_android_version]
# +
app_df['required_android_version'] = app_df['required_android_version'].str.replace(" and up",'')
app_df['required_android_version'] = app_df['required_android_version'].str.replace("Varies with device",'4.0')
app_df['required_android_version'] = app_df['required_android_version'].replace("4.4W",'4.4')
app_df['required_android_version'] = app_df['required_android_version'].replace("2.3 - 2.3.2",'2.3')
app_df['required_android_version'] = app_df['required_android_version'].replace("4.0.3",'4.0')
app_df['required_android_version'] = app_df['required_android_version'].replace("2.0.1",'2.0')
app_df['required_android_version'] = app_df['required_android_version'].replace("2.3.3",'2.3')
app_df['required_android_version'] = pd.to_numeric(app_df['required_android_version'])
# -
# #### Dropping unnecessary columns
app_df = app_df.drop(['icon','screenshots','video','histogram','description_html','developer_address',
'developer_email'],axis=1)
app_df.info()
pd.set_option('display.max_columns', 31) # Set option to disploy hidden columns
app_df.head(5)
# ## <br><br><br><br> Data Visualization and Observation
# +
grp_content = app_df.groupby(['flit_content']) # categorizing data by Content Rating
cont_cat_labels = tuple(k for k, kdf in grp_content) # list of content type
explode = [0.09 for i in range(len(cont_cat_labels))] # explode for pie chart
colors = ['red','lightcoral','yellowgreen', 'gold', 'skyblue','black']
sizes = [len(kdf) for k, kdf in grp_content] # size of each grouped data of Content Rating
cont_cat_labels_percent = list(np.round((np.array(sizes) * 100/sum(np.array(sizes))),1)) # percentange for each size of Content Rating
# Concatinating the percentage with list of content type
x = [x+" :- " for x in list(cont_cat_labels)]
y = [str(x)+ " %" for x in cont_cat_labels_percent]
labels = [x[i]+y[i] for i in range(len(cont_cat_labels))]
# plotting the pie chart
plt.figure(dpi=100)
plt.xlabel('Apps by Content Rating',fontsize=18)
patches, texts = plt.pie(sizes, labels = cont_cat_labels, explode = explode, colors = colors,
shadow=True,startangle = 90 ) # , autopct='%1.2f%%'
plt.legend(patches, labels, loc="lower right")
plt.axis(xmax = 2.6)
plt.tight_layout()
plt.show()
# -
# The apps are divided into six categories on the basis of their content ratings.It shows that 78% of the apps are available for all age groups, 13.4% apps are specially for teens, 4.9% apps are for age group above 10 but not teen, 3.6% apps are for age group above 17. There are no Adult Content Apps available on Play Store and non of the apps are left unrated.
# <br><br><br><br>
# generating pie chart for free and paid apps datasets
plt.figure(dpi=120) # setting figure dots per inch
plt.title('Free & Paid Apps Ratio Available in Play Store') # title of the pie chart
plt.xlabel('Free vs. Paid Apps',fontsize=10)
plt.pie([(app_df['free']==True).sum(), (app_df['free']==False).sum()], labels = ('Free','Paid'),
explode = (0.05, 0.05), colors = ['lightskyblue','lightcoral'],
shadow=True,startangle = 360, autopct='%1.2f%%') # plotting the pie chart
plt.show() # displaying the figure
# The ratio of free and paid apps are divided in 1:4. 1/4 of the total apps in the Play Stores are paid and remaining are free.<br><br><br><br>
# +
grp_category = app_df.groupby('flit_cat')
cat_lst = np.array([k for k, kdf in grp_category]) # list the content category-wise
clnum = np.array([len(grp_category.get_group(l)) for l in cat_lst]) # no. of downloads for each category-list
# Plotting data Categories Wise
plt.figure(figsize=(18, 10), dpi=150)
plt.style.use('seaborn-white')
plt.title('Category-wise Application Downloads',fontsize = 38)
plt.xlabel('Category of Applications',color='darkblue', fontsize = 22)
plt.ylabel('Number of Downloads (Category-Wise )', color='darkblue', fontsize = 22)
plt.xticks(rotation=90, fontsize = 14)
plt.plot(cat_lst, clnum)
plt.grid()
plt.show()
# -
# It seems that GAMES are downloaded the most and other categories apps are downloaded on an average. EDUCATION and PERSONALIZATION apps are downloaded above average.
# The least downloaded apps are BEAUTY and EVENTS apps.<br><br><br><br>
app_df['revenue'] = app_df['installs'] * app_df['price']
store_data_business = app_df.sort_values("revenue", ascending=False)
plt.figure(figsize=(8, 4), dpi=100)
plt.title('Top 10 Revenue Generating Apps',fontsize = 16)
plt.bar(store_data_business['title'].unique()[0:10],store_data_business['revenue'].unique()[0:10],
color=['lightcoral','yellowgreen', 'gold', 'skyblue','black','cyan'])
plt.xticks(rotation=90, fontsize = 8)
plt.xlabel('Name of Applications',color='darkblue', fontsize = 15)
plt.ylabel('Total Revenue', color='darkblue', fontsize = 15)
plt.show()
# Out of the Paid apps these top 10 apps generated most of the revenue for the developers.
# Minecraft is on the number one.<br><br><br><br>
# #### Top 10 Apps which hasn't updated till date by Developers
app_df['last_updated'] = app_df['updated'].apply(lambda x:date.today()-datetime.date(x))
pd.DataFrame(app_df.sort_values(['last_updated'],ascending=False,inplace=False)[['title','flit_cat','last_updated']][0:10])
# Plotting Last Update Categories Wise
cat_app_upd = dict(app_df.groupby(['flit_cat','last_updated']).sum().index.get_values())
plt.figure(figsize=(12, 8), dpi=100)
plt.style.use('seaborn-white')
plt.title('Category-Wise Apps Last Update',fontsize = 20)
plt.ylabel('Apps Categories',color='darkblue', fontsize = 12)
plt.xlabel('Number of Years', color='darkblue', fontsize = 12)
plt.barh(list(cat_app_upd.keys()),[x.days/365 for x in list(cat_app_upd.values())])
plt.grid()
plt.show()
# The GAMES, PERSONALIZATION and FINANCE Apps are least updated, whereas DATING apps are most frequently updated.<br><br><br><br>
# #### <br><br><br> Top 10 on the basis of Price, Download size and Developers(no. of apps)
top10_apps = {'Cost-Wise top 10 Apps':list(app_df.sort_values(['price'],ascending=False,inplace=False)['title'].unique()[0:10]),
'Download Size-Wise top 10 Apps':list(app_df.sort_values(['size'],ascending=False,inplace=False)['title'].unique()[0:10]),
'Apps-Wise Top 10 Developers':list(app_df.groupby('developer').count().sort_values(['developer_url'], ascending = False, inplace = False).index[0:10])}
pd.DataFrame(top10_apps, index=[x for x in range(1,11)])
# ### <br><br><br> In-App Purchase:- Free vs Paid Apps
# +
## Excluding Category:- GAME, because it has highest In-App purchase
plt.figure(figsize=(16,12))
# Free Apps
plt.subplot(2,2,1)
iap_df = app_df[app_df['iap'] == True]
iap_df = iap_df[iap_df['free'] == True]
iap_df = iap_df[iap_df['flit_cat'] != "GAME"]
iap_df.groupby(['flit_cat'])['free'].count().plot(kind="bar")
plt.title('Free Apps with In-App Purchase')
plt.xlabel('App Categories')
plt.subplot(2,2,2)
iap_df = app_df[app_df['iap'] == False]
iap_df = iap_df[iap_df['free'] == True]
iap_df = iap_df[iap_df['flit_cat'] != "GAME"]
iap_df.groupby(['flit_cat'])['free'].count().plot(kind="bar")
plt.title('Free Apps with No In-App Purchase')
plt.xlabel('App Categories')
# Paid Apps
plt.subplot(2,2,3)
iap_df = app_df[app_df['iap'] == True]
iap_df = iap_df[iap_df['free'] == False]
iap_df = iap_df[iap_df['flit_cat'] != "GAME"]
iap_df.groupby(['flit_cat'])['free'].count().plot(kind="bar")
plt.title('Paid Apps with In-App Purchase')
plt.xlabel('App Categories')
plt.subplot(2,2,4)
iap_df = app_df[app_df['iap'] == False]
iap_df = iap_df[iap_df['free'] == False]
iap_df = iap_df[iap_df['flit_cat'] != "GAME"]
iap_df.groupby(['flit_cat'])['free'].count().plot(kind="bar")
plt.title('Paid Apps with No In-App Purchase')
plt.xlabel('App Categories')
plt.tight_layout()
plt.show()
# -
# Excluding Games, the Free Apps has more in-app purchases than Paid Apps. Very few of Paid Apps are having in-app purchase. The ratio of No in-app purchase in Free Apps are comaratively higher with respect to in-app purchases in Free Apps.
# ### Minimum App Version Requirement Distribution
plt.figure(figsize=(14,6))
sns.countplot(app_df['required_android_version'])
plt.show()
# The minimum android version requirement for most of the app are v4.1 and v4.0 and up. Yet some of the apps can run on very old android version which is a contradiction for android upgrade.
| Assignments/Python/Google Play Store Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "notes"}
# # Introduction: Digital Controller Design
#
# In this section we will discuss converting continuous time models into discrete time (or difference equation) models. We will also introduce the z-transform and show how to use it to analyze and design controllers for discrete time systems.
#
# ## Introduction
#
# The figure below shows the typical continuous feedback system that we have been considering so far in this tutorial. Almost all of the continuous controllers can be built using analog electronics.
# 
# The continuous controller, enclosed in the dashed square, can be replaced by a digital controller, shown below, that performs same control task as the continuous controller. The basic difference between these controllers is that the digital system operates on discrete signals (or samples of the sensed signal) rather than on continuous signals.
# 
# Different types of signals in the above digital schematic can be represented by the following plots.
# 
# The purpose of this Digital Control Tutorial is to show you how to use Python to work with discrete functions either in transfer function or state-space form to design digital control systems.
# + [markdown] slideshow={"slide_type": "notes"}
# ### Zero-Hold Equivalence
#
# In the above schematic of the digital control system, we see that the digital control system contains both discrete and the continuous portions. When designing a digital control system, we need to find the discrete equivalent of the continuous portion so that we only need to deal with discrete functions. For this technique, we will consider the following portion of the digital control system and rearrange as follows.
# 
#
# 
# The **clock** connected to the *D/A and A/D converters* supplies a pulse every T seconds and each D/A and A/D sends a signal only when the pulse arrives. The purpose of having this pulse is to require that Hzoh(z) have only samples u(k) to work on and produce only samples of output y(k); thus, Hzoh(z) can be realized as a discrete function. The philosophy of the design is the following. We want to find a discrete function Hzoh(z) so that for a piecewise constant input to the continuous system H(s), the sampled output of the continuous system equals the discrete output. Suppose the signal u(k) represents a sample of the input signal. There are techniques for taking this sample u(k) and holding it to produce a continuous signal uhat(t). The sketch below shows that the uhat(t) is held constant at u(k) over the interval kT to (k+1)T. This operation of holding uhat(t) constant over the sampling time is called zero-order hold.
# 
# The zero-order held signal uhat(t) goes through H2(s) and A/D to produce the output y(k) that will be the piecewise same signal as if the discrete signal u(k) goes through Hzoh(z) to produce the discrete output y(k).
# 
# Now we will redraw the schematic, placing Hzoh(z) in place of the continuous portion.
# 
# By placing Hzoh(z), we can design digital control systems dealing with only discrete functions. *Note:* There are certain cases where the discrete response does not match the continuous response due to a hold circuit implemented in digital control systems. For information, see Lagging effect associated with the hold.
# + [markdown] slideshow={"slide_type": "notes"}
# ### Conversion Using c2d
#
# There is a control toolbox function called `control.c2d` that converts a given continuous system (either in transfer function or state-space form) to a discrete system using the zero-order hold operation explained above.
#
# The sampling time (Ts in sec/sample) should be smaller than 1/(30*BW), where BW is the closed-loop bandwidth frequency.
# + [markdown] slideshow={"slide_type": "notes"}
# ## Example: Mass-Spring-Damper
#
# ### Transfer Function
#
# Suppose you have the following continuous transfer function
#
# $$
# \frac{X(s)}{F(s)} = \frac{1}{Ms^2+bs+k}
# $$
#
# Assuming the closed-loop bandwidth frequency is greater than 1 rad/sec, we will choose the sampling time (Ts) equal to 1/100 sec. Now, create an new m-file and enter the following commands.
# +
import control
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from IPython.display import Latex, Math, display
s = control.TransferFunction.s
# -
# %matplotlib inline
# Generate Larger more readable plots
# Double click on images to make them full scale.
sns.set(
rc={
"axes.labelsize": 8,
"axes.titlesize": 8,
"figure.figsize": (4 * 1.618, 4),
"figure.dpi": 200,
}
)
# + slideshow={"slide_type": "notes"}
M = 1
b = 10
k = 20
sys = 1 / (M * s ** 2 + b * s + k)
display(Math("$$\\frac{X(s)}{F(s)}=" + sys._repr_latex_().lstrip("$")))
# + slideshow={"slide_type": "notes"}
Ts = 1 / 100
sys_d = control.c2d(sys, Ts, "zoh")
display(Math("$$ \\frac{X(z)}{F(z)} = " + sys_d._repr_latex_().lstrip("$")))
# + [markdown] slideshow={"slide_type": "notes"}
# ### State-Space
#
# The continuous time state-space model is as follows:
#
# $$
# \mathbf{\dot{x}} = \left[ \begin{array}{c} \dot{x} \\ \ddot{x} \end{array} \right] = \left[ \begin{array}{cc} 0 & 1 \\ -\frac{k}{m} & -\frac{b}{m} \end{array} \right] \left[ \begin{array}{c} x \\ \dot{x} \end{array} \right] + \left[ \begin{array}{c} 0 \\ \frac{1}{m} \end{array} \right] F(t)
# $$
#
#
#
# $$
# y = \left[ \begin{array}{cc} 1 & 0 \end{array} \right] \left[ \begin{array}{c} x \\ \dot{x} \end{array} \right]
# $$
#
# All constants are the same as before. The following m-file converts the above continuous state-space to discrete state-space.
# + slideshow={"slide_type": "notes"}
A = np.array([[0, 1], [-k / M, -b / M]])
B = np.array([[0], [1 / M]])
C = np.array([[1, 0]])
D = np.array([[0]])
Ts = 1 / 100
sys = control.ss(A, B, C, D)
sys
# + slideshow={"slide_type": "notes"}
sys_d = control.c2d(sys, Ts, "zoh")
sys_d
# + [markdown] slideshow={"slide_type": "notes"}
# From these matrices, the discrete state-space can be written as
#
# $$
# \left[ \begin{array}{c} x(k) \\ v(k) \end{array} \right] = \left[ \begin{array}{cc} 0.9990 & 0.0095 \\ -0.1903 & 0.9039 \end{array} \right] \left[ \begin{array}{c} x(k-1) \\ v(k-1) \end{array} \right] + \left[ \begin{array}{c} 0 \\ 0.0095 \end{array} \right] F(k-1)
# $$
#
#
#
# $$
# y(k-1) = \left[ \begin{array}{cc} 1 & 0 \end{array} \right] \left[ \begin{array}{c} x(k-1) \\ v(k-1) \end{array} \right]
# $$
#
# Now you have the discrete time state-space model.
# + [markdown] slideshow={"slide_type": "notes"}
# ### Stability and Transient Response
#
# For continuous systems, we know that certain behaviors results from different pole locations in the s-plane. For instance, a system is unstable when any pole is located to the right of the imaginary axis. For discrete systems, we can analyze the system behaviors from different pole locations in the z-plane. The characteristics in the z-plane can be related to those in the s-plane by the expression
#
# $$
# z = e^{sT}
# $$
#
#
# * T = Sampling time (sec/sample)
# * s = Location in the s-plane
# * z = Location in the z-plane The figure below shows the mapping of lines of constant damping ratio (zeta) and natural frequency (Wn) from the s-plane to the z-plane using the expression shown above.
# 
# If you noticed in the z-plane, the stability boundary is no longer imaginary axis, but is the unit circle |z|=1. The system is stable when all poles are located inside the unit circle and unstable when any pole is located outside. For analyzing the transient response from pole locations in the z-plane, the following three equations used in continuous system designs are still applicable.
#
# $$
# \zeta \omega_n \geq \frac{4.6}{Ts}
# $$
#
#
#
# $$
# \omega_n \geq \frac{1.8}{Tr}
# $$
#
#
#
# $$
# \zeta = \frac{-\ln(\%OS/100)}{\sqrt{\pi^2+\ln(\%OS/100)^2}}
# $$
#
# where,
#
# - $\zeta$ = Damping ratio
# - $\omega_n$ = natural frequency (rad/sec)
# - $T_s$ = 1% settling time
# - $T_r$ = 10-90% rise time
# - $M_p$ = maximum overshoot
#
#
# **Important**: The natural frequency (Wn) in z-plane has the unit of rad/sample, but when you use the equations shown above, the Wn must be in the unit of rad/sec. Suppose we have the following discrete transfer function
#
# $$
# \frac{Y(z)}{F(z)} = \frac{1}{z^2-0.3z+0.5}
# $$
#
# Create an new m-file and enter the following commands. Running this m-file in the command window gives you the following plot with the lines of constant damping ratio and natural frequency.
# + slideshow={"slide_type": "notes"}
numDz = [1]
denDz = [1, -0.3, 0.5]
sys = control.TransferFunction(numDz, denDz, -1)
sys
# + slideshow={"slide_type": "notes"}
poles, zeros = control.pzmap(sys)
plt.gca().axis([-1, 1, -1, 1])
# + [markdown] slideshow={"slide_type": "notes"}
# From this plot, we see poles are located approximately at the natural frequency of $\frac{9\pi}{20T}$ (rad/sample) and the damping ratio of 0.25. Assuming that we have a sampling time of 1/20 sec (which leads to Wn = 28.2 rad/sec) and using three equations shown above, we can determine that this system should have the rise time of 0.06 sec, a settling time of 0.65 sec and a maximum overshoot of 45% (0.45 more than the steady-state value). Let's obtain the step response and see if these are correct. Add the following commands to the above cell and rerun it in the notebook. You should get the following step response.
# + slideshow={"slide_type": "notes"}
sys = control.TransferFunction(numDz, denDz, 1 / 20)
sys
# + slideshow={"slide_type": "notes"}
T, yout = control.step_response(sys=sys)
plt.plot(T, yout)
plt.xlabel("Time (s)")
plt.title("Step Response");
# + [markdown] slideshow={"slide_type": "notes"}
# As you can see from the plot, the rise time, settling time and overshoot came out to be what we expected. This shows how you can use the locations of poles and the above three equations to analyze the transient response of the system.
#
# ## Discrete Root Locus
#
# The root-locus is the locus of points where roots of characteristic equation can be found as a single gain is varied from zero to infinity. The characteristic equation of an unity feedback system is
#
# $$
# 1+KG(z)Hzoh(z) = 0
# $$
#
# where G(z) is the compensator implemented in the digital controller and Hzoh(z) is the plant transfer function in z. The mechanics of drawing the root-loci are exactly the same in the z-plane as in the s-plane. Recall from the continuous Root-Locus Tutorial, we used the Python function called sgrid to find the root-locus region that gives an acceptable gain (K). For the discrete root-locus analysis, we will use the function zgrid that has the same characteristics as sgrid. The command zgrid(zeta, Wn) draws lines of constant damping ratio (zeta) and natural frequency (Wn). Suppose we have the following discrete transfer function
#
# $$
# \frac{Y(z)}{F(z)} = \frac{z-0.3}{z^2-1.6z+0.7}
# $$
#
# and the requirements are a damping ratio greater than 0.6 and a natural frequency greater than 0.4 rad/sample (these can be found from design requirements, sampling time (sec/sample) and three equations shown in the previous section). The following commands draw the root-locus with the lines of constant damping ratio and natural frequency. Create an new m-file and enter the following commands. Running this m-file should give you the following root-locus plot.
# + slideshow={"slide_type": "notes"}
numDz = [1, -0.3]
denDz = [1, -1.6, 0.7]
sys = control.TransferFunction(numDz, denDz, -1)
rlist, klist = control.rlocus(sys)
# -
# From this plot, we can see that the system is stable for some values of $K$ since there are portions of the root locus where both branches are located inside the unit circle. Also, we can observe two dotted lines representing the constant damping ratio and natural frequency. The natural frequency is greater than 0.3 outside the constant-Wn line, and the damping ratio is greater than 0.4 inside the constant-zeta line. In this example, portions of the generated root-locus are within in the desired region. Therefore, a gain ($K$) chosen to place the two closed-loop poles on the loci within the desired region should provide us a response that satisfies the given design requirements.
| Introduction/Introduction_ControlDigital.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Waveguide array
#
# To define a waveguide array we need to define:
#
# - wg_gaps
# - widths
# - the material functions or refractive indices of box, waveguide and clad
# - thickness of each material
# - x and y_steps for discretizing the structure
# - sidewall angle
# - wavelength that can be used in case the refractive index are a function of the
# wavelength
#
# Where all units are in um
#
# ```
# __________________________________________________________
#
# clad_thickness
# widths[0] wg_gaps[0] widths[1]
# ___________ ___________ _ _ _ _ _ _
# | | | |
# _____| |____________| |____ |
# wg_heigth
# slab_thickness |
# ________________________________________________ _ _ _ _ _
#
# sub_thickness
# __________________________________________________________
#
# ```
#
import modes as ms
wg = ms.waveguide_array(widths=[0.5]*2, wg_gaps=0.2)
wg
# +
# ms.waveguide_array?
# -
wg_slab = ms.waveguide_array(widths=[0.5]*2, wg_gaps=0.2, slab_thickness=0.09)
wg_slab
s = ms.mode_solver_full(wg=wg_slab, plot=True, fields_to_write=('Ex'))
| docs/notebooks/03_waveguide_array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Practice Exercise: Importing data & Exploring data (manipulation)
# ## Context:
# - The data is about NBA (National Basketball Association) games from 2004 season to Dec, 2020.
# - We'll be focusing on practicing importing data and data manipulation techniques learned in the course. But the dataset is also popular to be used for predicting NBA games winners.
# - We've made minor changes on the data to fit this exercise, such as changing the column names. Check out the original source if you are interested in using this data for other purposes (https://www.kaggle.com/nathanlauga/nba-games)
# ## Dataset Description:
#
# We'll work on two datasets (in two separate csv files):
#
# - **games**: each game from 2004 season to Dec 2020, including information about the two teams in each game, and some details like number of points, etc
# - **teams**: information about each team played in the games
#
# Assume we want to study the game level data, but with detailed information about each team. We'll need to combine these two datasets together.
# ## Objective:
# - Load/examine/subset/rename/change dtypes of columns for each individual dataset
# - Combine them into a single dataset, and export it
# - Explore the final dataset by subsetting or sorting
# ### 1. Import the libraries
# ### 2. Load the data in `games.csv` as a DataFrame called `games`
#
# Save the csv file under the same directory as the notebook if not typing the full path.
# ### 3. Look at the first 5 rows of the DataFrame
# ### 4. Look at the columns of the DataFrame
# ### 5. Reassign `games` as its subset of the columns 'GAME_DATE', 'GAME_STATUS_TEXT', 'TEAM_ID_home', 'TEAM_ID_away', 'POINTS_home', 'POINTS_away', 'HOME_TEAM_WINS'
#
# We'll only keep some columns about the games
# ### 6. Look at the new `games` DataFrame's first 5 rows, and info summary
# ### 7. Convert `GAME_DATE` to a `datetime` dtype
# ### 8. Convert `GAME_STATUS_TEXT` to a `string` dtype
# ### 9. Look at the info summary of the DataFrame to verify the changes
# ### 10. Load the data in `teams.csv` as a DataFrame called `teams`, and look at its first 5 rows, and its columns
# ### 11. Reassign `teams` as a subset of its columns 'TEAM_ID', 'CITY', 'NICKNAME', and look at its first 5 rows and info summary
#
# We'll only keep some columns about the teams
# ### 12. Convert both columns `CITY` and `NICKNAME` to a `string` dtype
# ### 13. Verify the changes with the `dtypes` attribute
# ### 14. Print out the first two rows of `games` and `teams`, how can we combine them?
# *Hint:*
#
# *Within the `games` DataFrame, there are two columns `TEAM_ID_home` and `TEAM_ID_away`. This is because each game involves two teams playing against each other. The team that played in its own location, is called the 'home' team, the team that played outside its location, is called the 'away' team. Each game has one 'home' team and one 'away' team.*
#
# *While the `teams` DataFrame stores the information about each team, the identifier for each team is the column `TEAM_ID`.*
# *We can merge the two DataFrames based on:*
#
# - *`TEAM_ID_home` in `games` and `TEAM_ID` in `teams`: to get the team information for the 'home' team*
# - *`TEAM_ID_away` in `games` and `TEAM_ID` in `teams`: to get the team information for the 'away' team*
# ### 15. Merge (inner) `games` and `teams` based on 'TEAM_ID_home' and 'TEAM_ID', call the merged DataFrame `games_with_home_team`
# ### 16. Print out the first 5 rows of the new DataFrame
#
# Since we used the column `TEAM_ID_home` when merging, the two columns `CITY` and `NICKNAME` are storing the city and nickname of the 'home' team.
# So let's rename them.
# ### 17. Rename the column `CITY` as 'city_home', `NICKNAME` as 'nickname_home'
# ### 18. Merge (inner) `games_with_home_team` and `teams` based on 'TEAM_ID_away' and 'TEAM_ID', call the merged DataFrame `games_with_both_teams`
# ### 19. Print out the first two rows of the new DataFrame
#
# Since we used the column `TEAM_ID_away` when merging, the two columns `CITY` and `NICKNAME` are storing the city and nickname of the 'away' team.
# So let's rename them.
# ### 20. Rename the column `CITY` as 'city_away', `NICKNAME` as 'nickname_away'
# ### 21. Print out the first 5 rows of the new DataFrame
# You probably have noticed that there are two columns called `TEAM_ID_x` and `TEAM_ID_y`. This is because we merged the DataFrame twice with the `teams` DataFrame. So the `TEAM_ID` column from `teams` is added twice to the merged DataFrame.
#
# To avoide duplicate column names, `pandas` added the suffixes of '_x' and '_y' to distinguish them. But due to inner joins, `TEAM_ID_x` is the same as `TEAM_ID_home`, and `TEAM_ID_y` is the same as `TEAM_ID_away`.
# ### The team ID columns are not needed after the merge of the DataFrames.
#
# ### The below code is provided to drop the columns 'TEAM_ID_home', 'TEAM_ID_away', 'TEAM_ID_x', 'TEAM_ID_y' from `games_with_both_teams`. We'll learn about this `drop` method in a later section
games_with_both_teams = games_with_both_teams.drop(columns=['TEAM_ID_home', 'TEAM_ID_away', 'TEAM_ID_x', 'TEAM_ID_y'])
games_with_both_teams.head()
# ### 22. Make a copy of `games_with_both_teams` and assign it as `games`
# ### 23. Change the column names in `games` to all lowercase
# ### 24. Print out the columns of `games` to verify the changes
# ### 25. Print out the columns of `games_with_both_teams` to verify that the original DataFrame wasn't impacted by the copy
# ### 26. Check the dimensionality of `games`
# ### 27. Export `games` as a csv file called 'games_transformed.csv', and open the csv file to look at it
#
# Feel free to test the difference of the csv files with or without the argument `index=False`
# ### 28. Select all the columns of 'number' dtypes from `games`
# ### 29. Select all the columns NOT of 'number' dtypes from `games`
# ### 30. Print out the first 5 rows of `games` as a reference
# ### 31. Select the row with label 0
# ### 32. Select the row with integer position 0
# ### 33. Set the column `game_date` as the index of DataFrame `games`
# ### 34. Print out the index of `games` to verify the changes
# ### 35. Select the rows with label '2020-12-18'
# ### 36. Select the rows with labels from '2020-12-18' to '2020-12-19'
# ### 37. Select the rows with labels of '2020-12-18' and '2019-12-18'
# ### 38. Select the rows with `points_home` greater than 150
# ### 39. Select the rows with `points_home` greater than 150, and `home_team_wins` not being 1
# ### 40. Select the rows with `points_home` greater than 150, and `home_team_wins` not being 1, as well as the columns `home_team_wins` and `points_home`
# ### 41. Reset the index of `games` back to default and verify the changes
# ### 42. Add a new column called `points_total`, as the sum of columns `points_home` and `points_away`
# ### 43. Verify the changes by printing out the three columns `points_home`, `points_away`, `points_total`
# ### 44. Print out the 3 rows with the largest `points_total` using the `nlargest` method
# ### 45. Sort the DataFrame `games` by its `points_total` column in ascending order
#
# Don't forget to reassign the sorted result back to `games`
# ### 46. Print out the last 3 rows of the sorted DataFrame
#
# Verify that it's the same three rows as the previous example (`nlargest`)
# ### 47. Given that the DataFrame is sorted by `points_total`, select the row with the smallest `points_total` using `iloc`
# ### 48. Select the rows with the second, and third smallest `points_total` using `iloc`
# ### 49. Select a subset including the first 4 rows, and the first 5 columns using `iloc`
| Question_practice_exercise+(importing_manipulation).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Lab-02 Linear regression
# 학습목표
#
# 선형회귀(Linear Regression)에 대해 알아본다.
# 핵심키워드
#
# * 선형회귀(Linear Regression)
# * 평균 제곱 오차(Mean Squared Error)
# * 경사하강법(Gradient descent)
import torch
# ## Data Definition
x_train = torch.FloatTensor([[1],[2],[3]]) # 공부 시간
y_train = torch.FloatTensor([[2],[4],[6]]) # 점수
# Weight와 bias를 0으로 초기화: 항상 처음에는 모델이 0을 출력
W = torch.zeros(1, requires_grad = True) # requires_grad = True <- 학습할 것이라고 명시
b = torch.zeros(1, requires_grad = True)
hypothesis = x_train * W + b
# ## compute loss할 때의 loss함수 정의
loss = torch.mean((hypothesis - y_train)**2)
# ## Computed Loss를 활용해서 모델 개선시키기
from torch import optim
# +
# torch.optim 라이브러리 사용
optimizer = optim.SGD([W, b], lr = 0.01) # W,b는 학습할 텐서들.
# optimizer을 통해 학습시킬 때 항상 붙어다니는 3줄
optimizer.zero_grad() # zero_grad()로 gradient 초기화
loss.backward() # backward()로 gradient 계산
optimizer.step() # step()으로 개선
# -
# ## Full Training Code
import torch
from torch import optim
# 한번만!
# Data 정의
x_train = torch.FloatTensor([[1],[2],[3]]) # 공부 시간
y_train = torch.FloatTensor([[2],[4],[6]]) # 점수
# 초기 파라미터 정의(모델 초기화)
W = torch.zeros(1, requires_grad = True) # requires_grad = True <- 학습할 것이라고 명시
b = torch.zeros(1, requires_grad = True)
# Optimizer(파라미터 개선 규칙) 정의
optimizer = optim.SGD([W, b], lr = 0.01)
# 반복!
epochs_nb = 1000
for epoch in range(1, epochs_nb+1):
# loss 구하기: 예측 -> Loss(cost) 계산
hypothesis = x_train * W + b
loss = torch.mean((hypothesis - y_train)**2) # 텐서들이므로 x = [1,2,3], y = [1,2,3] 통째로 연산
# 구해진 loss값과 optimizer을 토대로 파라미터 개선하기
optimizer.zero_grad()
loss.backward()
optimizer.step()
# W, b가 각각 최적의 수로 수렴함!
| Lab-02 Linear regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests, pandas as pd, numpy as np
from requests import session
from bs4 import BeautifulSoup
manual=pd.read_excel('manual_manual.xlsx').set_index('Cégnév')
data=pd.read_excel('data.xlsx').set_index('Cégnév')
sectors=pd.read_excel('valid_manual.xlsx')
len(data), len(manual)
data=pd.concat([data,manual])
data['Sector']=sectors.set_index(0)[1]
# Bring in 2014 data
manual4=pd.read_excel('../2018/manual_manual.xlsx').set_index('Cégnév')
data4=pd.read_excel('../2018/data.xlsx').set_index('Cégnév')
sectors4=pd.read_excel('../2018/valid_manual.xlsx')
len(data4), len(manual4)
data4=pd.concat([data4,manual4])
data4['Sector4']=sectors4.set_index(0)[1]
data=data.join(data4[['Alkalmazottak száma 2014','Árbevétel 2014 (RON)']])
repl={'udvarhelyszek':'Udvarhelyszék',
'csikszek':'Csíkszék',
'gyergyoszek':'Gyergyószék',
'marosszek':'Marosszék',
'also-haromszek':'Alsó-háromszék',
'felso-haromszek':'Felső-háromszék'}
for r in repl:
data=data.replace(r,repl[r])
data.to_excel('export1.xlsx')
d1=data[['Régió', 'Latitude', 'Longitude', 'Sector']]
d1.columns=['Régió', 'Hosszúság', 'Szélesség', 'Iparág']
d1.to_excel('d1.xlsx')
data.columns
ds=[]
ds1=[]
ds2=[]
for i in range(2014,2019):
g='Alkalmazottak száma'
c=g+' '+str(i)
print(c)
df=data[[c]]
df.columns=[g]
df[g]=df[g].fillna('').str.replace(u'\xa0','').str.replace('-','')
df['Év']=i
ds1.append(df)
ds.append(d1.join(df))
g='Árbevétel'
c=g+' '+str(i)+' (RON)'
print(c)
df=data[[c]]
df.columns=[g]
df[g]=df[g].fillna('').str.replace(u'\xa0','').str.replace('-','')
df['Év']=i
ds2.append(df)
df=pd.concat(ds,sort=False)
df1=pd.concat(ds1,sort=False)
df2=pd.concat(ds2,sort=False)
df1.to_excel('df1.xlsx')
df2.to_excel('df2.xlsx')
df1=df1.set_index('Év',append=True)
df2=df2.set_index('Év',append=True)
df=df1.join(df2).replace('',np.nan).astype(float)
df['Bevétel']=np.round(df['Árbevétel']/1000000.0,3)
df['Bevétel/Alkalmazott']=np.round(df['Bevétel']/df['Alkalmazottak száma'],3)
data2=df.reset_index().join(d1,on='Cégnév').set_index('Cégnév')
# Normalize names
#[' '.join([j.title() if len(j)>3 else j for j in i.split(' ')]) for i in data2.index.str.replace(' SA','').str.replace(' SRL','').str.replace(' ROMANIA ','')]
data2.index=data2.index.str.replace(' SA','').str.replace(' SRL','').str.replace(' ROMANIA','')
data2.to_excel('export2.xlsx')
# Calculate percentages
totals=data2.groupby(['Régió','Év']).sum()[['Alkalmazottak száma','Bevétel']]
totals.columns=['TAlkalmazottak száma','TBevétel']
totals=totals.unstack()
totals=(np.round(totals/totals.sum()*100,0)).stack().astype(str)
totals['TAlkalmazottak száma']=totals['TAlkalmazottak száma'].str[:-2]+'%'
totals['TBevétel']=totals['TBevétel'].str[:-2]+'%'
totals2=data2.groupby(['Cégnév','Év']).sum()[['Alkalmazottak száma','Bevétel']]
totals2.columns=['TAlkalmazottak száma','TBevétel']
totals2=totals2.unstack()
totals2=(np.round(totals2/totals2.sum()*100,1)).stack().astype(str)
totals2['TAlkalmazottak száma']=totals2['TAlkalmazottak száma'].str[:]+'%'
totals2['TBevétel']=totals2['TBevétel'].str[:]+'%'
totals2.columns=['T2Alkalmazottak száma','T2Bevétel']
totals2.loc['AZOMURES']
totals3=data2.groupby(['Cégnév','Év','Régió']).sum()[['Alkalmazottak száma','Bevétel']]
totals3.columns=['TAlkalmazottak száma','TBevétel']
totals3=totals3.unstack().unstack()
totals3=(np.round(totals3/totals3.sum()*100,1)).stack().stack().astype(str)
totals3['TAlkalmazottak száma']=totals3['TAlkalmazottak száma'].str[:]+'%'
totals3['TBevétel']=totals3['TBevétel'].str[:]+'%'
totals3.columns=['T3Alkalmazottak száma','T3Bevétel']
totals3.loc['AZOMURES']
data22=data2.reset_index().set_index(['Régió','Év']).join(totals).reset_index()
data22=data22.set_index(['Cégnév','Év']).join(totals2).reset_index()
data22=data22.set_index(['Cégnév','Év','Régió']).join(totals3).reset_index()
data221=data22[['Régió','Év','Bevétel','TBevétel','T2Bevétel','T3Bevétel','Cégnév']]
data221['Régió %']=data221['TBevétel']+' '+data221['Régió']
data221['Cégnév %']=data221['Cégnév']+' | '+data221['T2Bevétel']+' | megyében '+data221['T3Bevétel']
data221=data221.set_index(['Cégnév %','Régió %','Év'])['Bevétel'].unstack()
data221['Type']='Árbevétel (millió RON)'
data221=data221.reset_index()
data222=data22[['Régió','Év','Alkalmazottak száma','TAlkalmazottak száma','T2Alkalmazottak száma','T3Alkalmazottak száma','Cégnév']]
data222['Régió %']=data222['TAlkalmazottak száma']+' '+data222['Régió']
data222['Cégnév %']=data222['Cégnév']+' | ' +data222['T2Alkalmazottak száma']+' | megyében '+data222['T3Alkalmazottak száma']
data222=data222.set_index(['Cégnév %','Régió %','Év'])['Alkalmazottak száma'].unstack()
data222['Type']='Alkalmazottak száma'
data222=data222.reset_index()
data223=pd.concat([data221,data222])
data223.to_excel('export22.xlsx')
colors22={'Alsó-háromszék': '#855c75',
'Csíkszék': '#526a83',
'Felső-háromszék': '#625377',
'Gyergyószék': '#af6458',
'Marosszék': '#736f4c',
'Udvarhelyszék': '#d9af6b'}
for i in data223['Régió %'].unique():
print(i,':',colors22[i[i.find(' ')+1:]])
# Simplify
data3=df.unstack()
data3.columns=data3.columns.get_level_values(1).astype(str)
data3=data3.reset_index().join(d1,on='Cégnév').set_index('Cégnév')
data3.index.name=None
data3.index=data3.index.str.replace(' SA','').str.replace(' SRL','').str.replace(' ROMANIA','')
data3.to_excel('export3.xlsx')
data3b=data3.copy()
data3b.columns=range(len(data3b.columns))
data3b=data3b[[14,20,21,22,23]]
data3b['Size']=data3b[14]**0.5+5
data3b.dropna(subset=[14]).to_excel('export3b.xlsx')
data4=data3.groupby(['Iparág','Régió']).sum()
data4b=data3.groupby(['Régió']).sum()
data4=data4.unstack().T
data4b=data4b.unstack().T
data4['Mind']=data4b
data4=data4.unstack().T
data4.reset_index().to_excel('export4.xlsx')
data4=data3.groupby(['Régió','Iparág']).sum()
data4b=data3.groupby(['Iparág']).sum()
data4=data4.unstack().T
data4b=data4b.unstack().T
data4['Mind']=data4b
data4=data4.unstack().T
data4.reset_index().to_excel('export4b.xlsx')
d1s=[]
for y in ['2014','2015','2016','2017','2018']:
d1=data4[[y]]
d1.columns=[0,1,'Bevétel',2]
d1=d1[['Bevétel']]
d1['Év']=y
d1s.append(d1)
d1s=pd.concat(d1s)
d1s.reset_index().to_excel('export5.xlsx')
d2s=d1s.reset_index().set_index(['Iparág','Év','Régió']).unstack()
d3s=d2s.reset_index().groupby(['Év']).sum()
d3s['Iparág']='Mind'
d3s=d3s.reset_index().set_index(['Iparág','Év'])
d6=pd.concat([d2s,d3s])['Bevétel']
d6.reset_index().to_excel('export6.xlsx')
d6T=d6.stack().reset_index().set_index(['Régió','Év','Iparág']).unstack().reset_index()
d6T[d6T['Régió']!='Mind'].to_excel('export6T.xlsx')
# +
regiok=[i for i in d6.columns if i!='Mind']
d6=d6.reset_index()
# -
for i in regiok:
d6[i]=np.round(d6[i]*100/d6['Mind'],0)
d6=d6.replace(0,np.nan).replace(1,np.nan).replace(2,np.nan)
d6.to_excel('export6b.xlsx')
| gazdasag/2019/3_formatter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="2k4ZB9yltJCa"
# Initialize a dictionary "emp_info" with below details
# In - emp_info['Tom']
# Out - {'email':'<EMAIL>', 'Phone': +1987654321, 'City': 'California'}
# In - emp_info['Kathy']
# Out - {'email':'<EMAIL>', 'Phone': +1887654321, 'City': 'New York'}
emp_info = {'Tom':{'email':'<EMAIL>', 'Phone': +1987654321, 'City': 'California'},
'Kathy':{'email':'<EMAIL>', 'Phone': +1887654321, 'City': 'New York'}}
print(emp_info['Tom'])
print(emp_info['Kathy'])
# + colab={} colab_type="code" id="BAUj-S0fqFWG"
# Create a dictionary out of below inputs
lst1 = ['emp1', 'emp2', 'emp3']
emp_key = ['e_name', 'e_id', 'e_sal']
emp1_val = ['John', 'SG101', '$10,000']
emp2_val = ['Smith', 'SG102', '$9,000']
emp3_val = ['Peter', 'SG103', '$9,500']
# d = {'emp1':{'e_name':'John', 'e_id':'SG101', 'e_sal':'$10,000'},
# 'emp2':{'e_name':'Smith', 'e_id':'SG102', 'e_sal':'$9,000'},
# 'emp3':{'e_name':'Peter', 'e_id':'SG103', 'e_sal':'$9,500'}}
d1 = {}
d2 = {}
d3 = {}
d = {}
for i,j in enumerate(emp_key):
d1[j] = emp1_val[i]
d2[j] = emp2_val[i]
d3[j] = emp3_val[i]
my_list = [d1,d2,d3]
for i,j in enumerate(lst1):
d[j] = my_list[i]
print(d)
# + colab={} colab_type="code" id="LxGZ3jbjrBkj"
# Acess the value of key 'history'
sampleDict = {
"class":{
"student":{
"name":"Mike",
"marks":{
"physics":70,
"history":80
}
}
}
}
print(sampleDict['class']['student']['marks']['history'])
# + colab={} colab_type="code" id="Vm8UZbyNrKID"
# Initialize dictionary with default values. Inputs are:-
employees = ['Kelly', 'Emma', 'John']
defaults = {"designation": 'Application Developer', "salary": 8000}
# d = {'Kelly': {'designation': 'Application Developer', 'salary': 8000},
# 'Emma': {'designation': 'Application Developer', 'salary': 8000},
# 'John': {'designation': 'Application Developer', 'salary': 8000}}
d = {}
for i in employees:
d[i] = {"designation": 'Application Developer', "salary": 8000}
print(d)
# + colab={} colab_type="code" id="nmFdbaHBrThC"
# In gene expression, mRNA is transcribed from a DNA template.
# The 4 nucleotide bases of A, T, C, G corresponds to the U, A, G, C bases of the mRNA.
# Write a function that returns the mRNA transcript given the sequence of a DNA strand.
# Use a dictionary to provide the mapping of DNA to RNA bases.
mRNA = list('UAGC')
dna = list('ATCG')
d = dict(zip(dna,mRNA))
print(d)
# + colab={} colab_type="code" id="rop1A1pQw6Uk"
# Write a function which takes a word as input and returns a dictionary with letters as key and no of time letters are repeated as value.
# In - count_letter('google.com')
# Out - {'g': 2, 'o': 3, 'l': 1, 'e': 1, '.': 1, 'c': 1, 'm': 1}
word = input('Enter the word: ')
d = {}
for i in word:
d[i] = word.count(i)
print(d)
# + colab={} colab_type="code" id="vAwGIoxdw856"
# A DNA strand consisting of the 4 nucleotide bases is usually represented with a string of letters: A,T, C, G.
# Write a function that computes the base composition of a given DNA sequence.
# In - baseComposition("CTATCGGCACCCTTTCAGCA")
# Out - {'A': 4, 'C': 8, 'T': 5, 'G': 3 }
# In - baseComposition("AGT")
# Out - {'A': 1, 'C': 0, 'T': 1, 'G': 1 }
seq = input('Enter the sequence: ')
dna = list('ATCG')
d = {key: 0 for key in dna}
for i in seq:
d[i] = seq.count(i)
print(d)
# + colab={} colab_type="code" id="iVNnSsvaxMDo"
# [MCQ] Suppose "d" is an empty dictionary, which statement does not assign "d" with {"Name":"Tom"}?
# 1. d = {"Name": "Tom" }
# 2. d["Name"] = "Tom"
# 3. d.update({"Name": "Tom" })
# 4. d.setdefault("Name", "Tom")
# 5. None of the above. This is the answer
# + colab={} colab_type="code" id="tOgsXi2axnhF"
# [MCQ] d = {"a":1, "b":2}. Which of the statements returns [1,2]?
# 1. d.keys()
# 2. d.values() This is the answer
# 3. d.items()
# 4. d.popitem()
# 5. None of the above.
# + colab={} colab_type="code" id="BeNNopkDzK0k"
# [MCQ] Which of the following declarations is not valid for 'dict' type?
# 1. d = {"Name": "Tom" }
# 2. d = { (1,3,4): 4.5 }
# 3. d = { ["First", "Last"]: (1,3) } This one
# 4. d = { 1: 0.4 }
# 5. None of the above
# + colab={} colab_type="code" id="7kPbKcfMzTHP"
# Write a function reverseLookup(dictionary, value) that takes in a dictionary
# and a value as arguments and returns a sorted list of all keys that contains the value.
# The function will return an empty list if no match is found.
# In - reverseLookup({'a':1, 'b':2, 'c':2}, 1)
# Out - ['a']
# In - reverseLookup({'a':1, 'b':2, 'c':2}, 2)
# Out - ['b', 'c']
# In - reverseLookup({'a':1, 'b':2, 'c':2}, 3)
# Out - []
d = eval(input("Enter the dict: "))
k = int(input('Enter the integer: '))
my_list = []
for i,j in d.items():
#print(i,j)
if j == k:
my_list.append(i)
print(my_list)
# + colab={} colab_type="code" id="s87Rvg2HDFYp"
# Write a function invertDictionary(d) that takes in a dictionary as argument and return a dictionary that inverts the keys and the values of the original dictionary.
# In - invertDictionary({'a':1, 'b':2, 'c':3, 'd':2})
# Out - {1: ['a'], 2: ['b', 'd'], 3: ['c']}
# In - invertDictionary({'a':3, 'b':3, 'c':3})
# Out - {3: ['a', 'c', 'b']}
# In - invertDictionary({'a':2, 'b':1, 'c':2, 'd':1})
# Out - {1: ['b', 'd'], 2: ['a', 'c']}
mydict =eval(input("Enter the dict: "))
new_dict = {}
for i in mydict:
if mydict[i] not in new_dict:
new_dict[mydict[i]] = list(i)
else:
new_dict[mydict[i]].append(i)
new_dict
# + colab={} colab_type="code" id="DgYyaZU6Oz7E"
# Write a function that converts a sparse vector into a dictionary as described above.
# In - convertVector([1, 0, 0, 2, 0, 0, 0, 3, 0, 0, 0, 0, 4])
# Out - {0: 1, 3: 2, 7: 3, 12: 4}
# In - convertVector([1, 0, 1 , 0, 2, 0, 1, 0, 0, 1, 0])
# Out - {0: 1, 2: 1, 4: 2, 6: 1, 9: 1}
# In - convertVector([0, 0, 0, 0, 0])
# Out - {}
my_list = eval(input('Enter the list: '))
my_dict = {}
for i in my_list:
if i != 0:
my_dict[my_list.index(i)] = i
my_dict
# + colab={} colab_type="code" id="ETCmEcflTGCI"
# Write a function that converts a dictionary back to its sparse vector representation.
# In - convertDictionary(c)
# Out - [1, 0, 0, 2, 0, 0, 0, 3, 0, 0, 0, 0, 4]
# In - convertDictionary({0: 1, 2: 1, 4: 2, 6: 1, 9: 1})
# Out - [1, 0, 1, 0, 2, 0, 1, 0, 0, 1]
# In - convertDictionary({})
# Out - []
my_dict = eval(input("Enter the dict: "))
maximum = max(list(my_dict.keys()))
my_list = [0]*(maximum+1)
for i in my_dict:
my_list[i] = my_dict[i]
my_list
# + colab={} colab_type="code" id="tTANm2QKZXh3"
# Given a Python dictionary, Change Brad’s salary to 8500
sampleDict = {
'emp1': {'name': 'Jhon', 'salary': 7500},
'emp2': {'name': 'Emma', 'salary': 8000},
'emp3': {'name': 'Brad', 'salary': 6500}
}
# Expected Output
# sampleDict = {
# 'emp1': {'name': 'Jhon', 'salary': 7500},
# 'emp2': {'name': 'Emma', 'salary': 8000},
# 'emp3': {'name': 'Brad', 'salary': 8500}
# }
sampleDict['emp3']['salary'] = 8500
sampleDict
# + colab={} colab_type="code" id="8dUHXiCVZmUx"
# Get the key corresponding to the minimum value from the following dictionary
sampleDict = {
'Physics': 82,
'Math': 65,
'history': 75
}
# Expected Output
# Math
minimum = min(list(sampleDict.values()))
for i in sampleDict:
if sampleDict[i] == minimum:
print(i)
# + colab={} colab_type="code" id="AQe5QNGdZ7aJ"
# Rename key city to location in the following dictionary
sampleDict = {
"name": "Kelly",
"age":25,
"salary": 8000,
"city": "New york"
}
# Expected Output
# {
# "name": "Kelly",
# "age":25,
# "salary": 8000,
# "location": "New york"
# }
x = sampleDict['city']
del sampleDict['city']
sampleDict['location'] = x
print(sampleDict)
# + colab={} colab_type="code" id="e19_ddO9aNNw"
# Check if a value 200 exists in a dictionary
sampleDict = {'a': 100, 'b': 200, 'c': 300}
# Expected Output: True
for i in sampleDict:
if sampleDict[i] == 200:
print(True)
# + colab={} colab_type="code" id="yphwjc1gabMa"
# Delete set of keys from Python Dictionary
sampleDict = {
"name": "Kelly",
"age":25,
"salary": 8000,
"city": "New york"
}
keysToRemove = ["name", "salary"]
# Expected Output:
# {'city': 'New york', 'age': 25}
for i in keysToRemove:
sampleDict.pop(i)
sampleDict
# -
| Pranjal/Dictionary_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false
# # 4. Create reports <a name="reports"></a>
#
# This section introduces different ways to save query results:<br>
#
# - [4.1 - Export results as a table (<i>alminer.save_table</i>)](#4.1-Export-results-as-a-table)<br>
# - [4.2 - Save overview plots for each target (<i>alminer.save_source_reports</i>)](#4.2-Save-overview-plots)<br>
#
#
# + [markdown] deletable=false editable=false
# <h3>Load libraries & create a query</h3>
#
# To explore these options, we will first query the archive using one of the methods presented in the previous section and use the results in the remainder of this tutorial.
# + deletable=false editable=false
import alminer
observations = alminer.keysearch({'science_keyword':['Galaxy chemistry']},
print_targets=False)
# + [markdown] deletable=false editable=false
# ## 4.1 Export results as a table
#
# The [<code>alminer.save_table</code>](../pages/api.rst#alminer.save_table) function writes the provided DataFrame to a table in CSV format in the 'tables' folder within the current working directory. If the 'tables' folder does not exist, it will be created.
#
# + [markdown] deletable=false editable=false
# ### Example 4.1.1: save query results as a table
# + deletable=false editable=false
alminer.save_table(observations, filename="galaxy_chemistry")
# + [markdown] deletable=false editable=false
# ## 4.2 Save overview plots
#
# The [<code>alminer.save_source_reports</code>](../pages/api.rst#alminer.save_source_reports) function creates overview plots of observed frequencies, angular resolution, LAS, frequency and velocity resolutions for each source in the provided DataFrame and saves them in PDF format in the 'reports' folder in the current working directory. If the 'reports' folder does not exist, it will be created. The reports are named after the target names.
#
# <u>Note</u>: Currently, the grouping is done based on ALMA target names, so the same source with a slighly different naming schemes will be treated as separate targets.
#
# + [markdown] deletable=false editable=false
# ### Example 4.2.1: save overview plots of each target with CO lines marked
# + [markdown] deletable=false editable=false
# Let's first narrow down our large query to a smaller subset to only a range of frequencies (Band 3) and angular resolutions < 0.5":
# + deletable=false editable=false
selected = observations[(observations["min_freq_GHz"] > 80.0) &
(observations["max_freq_GHz"] < 115.0) &
(observations["ang_res_arcsec"] < 0.5)]
alminer.summary(selected)
# + [markdown] deletable=false editable=false
# Now we can create and save plots for each source, with CO and its isotopologues marked:
# + deletable=false editable=false
alminer.save_source_reports(selected, mark_CO=True)
| docs/tutorials/4_create_reports.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,.md//md
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3 Keeping track of distance – how far have we come?
#
# One of the main problems a robot has is locating itself in its environment: the simulated robot doesn’t know where it is in the simulated world. Within the simulator, some `x` and `y` coordinate values are used by the simulator to keep track of the simulated robot’s location, and an angle parameter records its orientation. This information is then used to draw the robot on the background canvas.
#
# Typically when using a simulator, we try not to cheat by providing the robot direct access to simulator state information. That would be rather like you going outside, raising your arms to the sky, shouting ‘*Where am I?*’ and the universe responding with your current location.
#
# Instead, we try to ensure that the information on which the robot makes its decisions comes from its own internal reasoning and any sensor values it has access to.
#
# At this point, we could create a magical ‘simulated-GPS’ sensor that allows the robot to identify its location from the simulator’s point of view; but in the real world we can’t always guarantee that external location services are available. For example, GPS doesn’t work indoors or underground, or even in many cities where line-of-sight access to four or more GPS satellites is not available.
#
# Instead, we often have to rely on other sensors to help us identify our robot’s location, at least in a relative sense to where it has been previously. So that’s what we’ll be exploring in this notebook.
#
# Load in the simulator in the normal way, and then we’ll begin.
# +
from nbev3devsim.load_nbev3devwidget import roboSim, eds
# %load_ext nbev3devsim
# -
# ## 3.1 Motor tachometer or ‘rotation sensor’ data
#
# We can get a sense of how far the robot has travelled by logging the `.position` of each motor as recorded by its internal tachometer. Inside a real motor, a rotary encoder can be used to detect rotation of the motor. When the motor turns in one direction, the count goes up; when it turns in the other direction, it goes down.
#
# We’ll be looking at the logged data using notebook tools, so let’s take the precaution of clearing the datalog and also setting up a default chart object:
# %sim_data --clear
# %sim_magic --chart
# In the chart, make sure the *Wheel left* and *Wheel right* traces are selected.
# As well as each sensor value, we will also capture the simulator ‘clock time’.
#
# When running the simulator, you may have noticed that the simulator sometimes seems to slow down, perhaps because your computer processor is interrupted by having to commit resource to some other task. Inside the simulator, however, is an internal clock that counts simulator steps. Depending on how much work the simulator has to do to calculate updates in each step of the simulator run, this may take more or less ‘real time’ as measured by a clock on your wall (or, more likely, the clock on your mobile phone!).
#
# By logging the simulator step time using the preloaded `get_clock()` function, we can create a more accurate plot of the sensor values at each step of simulator time, irrespective of how long that step took to calculate in the real world.
# + [markdown] tags=["alert-success"]
# In his book *New Dark Age*, artist <NAME> describes the evolution of weather forecasting based on mathematical models:
#
# >In 1950, a team of meteorologists assembled at Aberdeen in order to perform the first automated twenty-four-hour weather forecast.... For this project, the boundaries of the world were the edges of the continental United States; a grid separated it into fifteen rows and eighteen columns. The calculation programmed into the machine consisted of sixteen successive operations, each of which had to be carefully planned and punched into cards, and which in turn output a new deck of cards that had to be reproduced, collated, and sorted. The meteorologists worked in eight-hour shifts, supported by programmers, and the entire run absorbed almost five weeks, 100,000 IBM punch cards, and a million mathematical operations. But when the experimental logs were examined, <NAME>, the director of the experiment, discovered that the actual computational time was almost exactly twenty-four hours. ‘One has reason to hope’, he wrote, that ‘Richardson’s dream of advancing computation faster than the weather may soon be realised’.
#
# *Historical note: <NAME> was an early pioneer of weather forecasting, whose story is also summarised by Bridle.*
#
# As Bridle observes, the number crunching required to perform a weather forecast requires the solution of lots of complex mathematical equations, so much so that the earliest computers might take days to make a 24-hour weather forecast. If you’re in the habit of checking online weather reports just before you set out for the day, they wouldn’t be much use if they took the next three days to compute. Although our simulator is a simple one, at times it may still take more computational resource than is available to the program for it to compute a single second of time in the simulated world in less than a second of real-world time.
#
# Reference: <NAME>. (2018) *New Dark Age: Technology and the End of the Future*, London: Verso Books.
# + [markdown] activity=true
# ### 3.1.1 Activity – Keeping track of motor position data
#
# The following program logs the position count as we drive the robot forwards, backwards, wait a while, then turn on the spot slowly one way, then quickly the other. I have also instrumented it so that the simulated robot says aloud what it is about to do next as the program progresses.
#
# For convenience, let’s clear the datalog again just here:
# + activity=true
# %sim_data --clear
# + [markdown] activity=true
# Download the program, and then enable the *Chart display* from the simulator toggle display buttons.
#
# Ensure the *Left wheel* and *Right wheel* traces are selected, then run the program from the *Simulator controls* or keyboard shortcut (`R`).
#
# Observe what happens to the wheel position counts as the robot progresses.
# + activity=true
# %%sim_magic_preloaded -pC -b Empty_Map -x 250 -y 550
tank_steering = MoveSteering(OUTPUT_B, OUTPUT_C)
tank_steering.left_motor.position = str(0)
tank_steering.right_motor.position = str(0)
#tank_steering.on(DIRECTION, SPEED)
from time import sleep
SAMPLE_TIME = 0.1
def datalogger():
"""Simple datalogging function."""
print('Wheel_left: {} {}'.format(tank_steering.left_motor.position, get_clock()))
print('Wheel_right: {} {}'.format(tank_steering.right_motor.position, get_clock()))
sleep(SAMPLE_TIME)
# Move forwards quickly
say("Forwards quickly")
tank_steering.on(0, 40)
for i in range(20):
datalogger()
# Move forwards slowly
say("Forwards slowly")
tank_steering.on(0, 5)
for i in range(40):
datalogger()
# Move backwards at intermediate speed
say("Backwards medium")
tank_steering.on(0, -20)
for i in range(20):
datalogger()
# Stop awhile
say("Stop awhile")
tank_steering.on(0, 0)
for i in range(20):
datalogger()
# Turn slowly on the spot
say("On the spot")
tank_steering.on(-100, 5)
for i in range(20):
datalogger()
# Turn more quickly on the spot the other way
say("And back")
tank_steering.on(100, 50)
for i in range(20):
datalogger()
say("All done")
# + [markdown] student=true
# *Record your observations about the chart data trace here. Make your own notes to describe how the behaviour of the different motor position chart traces explains the observed behaviour of the robot.*
# -
# ### 3.1.2 Viewing the data in a notebook chart
#
# The chart display in the simulator uses ‘sample number’ along the horizontal *x*-axis to log the data. This can result in some misleading traces as we can currently only add one sensor value to each sample.
#
# To plot the data more accurately, we need to plot the samples as a proper time series, with the sample timestamp as the *x*-coordinate.
#
# We can do that by charting the data from the datalog directly in the notebook.
#
# Retrieve the data from the datalog, and preview it:
# +
#Grab the logged data into a pandas dataframe
# df = %sim_data
#Preview the first few rows of the dataset
df.head()
# -
# Now we can chart the data:
# +
# Load in the seaborn charting package
import seaborn as sns
# Generate a line chart from the datalog dataframe
ax = sns.lineplot(x="time",
y="value",
# The hue category defines line color
hue='variable',
data=df);
# -
# Here’s a stylised impression of what my chart looked like:
#
# 
#
# You’ll notice that I have added some vertical grey lines to my chart to indicate different areas of the chart, as well as some simple labels identifying each area.
#
# Annotating charts can often help us make more sense of them when we try to read them. In creating such charts there is often a balance between making a ‘production quality’ chart that you could share with other people as part of a formal report (or formal teaching materials!) and a ‘good enough for personal use’ feel for your own reference.
#
# In this regard, you may also notice that the chart shown above has an informal, stylised feel to it. The chart really was created from the data I collected, but I then styled it using an XKCD chart theme to differentiate it from the charts generated within the notebook.
# + [markdown] heading_collapsed=true tags=["optional-extra"]
# ### 3.1.3 Create your own annotated chart (optional)
#
# *Click on the arrow in the sidebar or run this cell to reveal how to create the annotated chart.*
# + [markdown] hidden=true tags=["optional-extra"]
# I used the following code to create the annotated chart. I manually set the horizontal *x*-axis values where I wanted the vertical lines to appear.
# + hidden=true tags=["optional-extra"]
# Manually set x-axis coordinates for vertical lines
forward_fast = 0.3
forward_slow = 1.6
backwards_medium = 4.1
stop_awhile = 5.5
on_the_spot = 6.7
and_back = 8.0
import warnings
warnings.filterwarnings('ignore', '.*missing from current font.*',)
# We need some additional tools from matplotlib
import matplotlib.pyplot as plt
# I am using the xkcd theme.
# This gives the chart an informal
# or 'indicative example' feel
with plt.xkcd():
# Generate a line chart from the datalog dataframe
ax = sns.lineplot(x="time",
y="value",
# The hue category defines line color
hue='variable',
data=df)
# Move the legend outside the plotting area.
# The prevents it from overlapping areas of the plot
ax.legend( bbox_to_anchor=(1.0, 0.5))
o = 0.3 #offset text from line
line_colour = 'lightgrey'
def plot_line_label(x, label):
"""Annotate boundaried areas."""
# Create a vertical line
plt.axvline(x, c=line_colour)
# Create a text label
plt.text(x+o, 20, label)
# Add lines and labels to the chart
plot_line_label(forward_fast, 'Fwd\nfast')
plot_line_label(forward_slow, 'Fwd\nslow')
plot_line_label(backwards_medium, 'Back\nmed')
plot_line_label(stop_awhile, 'Stop\nawhile')
plot_line_label(on_the_spot, 'On the\nspot')
plot_line_label(and_back, 'And\nback');
# We can save the image as a file if required.
# Increase the figure size
#plt.figure(figsize=(12,8))
# Nudge the margins so we don't cut off labels
#plt.subplots_adjust(left=0.1, bottom=0.1,
# right=0.7, top=0.8)
# Save the image file
#plt.savefig('position_time.png')
# + [markdown] activity=true
# ### 3.1.4 Activity – Observing motor position counts for different motor actions (optional)
#
# How do the position counts vary for each wheel if the robot is driving forwards in a gentle curve, or a tight turn?
#
# For example, we might create such turns using the following steering commands:
#
# ```python
# # Graceful forwards left
# tank_steering.on(-30, 20)
#
# # Tighter turn forwards and to the right <!-- JD: but this is the same as the code above for a 'graceful left turn'. Needs to be something like 40, 100 -->
# tank_steering.on(-30, 20)
# ```
#
# Feel free to make your own predictions, or run a program, grab the data and analyse it yourself. If you do run your own experiment(s), then remember to clear the datalog before running your data-collecting code in the simulator.
# -
# ## 3.2 Measuring how far the robot has travelled
#
# The wheel `position` data corresponds to an angular measure, that is, how far the wheel has turned.
# + [markdown] activity=true
# ### 3.2.1 Activity – Driving the robot for a fixed number of wheel rotations
#
# Use the following program to drive the robot for a fixed number of rotations and observe how the position count increases. Based on your observations, make a note of what you think the position count actually measures.
# + activity=true
# %%sim_magic_preloaded
tank_steering= MoveSteering(OUTPUT_B, OUTPUT_C)
def reporter(last_position=0):
position = int(tank_steering.left_motor.position)
diff = position - last_position
print('Current {}, diff {}'.format(position, diff))
say('Diff {}'.format(diff))
return position
tank_steering.on_for_rotations(0, 10, 1)
last_position = reporter()
tank_steering.on_for_rotations(0, 10, 1)
last_position = reporter(last_position)
tank_steering.on_for_rotations(0, 10, 1)
reporter(last_position)
# + [markdown] student=true
# *Based on your observations of position counts for the number of wheel rotations travelled, what do you think the position value measures? Bear in mind that the simulation, like the real world, may have sources of noise that affect the actual values recorded, rather than ‘ideal’ ones.*
#
# *Record your impressions here.*
# + [markdown] activity=true heading_collapsed=true
# #### Example observations
#
# *Click the arrow in the sidebar or run this cell to reveal my observations.*
# + [markdown] activity=true hidden=true
# When I ran the program, I got counts between 365 and 380 for each rotation, depending in part on the speed I set the wheels to run at.
#
# The simulator actually runs in steps, with 30 steps per simulated second. This means that at 20% speed, the wheel will turn approximately 6 to 7 degrees each step. By the time the simulator detects that the wheel has reached *at least* 360 degrees (i.e. completed one rotation), it may already have exceeded that amount of turn; so the stopping condition for the `.on_for_rotations` function, which is based on observing the turned angle, may actually stop the motors after more than one rotation.
#
# So notwithstanding the values we get for the position count after a single rotation, the position is actually measured in degrees.
# + [markdown] activity=true
# ## 3.3 Controlling the distance travelled
#
# If we can continuously monitor the distance we have travelled, then we can use that as part of a control strategy in a non-blocking way: rather than tell the motors to turn on for N rotations, we can just turn them on, and then take a particular action when they have turned far enough *as we have measured them*.
# + [markdown] activity=true
# ### 3.3.1 Activity – Are we there yet?
#
# In this activity, you will experiment with driving the robot over a fixed distance.
#
# Nominally, the wheel diameter is set in the robot configuration file to `56`, that is, 56 mm, so just under six centimetres.
#
# What this means is that we can drive the robot forward a specified distance.
#
# The bands shown on the *Coloured_bands* background are 60 cm high.
#
# See if you can write a program that drives the robot exactly the length of one of the bands by monitoring the `position` value of one of the motors as the robot drives in a straight line.
#
# How accurately can you cover the distance? (Don’t panic, or waste too much time, if you can’t...). Comment on your results.
#
# __Hint: how many degrees will the wheel need to turn for the wheel to turn 60 cm?__
# + [markdown] activity=true tags=["alert-danger"]
# __Warning: if you use a loop then put something in it that uses up simulator time and progresses the clock, such as a short `sleep(0.1)` command, otherwise you may find that your simulator Python program gets stuck and hangs the browser.__
#
# *If that happens try to reload the notebook in the browser.*
#
# *If that doesn’t work, from the notebook home page, try to shutdown the notebook.*
#
# *If that is also stuck, restart your browser.*
#
# *If things are still not working properly, you will need to restart the container from Docker dashboard or the command line: `docker restart tm129vce`.*
# + [markdown] activity=true
# *Note: the `.position` value is returned as a string and should be converted to an integer (`int`) if you want to use it numerically.*
# + [markdown] student=true
# *How many degrees does the wheel need to turn? Record your calculation and result here.*
#
# *Remember that you can use a code cell as a interactive calculator. You can access pi as a number by importing `from math import pi`.*
# + activity=true
# %%sim_magic_preloaded -b Coloured_bands -p -x 1100 -y 200 -a 90
from time import sleep
# Your code here
# + [markdown] student=true
# *Record your notes and observations here about how effectively the robot performed the desired task.*
# + [markdown] activity=true heading_collapsed=true
# #### Example observations
#
# *Click on the arrow in the sidebar or run this cell to reveal some example observations.*
# + [markdown] activity=true hidden=true
# The `position` counter reports the number of degrees turned by the wheel, so let’s start by finding out how many degrees we need to turn the wheel to travel 60 cm.
#
# Recall that the wheel diameter is 56 mm.
# + activity=true hidden=true
from math import pi
distance = 60 #cm
# 56mm is 5.6cm
circumference = 5.6 * pi
no_of_turns = distance / circumference
no_of_degrees = no_of_turns * 360
int(no_of_degrees)
# + [markdown] activity=true hidden=true
# We can use this calculation in a program that drives the robot forward until the wheels have turned by the desired amount, and then stops.
#
# Note that the speed of the robot may affect how accurately the robot can perform the task, bearing in mind the comment earlier about how the simulator uses quite crude discrete time steps to animate the world.
# + activity=true hidden=true
# %%sim_magic_preloaded -b Coloured_bands -p -x 1100 -y 200 -a 90
from time import sleep
tank_steering= MoveSteering(OUTPUT_B, OUTPUT_C)
# Go Forwards
DIRECTION = 0
SPEED = 20
tank_steering.on(DIRECTION, SPEED)
# Do the math...
from math import pi
distance = 60 #cm
# 56mm is 5.6cm
circumference = 5.6 * pi
no_of_turns = distance / circumference
no_of_degrees = no_of_turns * 360
while int(tank_steering.left_motor.position) < no_of_degrees:
sleep(0.01)
print("Position: {}".format(tank_steering.left_motor.position))
# + [markdown] activity=true hidden=true
# When I ran the program, it did pretty well, running between the lines and stopping maybe just a fraction too long.
# -
# ## 3.4 Measuring the width of a coloured track
#
# One of the activities in the Open University [T176 *Engineering* residential school](http://www.open.ac.uk/jobs/residential-schools/modules/modules-summer-schools/txr120-engineering-active-introduction) is a robotics challenge to recreate a test track that depicts several coloured bands of various widths purely from data logged by an EV3 Lego robot.
#
# Let’s try a related, but slightly simpler challenge: identifying the width of the track that is displayed on the *Loop* background.
# + [markdown] activity=true
# ### 3.4.1 Using logged data to take measurements from the simulated world (optional)
#
# In this activity, you will use data logged by the robot to learn something about the structure of the world it is operating in.
#
# Clear the data from the datalog:
# + activity=true
# %sim_data --clear
# + [markdown] activity=true
# Then download and run the following program in the simulator to drive the robot over the test track.
# + activity=true
# %%sim_magic_preloaded -C -b Loop
from time import sleep
SAMPLE_TIME = 0.1
tank_steering= MoveSteering(OUTPUT_B, OUTPUT_C)
# Define a light sensor
colorLeft = ColorSensor(INPUT_2)
def datalogger():
"""Simple datalogging function."""
print('Wheel_left: {} {}'.format(tank_steering.left_motor.position, get_clock()))
print('Light_left: {} {}'.format(colorLeft.full_reflected_light_intensity, get_clock()))
sleep(SAMPLE_TIME)
tank_steering.on(0, 20)
while int(tank_steering.left_motor.position)<1000:
datalogger()
# + [markdown] activity=true
# From a chart display of the data, such as the one that is generated if you run the code cell below, how might you identify the width of the black line?
#
# *Hint: in the interactive plotly chart, if you hover over the chart to raise the plotly toolbar then you can select `Compare data on hover` to report the line values for a particular x-axis value; the `Toggle Spike Lines` view will also show dynamic crosslines highlighting the current x- and y-values.*
# + activity=true
import pandas as pd
pd.options.plotting.backend = "plotly"
# Grab the logged data into a pandas dataframe
# df_line = %sim_data
# Generate a line chart from the datalog dataframe
df_line.plot( x = 'time', y = 'value', color='variable')
# + [markdown] student=true
# *How can you use the logged data displayed in the chart above, or otherwise, to work out how wide the black line is? What other information, if any, do you need in order to express this as a distance in (simulated) metres?*
#
# *Record your observations here.*
# + [markdown] activity=true heading_collapsed=true
# #### Example observations
# *Click on the arrow in the sidebar or run this cell to reveal some example observations.*
# + [markdown] activity=true hidden=true
# The chart shows the increasing trace from the position sensor and another trace for the light sensor. The light sensor value dips from 100 to 0 as the robot goes over the black line, then goes back to 100.
#
# The horizontal *x*-axis is simulator time. If we take the position count reading at the same time that the robot detects the black line, and again at the same time that the robot crosses back onto the white background, then we can subtract the first position value from the second to give use the distance travelled by the robot.
#
# To convert this to simulated metres, we would need to know what distance is travelled for a motor position increment value of 1.
#
# If the position count is an angular measure (for example, degrees of wheel turn), then we could calculate the distance travelled as:
#
# `wheel_circumference * degrees_turned / 360`
#
# since the distance travelled by the wheel in one complete turn is the same as its circumference.
#
# We could calculate the circumference as:
#
# `wheel_circumference = 2 * wheel_radius * pi`
#
# or:
#
# `wheel_circumference = wheel_diameter * pi`
#
# both of which could be measured from the robot *if* we had physical access to it.
# -
# ## 3.5 Summary
#
# In this notebook you have seen how the motor `position` tachometer can be used to record how far, in fractions of a turn, each motor has turned. Moving forwards increases the motor tacho counts, whereas reversing reduces them both; turning increases one whilst reducing the other.
#
# Tacho counts are very useful for providing an indicative feel for how a robot has travelled, but they may not be particularly accurate. As with many data traces, *trends* and *differences* often tell us much of what we need to know.
#
# Through working with the motors and sensors at quite a low level, you have also started to learn how the implementation of the simulator itself may affect the performance of our programs. In certain cases, we may even have to do things in the program code that are there simply to accommodate some ‘feature’ of the way the simulator is implemented that would not occur in the real robot. In the physical world, time flows continuously of its own accord – in real time! In the simulator, we simulate it in discrete steps, which may even take longer to compute than the amount of time the step is supposed to represent.
#
# In the next notebook, you will review another sensor that helps our robot know where it’s going: the gyro sensor.
| content/06. Where in the world are we/06.3 Keeping track of distance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: base_py39
# language: python
# name: base_py39
# ---
# # AE vs VAE for MNIST
# **All sources can be found in the notebook called ECG AE Anomaly detection.ipynb**
# I don't have access to torchvision because my cuda is too recent. So I had to do the dataloading myself. You probably don't have to.
#
#
# We are going to consider the MNIST dataset as tabular data of 784 features (28*28 pixels =784).Indeed we are going to use mulilayer perceptron and not convolutional layers here. First because we actually don't really need it, second because it is easier, third because this is the kind of data I am actually going to work with in the projects.
#
# The auto encoder will look like 784->400->50->400->784 : so we are going to represent our images (vector of length 784) into a 50 dimensional latent space, using 2 perceptrons as an encoder and 2 perceptrons as a decoder.
# +
from sklearn.datasets import fetch_openml
from sklearn.utils import check_random_state
from sklearn.model_selection import train_test_split
train_samples = 60000
X, y = fetch_openml('mnist_784', version=1, return_X_y=True, as_frame=False)
random_state = check_random_state(0)
permutation = random_state.permutation(X.shape[0])
X = X[permutation]
y = y[permutation]
X = X.reshape((X.shape[0], -1))
X_train, X_test, y_train, y_test = train_test_split(
X, y, train_size=train_samples, test_size=10000)
# -
# # Pyro VAE
# +
import os
import numpy as np
import torch
import torch.nn as nn
import pyro
import pyro.distributions as dist
from pyro.infer import SVI, Trace_ELBO
from pyro.optim import Adam
# +
import torch; torch.manual_seed(0)
import torch.nn.functional as F
import torch.utils
import torch.distributions
import numpy as np
import matplotlib.pyplot as plt; plt.rcParams['figure.dpi'] = 200
# +
assert pyro.__version__.startswith('1.7.0')
pyro.distributions.enable_validation(False)
pyro.set_rng_seed(0)
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# +
class Decoder_pyro(nn.Module):
def __init__(self, z_dim, hidden_dim):
super().__init__()
# setup the two linear transformations used
self.fc1 = nn.Linear(z_dim, hidden_dim)
self.fc21 = nn.Linear(hidden_dim, 784)
# setup the non-linearities
self.softplus = nn.Softplus()
self.sigmoid = nn.Sigmoid()
def forward(self, z):
# define the forward computation on the latent z
# first compute the hidden units
hidden = self.softplus(self.fc1(z))
# return the parameter for the output Bernoulli
# each is of size batch_size x 784
loc_img = self.sigmoid(self.fc21(hidden))
return loc_img
class Encoder_pyro(nn.Module):
def __init__(self, z_dim, hidden_dim):
super().__init__()
# setup the three linear transformations used
self.fc1 = nn.Linear(784, hidden_dim)
self.fc21 = nn.Linear(hidden_dim, z_dim)
self.fc22 = nn.Linear(hidden_dim, z_dim)
# setup the non-linearities
self.softplus = nn.Softplus()
def forward(self, x):
# define the forward computation on the image x
# first shape the mini-batch to have pixels in the rightmost dimension
x = x.reshape(-1, 784)
# then compute the hidden units
hidden = self.softplus(self.fc1(x))
# then return a mean vector and a (positive) square root covariance
# each of size batch_size x z_dim
z_loc = self.fc21(hidden)
z_scale = torch.exp(self.fc22(hidden))
return z_loc, z_scale
class VAE(nn.Module):
# by default our latent space is 50-dimensional
# and we use 400 hidden units
def __init__(self, z_dim=50, hidden_dim=400, use_cuda=False):
super().__init__()
# create the encoder and decoder networks
self.encoder = Encoder_pyo(z_dim, hidden_dim)
self.decoder = Decoder_pyro(z_dim, hidden_dim)
if use_cuda:
# calling cuda() here will put all the parameters of
# the encoder and decoder networks into gpu memory
self.cuda()
self.use_cuda = use_cuda
self.z_dim = z_dim
# define the model p(x|z)p(z)
def model(self, x):
# register PyTorch module `decoder` with Pyro
pyro.module("decoder", self.decoder)
with pyro.plate("data", x.shape[0]):
# setup hyperparameters for prior p(z)
z_loc = x.new_zeros(torch.Size((x.shape[0], self.z_dim)))
z_scale = x.new_ones(torch.Size((x.shape[0], self.z_dim)))
# sample from prior (value will be sampled by guide when computing the ELBO)
z = pyro.sample("latent", dist.Normal(z_loc, z_scale).to_event(1))
# decode the latent code z
loc_img = self.decoder(z)
# score against actual images(I now understand why they ask to rescale the image but still don't get
# why thsy use bernoulli : it can't be bernouilli. The only answer I found on internet was : it work
# best that way....)
pyro.sample("obs", dist.Bernoulli(loc_img).to_event(1), obs=x.reshape(-1, 784))
# define the guide (i.e. variational distribution) q(z|x)
def guide(self, x):
# register PyTorch module `encoder` with Pyro
pyro.module("encoder", self.encoder)
with pyro.plate("data", x.shape[0]):
# use the encoder to get the parameters used to define q(z|x)
z_loc, z_scale = self.encoder(x)
# sample the latent code z
pyro.sample("latent", dist.Normal(z_loc, z_scale).to_event(1))
# define a helper function for reconstructing images
def reconstruct_img(self, x):
# encode image x
z_loc, z_scale = self.encoder(x)
# sample in latent space
z = dist.Normal(z_loc, z_scale).sample()
# decode the image (note we don't sample in image space)
loc_img = self.decoder(z)
return loc_img
# -
from torch.utils.data import Dataset
class Mnist_dataset(Dataset):
def __init__(self,images,targets):
self.images=images
self.targets=targets
def __len__(self):
return len(self.images)
def __getitem__(self, item):
image=self.images[item]
re_image=image/255
target=int(self.targets[item])
return {'re_image': torch.tensor(re_image.reshape(28,28)[np.newaxis,:,:], dtype=torch.float32),
'target': torch.tensor(target, dtype=torch.long)
}
# +
def train(svi, train_loader, use_cuda=False):
# initialize loss accumulator
epoch_loss = 0.
# do a training epoch over each mini-batch x returned
# by the data loader
for d in train_loader:
# if on GPU put mini-batch into CUDA memory
x=d['re_image']
if use_cuda:
x = x.cuda()
# do ELBO gradient and accumulate loss
epoch_loss += svi.step(x)
# return epoch loss
normalizer_train = len(train_loader.dataset)
total_epoch_loss_train = epoch_loss / normalizer_train
return total_epoch_loss_train
# +
LEARNING_RATE = 1.0e-3
USE_CUDA = True
NUM_EPOCHS = 20
# +
i_dataset = Mnist_dataset(images=X_train,targets=y_train)
i_data_loader = torch.utils.data.DataLoader(i_dataset,batch_size=128,num_workers=16)
# +
pyro.clear_param_store()
# setup the VAE
vae = VAE(use_cuda=USE_CUDA)
# setup the optimizer
adam_args = {"lr": LEARNING_RATE}
optimizer = Adam(adam_args)
# setup the inference algorithm
svi = SVI(vae.model, vae.guide, optimizer, loss=Trace_ELBO())
train_elbo = []
test_elbo = []
# training loop
for epoch in range(NUM_EPOCHS):
total_epoch_loss_train = train(svi, i_data_loader, use_cuda=USE_CUDA)
train_elbo.append(-total_epoch_loss_train)
#print("[epoch %03d] average training loss: %.4f" % (epoch, total_epoch_loss_train))
# -
def loss_plot_pyro(loss):
fig=plt.figure(figsize=(5,5))
plt.plot(np.arange(len(loss)),loss)
plt.xlabel('Epochs')
plt.ylabel('ELBO')
plt.show()
loss_plot_pyro(train_elbo)
# +
import umap.umap_ as umap
def plot_latent_var_pyro(autoencoder, data,nei, num_batches=100):
stack=[]
stacky=[]
for i, d in enumerate(data):
x=d['re_image']
y=d['target'].to('cpu').detach().numpy().tolist()
z,sigma = autoencoder.encoder(x.to(device))
z = z.to('cpu').detach().numpy().tolist()
stack.extend(z)
stacky.extend(y)
#umaper = umap.UMAP(n_components=2,n_neighbors=nei)
#x_umap = umaper.fit_transform(z)
#plt.scatter(z[:, 0], z[:, 1], c=y, cmap='tab10')
if i > num_batches:
umaper = umap.UMAP(n_components=2,n_neighbors=nei)
x_umap = umaper.fit_transform(stack)
plt.scatter(x_umap[:, 0], x_umap[:, 1],s=2, c=stacky, cmap='tab10')
plt.colorbar()
plt.xlabel('UMAP 1')
plt.ylabel('UMAP 2')
break
# -
plot_latent_var_pyro(vae, i_data_loader,10)
# # Pytorch AE multilayer perceptron
class Encoder(nn.Module):
def __init__(self, latent_dims):
super(Encoder, self).__init__()
self.linear1 = nn.Linear(784, 400)
self.linear2 = nn.Linear(400, latent_dims)
def forward(self, x):
x = torch.flatten(x, start_dim=1)
x = F.relu(self.linear1(x))
return self.linear2(x)
class Decoder(nn.Module):
def __init__(self, latent_dims):
super(Decoder, self).__init__()
self.linear1 = nn.Linear(latent_dims, 400)
self.linear2 = nn.Linear(400, 784)
def forward(self, z):
z = F.relu(self.linear1(z))
z = torch.sigmoid(self.linear2(z))
return z.reshape((-1, 1, 28, 28))
class Autoencoder(nn.Module):
def __init__(self, latent_dims):
super(Autoencoder, self).__init__()
self.encoder = Encoder(latent_dims)
self.decoder = Decoder(latent_dims)
def forward(self, x):
z = self.encoder(x)
return self.decoder(z)
keep_loss=[]
def train(autoencoder, data, epochs=20):
opt = torch.optim.Adam(autoencoder.parameters())
for epoch in range(epochs):
loss_epo=[]
for d in data:
x=d['re_image']
x = x.to(device) # GPU
opt.zero_grad()
x_hat = autoencoder(x)
loss = ((x - x_hat)**2).mean()
loss.backward()
opt.step()
loss_epo.append(loss.cpu().detach().numpy())
keep_loss.append(np.mean(loss_epo))
return autoencoder
# +
latent_dims = 50
autoencoder = Autoencoder(latent_dims).to(device) # GPU
i_dataset = Mnist_dataset(images=X_train,targets=y_train)
i_data_loader = torch.utils.data.DataLoader(i_dataset,batch_size=128,num_workers=16)
autoencoder = train(autoencoder, i_data_loader)
# +
import umap.umap_ as umap
def plot_latent(autoencoder, data,nei, num_batches=100):
stack=[]
stacky=[]
for i, d in enumerate(data):
x=d['re_image']
y=d['target'].to('cpu').detach().numpy().tolist()
z = autoencoder.encoder(x.to(device))
z = z.to('cpu').detach().numpy().tolist()
stack.extend(z)
stacky.extend(y)
#umaper = umap.UMAP(n_components=2,n_neighbors=nei)
#x_umap = umaper.fit_transform(z)
#plt.scatter(z[:, 0], z[:, 1], c=y, cmap='tab10')
if i > num_batches:
umaper = umap.UMAP(n_components=2,n_neighbors=nei)
x_umap = umaper.fit_transform(stack)
plt.scatter(x_umap[:, 0], x_umap[:, 1],s=2, c=stacky, cmap='tab10')
plt.colorbar()
plt.xlabel('UMAP 1')
plt.ylabel('UMAP 2')
break
def loss_plot(loss):
fig=plt.figure(figsize=(5,5))
plt.plot(np.arange(len(loss)),loss)
plt.xlabel('Epochs')
plt.ylabel('MSE')
plt.show()
# -
loss_plot(keep_loss)
plot_latent(autoencoder, i_data_loader,100)
| ECG_AE_anomaly_detection/AE comparison for Mnist-Short.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
import tensorflow as tf
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
import sys
if not '../' in sys.path: sys.path.append('../')
import pandas as pd
from utils import data_utils
from model_config import config
from ved_detAttn import VarSeq2SeqDetAttnModel
# +
if config['experiment'] == 'qgen':
print('[INFO] Preparing data for experiment: {}'.format(config['experiment']))
train_data = pd.read_csv(config['data_dir'] + 'df_qgen_train.csv')
val_data = pd.read_csv(config['data_dir'] + 'df_qgen_val.csv')
test_data = pd.read_csv(config['data_dir'] + 'df_qgen_test.csv')
input_sentences = pd.concat([train_data['answer'], val_data['answer'], test_data['answer']])
output_sentences = pd.concat([train_data['question'], val_data['question'], test_data['question']])
true_test = test_data['question']
input_test = test_data['answer']
filters = '!"#$%&()*+,./:;<=>?@[\\]^`{|}~\t\n'
w2v_path = config['w2v_dir'] + 'w2vmodel_qgen.pkl'
elif config['experiment'] == 'dialogue':
train_data = pd.read_csv(config['data_dir'] + 'df_dialogue_train.csv')
val_data = pd.read_csv(config['data_dir'] + 'df_dialogue_val.csv')
test_data = pd.read_csv(config['data_dir'] + 'df_dialogue_test.csv')
input_sentences = pd.concat([train_data['line'], val_data['line'], test_data['line']])
output_sentences = pd.concat([train_data['reply'], val_data['reply'], test_data['reply']])
true_test = test_data['reply']
input_test = test_data['line']
filters = '!"#$%&()*+/:;<=>@[\\]^`{|}~\t\n'
w2v_path = config['w2v_dir'] + 'w2vmodel_dialogue.pkl'
else:
print('Invalid experiment name specified!')
# +
print('[INFO] Tokenizing input and output sequences')
x, input_word_index = data_utils.tokenize_sequence(input_sentences,
filters,
config['encoder_num_tokens'],
config['encoder_vocab'])
y, output_word_index = data_utils.tokenize_sequence(output_sentences,
filters,
config['decoder_num_tokens'],
config['decoder_vocab'])
print('[INFO] Split data into train-validation-test sets')
x_train, y_train, x_val, y_val, x_test, y_test = data_utils.create_data_split(x,
y,
config['experiment'])
encoder_embeddings_matrix = data_utils.create_embedding_matrix(input_word_index,
config['embedding_size'],
w2v_path)
decoder_embeddings_matrix = data_utils.create_embedding_matrix(output_word_index,
config['embedding_size'],
w2v_path)
# Re-calculate the vocab size based on the word_idx dictionary
config['encoder_vocab'] = len(input_word_index)
config['decoder_vocab'] = len(output_word_index)
# -
model = VarSeq2SeqDetAttnModel(config,
encoder_embeddings_matrix,
decoder_embeddings_matrix,
input_word_index,
output_word_index)
# +
if config['load_checkpoint'] != 0:
checkpoint = config['model_checkpoint_dir'] + str(config['load_checkpoint']) + '.ckpt'
else:
checkpoint = tf.train.get_checkpoint_state(os.path.dirname('models/checkpoint')).model_checkpoint_path
preds = model.predict(checkpoint,
x_test,
y_test,
true_test,
)
# -
count = 100
model.show_output_sentences(preds[:count],
y_test[:count],
input_test[:count],
true_test[:count],
)
model.get_diversity_metrics(checkpoint, x_test, y_test)
| ved_detAttn/predict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: Scorecard with continuous target
# In this tutorial, we show that the use of scorecards is not limited to binary classification problems. We develop a scorecard using the Huber regressor as an estimator. The dataset for this tutorial is https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_california_housing.html.
# +
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.datasets import fetch_california_housing
from sklearn.linear_model import HuberRegressor
from optbinning import BinningProcess
from optbinning import Scorecard
# -
# Load the dataset.
# +
data = fetch_california_housing()
target = "target"
variable_names = data.feature_names
df = pd.DataFrame(data.data, columns=variable_names)
df[target] = data.target
# -
df.head()
# Then, we instantiate a ``BinningProcess`` object class with variable names.
binning_process = BinningProcess(variable_names)
# We select a robust linear model as an estimator.
estimator = HuberRegressor(max_iter=200)
# Finally, we instantiate a ``Scorecard`` class with the target name, a binning process object, and an estimator. In addition, we want to apply a scaling method to the scorecard points. Also, we select the reverse scorecard mode, so the score increases as the average house value increases.
scorecard = Scorecard(binning_process=binning_process, target=target,
estimator=estimator, scaling_method="min_max",
scaling_method_params={"min": 0, "max": 100},
reverse_scorecard=True)
scorecard.fit(df)
# Similar to other objects in OptBinning, we can print overview information about the options settings, problems statistics, and the number of selected variables after the binning process.
scorecard.information(print_level=2)
# Two scorecard styles are available: ``style="summary"`` shows the variable name, and their corresponding bins and assigned points; ``style="detailed"`` adds information from the corresponding binning table.
scorecard.table(style="summary")
scorecard.table(style="detailed")
# Compute score and predicted target using the fitted estimator.
score = scorecard.score(df)
y_pred = scorecard.predict(df)
# The following plot shows a perfect linear relationship between the score and average the house value.
plt.scatter(score, df[target], alpha=0.01, label="Average house value")
plt.plot(score, y_pred, label="Huber regression", linewidth=2, color="orange")
plt.ylabel("Average house value (unit=100,000)")
plt.xlabel("Score")
plt.legend()
plt.show()
| doc/source/tutorials/tutorial_scorecard_continuous_target.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import display, Image
# + [markdown] nbpresent={"id": "29b9bd1d-766f-4422-ad96-de0accc1ce58"}
# # Lab 4 - Convolutional Neural Network with MNIST
#
# This lab corresponds to Module 4 of the "Deep Learning Explained" course. We assume that you have successfully Lab 1 to download the MNIST data.
#
# We will train a Convolutional Neural Network (CNN) on MNIST data.
#
# ## Introduction
#
# A [convolutional neural network](https://en.wikipedia.org/wiki/Convolutional_neural_network) (CNN, or ConvNet) is a type of [feed-forward](https://en.wikipedia.org/wiki/Feedforward_neural_network) artificial neural network made up of neurons that have learnable weights and biases, very similar to ordinary multi-layer perceptron (MLP) networks introduced in Module 3. The CNNs take advantage of the spatial nature of the data.
#
# In nature, we perceive different objects by their shapes, size and colors. For example, objects in a natural scene are typically composed of edges, corners/vertices (defined by two of more edges), color patches etc. These primitives are often identified using different detectors (e.g., edge detection, color detector) or combination of detectors interacting to facilitate image interpretation (object classification, region of interest detection, scene description etc.) in real world vision related tasks. These detectors are also known as filters.
#
# Convolution is a mathematical operator that takes an image and a filter as input and produces a filtered output (representing say egdges, corners, colors etc in the input image). Historically, these filters are a set of weights that were often hand crafted or modeled with mathematical functions (e.g., [Gaussian](https://en.wikipedia.org/wiki/Gaussian_filter) / [Laplacian](http://homepages.inf.ed.ac.uk/rbf/HIPR2/log.htm) / [Canny](https://en.wikipedia.org/wiki/Canny_edge_detector) filter). The filter outputs are mapped through non-linear activation functions mimicking human brain cells called [neurons](https://en.wikipedia.org/wiki/Neuron).
#
# Convolutional networks provide a machinery to learn these filters from the data directly instead of explicit mathematical models and have been found to be superior (in real world tasks) compared to historically crafted filters. With convolutional networks, the focus is on learning the filter weights instead of learning individually fully connected pair-wise (between inputs and outputs) weights. In this way, the number of weights to learn is reduced when compared with the traditional MLP networks from the previous tutorials. In a convolutional network, one learns several filters ranging in number from single digits to thousands depending on the network complexity.
#
# Many of the CNN primitives have been shown to have a conceptually parallel components in brain's [visual cortex](https://en.wikipedia.org/wiki/Visual_cortex). A neuron in the visual cortex will emit responses when a certain region of its input cells are stimulated. This region is known as the receptive field (RF) of the neuron.
#
# Equivalently, in a convolution layer, the input region corresponding to the filter dimensions at certain locations in the input layer can be considered as the receptive field of the nodes in the convolutional layer. Popular deep CNNs or ConvNets (such as [AlexNet](https://en.wikipedia.org/wiki/AlexNet), [VGG](https://arxiv.org/abs/1409.1556), [Inception](http://www.cv-foundation.org/openaccess/content_cvpr_2015/papers/Szegedy_Going_Deeper_With_2015_CVPR_paper.pdf), [ResNet](https://arxiv.org/pdf/1512.03385v1.pdf)) that are used for various [computer vision](https://en.wikipedia.org/wiki/Computer_vision) tasks have many of these architectural primitives (inspired from biology).
#
# We will introduce the convolution operation and gain familiarity with the different parameters in CNNs.
#
# **Problem**:
# We will continue to work on the same problem of recognizing digits in MNIST data. The MNIST data comprises of hand-written digits with little background noise.
# -
# Figure 1
Image(url= "http://3.bp.blogspot.com/_UpN7DfJA0j4/TJtUBWPk0SI/AAAAAAAAABY/oWPMtmqJn3k/s1600/mnist_originals.png", width=200, height=200)
# **Goal**:
# Our goal is to train a classifier that will identify the digits in the MNIST dataset.
#
# **Approach**:
#
# The same 5 stages we have used in the previous labs are applicable: Data reading, Data preprocessing, Creating a model, Learning the model parameters and Evaluating (a.k.a. testing/prediction) the model.
#
# We will experiment with two models with different architechtural components.
# + nbpresent={"id": "138d1a78-02e2-4bd6-a20e-07b83f303563"}
from __future__ import print_function # Use a function definition from future version (say 3.x from 2.7 interpreter)
import matplotlib.image as mpimg
import matplotlib.pyplot as plt
import numpy as np
import os
import sys
import time
import cntk as C
# %matplotlib inline
# -
# In the block below, we check if we are running this notebook in the CNTK internal test machines by looking for environment variables defined there. We then select the right target device (GPU vs CPU) to test this notebook. In other cases, we use CNTK's default policy to use the best available device (GPU, if available, else CPU).
# Select the right target device when this notebook is being tested:
if 'TEST_DEVICE' in os.environ:
if os.environ['TEST_DEVICE'] == 'cpu':
C.device.try_set_default_device(C.device.cpu())
else:
C.device.try_set_default_device(C.device.gpu(0))
# Test for CNTK version
if not C.__version__ == "2.0":
raise Exception("this notebook is designed to work with 2.0. Current Version: " + C.__version__)
# ## Data reading
# In this section, we will read the data generated in Lab1_MNIST_DataLoader.
#
# We are using the MNIST data that you have downloaded using the Lab1_MNIST_DataLoader notebook. The dataset has 60,000 training images and 10,000 test images with each image being 28 x 28 pixels. Thus the number of features is equal to 784 (= 28 x 28 pixels), 1 per pixel. The variable `num_output_classes` is set to 10 corresponding to the number of digits (0-9) in the dataset.
#
# In previous labs, as shown below, we have always flattened the input image into a vector. With convoultional networks, we do **not** flatten the image - we preserve its 2D shape..
#
# **Input Dimensions**:
#
# In convolutional networks for images, the input data is often shaped as a 3D matrix (number of channels, image width, image height), which preserves the spatial relationship between the pixels. In the MNIST data, the image is a single channel (grayscale) data, so the input dimension is specified as a (1, image width, image height) tuple.
#
# 
#
# Natural scene color images are often presented as Red-Green-Blue (RGB) color channels. The input dimension of such images are specified as a (3, image width, image height) tuple. If one has RGB input data as a volumetric scan with volume width, volume height and volume depth representing the 3 axes, the input data format would be specified by a tuple of 4 values (3, volume width, volume height, volume depth). In this way CNTK enables specification of input images in arbitrary higher-dimensional space.
# +
# Ensure we always get the same amount of randomness
np.random.seed(0)
C.cntk_py.set_fixed_random_seed(1)
C.cntk_py.force_deterministic_algorithms()
# Define the data dimensions
input_dim_model = (1, 28, 28) # images are 28 x 28 with 1 channel of color (gray)
input_dim = 28*28 # used by readers to treat input data as a vector
num_output_classes = 10
# -
# ## Data reading
#
# There are different ways one can read data into CNTK. The easiest way is to load the data in memory using NumPy / SciPy / Pandas readers. However, this can be done only for small data sets. Since deep learning requires large amount of data we have chosen in this course to show how to leverage built-in distributed readers that can scale to terrabytes of data with little extra effort.
#
# We are using the MNIST data you have downloaded using Lab 1 DataLoader notebook. The dataset has 60,000 training images and 10,000 test images with each image being 28 x 28 pixels. Thus the number of features is equal to 784 (= 28 x 28 pixels), 1 per pixel. The variable `num_output_classes` is set to 10 corresponding to the number of digits (0-9) in the dataset.
#
# In Lab 1, the data was downloaded and written to 2 CTF (CNTK Text Format) files, 1 for training, and 1 for testing. Each line of these text files takes the form:
#
# |labels 0 0 0 1 0 0 0 0 0 0 |features 0 0 0 0 ...
# (784 integers each representing a pixel)
#
# We are going to use the image pixels corresponding the integer stream named "features". We define a `create_reader` function to read the training and test data using the [CTF deserializer](https://cntk.ai/pythondocs/cntk.io.html?highlight=ctfdeserializer#cntk.io.CTFDeserializer). The labels are [1-hot encoded](https://en.wikipedia.org/wiki/One-hot). Refer to Lab 1 for data format visualizations.
# Read a CTF formatted text (as mentioned above) using the CTF deserializer from a file
def create_reader(path, is_training, input_dim, num_label_classes):
ctf = C.io.CTFDeserializer(path, C.io.StreamDefs(
labels=C.io.StreamDef(field='labels', shape=num_label_classes, is_sparse=False),
features=C.io.StreamDef(field='features', shape=input_dim, is_sparse=False)))
return C.io.MinibatchSource(ctf,
randomize = is_training, max_sweeps = C.io.INFINITELY_REPEAT if is_training else 1)
# +
# Ensure the training and test data is available
# We search in two locations in the toolkit for the cached MNIST data set.
data_found=False # A flag to indicate if train/test data found in local cache
for data_dir in [os.path.join("..", "Examples", "Image", "DataSets", "MNIST"),
os.path.join("data", "MNIST")]:
train_file=os.path.join(data_dir, "Train-28x28_cntk_text.txt")
test_file=os.path.join(data_dir, "Test-28x28_cntk_text.txt")
if os.path.isfile(train_file) and os.path.isfile(test_file):
data_found=True
break
if not data_found:
raise ValueError("Please generate the data by completing Lab1_MNIST_DataLoader")
print("Data directory is {0}".format(data_dir))
# -
# <a id='#Model Creation'></a>
# ## CNN Model Creation
#
# CNN is a feedforward network made up of bunch of layers in such a way that the output of one layer becomes the input to the next layer (similar to MLP). In MLP, all possible pairs of input pixels are connected to the output nodes with each pair having a weight, thus leading to a combinatorial explosion of parameters to be learnt and also increasing the possibility of overfitting ([details](http://cs231n.github.io/neural-networks-1/)). Convolution layers take advantage of the spatial arrangement of the pixels and learn multiple filters that significantly reduce the amount of parameters in the network ([details](http://cs231n.github.io/convolutional-networks/)). The size of the filter is a parameter of the convolution layer.
#
# In this section, we introduce the basics of convolution operations. We show the illustrations in the context of RGB images (3 channels), eventhough the MNIST data we are using a grayscale image (single channel).
#
# 
#
# ### Convolution Layer
#
# A convolution layer is a set of filters. Each filter is defined by a weight (**W**) matrix, and bias ($b$).
#
# 
#
# These filters are scanned across the image performing the dot product between the weights and corresponding input value (${x}$). The bias value is added to the output of the dot product and the resulting sum is optionally mapped through an activation function. This process is illustrated in the following animation.
Image(url="https://www.cntk.ai/jup/cntk103d_conv2d_final.gif", width= 300)
# Convolution layers incorporate the following key features:
#
# - Instead of being fully-connected to all input nodes , each convolution node is **locally-connected** to a subset of input nodes localized to a smaller input region, also referred to as receptive field (RF). The figure above illustrates a small 3 x 3 regions in the image as the RF region. In the case of an RGB, image there would be 3 such 3 x 3 regions, one for each of the 3 color channels.
#
#
# - Instead of having a single set of weights (as in a Dense layer), convolutional layers have multiple sets (shown in figure with multiple colors), called **filters**. Each filter detects features within each possible RF in the input image. The output of the convolution is a set of `n` sub-layers (shown in the animation below) where `n` is the number of filters (refer to the above figure).
#
#
# - Within a sublayer, instead of each node having its own set of weights, a single set of **shared weights** are used by all nodes in that sublayer. This reduces the number of parameters to be learnt and help reduce the risk of overfitting. This also opens the door for several aspects of deep learning which has enabled very practical solutions to be built:
# - Handling larger images (say 512 x 512)
# - Trying larger filter sizes (corresponding to a larger RF) say 11 x 11
# - Learning more filters (say 128)
# - Explore deeper architectures (100+ layers)
# - Achieve translation invariance (the ability to recognize a feature independent of where they appear in the image).
# ### Strides and Pad parameters
#
# **How are filters positioned?** In general, the filters are arranged in overlapping tiles, from left to right, and top to bottom. Each convolution layer has a parameter to specify the `filter_shape`, specifying the width and height of the filter. There is a parameter (`strides`) that controls the how far to step to right when moving the filters through multiple RF's in a row, and how far to step down when moving to the next row. The boolean parameter `pad` controls if the input should be padded around the edges to allow a complete tiling of the RF's near the borders.
#
# The animation above shows the results with a `filter_shape` = (3, 3), `strides` = (2, 2) and `pad` = False. The two animations below show the results when `pad` is set to True. First, with a stride of 2 and second having a stride of 1.
# Note: the shape of the output (the teal layer) is different between the two stride settings. In many problems, the stride and pad values are chosen to control the size of output layer.
# +
# Plot images with strides of 2 and 1 with padding turned on
images = [("https://www.cntk.ai/jup/cntk103d_padding_strides.gif" , 'With stride = 2'),
("https://www.cntk.ai/jup/cntk103d_same_padding_no_strides.gif", 'With stride = 1')]
for im in images:
print(im[1])
display(Image(url=im[0], width=200, height=200))
# -
# ## Building our CNN models
#
# We define two containers. One for the input MNIST image and the second one being the labels corresponding to the 10 digits. When reading the data, the reader automatically maps the 784 pixels per image to a shape defined by the `input_dim_model` tuple (in this example it is set to (1, 28, 28)).
x = C.input_variable(input_dim_model)
y = C.input_variable(num_output_classes)
# The first model we build is a simple convolution only network. Here we have two convolutional layers. Since, our task is to detect the 10 digits in the MNIST database, the output of the network should be a vector of length 10, 1 element corresponding to each output class. This is achieved by projecting the output of the last convolutional layer using a dense layer with the output being `num_output_classes`. We have seen this before with Logistic Regression and MLP where features were mapped to the number of classes in the final layer. Also, note that since we will be using the `softmax` operation that is combined with the `cross entropy` loss function during training (see a few cells below), the final dense layer has no activation function associated with it.
#
# The following figure illustrates the model we are going to build. Note the parameters in the model below are to be experimented with. These are often called network hyperparameters. Increasing the filter shape leads to an increase in the number of model parameters, increases the compute time and helps the model better fit to the data. However, one runs the risk of [overfitting](https://en.wikipedia.org/wiki/Overfitting). Typically, the number of filters in the deeper layers are more than the number of filters in the layers before them. We have chosen 8, 16 for the first and second layers, respectively. These hyperparameters should be experimented with during model building.
#
# 
# +
# function to build model
def create_model(features):
#with C.layers.default_options(init=C.glorot_uniform(), activation=C.sigmoid):
#with C.layers.default_options(init=C.glorot_uniform(), activation=C.tanh):
#with C.layers.default_options(init=C.glorot_uniform(), activation=C.leaky_relu):
with C.layers.default_options(init=C.glorot_uniform(), activation=C.relu):
h = features
h = C.layers.Convolution2D(filter_shape=(5,5),
num_filters=8,
strides=(2,2),
pad=True, name='first_conv')(h)
h = C.layers.Convolution2D(filter_shape=(5,5),
num_filters=16,
strides=(2,2),
pad=True, name='second_conv')(h)
r = C.layers.Dense(num_output_classes, activation=None, name='classify')(h)
return r
# -
# Let us create an instance of the model and inspect the different components of the model. `z` will be used to represent the output of a network. In this model, we use the `relu` activation function. Note: using the `C.layers.default_options` is an elegant and concise way to build models. This is key to minimizing modeling errors, saving precious debugging time.
# +
# Create the model
z = create_model(x)
# Print the output shapes / parameters of different components
print("Output Shape of the first convolution layer:", z.first_conv.shape)
print("Output Shape of the Second convolution layer:", z.second_conv.shape)
print("Output Shape of the Classify layer:", z.classify.shape)
print("Bias value of the last dense layer:", z.classify.b.value)
# -
# Understanding the number of learnable parameters in a model is key to deep learning since there is a dependency between the number of parameters and the amount of data one needs to have to train the model.
#
# You need more data for a model that has a larger number of parameters to prevent overfitting. In other words, with a fixed amount of data, one has to constrain the number of parameters. There is no golden rule between the amount of data one needs for a model. However, there are ways one can boost performance of model training with [data augmentation](https://deeplearningmania.quora.com/The-Power-of-Data-Augmentation-2).
# Number of parameters in the network
C.logging.log_number_of_parameters(z)
# **Understanding Parameters**:
#
#
# Our model has 2 convolution layers each having a weight and bias. This adds up to 4 parameter tensors. Additionally the dense layer has weight and bias tensors. Thus, the 6 parameter tensors.
#
# Remember that in a convolutional layer, the number of parameters is not dependent on the number of nodes, only on the shared weights and bias of each filter.
#
# Let us now count the number of parameters:
# - *First convolution layer*: There are 8 filters each of size (1 x 5 x 5) where 1 is the number of channels in the input image. This adds up to 200 values in the weight matrix and 8 bias values.
#
#
# - *Second convolution layer*: There are 16 filters each of size (8 x 5 x 5) where 8 is the number of channels in the input to the second layer (= output of the first layer). This adds up to 3200 values in the weight matrix and 16 bias values.
#
#
# - *Last dense layer*: There are 16 x 7 x 7 input values and it produces 10 output values corresponding to the 10 digits in the MNIST dataset. This corresponds to (16 x 7 x 7) x 10 weight values and 10 bias values.
#
# Adding these up gives the 11274 parameters in the model.
# **Knowledge Check**: Does the dense layer shape align with the task (MNIST digit classification)?
#
# ** Suggested Explorations **
# - Try printing shapes and parameters of different network layers,
# - Record the training error you get with `relu` as the activation function,
# - Now change to `sigmoid` as the activation function and see if you can improve your training error.
# - Different supported activation functions can be [found here][]. Which activation function gives the least training error?
#
# [found here]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
# ### Learning model parameters
#
# We use the `softmax` function to map the accumulated evidences or activations to a probability distribution over the classes (Details of the [softmax function][] and other [activation][] functions).
#
# [softmax function]: http://cntk.ai/pythondocs/cntk.ops.html#cntk.ops.softmax
#
# [activation]: https://github.com/Microsoft/CNTK/wiki/Activation-Functions
# ## Training
#
# We minimize the cross-entropy between the label and predicted probability by the network. Since we are going to build more than one model, we will create a few helper functions.
def create_criterion_function(model, labels):
loss = C.cross_entropy_with_softmax(model, labels)
errs = C.classification_error(model, labels)
return loss, errs # (model, labels) -> (loss, error metric)
# Next we will need a helper function to perform the model training. First let us create additional helper functions that will be needed to visualize different functions associated with training.
# +
# Define a utility function to compute the moving average sum.
# A more efficient implementation is possible with np.cumsum() function
def moving_average(a, w=5):
if len(a) < w:
return a[:] # Need to send a copy of the array
return [val if idx < w else sum(a[(idx-w):idx])/w for idx, val in enumerate(a)]
# Defines a utility that prints the training progress
def print_training_progress(trainer, mb, frequency, verbose=1):
training_loss = "NA"
eval_error = "NA"
if mb%frequency == 0:
training_loss = trainer.previous_minibatch_loss_average
eval_error = trainer.previous_minibatch_evaluation_average
if verbose:
print ("Minibatch: {0}, Loss: {1:.4f}, Error: {2:.2f}%".format(mb, training_loss, eval_error*100))
return mb, training_loss, eval_error
# -
# ### Configure training
#
# Previously we have described the concepts of `loss` function, the optimizers or [learners](https://cntk.ai/pythondocs/cntk.learners.html) and the associated machinery needed to train a model. Please refer to earlier labs for gaining familiarility with these concepts. Here we combine model training and testing in a helper function below.
#
def train_test(train_reader, test_reader, model_func, num_sweeps_to_train_with=10):
# Instantiate the model function; x is the input (feature) variable
# We will scale the input image pixels within 0-1 range by dividing all input value by 255.
model = model_func(x/255)
# Instantiate the loss and error function
loss, label_error = create_criterion_function(model, y)
# Instantiate the trainer object to drive the model training
learning_rate = 0.1
lr_schedule = C.learning_rate_schedule(learning_rate, C.UnitType.minibatch)
learner = C.sgd(z.parameters, lr_schedule)
trainer = C.Trainer(z, (loss, label_error), [learner])
# Initialize the parameters for the trainer
#minibatch_size = 64
minibatch_size = 16
num_samples_per_sweep = 60000
num_minibatches_to_train = (num_samples_per_sweep * num_sweeps_to_train_with) / minibatch_size
# Map the data streams to the input and labels.
input_map={
y : train_reader.streams.labels,
x : train_reader.streams.features
}
# Uncomment below for more detailed logging
training_progress_output_freq = 500
# Start a timer
start = time.time()
for i in range(0, int(num_minibatches_to_train)):
# Read a mini batch from the training data file
data=train_reader.next_minibatch(minibatch_size, input_map=input_map)
trainer.train_minibatch(data)
print_training_progress(trainer, i, training_progress_output_freq, verbose=1)
# Print training time
print("Training took {:.1f} sec".format(time.time() - start))
# Test the model
test_input_map = {
y : test_reader.streams.labels,
x : test_reader.streams.features
}
# Test data for trained model
test_minibatch_size = 512
num_samples = 10000
num_minibatches_to_test = num_samples // test_minibatch_size
test_result = 0.0
for i in range(num_minibatches_to_test):
# We are loading test data in batches specified by test_minibatch_size
# Each data point in the minibatch is a MNIST digit image of 784 dimensions
# with one pixel per dimension that we will encode / decode with the
# trained model.
data = test_reader.next_minibatch(test_minibatch_size, input_map=test_input_map)
eval_error = trainer.test_minibatch(data)
test_result = test_result + eval_error
# Average of evaluation errors of all test minibatches
print("Average test error: {0:.2f}%".format(test_result*100 / num_minibatches_to_test))
# <a id='#Run the trainer'></a>
# ### Run the trainer and test model
#
# We are now ready to train our convolutional neural net.
# +
def do_train_test():
global z
z = create_model(x)
reader_train = create_reader(train_file, True, input_dim, num_output_classes)
reader_test = create_reader(test_file, False, input_dim, num_output_classes)
train_test(reader_train, reader_test, z)
# Only For part 1, for part 2 use the network with padding below
# do_train_test()
# -
# Note, the average test error is very comparable to our training error indicating that our model has good "out of sample" error a.k.a. [generalization error](https://en.wikipedia.org/wiki/Generalization_error). This implies that our model can very effectively deal with previously unseen observations (during the training process). This is key to avoid [overfitting](https://en.wikipedia.org/wiki/Overfitting).
#
# Let us check what is the value of some of the network parameters. We will check the bias value of the output dense layer. Previously, it was all 0. Now you see there are non-zero values, indicating that a model parameters were updated during training.
# +
# Works w only if we invoke do_train_test() in the above cell
#print("Bias value of the last dense layer:", z.classify.b.value)
# -
# ## Evaluation / Prediction
# We have so far been dealing with aggregate measures of error. Let us now get the probabilities associated with individual data points. For each observation, the `eval` function returns the probability distribution across all the classes. The classifier is trained to recognize digits, hence has 10 classes. First let us route the network output through a `softmax` function. This maps the aggregated activations across the network to probabilities across the 10 classes.
out = C.softmax(z)
# Let us test a small minibatch sample from the test data.
# +
# Read the data for evaluation
reader_eval=create_reader(test_file, False, input_dim, num_output_classes)
eval_minibatch_size = 25
eval_input_map = {x: reader_eval.streams.features, y:reader_eval.streams.labels}
data = reader_eval.next_minibatch(eval_minibatch_size, input_map=eval_input_map)
img_label = data[y].asarray()
img_data = data[x].asarray()
# reshape img_data to: M x 1 x 28 x 28 to be compatible with model
img_data = np.reshape(img_data, (eval_minibatch_size, 1, 28, 28))
predicted_label_prob = [out.eval(img_data[i]) for i in range(len(img_data))]
# -
# Find the index with the maximum value for both predicted as well as the ground truth
pred = [np.argmax(predicted_label_prob[i]) for i in range(len(predicted_label_prob))]
gtlabel = [np.argmax(img_label[i]) for i in range(len(img_label))]
print("Label :", gtlabel[:25])
print("Predicted:", pred)
# Let us visualize some of the results
# +
# Plot a random image
sample_number = 5
plt.imshow(img_data[sample_number].reshape(28,28), cmap="gray_r")
plt.axis('off')
img_gt, img_pred = gtlabel[sample_number], pred[sample_number]
print("Image Label: ", img_pred)
# -
# ## Pooling Layer
#
# Often times, one needs to control the number of parameters especially when having deep networks. For every layer of the convolution layer output (each layer, corresponds to the output of a filter), one can have a pooling layer. Pooling layers are typically introduced to:
# - Reduce the shape of the current layer (speeding up the network),
# - Make the model more tolerant to changes in object location in the image. For example, even when a digit is shifted to one side of the image instead of being in the middle, the classifer would perform the classification task well.
#
# The calculation on a pooling node is much simpler than a normal feedforward node. It has no weight, bias, or activation function. It uses a simple aggregation function (like max or average) to compute its output. The most commonly used function is "max" - a max pooling node simply outputs the maximum of the input values corresponding to the filter position of the input. The figure below shows the input values in a 4 x 4 region. The max pooling window size is 2 x 2 and starts from the top left corner, and uses a stride of 2x2. The maximum value within the window becomes the output of the region. Every time the model is shifted by the amount specified by the stride parameter (as shown in the figure below) and the maximum pooling operation is repeated.
# 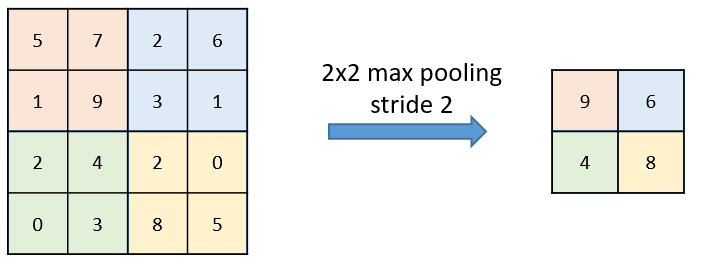
# Another alternative is average pooling, which emits that average value instead of the maximum value. The two different pooling opearations are summarized in the animation below.
# +
# Plot images with strides of 2 and 1 with padding turned on
images = [("https://www.cntk.ai/jup/c103d_max_pooling.gif" , 'Max pooling'),
("https://www.cntk.ai/jup/c103d_average_pooling.gif", 'Average pooling')]
for im in images:
print(im[1])
display(Image(url=im[0], width=200, height=200))
# -
# # Typical convolution network
#
# 
#
# A typical CNN contains a set of alternating convolution and pooling layers followed by a dense output layer for classification. You will find variants of this structure in many classical deep networks (VGG, AlexNet etc). This is in contrast to the MLP network we used in Lab 3, which consisted of 2 dense layers followed by a dense output layer.
#
# The illustrations are presented in the context of 2-dimensional (2D) images, but the concept and the CNTK components can operate on any dimensional data. The above schematic shows 2 convolution layer and 2 pooling layers. A typical strategy is to increase the number of filters in the deeper layers while reducing the spatial size of each intermediate layers.
# ## Task: Create a network with Average Pooling
#
# Typical convolutional networks have interlacing convolution and pooling layers. The previous model had only convolution layer. In this section, you will create a model with the following architecture.
#
# 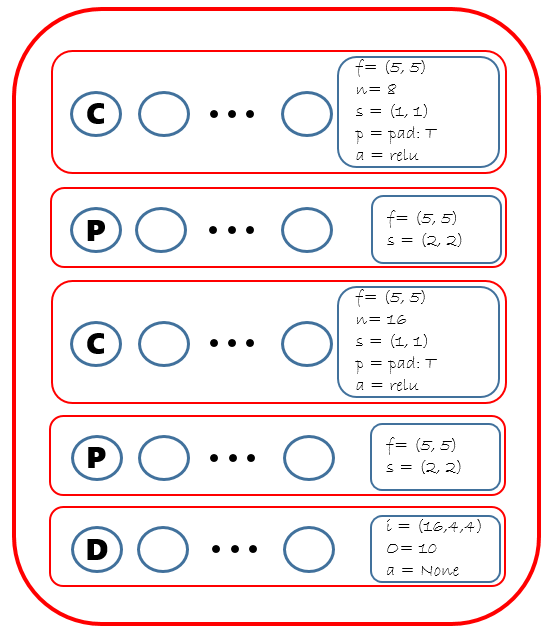
#
# You will use the CNTK [Average Pooling](https://cntk.ai/pythondocs/cntk.layers.layers.html#cntk.layers.layers.AveragePooling) function to achieve this task. You will edit the `create_model` function below and add the Average Pooling operation.
#
#
# +
# Modify this model
def create_model(features):
#with C.layers.default_options(init = C.glorot_uniform(), activation = C.relu):
with C.layers.default_options(init = C.glorot_uniform(), activation = C.leaky_relu):
#with C.layers.default_options(init = C.glorot_uniform(), activation = C.sigmoid):
#with C.layers.default_options(init = C.glorot_uniform(), activation = C.tanh):
h = features
h = C.layers.Convolution2D(filter_shape=(5,5),
num_filters=8,
strides=(1,1),
pad=True, name='first_conv')(h)
h = C.layers.AveragePooling(filter_shape = (5, 5), strides = (2, 2))(h)
#h = C.layers.MaxPooling(filter_shape = (5, 5), strides = (2, 2))(h)
h = C.layers.Convolution2D(filter_shape=(5,5),
num_filters=16,
strides=(1,1),
pad=True, name='second_conv')(h)
h = C.layers.AveragePooling(filter_shape = (5, 5), strides = (2, 2))(h)
#h = C.layers.MaxPooling(filter_shape = (5, 5), strides = (2, 2))(h)
r = C.layers.Dense(num_output_classes, activation = None, name='classify')(h)
return r
#Uncommented after the last one without pooling is commented out
do_train_test()
# Average Test Error, with activation as Sigmoid is 3.96%
# Average Test Error, with Activation as Tanh is 1.58 %
# Average Test Error, with activation as Relu is 1.20 %
# Average test Error with activation as leaky relu is 1.12 %
# -
# **Knowledge Check**: How many parameters do we have in this second model (as shown in the figure)?
#
#
# **Suggested Explorations**
# - Add average pooling layer after each of the two convolution layer. Use the parameters as shown in the figure.
# - Does use of LeakyRelu help improve the error rate?
# - What percentage of the parameter does the last dense layer contribute w.r.t. the overall number of parameters for (a) purely two convolutional layer and (b) alternating 2 convolutional and average pooling layers
C.logging.log_number_of_parameters(create_model(x))
from PIL import Image
in_image = np.array(Image.open('MysteryNumberD.bmp').convert('L'))
print("Predicted number is " + str(np.argmax(z.eval(in_image.reshape(1, 28, 28) / 255))))
plt.imshow(in_image, cmap = 'gray')
| 5 - Deep Learning Explained/hw3/Lab4_ConvolutionalNeuralNetwork.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # `truevoice-intent` Dataset
# This notebook performs a preliminary exploration of the `truevoice-intent` dataset which was provided by [TrueVoice](http://www.truevoice.co.th/) through Nattapote Kuslasayanon. The texts are transcribed from customer service phone calls to a mobile phone service provider using Truevoice's [Mari](http://www.truevoice.co.th/en/true-voice-mari/). This dataset is a part of [pyThaiNLP](https://github.com/PyThaiNLP/) Thai text [classification-benchmarks](https://github.com/PyThaiNLP/classification-benchmarks). `texts` column contains raw texts and `texts_deepcut` column contains those segmented by [deepcut](https://github.com/rkcosmos/deepcut).
#
# The benchmark features a set of **three multi-class classification tasks** for `action`, `object`, and `destination` of all the calls. Performance metrics are macro- and micro-averaged accuracy, F1 score, precision and recall.
# +
import pandas as pd
import numpy as np
from pythainlp import word_tokenize
from ast import literal_eval
from tqdm import tqdm_notebook
from collections import Counter
#viz
from plotnine import *
import matplotlib.pyplot as plt
import seaborn as sns
# +
# #snippet to install thai font in matplotlib from https://gist.github.com/korakot/9d7f5db632351dc92607fdec72a4953f
import matplotlib
# # !wget https://github.com/Phonbopit/sarabun-webfont/raw/master/fonts/thsarabunnew-webfont.ttf
# # !cp thsarabunnew-webfont.ttf /usr/local/lib/python3.6/dist-packages/matplotlib/mpl-data/fonts/ttf/
# # !cp thsarabunnew-webfont.ttf /usr/share/fonts/truetype/
matplotlib.font_manager._rebuild()
matplotlib.rc('font', family='TH Sarabun New')
# -
# ## Data Cleaning
# From the original `.xlsx` files we cleaned the datasets and saved them as `.csv`.
# +
# all_df = pd.read_excel('mari_deepcut_train.xlsx')
# test_df = pd.read_excel('mari_deepcut_test.xlsx')
# #select only relevant columns
# selected = ['Sentence Utterance', 'Sentence Utterance (deep cut)',
# 'Action', 'Object', 'Destination']
# all_df = all_df[selected]
# test_df = test_df[selected]
# all_df.columns = ['texts', 'texts_deepcut', 'action', 'object', 'destination']
# test_df.columns = ['texts', 'texts_deepcut', 'action', 'object', 'destination']
# for i in range(2,5):
# all_df.iloc[:,i] = all_df.iloc[:,i].map(lambda x: x.lower())
# test_df.iloc[:,i] = test_df.iloc[:,i].map(lambda x: x.lower())
# all_df.to_csv('mari_train.csv',index=False)
# test_df.to_csv('mari_test.csv',index=False)
# -
# ## Load Data
all_df = pd.read_csv('mari_train.csv')
print(all_df.shape)
all_df.tail()
# ## Labels
# ### Label Distribution
# +
action_labels = pd.DataFrame(all_df.action.value_counts()).reset_index()
action_labels.columns = ['label','cnt']
action_labels['per'] = action_labels.cnt / np.sum(action_labels.cnt)
action_labels = action_labels.sort_values('cnt',ascending=False)
for i,row in action_labels.iterrows():
action_labels.iloc[i,0] = f"{i}_{row['label']}"
print(action_labels.shape)
g = (ggplot(action_labels,aes(x='label', y='per', fill='label')) + geom_bar(stat='identity') +
theme_minimal() + scale_y_continuous(labels = lambda x: np.round(x*100,0)) +
xlab('Action Labels') + ylab('Share of Samples (%)') +
geom_text(aes(x='label',y='per+0.05',label='round(per*100,1)')) +
theme(axis_text_x = element_blank(),legend_title=element_blank()))
g
# +
object_labels = pd.DataFrame(all_df.object.value_counts()).reset_index()
object_labels.columns = ['label','cnt']
object_labels['per'] = object_labels.cnt / np.sum(object_labels.cnt)
object_labels = object_labels.sort_values('cnt',ascending=False)
for i,row in object_labels.iterrows():
object_labels.iloc[i,0] = f"{i:02}_{row['label']}"
print(object_labels.shape)
g = (ggplot(object_labels,aes(x='label', y='per', fill='label')) + geom_bar(stat='identity') +
theme_minimal() + scale_y_continuous(labels = lambda x: np.round(x*100,0)) +
xlab('Object Labels') + ylab('Share of Samples (%)') +
geom_text(aes(x='label',y='per+0.01',label='round(per*100,1)',size=0.7)) +
theme(axis_text_x = element_blank(),legend_title=element_blank(),legend_position='bottom'))
g
# +
destination_labels = pd.DataFrame(all_df.destination.value_counts()).reset_index()
destination_labels.columns = ['label','cnt']
destination_labels['per'] = destination_labels.cnt / np.sum(destination_labels.cnt)
destination_labels = destination_labels.sort_values('cnt',ascending=False)
for i,row in destination_labels.iterrows():
destination_labels.iloc[i,0] = f"{i}_{row['label']}"
print(destination_labels.shape)
g = (ggplot(destination_labels,aes(x='label', y='per', fill='label')) + geom_bar(stat='identity') +
theme_minimal() + scale_y_continuous(labels = lambda x: np.round(x*100,0)) +
xlab('Destination Labels') + ylab('Share of Samples (%)') +
geom_text(aes(x='label',y='per+0.05',label='round(per*100,1)')) +
theme(axis_text_x = element_blank(),legend_title=element_blank()))
g
# -
# ### Label Association
all_df[['action','destination']].pivot_table(index='action',
columns='destination',
aggfunc=len, fill_value=0)
all_df[['object','action']].pivot_table(index='object',
columns='action',
aggfunc=len, fill_value=0)
all_df[['object','destination']].pivot_table(index='object',
columns='destination',
aggfunc=len, fill_value=0)
# ## Texts
from sklearn.feature_extraction.text import CountVectorizer
texts_cnt = CountVectorizer(tokenizer=lambda x: x.split())
texts_mat = texts_cnt.fit_transform(all_df.texts_deepcut)
texts_mat.shape
# ### Word Count Distribution
texts_wc = pd.DataFrame(texts_mat.sum(axis=1))
texts_wc.columns = ['word_count']
g = (ggplot(texts_wc, aes(x='word_count')) + geom_histogram(bins=30) + theme_minimal() +
xlab('Text Word Count') + ylab('Number of Samples') +
geom_vline(xintercept = np.mean(texts_wc.word_count), color='red'))
print(f'Average Text Word Count: {np.mean(texts_wc.word_count)}')
g
all_df['wc'] = texts_wc['word_count']
action_agg = all_df[['action','wc']].groupby('action').mean().reset_index()\
.sort_values('wc',ascending=False)
for i,row in action_agg.iterrows():
action_agg.iloc[i,0] = f"{i:02}_{row['action']}"
g = (ggplot(action_agg,aes(x='action', y='wc', fill='action')) + geom_bar(stat='identity') +
theme_minimal() + scale_y_continuous(labels = lambda x: np.round(x*100,0)) +
xlab('Action Labels') + ylab('Mean Word Count') +
geom_text(aes(x='action',y='wc+1',label='round(wc)')) +
theme(axis_text_x = element_blank(),legend_title=element_blank()))
g
all_df['wc'] = texts_wc['word_count']
object_agg = all_df[['object','wc']].groupby('object').mean().reset_index()\
.sort_values('wc',ascending=False)
for i,row in object_agg.iterrows():
object_agg.iloc[i,0] = f"{i:02}_{row['object']}"
g = (ggplot(object_agg,aes(x='object', y='wc', fill='object')) + geom_bar(stat='identity') +
theme_minimal() + scale_y_continuous(labels = lambda x: np.round(x*100,0)) +
xlab('Object Labels') + ylab('Mean Word Count') +
geom_text(aes(x='object',y='wc+1',label='round(wc)'), size = 7) +
theme(axis_text_x = element_blank(),legend_title=element_blank(),legend_position='bottom'))
g
all_df['wc'] = texts_wc['word_count']
destination_agg = all_df[['destination','wc']].groupby('destination').mean().reset_index()\
.sort_values('wc',ascending=False)
for i,row in destination_agg.iterrows():
destination_agg.iloc[i,0] = f"{i:02}_{row['destination']}"
g = (ggplot(destination_agg,aes(x='destination', y='wc', fill='destination')) + geom_bar(stat='identity') +
theme_minimal() + scale_y_continuous(labels = lambda x: np.round(x*100,0)) +
xlab('Destination Labels') + ylab('Mean Word Count') +
geom_text(aes(x='destination',y='wc+1',label='round(wc)')) +
theme(axis_text_x = element_blank(),legend_title=element_blank()))
g
# ### Word Frequency
texts_top = pd.DataFrame({
'vocab': list(texts_cnt.get_feature_names()),
'cnt': np.asarray(texts_mat.sum(axis=0)).squeeze()}).sort_values('cnt',ascending=False)
texts_top = texts_top.reset_index(drop=True).reset_index()
g = (ggplot(texts_top.iloc[:500,:], aes(x='index',y='cnt')) +
geom_bar(stat='identity') +
theme_minimal() +
xlab('Word Frequency') + ylab('Number of Words') +
geom_vline(xintercept = np.mean(texts_top.cnt), color='red'))
print(f'Mean Word Frequency: {np.mean(texts_top.cnt)}')
g
texts_top.head(10)
texts_top.tail(10)
# ## Text Feature Correlations
from class_features import *
from sklearn.feature_extraction.text import TfidfVectorizer
show_classfeats(
df=all_df,
vectorizer=TfidfVectorizer,
analyzer=lambda x: x.split(),
text_col='texts_deepcut',
class_col='action'
)
show_classfeats(
df=all_df,
vectorizer=TfidfVectorizer,
analyzer=lambda x: x.split(),
text_col='texts_deepcut',
class_col='object',
nrow=7,
ncol=4
)
show_classfeats(
df=all_df,
vectorizer=TfidfVectorizer,
analyzer=lambda x: x.split(),
text_col='texts_deepcut',
class_col='destination'
)
| exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Grid Search (Mitigation Algorithm) with Adult/Census Data
# Census dataset is used to predict if an individual's income is below or above 50k per year.
# +
from fairlearn.widget import FairlearnDashboard
from sklearn.model_selection import train_test_split
from fairlearn.reductions import GridSearch
from fairlearn.reductions import DemographicParity, ErrorRate
from sklearn.preprocessing import LabelEncoder, StandardScaler
from sklearn.linear_model import LogisticRegression
import pandas as pd
from sklearn.datasets import fetch_openml
# -
data = fetch_openml(data_id=1590, as_frame=True)
X_raw = data.data
Y = (data.target == '>50K') * 1
X_raw
Y
# +
#in this setup, gender is the sensitive data and we are dropping that column from the dataset.
A = X_raw["sex"]
X = X_raw.drop(labels=['sex'], axis=1)
X = pd.get_dummies(X)
sc = StandardScaler()
X_scaled = sc.fit_transform(X)
X_scaled = pd.DataFrame(X_scaled, columns=X.columns)
le = LabelEncoder()
Y = le.fit_transform(Y)
Y
# +
#train-test split
X_train, X_test, Y_train, Y_test, A_train, A_test = train_test_split(X_scaled,
Y,
A,
test_size=0.2,
random_state=0,
stratify=Y)
# Work around indexing bug
X_train = X_train.reset_index(drop=True)
A_train = A_train.reset_index(drop=True)
X_test = X_test.reset_index(drop=True)
A_test = A_test.reset_index(drop=True)
# -
# ### Training a fairness-unaware predictor
#logistic regression
unmitigated_predictor = LogisticRegression(solver='liblinear', fit_intercept=True)
unmitigated_predictor.fit(X_train, Y_train)
#Display of dashboard to investigate disparity
FairlearnDashboard(sensitive_features=A_test, sensitive_feature_names=['sex'],
y_true=Y_test,
y_pred={"unmitigated": unmitigated_predictor.predict(X_test)})
# __Result:__ Despite the fact that we removed the feature from the training data, our predictor still discriminates based on sex. This demonstrates that simply ignoring a sensitive feature when fitting a predictor rarely eliminates unfairness. There will generally be enough other features correlated with the removed feature to lead to disparate impact.
#
# ### Disparity mitigation with GridSearch algorithm
# The user supplies a standard ML estimator to this algorithm, which is treated as a blackbox. GridSearch works by generating a sequence of relabellings and reweightings, and trains a predictor for each. Fairness metric is chosen as _demographic parity_.
# +
#takes ~5 mins to run
sweep = GridSearch(LogisticRegression(solver='liblinear', fit_intercept=True),
constraints=DemographicParity(),
grid_size=71)
#grid size gives the number of predictors calculated (The number of Lagrange multipliers to generate in the grid)
sweep.fit(X_train, Y_train,
sensitive_features=A_train)
predictors = sweep.predictors_
predictors
# -
# We could load these predictors into the Fairness dashboard now. However, the plot would be somewhat confusing due to their number. In this case, we are going to remove the predictors which are dominated in the error-disparity space by others from the sweep (note that the disparity will only be calculated for the sensitive feature; other potentially sensitive features will not be mitigated). In general, one might not want to do this, since there may be other considerations beyond the strict optimization of error and disparity (of the given sensitive feature). After eliminating the _dominated_ models, we can put the _dominant_ models into the Fairness dashboard, along with the unmitigated model.
# +
errors, disparities = [], []
for m in predictors:
def classifier(X): return m.predict(X)
error = ErrorRate()
#load_data loads the specified data into the object.
error.load_data(X_train, pd.Series(Y_train), sensitive_features=A_train)
disparity = DemographicParity()
disparity.load_data(X_train, pd.Series(Y_train), sensitive_features=A_train)
errors.append(error.gamma(classifier)[0])
disparities.append(disparity.gamma(classifier).max())
all_results = pd.DataFrame({"predictor": predictors, "error": errors, "disparity": disparities})
all_results
# -
non_dominated = []
#itertuples() method will return an iterator yielding a named tuple for each row in the DataFrame. The first element of the tuple will be the row’s corresponding index value, while the remaining values are the row values.
for row in all_results.itertuples():
errors_for_lower_or_eq_disparity = all_results["error"][all_results["disparity"] <= row.disparity]
if row.error <= errors_for_lower_or_eq_disparity.min():
non_dominated.append(row.predictor)
non_dominated
# +
dashboard_predicted = {"unmitigated": unmitigated_predictor.predict(X_test)}
for i in range(len(non_dominated)):
key = "dominant_model_{0}".format(i)
value = non_dominated[i].predict(X_test)
dashboard_predicted[key] = value
FairlearnDashboard(sensitive_features=A_test, sensitive_feature_names=['sex'],
y_true=Y_test,
y_pred=dashboard_predicted)
# +
#check the difference between the feature coefficients assigned by the GridSearch algorithm
print(non_dominated[0].coef_)
print(non_dominated[2].coef_)
| Microfost Fairlearn 2 (Adult-Census dataset using GridSearch Algorithm).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
from scipy import stats
import pandas as pd
import matplotlib.pyplot as plt
# ### Python recipe for performing the 1-sample t-test
#
# 
#
# ### Error bar represents standard deviation
# +
# we will draw numbers from a random normal distribution - set mean and standard deviation
mymean=FILL
mystd=FILL
myN=FILL
# if you want to draw the same random numbers or compare with a friend then
# set the random number seed the same
np.random.seed(12345)
# make sure you understand all the arguments
myrand=np.random.normal(loc=mymean,scale=mystd,size=myN)
# just print out some basic descriptive stats
print myrand
print myrand.mean()
print myrand.std(ddof=1)
# this is the s.e.m. - showed formula in class
print stats.sem(myrand,ddof=1)
# +
# here is the hand calculation for the t-score, based on our formula
t=(myrand.mean()-mymean)/stats.sem(myrand)
print t
# +
# the function stats.t.sf is the lookup value - given a t-score it will
# provide you the 1-sided p-value. Feed the abs value of t-score and multiple by 2
# to obtain the standard 2-sided p-value!
stats.t.sf(np.abs(t),myrand.size-1)*2
# +
# you can also just use this function to directly output the t/p values
stats.ttest_1samp(myrand,mymean)
# +
# and this is how you store the output in two separate variables
[tcalc,p]=stats.ttest_1samp(myrand,mymean)
# -
# ## Calculation of 95% CI manually and with python tools
#
# The CI is given by: $ \bar x \pm t^* * s.e.m.$
#
# * _t*_ is the critical t-value at a given combination of P and (N-1)
# * Take care with 1-sided vs. 2-sided. For the majority of examples you will want to use 2-sided
## Inverse lookup of critical t value at P=0.05 (2-sided) for our system above
tcrit=stats.t.ppf(1-.025,myN-1)
print tcrit
## calculation of the 95% CI w/above formula
my95CI=stats.sem(myrand)*tcrit
print my95CI
## we can use a python stats tool to do this dirty work for us
## usage:
## stats.t.interval($CI, d.o.f., loc=mean, scale=sem)
## the output are the Lower and Upper bounds of the 95% CI.
[L,U]=stats.t.interval(.95,myN-1,loc=myrand.mean(), scale=stats.sem(myrand))
print L,U
print (U-L)/2
## Let me convince you my hand calc matches the stats tool
print myrand.mean()-(U-L)/2
print myrand.mean()-my95CI
| 2017/DSMCER_content/PValuesW4L1/W4L1skel.ipynb |