code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# language: python
# name: python37364bit4db73d59933341feaa47a8db6a2db2c7
# ---
import pandas as pd
import numpy as np
import lib.data_processing
import importlib
importlib.reload(lib.data_processing)
TRAIN_FP = 'data/bias_data/WNC/biased.word.train'
TEST_FP = 'data/bias_data/WNC/biased.word.test'
# +
wnc_train = lib.data_processing.raw_data(TRAIN_FP, 3, 4)
wnc_train_df = wnc_train.add_miss_word_col(dtype='df')
wnc_test = lib.data_processing.raw_data(TEST_FP, 3, 4)
wnc_test_df = wnc_test.add_miss_word_col(dtype='df')
# -
wnc_test_df.head(5)
sample = lib.data_processing.raw_data('data/bias_data/real_world_samples/ibc_right', 2, 3)
sample.add_miss_word_col(dtype='df').head(5)
| WNC Data Pull.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.12 64-bit (''onnx'': conda)'
# name: python3
# ---
# <a id='StartingPoint'></a>
# # ONNX classification example
#
# Sharing DL models between frameworks or programming languages is possible with Open Neural Network Exchange (ONNX for short).
#
# This notebook starts from an onnx model exported from MATLAB and uses it in Python.
# On MATLAB a GoogleNet model pre-trained on ImageNet was loaded and saved to onnx file format through a one-line command [exportONNXNetwork(net,filename)](https://www.mathworks.com/help/deeplearning/ref/exportonnxnetwork.html).
#
# The model is then loaded here, as well as the data to evaluate (some images retrieved from google).
# Images are preprocessed to the desired format np.array with shape (BatchSize, numChannels, width, heigh), and the model is applied to classify the images and get the probabilities of the classifications.
# ## Input Variables
#
# The why for the existence of each variable is explained [bellow](#ModelDef), near the model definition.
# +
# %%time
# model vars
modelPath = 'D:/onnxStartingCode/model/preTrainedImageNetGooglenet.onnx'
labelsPath = 'D:/onnxStartingCode/model/labels.csv'
hasHeader = 1
#data vars
image_folder = 'D:/onnxStartingCode/ImageFolder/'
EXT = ("jfif","jpg")
# -
# ## Import Modules
#
# Let's start by importing all the needed modules
#
# [back to top](#StartingPoint)
# %%time
import onnx
import numpy as np
from PIL import Image
import os as os
import matplotlib.pyplot as plt
from onnxruntime import InferenceSession
import csv
# <a id='ModelDef'></a>
# ## Define Model and Data functions
#
# We need to define functions to retrieve the Classifier and Data array.
#
# To load the model we need the path to the file that stores it and the pat to the file that stores the labels. Finnaly, the parameter hasHeader defines the way the firs row of the labelsFile is treated, as header or ar a label.
# The labelsPath is required here becuase the model here used does not contain label information, so an external csv file needs to be read.
#
# [back to top](#StartingPoint)
# +
# %%time
def loadmodel(modelPath,labelsPath,hasHeader):
# define network
# load and check the model
# load the inference module
onnx.checker.check_model(modelPath)
sess = InferenceSession(modelPath)
# Determine the name of the input and output layers
inname = [input.name for input in sess.get_inputs()]
outname = [output.name for output in sess.get_outputs()]
# auxiliary function to load labels file
def extractLabels( filename , hasHeader ):
file = open(filename)
csvreader = csv.reader(file)
if (hasHeader>0):
header = next(csvreader)
#print(header)
rows = []
for row in csvreader:
rows.append(row)
#print(rows)
file.close()
return rows
# Get labels
labels = extractLabels(labelsPath,hasHeader)
# Extract information on the inputSize =(width, heigh) and numChannels = 3(RGB) or 1(Grayscale)
for inp in sess.get_inputs():
inputSize = inp.shape
numChannels = inputSize[1]
inputSize = inputSize[2:4]
return sess,inname,outname,numChannels,inputSize,labels
def getData(image_folder,EXT,inputSize):
def getImagesFromFolder(EXT):
imageList = os.listdir(image_folder)
if (not(isinstance(EXT, list)) and not(isinstance(EXT,tuple))):
ext = [EXT]
fullFilePath = [os.path.join(image_folder, f)
for ext in EXT for f in imageList if os.path.isfile(os.path.join(image_folder, f)) & f.endswith(ext)]
return fullFilePath
def imFile2npArray(imFile,inputSize):
data = np.array([
np.array(
Image.open(fname).resize(inputSize),
dtype=np.int64)
for fname in fullFilePath
])
X=data.transpose(0,3,1,2)
X = X.astype(np.float32)
return X, data
fullFilePath = getImagesFromFolder(EXT)
X, data = imFile2npArray(fullFilePath,inputSize)
return X,data,fullFilePath
# -
# ## Run loading functions to get model and data
#
# * get full filename of all files in a gives directory that end with a given ext (might be an array of EXTENSIONS)
# * load data into numpy arrays for future use:
# * to plot data has to have shape = (x,y,3)
# * the model here presented requires data with shape (3,x,y)
# * two data arrays are then exported, data for ploting and X for classification
#
# [back to top](#StartingPoint)
# +
# %%time
# run code
sess,inname,outname,numChannels,inputSize,labels = loadmodel(modelPath,labelsPath,hasHeader)
X,data,fullFilePath = getData(image_folder,EXT,inputSize)
print("inputSize: " + str(inputSize))
print("numChannels: " + str(numChannels))
print("inputName: ", inname[0])
print("outputName: ", outname[0])
# -
# ## Define a functions to load all data
#
# 1. get full filename of all files in a gives directory that end with a given ext (might be an arrat of EXT)
# 2. load data into numpy arrays for future use:
# * to plot data has to have shape = (x,y,3)
# * the model here presented requires data with shape (3,x,y)
# * two data arrays are then exported, data for ploting and X for classification
#
# [back to top](#StartingPoint)
# ## Classification
#
# [back to top](#StartingPoint)
# +
# %%time
#data_output = sess.run(outname, {inname: X[0]})
out = sess.run(None, {inname[0]: X})
out=np.asarray(out[0])
print(out.shape)
IND = []
PROB= []
for i in range(out.shape[0]):
ind=np.where(out[i] == np.amax(out[i]))
IND.append(ind[0][0])
PROB.append(out[i,ind[0][0]])
l = [labels[ind] for ind in IND]
print([labels[ind] for ind in IND])
print(IND)
print(PROB)
# -
# ## Plot some examples
#
# [back to top](#StartingPoint)
# +
# %%time
plt.figure(figsize=(10,10))
if data.shape[0]>=6:
nPlots=6
subArray=[2,3]
else:
nPlots=data.shape[0]
subArray = [1, nPlots]
for i in range(nPlots):
plt.subplot(subArray[0],subArray[1],i+1)
plt.imshow(data[i])
plt.axis('off')
plt.title(l[i][0] + ' --- ' + str(round(100*PROB[i])) + '%')
plt.show()
# -
[back to top](#StartingPoint)
| ONNXclassifier/onnxClassify.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 1
#
# # Used Vehicle Price Prediction
# ## Introduction
#
# - 1.2 Million listings scraped from TrueCar.com - Price, Mileage, Make, Model dataset from Kaggle: [data](https://www.kaggle.com/jpayne/852k-used-car-listings)
# - Each observation represents the price of an used car
# %matplotlib inline
import pandas as pd
data = pd.read_csv('https://github.com/albahnsen/PracticalMachineLearningClass/raw/master/datasets/dataTrain_carListings.zip')
data.head()
data.shape
data.Price.describe()
data.plot(kind='scatter', y='Price', x='Year')
data.plot(kind='scatter', y='Price', x='Mileage')
data.columns
# # Exercise P1.1 (50%)
#
# Develop a machine learning model that predicts the price of the of car using as an input ['Year', 'Mileage', 'State', 'Make', 'Model']
#
# Submit the prediction of the testing set to Kaggle
#
# #### Evaluation:
# - 25% - Performance of the model in the Kaggle Private Leaderboard
# - 25% - Notebook explaining the modeling process
#
data_test = pd.read_csv('https://github.com/albahnsen/PracticalMachineLearningClass/raw/master/datasets/dataTest_carListings.zip', index_col=0)
data_test.head()
data_test.shape
# # Exercise P1.2 (50%)
#
# Create an API of the model.
#
# Example:
# 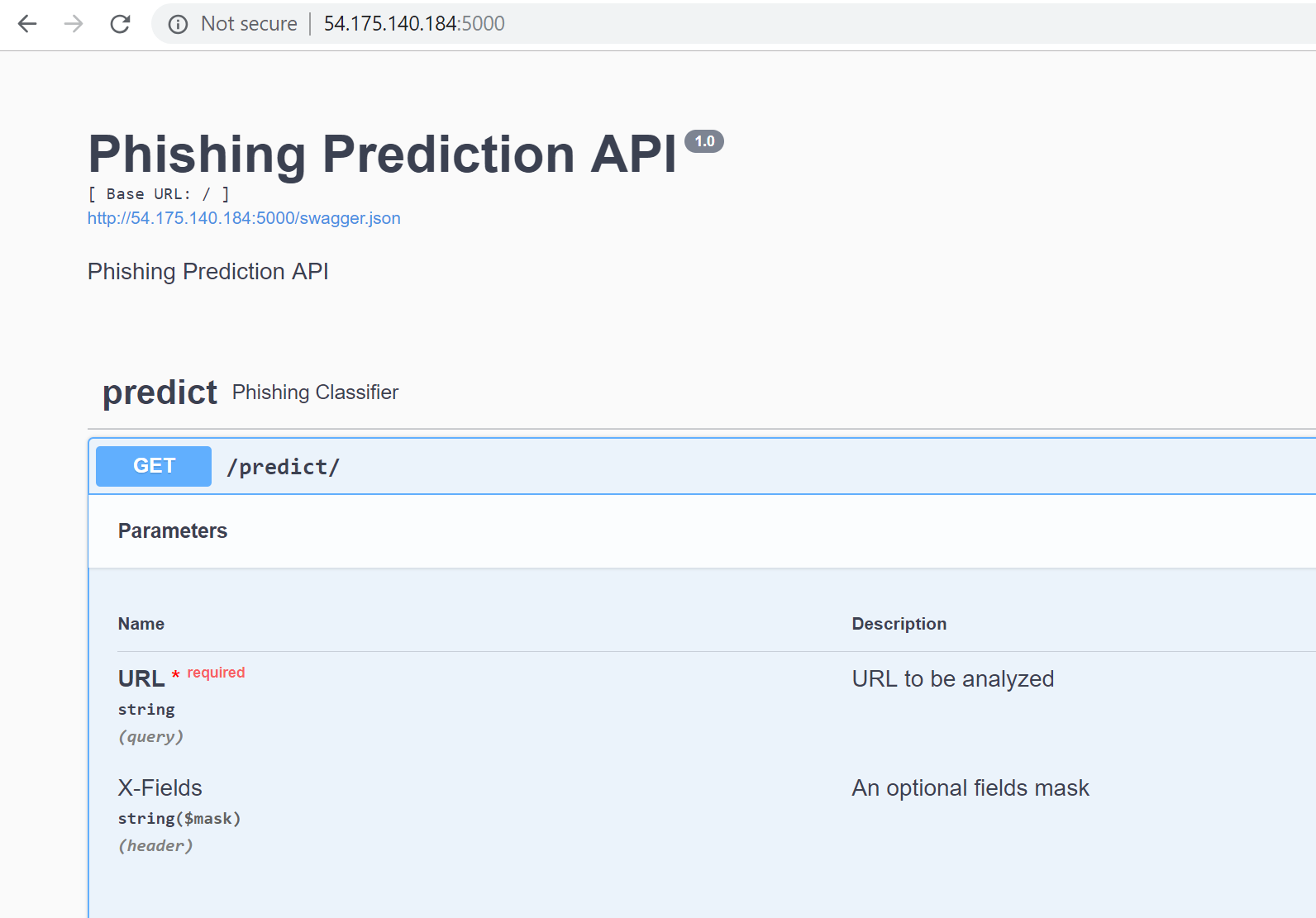
#
# #### Evaluation:
# - 40% - API hosted on a cloud service
# - 10% - Show screenshots of the model doing the predictions on the local machine
#
| exercises/P1-UsedVehiclePricePrediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot as plt
# +
img = cv2.imread("../data/train_p_1/1.png")
plt.imshow(img)
plt.show()
# -
def filter(image, kernel):
kernel = kernel / (np.sum(kernel) if np.sum(kernel) != 0 else 1)
last_image = np.zeros_like(image, dtype=np.uint8)
for i in range(3):
use_im = image[:, :, i]
use_im = cv2.filter2D(use_im.astype(float), -1, kernel)
# print(image.max(), image.min())
last_image[:, :, i] = use_im
return last_image
# +
kernel = np.array([
[-1, -1, -1],
[-1, 8, -1],
[-1, -1, -1]
], np.float32)
fil_img = filter(img, kernel)
plt.imshow(fil_img)
plt.show()
# +
th_img = cv2.threshold(cv2.cvtColor(img, cv2.COLOR_BGR2GRAY), 146, 255, cv2.THRESH_BINARY)[1]
plt.imshow(th_img)
# -
#cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
th_img = cv2.threshold(cv2.cvtColor(cv2.threshold(img,240, 255, cv2.THRESH_BINARY)[1], cv2.COLOR_BGR2GRAY), 2, 255, cv2.THRESH_BINARY)[1]
plt.imshow(th_img)
plt.show()
countours, hierachy = cv2.findContours(th_img, cv2.RETR_LIST, cv2.CHAIN_APPROX_NONE)
def max_countour(countours):
count_max = 0
max_idx = -1
for i, cnt in enumerate(countours):
# count_max = max(cnt.shape[0], count_max)
if cnt.shape[0] > count_max:
count_max = cnt.shape[0]
max_idx = i
return countours[i]
# +
img_contour = cv2.drawContours(img, [max_countour(countours)], -1, (0, 255, 0), 5)
plt.imshow(img_contour)
# -
[max_countour(countours)]
mu = cv2.moments(max_countour(countours), False)
# +
x,y= int(mu["m10"]/mu["m00"]) , int(mu["m01"]/mu["m00"])
cv2.circle(img, (x,y), 4, 100, 4, 4)
plt.imshow(img)
plt.show()
# -
x, y
| notebooks/filter_crop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KeisukeShimokawa/CarND-Advanced-Lane-Lines/blob/master/lesson18_create_movie.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="-H6grmIa7ONk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f1aea510-b6a8-42f4-e455-b2fabd884af4"
# !wget https://www.dropbox.com/s/uflwm2ii7sivkv8/run1.tar.gz -O run1.tar.gz
# !tar -zxvf run1.tar.gz
# !rm run1.tar.gz
# + id="IANdQFpN7OKW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 122} outputId="ebc07ef3-c798-4826-8ae1-9b1ea07d23e8"
from moviepy.editor import ImageSequenceClip
import argparse
import os
# + id="IqudRxxx7OG-" colab_type="code" colab={}
IMAGE_EXT = ['jpeg', 'gif', 'png', 'jpg']
# + id="zV3zHMl07ODJ" colab_type="code" colab={}
image_folder = 'run1'
fps = 50
# + id="6UznCLPk8WJq" colab_type="code" colab={}
#convert file folder into list firltered for image file types
image_list = sorted([os.path.join(image_folder, image_file)
for image_file in os.listdir(image_folder)])
# + id="bN8ejRNH8rY3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="9ba9711a-e923-4195-9b31-3759f70aa467"
image_list[:10]
# + id="SxqsPMoi83J-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="2390867d-9450-4248-ef13-891a756b531b"
image_list = [image_file for image_file in image_list if os.path.splitext(image_file)[1][1:].lower() in IMAGE_EXT]
image_list[:10]
# + id="7woYK6E-87gG" colab_type="code" colab={}
#two methods of naming output video to handle varying environemnts
video_file_1 = image_folder + '.mp4'
video_file_2 = image_folder + 'output_video.mp4'
# + id="V13SzTtZ9CgW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="f1d5e928-2c59-4d24-fe84-ff3b5be0159a"
clip = ImageSequenceClip(image_list, fps=fps)
try:
clip.write_videofile(video_file_1)
except:
clip.write_videofile(video_file_2)
# + id="_4pZ-Uei7LoQ" colab_type="code" colab={}
| lesson18_create_movie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
class hashSet(object):
def __init__(self, numBuckets):
'''
numBuckets: int. The number of buckets this hash set will have.
Raises ValueError if this value is not an integer, or if it is not greater than zero.
Sets up an empty hash set with numBuckets number of buckets.
'''
if type(numBuckets) != int or numBuckets <= 0:
raise ValueError
self.NUM_BUCKETS = numBuckets
self.set = []
for i in range(self.NUM_BUCKETS):
self.set.append([])
def hashValue(self, e):
'''
e: an integer
returns: a hash value for e, which is simply e modulo the number of
buckets in this hash set. Raises ValueError if e is not an integer.
'''
if type(e) != int:
raise ValueError
return e % self.NUM_BUCKETS
def member(self, e):
'''
e: an integer
Returns True if e is in self, and False otherwise. Raises ValueError if e is not an integer.
'''
if type(e) != int:
raise ValueError
return e in self.set[self.hashValue(e)]
def insert(self, e):
'''
e: an integer
Inserts e into the appropriate hash bucket. Raises ValueError if e is not an integer.
'''
if type(e) != int:
raise ValueError
if e not in self.set[self.hashValue(e)]:
self.set[self.hashValue(e)].append(e)
def remove(self, e):
'''
e: is an integer
Removes e from self
Raises ValueError if e is not in self or if e is not an integer.
'''
if type(e) != int or not self.member(e):
raise ValueError
find = self.set[self.hashValue(e)]
del find[find.index(e)]
def getNumBuckets(self):
'''
Returns number of buckets.
'''
return self.NUM_BUCKETS
def __str__(self):
'''
Returns string representation of data inside hash set.
'''
result = ""
bucketNr = 0
for bucket in self.set:
result += str(bucketNr) + ": " + ",".join(str(e) for e in bucket) + "\n"
bucketNr += 1
return result
hs = hashSet(10)
hs.insert(100)
hs.insert(99)
hs.insert(98)
hs.insert(92)
hs.insert(93)
hs.insert(94)
hs.insert(95)
hs.insert(96)
hs.insert(97)
hs.insert(91)
hs.insert(31)
print str(hs)
# -
a = [1,2,3]
| Haseeb DS/HashSet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# from __future__ import print_statement
# import time
# pip install swagger_client
# import swagger_client
# from swagger_client.rest import ApiException
# from pprint import pprint
import pandas as pd
from pandas import json_normalize
import numpy as np
import requests
import json
import ipywidgets as widgets
import spacy
import math
import numpy as np
import scattertext as st
from sklearn import manifold
from sklearn.metrics import euclidean_distances
# +
# pip install json_normalize
# +
# dir(requests.get("https://api.nal.usda.gov/fdc/v1/food/534358?api_key=<KEY>"))
# +
# food = json.loads(requests.get("https://api.nal.usda.gov/fdc/v1/food/534358?api_key=<KEY>").content)
# +
# for item in food.items():
# print(item)
# -
# # Direct ID pulls
food = pd.read_csv('complementaryFiles/food.csv')
food.fillna('N/A',inplace=True)
# +
# food[food['description'].str.contains('Salmon')]
# +
food[(food['description'].str.contains('salmon'))&(food['description'].str.contains('Alaska')&(food['description'].str.contains('sockeye')))]
# salmonLst = list(food[(food['description'].str.contains('salmon'))&(food['description'].str.contains('Alaska'))]['fdc_id'].unique())
# salmonLst
salmonLst = list(food[food['description'].str.contains('salmon')]['fdc_id'])
# salmonLst
# +
# json.loads(requests.get(f'https://api.nal.usda.gov/fdc/v1/foods?&fdcIds=167614,167615&api_key=<KEY>').content)
# -
salmonLst = str(salmonLst).replace('[','').replace(']','')
# +
apiKey = '<KEY>'
# salmon = json.loads(requests.get(f'https://api.nal.usda.gov/fdc/v1/foods?&fdcIds=167614,167615&api_key=<KEY>').content)
salmon = json.loads(requests.get(f'https://api.nal.usda.gov/fdc/v1/foods?&fdcIds={salmonLst}&api_key={apiKey}').content)
#survey food
# salmon = json.loads(requests.get(f'https://api.nal.usda.gov/fdc/v1/foods?&fdcIds=1098960&api_key=<KEY>').content)
df = pd.DataFrame(salmon)
df = df.dropna(subset=['foodPortions'])
# df = df[df['inputFoods'].map(lambda d: len(d)) > 0]
df = df.reset_index()
df = df.drop('index',axis=1)
# df = pd.DataFrame(df.iloc[23:24])
df.head()
# +
current_food = df['description'].values[0]
df['foodNutrients'][0] #hidden dataframe with all of the nutrient data
macros = ['Protein','Carbohydrates','Fat','Total lipid (fat)']
json_normalize(df['nutrientConversionFactors'][0][0])
json_normalize(df['foodCategory'][0])
len(list(json_normalize(df['foodNutrients'][0]).columns))
#legacy food iteration
for x in range(df.shape[0]):
col_len = len(list(json_normalize(df['foodNutrients'][x]).columns))
if col_len == 17:
nutrient_df = json_normalize(df['foodNutrients'][x])
# display(nutrient_df)
else:
pass
#survey food
nutrient_df = json_normalize(df['foodNutrients'][0])
# display(nutrient_df)
macro_df = nutrient_df[nutrient_df['nutrient.name'].isin(macros)]
conv_factors = json_normalize(df['nutrientConversionFactors'][0])
portion_df = json_normalize(df['foodPortions'][0])
# display(portion_df.head())
input_df = json_normalize(df['inputFoods'][0])
protein = macro_df[macro_df['nutrient.name']=='Protein']['amount'].values[0]
calories = nutrient_df[nutrient_df['nutrient.unitName']=='kcal']['amount'].values[0]
# input_grams = input_df['amount'].values[0]
portion_grams = portion_df['gramWeight'].values[0]
# portion_ounces = portion_df[portion_df['portionDescription'].str.contains('oz')]['portionDescription'].values[0].split()[0]
print(f'Data for {current_food}')
print('===================')
print('===================')
print('Protein per serving')
print(protein)
print('===================')
print('Calories per serving')
print(calories)
# -
# # PDF generation
# +
# pip install reportlab
# +
#table gen - https://pythonguides.com/create-and-modify-pdf-file-in-python/
from reportlab.pdfgen import canvas
# c = canvas.Canvas("salmon.pdf")
from reportlab.lib import colors
from reportlab.lib.pagesizes import letter, inch
from reportlab.platypus import SimpleDocTemplate, Table, TableStyle
# # move the origin up and to the left
# c.translate(inch,inch)
# c.setFont("Helvetica", 20)
# c.setFillColorRGB(0,0,0)
# c.drawString(.5*inch, 9*inch, f'Data for {current_food}')
# c.drawString(.5*inch, 8*inch, "========================")
# c.drawString(.5*inch, 7*inch, "========================")
# c.drawString(.5*inch, 6*inch, f'Protein per serving = {protein}')
# c.drawString(.5*inch, 5*inch, "========================")
# c.drawString(.5*inch, 4*inch, f'Calories per serving = {calories}')
# c.drawString(.5*inch, 3*inch, "========================")
# creating a pdf file to add tables
my_doc = SimpleDocTemplate("salmon.pdf", pagesize = letter)
my_obj = []
# defining Data to be stored on table
my_data = [
["Food", "Calories", "Protein"],
[current_food, calories, protein],
]
# Creating the table with 5 rows
my_table = Table(my_data, 1 * [2.5 * inch], 2 * [0.5 * inch])
# setting up style and alignments of borders and grids
my_table.setStyle(
TableStyle(
[
("ALIGN", (1, 1), (0, 0), "LEFT"),
("VALIGN", (-1, -1), (-1, -1), "TOP"),
("ALIGN", (-1, -1), (-1, -1), "RIGHT"),
("VALIGN", (-1, -1), (-1, -1), "TOP"),
("INNERGRID", (0, 0), (-1, -1), 1, colors.gray),
# ("BOX", (0, 0), (-1, -1), 2, colors.black),
]
)
)
my_obj.append(my_table)
my_doc.build(my_obj)
# c.showPage()
# c.save()
# -
# ## Search API functionality
cheddar = json.loads(requests.get("https://api.nal.usda.gov/fdc/v1/foods/search?api_key=IDmcqSbD991dUudZIXjztGVAAKnMwhvABtxzlvOQ&query=Cheddar%20Cheese").content)
cheese = json.loads(requests.get("https://api.nal.usda.gov/fdc/v1/foods/search?api_key=IDmcqSbD991dUudZIXjztGVAAKnMwhvABtxzlvOQ&query=Cheese").content)
for key, value in cheese.items():
if key == 'foods':
# ched = pd.DataFrame(columns = ['fdcId', 'description', 'lowercaseDescription', 'dataType', 'gtinUpc','publishedDate',
# 'brandOwner', 'brandName', 'ingredients','marketCountry', 'foodCategory', 'allHighlightFields', 'score','foodNutrients'])
df=[]
for val in value:
# for key,value in val.items():
# print(key,value)
df.append(pd.DataFrame(val))
else:
pass
df = pd.concat(df,sort=False)
df
# +
#### lets use k nearest neighbors on macros !!!! ######
# -
from pandas.io.json import json_normalize
nutrients = json_normalize(df['foodNutrients'])
nutrients[nutrients['nutrientName']=='Protein']
df[df['fdcId']==1497465]['foodNutrients']
# +
sp = spacy.load('en_core_web_sm')
output = widgets.Output()
from IPython.display import display
def clicked(b):
output.clear_output()
with output:
_norm = True
_sortby = 'name'
_query = querybox.value
if (normalizedradio.value == "false"):
_norm = False
if (sortradio.value == 'score'):
_sortby = 'score'
if (_query == ""):
print("please enter a query")
else:
drawTilebars(_query,normalized=_norm,sortby=_sortby).display()
querybox = widgets.Text(description='Query:')
searchbutton = widgets.Button(description="Search")
normalizedradio = widgets.RadioButtons(description="Normalized?",options=['true', 'false'])
sortradio = widgets.RadioButtons(description="Sort by",options=['name', 'score'])
searchbutton.on_click(clicked)
normalizedradio.observe(clicked)
sortradio.observe(clicked)
list_widgets = [widgets.VBox([widgets.HBox([querybox,searchbutton]),
widgets.HBox([normalizedradio,sortradio])])]
accordion = widgets.Accordion(children=list_widgets)
accordion.set_title(0,"Search Controls")
# display(accordion,output)
display(accordion)
# -
| fdcAPI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import zipfile
import numpy as np
with zipfile.ZipFile("Soccer Data\matches.zip") as Z:
with Z.open("matches_England.json") as f:
df = pd.read_json(f)
point_data = list()
result = {1 : "draw", 0 : "lost", 3: "win"}
for i in range(len(df)):
gameweek = df.iloc[i].gameweek
label = df.iloc[i].label
[[home_team, away_team], [home_score, away_score]] = [[o.strip() for o in s.split('-')] for s in label.split(',')]
home_score = int(home_score)
away_score = int(away_score)
if home_score > away_score:
home_point = 3
away_point = 0
if away_score > home_score:
away_point = 3
home_point = 0
if away_score == home_score:
home_point = 1
away_point = 1
point_data.append([gameweek, home_team, home_point, 'home', result[home_point]])
point_data.append([gameweek, away_team, away_point, 'away', result[away_point]])
point_df = pd.DataFrame(point_data, columns=['gameweek', 'team', 'point', 'home_away', 'result'])
point_df
point_df.to_csv("point.csv")
import matplotlib.pyplot as plt
team_table = point_df.pivot(index= 'gameweek', columns='team', values=['point']).cumsum().fillna(method = 'backfill').fillna(method='ffill')
plt.figure(figsize=[20,12])
colormap = plt.cm.gist_ncar
color = [colormap(i) for i in np.linspace(0, 0.9, len(team_table.columns))]
[plt.plot(team_table.iloc[:,i], color = color[i]) for i in range(len(team_table.columns))]
plt.legend([team_table.columns[i][1] for i in range(len(team_table.columns))], fontsize=12)
plt.xticks(team_table.index)
plt.xlabel("Weeks", fontsize=16)
plt.ylabel("Points", fontsize=16)
plt.show()
teams = ['Arsenal', 'Chelsea', 'Liverpool', 'Manchester United', 'Manchester City']
point_df_selected = point_df[[t in teams for t in point_df['team']]]
tab = pd.crosstab(index=[point_df_selected['team'],point_df_selected['home_away']], columns=point_df_selected['result'])
tab
from scipy.stats import chi2_contingency
chi2_contingency(tab.iloc[4:6,:].values)
point_df_selected
teams_df = pd.read_json('soccer data/teams.json', orient = 'values')
teams_df
coaches_df = pd.read_json('soccer data/coaches.json', orient = 'values')
coaches_df
coaches_teams_df = pd.merge(left=teams_df, right=coaches_df,
left_on='wyId', right_on='currentTeamId',
how='inner')[['name', 'birthDate', 'shortName']].groupby('name').agg('max', on = 'birthDate').sort_values(by='birthDate', ascending = False)
now = pd.Timestamp('now')
age = (now - pd.to_datetime(coaches_teams_df['birthDate'], yearfirst=True)).astype('<m8[Y]')
coaches_teams_df['age'] = age
print(coaches_teams_df.head(10))
plt.hist(age, density = True, edgecolor='black', linewidth=1.2)
plt.xlabel('Age', fontsize=16)
plt.title('Histogram of Coaches Ages')
events_df = pd.DataFrame()
with zipfile.ZipFile("Soccer Data\events.zip") as Z:
for name in Z.namelist():
with Z.open(name) as f:
df_temp = pd.read_json(f)#[['playerId', 'matchId', 'eventName', 'tags']]
events_df = pd.concat([events_df, df_temp])
print("file " + name + " is loaded")
break
passes_df = events_df[['playerId', 'matchId', 'eventName', 'tags']]
passes_df.head()
passes_df = passes_df.loc[passes_df.eventName == 'Pass']
passes_df['pass_success'] = [str(t).find('1801') != -1 for t in passes_df.tags]
passes_df.drop(columns=['tags','eventName'], inplace = True)
passes_df.head()
passes_df = passes_df.groupby(['playerId', 'matchId'], as_index = False, group_keys = False).agg(['sum','count'] , on='pass_success').reset_index()
passes_df.columns = ['playerId', 'matchId', 'sum', 'count']
passes_df.head()
# +
#plt.hist(df['pass_success']['count'], bins=100)
# -
passes_df = passes_df.loc[passes_df['count'] > 100]
passes_df.head()
passes_df.drop(columns = ['matchId'], inplace = True)
passes_df = passes_df.groupby('playerId').agg('sum', level = 0, on = ['sum', 'count']).reset_index()
passes_df.head()
passes_df['ratio'] = passes_df['sum']/passes_df['count']*100
passes_df.head()
passes_top10 = passes_df.sort_values('ratio', ascending=False).head(10)
passes_top10
players_df = pd.read_json('soccer data\players.json')
players_df.head(3)
players_name = players_df[['firstName','middleName','lastName', 'wyId']].copy()
players_name['fullName'] = players_name['firstName'] + ' ' + players_name['middleName'] + ' ' + players_name['lastName']
players_name.head()
players_name.drop(columns = ['firstName', 'middleName', 'lastName'], inplace = True)
players_name.head()
passes_top10 = pd.merge(left=passes_top10, right=players_name, left_on='playerId', right_on='wyId', how='left')
passes_top10[['fullName','ratio']]
airduels_df = events_df[['playerId', 'matchId', 'eventName', 'subEventName', 'tags']]
airduels_df.head()
airduels_df = airduels_df.loc[airduels_df.subEventName == 'Air duel']
airduels_df = airduels_df.loc[airduels_df.eventName == 'Duel']
airduels_df['duel_success'] = [str(t).find('1801') != -1 for t in airduels_df.tags]
airduels_df.drop(columns=['tags','eventName', 'subEventName'], inplace = True)
airduels_df.head()
airduels_df = airduels_df.groupby(['playerId', 'matchId'], as_index = False, group_keys = False).agg(['sum','count'] , on='duel_success').reset_index()
airduels_df.columns = ['playerId', 'matchId', 'sum', 'count']
airduels_df.head()
airduels_df = airduels_df.loc[airduels_df['count'] > 5]
airduels_df.head()
players_height = players_df[['height', 'wyId']].copy()
players_height.head()
airduels_height = pd.merge(left=airduels_df, right=players_height, left_on='playerId', right_on='wyId', how='inner')[['height', 'sum','count']]
airduels_height = airduels_height.groupby(pd.cut(airduels_height["height"], np.arange(155, 210, 5))).sum(on = ['sum' , 'count'])
airduels_height.drop(columns='height', inplace = True)
airduels_height.reset_index()
airduels_height['ratio'] = airduels_height['sum']/airduels_height['count']*100
plt.figure(figsize=(15,7))
plt.scatter(range(len(airduels_height)), airduels_height['ratio'].values, c = range(len(airduels_height)), cmap = 'YlOrRd')
plt.xticks(range(len(airduels_height)), airduels_height.index)
# ## CQ1
events_df.head()
goals_df = events_df[['playerId', 'eventSec','teamId','tags','eventName', 'matchPeriod']]
goals_df.head()
tags101 = [str(t).find(' 101') != -1 for t in goals_df['tags']]
goals_df = goals_df.loc[tags101]
goals_df.head()
goals_df = goals_df.loc[goals_df['eventName'] != 'Save attempt']
goals_df.head()
goals_df['eventMin'] = goals_df['eventSec']//60 + 1
goals_df.head()
time_slots = [str(t) for t in pd.cut(goals_df['eventMin'], np.arange(0, 60, 9))]
goals_df['time_slot'] = time_slots
goals_df.head()
res = goals_df.groupby(['matchPeriod', 'time_slot']).count()[['playerId']]
res
res_plot = res.plot(kind='bar', legend=False)
res1 = goals_df.groupby(['teamId', 'time_slot', 'matchPeriod']).count()[['playerId']].reset_index()
res1.columns = ['teamId','time_slot','matchPeriod','scores']
res2 = res1.loc[res1['time_slot'] == '(36, 45]']
res3 = res2.loc[[str(t).find('2H') != -1 for t in res2['matchPeriod']]]
asd = pd.merge(left = res3, right=teams_df, left_on='teamId', right_on='wyId')[['time_slot','matchPeriod','scores','officialName']]
asd.max()
goals_df.head()
r0 = goals_df.groupby(['time_slot','playerId']).count().reset_index()[['time_slot','playerId','tags']]
r0.columns = ['time_slot','playerId','scores']
r0.head()
r1 = r0.groupby('playerId').count().reset_index()[['playerId','time_slot']]
r1.columns = ['playerId', 'nslot_covered']
r1.sort_values(by = 'nslot_covered', ascending=False)
events_df.head()
pd.unique(events_df['eventName'])
# ## RCQ2
with zipfile.ZipFile("Soccer Data\matches.zip") as Z:
with Z.open('matches_Spain.json') as f:
matches_df = pd.read_json(f)
with zipfile.ZipFile("Soccer Data\events.zip") as Z:
with Z.open('events_Spain.json') as f:
events_spain_df = pd.read_json(f)
events_spain_df.iloc[594533,:]
barcelona_mardrid_id = 2565907 #Barcelona - Real Madrid
CR7_id = 3359 #CR7
LM_id = 3322 #Messi
def event_coordinate(coordinate):
[[_,y_start],[_,x_start],[_,y_end],[_,x_end]] = [i.split(': ') for i in str(coordinate).replace('[','').replace(']','').replace('{','').replace('}','').split(',')]
return int(x_start)/100*130, int(y_start)/100*90, int(x_end)/100*130, int(y_end)/100*90
barcelona_madrid_df = events_spain_df[['eventName','matchId','positions','playerId']].loc[
events_spain_df['eventName'].isin(['Pass', 'Duel','Free Kick','Shot']) &
events_spain_df['matchId'].isin([barcelona_mardrid_id]) &
events_spain_df['playerId'].isin([CR7_id])]
xy_CR7 = barcelona_madrid_df['positions'].apply(event_coordinate)
xy_CR7 = xy_CR7.loc[[i[2] != 0 and i[3] != 0 for i in xy_CR7]]
barcelona_madrid_df = events_spain_df[['eventName','matchId','positions','playerId']].loc[
events_spain_df['eventName'].isin(['Pass', 'Duel','Free Kick','Shot']) &
events_spain_df['matchId'].isin([barcelona_mardrid_id]) &
events_spain_df['playerId'].isin([LM_id])]
xy_LM = barcelona_madrid_df['positions'].apply(event_coordinate)
xy_LM = xy_CR7.loc[[i[2] != 0 and i[3] != 0 for i in xy_LM]]
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.patches import Arc
import seaborn as sns
#Create figure
def plot_pitch():
fig=plt.figure()
fig.set_size_inches(7, 5)
ax=fig.add_subplot(1,1,1)
#Pitch Outline & Centre Line
plt.plot([0,0],[0,90], color="black")
plt.plot([0,130],[90,90], color="black")
plt.plot([130,130],[90,0], color="black")
plt.plot([130,0],[0,0], color="black")
plt.plot([65,65],[0,90], color="black")
#Left Penalty Area
plt.plot([16.5,16.5],[65,25],color="black")
plt.plot([0,16.5],[65,65],color="black")
plt.plot([16.5,0],[25,25],color="black")
#Right Penalty Area
plt.plot([130,113.5],[65,65],color="black")
plt.plot([113.5,113.5],[65,25],color="black")
plt.plot([113.5,130],[25,25],color="black")
#Left 6-yard Box
plt.plot([0,5.5],[54,54],color="black")
plt.plot([5.5,5.5],[54,36],color="black")
plt.plot([5.5,0.5],[36,36],color="black")
#Right 6-yard Box
plt.plot([130,124.5],[54,54],color="black")
plt.plot([124.5,124.5],[54,36],color="black")
plt.plot([124.5,130],[36,36],color="black")
#Prepare Circles
centreCircle = plt.Circle((65,45),9.15,color="black",fill=False)
centreSpot = plt.Circle((65,45),0.8,color="black")
leftPenSpot = plt.Circle((11,45),0.8,color="black")
rightPenSpot = plt.Circle((119,45),0.8,color="black")
#Draw Circles
ax.add_patch(centreCircle)
ax.add_patch(centreSpot)
ax.add_patch(leftPenSpot)
ax.add_patch(rightPenSpot)
#Prepare Arcs
leftArc = Arc((11,45),height=18.3,width=18.3,angle=0,theta1=310,theta2=50,color="black")
rightArc = Arc((119,45),height=18.3,width=18.3,angle=0,theta1=130,theta2=230,color="black")
#Draw Arcs
ax.add_patch(leftArc)
ax.add_patch(rightArc)
#Tidy Axes
plt.axis('off')
plot_pitch()
x_coord = [i[0] for i in xy_CR7]
y_coord = [i[1] for i in xy_CR7]
sns.kdeplot(x_coord, y_coord, shade = "True", color = "green", n_levels = 30, shade_lowest = False)
ply.title('asdasd')
plt.show()
plot_pitch()
x_coord = [i[0] for i in xy_LM]
y_coord = [i[1] for i in xy_LM]
sns.kdeplot(x_coord, y_coord, shade = "True", color = "green", n_levels = 30, shade_lowest = False)
plt.show()
with zipfile.ZipFile("Soccer Data\matches.zip") as Z:
with Z.open('matches_Italy.json') as f:
matches_df = pd.read_json(f)
with zipfile.ZipFile("Soccer Data\events.zip") as Z:
with Z.open('events_Italy.json') as f:
events_italy_df = pd.read_json(f)
juventus_napoli_id = 2576295 #Barcelona - Real Madrid
Jorg_id = 21315 # Jorginho
Pjan_id = 20443 # <NAME>
juventus_napoli_df = events_italy_df[['eventName','matchId','positions','playerId']].loc[
events_italy_df['eventName'].isin(['Pass', 'Duel','Free Kick','Shot']) &
events_italy_df['matchId'].isin([juventus_napoli_id]) &
events_italy_df['playerId'].isin([Jorg_id])]
xy_Jorg = juventus_napoli_df['positions'].apply(event_coordinate)
xy_Jorg = xy_Jorg.loc[[i[2] != 0 and i[3] != 0 for i in xy_Jorg]]
plot_pitch()
x_coord = [i[0] for i in xy_Jorg]
y_coord = [i[1] for i in xy_Jorg]
sns.kdeplot(x_coord, y_coord, shade = "True", color = "green", n_levels = 30, shade_lowest = False)
for xy in xy_Jorg:
plt.annotate(xy = [xy[2],xy[3]], arrowprops=dict(arrowstyle="->",connectionstyle="arc3", color = "blue"),s ='',
xytext = [xy[0],xy[1]])
juventus_napoli_df = events_italy_df[['eventName','matchId','positions','playerId']].loc[
events_italy_df['eventName'].isin(['Pass', 'Duel','Free Kick','Shot']) &
events_italy_df['matchId'].isin([juventus_napoli_id]) &
events_italy_df['playerId'].isin([Pjan_id])]
xy_Pjan = juventus_napoli_df['positions'].apply(event_coordinate)
xy_Pjan = xy_Jorg.loc[[i[2] != 0 and i[3] != 0 for i in xy_Pjan]]
plot_pitch()
#plt.title('asdasd')
x_coord = [i[0] for i in xy_Pjan]
y_coord = [i[1] for i in xy_Pjan]
sns.kdeplot(x_coord, y_coord, shade = "True", color = "green", n_levels = 30, shade_lowest = False)
for xy in xy_Pjan:
plt.annotate(xy = [xy[2],xy[3]], arrowprops=dict(arrowstyle="->",connectionstyle="arc3", color = "blue"),s ='',
xytext = [xy[0],xy[1]])
# +
#events_italy_df
# +
#events_italy_df.loc[events_italy_df['eventId'] == 2]
# -
| HW2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python: Advanced
# ## Classes
#
# **Classes** are like blueprints for creating objects.
#
# **Instances** are what is built using the class.
#
# The `__init__()` function is found in all **classes**. It is used to assign values to object properties, or other operations when the *object is being created*.
#
# `self` is used to reference the current instance of the class, and is used to access variables that belong to the class. *it doesn't need to be named self*
#
# Classes can contain **methods** (in object functions)
#
# +
class car:
def __init__(self, make, topspeed):
self.make = make
self.topspeed = topspeed
blueCar = car("Ford", "150")
print(blueCar.make)
print(blueCar.topspeed)
# -
# add a method to the class
# +
class animal:
def __init__(self, name, weight, habitat):
self.name = name
self.weight = weight
self.habitat = habitat
def sayHello(self):
print("Hello I am a " + self.name)
tiger = animal("tiger", 200, "forest")
tiger.sayHello()
# -
# modifying object parameters is similar to normal python sytax
# +
tiger.weight = 220
print(tiger.weight)
del tiger.habitat
print(tiger.__dict__) # quick investigation
# -
# ### Inheritance
# Inheritance allows us to define a class that inherits methods and properties from another class.
#
# **parent** and **child** classes will be used.
#
#
# +
# Parent class
class animal:
def __init__(self, name, weight, habitat):
self.name = name
self.weight = weight
self.habitat = habitat
def sayHello(self):
print("Hello I am a " + self.name)
# Child class
class fish(animal):
pass # used for empty classes
shark = fish("Great White", "1000", "tropical ocean")
print(shark.name)
# -
# if we were to add the `__init__()` function, the child overrides that of the parents class.
#
# To keep it, add a call to the parents `__init__()` directly or use `super()`
# +
class animal:
def __init__(self, name, weight, habitat):
self.name = name
self.weight = weight
self.habitat = habitat
# explicitly calling
class bird(animal):
def __init__(self, name, weight, habitat):
animal.__init__(self, name, weight, habitat)
# using super()
class mammal(animal):
def __init__(self, name, weight, habitat):
super().__init__(name, weight, habitat)
eagle = bird("Golden", 2, "mountains")
print(eagle.__dict__)
bear = mammal("Grizzly", 500, "forest")
print(bear.__dict__)
# -
# properties can also be added at the inheritance stage
# +
class animal:
def __init__(self, name, weight, habitat):
self.name = name
self.weight = weight
self.habitat = habitat
class insect(animal):
def __init__(self, name, weight, habitat, colonySize):
super().__init__(name, weight, habitat)
self.colonySize = colonySize
self.exoskeleton = True # you can also add properties that aren't assigned
def printColonySize(self):
print("The ", self.name, " has a colony size of ", self.colonySize)
ant = insect("fire ant", 0.001, "tropical forest", 100000)
ant.printColonySize()
ant.__dict__
# -
# ### Iterators
# Is an object that can be iterated upon.
#
# Specifically consitsting of the metods `__iter__()` and `__next__()`
#
# `__iter__()` - returns the iterator object
#
# `__next__()` - returns the next item in the sequence
#
#
# +
class numbers:
def __iter__(self):
self.a = 1
return self
def __next__(self):
x = self.a
self.a += 1
return x
myclass = numbers()
myiter = iter(myclass)
print(next(myiter))
print(next(myiter))
print(next(myiter))
# -
# to stop the iteration use `StopIteration`
# +
class numbers:
def __iter__(self):
self.a = 1
return self
def __next__(self):
if self.a <= 10:
x = self.a
self.a += 1
return x
else:
raise StopIteration
myclass = numbers()
myiter = iter(myclass)
for x in myiter:
print(x)
# -
# ## Class Concepts
#
# classes have a number of key concepts
# * inheritance - creating a new class from a parent
# * encapsulation - being unable to effect the core data, unless a function specifices this
# * Polymorphism - to use a common interface for multiple forms
# ### Inheritance
# +
# Parent class
class animal:
def __init__(self, species, habitat, size):
self.species = species
self.habitat = habitat
self.size = size
def __str__(self):
return "I am a " + self.species
def sayHelloSpecies(self):
print("hello I am a " + self.species)
# child class
class fish(animal):
def __init__(self, species, habitat, size):
super().__init__(species, habitat, size)
def fishSpeak(self):
print("bubble bubble bubble")
skate = fish("skate", "north sea", 4)
skate.sayHelloSpecies()
skate.fishSpeak()
print(skate)
# -
# ### Encapsulation
#
# note how the `__maxprice` cannot be changed without the use of the specificed function.
# +
class Computer:
def __init__(self):
self.__maxprice = 900
def sell(self):
print("selling at price {}".format(self.__maxprice))
def setMaxPrice(self, price):
self.__maxprice = price
c = Computer()
c.sell()
# change the price - doesn't work! good
c.__maxprice = 9
c.sell()
# using setter function - works
c.setMaxPrice(1000)
c.sell()
# -
# ### Polymorphism
#
# We can have multiple common classes, that can be used by other functions/class/etc.
#
# the `flying_test()` function uses the `fly()` method in both the parrot and emu classes.
# +
# parent
class animal:
def __init__(self, species, habitat, size):
self.species = species
self.habitat = habitat
self.size = size
class parrot(animal):
def __init__(self, species, habitat, size):
super().__init__(species, habitat, size)
def fly(self):
print("can fly")
class emu(animal):
def __init__(self, species, habitat, size):
super().__init__(species, habitat, size)
def fly(self):
print("cannot fly")
# define a common test
def flying_test(birdSpecies):
birdSpecies.fly()
# create object and run tests
polly = parrot("parrot", "tropics", 1)
paul = emu("emu", "tropics", 67)
flying_test(polly)
flying_test(paul)
# -
# ## Custom iterators
# uses `__iter__()` and `__next__()` methods
#
# `__iter__()` - returns the iterator object
#
# `__next__()` - returns the next item in the sequence
# it also requires use of `StopIteration`
#
# +
class powerTwo:
def __init__(self, max=0):
self.max = max
def __iter__(self):
self.n = 0
return self
def __next__(self):
if self.n <= self.max:
result = 2 ** self.n
self.n += 1
return result
else:
raise StopIteration
for i in powerTwo(3):
print(i)
# -
# ## Generators
# Are a simple way of creating iterations
#
# `yield` is used at least once in a function. It has the same effect as the `return` function, but pauses the functions state, allowing it to be iterable.
#
# `__iter__()` and `__next__()` methods are automatically initiated
#
# They are:
# * memory efficient
# * easier to understand
#
# +
def powerTwoAlt(max = 0):
n = 0
while n <= max:
yield 2 ** n
n += 1
for i in powerTwoAlt(3):
print(i)
# -
# An alternative is in the style of a list comprehension, but uses `()` over `[]`
# +
# list comprehension
listPowerTwo = [2 ** x for x in range(0, 4)]
# generator
generatorPowerTwo = (2 ** x for x in range(0, 4))
print(listPowerTwo)
print(generatorPowerTwo)
for i in generatorPowerTwo:
print(i)
# -
# ## Closure
# if a function has a *nested* function with a value that must be *referenced in the enclosed function* then closure should be used.
#
# It is the concept of returning the nested function
#
# +
def makePrinter(msg):
def printer():
print(msg)
def doSomethingElse():
return 0
return printer
testPrint = makePrinter("hello there")
testPrint()
# -
# ## Decorators
# adds functionallity to existing code - *metacoding*
#
# decorators act as a wrapper
#
# Divider example, we are aiming to add functionality to:
#
# ```python
# def divide(a, b):
# return a / b
# ```
# +
# the decorator function
def describeDivide(func):
def innerDivide(a, b):
print("I am going to divide ", a, " and ", b)
return func(a, b) # returns answer
return innerDivide # returns statement
@describeDivide
def divide(a, b):
return a / b
divide(10, 2)
# -
# Decorators can be **chained together**, making the code more modular
#
# in the below example:
#
# `args` is the tuple of positional arguments
#
# `kwargs` is the dictionary of keyword arguments
#
# thus...
#
# `function(*args, **kwargs)` is th ultimate python wildcard
#
# **pseudocode for decorators**
# 1. define function
# 2. define inner function allowing everything to pass
# 3. print `*`s / `%`s
# 4. Let the function run
# 5. print `*`s / `%`s
# 6. exit
#
#
# +
def addStars(func):
def inner(*args, **kwargs):
print("*" * 30)
func(*args, **kwargs)
print("*" * 30)
return inner
def addPercentage(func):
def inner(*args, **kwargs):
print("%" * 30)
func(*args, **kwargs)
print("%" * 30)
return inner
# now chain them
@addStars
@addPercentage
def printBanner(msg):
print(msg)
printBanner("Hello I am a banner")
# -
| programming_notes/_build/jupyter_execute/python_advanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Линейная регрессия и основные библиотеки Python для анализа данных и научных вычислений
# Это задание посвящено линейной регрессии. На примере прогнозирования роста человека по его весу Вы увидите, какая математика за этим стоит, а заодно познакомитесь с основными библиотеками Python, необходимыми для дальнейшего прохождения курса.
# **Материалы**
#
# - Лекции данного курса по линейным моделям и градиентному спуску
# - [Документация](http://docs.scipy.org/doc/) по библиотекам NumPy и SciPy
# - [Документация](http://matplotlib.org/) по библиотеке Matplotlib
# - [Документация](http://pandas.pydata.org/pandas-docs/stable/tutorials.html) по библиотеке Pandas
# - [Pandas Cheat Sheet](http://www.analyticsvidhya.com/blog/2015/07/11-steps-perform-data-analysis-pandas-python/)
# - [Документация](http://stanford.edu/~mwaskom/software/seaborn/) по библиотеке Seaborn
# ## Задание 1. Первичный анализ данных c Pandas
# В этом заданиии мы будем использовать данные [SOCR](http://wiki.stat.ucla.edu/socr/index.php/SOCR_Data_Dinov_020108_HeightsWeights) по росту и весу 25 тысяч подростков.
# **[1].** Если у Вас не установлена библиотека Seaborn - выполните в терминале команду *conda install seaborn*. (Seaborn не входит в сборку Anaconda, но эта библиотека предоставляет удобную высокоуровневую функциональность для визуализации данных).
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# Считаем данные по росту и весу (*weights_heights.csv*, приложенный в задании) в объект Pandas DataFrame:
data = pd.read_csv('weights_heights.csv', index_col='Index')
# Чаще всего первое, что надо надо сделать после считывания данных - это посмотреть на первые несколько записей. Так можно отловить ошибки чтения данных (например, если вместо 10 столбцов получился один, в названии которого 9 точек с запятой). Также это позволяет познакомиться с данными, как минимум, посмотреть на признаки и их природу (количественный, категориальный и т.д.).
#
# После этого стоит построить гистограммы распределения признаков - это опять-таки позволяет понять природу признака (степенное у него распределение, или нормальное, или какое-то еще). Также благодаря гистограмме можно найти какие-то значения, сильно не похожие на другие - "выбросы" в данных.
# Гистограммы удобно строить методом *plot* Pandas DataFrame с аргументом *kind='hist'*.
#
# **Пример.** Построим гистограмму распределения роста подростков из выборки *data*. Используем метод *plot* для DataFrame *data* c аргументами *y='Height'* (это тот признак, распределение которого мы строим)
data.plot(y='Height', kind='hist',
color='red', title='Height (inch.) distribution')
# Аргументы:
#
# - *y='Height'* - тот признак, распределение которого мы строим
# - *kind='hist'* - означает, что строится гистограмма
# - *color='red'* - цвет
# **[2]**. Посмотрите на первые 5 записей с помощью метода *head* Pandas DataFrame. Нарисуйте гистограмму распределения веса с помощью метода *plot* Pandas DataFrame. Сделайте гистограмму зеленой, подпишите картинку.
data.head()
data.plot(y='Weight', kind='hist',
color='green', title='Weight (pounds) distribution')
# Один из эффективных методов первичного анализа данных - отображение попарных зависимостей признаков. Создается $m \times m$ графиков (*m* - число признаков), где по диагонали рисуются гистограммы распределения признаков, а вне диагонали - scatter plots зависимости двух признаков. Это можно делать с помощью метода $scatter\_matrix$ Pandas Data Frame или *pairplot* библиотеки Seaborn.
#
# Чтобы проиллюстрировать этот метод, интересней добавить третий признак. Создадим признак *Индекс массы тела* ([BMI](https://en.wikipedia.org/wiki/Body_mass_index)). Для этого воспользуемся удобной связкой метода *apply* Pandas DataFrame и lambda-функций Python.
def make_bmi(height_inch, weight_pound):
METER_TO_INCH, KILO_TO_POUND = 39.37, 2.20462
return (weight_pound / KILO_TO_POUND) / \
(height_inch / METER_TO_INCH) ** 2
data['BMI'] = data.apply(lambda row: make_bmi(row['Height'],
row['Weight']), axis=1)
# **[3].** Постройте картинку, на которой будут отображены попарные зависимости признаков , 'Height', 'Weight' и 'BMI' друг от друга. Используйте метод *pairplot* библиотеки Seaborn.
sns.pairplot(data)
# Часто при первичном анализе данных надо исследовать зависимость какого-то количественного признака от категориального (скажем, зарплаты от пола сотрудника). В этом помогут "ящики с усами" - boxplots библиотеки Seaborn. Box plot - это компактный способ показать статистики вещественного признака (среднее и квартили) по разным значениям категориального признака. Также помогает отслеживать "выбросы" - наблюдения, в которых значение данного вещественного признака сильно отличается от других.
# **[4]**. Создайте в DataFrame *data* новый признак *weight_category*, который будет иметь 3 значения: 1 – если вес меньше 120 фунтов. (~ 54 кг.), 3 - если вес больше или равен 150 фунтов (~68 кг.), 2 – в остальных случаях. Постройте «ящик с усами» (boxplot), демонстрирующий зависимость роста от весовой категории. Используйте метод *boxplot* библиотеки Seaborn и метод *apply* Pandas DataFrame. Подпишите ось *y* меткой «Рост», ось *x* – меткой «Весовая категория».
# +
def weight_category(weight):
pass
return (1 if weight < 120 else (3 if weight >= 150 else 2))
data['weight_cat'] = data['Weight'].apply(weight_category)
sns_boxplot = sns.boxplot(x='weight_cat', y='Height', data=data)
sns_boxplot.set(xlabel='Весовая категория', ylabel='Рост');
# -
# **[5].** Постройте scatter plot зависимости роста от веса, используя метод *plot* для Pandas DataFrame с аргументом *kind='scatter'*. Подпишите картинку.
data.plot(x='Weight', y='Height', kind='scatter')
# ## Задание 2. Минимизация квадратичной ошибки
# В простейшей постановке задача прогноза значения вещественного признака по прочим признакам (задача восстановления регрессии) решается минимизацией квадратичной функции ошибки.
#
# **[6].** Напишите функцию, которая по двум параметрам $w_0$ и $w_1$ вычисляет квадратичную ошибку приближения зависимости роста $y$ от веса $x$ прямой линией $y = w_0 + w_1 * x$:
# $$error(w_0, w_1) = \sum_{i=1}^n {(y_i - (w_0 + w_1 * x_i))}^2 $$
# Здесь $n$ – число наблюдений в наборе данных, $y_i$ и $x_i$ – рост и вес $i$-ого человека в наборе данных.
def error(w0, w1):
error = 0
for i in range(1, data.shape[0]):
error += (data['Height'][i] - (w0 + w1 * data['Weight'][i])) ** 2
return error
# Итак, мы решаем задачу: как через облако точек, соответсвующих наблюдениям в нашем наборе данных, в пространстве признаков "Рост" и "Вес" провести прямую линию так, чтобы минимизировать функционал из п. 6. Для начала давайте отобразим хоть какие-то прямые и убедимся, что они плохо передают зависимость роста от веса.
#
# **[7].** Проведите на графике из п. 5 Задания 1 две прямые, соответствующие значениям параметров ($w_0, w_1) = (60, 0.05)$ и ($w_0, w_1) = (50, 0.16)$. Используйте метод *plot* из *matplotlib.pyplot*, а также метод *linspace* библиотеки NumPy. Подпишите оси и график.
# +
def f(w0, w1, x):
return w0 + w1 * x
data.plot(x='Weight', y='Height', kind='scatter');
x = np.linspace(60, 180, 6)
y = f(60, 0.05, x)
plt.plot(x, y)
y = f(50, 0.16, x)
plt.plot(x, y)
plt.xlabel('Weight')
plt.ylabel('Height')
plt.title('First approximation');
# -
# Минимизация квадратичной функции ошибки - относительная простая задача, поскольку функция выпуклая. Для такой задачи существует много методов оптимизации. Посмотрим, как функция ошибки зависит от одного параметра (наклон прямой), если второй параметр (свободный член) зафиксировать.
#
# **[8].** Постройте график зависимости функции ошибки, посчитанной в п. 6, от параметра $w_1$ при $w_0$ = 50. Подпишите оси и график.
w1 = np.linspace(-5, 5, 200)
y = error(50, w1)
plt.plot(w1, y)
plt.xlabel('w1')
plt.ylabel('error(50, w1)')
plt.title('Error function (w0 fixed, w1 variates)');
# Теперь методом оптимизации найдем "оптимальный" наклон прямой, приближающей зависимость роста от веса, при фиксированном коэффициенте $w_0 = 50$.
#
# **[9].** С помощью метода *minimize_scalar* из *scipy.optimize* найдите минимум функции, определенной в п. 6, для значений параметра $w_1$ в диапазоне [-5,5]. Проведите на графике из п. 5 Задания 1 прямую, соответствующую значениям параметров ($w_0$, $w_1$) = (50, $w_1\_opt$), где $w_1\_opt$ – найденное в п. 8 оптимальное значение параметра $w_1$.
# +
from scipy.optimize import minimize_scalar
def error_50(w1):
return error(50, w1)
res = minimize_scalar(error_50)
w1_opt = res.x
# +
data.plot(x='Weight', y='Height', kind='scatter');
x = np.linspace(60, 180, 6)
y = f(50, w1_opt, x)
plt.plot(x, y)
plt.xlabel('Weight')
plt.ylabel('Height')
plt.title('Second approximation (w0 fixed, w1 optimized)');
# -
# При анализе многомерных данных человек часто хочет получить интуитивное представление о природе данных с помощью визуализации. Увы, при числе признаков больше 3 такие картинки нарисовать невозможно. На практике для визуализации данных в 2D и 3D в данных выделаяют 2 или, соответственно, 3 главные компоненты (как именно это делается - мы увидим далее в курсе) и отображают данные на плоскости или в объеме.
#
# Посмотрим, как в Python рисовать 3D картинки, на примере отображения функции $z(x,y) = sin(\sqrt{x^2+y^2})$ для значений $x$ и $y$ из интервала [-5,5] c шагом 0.25.
from mpl_toolkits.mplot3d import Axes3D
# Создаем объекты типа matplotlib.figure.Figure (рисунок) и matplotlib.axes._subplots.Axes3DSubplot (ось).
# +
fig = plt.figure()
ax = fig.gca(projection='3d') # get current axis
# Создаем массивы NumPy с координатами точек по осям X и У.
# Используем метод meshgrid, при котором по векторам координат
# создается матрица координат. Задаем нужную функцию Z(x, y).
X = np.arange(-5, 5, 0.25)
Y = np.arange(-5, 5, 0.25)
X, Y = np.meshgrid(X, Y)
Z = np.sin(np.sqrt(X**2 + Y**2))
# Наконец, используем метод *plot_surface* объекта
# типа Axes3DSubplot. Также подписываем оси.
surf = ax.plot_surface(X, Y, Z)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
plt.show()
# -
# **[10].** Постройте 3D-график зависимости функции ошибки, посчитанной в п.6 от параметров $w_0$ и $w_1$. Подпишите ось $x$ меткой «Intercept», ось $y$ – меткой «Slope», a ось $z$ – меткой «Error».
# +
fig = plt.figure()
ax = fig.gca(projection='3d') # get current axis
w0 = np.arange(0, 100, 1)
w1 = np.arange(-5, 5, 0.1)
w0, w1 = np.meshgrid(w0, w1)
err_func = error(w0, w1)
surf = ax.plot_surface(w0, w1, err_func)
ax.set_xlabel('Intercept')
ax.set_ylabel('Slope')
ax.set_zlabel('Error')
plt.show()
# -
# **[11].** С помощью метода *minimize* из scipy.optimize найдите минимум функции, определенной в п. 6, для значений параметра $w_0$ в диапазоне [-100,100] и $w_1$ - в диапазоне [-5, 5]. Начальная точка – ($w_0$, $w_1$) = (0, 0). Используйте метод оптимизации L-BFGS-B (аргумент method метода minimize). Проведите на графике из п. 5 Задания 1 прямую, соответствующую найденным оптимальным значениям параметров $w_0$ и $w_1$. Подпишите оси и график.
# +
from scipy.optimize import minimize
def error_func(w):
return error(w[0], w[1])
minimum = minimize(error_func, [0, 0], method='L-BFGS-B')
w0_opt = minimum.x[0]
w1_opt = minimum.x[1]
# +
data.plot(x='Weight', y='Height', kind='scatter');
x = np.linspace(60, 180, 6)
y = f(w0_opt, w1_opt, x)
plt.plot(x, y)
plt.xlabel('Weight')
plt.ylabel('Height')
plt.title('Third approximation (params are optimized)');
# -
# ## Критерии оценки работы
# - Выполняется ли тетрадка IPython без ошибок? (15 баллов)
# - Верно ли отображена гистограмма распределения роста из п. 2? (3 балла). Правильно ли оформлены подписи? (1 балл)
# - Верно ли отображены попарные зависимости признаков из п. 3? (3 балла). Правильно ли оформлены подписи? (1 балл)
# - Верно ли отображена зависимость роста от весовой категории из п. 4? (3 балла). Правильно ли оформлены подписи? (1 балл)
# - Верно ли отображен scatter plot роста от веса из п. 5? (3 балла). Правильно ли оформлены подписи? (1 балл)
# - Правильно ли реализована функция подсчета квадратичной ошибки из п. 6? (10 баллов)
# - Правильно ли нарисован график из п. 7? (3 балла) Правильно ли оформлены подписи? (1 балл)
# - Правильно ли нарисован график из п. 8? (3 балла) Правильно ли оформлены подписи? (1 балл)
# - Правильно ли используется метод minimize\_scalar из scipy.optimize? (6 баллов). Правильно ли нарисован график из п. 9? (3 балла) Правильно ли оформлены подписи? (1 балл)
# - Правильно ли нарисован 3D-график из п. 10? (6 баллов) Правильно ли оформлены подписи? (1 балл)
# - Правильно ли используется метод minimize из scipy.optimize? (6 баллов). Правильно ли нарисован график из п. 11? (3 балла). Правильно ли оформлены подписи? (1 балл)
| supervised-learning/linreg-height-weight/peer_review_linreg_height_weight.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../sample/')
from ann import NeuralNetwork
sys.path.append('../../') # put `little_mcmc/` in `../../`.
import little_mcmc.sample.mcmc as mc
import numpy as np
from random import gauss, random
from time import time
import matplotlib.pyplot as plt
# -
# Typing Hint:
# +
from typing import List, Tuple, Mapping
Array = np.array(List[float])
Value = float
# -
# ### Set global variables:
# +
# Ann
net_size = [5, 1]
input_size = 1
# effective_dim = num_of_connections
effective_dim = np.prod(net_size) * (input_size * net_size[0])
print('effective_dim: ', effective_dim)
step_length = 0.5
chain_num = 1 #* effective_dim
tolerence = 0.00
max_break_counter = 100 * effective_dim
# -
# ### Set error-function as, e.g.,
def error_function(outputs: Array, targets: Array) -> float:
assert len(outputs) == len(targets)
return 0.5 * np.sum(np.square(outputs - targets)) / len(outputs)
# ## Aim: fit `[0.5 * sin(x) for x in np.linspace(-7, 7, 20)]`.
# ### Preparing for MCMC
def random_move(net: NeuralNetwork, step_length=step_length) -> NeuralNetwork:
result_net = net.copy()
for layer in result_net.layers:
for perceptron in layer:
perceptron.weights = ( perceptron.weights
+ np.array([ gauss(0, 1) * step_length
for _ in range(len(perceptron.weights))])
)
return result_net
# +
init_net = NeuralNetwork(net_size, input_size)
print(init_net.weights)
print()
new_net = random_move(init_net, step_length=1)
print(new_net.weights)
# -
# Function to maximize:
# +
# Input
x = np.linspace(-7, 7, 20)
# Target
y = np.sin(x) * 0.5
def f(net: NeuralNetwork, inputs=x, targets=y) -> Value:
outputs = np.array([net.output([__])[0] for __ in inputs])
erf = error_function(outputs, targets)
return -1 * np.log(erf)
# + active=""
# x = np.array([1, 0])
# y = np.array([2, -5])
#
# error_function(x, y)
# -
# ### Do MCMC
# +
# Do mcmc
chain_list = []
t_begin = time()
for step in range(chain_num):
init_net = NeuralNetwork(net_size, input_size)
net_chain = mc.single_chain_mcmc(
f, random_move, init_net,
tolerence=tolerence,
max_break_counter=max_break_counter,
iterations = 10 ** 10
)
chain_list.append(net_chain)
bc = mc.best_chain(chain_list)
best_net = bc[-1][0]
ef_value = bc[-1][1]
t_end = time()
print('time spent by fitting: ', t_end - t_begin)
# -
# ### Show Result
# +
error_function_values = [__[1] for __ in bc]
plt.plot([i for i in range(len(error_function_values))],
error_function_values)
plt.xlabel('chain_node of best_chain')
plt.ylabel('-1 * log(error_function_value)')
plt.show()
plot_x = np.linspace(-6.0,6.0,150)
plot_y_ann = [best_net.output([__]) for __ in plot_x]
plot_y_target = np.sin(plot_x) * 0.5
plt.plot(plot_x, plot_y_target, '-',
plot_x, plot_y_ann, '.', alpha = 0.3
)
plt.legend(['train target', 'net output'])
plt.show()
# -
# **Far from good.**
# Further exame
# +
test_net = random_move(best_net, 0.01)
print('test: ', f(test_net))
print('best: ', f(best_net))
print('test of relative gain: ', mc.relative_gain(f(best_net), f(test_net)))
# -
# ### Print out Fitted Weights
# +
for i in range(len(best_net.layers[0])):
print(best_net.layers[0][i].weights)
print()
for i in range(len(best_net.layers[1])):
print(best_net.layers[1][i].weights)
| test/test_mcmc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# Three groups: microsatellite instability high (MSI-H), microsatellite instability low (MSI-L) and microsatellite stable (MSS).
#
# Two sets of measurements: 7_marker, 5_marker. The original report consider one sample as MSI if both indicate positive.
# +
import pandas as pd
import matplotlib as mpl
import numpy as np
import matplotlib.pyplot as plt
import scipy.stats
# mpl.rcParams['figure.dpi'] = 600
# -
df = pd.read_excel("41591_2016_BFnm4191_MOESM27_ESM.xlsx", index=1)
df
df[df['Tumor Type'] == 'COAD']['MOSAIC classification'].value_counts().plot(kind="bar")
# +
f = lambda x: x[0:3]
msi_mask = (df['MOSAIC classification'] == 'MSI-H') & (df['Tumor Type'] == 'COAD')
mss_mask = (df['MOSAIC classification'] == 'MSS') & (df['Tumor Type'] == 'COAD')
msi_patient_id = df['Sample Name'][msi_mask].to_frame()
mss_patient_id = df['Sample Name'][mss_mask].to_frame()
msi_patient_id['microsatellite'] = 'MSI-H'
mss_patient_id['microsatellite'] = 'MSS'
microsatellite_label_df = msi_patient_id.append(mss_patient_id)
# -
data = pd.read_csv("TCGA.Kallisto.fullIDs.cibersort.relative.tsv", sep="\t")
data["SampleID"] = data["SampleID"].apply(lambda x: x.replace('.', '-'))
data["PatientID"] = data["SampleID"].apply(lambda x: '-'.join(x.split('-')[0:3]))
merged = data.merge(microsatellite_label_df, left_on="PatientID", right_on='Sample Name')
merged.microsatellite.value_counts().plot(kind="bar")
# +
cell_types = ['B.cells.naive', 'B.cells.memory', 'Plasma.cells', 'T.cells.CD8',
'T.cells.CD4.naive', 'T.cells.CD4.memory.resting',
'T.cells.CD4.memory.activated', 'T.cells.follicular.helper',
'T.cells.regulatory..Tregs.', 'T.cells.gamma.delta', 'NK.cells.resting',
'NK.cells.activated', 'Monocytes', 'Macrophages.M0', 'Macrophages.M1',
'Macrophages.M2', 'Dendritic.cells.resting',
'Dendritic.cells.activated', 'Mast.cells.resting',
'Mast.cells.activated', 'Eosinophils', 'Neutrophils']
#merged['Leukocytes.all'] = merged[cell_types].sum(1)
merged['T.cells.all'] = merged[['T.cells.CD8',
'T.cells.CD4.naive',
'T.cells.CD4.memory.resting',
'T.cells.CD4.memory.activated',
'T.cells.follicular.helper',
'T.cells.regulatory..Tregs.',
'T.cells.gamma.delta']].sum(1)
merged['B.cells.all'] = merged[['B.cells.naive', 'B.cells.memory']].sum(1)
merged['Nk.cells.all'] = merged[['NK.cells.resting', 'NK.cells.activated']].sum(1)
merged['Macrophages.all'] = merged[['Macrophages.M0', 'Macrophages.M1', 'Macrophages.M2']].sum(1)
merged['Dendritic.cells.all'] = merged[['Dendritic.cells.resting', 'Dendritic.cells.activated']].sum(1)
merged['Mast.cells.all'] = merged[['Mast.cells.resting', 'Mast.cells.activated']].sum(1)
augmented_cell_types = cell_types + ['T.cells.all', 'B.cells.all', 'Nk.cells.all', 'Macrophages.all',
'Dendritic.cells.all', 'Mast.cells.all']
merged
# -
mss = merged[merged.microsatellite == 'MSS'][augmented_cell_types]
msi = merged[merged.microsatellite == 'MSI-H'][augmented_cell_types]
mss
# # Results
#
# ## MSS
# +
a = 0.05
sanitize = lambda x: 0.0 if x < 0 else 1.0 if x > 1 else x
res_mss = pd.DataFrame(index = cell_types, columns = ['mean', 'mean lower', 'mean upper', 'sd', 'sd lower', 'sd upper'])
n = mss.shape[0];
# Mean
res_mss['mean'] = mss.mean(axis=0)
res_mss['sd'] = mss.std(ddof=1, axis=0)
# Mean CI
err = scipy.stats.t.ppf(1 - a / 2, n - 1) * res_mss['sd'] / np.sqrt(n)
res_mss['mean lower'] = (res_mss['mean'] - err).apply(sanitize)
res_mss['mean upper'] = (res_mss['mean'] + err).apply(sanitize)
# Standard deviation CI
res_mss['sd lower'] = np.sqrt((n - 1) * res_mss['sd'] ** 2 / scipy.stats.chi2.ppf(1 - a / 2, n - 1))
res_mss['sd upper'] = np.sqrt((n - 1) * res_mss['sd'] ** 2 / scipy.stats.chi2.ppf(a / 2, n - 1))
res_mss
# -
# ## MSI
# +
a = 0.05
sanitize = lambda x: 0.0 if x < 0 else 1.0 if x > 1 else x
res_msi = pd.DataFrame(index = cell_types, columns = ['mean', 'mean lower', 'mean upper', 'sd', 'sd lower', 'sd upper'])
n = msi.shape[0];
# Mean
res_msi['mean'] = msi.mean(axis=0)
res_msi['sd'] = msi.std(ddof=1, axis=0)
# Mean CI
err = scipy.stats.t.ppf(1 - a / 2, n - 1) * res_msi['sd'] / np.sqrt(n)
res_msi['mean lower'] = (res_msi['mean'] - err).apply(sanitize)
res_msi['mean upper'] = (res_msi['mean'] + err).apply(sanitize)
# Standard deviation CI
res_msi['sd lower'] = np.sqrt((n - 1) * res_msi['sd'] ** 2 / scipy.stats.chi2.ppf(1 - a / 2, n - 1))
res_msi['sd upper'] = np.sqrt((n - 1) * res_msi['sd'] ** 2 / scipy.stats.chi2.ppf(a / 2, n - 1))
res_msi
# -
# ## Figures
# +
x_map = {v: i for i, v in enumerate(augmented_cell_types)}
offset_map = {
'mss': -0.15,
'msi': 0.15}
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
color_map = {'mss': colors[0],
'msi': colors[1]}
fig = plt.figure(figsize=(10, 3))
ax = fig.add_subplot(1, 1, 1)
n = mss.shape[0];
mean = mss.mean(axis=0)
sd = mss.std(ddof=1, axis=0)
err = scipy.stats.t.ppf(1 - a / 2, n - 1) * sd / np.sqrt(n)
x = [v + offset_map['mss'] for v in x_map.values()]
ax.errorbar(x, mean, yerr=sd, fmt='.', color = color_map['mss'], ecolor = 'darkgray', elinewidth=3.0)
ax.errorbar(x, mean, yerr=err, fmt="None", color = color_map['mss'], ecolor = 'black', elinewidth=1.0)
n = msi.shape[0];
mean = msi.mean(axis=0)
sd = msi.std(ddof=1, axis=0)
err = scipy.stats.t.ppf(1 - a / 2, n - 1) * sd / np.sqrt(n)
x = [v + offset_map['msi'] for v in x_map.values()]
ax.errorbar(x, mean, yerr=sd, fmt='.', color = color_map['msi'], ecolor = 'darkgray', elinewidth=3.0)
ax.errorbar(x, mean, yerr=err, fmt="None", color = color_map['msi'], ecolor = 'black', elinewidth=1.0)
# Hide the right and top spines
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Only show ticks on the left and bottom spines
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
current_ylim = ax.get_ylim()
for v in x_map.values():
if v % 2 == 0:
ax.fill_between([v - 0.5, v + 0.5], current_ylim[0], current_ylim[1], facecolor='lightgray', alpha=0.2)
ax.set_ylim(current_ylim)
ax.set_xticks(list(x_map.values()))
ax.tick_params(axis='x', which = 'both', labelbottom=None)
ax.set_ylabel('Abundance')
ax.set_xlim(-0.5, len(x_map) - 0.5)
# Ticks
ax.tick_params(axis='x', which = 'both', labelbottom=True)
ax.set_xticks(list(x_map.values()))
ax.set_xticklabels(list(x_map.keys()), rotation=30, ha='right')
legend_elements = [mpl.lines.Line2D([0], [0], marker='.', color='w', markerfacecolor=color_map[i], label=i, markersize=15)
for i in color_map]
ax.legend(handles=legend_elements, loc='upper left')
# -
len(cell_types)
| python/statistics/.ipynb_checkpoints/TCGA-MSI-MSS-relative-figure-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example powerbook usage
#
# In this example, we're going to generate some figures and some text, and compile them into a nice Powerpoint slideshow as we go. Every time you rerun this notebook, a new Powerpoint will be generated with the latest analysis!
# # !pip3 install python-pptx
import powerbook
import matplotlib.pyplot as plt
import numpy as np
# First, we create the Powerbook. You could optionally pass an existing slideshow pptx file to the Powerbook constructor in order to _append_ slides to the end of an existing show, with `p = powerbook.Powerbook("my_cool_slideshow.pptx")`, but here we're creating one from scratch.
P = powerbook.Powerbook()
# We can add a title slide like so:
P.add_title_slide(title="My Fascinating Discoveries", subtitle="By <NAME>")
# Next we'll start our analysis:
my_data = np.random.randint(0, 255, size=(100, 2))
f = plt.figure(figsize=(5, 2.5), dpi=150)
plt.scatter(my_data[:,0], my_data[:, 1])
plt.xlabel("Voominess ($\phi/g^2$)")
plt.ylabel("Zoopiness (zph)")
# Wow. Now _that_ is an important figure! Let's add it to our powerbook.
P.add_image_slide(title="Get a load of this dataset!", image=f)
# Yes indeed! We can pass a `matplotlib.Figure` directly into Powerbook.
#
# Let's see how our slideshow is going:
len(P.slides)
# Nice! Let's add a final discussion slide to wrap things up:
P.add_text_slide(
"Discussion",
"""
These data are super interesting.
* I wonder what it means!
* Can't wait to do more science:
* Tomorrow
* Next week
* (maybe less on the weekend)
Thanks for coming to my TED Talk!
"""
)
# And finally, we can save this document out to disk:
P.save("MyAmazingSlideshow.pptx")
| examples/basic-example/Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import pickle
import datetime
from sklearn.model_selection import train_test_split
# %matplotlib inline
# %config InlineBackend.figure_formats = {'png','retina'}
def import_data():
df = pd.read_csv('data/final_df2.csv')
df = df.iloc[:,1:]
return df
# +
def fulldf(df):
df.snowfall = df.snowfall.replace(["T"," T"],0.05)
df.preciptotal = df.preciptotal.replace(["T"," T"],0.005)
df.depart = pd.to_numeric(df.depart, errors='coerce')
df = type_change_numeric(df,[ 'store_nbr', 'item_nbr', 'units', 'station_nbr', 'tmax', 'tmin',
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
'avgspeed', 'resultspeed', 'resultdir' ])
df['date'] = pd.to_datetime(df['date'])
df["day_of_year"] = df['date'].dt.dayofyear
df["year"] = df['date'].dt.year
df["month"] = df["date"].dt.month
for idx in range(5, 8):
df.iloc[:,idx].fillna(df.groupby(["day_of_year","store_nbr"])[df.columns[idx]].\
transform('mean'), inplace=True)
for idx in range(16, 23):
df.iloc[:,idx].fillna(df.groupby(["day_of_year","store_nbr"])[df.columns[idx]].\
transform('mean'), inplace=True)
add_depart1(df)
return df
# for column in item37.columns:
# item37[column].interpolate()
# -
def type_change_numeric(df, ls = []):
#ls에 있는 column name은 numeric형으로 바꾸지 않는다.
cols = df.columns
for i in cols:
if i in ls:
#df = df.replace(["M",None], '')
df.snowfall = df.snowfall.replace(["T"," T"],0.05)
df.preciptotal = df.preciptotal.replace(["T"," T"],0.005)
df[i] = pd.to_numeric(df[i], errors='coerce')
return df
def add_depart1(x):
x.depart.fillna(x.tavg - x.groupby(["day_of_year","store_nbr"])["tavg"].transform('mean'),inplace = True)
x.depart = x.depart.round(2)
return x.sort_values(["store_nbr","date","item_nbr"])
def reorder_df(df):
#Column 정렬 (y값을 마지막으로 ) 후 FG+ -> FG2 변환 (formula에 인식시키기위해 )
new_order = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin',
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
'avgspeed', 'resultspeed', 'TS', 'GR', 'RA', 'DZ', 'SN',
'SG', 'GS', 'PL', 'FG+', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ',
'MI', 'PR', 'BC', 'BL', 'VC', 'day_of_year', 'year', 'month', 'units' ]
df = df[new_order]
df.columns = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin',
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'snowfall', 'preciptotal', 'stnpressure', 'sealevel',
'avgspeed', 'resultspeed', 'TS', 'GR', 'RA', 'DZ', 'SN',
'SG', 'GS', 'PL', 'FG2', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ',
'MI', 'PR', 'BC', 'BL', 'VC', 'day_of_year', 'year', 'month', 'units' ]
return df
def add_cat_columns(df):
df['date'] = pd.to_datetime(df['date'])
# df['sunrise'] = pd.to_datetime(df['sunrise'], format='%H%M')
# df['sunset'] = pd.to_datetime(df['sunset'], format='%H%M')
blackfriday = ["2012-11-21","2012-11-22","2012-11-23", "2012-11-24","2012-11-25",
"2013-11-27","2013-11-28", "2013-11-29","2013-11-30","2013-11-31",
"2014-11-26", "2014-11-27", "2014-11-28","2014-11-29","2014-11-30"]
df["week_day_name"] = df['date'].dt.weekday_name
df['is_week'] = False
df.is_week[df['week_day_name'] == 'Sunday'] = True
df.is_week[df['week_day_name'] == 'Saturday'] = True
df.is_week[df['week_day_name'] == 'Friday'] = True
df["is_blackfriday"] = df.date.apply(lambda x : str(x)[:10] in blackfriday).astype(int)
holiday = ["2012-01-02","2012-01-16","2012-02-14", "2012-02-20",\
"2012-05-28","2012-07-04","2012-09-03", "2012-10-08",\
"2012-11-12", "2012-12-24","2012-12-25", "2012-12-31",\
"2013-01-01","2013-01-21", "2013-02-14",\
"2013-05-27", "2013-07-04", "2013-09-02", \
"2013-11-24", "2013-11-25","2013-12-24", "2013-12-31",\
"2014-01-01", "2014-01-20", "2014-02-14",\
"2014-05-26", "2014-07-04", "2014-09-01", "2014-10-13",\
"2014-11-11", "2014-12-24", "2014-12-25", "2014-12-31"]
df["is_holiday"] = df.date.apply(lambda x : str(x)[:10] in holiday).astype(int)
df['hardrain'] = [(((4 if i > 8 else 3) if i > 6 else 2) if i > 1 else 1) if i > 0 else 0 for i in df['preciptotal']]
# rain 1 snow 2
# 득정조건열 추가 ()
df['hardsnow'] = [(( 3 if i > 3.5 else 2) if i > 1 else 1) if i > 0 else 0 for i in df['snowfall']]
df['log_units'] = df.units.apply(lambda x: np.log(x + 1)).astype(float)
return df
def df_sampling(df):
new_order = ['date', 'store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin', 'tavg',
'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'preciptotal', 'stnpressure', 'sealevel', 'avgspeed',
'resultspeed', 'TS', 'GR', 'RA', 'DZ', 'SN', 'SG', 'GS',
'PL', 'FG2', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', 'SQ', 'FZ', 'MI', 'PR',
'BC', 'BL', 'VC', 'day_of_year', 'year', 'month',
'week_day_name', 'is_week', 'is_holiday', 'is_blackfriday', 'hardrain', 'hardsnow',
'log_units', 'units' ]
df = df[new_order]
X, y = df.iloc[:,:-1], df.units
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3, random_state = 2018)
train = pd.concat([X_train, y_train] ,axis = 1)
train = train.sort_values(by=['date', 'store_nbr', 'item_nbr']).reset_index(drop= True)
X_test = X_test.sort_index()
y_train = y_train.sort_index()
y_test = y_test.sort_index()
return train, y_train, X_test ,y_test
data = import_data()
data_t = fulldf(data)
data_t = reorder_df(data_t)
data_t = add_cat_columns(data_t)
del data_t['snowfall']
data_t = data_t.dropna()
train0, y_train, X_test, y_test = df_sampling(data_t)
train = sm.add_constant(train0)
results_OLS_stormy
params
len(results_OLS.params.values)
results_OLS_stormy.params.values = np.array(range(1,149))
train.columns
model_OLS_stormy = sm.OLS.from_formula('log_units ~ C(item_nbr)+C(store_nbr) + C(is_week) + C(is_holiday) + C(is_blackfriday) + C(0', data = train)
results_OLS = model_OLS.fit()
print(results_OLS.summary())
sns.jointplot(results_OLS.predict(X_test), np.log(y_test + 1))
plt.show()
# +
# hat = influence.hat_matrix_diag
# plt.stem(hat)
# plt.show()
# +
sm.graphics.influence_plot(results_OLS, plot_alpha=0.3)
plt.show()
# -
influence = results_OLS.get_influence()
cooks_d2, pvals = influence.cooks_distance
fox_cr = 4 / (len(y_train) - 2)
idx = np.where(cooks_d2 > fox_cr)[0]
X_new.columns
def remove_outlier(df, idx):
idx_t = list(set(df.index) - set(idx))
X_new = df.T[idx_t].T
X_new.const = X_new.const.astype(int)
X_new['date'] = pd.to_datetime(X_new['date'])
X_new["day_of_year"] = X_new['date'].dt.dayofyear
X_new["year"] = X_new['date'].dt.year
X_new["month"] = X_new["date"].dt.month
X_new.store_nbr = X_new.store_nbr.astype(int)
X_new.staion_nbr = X_new.staion_nbr.astype(int)
X_new.tmax = X_new.tmax.astype(float)
X_new.tmin = X_new.tmax.astype(float)
X_new.tavg = X_new.tmax.astype(float)
numeric_columns = ['const','store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin', \
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'preciptotal', \
'stnpressure', 'sealevel', 'avgspeed', 'resultspeed', 'day_of_year', 'year', \
'month', 'week_day_name', 'is_week', 'is_holiday', 'is_blackfriday', \
'hardrain', 'hardsnow', 'log_units', 'units']
boo_columns = ['TS', 'GR', 'RA', \
'DZ', 'SN', 'SG', 'GS', 'PL', 'FG2', 'FG', 'BR', 'UP', 'HZ', 'FU', 'DU', \
'SQ', 'FZ', 'MI', 'PR', 'BC', 'BL', 'VC',]
for col in numeric_columns:
X_new[col] = pd.to_numeric(X_new[col], errors='coerce')
for col in boo_columns:
X_new[col] = X_new[col].astype(bool)
blackfriday = ["2012-11-21","2012-11-22","2012-11-23", "2012-11-24","2012-11-25",
"2013-11-27","2013-11-28", "2013-11-29","2013-11-30","2013-11-31",
"2014-11-26", "2014-11-27", "2014-11-28","2014-11-29","2014-11-30"]
X_new["week_day_name"] = X_new['date'].dt.weekday_name
X_new['is_week'] = False
X_new.is_week[X_new['week_day_name'] == 'Sunday'] = True
X_new.is_week[X_new['week_day_name'] == 'Saturday'] = True
X_new.is_week[X_new['week_day_name'] == 'Friday'] = True
X_new["is_blackfriday"] = X_new.date.apply(lambda x : str(x)[:10] in blackfriday).astype(int)
holiday = ["2012-01-02","2012-01-16","2012-02-14", "2012-02-20",\
"2012-05-28","2012-07-04","2012-09-03", "2012-10-08",\
"2012-11-12", "2012-12-24","2012-12-25", "2012-12-31",\
"2013-01-01","2013-01-21", "2013-02-14",\
"2013-05-27", "2013-07-04", "2013-09-02", \
"2013-11-24", "2013-11-25","2013-12-24", "2013-12-31",\
"2014-01-01", "2014-01-20", "2014-02-14",\
"2014-05-26", "2014-07-04", "2014-09-01", "2014-10-13",\
"2014-11-11", "2014-12-24", "2014-12-25", "2014-12-31"]
X_new["is_holiday"] = X_new.date.apply(lambda x : str(x)[:10] in holiday).astype(int)
X_new['hardrain'] = [(((4 if i > 8 else 3) if i > 6 else 2) if i > 1 else 1) if i > 0 else 0 for i in X_new['preciptotal']]
# rain 1 snow 2
# 득정조건열 추가 ()
X_new['log_units'] = X_new.units.apply(lambda x: np.log(x + 1)).astype(float)
return X_new
# +
X_new.dtypes
# -
X_new.columns
model_OLS2 = sm.OLS.from_formula('log_units ~ C(item_nbr)+C(store_nbr) + C(is_week) + C(is_holiday) + C(FG2) + C(is_blackfriday) + 0', data = X_new)
results_OLS2 = model_OLS2.fit()
print(results_OLS2.summary())
X_new[X_new.hardrain > 2]
model_OLS3 = sm.OLS.from_formula('log_units ~ C(item_nbr) + C(store_nbr) + scale(preciptotal) ++ C(is_week) + C(is_holiday) + + C(TS) + C(DZ) + C(PL) + C(HZ) + C(FU) + C(FZ) + C(is_blackfriday) +C(hardrain) + C(hardsnow) + 0', data = X_new)
results_OLS3 = model_OLS3.fit()
print(results_OLS3.summary())
data_T
data2 = import_data()
data_t2 = fulldf(data2)
data_t2 = reorder_df(data_t2)
data_t2 = add_cat_columns(data_t2)
data_t2.columns
data_t2.isnull().any()
results_OLS3.predict(data_t2)
plt.scatter(X.)
influence.
data_t = fulldf(data)
len()
data_t = fulldf(data)
data_t = reorder_df(data_t)
data_t = add_cat_columns(data_t)
data_t = data_t.dropna()
data_t.dtypes
# +
model_OLS_stormy = sm.OLS.from_formula('log_units ~ C(item_nbr) + C(store_nbr) + scale(preciptotal) + C(FZ) + C(is_blackfriday) +C(hardrain) + C(hardsnow) + 0', data = data_t)
results_OLS_stormy = model_OLS_stormy.fit()
print(results_OLS_stormy.summary())
# -
cooks_d2, pvals = influence.cooks_distance
fox_cr = 4 / (len(y_train) - 2)
idx = np.where(cooks_d2 > fox_cr)[0]
model_OLS4 = sm.OLS.from_formula('log_units ~ C(item_nbr)+C(store_nbr) + C(is_week) + C(is_holiday) + C(FG2) + C(is_blackfriday) + 0', data = X_test)
results_OLS4 = model_OLS4.fit()
print(results_OLS4.summary())
influence2 = results_OLS4.get_influence()
cooks_d2, pvals = influence2.cooks_distance
fox_cr = 4 / (len(y_test) - 2)
idx2 = np.where(cooks_d2 > fox_cr)[0]
len(X_new), len()
influence3 = results_OLS4.get_influence()
hat = influence3.hat_matrix_diag
X0 = results_OLS4.predict(X_test).reindex()
y0 = np.log(y_test + 1).reindex()
plt.scatter(X0, y0 , alpha=0.5)
plt.scatter(X0[idx2], y0[idx2], s=20, c="r")
results_OLS4.params
ax = plt.subplot()
plt.scatter(X0, y0)
idx_n = hat > 0.05
plt.scatter(X0[idx], y0[idx], s=20, c="r", alpha=0.5)
plt.show()
full_frame = pd.read_csv('data/data.csv')
del full_frame['Unnamed: 0']
len(full_frame)
full_frame.snowfall = full_frame.snowfall.replace(["T"," T"],0.05)
full_frame.preciptotal = full_frame.preciptotal.replace(["T"," T"],0.005)
full_frame.snowfall = full_frame.snowfall.replace(["M"," M"], 0)
full_frame.preciptotal = full_frame.preciptotal.replace(["M"," M"], 0)
ls = ['store_nbr', 'item_nbr', 'station_nbr', 'tmax', 'tmin', \
'tavg', 'depart', 'dewpoint', 'wetbulb', 'heat', 'cool', 'preciptotal', \
'stnpressure', 'sealevel', 'avgspeed', 'resultspeed', 'units' ]
cols = full_frame.columns
for i in cols:
if i in ls:
#df = df.replace(["M",None], '')
# full_frame.snowfall = full_frame.snowfall.replace(["T"," T"],0.05)
# full_frame.preciptotal = full_frame.preciptotal.replace(["T"," T"],0.005)
full_frame[i] = pd.to_numeric(full_frame[i], errors='coerce')
full_frame['snowfall'] = pd.to_numeric(full_frame['snowfall'], errors='coerce')
full_frame.dtypes
full_frame.head()
full_frame['date'] = pd.to_datetime(full_frame['date'])
full_frame["day_of_year"] = full_frame['date'].dt.dayofyear
full_frame["year"] = full_frame['date'].dt.year
full_frame["month"] = full_frame["date"].dt.month
# +
blackfriday = ["2012-11-21","2012-11-22","2012-11-23", "2012-11-24","2012-11-25",
"2013-11-27","2013-11-28", "2013-11-29","2013-11-30","2013-11-31",
"2014-11-26", "2014-11-27", "2014-11-28","2014-11-29","2014-11-30"]
full_frame["week_day_name"] = full_frame['date'].dt.weekday_name
full_frame['is_week'] = False
full_frame.is_week[full_frame['week_day_name'] == 'Sunday'] = True
full_frame.is_week[full_frame['week_day_name'] == 'Saturday'] = True
full_frame.is_week[full_frame['week_day_name'] == 'Friday'] = True
full_frame["is_blackfriday"] = full_frame.date.apply(lambda x : str(x)[:10] in blackfriday).astype(int)
holiday = ["2012-01-02","2012-01-16","2012-02-14", "2012-02-20",\
"2012-05-28","2012-07-04","2012-09-03", "2012-10-08",\
"2012-11-12", "2012-12-24","2012-12-25", "2012-12-31",\
"2013-01-01","2013-01-21", "2013-02-14",\
"2013-05-27", "2013-07-04", "2013-09-02", \
"2013-11-24", "2013-11-25","2013-12-24", "2013-12-31",\
"2014-01-01", "2014-01-20", "2014-02-14",\
"2014-05-26", "2014-07-04", "2014-09-01", "2014-10-13",\
"2014-11-11", "2014-12-24", "2014-12-25", "2014-12-31"]
full_frame["is_holiday"] = full_frame.date.apply(lambda x : str(x)[:10] in holiday).astype(int)
full_frame['hardrain'] = [(((4 if i > 8 else 3) if i > 6 else 2) if i > 1 else 1) if i > 0 else 0 for i in full_frame['preciptotal']]
# rain 1 snow 2
# 득정조건열 추가 ()
full_frame['hardsnow'] = [(( 3 if i > 3.5 else 2) if i > 1 else 1) if i > 0 else 0 for i in full_frame['snowfall']]
full_frame['log_units'] = full_frame.units.apply(lambda x: np.log(x + 1)).astype(float)
# -
full_frame.preciptotal.max()
full_frame.iloc[0,:]
results_OLS3.predict(full_frame)
| team_project/OLS-almostfinal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lab 3: Functions and matrix algebra
# **Data Science for Biologists** • University of Washington • BIOL 419/519 • Winter 2019
#
# Course design and lecture material by [<NAME>](https://github.com/bwbrunton) and [<NAME>](https://github.com/kharris/). Lab design and materials by [<NAME>](https://github.com/eleanorlutz/), with helpful comments and suggestions from Bing and Kam.
#
# ### Table of Contents
# 1. Matrix algebra in Python
# 2. Review of functions
# 3. Bonus exercises
#
# ### Helpful Resources
# - [A Primer on Matrices](https://see.stanford.edu/materials/lsoeldsee263/Additional1-notes-matrix-primer.pdf) by <NAME>
# - [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by <NAME>
# - [Python Basics Cheat Sheet](https://datacamp-community-prod.s3.amazonaws.com/e30fbcd9-f595-4a9f-803d-05ca5bf84612) by Python for Data Science
# - [Jupyter Notebook Cheat Sheet](https://datacamp-community-prod.s3.amazonaws.com/48093c40-5303-45f4-bbf9-0c96c0133c40) by Python for Data Science
# - [Numpy Cheat Sheet](https://datacamp-community-prod.s3.amazonaws.com/e9f83f72-a81b-42c7-af44-4e35b48b20b7) by Python for Data Science
#
# ## Lab 3 Part 1: Matrix algebra in Python
#
# In Python we can do matrix algebra without having to calculate each value by hand. Given the numpy arrays `A` and `B`, `A+B` or `A-B` returns the elementwise addition or subtraction, and `A*B` returns the elementwise multiplication. *Elementwise* operations are performed on two matrices of the exact same shape, and produces an output matrix that also has the same dimensions. In elementwise addition, each value from matrix `A` is added to the single value in matrix `B` that is in the same position. Similarly, elementwise multiplication multiplies each value in `A` by the corresponding value in `B`.
#
# The elementwise addition for the following two matrices `A` and `B`, (calculated in Python as `A+B` ), looks like this:
# 
#
# A **matrix product** is different from elementwise matrix multiplication. A matrix product calculates the product of each value in the *rows* of the first matrix against the *columns* of the second matrix. Matrix products can only be calculated if the **number of columns in the first matrix equals the number of rows of the second matrix**.
#
# The matrix product for `A` and `B` looks like this:
# 
#
# To calculate the matrix product for `A` and `B` in Python, we use the function `np.dot(A, B)`:
import numpy as np
# +
A = np.array([[0, 1],
[2, 3]])
B = np.array([[4, 5],
[6, 7]])
print(np.dot(A,B))
# -
# Although Python runs all of these calculations without requiring our input, we can also calculate the matrix product the long way in Python. This is probably not something you'll need to do in your research, but it's a good learning exercise to practice array indexing and matrix algebra.
#
# To use array indexing to calculate each value in the `AB` matrix product, this is what we need to type in Python:
output = np.zeros([2, 2])
output[0, 0] = A[0, 0]*B[0, 0] + A[0, 1]*B[1, 0]
output[0, 1] = A[0, 0]*B[0, 1] + A[0, 1]*B[1, 1]
output[1, 0] = A[1, 0]*B[0, 0] + A[1, 1]*B[1, 0]
output[1, 1] = A[1, 0]*B[0, 1] + A[1, 1]*B[1, 1]
print(output)
# **Exercise 1:** To practice indexing and matrix algebra, go through a similar process for the equation `A + B`. Your code should create a new matrix called `output` of the correct size, and then fill each value in the `output` matrix individually using indexing. Check that your answer matches the Numpy matrix addition `A+B`.
# +
A = np.array([[0, 1],
[2, 3]])
B = np.array([[4, 5],
[6, 7]])
# Your code here
# -
print(A + B)
# ## Lab 3 Part 2: Review of functions
#
# Functions enclose a set of operations into a package that can be easily reused. We have actually already been using functions imported from other libraries many times throughout the quarter. For example, `np.mean(x)` is a function that returns the mean of variable `x` and `np.median(x)` is another function that returns the median. Functions can do many things, from calculating statistics to plotting figures. In the Matplotlib library, `plt.hist(x)` is a function that returns a histogram of values in variable `x`.
#
# If you were to write your own version of the `np.sum()` function, it might look something like this:
# Define a function that sums a list of numbers.
def sum_values(values):
sum_of_values = 0
for value in values:
sum_of_values = sum_of_values + value
return sum_of_values
# After defining the function by running the code block above, we can use our function to calculate the sum of any list of values:
print(sum_values([0, 1, 2]))
print(sum_values([-2, -5, -6]))
print(sum_values([100, 100, 100]))
print(sum_values([2]))
# Similarly, we can write our own function to find the length of a list, instead of using the built-in `len()` function:
# Define a function that counts the number of values in a list of numbers.
def count_values(values):
count_of_values = 0
for value in values:
count_of_values = count_of_values + 1
return count_of_values
print(count_values([0, 1, 2]))
print(count_values([-2, -5, -6]))
print(count_values([100, 100, 100]))
print(count_values([2]))
# **Exercise 2:** Write a new function called `my_mean` that takes a list of numbers called `values` as an input, and returns the mean. Your function should use the previous two functions, `sum_values` and `count_values`. Test your function using the code block given below. All five statements should print `True` if you have written your function correctly.
print(my_mean([1, 3, 2, 0]) == 1.5)
print(my_mean([1, 3, 2]) == 2)
print(my_mean([-3, -4, -7, -8, -9, -10]) == -7)
print(my_mean([1]) == 1)
print(my_mean([1, -1]) == 0)
# **Exercise 3:** Write a new function called `my_median` that takes a list of numbers as an input, and returns the median of the set of values. You should not use the builtin function `np.median()`. Test your function using the code block given below. All five statements should print `True` if you have written your function correctly.
print(my_median([1, 3, 2, 0]) == 1.5)
print(my_median([1, 3, 2]) == 2)
print(my_median([-3, -4, -7, -8, -10, -10]) == -7.5)
print(my_median([1]) == 1)
print(my_median([1, -1]) == 0)
# ## Lab 3 Bonus exercises
#
# **Bonus Exercise 1:** Write a function that calculates the elementwise sum of any two Numpy arrays. Your function should also check that both matrices are the same size, and print an error message if this is not the case. It may be helpful to use the code you wrote in Exercise 1 for reference.
# +
# Check that your function works as expected:
A = np.array([[0, 1],
[2, 3]])
B = np.array([[4, 5],
[6, 7]])
print(elementwise_sum(A,B))
# -
# **Bonus Exercise 2:** Write a function that calculates the matrix product of any two Numpy arrays. Your function should also check that multiplication is possible, and print an error message if the matrices are not the correct shape.
# +
# Check that your function works as expected:
A = np.array([[0, 1],
[2, 3]])
B = np.array([[4, 5],
[6, 7]])
print(matrix_product(A,B))
# -
| Lab_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Undistort & Transform
#
# Now lets combine everything we've learned so far.
#
# Pass in your image into this function
# Write function to do the following steps
# 1) Undistort using mtx and dist
# 2) Convert to grayscale
# 3) Find the chessboard corners
# 4) If corners found:
# a) draw corners
# b) define 4 source points src = np.float32([[,],[,],[,],[,]])
# #Note: you could pick any four of the detected corners
# # as long as those four corners define a rectangle
# #One especially smart way to do this would be to use four well-chosen
# # corners that were automatically detected during the undistortion steps
# #We recommend using the automatic detection of corners in your code
# c) define 4 destination points dst = np.float32([[,],[,],[,],[,]])
# d) use cv2.getPerspectiveTransform() to get M, the transform matrix
# e) use cv2.warpPerspective() to warp your image to a top-down view
#
# +
import pickle
import cv2
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# First lets read in the camera matrix we made earlier
dist_pickle = pickle.load(open("camera_cal/dist_pickle.p","rb"))
mtx = dist_pickle['mtx']
dist = dist_pickle["dist"]
# Pass your image
img = cv2.imread('camera_cal/test_image.jpg')
nx = 9 # numbers of corners on X
ny = 6 # numbers of corners on Y
# Write function to do the following steps
def corners_unwarp(img,nx,ny,mtx,dist):
#1) Undistort using mtx and dist
img2 = cv2.undistort(img,mtx,dist,None,mtx)
#2) Convert to grayscale
gray = cv2.cvtColor(img2,cv2.COLOR_RGB2GRAY)
#3) Find the chessboard corners
ret,corners = cv2.findChessboardCorners(gray,(nx,ny),None)
#4) If corners found:
if ret == True:
#a) draw corners
img2 = cv2.drawChessboardCorners(img2,(nx,ny),corners,ret)
#b) define 4 source points src = np.float32([[,],[,],[,],[,]])
#Note: you could pick any four of the detected corners
# as long as those four corners define a rectangle
#One especially smart way to do this would be to use four well-chosen
# corners that were automatically detected during the undistortion steps
#We recommend using the automatic detection of corners in your code
src = np.float32(corners[0],corners[nx-1],corners[ny*nx -nx],corners[ny*nx -1])
#c) define 4 destination points dst = np.float32([[,],[,],[,],[,]])
h,w = img.shape[:2]
dst = np.float32([[100,100],[w-100,100],[100,h-100],[w-100,h-100]])
#d) use cv2.getPerspectiveTransform() to get M, the transform matrix
M = cv2.getPerspectiveTransform(src, dst)
#e) use cv2.warpPerspective() to warp your image to a top-down view
warped = cv2.warpPerspective(img2 ,M, (w,h))
return warped, M
top_down ,perspective_M = corners_unwarp(img,nx,ny,mtx,dist)
f,(ax1,ax2) = plt.subplots(1,2,figsize=(20,20))
ax1.set_title('Source')
ax1.imshow(img)
ax2.set_title('Transformed')
ax2.imshow(top_down)
| 02-Camera-Calibration/.ipynb_checkpoints/Undistort&Transform-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.0 ('dhiraj_ml_march')
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
df=pd.read_csv(r"C:\Users\dhire\Documents\Machine_learning_Inuron\ML_Live_Class\data\mouse_viral_study.csv")
df.head()
df.columns
sns.scatterplot(x='Med_1_mL',y='Med_2_mL',data=df,hue='Virus Present')
x=df.drop('Virus Present',axis=1)
y=df['Virus Present']
from sklearn.svm import SVC
a=SVC()
a.get_params()
model=SVC(kernel='poly',C=1000)
model.fit(x,y)
# +
# CODE SOURCE IS DIRECTLY FROM DOCUMENTATION
# https://scikit-learn.org/stable/auto_examples/svm/plot_separating_hyperplane.html
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
def plot_svm_boundary(model,X,y):
X = X.values
y = y.values
# Scatter Plot
plt.scatter(X[:, 0], X[:, 1], c=y, s=30,cmap='seismic')
# plot the decision function
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
xx = np.linspace(xlim[0], xlim[1], 30)
yy = np.linspace(ylim[0], ylim[1], 30)
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
Z = model.decision_function(xy).reshape(XX.shape)
# plot decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
ax.scatter(model.support_vectors_[:, 0], model.support_vectors_[:, 1], s=100,
linewidth=1, facecolors='none', edgecolors='k')
plt.show()
plot_svm_boundary(model, x, y)
# -
model=SVC(kernel='linear',C=0.05)
model.fit(x,y)
# +
# CODE SOURCE IS DIRECTLY FROM DOCUMENTATION
# https://scikit-learn.org/stable/auto_examples/svm/plot_separating_hyperplane.html
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
def plot_svm_boundary(model,X,y):
X = X.values
y = y.values
# Scatter Plot
plt.scatter(X[:, 0], X[:, 1], c=y, s=30,cmap='seismic')
# plot the decision function
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
xx = np.linspace(xlim[0], xlim[1], 30)
yy = np.linspace(ylim[0], ylim[1], 30)
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
Z = model.decision_function(xy).reshape(XX.shape)
# plot decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
ax.scatter(model.support_vectors_[:, 0], model.support_vectors_[:, 1], s=100,
linewidth=1, facecolors='none', edgecolors='k')
plt.show()
plot_svm_boundary(model, x, y)
# -
model=SVC(kernel='rbf',C=1)
model.fit(x,y)
# +
# CODE SOURCE IS DIRECTLY FROM DOCUMENTATION
# https://scikit-learn.org/stable/auto_examples/svm/plot_separating_hyperplane.html
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
def plot_svm_boundary(model,X,y):
X = X.values
y = y.values
# Scatter Plot
plt.scatter(X[:, 0], X[:, 1], c=y, s=30,cmap='seismic')
# plot the decision function
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
xx = np.linspace(xlim[0], xlim[1], 30)
yy = np.linspace(ylim[0], ylim[1], 30)
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
Z = model.decision_function(xy).reshape(XX.shape)
# plot decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
ax.scatter(model.support_vectors_[:, 0], model.support_vectors_[:, 1], s=100,
linewidth=1, facecolors='none', edgecolors='k')
plt.show()
plot_svm_boundary(model, x, y)
# -
model=SVC(kernel='rbf',C=1000)
model.fit(x,y)
# +
# CODE SOURCE IS DIRECTLY FROM DOCUMENTATION
# https://scikit-learn.org/stable/auto_examples/svm/plot_separating_hyperplane.html
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
def plot_svm_boundary(model,X,y):
X = X.values
y = y.values
# Scatter Plot
plt.scatter(X[:, 0], X[:, 1], c=y, s=30,cmap='seismic')
# plot the decision function
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
xx = np.linspace(xlim[0], xlim[1], 30)
yy = np.linspace(ylim[0], ylim[1], 30)
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
Z = model.decision_function(xy).reshape(XX.shape)
# plot decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
ax.scatter(model.support_vectors_[:, 0], model.support_vectors_[:, 1], s=100,
linewidth=1, facecolors='none', edgecolors='k')
plt.show()
plot_svm_boundary(model, x, y)
# +
from sklearn.model_selection import GridSearchCV
svm_model = SVC()
param_grid = {'C': [0.01,0.1, 1], 'kernel': ['linear', 'rbf', 'sigmoid', 'poly']}
# -
grid_model = GridSearchCV(svm_model, param_grid)
grid_model.fit(x,y)
grid_model.best_params_
model=SVC(kernel='linear',C=0.01)
model.fit(x,y)
# +
# CODE SOURCE IS DIRECTLY FROM DOCUMENTATION
# https://scikit-learn.org/stable/auto_examples/svm/plot_separating_hyperplane.html
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
def plot_svm_boundary(model,X,y):
X = X.values
y = y.values
# Scatter Plot
plt.scatter(X[:, 0], X[:, 1], c=y, s=30,cmap='seismic')
# plot the decision function
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
xx = np.linspace(xlim[0], xlim[1], 30)
yy = np.linspace(ylim[0], ylim[1], 30)
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
Z = model.decision_function(xy).reshape(XX.shape)
# plot decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
ax.scatter(model.support_vectors_[:, 0], model.support_vectors_[:, 1], s=100,
linewidth=1, facecolors='none', edgecolors='k')
plt.show()
plot_svm_boundary(model, x, y)
# -
| SVM/SVM_project1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Cart-pole Balancing Model with Amazon SageMaker and Ray
#
# ---
# ## Introduction
#
# In this notebook we'll start from the cart-pole balancing problem, where a pole is attached by an un-actuated joint to a cart, moving along a frictionless track. Instead of applying control theory to solve the problem, this example shows how to solve the problem with reinforcement learning on Amazon SageMaker and Ray RLlib. You can choose either TensorFlow or PyTorch as your underlying DL framework.
#
# (For a similar example using Coach library, see this [link](../rl_cartpole_coach/rl_cartpole_coach_gymEnv.ipynb). Another Cart-pole example using Coach library and offline data can be found [here](../rl_cartpole_batch_coach/rl_cartpole_batch_coach.ipynb).)
#
# 1. *Objective*: Prevent the pole from falling over
# 2. *Environment*: The environment used in this exmaple is part of OpenAI Gym, corresponding to the version of the cart-pole problem described by Barto, Sutton, and Anderson [1]
# 3. *State*: Cart position, cart velocity, pole angle, pole velocity at tip
# 4. *Action*: Push cart to the left, push cart to the right
# 5. *Reward*: Reward is 1 for every step taken, including the termination step
#
# References
#
# 1. AG Barto, RS Sutton and CW Anderson, "Neuronlike Adaptive Elements That Can Solve Difficult Learning Control Problem", IEEE Transactions on Systems, Man, and Cybernetics, 1983.
# ## Pre-requisites
#
# ### Imports
#
# To get started, we'll import the Python libraries we need, set up the environment with a few prerequisites for permissions and configurations.
import sagemaker
import boto3
import sys
import os
import glob
import re
import subprocess
import numpy as np
from IPython.display import HTML
import time
from time import gmtime, strftime
sys.path.append("common")
from misc import get_execution_role, wait_for_s3_object
from docker_utils import build_and_push_docker_image
from sagemaker.rl import RLEstimator, RLToolkit, RLFramework
# ### Setup S3 bucket
#
# Set up the linkage and authentication to the S3 bucket that you want to use for checkpoint and the metadata.
# + tags=[]
sage_session = sagemaker.session.Session()
s3_bucket = sage_session.default_bucket()
s3_output_path = 's3://{}/'.format(s3_bucket)
print("S3 bucket path: {}".format(s3_output_path))
# -
# ### Define Variables
#
# We define variables such as the job prefix for the training jobs *and the image path for the container (only when this is BYOC).*
# create a descriptive job name
job_name_prefix = 'rl-cartpole-ray'
# ### Configure where training happens
#
# You can train your RL training jobs using the SageMaker notebook instance or local notebook instance. In both of these scenarios, you can run the following in either local or SageMaker modes. The local mode uses the SageMaker Python SDK to run your code in a local container before deploying to SageMaker. This can speed up iterative testing and debugging while using the same familiar Python SDK interface. You just need to set `local_mode = True`.
# + tags=["parameters"]
# run in local_mode on this machine, or as a SageMaker TrainingJob?
local_mode = False
if local_mode:
instance_type = 'local'
else:
# If on SageMaker, pick the instance type
instance_type = "ml.c5.2xlarge"
# -
# ### Create an IAM role
#
# Either get the execution role when running from a SageMaker notebook instance `role = sagemaker.get_execution_role()` or, when running from local notebook instance, use utils method `role = get_execution_role()` to create an execution role.
# + tags=[]
try:
role = sagemaker.get_execution_role()
except:
role = get_execution_role()
print("Using IAM role arn: {}".format(role))
# -
# ### Install docker for `local` mode
#
# In order to work in `local` mode, you need to have docker installed. When running from you local machine, please make sure that you have docker and docker-compose (for local CPU machines) and nvidia-docker (for local GPU machines) installed. Alternatively, when running from a SageMaker notebook instance, you can simply run the following script to install dependenceis.
#
# Note, you can only run a single local notebook at one time.
# only run from SageMaker notebook instance
if local_mode:
# !/bin/bash ./common/setup.sh
# ## Use docker image
#
# We are using the latest public docker image for RLlib from the [Amazon SageMaker RL containers repository](https://github.com/aws/sagemaker-rl-container).
#
# + tags=[]
# %%time
cpu_or_gpu = 'gpu' if instance_type.startswith('ml.p') else 'cpu'
aws_region = boto3.Session().region_name
framework = 'tf' # change to 'torch' for PyTorch training
custom_image_name = "462105765813.dkr.ecr.%s.amazonaws.com/sagemaker-rl-ray-container:ray-0.8.5-%s-%s-py36" % (aws_region, framework, cpu_or_gpu)
custom_image_name
# -
# ## Write the Training Code
#
# The training code is written in the file “train-rl-cartpole-ray.py” which is uploaded in the /src directory.
# First import the environment files and the preset files, and then define the main() function.
#
# **Note**: If PyTorch is used, plese update the above training code and set `use_pytorch` to `True` in the config.
# + tags=[]
# !pygmentize src/train-rl-cartpole-ray.py
# -
# ## Train the RL model using the Python SDK Script mode
#
# If you are using local mode, the training will run on the notebook instance. When using SageMaker for training, you can select a GPU or CPU instance. The RLEstimator is used for training RL jobs.
#
# 1. Specify the source directory where the environment, presets and training code is uploaded.
# 2. Specify the entry point as the training code
# 3. Specify the custom image to be used for the training environment.
# 4. Define the training parameters such as the instance count, job name, S3 path for output and job name.
# 5. Define the metrics definitions that you are interested in capturing in your logs. These can also be visualized in CloudWatch and SageMaker Notebooks.
# + tags=["parameters"]
train_instance_count = 1
# + tags=[]
# %%time
metric_definitions = RLEstimator.default_metric_definitions(RLToolkit.RAY)
estimator = RLEstimator(entry_point="train-rl-cartpole-ray.py",
source_dir='src',
dependencies=["common/sagemaker_rl"],
image_uri=custom_image_name,
role=role,
debugger_hook_config=False,
instance_type=instance_type,
instance_count=train_instance_count,
output_path=s3_output_path,
base_job_name=job_name_prefix,
metric_definitions=metric_definitions,
hyperparameters={
# Attention scientists! You can override any Ray algorithm parameter here:
#"rl.training.config.horizon": 5000,
#"rl.training.config.num_sgd_iter": 10,
}
)
estimator.fit(wait=local_mode)
job_name = estimator.latest_training_job.job_name
print("Training job: %s" % job_name)
# -
# ## Visualization
#
# RL training can take a long time. So while it's running there are a variety of ways we can track progress of the running training job. Some intermediate output gets saved to S3 during training, so we'll set up to capture that.
# + tags=[]
print("Job name: {}".format(job_name))
s3_url = "s3://{}/{}".format(s3_bucket,job_name)
intermediate_folder_key = "{}/output/intermediate/".format(job_name)
intermediate_url = "s3://{}/{}".format(s3_bucket, intermediate_folder_key)
print("S3 job path: {}".format(s3_url))
print("Intermediate folder path: {}".format(intermediate_url))
tmp_dir = "/tmp/{}".format(job_name)
os.system("mkdir {}".format(tmp_dir))
print("Create local folder {}".format(tmp_dir))
# -
# ### Fetch videos of training rollouts
# Videos of certain rollouts get written to S3 during training. Here we fetch the last 10 videos from S3, and render the last one.
recent_videos = wait_for_s3_object(
s3_bucket, intermediate_folder_key, tmp_dir,
fetch_only=(lambda obj: obj.key.endswith(".mp4") and obj.size>0),
limit=10, training_job_name=job_name)
last_video = sorted(recent_videos)[-1] # Pick which video to watch
os.system("mkdir -p ./src/tmp_render/ && cp {} ./src/tmp_render/last_video.mp4".format(last_video))
HTML('<video src="./src/tmp_render/last_video.mp4" controls autoplay></video>')
# ### Plot metrics for training job
# We can see the reward metric of the training as it's running, using algorithm metrics that are recorded in CloudWatch metrics. We can plot this to see the performance of the model over time.
# +
# %matplotlib inline
from sagemaker.analytics import TrainingJobAnalytics
if not local_mode:
df = TrainingJobAnalytics(job_name, ['episode_reward_mean']).dataframe()
num_metrics = len(df)
if num_metrics == 0:
print("No algorithm metrics found in CloudWatch")
else:
plt = df.plot(x='timestamp', y='value', figsize=(12,5), legend=True, style='b-')
plt.set_ylabel('Mean reward per episode')
plt.set_xlabel('Training time (s)')
else:
print("Can't plot metrics in local mode.")
# -
# ### Monitor training progress
# You can repeatedly run the visualization cells to get the latest videos or see the latest metrics as the training job proceeds.
# ## Evaluation of RL models
#
# We use the last checkpointed model to run evaluation for the RL Agent.
#
# ### Load checkpointed model
#
# Checkpointed data from the previously trained models will be passed on for evaluation / inference in the checkpoint channel. In local mode, we can simply use the local directory, whereas in the SageMaker mode, it needs to be moved to S3 first.
# +
if local_mode:
model_tar_key = "{}/model.tar.gz".format(job_name)
else:
model_tar_key = "{}/output/model.tar.gz".format(job_name)
local_checkpoint_dir = "{}/model".format(tmp_dir)
wait_for_s3_object(s3_bucket, model_tar_key, tmp_dir, training_job_name=job_name)
if not os.path.isfile("{}/model.tar.gz".format(tmp_dir)):
raise FileNotFoundError("File model.tar.gz not found")
os.system("mkdir -p {}".format(local_checkpoint_dir))
os.system("tar -xvzf {}/model.tar.gz -C {}".format(tmp_dir, local_checkpoint_dir))
print("Checkpoint directory {}".format(local_checkpoint_dir))
# -
if local_mode:
checkpoint_path = 'file://{}'.format(local_checkpoint_dir)
print("Local checkpoint file path: {}".format(local_checkpoint_dir))
else:
checkpoint_path = "s3://{}/{}/checkpoint/".format(s3_bucket, job_name)
if not os.listdir(local_checkpoint_dir):
raise FileNotFoundError("Checkpoint files not found under the path")
os.system("aws s3 cp --recursive {} {}".format(local_checkpoint_dir, checkpoint_path))
print("S3 checkpoint file path: {}".format(checkpoint_path))
# +
# %%time
estimator_eval = RLEstimator(entry_point="evaluate-ray.py",
source_dir='src',
dependencies=["common/sagemaker_rl"],
image_uri=custom_image_name,
role=role,
instance_type=instance_type,
instance_count=1,
base_job_name=job_name_prefix + "-evaluation",
hyperparameters={
"evaluate_episodes": 10,
"algorithm": "PPO",
"env": 'CartPole-v1'
}
)
estimator_eval.fit({'model': checkpoint_path})
job_name = estimator_eval.latest_training_job.job_name
print("Evaluation job: %s" % job_name)
# -
# # Model deployment
#
# Now let us deploy the RL policy so that we can get the optimal action, given an environment observation.
#
# **Note**: Model deployment is supported for TensorFLow only at current stage.
#
# STOP HERE IF PYTORCH IS USED.
# +
from sagemaker.tensorflow.model import TensorFlowModel
model = TensorFlowModel(model_data=estimator.model_data,
framework_version='2.1.0',
role=role)
predictor = model.deploy(initial_instance_count=1,
instance_type=instance_type)
# +
# ray 0.8.5 requires all the following inputs
# 'prev_action', 'is_training', 'prev_reward' and 'seq_lens' are placeholders for this example
# they won't affect prediction results
# Number of different values stored in at any time in the current state for the Cartpole example.
CARTPOLE_STATE_VALUES = 4
input = {"inputs": {'observations': np.ones(shape=(1, CARTPOLE_STATE_VALUES)).tolist(),
'prev_action': [0, 0],
'is_training': False,
'prev_reward': -1,
'seq_lens': -1
}
}
# +
result = predictor.predict(input)
result['outputs']['actions_0']
# -
# ### Clean up endpoint
predictor.delete_endpoint()
| reinforcement_learning/rl_cartpole_ray/rl_cartpole_ray_gymEnv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# +
import os
import pandas as pd
import numpy as np
import urllib2
from bs4 import BeautifulSoup
data_location = os.path.join("..","data","external")
# Read in info needed for strings we need to search urls for
file_path = os.path.join(data_location, "MCCVB-locations-members.xlsx")
locations = pd.read_excel(file_path, sheetname = "locations")
# Read in urls data
file_path = os.path.join(data_location, "FY 16 - 17 PR Report Monthly Master.xlsx")
month_sheet = pd.read_excel(file_path, sheetname = "October 2016")
#Headers
hdr = {'User-Agent': 'Mozilla/5.0 (X11; Linux x86_64) AppleWebKit/537.11 (KHTML, like Gecko) Chrome/23.0.1271.64 Safari/537.11',
'Accept': 'text/html,application/xhtml+xml,application/xml;q=0.9,*/*;q=0.8',
'Accept-Charset': 'ISO-8859-1,utf-8;q=0.7,*;q=0.3',
'Accept-Encoding': 'none',
'Accept-Language': 'en-US,en;q=0.8',
'Connection': 'keep-alive'}
month_sheet_v1 = month_sheet[~pd.isnull(month_sheet['URL'])]
month_sheet_v1 = month_sheet_v1[~(month_sheet_v1['URL'].str.contains('n/a'))]
month_sheet_v1 = month_sheet_v1[~(month_sheet_v1['URL'].str.contains('burrellesluce.com'))]
urls = month_sheet_v1['URL']
# select 10 for simplicity
def processUrl(url):
query = urllib2.Request(url, None, hdr)
html = urllib2.urlopen(query).read()
soup = BeautifulSoup(html)
# kill all script and style elements
for script in soup(["script", "style"]):
script.extract() # rip it out
# get text
text = soup.get_text(separator=' ')
# break into lines and remove leading and trailing space on each
lines = (line.strip() for line in text.splitlines())
# break multi-headlines into a line each
chunks = (phrase.strip() for line in lines for phrase in line.split(" "))
# drop blank lines
text = '\n'.join(chunk for chunk in chunks if chunk)
return text
row = {}
i=0
for url in urls:
i+= 1
print "URL %s :: Processing: %s" % (i, url)
try:
text = processUrl(url)
except:
print "Please check the following URL it was not processed: %s " % (url)
location_hits = []
location_hits_string = ''
if len(text) < 5:
location_hits.append('No locations found, please check')
else:
for index, location in locations.iterrows():
if location[0] in text:
print location[0]
location_hits.append(location[0])
location_hits_string = ','.join(location_hits)
url = str(url)
row[url] = location_hits_string
frame = pd.DataFrame(row.items(), columns = ['URL','Juris_mention_auto'])
frame_out = pd.merge(month_sheet, frame, on = ['URL'], how = 'left')
writer = pd.ExcelWriter('FY 16-17 PR TESTING.xlsx')
frame_out.to_excel(writer, 'October 2016', index = False)
# -
frame_out
| notebooks/PR-MVP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SEP Tutorial
import numpy as np
import sep
# Import Astropy to use as a FITS analyzer
# +
import astropy.io
from astropy.io import fits
import matplotlib.pyplot as plt
from matplotlib import rcParams
# %matplotlib inline
rcParams['figure.figsize'] = [10.,8.]
# -
# Open the FITS file and read it as a 2-d numpy array by indexing the image list
image = fits.open("image.fits")
data = image[0].data
# Display the image using the mean and standard deviation
m, s = np.mean(data), np.std(data)
plt.imshow(data, interpolation='nearest', cmap='gray',vmin=m-s,vmax=m+s, origin='lower')
plt.colorbar();
plt.savefig('image_raw.png')
# Determine a spatially varying background on the image file
bkg = sep.Background(data)
# Determine the 'global' mean and noise of the background created
print(bkg.globalback)
print(bkg.globalrms)
# Convert the background to be a 2-d array -- the same size as the original image
bkg_image = np.array(bkg)
# Display the background
plt.imshow(bkg_image, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
plt.savefig('image_bkg.png')
# Convert the background noise to be a 2-d array -- the same size as the original image and the background
bkg_rms = bkg.rms()
# Display the background noise
plt.imshow(bkg_rms, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
plt.savefig('image_bkgnoise.png')
# Subtract the background from the data array in order for sources to be detected
data_sub = data - bkg
# Run object detection with the detection threshold to be 1.5 times the global background RMS
objects = sep.extract(data_sub, 1.5, err=bkg.globalrms)
# Display the objects detected
len(objects)
# Plot the new background-subtracted image and a red ellipse for each object detected
# +
from matplotlib.patches import Ellipse
fig, ax = plt.subplots()
m, s = np.mean(data_sub), np.std(data_sub)
im = ax.imshow(data_sub,interpolation='nearest', cmap='gray',
vmin=m-s, vmax=m+s, origin='lower')
for i in range(len(objects)):
e = Ellipse(xy=(objects['x'][i], objects['y'][i]),
width=6*objects['a'][i],
height=6*objects['b'][i],
angle=objects['theta'][i] * 180. / np.pi)
e.set_facecolor('none')
e.set_edgecolor('red')
ax.add_artist(e)
plt.savefig('image_sources.png')
# -
# Display the field types of the objects
objects.dtype.names
# Perform simple circular aperture photometry with 3 pixel radii at the object locations
flux, fluxerr, flag = sep.sum_circle(data_sub, objects['x'], objects['y'],
3.0, err=bkg.globalrms, gain=1.0)
# Display the first 10 results
for i in range(10):
print("object {:d}: flux = {:f} +/- {:f}".format(i, flux[i], fluxerr[i]))
# Completed by <NAME>
| SEP.TUTORIAL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Learning: Fundamentals (cont.)
# ## 5. Neural Networks: Practicality
# As shown in the previous sections, training a neural network revolves around the following:
# * _Layers_, which are combined into a _network_ (or _model_).
# * The _input data_ and correspoding _targets_.
# * The _loss function_, which defines the feedback signal used for learning.
# * The _optimizer_, which determines how leaening proceeds.
#
# This can be shown in a visualization as below; a network, composed of layers that are chained together, maps the input data to predictions. The loss function then compares these predictions to the targets, producing a loss value; a measure of how well the network's predictions match what was expected. The optimizer uses this loss value to update the network's weights.
#
# 
# ### 5a. Layers: the building blocks of deep learning
#
# This is the fundamental data structure in neural networks, and is a data-processing module that takes as input one or more tensors and outputs one or more tensors. Some layers are stateless, but more frequently layers have a state; the layer's _weights_, one or several tensors learned with stochastic gradient descent, which together contain the network's _knowledge_. Different layers are appropriate for different tensor formats and different types of data processing. For instance, simple vector data, stored in 2D tensors of shape _(samples, features)_ is often processed by _densely connected_ layers, also called _fully connected_ layers. Sequence data stored in 3D tensors of shape _(samples, timesteps, features)_ is typically processed by _recurrent_layers such as an LSTM layer. Image data, stored in 4D tensors is usually processed by 2D convolution layers.
#
# The notion of _layer compatibility_ here refers specifically to the fact that every layer will only accept input tensors of a certain shape and will return output tensors of a certain shape. Consider the example below, where we're creating a layer that will only accept as input 2D tensors where the first dimension is 784 (axis 0, the batch dimension, is specified, and this any value would be accepted). This layer will return a tensor where the first dimension has been transformed to be 32:
from keras import layers
layer = layers.Dense(32, input_shape=(784,))
# Thus, this layer can only be connected to a downstream layer that expects 32-dimensional vectors as its input. When using Keras you don't have to worry about compatibility, because the layers you add to your models are dynamically built to match the shape of the incoming layer. For instance, in the example below, the second layer will not receive an input shape argument but instead, it automatically inferred its input shape as being the output shape of the layer that came before it:
# +
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(32, input_shape=(784,)))
model.add(layers.Dense(32))
# -
# ### 5b. Models: networks of layers
# A deep-learning model is a directed, acyclic graph of layers, and the most common instance is a linear stack of layers, mapping a single input to a single output. But as we move along, we'll see a much broader variety of network topologies, the common ones being:
# * Two-branch networks
# * Multihead networks
# * Inception blocks
#
# The topology of a network defines a _hypothesis space_, and by choosing a network topology, you constrain your space of possibilities to a specific series of tensor operations, mapping input data to output data. What you'll then be searching for is a good set of values for the weight tensors involved in these tensor operations. Picking the right network architecture is more art than a science, and although there are some best practices and principles you can rely on, only practice can help you become a proper neural-network architect.
#
#
# ### 5c. Loss functions and optimizers: keys to configuring the learning process
# Once the network architecture is defined, you'll still have to choose two more things:
# * _Loss function (objective function)_ - The quantity that will be minimized during training, and it represents a measure of success for the task at hand.
# * _Optimizer_ - Determines how the network will be updated based on the loss function, and it implements a specific variant of stochastic gradient descent (SGD).
#
# A neural network that has multiple outputs may have multiple loss functions (one per output), but the gradient-descent process must be based on a single scalar loss value; so for multiloss networks, all losses are combined (via averaging) into a single scalar quantity. Choosing the right loss function for the right problem is extremely important; your network will take any shortcut it can, to minimize the loss, so if teh loss function doesn't fully correlate with success for the task at hand, your network will end up doing things you may not have wanted.
#
# Fortunately, when it comes to common problems such as classification, regression, and sequence prediction, there are simple guidelines you can follow to choose the correct loss. For instance, you'll use binary corssentropy for a two-class classification problem, categorical crossentropy for a many-class classification problem, meaan-squared error for a regression problem, connectionist temporal classification (CTC) for a sequence-learning problem, and so on. Only when you're working on truly new research problems will you have to develop your own objective functions.
# ## 6. Keras Introduction
# Keras was initially developed for researchers with the aim of enabling fast experimentation, and some of the key features include:
# * It allows teh same code to run seamlessly on CPU/GPU.
# * It has a user-friendly API that makes it easy to quickly prototype deep-learning models.
# * It has built-in support for CNNs, RNNs, and any combination of both.
# * It supports arbitrary network architectures; multi-input/output models, layer sharing, model sharing, and so on. This means that Keras is appropriate for building essentially any deep-learning model, from a generative adversarial network to a neural Turing machine.
#
# Keras is a model-level library, providing high-level building blocks for developing DL models, and it doesn't handle low-level operations such as tensor manipulation and differentiation. Instead, it relies on a specialized, well-optimized tensor library to do so, serving as the _backend engine_ of Keras. The typical Keras workflow looks like this:
# 1. Define your training data; input tensors and output targets.
# 2. Define a network of layers (or _model_) that maps your inputs to your targets.
# 3. Configure the learning process by choosing a loss function, an optimizer, and some metrics to monitor.
# 4. Iterate on your training data by calling the _fit()_ method of your model.
#
# There are two ways to define a model; using the _Sequential_ class (only for linear stacks of layers, which is the most common network architecture) or the _Functional_ API (for directed acyclic graphs of layers, which lets you build completely arbitrary architectures). Below is an example of a model defined using the _Functinoal_ API:
input_tensor = layers.Input(shape=(784,))
x = layers.Dense(32, activation="relu")(input_tensor)
output_tensor = layers.Dense(10, activation="softmax")(x)
model = models.Model(inputs=input_tensor, outputs=output_tensor)
# With the functional API, you're manipulating the data tensors that the model processes and applying layers to this tensor as if they were functions. Once your model architecture is defined, it doesn't matter whether you used a sequential or functional API, all other steps are the same. The learning process is configured in the compilation step, where you specify the optimizer and loss function(s) that the model should use, as well as the metrics you want to monitor during training:
from keras import optimizers
model.compile(
optimizer=optimizers.RMSprop(lr=0.001),
loss="mse",
metrics=["accuracy"]
)
# Finally, the learning process consists of passing numpy arrays of input data (and the corresponding target data) to the model via the _fit()_ method, similar to what you would do in sklearn and other libraries:
# +
# model.fit(
# input_tensor,
# target_tensor,
# batch_size=128,
# epochs=10
# )
# -
# ### 6a. Classifying Movie Reviews: binary classification
# Here, we'll classify movie reviews as possitive or negative, based on the text content of the reviews. We'll work with the IMDB dataset which has 50k reviews, split into 25k for training and testing on each set, containing 50% negative and 50% positive reviews. First we'll load the dataset which comes with Keras:
from keras.datasets import imdb
(train_data, train_labels),(test_data, test_labels) = imdb.load_data(num_words=10000)
# The argument *num_words=10000* means you'll only keep the top 10k most frequently occurring words in the training data, and rare words will be discarded. This allows us to work with vector data of manageable size. For kicks, here's how you can quickly decode one of these reviews back to English words:
# +
# get mapping of words to an integer index
word_index = imdb.get_word_index()
# map integer indices to words
reverse_word_index = dict(
[(value, key) for (key,value) in word_index.items()]
)
# decode the review
decoded_review = " ".join(
[reverse_word_index.get(i-3, "?") for i in train_data[0]]
)
# -
# Also, you can't feed lists of integers into a neural network; you have to turn your lists into tensors, and there are two ways to do that:
# * Pad your lists so that they all have the same length, turn them into an integer tensor of shape *(samples, word_indices)*, and then use as the first layer in your network a layer capable of handling such integer tensors (the _Embedding_ layer).
# * One-hot encode your lists to turn them into vectors of 0s and 1s. This would mean, for instance, turning the sequence [3, 5] into 10,000-dimensional vector that would be all 0s except for indices 3 and 5, which would be 1s. Then you could use as the first layer in your network a _Dense_ layer, capable of handling floating-point vector data.
# +
# encoding integer sequences into a binary matrix
import numpy as np
def vectorize_sequences(sequences, dimension=10000):
# create an all zeros matrix of shape (len(sequences), dimension)
results = np.zeros((len(sequences), dimension))
for i, sequence in enumerate(sequences):
# set specific indices of results[i] to 1s
results[i,sequence] = 1.
return results
# vectorize data
x_train = vectorize_sequences(train_data)
x_test = vectorize_sequences(test_data)
# vectorize labels
y_train = np.asarray(train_labels).astype("float32")
y_test = np.asarray(test_labels).astype("float32")
# -
# Now onto building the network, the input data is vectors and the labels are scalars, and a type of network that performs well on such a problem is a simple stack of fully connected layers with relu activations: _Dense(16, activation="relu")_. Having 16 hidden units means the weight matrix W will have shape *(input_dimension, 16)*, and you can intuitively understand the dimensionality of your representation space as "how much freedom you're allowing the network to have when learning internal representations". Having more hidden units (a higher-dimensional representation space) allows your network to learn more-complex representations, but it makes the network more computationally expensive and may lead to learning unwanted patterns (patterns that will improve performance on the training data but not on the test data).
#
# Finally you'll need to choose a loss function and an optimizer. Because we're facing a binary classification problem and the output of the network is a probability (you end the network with a single-unit layer with a sigmoid function), it's best to use the *binary_crossentropy* loss. Crossentropy is a quantity from the field of information theory that measures the distance between probability distributions or, in this case, between the ground-truth distribution and your distribution. We'll also monitor the accuracy during training:
# +
from keras import models
from keras import layers
model = models.Sequential()
model.add(layers.Dense(16, activation="relu", input_shape=(10000,)))
model.add(layers.Dense(16, activation="relu"))
model.add(layers.Dense(1, activation="sigmoid"))
model.compile(
optimizer="rmsprop",
loss="binary_crossentropy",
metrics=["accuracy"]
)
# -
# In order to monitor the accuracy of the model during training on data it has never seen before, you'll create a validation set by setting apart 10k samples from the original training data, and train the model on 20 epochs in mini-batches of 512 samples. At the same time, you'll monitor loss and accuracy on the 10k samples that you set apart. You do so by passing the validation data as the *validation_data* argument:
# +
x_val = x_train[:10000]
partial_x_train = x_train[10000:]
y_val = y_train[:10000]
partial_y_train = y_train[10000:]
history = model.fit(
partial_x_train,
partial_y_train,
epochs=20,
batch_size=512,
validation_data=(x_val,y_val)
)
# +
# plotting training + validation loss
import matplotlib.pyplot as plt
history_dict = history.history
acc = history_dict["acc"]
loss_values = history_dict["loss"]
val_loss_values = history_dict["val_loss"]
epochs = range(1, len(acc) + 1)
plt.plot(epochs, loss_values, "bo", label="Training loss")
plt.plot(epochs, val_loss_values, "b", label="Validation loss")
plt.title("Training and validation loss")
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
# +
# plot training + validation accuracy
plt.clf() # clear plots
val_acc = history_dict["val_acc"]
plt.plot(epochs, acc, "bo", label="Training acc.")
plt.plot(epochs, val_acc, "b", label="Validation acc")
plt.title("Training and validation accuracy")
plt.xlabel("Epochs")
plt.ylabel("Loss")
plt.legend()
plt.show()
# -
# As you can see, the training loss decreases with every epoch and the training accuracy increases with every epoch. In this case, to mitigate against overfitting the model, we could stop training after three epochs and evaluate the data (the accuracy should be +2%).
#
# After having trained the network, you'll can generate the likelihood of reviews being positive by using the _predict_ method:
model.predict(x_test)
# >= 0.99 is confident
# <= 0.01 not-confident
# others not thaat sure
# The following experiments will help convince you about the neural net architecture, but some improvements could be made such as:
# * We used two hidden layers. Try using one or three hidden layers, and see how doing so affects validation and test accuracy.
# * Try using layers with more/fewer hidden units; 32, 64, and so on.
# * Try using the _mse_ loss function instead of *binary_crossentropy*.
# * Try using the _tanh_ activation function instead of _relu_.
# ### 6b. Classifying Newswires: multiclass classification
| _archived/keras/keras_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Reading and Interpreting the Dataset
#
# The following section is using pandas to read the dataframe of the 'labeled_and_scored_comments.csv' downloaded from the Data Bias assignment. This is a dataset of Wikipedia comments made available by Jigsaw, a subsidiary of Google that created the Perspective tool. The dataset includes a unique comment id, the text of the comment, and a series of binary labels applied by human raters: "toxic," "severe_toxic," "obscene," "threat," "insult," and "identity_hate." The "score" column was appended by Professor Engler, which represents the toxicity score assigned to the comment text by the live version of the Perspective API. The data is available under a CC0 license.
# +
import pandas as pd
import time
df = pd.read_csv('labeled_and_scored_comments.csv')
# -
df.sort_values(['score'])
# #### Describe the Dataset
#
# The data description notes an n value of 41338, which is sufficiently large.
df.describe()
pip install --upgrade google-api-python-client
# +
from googleapiclient.discovery import build
import json
def get_toxicity_score(comment):
client = build(
"commentanalyzer",
"v1alpha1",
developerKey=API_KEY,
discoveryServiceUrl="https://commentanalyzer.googleapis.com/$discovery/rest?version=v1alpha1",
static_discovery=False,
)
analyze_request = {
'comment': { 'text': comment },
'requestedAttributes': {'TOXICITY': {}}
}
response = client.comments().analyze(body=analyze_request).execute()
toxicity_score = response["attributeScores"]["TOXICITY"]["summaryScore"]["value"]
return toxicity_score
# -
# ## False Positives
#
# The context of the following comments aren't posed in a toxic or negative manner, but the resulting toxicity score calculated is high.
get_toxicity_score("i'm so hungry i could eat a pig")
get_toxicity_score("hey bitch i love you")
get_toxicity_score("she's serving cunt")
# ### Finding an example of a comment with a high score that isn't necessarily presented in a toxic manner.
df[df['comment_text'].str.contains("bitch")].sort_values(['score'])
print(df.iloc[40381]['comment_text'])
print(df.iloc[40381]['score'])
| Data Bias Assignment (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img src="./assets/illustration.png" style="width: 1000px;"/>
# <script>
# // AUTORUN ALL CELLS ON NOTEBOOK-LOAD!
# require(
# ['base/js/namespace', 'jquery'],
# function(jupyter, $) {
# $(jupyter.events).on("kernel_ready.Kernel", function () {
# console.log("Auto-running all cells-below...");
# jupyter.actions.call('jupyter-notebook:run-all-cells-below');
# jupyter.actions.call('jupyter-notebook:save-notebook');
# });
# }
# );
# </script>
# + [markdown] tags=[]
# ## Supplementary material
#
# <p style="color:rgb(255,0,0);">Please cite the source when using these data:</p>
# <NAME>., <NAME>., <NAME>. and <NAME>. Relative continent/mid-ocean ridge elevation: a reference case for isostasy in geodynamics. Earth-Science Reviews (Submission January 2022)<br><br>
#
#
# This Repository allows:
# - Displaying data and computing statistics on elevation of [continents](topo_cont.ipynb) and [mid-ocean ridges](topo_MOR.ipynb)
#
# - [Displaying](thermo.ipynb) and downloading thermodynamic solutions (raw data and grids of density, melt fraction,...)
#
# - Computing a basic 1-D isostatic balance for a reference case: [reference cases](isostasy_ref.ipynb), [calibration of crustal and mantle densities](isostasy_calib.ipynb), and [calibration of the reference mantle density with temperature dependent only densities](isostasy_calib2.ipynb)
#
# While you can run these notebooks on Binder (except topo_cont.ipynb that requires too much RAM), only a part of the data are accessible from there.
# Full data are localized on FigShare with the same structure as on the github repository.
# Therefore, we encourage you to download the full dataset on FigShare and to run `jupyter-lab` locally in order to be able to display everything ([README.md](https://github.com/tth030/SM_ESR_isostasy) for instructions).
#
# ## What are the data used to analyse Earth topography?
#
# <strong>Disclaimer</strong>: Some files provided here (data/) comes from a preliminary filtering using a command that is described in each binary header (can be read using `ncinfo` or `gmt gmtinfo`). Links provided here will give you access directly to the raw data or to a contact email. <p style="color:rgb(255,0,0);">Please cite each specific source when using these data.</p>
#
# - The following data can be display with/without mask using the notebook [topo_cont.ipynb](topo_cont.ipynb)
#
# - ETOPO1
# - https://www.ngdc.noaa.gov/mgg/global/
# - NOAA National Geophysical Data Center. 2009: ETOPO1 1 Arc-Minute Global Relief Model. NOAA National Centers for Environmental Information. Accessed [date]
# - <NAME>. and <NAME>, 2009. ETOPO1 1 Arc-Minute Global Relief Model: Procedures, Data Sources and Analysis. NOAA Technical Memorandum NESDIS NGDC-24. National Geophysical Data Center, NOAA. doi:10.7289/V5C8276M [access date]
#
#
# - Seafloor ages and spreading rate
# - https://www.earthbyte.org/age-spreading-rates-and-spreading-asymmetry-of-the-worlds-ocean-crust/
# - <NAME>., <NAME>, <NAME>, and <NAME> 2008. Age, spreading rates and spreading symmetry of the world's ocean crust, Geochem. Geophys. Geosyst., 9, Q04006, <a href="https://agupubs.onlinelibrary.wiley.com/doi/10.1029/2007GC001743">doi:10.1029/2007GC001743</a>
#
#
# - Horizontal strain rate
# - https://gsrm2.unavco.org/intro/intro.html
# - <NAME>., <NAME>, <NAME>, 2014, A geodetic plate motion and Global Strain Rate Model, Geochemistry, Geophysics, Geosystems, 15, 3849-3889, <a href="https://agupubs.onlinelibrary.wiley.com/doi/full/10.1002/2014GC005407">https://doi.org/10.1002/2014GC005407</a>
#
#
# - High resolution bathymetry data
# - https://www.gmrt.org/about/
# - <NAME>., et al. (2009), Global Multi-Resolution Topography synthesis, Geochem. Geophys. Geosyst., 10, Q03014, <a href="https://agupubs.onlinelibrary.wiley.com/doi/full/10.1029/2008GC002332">doi:10.1029/2008GC002332</a>
#
#
# - Lithospheric thickness (average based on several seismological estimates)
# - Ste<NAME>. and <NAME>. A comparison of lithospheric thickness models, Tectonophysics, 746 (2018), pp. 325-338, <a href="https://www.sciencedirect.com/science/article/pii/S004019511630316X?via%3Dihub">doi.org/10.1016/j.tecto.2016.08.001</a>
#
#
# - Age of the lithosphere from seismological analysis
# - <NAME>., <NAME>. Worldwide distribution of ages of the continental lithosphere derived from a global seismic tomographic model, Lithos, 109 (2009), pp. 125-130, <a href="https://www.sciencedirect.com/science/article/pii/S0024493708002582">doi.org/10.1016/j.lithos.2008.10.023</a>
#
#
# - Hot spots list
# - http://www.mantleplumes.org/P%5E4/P%5e4Chapters/MorganP4ElectronicSupp1.pdf or
# - https://gsapubs.figshare.com/articles/book/Supplemental_material_Plate_velocities_in_the_hotspot_reference_frame/12337703
# - <NAME>. and <NAME>. Plate velocities in the hotspot reference frame, Plates, Plumes and Planetary Processes, <NAME>, <NAME>, (2007), <a href="https://pubs.geoscienceworld.org/gsa/books/book/618/chapter/3805271/Plate-velocities-in-the-hotspot-reference-frame">doi.org/10.1130/2007.2430(04)</a>
#
#
# ## PerpleX | Thermodynamic calculations
#
# - [PerpleX](https://www.perplex.ethz.ch/)
# - In order to generate the phase equilbria in the mantle (including supra-solidus conditions) we use the thermodynamic dataset from Jennings and Holland (2015). Phase diagrams are produced using [Perple X 6.8.4](https://www.perplex.ethz.ch/perplex_updates.html) and the hp622.dat thermodynamic database for pure species and end-members (Holland and Powell, 2011). We use the following set of solution phases: O(JH), Sp(JH), Pl(JH),Melt(JH), Grt(JH), Opx(JH), Cpx(JH), Eskol(C), Ring(H), for olivine, spinel, plagioclase, melt, garnet, orthopyroxene, clinopyroxene, eskolaite, and ringwoodite, respectively (See paper for details).
#
#
# - What are inputfiles and output files?
# - Input file: *.dat
# - Output files: *.phm (data) *.ps (phase map)
#
#
# - P-T grids (density,melting,)
# - Files *.bin are built here and used for display and for loading in the 2-D thermo-mechanical code
# - See [Building grids and display key variables](thermo.ipynb)
#
# -
| start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Learning Objectives
#
# - What is Generator in Python, why do we need it
#
# - Learn about Abstract Method, Class Method and Static Method
#
# - Learn about functional programming and decorators in Python
#
#
# ## Iterator - Generator
#
# - Any function that uses the ‘yield’ statement is the generator
#
# - Each yield temporarily suspends processing, remem- bering the location execution state
#
# - When the generator iteration resumes, it picks-up where it left-off
# +
def generator_ex(ls):
s = 0
for i in ls:
s += i
yield s
G = generator_ex([1, 2, 3, 4])
for item in G:
print(item)
# -
for item in G:
print(item)
# ### Explain why nothing will be printed for above?
G = generator_ex([1, 2, 3, 4])
print(next(G))
print(next(G))
iter_ex = [1, 2, 3, 4]
print(next(iter_ex))
iter_ex = iter((1, 2, 3, 4))
print(next(iter_ex))
print(next(iter_ex))
# +
G = generator_ex(iter((1, 2, 3, 4)))
for item in G:
print(item)
# -
a = range(1, 4)
print(type(a))
print(a[0])
# ## Generator Expressions
b = (x*x for x in range(10))
print(type(b))
print(b[0])
for i in b:
print(next(b))
print(next(b))
print(sum(b))
c = (x*x for x in iter([1, 2, 3, 4]))
for i in c:
print(i)
# +
def firstn(ls):
n = len(ls)
ls = iter(ls)
S = 0
for _ in range(n):
S = S + next(ls)
yield S
G = firstn(range(100000))
for i in G:
print(i)
# -
# ## Resources:
#
# - https://nvie.com/posts/iterators-vs-generators/
#
# ## How to check memory and time
#
# - Let's assume that I want to find n**2 for all numbers smaller than 20000000
# +
import time, psutil, gc
import sys
# gc.collect()
# mem_before = psutil.virtual_memory()[3]
# time1 = time.time()
x = (i**2 for i in range(20000000))
sys.getsizeof(x)
time2 = time.time()
# mem_after = psutil.virtual_memory()[3]
print('Used Mem = {}'.format(sys.getsizeof(x)/1024**2)) # convert Byte to Megabyte
print('Calculation time = {}'.format(time2 - time1))
# -
# ## Abstract Methods
#
# - Abstract methods are the methods which does not contain any implemetation
#
# - But the child-class need to implement these methods. Otherwise error will be reported
#
# +
from abc import ABC, abstractmethod
class AbstractOperation(ABC):
def __init__(self, operand_a, operand_b):
self.operand_a = operand_a
self.operand_b = operand_b
super(AbstractOperation, self).__init__()
@abstractmethod
def execute(self):
pass
class AddOperation(AbstractOperation):
def execute(self):
return self.operand_a + self.operand_b
class SubtractOperation(AbstractOperation):
def execute(self):
return self.operand_a - self.operand_b
class MultiplyOperation(AbstractOperation):
def execute(self):
return self.operand_a * self.operand_b
class DivideOperation(AbstractOperation):
def execute(self):
return self.operand_a / self.operand_b
operation = AddOperation(1, 2)
print(operation.execute())
operation = SubtractOperation(8, 2)
print(operation.execute())
operation = MultiplyOperation(8, 2)
print(operation.execute())
operation = DivideOperation(8, 2)
print(operation.execute())
# -
# ## Classmethod, Staticmethod
# +
from datetime import date
# random Person
class Person:
def __init__(self, name, age):
self.name = name
self.age = age
@classmethod
def fromBirthYear(cls, name, birthYear):
return cls(name, date.today().year - birthYear)
def display(self):
print(self.name + "'s age is: " + str(self.age))
person = Person('Adam', 19)
person.display()
person1 = Person.fromBirthYear('John', 1985)
person1.display()
# +
from datetime import date
# random Person
class Person:
def __init__(self, name, age):
self.name = name
self.age = age
@staticmethod
def from_fathers_age(name, father_age, father_person_age_diff):
return Person(name, date.today().year - father_age + father_person_age_diff)
@classmethod
def from_birth_year(cls, name, birth_year):
return cls(name, date.today().year - birth_year)
def display(self):
print(self.name + "'s age is: " + str(self.age))
class Man(Person):
sex = 'Male'
man = Man.from_birth_year('John', 1985)
print(isinstance(man, Man))
man1 = Man.from_fathers_age('John', 1965, 20)
print(isinstance(man1, Man))
# +
class ClassGrades:
def __init__(self, grades):
self.grades = grades
@classmethod
def from_csv(cls, grade_csv_str):
grades = list(map(int, grade_csv_str.split(', ')))
cls.validate(grades)
return cls(grades)
@staticmethod
def validate(grades):
for g in grades:
if g < 0 or g > 100:
raise Exception()
try:
# Try out some valid grades
class_grades_valid = ClassGrades.from_csv('90, 80, 85, 94, 70')
print('Got grades:', class_grades_valid.grades)
# Should fail with invalid grades
class_grades_invalid = ClassGrades.from_csv('92, -15, 99, 101, 77, 65, 100')
print(class_grades_invalid.grades)
except:
print('Invalid!')
# +
class Date(object):
def __init__(self, day=0, month=0, year=0):
self.day = day
self.month = month
self.year = year
@classmethod
def from_string(cls, date_as_string):
day, month, year = map(int, date_as_string.split('-'))
date1 = cls(day, month, year)
return date1
date2 = Date.from_string('11-09-2012')
print(date2.__dict__)
# +
class UniqueIdentifier(object):
value = 0
def __init__(self, name):
self.name = name
@classmethod
def produce(cls):
instance = cls(cls.value)
cls.value += 1
return instance
class FunkyUniqueIdentifier(UniqueIdentifier):
@classmethod
def produce(cls):
instance = super(FunkyUniqueIdentifier, cls).produce()
instance.name = "Funky %s" % instance.name
return instance
x = UniqueIdentifier.produce()
y = FunkyUniqueIdentifier.produce()
print(x.__dict__)
print(y.__dict__)
print(type(x))
print(type(y))
print(x.value)
print(x.name)
print(y.name)
# -
# ## Class Variable, object variable
# empCount is a class variable
class Employee:
'Common base class for all employees'
empCount = 0
def __init__(self, name, salary):
self.name = name
self.salary = salary
Employee.empCount += 1
def displayCount(self):
print("Total Employee %d" % Employee.empCount)
def displayEmployee(self):
print("Name : ", self.name, ", Salary: ", self.salary)
# "This would create first object of Employee class"
emp1 = Employee("Zara", 2000)
# "This would create second object of Employee class"
emp2 = Employee("Manni", 5000)
print("Total Employee %d" % Employee.empCount)
# ### But if we change the code to this:
# +
class Employee:
'Common base class for all employees'
empCount = 0
def __init__(self, name, salary):
self.name = name
self.salary = salary
self.empCount += 1
def displayCount(self):
print("Total Employee %d" % self.empCount)
def displayEmployee(self):
print("Name : ", self.name, ", Salary: ", self.salary)
# "This would create first object of Employee class"
emp1 = Employee("Zara", 2000)
# "This would create second object of Employee class"
emp2 = Employee("Manni", 5000)
print(Employee.empCount)
print(emp1.empCount)
# -
# ### Although the second code is running without any error but does not satisfy our intention
# ## Another Example for class variable and object variable
# +
class A:
i = 1
def __init__(self):
self.i = 2
print(A.i)
print(A().i)
# -
# ## Method, Classmethod, Staticmethod
#
# Assume the class is written for addition
#
# - Method : it uses the instance variable (self.x) for addition, which is set by __init__ function
#
# - classmethod : it uses class variable for addition
#
# - staticmethod : it uses the value of x which is defined in main program (i.e. outside the class)
# +
#Resource: https://media.readthedocs.org/pdf/pythonguide/latest/pythonguide.pdf
# below x will be used by static method
# if we do not define it, the staticmethod will generate error.
x = 20
class Add(object):
x = 9 # class variable
def __init__(self, x):
self.x = x # instance variable
def addMethod(self, y):
print("method:", self.x + y)
@classmethod
# as convention, cls must be used for classmethod, instead of self
def addClass(cls, y):
print("classmethod:", cls.x + y)
@staticmethod
def addStatic(y):
print("staticmethod:", x + y)
def main(): # method
m = Add(x=4) # or m=Add(4)
# for method, above x = 4, will be used for addition
m.addMethod(10) # method : 14
# classmethod
c = Add(4)
# for class method, class variable x = 9, will be used for addition
c.addClass(10) # classmethod : 19
# for static method, x=20 (at the top of file), will be used for addition
s = Add(4)
s.addStatic(10) # staticmethod : 30
main()
# -
# ## Decorator
#
# - Decorator is a function that creates a wrapper around another function
#
# - This wrapper adds some additional functionality to existing code
# +
def addOne(myFunc):
def addOneInside(x):
print("adding One")
return myFunc(x) + 1
return addOneInside
def subThree(x):
return x - 3
result = addOne(subThree)
print(subThree(5))
print(result(5))
# +
@addOne
def subThree(x):
return x - 3
print(subThree(5))
# +
def memoize(f):
memo = {}
def helper(x):
if x not in memo:
memo[x] = f(x)
return memo[x]
return helper
def fib(n):
if n == 0:
return 0
elif n == 1:
return 1
else:
return fib(n-1) + fib(n-2)
result = memoize(fib)
print(result(40))
# -
# ### Explain why the above code is slow
# +
def memoize(f):
memo = {}
def helper(x):
if x not in memo:
memo[x] = f(x)
return memo[x]
return helper
def fib(n):
if n == 0:
return 0
elif n == 1:
return 1
else:
return fib(n - 1) + fib(n - 2)
fib = memoize(fib)
print(fib(40))
# +
@memoize
def fib(n):
if n == 0:
return 0
elif n == 1:
return 1
else:
return fib(n - 1) + fib(n - 2)
print(fib(40))
# +
def call(*argv, **kwargs):
def call_fn(fn):
return fn(*argv, **kwargs)
return call_fn
#@call(5)
def table(n):
value = []
for i in range(n):
value.append(i*i)
return value
table = call(5)(table)
print(table)
# +
def call(*argv, **kwargs):
def call_fn(fn):
return fn(*argv, **kwargs)
return call_fn
@call(5)
def table(n):
value = []
for i in range(n):
value.append(i*i)
return value
# table = call(5)(table)
print(table)
# print(len(table), table[3])
# -
# ## Functional Programming
# +
def cal(f, x, y):
return f(x, y)
def addition(x, y):
return x + y
def subtraction(x, y):
return x - y
print(cal(addition, 3, 2))
print(cal(subtraction, 3, 2))
# +
def next_(n, x):
return (x+n/x)/2
n= 2
f= lambda x: next_(n, x)
a0= 1.0
print([round(x, 4) for x in (a0, f(a0), f(f(a0)), f(f(f(a0))))])
# +
m = 4
def repeat(f, a):
# global m
# need global m if want to change m in the function (m = m)
for _ in range(m):
yield a
a = f(a)
for i in repeat(f, 1):
print(i)
# -
# ### Global Variable
# +
x = 10
def mathEx(a, b):
""" calculate (a+b)*x """
global x
x = x - 1
c=(a+b)*x
return c
print(mathEx(1,2))
# -
| Lessons/.ipynb_checkpoints/advance_python-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# Cargar funciones de la librería de python data analysis
import pandas as pd
# Leer csv con datos y cargar en el dataframe data
data = pd.read_csv("data/creditcard.csv")
# Preview de las 5 primeras filas de data, total filas: 284807
data.head()
# -
# total confirmado como fraude (class=1): 492
data[data['Class']==1]
# +
from sklearn.ensemble import IsolationForest
# train con % de registros con Class=0
n = 1000
X_train = data[data["Class"]==0].iloc[0:n,0:30]
# test1 492 registros con Class=1, y_test1 colocamos -1 para poder comparar con score de clasificacion
# test0 500 registros con Class=0 (disjunto de train), y_test0 colocamos 0 para poder comparar con score de clasificacion
X_test1 = data[data["Class"]==1].iloc[:,0:30]
y_test1 = data[data["Class"]==1].iloc[:,30:31]
y_test1['Class']=-1
X_test0 = data[data["Class"]==0].iloc[n:n+500,0:30]
y_test0 = data[data["Class"]==0].iloc[n:n+500,30:31]
y_test0['Class']=1
# unificar test
X_test = pd.concat([X_test1,X_test0], axis=0)
y_test = pd.concat([y_test1,y_test0], axis=0)
# fit the model
clf = IsolationForest(behaviour='new', max_samples=100, n_estimators=100,
random_state=1, contamination='auto')
clf.fit(X_train)
y_pred_test = clf.predict(X_test)
# +
dtest = pd.DataFrame(zip(y_test['Class'],y_pred_test),
columns=['actual test','score pred test'])
#dtest[dtest['score pred test']==-1]
# -
from sklearn import metrics
print("Accuracy:",metrics.accuracy_score(dtest['actual test'], dtest['score pred test']))
metrics.precision_recall_fscore_support(dtest['actual test'], dtest['score pred test'], average=None)
pd.crosstab(dtest['actual test'], dtest['score pred test'],
rownames=['actual'],
colnames=['pred'], margins=False, margins_name="Total")
| ScriptsAndData/.ipynb_checkpoints/isolationForest-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import Libraries
import numpy as np
import pandas as pd
from pandas.tools.plotting import scatter_matrix
import matplotlib.pyplot as plt
from sklearn import model_selection
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
import seaborn as sns
sns.set()
# Load Dataset
dataset = pd.read_csv('sensor_data.csv', usecols=['datetime', 'temperature', 'co2', 'light', 'noise'])
# -
df = pd.read_csv('sensor_data.csv', index_col='datetime', parse_dates=True)
df.info()
# Drop location columns
df.drop(df.columns[[0,1]], axis=1, inplace=True)
df.info()
df.drop(df.columns[[-1, -2]], axis=1, inplace=True)
df.info()
df.drop(df.columns[[1]], axis=1, inplace=True)
df.info()
# +
# Shape
print(dataset.shape)
# +
# Head
dataset.head(20)
# +
# Descriptions
print(dataset.describe())
# +
# Eliminate Outliers
dataset = dataset [dataset.temperature > 0]
dataset = dataset [dataset.light > 0]
dataset = dataset [dataset.noise > 0]
dataset = dataset [dataset.co2 != 2]
print(dataset.describe())
# +
# Create copy of temperature data
temp_data = df[['temperature']].copy()
# -
temp_data.info()
# Delete the 2 -999 error values
temp_data = temp_data[temp_data['temperature'] != -999]
# +
# Convert dataframe to PeriodIndex of temperature data
temp_period = temp_data.to_period(freq='D')
temp_period.info()
# +
# Box-and-whisker plots for daily temperature
fig, ax = plt.subplots(figsize=(16,10))
sns.boxplot(x=temp_period.index, y='temperature', data=temp_period, ax=ax)
ax.set_title('Daily Temperature Data', fontsize=24)
ax.set_xlabel('Date', fontsize=16)
ax.set_ylabel('Temperature (Celcius)')
plt.show()
# +
# Normalize time series data
from sklearn.preprocessing import MinMaxScaler
# load the dataset and print the first 5 rows
series = temp_period
print(series.head())
# prepare data for normalization
values = series.values
values = values.reshape((len(values), 1))
# train the normalization
scaler = MinMaxScaler(feature_range=(0, 1))
scaler = scaler.fit(values)
print('Min: %f, Max: %f' % (scaler.data_min_, scaler.data_max_))
# normalize the dataset and print the first 5 rows
normalized = scaler.transform(values)
for i in range(5):
print(normalized[i])
# inverse transform and print the first 5 rows
inversed = scaler.inverse_transform(normalized)
for i in range(5):
print(inversed[i])
# +
# Plotting the results
series.hist()
plt.show()
# +
# Lag Plot
from pandas.tools.plotting import lag_plot
lag_plot(series)
plt.show()
# +
# Autocorrelation plot
from pandas.tools.plotting import autocorrelation_plot
autocorrelation_plot(series)
plt.show()
# -
# %matplotlib inline
series.plot()
norma = pd.DataFrame(normalized)
norma.plot()
# +
# Linear Regression Before Normalization
from sklearn.linear_model import LinearRegression
estimator = LinearRegression()
sen = pd.read_csv('dataset2.csv')
sen.datetime = pd.to_datetime(sen.datetime)
#sen.set_index('datetime')
sen = sen[['datetime', 'temperature', 'count_total']]
X = sen[['temperature']]
Y = sen['count_total']
model = estimator.fit(X, Y)
# Put a random number to try the model
print(model.predict([[24]]))
# +
# K-Nearest Neighbour Regression Before Normalization
from sklearn.neighbors import KNeighborsRegressor
estimator = KNeighborsRegressor()
sen = sen[['datetime', 'temperature', 'count_total']]
X = sen[['temperature']]
Y = sen['count_total']
model=estimator.fit(X, Y)
# Put a random number to try the model
print(model.predict([[24]]))
# +
# Random Forest Regression Before Normalization
from sklearn.ensemble import RandomForestRegressor
estimator = RandomForestRegressor()
sen = sen[['datetime', 'temperature', 'count_total']]
X = sen[['temperature']]
Y = sen['count_total']
model = estimator.fit(X, Y)
# Put a random number to try the model
print(model.predict([[24]]))
# +
# Linear Regression Classification
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import classification_report
from sklearn.model_selection import train_test_split
X = sen[['temperature']]
y = sen['count_total']
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size = 0.2)
model = LogisticRegression()
model.fit(X_train, y_train)
expected = y_test
predicted = model.predict(X_test)
print(classification_report(expected, predicted))
# +
# K-Nearest Neighbour Classification
from sklearn.neighbors import KNeighborsClassifier
X = sen[['temperature']]
y = sen['count_total']
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size = 0.2)
model = KNeighborsClassifier(20)
model.fit(X_train, y_train)
expected = model.predict(X_test)
print(classification_report(expected, predicted))
# +
# Random Forest Classification
from sklearn.ensemble import RandomForestClassifier
X = sen[['temperature']]
y = sen['count_total']
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size = 0.2)
model = RandomForestClassifier()
model.fit(X_train, y_train)
expested = y_test
predicted = model.predict(X_test)
print(classification_report(expected, predicted))
# -
| data output/data until 05-13-17 + night office data/Machine learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Clustering comparative analysis
#
# **Original code**: [<NAME>](https://github.com/akorkovelos)<br>
# **Conceptualization & Methodological review** : [<NAME>](https://github.com/akorkovelos)<br>
# **Updates, Modifications**: [<NAME>](https://github.com/akorkovelos)<br>
# ## Importing packages
# +
# Operational
import os
import tkinter as tk
from tkinter import filedialog, messagebox
root = tk.Tk()
root.withdraw()
root.attributes("-topmost", True)
# Numerical
import geopandas as gpd
import pandas as pd
import numpy as np
import math
# Mapping
import contextily as ctx
from ipyleaflet import *
# Graphs and plotting
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# Coordinate system in EPSG registry
proj_crs = "epsg:32736"
def_crs = "epsg:4326"
# ## Import & prepare building footprint dataset
# #### Importing dataset
# +
# Define path and name of the geometry file
path = r"C:\Users\alexl\Dropbox\Self-employment\WBG\Work\GEP\clustering_process\DBSCAN_Clustering\Sample_Output"
path_KTH = r"C:\Users\alexl\Dropbox\Self-employment\WBG\Work\GEP\clustering_process\DBSCAN_Clustering\Sample_Input"
name_shp_HDBSCAN_self = "Sample_Concave_clustered_buildings_HDBSCAN_self_tuned.shp"
name_shp_HDBSCAN_auto = "Sample_Concave_clustered_buildings_HDBSCAN_auto_tuned.shp"
name_shp_DBSCAN = "Sample_Concave_clustered_buildings_DBSCAN.shp"
name_shp_True = "Sample_Concave_clustered_buildings_true.shp"
name_shp_KTH = "Kisoro_KTH_clusters.gpkg"
# -
# Create a new geo-dataframe(s)
geom_gdf_HDBSCAN_self = gpd.read_file(path + "\\" + name_shp_HDBSCAN_self)
geom_gdf_HDBSCAN_auto = gpd.read_file(path + "\\" + name_shp_HDBSCAN_auto)
geom_gdf_DBSCAN = gpd.read_file(path + "\\" + name_shp_DBSCAN)
geom_gdf_True = gpd.read_file(path + "\\" + name_shp_True)
geom_gdf_KTH = gpd.read_file(path_KTH + "\\" + name_shp_KTH)
# +
m = Map(
basemap=basemap_to_tiles(basemaps.OpenStreetMap.HOT),
center=(buildings.geometry.centroid[0].y, buildings.geometry.centroid[0].x),
zoom=13)
building_data = GeoData(geo_dataframe = buildings,
style={'color': 'black', 'opacity':3, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'Buildings')
DBSCAN_geom = GeoData(geo_dataframe = geom_gdf_DBSCAN,
style={'color': 'black', 'fillColor': '#3366cc', 'opacity':0.05, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'DBSCAN')
HDBSCAN_geom_self = GeoData(geo_dataframe = geom_gdf_HDBSCAN_self,
style={'color': 'black', 'fillColor': '#3366cc', 'opacity':0.05, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'HDBSCAN self-tuned')
HDBSCAN_geom_auto = GeoData(geo_dataframe = geom_gdf_HDBSCAN_auto,
style={'color': 'black', 'fillColor': '#3366cc', 'opacity':0.05, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'HDBSCAN auto-tuned')
KTH_geom = GeoData(geo_dataframe = geom_gdf_KTH,
style={'color': 'black', 'fillColor': '#3366cc', 'opacity':0.05, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'KTH clusters')
true_geom = GeoData(geo_dataframe = geom_gdf_True,
style={'color': 'black', 'fillColor': '#3366cc', 'opacity':0.05, 'weight':1.9, 'dashArray':'2', 'fillOpacity':0.6},
hover_style={'fillColor': 'red' , 'fillOpacity': 0.2},
name = 'true geometries')
m.add_layer(building_data)
m.add_layer(HDBSCAN_geom_self)
m.add_layer(HDBSCAN_geom_auto)
m.add_layer(DBSCAN_geom)
m.add_layer(KTH_geom)
m.add_layer(true_geom)
m.add_control(LayersControl())
m
# -
| Comparative_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pablocarreira-py39] *
# language: python
# name: conda-env-pablocarreira-py39-py
# ---
# Art. 38 A água potável deve estar em conformidade com o padrão organoléptico de potabilidade expresso no Anexo 11 e demais disposições deste Anexo.
#
# Paragráfo único. Para os parâmetros ferro e manganês são permitidos valores superiores ao VMPs estabelecidos no Anexo 11, desde que sejam observados os seguintes critérios:
#
# I - os elementos ferro e manganês estejam complexados com produtos químicos comprovadamente de baixo risco à saúde, conforme preconizado no Inciso VIII do Art. 14 e nas normas da ABNT; e
# II - as concentrações de ferro e manganês não ultrapassem 2,4 e 0,4 mg/L, respectivamente.
import os
import re
import sys
import pprint
import pandas as pd
from scipy.stats import gmean
from dateutil.relativedelta import relativedelta
from paths import *
# Parameters
cod_ibge = '3548906' # São Carlos
cod_ibge = '3526902' # Limeira
cod_ibge = '3501608' # Americana
# O Anexo 11 lista esses parâmetros:
#
#
# parametro_descricao
# - Alumínio
# - Amônia (como N)
# - Cloreto
# - Cor Aparente
# - 1,2 diclorobenzeno
# - 1,4 diclorobenzeno
# - Dureza total
# - Ferro
# - Gosto e Odor
# - Manganês
# - Monoclorobenzeno
# - Sódio
# - Sólidos Dissolvidos Totais
# - Sulfato
# - Sulfeto de Hidrogênio
# - Turbidez
# - Zinco
#
#
# Todos, exceto Turbidez, encontram-se na tabela de "Controle Semestral"
# <br>
#
# # Tabelas do Controle
# <br>
#
# ## Parâmetros Básicos
# Read Table
df_bruta = pd.read_excel(
os.path.join(output_path, str(cod_ibge), 'dados brutos', 'controle', 'controle_mensal_parametros_basicos.xlsx')
)
# Filtra Apenas SAAs
df = df_bruta.loc[df_bruta['Tipo Da Forma De Abastecimento'] == 'SAA'].copy()
# Filtra Apenas Último Ano
df = df[df['Ano De Referência'] == max(df['Ano De Referência'])].copy()
set(df['Parâmetro'])
# <br>
#
# ## Controle Semestral
# Read Table
df_bruta = pd.read_excel(
os.path.join(output_path, str(cod_ibge), 'dados brutos', 'controle', 'controle_semestral.xlsx')
)
# Filtra Apenas SAAs
df = df_bruta.loc[df_bruta['Tipo Da Forma De Abastecimento'] == 'SAA']
# +
#df.info()
#list(df.columns)
# -
# Filtra Apenas Último Ano
df = df[df['Ano De Referência'] == max(df['Ano De Referência'])].copy()
set(df['Parâmetro'])
# <br>
#
# # Análises
# # Lixos
#df = df[df['Parâmetro'] == 'Escherichia coli'].copy()
df = df[df['Parâmetro'].str.contains('Cloro')].copy()
df.head()
set(df['Ponto De Monitoramento'])
df = df[df['Ponto De Monitoramento'] == 'SAÍDA DO TRATAMENTO'].copy()
df.head()
df = df[['Ano De Referência', 'Mês De Referência', 'Campo', 'Valor']].copy()
df = df.sort_values(by=['Ano De Referência', 'Mês De Referência', 'Campo']).copy()
df.head()
# Americana não tinha amostras no Ponto de captação....
#
# {'SAÍDA DO TRATAMENTO', 'SISTEMA DE DISTRIBUIÇÃO'}
df['Valor'] = df['Valor'].astype(str).str.replace(',','.')
df['Valor'] = df['Valor'].astype(float).fillna(0.0)
df.head()
# <br>
#
# Fazer gráfico multidimensional...
# Gráfico...
| test/34_art38_organoleptico_CONTROLE-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# # Time series / date functionality in Pandas
# * Pandas was developed for financial modelling (<NAME>, AQR Capital)
# * Handling time series therefore comprises an integral part of the package
# * We're going to look at two different concepts in Pandas
# - 1) Timestamps
# - 2) Timedeltas
# ## 1) Timestamp objects
# * Pandas has built-in TimeStamp objects
# * An array of TimeStamp objects is a DateTimeIndex
# * The datatype of a TimeStamp object is datetime64
# * There are two main methods of creating Timestamps or a DateTimeIndex:
# - 1) `pd.to_datetime()`
# - 2) `pd.date_range()`
# ### 1.1) `pd.to_datetime()`
#
# - You can use a lot of different formats to convert a string into a Timestamp
# +
# Convert todays day in a pd.Timestamp
today = '29 March 2021'
# -
type(today)
today = pd.to_datetime(today)
pd.to_datetime('10/03/2021', dayfirst=True)
# - You could also pass a `pd.Series` or a `pd.DataFrame` into `pd.to_datetime()` if the values are convertable to a Timestamp.
# +
# Create a list of dates, e.g. [today, tomorrow]
datetime_index = pd.to_datetime(['29/03/2021', '30-03-2021'])
# -
datetime_index
datetime_index[0]
# Convert it to a pd.DatetimeIndex
# ### 1.2) `pd.date_range()`
#
# What happens if you want to create a range of dates?
# You can use `pd.date_range()` to create a DateTimeIndex (array of Timestamps):
# +
# On which date did you start the bootcamp?
start_date = '15 March 2021'
# +
# On which date are you going to graduate?
end_date = '10 June 2021'
# +
# Create a DatetimeIndex from start to end
dt_range = pd.date_range(start=start_date, end=end_date, freq='D')
# -
dt_range
# Convert dt_range into a pd.Series
pd.Series(dt_range)
# ### 1.3) This is all nice and fine, but why do we care?
#
# The reason we care for Timestamps in pandas is because they allow us to perform time related actions on the data.
# Several things you can do with a timestamp:
#
# - extract the hour
# - extract the day
# - extract the month
# - extract the year
# - Slice a DataFrame or Series if the DateTimeIndex is the Index of the DataFrame
# - calculate time differences
# How could that be useful?
# Let us look at the data for this week and think about what we could do with this functionality:
#
# - Extract weekdays from the Timestamp
# - Extract the hour of the day from the Timestamp
# - Create subsets of the data
# - ...
# +
# Load this weeks training data
df = pd.read_csv('./data/train.csv', parse_dates=True, index_col=0)
# parse_dates=True will try to interpret the index_col as a pd.DatetimeIndex
df.head()
# +
# Inspect the type of the df.index
type(df.index)
# +
# Until which date do we have data?
df.tail()
# -
df.index.max()
# +
# Slice the pd.DataFrame by one day / year
df['2011-01-01']
# -
# Slice the pd.DataFrame by one date + datetime
df.loc['2011-01-01 01']
# +
# Extract information about the date or time
df['hour'] = df.index.hour
# -
df.shape
df.head()
df.index.day_name()
# Can we use these datetime functionalities if the array of timestamps is not in the index
df_reindexed = df.reset_index()
df_reindexed.head()
df_reindexed.datetime[0]
df_reindexed.datetime.dt.day
# +
# df_reindexed[df_reindexed.datetime == '2011-01-01']
# -
# ### 1.4) `df.between_time()`
df.between_time(start_time='22:00', end_time='05:00')
# ## 2) Timedelta objects
#
# Pandas has built-in Timedelta objects
# * An array of Timedelta objects is a TimedeltaIndex
# * The datatype of a Timedelta object is timedelta64
# * There are three methods of creating Timedeltas or a TimedeltaIndex:
# - 1) `pd.to_timedelta()`
# - 2) `pd.timedelta_range()`
# - 3) Subtract two pd.Timestamp objects
# +
# Calculate the Timedelta between the last and the first observation of our data
time_range = df.index.max() - df.index.min()
# -
time_range.seconds
# ## 3) Other concepts: `resample()`, `shift()` and `rolling()`
#
# You will see these concepts in later points of the course.
| week_03/TimeData_Pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 我們使用最基礎的 Gradient Descent 方法來找到y=f(x)=Wx+b 當中的 W 和 b , 用來預測 PokenMon 升級後的 CP 值
# * 主要的學習教材為台大電機系李弘毅教授的教材 : http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2017/Lecture/Linear%20Regression.mp4
# * 這問題可以視為 Regression 問題
# * Gradient Descent 的推導會使用到基礎的微分觀念,不了解的話可以參考 : http://www.amath.nchu.edu.tw/~tdoc/4_2.htm
# * 實驗資料可以從這裡下載 : https://www.openintro.org/stat/data/?data=pokemon
# ## 直觀推導方式如下:
# * $\hat y$ 為 Traing Data 當的答案,也就是範例中的 $newcp $
# * $x$ 為 $cp$ 值,而我們要找到一個 $f(x)$ 能準確預測 $newcp$,故令 $y=f(x)=w*x+b $
# * 先定義 Lose Function : $L= (\hat y-f(x))^2 => L(w,b) = (\hat y - (b+w*x))^2 $
# * 我們想要將 Lose 值降到最低,直觀的想法是找出 w,b 對於 Lose Function 的變化影響,如果值變化是往變大的方向我們就減上變化量,更數學的說就是看 w,b 的微小的變化對於 Lose 值的變化方向,然後進行反方向的更新。
# * 使用泰勒展開式來進行 Gradient Descent 的推導 : http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2017/Lecture/Gradient%20Descent.pdf
# * 基於上述直觀的想法,我們對 Lose Function 分別對 w,b 作偏微分,再將此偏微分的結果用來減掉原來的 w,b
#
# ## 最終推導的結果如下圖
# <img align="left" width="30%" src="./imgs/GradientDescent.png" />
# ## 我們先取前 50 筆資料,作為實驗素材
# +
import numpy as np
import pandas as pd
pd = pd.read_csv("./pokemon.csv")
newcp = np.array(pd['cp_new'])
cp = np.array(pd['cp'])
cp = cp[:50]
newcp = newcp[:50]
import matplotlib.pyplot as plt
# plt.plot(newcp,cp)
plt.scatter(cp, newcp)
# plt.ylabel('some numbers')
plt.show()
# -
# ## 實作 Gradient Descent 的公式
# * 這裡有作一點變形,使用的方法為 Stochastic Gradient Descent
# * 用比較白話的說就是,並不是拿所有的 Training Data 的 Lose Value 和來算 Gradient ,而是針對每一個 X 就算一次並進行 Gradient Descent 的 update
# * 除了使用 Stochastic Gradient Descent 方法之中,還使用了 Regularization 的技巧,避免 w 值過大,造成 f(x) 不夠平滑。
w = 1
b = 1
n = len(cp)
r= 0.000001
for i in range(100000):
dw = 0
db = 0
error = 0
for x , y in zip(cp,newcp):
# 加上 2*w 是考量到 Regularization
dw = -2*x*(y- (w*x+b)) + 2*w
db = -2*(y- (w*x+b))
# print dw,db
y_head = w*x + b
w = w - dw*r
b = b - db*r
error +=(y_head-y)*(y_head-y)
if i % 10000 ==0 :
print("w={:0.3f}, b={:0.3f}, error={:0.3f} ".format(w,b,error/n))
# +
y_head = []
for x in cp :
predict = w*x +b
y_head.append(predict)
plt.scatter(y_head, newcp)
plt.show()
# -
| GradientDescent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="HJ0S9kEasyjc"
# ## Imports and setup
# + id="a60ZN1XJqjqs"
# !pip install -q tensorflow-addons
# + id="KclKS2uSqsTn"
from tensorflow.keras import layers
import tensorflow_addons as tfa
from tensorflow import keras
import tensorflow as tf
from datetime import datetime
import matplotlib.pyplot as plt
import numpy as np
import random
# Setting seeds for reproducibility.
SEED = 42
tf.random.set_seed(SEED)
np.random.seed(SEED)
random.seed(SEED)
# + [markdown] id="2CAdE6uZs1bg"
# ## Hyperparameters
# + id="nGNq1YE9quPL"
# DATA
BUFFER_SIZE = 1024
BATCH_SIZE = 256
AUTO = tf.data.AUTOTUNE
INPUT_SHAPE = (32, 32, 3)
NUM_CLASSES = 10
# OPTIMIZER
LEARNING_RATE = 5e-3
WEIGHT_DECAY = 1e-4
# TRAINING
EPOCHS = 100
# AUGMENTATION
IMAGE_SIZE = 48 # We'll resize input images to this size.
PATCH_SIZE = 6 # Size of the patches to be extract from the input images.
NUM_PATCHES = (IMAGE_SIZE // PATCH_SIZE) ** 2
# ENCODER and DECODER
LAYER_NORM_EPS = 1e-6
ENC_PROJECTION_DIM = 128
ENC_NUM_HEADS = 4
ENC_LAYERS = 3
ENC_TRANSFORMER_UNITS = [
ENC_PROJECTION_DIM * 2,
ENC_PROJECTION_DIM,
] # Size of the transformer layers.
# + [markdown] id="82XzhKTus3Ol"
# ## CIFAR-10 dataset loading and preparation
# + id="fNENQQyJqwDY"
(x_train, y_train), (x_test, y_test) = keras.datasets.cifar10.load_data()
(x_train, y_train), (x_val, y_val) = (
(x_train[:40000], y_train[:40000]),
(x_train[40000:], y_train[40000:]),
)
print(f"Training samples: {len(x_train)}")
print(f"Validation samples: {len(x_val)}")
print(f"Testing samples: {len(x_test)}")
# + id="UkKfe5hWq19S"
def get_train_augmentation_model():
model = keras.Sequential(
[
layers.Rescaling(1 / 255.0),
layers.Resizing(INPUT_SHAPE[0] + 20, INPUT_SHAPE[0] + 20),
layers.RandomCrop(IMAGE_SIZE, IMAGE_SIZE),
layers.RandomFlip("horizontal"),
],
name="train_data_augmentation",
)
return model
def get_test_augmentation_model():
model = keras.Sequential(
[
layers.Rescaling(1 / 255.0),
layers.Resizing(IMAGE_SIZE, IMAGE_SIZE),
],
name="test_data_augmentation",
)
return model
# + id="LlCeqaYaqxWu"
def prepare_data(images, labels, is_train=True):
if is_train:
augmentation_model = get_train_augmentation_model()
else:
augmentation_model = get_test_augmentation_model()
dataset = tf.data.Dataset.from_tensor_slices((images, labels))
if is_train:
dataset = dataset.shuffle(BUFFER_SIZE)
dataset = dataset.batch(BATCH_SIZE).map(
lambda x, y: (augmentation_model(x), y), num_parallel_calls=AUTO
)
return dataset.prefetch(AUTO)
train_ds = prepare_data(x_train, y_train)
val_ds = prepare_data(x_train, y_train, is_train=False)
test_ds = prepare_data(x_test, y_test, is_train=False)
# + [markdown] id="cXWdbsOjs6ek"
# ## Patchify layers
# + id="H5rZ-br7q3lL"
class Patches(layers.Layer):
def __init__(self, patch_size=PATCH_SIZE):
super(Patches, self).__init__()
self.patch_size = patch_size
def call(self, images):
batch_size = tf.shape(images)[0]
patches = tf.image.extract_patches(
images=images,
sizes=[1, self.patch_size, self.patch_size, 1],
strides=[1, self.patch_size, self.patch_size, 1],
rates=[1, 1, 1, 1],
padding="VALID",
)
patch_dims = patches.shape[-1]
patches = tf.reshape(patches, [batch_size, -1, patch_dims])
return patches
# + id="hJ40Eh_9q9F-"
class PatchEncoder(layers.Layer):
def __init__(self, num_patches=NUM_PATCHES, projection_dim=ENC_PROJECTION_DIM):
super(PatchEncoder, self).__init__()
self.num_patches = num_patches
self.projection = layers.Dense(units=projection_dim)
self.position_embedding = layers.Embedding(
input_dim=num_patches, output_dim=projection_dim
)
def call(self, patch):
positions = tf.range(start=0, limit=self.num_patches, delta=1)
encoded = self.projection(patch) + self.position_embedding(positions)
return encoded
# + [markdown] id="HSgz58Uys_wN"
# ## ViT model utility
# + id="VGRHNeUW7wWN"
def mlp(x, dropout_rate, hidden_units):
for units in hidden_units:
x = layers.Dense(units, activation=tf.nn.gelu)(x)
x = layers.Dropout(dropout_rate)(x)
return x
# + id="WDtU--GgrPGG"
def create_vit_classifier():
inputs = layers.Input(shape=(IMAGE_SIZE, IMAGE_SIZE, 3))
# Create patches.
patches = Patches()(inputs)
# Encode patches.
encoded_patches = PatchEncoder()(patches)
# Create multiple layers of the Transformer block.
for _ in range(ENC_LAYERS):
# Layer normalization 1.
x1 = layers.LayerNormalization(epsilon=LAYER_NORM_EPS)(encoded_patches)
# Create a multi-head attention layer.
attention_output = layers.MultiHeadAttention(
num_heads=ENC_NUM_HEADS, key_dim=ENC_PROJECTION_DIM, dropout=0.1
)(x1, x1)
# Skip connection 1.
x2 = layers.Add()([attention_output, encoded_patches])
# Layer normalization 2.
x3 = layers.LayerNormalization(epsilon=LAYER_NORM_EPS)(x2)
# MLP.
x3 = mlp(x3, hidden_units=ENC_TRANSFORMER_UNITS, dropout_rate=0.1)
# Skip connection 2.
encoded_patches = layers.Add()([x3, x2])
# Create a [batch_size, projection_dim] tensor.
representation = layers.LayerNormalization(epsilon=LAYER_NORM_EPS)(encoded_patches)
representation = layers.GlobalAveragePooling1D()(representation)
# Classify outputs.
outputs = layers.Dense(NUM_CLASSES, activation="softmax")(representation)
# Create the Keras model.
model = keras.Model(inputs=inputs, outputs=outputs)
return model
# + [markdown] id="ihd604hFtCcg"
# ## LR scheduler
# + id="v2wkvqhlsQ-M"
# Some code is taken from:
# https://www.kaggle.com/ashusma/training-rfcx-tensorflow-tpu-effnet-b2.
class WarmUpCosine(keras.optimizers.schedules.LearningRateSchedule):
def __init__(
self, learning_rate_base, total_steps, warmup_learning_rate, warmup_steps
):
super(WarmUpCosine, self).__init__()
self.learning_rate_base = learning_rate_base
self.total_steps = total_steps
self.warmup_learning_rate = warmup_learning_rate
self.warmup_steps = warmup_steps
self.pi = tf.constant(np.pi)
def __call__(self, step):
if self.total_steps < self.warmup_steps:
raise ValueError("Total_steps must be larger or equal to warmup_steps.")
cos_annealed_lr = tf.cos(
self.pi
* (tf.cast(step, tf.float32) - self.warmup_steps)
/ float(self.total_steps - self.warmup_steps)
)
learning_rate = 0.5 * self.learning_rate_base * (1 + cos_annealed_lr)
if self.warmup_steps > 0:
if self.learning_rate_base < self.warmup_learning_rate:
raise ValueError(
"Learning_rate_base must be larger or equal to "
"warmup_learning_rate."
)
slope = (
self.learning_rate_base - self.warmup_learning_rate
) / self.warmup_steps
warmup_rate = slope * tf.cast(step, tf.float32) + self.warmup_learning_rate
learning_rate = tf.where(
step < self.warmup_steps, warmup_rate, learning_rate
)
return tf.where(
step > self.total_steps, 0.0, learning_rate, name="learning_rate"
)
# + id="5PtvlXWYsXbD"
total_steps = int((len(x_train) / BATCH_SIZE) * EPOCHS)
warmup_steps = int(total_steps * 0.15)
scheduled_lrs = WarmUpCosine(
learning_rate_base=LEARNING_RATE,
total_steps=total_steps,
warmup_learning_rate=0.0,
warmup_steps=warmup_steps,
)
# + [markdown] id="bnQzF-v3tE0X"
# ## Train and evaluate model
# + id="HT-gf8mpsZ5F"
optimizer = tfa.optimizers.AdamW(
learning_rate=scheduled_lrs,
weight_decay=WEIGHT_DECAY
)
vit_model = create_vit_classifier()
vit_model.compile(
optimizer=optimizer,
loss="sparse_categorical_crossentropy",
metrics=["accuracy"]
)
vit_model.fit(train_ds, validation_data=val_ds, epochs=EPOCHS)
loss, accuracy = vit_model.evaluate(test_ds)
accuracy = round(accuracy * 100, 2)
print(f"Accuracy on the test set: {accuracy}%.")
# + id="lfbsfkGv7wWP"
vit_model.save(f"classification_vit_model@acc_{accuracy}", include_optimizer=False)
# + [markdown] id="0ZcA2gkN7wWQ"
# ## References
#
# * https://keras.io/examples/vision/image_classification_with_vision_transformer/
| regular-classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/CT-LU/Notes-of-Clean-Code-in-Python/blob/main/Clean_Code_in_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="d0b8AWYcc6BB"
# # unittest
# + id="6x8hVuQyCORm"
import unittest
def division(a, b):
return a / b
class MyTest(unittest.TestCase):
def test_upper(self):
self.assertEqual('foo'.upper(), 'FOO')
def test_isupper(self):
self.assertTrue('FOO'.isupper())
self.assertFalse('Foo'.isupper())
def test_split(self):
s = 'hello world'
self.assertEqual(s.split(), ['hello', 'world'])
# check that s.split fails when the separator is not a string
#s.split(2)
with self.assertRaises(TypeError):
s.split(2) # TypeError: must be str or None, not int
def test_raise(self): # 通過regex 'by zero' 匹配除零異常
#division(1, 0) # ZeroDivisionError: division by zero
self.assertRaises(Exception, division, 1, 0)
self.assertRaisesRegex(Exception, "by zero", division, 1, 0)
if __name__ == '__main__':
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + id="qYUotAhAqLrT"
import unittest
import time
from unittest import TestCase
from unittest.mock import patch
def sum(a, b):
time.sleep(100) # 測試sum要花狠久
return a + b
class ClassName1: pass
class ClassName2: pass
class TestCalculator(TestCase):
@patch('__main__.sum', return_value=5) #直接patch sum函數return 5
def test_sum(self, sum):
self.assertEqual(sum(2, 3), 5)
self.assertEqual(sum(333, 3), 5) #無論sum傳什麼都會是return 5
@patch('__main__.ClassName2')
@patch('__main__.ClassName1')
def test_patch(self, MockClass1, MockClass2): #patch 兩個Mock
MockClass1.return_value = 1
MockClass2.return_value = 2
print(ClassName1())
print(ClassName2())
assert MockClass1 is ClassName1
assert MockClass2 is ClassName2
assert MockClass1.called
assert MockClass2.called
if __name__ == '__main__':
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + id="eXipcLKme0kg"
import unittest
import time
from unittest import TestCase
from unittest.mock import patch
from unittest.mock import Mock
class TestCalculator(TestCase):
m = Mock()
m.return_value = 3 # 等同 m = Mock(return_value=42)
m.foo = 42
m.configure_mock(bar='baz') # 等同 m.bar = 'baz'
print(m()) # 3
print(m.foo) # 42
print(m.bar) # baz
if __name__ == '__main__':
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + id="c37hQTpNzqeY"
from unittest.mock import Mock
myMethod = Mock() # mock 一個函數
myMethod.return_value = 3 # 函數return 3
print(myMethod(1, 'a', foo='bar')) # 呼叫函數傳入三個參數 會得到return 3
myMethod.assert_called_with(1, 'a', foo='bar') # 傳入三個參數(1, 'a', foo='bar') 被呼叫過會是true
print(myMethod()) # 再呼叫一次
print(myMethod.call_count) # 2 被呼叫過2次
# + id="H4mdUSdU2qeo"
myMethod.side_effect = KeyError("Hi Hi Key Error")
#myMethod("abc") # KeyError: 'Hi Hi Key Error'
new_mock = Mock(side_effect=KeyboardInterrupt("Error by ctrl + c"))
#new_mock() # KeyboardInterrupt: Error by ctrl + c
def for_side(*args, **kwargs):
print('args: ', args)
print('kwargs: ', kwargs)
myMethod.side_effect = for_side
myMethod('dsf', **{"a": 1, "bn": 2})
# args: ('dsf',)
# kwargs: {'a': 1, 'bn': 2}
myMethod('dsf', {"a": 1, "bn": 2})
#args: ('dsf', {'a': 1, 'bn': 2})
#kwargs: {}
# + id="69B_PW-c56Fu"
import unittest
class Person:
def __init__(self):
self.__age = 10
def get_fullname(self, first_name, last_name):
return first_name + ' ' + last_name
def get_age(self):
return self.__age
@staticmethod
def get_class_name():
return Person.__name__
class PersonTest(unittest.TestCase):
def setUp(self):
self.p = Person()
def test_should_get_age(self):
self.p.get_age = Mock(side_effect=[10, 11, 12]) # 摸擬每次呼叫get_age會得到的結果
self.assertEqual(self.p.get_age(), 10)
self.assertEqual(self.p.get_age(), 11)
self.assertEqual(self.p.get_age(), 12)
def test_should_get_fullname(self):
'''
side_effect摸擬get_fullname,用lambda吃兩個參數,return回dict中的value
'''
values = {('James', 'Harden'): '<NAME>', ('Tracy', 'Grady'): '<NAME>'}
self.p.get_fullname = Mock(side_effect=lambda x, y: values[(x, y)])
self.assertEqual(self.p.get_fullname('James', 'Harden'), '<NAME>')
self.assertEqual(self.p.get_fullname('Tracy', 'Grady'), '<NAME>')
if __name__ == '__main__':
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + [markdown] id="8kr-jwxFZVih"
# # **Chapter 2 Pythonic Code**
# + [markdown] id="aJKayzZti3z8"
# ## Indexes and slices
# + id="2CCO1qNdi-27"
"""Indexes and slices
Getting elements by an index or range
"""
import doctest
def index_last():
"""
>>> my_numbers = (4, 5, 3, 9)
>>> my_numbers[-1]
9
>>> my_numbers[-3]
5
"""
def get_slices():
"""
>>> my_numbers = (1, 1, 2, 3, 5, 8, 13, 21)
>>> my_numbers[2:5]
(2, 3, 5)
>>> my_numbers[:3]
(1, 1, 2)
>>> my_numbers[3:]
(3, 5, 8, 13, 21)
>>> my_numbers[::]
(1, 1, 2, 3, 5, 8, 13, 21)
>>> my_numbers[1:7:2]
(1, 3, 8)
>>> interval = slice(1, 7, 2)
>>> my_numbers[interval]
(1, 3, 8)
>>> interval = slice(None, 3)
>>> my_numbers[interval] == my_numbers[:3]
True
"""
def main():
index_last()
get_slices()
fail_count, _ = doctest.testmod(verbose=True)
#raise SystemExit(fail_count)
if __name__ == "__main__":
main()
# + [markdown] id="tM8F0fF5r6d4"
# ## Creating your own sequences
# + id="B_-f59I0sF6l"
"""Clean Code in Python - Chapter 2: Pythonic Code
可以使用pythonic的統一存取方式(magic function),
實作自己的class也應該要先思考,實作magic function來存取
"""
class Items:
def __init__(self, *values):
self._values = list(values)
def __len__(self):
return len(self._values)
def __getitem__(self, item):
return self._values.__getitem__(item)
def main():
a_item = Items(10, 1, 'hello item !!')
print(a_item[-1])
if __name__ == "__main__":
main()
# + [markdown] id="SNhuXjOCin7j"
# ## Context Managers
# 三種with obj: ... 的實作方式
# + id="mMcJkR81ZpXx"
import contextlib
run = print
def stop_database():
run("systemctl stop postgresql.service")
def start_database():
run("systemctl start postgresql.service")
'''
method 1: 利用__magic__ ,__enter__會return給as,__exit__是結束with
'''
class DBHandler:
def __enter__(self):
stop_database()
return self
def __exit__(self, exc_type, ex_value, ex_traceback):
start_database()
def db_backup():
run("pg_dump database")
'''
method 2: contextlib.contextmanager,裝飾yield產生器函式
'''
@contextlib.contextmanager
def db_handler():
stop_database()
yield
start_database()
'''
method 3: 實作contextlib.ContextDecorator裝飾器class, 就可以不用with
但是,無法在環境管理器中就拿不到 as obj,原則上裝飾器不曉得發生什麼事
'''
class dbhandler_decorator(contextlib.ContextDecorator):
def __enter__(self):
stop_database()
def __exit__(self, ext_type, ex_value, ex_traceback):
start_database()
@dbhandler_decorator()
def offline_backup():
run("pg_dump database")
def main():
with DBHandler():
db_backup()
with db_handler():
db_backup()
offline_backup() #第三種沒有with obj
'''
想乎略錯誤, 例如檔案不存在就乎略
'''
with contextlib.suppress(FileNotFoundError):
with open("1.txt") as f:
for line in f:
print(line)
if __name__ == "__main__":
main()
# + [markdown] id="1gPSpGzaydiv"
# ## Properties
# 約定成俗的private,想要存取它,要實作property
# + id="EsQjWaEjyiE4"
import re
EMAIL_FORMAT = re.compile(r"[^@]+@[^@]+\.[^@]+")
def is_valid_email(potentially_valid_email: str):
return re.match(EMAIL_FORMAT, potentially_valid_email) is not None
class User:
def __init__(self, username):
self.username = username
self._email = None #private
@property
def email(self): # read _email
return self._email
@email.setter # set _email
def email(self, new_email):
if not is_valid_email(new_email):
raise ValueError(
f"Can't set {new_email} as it's not a valid email"
)
self._email = new_email
def main():
u1 = User('jsmith')
u1.email = '<EMAIL>'
print(u1.email)
#u1.email = 'jsmith@' #raise error
if __name__ == "__main__":
main()
# + [markdown] id="2-jVn1FoirYP"
# ## Iterable objects
# + id="bODaXH1Yiw0X"
'''
想要for in,有兩種可能,
1 __len__ and __getitem__
2 __next__ or __iter__
'''
from datetime import timedelta, date
'''
這種作法只能使用一次loop
'''
class DateRangeIterable:
"""An iterable that contains its own iterator object."""
def __init__(self, start_date, end_date):
self.start_date = start_date
self.end_date = end_date
self._present_day = start_date
def __iter__(self):
return self #傳回自己成為iter物件
def __next__(self): # 執行一次for loop後, _present_day會更新成最後一天
if self._present_day >= self.end_date:
raise StopIteration
today = self._present_day
self._present_day += timedelta(days=1)
return today
'''
在__iter__中使用yeild生出generator, 可以重新loop
'''
class DateRangeContainerIterable:
"""An range that builds its iteration through a generator."""
def __init__(self, start_date, end_date):
self.start_date = start_date
self.end_date = end_date
def __iter__(self):
current_day = self.start_date
while current_day < self.end_date:
yield current_day #使用yield替代__next__
current_day += timedelta(days=1)
'''
就最直觀的create your own sequences
'''
class DateRangeSequence:
"""An range created by wrapping a sequence."""
def __init__(self, start_date, end_date):
self.start_date = start_date
self.end_date = end_date
self._range = self._create_range()
def _create_range(self):
days = []
current_day = self.start_date
while current_day < self.end_date:
days.append(current_day)
current_day += timedelta(days=1)
return days
def __getitem__(self, day_no):
return self._range[day_no]
def __len__(self):
return len(self._range)
def main():
for day in DateRangeIterable(date(2018, 1, 1), date(2018, 1, 5)):
print(day)
r = DateRangeIterable(date(2018, 1, 1), date(2018, 1, 5))
next(r)
next(r)
next(r)
next(r)
#next(r) #should raise stop
r1 = DateRangeIterable(date(2018, 1, 1), date(2018, 1, 5))
print(" , ".join(map(str, r1)))
#max(r1) #should raise stop
'''
前面這樣用iter物件執行一圈完就走到底不會從頭來,
讓我們使用下一個implementation
'''
r1 = DateRangeContainerIterable(date(2018, 1, 1), date(2018, 1, 5))
print(" , ".join(map(str, r1)))
print(max(r1))
'''
每個for都會藉__iter__建立新的generator
'''
s1 = DateRangeSequence(date(2018, 1, 1), date(2018, 1, 5))
for day in s1:
print(day)
'''
用iter,generator都是時間換空間,用list就是O(1),空間換時間
'''
if __name__ == "__main__":
main()
# + [markdown] id="QTiPVa5TpJtc"
# ## Container objects
# + id="SK3F2Rt7pPiq"
'''
作一個mask地圖
'''
import numpy as np
class Boundaries:
def __init__(self, width, height):
self.width = width
self.height = height
def __contains__(self, coord):
x, y = coord
return 0 <= x < self.width and 0 <= y < self.height
class Grid:
def __init__(self, width, height):
self.width = width
self.height = height
self.map = np.zeros([width, height])
self.limits = Boundaries(width, height)
def __contains__(self, coord):
return coord in self.limits #pythonic易讀因為實作了__contains__
def __setitem__(self, coord, value):
self.map[coord] = value
def mark_coordinate(grid, coord):
x, y = coord
if 0 <= x < grid.width and 0 <= y < grid.height:
grid[coord] = 1
if coord in grid: #pythonic易讀因為實作了__contains__
grid[coord] = 1
def main():
'''
element in container會是另一種magic的方式container.__contains__(element)
實作magic __contains__讓程式一致性易讀
'''
grid = Grid(640, 480)
mark_coordinate(grid, (1,2))
if __name__ == "__main__":
main()
# + [markdown] id="wjeXHjCqBHYf"
# ## Dynamic attributes for objects
# + id="fOPzH7gYBKPN"
class DynamicAttributes:
def __init__(self, attribute):
self.attribute = attribute
def __getattr__(self, attr): #沒有這個attribute就會呼叫它
if attr.startswith("fallback_"):
name = attr.replace("fallback_", "")
return f"[fallback resolved] {name}"
raise AttributeError(
f"{self.__class__.__name__} has no attribute {attr}"
)
def main():
dyn = DynamicAttributes("value")
print(dyn.attribute)
#'value'
print(dyn.fallback_test)
#'[fallback resolved] test'
dyn.__dict__["fallback_new"] = "new value"
'''
this call would be the same as running dyn.fallback_new = "new value"
'''
print(dyn.fallback_new)
#'new value'
'''
The syntax of getattr() method is:
getattr(object, name[, default])
The above syntax is equivalent to:
object.name
'''
print(getattr(dyn, "something", "default"))
#'default'
if __name__ == "__main__":
main()
# + [markdown] id="5aLY1RaqIjws"
# ## Callable objects
# + id="C3dklQBzImsY"
from collections import defaultdict, namedtuple
class CallCount:
def __init__(self):
self._counts = defaultdict(int) #dict的key可以是int
def __call__(self, argument): #functor呼叫加入新的key, 同時value + 1
self._counts[argument] += 1
return self._counts[argument]
def main():
cc = CallCount()
print(cc(1))
#1
print(cc(2))
#1
print(cc(1))
#2
print(cc(1))
#3
print(cc("something"))
#1
print(cc("something"))
#2
print(callable(cc))
#True
if __name__ == "__main__":
main()
# + [markdown] id="fYHAcVjzx5YG"
# ## Caveats in Python
# + id="vI1nQHj597kA"
import unittest
from datetime import datetime
class LoginEventSerializer:
def __init__(self, event):
self.event = event
def serialize(self) -> dict:
return {
"username": self.event.username,
"password": <PASSWORD>**",
"ip": self.event.ip,
"timestamp": self.event.timestamp.strftime("%Y-%m-%d %H:%M"),
}
class LoginEvent:
SERIALIZER = LoginEventSerializer
def __init__(self, username, password, ip, timestamp):
self.username = username
self.password = password
self.ip = ip
self.timestamp = timestamp
def serialize(self) -> dict:
return self.SERIALIZER(self).serialize()
class TestLoginEventSerialized(unittest.TestCase):
def test_serializetion(self):
event = LoginEvent(
"username", "password", "127.0.0.1", datetime(2016, 7, 20, 15, 45)
)
expected = {
"username": "username",
"password": <PASSWORD>**",
"ip": "127.0.0.1",
"timestamp": "2016-07-20 15:45",
}
self.assertEqual(event.serialize(), expected)
if __name__ == "__main__":
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + id="GCZIcj8Kx-iP"
from collections import UserList
def wrong_user_display(user_metadata: dict = {"name": "John", "age": 30}):
name = user_metadata.pop("name")
age = user_metadata.pop("age")
return f"{name} ({age})"
def user_display(user_metadata: dict = None):
user_metadata = user_metadata or {"name": "John", "age": 30}
name = user_metadata.pop("name")
age = user_metadata.pop("age")
return f"{name} ({age})"
class BadList(list): #list是CPython的實作, 直接繼承會把某些方法蓋掉
def __getitem__(self, index):
value = super().__getitem__(index)
if index % 2 == 0:
prefix = "even"
else:
prefix = "odd"
return f"[{prefix}] {value}"
class GoodList(UserList): #UserList按自己定制
def __getitem__(self, index):
value = super().__getitem__(index)
if index % 2 == 0:
prefix = "even"
else:
prefix = "odd"
return f"[{prefix}] {value}"
def main():
### mutable default arguments
print(wrong_user_display())
print(wrong_user_display({"name": "Jane", "age": 25}))
#print(wrong_user_display()) #keyError
print(user_display())
print(user_display({"name": "Jane", "age": 25}))
print(user_display()) #it works
### Extending built-in types
b1 = BadList((0, 1, 2, 3, 4, 5))
print(b1[0])
print(b1[1])
#print("".join(b1)) #TypeError
'''
join function 會試著iterate(run a for loop over)這個list,
但是預期是string,但我們已經改成output string
'''
g1 = GoodList((0, 1, 2))
print(g1[0])
print(g1[1])
print(";".join(g1)) #
if __name__ == "__main__":
main()
# + [markdown] id="fL3xhHky7Dj4"
# # Chapter 3 General Traits of Good Code
# + [markdown] id="W4x0P2ng9BhP"
# ## Handle exceptions at the right level of abstraction
# * 例外不要當成處理商業邏輯的go to
# * 函式只應做一件事情,這條原則也包括例外
# + id="5Z5B2bzZ9IPJ"
import logging
import unittest
from unittest.mock import Mock, patch
import time
logger = logging.getLogger(__name__)
class Connector:
"""Abstract the connection to a database."""
def connect(self):
"""Connect to a data source."""
return self
@staticmethod
def send(data):
return data
class Event:
def __init__(self, payload):
self._payload = payload
def decode(self):
return f"decoded {self._payload}"
#主要是deliver_event, 看它
class DataTransport:
"""An example of an object badly handling exceptions of different levels."""
retry_threshold: int = 5
retry_n_times: int = 3
def __init__(self, connector):
self._connector = connector
self.connection = None
'''
將event解碼後,傳輸data,它有兩種exception
'''
def deliver_event(self, event):
try:
self.connect()
data = event.decode()
self.send(data) # send f"decoded {self._payload}"
except ConnectionError as e: #ConnectionError是處理connect()沒有問題
logger.info("connection error detected: %s", e)
raise
except ValueError as e: #Value應該是decode要處理的,不該放在這裡
logger.error("%r contains incorrect data: %s", event, e)
raise
def connect(self):
for _ in range(self.retry_n_times):
try:
self.connection = self._connector.connect() #組合關係的_connector負責connect
except ConnectionError as e:
logger.info(
"%s: attempting new connection in %is",
e,
self.retry_threshold,
)
time.sleep(self.retry_threshold) # 這裡用time.sleep後再重連
else:
return self.connection
raise ConnectionError(
f"Couldn't connect after {self.retry_n_times} times"
)
def send(self, data):
return self.connection.send(data)
class FailsAfterNTimes:
'''
init可指定次數,跟例外處理
'''
def __init__(self, n_times: int, with_exception) -> None:
self._remaining_failures = n_times
self._exception = with_exception
def connect(self):
self._remaining_failures -= 1
if self._remaining_failures >= 0: # 可以重連的次數用光
raise self._exception
return self
def send(self, data):
return data
@patch("time.sleep", return_value=0) #patch time.sleep 成為sleep參數這個mock,且直接設定回傳0
class TestTransport(unittest.TestCase):
def test_connects_after_retries(self, sleep):
data_transport = DataTransport(
FailsAfterNTimes(2, with_exception=ConnectionError)
)
data_transport.send = Mock() # Mock一個send函數
data_transport.deliver_event(Event("test"))
data_transport.send.assert_called_once_with("decoded test") # 驗証否send呼叫過一次且參數是'decoded test'
assert (
sleep.call_count == DataTransport.retry_n_times - 1 # if false, assertionError is raised
), sleep.call_count # if 0, assertionError
def test_connects_directly(self, sleep):
connector = Mock()
data_transport = DataTransport(connector)
data_transport.send = Mock()
data_transport.deliver_event(Event("test"))
connector.connect.assert_called_once() # 驗証組合關係的connector.connect
assert sleep.call_count == 0
def test_connection_error(self, sleep):
data_transport = DataTransport(
Mock(connect=Mock(side_effect=ConnectionError))
) #組合關係connector用一個Mock摸擬, 且connector.connect也用一個Mock的side_effect摸擬Exception
self.assertRaisesRegex(
ConnectionError,
"Couldn't connect after \d+ times", #驗証這個expression的Exception
data_transport.deliver_event,
Event("connection error"),
)
assert sleep.call_count == DataTransport.retry_n_times
def test_error_in_event(self, sleep):
data_transport = DataTransport(Mock())
event = Mock(decode=Mock(side_effect=ValueError)) #decode摸擬一個ValueError
with patch("__main__.logger.error"): #patch
self.assertRaises(ValueError, data_transport.deliver_event, event)
assert not sleep.called
if __name__ == "__main__":
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + [markdown] id="J0zEbi9qNHtm"
# # Chapter 5 Using Decorators to Improve Our Code
# + [markdown] id="D_Y82J9wOHiF"
# ## Decorate functions
# + id="NAGyYuCgN7S_"
"""
Creating a decorator to be applied over a function.
"""
from functools import wraps
from unittest import TestCase, main, mock
import logging
class ControlledException(Exception):
"""A generic exception on the program's domain."""
def retry(operation):
@wraps(operation) #先別管@wraps
def wrapped(*args, **kwargs):
last_raised = None
RETRIES_LIMIT = 3
for _ in range(RETRIES_LIMIT):
try:
return operation(*args, **kwargs)
except ControlledException as e:
logging.info("retrying %s", operation.__qualname__)
last_raised = e
raise last_raised
return wrapped
class OperationObject:
"""A helper object to test the decorator."""
def __init__(self):
self._times_called: int = 0
def run(self) -> int:
"""Base operation for a particular action"""
self._times_called += 1
return self._times_called
def __str__(self):
return f"{self.__class__.__name__}()"
__repr__ = __str__
class RunWithFailure:
def __init__(
self,
task: "OperationObject",
fail_n_times: int = 0,
exception_cls=ControlledException,
):
self._task = task
self._fail_n_times = fail_n_times
self._times_failed = 0
self._exception_cls = exception_cls
def run(self):
called = self._task.run()
if self._times_failed < self._fail_n_times:
self._times_failed += 1
raise self._exception_cls(f"{self._task!s} failed!")
return called
@retry #語法糖,實際是執行 run_operation = retry(run_operation)
def run_operation(task):
"""Run a particular task, simulating some failures on its execution."""
return task.run()
class RetryDecoratorTest(TestCase):
def setUp(self):
self.info = mock.patch("logging.info").start()
def tearDown(self):
self.info.stop()
def test_fail_less_than_retry_limit(self):
"""Retry = 3, fail = 2, should work"""
task = OperationObject()
failing_task = RunWithFailure(task, fail_n_times=2)
times_run = run_operation(failing_task)
self.assertEqual(times_run, 3)
self.assertEqual(task._times_called, 3)
def test_fail_equal_retry_limit(self):
"""Retry = fail = 3, will fail"""
task = OperationObject()
failing_task = RunWithFailure(task, fail_n_times=3)
with self.assertRaises(ControlledException):
run_operation(failing_task)
def test_no_failures(self):
task = OperationObject()
failing_task = RunWithFailure(task, fail_n_times=0)
times_run = run_operation(failing_task)
self.assertEqual(times_run, 1)
self.assertEqual(task._times_called, 1)
if __name__ == "__main__":
main(argv=['first-arg-is-ignored'], exit=False)
# + [markdown] id="xf5Sbq2TO9r2"
# ## Decorate classes
# 當要開始擴展下面的例子時,就會有三個缺點
# * 過多class,事件數量增加,就要不同的serialize來對應
# * 無法reuse,屁如說,有一個新的event也需要hide password
# * boilerplate(指許多地方重覆出現只改少量code,冗),serialize呼叫會在不同類出現
# + id="ey7eyff3PBYC"
import unittest
from datetime import datetime
class LoginEventSerializer:
def __init__(self, event):
self.event = event
def serialize(self) -> dict:
return {
"username": self.event.username,
"password": "**redacted**",
"ip": self.event.ip,
"timestamp": self.event.timestamp.strftime("%Y-%m-%d %H:%M"),
}
class LoginEvent:
SERIALIZER = LoginEventSerializer
def __init__(self, username, password, ip, timestamp):
self.username = username
self.password = password
self.ip = ip
self.timestamp = timestamp
def serialize(self) -> dict:
return self.SERIALIZER(self).serialize()
class TestLoginEventSerialized(unittest.TestCase):
def test_serializetion(self):
event = LoginEvent(
"username", "password", "127.0.0.1", datetime(2016, 7, 20, 15, 45)
)
expected = {
"username": "username",
"password": <PASSWORD>**",
"ip": "127.0.0.1",
"timestamp": "2016-07-20 15:45",
}
self.assertEqual(event.serialize(), expected)
if __name__ == "__main__":
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + [markdown] id="vfm4X7P8T3Je"
# 讓我們用類別裝飾器來改善上面的例子
# + id="ulHQk6cMTckT"
'''
Reimplement the serialization of the events by applying a class decorator.
'''
import unittest
from datetime import datetime
'''
reuse呼叫func抽出來, 給EventSerializer專門轉Event的 attributes
'''
def hide_field(field) -> str:
return "**redacted**"
def format_time(field_timestamp: datetime) -> str:
return field_timestamp.strftime("%Y-%m-%d %H:%M")
def show_original(event_field):
return event_field
class EventSerializer:
"""Apply the transformations to an Event object based on its properties and
the definition of the function to apply to each field.
"""
def __init__(self, serialization_fields: dict) -> None:
"""Created with a mapping of fields to functions.
Example::
>>> serialization_fields = {
... "username": str.upper,
... "name": str.title,
... }
Means that then this object is called with::
>>> from types import SimpleNamespace
>>> event = SimpleNamespace(username="usr", name="name")
>>> result = EventSerializer(serialization_fields).serialize(event)
Will return a dictionary where::
>>> result == {
... "username": event.username.upper(),
... "name": event.name.title(),
... }
True
"""
self.serialization_fields = serialization_fields
def serialize(self, event) -> dict:
"""Get all the attributes from ``event``, apply the transformations to
each attribute, and place it in a dictionary to be returned.
"""
return {
field: transformation(getattr(event, field)) for field, transformation in self.serialization_fields.items()
}
class Serialization:
"""A class decorator created with transformation functions to be applied
over the fields of the class instance.
"""
def __init__(self, **transformations):
"""The ``transformations`` dictionary contains the definition of how to
map the attributes of the instance of the class, at serialization time.
"""
self.serializer = EventSerializer(transformations)
def __call__(self, event_class):
"""Called when being applied to ``event_class``, will replace the
``serialize`` method of this one by a new version that uses the
serializer instance.
"""
def serialize_method(event_instance): #event_instance就是class被裝飾產生的instance
return self.serializer.serialize(event_instance)
event_class.serialize = serialize_method #裝飾event_class新增serialize函式`
return event_class #return裝飾的class
'''
Serialization裝飾器帶dict參數, 裝飾event_class
'''
@Serialization(
username=str.lower,
password=hide_field,
ip=show_original,
timestamp=format_time,
)
class LoginEvent:
def __init__(self, username, password, ip, timestamp):
self.username = username
self.password = password
self.ip = ip
self.timestamp = timestamp
class TestLoginEventSerialized(unittest.TestCase):
def test_serialization(self):
event = LoginEvent(
"UserName", "password", "127.0.0.1", datetime(2016, 7, 20, 15, 45)
)
expected = {
"username": "username",
"password": <PASSWORD>**",
"ip": "127.0.0.1",
"timestamp": "2016-07-20 15:45",
}
self.assertEqual(event.serialize(), expected)
'''
event被裝飾才有serialize function
'''
if __name__ == "__main__":
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + [markdown] id="f--LOd0IfxYI"
# 用dataclass省去__init__ boilerplate
# + id="-hEZtOetOPL8"
"""
Class decorators.
Reimplement the serialization of the events by applying a class decorator.
Use the @dataclass decorator.
This code only works in Python 3.7+
"""
import sys
import unittest
from datetime import datetime
#from decorator_class_2 import (
# Serialization,
# format_time,
# hide_field,
# show_original,
#)
try:
from dataclasses import dataclass
except ImportError:
def dataclass(cls):
return cls
@Serialization(
username=show_original,
password=hide_field,
ip=show_original,
timestamp=format_time,
)
@dataclass #用它省去__init__ boilerplate
class LoginEvent:
username: str
password: str
ip: str
timestamp: datetime
class TestLoginEventSerialized(unittest.TestCase):
@unittest.skipIf(
sys.version_info[:3] < (3, 7, 0), reason="Requires Python 3.7+ to run"
)
def test_serializetion(self):
event = LoginEvent(
"username", "password", "127.0.0.1", datetime(2016, 7, 20, 15, 45)
)
expected = {
"username": "username",
"password": <PASSWORD>**",
"ip": "127.0.0.1",
"timestamp": "2016-07-20 15:45",
}
self.assertEqual(event.serialize(), expected)
if __name__ == "__main__":
unittest.main(argv=['first-arg-is-ignored'], exit=False)
# + [markdown] id="HZmmjKqOUAW1"
# ## Passing arguments to decorators
#
# + id="A44i1LM-mr2H"
from functools import wraps
RETRIES_LIMIT = 3
'''
實作裝飾器函式帶參數,可以設定retry次數
'''
def with_retry(retries_limit=RETRIES_LIMIT, allowed_exceptions=None):
allowed_exceptions = allowed_exceptions or (ControlledException,)
def retry(operation):
@wraps(operation)
def wrapped(*args, **kwargs):
last_raised = None
for _ in range(retries_limit):
try:
return operation(*args, **kwargs)
except allowed_exceptions as e:
logging.warning(
"retrying %s due to %s", operation.__qualname__, e
)
last_raised = e
raise last_raised
return wrapped
return retry
@with_retry()
def run_operation(task):
return task.run()
@with_retry(retries_limit=5)
def run_with_custom_retries_limit(task):
return task.run()
@with_retry(allowed_exceptions=(AttributeError,))
def run_with_custom_exceptions(task):
return task.run()
@with_retry(
retries_limit=4, allowed_exceptions=(ZeroDivisionError, AttributeError)
)
def run_with_custom_parameters(task):
return task.run()
# + id="SMri4w6WpdHm"
RETRIES_LIMIT = 3
'''
同的帶參數retry,用class裝飾器實作
函式實作的裝飾器,層數較多,class較清晰
__init__負責裝飾器參數, __call__處理被裝飾的函數
'''
class WithRetry:
def __init__(self, retries_limit=RETRIES_LIMIT, allowed_exceptions=None):
self.retries_limit = retries_limit
self.allowed_exceptions = allowed_exceptions or (ControlledException,)
def __call__(self, operation):
@wraps(operation)
def wrapped(*args, **kwargs):
last_raised = None
for _ in range(self.retries_limit):
try:
return operation(*args, **kwargs)
except self.allowed_exceptions as e:
logger.info(
"retrying %s due to %s", operation.__qualname__, e
)
last_raised = e
raise last_raised
return wrapped
@WithRetry()
def run_operation(task):
return task.run()
@WithRetry(retries_limit=5)
def run_with_custom_retries_limit(task):
return task.run()
@WithRetry(allowed_exceptions=(AttributeError,))
def run_with_custom_exceptions(task):
return task.run()
@WithRetry(
retries_limit=4, allowed_exceptions=(ZeroDivisionError, AttributeError)
)
def run_with_custom_parameters(task):
return task.run()
# + [markdown] id="wnkYIbEbrfp-"
# ## Good uses for decorators
# * Transforming parameters: 對傳入的參數做一些前處理
# * Tracing code: log執行過的函式足跡
# * Validate parameters
# * Implement retry operations
# * Simplify classes by moving some (repetitive) logic into decorators: 抽象出不變的部份給decorator
# + [markdown] id="ZmVt69sAszSR"
# ## Effective decorators – avoiding common mistakes
#
# + id="3uiY9PtJsjdK"
def trace_decorator(function):
def wrapped(*args, **kwargs):
logging.info("running %s", function.__qualname__)
return function(*args, **kwargs)
return wrapped
@trace_decorator
def process_account(account_id):
"""Process an account by Id."""
logging.info("processing account %s", account_id)
help(process_account) #想取得process_account, 它會變成是wrapped
process_account.__qualname__
#顯示函數的名字、類別、模組等位址
#發現看到都是都不是wrapped,如果要trace函式的足跡狠麻煩
'''
修正它,狠簡單的
'''
def trace_decorator(function):
@wraps(function)
def wrapped(*args, **kwargs):
logging.info("running %s", function.__qualname__)
return function(*args, **kwargs)
return wrapped
@trace_decorator
def process_account(account_id):
"""Process an account by Id."""
logging.info("processing account %s", account_id)
help(process_account)
process_account.__qualname__
# + id="KAhjnOR8c1xh"
import time
from functools import wraps
'''
試範怎麼誤用,量elapsed time, 當decorator被import或是被呼叫展開了
'''
def traced_function_wrong(function):
"""An example of a badly defined decorator."""
start_time = time.time()
@wraps(function)
def wrapped(*args, **kwargs):
print("started execution of %s" % function)
result = function(*args, **kwargs)
print("function %s took %.2fs"% (function, time.time() - start_time))
return result
return wrapped
@traced_function_wrong
def process_with_delay(callback, delay=0):
print("sleep(%d)"% delay)
return callback
'''
正確使用的話應該把time函式都放入wraps
'''
def traced_function(function):
@wraps(function)
def wrapped(*args, **kwargs):
print("started execution of %s" % function)
start_time = time.time()
result = function(*args, **kwargs)
print("function %s took %.2fs"% (function, time.time() - start_time))
return result
return wrapped
@traced_function
def call_with_delay(callback, delay=0):
print("sleep(%d)" % delay)
return callback
def a_callback():
f'cb'
def main():
a_fun = process_with_delay
b_fun = call_with_delay
time.sleep(2)
a_fun(a_callback)
print("------------------------------")
b_fun(a_callback)
if __name__ == "__main__":
main()
# + [markdown] id="saCXZYkOzHub"
# 我想利用side effect的話,
#
# + id="Lu7qJFVDzFth"
'''
>>> from decorator_side_effects_2 import EVENTS_REGISTRY
>>> EVENTS_REGISTRY
{'UserLoginEvent': decorator_side_effects_2.UserLoginEvent,
'UserLogoutEvent': decorator_side_effects_2.UserLogoutEvent}
import完後,我就會知道event table裡共有哪些events
'''
EVENTS_REGISTRY = {}
def register_event(event_cls):
"""Place the class for the event into the registry to make it accessible in
the module.
"""
EVENTS_REGISTRY[event_cls.__name__] = event_cls
return event_cls
class Event:
"""A base event object"""
class UserEvent:
TYPE = "user"
@register_event
class UserLoginEvent(UserEvent):
"""Represents the event of a user when it has just accessed the system."""
@register_event
class UserLogoutEvent(UserEvent):
"""Event triggered right after a user abandoned the system."""
def test():
"""
>>> sorted(EVENTS_REGISTRY.keys()) == sorted(('UserLoginEvent', 'UserLogoutEvent'))
True
"""
# + [markdown] id="JrXOaZBV85rz"
# ## Creating decorators that will always work
# 先舉一個例子是要query db,給一個字串
# + id="x9LVA_Qu9D6e"
from functools import wraps
class DBDriver:
def __init__(self, dbstring):
self.dbstring = dbstring
def execute(self, query):
return f"query {query} at {self.dbstring}"
def inject_db_driver(function):
"""This decorator converts the parameter by creating a ``DBDriver``
instance from the database dsn string.
"""
@wraps(function)
def wrapped(dbstring):
return function(DBDriver(dbstring))
return wrapped
@inject_db_driver
def run_query(driver):
return driver.execute("test_function")
class DataHandler:
"""The decorator will not work for methods as it is defined."""
#沒法重利用inject_db_driver,因為class參數多了self
@inject_db_driver
def run_query(self, driver):
return driver.execute(self.__class__.__name__)
def main():
print(run_query('test okay'))
#print(DataHandler().run_query("test fails")) #TypeError
if __name__ == "__main__":
main()
# + id="0X0p5dHnhF0a"
from functools import wraps
from types import MethodType
class DBDriver:
def __init__(self, dbstring):
self.dbstring = dbstring
def execute(self, query):
return f"query {query} at {self.dbstring}"
class inject_db_driver:
"""Convert a string to a DBDriver instance and pass this to the wrapped
function.
"""
def __init__(self, function):
self.function = function
wraps(self.function)(self)
def __call__(self, dbstring):
return self.function(DBDriver(dbstring))
def __get__(self, instance, owner):
if instance is None: #這裡先借助chapter06的描述器, 執行class method
return self
return self.__class__(MethodType(self.function, instance))
@inject_db_driver
def run_query(driver):
return driver.execute("test_function_2")
class DataHandler:
@inject_db_driver
def run_query(self, driver):
return driver.execute("test_method_2")
def main():
print(run_query('test okay'))
print(DataHandler().run_query("fix test fails"))
if __name__ == "__main__":
main()
# + [markdown] id="TzyFqnBRmM3w"
# ## The DRY principle with decorators
# DRY Don't repeat yourself
# * Do not create the decorator in the first place from scratch. Wait until the pattern emerges and the abstraction for the decorator becomes clear, and then refactor. 不要從零開始就亂做decorator,等設計模式跟可以抽象再來重構decorator
# * Consider that the decorator has to be applied several times (at least three times)
# before implementing it. 可以重覆利用decorator三次以上
# * Keep the code in the decorators to a minimum. 實持decorator的程式輕量簡潔
# + [markdown] id="0cGEn0fwoZGC"
# ## Decorators and separation of concerns
# + id="vyV8K_l0ojAu"
'''
這個例子的decorator要log也要量時間
'''
import functools
import time
def traced_function(function):
@functools.wraps(function)
def wrapped(*args, **kwargs):
logging.info("started execution of %s", function.__qualname__)
start_time = time.time()
result = function(*args, **kwargs)
logging.info(
"function %s took %.2fs",
function.__qualname__,
time.time() - start_time,
)
return result
return wrapped
@traced_function
def operation1():
time.sleep(2)
logging.info("running operation 1")
return 2
# + id="Ng7Y03bzofWt"
import time
from functools import wraps
import logging
'''
把log和量時間分開成兩個decorator
'''
def log_execution(function):
@wraps(function)
def wrapped(*args, **kwargs):
logging.info("started execution of %s", function.__qualname__)
return function(*kwargs, **kwargs)
return wrapped
def measure_time(function):
@wraps(function)
def wrapped(*args, **kwargs):
start_time = time.time()
result = function(*args, **kwargs)
logging.info(
"function %s took %.2f",
function.__qualname__,
time.time() - start_time,
)
return result
return wrapped
@measure_time
@log_execution
def operation():
time.sleep(3)
logging.info("running operation...")
return 33
| Clean_Code_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
import pandas as pd
# + jupyter={"outputs_hidden": true} tags=[]
df_mon = pd.read_csv('C:/Users/andre/Desktop/Montercarlosimulation/monday.csv', parse_dates=True, index_col=0, sep=';')
df_mon
# + jupyter={"outputs_hidden": true} tags=[]
df_tue= pd.read_csv('C:/Users/andre/Desktop/Montercarlosimulation/tuesday.csv', parse_dates=True, index_col=0, sep=';')
df_tue
# + jupyter={"outputs_hidden": true} tags=[]
df_wed= pd.read_csv('C:/Users/andre/Desktop/Montercarlosimulation/wednesday.csv', parse_dates=True, index_col=0, sep=';')
df_wed
# + jupyter={"outputs_hidden": true} tags=[]
df_thu= pd.read_csv('C:/Users/andre/Desktop/Montercarlosimulation/thursday.csv', parse_dates=True, index_col=0, sep=';')
df_thu
# + jupyter={"outputs_hidden": true} tags=[]
df_fri= pd.read_csv('C:/Users/andre/Desktop/Montercarlosimulation/friday.csv', parse_dates=True, index_col=0, sep=';')
df_fri
# -
df_cos_1= df_fri.loc[df_fri['customer_no']<5]
df_cos=df_fri.groupby("location").count()
df_cos
df_cos_1
df_fri['location'].hist();
df= pd.concat([df_mon,df_tue,df_wed,df_thu,df_fri])
df
# a) Total number of customer in each section for the whole week
df_cos=df.groupby("location").count()
df_cos
# + tags=[]
# -
df['customer_ID'] = df['customer_no'].astype(str) +'_'+ df.index.astype(str)
df
df['Day']=df.index.day
df
# b) Total number of customers in each section over time.
df_cos_ot=df.groupby(['Day','location']).count()
df_cos_ot.iloc[:,1]
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
df_cos_ot
# -
# c) Number of customers at checkout over time (Day)
df_checkout= df[df['location']=='checkout'].groupby('Day').count()
df_checkout['location']
# + tags=[]
# + tags=[]
df
# -
# d) calculate the time each customer spent in the market
df['Day_customer']=df['Day'].astype(str)+'_'+df['customer_no'].astype(str)
df
df2= df.sort_values('Day_customer')
df2['Day_customer']
df3=df2.groupby('Day_customer').max()
df3
import numpy as np
# + tags=[]
df['hour'] = df.index.strftime('%H:00')
df['date'] = df.index.date
df['date_h'] = df[['date', 'hour']].astype(str).apply(' '.join,1)
df.drop(['hour','date'], axis=1,inplace=True)
df
# -
df['time']=df.index
df.groupby('Day_customer')['time'].agg(np.ptp)
pd.DatetimeIndex(df['time'])
df['time_hour'] = df['Day'].astype(str) + '_' + pd.DatetimeIndex(df['time']).hour.astype(str)
df
df.groupby('time_hour').count()
# +
df3=df.groupby('time_hour')['customer_no'].value_counts()
# -
df3.groupby('time_hour').count().head(30)
df_sort=df.sort_values(by=['Day_customer','time'])
df_sort.head(30)
df_sort.iloc[1,7]
df_sort.shape
df_sort
# + jupyter={"source_hidden": true} tags=[]
#df_sort_2= df_sort.drop(df_sort.loc[df_sort['location']=='checkout'])
#df_sort_3=df_sort.loc[df_sort["location"]!="checkout"]
#df_sort_3
# -
# df_time = df.set_index('time')
# +
# df_sort.set_index('time')
# + tags=[]
df_sort_r=df_sort.groupby('Day_customer').resample('1min').ffill()
# + tags=[]
df_sort_r.head(30)
# -
df_sort_r['after']=df_sort_r['location'].shift(-1)
df_sort_r['time'].min()
# + tags=[]
df_sort_r.head(30)
# -
df_sort
# + tags=[]
df_sort_2=df_sort.groupby(['Day_customer'])['time'].min()
# -
df_sort_r
this_was_WAY_easier = df_sort_r[df_sort_r['location']=='checkout']['after']
#we need to drop nans then
this_was_WAY_easier.value_counts(normalize=True)
this_was_WAY_easier
df_sort_r
# ## MASTER MIND MATRIX
MMM = pd.crosstab(df_sort_r['after'],df_sort_r['location'], normalize=0)
MMM
# MMM Master Mind Matrix
type(MMM)
MMM['checkout']=[1,0,0,0,0]
MMM
# ## Random Choices
import random
thank_you_andres = this_was_WAY_easier.value_counts(normalize=True)
thank_you_andres
type(thank_you_andres)
initial_state_andres = random.choices(list(thank_you_andres.index), weights=thank_you_andres)
initial_state_andres
initial_state_andres
df_sort_r
# +
import random
import pandas as pd
from faker import Faker
f=Faker()
name= f.name()
class Customer:
"""
a single customer that moves through the supermarket
in a MCMC simulation
"""
def is_active(self):
"""Returns False if a customer has reached the checkout,other wise returns True """
if self.state == 'checkout':
return False
else:
return True
def __init__(self, transition_probs=0, budget=100):
self.name = f.name()
self.state = ''.join(initial_state_andres)
self.budget = budget
self.transition_probs = MMM
def __repr__(self):
return f'<Customer {self.name} in {self.state}>'
def next_state(self):
"""
propagates the customer to the next state , returns nothing
:return: nothing
"""
print(self.transition_probs.index)
print('---------')
print(self.transition_probs[self.state])
self.state = random.choices(list(self.transition_probs.index), weights=self.transition_probs[self.state])[0]
def next_state_p(self):
self.state = random.choices(list(self.transition_probs.index), weights=self.transition_probs[self.state])[0]
return self.state
cust1 = Customer()
cust2 = Customer()
print(cust1.name, cust1.state)
print(cust2.name, cust2.budget)
cust1.next_state()
print('--------------------')
print(cust1.name, cust1.state)
# -
customer3=Customer()
customer3.next_state()
customer3.next_state_p()
# + tags=[]
total_customers=[]
for x in range(100):
Cus= Customer()
row=[]
for i in range(20):
row.append(Cus.next_state_p())
total_customers.append(row)
Path=pd.DataFrame(total_customers)
Path
# +
Path_T= Path.T
Path_T
# -
Path_T.mode()
Path.value_counts()
| Project_08 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from sklearn import datasets
from sklearn.model_selection import cross_val_predict
from sklearn import linear_model
import matplotlib.pyplot as plt
lr = linear_model.LinearRegression()
boston = datasets.load_boston()
y = boston.target
# cross_val_predict returns an array of the same size as `y` where each entry
# is a prediction obtained by cross validation:
predicted = cross_val_predict(lr, boston.data, y, cv=10)
fig, ax = plt.subplots()
ax.scatter(y, predicted, edgecolors=(0, 0, 0))
ax.plot([y.min(), y.max()], [y.min(), y.max()], 'k--', lw=4)
ax.set_xlabel('Measured')
ax.set_ylabel('Predicted')
plt.show()
| statistics-notebook/tests/notebooks/scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Science Internship at Widhya
# ## Mission: Instagram Post Reach Predicition
# #### Importing required liberaries
import pandas as pd
import numpy as np
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
# #### Reading and Previewing dataset
df=pd.read_excel('insta.xlsx')
df.head()
# #### Removing unwanted features
df = df.drop(['S.No','USERNAME','Caption','Hashtags'],axis=1)
df
# #### Data preprocessing
df['Time']=df['Time since posted'].str.split(" ",n=1,expand=True)[0].astype(int)
df.head()
df.drop(['Time since posted'],axis=1)
df = df[['Followers','Time','Likes']]
df.head()
# #### Dividing and Spliting data into traing and testing set
df1 = df.values
X, y = df1[:,:-1], df1[:,-1]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.20, random_state=42)
# #### Feature scaling
scaled_features = StandardScaler().fit_transform(X_train, X_test)
# #### Model Building
l_reg=LinearRegression()
l_reg.fit(X_train,y_train)
print("Training completed")
# #### Training and Testing Score achived by model
print('Training Score: ', round(l_reg.score(X_train,y_train)*100,2),'%')
print('Testing Score: ', round(l_reg.score(X_test,y_test)*100,2))
# #### Prediction on test set
y_predict=l_reg.predict(X_test)
# #### Accuracy acchieved by Model
from sklearn import metrics
acc=metrics.r2_score(y_test,y_predict)
print("Accuracy Score of Model: ",round(acc*100,2),'%')
# #### Model Evaluation
from sklearn import metrics
print('Mean Absolute Error:',round(metrics.mean_absolute_error(y_test,y_predict),2))
print('Mean Squared Error:',round(metrics.mean_squared_error(y_test,y_predict),2))
print('Root Mean Squared Error:',round(np.sqrt(metrics.mean_squared_error(y_test,y_predict)),2))
print('Explained Variance Score:',round(metrics.explained_variance_score(y_test,y_predict),2))
# #### Sample Predicition
# >Prediction for the post's reach for someone who has 300 followers and after 10 hours
print(l_reg.predict([[300,10]]))
# >Prediction for the post's reach for someone who has 400 followers and after 5 hours
print(l_reg.predict([[400,5]]))
# ### Thank you :)
| Instagram Post Reach Predicition/Instagram_Post_Reach_Prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest
#
# Before we get into the guts of Random Forest Classification, lets get familiar with 2 concepts:
# - Bootstrapping
# - Bagging
#
# ## Bootstrapping
#
# **What is it ?**
# It is a method of accurately determining a _statistic_. These Statistics are aggregate quantities that cannot be directly observed. For example, the _mean_ of test scores of students in a class. The quantity is _accurately determined_ by minimizing as much as possible.
#
# **How do we do it?**
# Consider a school with a 1000 students. One can easily compute the mean math score of all students $X_{mean}$ with a simple formula.
#
# $$X_{mean} = \frac{\sum_{i=1}^{1000} x_{i}}{1000}$$
#
# This is fantastic. However, computing the mean of a population may not yeild useful results. Lets take a really simple example of this in action. Say we have the test scores of 10 students as follows.
#
# ```
# [10, 25, 36, 37, 32, 85, 49, 97, 24, 23]
# ```
#
# The mean of this computed from the formula above is $41.8$ . Lets see how different the mean is when bootstrapping. In bootstrapping, we perform the following steps:
# 1. Make $M$ of subsamples from the population (repetition is allowed)
# 2. Calculate the mean of each subsaple clutter.
# 3. Calculate the mean of each of those means. Mean of the means!
#
# Considering a population of size $N$ divided into $M$ subsamples with repetition, the general formula for computing the new mean is as follows:
#
# $$X_{Bmean} = \frac{\sum_{i=1}^{M}\sum_{j=1}^{len(M_{i})} x_{i,j}}{M}$$
#
# Say we create 6 subsamples in the following way:
# ```
# [10, 36, 32, 49]
# [25, 32, 97, 10]
# [36, 85, 24]
# [97, 23, 24, 25]
# [85, 10, 36]
# [23, 32, 24, 25, 36]
# ```
# The mean for each is $31.75, 41, 48.33, 42.25, 43.66, 28$. Taking $M = 6$ in the equation above, the bootstrapped mean becomes $39.165$. The difference with the original mean $41.8$ may seem trivial. However, this is just a simple example to illustrate bootstrapping. Real applications would involve the construction of hundreds of sample clutters from a population. Bootstrapped quantities better represent the statistic of the population. Greater the number of equally distributed subsamples can yield more accurate statistics.
#
#
# ## Bagging
#
#
| notebooks/.ipynb_checkpoints/Random Forest-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model Agnostic explainability with KernelExplainers
# ## CHAPTER 07 - *Practical exposure of using SHAP in ML*
#
# From **Applied Machine Learning Explainability Techniques** by [**<NAME>**](https://www.linkedin.com/in/aditya-bhattacharya-b59155b6/), published by **Packt**
# ### Objective
#
# We will explore the usage of KernelExplainers in SHAP based on the Kernel SHAP algorithm used for model agnostic explainability. We will use the same dataset with almost similar approach as seen with the [TreeExplainer tutorial](https://github.com/PacktPublishing/Applied-Machine-Learning-Explainability-Techniques/blob/main/Chapter07/TreeExplainers.ipynb). So, the initial part of the tutorial might appear to be exactly the same, but the model explainability is executed with KernelExplainers in SHAP. Please check out *Chapter 7 - Practical exposure of using SHAP in ML* for other interesting approaches of using SHAP in practice.
# ### Installing the modules
# Install the following libraries in Google Colab or your local environment, if not already installed.
# !pip install --upgrade numpy pandas matplotlib seaborn sklearn lightgbm shap
# ### Loading the modules
# +
import warnings
warnings.filterwarnings('ignore')
import numpy as np
np.random.seed(123)
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix, roc_auc_score
from sklearn.preprocessing import LabelEncoder
import lightgbm as lgb
import shap
print(f"Shap version used: {shap.__version__}")
# Initialize JS visualization for the notebook
shap.initjs()
# -
# ### About the dataset
# **German Credit Risk** | [Kaggle](https://www.kaggle.com/uciml/german-credit)
#
# The dataset that we will be using for this tutorial for used for classifying Good Risk and Bad Risk. The dataset is a derived dataset taken from Kaggle. The original source of the dataset is [UCI](https://archive.ics.uci.edu/ml/datasets/statlog+(german+credit+data)) which contains 1000 entries with 20 categorial or symbolic features prepared by Professor Dr. <NAME>, Institut f"ur Statistik und "Okonometrie Universit"at Hamburg, such that each entry represents a person who takes a credit by a bank. Each applicant is categorized as good or bad credit risks according to the set of features present in the dataset. But the current dataset is a simplified one with less number of features and the dataset has been added in the project repository to make it more accessible. The following are the key attributes present in dataset according to the Kaggle source:
# - Age (numeric)
# - Sex (text: male, female)
# - Job (numeric: 0 - unskilled and non-resident, 1 - unskilled and resident, 2 - skilled, 3 - highly skilled)
# - Housing (text: own, rent, or free)
# - Saving accounts (text - little, moderate, quite rich, rich)
# - Checking account (numeric, in DM - Deutsch Mark)
# - Credit amount (numeric, in DM)
# - Duration (numeric, in month)
# - Purpose (text: car, furniture/equipment, radio/TV, domestic appliances, repairs, education, business, vacation/others)
# ### Loading the data
data = pd.read_csv('datasets/german_credit_data.csv', index_col=0)
data.head()
data.shape
data.columns
data.info()
num_features = ['Age', 'Credit amount', 'Duration']
cat_features = ['Sex', 'Job', 'Housing', 'Saving accounts', 'Checking account','Purpose']
data[num_features].describe()
# From the dataset info, we can see that the features *Saving accounts* and *Checking account* does have some missing values. The percentage of missing values also seems to be higher for ignoring the entire row of records and hence we might need to apply data imputation.
sns.displot(
data=data.isna().melt(value_name="missing"),
y="variable",
hue="missing",
multiple="fill",
aspect=1.5,
palette='seismic'
)
plt.show()
missing_features = ['Saving accounts', 'Checking account']
data[missing_features].isna().sum()/1000 * 100
print(data[missing_features[0]].value_counts())
print(data[missing_features[1]].value_counts())
# For the features having moderately high amount of missing values, we cannot completely ignore the records or drop the feature. So, we need to definitely apply data imputation. For simplifying the data imputation technique, we will impute these categorical features with another "Unknown" category, but ideally there needs to some experimentation and the thought process should be more robust to select other types of imputation. But like the previous tutorials, our goal is to focus on the model explainability part and not towards building robust ML models. Even if the model is not good enough, we can focus on using SHAP to explain the same. But to proceed with the explainability process, we will apply median imputation and build a baseline ML model using a tree ensemble algorithm.
data.fillna('Unknown', inplace=True)
print(data[missing_features].isna().sum()/1000 * 100)
print(data[missing_features[0]].value_counts())
print(data[missing_features[1]].value_counts())
# Check for duplicate values
data.duplicated().any()
# We will use **Light Gradient Boosting Machine (LGBM) algorithm** (https://lightgbm.readthedocs.io/en/latest/) for the ML modelling part, which can directly use categorical variables, and hence we will not go for One-Hot Encoding. Otherwise, it is essential to apply OneHot Encoding before proceeding.
# Label Encoding output variable
le = LabelEncoder()
for feat in ['Sex', 'Housing', 'Saving accounts','Checking account', 'Purpose','Risk']:
le.fit(data[feat])
data[feat] = le.transform(data[feat])
classes = list(le.classes_)
print(classes)
data.head()
# For this tutorial, we will stop at here for the data pre-processing or feature engineering part. We will not perform any other complex feature engineering, data normalization or any other transformation as the primary objective is to focus on model explainability using SHAP KernelExplainers. So, let's proceed building a baseline modell using the **Light Gradient Boosting Machine (LGBM) algorithm** (https://lightgbm.readthedocs.io/en/latest/) which will be considered as a blac-box model. Let us proceed!
# ### Training the model
features = data.drop(columns=['Risk'])
labels = data['Risk']
# Dividing the data into training-test set with 80:20 split ratio
x_train,x_test,y_train,y_test = train_test_split(features,labels,test_size=0.2, random_state=123)
x_train.columns
# Create LightGBM Dataset
data_train = lgb.Dataset(x_train, label=y_train, categorical_feature=cat_features)
data_test = lgb.Dataset(x_test, label=y_test, categorical_feature=cat_features)
# Define the model configurations as a dictionary
params = {
'boosting_type': 'gbdt',
'objective': 'binary',
'metric': 'auc',
'num_leaves': 20,
'learning_rate': 0.05,
'feature_fraction': 0.9,
'bagging_fraction': 0.8,
'bagging_freq': 5,
'verbose': -1,
'lambda_l1': 1,
'lambda_l2': 1,
'seed': 123
}
model = lgb.train(params,
data_train,
num_boost_round=100,
verbose_eval=100,
valid_sets=[data_test, data_train])
y_pred = model.predict(x_test)
y_pred = [1 if y > 0.5 else 0 for y in y_pred]
print(f'Accuracy for the baseline model is: {accuracy_score(y_test, y_pred)}')
# Thus, we can see a model which is decent, but it is definitely over-fitting. Now, we would like to focus on the model explainability using SHAP KernelExplainers.
# ### Model Agnostic Explainability using SHAP KernelExplainers
# %%time
# Building a SHAP Explainer model
explainer = shap.KernelExplainer(model.predict, x_train)
shap_values = explainer.shap_values(x_test, nsamples=100)
# Global explainability with summary plots
shap.summary_plot(shap_values, x_test, plot_type='violin', show=False)
plt.gcf().axes[-1].set_box_aspect(10)
# KernelExplainers work with any models. Kernel SHAP algorithm used in KernelExplainers use a specially weighted local linear regression to approximate SHAP values. It is model-agnostic as it does not make assumption about any model.
# Local explainability with force plots
shap.force_plot(explainer.expected_value, shap_values[1], x_test.iloc[0,:])
# Force plots can be used for local interpretability. Certain features shown in the red side are used to push the prediction value higher, while the features shown in blue tries to lower the prediction value and negatively impact the model's decision making process.
shap.force_plot(explainer.expected_value, shap_values, x_test)
# Force plots can also be used for global interpretability. But personally I think these are quite difficult to interpret! In fact, I find decision plots much easier to understand and provides a better explainability.
shap.decision_plot(explainer.expected_value, shap_values[1], x_test.iloc[0,:])
# The deviation illustrated in decision plots makes it easier to comprehend such a plot and actionable insights can be drawn by comparing the difference between the local value and the mean values as shown in the plot.
# ## Final Thoughts
# KernelExplainers are very helpful as these are model agnostic and easy to use just like other SHAP explainers. But KernelExplainers can be slow. I would recommend you to look at other examples as provided in the project repository for SHAP: https://github.com/slundberg/shap/tree/master/notebooks/tabular_examples/model_agnostic
# ## Reference
# 1. German Credit Risk Dataset - https://www.kaggle.com/uciml/german-credit
# 2. SHAP GitHub Project - https://github.com/slundberg/shap
# 3. SHAP Documentations - https://shap.readthedocs.io/en/latest/index.html
# 4. Some of the utility functions and code are taken from the GitHub Repository of the author - <NAME> https://github.com/adib0073
| Chapter07/KernelExplainers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mask Generation with OpenCV
#
# ## Introduction
#
# In the paper they generate irregular masks by using occlusion/dis-occlusion between two consecutive frames of videos, as described in [this paper](https://lmb.informatik.uni-freiburg.de/Publications/2010/Bro10e/sundaram_eccv10.pdf).
#
# Instead we'll simply be using OpenCV to generate some irregular masks, which will hopefully perform just as well. We've implemented this in the function `_generate_mask`, which is located in the `util.py` file in the libs directory
# +
import os
import itertools
import matplotlib
import matplotlib.pyplot as plt
# Change to root path
if os.path.basename(os.getcwd()) != 'PConv-Keras':
os.chdir('..')
# Import function that generates random masks
from libs.util import MaskGenerator
# -
# %matplotlib inline
# Let us review of the code of this function
??MaskGenerator._generate_mask
# Let's create some output samples with this function to see what it does
# +
# Instantiate mask generator
mask_generator = MaskGenerator(512, 512, 3, rand_seed=42)
# Plot the results
_, axes = plt.subplots(5, 5, figsize=(20, 20))
axes = list(itertools.chain.from_iterable(axes))
for i in range(len(axes)):
# Generate image
img = mask_generator.sample()
# Plot image on axis
axes[i].imshow(img*255)
# -
# I've also implemented a function which will load masks from a directory, and potentially augments them. This can be used to load masks published with the original paper. E.g. get the original training and testing masks from http://masc.cs.gmu.edu/wiki/partialconv, and save them to `data/masks/train/` and `data/masks/test/`, respectively.
# No need to run the next cell. This is only needed if you want to load the masks from a directory (as done in original paper). Since we will use the masks generated with OpenvCV, we leave the next cell un-executed.
# +
# Instantiate mask generator
mask_generator = MaskGenerator(512, 512, 3, rand_seed=42, filepath='./data/masks/train')
# Plot the results
_, axes = plt.subplots(5, 5, figsize=(20, 20))
axes = list(itertools.chain.from_iterable(axes))
for i in range(len(axes)):
# Generate image
img = mask_generator.sample()
# Plot image on axis
axes[i].imshow(img*255)
# -
| notebooks/Step1 - Mask Generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# # %%
# # Plots on selected model
#
# * model: (50t,600,10)
# * scale: 125
# +
import pandas as pd
import numpy as np
from helpers import potus
import glob
import re
import os
import matplotlib.pyplot as plt
import seaborn as sns
# -
import imp
imp.reload(potus)
# ## (50t,600,10)
#
# Load results for scales 125 and 25.
# +
exp = 'potus_50t_600_10'
scales = [25]
result = potus.gridExpResult(exp, scales)
print(len(result.df))
print(len(result.settings))
display(result.settings)
display(result.df.groupby('Nw').speech.count())
# -
# # Prepare ts object
#
#
# a pandas data frame with a date column set as index. date is converted using `pd.to_datetime`
result.df.date = pd.to_datetime(result.df.date)
ts = result.df[['date', 'file_name', 'speaker', 'title', 'speech', 'novelty', 'transience', 'resonance', 'speech_len', 'probs', 'token_count', 'raw_tuple']].set_index('date')
ts.head()
ts.loc['2008-01-01':'2018-12-31']
# ## now average across all scales to arrive at mean metrics for each speech
# +
many_window_values = ts.pivot_table(index='file_name', values = ['novelty', 'transience', 'resonance'], aggfunc='mean')
ts = ts.drop_duplicates(subset=['file_name'])
ts = ts[['file_name', 'speaker', 'title', 'speech', 'speech_len', 'probs', 'token_count', 'raw_tuple']]
ts['date'] = ts.index
ts = pd.merge(ts,
many_window_values,
on='file_name',
how='left')
ts = ts.set_index('date')
print(ts.head())
# +
# best that we remove the n speeches at each end, since they do not have complete metrics
n = 25
e = 962-n
df = ts[n:e]
# -
# # plots
sns.set(rc={'figure.figsize':(11, 4)})
plot_cols = ['novelty', 'transience', 'resonance']
df[plot_cols].plot(
figsize=(11, 9),
color=['red', 'blue', 'green'],alpha=0.7,
subplots=True)
# +
import seaborn as sns
import matplotlib.pyplot as plt
cmap = plt.cm.get_cmap('OrRd')
# between 0 and 1, 0 for the leftmost color of the range, 1 for the rightmost, upper > lower
upper_color = 1.0
lower_color = 0.2
num_colors = 15 # len(self.data.columns)
colors = cmap(np.linspace(lower_color, upper_color, num_colors))
plt.figure(figsize=(6,6))
b = sns.kdeplot(x=df['novelty'], y=df['transience'], fill=True, colors=colors, thresh=0.015, alpha=1)
b.set_ylabel("Transience",fontsize=20)
b.set_xlabel("Novelty",fontsize=20)
b.tick_params(labelsize=12)
y_lim = plt.ylim()
x_lim = plt.xlim()
plt.plot(x_lim, y_lim, color = '#dadada', alpha = 0.5)
# -
plt.figure(figsize=(6,6))
b = sns.kdeplot(x=df['novelty'], y=df['resonance'], fill=True, colors=colors, thresh=0.015, alpha=1)
b.set_ylabel("Resonance",fontsize=20)
b.set_xlabel("Novelty",fontsize=20)
b.tick_params(labelsize=12)
plt.ylim(-4,4)
y_lim = [0,0]
x_lim = plt.xlim()
plt.plot(x_lim, y_lim, color = '#dadada', alpha = 0.5)
# +
import seaborn as sns
import matplotlib.pyplot as plt
cmap = plt.cm.get_cmap('Oranges')
# between 0 and 1, 0 for the leftmost color of the range, 1 for the rightmost, upper > lower
upper_color = 1
lower_color = 0.4
num_colors = 15 # len(self.data.columns)
colors = cmap(np.linspace(lower_color, upper_color, num_colors))
plt.figure(figsize=(6,6))
sns.relplot(x='novelty', y='transience', data=df, alpha=0.8, color = 'orange')
b = sns.kdeplot(x=df['novelty'], y=df['transience'], fill=True, colors=colors, thresh=0.3, alpha=0.4)
b.set_ylabel("Transience",fontsize=20)
b.set_xlabel("Novelty",fontsize=20)
b.tick_params(labelsize=12)
y_lim = plt.ylim()
x_lim = plt.xlim()
plt.plot(x_lim, y_lim, color = '#dadada', alpha = 0.5)
# +
import seaborn as sns
import matplotlib.pyplot as plt
cmap = plt.cm.get_cmap('Oranges')
# between 0 and 1, 0 for the leftmost color of the range, 1 for the rightmost, upper > lower
upper_color = 1
lower_color = 0.4
num_colors = 15 # len(self.data.columns)
colors = cmap(np.linspace(lower_color, upper_color, num_colors))
plt.figure(figsize=(6,6))
sns.relplot(x='novelty', y='resonance', data=df, alpha=0.8, color = 'orange')
b = sns.kdeplot(x=df['novelty'], y=df['resonance'], fill=True, colors=colors, thresh=0.3, alpha=0.4)
b.set_ylabel("Resonance",fontsize=20)
b.set_xlabel("Novelty",fontsize=20)
b.tick_params(labelsize=12)
plt.ylim(-4.5,4.5)
y_lim = [0,0]
x_lim = plt.xlim()
plt.plot(x_lim, y_lim, color = '#dadada', alpha = 0.5)
# -
print(df.sort_values('resonance', ascending=False)[:10][['speaker', 'title', 'resonance', 'novelty']])
print(df.sort_values('resonance', ascending=True)[:10][['speaker', 'title', 'resonance', 'novelty']])
df.loc['1913-03-04']['probs']
# # That's enough; didn't use the rest from here onwards
ts[plot_cols].plot(
marker='.',
alpha=.5,
linestyle='None',
figsize=(11, 9),
subplots=True)
ts.query('speaker == "obama"').plot()
# +
# monthly resampling,
ts_monthly = ts.resample('M').max() #mean or other aggregate
# -
ts_monthly.plot(
#marker='.',
#linestyle='None',
figsize=(11, 9),
subplots=True)
| Insights_potus_kld_plots_and_leaders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from abc import ABC, abstractmethod
from anytree import AnyNode, RenderTree
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
np.random.seed(1)
# -
# ## Exercise 1
# #### Abstract Base Class
# Every subclass of "Expression" must implement these functions
class Expression(ABC):
@abstractmethod
def __str__(self):
pass
@abstractmethod
def eval(self):
pass
@abstractmethod
def derivative(self, var):
pass
@abstractmethod
def simplify(self):
pass
# #### Constant Class
class Const(Expression):
def __init__(self, value):
self.value = value
def __str__(self):
return str(self.value)
def eval(self):
return self.value
def derivative(self, var):
return Const(0)
def simplify(self):
return self
# #### Variable Class
class Var(Expression):
def __init__(self, name, value=None):
self.name = name
self.value = value
def __str__(self):
return self.name
def eval(self):
if self.value is not None:
return self.value
else:
try:
return env[self.name]
except:
raise Exception(f"Variable {self.name} not initialized!")
def derivative(self, var):
if self.name == var:
return Const(1)
else:
return Const(0)
def simplify(self):
return self
# #### Sum Class
class Sum(Expression):
def __init__(self, left, right):
self.left = left
self.right = right
def __str__(self):
return f"({str(self.left)}+{str(self.right)})"
def eval(self):
return self.left.eval() + self.right.eval()
def derivative(self, var):
return Sum(self.left.derivative(var), self.right.derivative(var))
def simplify(self):
sx = self.left.simplify()
sy = self.right.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
try:
sy_val = sy.eval()
except:
sy_val = None
if sx_val == 0:
return sy
elif sy_val == 0:
return sx
elif isinstance(sx, Const) and isinstance(sy, Const):
return Const(sx_val + sy_val)
else:
return Sum(sx, sy)
# #### Subtraction Class
class Sub(Expression):
def __init__(self, left, right):
self.left = left
self.right = right
def __str__(self):
return f"({str(self.left)}-{str(self.right)})"
def eval(self):
return self.left.eval() - self.right.eval()
def derivative(self, var):
return Sub(self.left.derivative(var), self.right.derivative(var))
def simplify(self):
sx = self.left.simplify()
sy = self.right.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
try:
sy_val = sy.eval()
except:
sy_val = None
if sy_val == 0:
return sx
elif isinstance(sx, Const) and isinstance(sy, Const):
return Const(sx_val - sy_val)
elif sx_val == 0:
return Mul(Const(-1), sy)
else:
return Sub(sx, sy)
# #### Multiplication Class
class Mul(Expression):
def __init__(self, left, right):
self.left = left
self.right = right
def __str__(self):
return f"({str(self.left)}*{str(self.right)})"
def simplify(self):
if any((self.left.eval() == 0, self.right.eval() == 0)):
return Const(0)
else:
return self
def eval(self):
return self.left.eval() * self.right.eval()
def derivative(self, var):
return Sum(
Mul(self.left.derivative(var), self.right),
Mul(self.left, self.right.derivative(var)),
)
def simplify(self):
sx = self.left.simplify()
sy = self.right.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
try:
sy_val = sy.eval()
except:
sy_val = None
if sx_val == 0 or sy_val == 0:
return Const(0)
if sx_val == 1:
return sy
if sy_val == 1:
return sx
elif isinstance(sx, Const) and isinstance(sy, Const):
return Const(sx_val * sy_val)
else:
return Mul(sx, sy)
# #### Division Class
class Div(Expression):
def __init__(self, left, right):
self.left = left
self.right = right
def __str__(self):
return f"({str(self.left)}/{str(self.right)})"
def eval(self):
return self.left.eval() / self.right.eval()
def derivative(self, var):
dy = self.right.derivative(var)
return Div(
Sub(
Mul(
self.left.derivative(var),
self.right,
Mul(self.left, dy),
Mul(dy, dy),
)
)
)
def simplify(self):
sx = self.left.simplify()
sy = self.right.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
try:
sy_val = sy.eval()
except:
sy_val = None
if sx_val == 0:
return Const(0)
if sy_val == 0:
raise Exception("division by zero")
if sy_val == 1:
return sx
elif isinstance(sx, Const) and isinstance(sy, Const):
return Const(sx_val / sy_val)
else:
return Div(sx, sy)
# #### Exponential Function Class
class Exp(Expression):
def __init__(self, value):
self.value = value
def __str__(self):
return f"exp({str(self.value)})"
def eval(self):
return np.exp(self.value.eval())
def derivative(self, var):
return Mul(Exp(self.value), self.value.derivative(var))
def simplify(self):
sx = self.value.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
if sx_val == 0:
return Const(1)
elif isinstance(sx, Const):
return Const(np.exp(sx_val))
else:
return Exp(sx)
# #### Logarithm Class
class Log(Expression):
def __init__(self, value):
self.value = value
def __str__(self):
return f"log({str(self.value)})"
def eval(self):
return np.log(self.value.eval())
def derivative(self, var):
return Div(self.value.derivative(var), self.value)
def simplify(self):
sx = self.value.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
if sx_val <= 0:
raise Exception("logarithm of non-positive number")
elif isinstance(sx, Const):
return Const(np.log(sx_val))
else:
return Log(sx)
# #### Hyperbolic Tangent Class
class Tanh(Expression):
def __init__(self, value):
self.value = value
def __str__(self):
return f"tanh({str(self.value)})"
def eval(self):
return np.tanh(self.value.eval())
def derivative(self, var):
tx = Tanh(self.value)
return Mul(Sub(Const(1), Mul(tx, tx)), self.value.derivative(var))
def simplify(self):
sx = self.value.simplify()
try:
sx_val = sx.eval()
except:
sx_val = None
if sx_val == 0:
return Const(0)
elif isinstance(sx, Const):
return Const(np.tanh(sx_val))
else:
return Tanh(sx)
# #### Define a computation graph
# +
x = Var("x")
y = Var("y")
z = Sum(Mul(Mul(Const(4), x), y), Exp(Mul(Const(-1), y)))
# -
# #### Print the graph
print(z)
# #### Visualize as a tree
def build_tree(expr):
class_name = expr.__class__.__name__
if isinstance(expr, Const):
return AnyNode(label=class_name, value=expr.value)
if isinstance(expr, Var):
return AnyNode(label=class_name, value=expr.value, name=expr.name)
if "left" in expr.__dir__():
children = [build_tree(expr.left),build_tree(expr.right)]
else:
children = [build_tree(expr.value)]
return AnyNode(label=class_name, children=children)
z_tree = build_tree(z)
print(RenderTree(z_tree))
# #### Computing the value of an expression
x = Var("x", 2)
y = Var("y", 3)
z = Sum(Mul(Mul(Const(4), x), y), Exp(Mul(Const(-1), y)))
z.eval()
# #### Differentiating an expression
dz_dx = z.derivative("x")
print(dz_dx)
# #### Simplify the derivative graph
dz_dx_simple = dz_dx.simplify()
print(dz_dx_simple)
print(dz_dx_simple.derivative("y").simplify())
# ### Training a network
# #### Forward pass
# +
x1 = Var("x1")
x2 = Var("x2")
b1 = Var("b1")
w11 = Var("w11")
w21 = Var("w21")
b2 = Var("b2")
w12 = Var("w12")
w22 = Var("w22")
z1_out = Tanh(Sum(b1, Sum(Mul(x1, w11), Mul(x2, w21))))
z2_out = Tanh(Sum(b2, Sum(Mul(x1, w12), Mul(x2, w22))))
c = Var("c")
u1 = Var("u1")
u2 = Var("u2")
f_in = Sum(c, Sum(Mul(z1_out, u1), Mul(z2_out, u2)))
f_out = Div(Const(1), Sum(Const(1), Exp(Mul(Const(-1), f_in))))
# -
print(f_out)
f_out_tree = build_tree(f_out)
print(RenderTree(f_out_tree))
# set variable values through environment:
env = {
"b1": 1.543385,
"w11": 3.111573,
"w12": -2.808800,
"b2": 1.373085,
"w21": 3.130452,
"w22": -2.813466,
"c": -4.241453,
"u1": 4.036489,
"u2": 4.074885,
"x1": 1,
"x2": -1,
}
z1_out.eval()
# #### Loss
# +
y = Var("y")
loss = Sum(Sub(f_in, Mul(f_in, y)), Log(Sum(Const(1), Exp(Mul(Const(-1), f_in)))))
# -
print(loss)
# #### Gradients
# Note: when variable values can be updated from "outside" (i.e., through the env dict)
# simplification should not be used in the way it is implemented here (a simplification could only be valid
# for a given value of the variable, and changing this value after the fact could thus make the simplification invalid)
param_names = ("b1", "w11", "w12", "b2", "w21", "w22", "c", "u1", "u2")
gradient_graphs = list(map(lambda x: loss.derivative(x).simplify(), param_names))
print(gradient_graphs[1])
# #### Training data
train_data = pd.DataFrame(
data=np.array([[0, 1, 0, -1, 0], [0, 0, -1, 0, 1], [1, 0, 0, 0, 0]]).T,
columns=["x1", "x2", "y"],
)
train_data
# #### Initialization
def glorot_initialization():
b = np.sqrt(6 / 4)
return np.random.uniform(-b, b, 1)[0]
for p in param_names:
if "b" in p or "c" in p:
env[p] = 0
else:
env[p] = glorot_initialization()
env
# #### Training loop
losses = []
epoch_losses = []
for e in range(250):
epoch_loss = 0
epoch_parameters_grads = np.zeros(len(param_names))
for j in range(5):
env["x1"] = train_data.x1[j]
env["x2"] = train_data.x2[j]
env["y"] = train_data.y[j]
this_loss = loss.eval()
epoch_loss += this_loss
losses.append(this_loss)
gradients = list(map(lambda x: loss.derivative(x).eval(), param_names))
# alternative 1 - stochastic gradient descent
# for p, grad in zip(param_names, gradients):
# env[p] = env[p] - 0.5 * grad
# end alternative 1
# alternative 2 - gradient descent
for i, grad in enumerate(gradients):
epoch_parameters_grads[i] = epoch_parameters_grads[i] + grad / 5
for i, p in enumerate(param_names):
env[p] = env[p] - 2.5 * epoch_parameters_grads[i]
# end alternative 2
epoch_losses.append(epoch_loss)
# #### Visualization
plt.figure()
plt.scatter(range(len(losses)), losses, s=2)
plt.ylabel("loss of training sample")
plt.xlabel("training sample")
plt.show()
plt.figure()
plt.plot(epoch_losses)
plt.ylabel("epoch loss")
plt.xlabel("epoch")
plt.show()
# #### Predictions
for j in range(5):
env["x1"] = train_data.x1[j]
env["x2"] = train_data.x2[j]
env["y"] = train_data.y[j]
pred = f_out.eval()
print(f'Sample {j}, label {env["y"]}, pred {pred}')
| topic_02/lab5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ricklon/jupyter-rpg/blob/main/cyberpunk2020_character_sheet_and_rolldice.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UHuQCinlyemW"
#
# # Cyberpunk 2020 Character
# <NAME>
#
# Rocker: Controllerist
#
# inspired by:
#
# http://vircadesproject.blogspot.com/2014/11/cyberpunk-2020-pregens-rockerboys.html
#
#
#
# + id="vLZNCrlMbV-T"
# Setup python
#imports
import re
import numpy as np
from pprint import pprint
import pandas as pd
import json
# + id="ra6rehYvw4eY"
career_skills = {
"Charismatic Leadership (COOL)": 6,
"Awareness/Notice (INT)": 5,
"Perform (EMP)": 2,
"Wardrobe & Style (ATTR)": 3,
"Composition (INT)": 6,
"Streetwise (COOL)": 7,
"Play Instrument (TECH)": 6,
"Seduction (EMP)": 5
}
# + id="v2j_ePYDxCMX"
pickup_skills= {
"Martial Arts (Animal Kung Fu) (REF)": 7,
"Intimidate (COOL)": 6,
"Oratory (COOL)": 1,
"Resist Torture/Drugs (COOL)": 2,
"Mathematics (INT)": 1,
"Submachine Gun (REF)": 2,
"Drive (REF)": 1
}
# + id="iN4RlqRdw1VK"
attributes = {
"INT": 7,
"REF": 7,
"COOL": 8,
"TECH": 7,
"LK": 4,
"ATT": {"cur":6, "max":6 },
"MA": 7,
"EMP": 6,
"BT": 8,
"BT_modifier": -3,
"SAVE": 8,
"BTM bt_modifier": -3,
"humanity" : 60,
"run": "movement_allowance * 3, 7 * 3 = 21",
"leap": "run/4 round down, floor(21/4) = 5",
"lift": "40 * BT, 40*8=320kg",
"reputation": 5
}
# + id="ZFfPSBD96MvE"
# + id="kG1njzqvxOMp"
equipment = ["Cellphone", "Mic", "Mixing Deck with Smartlink", "SUV", "$200","SP 10 Armour T-Shirt", "varied wardrobe"
]
# + id="b3z0_qTvxTE0"
cyberware = {"item1":["Basic Processor, Smartgun Link, Machine/Tech Link, Interface Plugs, Audio Recorder Voice Synthesiser, AudioVox (5d6+6 Humanity Cost)"],
"item2":["Cyberaudio with Enhanced Hearing Range, Amplified Hearing, Sound Editing, Digital Recording Link, Level Damper (2d6+4.5 Humanity Cost)"]}
# + id="yaN-dK66bcUO"
character_sheet = {"Handle" : "<NAME>",
"Role" : "Rocker",
"attributes": attributes,
"career_skills": career_skills,
"pickup_skills": pickup_skills,
"equipment" : equipment,
"cyberware": cyberware,
}
# + colab={"base_uri": "https://localhost:8080/"} id="Ut3V99okzjKr" outputId="e6bd0468-5fc4-4bbd-a4ac-3c5e75e636a2"
print(json.dumps(character_sheet, indent=2))
# + colab={"base_uri": "https://localhost:8080/"} id="OIzz4SxE6Q5U" outputId="c1b833b8-73ba-44f2-c97e-8a311a115f1f"
pprint(character_sheet)
# + id="kaPThtrz-W2O"
#Dice Roll Examples:
def d6(n):
return np.random.randint(1, 7, size=n)
def d10(n):
return np.random.randint(1, 11, size=n)
# + id="ZFbzxw_sb-FN"
#skill checks
def check_awareness():
return d10(1)[0] + character_sheet["attributes"]["INT"] + character_sheet["career_skills"]["Awareness/Notice (INT)"]
def check_strength():
return d10(1)[0] + character_sheet["attributes"]["BT"]
def check_cool():
return d10(1)[0] + character_sheet["attributes"]["COOL"]
# + colab={"base_uri": "https://localhost:8080/"} id="OGLTRDb1A6MF" outputId="524d776a-e772-41fb-a32a-aaa8dfbf522f"
print(f'd6(36): {d6(36)}')
print(f'd10(36): {d10(36)}')
# + id="IFme27GlEdVO"
from IPython.display import SVG, display
def show_svg():
display(SVG(url='https://upload.wikimedia.org/wikipedia/commons/e/e6/Cyberpunk_2077_logo.svg'))
# + colab={"base_uri": "https://localhost:8080/", "height": 192} id="M31Mw7A-Ef7Q" outputId="2003f7ed-4a35-4a75-f246-c722a04372d4"
show_svg()
# + colab={"base_uri": "https://localhost:8080/", "height": 221} id="3iqwAISZFXFd" outputId="f2788910-bc43-4fd8-a33e-942861eeabba"
display(SVG("""<svg version="1.1"
baseProfile="full"
width="300" height="200"
xmlns="http://www.w3.org/2000/svg">
<rect width="100%" height="100%" fill="red" />
<circle cx="150" cy="100" r="80" fill="green" />
<text x="150" y="125" font-size="60" text-anchor="middle" fill="white">Cyberpunk 2020</text>
</svg>"""))
# + id="P3ftDvWnGFzb"
# + colab={"base_uri": "https://localhost:8080/"} id="m0gdXLC0GFm_" outputId="58bc710e-b651-4118-9522-65e94cccf35d"
d10(1)
# + colab={"base_uri": "https://localhost:8080/"} id="o8mpy-CFPF0J" outputId="45dfe889-3fac-4a35-dc32-edf5646803ee"
d10(1)[0] +5 + 7
# + id="dbEjfMNKPY8o"
roll = 14
# + [markdown] id="a9UfglV6f1RK"
# # Game Play: Session 1
#
# + [markdown] id="IRvfkMMWKSQT"
# Characeters:
#
# IV: Jean
#
# Ted: Eric
#
# The Club: Nuclear Eruption
#
# Meet you at the Nuke
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="cDRjsw_eaE9n" outputId="03fbefd2-8a9f-4a92-9776-870a516b204c"
# awarness check +5 + 7
d10(1)[0] + character_sheet["attributes"]["INT"] + character_sheet["career_skills"]["Awareness/Notice (INT)"]
# + colab={"base_uri": "https://localhost:8080/"} id="I2bqAGBZcJFT" outputId="5ab95b1c-0d49-47ce-eeee-085acf6f4ae9"
check_awareness()
# + [markdown] id="z7neXF_CgWE5"
# Discoverd that I was in a box. I can here my friends nearby.
#
# + [markdown] id="TOGjBLxcgdDN"
# "I gotta get out of this box."
#
# + colab={"base_uri": "https://localhost:8080/"} id="OZFnrM3dh725" outputId="94c98036-21b9-43ed-da7a-a33046b888e0"
# streng check
d10(1)[0] + 8
# + colab={"base_uri": "https://localhost:8080/"} id="HLWAx1zxiAyR" outputId="268576c9-e89f-466d-818d-739640482978"
check_strength()
# + colab={"base_uri": "https://localhost:8080/"} id="Kk2mEIR4jcc_" outputId="d7d5cea9-26ca-45fb-9f7b-f8ad63780d17"
d10(1)[0] + 8
# + colab={"base_uri": "https://localhost:8080/"} id="q2GItNkukezt" outputId="59e8dc34-eb6f-4541-e3ae-7ed9f5bdee32"
# looking out over box
check_awareness()
| cyberpunk2020_character_sheet_and_rolldice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Solution Notebook
# ## Problem: Determine the height of a tree.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Is this a binary tree?
# * Yes
# * Can we assume we already have a Node class with an insert method?
# * Yes
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * 5 -> 1
# * 5, 2, 8, 1, 3 -> 3
# ## Algorithm
#
# We'll use a recursive algorithm.
#
# * If the current node is None, return 0
# * Else, return 1 + the maximum height of the left or right children
#
# Complexity:
# * Time: O(n)
# * Space: O(h), where h is the height of the tree
# ## Code
# %run ../bst/bst.py
class BstHeight(Bst):
def height(self, node):
if node is None:
return 0
return 1 + max(self.height(node.left),
self.height(node.right))
# ## Unit Test
# +
# %%writefile test_height.py
from nose.tools import assert_equal
class TestHeight(object):
def test_height(self):
bst = BstHeight(Node(5))
assert_equal(bst.height(bst.root), 1)
bst.insert(2)
bst.insert(8)
bst.insert(1)
bst.insert(3)
assert_equal(bst.height(bst.root), 3)
print('Success: test_height')
def main():
test = TestHeight()
test.test_height()
if __name__ == '__main__':
main()
# -
# %run -i test_height.py
| graphs_trees/tree_height/height_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from IPython.core.interactiveshell import InteractiveShell
from IPython.display import display, set_matplotlib_formats
InteractiveShell.ast_node_interactivity = "all"
# %load_ext autoreload
# %autoreload 2
# Run once
# #!pip install -Uq pystan==2.19.1.1 prophet
# !pip install -Uq statsmodels
import os
import re
import time
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import investpy
from datetime import date, datetime
from pathlib import Path
from dateutil.relativedelta import relativedelta
from dataclasses import dataclass, field
from prophet import Prophet
from sklearn.metrics import mean_absolute_error
# %matplotlib inline
# %config InlineBackend.figure_formats = ['retina']
pd.set_option('display.max_rows', 500)
# +
COUNTRY = "malaysia"
DATASET_FOLDER = "test_data"
TARGET_DATASET_FOLDER = f"{DATASET_FOLDER}/{COUNTRY}"
STOCKS_DATASET = f"{TARGET_DATASET_FOLDER}/stocks.csv"
STOCKS_INFO_DATASET = f"{TARGET_DATASET_FOLDER}/stocks_info.csv"
STOCKS_FINANCE_DATASET = f"{TARGET_DATASET_FOLDER}/stocks_finance.csv"
STOCKS_DIVIDENDS_DATASET = f"{TARGET_DATASET_FOLDER}/stocks_dividends.csv"
STOCKS_SELECTED = f"{TARGET_DATASET_FOLDER}/stocks_selected.csv"
STOCKS_HISTORICAL_PRICES = f"{TARGET_DATASET_FOLDER}/stocks_historcal_prices.csv"
FROM_DATE = '1/1/2015'
TO_DATE = datetime.strftime(date.today(), '%d/%m/%Y')
# -
def save_csv(df, file_name, index=True):
df.to_csv(file_name, header=True, index=index)
df_stocks_selected = pd.read_csv(STOCKS_SELECTED)
display(df_stocks_selected.head(10))
# +
def get_stock_historical_prices(symbol, country):
try:
return investpy.get_stock_historical_data(symbol, country, FROM_DATE, TO_DATE)
except:
return None
def download_historical_prices(df):
df_history = None
count = 0
for _, row in df.iterrows():
count = count + 1
print(f"{count}/{len(df)}: {row.Symbol}")
df_stock = get_stock_historical_prices(row.Symbol, COUNTRY)
if df_stock is None:
continue
df_stock['Symbol'] = row.Symbol
if df_history is None:
df_history = df_stock
else:
df_history = df_history.append(df_stock)
if count % 10 == 0:
save_csv(df_history, STOCKS_HISTORICAL_PRICES)
time.sleep(3)
save_csv(df_history, STOCKS_HISTORICAL_PRICES)
# + tags=[]
#download_historical_prices(df_stocks_selected)
# -
# ## Forecast Changes
# + tags=[]
df_history = pd.read_csv(STOCKS_HISTORICAL_PRICES, parse_dates=['Date'])
display(df_history.info())
# +
# Try with 1 stock
df_stock = df_history[df_history['Symbol'] == 'MBBM'].copy()
df_stock['Close'].plot(figsize = (15,7))
df_stock.set_index('Date', inplace=True)
df_stock.sort_values('Date', inplace=True)
df_stock.drop(columns=['Open', 'High', 'Volume', 'Currency', 'Symbol', 'Low'], inplace=True)
df_stock.head(10)
# -
# Aggregate weekly
df_week = df_stock.resample('w').mean()
df_week = df_week[['Close']]
df_week.head(10)
# Take the log to of the returns to normalize large fluctuations.
df_week['weekly_ret'] = np.log(df_week['Close']).diff()
df_week.head(10)
# drop null rows
df_week.dropna(inplace=True)
df_week.weekly_ret.plot(kind='line', figsize=(12, 6))
udiff = df_week.drop(['Close'], axis=1)
udiff.head()
# +
# Stationarity test
import statsmodels.api as sm
from statsmodels.tsa.stattools import adfuller
rolmean = udiff.rolling(20).mean()
rolstd = udiff.rolling(20).std()
plt.figure(figsize=(12, 6))
orig = plt.plot(udiff, color='blue', label='Original')
mean = plt.plot(rolmean, color='red', label='Rolling Mean')
std = plt.plot(rolstd, color='black', label = 'Rolling Std Deviation')
plt.title('Rolling Mean & Standard Deviation')
plt.legend(loc='best')
plt.show(block=False)
# +
# Perform Dickey-Fuller test
dftest = sm.tsa.adfuller(udiff.weekly_ret, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic', 'p-value', '#Lags Used', 'Number of Observations Used'])
for key, value in dftest[4].items():
dfoutput['Critical Value ({0})'.format(key)] = value
# p < 0.05 reject null hypothesis. Data is stationary
dfoutput
# -
# ACF and PACF
from statsmodels.graphics.tsaplots import plot_acf
# the autocorrelation chart provides just the correlation at increasing lags
fig, ax = plt.subplots(figsize=(12,5))
plot_acf(udiff.values, lags=10, ax=ax)
plt.show()
# +
from statsmodels.graphics.tsaplots import plot_pacf
fig, ax = plt.subplots(figsize=(12,5))
plot_pacf(udiff.values, lags=10, ax=ax)
plt.show()
# +
from statsmodels.tsa.arima.model import ARIMA
from statsmodels.tsa.arima_model import ARMA
# Notice that you have to use udiff - the differenced data rather than the original data.
ar1 = ARIMA(tuple(udiff.values), order = (3,1,1)).fit()
ar1.summary()
# +
# https://colab.research.google.com/drive/1JmY9kbt_Irq3jv55aD4yyR_jFRY0BtKf#scrollTo=Tr_sUw_sHLRg
# https://towardsdatascience.com/time-series-forecasting-predicting-stock-prices-using-an-arima-model-2e3b3080bd70
# https://medium.com/@derejeabera/stock-price-prediction-using-arima-model-251ddb4ee52a
# 25873099.pdf research gate
# https://www.freecodecamp.org/news/algorithmic-trading-in-python/
# https://classroom.udacity.com/courses/ud501/lessons/4242038556/concepts/41998985520923
# https://www.coursera.org/learn/introduction-trading-machine-learning-gcp/lecture/Zng9E/ar-auto-regressive
# -
plt.figure(figsize=(12, 8))
plt.plot(udiff.values, color='blue')
preds = ar1.fittedvalues
plt.plot(preds, color='red')
plt.show()
# +
steps = 2
forecast = ar1.forecast(steps=steps)[0]
plt.figure(figsize=(12, 8))
plt.plot(udiff.values, color='blue')
preds = ar1.fittedvalues
plt.plot(preds, color='red')
plt.plot(pd.DataFrame(np.array([preds[-1],forecast[0]]).T,index=range(len(udiff.values)+1, len(udiff.values)+3)), color='green')
plt.plot(pd.DataFrame(forecast,index=range(len(udiff.values)+1, len(udiff.values)+1+steps)), color='green')
plt.title('Display the predictions with the ARIMA model')
plt.show()
# -
| nbs/02_02_stocks_time_series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
data = [json.loads(line) for line in open('./face_recognition_annotations.json', 'r')]
print(data[0]['_via_img_metadata']['img_001.jpeg43329']['regions'])
print(data[0]['_via_img_metadata']['img_001.jpeg43329']['regions'][0])
widths_list = []
heights_list = []
for key, value in data[0]['_via_img_metadata'].items():
local_w = [k['shape_attributes']['width'] for k in value['regions']]
widths_list.extend(local_w)
local_h = [k['shape_attributes']['height'] for k in value['regions']]
heights_list.extend(local_h)
print(widths_list)
print(heights_list)
print(zip(widths_list,heights_list))
X = [[key,value] for key, value in zip(widths_list,heights_list)]
print(X)
from sklearn.cluster import KMeans
import numpy as np
X = np.array(X)
kmeans = KMeans(n_clusters=4, random_state=0).fit(X)
kmeans.labels_
kmeans.cluster_centers_
# ### The 4 bounding box dimensions ([width, height]) are shown below:
#
# - [ 54.568 , 83.512 ]
# - [106.796875 , 174.09375 ]
# - [ 74.10869565, 124.11594203]
# - [124.83333333, 263.83333333]
# *Disclaimer: The contents of this notebook are used for educational purposes i.e. for learning and research.*
| Phase1/Project19/k-means - bounding boxes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.7 64-bit (''3.8.7'': pyenv)'
# language: python
# name: python3
# ---
# # Interaction Grounded Learning
import torch
from torch import nn, optim
import matplotlib.pyplot as plt
from torchvision.datasets import MNIST
# ## MNIST Digit-ID Environment
class MNISTEnv(object):
def __init__(self):
self.mnist = MNIST('./data', False, download=True)
self.mnist.data = self.mnist.data / 255.0
self.x_size = 784
self.y_size = 784
self.a_size = 10
# Cache the indecies of all 0 and 1 digit images
self.zeros = (self.mnist.targets == 0).nonzero().view(-1)
self.ones = (self.mnist.targets == 1).nonzero().view(-1)
self.x = None
self.done = False
def observe(self):
# Sample a random digit to provide as the context
self.id = torch.randint(0, self.mnist.data.shape[0], [1])
self.x = self.mnist.data[self.id]
self.done = False
return self.x.view(1, -1)
def step(self, action):
if not self.done:
self.done = True
# Sample a random 0 or 1 image to provie as feedback
if (action == self.mnist.targets[self.id]).all():
one_idx = self.ones[torch.randint(0, len(self.ones), [1])]
return self.mnist.data[one_idx].view(1, -1), torch.tensor([1.])
else:
zero_idx = self.zeros[torch.randint(0, len(self.zeros), [1])]
return self.mnist.data[zero_idx].view(1, -1), torch.tensor([0.])
else:
raise Warning("Trial already complete.")
# ## Linear Policy & Decoder Function
class Agent(nn.Module):
def __init__(self, x_size, a_size, y_size):
super(Agent, self).__init__()
self.x_size = x_size
self.a_size = a_size
self.y_size = y_size
self.policy = nn.Sequential(nn.Linear(x_size, a_size), nn.Softmax(-1))
self.decoder = nn.Sequential(nn.Linear(y_size, 1), nn.Sigmoid())
def evaluate(self, x, random=False):
pi = self.policy(x)
pi_dist = torch.distributions.categorical.Categorical(pi)
if random:
a = torch.randint(0, self.a_size, [1])
else:
a = pi_dist.sample()
# Return sampled action and policy probability for action
return a, pi[:,a]
def decode(self, y):
r = self.decoder(y)
return r
# ## Agent Training Process
# +
# Hyperparameters
epochs = 1000
num_trials = 32
learning_rate = 5e-4
# Create environment
env = MNISTEnv()
# -
def collect_batch(random, num_trials, env, agent):
experiences = []
for _ in range(num_trials):
x = env.observe()
a, p = agent.evaluate(x, random)
y, tr = env.step(a)
r = agent.decode(y)
experiences.append([x, a, p, y, r, tr])
x_s, a_s, p_s, y_s, lr_s, tr_s = zip(*experiences)
x_s = torch.stack(x_s).squeeze(1)
a_s = torch.stack(a_s).view(-1)
p_s = torch.stack(p_s).view(-1)
y_s = torch.stack(y_s).squeeze(1)
lr_s = torch.stack(lr_s).view(-1)
tr_s = torch.stack(tr_s).view(-1)
return p_s, lr_s, tr_s
# ## Train IGL Agent
# +
# Initialize agent and optimizer
agent = Agent(env.x_size, env.a_size, env.y_size)
optimizer = optim.Adam(agent.parameters(), lr=learning_rate)
rewards_exploit = []
rewards_explore = []
for i in range(epochs):
optimizer.zero_grad()
# Collect batches of explore and exploit trials
exploit_p_s, exploit_lr_s, exploit_trs = collect_batch(False, num_trials, env, agent)
_, explore_lr_s, explore_trs = collect_batch(True, num_trials, env, agent)
## Calculate loss function ##
# Policy-gradient with advantage is used to make exploit policy "good"
exploit_adv = (exploit_lr_s - torch.mean(exploit_lr_s)) / (torch.std(exploit_lr_s) + 1e-10)
exploit_loss = torch.mean(-torch.log(exploit_p_s) * exploit_adv)
# Learned rewards of explore policy are minimized to make it "bad"
explore_loss = torch.mean(explore_lr_s)
# Losses are combined
loss = exploit_loss + explore_loss
# Optimize model
loss.backward()
optimizer.step()
# Track rewards
rewards_exploit.append([torch.mean(exploit_lr_s.detach()), torch.mean(exploit_trs.detach())])
rewards_explore.append([torch.mean(explore_lr_s.detach()), torch.mean(explore_trs.detach())])
# -
# ## Train Baseline CB Agent
# +
# Initialize environment, agent, and optimizer
env = MNISTEnv()
agent = Agent(env.x_size, env.a_size, env.y_size)
optimizer = optim.Adam(agent.parameters(), lr=learning_rate)
rewards_cb = []
for i in range(epochs):
optimizer.zero_grad()
# Collect batches of explore and exploit trials
exploit_p_s, _, exploit_trs = collect_batch(False, num_trials, env, agent)
# Policy-gradient with advantage using true reward function
exploit_adv = (exploit_trs - torch.mean(exploit_trs)) / (torch.std(exploit_trs) + 1e-10)
loss = torch.mean(-torch.log(exploit_p_s) * exploit_adv)
# Optimize model
loss.backward()
optimizer.step()
# Track rewards
rewards_cb.append(torch.mean(exploit_trs.detach()))
# -
# ## Plot Results
# +
smoothing_var = 5
exploit_pred, exploit_true = zip(*rewards_exploit)
explore_pred, explore_true = zip(*rewards_explore)
exploit_pred = torch.stack(exploit_pred).view(-1, 5).mean(-1)
explore_pred = torch.stack(explore_pred).view(-1, 5).mean(-1)
exploit_true = torch.stack(exploit_true).view(-1, 5).mean(-1)
explore_true = torch.stack(explore_true).view(-1, 5).mean(-1)
rewards_cb = torch.stack(rewards_cb).view(-1, smoothing_var).mean(-1)
# -
fig, axs = plt.subplots(1,2, figsize=(15, 5), dpi=200)
axs[0].plot(exploit_pred, label="IGL-Exploit")
axs[0].plot(explore_pred, label="IGL-Explore")
axs[0].set_title("Learned Reward")
axs[0].legend()
axs[1].plot(exploit_true, label="IGL-Exploit")
axs[1].plot(explore_true, label="IGL-Explore")
axs[1].plot(rewards_cb, label="CB-Exploit")
axs[1].set_title("True Reward")
axs[1].legend()
| learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Python 2]
# language: python
# name: conda-env-Python 2-py
# ---
import pandas as pd
# +
from numpy.linalg import inv
import numpy as np
from scipy.linalg import eig
from sklearn.datasets import make_blobs
from sklearn.metrics import pairwise_distances
from diffmaps_util import *
# -
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib nbagg
df = pd.read_csv('annulus.csv')
df = df[['X', 'Y']]
df.head()
X = df.as_matrix()
plt.scatter(X[:,0], X[:,1])
plt.show()
# +
L = k(X, .1)
print 'L shape %s %s' % L.shape
D = diag(L)
print 'D shape %s %s' % D.shape
M = inv(D).dot(L)
print 'M shape %s %s' % M.shape
w, v = eig(M)
w = np.abs(w.real)
v = v.real
print 'eigenvalue shape %s' % w.shape
print 'eigenvector shape %s %s' % v.shape
# -
w, v = sort_eigens(w, v)
print 'eigenvalue shape %s' % w.shape
print 'eigenvector shape %s %s' % v.shape
psi = v / v[:,0]
psi.shape
diffmap = (w.reshape(-1,1) * psi.T).T[:,1:]
diffmap.shape
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
diffmap = (w.reshape(-1,1) * psi.T).T[:,1:]
diffmap.shape
ax.scatter(diffmap[:,0], diffmap[:,1], diffmap[:,2])
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_zlabel('z')
ax.set_title('T = 1')
| subjects/diffusion maps/demos/Diffusion Maps - Annulus Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py36spider
# language: python
# name: py36spider
# ---
# ### 载入环境
from damndata.damn_geoSpider import geoautonavi as amaPoi
import pandas as pd
# ### 参数设置
# ###### 说明**
# 详细文档: https://lbs.amap.com/api/webservice/guide/api/search
#
# http接口: 'resapi.amap.com/v3/place/'
#
# 检索模式:(地理参数 'geo')
#
# 关键字搜索: 'text?' >>> 'city=' (citycode|adcode)
#
# 周边搜索: 'around?' >>> 'location='、'radius='
#
# 多边形搜索: 'polygon?' >>> 'polygon='
#
# 参数: key、'geo'、types|keywords
http = 'restapi.amap.com/v3/place/around?'
key = 'key=f9257dab1e8214b074587fe16484cb1e'
para_loc = '&location=120.233851,30.167682&radius=800'
para_type = '&types=050000|070000|090000|120300|141200|160000|170100|170200'
# ### 爬取
# +
URL=http+key+para_loc+para_type+'&output=json&offset=25&extensions=all&page='
pois=amaPoi.getpois(URL) #请求数据
filePath = 'E:/zixunHUANG/2019-2021_Project/202007_FridaySalon/week200724/test.csv'#设定文件路径
amaPoi.write_to_csv(pois,filePath) #写入本地
print('Done!!!')
# -
# ### 读入爬取结果
df=pd.read_csv(filePath,index_col='Unnamed: 0')
df
# ### 定制自己的POI请求程序
type_list=['050000','060000','070000']
for i,typei in enumerate(type_list):
print(i,typei)
URL_i=http+key+para_loc+typei+'&output=json&offset=25&extensions=all&page='
pois_i=amaPoi.getpois(URL_i)
filePath_i = 'E:/zixunHUANG/2019-2021_Project/202007_FridaySalon/week200724/'+str(i).zfill(2)+'.csv'
amaPoi.write_to_csv(pois_i,filePath_i)
print('Done!!!')
| examples/.ipynb_checkpoints/scrapCode-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math as m
def add(a,b):
return a + b
def diff(a,b):
return a - b
def prod(a,b):
return a * b
def div(a,b):
try:
return a / b
except ZeroDivisionError as e:
print(e)
return div(float(input("Enter Dividend: ")), float(input("Enter Divisor: ")))
def sq_root(a,b):
try:
return m.sqrt(a),m.sqrt(b)
except ValueError as e:
print(e)
return sq_root(float(input("Enter a:")),float(input("Enter b:")))
def mod(a,b):
try:
return a % b
except ZeroDivisionError as e:
print(e)
return mod(float(input("Enter Dividend: ")), float(input("Enter Divisor: ")))
def factorial(n):
try:
if n < 0:
raise ValueError("Cannot take a factorial of negative number")
else:
if n == 0:
return 1
else:
return n*factorial(n-1)
except ValueError as e:
print(e)
return factorial(int(input("Enter a valid number: ")))
def enter_values():
try:
a = float(input("Enter a: "))
b = float(input("Enter b: "))
return a, b
except ValueError as e:
print(e)
print("Enter valid numbers: ")
return enter_values()
a, b = enter_values()
print("Sum:",add(a,b))
print("Difference:",diff(a,b))
print("Product:",prod(a,b))
print("Division:",div(a,b))
print("Modulus:",mod(a,b))
print("Square Root:",sq_root(a,b))
print("Factorial:",factorial(a))
# -
# def factorial(n):
# if n == 0:
# return 1
# else:
# return n*factorial(n-1)
#
# print(factorial(3))
| Topic Wise Problems/Functions/Functions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logic: `logic.py`; Chapters 6-8
# This notebook describes the [logic.py](https://github.com/aimacode/aima-python/blob/master/logic.py) module, which covers Chapters 6 (Logical Agents), 7 (First-Order Logic) and 8 (Inference in First-Order Logic) of *[Artificial Intelligence: A Modern Approach](http://aima.cs.berkeley.edu)*. See the [intro notebook](https://github.com/aimacode/aima-python/blob/master/intro.ipynb) for instructions.
#
# We'll start by looking at `Expr`, the data type for logical sentences, and the convenience function `expr`. We'll be covering two types of knowledge bases, `PropKB` - Propositional logic knowledge base and `FolKB` - First order logic knowledge base. We will construct a propositional knowledge base of a specific situation in the Wumpus World. We will next go through the `tt_entails` function and experiment with it a bit. The `pl_resolution` and `pl_fc_entails` functions will come next. We'll study forward chaining and backward chaining algorithms for `FolKB` and use them on `crime_kb` knowledge base.
#
# But the first step is to load the code:
from utils import *
from logic import *
# ## Logical Sentences
# The `Expr` class is designed to represent any kind of mathematical expression. The simplest type of `Expr` is a symbol, which can be defined with the function `Symbol`:
Symbol('x')
# Or we can define multiple symbols at the same time with the function `symbols`:
(x, y, P, Q, f) = symbols('x, y, P, Q, f')
# We can combine `Expr`s with the regular Python infix and prefix operators. Here's how we would form the logical sentence "P and not Q":
P & ~Q
# This works because the `Expr` class overloads the `&` operator with this definition:
#
# ```python
# def __and__(self, other): return Expr('&', self, other)```
#
# and does similar overloads for the other operators. An `Expr` has two fields: `op` for the operator, which is always a string, and `args` for the arguments, which is a tuple of 0 or more expressions. By "expression," I mean either an instance of `Expr`, or a number. Let's take a look at the fields for some `Expr` examples:
# +
sentence = P & ~Q
sentence.op
# -
sentence.args
P.op
P.args
# +
Pxy = P(x, y)
Pxy.op
# -
Pxy.args
# It is important to note that the `Expr` class does not define the *logic* of Propositional Logic sentences; it just gives you a way to *represent* expressions. Think of an `Expr` as an [abstract syntax tree](https://en.wikipedia.org/wiki/Abstract_syntax_tree). Each of the `args` in an `Expr` can be either a symbol, a number, or a nested `Expr`. We can nest these trees to any depth. Here is a deply nested `Expr`:
3 * f(x, y) + P(y) / 2 + 1
# ## Operators for Constructing Logical Sentences
#
# Here is a table of the operators that can be used to form sentences. Note that we have a problem: we want to use Python operators to make sentences, so that our programs (and our interactive sessions like the one here) will show simple code. But Python does not allow implication arrows as operators, so for now we have to use a more verbose notation that Python does allow: `|'==>'|` instead of just `==>`. Alternately, you can always use the more verbose `Expr` constructor forms:
#
# | Operation | Book | Python Infix Input | Python Output | Python `Expr` Input
# |--------------------------|----------------------|-------------------------|---|---|
# | Negation | ¬ P | `~P` | `~P` | `Expr('~', P)`
# | And | P ∧ Q | `P & Q` | `P & Q` | `Expr('&', P, Q)`
# | Or | P ∨ Q | `P`<tt> | </tt>`Q`| `P`<tt> | </tt>`Q` | `Expr('`|`', P, Q)`
# | Inequality (Xor) | P ≠ Q | `P ^ Q` | `P ^ Q` | `Expr('^', P, Q)`
# | Implication | P → Q | `P` <tt>|</tt>`'==>'`<tt>|</tt> `Q` | `P ==> Q` | `Expr('==>', P, Q)`
# | Reverse Implication | Q ← P | `Q` <tt>|</tt>`'<=='`<tt>|</tt> `P` |`Q <== P` | `Expr('<==', Q, P)`
# | Equivalence | P ↔ Q | `P` <tt>|</tt>`'<=>'`<tt>|</tt> `Q` |`P <=> Q` | `Expr('<=>', P, Q)`
#
# Here's an example of defining a sentence with an implication arrow:
~(P & Q) |'==>'| (~P | ~Q)
# ## `expr`: a Shortcut for Constructing Sentences
#
# If the `|'==>'|` notation looks ugly to you, you can use the function `expr` instead:
expr('~(P & Q) ==> (~P | ~Q)')
# `expr` takes a string as input, and parses it into an `Expr`. The string can contain arrow operators: `==>`, `<==`, or `<=>`, which are handled as if they were regular Python infix operators. And `expr` automatically defines any symbols, so you don't need to pre-define them:
expr('sqrt(b ** 2 - 4 * a * c)')
# For now that's all you need to know about `expr`. If you are interested, we explain the messy details of how `expr` is implemented and how `|'==>'|` is handled in the appendix.
# ## Propositional Knowledge Bases: `PropKB`
#
# The class `PropKB` can be used to represent a knowledge base of propositional logic sentences.
#
# We see that the class `KB` has four methods, apart from `__init__`. A point to note here: the `ask` method simply calls the `ask_generator` method. Thus, this one has already been implemented and what you'll have to actually implement when you create your own knowledge base class (if you want to, though I doubt you'll ever need to; just use the ones we've created for you), will be the `ask_generator` function and not the `ask` function itself.
#
# The class `PropKB` now.
# * `__init__(self, sentence=None)` : The constructor `__init__` creates a single field `clauses` which will be a list of all the sentences of the knowledge base. Note that each one of these sentences will be a 'clause' i.e. a sentence which is made up of only literals and `or`s.
# * `tell(self, sentence)` : When you want to add a sentence to the KB, you use the `tell` method. This method takes a sentence, converts it to its CNF, extracts all the clauses, and adds all these clauses to the `clauses` field. So, you need not worry about `tell`ing only clauses to the knowledge base. You can `tell` the knowledge base a sentence in any form that you wish; converting it to CNF and adding the resulting clauses will be handled by the `tell` method.
# * `ask_generator(self, query)` : The `ask_generator` function is used by the `ask` function. It calls the `tt_entails` function, which in turn returns `True` if the knowledge base entails query and `False` otherwise. The `ask_generator` itself returns an empty dict `{}` if the knowledge base entails query and `None` otherwise. This might seem a little bit weird to you. After all, it makes more sense just to return a `True` or a `False` instead of the `{}` or `None` But this is done to maintain consistency with the way things are in First-Order Logic, where, an `ask_generator` function, is supposed to return all the substitutions that make the query true. Hence the dict, to return all these substitutions. I will be mostly be using the `ask` function which returns a `{}` or a `False`, but if you don't like this, you can always use the `ask_if_true` function which returns a `True` or a `False`.
# * `retract(self, sentence)` : This function removes all the clauses of the sentence given, from the knowledge base. Like the `tell` function, you don't have to pass clauses to remove them from the knowledge base; any sentence will do fine. The function will take care of converting that sentence to clauses and then remove those.
# ## Wumpus World KB
# Let us create a `PropKB` for the wumpus world with the sentences mentioned in `section 7.4.3`.
wumpus_kb = PropKB()
# We define the symbols we use in our clauses.<br/>
# $P_{x, y}$ is true if there is a pit in `[x, y]`.<br/>
# $B_{x, y}$ is true if the agent senses breeze in `[x, y]`.<br/>
P11, P12, P21, P22, P31, B11, B21 = expr('P11, P12, P21, P22, P31, B11, B21')
# Now we tell sentences based on `section 7.4.3`.<br/>
# There is no pit in `[1,1]`.
wumpus_kb.tell(~P11)
# A square is breezy if and only if there is a pit in a neighboring square. This has to be stated for each square but for now, we include just the relevant squares.
wumpus_kb.tell(B11 | '<=>' | ((P12 | P21)))
wumpus_kb.tell(B21 | '<=>' | ((P11 | P22 | P31)))
# Now we include the breeze percepts for the first two squares leading up to the situation in `Figure 7.3(b)`
wumpus_kb.tell(~B11)
wumpus_kb.tell(B21)
# We can check the clauses stored in a `KB` by accessing its `clauses` variable
wumpus_kb.clauses
# We see that the equivalence $B_{1, 1} \iff (P_{1, 2} \lor P_{2, 1})$ was automatically converted to two implications which were inturn converted to CNF which is stored in the `KB`.<br/>
# $B_{1, 1} \iff (P_{1, 2} \lor P_{2, 1})$ was split into $B_{1, 1} \implies (P_{1, 2} \lor P_{2, 1})$ and $B_{1, 1} \Longleftarrow (P_{1, 2} \lor P_{2, 1})$.<br/>
# $B_{1, 1} \implies (P_{1, 2} \lor P_{2, 1})$ was converted to $P_{1, 2} \lor P_{2, 1} \lor \neg B_{1, 1}$.<br/>
# $B_{1, 1} \Longleftarrow (P_{1, 2} \lor P_{2, 1})$ was converted to $\neg (P_{1, 2} \lor P_{2, 1}) \lor B_{1, 1}$ which becomes $(\neg P_{1, 2} \lor B_{1, 1}) \land (\neg P_{2, 1} \lor B_{1, 1})$ after applying De Morgan's laws and distributing the disjunction.<br/>
# $B_{2, 1} \iff (P_{1, 1} \lor P_{2, 2} \lor P_{3, 2})$ is converted in similar manner.
# ## Inference in Propositional Knowlwdge Base
# In this section we will look at two algorithms to check if a sentence is entailed by the `KB`. Our goal is to decide whether $\text{KB} \vDash \alpha$ for some sentence $\alpha$.
# ### Truth Table Enumeration
# It is a model-checking approach which, as the name suggests, enumerates all possible models in which the `KB` is true and checks if $\alpha$ is also true in these models. We list the $n$ symbols in the `KB` and enumerate the $2^{n}$ models in a depth-first manner and check the truth of `KB` and $\alpha$.
# %psource tt_check_all
# Note that `tt_entails()` takes an `Expr` which is a conjunction of clauses as the input instead of the `KB` itself. You can use the `ask_if_true()` method of `PropKB` which does all the required conversions. Let's check what `wumpus_kb` tells us about $P_{1, 1}$.
wumpus_kb.ask_if_true(~P11), wumpus_kb.ask_if_true(P11)
# Looking at Figure 7.9 we see that in all models in which the knowledge base is `True`, $P_{1, 1}$ is `False`. It makes sense that `ask_if_true()` returns `True` for $\alpha = \neg P_{1, 1}$ and `False` for $\alpha = P_{1, 1}$. This begs the question, what if $\alpha$ is `True` in only a portion of all models. Do we return `True` or `False`? This doesn't rule out the possibility of $\alpha$ being `True` but it is not entailed by the `KB` so we return `False` in such cases. We can see this is the case for $P_{2, 2}$ and $P_{3, 1}$.
wumpus_kb.ask_if_true(~P22), wumpus_kb.ask_if_true(P22)
# ### Proof by Resolution
# Recall that our goal is to check whether $\text{KB} \vDash \alpha$ i.e. is $\text{KB} \implies \alpha$ true in every model. Suppose we wanted to check if $P \implies Q$ is valid. We check the satisfiability of $\neg (P \implies Q)$, which can be rewritten as $P \land \neg Q$. If $P \land \neg Q$ is unsatisfiable, then $P \implies Q$ must be true in all models. This gives us the result "$\text{KB} \vDash \alpha$ <em>if and only if</em> $\text{KB} \land \neg \alpha$ is unsatisfiable".<br/>
# This technique corresponds to <em>proof by <strong>contradiction</strong></em>, a standard mathematical proof technique. We assume $\alpha$ to be false and show that this leads to a contradiction with known axioms in $\text{KB}$. We obtain a contradiction by making valid inferences using inference rules. In this proof we use a single inference rule, <strong>resolution</strong> which states $(l_1 \lor \dots \lor l_k) \land (m_1 \lor \dots \lor m_n) \land (l_i \iff \neg m_j) \implies l_1 \lor \dots \lor l_{i - 1} \lor l_{i + 1} \lor \dots \lor l_k \lor m_1 \lor \dots \lor m_{j - 1} \lor m_{j + 1} \lor \dots \lor m_n$. Applying the resolution yeilds us a clause which we add to the KB. We keep doing this until:
# <ul>
# <li>There are no new clauses that can be added, in which case $\text{KB} \nvDash \alpha$.</li>
# <li>Two clauses resolve to yield the <em>empty clause</em>, in which case $\text{KB} \vDash \alpha$.</li>
# </ul>
# The <em>empty clause</em> is equivalent to <em>False</em> because it arises only from resolving two complementary
# unit clauses such as $P$ and $\neg P$ which is a contradiction as both $P$ and $\neg P$ can't be <em>True</em> at the same time.
# %psource pl_resolution
pl_resolution(wumpus_kb, ~P11), pl_resolution(wumpus_kb, P11)
pl_resolution(wumpus_kb, ~P22), pl_resolution(wumpus_kb, P22)
# ## First-Order Logic Knowledge Bases: `FolKB`
#
# The class `FolKB` can be used to represent a knowledge base of First-order logic sentences. You would initialize and use it the same way as you would for `PropKB` except that the clauses are first-order definite clauses. We will see how to write such clauses to create a database and query them in the following sections.
# ## Criminal KB
# In this section we create a `FolKB` based on the following paragraph.<br/>
# <em>The law says that it is a crime for an American to sell weapons to hostile nations. The country Nono, an enemy of America, has some missiles, and all of its missiles were sold to it by Colonel West, who is American.</em><br/>
# The first step is to extract the facts and convert them into first-order definite clauses. Extracting the facts from data alone is a challenging task. Fortnately we have a small paragraph and can do extraction and conversion manually. We'll store the clauses in list aptly named `clauses`.
clauses = []
# <em>“... it is a crime for an American to sell weapons to hostile nations”</em><br/>
# The keywords to look for here are 'crime', 'American', 'sell', 'weapon' and 'hostile'. We use predicate symbols to make meaning of them.
# <ul>
# <li>`Criminal(x)`: `x` is a criminal</li>
# <li>`American(x)`: `x` is an American</li>
# <li>`Sells(x ,y, z)`: `x` sells `y` to `z`</li>
# <li>`Weapon(x)`: `x` is a weapon</li>
# <li>`Hostile(x)`: `x` is a hostile nation</li>
# </ul>
# Let us now combine them with appropriate variable naming depict the meaning of the sentence. The criminal `x` is also the American `x` who sells weapon `y` to `z`, which is a hostile nation.
#
# $\text{American}(x) \land \text{Weapon}(y) \land \text{Sells}(x, y, z) \land \text{Hostile}(z) \implies \text{Criminal} (x)$
clauses.append(expr("(American(x) & Weapon(y) & Sells(x, y, z) & Hostile(z)) ==> Criminal(x)"))
# <em>"The country Nono, an enemy of America"</em><br/>
# We now know that Nono is an enemy of America. We represent these nations using the constant symbols `Nono` and `America`. the enemy relation is show using the predicate symbol `Enemy`.
#
# $\text{Enemy}(\text{Nono}, \text{America})$
clauses.append(expr("Enemy(Nono, America)"))
# <em>"Nono ... has some missiles"</em><br/>
# This states the existance of some missile which is owned by Nono. $\exists x \text{Owns}(\text{Nono}, x) \land \text{Missile}(x)$. We invoke existential instantiation to introduce a new constant `M1` which is the missile owned by Nono.
#
# $\text{Owns}(\text{Nono}, \text{M1}), \text{Missile}(\text{M1})$
clauses.append(expr("Owns(Nono, M1)"))
clauses.append(expr("Missile(M1)"))
# <em>"All of its missiles were sold to it by Colonel West"</em><br/>
# If Nono owns something and it classifies as a missile, then it was sold to Nono by West.
#
# $\text{Missile}(x) \land \text{Owns}(\text{Nono}, x) \implies \text{Sells}(\text{West}, x, \text{Nono})$
clauses.append(expr("(Missile(x) & Owns(Nono, x)) ==> Sells(West, x, Nono)"))
# <em>"West, who is American"</em><br/>
# West is an American.
#
# $\text{American}(\text{West})$
clauses.append(expr("American(West)"))
# We also know, from our understanding of language, that missiles are weapons and that an enemy of America counts as “hostile”.
#
# $\text{Missile}(x) \implies \text{Weapon}(x), \text{Enemy}(x, \text{America}) \implies \text{Hostile}(x)$
clauses.append(expr("Missile(x) ==> Weapon(x)"))
clauses.append(expr("Enemy(x, America) ==> Hostile(x)"))
# Now that we have converted the information into first-order definite clauses we can create our first-order logic knowledge base.
crime_kb = FolKB(clauses)
# ## Inference in First-Order Logic
# In this section we look at a forward chaining and a backward chaining algorithm for `FolKB`. Both the aforementioned algorithms rely on a process called <strong>unification</strong>, a key component of all first-order inference algorithms.
# ### Unification
# We sometimes require finding substitutions that make different logical expressions look identical. This process, called unification, is done by the `unify` algorithm. It takes as input two sentences and returns a <em>unifier</em> for them if one exists. A unifier is a dictionary which stores the substitutions required to make the two sentences identical. It does so by recursively unifying the components of a sentence, where the unification of a variable symbol `var` with a constant symbol `Const` is the mapping `{var: Const}`. Let's look at a few examples.
unify(expr('x'), 3)
unify(expr('A(x)'), expr('A(B)'))
unify(expr('Cat(x) & Dog(Dobby)'), expr('Cat(Bella) & Dog(y)'))
# In cases where there is no possible substitution that unifies the two sentences the function return `None`.
print(unify(expr('Cat(x)'), expr('Dog(Dobby)')))
# We also need to take care we do not unintentionally use same variable name. Unify treats them as a single variable which prevents it from taking multiple value.
print(unify(expr('Cat(x) & Dog(Dobby)'), expr('Cat(Bella) & Dog(x)')))
# ### Forward Chaining Algorithm
# We consider the simple forward-chaining algorithm presented in <em>Figure 9.3</em>. We look at each rule in the knoweldge base and see if the premises can be satisfied. This is done by finding a substitution which unifies the each of the premise with a clause in the `KB`. If we are able to unify the premises the conclusion (with the corresponding substitution) is added to the `KB`. This inferencing process is repeated until either the query can be answered or till no new sentences can be aded. We test if the newly added clause unifies with the query in which case the substitution yielded by `unify` is an answer to the query. If we run out of sentences to infer, this means the query was a failure.
#
# The function `fol_fc_ask` is a generator which yields all substitutions which validate the query.
# %psource fol_fc_ask
# Let's find out all the hostile nations. Note that we only told the `KB` that Nono was an enemy of America, not that it was hostile.
answer = fol_fc_ask(crime_kb, expr('Hostile(x)'))
print(list(answer))
# The generator returned a single substitution which says that Nono is a hostile nation. See how after adding another enemy nation the generator returns two substitutions.
crime_kb.tell(expr('Enemy(JaJa, America)'))
answer = fol_fc_ask(crime_kb, expr('Hostile(x)'))
print(list(answer))
# <strong><em>Note</em>:</strong> `fol_fc_ask` makes changes to the `KB` by adding sentences to it.
# ### Backward Chaining Algorithm
# This algorithm works backward from the goal, chaining through rules to find known facts that support the proof. Suppose `goal` is the query we want to find the substitution for. We find rules of the form $\text{lhs} \implies \text{goal}$ in the `KB` and try to prove `lhs`. There may be multiple clauses in the `KB` which give multiple `lhs`. It is sufficient to prove only one of these. But to prove a `lhs` all the conjuncts in the `lhs` of the clause must be proved. This makes it similar to <em>And/Or</em> search.
# #### OR
# The <em>OR</em> part of the algorithm comes from our choice to select any clause of the form $\text{lhs} \implies \text{goal}$. Looking at all rules's `lhs` whose `rhs` unify with the `goal`, we yield a substitution which proves all the conjuncts in the `lhs`. We use `parse_definite_clause` to attain `lhs` and `rhs` from a clause of the form $\text{lhs} \implies \text{rhs}$. For atomic facts the `lhs` is an empty list.
# %psource fol_bc_or
# #### AND
# The <em>AND</em> corresponds to proving all the conjuncts in the `lhs`. We need to find a substitution which proves each <em>and</em> every clause in the list of conjuncts.
# %psource fol_bc_and
# Now the main function `fl_bc_ask` calls `fol_bc_or` with substitution initialized as empty. The `ask` method of `FolKB` uses `fol_bc_ask` and fetches the first substitution returned by the generator to answer query. Let's query the knowledge base we created from `clauses` to find hostile nations.
# Rebuild KB because running fol_fc_ask would add new facts to the KB
crime_kb = FolKB(clauses)
crime_kb.ask(expr('Hostile(x)'))
# You may notice some new variables in the substitution. They are introduced to standardize the variable names to prevent naming problems as discussed in the [Unification section](#Unification)
# ## Appendix: The Implementation of `|'==>'|`
#
# Consider the `Expr` formed by this syntax:
P |'==>'| ~Q
# What is the funny `|'==>'|` syntax? The trick is that "`|`" is just the regular Python or-operator, and so is exactly equivalent to this:
(P | '==>') | ~Q
# In other words, there are two applications of or-operators. Here's the first one:
P | '==>'
# What is going on here is that the `__or__` method of `Expr` serves a dual purpose. If the right-hand-side is another `Expr` (or a number), then the result is an `Expr`, as in `(P | Q)`. But if the right-hand-side is a string, then the string is taken to be an operator, and we create a node in the abstract syntax tree corresponding to a partially-filled `Expr`, one where we know the left-hand-side is `P` and the operator is `==>`, but we don't yet know the right-hand-side.
#
# The `PartialExpr` class has an `__or__` method that says to create an `Expr` node with the right-hand-side filled in. Here we can see the combination of the `PartialExpr` with `Q` to create a complete `Expr`:
partial = PartialExpr('==>', P)
partial | ~Q
# This [trick](http://code.activestate.com/recipes/384122-infix-operators/) is due to [Ferdinand Jamitzky](http://code.activestate.com/recipes/users/98863/), with a modification by [<NAME>](https://github.com/Chipe1),
# who suggested using a string inside the or-bars.
#
# ## Appendix: The Implementation of `expr`
#
# How does `expr` parse a string into an `Expr`? It turns out there are two tricks (besides the Jamitzky/Vedant trick):
#
# 1. We do a string substitution, replacing "`==>`" with "`|'==>'|`" (and likewise for other operators).
# 2. We `eval` the resulting string in an environment in which every identifier
# is bound to a symbol with that identifier as the `op`.
#
# In other words,
expr('~(P & Q) ==> (~P | ~Q)')
# is equivalent to doing:
P, Q = symbols('P, Q')
~(P & Q) |'==>'| (~P | ~Q)
# One thing to beware of: this puts `==>` at the same precedence level as `"|"`, which is not quite right. For example, we get this:
P & Q |'==>'| P | Q
# which is probably not what we meant; when in doubt, put in extra parens:
(P & Q) |'==>'| (P | Q)
# ## Examples
from notebook import Canvas_fol_bc_ask
canvas_bc_ask = Canvas_fol_bc_ask('canvas_bc_ask', crime_kb, expr('Criminal(x)'))
# # Authors
#
# This notebook by [<NAME>](https://github.com/chiragvartak) and [<NAME>](https://github.com/norvig).
#
#
| logic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="k0dvbuCzd5qK"
# Model Building
from sklearn.feature_selection import SelectKBest, f_regression
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.metrics import mean_absolute_error, mean_squared_error
from sklearn.pipeline import make_pipeline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# For running this notebook either locally or in colab
import sys
# + id="wz7tyC22gLEA"
# %%capture
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Applied-Modeling/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="eK1JmTLj8wFC" colab={"base_uri": "https://localhost:8080/"} outputId="692f2098-b97f-4522-c298-40e26568a18e"
# For encoding categorical data
from category_encoders import OneHotEncoder
# + [markdown] id="c9XvBGFMguJ1"
# # NYC Rent
#
# **GOAL:** Improve our model for predicting NYC rent prices.
#
# **Objectives**
#
# - Do one-hot encoding of categorical features
# - Do univariate feature selection
# - Use scikit-learn to fit Ridge Regression models
# + [markdown] id="0kaV7oNahJLj"
# # I. Wrangle Data
# + [markdown] id="iOecYDRyx5m-"
# Create **wrangle function** for **reproducibility**.
# + id="m9Dm3h3Ux5AI"
def wrangle(filepath):
df = pd.read_csv(filepath, parse_dates = ['created'], index_col = 'created')
# Remove outlier observations
df = df[(df['price'] >= np.percentile(df['price'], 0.5)) &
(df['price'] <= np.percentile(df['price'], 99.5)) &
(df['latitude'] >= np.percentile(df['latitude'], 0.05)) &
(df['latitude'] < np.percentile(df['latitude'], 99.95)) &
(df['longitude'] >= np.percentile(df['longitude'], 0.05)) &
(df['longitude'] <= np.percentile(df['longitude'], 99.95))]
#df = df.drop(columns=['description', 'display_address', 'street_address'], inplace = True)
drop_cols = [col for col in df.select_dtypes('object') if df[col].nunique() > 10]
df.drop(columns = drop_cols, inplace=True)
return df
df = wrangle(DATA_PATH+'apartments/renthop-nyc.csv')
# + [markdown] id="eC-7iMbK9K9m"
# Feature Engineering and Visualization
# + colab={"base_uri": "https://localhost:8080/", "height": 440} id="fHW9kB8S9Ih6" outputId="9dfd0bfb-dc7b-4775-965f-38727348f39d"
df.head()
# + [markdown] id="Fr-x5LJo0pRc"
# # II. Split Data
#
# Split **target vector** from **feature matrix**.
# + id="r2TXo5xLo662"
target = 'price'
y = df[target]
X = df.drop(columns=target)
# + [markdown] id="ZMoRDMuT7A7q"
# Split data into **training** and **test** sets.
#
# (Use data from April & May 2016 to train. Use data from June 2016 to test.)
# + id="7w92aZ7co9CA"
cutoff = '2016-06-24 07:54:24'
mask = X.index < cutoff
X_train, y_train = X.loc[mask], y.loc[mask]
X_test, y_test = X.loc[~mask],y.loc[~mask]
# + [markdown] id="-x9lFKmupRWk"
# # III. Establish Baseline
# + [markdown] id="d1BOKNBc7Jnf"
# **Note:** This is a **regression** problem because we're predictiong the continuous value `'price'`.
# + id="-XNZHgf-7XQc" colab={"base_uri": "https://localhost:8080/"} outputId="ffa4de7d-6e80-43d1-ebc4-8a00ea136aad"
y_pred = [y_train.mean()] * len(y_train)
MAE = mean_absolute_error(y_train, y_pred)
print('Baseline MAE:', mean_absolute_error(y_train, y_pred))
# + [markdown] id="Xea7aFidqPoV"
# # IV. Build Models
# Before we build our model, we need to **encode** our categorical features
# -for high-dimensional data, your have a few options first is **dimensionality reduction** using SelectKBest,
# + id="2IXT_NJSVmnp"
# + [markdown] id="TFc7aAwylBwv"
# **Question:** How can we represent *categorical* features numerically so that we can use them to train our model?
# + [markdown] id="d4vt3ULnbapJ"
# We represent the categorical variable by encoding
# + colab={"base_uri": "https://localhost:8080/"} id="4oz_OMUeE3G8" outputId="2ab3f24b-a08d-4543-8f7e-89aa7c902fd4"
#instatiate Category coders works better
transformer = OneHotEncoder(use_cat_names = True)
# Fit the Transformer to the training data
transformer.fit(X_train)
#transform the data
XT_train = transformer.transform(X_train)
XT_test = transformer.transform(X_test)
XT_train.shape
# + id="u_ycWhZ5H1iq"
#Working on Another transformer
from sklearn.feature_selection import SelectKBest
transformer_2 = SelectKBest(k = 15) # K is a hyperparameter
transformer_2.fit(XT_train, y_train)
#Step:
XTT_train = transformer_2.transform(XT_train)
XTT_test = transformer_2.transform(XT_test)
# + [markdown] id="35SyuZhQQqNu"
# **Model 1:** Regular ol' linear regression
# + id="OOsVETc19TXY" colab={"base_uri": "https://localhost:8080/"} outputId="f33912a0-2d0a-4a7a-eb5d-d6b48d736a7b"
#instantaite our predictor
model = LinearRegression()
# fit our model
model.fit(XTT_train, y_train)
# + [markdown] id="gMjtQqeVQufl"
# **Model 2:** Ridge regression (regularization)
# + id="ZRs-fAaeQt1e" colab={"base_uri": "https://localhost:8080/"} outputId="48184754-63b5-47ae-c603-0f56af3c4fef"
model_r = Ridge(alpha=1)
model_r.fit(XT_train, y_train) # This is the OHE data w/ no SelectKBEst
# + [markdown] id="J2N4D4VFSiks"
# **Model 3:** `SelectKBest`
# + id="OiJ7pnJRSh-N"
model_lr_skb = ...
# + [markdown] id="rHhIH6VE0iRH"
# # V. Check Metrics
# + id="hDj8LEUVOdGs" colab={"base_uri": "https://localhost:8080/"} outputId="23d7e696-1f5d-4551-8787-def5af77336d"
print('LR Training MAE', mean_absolute_error(y_train, model.predict(XTT_train)))
print('LR Test MAE', mean_absolute_error(y_test, model.predict(XTT_test)))
# + id="BXrqHj_lHPum" colab={"base_uri": "https://localhost:8080/"} outputId="63127a9f-7d72-4026-a5dc-a752be68455c"
print('Ridge Training MAE', mean_absolute_error(y_train, model_r.predict(XT_train)))
print('Ridge Test MAE', mean_absolute_error(y_test, model_r.predict(XT_test)))
# + id="ZJUhiKt7TrW5"
# + [markdown] id="uI9K2OXDWThQ"
# # Communicate results
# + id="84xIPGtUBbsA"
# + id="RlAyipCNav1H"
# + id="kGaaKSUXaeGo"
| module3-ridge-regression/213_guided_project_notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
from collections import defaultdict, Counter
from matplotlib import pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all'
# -
data = \
[[['<NAME>'],
['<NAME>', '<NAME>', 'Холмин'],
['<NAME>', 'Холмин', 'Алексей'],
['Холмин', 'Алексей'],
['Холмин'],
['Холмин', 'Диковский', '<NAME>'],
['Холмин', '<NAME>'],
['Холмин'],
['Холмин', '<NAME>']],
[['Лакей', 'Холмин'],
['Холмин'],
['Холмин', 'Диковский'],
['Холмин', 'Диковский', 'Лакей', 'Алексей'],
['Холмин', 'Диковский'],
['Диковский', '<NAME>'],
['Холмин'],
['Холмин', 'Алексей'],
['<NAME>', 'Холмин'],
['<NAME>', 'Холмин', 'Лидия Григорьевна', 'Диковский']],
[['Холмин', 'Е<NAME>'],
['Холмин'],
['Холмин', 'Е<NAME>'],
['Холмин', 'Акулина'],
['Холмин', 'Акулина', 'Диковский'],
['Холмин', 'Диковский'],
['Холмин', 'Диковский', 'Е<NAME>'],
['Холмин', 'Диковский'],
['Холмин', 'Диковский', 'Акулина'],
['Холмин', 'Диковский'],
['Диковский'],
['Холмин', 'Диковский'],
['Холмин', 'Диковский', '<NAME>'],
['Холмин', '<NAME>'],
['Холмин', 'Е<NAME>горьевна', 'Акулина'],
['Холмин', 'Е<NAME>ригорьевна']],
[['Е<NAME>', 'Кормилица'],
['Холмин', '<NAME>'],
['Холмин', 'Е<NAME>горьевна', 'Лидия Григорьевна'],
['Лидия Григорьевна', 'Холмин', 'Елена Григорьевна'],
['Лидия Григорьевна', 'Елена Григорьевна'],
['Лидия Григорьевна', 'Елена Григорьевна', 'Холмин']],
[['Лидия Григорьевна',
'Диковский',
'Старенький князь',
'Фельетонист',
'Юноша'],
['<NAME>', 'Диковский'],
['<NAME>', 'Диковский', 'Алексей', '<NAME>'],
['Фельетонист'],
['<NAME>'],
['Алексей', 'Холмин'],
['Холмин'],
['Холмин', '<NAME>'],
['Холмин'],
['Холмин', 'Диковский'],
['Холмин'],
['Холмин', '<NAME>', 'Диковский'],
['Холмин', '<NAME>'],
['Холмин',
'<NAME>',
'Диковский',
'Старенький князь',
'Юноша',
'Фельетонист',
'Алексей',
'<NAME>']]]
# ## Матрица
drama_heroes = [hero for act in data for scene in act for hero in scene]
drama_heroes = list(sorted(set(drama_heroes)))
matrices = [np.array([[1 if hero in scene else 0 for scene in act] for hero in drama_heroes]) for act in data]
# +
def rome(dec):
return {1: 'I', 2: 'II', 3: 'III', 4: 'IV', 5: 'V', 6: 'VI'}[dec]
def col(act, scene):
return rome(act) + '.' + str(scene)
def cols():
return [col(n_act+1, n_scene+1) for n_act, act in enumerate(matrices) for n_scene in range(len(act[0]))]
matrix = np.hstack(matrices)
pd.DataFrame(matrix, index=drama_heroes, columns=cols())
# -
# ## Мобильность
# +
def mob(mat):
return (np.sum(np.abs(mat[:, 1:] - mat[:, :-1]), axis=1) - 1) / (len(mat[0])-1)
index = list(range(1, len(matrices) + 1)) + ['пьеса']
mobility = list(map(mob, matrices + [matrix]))
mobility = pd.DataFrame(mobility, index=index, columns=drama_heroes).round(2).transpose()
mobility[mobility < 0] = ''
mobility
# -
# ## Плотность
def dens(matrix):
# print(np.count_nonzero(matrix), '/', matrix.size, sep='', end=', ')
return np.count_nonzero(matrix)/matrix.size
# +
print('Плотность всей пьесы:', round(dens(matrix), 2))
act_d = list(map(dens, matrices))
act_d_ = np.array(act_d).reshape(-1, 1)
scene_d = [[dens(c) for c in m.T] for m in matrices]
scene_d_ = pd.DataFrame(scene_d).values # to fill with nans
columns = list(range(1, len(scene_d_[0]) + 1)) + ['действия']
index = list(range(1, len(data)+1))
density = pd.DataFrame(np.hstack((scene_d_, act_d_)), columns=columns, index=index)
density.round(2).fillna('')
# +
from itertools import accumulate
plt.subplots(figsize=(16, 5));
flat = [tup for act in scene_d for tup in enumerate(act, start=1)]
ticks, values = zip(*flat)
plt.plot(values);
plt.xticks(range(len(ticks)), ticks);
borders = np.array(list(accumulate(map(len, [[]] + scene_d))))
plt.scatter(borders[:-1]-0.5, act_d, c='red');
for x in borders[:-1]:
plt.axvline(x-0.5, c='gray', ls='--');
plt.xlabel('явление');
plt.ylabel('плотность');
plt.grid();
# -
plt.plot(act_d)
plt.grid();
plt.xlabel('действие');
plt.ylabel('плотность');
plt.xticks(range(len(act_d)+1), range(1, len(act_d)+1));
plt.xlim([0, len(act_d)-1]);
# ## Расстояние
def dist(mat):
n = len(drama_heroes)
dist = np.zeros((n, n))*np.nan
for i in range(n-1):
for j in range(i+1, n):
if np.sum(mat[i]) != 0 and np.sum(mat[j]) != 0:
dist[i][j] = dist[j][i] = np.sum(np.abs(mat[i] - mat[j]))
return dist/len(mat[0])
act_d = [dist(m) for m in matrices]
drama_d = dist(matrix)
distances = [pd.DataFrame(d, index=drama_heroes, columns=drama_heroes).stack() for d in act_d + [drama_d]]
distances = pd.concat(distances, axis=1)
distances.columns = list(range(1, len(act_d) + 1)) + ['пьеса']
distances.round(2).fillna('')
plt.subplots(figsize=(14, 10))
sns.heatmap(drama_d, vmin=0, vmax=1, annot=True, xticklabels=drama_heroes, yticklabels=drama_heroes, fmt='.2f')
plt.title('по всей пьесе');
# ## Абстрактные типы отношений (= соп., <> альт., > или < дом.)
for i, j in zip(*np.where(drama_d == 0)):
if i != j and i < j:
p = np.sum(matrix[i])
q = np.sum(matrix[j])
s = '=' if p == q else ('>' if p > q else '<')
print(drama_heroes[i], s, drama_heroes[j])
# ## Ранг
rank = [np.sum(np.sum(matrix, axis=0)[np.where(matrix[i] != 0)[0]] - 1) for i in range(len(drama_heroes))]
df = pd.DataFrame(dict(zip(drama_heroes, rank)), index=['ранг']).sort_values(by='ранг', axis=1, ascending=False)
plt.scatter(range(len(drama_heroes)), df.values);
plt.xticks(range(len(drama_heroes)), df.columns, rotation=90);
plt.grid();
df
# ## Абсолютные относительные частоты
# +
def freq(mat):
return np.sum(mat, axis=1)/len(mat[0])
index = list(range(1, len(matrices) + 1)) + ['пьеса']
freq_abs = list(map(freq, matrices + [matrix]))
freq_abs = pd.DataFrame(freq_abs, index=index, columns=drama_heroes).round(2).transpose()
freq_abs[freq_abs == 0] = ''
freq_abs.sort_values(by='пьеса', ascending=False)
# -
# ## Условные относительные частоты
# +
def rel_freq(mat):
busyness = np.sum(mat, axis=1)
freq_rel = np.zeros((len(drama_heroes), len(drama_heroes)))
for i in range(len(mat)):
for j in range(len(mat)):
if busyness[j] == 0 or i == j:
freq_rel[i, j] = np.nan
else:
freq_rel[i, j] = np.count_nonzero(mat[i] + mat[j] - 2 == 0)/busyness[j]
return freq_rel
for i, m in enumerate(matrices):
print('Действие', i+1)
pd.DataFrame(rel_freq(m), index=drama_heroes, columns=drama_heroes).round(2).fillna('')
print('По всей пьесе')
pd.DataFrame(rel_freq(matrix), index=drama_heroes, columns=drama_heroes).round(2).fillna('')
# +
for i, m in enumerate(matrices):
plt.subplots(figsize=(14, 10))
sns.heatmap(rel_freq(m), vmin=0, vmax=1, annot=True, xticklabels=drama_heroes, yticklabels=drama_heroes, fmt='.2f')
plt.title(r"$P_{1,2}$ в действии " + str(i+1));
plt.subplots(figsize=(14, 10))
sns.heatmap(rel_freq(matrix), vmin=0, vmax=1, annot=True, xticklabels=drama_heroes, yticklabels=drama_heroes, fmt='.2f')
plt.title(r"$P_{1,2}$ по всей пьесе");
# -
freq_abs = freq(matrix).reshape((-1, 1))
freq_rel = rel_freq(matrix)
freq_ = freq_abs - freq_rel
freq_[np.diag_indices(len(freq_[0]))] = freq_abs.ravel()
plt.subplots(figsize=(14, 10))
sns.heatmap(freq_, annot=True, xticklabels=drama_heroes, yticklabels=drama_heroes, fmt='.2f', vmin=-1, vmax=1)
plt.title(r"$P_1 – P_{1,2}$, на диагонали абсолютные частоты");
| notebooks/Блуждающие огни.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
mean_df = pd.read_csv('data/opendata-aggregated-means.csv', index_col=0, sep='\t')
std_df = pd.read_csv('data/opendata-aggregated-std.csv', index_col=0, sep='\t')
n_df = pd.read_csv('data/opendata-aggregated-n.csv', index_col=0, sep='\t')
mean_df.shape, std_df.shape, n_df.shape
mean_df.head(3)
stderr_df = std_df / np.sqrt(n_df)
stderr_df.head(3)
# +
measures = {
1 : ["playful","serious"],
2 : ["shy","bold"],
3 : ["cheery","sorrowful"],
4 : ["masculine","feminine"],
5 : ["charming","awkward"],
6 : ["lewd","tasteful"],
7 : ["intellectual","physical"],
8 : ["strict","lenient"],
9 : ["refined","rugged"],
10 : ["trusting","suspicious"],
11 : ["innocent","worldly"],
12 : ["artistic","scientific"],
13 : ["stoic","expressive"],
14 : ["cunning","honorable"],
15 : ["orderly","chaotic"],
16 : ["normal","weird"],
17 : ["competitive","cooperative"],
18 : ["tense","relaxed"],
19 : ["brave","careful"],
20 : ["spiritual","skeptical"],
21 : ["unlucky","fortunate"],
22 : ["ferocious","pacifist"],
23 : ["modest","flamboyant"],
24 : ["dominant","submissive"],
25 : ["forgiving","vengeful"],
26 : ["wise","foolish"],
27 : ["impulsive","cautious"],
28 : ["loyal","traitorous"],
29 : ["creative","conventional"],
30 : ["curious","apathetic"],
31 : ["rude","respectful"],
32 : ["diligent","lazy"],
33 : ["lustful","chaste"],
34 : ["chatty","reserved"],
35 : ["emotional","logical"],
36 : ["moody","stable"],
37 : ["dunce","genius"],
38 : ["arrogant","humble"],
39 : ["heroic","villainous"],
40 : ["attractive","repulsive"],
41 : ["rational","whimsical"],
42 : ["mischievous","well behaved"],
43 : ["aloof","obsessed"],
44 : ["indulgent","sober"],
45 : ["kinky","vanilla"],
46 : ["straightforward","cryptic"],
47 : ["spontaneous","deliberate"],
48 : ["libertarian","socialist"],
49 : ["scheduled","spontaneous"],
50 : ["works hard","plays hard"],
51 : ["reasoned","instinctual"],
52 : ["focused on the present","focused on the future"],
53 : ["empirical","theoretical"],
54 : ["open","guarded"],
55 : ["methodical","astonishing"],
56 : ["mighty","puny"],
57 : ["bossy","meek"],
58 : ["barbaric","civilized"],
59 : ["gregarious","private"],
60 : ["quiet","loud"],
61 : ["political","nonpolitical"],
62 : ["confident","insecure"],
63 : ["democratic","authoritarian"],
64 : ["debased","pure"],
65 : ["fast","slow"],
66 : ["frugal","lavish"],
67 : ["ludicrous","sensible"],
68 : ["orange","purple"],
69 : ["tall","short"],
70 : ["young","old"],
71 : ["down2earth","head@clouds"],
72 : ["extrovert","introvert"],
73 : ["open to new experinces","uncreative"],
74 : ["calm","anxious"],
75 : ["disorganized","self-disciplined"],
76 : ["quarrelsome","warm"],
77 : ["nerd","jock"],
78 : ["lowbrow","highbrow"],
79 : ["selfish","altruistic"],
80 : ["autistic","neurotypical"],
81 : ["angelic","demonic"],
82 : ["hesitant","decisive"],
83 : ["devout","heathen"],
84 : ["cruel","kind"],
85 : ["direct","roundabout"],
86 : ["mathematical","literary"],
87 : ["blue-collar","ivory-tower"],
88 : ["slovenly","stylish"],
89 : ["playful","shy"],
90 : ["serious","bold"],
91 : ["charming","trusting"],
92 : ["awkward","suspicious"],
93 : ["hipster","basic"],
94 : ["coordinated","clumsy"],
95 : ["funny","humorless"],
96 : ["politically correct","edgy"],
97 : ["rich","poor"],
98 : ["hard","soft"],
99 : ["remote","involved"],
100 : ["metaphorical","literal"],
101 : ["biased","impartial"],
102 : ["mundane","extraordinary"],
103 : ["tirewsome","interesting"],
104 : ["smooth","rough"],
105 : ["spicy","mild"],
106 : ["enslaved","emancipated"],
107 : ["optimistic","pessimistic"],
108 : ["sickly","healthy"],
109 : ["luddite","technophile"],
110 : ["vain","demure"],
111 : ["high-tech","low-tech"],
112 : ["flexible","rigid"],
113 : ["cosmopolitan","provincial"],
114 : ["arcane","mainstream"],
115 : ["outlaw","sheriff"],
116 : ["pronatalist","child free"],
117 : ["sad","happy"],
118 : ["jealous","compersive"],
119 : ["bitter","sweet"],
120 : ["resigned","resistant"],
121 : ["sarcastic","genuine"],
122 : ["human","animalistic"],
123 : ["sporty","bookish"],
124 : ["moderate","extreme"],
125 : ["angry","good-humored"],
126 : ["depressed","bright"],
127 : ["self-conscious","self-assured"],
128 : ["vulnerable","armoured"],
129 : ["warm","cold"],
130 : ["assertive","passive"],
131 : ["active","slothful"],
132 : ["imaginative","practical"],
133 : ["adventurous","stick-in-the-mud"],
134 : ["obedient","rebellious"],
135 : ["competent","incompetent"],
136 : ["unambitious","driven"],
137 : ["simple","complicated"],
138 : ["proletariat","bourgeoisie"],
139 : ["alpha","beta"],
140 : ["'right-brained'","'left-brained'"],
141 : ["thick-skinned","sensitive"],
142 : ["charismatic","uninspiring"],
143 : ["feisty","gracious"],
144 : ["eloquent","unpolished"],
145 : ["high IQ","low IQ"],
146 : ["insider","outsider"],
147 : ["morning lark","night owl"],
148 : ["thin","thick"],
149 : ["sheeple","conspiracist"],
150 : ["neat","messy"],
151 : ["vague","precise"],
152 : ["philosophical","real"],
153 : ["modern","historical"],
154 : ["judgemental","accepting"],
155 : ["average","deviant"],
156 : ["gossiping","confidential"],
157 : ["official","backdoor"],
158 : ["scholarly","crafty"],
159 : ["leisurely","hurried"],
160 : ["explorer","builder"],
161 : ["captain","first-mate"],
162 : ["mysterious","unambiguous"],
163 : ["independent","codependent"],
164 : ["family-first","work-first"],
165 : ["scruffy","manicured"],
166 : ["wild","tame"],
167 : ["prestigious","disreputable"],
168 : ["scandalous","proper"],
169 : ["unprepared","hoarder"],
170 : ["sheltered","street-smart"],
171 : ["open-minded","close-minded"],
172 : ["permanent","transient"],
173 : ["dramatic","no-nonsense"],
174 : ["apprentice","master"],
175 : ["straight","queer"],
176 : ["androgynous","gendered"],
177 : ["repetitive","varied"],
178 : ["patient","impatient"],
179 : ["poisonous","nurturing"],
180 : ["creepy","disarming"],
181 : ["inspiring","cringeworthy"],
182 : ["soulless","soulful"],
183 : ["hard","soft"],
184 : ["beautiful","ugly"],
185 : ["domestic","industrial"],
186 : ["juvenile","mature"],
187 : ["idealist","realist"],
188 : ["nihilist","existentialist"],
189 : ["objective","subjective"],
190 : ["theist","atheist"],
191 : ["classical","avant-garde"],
192 : ["utilitarian","decorative"],
193 : ["generalist","specialist"],
194 : ["multicolored","monochrome"],
195 : ["complimentary","insulting"],
196 : ["individualist","communal"],
197 : ["equitable","hypocritical"],
198 : ["traditional","unorthodox"],
199 : ["workaholic","slacker"],
200 : ["resourceful","helpless"],
201 : ["crazy","sane"],
202 : ["anarchist","statist"],
203 : ["cool","dorky"],
204 : ["important","irrelevant"],
205 : ["noob","pro"],
206 : ["deranged","reasonable"],
207 : ["rural","urban"],
208 : ["introspective","not introspective"],
209 : ["city-slicker","country-bumpkin"],
210 : ["western","eastern"],
211 : ["mad","glad"],
212 : ["social","reclusive"],
213 : ["studious","goof-off"],
214 : ["slugabed","go-getter"],
215 : ["penny-pincher","overspender"],
216 : ["liberal","conservative"],
217 : ["unassuming","pretentious"],
218 : ["persistent","quitter"],
219 : ["hedonist","monastic"],
220 : ["patriotic","unpatriotic"],
221 : ["tactful","indiscreet"],
222 : ["wholesome","salacious"],
223 : ["joyful","miserable"],
224 : ["zany","regular"],
225 : ["alert","oblivious"],
226 : ["feminist","sexist"],
227 : ["racist","egalitarian"],
228 : ["abstract","concrete"],
229 : ["formal","intimate"],
230 : ["resolute","wavering"],
231 : ["deep","shallow"],
232 : ["valedictorian","drop out"],
233 : ["minimalist","pack rat"],
234 : ["trash","treasure"],
235 : ["🥰","🙃"],
236 : ["🥴","🥳"],
237 : ["😎","🧐"],
238 : ["😭","😀"],
239 : ["🤑","🤠"],
240 : ["👽","🤡"],
241 : ["💔","💝"],
242 : ["🤖","👻"],
243 : ["💩","🌟"],
244 : ["💪","🧠"],
245 : ["🙋‍♂️","🙅‍♂️"],
246 : ["👨‍⚕️","👨‍🔧"],
247 : ["👩‍🔬","👩‍🎤"],
248 : ["😈","😇"],
249 : ["🤔","🤫"],
250 : ["🐘","🐀"],
251 : ["🐮","🐷"],
252 : ["🐴","🦄"],
253 : ["🐩","🐒"],
254 : ["🚴","🏋️‍♂️"],
255 : ["🤺","🏌"],
256 : ["💃","🧕"],
257 : ["🧙","👨‍🚀"],
258 : ["🐐","🦒"],
259 : ["🦇","🐿"],
260 : ["😬","😏"],
261 : ["🤐","😜"],
262 : ["🤣","😊"],
263 : ["🧗","🛌"],
264 : ["🥾","👟"],
265 : ["🎩","🧢"],
266 : ["📈","📉"],
267 : ["stinky","fresh"],
268 : ["legit","scrub"],
269 : ["self-destructive","self-improving"],
270 : ["French","Russian"],
271 : ["German","English"],
272 : ["Italian","Swedish"],
273 : ["Greek","Roman"],
274 : ["traumatized","flourishing"],
275 : ["sturdy","flimsy"],
276 : ["macho","metrosexual"],
277 : ["claustrophobic","spelunker"],
278 : ["offended","chill"],
279 : ["rhythmic","stuttering"],
280 : ["musical","off-key"],
281 : ["lost","enlightened"],
282 : ["masochistic","pain-avoidant"],
283 : ["efficient","overprepared"],
284 : ["oppressed","privileged"],
285 : ["sunny","gloomy"],
286 : ["vegan","cannibal"],
287 : ["loveable","punchable"],
288 : ["slow-talking","fast-talking"],
289 : ["believable","poorly-written"],
290 : ["vibrant","geriatric"],
291 : ["consistent","variable"],
292 : ["dispassionate","romantic"],
293 : ["linear","circular"],
294 : ["intense","lighthearted"],
295 : ["knowledgeable","ignorant"],
296 : ["fixable","unfixable"],
297 : ["exuberant","subdued"],
298 : ["secretive","open-book"],
299 : ["perceptive","unobservant"],
300 : ["folksy","presidential"],
301 : ["corporate","freelance"],
302 : ["sleepy","frenzied"],
303 : ["loose","tight"],
304 : ["narcissistic","low self esteem"],
305 : ["poetic","factual"],
306 : ["melee","ranged"],
307 : ["giggling","chortling"],
308 : ["whippersnapper","sage"],
309 : ["tailor","blacksmith"],
310 : ["hunter","gatherer"],
311 : ["experimental","reliable"],
312 : ["moist","dry"],
313 : ["trolling","triggered"],
314 : ["tattle-tale","f***-the-police"],
315 : ["punk rock","preppy"],
316 : ["realistic","fantastical"],
317 : ["trendy","vintage"],
318 : ["factual","exaggerating"],
319 : ["good-cook","bad-cook"],
320 : ["comedic","dramatic"],
321 : ["OCD","ADHD"],
322 : ["interrupting","attentive"],
323 : ["exhibitionist","bashful"],
324 : ["badass","weakass"],
325 : ["gamer","non-gamer"],
326 : ["random","pointed"],
327 : ["epic","deep"],
328 : ["serene","pensive"],
329 : ["bored","interested"],
330 : ["envious","prideful"],
331 : ["ironic","profound"],
332 : ["sexual","asexual"],
333 : ["🥵","🥶"],
334 : ["🎃","💀"],
335 : ["🏀","🎨"],
336 : ["clean","perverted"],
337 : ["empath","psychopath"],
338 : ["haunted","blissful"],
339 : ["entitled","grateful"],
340 : ["ambitious","realistic"],
341 : ["stuck-in-the-past","forward-thinking"],
342 : ["fire","water"],
343 : ["earth","air"],
344 : ["lover","fighter"],
345 : ["overachiever","underachiever"],
346 : ["Coke","Pepsi"],
347 : ["twitchy","still"],
348 : ["freak","normie"],
349 : ["thinker","doer"],
350 : ["hard-work","natural-talent"],
351 : ["stingy","generous"],
352 : ["stubborn","accommodating"],
353 : ["extravagant","thrifty"],
354 : ["demanding","unchallenging"],
355 : ["two-faced","one-faced"],
356 : ["plastic","wooden"],
357 : ["neutral","opinionated"],
358 : ["chivalrous","businesslike"],
359 : ["high standards","desperate"],
360 : ["on-time","tardy"],
361 : ["everyman","chosen one"],
362 : ["jealous","opinionated"],
363 : ["protagonist","antagonist"],
364 : ["devoted","unfaithful"],
365 : ["fearmongering","reassuring"],
366 : ["common sense","analysis"],
367 : ["unemotional","emotional"],
368 : ["rap","rock"],
369 : ["genocidal","not genocidal"],
370 : ["cat person","dog person"],
371 : ["indie","pop"],
372 : ["cultured","rustic"],
373 : ["tautology","oxymoron"],
374 : ["bad boy","white knight"],
375 : ["princess","queen"],
376 : ["hypochondriac","stoic"],
377 : ["yes-man","contrarian"],
378 : ["giving","receiving"],
379 : ["chic","cheesy"],
380 : ["celebrity","boy/girl-next-door"],
381 : ["goth","flower child"],
382 : ["summer","winter"],
383 : ["frank","sugarcoated"],
384 : ["naive","paranoid"],
385 : ["gullible","cynical"],
386 : ["motivated","unmotivated"],
387 : ["radical","centrist"],
388 : ["monotone","expressive"],
389 : ["love-focused","money-focused"],
390 : ["transparent","machiavellian"],
391 : ["timid","cocky"],
392 : ["concise","long-winded"],
393 : ["picky","always down"],
394 : ["proactive","reactive"],
395 : ["prudish","flirtatious"],
396 : ["innocent","jaded"],
397 : ["touchy-feely","distant"],
398 : ["muddy","washed"],
399 : ["quirky","predictable"],
400 : ["never cries","often crying"],
}
# -
len(measures)
label_columns = [','.join(measures[int(column.replace('F', ''))]) for column in mean_df.columns ]
len(label_columns)
labeled_df = pd.DataFrame(mean_df)
labeled_df.columns = label_columns
labeled_df.head(5)
manual_labels_df = pd.read_csv('data/image_labels_manual.csv', index_col=0)
manual_labels_df.head(5)
manual_labels_df.loc['A1.jpg']
# +
labels = list()
for value in labeled_df.index.values:
source_labels = manual_labels_df.loc[value.replace('/', '') + ".jpg"]
if len(source_labels) == 0:
continue
labels.append({
'id': value,
'race.label': source_labels['race.manual'],
'gender.label': source_labels['gender.manual']}
)
consolidated_df = pd.concat([labeled_df, pd.DataFrame(labels).set_index('id')], axis=1)
consolidated_df.head(5)
# +
drop_invalid_race = consolidated_df[consolidated_df['race.label'] != 'unknown']
drop_invalid_gender = drop_invalid_race[drop_invalid_race['gender.label'] != 'unknown']
drop_invalid_gender.to_csv('data/opendata-with-labels.csv')
# -
consolidated_df.loc['A/1']['empath,psychopath']
drop_invalid_gender.shape
| EDA-OpenData.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Wavelet Features
# ## Data Preparation
# +
import os
import sys
from sklearn.model_selection import train_test_split
import numpy
import pandas
from keras.utils import to_categorical
WORKING_DIR_PATH = globals()['_dh'][0]
WORKING_DIR_PARENT_PATH = os.path.dirname(WORKING_DIR_PATH)
sys.path.insert(1, WORKING_DIR_PARENT_PATH)
from custom_module.utilities import *
# -
# ? tf.feature_column
tf.__version__
import sklearn; sklearn.__version__
# load dataset
features = pandas.read_csv(MOUNTED_DATASET_PATH + '/data/cleaned_features_1.csv', index_col=0)
features
# shuffle dataset
features = features.reindex(numpy.random.permutation(features.index))
features
# split dataset into target (y) and predictors (x)
y = features.loc[:,'genre_label']
X = features.loc[:, features.columns.difference(['genre_label'])]
# get wavelet predictors
wavelet_predictors = X.filter(regex=(r'.+_db[458]{1}_.+'))
wavelet_predictor_labels = wavelet_predictors.columns.values
wavelet_predictors
# get wavelet predictor labels with and without outliers
wavelet_predictor_labels_wf_outliers= numpy.asarray([
'25th_percentile_db8_cA7', '5th_percentile_db8_cA7', '5th_percentile_db8_cD7',
'75th_percentile_db8_cA7', 'mcr_db4_cD1', 'mcr_db4_cD2', 'mcr_db8_cD2',
'mcr_db8_cD1', 'mcr_db5_cD2','mcr_db5_cD1','zcr_db8_cD1'])
wavelet_predictor_labels_wo_outliers = [i for i in wavelet_predictor_labels \
if i not in wavelet_predictor_labels_wf_outliers]
# +
# Create standardization and normalization pipelines
preprocess_sp = standardization_pipeline(
wavelet_predictor_labels,
wavelet_predictor_labels_wf_outliers,
wavelet_predictor_labels_wo_outliers)
preprocess_np = normalization_pipeline(
wavelet_predictor_labels)
# -
# split into sets: training 80% & testing 20% of total dataset
X_train, X_test, y_train, y_test = train_test_split(
wavelet_predictors, y, test_size=0.2, shuffle=True)
_, ncols = X_train.shape
ncols
# one hot encoding our labels
y_train = to_categorical(y_train, 3)
y_test = to_categorical(y_test, 3)
y_train.shape, y_test.shape
# ## Predictive Modelling
param_dist = {
'clf__n_hidden': [1,2,3,4,5],
'clf__activation': ['relu', 'elu', 'selu'],
'clf__optimizer': ['rmsprop','adam','adagrad'],
'clf__kernel_initializer':['glorot_uniform','normal','uniform'],
'clf__lr': [3, 1, .3, .1, .03, .01, .003, .001, .0003, .0001],
'clf__units': numpy.arange(4,100)
}
# ### FNN With Standardized Input
(fnn1_best_score, fnn1_best_params, fnn1_best_estimator) = \
train_model(X_train, y_train, 'fnn_1', ncols, set_shape_create_model,
pipeline_estimator_sp, param_dist)
# ### FNN With Normalized Input
(fnn2_best_score, fnn2_best_params, fnn2_best_estimator) = \
train_model(X_train, y_train, 'fnn_2', ncols, set_shape_create_model,
pipeline_estimator_np, param_dist)
# ### CNN with Standardized Input
param_dist = {
'clf__n_hidden': [1, 2, 3],
'clf__activation': ['relu', 'elu', 'selu'],
'clf__optimizer': ['rmsprop','adam','adagrad'],
'clf__kernel_initializer':['glorot_uniform','normal','uniform'],
'clf__units': numpy.arange(1,100),
'clf__filters': [16, 32, 64],
# 'clf__kernel_size': [3,4,5],
'clf__dropout':[.1, .15, .2, .25],
'clf__lr': [3, 1, .3, .1, .03, .01, .003, .001, .0003, .0001],
}
(cnn1_best_score, cnn1_best_params, cnn1_best_estimator) = \
train_model(X_train, y_train, 'cnn_1', ncols, set_shape_create_cnn_model,
pipeline_estimator_sp, param_dist)
# ### CNN With Normalized Input
(cnn2_best_score, cnn2_best_params, cnn2_best_estimator) = \
train_model(X_train, y_train, 'cnn_2', ncols, set_shape_create_cnn_model,
pipeline_estimator_np, param_dist)
| feature_extraction_deep_learning/notebooks/06_data_prep_predictive_modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Berea Sandstone Simulation Using PoreSpy and OpenPNM
# The example explains effective permeabilty calculations using PoreSpy and OpenPNM software. The simulation is performed on X-ray tomography image of [BereaSandstone](https://www.imperial.ac.uk/earth-science/research/research-groups/perm/research/pore-scale-modelling/micro-ct-images-and-networks/berea-sandstone/). The calculated effective permeablity value can compared with value report in [Dong et al](https://www.semanticscholar.org/paper/Pore-network-extraction-from-images.-Dong-Blunt/31fbb0362bd02e483c8b1f19f944f9bf15095a80).
# ### Start by importing the necessary packages
import os
import imageio
import scipy as sp
import numpy as np
import openpnm as op
import porespy as ps
import matplotlib.pyplot as plt
np.set_printoptions(precision=4)
np.random.seed(10)
# %matplotlib inline
# ### Load BreaSandstone Image file
# Give path to image file and load the image. Please note image should be binarized or in boolean format before performing next steps.
path = '../../_fixtures/ICL-Sandstone(Berea)/'
file_format = '.tif'
file_name = 'Berea'
file = file_name + file_format
fetch_file = os.path.join(path, file)
im = imageio.mimread(fetch_file)
im = ~np.array(im, dtype=bool)[:250, :250, :250] # Make image a bit smaller
# ### Confirm image and check image porosity
# Be patient, this might take ~30 seconds (depending on your CPU)
# NBVAL_IGNORE_OUTPUT
fig, ax = plt.subplots(1, 3, figsize=(12,5))
ax[0].imshow(im[:, :, 100]);
ax[1].imshow(ps.visualization.show_3D(im));
ax[2].imshow(ps.visualization.sem(im));
ax[0].set_title("Slice No. 100 View");
ax[1].set_title("3D Sketch");
ax[2].set_title("SEM View");
print(ps.metrics.porosity(im))
# ### Extract pore network using SNOW algorithm in PoreSpy
# The SNOW algorithm (an accronym for Sub-Network from an Oversegmented Watershed) was presented by [Gostick](https://journals.aps.org/pre/abstract/10.1103/PhysRevE.96.023307). The algorithm was used to extract pore network from BereaSandstone image.
# NBVAL_IGNORE_OUTPUT
resolution = 5.345e-6
net = ps.networks.snow(im=im, voxel_size=resolution)
# ### Import network in OpenPNM
# The output from the SNOW algorithm above is a plain python dictionary containing all the extracted pore-scale data, but it is NOT yet an OpenPNM network. We need to create an empty network in OpenPNM, then populate it with the data from SNOW:
pn, geo = op.io.PoreSpy.import_data(net)
# Now we can print the network to see how the transferred worked.
#
# > Note to developers: We need to ignore the output of the following cell since the number of pores differs depending on whether the code is run on a windows or linux machine.
# NBVAL_IGNORE_OUTPUT
print(pn)
# ### Check network health
# Remove isolated pores or cluster of pores from the network by checking it network health. Make sure ALL keys in network health functions have no value.
h = pn.check_network_health()
op.topotools.trim(network=pn, pores=h['trim_pores'])
h = pn.check_network_health()
print(h)
# ### Assign phase
# In this example air is considered as fluid passing through porous channels.
air = op.phases.Air(network=pn)
# ### Assign physics
phys_air = op.physics.Standard(network=pn, phase=air, geometry=geo)
# ### Assign Algorithm and boundary conditions
# Select stokes flow algorithm for simulation and assign dirichlet boundary conditions in top and bottom faces of the network.
perm = op.algorithms.StokesFlow(network=pn)
perm.setup(phase=air)
perm.set_value_BC(pores=pn.pores('top'), values=0)
perm.set_value_BC(pores=pn.pores('bottom'), values=101325)
perm.run()
air.update(perm.results())
# ### Calculate effective permeability
# Caclulate effective permeablity using hagen poiseuille equation. Use cross section area and flow length manually from image dimension.
resolution = 5.345e-6
Q = perm.rate(pores=pn.pores('bottom'), mode='group')[0]
A = (im.shape[0] * im.shape[1]) * resolution**2
L = im.shape[2] * resolution
mu = air['pore.viscosity'].max()
delta_P = 101325 - 0
K = Q * L * mu / (A * delta_P)
# > Note to developers: We need to ignore the output of the following cell since the results are slightly different on different platforms (windows vs linux)
# NBVAL_IGNORE_OUTPUT
print(f'The value of K is: {K/0.98e-12*1000:.2f} mD')
| examples/notebooks/networks/extraction/predicting_effective_permeability_of_berea.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext sql
# %config SqlMagic.feedback = False
# %matplotlib inline
import pandas as pd
# Get env vars from local.env
# host = %env DB_HOSTNAME
# user = %env DB_USERNAME
# password = %env DB_PASSWORD
# db = %env DB_NAME
# Connection URL to our local MySQL DB
# %sql mysql+mysqldb://{user}:{password}@{host}/{db}?charset=utf8
lane_county_fips = '41039'
lane_county_fips_like = '41039%'
# -
# %sql DELETE FROM submissions WHERE county != :lane_county_fips
# %sql DELETE FROM boundaries WHERE boundary_type = 'region' AND boundary_id != 'OR'
# %sql DELETE FROM boundaries WHERE boundary_type = 'county' AND boundary_id != :lane_county_fips
# %sql DELETE FROM boundaries WHERE boundary_type = 'census_tract' AND boundary_id not like :lane_county_fips_like
# %sql DELETE FROM boundaries WHERE boundary_type = 'census_block' AND boundary_id not like :lane_county_fips_like
# %sql DELETE FROM boundaries WHERE boundary_type = 'zip_code' AND boundary_id not like '974%'
# %sql DELETE FROM stats_caches WHERE stat_type = 'region' AND stat_id != '41'
# %sql DELETE FROM stats_caches WHERE stat_type = 'county' AND stat_id != '41039'
# %sql DELETE FROM stats_caches WHERE stat_type = 'zip_code' AND stat_id not like '974%'
# %sql DELETE FROM stats_caches WHERE stat_type = 'census_block' AND stat_id not like '41039%'
# %sql DELETE FROM stats_caches WHERE stat_type = 'census_tract' AND stat_id not like '41039%'
# %sql SHOW tables
# %sql DESCRIBE boundaries
# %sql SELECT boundary_type, count(boundary_id) FROM boundaries GROUP by boundary_type
# %sql SELECT stat_type, count(stat_id) FROM stats_caches GROUP by stat_type
# %sql SELECT count(*) FROM boundaries WHERE boundary_type = 'county'
# %sql SELECT count(*) FROM submissions WHERE county = :lane_county_fips
# +
# submissions_by_date = %sql SELECT test_date, count(id) as submissions FROM submissions WHERE county = :lane_county_fips GROUP BY test_date ORDER BY test_date DESC
submissions_by_date = submissions_by_date.DataFrame()
submissions_by_date['test_date'] = pd.to_datetime(submissions_by_date['test_date'])
submissions_by_date.set_index('test_date', inplace=True)
submissions_by_date.plot(figsize=(20,10))
# -
| notebooks/Narrow to Lane County.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LambdaMART in Python
#
# This is an implementation of LambdaMART in Python using sklearn and pandas. This is for educational purposes.
#
# But a secondary goal in getting this into Python is to more easily hack the algorithm to try new ideas. For example, this [blog article on two-sided marketplaces](https://opensourceconnections.com/blog/2017/07/04/optimizing-user-product-match-economies/), perhaps as more of an online algorithm (retiring old trees in the ensemble, adding new ones over time), perhaps with different model architectures in the ensemble (BERTy transformery things?) but all that preserve some of the nice things about LambdaMART (directly optimizing a list-wise metric)
#
# This is adapted from [RankLib](https://github.com/o19s/RankyMcRankFace/blob/master/src/ciir/umass/edu/learning/tree/LambdaMART.java#L444) based on [this paper](https://www.microsoft.com/en-us/research/wp-content/uploads/2016/02/MSR-TR-2010-82.pdf) from Microsoft Research.
#
# ## Table of Contents
#
# 1. [Setup - TheMovieDB corpus and log Elasticsearch features from TMDB](#Part-Zero---Setup---Get-TheMovieDB-Corpus-and-Log-Simple-Features) - plumbing to interact with Elasticsearch Learning to Rank to log a few basic features for our exploration
# 2. [Pairwise swapping](#Part-One---Collect-pair-wise-DCG-diffs) - here we demonstrate the core operation of LambdaMART - pairwise swapping of pairs and examining DCG (or another ranking metric) impact
# 3. [Scale to learn errors, not just swaps](#Part-Two---Compute-the-swaps-but-scaled-to-current-model's-error) - here we show how LambdaMART isn't just about learning the pairwise DCG difference of a swap, but the error currently in the model at predicting the DCG impact of that swap
# 4. [Weigh predictions](#Part-Three---Weigh-each-leaf's-predictions) - here we weigh the predictions of the next model in the ensemble based on how much DCG remains to be learned.
# 5. [Putting it all together](#Part-Four---Putting-it-all-together,-from-the-top!) - the full algorithm in one place. You can also compare this notebook's output and learning to Ranklib.
#
# ---
#
# 6. [A Pandas version!](#5.-Pure-Pandas-Implementation?) -- walking through a faster version computing the per-tree training data using Pandas - a much for useful toy example.
#
# ## Known Issues
#
# I'm still learning the nooks and crannies of the algorithm. So there are some known issues as this is actively being developed.
#
# 1. **Performance** - a single training round takes about 9 seconds. There's room for improvement in the hot part of the loop (dcg computation and swapping)
# ## Part Zero - Setup - Get TheMovieDB Corpus and Log Simple Features
#
# In this step we download TheMovieDB Corpus and log some featurs (title and overview BM25). At the end we have a simple dataframe
from ltr.client import ElasticClient
client = ElasticClient()
# ### Download and index TMDB corpus and training set
#
# Download [TheMovieDB](http://themoviedb.org) corpus and small toy training set with 40 queries labeled.
# +
from ltr import download
corpus='http://es-learn-to-rank.labs.o19s.com/tmdb.json'
judgments='http://es-learn-to-rank.labs.o19s.com/title_judgments.txt'
download([corpus, judgments], dest='data/');
from ltr.index import rebuild
from ltr.helpers.movies import indexable_movies
movies=indexable_movies(movies='data/tmdb.json')
rebuild(client, index='tmdb', doc_src=movies)
# -
# ### Log two features - title & overview
#
# Using the Elasticsearch Learning to Rank plugin, we:
#
# 1. Log two features: title and overview bm25
# 2. Create a pandas dataframe containing the labels and features
# +
from ltr.log import FeatureLogger
from ltr.judgments import judgments_open
from itertools import groupby
from ltr.judgments import judgments_to_dataframe
client.reset_ltr(index='tmdb')
config = {"validation": {
"index": "tmdb",
"params": {
"keywords": "rambo"
}
},
"featureset": {
"features": [
{ #1
"name": "title_bm25",
"params": ["keywords"],
"template": {
"match": {"title": "{{keywords}}"}
}
},
{ #2
"name": "overview_bm25",
"params": ["keywords"],
"template": {
"match": {"overview": "{{keywords}}"}
}
}
]
}}
client.create_featureset(index='tmdb', name='movies', ftr_config=config)
# Log features for each query
ftr_logger=FeatureLogger(client, index='tmdb', feature_set='movies')
with judgments_open('data/title_judgments.txt') as judgment_list:
for qid, query_judgments in groupby(judgment_list, key=lambda j: j.qid):
ftr_logger.log_for_qid(judgments=query_judgments,
qid=qid,
keywords=judgment_list.keywords(qid))
# Convert to Pandas Dataframe
judgments = judgments_to_dataframe(ftr_logger.logged, unnest=False)
# -
# ### Examine judgments dataframe
#
# In the dataframe we have a set of (query, document, grade) that label how relevant a document (movie) is for each query.
#
# * qid - 'query id' - a unique identifier for this query
# * docId - an identifier for the document (here movie) being labeled
# * grade - how relevant a movie is on a 0-4 scale
# * keywords - the query keywords that go along with the query id
# * features - the two features we logged, 0th is title_bm25, 1st is overview_bm25
judgments
# ## Part One - Collect pair-wise DCG diffs
#
# The first-pass iteration of LambdaMART, for each query, we examine the DCG\* impact of swapping each result with another result in the listing.
#
# \* replace DCG with your metric of interest: MAP, Precision@N, etc
# +
from math import log, exp
import numpy as np
def rank_with_swap(ranked_list, rank1=0, rank2=0):
""" Set the display rank of positions given the provided swap """
ranked_list['display_rank'] = ranked_list.index.to_series()
if rank1 != rank2:
ranked_list.loc[rank1, 'display_rank'] = rank2
ranked_list.loc[rank2, 'display_rank'] = rank1
return ranked_list
def dcg(ranked_list, at=10):
"""Given a list, compute DCG --
uses same variant as lambdamart 2**grade / log2(displayrank)
"""
ranked_list['discount'] = 1 / np.log2(2 + ranked_list['display_rank'])
ranked_list['gain'] = (2**ranked_list['grade'] - 1) * ranked_list['discount'] # TODO - precompute gain on swapping
return sum(ranked_list['gain'].head(at))
def compute_swaps(query_judgments, axis, metric=dcg, at=10):
"""Compute the 'lambda' the DCG impact of every query result swapped with every-other query result"""
# Sort to see ideal ordering
# This isn't strictly nescesarry, but it's helpful to understand the algorithm
query_judgments = query_judgments.sort_values('grade', kind='stable', ascending=False).reset_index()
# Instead of explicitly 'swapping' we just swap the 'display_rank' - where
# in the final ranking this would be placed. We can easily use that to compute DCG
query_judgments['display_rank'] = query_judgments.index.to_series()
query_judgments['dcg'] = metric(query_judgments, at=at)
best_dcg = query_judgments.loc[0, 'dcg']
query_judgments['lambda'] = 0.0
# TODO - redo inner body as
for better in range(0,len(query_judgments)):
for worse in range(0,len(query_judgments)):
if better > at and worse > at:
break
if query_judgments.loc[better, 'grade'] > query_judgments.loc[worse, 'grade']:
query_judgments = rank_with_swap(query_judgments, better, worse)
query_judgments['dcg'] = metric(query_judgments, at=at)
dcg_after_swap = query_judgments.loc[0, 'dcg']
delta = abs(best_dcg - dcg_after_swap)
if delta > 0.0:
# Add delta to better's lambda (-delta to worse's lambda)
query_judgments.loc[better, 'lambda'] += delta
query_judgments.loc[worse, 'lambda'] -= delta
# print(query_judgments[['keywords', 'docId', 'grade', 'lambda', 'features']])
return query_judgments
# For each query, compute lambdas
# # %prun -s cumulative lambdas_per_query = judgments.groupby('qid').apply(compute_swaps, axis=1)
# judgments
lambdas_per_query = judgments.groupby('qid').apply(compute_swaps, axis=1)
lambdas_per_query
# -
# ### Look at Precision instead of DCG
#
# We can really use any ranking metric to achieve goals important to our product. This includes potentially ones we invent or come up with ourselves!
# +
def precision(ranked_list, max_grade=4.0, at=10):
"""Given a list, compute simple precision. Really this is cumalitive gain."""
above_n = ranked_list[ranked_list['display_rank'] < at]
if (max_grade * at) == 0.0:
print("0")
return 0.0
return float(sum(above_n['grade'])) / (max_grade * at)
lambdas_per_query_prec = judgments.groupby('qid').apply(compute_swaps, axis=1, metric=precision)
lambdas_per_query_prec.loc[5, :]
# -
# ### Fit a model on the lambdas
#
# The core operation is fitting an operation on the lambdas (the accumulated pairwise differences)
train_set = lambdas_per_query[['lambda', 'features']]
train_set
# +
from sklearn.tree import DecisionTreeRegressor, plot_tree
tree = DecisionTreeRegressor()
tree.fit(train_set['features'].tolist(), train_set['lambda'])
# -
# ### DCG-based Lambda Predictions
#
# We show predicting some known examples. In the first case, strong title and overview scores. In the second case, no title or overview scores
tree.predict([[11.1, 10.08]])
tree.predict([[0.0, 0.0]])
# It's more typical we would restrict the complexity of each tree in the ensemble. We can dump the tree see [understanding sklearn's tree structure](https://scikit-learn.org/stable/auto_examples/tree/plot_unveil_tree_structure.html#sphx-glr-auto-examples-tree-plot-unveil-tree-structure-py)
tree = DecisionTreeRegressor(max_leaf_nodes=4)
tree.fit(train_set['features'].tolist(), train_set['lambda'])
plot_tree(tree)
# ## Part Two - Compute the swaps but _scaled_ to current model's error
#
# LambdaMART is an _ensemble_ model. It's not just about the first model, but collecting a series of models where each model makes a gradual improvement on the current model. The technique used is known as [Gradient Boosting]()
#
# To build a model that compensates for the current model's error, we scale the next set of dependent vars to predict based on the correctness of the existing model in ranking. In this way, we eliminate where the model currently does a good job (no need to learn these) and leave in places where the model isn't doing a good job (this is where. we want ot learn)
# +
learning_rate = 0.1
judgments['last_prediction'] = tree.predict(judgments['features'].tolist()) * learning_rate
def compute_swaps_scaled(query_judgments, axis, metric=dcg, at=10):
"""Compute the 'lambda' the DCG impact of every query result swapped with every-other query result
"""
# Important - stable sort. Otherwise DCG swaps get kind of wonky due to position discounts
query_judgments = query_judgments.sort_values('last_prediction', ascending=False, kind='stable').reset_index()
# Instead of explicitly 'swapping' we just swap the 'display_rank' - where
# in the final ranking this would be placed. We can easily use that to compute DCG
query_judgments['display_rank'] = query_judgments.index.to_series()
query_judgments['dcg'] = metric(query_judgments, at=at)
best_dcg = query_judgments.loc[0, 'dcg']
query_judgments['lambda'] = 0.0
query_judgments['delta'] = 0.0
for better in range(0,len(query_judgments)):
for worse in range(better+1,len(query_judgments)):
if better > at and worse > at:
break
if query_judgments.loc[better, 'grade'] > query_judgments.loc[worse, 'grade']:
swap_judgments = rank_with_swap(query_judgments, better, worse)
dcg_after_swap = metric(swap_judgments, at=at)
delta = abs(best_dcg - dcg_after_swap)
if delta > 0.0:
# --------------
# NEW!
model_score_diff = query_judgments.loc[better, 'last_prediction'] - query_judgments.loc[worse, 'last_prediction']
rho = 1.0 / (1.0 + exp(model_score_diff))
# --------------
# rho works as follows
#
# model ranks rho
# better higher than worse approaches 0 <-- model currently doing well!
# better same as worse. 0.5
# worse higher than better approaches 1 <-- model currently doing poorly!
#
query_judgments.loc[better, 'delta'] += delta
# Use rho to scale the lambdas
query_judgments.loc[better, 'lambda'] += delta * rho
query_judgments.loc[worse, 'lambda'] -= delta * rho
query_judgments.loc[worse, 'delta'] -= delta
return query_judgments
judgments = judgments_to_dataframe(ftr_logger.logged, unnest=False)
judgments['last_prediction'] = 0.0
lambdas_per_query = judgments.groupby('qid').apply(compute_swaps_scaled, axis=1)
#
lambdas_per_query
# -
# Zero in on 2 swapped by each result worse than it in query `ramba`
#
# ```
# better_grade worse_grade, model_score_diffs, rho, dcg_delta
# 2 1 0.758706128029972 0.31892724571177816 0.02502724555344038
# 2 1 0.981162516523929 0.2726611758805124 0.04377062727053094
# 2 0 1.142439045359345 0.24187283082372052 0.11698017724693699
# 2 0 1.142439045359345 0.24187283082372052 0.14090604137532914
# 2 1 1.142439045359345 0.24187283082372052 0.08043118677314176
# 2 1 1.142439045359345 0.24187283082372052 0.17784892734690594
# 2 1 1.142439045359345 0.24187283082372052 0.19254512210920538
# 2 1 1.142439045359345 0.24187283082372052 0.20543079947538878
# 2 1 1.142439045359345 0.24187283082372052 0.21685570619866112
# 2 0 1.142439045359345 0.24187283082372052 0.22708156316293504
# 2 0 1.142439045359345 0.24187283082372052 0.23630863576764227
# 2 0 1.142439045359345 0.24187283082372052 0.24469312448475122
# 2 0 1.142439045359345 0.24187283082372052 0.2523588995024131
# 2 0 1.142439045359345 0.24187283082372052 0.25940563061320177
# 2 0 1.142439045359345 0.24187283082372052 0.2659145510893115
# ...
# 2 0 1.142439045359345 0.24187283082372052 0.34214193097965406
# ```
#
# Summing all the model score diffs, we see those are rather high. This results in a high-ish rho between (for each value here 0.25-0.31). So each dcg_delta is added to the model.
#
# What's the intuition here? The model hasn't entirely nailed this example, the model feels there's more 'dcg_delta' to learn to push it away from those less relevant results.
# ### Zooming out to more of `rambo`
#
# We see a similar pattern in results with mediocre grades (2 and 3) where the resulting rho-scaled lambda's are higher than you might expect. The model's happy with the position of 0, but the ranking of other results could be separated more. The model diff should be higher when compared to the dcg diff to push the middling results away from the irrelevant result.
#
# So the next tree learns these lambdas using the resulting features moreso than other results.
lambdas_per_query.loc[1, :][['keywords', 'display_rank', 'grade', 'last_prediction', 'delta', 'lambda', 'features']].sort_values('last_prediction', ascending=False)
# +
from sklearn.tree import DecisionTreeRegressor
train_set = lambdas_per_query[['lambda', 'features']]
train_set
tree2 = DecisionTreeRegressor()
tree2.fit(train_set['features'].tolist(), train_set['lambda'])
# -
# ### More 'oomph' in second tree for the last tree's error cases
#
# We see in the following lambdas our next tree learns more about the areas the last model seemed to need correction.
#
# The first example is well covered by the first tree.
tree2.predict([[11.6, 10.08]])
# The second example reflects some of the middling ranked results
tree2.predict([[0.0, 6.869545]])
# ## Part Three - Weigh each leaf's predictions
#
# Because we're dealing with trees, each leaf corresponds to a set of examples that have been grouped to this node. In addition to per-swap 'rho' we also care about a per-swap 'weight', referred to in gradient boosting as 'gamma'.
#
# Gamma means picking a weight for this sub-model that best predicts the final function.
#
# First we group by the paths in the tree to uniquely identify each leaf
# +
def compute_swaps_scaled_with_weights(query_judgments, axis, metric=dcg, at=10):
"""Compute the 'lambda' the DCG impact of every query result swapped with every-other query result
"""
# Sort to see ideal ordering
# This isn't strictly nescesarry, but it's helpful to understand the algorithm
query_judgments = query_judgments.sort_values('last_prediction', ascending=False, kind='stable').reset_index()
# Instead of explicitly 'swapping' we just swap the 'display_rank' - where
# in the final ranking this would be placed. We can easily use that to compute DCG
query_judgments['display_rank'] = query_judgments.index.to_series()
query_judgments['train_dcg'] = query_judgments['dcg'] = metric(query_judgments, at=at)
train_dcg = query_judgments.loc[0, 'dcg']
qid = query_judgments.loc[0, 'qid']
keywords = query_judgments.loc[0, 'keywords']
query_judgments['lambda'] = 0.0
query_judgments['weight'] = 0.0
for better in range(0,len(query_judgments)):
for worse in range(0,len(query_judgments)):
if better > at and worse > at:
return query_judgments
if query_judgments.loc[better, 'grade'] > query_judgments.loc[worse, 'grade']:
query_judgments = rank_with_swap(query_judgments, better, worse)
query_judgments['dcg'] = metric(query_judgments, at=at)
dcg_after_swap = query_judgments.loc[0, 'dcg']
delta = abs(train_dcg - dcg_after_swap)
if delta != 0.0:
last_model_score_diff = query_judgments.loc[better, 'last_prediction'] - query_judgments.loc[worse, 'last_prediction']
rho = 1.0 / (1.0 + exp(last_model_score_diff))
assert(delta >= 0.0)
assert(rho >= 0.0)
query_judgments.loc[better, 'lambda'] += delta * rho
query_judgments.loc[worse, 'lambda'] -= delta * rho
# --------------
# NEW!
# last_model_score_diff rho weight
# 0.0 0.5 0.25 (max possible value)
# 100.0 0.0000 0.0 (max possible value)
#
# If the current model has an ambiguous prediction, we include more of the delta in the weight
# If the current model has a strong prediction, weight approaches 0
query_judgments.loc[better, 'weight'] += rho * (1.0 - rho) * delta;
query_judgments.loc[worse, 'weight'] += rho * (1.0 - rho) * delta;
#
# These will be used to rescale each decision tree node's predictions
# If many results in a leaf node have last model score ~ ambiguous
# the resulting model will have a high denominator ~ (1 / deltaDCG)
# If many results in a leaf node have last model score - not ambiguous, positive
# the resulting model will have a low denominator
#
# Apparently we want to cancel out the deltas if last model was ambiguous?
# ---------------
return query_judgments
# Convert to Pandas Dataframe
judgments = judgments_to_dataframe(ftr_logger.logged, unnest=False)
judgments['last_prediction'] = 0
lambdas_per_query = judgments.groupby('qid').apply(compute_swaps_scaled_with_weights, axis=1)
lambdas_per_query = lambdas_per_query.drop('qid', axis=1).reset_index().drop(['level_1', 'index'], axis=1)
lambdas_per_query
# +
from sklearn.tree import DecisionTreeRegressor
train_set = lambdas_per_query[['lambda', 'features']]
train_set
tree3 = DecisionTreeRegressor(max_leaf_nodes=4)
tree3.fit(train_set['features'].tolist(), train_set['lambda'])
# -
# ### Label each row with its unique prediction (ie tree path)
# +
def tree_paths(tree, X):
paths_as_array = tree.decision_path(X).toarray()
paths = ["".join(item) for item in paths_as_array.astype(str)]
return paths
lambdas_per_query['path'] = tree_paths(tree3, train_set['features'].tolist())
# -
# ### Override outputs using our own weighted average
#
# The typical decision tree uses either the [median or mean of the target values](https://scikit-learn.org/stable/modules/tree.html#regression-criteria) classified to a given leaf node as the prediction. However, in the case of lambdaMART, we want to use a weighted average that accounts for how much of the DCG error out there has been accounted for. Thus the psuedoresponses are summed and divided by the remaining error DCG.
#
# rho=0, then an example is weighed by `1/0.25*deltaNDCG` as there's a lot of outstanding DCG error left.
lambdas_per_query.groupby('path')['weight'].sum()
lambdas_per_query.groupby('path')['lambda'].sum()
round_predictions = lambdas_per_query.groupby('path')['lambda'].sum() / lambdas_per_query.groupby('path')['weight'].sum()
round_predictions.to_dict()
plot_tree(tree3)
# ### Wrap the original tree, and provide new value
#
# Instead of directly using the provided decision tree, we want to use our prediction for each leaf. This `OverridenDecisionTree` takes the original tree, looks up the prediction path in the original tree, but uses our values for the predicted variable instead of what's here.
# +
class OverridenRegressionTree:
def __init__(self, predictions, tree):
self.predictions = predictions
self.tree = tree
def predict(self, X, use_original=False):
if use_original:
return self.predict(X)
path = self.tree.decision_path(X).toarray().astype(str)
path = "".join(path[0])
paths_as_array = self.tree.decision_path(X).toarray()
paths = ["".join(item) for item in paths_as_array.astype(str)]
predictions = self.predictions[paths]
# Any NaN predictions is a red flag, debug
if np.any(predictions.isnull()):
print(predictions[predictions.isnull()])
print(pd.DataFrame(X)[predictions.isnull().reset_index(drop=True)])
raise AssertionError("No prediction should be NaN")
return np.array(self.predictions[paths].tolist())
override_tree = OverridenRegressionTree(predictions = round_predictions, tree=tree3)
override_tree.predict([[0.0, 6.869545], [10.0, 10.0]])
# -
# ## Part Four - Putting it all together, from the top!
#
# Now we can put together the full lambdamart algorithm that
#
# 1. Uses pair-wise swaps on our our metric (ie DCG) to generate decision tree predictors (the 'lambdas')
# 2. Focuses in on predicting where current model makes the wrong call when ranking by DCG
# 3. Predicts using a weighted average, weighed by 1 / (remaining DCG)
#
# TODOs / known issues
# * While DCG converges, it does sometimes wander a tad, so there might be more room for improvement
# * Speeding up the inner loop of the `compute_swaps_scaled_with_weights` that must run for ever query's swaps
# +
from sklearn.tree import DecisionTreeRegressor
import pandas as pd
def predict(ensemble, X, learning_rate=0.1):
prediction = 0
for tree in ensemble:
prediction += tree.predict(X) * learning_rate
return prediction.rename('prediction')
def tree_paths(tree, X):
paths_as_array = tree.decision_path(X).toarray()
paths = ["".join(item) for item in paths_as_array.astype(str)]
return paths
ensemble=[]
def lambda_mart(judgments, rounds=20, learning_rate=0.1, max_leaf_nodes=8, metric=dcg):
print(judgments.columns)
# Convert to Pandas Dataframe
lambdas_per_query = judgments.copy()
lambdas_per_query['last_prediction'] = 0.0
for i in range(0, rounds):
print(f"round {i}")
# ------------------
#1. Build pair-wise predictors for this round
lambdas_per_query = lambdas_per_query.groupby('qid').apply(compute_swaps_scaled_with_weights,
axis=1, metric=dcg)
# ------------------
#2. Train a regression tree on this round's lambdas
features = lambdas_per_query['features'].tolist()
tree = DecisionTreeRegressor(max_leaf_nodes=max_leaf_nodes)
tree.fit(features, lambdas_per_query['lambda'])
# ------------------
#3. Reweight based on LambdaMART's weighted average
# Add each tree's paths
lambdas_per_query['path'] = tree_paths(tree, features)
predictions = lambdas_per_query.groupby('path')['lambda'].sum() / lambdas_per_query.groupby('path')['weight'].sum()
predictions = predictions.fillna(0.0) # for divide by 0
# -------------------
#4. Add to ensemble, recreate last prediction
new_tree = OverridenRegressionTree(predictions=predictions, tree=tree)
ensemble.append(new_tree)
next_predictions = new_tree.predict(features)
lambdas_per_query['last_prediction'] += (next_predictions * learning_rate)
print("Train DCGs")
print("mean ", lambdas_per_query['train_dcg'].mean())
print("median ", lambdas_per_query['train_dcg'].median())
print("----------")
# Reset the dataframe for further processing
lambdas_per_query = lambdas_per_query.drop('qid', axis=1).reset_index().drop(['level_1', 'index'], axis=1)
judgments = judgments_to_dataframe(ftr_logger.logged, unnest=False)
lambdas_per_query = lambda_mart(judgments=judgments, rounds=50, max_leaf_nodes=10, learning_rate=0.1, metric=dcg)
# -
lambdas_per_query.iloc[282]
ensemble[0].predictions
plot_tree(ensemble[0].tree)
# ## Compare to ranklib output
# +
from ltr.ranklib import train
trainLog = train(client,
training_set=ftr_logger.logged,
index='tmdb',
trees=10,
featureSet='movies',
modelName='title')
print("Every N rounds of ranklib")
trainLog.trainingLogs[0].rounds
# -
# ## Examine queries we learned
#
# Try out some queries, look at the final model prediction `last_prediction` compare to the correct ordering `grade`.
lambdas_per_query[lambdas_per_query['qid'] == 2]
# # 5. Pure Pandas Implementation?
#
# Can we make it faster by vectorizing with pandas?
#
# Turns out Yes!
# +
lambdas_per_query = judgments.copy()
lambdas_per_query['last_prediction'] = 0.0
lambdas_per_query.sort_values(['qid', 'last_prediction'], ascending=[True, False], kind='stable')
# +
lambdas_per_query.sort_values(['qid', 'last_prediction'], ascending=[True, False], kind='stable')
lambdas_per_query['display_rank'] = lambdas_per_query.groupby('qid').cumcount()
#TBD - How do generalize this?
lambdas_per_query['discount'] = 1 / np.log2(2 + lambdas_per_query['display_rank'])
lambdas_per_query['gain'] = (2**lambdas_per_query['grade'] - 1) # * lambdas_per_query['discount']
lambdas_per_query[['qid', 'display_rank', 'discount', 'grade', 'gain']]
# -
# ## Pairwise deltas
#
# Delta captures pair-wise difference of the ranking metric (ie DCG)
#
# each group paired with each other group
swaps = lambdas_per_query.merge(lambdas_per_query, on='qid', how='outer')
# changes[j][i] = changes[i][j] = (discount(i) - discount(j)) * (gain(rel[i]) - gain(rel[j]));
swaps['delta'] = np.abs((swaps['discount_x'] - swaps['discount_y']) * (swaps['gain_x'] - swaps['gain_y']))
swaps[['qid', 'display_rank_x', 'display_rank_y', 'delta']]
# ## Pairwise rhos
#
# Rho captures pair-wise difference of the current model's prediction.
#
swaps['rho'] = 1 / (1 + np.exp(swaps['last_prediction_x'] - swaps['last_prediction_y']))
swaps[['qid', 'display_rank_x', 'display_rank_y', 'delta', 'last_prediction_x', 'last_prediction_y', 'rho']]
# ## Compute lambdas
#
# For every row where grade_x > grade_y, compute `delta*rho`
swaps['lambda'] = 0
slice_x_better =swaps[swaps['grade_x'] > swaps['grade_y']]
swaps.loc[swaps['grade_x'] > swaps['grade_y'], 'lambda'] = slice_x_better['delta'] * slice_x_better['rho']
swaps[['qid', 'display_rank_x', 'display_rank_y', 'delta', 'last_prediction_x', 'last_prediction_y', 'rho', 'lambda']]
# ## Get per-key lambdas
#
# We merge back together the xs minuse the ys
# Better minus worse
lambdas_x = swaps.groupby(['qid', 'display_rank_x'])['lambda'].sum().rename('lambda')
lambdas_y = swaps.groupby(['qid', 'display_rank_y'])['lambda'].sum().rename('lambda')
lambdas = lambdas_x - lambdas_y
lambdas
lambdas_per_query = lambdas_per_query.merge(lambdas, left_on=['qid', 'display_rank'], right_on=['qid', 'display_rank_x'], how='left')
lambdas_per_query[['qid', 'docId', 'grade', 'features', 'lambda']]
lambdas_per_query.merge(lambdas, left_on=['qid', 'display_rank'], right_on=['qid', 'display_rank_x'], how='left')
# +
#2. Train a regression tree on this round's lambdas
features = lambdas_per_query['features'].tolist()
tree = DecisionTreeRegressor(max_leaf_nodes=10)
tree.fit(features, lambdas_per_query['lambda'])
tree
# -
ensemble.append(tree)
def compute_lambdas(lambdas_per_query):
lambdas_per_query = lambdas_per_query.sort_values(['qid', 'last_prediction'], ascending=[True, False], kind='stable')
lambdas_per_query['display_rank'] = lambdas_per_query.groupby('qid').cumcount()
#TBD - How do generalize this to any metric?
lambdas_per_query['discount'] = 1 / np.log2(2 + lambdas_per_query['display_rank'])
lambdas_per_query['gain'] = (2**lambdas_per_query['grade'] - 1)
# swaps dataframe holds each pair-wise swap computed (shrink columns for memory?)
# Optimization of swaps = lambdas_per_query.merge(lambdas_per_query, on='qid', how='outer')
# to limit to just needed columns
to_swap = lambdas_per_query[['qid', 'display_rank', 'grade', 'last_prediction', 'discount', 'gain']]
#to_swap = lambdas_per_query
swaps = to_swap.merge(to_swap, on='qid', how='outer')
# delta - delta in DCG due to swap
swaps['delta'] = np.abs((swaps['discount_x'] - swaps['discount_y']) * (swaps['gain_x'] - swaps['gain_y']))
# rho - based on current model prediction delta
swaps['rho'] = 1 / (1 + np.exp(swaps['last_prediction_x'] - swaps['last_prediction_y']))
# If you want to be pure gradient boosting, weight reweights each models prediction
# I haven't found this to matter in practice
swaps['weight'] = swaps['rho'] * (1.0 - swaps['rho']) * swaps['delta']
# Compute lambdas (the next model in ensemble's predictors) when grade_x > grade_y
swaps['lambda'] = 0
slice_x_better =swaps[swaps['grade_x'] > swaps['grade_y']]
swaps.loc[swaps['grade_x'] > swaps['grade_y'], 'lambda'] = slice_x_better['delta'] * slice_x_better['rho']
# accumulate lambdas and add back to model
lambdas_x = swaps.groupby(['qid', 'display_rank_x'])['lambda'].sum().rename('lambda')
lambdas_y = swaps.groupby(['qid', 'display_rank_y'])['lambda'].sum().rename('lambda')
weights_x = swaps.groupby(['qid', 'display_rank_x'])['weight'].sum().rename('weight')
weights_y = swaps.groupby(['qid', 'display_rank_y'])['weight'].sum().rename('weight')
weights = weights_x + weights_y
lambdas = lambdas_x - lambdas_y
lambdas_per_query = lambdas_per_query.merge(lambdas,
left_on=['qid', 'display_rank'],
right_on=['qid', 'display_rank_x'],
how='left')
lambdas_per_query = lambdas_per_query.merge(weights,
left_on=['qid', 'display_rank'],
right_on=['qid', 'display_rank_x'],
how='left')
return lambdas_per_query
# +
from sklearn.tree import DecisionTreeRegressor
import pandas as pd
ensemble=[]
def lambda_mart_pure(judgments, rounds=20, learning_rate=0.1, max_leaf_nodes=8, metric=dcg):
print(judgments.columns)
# Convert to Pandas Dataframe
lambdas_per_query = judgments.copy()
lambdas_per_query['last_prediction'] = 0.0
for i in range(0, rounds):
print(f"round {i}")
# ------------------
#1. Build pair-wise predictors for this round
lambdas_per_query = compute_lambdas(lambdas_per_query)
# ------------------
#2. Train a regression tree on this round's lambdas
features = lambdas_per_query['features'].tolist()
tree = DecisionTreeRegressor(max_leaf_nodes=max_leaf_nodes)
tree.fit(features, lambdas_per_query['lambda'])
# ------------------
#3. Reweight based on LambdaMART's weighted average
# Add each tree's paths
lambdas_per_query['path'] = tree_paths(tree, features)
predictions = lambdas_per_query.groupby('path')['lambda'].sum() / lambdas_per_query.groupby('path')['weight'].sum()
predictions = predictions.fillna(0.0) # for divide by 0
# -------------------
#4. Add to ensemble, recreate last prediction
new_tree = OverridenRegressionTree(predictions=predictions, tree=tree)
ensemble.append(new_tree)
next_predictions = new_tree.predict(features)
lambdas_per_query['last_prediction'] += (next_predictions * learning_rate)
print(lambdas_per_query.loc[0, ['grade', 'last_prediction']])
print("Train DCGs")
lambdas_per_query['discounted_gain'] = lambdas_per_query['gain'] * lambdas_per_query['discount']
dcg = lambdas_per_query[lambdas_per_query['display_rank'] < 10].groupby('qid')['discounted_gain'].sum().mean()
print("mean ", dcg)
print("----------")
lambdas_per_query = lambdas_per_query.drop(['lambda', 'weight'], axis=1)
return lambdas_per_query
judgments = judgments_to_dataframe(ftr_logger.logged, unnest=False)
lambdas_per_query = lambda_mart_pure(judgments=judgments, rounds=50, max_leaf_nodes=10, learning_rate=0.01, metric=dcg)
# -
judgments = judgments_to_dataframe(ftr_logger.logged, unnest=False)
# %prun -s cumtime lambdas_per_query = lambda_mart_pure(judgments=judgments, rounds=50, max_leaf_nodes=10, learning_rate=0.01, metric=dcg)
# +
from ltr.ranklib import train
# %prun -s cumtime trainLog = train(client, training_set=ftr_logger.logged, index='tmdb', trees=10, featureSet='movies', modelName='title')
print("Every N rounds of ranklib")
trainLog.trainingLogs[0].rounds
# -
class OverridenRegressionTree:
def __init__(self, predictions, tree):
self.predictions = predictions
self.tree = tree
def predict(self, X, use_original=False):
if use_original:
return self.predict(X)
path = self.tree.decision_path(X).toarray().astype(str)
path = "".join(path[0])
paths_as_array = self.tree.decision_path(X).toarray()
paths = ["".join(item) for item in paths_as_array.astype(str)]
predictions = self.predictions[paths]
# Any NaN predictions is a red flag, debug
if np.any(predictions.isnull()):
print(predictions[predictions.isnull()])
print(pd.DataFrame(X)[predictions.isnull().reset_index(drop=True)])
raise AssertionError("No prediction should be NaN")
return np.array(self.predictions[paths].tolist())
| notebooks/elasticsearch/tmdb/lambda-mart-in-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # jQueryUI tests
#
# These tests verify that JQueryUI loads with CSS styling when requested.
# + deletable=true editable=true
from jp_proxy_widget import notebook_test_helpers
validators = notebook_test_helpers.ValidationSuite()
# + deletable=true editable=true
import jp_proxy_widget
from jp_proxy_widget import js_context
class JQueryUIDialogTestWidget(jp_proxy_widget.JSProxyWidget):
"A jQueryUI dialog with custom styling that reports back to Python when it is closed."
def __init__(self, *pargs, **kwargs):
super(JQueryUIDialogTestWidget, self).__init__(*pargs, **kwargs)
self.is_open = "unknown"
self.check_jquery()
random_style = """
.dialog-random-style-for-testing2 {
color: rgb(216, 50, 61);
background-color: rgb(200, 218, 61);
}
"""
self.load_css_text("dialog-random-stylesheet", random_style)
self.js_init("""
// Add dialog as a child to the widget element.
element.empty();
var dialog = $('<div><b class="dialog-random-style-for-testing2">Hello from jQueryUI</b></div>')
.appendTo(element);
// Add some callables to the element accessible from Python.
element.report_status = function() {
// Report whether the dialog is now open back to Python.
is_open_callback(dialog.dialog("isOpen"))
};
element.close = function () {
// Close the dialog.
dialog.dialog("close");
};
// Open the dialog and have it report its open status when it's closed.
dialog.dialog({close: element.report_status});
element.report_status();
""", is_open_callback=self.is_open_callback)
def is_open_callback(self, value):
"Python callback which remembers whether the dialog reported itself open or not."
self.is_open = value
# + deletable=true editable=true
# Create and display the dialog.
test_dialog = JQueryUIDialogTestWidget()
test_dialog
# + deletable=true editable=true
# NOTE: Click here to focus on the cell (otherwise "enter" with the dialog focused will close the dialog).
test_dialog.is_open
# + deletable=true editable=true
# Close the dialog by calling the element.close() method.
test_dialog.element.close()
# + deletable=true editable=true
def validate_test_dialog():
assert test_dialog.is_open == False
print('test_dialog apparently ok!')
validators.add_validation(test_dialog, validate_test_dialog)
# + deletable=true editable=true
test_slider_css = """
#test-slider-custom-handle {
width: 3em;
height: 1.6em;
top: 50%;
margin-top: -.8em;
text-align: center;
line-height: 1.6em;
}
#test-slider-container {
width: 50em;
height: 10em;
}
"""
class JQueryUISliderTestWidget(jp_proxy_widget.JSProxyWidget):
"A slider adapted from http://jqueryui.com/slider/#custom-handle"
def __init__(self, *pargs, **kwargs):
super(JQueryUISliderTestWidget, self).__init__(*pargs, **kwargs)
self.slider_value = "unknown"
self.load_css_text("test-slider-stylesheet", test_slider_css)
self.js_init("""
element.html('<div class="test-slider-container"></div>');
var slider_div = $("<div></div>").appendTo(element);
var handle = $('<div class="test-slider-custom-handle ui-slider-handle"></div>')
.width("30px")
.appendTo(slider_div);
slider_div.slider({
min: -50,
max: 65,
value: 43,
create: function() {
//handle.text( element.slider( "value" ) ); // causes error
},
slide: function( event, ui ) {
handle.text( ui.value );
},
change: function( event, ui ) {
handle.text( ui.value );
report_change( ui.value );
}
});
element.height("50px").width("300px").css("background-color", "cornsilk");
slider_div.width("250px");
slider_div.position({
my: "center",
at: "center",
of: element
});
element.set_value = function(value) {
slider_div.slider("value", value);
};
element.set_value(44);
""", report_change=self.report_change)
def report_change(self, value):
self.slider_value = value
# + deletable=true editable=true
slider_widget = JQueryUISliderTestWidget()
slider_widget
# + deletable=true editable=true
slider_widget.slider_value
#slider_widget.print_status()
# + deletable=true editable=true
slider_widget.element.set_value(-15)
def validate_slider_widget():
assert slider_widget.slider_value == -15
print('slider_widget apparently ok!')
validators.add_validation(slider_widget, validate_slider_widget)
# + deletable=true editable=true
# + deletable=true editable=true
# This should be the last cell
validators.run_all_in_widget()
| notebooks/notebook_tests/jqueryui test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3. Model tests
# ## 3.04 Example: GradientBoost
# Process outline
# 1. Prepare time delay features
# 2. Spatial aggregation
# - Area weighted sum for precipitation features
# - Area mean for all other features
# 3. Define the model
# 4. Train the model
# - Reshape the input dataset to the model specific `X` and `y` arrays
# - Actually train on `X` and `y`
# +
import sys
sys.path.append("../../")
print(sys.executable)
import numpy as np
import datetime as dt
import pandas as pd
import matplotlib.pyplot as plt
import dask
dask.config.set(scheduler='threads')
import xarray as xr
from python.aux.utils_floodmodel import add_time, generate_prediction_array, remove_outlier, multi_forecast_case_study
from python.aux.plot import plot_multif_prediction
import joblib
from sklearn.pipeline import Pipeline
from dask_ml.preprocessing import StandardScaler
from dask_ml.decomposition import PCA
from sklearn.ensemble import GradientBoostingRegressor
import matplotlib
matplotlib.rcParams.update({'font.size': 14})
# -
# ## Loading the data
# Sample dataset contained in the git repository.
#
# As you are reading these lines, you opened the notebook in the `./docs/` folder of the main repository directory. To access the sample dataset that was delivered to you with the code, step outside the current directory (`../`) and enter the `data/` folder. The names of the sample datasets are `smallsampledata-era5.nc` and `smallsampledata-glofas.nc`, both in netCDF format, a user-friendly format that keeps the file size low and stores meta-data within the file.
#
# We use `xarray` to access the files, as it provides us with a very powerful interface to work with the data.
#
# The features are already prepared and ready to use, hence why they can be loaded directly from the `features_xy.nc` file.
#
# Let's open the files and see what's in there:
# load data
features = xr.open_dataset('../../data/features_xy.nc')
y = features['dis']
X = features.drop(['dis', 'dis_diff'])
features
# Some control parameters for the procedure:
#
# * `add_shift_switch`: Shift and add shifted values as new features for multi-day forecasts. (only needed if forecasting multiple days at once => currently only implemented with nns!)
# * `dis_shift_1`: Add the discharge from the last day as additional feature for testing purposes.
# * `abs_vals_switch`: Model is trained on absolute discharge values if True and on differences/change in discharge else.
dis_shift_switch = False
abs_vals_switch = False
# Begin with some short preprocessing..
if dis_shift_switch:
dis_shift_1 = y.shift(time=1)
X = X.assign({'dis-1': dis_shift_1})
X_base = X.to_array(dim='features').T.copy()
y_base = y.copy()
# Next, we split the dataset into three parts for
# - training (the longest period for estimating the free parameters within the model),
# - validation (for tuning hyperparameters) and
# - testing (for the model comparison).
period_train = dict(time=slice(None, '2005'))
period_valid = dict(time=slice('2006', '2011'))
period_test = dict(time=slice('2012', '2016'))
X_train, y_train = X_base.loc[period_train], y_base.loc[period_train]
X_valid, y_valid = X_base.loc[period_valid], y_base.loc[period_valid]
X_test, y_test = X_base.loc[period_test], y_base.loc[period_test]
X_train.shape, y_train.shape
# +
time = y_train.time
Xda = X_train.chunk(dict(time=-1, features=-1)).dropna(dim='time').to_pandas()
if abs_vals_switch:
# train on absolute values
yda = y_train.to_pandas().loc[Xda.index]
# removing outlier and removing same parts from X
yda = remove_outlier(yda)
Xda = Xda.loc[yda.index]
else:
# train on change in discharge values
yda = y_train.diff(dim='time').to_pandas().loc[Xda.index]
# removing outlier and removing same parts from X
yda = remove_outlier(yda)
Xda = Xda.loc[yda.index]
# -
# # Define boosting regressor
# +
model = GradientBoostingRegressor(n_estimators=200,
learning_rate=0.1,
max_depth=5,
random_state=0,
# loss='ls'
)
pipe = Pipeline([('scaler', StandardScaler()),
#('pca', PCA(n_components=6)),
('model', model),], verbose=True)
X_fit = Xda.copy()
y_fit = yda.copy()
pipe.fit(X_fit, y_fit)
# -
# ### in sample test for the predicted change, to verify functionality
Xda_insample = Xda.copy()
insample_check = pipe.predict(Xda_insample)
insample_check = add_time(insample_check, Xda.index, name='forecast')
insample_check.to_pandas().plot(linewidth=0.5)
yda.plot(linestyle='--', linewidth=0.5)
# Set the frequency at which forecast are started.
# prediction start from every nth day
# if in doubt, leave n = 1 !!!
n = 1
X_pred = X_valid[::n].copy()
y_pred = pipe.predict(X_pred)
y_pred = add_time(y_pred, X_pred.time, name='forecast')
print(y_pred)
import matplotlib
matplotlib.rcParams.update({'font.size': 14})
multif = generate_prediction_array(y_pred, y, forecast_range=14)
plot_multif_prediction(multif, y, forecast_range=14, title='14-day forecast - Validation period - GradientBoostingRegressor');
plt.savefig('validation_period_gradboost.png', dpi=600, bbox_inches='tight')
# #### Check the skill of a 14-day persistence forecast:
# +
forecast_range = 14
y_o_pers = y_valid
# persistence
y_m_pers = y_valid.copy()
for i in range(1, forecast_range):
y_m_pers.loc[y_valid.time[i::forecast_range]] = y_valid.shift(time=i)[i::forecast_range].values
rmse = np.sqrt(np.nanmean((y_m_pers - y_o_pers)**2))
nse = 1 - np.sum((y_m_pers - y_o_pers)**2)/(np.sum((y_o_pers - np.nanmean(y_o_pers))**2))
print(f"Persistence {forecast_range}-day forecast: RMSE={round(float(rmse), 2)}; NSE={round(float(nse.values), 2)}")
# -
# Now, lets run a few different settings and compare the results:
# | n_estimators | learning_rate | max_depth | RMSE | NSE |
# | :-----: | :-----: | :-----: | :-----: | :-----: |
# | 10 | 0.05 | 1 | 197.41 | 0.8
# | 10 | 0.05 | 2 | 191.4 | 0.81
# | 10 | 0.05 | 3 | 186.96 | 0.82
# | 10 | 0.05 | 4 | 182.56 | 0.83
# | 10 | 0.05 | 5 | 179.38 | 0.83
# | 10 | 0.05 | 6 | 177.28 | 0.84
# | 10 | 0.05 | 7 | 174.66 | 0.84
# | 10 | 0.05 | 8 | 174.55 | 0.84
# | 10 | 0.05 | 9 | 172.26 | 0.85
# | 10 | 0.05 | 10 | 172.66 | 0.84
# | 10 | 0.05 | 15 | 172.49 | 0.85
# | 10 | 0.1 | 1 | 192.42 | 0.81
# | 10 | 0.1 | 2 | 183.21 | 0.83
# | 10 | 0.1 | 3 | 176.93 | 0.84
# | 10 | 0.1 | 4 | 170.17 | 0.85
# | 10 | 0.1 | 5 | 164.83 | 0.86
# | 10 | 0.1 | 6 | 162.34 | 0.86
# | 10 | 0.1 | 7 | 160.55 | 0.87
# | 10 | 0.1 | 8 | 158.55 | 0.87
# | 10 | 0.1 | 9 | 155.32 | 0.87
# | 10 | 0.1 | 10 | 158.16 | 0.87
# | 10 | 0.1 | 15 | 159.2 | 0.87
# | 10 | 0.2 | 1 | 184.54 | 0.82
# | 10 | 0.2 | 2 | 173.02 | 0.84
# | 10 | 0.2 | 3 | 165.05 | 0.86
# | 10 | 0.2 | 4 | 156.02 | 0.87
# | 10 | 0.2 | 5 | 148.13 | 0.89
# | 10 | 0.2 | 6 | 147.61 | 0.89
# | 10 | 0.2 | 7 | 147.86 | 0.89
# | 10 | 0.2 | 8 | 149.23 | 0.88
# | 10 | 0.2 | 9 | 147.06 | 0.89
# | 10 | 0.2 | 10 | 148.5 | 0.89
# | 10 | 0.2 | 15 | 156.1 | 0.87
# | 50 | 0.05 | 1 | 183.64 | 0.82
# | 50 | 0.05 | 2 | 170.91 | 0.85
# | 50 | 0.05 | 3 | 160.38 | 0.87
# | 50 | 0.05 | 4 | 151.42 | 0.88
# | 50 | 0.05 | 5 | 148.28 | 0.89
# | 50 | 0.05 | 6 | 143.87 | 0.89
# | 50 | 0.05 | 7 | 144.23 | 0.89
# | 50 | 0.05 | 8 | 144.34 | 0.89
# | 50 | 0.05 | 9 | 143.31 | 0.89
# | 50 | 0.05 | 10 | 147.52 | 0.89
# | 50 | 0.05 | 15 | 154.18 | 0.88
# | 50 | 0.1 | 1 | 174.74 | 0.84
# | 50 | 0.1 | 2 | 159.71 | 0.87
# | 50 | 0.1 | 3 | 149.85 | 0.88
# | 50 | 0.1 | 4 | 144.3 | 0.89
# | 50 | 0.1 | 5 | 138.97 | 0.9
# | 50 | 0.1 | 6 | 137.74 | 0.9
# | 50 | 0.1 | 7 | 140.0 | 0.9
# | 50 | 0.1 | 8 | 139.16 | 0.9
# | 50 | 0.1 | 9 | 137.26 | 0.9
# | 50 | 0.1 | 10 | 143.42 | 0.89
# | 50 | 0.1 | 15 | 150.03 | 0.88
# | 50 | 0.2 | 1 | 164.16 | 0.86
# | 50 | 0.2 | 2 | 151.27 | 0.88
# | 50 | 0.2 | 3 | 142.9 | 0.89
# | 50 | 0.2 | 4 | 138.33 | 0.9
# | 50 | 0.2 | 5 | 134.38 | 0.91
# | 50 | 0.2 | 6 | 135.02 | 0.91
# | 50 | 0.2 | 7 | 140.31 | 0.9
# | 50 | 0.2 | 8 | 141.95 | 0.9
# | 50 | 0.2 | 9 | 141.16 | 0.9
# | 50 | 0.2 | 10 | 146.41 | 0.89
# | 50 | 0.2 | 15 | 154.09 | 0.88
# | 100 | 0.05 | 1 | 174.64 | 0.84
# | 100 | 0.05 | 2 | 159.49 | 0.87
# | 100 | 0.05 | 3 | 149.45 | 0.88
# | 100 | 0.05 | 4 | 143.9 | 0.89
# | 100 | 0.05 | 5 | 142.6 | 0.89
# | 100 | 0.05 | 6 | 138.71 | 0.9
# | 100 | 0.05 | 7 | 138.71 | 0.9
# | 100 | 0.05 | 8 | 138.32 | 0.9
# | 100 | 0.05 | 9 | 138.41 | 0.9
# | 100 | 0.05 | 10 | 144.13 | 0.89
# | 100 | 0.05 | 15 | 153.26 | 0.88
# | 100 | 0.1 | 1 | 164.5 | 0.86
# | 100 | 0.1 | 2 | 151.08 | 0.88
# | 100 | 0.1 | 3 | 144.4 | 0.89
# | 100 | 0.1 | 4 | 138.96 | 0.9
# | 100 | 0.1 | 5 | 134.78 | 0.91
# | 100 | 0.1 | 6 | 136.09 | 0.9
# | 100 | 0.1 | 7 | 138.78 | 0.9
# | 100 | 0.1 | 8 | 139.74 | 0.9
# | 100 | 0.1 | 9 | 137.79 | 0.9
# | 100 | 0.1 | 10 | 143.69 | 0.89
# | 100 | 0.1 | 15 | 149.92 | 0.88
# | 100 | 0.2 | 1 | 157.8 | 0.87
# | 100 | 0.2 | 2 | 143.58 | 0.89
# | 100 | 0.2 | 3 | 139.44 | 0.9
# | 100 | 0.2 | 4 | 134.87 | 0.91
# | 100 | 0.2 | 5 | 133.99 | 0.91
# | 100 | 0.2 | 6 | 137.64 | 0.9
# | 100 | 0.2 | 7 | 140.78 | 0.9
# | 100 | 0.2 | 8 | 143.56 | 0.89
# | 100 | 0.2 | 9 | 142.34 | 0.89
# | 100 | 0.2 | 10 | 146.95 | 0.89
# | 100 | 0.2 | 15 | 154.09 | 0.88
# | 200 | 0.05 | 1 | 164.87 | 0.86
# | 200 | 0.05 | 2 | 152.16 | 0.88
# | 200 | 0.05 | 3 | 142.79 | 0.89
# | 200 | 0.05 | 4 | 138.14 | 0.9
# | 200 | 0.05 | 5 | 138.43 | 0.9
# | 200 | 0.05 | 6 | 136.32 | 0.9
# | 200 | 0.05 | 7 | 137.88 | 0.9
# | 200 | 0.05 | 8 | 138.56 | 0.9
# | 200 | 0.05 | 9 | 139.12 | 0.9
# | 200 | 0.05 | 10 | 144.56 | 0.89
# | 200 | 0.05 | 15 | 153.25 | 0.88
# | 200 | 0.1 | 1 | 157.82 | 0.87
# | 200 | 0.1 | 2 | 144.66 | 0.89
# | 200 | 0.1 | 3 | 141.18 | 0.9
# | 200 | 0.1 | 4 | 137.15 | 0.9
# | 200 | 0.1 | 5 | 133.8 | 0.91
# | 200 | 0.1 | 6 | 138.13 | 0.9
# | 200 | 0.1 | 7 | 139.21 | 0.9
# | 200 | 0.1 | 8 | 141.52 | 0.9
# | 200 | 0.1 | 9 | 138.44 | 0.9
# | 200 | 0.1 | 10 | 144.09 | 0.89
# | 200 | 0.1 | 15 | 149.92 | 0.88
# | 200 | 0.2 | 1 | 153.71 | 0.88
# | 200 | 0.2 | 2 | 138.04 | 0.9
# | 200 | 0.2 | 3 | 136.03 | 0.9
# | 200 | 0.2 | 4 | 135.76 | 0.9
# | 200 | 0.2 | 5 | 134.93 | 0.91
# | 200 | 0.2 | 6 | 138.36 | 0.9
# | 200 | 0.2 | 7 | 142.07 | 0.9
# | 200 | 0.2 | 8 | 144.82 | 0.89
# | 200 | 0.2 | 9 | 141.88 | 0.9
# | 200 | 0.2 | 10 | 146.97 | 0.89
# | 200 | 0.2 | 15 | 154.09 | 0.88
# | 300 | 0.05 | 1 | 160.46 | 0.87
# | 300 | 0.05 | 2 | 149.26 | 0.88
# | 300 | 0.05 | 3 | 139.02 | 0.9
# | 300 | 0.05 | 4 | 135.69 | 0.9
# | 300 | 0.05 | 5 | 137.48 | 0.9
# | 300 | 0.05 | 6 | 136.74 | 0.9
# | 300 | 0.05 | 7 | 138.85 | 0.9
# | 300 | 0.05 | 8 | 139.89 | 0.9
# | 300 | 0.05 | 9 | 139.9 | 0.9
# | 300 | 0.05 | 10 | 145.1 | 0.89
# | 300 | 0.05 | 15 | 153.26 | 0.88
# | 300 | 0.1 | 1 | 155.08 | 0.87
# | 300 | 0.1 | 2 | 141.9 | 0.9
# | 300 | 0.1 | 3 | 139.95 | 0.9
# | 300 | 0.1 | 4 | 137.32 | 0.9
# | 300 | 0.1 | 5 | 134.8 | 0.91
# | 300 | 0.1 | 6 | 138.89 | 0.9
# | 300 | 0.1 | 7 | 139.32 | 0.9
# | 300 | 0.1 | 8 | 142.71 | 0.89
# | 300 | 0.1 | 9 | 138.2 | 0.9
# | 300 | 0.1 | 10 | 144.24 | 0.89
# | 300 | 0.1 | 15 | 149.92 | 0.88
# | 300 | 0.2 | 1 | 151.95 | 0.88
# | 300 | 0.2 | 2 | 136.58 | 0.9
# | 300 | 0.2 | 3 | 135.88 | 0.9
# | 300 | 0.2 | 4 | 136.42 | 0.9
# | 300 | 0.2 | 5 | 135.36 | 0.9
# | 300 | 0.2 | 6 | 139.76 | 0.9
# | 300 | 0.2 | 7 | 142.15 | 0.89
# | 300 | 0.2 | 8 | 144.55 | 0.89
# | 300 | 0.2 | 9 | 141.74 | 0.9
# | 300 | 0.2 | 10 | 147.04 | 0.89
# | 300 | 0.2 | 15 | 154.09 | 0.88
# | --------------- | --------------- | --------------- | --------------- | --------------- |
# | persistence 14-day forecast | | | 251.09 | 0.77 |
#
#
#
# Concluding, the best setting is `n_estimators=200`, `learning_rate=0.1` and `max_depth=5`, although, as already mentioned, the differences between on the upper end is quite marginal and hence, is not that critical to small deviations. As can be seen, the model is baseline better than the persistence, no matter the setting, which shows that there is at least some skill in using it. It should also be noted, that the persistence forecast for such an application is in general not that bad, because absolute values of discharge are validated and the day-to-day change is usually one magnitude lower than the absoluate value, resulting in quite a good skill for the persistence forecast (the major exception being strong flooging events, where most of the time long periods of precipitation lead to day-to-day changes of the possibly almost the same order of magnitude as the absolute values.
# ## Prediction for the test data (used in the model evaluation notebook for comparison)
# Note that the optimized model setting is already updated in the model call far above.
# +
# prediction start from every nth day
# if in doubt, leave n = 1 !!!
n = 1
X_pred = X_test[::n].copy()
y_pred = pipe.predict(X_pred)
y_pred = add_time(y_pred, X_pred.time, name='forecast')
multif_test = generate_prediction_array(y_pred, y, forecast_range=14)
plot_multif_prediction(multif_test, y, forecast_range=14, title='Setting: GradientBoostingRegressor: n_estimators=200; learning_rate=0.1; max_depth=5')
# -
# ## Case study: May/June 2013; Flooding event at Krems (Danube river)
# Drop long time feature not available for the period of interest and do the same preparation steps as before.
X_multif_fin, X_multifr_fin, y_case_fin = multi_forecast_case_study(pipe, X_test, y)
# +
fig, ax = plt.subplots(figsize=(15, 5))
frerun_c = ['silver', 'darkgray', 'gray', 'dimgray']
y_case_fin.to_pandas().plot(ax=ax, label='reanalysis', linewidth=4)
run = 0
for i in X_multifr_fin.num_of_forecast:
X_multif_fin.sel(num_of_forecast=i).to_pandas().T.plot(ax=ax, label='forecast',
linewidth=2, color='tab:cyan')
X_multifr_fin.sel(num_of_forecast=i).to_pandas().T.plot(ax=ax, label='frerun', linewidth=1,
linestyle='--', color=frerun_c[run])
run += 1
ax.set_ylabel('river discharge [m$^3$/s]')
plt.legend(['reanalysis', 'gradient boosting regressor', 'GloFAS 05-18', 'GloFAS 05-22', 'GloFAS 05-25', 'GloFAS 05-29'])
plt.title('GradientBoostingRegressor: n_estimators=200; learning_rate=0.1; max_depth=5 case study May/June 2013');
plt.savefig('gradboost_case.png', dpi=600, bbox_inches='tight')
# -
# Save the postprocessed model forecast, as well as the forecast rerun and the reanalysis data.
X_multif_fin.to_netcdf('../../data/models/GradientBoost/gradient_boost_result_case_study.nc', mode='w')
# Save data of the test period for the model comparison.
multif_test.to_netcdf('../../data/models/GradientBoost/gradient_boost_result_test_period.nc', mode='w')
| notebooks/3_model_tests/3.04_GradientBoost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import math
import os
from matplotlib import pyplot as plt
from PyRadioLoc.Utils.GeoUtils import GeoUtils
# %run utils.ipynb
FILES = [
'./result/Resultados_Equipe4_Metodo_1.csv', # apenas medições
'./result/Resultados_Equipe4_Metodo_2.csv', # fingerprint 20x20
'./result/Resultados_Equipe4_Metodo_3.csv', # fingerprint 20x20 + medições
'./result/Resultados_Equipe4_Metodo_4.csv', # apenas random forest
'./result/Resultados_Equipe4_Metodo_5.csv', # apenas MLP
'./result/Resultados_Equipe4_Metodo_6.csv', # Fingerprint + random forest (sem otimização)
'./result/Resultados_Equipe4_Metodo_7.csv', # Fingerprint + random forest (com otimização - grid)
'./result/Resultados_Equipe4_Metodo_8.csv', # k-NN
]
METHODS = [x.split("_")[-1].split(".")[0] for x in FILES] # lista contendo os metodos que serão usados
db = pd.read_csv('database/LocTreino_Equipe_4.csv')
erbs = pd.read_csv('database/Bts.csv')
errorLocation = pd.read_csv('result/erros.csv')
ERROS_CSV_FILE = './result/erros.csv'
# # 1. Análise dos dados
data = pd.read_csv("./database/LocTreino_Equipe_4.csv")
round(data.describe(), 4)
# # 2. Histograma dos erros (em metros)
for i in METHODS:
errorLocation = pd.read_csv(ERROS_CSV_FILE)
plt.title("Histograma de Erros de Localização Método" + str(i))
histogram_example = plt.hist(errorLocation[str(i)], bins='auto')
plt.xlabel('Metros')
plt.ylabel('Amostras')
plt.show()
# # 3. Boxplot dos erros (em metros)
for i in METHODS:
boxplotErrorLocationM1 = errorLocation.boxplot(column=[str(i)], notch = True, patch_artist = True)
plt.title("Boxplot de Erros de Localização Método "+ str(i))
plt.xlabel('Método '+ str(i))
plt.ylabel('Metros')
plt.show()
# # 4. Mapa de comparação - posições preditas vs. posições reais
for i in METHODS:
dbTeorico = pd.read_csv('result/Resultados_Equipe4_Metodo_'+ str(i) +'.csv')
testMap = merge_on_pontoId(dbTeorico, db)
cols = ['pontoId','rssi_1_1', 'rssi_1_2', 'rssi_1_3', 'rssi_2_1', 'rssi_2_2', 'rssi_2_3', 'rssi_3_1', 'rssi_3_2', 'rssi_3_3', 'delay_1','delay_2','delay_3']
testMap = testMap.drop(cols, axis=1)
fig, ax = plt.subplots()
ax.scatter(testMap['lon_pred'], testMap['lat_pred'], s = 30, color = 'green', alpha = '1')
ax.scatter(testMap['lon'], testMap['lat'], s = 30, color = 'blue', alpha = 1)
ax.scatter(erbs['lon'], erbs['lat'], s = 70, color = 'red', alpha = 1)
ax.set(xlabel = 'lon', ylabel = 'lat')
plt.axis([-34.961,-34.944,-8.060,-8.043])
plt.gcf().set_size_inches((12,12))
plt.title('Método ' + str(i))
plt.show()
# # 5. Erros de localização médio, mínimo, máximo e desvio padrão
# +
def calculate_errors(df_real, df_predict):
"""
Retorna uma lista contendo os erros em metros de todos os pontos no real e na predição
"""
df_merged = merge_on_pontoId(df_predict, df_real)
errors = []
for idx, point in df_merged.iterrows():
distance = GeoUtils.distanceInKm(point.lat, point.lon, point.lat_pred, point.lon_pred)
errors.append(round(distance * 1000, 2))
return errors
def calculate_theorical_errors():
df_real = pd.read_csv("./database/LocTreino_Equipe_4.csv")
dict_errors = {}
for file, method in zip(FILES,METHODS):
df_predict = pd.read_csv(file)
dict_errors[method] = calculate_errors(df_real, df_predict)
return pd.DataFrame(dict_errors)
# -
# Gera o dataframe contendo os erros
errors_df = calculate_theorical_errors()
round(errors_df.describe(), 2)
errors_df.to_csv(ERROS_CSV_FILE, index=False)
| results.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Objective
#
# # Packages
import pandas as pd
import numpy as np
import scipy.stats as stats
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import datetime
from itertools import compress
from matplotlib.ticker import PercentFormatter
from matplotlib_venn import venn3, venn2, venn2_circles
# %matplotlib inline
# # Datasets
record = pd.read_pickle('../primary_care/records2.pkl')
pri_cli = pd.read_csv('../primary_care/gp_clinical.txt', sep = '\t', encoding='ISO-8859-1')
diag = pd.read_excel('../primary_care/diagnosis_cvd.xlsx')
# # Specifying dataset
rec = record[(record['discrepancy'] == False) & (record['event_int'] > 0)]
patients = list(rec['eid'].values)
pri = pri_cli[pri_cli['eid'].isin(patients)].reset_index()
# # Diagnosis
diag
diag['diagnosis'].unique()
diagnosis_codes = {
'hyperlipidaemia': {
'read2': list(diag[diag['diagnosis'] == 'hyperlipidaemia']['READV2_CODE']),
'read3': list(diag[diag['diagnosis'] == 'hyperlipidaemia']['READV3_CODE'])
},
'hypertension': {
'read2': list(diag[diag['diagnosis'] == 'hypertension']['READV2_CODE']),
'read3': list(diag[diag['diagnosis'] == 'hypertension']['READV3_CODE'])
},
'PAD': {
'read2': list(diag[diag['diagnosis'] == 'PAD']['READV2_CODE']),
'read3': list(diag[diag['diagnosis'] == 'PAD']['READV3_CODE'])
},
'CKD': {
'read2': list(diag[diag['diagnosis'] == 'CKD']['READV2_CODE']),
'read3': list(diag[diag['diagnosis'] == 'CKD']['READV3_CODE'])
},
'diabetes': {
'read2': list(diag[diag['diagnosis'] == 'diabetes']['READV2_CODE']),
'read3': list(diag[diag['diagnosis'] == 'diabetes']['READV3_CODE'])
},
'all': {
'read2': list(diag['READV2_CODE']),
'read3': list(diag['READV3_CODE'])
}
}
diag_pri = pri[pri['read_2'].isin(diagnosis_codes['all']['read2'])| pri['read_3'].isin(diagnosis_codes['all']['read3'])]
diag_pri.drop('index', axis = 1, inplace = True)
diag_pri.drop('event_dt', axis = 1, inplace = True)
diag_pri.drop_duplicates(keep = 'first', inplace = True)
new_diag_pri = diag_pri.groupby('eid').agg(list).reset_index()
new_diag_pri
type(new_diag_pri.iloc[0]['read_2'][0])
# # diabetes
diabetes = pri[(pri['read_2'].isin(diagnosis_codes['diabetes']['read2']))|(pri['read_3'].isin(diagnosis_codes['diabetes']['read3']))]
diabetes_df = diabetes.groupby('eid').agg(list).reset_index()
diabetics = list(diabetes_df.eid)
# # CKD
CKD = pri[(pri['read_2'].isin(diagnosis_codes['CKD']['read2']))|(pri['read_3'].isin(diagnosis_codes['CKD']['read3']))]
CKD_df = CKD.groupby('eid').agg(list).reset_index()
chronic = list(CKD_df.eid)
# # PAD
PAD = pri[(pri['read_2'].isin(diagnosis_codes['PAD']['read2']))|(pri['read_3'].isin(diagnosis_codes['PAD']['read3']))]
PAD_df = PAD.groupby('eid').agg(list).reset_index()
peripheral = list(PAD_df.eid)
# # Hypertension
hypertension = pri[(pri['read_2'].isin(diagnosis_codes['hypertension']['read2']))|(pri['read_3'].isin(diagnosis_codes['hypertension']['read3']))]
hypertension_df = hypertension.groupby('eid').agg(list).reset_index()
hypertensives = list(hypertension_df.eid)
# # Hyperlipidaemia
hyperlipid = pri[(pri['read_2'].isin(diagnosis_codes['hyperlipidaemia']['read2']))|(pri['read_3'].isin(diagnosis_codes['hyperlipidaemia']['read3']))]
hyperlipid_df = hyperlipid.groupby('eid').agg(list).reset_index()
hyperchol= list(hyperlipid_df.eid)
# # Gathering
rec['diabetic'] = ['diabetic' if x in diabetics else "" for x in rec['eid'] ]
rec['CKD'] = ['CKD' if x in chronic else "" for x in rec['eid']]
rec['PAD'] = ['peripheral' if x in peripheral else "" for x in rec['eid']]
rec['hypertension'] = ['hypertensives' if x in hypertensives else "" for x in rec['eid']]
rec['hyperlipidaemia'] = ['hyperchol' if x in hyperchol else "" for x in rec['eid']]
concern = ['diabetic', 'CKD', 'PAD', 'hypertension', 'hyperlipidaemia']
for x in concern:
print(rec[x].value_counts())
rec['noconcern'] = rec['diabetic'] + rec['CKD'] + rec['PAD'] + rec['hypertension'] + rec['hyperlipidaemia']
temp = pd.DataFrame(rec.noconcern.value_counts())
temp.to_csv('../primary_care/rec_diagnosis.csv')
| notebooks/diagnosis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # MAT281 - Laboratorio N°02
#
#
# <a id='p1'></a>
#
# ## Problema 01
#
# Una **media móvil simple** (SMA) es el promedio de los últimos $k$ datos anteriores, es decir, sea $a_1$,$a_2$,...,$a_n$ un arreglo $n$-dimensional, entonces la SMA se define por:
#
# $$\displaystyle sma(k) =\dfrac{1}{k}(a_{n}+a_{n-1}+...+a_{n-(k-1)}) = \dfrac{1}{k}\sum_{i=0}^{k-1}a_{n-i} $$
#
#
# Por otro lado podemos definir el SMA con una venta móvil de $n$ si el resultado nos retorna la el promedio ponderado avanzando de la siguiente forma:
#
# * $a = [1,2,3,4,5]$, la SMA con una ventana de $n=2$ sería:
# * sma(2) = [promedio(1,2), promedio(2,3), promedio(3,4), promedio(4,5)] = [1.5, 2.5, 3.5, 4.5]
#
#
# * $a = [1,2,3,4,5]$, la SMA con una ventana de $n=3$ sería:
# * sma(3) = [promedio(1,2,3), promedio(2,3,4), promedio(3,4,5)] = [2.,3.,4.]
#
#
# Implemente una función llamada `sma` cuyo input sea un arreglo unidimensional $a$ y un entero $n$, y cuyo ouput retorne el valor de la media móvil simple sobre el arreglo de la siguiente forma:
#
# * **Ejemplo**: *sma([5,3,8,10,2,1,5,1,0,2], 2)* = $[4. , 5.5, 9. , 6. , 1.5, 3. , 3. , 0.5, 1. ]$
#
# En este caso, se esta calculando el SMA para un arreglo con una ventana de $n=2$.
#
# **Hint**: utilice la función `numpy.cumsum`
# importar librerias
import numpy as np
# ### Definir Función
def sma(a,n):
"""
sma(a,n)
Calculo de la media móvil simple (SMA) del arreglo a.
Parameters
----------
a : array
Arreglo al cual se le calculara el SMA.
n : int
Numero de terminos utilizados para calcular el SMA.
Returns
-------
output : float array
Arreglo con los valores del SMA del arreglo a.
"""
l=np.size(a)
list=[]#lista vacia
for i in range(0,l):
if i+n-1<l:
c=a[i:i+n]#se toman los terminos que se van a sumar
b=np.cumsum(c, dtype=float)#se generan las sumas de los terminos
k=b[len(b)-1]/n#k es el valor de sma(k) para los terminos de c
list.append(k)#a la lista se le agrega k que el sma(k)
r=np.array(list)#se transforma la lista en un arreglo
return r
# ### Verificar ejemplos
# +
# ejemplo 01
a = np.array([1,2,3,4,5])
np.testing.assert_array_equal(
sma(a,2),
np.array([1.5, 2.5, 3.5, 4.5])
)
# +
# ejemplo 02
a = np.array([5,3,8,10,2,1,5,1,0,2])
np.testing.assert_array_equal(
sma(a,2),
np.array([4. , 5.5, 9. , 6. , 1.5, 3. , 3. , 0.5, 1. ])
)
# -
# <a id='p2'></a>
#
# ## Problema 02
#
# La función **strides($a,n,p$)**, corresponde a transformar un arreglo unidimensional $a$ en una matriz de $n$ columnas, en el cual las filas se van construyendo desfasando la posición del arreglo en $p$ pasos hacia adelante.
#
# * Para el arreglo unidimensional $a$ = [ 1, 2, 3, 4, 5, 6, 7, 8, 9, 10], la función strides($a,4,2$), corresponde a crear una matriz de $4$ columnas, cuyos desfaces hacia adelante se hacen de dos en dos.
#
# El resultado tendría que ser algo así:$$\begin{pmatrix}
# 1& 2 &3 &4 \\
# 3& 4&5&6 \\
# 5& 6 &7 &8 \\
# 7& 8 &9 &10 \\
# \end{pmatrix}$$
#
#
# Implemente una función llamada `strides(a,n,p)` cuyo input sea:
# * $a$: un arreglo unidimensional,
# * $n$: el número de columnas,
# * $p$: el número de pasos hacia adelante
#
# y retorne la matriz de $n$ columnas, cuyos desfaces hacia adelante se hacen de $p$ en $p$ pasos.
#
# * **Ejemplo**: *strides($a$,4,2)* =$\begin{pmatrix}
# 1& 2 &3 &4 \\
# 3& 4&5&6 \\
# 5& 6 &7 &8 \\
# 7& 8 &9 &10 \\
# \end{pmatrix}$
#
# ### Definir Función
def strides(a,n,p):
"""
strides(a,n,p)
Transformacion del arreglo a en una matriz de 𝑛 columnas,
donde las filas se van construyendo desfasando la posición del arreglo en 𝑝 pasos hacia adelante.
Parameters
----------
a : array
Arreglo que se transformara a matriz.
n : int
Tamaño de las filas y columnas de la matriz.
p : int
Desfase para los primeros terminos de cada fila de la matriz
Returns
-------
output : matrix
Matriz hecha del arreglo a, tamaño nxn con un desfase de p en el primer termino de cada fila.
"""
l=np.size(a)
i=0
list_provisoria=[]#lista vacia
while i<l:
k=i
if np.size(a[i:i+n])==n:#se revisa si el arreglo tiene el largo de n
list_provisoria.append(a[i:i+n])# se agrega el arreglo a la lista
else:
break
i+=p
r = np.array(list_provisoria)#se transforma la lista en un arreglo
return r
# ### Verificar ejemplos
# +
# ejemplo 01
a = np.array([ 1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
n=4
p=2
np.testing.assert_array_equal(
strides(a,n,p),
np.array([
[ 1, 2, 3, 4],
[ 3, 4, 5, 6],
[ 5, 6, 7, 8],
[ 7, 8, 9, 10]
])
)
# -
# <a id='p3'></a>
#
# ## Problema 03
#
#
# Un **cuadrado mágico** es una matriz de tamaño $n \times n$ de números enteros positivos tal que
# la suma de los números por columnas, filas y diagonales principales sea la misma. Usualmente, los números empleados para rellenar las casillas son consecutivos, de 1 a $n^2$, siendo $n$ el número de columnas y filas del cuadrado mágico.
#
# Si los números son consecutivos de 1 a $n^2$, la suma de los números por columnas, filas y diagonales principales
# es igual a : $$\displaystyle M_{n} = \dfrac{n(n^2+1)}{2}$$
# Por ejemplo,
#
# * $A= \begin{pmatrix}
# 4& 9 &2 \\
# 3& 5&7 \\
# 8& 1 &6
# \end{pmatrix}$,
# es un cuadrado mágico.
#
# * $B= \begin{pmatrix}
# 4& 2 &9 \\
# 3& 5&7 \\
# 8& 1 &6
# \end{pmatrix}$, no es un cuadrado mágico.
#
# Implemente una función llamada `es_cudrado_magico` cuyo input sea una matriz cuadrada de tamaño $n$ con números consecutivos de $1$ a $n^2$ y cuyo ouput retorne *True* si es un cuadrado mágico o 'False', en caso contrario
#
# * **Ejemplo**: *es_cudrado_magico($A$)* = True, *es_cudrado_magico($B$)* = False
#
# **Hint**: Cree una función que valide la mariz es cuadrada y que sus números son consecutivos del 1 a $n^2$.
# ### Definir Función
def es_cudrado_magico(M):
"""
es_cudrado_magico(M)
Revisa si la matriz M es un cuadrado mágico.
Parameters
----------
M : matriz
Matriz a revisar si es cuadrado mágico.
Returns
-------
output : bool
True or False.
"""
if M.shape[0]!=M.shape[1]:#revisa que la matriz sea cuadrada
return False
else:
p=[]
for i in range(0,M.shape[0]):#pone los elementos en una lista
for k in range(0,M.shape[0]):
p.append(M[i][k])
p.sort()
for i in range(0,len(p)):#revisa que todos los numeros sean unicos de 1 a n^2
if p[i]!=i+1:
return False
for i in range(0,M.shape[0]):#revisa que ningun termino sea mayor a n^2
for k in range (0,M.shape[0]):
if M[i][k]>M.shape[0]**2:
return False
M_n=M.shape[0]*(M.shape[0]**2+1)/2#calcula valor M_n con el que se comparan las sumas
for i in range(0,M.shape[0]):#suma las filas y revisa si dan M_n
v=0
for k in range (0,M.shape[0]):
v+=M[i][k]
if v!=M_n:#si alguna fila no suma M_n devuelve falso
return False
for k in range(0,M.shape[0]):#suma las columnas y revisa si dan M_n
v=0
for i in range (0,M.shape[0]):
v+=M[i][k]
if v!=M_n:#si alguna columna no suma M_n devuelve falso
return False
for i in range(0,M.shape[0]):#suma la diagonal principal y revisa si dan M_n
v=0
v+=M[i][i]
if i==M.shape[0]:
if v!=M_n:#si la diagonal principal no suma M_n devuelve falso
return False
v=0
for i in range (0,M.shape[0]):#suma la diagonal inversa (M_13,M_22,M_31) y revisa si dan M_n
k=0
v+=M[i][M.shape[0]-1-k]
k+=1
if v!=M_n:#si la diagonal inversa no suma M_n devuelve falso
return False
return True
# ### Verificar ejemplos
# ejemplo 01
A = np.array([[4,9,2],[3,5,7],[8,1,6]])
assert es_cudrado_magico(A) == True, "ejemplo 01 incorrecto"
# ejemplo 02
B = np.array([[4,2,9],[3,5,7],[8,1,6]])
assert es_cudrado_magico(B) == False, "ejemplo 02 incorrecto"
| labs/lab_02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## k-近傍分類器
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from sklearn import neighbors, datasets
# %matplotlib inline
input_file = './data/data.txt'
data = np.loadtxt(input_file, delimiter=',')
X, y = data[:, :-1], data[:, -1].astype(np.int)
plt.figure()
plt.title('Input data')
marker_shapes = 'v^os'
for i in range(X.shape[0]):
plt.scatter(X[i, 0], X[i, 1], marker=marker_shapes[y[i]],
s=75, edgecolor='black', facecolor='none')
plt.show()
# +
num_neighbors = 12
classifier = neighbors.KNeighborsClassifier(num_neighbors, weights='distance')
classifier.fit(X, y)
step_size = 0.01
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
x_values, y_values = np.meshgrid(np.arange(x_min, x_max, step_size), np.arange(y_min, y_max, step_size))
output = classifier.predict(np.c_[x_values.ravel(), y_values.ravel()])
output = output.reshape(x_values.shape)
plt.figure()
plt.pcolormesh(x_values, y_values, output, cmap=cm.Blues)
for i in range(X.shape[0]):
plt.scatter(X[i, 0], X[i, 1], marker=marker_shapes[y[i]],
s=50, edgecolor='black', facecolor='none')
plt.xlim(x_values.min(), x_values.max())
plt.ylim(y_values.min(), y_values.max())
plt.title('K Nearest neighbors classifier model boundaries')
plt.show()
# +
test_datapoint = [5.1, 3.6]
plt.figure()
plt.title('Test datapoint')
for i in range(X.shape[0]):
plt.scatter(X[i, 0], X[i, 1], marker=marker_shapes[y[i]],
s=75, edgecolor='black', facecolor='none')
plt.scatter(test_datapoint[0], test_datapoint[1], marker='x',
linewidth=6, s=200, facecolor='black')
plt.show()
# +
_, indices = classifier.kneighbors([test_datapoint])
indices = indices.astype(np.int)[0]
plt.figure()
plt.title('K Nearest Neighbors')
for i in indices:
plt.scatter(X[i, 0], X[i, 1], marker=marker_shapes[y[i]],
linewidth=3, s=100, facecolor='black')
plt.scatter(test_datapoint[0], test_datapoint[1], marker='x',
linewidth=6, s=200, facecolor='black')
for i in range(X.shape[0]):
plt.scatter(X[i, 0], X[i, 1], marker=marker_shapes[y[i]],
s=75, edgecolor='black', facecolor='none')
plt.show()
# +
print('K-Nearest Neighbours:')
for i in indices:
print('({},{}) -> {}'.format(X[i, 0], X[i, 1], y[i]))
print("Predicted output:", classifier.predict([test_datapoint])[0])
# -
| Ex/Chapter5/Chapter5-3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ECE 590, Fall 2019
# ## Problem Set 1
# * ### __Important :__ You are only allowed to use the Python built in function for generating uniform random variables.
#
# ## Full name: <NAME>
#
# ### Problem 1 (Exponential distribution)
import numpy as np
import matplotlib.pyplot as plt
import math
# Put your code here
uniform = np.random.uniform(0,1,100000)
parameters = [0.1, 1, 10]
plt.figure(figsize=(15, 5))
for i, lamb in enumerate(parameters):
samples = -(1/lamb) * np.log(1 - uniform)
plt.subplot(1, 3, i+1)
plt.title('Exponential Distribution with $\lambda$ =' + str(lamb))
plt.hist(samples, bins = np.arange(min(samples), max(samples), 0.01))
# ### Problem 2 (Gamma distribution)
# Put your code here
betas = [0.1, 1, 10]
def getExponential(beta, x):
X = -beta * np.log(1 - x)
return X
def getGamma(k, beta):
gamma = 0
for i in range(k):
uniform = np.random.uniform(0,1,100000)
X = getExponential(beta, uniform)
gamma += X
return gamma
plt.figure(figsize=(15, 5))
for i, beta in enumerate(betas):
samples = getGamma(5, beta)
plt.subplot(1, 3, i+1)
plt.title('Gamma Distribution with beta =' + str(beta))
plt.hist(samples, bins = np.arange(min(samples), max(samples), 0.01))
# ### Problem 3 (Beta distribution)
# Put your code here
alpha_1s = [5, 10]
alpha_2s = [16, 11]
def getBeta(alpha1, alpha2, sample_size):
n = alpha1 + alpha2 - 1
k = alpha1
data = np.zeros((n, sample_size))
for i in range(n):
data[i, :] = np.random.uniform(0,1,sample_size)
data = np.sort(data, axis = 0) # Order statistics, exactly k-1 of the others < x
return data[k, :]
plt.figure(figsize=(10, 5))
for (i,alpha_1), alpha_2 in zip(enumerate(alpha_1s), alpha_2s):
samples = getBeta(alpha_1, alpha_2, 100000)
plt.subplot(1, 2, i+1)
plt.title('Beta Distribution with parameters (%d, %d)' % (alpha_1, alpha_2))
plt.hist(samples, bins = np.arange(min(samples), max(samples), 0.01))
# ### Problem 4 (Dirichlet distribution)
# ### 1. Gamma
# Put your code here
def getDirichlet(N, alpha, sample_size):
data = np.zeros((N, sample_size))
for i in range(N):
data[i,:] = getGamma(alpha, 1/10)
data_sum = np.sum(data, axis = 0)
data_dirich = data/data_sum
return data_dirich
data_dirich = getDirichlet(3, 10, 100000)
plt.hist2d(data_dirich[0,:], data_dirich[1,:], bins=[np.arange(min(data_dirich[0,:]), max(data_dirich[0,:]), 0.01),\
np.arange(min(data_dirich[1,:]), max(data_dirich[1,:]), 0.01)] )
pass
# ### 2. Stick method
def stick_breaking(alpha1, k, sample_size):
final_beta = np.zeros((k, sample_size))
alpha2 = 2 * alpha1
betas_1 = getBeta(alpha1, alpha2, sample_size)
final_beta[0,:] = betas_1
alpha2 = alpha1
betas_2 = getBeta(alpha1, alpha2, sample_size)
remaining_pieces = 1 - betas_1
p = betas_2 * remaining_pieces
final_beta[1,:] = p
remaining_pieces = (1 - betas_1) * (1 - betas_2)
p = remaining_pieces
final_beta[2,:] = p
return final_beta
final_beta = stick_breaking(10, 3, 100000)
plt.hist2d(final_beta[0,:], final_beta[1,:], bins=[np.arange(min(final_beta[0,:]), max(final_beta[0,:]), 0.01),\
np.arange(min(final_beta[1,:]), max(final_beta[1,:]), 0.01)] )
pass
| HW1_Jupyter_template_ECE590_Bingying_Liu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import climin
from functools import partial
import warnings
import os
sys.path.append('..')
import numpy as np
from scipy.stats import multinomial
from scipy.linalg.blas import dtrmm
import GPy
from GPy.util import choleskies
from GPy.core.parameterization.param import Param
from GPy.kern import Coregionalize
from GPy.likelihoods import Likelihood
from GPy.util import linalg
from likelihoods.bernoulli import Bernoulli
from likelihoods.gaussian import Gaussian
from likelihoods.categorical import Categorical
from likelihoods.hetgaussian import HetGaussian
from likelihoods.beta import Beta
from likelihoods.gamma import Gamma
from likelihoods.exponential import Exponential
from hetmogp.util import draw_mini_slices
from hetmogp.het_likelihood import HetLikelihood
from hetmogp.svmogp import SVMOGP
from hetmogp import util
from hetmogp.util import vem_algorithm as VEM
import matplotlib.pyplot as plt
from matplotlib.pyplot import gca
from matplotlib import rc, font_manager
from matplotlib import rcParams
from matplotlib2tikz import save as tikz_save
warnings.filterwarnings("ignore")
os.environ['PATH'] = os.environ['PATH'] + ':/usr/texbin'
# +
M = 8 # number of inducing points
Q = 2 # number of latent functions
# Heterogeneous Likelihood Definition
likelihoods_list = [Gaussian(sigma=1.), Bernoulli()] # Real + Binary
likelihood = HetLikelihood(likelihoods_list)
Y_metadata = likelihood.generate_metadata()
D = likelihood.num_output_functions(Y_metadata)
W_list, _ = util.random_W_kappas(Q, D, rank=1, experiment=True)
X1 = np.sort(np.random.rand(600))[:, None]
X2 = np.sort(np.random.rand(500))[:, None]
X = [X1, X2]
# +
# True U and F functions
def experiment_true_u_functions(X_list):
u_functions = []
for X in X_list:
u_task = np.empty((X.shape[0],2))
u_task[:,0,None] = 4.5*np.cos(2*np.pi*X + 1.5*np.pi) - \
3*np.sin(4.3*np.pi*X + 0.3*np.pi) + \
5*np.cos(7*np.pi * X + 2.4*np.pi)
u_task[:,1,None] = 4.5*np.cos(1.5*np.pi*X + 0.5*np.pi) + \
5*np.sin(3*np.pi*X + 1.5*np.pi) - \
5.5*np.cos(8*np.pi * X + 0.25*np.pi)
u_functions.append(u_task)
return u_functions
def experiment_true_f_functions(true_u, X_list):
true_f = []
W = W_lincombination()
# D=1
for d in range(2):
f_d = np.zeros((X_list[d].shape[0], 1))
for q in range(2):
f_d += W[q][d].T*true_u[d][:,q,None]
true_f.append(f_d)
return true_f
# True Combinations
def W_lincombination():
W_list = []
# q=1
Wq1 = np.array(([[-0.5],[0.1]]))
W_list.append(Wq1)
# q=2
Wq2 = np.array(([[-0.1],[.6]]))
W_list.append(Wq2)
return W_list
# +
# True functions values for inputs X
trueU = experiment_true_u_functions(X)
trueF = experiment_true_f_functions(trueU, X)
# Generating training data Y (sampling from heterogeneous likelihood)
Y = likelihood.samples(F=trueF, Y_metadata=Y_metadata)
# +
# Plot true parameter functions PFs (black) and heterogeneous data (blue, orange)
plt.figure(figsize=(10, 6))
Ntask = 2
for t in range(Ntask):
plt.plot(X[t],trueF[t],'k-', alpha=0.75)
plt.plot(X[t],Y[t],'+')
plt.show()
# +
# Generating missing data (gap)
X2test = X[1][np.r_[351:450],:]
Y2test = Y[1][np.r_[351:450],:]
X2train_s1 = X[1][1:351,:]
X2train_s2 = X[1][450:,:]
X2 = np.delete(X2, np.s_[np.r_[351:450]],0)
Y2 = np.delete(Y[1], np.s_[np.r_[351:450]],0)
X = [X1, X2]
Y = [Y[0], Y2]
# +
# Plot gap
plt.figure(figsize=(10, 6))
Ntask = 2
for t in range(Ntask):
plt.plot(X[t],Y[t],'b+')
plt.plot(X2test, Y2test,'r+')
plt.show()
# +
# KERNELS
ls_q = np.array(([.05]*Q))
var_q = np.array(([.5]*Q))
kern_list = util.latent_functions_prior(Q, lenghtscale=ls_q, variance=var_q, input_dim=1)
# # INDUCING POINTS
Z = np.linspace(0, 1, M)
Z = Z[:, np.newaxis]
# -
# MODEL and INFERENCE
model = SVMOGP(X=X, Y=Y, Z=Z, kern_list=kern_list, likelihood=likelihood, Y_metadata=Y_metadata)
model = VEM(model, stochastic=False, vem_iters=5, optZ=True, verbose=False, verbose_plot=False, non_chained=True)
def plot_figure_gaplatex(model, Xtrain, Ytrain, Xtest, Ytest):
sorted_Xtrain0 = np.sort(Xtrain[0])
m_f_gaussian, v_f_gaussian = model.predictive_new(sorted_Xtrain0, output_function_ind=0)
m_f_gaussian_upper = m_f_gaussian + 2*np.sqrt(v_f_gaussian)
m_f_gaussian_lower = m_f_gaussian - 2*np.sqrt(v_f_gaussian)
sorted_Xtrain1_s1 = np.sort(X2train_s1)
m_f_ber, v_f_ber = model.predictive_new(sorted_Xtrain1_s1, output_function_ind=1)
m_f_gp_upper = m_f_ber + 2*np.sqrt(v_f_ber)
m_f_gp_lower = m_f_ber - 2*np.sqrt(v_f_ber)
m_ber_gp_s1 = np.exp(m_f_ber)/ (1 + np.exp(m_f_ber))
m_ber_gp_upper_s1 = np.exp(m_f_gp_upper)/ (1 + np.exp(m_f_gp_upper))
m_ber_gp_lower_s1 = np.exp(m_f_gp_lower)/ (1 + np.exp(m_f_gp_lower))
sorted_Xtrain1_s2 = np.sort(X2train_s2)
m_f_ber, v_f_ber = model.predictive_new(sorted_Xtrain1_s2, output_function_ind=1)
m_f_gp_upper = m_f_ber + 2*np.sqrt(v_f_ber)
m_f_gp_lower = m_f_ber - 2*np.sqrt(v_f_ber)
m_ber_gp_s2 = np.exp(m_f_ber)/ (1 + np.exp(m_f_ber))
m_ber_gp_upper_s2 = np.exp(m_f_gp_upper)/ (1 + np.exp(m_f_gp_upper))
m_ber_gp_lower_s2 = np.exp(m_f_gp_lower)/ (1 + np.exp(m_f_gp_lower))
sorted_Xtest = np.sort(Xtest)
m_pred_ber, v_pred_ber = model.predictive_new(sorted_Xtest, output_function_ind=1)
m_pred_gp_upper = m_pred_ber + 2*np.sqrt(v_pred_ber)
m_pred_gp_lower = m_pred_ber - 2*np.sqrt(v_pred_ber)
m_pred_gp = np.exp(m_pred_ber)/ (1 + np.exp(m_pred_ber))
m_pred_gp_upper = np.exp(m_pred_gp_upper)/ (1 + np.exp(m_pred_gp_upper))
m_pred_gp_lower = np.exp(m_pred_gp_lower)/ (1 + np.exp(m_pred_gp_lower))
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
fig_gap_gaussian = plt.figure(figsize=(10, 5))
plt.plot(Xtrain[0], Ytrain[0], 'x', color='blue', markersize=10, alpha=0.1)
plt.plot(sorted_Xtrain0, m_f_gaussian, 'b-', linewidth=4, alpha=1)
plt.plot(sorted_Xtrain0, m_f_gaussian_upper, 'b-', linewidth=4, alpha=0.25)
plt.plot(sorted_Xtrain0, m_f_gaussian_lower, 'b-', linewidth=4, alpha=0.25)
plt.title(r'Output 1: Gaussian Regression')
plt.ylabel(r'Real Output')
plt.xlabel(r'Real Input')
plt.xlim(0,1)
plt.show()
fig_gap_bernoulli = plt.figure(figsize=(10, 5))
plt.plot(X2, Y2, 'x', color='blue', markersize=10, alpha=0.1)
plt.plot(Xtest, Ytest, 'x', color='red', markersize=10, alpha=0.1)
plt.plot(sorted_Xtrain1_s1, m_ber_gp_s1, 'b-', linewidth=4, alpha=1)
plt.plot(sorted_Xtrain1_s1, m_ber_gp_upper_s1, 'b-', linewidth=4, alpha=0.25)
plt.plot(sorted_Xtrain1_s1, m_ber_gp_lower_s1, 'b-', linewidth=4, alpha=0.25)
plt.plot(sorted_Xtrain1_s2, m_ber_gp_s2, 'b-', linewidth=4, alpha=1)
plt.plot(sorted_Xtrain1_s2, m_ber_gp_upper_s2, 'b-', linewidth=4, alpha=0.25)
plt.plot(sorted_Xtrain1_s2, m_ber_gp_lower_s2, 'b-', linewidth=4, alpha=0.25)
plt.plot(sorted_Xtest, m_pred_gp, 'r-', linewidth=4, alpha=1)
plt.plot(sorted_Xtest, m_pred_gp_upper, 'r-', linewidth=4, alpha=0.25)
plt.plot(sorted_Xtest, m_pred_gp_lower, 'r-', linewidth=4, alpha=0.25)
plt.title(r'Output 2: Binary Classification')
plt.ylabel(r'Binary Output')
plt.xlabel(r'Real Input')
plt.xlim(0,1)
plt.show()
plot_figure_gaplatex(model, X, Y, X2test, Y2test)
| notebooks/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## icepyx PyPI Statistics
# Use PyPIStats library to get data on PyPI downloads of icepyx (or any other package)
#
# See the [pypistats website](https://github.com/hugovk/pypistats) for potential calls, options, and formats (e.g. markdown, rst, html, json, numpy, pandas)
#
# **Note: currently this needs to be run manually (should be able to run all cells) and the changes committed.**
import os
import pypistats
import pandas as pd
# # !pip install --upgrade "pypistats[pandas]" # may need this if pypistats wasn't installed with it
# Note: a numpy version is also available
# +
cwd = os.getcwd()
trackpath= cwd + '/' # '/doc/source/tracking/pypistats/'
downloadfn = "downloads_data.csv"
sysdownloadfn = "sys_downloads_data.csv"
# +
downloads = pypistats.overall("icepyx", total=True, format="pandas").drop(columns=['percent'])
downloads = downloads[downloads.category != "Total"]
# try:
exist_downloads = pd.read_csv(trackpath+downloadfn)#.drop(columns=['percent'])
# exist_downloads = exist_downloads[exist_downloads.category != "Total"]
dl_data = downloads.merge(exist_downloads, how='outer',
on=['category','date','downloads']).reindex()
# except:
# dl_data = downloads
dl_data.to_csv(trackpath+downloadfn, index=False)
# +
sysdownloads = pypistats.system("icepyx", total=True, format="pandas").drop(columns=['percent'])
sysdownloads = sysdownloads[sysdownloads.category != "Total"]
# try:
exist_sysdownloads = pd.read_csv(trackpath+sysdownloadfn)#.drop(columns=['percent'])
# exist_sysdownloads = exist_sysdownloads[exist_sysdownloads.category != "Total"]
exist_sysdownloads['category'] = exist_sysdownloads['category'].fillna("null")
sysdl_data = sysdownloads.merge(exist_sysdownloads, how='outer',
on=['category','date','downloads']).reindex()
# except:
# dl_data = sysdownloads
sysdl_data.to_csv(trackpath+sysdownloadfn, index=False)
# +
dl_data = dl_data.groupby("category").get_group("without_mirrors").sort_values("date")
chart = dl_data.plot(x="date", y="downloads", figsize=(10, 2),
label="Number of PyPI Downloads")
chart.figure.show()
chart.figure.savefig(trackpath+"downloads.svg")
# -
| doc/source/tracking/pypistats/get_pypi_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
print(plt.style.available)
mpl.style.use('ggplot')
plt.plot([1, 4, 9, 16],'rs--')
plt.show()
mpl.style.use('default')
plt.plot([1, 4, 9, 16], 'rs--')
plt.show()
mpl.style.use('seaborn-dark')
plt.plot([1, 4, 9, 16], 'rs--')
plt.show()
mpl.style.use('seaborn')
plt.plot([1, 4, 9, 16], 'rs--')
plt.show()
x = range(1000)
y = [i ** 2 for i in x]
plt.plot(x,y)
plt.show();
# %config InlineBackend.figure_format = 'retina'
plt.plot(x,y)
plt.show();
| python/matplotlib style custom ( retina style ).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# ## Why text is bad for you
# +
import pandas as pd
import re
pd.set_option('max_colwidth', 300)
# -
df = pd.read_csv('../data/wiki/wiki.csv.gz', encoding='utf8', index_col=None)
df['text'] = df.text.str.extract(r'^(.*?)\.', expand=False)
df.head()
# ## So what can we do?
import nltk
#nltk.download('all')
from nltk.tokenize import SpaceTokenizer
tokenizer = SpaceTokenizer()
tokenizer.tokenize('He takes long walks')
from nltk.stem.snowball import EnglishStemmer
stemmer = EnglishStemmer()
stemmer.stem('Walks')
# +
def tokenize_only(text):
tokens = tokenizer.tokenize(text)
# filter out any tokens not containing letters (e.g., numeric tokens, raw punctuation)
filtered_tokens = [token for token in tokens if re.search('[a-zA-Z]', token)]
return filtered_tokens
def tokenize_and_stem(text):
tokens = tokenize_only(text)
stems = map(stemmer.stem, tokens)
return stems
# -
# ## Ok.. but, how does that help me?
sample = df[df.name.isin(['4chan', '8chan', 'Aerosmith', 'Alabama', 'Texas'])]
sample
from sklearn.feature_extraction.text import TfidfVectorizer
tfidf_vectorizer = TfidfVectorizer(stop_words='english', min_df=2, tokenizer=tokenize_and_stem)
vec_text = tfidf_vectorizer.fit_transform(sample.text)
vec_text
pd.DataFrame(vec_text.toarray())
tfidf_vectorizer.get_feature_names()
tfidf_vectorizer.idf_
vec_df = pd.DataFrame(vec_text.toarray(), columns=tfidf_vectorizer.get_feature_names())
sample.reset_index(drop=True).join(vec_df)
| notebooks/401-WorkingWithText.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from datetime import date, timedelta
import pandas as pd
import numpy as np
from sklearn.metrics import mean_squared_error
import lightgbm as lgb
# -
# cd E:\Time-Series Data
df_train_train = pd.read_csv("train.csv", usecols=[1, 2, 3, 4, 5],
dtype={'onpromotion': bool},
converters={'unit_sales': lambda u: np.log1p(
float(u)) if float(u) > 0 else 0},
parse_dates=["date"],
skiprows=range(1, 66458909) # 2016-01-01
)
df_train.dtypes
df_train.head()
df_test = pd.read_csv("test.csv", usecols=[0, 1, 2, 3, 4],
dtype={'onpromotion': bool},
parse_dates=["date"] # , date_parser=parser
).set_index(
['store_nbr', 'item_nbr', 'date']
)
df_test.dtypes
df_test.head()
items = pd.read_csv("items.csv",
).set_index("item_nbr")
items.head()
items.dtypes
df_2017 = df_train[df_train.date.isin(
pd.date_range("2017-05-31", periods=7 * 11))].copy()
df_2017.head()
promo_2017_train = df_2017.set_index(
["store_nbr", "item_nbr", "date"])[["onpromotion"]].unstack(
level=-1).fillna(False)
promo_2017_train.head()
# + jupyter={"outputs_hidden": true}
promo_2017_train.columns = promo_2017_train.columns.get_level_values(1)
# -
promo_2017_test = df_test[["onpromotion"]].unstack(level=-1).fillna(False)
promo_2017_test.head()
promo_2017_test.columns = promo_2017_test.columns.get_level_values(1)
promo_2017_test.columns
promo_2017_test = promo_2017_test.reindex(promo_2017_train.index).fillna(False)
promo_2017_test.head()
promo_2017 = pd.concat([promo_2017_train, promo_2017_test], axis=1)
promo_2017.head()
df_2017 = df_2017.set_index(
["store_nbr", "item_nbr", "date"])[["unit_sales"]].unstack(
level=-1).fillna(0)
df_2017.head()
df_2017.columns = df_2017.columns.get_level_values(1)
df_2017.head()
df_2017.columns
items = items.reindex(df_2017.index.get_level_values(1))
items.head()
def get_timespan(df, dt, minus, periods):
return df[
pd.date_range(dt - timedelta(days=minus), periods=periods)
]
def prepare_dataset(t2017, is_train=True):
X = pd.DataFrame({
"mean_3_2017": get_timespan(df_2017, t2017, 3, 3).mean(axis=1).values,
"mean_7_2017": get_timespan(df_2017, t2017, 7, 7).mean(axis=1).values,
"mean_14_2017": get_timespan(df_2017, t2017, 14, 14).mean(axis=1).values,
"promo_14_2017": get_timespan(promo_2017, t2017, 14, 14).sum(axis=1).values
})
for i in range(16):
X["promo_{}".format(i)] = promo_2017[
t2017 + timedelta(days=i)].values.astype(np.uint8)
if is_train:
y = df_2017[
pd.date_range(t2017, periods=16)
].values
return X, y
return X
print("Preparing dataset...")
t2017 = date(2017, 6, 21)
t2017
X_l, y_l = [], []
for i in range(4):
delta = timedelta(days=7 * i)
X_tmp, y_tmp = prepare_dataset(
t2017 + delta
)
X_l.append(X_tmp)
y_l.append(y_tmp)
X_train = pd.concat(X_l, axis=0)
X_train.head()
y_train = np.concatenate(y_l, axis=0)
y_train
X_val, y_val = prepare_dataset(date(2017, 7, 26))
X_test = prepare_dataset(date(2017, 8, 16), is_train=False)
X_test.head()
print("Training and predicting models...")
params = {
'num_leaves': 2**5 - 1,
'objective': 'regression_l2',
'max_depth': 8,
'min_data_in_leaf': 50,
'learning_rate': 0.05,
'feature_fraction': 0.75,
'bagging_fraction': 0.75,
'bagging_freq': 1,
'metric': 'l2',
'num_threads': 4
}
X_train.head()
X_val.head()
MAX_ROUNDS = 1000
val_pred = []
test_pred = []
cate_vars = []
for i in range(16):
print("=" * 50)
print("Step %d" % (i+1))
print("=" * 50)
dtrain = lgb.Dataset(
X_train, label=y_train[:, i],
categorical_feature=cate_vars,
weight=pd.concat([items["perishable"]] * 4) * 0.25 + 1
)
dval = lgb.Dataset(
X_val, label=y_val[:, i], reference=dtrain,
weight=items["perishable"] * 0.25 + 1,
categorical_feature=cate_vars)
bst = lgb.train(
params, dtrain, num_boost_round=MAX_ROUNDS,
valid_sets=[dtrain, dval], early_stopping_rounds=50, verbose_eval=50
)
print("\n".join(("%s: %.2f" % x) for x in sorted(
zip(X_train.columns, bst.feature_importance("gain")),
key=lambda x: x[1], reverse=True
)))
val_pred.append(bst.predict(
X_val, num_iteration=bst.best_iteration or MAX_ROUNDS))
test_pred.append(bst.predict(
X_test, num_iteration=bst.best_iteration or MAX_ROUNDS))
print("Validation mse:", mean_squared_error(
y_val, np.array(val_pred).transpose()))
print("Making submission...")
y_test = np.array(test_pred).transpose()
y_test
df_preds = pd.DataFrame(
y_test, index=df_2017.index,
columns=pd.date_range("2017-08-16", periods=16)
).stack().to_frame("unit_sales")
df_preds.head()
df_preds.index.set_names(["store_nbr", "item_nbr", "date"], inplace=True)
df_preds.head()
submission = df_test[["id"]].join(df_preds, how="left").fillna(0)
submission.head()
submission["unit_sales"] = np.clip(np.expm1(submission["unit_sales"]), 0, 1000)
submission.head()
submission.to_csv('my_lgb.csv', float_format='%.4f', index=None)
| Machine Learning/0) Time-series/my_practice_lgbm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="5uoyav_d-V1s"
# # Projet préparez des données pour un organisme de santé publique
# -
# Our goal is to select the best products for a given type of products and given features.
# We will ask the user the parameters and give him a ranking of the products as well as a preview of the product (not always available).
#
# But before any ranking, we need to acquire the data and clean them.
# We will use the [OpenFoodFacts database](https://world.openfoodfacts.org/data) which is a open database under the Open Database License (ODbL).
# + id="Fo43CJ2vH0pB"
# %%capture
# To run the notebook
# %pip install jupyter
# To draw plots
# %pip install matplotlib
import matplotlib.pyplot as plt
from matplotlib.collections import LineCollection
# To draw plots in the notebook
# %matplotlib inline
# To manipulate dataFrames
# %pip install pandas
import pandas as pd
# To use quick functions (mainly on arrays)
# %pip install numpy
import numpy as np
# To plot prettiers graphs simpler
# %pip install seaborn
import seaborn as sns
sns.set()
# Allow to omit the warnings
# %pip install warnings
import warnings
warnings.filterwarnings(action='ignore')
# To print dataframes in a nice way
# %pip install dataframe_image
import dataframe_image as dfi
# To be able to encode dataframe to display them as pictures
# %pip install unicodedata
import unicodedata
# To use widgets to interact with the notebook
# %pip install ipywidgets
import ipywidgets as widgets
# To use data science models
# %pip install sklearn
from sklearn import linear_model
from sklearn.impute import SimpleImputer
# To make reports
# %pip install scipy
from scipy.cluster.hierarchy import dendrogram
from IPython.display import HTML
# -
def export_df(df, name, cols = 20, rows = 20):
df = df[:cols][:rows]
df = df.applymap(lambda x : unicodedata.normalize('NFKD', str(x)).encode('ascii', 'ignore').decode())
dfi.export(df, name, max_cols=cols, max_rows=rows)
# +
url = 'https://s3-eu-west-1.amazonaws.com/static.oc-static.com/prod/courses/files/parcours-data-scientist/P2/fr.openfoodfacts.org.products.csv.zip'
# Download file from url
import urllib.request
urllib.request.urlretrieve(url, 'fr.openfoodfacts.org.products.csv.zip')
# Unzip file
import zipfile
with zipfile.ZipFile('fr.openfoodfacts.org.products.csv.zip', 'r') as zip_ref:
zip_ref.extractall()
data = pd.read_csv('./fr.openfoodfacts.org.products.csv', sep='\t')
# + id="r-9R7rcvydZZ"
# Deep copy of the original dataframe
df = data.copy()
# -
# After downloading and loading the data, let's see the shape of the data. First, we want to have a quick overview of the number of columns and rows and get some statistics about them.
# + colab={"base_uri": "https://localhost:8080/"} id="Yu8DsyWbMozZ" outputId="0056898c-1437-4946-f619-6222c6326f4b"
# Quick description of the data
print(f'The dataset contains {df.shape[1]} rows and {df.shape[0]} columns')
# + colab={"base_uri": "https://localhost:8080/", "height": 264} id="QekiMTiRJTzw" outputId="29303ffe-e2fb-4e84-a8c3-419d4104b612"
value_counts = df.dtypes.value_counts()
figure, ax = plt.subplots()
figure.set_size_inches(5,5)
plt.pie(value_counts.values, autopct='%1.0f%%')
plt.legend(labels = value_counts.index, loc = "best")
ax.set_title('Data type analysis')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 508} id="UGSOhV5Nd4wf" outputId="fe0c74cf-20ef-4bd2-cd8e-5f3806ca2f21"
print('Now, let\'s get a quick pic of the data (only the first 5 rows)')
export_df(df.head(), 'data_head.png')
# -
# 
# There seems to be a lot of missing values. But before making any cleaning of the data, we must see which are the proportion of missing values and what are the columns we can rely on.
# + [markdown] id="lV7o5Md2Sdaw"
# ## Missing values analysis
# + id="ZpO3rOkxJ9a2"
# For better understanding of the dataset completion, we try to display the arrangement of the missing values by using a heatmap
# Very heavy to compute
sns.set()
figure, ax = plt.subplots(figsize=(20, 5))
sns.heatmap(df.isna(), cbar=False, ax = ax)
ax.set_title('Missing values heatmap')
plt.show()
# -
# This heatmap is a great way to represent missing values. White parts indicates missing values.
#
# We can instantly see which columns are filled and which are not, for example, `code`, `creator name` and `created_datetime` are columns we can always count on but sadly they aren't very interesting for our analysis.
#
# Hopefully, we can see that a LOT of columns are named `[type]_100g` which means wheight over 100g of product could be a great mesure for our features ! However, a lot of them have are greatly hollow and we won't be able to use them.
#
# Even if this graph is convenient for our eyes, lets put a **totally arbitrary** threshold of missing value to see which columns we will allow us to use !
# + colab={"base_uri": "https://localhost:8080/"} id="4WC9Nq61Ku9F" outputId="9d946825-7819-4791-8e06-be6eb114c53d"
# After the visualization, we try to mesure the missing values
proportion = 100- (round(df.isna().sum()*100/df.shape[0],2)).sort_values(ascending=True)
print('Columns sorted by their completion rate (only the 10 first and 10 last):')
print(proportion[:10])
print('----------------')
print(proportion[-10:])
# + colab={"base_uri": "https://localhost:8080/"} id="-L17nJZLKqAx" outputId="445d3e93-8c4c-4193-d07b-1097dbbc3ac4"
threshold = 70 # Arbitrairy threshold to determine if a column is usable
print(f'We choose to keep the columns with more than {threshold}% of filled values, which are {len(proportion[proportion > threshold])} columns :')
print(proportion[proportion > threshold].index.to_list())
# -
# Finally, we chose to focus on columns that represent weight per 100g as they seem to have a great completion rate and seem relevant and consitent enough for our analysis.
#
# Now, let's enter the cleaning phase. This is a required step, because if each product are false or missing, we won't be able to evaluate them and rank them as the user would like.
# + [markdown] id="uLDiRkSFhrqb"
# # Data cleaning
# -
# First, we must remove the duplicates.
# A duplicate is when an identifier column (which is supposed to has only unique values) contains two similar values. This is the case for the `code` column, so we chose to use the `url` column instead.
# ## Duplicates
# + colab={"base_uri": "https://localhost:8080/"} id="V8CgqFrfwKIK" outputId="078097b0-c808-40f4-99a6-5f061a68422f"
df_nan_code = pd.isna(df['code'])
df_nan_url = pd.isna(df['url'])
df_code_possible = df[df_nan_code != df_nan_url]
print(f"There are {df_code_possible.shape[0]} rows where we can compute either the code or url columns.")
# The purpose of this tool is to redirect the user to the openfoodfacts website, if the url is missing and we cannot reconstruct it from the code, we choose to remove the row.
df = df.drop(df[df_nan_code].index)
# Ultimately, even if the code columns contains duplicates, as we choose to use the url column as Id, this is not useful anymore to clean the code column.
df_duplicated_url = df[df['url'].duplicated()].sort_values('url')
print(f"There are {df_duplicated_url.shape[0]} rows where urls are duplicated. We deleted them.")
df = df.drop(df_duplicated_url.index)
# -
# But unfortunately, duplicates aren't the worst part of data cleaning.
# Now, let's head to the incoherent data section.
# ## Incoherent values
# + [markdown] id="G2V660dSAC8F"
# In this chapter, we will focus on selection of incoherent data to remove them from the dataset.
# Mainly, we will sum every 'on 100g' columns to see if it sums above 100g. If this is the case, we cannot determine which one (or which one**s**) of the addends is/are the outlier(s) and we must delete the entire row.
#
# Another problem is that, most of the 'on 100g' columns are included in anothers. For exemple, it would be an error to sum 'saturated_fat_100g' and 'fat_100g' as 'saturated_fat_100g' is included in 'fat_100g'.
# We must try to understand which columns contains the others.
# For this exercice, I needed to do a lot of reasearch on biochemistry and nutrition.
#
# I recommend you to look at the dev notebook to see how I did it, because it was a complex process and I won't go into details here.
# + id="OFQjpDzjCWt_"
# Using our reasearch, this is the dictionnary we came with
dict_feature_combinations = {
'fat_100g':
{
'cholesterol_100g':{},
'saturated-fat_100g':{
'caprylic-acid_100g':{},
'lauric-acid_100g':{},
'myristic-acid_100g':{},
'palmitic-acid_100g':{},
'stearic-acid_100g':{},
'arachidic-acid_100g':{},
'behenic-acid_100g':{},
'lignoceric-acid_100g':{},
'cerotic-acid_100g':{},
'montanic-acid_100g':{},
'melissic-acid_100g':{},
'butyric-acid_100g':{},
'caproic-acid_100g':{},
'capric-acid_100g':{}
},
'monounsaturated-fat_100g':{
'omega-9-fat_100g':{
'oleic-acid_100g':{},
'elaidic-acid_100g':{},
'gondoic-acid_100g':{},
'mead-acid_100g':{},
'erucic-acid_100g':{},
'nervonic-acid_100g':{}
}
},
'polyunsaturated-fat_100g':{
'omega-3-fat_100g':{
'alpha-linolenic-acid_100g':{},
'eicosapentaenoic-acid_100g':{},
'docosahexaenoic-acid_100g':{}
},
'omega-6-fat_100g':{
'linoleic-acid_100g':{},
'arachidonic-acid_100g':{},
'gamma-linolenic-acid_100g':{},
'dihomo-gamma-linolenic-acid_100g':{}
}
},
'trans-fat_100g':{}
},
'sugars_100g':
{
'carbohydrates_100g':{
'sucrose_100g':{},
'glucose_100g':{},
'fructose_100g':{},
'lactose_100g':{},
'maltose_100g':{}
}
},
'proteins_100g':{
'casein_100g':{},
}
}
# This function can either return every string of a dictionnary of return every columns for every levels (depending of the choosen option)
def multi_purpose_function(dico, option):
# Option 1: get_every_string_of_dict
# Option 2: get_levels_features
parent =''
level_list = [set(dico.keys())]
parent_list = [set()]
string_list = list(dico.keys())
# Prends toutes les clefs niveau 1
keys = list(dico.keys())
# Tant que la boite à clef à explorer n'est pas vide, on continue à explorer
while (keys != []):
# Pour chaque clef de la boite à clefs
for key in keys:
# On explore
# On décompose la clef pour avoir le multi level : On a une clef du type lvl1;lvl2;lvl3...
sublevel = dico
string_to_print = ''
level = 0
for key_level in key.split(';'):
string_to_print += '--'
level += 1
sublevel = sublevel[key_level]
if level >= len(level_list):
level_list.append(set())
if level >= len(parent_list):
parent_list.append(set())
# On retourne toutes les clefs (sans les dict([]).keys())
ajout_list = False
prochaines_clefs = list(sublevel.keys())
if len(prochaines_clefs) != 0:
parent = prochaines_clefs[0]
for key2 in prochaines_clefs:
keys.append(f"{key};{key2}")
string_list.append(key2)
level_list[level].add(key2)
if sublevel[key2] != dict([]):
level_list[level].add(key2)
else:
parent_list[level].add(key2)
# On retire la clef explorée de la boîte à clef
keys.remove(key)
for i in range(1,level):
parent_list[i] = parent_list[i].union(parent_list[i-1])
level_list[i] = level_list[i].union(parent_list[i-1])
if option == 1:
return string_list
if option == 2:
level_list.pop()
return level_list
# + id="kSklLdU3Ardg"
per_100g_features_list = []
# Get every 'for 100g' column
for index in data.columns:
if '100' in index and data[index].dtypes == 'float64':
per_100g_features_list.append(index)
# Here we delete the columns that doesn't seem to fit (most of them are just not weights so no possibility to sum them)
not_weight_on_100g_columns = [
'energy_100g',
'energy-from-fat_100g',
'carbon-footprint_100g',
'nutrition-score-fr_100g',
'nutrition-score-uk_100g',
'glycemic-index_100g',
'water-hardness_100g',
'ph_100g',
'collagen-meat-protein-ratio_100g',
]
for col in not_weight_on_100g_columns :
per_100g_features_list.remove(col)
# In order not to interfere with the process, every value over 100g is considered as NaN (so will cound as 0 in the sum).
# This is not a problem, as it will be replaced later in the outlier selection process.
for col in per_100g_features_list:
df.loc[df[col] > 100, [col]] = np.nan
# We select the columns that only are not composed of other columns (the deepest ones)
for feature in multi_purpose_function(dict_feature_combinations,1):
per_100g_features_list.remove(feature)
# We drop every column that we feel not to fit in the sum for different reasons
col_to_drop = [
'maltodextrins_100g', # Composed of glucose and fructose. Don't know where to put it.
'starch_100g', # Disn't find what it is.
'polyols_100g', # Organic component. Isn't a component of the food.
'serum-proteins_100g', # Protein coding gene. Don't know where to put it.
'nucleotides_100g', # Nucleic acid component. Don't know where to put it.
'beta-carotene_100g', # Precursor of the synthesis of vitamin A. We already study vitamin A.
'folates_100g', # Same as B9 vitamine
]
for col in col_to_drop:
per_100g_features_list.remove(col)
# These columns are the deepest ones
per_100g_invar_cols = per_100g_features_list
# + colab={"base_uri": "https://localhost:8080/", "height": 904} id="z471TTwa29Rd" outputId="8bc6cd75-5ef2-463e-8a44-7eb768e265eb"
# Time to do the sums
set_index_surcharge = set()
for i, colonne_list_variable in enumerate(multi_purpose_function(dict_feature_combinations,2)):
colonne_nom = 'somme_100g_n' + str(i)
# We sum the deepest one with the level 'columns'
colonne_list = per_100g_invar_cols + list(colonne_list_variable)
df[colonne_nom] = df[colonne_list].sum(axis=1)
set_index_surcharge = set_index_surcharge.union(set(df[(df[colonne_nom] > 100) | (df[colonne_nom] < 0)].index))
# When the weight is over 100g for 100g of product, we don't know what columns are wrong so we must delete the entire row
print('After calculating the sums over 100g of each product, we can delete the outlier rows :')
print(f"We delete {len(set_index_surcharge)} rows on {df.shape[0]} which makes {round(len(set_index_surcharge)*100/df.shape[0],2)}% of the initial dataset.")
print(f"There will be left {df.shape[0]-len(set_index_surcharge)} rows (which is enough to work with).")
df.drop(set_index_surcharge,inplace=True)
# -
# After cleaning the duplicates and the incoherent data, let's head to the outliers !
# ## Outliers
# [Outlier definition](https://www.wikiwand.com/en/Outlier): In statistics, an outlier is a data point that differs significantly from other observations.
#
# Outlier selection is a very important but very fastifious step in data cleaning. In other words, there is no need to remove values if we won't use them later.
# This is the reason why, in the first step, we will focus on electing the useful columns in order to clean them.
# + [markdown] id="anDYsa4yQF0M"
# ### Interesting features selection
# -
# We said earlier that we wanted to focus on columns above the chosen threshold and representing a weight on 100g.
#
# This is why we chose, as arbitrarly as for the threshold, to keep only the following columns :
# + id="4_0D1BZFQOcV"
# We gather the most interesting features (columns) and set options for them that will serve later
filter_features = pd.DataFrame(data=[
['fiber_100g',True, 0,50,0,20],
# ['cholesterol_100g',False],
['trans-fat_100g',False, 0,1,0,1],
['calcium_100g',True,0,2,0,2],
['iron_100g',True,0,0.2,0,0.04],
# ['energy_100g',True],
['proteins_100g',True,0,90,0,30],
# ['salt_100g',False],
# ['sodium_100g',False],
['salt_proc_100g',False,0,100,0,20],
['vitamins_count',True,0,11,0,11]
],
columns = ['feature', 'shouldIMaximiseIt', 'min_lim', 'max_lim', 'min_lim_arbitrary', 'max_lim_arbitrary'])
print(filter_features['feature'].tolist())
# -
# If you have a sharp eye, you'll have notice that we chose the column `vitamins_count`, which is not a weight on 100g, but that is NOT EVEN in the dataset !! You will also notice salt_proc_100g, is also a made up feature.
#
# This is because taking columns from the dataset is great, but we also should create our own variable in order to increase accuracy in our model and give more features to rank the products with.
# + [markdown] id="9pIu1NwiezE2"
# ### Artificial features creation
# + [markdown] id="873WkcFEe7sh"
#
# #### Counting the vitamins
#
# -
# Our first feature isn't that complicated to understand. Instead of leaving the vitamins columns away because they don't have enough data (remember the 70% threshold), we cound missing value as an abscence of vitamin and we sum for each product the number of vitamins.
# + id="XeGdARjYJxRc"
vitamin_columns = []
for index in data.columns:
if index[0] == 'v' : vitamin_columns.append(index)
# Also adding new vitamins that aren't labeled as 'vitamins'
# Vitamine B5
vitamin_columns.append('pantothenic-acid_100g')
# Vitamine B8
vitamin_columns.append('biotin_100g')
# Create a new column 'vitamins_count' that will count the number of vitamins in the product
vitamins_bool_isna = pd.notna(df[vitamin_columns])
df['vitamins_count'] = vitamins_bool_isna.sum(axis=1)
# + [markdown] id="hYD-NWILgCV4"
# The second artificial feature is more of a combination.
#
# In the initial dataset, they were two columns named `salt_100g` and `sodium_100g`. These two columns counts more or less the same thing, the salt proportion, but not the same component.
# We can find [here](https://www.wikiwand.com/en/Sodium_chloride) that salt (Sodium Chloride) is composed by 39.34% of sodium and 60.66% of chloride. So `salt = sodium * (100/39.34)`.
# In order to be able to use the right column, we chose to create a new column named `salt_proc_100g` which is the same value as the salt column if filled, or the calculation with the sodium in the other case.
# + id="TiC4l-c_iqcn"
salt_columns = ['salt_100g', 'sodium_100g']
# NaN values in the salt rows
rows_where_salt_na = df['salt_100g'].isna()
# but filled value in the sodium row
rows_where_sodium = df['sodium_100g'].notna()
rows_where_must_calculate_salt = df[rows_where_salt_na & rows_where_sodium].index.tolist()
def _fill_salt_proc_100g_column(x):
# If there is no salt but sodium, we return the operation, in the otehr case, we simply return the salt value
return x['sodium_100g'] * (100/39.34) if x.name in rows_where_must_calculate_salt else x['salt_100g']
df['salt_proc_100g'] = df.apply(lambda x: _fill_salt_proc_100g_column(x), axis=1)
# + [markdown] id="P4XkDl5QQx1f"
# ### Outliers selection in selected features
# + [markdown] id="uKY9WMexQ1B4"
# Now that we have the features we want to use for our ranking, we can start the detection of outliers.
# But what outliers means, for each feature ? Is 30g for 100g of fiber an outlier or not ?
# Well, if you're not a nutritionist or a biochemist, you'll probably not know and have to search by yourself.
#
# This is why I had to search on the web reliable sources for every low and high limits for each feature (and this is when I realised doing it for only a subset of the features was a great idea).
#
# Fortunately, I found a report of the [FoodData Central Dataset of the US department of Agriculture](https://fdc.nal.usda.gov/fdc-app.html#/) which contains a lot of products and their nutritional information. And this is how I achieve to gather every possible limits for each feature. Later, after the visualisation of the distribution of each feature (you'll see it later, don't worry), I also submitted some more 'arbritary' limits.
# -
export_df(filter_features.drop('shouldIMaximiseIt',axis=1), 'filter_features.png')
# 
# + id="Rc99b848c__r"
df_without_outliers = df.copy()
df_without_outliers_sharp = df.copy()
for index, feature in filter_features.iterrows() :
feature_name = feature['feature']
lim_bas_sharp = feature['min_lim_arbitrary']
lim_haut_sharp = feature['max_lim_arbitrary']
lim_bas = feature['min_lim']
lim_haut = feature['max_lim']
conditions = (df_without_outliers[feature_name] > lim_haut) | (df_without_outliers[feature_name] < lim_bas)
conditions_sharp = (df_without_outliers_sharp[feature_name] > lim_haut_sharp) | (df_without_outliers_sharp[feature_name] < lim_bas_sharp)
# Colonnes we will display over the plots
display_columns = [feature_name,'product_name','brands', 'code']
df_without_outliers.loc[conditions, feature['feature']] = np.nan
df_without_outliers_sharp.loc[conditions_sharp, feature['feature']] = np.nan
# + [markdown] id="ULR3FTYqV7Zh"
# ### Outliers visualization
# -
# This is the moment we've been waiting for, we'll able to see the distribution of our data and the removal of our outliers.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="7M-9KnMWV8x-" outputId="6d58535a-c22a-4277-f45b-c353ed7da984"
# We will visualize our outliers using boxplots
# Initialise the subplot function using number of rows and columns
figure, axis = plt.subplots(filter_features.shape[0], 3, figsize=(20, 20))
cols = ['Not cleaned', "Cleaned", "Sharply cleaned"]
figure.suptitle('Visualisation of the outliers', fontsize=20)
for ax, col in zip(axis[0], cols):
ax.set_title(col)
sns.set_theme(style="whitegrid")
for index, column in filter_features.iterrows():
# Keeping the oultiers
sns.boxplot(ax=axis[index,0], x=df[column['feature']])
# Removing the oultiers
sns.boxplot(ax=axis[index,1], x=df_without_outliers[column['feature']])
# Removing the oultiers with more fine limits
sns.boxplot(ax=axis[index,2], x=df_without_outliers_sharp[column['feature']])
figure.tight_layout()
plt.show()
# + [markdown] id="sX1RxKo3VdJg"
# Then, we had to drop the outliers, and chosing the limits. Do we prefer, the fine sharper one, or the one found on the Food Central dataset ?
#
#
# + id="DnJOdHXLVcbn"
# Shoud we use sharp cleaning ?
use_sharp_limits = True
print(f"We will use sharp limits ? {'Yes' if use_sharp_limits else 'No'}")
df = df_without_outliers_sharp if use_sharp_limits else df_without_outliers
# -
# But keep in mind that, unlike the incoherent values (the sums over 100g), here we know which value is the outlier and we can remove the value itself, not deleting the entire row. We just have to replace the outlier by a missing value and keep the other values in the same row.
# + [markdown] id="MFlvZdzVeg3w"
# ## Missing values treatment
#
#
# + [markdown] id="W3baZM1bep2l"
# After removing any duplicates, incoherent values and outliers, we now have to deal with missing values.
# Here, identifying them is not a problem because they all have the same value, np.nan.
# But the real problem is to replace them by a value that we will predict !
#
# We will use two methods of prediction :
#
# * The first is the regression method : more accurate but requires more data. The process isn't that hard, just give a lot of non-missing values to a model that will try to guess a linear equation between all the features and later will be able to predict your taget feature if you give it the same parameters as the training ones.
#
# e.g. : You want your model to give you the perfect proportion of ingredient to bake a cake for a given number of people. You'll give it the recipe for 4 people, 6 people, 8 people, etc... And it'll learn that it is proportionnal and when you'll need to bake for 20 people, it'll give you the perfect proportion of ingredient. But, it won't work if you don't tell it that you want to bake for 20 people. It won't be able to guess.
#
# * In the other hand, the second method, the imputation method, is simpler and won't require any other variable to guess. It'll just take the mean/median/best value of the column and copy it to the missing values. It is not very accurate because it doesn't take in account the other variable of this row, but it won't make a huge difference in the distribution.
#
# e.g. : A student have been missing at an exam and you want to predict his grade. You'll give the grades of the other students and it'll be able to predict his grade by doing the mean of the other grades.
#
# In order to have the most accurate data, we will do imputation only on subset of the dataset filtered by group (another variable).
#
#
# In priority, we want to use the first method, but as you can see, you need a lot of data because, firstly, the model need to understand the correlated variables (you don't need to know the hour of the day to bake your cake, but the number of people is important) and then it needs to have those data to predict the missing value.
# If we don't have those data, we'll use the second model.
# + id="GPeWhE6XRJUa"
def choose_most_related_features(df, target, nb, features, quiet = True):
if not quiet : print('\nChoose_most_related_features')
if target in features : features.remove(target)
features_list = features + [target]
corr = df[features_list].corr()
corr = corr.drop(target)
correlated_features = []
for i in range(nb):
feature_label = corr[target].abs().idxmax()
if not quiet : print(f"-- Feature selected n°{i+1} : {feature_label} with corr {round(corr[target][feature_label],3)*100}%")
correlated_features.append(feature_label)
corr = corr.drop(feature_label)
return correlated_features
# + id="pMwIrYW01iIn"
def make_imputation(df, target, method, quiet = True):
if not quiet : print('\nImputation')
df['pnns_groups_2'] = df['pnns_groups_2'].apply(lambda x : 'nan' if pd.isna(x) else x) # Do this because if not, it is impossible select the nan group
for group2 in df['pnns_groups_2'].unique():
sub_df = df[df['pnns_groups_2'] == str(group2)][target]
if not quiet : print(f"------ {group2} --> {sub_df.shape[0]} row and {sub_df.isna().values.sum()} imputations found !")
sub_df[sub_df.isna()]['target_imputed'] = 1
imputer = SimpleImputer(missing_values=np.nan, strategy=method)
imp_ser = imputer.fit_transform(sub_df.values.reshape(-1, 1))
df.loc[df['pnns_groups_2'] == group2, target] = imp_ser
return df
# + id="b56TG8jer97D"
def make_regression(df, features, target, quiet = True):
if not quiet : print('\nRegression')
# We split the dataset into two groups, the one where the target is filled (to train the regression) and the other where the target is missing (to make the prediction).
columns_used = features + [target]
train_df = df[columns_used]
train_df = train_df.dropna()
train_features = train_df[features]
train_target = train_df[target]
predict_df = df[df[target].isna()]
predict_df = predict_df[features].dropna()
if (predict_df.shape[0] == 0 and not quiet):
print('-- Not enough valid features to make any prediction. At least one feature in each prediction row is missing. Will do it by imputation.')
else:
if not quiet : print(f"-- {predict_df.shape[0]} rows eligible to prediction")
predict_features = predict_df[features]
X = train_features
y = train_target
regr = linear_model.LinearRegression()
if not quiet : print('---- fitting...')
regr.fit(X, y)
if not quiet : print('---- fitted')
predict_df[target] = regr.predict(predict_features)
df.loc[predict_df.index,target] = predict_df
return df
# + colab={"base_uri": "https://localhost:8080/"} id="oEtjNnnUp566" outputId="765a5062-2c4d-4210-b31c-c076c795d275"
nb_correlated_features = 5
interesting_features = proportion[proportion > threshold].index.tolist()
print('Doing the process of regression then imputation for each feature but only displaying the first one :')
for index, target in enumerate(filter_features['feature'].tolist()):
# On rajoute une colonne flag pour si la target a été imputed ou non.
# Ca sert pour amélriorer la regression.
# Ce flag est reset pour chaque nouvelle target
df['target_imputed'] = 0
if index == 0 : print(f"_________________________________________________________________\nFilter n°{index+1} : {target}")
most_related_features = choose_most_related_features(df, target , nb_correlated_features, interesting_features, quiet = False if index == 0 else True)
if index == 0 : print(f"\n{df[target].isna().sum()} targets left to predict")
regression_df = make_regression(df, most_related_features, target, quiet = False if index == 0 else True)
if index == 0 : print(f"\n{regression_df[target].isna().sum()} targets left to impute")
imputed_df = make_imputation(regression_df, target, 'mean', quiet = False if index == 0 else True) # Faire une moyenne en prenant la même catégorie de produits
assert imputed_df[target].isna().sum() == 0, f"imputation failed, there are still missing values in {target}"
df.loc[:,target] = imputed_df[target]
# -
# Ok ! The cleaning is done ! It has been a long process but now we can look at a cleaned and full dataset and try to make the ranking we wanted.
# First, let's take a look at the distribution of the features to see if there is some important behaviour to notice.
# + [markdown] id="AXYzruelW1rX"
# # Exploratory analysis of the cleaned dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 551} id="N7xJVN1lXgKA" outputId="1f611a09-2a33-4126-bbec-59d8604b93b0"
corr = df[interesting_features].corr()
# To show heatmap
fig, axs = plt.subplots(1,1,figsize=(5,5))
fig.suptitle('Feature correlation heatmap', fontsize=20)
sns.heatmap(corr)
plt.show()
# -
# An interesting fact is that ingredient_that_may_be_from_palm oil is greatly correlated with additives.
# This is not a correlation we exploit later but deserves to be noticed.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Yj37lbaueBAR" outputId="2966f576-95d5-4db2-b138-62cb4be7f155"
# Analysis of the data repartition on the interesting features columns after cleaning
# Initialise the subplot function using number of rows and columns
figure, axis = plt.subplots(filter_features.shape[0], 2, figsize=(20, 20))
figure.suptitle('Repartition of the data after cleaning')
cols = ['Histogram','Boxplot']
for ax, col in zip(axis[0], cols):
ax.set_title(col)
sns.set_theme(style="whitegrid")
for index, column in filter_features.iterrows():
# histograme
sns.histplot(ax=axis[index,0], x=df[column['feature']])
# Boxplot
sns.boxplot(ax=axis[index,1], x=df[column['feature']])
figure.tight_layout()
plt.show()
# + [markdown] id="PnTm8Rugoi49"
# We can notice that the linear regression created some outliers (eg : negative values of fiber). We know they were added with regression because we cleaned the oultiers before doing any prediction. Or, with cleaned data (over 0g), imputation is not able to predict negative value but linear regression is.
# + [markdown] id="yHKfnbacVT4v"
# ## Principal component analysis
# -
# After visualizing the distribution of the features, we'll briefly try to find some 'regrouping features' that will help us to reduce the dimension of our dataset. It is called PCA.
# + id="5m5lGwgCrYb4"
# Functions definition
# Taken from here https://github.com/stenier-oc/realisez-une-analyse-de-donnees-exploratoire/blob/master/functions.py
def display_circles(pcs, n_comp, pca, axis_ranks, labels=None, label_rotation=0, lims=None):
for d1, d2 in axis_ranks: # On affiche les 3 premiers plans factoriels, donc les 6 premières composantes
if d2 < n_comp:
# initialisation de la figure
fig, ax = plt.subplots(figsize=(7,6))
# détermination des limites du graphique
if lims is not None :
xmin, xmax, ymin, ymax = lims
elif pcs.shape[1] < 30 :
xmin, xmax, ymin, ymax = -1, 1, -1, 1
else :
xmin, xmax, ymin, ymax = min(pcs[d1,:]), max(pcs[d1,:]), min(pcs[d2,:]), max(pcs[d2,:])
# affichage des flèches
# s'il y a plus de 30 flèches, on n'affiche pas le triangle à leur extrémité
if pcs.shape[1] < 30 :
plt.quiver(np.zeros(pcs.shape[1]), np.zeros(pcs.shape[1]),
pcs[d1,:], pcs[d2,:],
angles='xy', scale_units='xy', scale=1, color="grey")
# (voir la doc : https://matplotlib.org/api/_as_gen/matplotlib.pyplot.quiver.html)
else:
lines = [[[0,0],[x,y]] for x,y in pcs[[d1,d2]].T]
ax.add_collection(LineCollection(lines, axes=ax, alpha=.1, color='black'))
# affichage des noms des variables
if labels is not None:
for i,(x, y) in enumerate(pcs[[d1,d2]].T):
if x >= xmin and x <= xmax and y >= ymin and y <= ymax :
plt.text(x, y, labels[i], fontsize='14', ha='center', va='center', rotation=label_rotation, color="blue", alpha=0.5)
# affichage du cercle
circle = plt.Circle((0,0), 1, facecolor='none', edgecolor='b')
plt.gca().add_artist(circle)
# définition des limites du graphique
plt.xlim(xmin, xmax)
plt.ylim(ymin, ymax)
# affichage des lignes horizontales et verticales
plt.plot([-1, 1], [0, 0], color='grey', ls='--')
plt.plot([0, 0], [-1, 1], color='grey', ls='--')
# nom des axes, avec le pourcentage d'inertie expliqué
plt.xlabel('F{} ({}%)'.format(d1+1, round(100*pca.explained_variance_ratio_[d1],1)))
plt.ylabel('F{} ({}%)'.format(d2+1, round(100*pca.explained_variance_ratio_[d2],1)))
plt.title("Cercle des corrélations (F{} et F{})".format(d1+1, d2+1))
plt.show(block=False)
def display_factorial_planes(X_projected, n_comp, pca, axis_ranks, labels=None, alpha=1, illustrative_var=None):
for d1,d2 in axis_ranks:
if d2 < n_comp:
# initialisation de la figure
fig = plt.figure(figsize=(7,6))
# affichage des points
if illustrative_var is None:
plt.scatter(X_projected[:, d1], X_projected[:, d2], alpha=alpha)
else:
illustrative_var = np.array(illustrative_var)
for value in np.unique(illustrative_var):
selected = np.where(illustrative_var == value)
plt.scatter(X_projected[selected, d1], X_projected[selected, d2], alpha=alpha, label=value)
plt.legend()
# affichage des labels des points
if labels is not None:
for i,(x,y) in enumerate(X_projected[:,[d1,d2]]):
plt.text(x, y, labels[i],
fontsize='14', ha='center',va='center')
# détermination des limites du graphique
boundary = np.max(np.abs(X_projected[:, [d1,d2]])) * 1.1
plt.xlim([-boundary,boundary])
plt.ylim([-boundary,boundary])
# affichage des lignes horizontales et verticales
plt.plot([-100, 100], [0, 0], color='grey', ls='--')
plt.plot([0, 0], [-100, 100], color='grey', ls='--')
# nom des axes, avec le pourcentage d'inertie expliqué
plt.xlabel('F{} ({}%)'.format(d1+1, round(100*pca.explained_variance_ratio_[d1],1)))
plt.ylabel('F{} ({}%)'.format(d2+1, round(100*pca.explained_variance_ratio_[d2],1)))
plt.title("Projection des individus (sur F{} et F{})".format(d1+1, d2+1))
plt.show(block=False)
def display_scree_plot(pca):
scree = pca.explained_variance_ratio_*100
plt.bar(np.arange(len(scree))+1, scree)
plt.plot(np.arange(len(scree))+1, scree.cumsum(),c="red",marker='o')
plt.xlabel("rang de l'axe d'inertie")
plt.ylabel("pourcentage d'inertie")
plt.title("Eboulis des valeurs propres")
plt.show(block=False)
def plot_dendrogram(Z, names):
plt.figure(figsize=(10,25))
plt.title('Hierarchical Clustering Dendrogram')
plt.xlabel('distance')
dendrogram(
Z,
labels = names,
orientation = "left",
)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="b8uxA-GXqO5z" outputId="802e7f33-54ff-48bc-9d62-844fb18f3f0f"
# PCA realisation
from sklearn import decomposition
from sklearn import preprocessing
pd.set_option('display.max_rows', 10)
pd.set_option('display.max_columns', 10)
floatInterrestingFeatures = [col for col in df[interesting_features].columns if df[col].dtypes == 'float64']
data = df[filter_features['feature']]
# Selection of the number of PCA components
n_comp = 2
# Centering and Reduction
std_scale = preprocessing.StandardScaler().fit(data.values)
X_scaled = std_scale.transform(data.values)
# PCA calculations
pca = decomposition.PCA(n_components=n_comp)
pca.fit(X_scaled)
# Cumulated inertia
display_scree_plot(pca)
# Correlation circles
pcs = pca.components_
display_circles(pcs, n_comp, pca, [(0,1),(2,3),(4,5)], labels = np.array(data.columns))
# # Data projection (doesn't work properly)
# X_projected = pca.transform(X_scaled)
# display_factorial_planes(X_projected, n_comp, pca, [(0,1),(2,3),(4,5)], labels = np.array(data.index))
# plt.show()
# -
# These are not really easy to interpret, so we won't use them.
#
# In the technical notebook, you'll also find ANOVA and fisher's test to test for hypothesis but we won't include them here as they don't serve our main goal (ranking products by feature.)
# # Ranking
# You've been waiting for too long ! Time to rank them all !
#
# Please read carefully the following instructions to ensure you know how to get what you want.
#
# * Do you want a demo ? Click the Auto button and the computer will choose random parameters.
# * Do you wand to tune your own filtering ? Please provide the name of the product you want to rank, the filters you want to rank and the number of products you want to rank and click Manual !
# +
# %%capture
from IPython.display import HTML, display
# %pip install ipywidgets
import ipywidgets as widgets
plt.ioff()
# -
class App:
def __init__(self, df, filter_features):
self._df = df
self._nb_products_max = 10
self._nb_products_min = 3
self._list_score_col_label = []
self._filter_features = filter_features
self._product_list = self._create_product_list()
self._product_list_size = 0
self._old_product = []
self.product = []
self._product_input = self._create_product_input()
self._old_filters = []
self.filters = []
self._filters_list = self._create_filters_list()
self._filters_input = self._create_filters_input()
self._old_nb_products = self._nb_products_min
self.nb_products = self._nb_products_min
self._nb_products_input = self._create_nb_products_input()
self._manual_button = self.create_button_manual()
self._input_container = widgets.HBox([self._manual_button, self._product_input, self._filters_input, self._nb_products_input])
self._auto_button = self.create_button_auto()
self._plot_container = widgets.Output()
self.container = widgets.VBox([self._auto_button, self._input_container, self._plot_container],layout=widgets.Layout())
self._update_app()
@classmethod
def create_class(cls, df, filter_features):
return cls(df, filter_features)
def _create_product_input(self):
product_input = widgets.Combobox(
placeholder='Choose a product (in the list or not)',
options= self._product_list,
description='Combobox:',
ensure_option=False,
disabled=False
)
return product_input
def _create_filters_input(self):
filters_input = widgets.SelectMultiple(value = [], options = self._filters_list, allow_duplicates = True, description = 'Filters (select multiple)', disabled = False)
return filters_input
def _create_nb_products_input(self):
nb_products_input = widgets.IntSlider(value=5, min=self._nb_products_min, max=self._nb_products_max, step=1, description='Number of products to rank', id='test')
return nb_products_input
def create_button_auto(self):
button = widgets.Button(description='Auto')
button.on_click(self._launch_auto)
return button
def create_button_manual(self):
button = widgets.Button(description='Manual')
button.on_click(self._launch_manual)
return button
def _create_filters_list(self):
return self._filter_features['feature'].tolist()
def _create_product_list(self):
sub_df_value_counts = self._df['product_name'].value_counts() > 50
return sub_df_value_counts[sub_df_value_counts.values == True].index.tolist()
def _launch_auto(self, _):
filtersList = self._filter_features['feature'].tolist()
self.filters = np.random.choice(filtersList, size=np.random.randint(1,7), replace=False).tolist()
self._filters_input.value = self.filters
random_product = np.random.choice(self._product_list, size=1, replace=False)[0]
self.product = random_product
self._product_input.value = random_product
self.nb_products = np.random.randint(self._nb_products_min, self._nb_products_max)
self._nb_products_input.value = self.nb_products
self._update_app()
def _launch_manual(self, _):
self.filters = self._filters_input.value
self.product = self._product_input.value
self.nb_products = self._nb_products_input.value
self._update_app()
def _ranking(self, df, product_name, filters, nb_products, filter_features):
selected_df = df
product_list = selected_df[selected_df['product_name'].str.contains(product_name, case=False, na=False)]
list_score_col_label = set()
for index, feature in enumerate(filters):
should_maximise = self._filter_features.loc[self._filter_features['feature'] == feature,('shouldIMaximiseIt')].iloc[0]
print(f"feature is {feature} and should I maximise it ? {should_maximise}")
product_list[feature + '_rank'] = product_list[feature].rank(ascending=should_maximise)
list_score_col_label.add(feature + '_rank')
self._list_score_col_label = list_score_col_label
product_list['sum_scores_rank'] = product_list[list_score_col_label].sum(axis=1)
product_list['multiple_rank'] = product_list['sum_scores_rank'].rank()
best = product_list.nlargest(nb_products, 'multiple_rank')
self._product_list_size = product_list.shape[0]
return best
def _update_app(self, *args):
if ((len(args) > 0) and (args[0]['name'] == '_property_lock')):
match str(args[0]['owner'])[:8]:
case 'IntSlide':
self.nb_products = args[0]['old']['value']
case 'Combobox':
self.product = args[0]['old']['value']
case 'SelectMu':
filtersIndex = args[0]['old']['index']
self.filters = []
for index in filtersIndex:
self.filters.append(self._filters_list[index])
if (len(self.filters) and (self.product) and (self.nb_products)):
best = self._ranking(self._df, '' if len(self.product) == 0 else self.product, self.filters, self.nb_products, self._filter_features)
self._old_filters = self.filters
self._old_product = self.product
self._old_nb_products = self.nb_products
with self._plot_container:
self._plot_container.clear_output(wait=True)
plt.figure(figsize=(20,30))
if len(self._list_score_col_label) > 1:
new_perc_cols = set()
self._list_score_col_label.add('multiple_rank')
# Pour que ce soit plus lisible dans le graphique, on va noter chaque rank en pourcentage (ex : 6ème sur 10 --> (6/10) * 100 = au dessus de 60% de l'échantillon)
for feature_rank in self._list_score_col_label:
col_name = feature_rank + '_perc'
best[col_name] = best[feature_rank]*100/self._product_list_size
new_perc_cols.add(col_name)
else : # Si il n'y a qu'une seule colonne, on préfere afficher la colonne en question plutôt qu'un ranking par rapport aux autres
new_perc_cols = self.filters
# On rajoutera aussi aux brand names le nom du produit
best['new_name'] = best['brands'] + '\n' + best['product_name']
# Avec sns
best_sns = best.melt(id_vars="new_name")
best_sns = best_sns.drop(best_sns[~best_sns['variable'].isin(new_perc_cols)].index).sort_values('value', ascending=False)
fig, axs = plt.subplots(1,1,figsize=(len(new_perc_cols)*5,5))
sns.barplot(x='new_name', y='value', hue='variable', data=best_sns, ax=axs)
axs.tick_params(axis='x', rotation=90, labelsize=20)
fig_title = f"Ranking of {self.product} by {self.filters}"
axs.set_title(fig_title,fontsize=30)
axs.set_xlabel("Product name",fontsize=20)
axs.set_ylabel("Ranking (in %)",fontsize=20)
axs.set_ylim(bottom = 0)
axs.tick_params()
plt.show()
# +
app = App.create_class(df, filter_features)
app.container
| PresentationNotebookForVoila/Projet3Voila.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
from tensorflow.keras.applications import mobilenet_v2,MobileNetV2
from tensorflow.keras.layers import Dense, BatchNormalization,GaussianDropout,Dropout,SeparableConv2D,MaxPooling2D,GlobalAveragePooling2D
from tensorflow.keras.models import save_model,Model,Sequential
from tensorflow.keras.preprocessing.image import ImageDataGenerator,load_img,img_to_array
from tensorflow.keras.losses import BinaryCrossentropy,MSE
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import Adam,RMSprop,Adagrad
import numpy as np
# -
train_path=r"dataset"
val_path=r"Z:\covid19-repo\validation_set"
# +
shape=150
seed=np.random.seed()
# +
data=ImageDataGenerator(featurewise_center=False,samplewise_center=False,
featurewise_std_normalization=False,
samplewise_std_normalization=False,
zca_whitening=False,zca_epsilon=1e-06,
rotation_range=30,width_shift_range=0.0,
height_shift_range=0.0,brightness_range=None,
shear_range=0.0,zoom_range=0.2,channel_shift_range=0.0,
fill_mode="nearest", cval=0.0,
horizontal_flip=True, vertical_flip=True,
rescale=1./255,
preprocessing_function=None, data_format=None,
validation_split=0.4,
dtype='float32')
train_data=data.flow_from_directory(train_path, target_size=(shape,shape,), color_mode="rgb",class_mode="categorical", batch_size=10, shuffle=False,seed=seed,
save_to_dir=None, save_prefix="",save_format="png",
follow_links=False,subset=None,
interpolation="nearest")
# -
train_data.class_indices
# +
valdata=ImageDataGenerator(
rescale=1./255,
dtype='float32')
val_data=data.flow_from_directory(val_path, target_size=(shape,shape,), color_mode="rgb",
classes=lis,class_mode="categorical", batch_size=10, shuffle=False,seed=seed,
save_to_dir=None, save_prefix="",save_format="png",
follow_links=False,subset=None,
interpolation="nearest")
# -
val_data.class_indices
# model
basemodel = MobileNetV2(input_shape=(shape,shape,3),include_top=False,weights='imagenet')
basemodel.trainable=False
x=basemodel.output
x=GlobalAveragePooling2D()(x)
x=Dense(1024,activation='relu')(x) #we add dense layers so that the model can learn more complex functions and classify for better results.
x=Dense(1024,activation='relu')(x) #dense layer 2
x=Dense(512,activation='relu')(x) #dense layer 3
preds=Dense(3,activation='softmax')(x)
model=Model(inputs=basemodel.input,outputs=preds)
model.summary()
epoch=30
opt=Adam(learning_rate=0.0001,decay=0.0001 / epoch)
model.compile(opt,loss='binary_crossentropy',metrics=['acc'])
from tensorflow.keras.utils import plot_model
plot_model(model)
hist=model.fit_generator(train_data,epochs=epoch,validation_data=val_data)
model.evaluate(val_data)
img=load_img(r"Z:\covid19-repo\dataset\Viral Pneumonia\Viral Pneumonia-831.png")
# +
import cv2
img=img_to_array(img)/255
img=cv2.resize(img,(150,150))
img=img.reshape(-1,150,150,3)
# -
img.shape
pred=model.predict(img)
pred
df=list(train_data.class_indices)
df[np.argmax(pred)]
model.save('covid_model.h5')
from tensorflow.keras.models import load_model
pp=sf.predict(img)
df[np.argmax(pp)]
# # tflite conversion
import tensorflow as tf
saf=load_model('mymodel.h5')
converter = tf.lite.TFLiteConverter.from_keras_model(sf)
tflite_model = converter.convert()
open("my_model.tflite", "wb").write(tflite_model)
interpreter = tf.lite.Interpreter('covid_model.tflite')
interpreter.get_tensor_details()
| covid transfer learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import staintools
import csv
import os
import glob
import re
from pandas import DataFrame, Series
from PIL import Image
import timeit
import time
import cv2
from matplotlib import pyplot as plt
import numpy as np
train_paths = ["/scratch/kk4ze/data_lowres_1100x1100/valid/EE/"]
# get images
images = {}
images_by_folder = {}
for train_path in train_paths:
images_by_folder[str(train_path)] = []
files = glob.glob(os.path.join(train_path, '*.jpg'))
for fl in files:
flbase = os.path.basename(fl)
flbase_noext = os.path.splitext(flbase)[0]
images[flbase_noext]=fl
images_by_folder[str(train_path)].append(flbase_noext)
# initialize stain and brightness normalizer
stain_normalizer = staintools.StainNormalizer(method='vahadane')
standardizer = staintools.BrightnessStandardizer()
# choose target image
target_image = staintools.read_image("/scratch/kk4ze/data_lowres_1100x1100/train/Celiac/C03-05_03_5901_4803_horiz__0.jpg")
standard_target_image = standardizer.transform(target_image)
stain_normalizer.fit(standard_target_image)
# +
# get destination path
path_change_map = {}
for key in list(images_by_folder.keys()):
temp = key.replace('data_lowres_1100x1100', 'data_lowres_1100x1100_augmented')
path_change_map[key] = temp
# +
for key in images_by_folder.keys():
for value in list(images_by_folder[key]):
# print(key)
# print(value)
# print (str(count) + ' ' + str(value))
source_img_path = str(key) + str(value) + '.jpg'
value=value.replace('-','.')
dest_img_path = str(path_change_map[key]) + str(value) + '.jpg'
# print(source_img_path)
img = staintools.read_image(source_img_path)
if (np.mean(img) > 240) or (np.mean(img) < 10):
continue
# standardize brightness
img_standard = standardizer.transform(img)
# transform the images
img_normalized = stain_normalizer.transform(img_standard)
# write image to path
# plt.imshow(img)
# plt.title('my picture')
# plt.show()
# plt.imshow(img_normalized)
# plt.title('my picture')
# plt.show()
cv2.imwrite(os.path.normpath(dest_img_path), img_normalized)
# -
path_change_map
| image_augmentaion/lowres/transform_lowres_images_1100_valid_EE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
from PIL import Image
import glob
import os
import random
import imageio
# +
final_gif_name = "colloboartion_01"
final_pieces = glob.glob("*_final_*.png")
grid_size = 9
widths = []
heights = []
for i in range(len(final_pieces)):
im = Image.open(final_pieces[i])
widths.append(im.size[0])
heights.append(im.size[1])
w = min(heights)
h = min(widths)
# resize loop
for i in range(len(final_pieces)):
im = Image.open(final_pieces[i])
if im.size[0] != w or im.size[1] != h:
wpercent = (w/float(im.size[0]))
hsize = int((float(im.size[1])*float(wpercent)))
img = im.resize((w,hsize), Image.ANTIALIAS)
img.save(final_pieces[i].replace(".jpg", "") + str(w) + '_resized.png')
final_pieces[i] = final_pieces[i].replace(".jpg", "") + str(w) + '_resized.png'
# create gif folder and subdirectories
os.mkdir(final_gif_name)
os.mkdir(final_gif_name + "/diagionals/")
os.mkdir(final_gif_name + "/single_rows/")
os.mkdir(final_gif_name + "/full_grids/")
# +
# diagionals - in order for now
# odd numbers are right to left, even numbers left to right
grid_size = 9
file_suffix = 0
for i in range(0, len(final_pieces), grid_size):
print(i)
images = [Image.open(x) for x in final_pieces[i:i+grid_size]]
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = sum(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset += im.size[0]
y_offset += im.size[1]
new_im.save(final_gif_name + "/diagionals/" + str(file_suffix) + '.png')
file_suffix += 1
flipped_new_im = new_im.transpose(Image.FLIP_LEFT_RIGHT)
flipped_new_im.save(final_gif_name + "/diagionals/" + str(file_suffix) + '.png')
file_suffix += 1
# -
os.mkdir(final_gif_name + "/diagionals_seq/")
# +
grid_size = 9
file_suffix = 0
for i in range(0, len(final_pieces)):
print(i)
images = [Image.open(x) for x in final_pieces[i:i+grid_size]]
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = sum(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset += im.size[0]
y_offset += im.size[1]
new_im.save(final_gif_name + "/diagionals_seq/" + str(file_suffix) + '.png')
file_suffix += 1
flipped_new_im = new_im.transpose(Image.FLIP_LEFT_RIGHT)
flipped_new_im.save(final_gif_name + "/diagionals_seq/" + str(file_suffix) + '.png')
file_suffix += 1
# -
os.mkdir(final_gif_name + "/partial_grids/")
# +
# grid pieces
file_suffix = 0
for i in range(0, len(final_pieces), grid_size):
print(i)
images = [Image.open(x) for x in final_pieces[0+i:grid_size*grid_size+1+i]]
widths, heights = zip(*(i.size for i in images))
# assuming same size images
total_width = widths[0]*grid_size
total_height = heights[0]*grid_size
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for i in range(len(images)):
new_im.paste(images[i], (x_offset,y_offset))
x_offset += im.size[0]
if i !=0 and i%9 == 0:
x_offset = 0
y_offset += im.size[1]
new_im.save(final_gif_name + "/partial_grids/" + str(file_suffix) + '.png')
file_suffix += 1
# +
# grids - images randomly shuffeled
#file_suffix = 0
num_grids = 25
for i in range(num_grids):
print(i)
random.shuffle(final_pieces)
images = [Image.open(x) for x in final_pieces[0:grid_size*grid_size+1]]
widths, heights = zip(*(i.size for i in images))
# assuming same size images
total_width = widths[0]*grid_size
total_height = heights[0]*grid_size
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for i in range(len(images)):
new_im.paste(images[i], (x_offset,y_offset))
x_offset += im.size[0]
if i !=0 and i%grid_size == 0:
x_offset = 0
y_offset += im.size[1]
new_im.save(final_gif_name + "/full_grids/" + str(file_suffix) + "_" + str(grid_size) + '_.png')
file_suffix += 1
# +
# make images 3x larger
three_times_larger_images = []
for i in range(len(final_pieces)):
im = Image.open(final_pieces[i])
wpercent = (w/float(im.size[0]))
hsize = int((float(im.size[1])*float(wpercent)))
img = im.resize((3*w,3*hsize), Image.ANTIALIAS)
img.save(final_pieces[i].replace(".jpg", "") + str(w) + '_resized_3_times.png')
three_times_larger_images.append(final_pieces[i].replace(".jpg", "") + str(w) + '_resized_3_times.png')
# -
os.mkdir(final_gif_name + "/three_by_three_gifs/")
# +
num_grids = 25
grid_size = 3
for i in range(num_grids):
print(i)
random.shuffle(three_times_larger_images)
images = [Image.open(x) for x in three_times_larger_images[0:grid_size*grid_size+1]]
widths, heights = zip(*(i.size for i in images))
# assuming same size images
total_width = widths[0]*grid_size
total_height = heights[0]*grid_size
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for i in range(len(images)):
new_im.paste(images[i], (x_offset,y_offset))
x_offset += im.size[0]
if i !=0 and i%grid_size == 0:
x_offset = 0
y_offset += im.size[1]
new_im.save(final_gif_name + "/three_by_three_gifs/" + str(file_suffix) + "_" + str(grid_size) + '_.png')
file_suffix += 1
flipped_new_im = new_im.transpose(Image.FLIP_LEFT_RIGHT)
flipped_new_im.save(final_gif_name + "/three_by_three_gifs/" + str(file_suffix) + "_" + str(grid_size) + '_.png')
file_suffix += 1
flipped_new_im = new_im.transpose(Image.FLIP_TOP_BOTTOM)
flipped_new_im.save(final_gif_name + "/three_by_three_gifs/" + str(file_suffix) + "_" + str(grid_size) + '_.png')
file_suffix += 1
flipped_new_im = flipped_new_im.transpose(Image.FLIP_LEFT_RIGHT)
flipped_new_im.save(final_gif_name + "/three_by_three_gifs/" + str(file_suffix) + "_" + str(grid_size) + '_.png')
file_suffix += 1
# -
os.mkdir(final_gif_name + "/grids_3x3/")
# +
num_grids = 2
grid_size = 3
for i in range(num_grids):
print(i)
random.shuffle(final_pieces)
images = [Image.open(x) for x in final_pieces[0:grid_size*grid_size+1]]
widths, heights = zip(*(i.size for i in images))
# assuming same size images
total_width = widths[0]*grid_size
total_height = heights[0]*grid_size
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for i in range(len(images)):
new_im.paste(images[i], (x_offset,y_offset))
x_offset += im.size[0]
if i !=0 and i%grid_size == 0:
x_offset = 0
y_offset += im.size[1]
new_im.save(final_gif_name + "/grids_3x3/" + str(file_suffix) + "_" + str(grid_size) + '_.png')
# wpercent = (w*float(new_im.size[0]))
# hsize = int((float(new_im.size[1])*float(wpercent)))
# resize_new_im = new_im.resize((w,hsize), Image.ANTIALIAS)
# resize_new_im.save(final_gif_name + "/grids_3x3/" + str(file_suffix) + "_" + str(grid_size) + '_resize_.png')
file_suffix += 1
# -
os.mkdir(final_gif_name + "/cropped_9x9/")
# +
# take a 9 x 9 grid, crop and then make it 3 X 3
nine_by_nine = glob.glob(final_gif_name + "/full_grids/*.png")
image_file_name = nine_by_nine[0]
im = Image.open(image_file_name)
im.crop((float(im.size[0])/3, float(im.size[1])/3, 0, 0))
im.show()
# +
# single_rows folder
# https://stackoverflow.com/questions/30227466/combine-several-images-horizontally-with-python
# One 9 x 9 grid created from people's responces
# diaginal images, cool but not what intended
images = [Image.open(x) for x in final_pieces[3:12]]
# not resizing - figure out
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = sum(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset += im.size[0]
# y_offset += im.size[1]
new_im.save('test_combo_43.png')
new_im.show()
# -
# ## Create GIF
# +
final_gif_name = "colloboartion_01"
full_grids = sorted(glob.glob(final_gif_name + "/full_grids/*.png"))
diagionals = sorted(glob.glob(final_gif_name + "/diagionals/*.png"))
rotations = sorted(glob.glob(final_gif_name + "/three_by_three_gifs/*.png"))[40:]
# images_in_order = full_grids[10:20] + diagionals[0:10] + rotations[20:32] + full_grids[:10]
# with imageio.get_writer(final_gif_name + '_monster_intial_composites.gif', mode='I') as writer:
# for filename in images_in_order:
# try:
# image = imageio.imread(filename)
# writer.append_data(image)
# except:
# print(filename)
# +
final_gif_name = "intial_iteration"
images_in_order = full_grids[0:19]
with imageio.get_writer(final_gif_name + '_grids_composites.gif', mode='I') as writer:
for filename in images_in_order:
try:
image = imageio.imread(filename)
writer.append_data(image)
except:
print(filename)
# -
len(images_in_order)
# +
fp_out = final_gif_name + "_all_composites_4.gif"
img, *imgs = [Image.open(f) for f in images_in_order]
img.save(fp=fp_out, format='GIF', append_images=imgs,
save_all=True, duration=150, loop=0)
# -
len(full_grids), len(diagionals), len(rotations)
# +
# https://stackoverflow.com/questions/30227466/combine-several-images-horizontally-with-python
# One grid_size x grid_size grid created from people's responces
# all of them horizonitial
grid_size = 9
images = [Image.open(x) for x in final_pieces[0+40:grid_size*grid_size+1+40]]
# not resizing - figure out
widths, heights = zip(*(i.size for i in images))
# assuming same size images
total_width = widths[0]*grid_size
total_height = heights[0]*grid_size
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for i in range(len(images)):
new_im.paste(images[i], (x_offset,y_offset))
x_offset += im.size[0]
if i !=0 and i%9 == 0:
x_offset = 0
y_offset += im.size[1]
new_im.save('test_combo_square_2_' + str(grid_size) + '_.png')
new_im.show()
# -
81%9
18%9
# +
# https://stackoverflow.com/questions/30227466/combine-several-images-horizontally-with-python
# One 9 x 1 grid created from people's responces
# all of them horizonitial
images = [Image.open(x) for x in final_pieces[0+10:9+10]]
# not resizing - figure out
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = max(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset += im.size[0]
# y_offset += im.size[1]
new_im.save('test_combo_9.png')
new_im.show()
# +
# https://stackoverflow.com/questions/30227466/combine-several-images-horizontally-with-python
# One 9 x 1 grid created from people's responces
# all of them horizonitial
images = [Image.open(x) for x in final_pieces[3:12]]
# not resizing - figure out
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = max(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset += im.size[0]
# y_offset += im.size[1]
new_im.save('test_combo_33.png')
new_im.show()
# +
# https://stackoverflow.com/questions/30227466/combine-several-images-horizontally-with-python
# One 9 x 9 diag created from people's responces
# diaginal images, cool but not what intended
images = [Image.open(x) for x in final_pieces[3:12]]
# not resizing - figure out
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = sum(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = 0
y_offset = 0
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset += im.size[0]
y_offset += im.size[1]
new_im.save('test_combo_583747.png')
new_im.show()
# +
# https://stackoverflow.com/questions/30227466/combine-several-images-horizontally-with-python
# One 9 x 9 grid created from people's responces
# diaginal images, cool but not what intended
images = [Image.open(x) for x in final_pieces[3:12]]
# not resizing - figure out
widths, heights = zip(*(i.size for i in images))
total_width = sum(widths)
total_height = sum(heights)
new_im = Image.new('RGB', (total_width, total_height))
x_offset = total_width - im.size[0]
y_offset = total_height - im.size[1]
for im in images:
new_im.paste(im, (x_offset,y_offset))
x_offset -= im.size[0]
y_offset -= im.size[1]
flipped_new_im = new_im.transpose(Image.FLIP_LEFT_RIGHT)
flipped_new_im.save('test_combo_diag_flipped_right_left.png')
flipped_new_im.show()
# -
widths.index(max(widths))
# +
# https://stackoverflow.com/questions/273946/how-do-i-resize-an-image-using-pil-and-maintain-its-aspect-ratio
# reszie large image
from PIL import Image
basewidth = w
img = Image.open(final_pieces[widths.index(max(widths))])
wpercent = (basewidth/float(img.size[0]))
hsize = int((float(img.size[1])*float(wpercent)))
img = img.resize((basewidth,hsize), Image.ANTIALIAS)
img.save(final_pieces[widths.index(max(widths))].replace(".jpg", "") + str(w) + '_resized.png')
img.show()
# -
| Create_Collorative_Piece_GIF_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/anmol-sinha-coder/DEmoClassi/blob/master/Age_Gender_Race_Emotion_GPU.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g--abrgzUbl3"
from google.colab import drive
drive.mount('/content/G_Drive')
# + id="pnLx-juX2pBA"
# ! git clone https://github.com/anmol-sinha-coder/DEmoClassi.git
# ! cp -ra DEmoClassi/{vision_utils,emotion_detection,multitask_rag,'setup.py'} ./
# ! pip install tensorboardX pytorch-ignite pillow
# ! unzip /content/G_Drive/MyDrive/ADNN/facial-expression-recognition-challenge.zip -d .
# ! tar -xzvf /content/G_Drive/MyDrive/ADNN/UTKFace/UTKFace.tar.gz -C .
# ! tar -xzvf /content/G_Drive/MyDrive/ADNN/UTKFace/crop_part1.tar.gz -C .
# ! tar -xzvf fer2013.tar.gz
# ! cp /content/G_Drive/MyDrive/ADNN/cv2_gpu/cv2.cpython-36m-x86_64-linux-gnu.so .
# + id="yyilh9MmRaqB"
import torch
import torchvision.transforms as transforms
from vision_utils.custom_torch_utils import load_model
from vision_utils.custom_architectures import SepConvModelMT, SepConvModel, initialize_model, PretrainedMT
from emotion_detection.evaluate import evaluate_model as eval_fer
from emotion_detection.fer_data_utils import *
from emotion_detection.train import run_fer
from multitask_rag.train import run_utk
from multitask_rag.utk_data_utils import get_utk_dataloader, split_utk
from multitask_rag.evaluate import evaluate_model as eval_utk
from multitask_rag.utk_data_utils import display_examples_utk
import numpy as np
import pandas as pd
import glob
import os
import tqdm
import random
import cv2
cv2.__version__
# + [markdown] id="DYFah_cf7nNx"
# ## Fer2013 dataset
# Fer2013 is a kaggle dataset which consists of a set of 48x48 grayscale images representing the following facial expressions :
# * 0 : Angry
# * 1 : Disgust
# * 2 : Fear
# * 3 : Happy
# * 4 : Sad
# * 5 : Surprise
# * 6 : Neutral
# + id="iKF7mvSQUT3F"
path_fer = './fer2013/fer2013.csv'
df_fer2013 = pd.read_csv(path_fer)
# + id="RIA39F-P4mYK"
display_examples_fer(df_fer2013, 0)
# + id="M6lPU88E4oeY"
display_examples_fer(df_fer2013, 1)
# + id="s1rir-Xl7Eyh"
display_examples_fer(df_fer2013, 2)
# + id="Gehh2Xu97Tr1"
display_examples_fer(df_fer2013, 3)
# + id="23PMhdF_7XeA"
display_examples_fer(df_fer2013, 4)
# + id="mzTTdxxa7cFU"
display_examples_fer(df_fer2013, 5)
# + id="fOsVwgyz7gzs"
display_examples_fer(df_fer2013, 6)
# + [markdown] id="RwOH5aRq8KUe"
# ## UTKFace dataset
# This is a dataset of cropped face images for the task of predicting the age, gender and race of a person.<br>
#
# **Age :** A number between 0 and 101 (representing the age of the person)<br>
#
# **Gender :**
# * 0 : Male
# * 1 : Female
#
# **Race :**
# * 0 : White
# * 1 : Black
# * 2 : Asian
# * 3 : Indian
# * 4 : Other
#
# + id="o_eWQCEV7imj"
path_utk = './UTKFace/'
# + id="zX9IIREl9odh"
display_examples_utk(path_utk, 'gender', 0)
# + id="1lXMGe9L-HDd"
display_examples_utk(path_utk, 'gender', 1)
# + id="g7uMJbmT9qTc"
display_examples_utk(path_utk, 'race', 0)
# + id="41kiL5qF9vdW"
display_examples_utk(path_utk, 'race', 1)
# + id="IlAB1hrY90iC"
display_examples_utk(path_utk, 'age', 10)
# + [markdown] id="CPHCrAPA-MZG"
# # Training
#
# Now that we have the data ready, let's move to the funniest part : model training!
# As I have two separate datasets (`Fer2013` for emotion detection and `UTKFace` for gender-race-age prediction) we'll
# have to train two separate models. For each of the two tasks I tested 3 different architectures :
# * A CNN based on Depthwise Separable Convolution
# * Finetuning a pretrained Resnet50
# * Finetuning a pretrained VGG19
#
# <hr size=10 color=black>
# + id="I6_DbSPh6Uen"
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device, torch.cuda.is_available()
# + id="1R2O1XyQCXtJ"
DATA_DIR = "./fer2013/fer2013.csv" # path to the csv file
BATCH_SIZE = 256 # size of batches
train_flag = 'Training' #`Usage` column in the csv file represents the usage of the data : train or validation or test
val_flag = 'PublicTest'
# + [markdown] id="1_UgNyaqogxD"
#
# ## (1)Training Emotion/Reaction detector
# ### [1.a] Depthwise Separable Convolution model
# First we need to create DataLoader objects which are handy Pytorch objects for yielding batches of data during training.
# Basically, what the following code does is :
# * read the csv file and convert the raw pixels into numpy arrays
# * Apply some pre-processing operations :
# * Histogram equalization
# * Add a channel dimension so that the image becomes 48x48x1 instead of 48x48
# * Convert the numpy array to a pytorch tensor
# + id="Wja9Wctg94b-"
# The transformations to apply
data_transforms = transforms.Compose([
HistEq(), # Apply histogram equalization
AddChannel(), # Add channel dimension to be able to apply convolutions
transforms.ToTensor()
])
train_dataloader = get_fer_dataloader(BATCH_SIZE, DATA_DIR, train_flag, data_transforms=data_transforms)
validation_dataloader = get_fer_dataloader(BATCH_SIZE, DATA_DIR, val_flag, data_transforms=data_transforms)
my_data_loaders = {
'train': train_dataloader,
'valid': validation_dataloader
}
backup_path = '/content/G_Drive/MyDrive/ADNN/Demography_Psychology/Separable_Convolutional_Model'
os.makedirs(backup_path, exist_ok=True) # create the directory if it doesn't exist
checkpoint = '/content/checkpoints/fer_sep_conv_histeq' # folder where to save checkpoints during training
os.makedirs(checkpoint, exist_ok=True)
# + id="3CXvacNY_XG6"
my_model = SepConvModel().to(device)
my_optimizer = torch.optim.Adam(my_model.parameters(), lr=1e-3)
# + id="L4IP0kztGlk7"
# Evaluation of model
run_fer(model=my_model, optimizer=my_optimizer, epochs=300,
log_interval=1, dataloaders=my_data_loaders,
dirname=checkpoint, filename_prefix='FER-Sep_Conv',
n_saved=1, log_dir=None, launch_tensorboard=False,
patience=50, resume_model=None, resume_optimizer=None,
backup_step=5, backup_path=backup_path,
n_epochs_freeze=0, n_cycle=None)
# + [markdown] id="ogF-GBWMsK4s"
# ### [1.b] Resnet-50
# Over the years there is a trend to go more deeper, to solve more complex tasks and to also increase /improve the classification/recognition accuracy. But, as we go deeper; the training of neural network becomes difficult and also the accuracy starts saturating and then degrades also. Residual Learning tries to solve both these problems.
#
# - In general, in a deep convolutional neural network, several layers are stacked and are trained to the task at hand.
# - The network learns several low OR mid OR high level features at the end of its layers.
# - In residual learning, instead of trying to learn some features, we try to learn some residual. Residual can be simply understood as subtraction of feature learned from input of that layer.
# - ResNet does this using shortcut connections (directly connecting input of nth layer to some (n+x)th layer.
# - It has proved that training this form of networks is easier than training simple deep convolutional neural networks and also the problem of degrading accuracy is resolved.
# + id="EHxOdq9IG0QW"
# The transformations to
data_transforms = transforms.Compose([
HistEq(), # Apply histogram equalization
ToRGB(),
transforms.ToTensor()
])
my_data_loaders = {
'train': get_fer_dataloader(BATCH_SIZE, DATA_DIR, train_flag, data_transforms=data_transforms),
'valid': get_fer_dataloader(BATCH_SIZE, DATA_DIR, val_flag, data_transforms=data_transforms)
}
backup_path = '/content/G_Drive/MyDrive/ADNN/Demography_Psychology/ResNet_Model'
os.makedirs(backup_path, exist_ok=True)
checkpoint = '/content/checkpoints/fer_resnet_adam_histeq'
os.makedirs(checkpoint, exist_ok=True)
# + id="ivIuNwvsibFk"
my_model, _ = initialize_model(model_name='resnet', feature_extract=True, num_classes=7, task='fer2013', use_pretrained=True, device=device)
# The optimizer must only track the parameters that are trainable (thus excluding frozen ones)
my_optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, my_model.parameters()), lr=1e-3)
# + id="MCVPh91ok_4a"
run_fer(model=my_model, optimizer=my_optimizer, epochs=200,
log_interval=1, dataloaders=my_data_loaders,
dirname=checkpoint, filename_prefix='FER-Resnet',
n_saved=1, log_dir=None, launch_tensorboard=False,
patience=75, val_monitor='acc', resume_model=None,
resume_optimizer=None, backup_step=5, backup_path=backup_path,
n_epochs_freeze=10, n_cycle=None)
# + [markdown] id="jXnJFNrotwRx"
# ### [1.c] VGG-19
# VGG19 is a variant of VGG model which in short consists of 19 layers (16 convolution layers, 3 Fully connected layer, 5 MaxPool layers and 1 SoftMax layer).
#
# - Conv3x3 (64)
# - Conv3x3 (64)
# - MaxPool
# - Conv3x3 (128)
# - Conv3x3 (128)
# - MaxPool
# - Conv3x3 (256)
# - Conv3x3 (256)
# - Conv3x3 (256)
# - Conv3x3 (256)
# - MaxPool
# - Conv3x3 (512)
# - Conv3x3 (512)
# - Conv3x3 (512)
# - Conv3x3 (512)
# - MaxPool
# - Conv3x3 (512)
# - Conv3x3 (512)
# - Conv3x3 (512)
# - Conv3x3 (512)
# - MaxPool
# - Fully Connected (4096)
# - Fully Connected (4096)
# - Fully Connected (1000)
# - SoftMax
#
# + id="AQ2-8a_lr0BE"
# The transformations to
data_transforms = transforms.Compose([
HistEq(),
ToRGB(),
transforms.ToTensor()
])
my_data_loaders = {
'train': get_fer_dataloader(256, DATA_DIR, train_flag, data_transforms=data_transforms),
'valid': get_fer_dataloader(512, DATA_DIR, val_flag, data_transforms=data_transforms)
}
my_model, _ = initialize_model(model_name='vgg', feature_extract=True, num_classes=7, task='fer2013', use_pretrained=True, device=device)
my_optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, my_model.parameters()), lr=1e-3)
backup_path = '/content/G_Drive/MyDrive/ADNN/Demography_Psychology/VGGNet_Model'
os.makedirs(backup_path, exist_ok=True)
checkpoint = '/content/checkpoints/fer_vgg_adam_histeq'
os.makedirs(checkpoint, exist_ok=True)
# + id="65Lxp79ItgI8"
run_fer(model=my_model, optimizer=my_optimizer, epochs=300,
log_interval=1, dataloaders=my_data_loaders,
dirname=checkpoint, filename_prefix='FER-VGGnet',
n_saved=1, log_dir=None, launch_tensorboard=False,
patience=100, val_monitor='acc',
resume_model=None, resume_optimizer=None,
backup_step=5, backup_path=backup_path,
n_epochs_freeze=20, n_cycle=None)
# + [markdown] id="-uzkfmdpxyX4"
# <hr>
#
# # (2)Training Age, Gender, Race detector
# + id="VUab7N1g1Qqa"
# ! mkdir logs/
list_images = glob.glob('/content/UTKFace/*jp*')
print(len(list_images))
# + id="qPsL3jDc2S_H"
# Labels are given in the image names. the image names format is the following : age_gender_race_date.
# for instance this image name 1_0_0_20161219140623097.jpg.chip.jpg suggests that the image corresponds
# to a person whose age is 1, gender is 0 (Male) and race is 0 (White). However there are few images for
# which the name is malfomed, so we remove them using the following code snippet :
# function to remove invalid images (that he filenames is not correctly formatted)
def get_invalid_images(root_path='/content/data/UTKFace/'):
list_files = glob.glob(os.path.join(root_path, '*.[jJ][pP]*'))
filenames = [path.split('/')[-1].split('_') for path in list_files]
print()
invalid_images = []
for i, im in enumerate(tqdm.tqdm(filenames)):
if im[0].isdigit() and im[1].isdigit() and im[2].isdigit():
continue
else:
invalid_images.append(list_files[i])
return invalid_images
invalid_images = get_invalid_images()
print(invalid_images)
for f in invalid_images: # removal of invalid images
os.remove(f)
# + [markdown] id="illW25ZHhxY2"
# ## [2.a] Depthwise Separable Convolution model
# + id="kEHh3YnT4C0e"
# split the dataset into train, test and validation sets
SRC_DIR = '/content/UTKFace/' # path to the folder containing all images
DEST_DIR = '/content/utk_split/' # path where to save the split dataset, 3 subdirectories will be created (train, valid and test)
SPLIT = 0.7 # ratio of the train set, the remaining (30%) will be split equally between validation and test sets
split_utk(SRC_DIR, DEST_DIR, SPLIT)
# + id="dcUzyRP84M8F"
data_transforms = transforms.Compose([
transforms.Resize((128, 128)),
transforms.ToTensor()
])
train_loader = get_utk_dataloader(batch_size=128, data_dir=DEST_DIR, data_transforms=data_transforms, flag='train')
val_loader = get_utk_dataloader(batch_size=128, data_dir=DEST_DIR, data_transforms=data_transforms, flag='valid')
my_data_loaders = {
'train': train_loader,
'valid': val_loader
}
backup_path = '/content/G_Drive/MyDrive/ADNN/Demography_Psychology/Separable_Convolutional_Model'
os.makedirs(backup_path, exist_ok=True)
checkpoint = '/content/checkpoints/utk_sep_conv_histeq' # folder where to save checkpoints during training
os.makedirs(checkpoint, exist_ok=True)
# + id="F9GLlytf63Gr"
my_model = SepConvModelMT(dropout=0.7, n_class=[1, 2, 5], n_filters=[64, 128, 256, 512], kernels_size=[3, 3, 3, 3]).to(device)
my_optimizer = torch.optim.Adam(my_model.parameters(), lr=1e-3)
# + id="h3wrciXQc6W2"
run_utk(my_model, my_optimizer, epochs=300,
log_interval=1, dataloaders=my_data_loaders,
dirname=checkpoint, filename_prefix='UTK-Sep_Conv',
n_saved=1, log_dir='/content/logs', launch_tensorboard=False,
patience=50, resume_model=None, resume_optimizer=None,
backup_step=5, backup_path=backup_path,
n_epochs_freeze=0, lr_after_freeze=None,
loss_weights=[1/10, 1/0.16, 1/0.44])
# + [markdown] id="NchhSJ5LiTOU"
# ## [2.b] ResNet-50 model
# + id="r3sjZrUgd0OZ"
backup_path = '/content/G_Drive/MyDrive/ADNN/Demography_Psychology/ResNet_Model'
os.makedirs(backup_path, exist_ok=True)
checkpoint = '/content/checkpoints/utk_resnet_adam_histeq'
os.makedirs(checkpoint, exist_ok=True)
# + id="b_9AfIr-kkqn"
my_model = PretrainedMT(model_name='resnet', feature_extract=True, use_pretrained=True).to(device)
my_optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, my_model.parameters()), lr=1e-3)
# + id="p3hhdwu5ko_S"
run_utk(my_model, my_optimizer, epochs=300, log_interval=1, dataloaders=my_data_loaders,
dirname=checkpoint, filename_prefix='UTK-Resnet', n_saved=1,
log_dir='/content/logs', launch_tensorboard=False, patience=50,
resume_model=None, resume_optimizer=None, backup_step=5, backup_path=backup_path,
n_epochs_freeze=10, n_cycle=None, lr_after_freeze=1e-4,
loss_weights=[1/10, 1/0.16, 1/0.44], lr_plot=True)
# + [markdown] id="si6J1mc4l4LH"
# ## [2.c] VGG-19 model
# + id="v7nl4O4El0ol"
backup_path = '/content/G_Drive/MyDrive/ADNN/Demography_Psychology/VGGNet_Model'
os.makedirs(backup_path, exist_ok=True)
checkpoint = '/content/checkpoints/utk_vgg_adam_histeq'
os.makedirs(checkpoint, exist_ok=True)
# + id="nhmn-EmZnT_U"
my_model = PretrainedMT(model_name='vgg', feature_extract=True, use_pretrained=True).to(device)
my_optimizer = torch.optim.Adam(filter(lambda p: p.requires_grad, my_model.parameters()), lr=1e-3)
# + id="iehIuf6eot5P"
run_utk(my_model, my_optimizer, epochs=300, log_interval=1, dataloaders=my_data_loaders,
dirname=checkpoint, filename_prefix='UTK-VGGnet', n_saved=1,
log_dir='/content/logs', launch_tensorboard=False, patience=50,
resume_model=None, resume_optimizer=None, backup_step=5, backup_path=backup_path,
n_epochs_freeze=10, n_cycle=None, lr_after_freeze=1e-4,
loss_weights=[1/10, 1/0.16, 1/0.44], lr_plot=True)
| Age_Gender_Race_Emotion_GPU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sys import argv
from math import log, exp
from random import randrange
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.ensemble import IsolationForest
from sklearn import metrics
from sklearn.preprocessing import MaxAbsScaler
from sklearn.utils import shuffle
from sklearn import preprocessing
#--[Basic Function]---------------------------------------------------------------------
#input decision_values, real_labels{1,-1}, #positive_instances, #negative_instances
#output [A,B] that minimize sigmoid likilihood
#refer to Platt's Probablistic Output for Support Vector Machines
def SigmoidTrain(deci, label, A = None, B = None, prior0=None,prior1=None):
#Count prior0 and prior1 if needed
if prior1==None or prior0==None:
prior1, prior0 = 0, 0
for i in range(len(label)):
if label[i] > 0:
prior1+=1
else:
prior0+=1
#Parameter Setting
maxiter=1000 #Maximum number of iterations
minstep=1e-10 #Minimum step taken in line search
sigma=1e-12 #For numerically strict PD of Hessian
eps=1e-5
length = len(deci)
#Construct Target Support
hiTarget=(prior1+1.0)/(prior1+2.0)
#print(hiTarget)
loTarget=1/(prior0+2.0)
length=prior1+prior0
t=[]
for i in range(length):
if label[i] > 0:
t.append(hiTarget)
else:
t.append(loTarget)
#print(np.mean(t))
#Initial Point and Initial Fun Value
A,B=0.0, log((prior0+1.0)/(prior1+1.0))
#print("A,B",A,B)
fval = 0.0
for i in range(length):
fApB = deci[i]*A+B
if fApB >= 0: # Positive class hence label will be +1
fval += t[i]*fApB + log(1+exp(-fApB))
else: # Negative class label will be -1
fval += (t[i] - 1)*fApB +log(1+exp(fApB))
for it in range(maxiter):
#Update Gradient and Hessian (use H' = H + sigma I)
h11=h22=sigma #Numerically ensures strict PD
h21=g1=g2=0.0
for i in range(length):
fApB = deci[i]*A+B
if (fApB >= 0):
p=exp(-fApB)/(1.0+exp(-fApB))
q=1.0/(1.0+exp(-fApB))
else:
p=1.0/(1.0+exp(fApB))
q=exp(fApB)/(1.0+exp(fApB))
d2=p*q
h11+=deci[i]*deci[i]*d2
h22+=d2
h21+=deci[i]*d2
d1=t[i]-p
g1+=deci[i]*d1
g2+=d1
#Stopping Criteria
if abs(g1)<eps and abs(g2)<eps:
break
#Finding Newton direction: -inv(H') * g
det=h11*h22-h21*h21
dA=-(h22*g1 - h21 * g2) / det
dB=-(-h21*g1+ h11 * g2) / det
gd=g1*dA+g2*dB
#Line Search
stepsize = 1
while stepsize >= minstep:
newA = A + stepsize * dA
newB = B + stepsize * dB
#New function value
newf = 0.0
for i in range(length):
fApB = deci[i]*newA+newB
if fApB >= 0:
newf += t[i]*fApB + log(1+exp(-fApB))
else:
newf += (t[i] - 1)*fApB +log(1+exp(fApB))
#Check sufficient decrease
if newf < fval + 0.0001 * stepsize * gd:
A, B, fval = newA, newB, newf
break
else:
stepsize = stepsize / 2.0
if stepsize < minstep:
print("line search fails",A,B,g1,g2,dA,dB,gd)
return [A,B]
if it>=maxiter-1:
print("reaching maximal iterations",g1,g2)
return (A,B,fval)
#reads decision_value and Platt parameter [A,B]
#outputs predicted probability
def SigmoidPredict(deci, AB):
A, B = AB
fApB = deci * A + B
if (fApB >= 0):
return exp(-fApB)/(1.0+exp(-fApB))
else:
return 1.0/(1+exp(fApB))
return prob
def Expectation(score, A, B):
t = []
for i in range(len(score)):
if A*score[i] + B >= 0:
t.append(1)
else:
t.append(0)
p = np.mean(t)
t[t == 0] = -1
#print(t)
return t
def EM(score, A_init, B_init, prior0, prior1, maxit=1000, tol=1e-8):
# Estimation of parameter(Initial)
flag = 0
A_cur = A_init
B_cur = B_init
A_new = 0.0
B_new = 0.0
# Iterate between expectation and maximization parts
for i in range(maxit):
#print(i)
if(i != 0):
(A_new, B_new) = SigmoidTrain(score,Expectation(score, A_cur, B_cur), A_cur, B_cur)
#print(A_new, B_new)
else:
t = []
for i in range(len(score)):
if A_cur*score[i] + B_cur >= 0:
t.append(1)
else:
t.append(0)
t[t == 0] = -1
(A_new, B_new) = SigmoidTrain(score,t, A_cur, B_cur, prior0, prior1)
#print(A_new, B_new)
# Stop iteration if the difference between the current and new estimates is less than a tolerance level
if(A_cur - A_new < tol and B_cur - B_new < tol):
flag = 1
#break
# Otherwise continue iteration
A_cur = A_new
B_cur = B_new
if(not flag):
print("Didn't converge\n")
return (A_cur, B_cur)
def SigmoidFitting(score, proportion):
fval = []
A = []
B = []
sorted_score = list(sorted(set(np.round(score, decimals = 2))))
for i in range(int(proportion*len(sorted_score))):
threshold = sorted_score[i]
print(threshold)
t = [1 if j <= threshold else -1 for j in score]
(a, b, f) = SigmoidTrain(score,t)
A.append(a)
B.append(b)
fval.append(f)
return(A,B,fval)
def SigmoidFittingGrid(score, proportion):
ngrid = score.shape[1]
fval = []
A = []
B = []
threshold = []
for param in range(ngrid):
a,b,f = SigmoidFitting(score[:,param], proportion)
fval.append(min(f))
A.append(a[np.argmin(f)])
B.append(b[np.argmin(f)])
threshold.append(score[np.argmin(f),param])
return A,B,fval,threshold
def ContextualForest(contexts, features, features_cat, features_num, gamma_range, ncontexts = None):
#contexts = shuffle(contexts)
if(not ncontexts):
ncontexts = contexts.shape[0]
context_scores = np.zeros((ncontexts,features.shape[0],len(gamma_range)))
for i in range(ncontexts):
context = list(contexts.iloc[i,])
print(context)
feature_names = list(features)
context_features = list()
behavioral_features = list()
for feat in feature_names:
c = feat.split("_")
if len(set(c).intersection(set(context))) > 0 or feat in context:
context_features.append(feat)
else:
behavioral_features.append(feat)
context_f = features[context_features]
behav_f = features[behavioral_features]
# Finding the categorical and numerical features in context features.
if(features_cat != None):
cat_names = list(features_cat)
cat_context = list(set(context_f).intersection(set(cat_names)))
context_f_cat = context_f[cat_context]
num_names = list(features_num)
num_context = list(set(context_f).intersection(set(num_names)))
context_f_num = context_f[num_context]
# Finding the distances of the context space
cat_context_distance = metrics.pairwise.cosine_similarity(np.array(context_f_cat))
# Scaling the numerical data using MaxAbsScaler
context_f_num_scaled = MaxAbsScaler().fit_transform(np.array(context_f_num))
# Zero mean and unit variance scaling
#context_f_num_scaled = preprocessing.scale(context_f_num)
#num_context_distance = metrics.pairwise.euclidean_distances(context_f_num_scaled)
#print("Cat distance",cat_context_distance)
for gamma in range(len(gamma_range)):
print(gamma_range[gamma])
num_context_distance = metrics.pairwise.rbf_kernel(context_f_num_scaled, gamma= gamma_range[gamma])
#print("Num distance",num_context_distance)
context_distance = np.minimum(cat_context_distance,num_context_distance)
#context_distance = num_context_distance
rng = np.random.RandomState(42)
clf = IsolationForest(max_samples=256, random_state=rng, smoothing = True)
clf.fit(behav_f, context_distance)
context_scores[i,:,gamma] = clf.decision_function(behav_f,distance=context_distance)
else:
num_names = list(features_num)
num_context = list(set(context_f).intersection(set(num_names)))
context_f_num = context_f[num_context]
# Finding the distances of the context space
# Scaling the numerical data using MaxAbsScaler
context_f_num_scaled = MaxAbsScaler().fit_transform(np.array(context_f_num))
# Zero mean and unit variance scaling
#context_f_num_scaled = preprocessing.scale(context_f_num)
#num_context_distance = metrics.pairwise.euclidean_distances(context_f_num_scaled)
#print("Cat distance",cat_context_distance)
for gamma in range(len(gamma_range)):
print(gamma_range[gamma])
num_context_distance = metrics.pairwise.rbf_kernel(context_f_num_scaled, gamma= gamma_range[gamma], )
#print("Num distance",num_context_distance)
#context_distance = np.minimum(cat_context_distance,num_context_distance)
context_distance = num_context_distance
rng = np.random.RandomState(42)
clf = IsolationForest(max_samples=256, random_state=rng, smoothing = True)
clf.fit(behav_f, context_distance)
context_scores[i,:,gamma] = clf.decision_function(behav_f,distance=context_distance)
return context_scores
# +
# Play Arena Test times of the implementation.
import numpy as np
import matplotlib.pyplot as plt
from sklearn.ensemble import IsolationForest
from sklearn import metrics
rng = np.random.RandomState(42)
# Generate train data
X = 0.3 * rng.randn(5000, 2)
X_train = np.r_[X + 2, X - 2]
# fit the iForest model
# distance = np.ones((X_train.shape[0], X_train.shape[0]))
# clf = IsolationForest(random_state=rng, smoothing = False)
# import timeit
# start_time = timeit.default_timer()
# clf.fit(X_train, distance)
# y_pred_train = clf.decision_function(X_train, distance)
# print("iForest:",timeit.default_timer() - start_time)
# # fit the cForest model
distance = metrics.pairwise.rbf_kernel(X_train, gamma= 10000)
clf = IsolationForest(random_state=rng, smoothing = True)
import timeit
start_time = timeit.default_timer()
clf.fit(X_train, distance)
y_pred_train = clf.decision_function(X_train, distance)
print("cForest:",timeit.default_timer() - start_time)
# +
# Loading the feature matrix and the context from the UnifiedMeasure.
import rpy2.robjects as robjects
from rpy2.robjects import pandas2ri
import pandas as pd
import numpy as np
pandas2ri.activate()
# Feat matrix 2 will need to be saved as data frame in the R script.
readRDS = robjects.r['readRDS']
df_feat = readRDS('features_pen.RDS')
df_feat = pandas2ri.ri2py_dataframe(df_feat)
#df_feat = pd.DataFrame(np.transpose(df_feat))
# Contexts are being stored as data frames in R.
df_contexts = readRDS('contextpens.RDS')
df_contexts = pandas2ri.ri2py_dataframe(df_contexts)
#df_contexts = pd.DataFrame(df_contexts)
# Loading typevar
df_typevar = readRDS('typevar_pen.RDS')
df_typevar = pandas2ri.ri2py_dataframe(df_typevar)
# Loading ground truth
labels = readRDS('labels_pen.RDS')
labels = pandas2ri.ri2py(labels)
len(labels)
# -
# Getting dummies for isolation forest input
#test = pd.get_dummies(df_feat)
df_typevar.head(10)
categorical = df_typevar[df_typevar['typevar'] == "categorical"].index.tolist()
other = df_typevar[df_typevar['typevar'] != "categorical"].index.tolist()
# adjusting for indices in py
categorical = [i-1 for i in categorical]
other = [i-1 for i in other]
# Handling for no categorical features.
if(len(categorical)):
df_feat_cat = df_feat.iloc[:,categorical]
df_feat_other = df_feat.iloc[:,other]
df_feat_cat = pd.get_dummies(df_feat_cat)
df_feat_all = pd.concat([df_feat_other, df_feat_all], axis=1)
else:
df_feat_other = df_feat.iloc[:,other]
df_feat_all = df_feat_other
df_feat_cat = None
# The full feature list in built and ready to be passed to iForest.
gamma_range = [0.0001,0.001,0.01,0.1,1,10,100,1000]
import timeit
start_time = timeit.default_timer()
scores = ContextualForest(df_contexts, df_feat_all, df_feat_cat, df_feat_other,
gamma_range = gamma_range)
print(timeit.default_timer() - start_time)
#np.savez("scores_r5.npz",scores)
np.savez("scores_all_pens.npz",scores)
#scores = np.load("scores_all_pens.npz")["arr_0"]
#fscores = np.median(scores, axis = 0)
scores #Each row corresponds to a context with varying gammas
# Aggregating the scores
fscores = (scores - np.amin(scores, axis = 1, keepdims= True))/(np.amax(scores,axis =1, keepdims=True) - np.amin(scores,axis = 1,
keepdims=True))
fscores = np.min(fscores, axis = 0)
fscores = np.array(fscores)
fscores
ABft = SigmoidFittingGrid(fscores, proportion = 0.1)
print(ABft)
A,B,f,t = ABft
A,B,fmin = A[np.argmin(f)],B[np.argmin(f)],min(f)
print("A:",A,"B:",B,"f:",f)
print("Threshold:",t[np.argmin(f)])
print("Gamma Chosen:",gamma_range[np.argmin(f)])
fgscores = fscores[:,np.argmin(f)]
ABf = SigmoidFitting(fscores, proportion = 0.2)
print(ABf)
A,B,f = ABf
A,B,f = A[np.argmin(f)],B[np.argmin(f)],min(f)
print("A:",A,"B:",B,"f:",f)
print("Threshold:",fscores[np.argmin(f)])
# +
import random
import numpy
from matplotlib import pyplot
inliers = fgscores[np.where(labels == 0)]
outliers = fgscores[np.where(labels == 1)]
pyplot.hist(inliers, alpha=0.5, label='inliers')
pyplot.legend(loc='upper right')
pyplot.show()
pyplot.hist(outliers, alpha=0.5, label='outliers')
pyplot.legend(loc='upper right')
pyplot.show()
import seaborn as sns
fig, ax = plt.subplots()
sns.distplot(outliers, hist=False, label = "outliers")
sns.distplot(inliers, hist=False, label = "inliers")
# -
fgscores[np.where(labels == 0)]
prob = []
for i in range(len(fscores)):
prob.append(SigmoidPredict(fgscores[i],(A,B)))
np.mean(prob)
#prob = (prob - np.min(prob))/(np.max(prob) - np.min(prob))
# +
import numpy as np
import matplotlib.pyplot as plt
#Fixing random state for reproducibility
np.random.seed(19680801)
# the histogram of the data
#n, bins, patches = plt.hist(prob, 50, normed=1, facecolor='g', alpha=0.75)
plt.scatter(fgscores, prob, c="g", alpha=0.5)
plt.xlim((0,1))
plt.ylim((0,1))
plt.ylabel('Probability Estimate')
plt.xlabel('Contextual iForest Score')
plt.title("Fraud Data")
# -
# No Context iForest
rng = np.random.RandomState(42)
clf = IsolationForest(max_samples=512, random_state=rng, smoothing = False)
distance = np.ones((len(df_feat_all), len(df_feat_all)), order = "f")
clf.fit(df_feat_all, distance)
ifscores = clf.decision_function(df_feat_all,distance)
# LoF analysis
from sklearn.neighbors import LocalOutlierFactor
clf = LocalOutlierFactor(n_neighbors=200)
y_pred = clf.fit_predict(df_feat_all)
lof_scores = clf.negative_outlier_factor_
# +
from sklearn.metrics import precision_recall_curve, auc
f, axes = plt.subplots(1, 1, figsize=(7, 5))
y_real = list(labels)
y_proba = []
fscores = (scores - np.amin(scores, axis = 1, keepdims= True))/(np.amax(scores,axis =1, keepdims=True) - np.amin(scores,axis = 1,
keepdims=True))
fscores = np.min(fscores, axis = 0)
fscores = np.array(fscores)
for i in range(len(gamma_range)):
precision, recall, _ = precision_recall_curve(labels, -1*fscores[:,i])
lab = 'Gamma: %f AUC=%.4f' % (gamma_range[i], auc(recall, precision))
axes.step(recall, precision, label=lab)
y_real.append(labels)
y_proba.append(-1*fscores[:,i])
precision, recall, _ = precision_recall_curve(labels, -1*ifscores)
y_real.append(labels)
y_proba.append(-1*ifscores)
lab = 'No Context iForest AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2, color='black')
precision, recall, _ = precision_recall_curve(labels, -1*lof_scores)
y_real.append(labels)
y_proba.append(-1*lof_scores)
lab = 'No Context LOF AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
axes.set_xlabel('Recall')
axes.set_ylabel('Precision')
axes.legend(loc='upper right', fontsize='small')
# +
from sklearn.metrics import precision_recall_curve, auc
f, axes = plt.subplots(1, 1, figsize=(7, 5))
y_real = list(labels)
y_proba = []
# fscores = np.min(scores, axis = 0)
# fscores = (fscores - np.amin(fscores, axis = 0))/(np.amax(fscores,axis =0) - np.amin(fscores,axis = 0))
# fscores = np.array(fscores)
precision, recall, _ = precision_recall_curve(labels, -1*fgscores)
lab = 'ContextualForest: AUC=%.4f' % ( auc(recall, precision))
axes.step(recall, precision, label=lab)
y_real.append(labels)
y_proba.append(-1*fgscores)
precision, recall, _ = precision_recall_curve(labels, -1*ifscores)
y_real.append(labels)
y_proba.append(-1*ifscores)
lab = 'iForest: AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2, color='black')
precision, recall, _ = precision_recall_curve(labels, -1*lof_scores)
y_real.append(labels)
y_proba.append(-1*lof_scores)
lab = 'LOF: AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2, color='red')
axes.set_xlabel('Recall')
axes.set_ylabel('Precision')
axes.legend(loc='upper right', fontsize='small')
# +
from sklearn.metrics import precision_recall_curve, auc
h, axes = plt.subplots(1, 1, figsize=(7, 5))
y_real = list(labels)
y_proba = []
# rng = np.random.RandomState(42)
# clf = IsolationForest(max_samples=256, random_state=rng, n_estimators=5400, smoothing = False)
# distance = np.ones((len(df_feat_all), len(df_feat_all)), order = "f")
# clf.fit(df_feat_all, distance)
# ifscores = clf.decision_function(df_feat_all,distance)
# precision, recall, _ = precision_recall_curve(labels, -1*ifscores)
# lab = 'iForest with 100*K estimators: AUC=%.4f' % ( auc(recall, precision))
# axes.step(recall, precision, label=lab, lw = 2)
# y_real.append(labels)
# y_proba.append(-1*ifscores)
rng = np.random.RandomState(42)
clf = IsolationForest(max_samples=256, random_state=rng, n_estimators=100, smoothing = False, max_features= 5)
distance = np.ones((len(df_feat_all), len(df_feat_all)), order = "f")
clf.fit(df_feat_all, distance)
ifscores = clf.decision_function(df_feat_all,distance)
precision, recall, _ = precision_recall_curve(labels, -1*ifscores)
y_real.append(labels)
y_proba.append(-1*ifscores)
lab = 'iForest 100 estimators: AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2, color='black')
# LoF analysis
from sklearn.neighbors import LocalOutlierFactor
clf = LocalOutlierFactor(n_neighbors=25)
y_pred = clf.fit_predict(df_feat_all)
lof_scores = clf.negative_outlier_factor_
precision, recall, _ = precision_recall_curve(labels, -1*lof_scores)
y_real.append(labels)
y_proba.append(-1*lof_scores)
lab = 'LOF (25 neighbors): AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
from sklearn.neighbors import LocalOutlierFactor
clf = LocalOutlierFactor(n_neighbors=50)
y_pred = clf.fit_predict(df_feat_all)
lof_scores = clf.negative_outlier_factor_
precision, recall, _ = precision_recall_curve(labels, -1*lof_scores)
y_real.append(labels)
y_proba.append(-1*lof_scores)
lab = 'LOF (50 neighbors): AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
# LoF analysis
from sklearn.neighbors import LocalOutlierFactor
clf = LocalOutlierFactor(n_neighbors=100)
y_pred = clf.fit_predict(df_feat_all)
lof_scores = clf.negative_outlier_factor_
precision, recall, _ = precision_recall_curve(labels, -1*lof_scores)
y_real.append(labels)
y_proba.append(-1*lof_scores)
lab = 'LOF (100 neighbors): AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
# LoF analysis
from sklearn.neighbors import LocalOutlierFactor
clf = LocalOutlierFactor(n_neighbors=150)
y_pred = clf.fit_predict(df_feat_all)
lof_scores = clf.negative_outlier_factor_
precision, recall, _ = precision_recall_curve(labels, -1*lof_scores)
y_real.append(labels)
y_proba.append(-1*lof_scores)
lab = 'LOF (150 neighbors): AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
# Elliptic Covariance
from sklearn.covariance import EllipticEnvelope
clf = EllipticEnvelope(contamination=0.1)
clf.fit(df_feat_all)
ell_scores = clf.decision_function(df_feat_all)
precision, recall, _ = precision_recall_curve(labels, -1*ell_scores)
y_real.append(labels)
y_proba.append(-1*ell_scores)
lab = 'Ellpitic Covaraince: AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
# One class SVM
from sklearn import svm
clf = svm.OneClassSVM(nu=0.95 * 0.1 + 0.05,
kernel="rbf", gamma=0.1)
clf.fit(df_feat_all)
svm_scores = clf.decision_function(df_feat_all)
precision, recall, _ = precision_recall_curve(labels, -1*svm_scores)
y_real.append(labels)
y_proba.append(-1*svm_scores)
lab = 'oSVM: AUC=%.4f' % (auc(recall, precision))
axes.step(recall, precision, label=lab, lw=2)
axes.set_xlabel('Recall')
axes.set_ylabel('Precision')
axes.legend(loc='upper right', fontsize='small')
# -
| code/Pens/IsoForest-Pens.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// [](https://lab.mlpack.org/v2/gh/mlpack/examples/master?urlpath=lab%2Ftree%2Fq_learning%2Fpendulum_dqn.ipynb)
//
// You can easily run this notebook at https://lab.mlpack.org/
//
// Here, we train a [Simple DQN](https://www.cs.toronto.edu/~vmnih/docs/dqn.pdf) agent to get high scores for the [Pendulum](https://gym.openai.com/envs/Pendulum-v0/) environment.
//
// We make the agent train and test on OpenAI Gym toolkit's GUI interface provided through a distributed infrastructure (TCP API). More details can be found [here](https://github.com/zoq/gym_tcp_api).
//
// A video of the trained agent can be seen in the end.
// ## Including necessary libraries and namespaces
#include <mlpack/core.hpp>
#include <mlpack/methods/ann/ffn.hpp>
#include <mlpack/methods/reinforcement_learning/q_learning.hpp>
#include <mlpack/methods/reinforcement_learning/q_networks/simple_dqn.hpp>
#include <mlpack/methods/reinforcement_learning/environment/env_type.hpp>
#include <mlpack/methods/reinforcement_learning/policy/greedy_policy.hpp>
#include <mlpack/methods/reinforcement_learning/training_config.hpp>
// Used to run the agent on gym's environment (provided externally) for testing.
#include <gym/environment.hpp>
// Used to generate and display a video of the trained agent.
#include "xwidgets/ximage.hpp"
#include "xwidgets/xvideo.hpp"
#include "xwidgets/xaudio.hpp"
using namespace mlpack;
using namespace mlpack::ann;
using namespace ens;
using namespace mlpack::rl;
// ## Initializing the agent
// Set up the state and action space.
DiscreteActionEnv::State::dimension = 3;
DiscreteActionEnv::Action::size = 3;
// Set up the network.
FFN<MeanSquaredError<>, GaussianInitialization> network(
MeanSquaredError<>(), GaussianInitialization(0, 1));
network.Add<Linear<>>(DiscreteActionEnv::State::dimension, 128);
network.Add<ReLULayer<>>();
network.Add<Linear<>>(128, DiscreteActionEnv::Action::size);
SimpleDQN<> model(network);
// Set up the policy and replay method.
GreedyPolicy<DiscreteActionEnv> policy(1.0, 1000, 0.1, 0.99);
RandomReplay<DiscreteActionEnv> replayMethod(32, 10000);
// Set up training configurations.
TrainingConfig config;
config.ExplorationSteps() = 100;
// Set up DQN agent.
QLearning<DiscreteActionEnv, decltype(model), AdamUpdate, decltype(policy)>
agent(config, model, policy, replayMethod);
// ## Preparation for training the agent
// +
// Set up the gym training environment.
gym::Environment env("gym.kurg.org", "4040", "Pendulum-v0");
// Initializing training variables.
std::vector<double> returnList;
size_t episodes = 0;
bool converged = true;
// The number of episode returns to keep track of.
size_t consecutiveEpisodes = 50;
// -
// Since the Pendulum environment has a continuous action space, we need to perform "discretization" of the action space.
//
// For that, we assume that our Q learning agent outputs 3 action values for our actions {0, 1, 2}. Meaning the actions given by the agent will either be `0`, `1`, or `2`.
//
// Now, we subtract `1.0` from the actions, which then becomes the input to the environment. This essentially means that we correspond the actions `0`, `1`, and `2` given by the agent, to the torque values `-1.0`, `0`, and `1.0` for the environment, respectively.
//
// This simple trick allows us to train a continuous action-space environment using DQN.
//
// Note that we have divided the action-space into 3 divisions here. But you may use any number of divisions as per your choice. More the number of divisions, finer are the controls available for the agent, and therefore better are the results!
// Function to train the agent on the Pendulum gym environment.
void Train(const size_t numSteps)
{
agent.Deterministic() = false;
std::cout << "Training for " << numSteps << " steps." << std::endl;
while (agent.TotalSteps() < numSteps)
{
double episodeReturn = 0;
env.reset();
do
{
agent.State().Data() = env.observation;
agent.SelectAction();
arma::mat action = {double(agent.Action().action) - 1.0};
env.step(action);
DiscreteActionEnv::State nextState;
nextState.Data() = env.observation;
replayMethod.Store(agent.State(), agent.Action(), env.reward, nextState,
env.done, 0.99);
episodeReturn += env.reward;
agent.TotalSteps()++;
if (agent.Deterministic() || agent.TotalSteps() < config.ExplorationSteps())
continue;
agent.TrainAgent();
} while (!env.done);
returnList.push_back(episodeReturn);
episodes += 1;
if (returnList.size() > consecutiveEpisodes)
returnList.erase(returnList.begin());
double averageReturn = std::accumulate(returnList.begin(),
returnList.end(), 0.0) /
returnList.size();
if(episodes % 4 == 0)
{
std::cout << "Avg return in last " << returnList.size()
<< " episodes: " << averageReturn
<< "\t Episode return: " << episodeReturn
<< "\t Total steps: " << agent.TotalSteps() << std::endl;
}
}
}
// ## Let the training begin
// Training the agent for a total of at least 5000 steps.
Train(5000)
// ## Testing the trained agent
// +
agent.Deterministic() = true;
// Creating and setting up the gym environment for testing.
gym::Environment envTest("gym.kurg.org", "4040", "Pendulum-v0");
envTest.monitor.start("./dummy/", true, true);
// Resets the environment.
envTest.reset();
envTest.render();
double totalReward = 0;
size_t totalSteps = 0;
// Testing the agent on gym's environment.
while (1)
{
// State from the environment is passed to the agent's internal representation.
agent.State().Data() = envTest.observation;
// With the given state, the agent selects an action according to its defined policy.
agent.SelectAction();
// Action to take, decided by the policy.
arma::mat action = {double(agent.Action().action) - 1.0};
envTest.step(action);
totalReward += envTest.reward;
totalSteps += 1;
if (envTest.done)
{
std::cout << " Total steps: " << totalSteps << "\t Total reward: "
<< totalReward << std::endl;
break;
}
// Uncomment the following lines to see the reward and action in each step.
// std::cout << " Current step: " << totalSteps << "\t current reward: "
// << totalReward << "\t Action taken: " << action;
}
envTest.close();
std::string url = envTest.url();
auto video = xw::video_from_url(url).finalize();
video
// -
// ## A little more training...
// Training the same agent for a total of at least 20000 steps.
Train(20000)
// # Final agent testing!
// +
agent.Deterministic() = true;
// Creating and setting up the gym environment for testing.
gym::Environment envTest("gym.kurg.org", "4040", "Pendulum-v0");
envTest.monitor.start("./dummy/", true, true);
// Resets the environment.
envTest.reset();
envTest.render();
double totalReward = 0;
size_t totalSteps = 0;
// Testing the agent on gym's environment.
while (1)
{
// State from the environment is passed to the agent's internal representation.
agent.State().Data() = envTest.observation;
// With the given state, the agent selects an action according to its defined policy.
agent.SelectAction();
// Action to take, decided by the policy.
arma::mat action = {double(agent.Action().action) - 1.0};
envTest.step(action);
totalReward += envTest.reward;
totalSteps += 1;
if (envTest.done)
{
std::cout << " Total steps: " << totalSteps << "\t Total reward: "
<< totalReward << std::endl;
break;
}
// Uncomment the following lines to see the reward and action in each step.
// std::cout << " Current step: " << totalSteps << "\t current reward: "
// << totalReward << "\t Action taken: " << action;
}
envTest.close();
std::string url = envTest.url();
auto video = xw::video_from_url(url).finalize();
video
| reinforcement_learning_gym/pendulum_dqn/pendulum_dqn.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # <center>Block 9: Optimal transport with entropic regularization</center>
# ### <center><NAME> (NYU)</center>
# ## <center>'math+econ+code' masterclass on optimal transport and economic applications</center>
# <center>© 2018-2020 by <NAME>. Support from NSF grant DMS-1716489 and <NAME>'s contribution are acknowledged.</center>
# ### Learning objectives
#
# * Entropic regularization
#
# * The log-sum-exp trick
#
# * The Iterated Proportional Fitting Procedure (IPFP)
# ### References
#
# * [OTME], Ch. 7.3
#
# * <NAME>, Computational Optimal Transport, Ch. 4.
# ### Entropic regularization of the optimal transport problem
#
# Consider the problem
#
# \begin{align*}
# \max_{\pi\in\mathcal{M}\left( p,q\right) }\sum_{ij}\pi_{ij}\Phi_{ij}-\sigma\sum_{ij}\pi_{ij}\ln\pi_{ij}
# \end{align*}
#
# where $\sigma>0$. The problem coincides with the optimal assignment problem when $\sigma=0$. When $\sigma\rightarrow+\infty$, the solution to this problem approaches the independent coupling, $\pi_{ij}=p_{i}q_{j}$.
#
# Later on, we will provide microfoundations for this problem, and connect it with a number of important methods in economics (BLP, gravity model, Choo-Siow...). For now, let's just view this as an extension of the optimal transport problem.
# ### Dual of the regularized problem
#
# Let's compute the dual by the minimax approach. We have
#
# \begin{align*}
# \max_{\pi\geq0}\min_{u,v}\sum_{ij}\pi_{ij}\left( \Phi_{ij}-u_{i}-v_{j}%
# -\sigma\ln\pi_{ij}\right) +\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}%
# \end{align*}
#
# thus
#
# \begin{align*}
# \min_{u,v}\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}+\max_{\pi\geq0}\sum_{ij}%
# \pi_{ij}\left( \Phi_{ij}-u_{i}-v_{j}-\sigma\ln\pi_{ij}\right)
# \end{align*}
#
# By FOC in the inner problem, one has $\Phi_{ij}-u_{i}-v_{j}-\sigma\ln \pi_{ij}-\sigma=0,$thus
#
# \begin{align*}
# \pi_{ij}=\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}-\sigma}{\sigma}\right)
# \end{align*}
#
# and $\pi_{ij}\left( \Phi_{ij}-u_{i}-v_{j}-\sigma\ln\pi_{ij}\right) =\sigma\pi_{ij}$, thus the dual problem is
#
# \begin{align*}
# \min_{u,v}\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}+\sigma\sum_{ij}\exp\left(
# \frac{\Phi_{ij}-u_{i}-v_{j}-\sigma}{\sigma}\right) .
# \end{align*}
#
# After replacing $v_{j}$ by $v_{j}+\sigma$, the dual is
#
# \begin{align*}
# \min_{u,v}\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}+\sigma\sum_{ij}\exp\left(
# \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right) -\sigma. \tag{V1}
# \end{align*}
# ### Another expression of the dual
#
# **Claim:** the problem is equivalent to
#
# <a name='V2'></a>
# \begin{align*}
# \min_{u,v}\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}+\sigma\log\sum_{i,j}
# \exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right) \tag{V2}
# \end{align*}
#
# Indeed, let us go back to the minimax expression
#
# \begin{align*}
# \min_{u,v}\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}+\max_{\pi\geq0}\sum_{ij}\pi_{ij}\left( \Phi_{ij}-u_{i}-v_{j}-\sigma\ln\pi_{ij}\right)
# \end{align*}
#
# we see that the solution $\pi$ has automatically $\sum_{ij}\pi_{ij}=1$; thus we can incorporate the constraint into
#
# \begin{align*}
# \min_{u,v}\sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}+\max_{\pi\geq0:\sum_{ij}\pi_{ij}=1}\sum_{ij}\pi_{ij}\left( \Phi_{ij}-u_{i}-v_{j}-\sigma\ln\pi_{ij}\right)
# \end{align*}
#
# which yields the [our desired result](#V2).
#
# [This expression](#V2) is interesting because, taking *any* $\hat{\pi}\in
# M\left( p,q\right)$, it reexpresses as
#
# \begin{align*}
# \max_{u,v}\sum_{ij}\hat{\pi}_{ij}\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right) -\log\sum_{ij}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# \end{align*}
#
# therefore if the parameter is $\theta=\left( u,v\right)$, observations are
# $ij$ pairs, and the likelihood of $ij$ is
#
# \begin{align*}
# \pi_{ij}^{\theta}=\frac{\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma
# }\right) }{\sum_{ij}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# }
# \end{align*}
#
# Hence, [our expression](#problem) will coincide with the maximum likelihood in this model.
# ### A third expression of the dual problem
#
# Consider
#
# <a name='V2'></a>
# \begin{align*}
# \min_{u,v} & \sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j} \\
# s.t. \quad & \sum_{i,j}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# =1
# \end{align*}
#
# It is easy to see that the solutions of this problem coincide with [version 2](#V2). Indeed, the Lagrange multiplier is forced to be one. In other words,
#
# \begin{align*}
# \min_{u,v} & \sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}\\
# s.t. \quad & \sigma\log\sum_{i,j}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma
# }\right) =0
# \end{align*}
# ### Small-temperature limit and the log-sum-exp trick
#
# Recall that when $\sigma\rightarrow0$, one has
#
# \begin{align*}
# \sigma\log\left( e^{a/\sigma}+e^{b/\sigma}\right) \rightarrow\max\left(
# a,b\right)
# \end{align*}
#
# Indeed, letting $m=\max\left( a,b\right)$,
#
# <a name='lse'></a>
# \begin{align*}
# \sigma\log\left( e^{a/\sigma}+e^{b/\sigma}\right) =m+\sigma\log\left(\exp\left( \frac{a-m}{\sigma}\right) +\exp\left( \frac{b-m}{\sigma}\right)\right),
# \end{align*}
# and the argument of the logarithm lies between $1$ and $2$.
#
# This simple remark is actually a useful numerical recipe called the *log-sum-exp trick*: when $\sigma$ is small, using [the formula above](#lse) to compute $\sigma\log\left( e^{a/\sigma}+e^{b/\sigma}\right)$ ensures the exponentials won't blow up.
#
# Back to the third expression, with $\sigma\rightarrow0$, one has
#
# \begin{align*}
# \min_{u,v} & \sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}\tag{V3}\\
# s.t. & \max_{ij}\left( \Phi_{ij}-u_{i}-v_{j}\right) =0\nonumber
# \end{align*}
#
# This is exactly equivalent with the classical Monge-Kantorovich expression
#
# \begin{align*}
# \min_{u,v} & \sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}\tag{V3}\\
# s.t. & \Phi_{ij}-u_{i}-v_{j}\leq0\nonumber
# \end{align*}
# Back to the third expression of the dual, with $\sigma\rightarrow0$, one has
#
# \begin{align*}
# \min_{u,v} & \sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}\tag{V3}\\
# s.t. & \max_{ij}\left( \Phi_{ij}-u_{i}-v_{j}\right) =0\nonumber
# \end{align*}
#
# This is exactly equivalent with the classical Monge-Kantorovich expression
#
# \begin{align*}
# \min_{u,v} & \sum_{i}u_{i}p_{i}+\sum_{j}v_{j}q_{j}\tag{V3}\\
# s.t. & \Phi_{ij}-u_{i}-v_{j}\leq0\nonumber
# \end{align*}
# ### Computation
#
# We can compute $\min F\left( x\right)$ by two methods:
#
# Either by gradient descent: $x\left( t+1\right) =x_{t}-\epsilon _{t}\nabla F\left( x_{t}\right) $. (Steepest descent has $\epsilon _{t}=1/\left\vert \nabla F\left( x_{t}\right) \right\vert $.)
#
# Or by coordinate descent: $x_{i}\left( t+1\right) =\arg\min_{x_{i}}F\left( x_{i},x_{-i}\left( t\right) \right)$.
#
# Why do these methods converge? Let's provide some justification. We will decrease $x_{t}$ by $\epsilon d_{t}$, were $d_{t}$ is normalized by $\left\vert d_{t}\right\vert _{p}:=\left( \sum_{i=1}^{n}d_{t}^{i}\right) ^{1/p}=1$. At first order, we have
#
# \begin{align*}
# F\left( x_{t}-\epsilon d_{t}\right) =F\left( x_{t}\right) -\epsilon d_{t}^{\intercal}\nabla F\left( x_{t}\right) +O\left( \epsilon^{1}\right).
# \end{align*}
#
# We need to maximize $d_{t}^{\intercal}\nabla F\left( x_{t}\right)$ over $\left\vert d_{t}\right\vert _{p}=1$.
#
# * For $p=2$, we get $d_{t}=\nabla F\left( x_{t}\right) /\left\vert \nabla F\left( x_{t}\right) \right\vert $
#
# * For $p=1$, we get $d_{t}=sign\left( \partial F\left( x_{t}\right)/\partial x^{i}\right) $ if $\left\vert \partial F\left( x_{t}\right) /\partial x^{i}\right\vert =\max_{j}\left\vert \partial F\left( x_{t}\right) /\partial x^{j}\right\vert $, $0$ otherwise.
#
# In our context, gradient descent is
#
# \begin{align*}
# u_{i}\left( t+1\right) & =u_{i}\left( t\right) -\epsilon\frac{\partial
# F}{\partial u_{i}}\left( u\left( t\right) ,v\left( t\right) \right)
# ,\text{ and }\\
# v_{j}\left( t+1\right) & =v_{j}\left( t\right) -\epsilon\frac{\partial
# F}{\partial v_{j}}\left( u\left( t\right) ,v\left( t\right) \right)
# \end{align*}
#
# while coordinate descent is
#
# \begin{align*}
# \frac{\partial F}{\partial u_{i}}\left( u_{i}\left( t+1\right)
# ,u_{-i}\left( t\right) ,v\left( t\right) \right) =0,\text{ and }
# \frac{\partial F}{\partial v_{j}}\left( u\left( t\right) ,v_{j}\left(
# t+1\right) ,v_{-j}\left( t\right) \right) =0.
# \end{align*}
# ### Gradient descent
#
# Gradient of objective function in version 1 of our problem:
#
# \begin{align*}
# \left( p_{i}-\sum_{j}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# ,q_{j}-\sum_{i}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# \right)
# \end{align*}
#
# Gradient of objective function in version 2
#
# \begin{align*}
# \left( p_{i}-\frac{\sum_{j}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma
# }\right) }{\sum_{ij}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# },q_{j}-\frac{\sum_{i}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right)
# }{\sum_{ij}\exp\left( \frac{\Phi_{ij}-u_{i}-v_{j}}{\sigma}\right) }\right)
# \end{align*}
# ### Coordinate descent
#
# Coordinate descent on objective function in version 1:
#
# \begin{align*}
# p_{i} & =\sum_{j}\exp\left( \frac{\Phi_{ij}-u_{i}\left( t+1\right)
# -v_{j}\left( t\right) }{\sigma}\right) ,\\
# q_{j} & =\sum_{i}\exp\left( \frac{\Phi_{ij}-u_{i}\left( t\right)
# -v_{j}\left( t+1\right) }{\sigma}\right)
# \end{align*}
#
# that is
#
# \begin{align*}
# \left\{
# \begin{array}
# [c]{c}
# u_{i}\left( t+1\right) =\sigma\log\left( \frac{1}{p_{i}}\sum_{j}\exp\left(
# \frac{\Phi_{ij}-v_{j}\left( t\right) }{\sigma}\right) \right) \\
# v_{j}\left( t+1\right) =\sigma\log\left( \frac{1}{q_{j}}\sum_{i}\exp\left(
# \frac{\Phi_{ij}-u_{i}\left( t\right) }{\sigma}\right) \right)
# \end{array}
# \right.
# \end{align*}
#
# this is called the Iterated Fitting Proportional Procedure (IPFP), or Sinkhorn's algorithm.
#
# Coordinate descent on objective function in version 2 does not yield a closed-form expression.
# ### IPFP, linear version
#
# Letting $a_{i}=\exp\left( -u_{i}/\sigma\right) $ and $b_{j}=\exp\left( -v_{j}/\sigma\right) $ and $K_{ij}=\exp\left( \Phi_{ij}/\sigma\right) $, one has $\pi_{ij}=a_{i}b_{j}K_{ij}$, and the procedure reexpresses as
#
# \begin{align*}
# \left\{
# \begin{array}
# [c]{l}%
# a_{i}\left( t+1\right) =p_{i}/\left( Kb\left( t\right) \right)
# _{i}\text{ and }\\
# b_{j}\left( t+1\right) =q_{j}/\left( K^{\intercal}a\left( t\right)
# \right) _{j}.
# \end{array}
# \right.
# \end{align*}
# ### The log-sum-exp trick
#
# The previous program is extremely fast, partly due to the fact that it involves linear algebra operations. However, it breaks down when $\sigma$ is small; this is best seen taking a log transform and returning to $u^{k}=-\sigma\log a^{k}$ and $v^{k}=-\sigma\log b^{k}$, that is
#
# \begin{align*}
# \left\{
# \begin{array}
# [c]{l}%
# u_{i}^{k}=\mu_{i}+\sigma\log\sum_{j}\exp\left( \frac{\Phi_{ij}-v_{j}^{k-1}%
# }{\sigma}\right) \\
# v_{j}^{k}=\zeta_{j}+\sigma\log\sum_{i}\exp\left( \frac{\Phi_{ij}-u_{i}^{k}%
# }{\sigma}\right)
# \end{array}
# \right.
# \end{align*}
#
# where $\mu_{i}=-\sigma\log p_{i}$ and $\zeta_{j}=-\sigma\log q_{j}$.
#
# One sees what may go wrong: if $\Phi_{ij}-v_{j}^{k-1}$ is positive in the exponential in the first sum, then the exponential blows up due to the small $\sigma$ at the denominator. However, the log-sum-exp trick can be used in order to avoid this issue.
#
# Consider
#
# \begin{align*}
# \left\{
# \begin{array}
# [c]{l}%
# \tilde{v}_{i}^{k}=\max_{j}\left\{ \Phi_{ij}-v_{j}^{k}\right\} \\
# \tilde{u}_{j}^{k}=\max_{i}\left\{ \Phi_{ij}-u_{i}^{k}\right\}
# \end{array}
# \right.
# \end{align*}
#
# (the indexing is not a typo: $\tilde{v}$ is indexed by $i$ and $\tilde{u}$ by $j$).
#
# One has
#
# \begin{align*}
# \left\{
# \begin{array}
# [c]{l}%
# u_{i}^{k}=\mu_{i}+\tilde{v}_{i}^{k-1}+\sigma\log\sum_{j}\exp\left( \frac
# {\Phi_{ij}-v_{j}^{k-1}-\tilde{v}_{i}^{k}}{\sigma}\right) \\
# v_{j}^{k}=\zeta_{j}+\tilde{u}_{j}^{k}+\sigma\log\sum_{i}\exp\left( \frac
# {\Phi_{ij}-u_{i}^{k}-\tilde{u}_{j}^{k}}{\sigma}\right)
# \end{array}
# \right.
# \end{align*}
#
# and now the arguments of the exponentials are always nonpositive, ensuring the exponentials don't blow up.
# ## Application
# We will return to our marriage example from Lecture 4. We will do this both using synthetic data and real data.
# +
library(gurobi)
library(Matrix)
library(tictoc)
nbX = 50
nbY = 30
tol = 1e-09
maxiter = 1e+06
# -
# Let's load up the `affinitymatrix.csv`, `Xvals.csv` and `Yvals.csv` that you will recall from Lecture 4. We will work on a smaller population, with `nbX` types of men and `nbY` types of women.
# +
thePath = paste0(getwd(),"/../data_mec_optim/marriage_personality-traits")
data_X = as.matrix(read.csv(paste0(thePath,"/Xvals.csv"), sep=",", header=TRUE)) # loads the data
Xvals = matrix(as.numeric(data_X[,1:10]), ncol=10)
data_Y = as.matrix(read.csv(paste0(thePath,"/Yvals.csv"), sep=",", header=TRUE)) # loads the data
Yvals = matrix(as.numeric(data_Y[,1:10]), ncol=10)
data_aff = as.matrix(read.csv(paste0(thePath,"/affinitymatrix.csv"),sep=",", header=TRUE)) # loads the data
A = matrix(as.numeric(data_aff[1:10, 2:(11)]), 10, 10)
sdX = apply(Xvals, 2, sd)
sdY = apply(Yvals, 2, sd)
mX = apply(Xvals, 2, mean)
mY = apply(Yvals, 2, mean)
Xvals = t((t(Xvals) - mX)/sdX)
Yvals = t((t(Yvals) - mY)/sdY)
Phi = (Xvals %*% A %*% t(Yvals))[1:nbX,1:nbY]
p = rep(1/nbX, nbX)
q = rep(1/nbY, nbY)
nrow = min(8, nbX)
ncol = min(8, nbY)
# -
# We are going to run a horse race between solving this problem using Gurobi and two IPFP algorithms. First Gurobi
# +
A1 = kronecker(matrix(1, 1, nbY), Diagonal(nbX))
A2 = kronecker(Diagonal(nbY), matrix(1, 1, nbX))
A = rbind2(A1, A2)
d = c(p, q)
tic()
result = gurobi(list(A = A, obj = c(Phi), modelsense = "max", rhs = d, sense = "="),
params = list(OutputFlag = 0))
toc()
if (result$status == "OPTIMAL") {
x = matrix(result$x, nrow = nbX)
u_gurobi = result$pi[1:nbX]
v_gurobi = result$pi[(nbX + 1):(nbX + nbY)]
val_gurobi = result$objval
} else {
stop("optimization problem with Gurobi.")
}
print(paste0("Value of the problem (Gurobi) = ", val_gurobi))
print(paste0("Sum(pi*Phi) (IPFP1) = ", sum(x * Phi)))
#print(u_gurobi[1:nrow] - u_gurobi[nrow])
#print(v_gurobi[1:ncol] + u_gurobi[nrow])
# -
# Next IPFP.
# +
sigma = 0.1
tic()
cont = TRUE
iter = 0
K = exp(Phi/sigma)
B = rep(1, nbY) # Guess B = vector of ones
while (cont) {
iter = iter + 1
A = p/c(K %*% B)
KA = c(t(A) %*% K)
error = max(abs(KA * B/q - 1))
if ((error < tol) | (iter >= maxiter)) {
cont = FALSE
}
B = q/KA
}
u = -sigma * log(A)
v = -sigma * log(B)
pi_1 = (K * A) * matrix(B, nbX, nbY, byrow = T)
val = sum(pi_1 * Phi) - sigma * sum(pi_1 * log(pi_1))
toc()
if (iter >= maxiter) {
print("Maximum number of iterations reached in IPFP1.")
} else {
print(paste0("IPFP1 converged in ", iter, " steps."))
print(paste0("Value of the problem (IPFP1) = ", val))
print(paste0("Sum(pi*Phi) (IPFP1) = ", sum(pi_1 * Phi)))
#print(u[1:nrow] - u[nrow])
#print(v[1:ncol] + u[nrow])
}
# -
# The following procedure, carried in the log-domain is mathematically equivalent to the previous implementation of the IPFP, but it is noticeably slower.
# +
sigma = 0.01
tic()
iter = 0
cont = TRUE
v = rep(0, nbY)
mu = -sigma * log(p)
nu = -sigma * log(q)
while (cont) {
iter = iter + 1
u = mu + sigma * log(apply(exp((Phi - matrix(v, nbX, nbY, byrow = T))/sigma), 1, sum))
KA = apply(exp((Phi - u)/sigma), 2, sum)
error = max(abs(KA * exp(-v/sigma)/q - 1))
if ((error < tol) | (iter >= maxiter)) {
cont = FALSE
}
v = nu + sigma * log(KA)
}
pi = exp((Phi - u - matrix(v, nbX, nbY, byrow = T))/sigma)
val = sum(pi * Phi) - sigma * sum((pi * log(pi))[which(pi != 0)])
toc()
if (iter >= maxiter) {
print("Maximum number of iterations reached in IPFP1bis.")
} else {
print(paste0("IPFP1_logs converged in ", iter, " steps."))
print(paste0("Value of the problem (IPFP1_logs) = ", val))
print(paste0("Sum(pi*Phi) (IPFP1_logs) = ", sum(pi * Phi)))
}
# -
# Both procedures above will break down when $\sigma$ is small, e.g. $\sigma=0.001$ (Try!). However if we modify the second procedure using the log-sum-exp trick, things work again:
# +
sigma = 0.001
tic()
iter = 0
cont = TRUE
v = rep(0, nbY)
mu = -sigma * log(p)
nu = -sigma * log(q)
uprec = -Inf
while (cont) {
# print(iter)
iter = iter + 1
vstar = apply(t(t(Phi) - v), 1, max)
u = mu + vstar + sigma * log(apply(exp((Phi - matrix(v, nbX, nbY, byrow = T) -
vstar)/sigma), 1, sum))
error = max(abs(u - uprec))
uprec = u
ustar = apply(Phi - u, 2, max)
v = nu + ustar + sigma * log(apply(exp((Phi - u - matrix(ustar, nbX, nbY, byrow = T))/sigma),
2, sum))
if ((error < tol) | (iter >= maxiter)) {
cont = FALSE
}
}
pi_2 = exp((Phi - u - matrix(v, nbX, nbY, byrow = T))/sigma)
val = sum(pi_2 * Phi) - sigma * sum(pi_2 * log(pi_2))
toc()
if (iter >= maxiter) {
print("Maximum number of iterations reached in IPFP2.")
} else {
print(paste0("IPFP2 converged in ", iter, " steps."))
print(paste0("Value of the problem (IPFP2) = ", val))
print(paste0("Sum(pi*Phi) (IPFP2) = ", sum(pi_2 * Phi)))
}
# -
| ipynb_r_mec_optim/B09_entropicregularization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Run the MICE imputation
#
# This is simply a jupyter notebook wrapper around the R Markdown file `MICE_step.Rmd`
# !R -e "rmarkdown::render('MICE_step.Rmd',output_file='output.html')"
| step3_MICE_imputation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Make Predictions with Linear Regression
# ## Mini-Lab: Linear Regression
# Welcome to your next mini-lab! Go ahead an run the following cell to get started. You can do that by clicking on the cell and then clickcing `Run` on the top bar. You can also just press `Shift` + `Enter` to run the cell.
# +
from datascience import *
import numpy as np
import otter
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
grader = otter.Notebook("m10_l1_tests")
# -
# For this mini-lab and the next mini-lab, we'll be looking at something a bit lighter when compared to COVID-19 data. We'll instead be looking (and trying to establish correlation) between the the various scores of students who took the SAT in 2014. Run the next cell to import this data.
sat = Table().read_table("../datasets/sat2014.csv").select("Critical Reading", "Math", "Writing", "Combined")
sat.show(5)
# Next we'll be recreating the standard set of statistical functions that will be used for linear regression. First up is the `standard_units` and `correlation` functions. The `standard_units` function converts an array of numbers into...well, standard units! The `correlation` function utilizes the `standard units` function in order find the correlation coefficient between two different arrays - the `x_array` and the `y_array`. Implement these functions below!
# +
def standard_units(array):
return ...
def correlation(x_array, y_array):
return ...
# -
grader.check("q1")
# Next up is the `slope` and `intercept` functions which calculate the slope and intercept between two arrays. Again, they take the `x_array` and `y_array` as input and utilize the `correlation` function that you implemented above. Continue implementing these functions in the cell below.
# +
def slope(x_array, y_array):
r = ...
return ...
def intercept(x_array, y_array):
a = ...
return ...
# -
grader.check("q2")
# Finally we'll be putting all of this together in order to predict values given a certain x! Fill in the missing code for the `regression_line` function. This function may seem a little strange - there's a function within a function! But don't worry too much about how it's strucutred, as long as `a` and `b` are assigned correctly, the rest of the lab should flow smoothly.
#
# *Note*: You may have noticed that we used functions inside functions before, specifically in the bootstrapping and hypothesis testing labs. These are examples of [higher-order functions](https://en.wikipedia.org/wiki/Higher-order_function)!
def regression_line(x_array, y_array):
a = ...
b = ...
def prediction_function(x_value):
return (a * x_value) + b
return prediction_function
grader.check("q3")
# Last but not least, we'll be setting up our regression-line so that we can start predicting points. Replace the `...` below with the columns present in the `sat` table that interest you. After doing this, run the cell below to set up our prediction function.
# +
x_array = ...
y_array = ...
predict = regression_line(x_array, y_array)
# -
# Now start predicting! Feel free to change around the columns above as well as the prediction below.
predict(720)
# Now that we have set up a prediction function, are there any limits to this function? For example, what if we input a score out of range? Does this data actually mean anything? What if the output is out of range, what would we do then? Linear regression is an amazing and powerful tool but like everything else in life it isn't perfect. Nonetheless, it's a basis of data science and rightfully so. Congratulations on finishing! Run the next cell to make sure that you passed all of the test cases.
grader.check_all()
| minilabs/make-predictions-with-linear-regression/linear_regression_minilab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# # Working with raster data in python
#
#
# ## Table of Contents
#
# 1. [About the dataset](#dataset)<br>
# 2. [Part 1 - Weather maps with netCDF4 and matplotlib](#part1)<br>
# 2.1. [Import packages](#import1)<br>
# 2.2. [Load gridded data with netCDF4](#load1)<br>
# 2.3. [Create a global map of the average temperature in January using matplotlib](#map1)<br>
#
# 3. [Part 2 - Weather maps with xarray and Cartopy](#part2)<br>
# 3.1. [Import packages](#import2)<br>
# 3.2. [Load gridded data with xarray](#load2)<br>
# 3.3. [Create maps using xarray](#map21)<br>
# 3.4. [Create maps using Cartoid](#map22)<br>
#
# <a id="dataset"></a>
# ## About the dataset
#
# With the gridded data from [CRU](http://www.cru.uea.ac.uk/data/) you will learn how to work with gridded historical data.
#
# The [dataset](https://crudata.uea.ac.uk/cru/data/temperature/#datdow) contains a 5° by 5° grid with absolute temperatures from 1961 to 1990. The data is represented in a [NetCDF](https://pro.arcgis.com/en/pro-app/help/data/multidimensional/what-is-netcdf-data.htm) format.
#
# Download the following file, and store it locally or in object-store when working on the [IBM Data Science Experience](https://datascience.ibm.com/) :
#
# * https://crudata.uea.ac.uk/cru/data/temperature/absolute.nc
#
#
# <a id="part1"></a>
# ## Part 1 - Weather maps with netCDF4 and matplotlib
#
#
# In the first half of this tutorial, we will see how to use Python's [netCDF4](https://unidata.github.io/netcdf4-python/netCDF4/index.html) module to extract data from the dataset.
#
#
#
#
# <a id="import1"></a>
# ### 1. Import packages
#
# Following is the explicit list of imports that we used through this notebook.
import pandas as pd
import requests, json
from io import StringIO
from netCDF4 import Dataset
import numpy as np
import scipy
import matplotlib
from pylab import *
from mpl_toolkits.basemap import Basemap, addcyclic, shiftgrid
# %matplotlib inline
# We first import the dataset with a helper function that uses the project token created per instructions mentioned above. Import the `absolute.nc` file locally or add the below code by clicking on `Insert to code` below the file under the file in object-store. Then load the data and explore the variables and dimensions of the file.
#
#
# +
# define the helper function
def download_file_to_local(project_filename, local_file_destination=None, project=None):
"""
Uses project-lib to get a bytearray and then downloads this file to local.
Requires a valid `project` object.
Args:
project_filename str: the filename to be passed to get_file
local_file_destination: the filename for the local file if different
Returns:
0 if everything worked
"""
project = project
# get the file
print("Attempting to get file {}".format(project_filename))
_bytes = project.get_file(project_filename).read()
# check for new file name, download the file
print("Downloading...")
if local_file_destination==None: local_file_destination = project_filename
with open(local_file_destination, 'wb') as f:
f.write(bytearray(_bytes))
print("Completed writing to {}".format(local_file_destination))
return 0
# -
download_file_to_local('absolute.nc', project=project)
cfile = "absolute.nc"
# <a id="load1"></a>
# ### 2. Load gridded data with netCDF4
#
# We then use netCDF4's *Dictionary* collection to analyse the data and its relations between the fields that consitute the netCDF file.
dataset = Dataset(cfile)
#
# To extract the data model version of the netCDF file, we use the *data_model* variable is used. The data model can be one of NETCDF3_CLASSIC, NETCDF4, NETCDF4_CLASSIC, NETCDF3_64BIT_OFFSET OR NETCDF3_63BIT_DATA.
#
print(dataset.data_model)
# *dimensions* returns a dictionary with variables names from the dataset mapped to instances of the Dimensions class. It provides the name of the variable along with its size.
print(dataset.dimensions)
# *variables* returns a dictionary that maps the variable names from the dataset as instances of *Variable* class.
print(dataset.variables)
# Below is an example of how variables from the dataset can be accessed as keys of the dictionary returned in the line above.
# +
lons = dataset.variables['lon'][:]
print("Shape of longitude data : ",np.shape(lons))
lats = dataset.variables['lat'][:]
print("Shape of latitude data : ",np.shape(lats))
time = dataset.variables['time'][:]
print("Shape of time data : ",np.shape(time))
temperature = dataset.variables['tem'][:,:,:]
print("Shape of temperature data : ",np.shape(temperature))
# -
# <a id="map1"></a>
# ### 3. Create a global map of the average temperature in January using matplotlib
#
#
# We will now see how matplotlib and its extensions can be used to plot 2D maps in Python. Here we use the matplotlib [basemap](https://matplotlib.org/basemap/users/intro.html) toolkit. To map the points on a 2D surface, basemap supports 24 different types of [projections](https://matplotlib.org/basemap/users/mapsetup.html). In this example Miller Projections is used. Miller projections are generally used for wall maps rather than as navigational maps. Details of Miller projections can be found [here](https://matplotlib.org/basemap/users/mill.html). llcrnrlon, llcrnrlat refer to longitude and latitude of lower left hand corner of the desired map domain(degrees) respectively. urcrnrlon, urcrnrlat refer to longitude and latitude of lower right hand corner of the desired map domain(degrees) respectively.
# +
# define the area to plot and projection to use
m =\
Basemap(llcrnrlon=-180,llcrnrlat=-60,urcrnrlon=180,urcrnrlat=80,projection='mill')
# covert the latitude, longitude and temperatures to raster coordinates to be plotted
t1=temperature[0,:,:]
t1,lon=addcyclic(t1,lons)
january,longitude=shiftgrid(180.,t1,lon,start=False)
x,y=np.meshgrid(longitude,lats)
px,py=m(x,y)
palette=cm.RdYlBu_r
rmin=-30.
rmax=30.
ncont=20
dc=(rmax-rmin)/ncont
vc=arange(rmin,rmax+dc,dc)
pal_norm=matplotlib.colors.Normalize(vmin = rmin, vmax = rmax, clip = False)
m.drawcoastlines(linewidth=0.5)
m.drawmapboundary(fill_color=(1.0,1.0,1.0))
cf=m.pcolormesh(px, py, january, cmap = palette)
cbar=colorbar(cf,orientation='horizontal', shrink=0.95)
cbar.set_label('Mean Temperature in January')
tight_layout()
# -
# *addcyclic* adds a column of longitude to a set of data. In the code below we see that the longitude array is added to an array containing temperature entries. *shiftgrid* moves all longitudes and data east or west. The *meshgrid* method returns co-ordinate matrictes from one dimentional coordinate arrays. In the code below, we use meshgrid to convert longitude and latitude arrays into x and y coordinate arrays.
# <a id="part2"></a>
# ## Part 2 - Weather maps with xarray and Cartopy
#
# In the second half of tutorial, we will see how to use [xarray](http://xarray.pydata.org/en/stable/) to process the netCDF data. xarray is useful with analysing multidimensional arrays. It shares functionalities from pandas and NumPy. xarray has proven to be a robust library to handle netCDF files.
#
# <a id="import2"></a>
# ### 1. Import packages
#
# Following snippet shows the required imports that needs to be done to be able to run the notebook.
import numpy as np
import pandas as pd
import xarray as xr
import matplotlib.pyplot as plt
import cartopy.crs as ccrs
# %matplotlib inline
# <a id="load2"></a>
# ### 2. Load gridded data with xarray
#
# We then open and load the dataset using xarray.
dataset = xr.open_dataset(cfile)
# xarray supports the following data structures :
#
# - *DataArray* which is a multidimensional array
# - *Dataset* which is a dictionaty of multiple DataArray objects.
#
# netCDF data is represented as a Dataset in xarray.
dataset.values
# *dims* returns the value of the x, y and z coordinates.
dataset.dims
# *coords* returns just the coordinates section from the *values* variable we saw above.
dataset.coords
#
# Since xarray is an extension to pandas, it offers a method which enables us to convert the dataset to a dataframe.
df = dataset.to_dataframe()
df.head()
df.describe()
# <a id="map21"></a>
# ### 3. Create maps using xarray
# xarray also supports plotting fuctionalities by extending the *matplotlib* library. DataArray objects can be plotted using xarray libraries. To plot Dataset objects, the relevant DataArrays or dimensions needs to be accessed.
dataset.mean(dim=['time','lon']).to_dataframe().plot()
dataset.tem[0].plot()
# <a id="map22"></a>
# ### 4. Create maps using Cartopy
#
# [Cartopy](https://scitools.org.uk/cartopy/docs/latest/) is one of the several plotting applications that are compatible with xarray. Few others are Seaborn, HoloViews and GeoViews.
#
# Below is a simple example of using cartopy to create visualizations. We compare the Molleweide projection vs the Miller projection. A complete list of projections can be found [here](https://scitools.org.uk/cartopy/docs/latest/crs/projections.html)
# +
f = plt.figure(figsize=(14,5))
plt.title("Molleweide VS Miller Projection for the month of January", fontsize=20)
plt.axis('off')
ax1 = f.add_subplot(1,2,1, projection = ccrs.Mollweide())
ax2 = f.add_subplot(1,2,2, projection = ccrs.Miller())
ax1.coastlines()
ax1.gridlines()
ax2.coastlines()
ax2.gridlines()
dataset.tem[0].plot(ax=ax1, transform=ccrs.PlateCarree())
dataset.tem[0].plot(ax=ax2, transform=ccrs.PlateCarree())
# -
# Following is a heat map comparing the intensity of temperatures between the month of January and June.
# +
proj = ccrs.Miller()
jan_june = dataset.tem.isel(time=[0,5])
months = ['January','June']
i = 0
p = jan_june.plot(transform=ccrs.PlateCarree(),
col='time', col_wrap=2,
aspect=dataset.dims['lon'] / dataset.dims['lat'],
subplot_kws={'projection': proj})
for ax in p.axes.flat:
ax.coastlines()
ax.gridlines()
ax.set_title(months[i])
i = i+1
# -
# ### Author
# <NAME> is a Data & AI Developer Advocate for IBM. She develops and presents talks and workshops about data science and AI. She is active in the local developer communities through attending, presenting and organising meetups. She has a background in climate science where she explored large observational datasets of carbon uptake by forests during her PhD, and global scale weather and climate models as a postdoctoral fellow.
#
# <NAME> is an Advisory Software Engineer with IBM where she currently publishes content that are related to machine learning and deep learning. She is also a full stack software developer, experienced in offering AI based solutions within the healthcare domain. Samaya has her Bachelor of Engineering in Computer Science from College of Engineering, Guindy and her Master of Science in Computer Science from University of Texas at Arlington. She is an ardent learner and a very passionate algorithm solver.
#
# Copyright © 2019 IBM. This notebook and its source code are released under the terms of the MIT License.
| notebooks/raster-data-using-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # I. INTRODUÇÃO
# ## Contexto
#
# Este notebook é resultado da minha prática com dados para treinar estatística com Python.
# + [markdown] colab_type="text" id="U6V7br1RjQKi"
# ## Fonte dos Dados e Variáveis utilizadas
#
# Notebook 1
# -
# ## Objetivos
#
# **Objetivo principal:** treino, prática e aprendizado de componentes de estatística.
# # II. Desenvolvimento
# + [markdown] colab_type="text" id="8Rw70V6ojQKk"
# ### Bibliotecas importadas
# + colab={} colab_type="code" id="8QgOpE-wjQKl"
import pandas as pd
import numpy as np
from scipy.stats import norm
# -
# ### Extração de dados
# + colab={} colab_type="code" id="ufRZ7TLwjQKo"
dados = pd.read_csv('dataset.csv')
# -
# ### Breve visualização dos dados
# Visualizando o início
dados.head(3)
# ## Partes anteriores
#
# ### Parte I: Classificação (notebook 1)
# ### Parte II: Frequências (notebook 1)
# ### Parte III: Medidas de Tendência Central (notebook 2)
# ### Parte IV: Medidas Separatrizes (notebook 2)
# ### Parte V: Distribuições de probabilidade I (notebook 3)
# ### Parte V: Distribuições de probabilidade II (notebook 3)
# ## Parte VII: Amostragem
# **População finita** permite a contagem de seus elementos, como por exemplo o número de clientes de uma empresa. Enquanto que uma **População infinita** não é possível contar seus elementos, como por exemplo a quantidade de porções que se pode extrair da água do mar, ou mesmo uma população finita mas com muitos dados.
#
# **Amostra** é o subconjunto representativo da população. Seus **parâmetros** são seus atributos numéricos, como média, variância e desvio padrão.
#
# Utiliza-se amostras em população infinitas, para resultados rápidos, custo elevado e testes destrutivos.
# ### 7.1. Amostragem Aleatória simples
#
# Cada elemento tem a mesma chance de ser selecionado.
# +
amostra = dados.sample(n = 100, random_state = 101)
print(f'Tamanho da população:\t {dados.shape[0]}')
print(f'Tamanho da amostra:\t {amostra.shape[0]}')
# -
# Parametros
print(f'Média da população:\t {dados.Renda.mean()}')
print(f'Média da amostra:\t {amostra.Renda.mean()}')
# Parametros próximos aos originais, da população
print('Proporção de cada sexo na populacao:')
print(dados.Sexo.value_counts(normalize = True))
print('\nProporção de cada sexo na amostra:')
print(amostra.Sexo.value_counts(normalize = True))
# +
# Aumentando a amostra
amostra = dados.sample(n = 1000, random_state = 101)
print(f'Média da população:\t {dados.Renda.mean()}')
print(f'Média da amostra:\t {amostra.Renda.mean()}')
print('\nProporção de cada sexo na populacao:')
print(dados.Sexo.value_counts(normalize = True))
print('\nProporção de cada sexo na amostra:')
print(amostra.Sexo.value_counts(normalize = True))
# -
# ### 7.2. Teorema do limite central
# + [markdown] colab_type="text" id="QpgTvbISG3GU"
# O **Teorema do Limite Central** afirma que, com o aumento do tamanho da amostra, a distribuição das médias amostrais se aproxima de uma distribuição normal com média igual à média da população e desvio padrão igual ao desvio padrão da variável original dividido pela raiz quadrada do tamanho da amostra. Este fato é assegurado para $n$ maior ou igual a 30.
#
# O desvio padrão das médias amostrais é conhecido como **erro padrão da média**. Fórmula:
#
# $$\sigma_\bar{x} = \frac{\sigma}{\sqrt{n}}$$
#
# -
# **Exemplo:**
# + colab={} colab_type="code" id="S3q73XpnizTi"
# Definindo quantidades
n = 2000 # numero de dados obtidos da população
total_de_amostras = 1500 # total de amostras de 2000 unidades
amostras = pd.DataFrame() # criando um dataframe
# Preenchendo o dataframe de amostras - 1500 amostras com 2mil elementos cada
for i in range(total_de_amostras):
_ = dados.Idade.sample(n)
_.index = range(0, len(_))
amostras['amostra_' + str(i)] = _
# + colab={"base_uri": "https://localhost:8080/", "height": 1989} colab_type="code" id="tXa7XKhKTyMu" outputId="2fe05b86-ecc8-48af-be13-17e8c7c235ce"
amostras
# + [markdown] colab_type="text" id="JfgEBDx0jDzf"
# O Teorema do Limite Central afirma que, **com o aumento do tamanho da amostra, a distribuição das médias amostrais se aproxima de uma distribuição normal** com média igual à média da população.
# -
# Graficos para comparação
dados.Idade.hist()
amostras.mean().hist()
# Valores
print(f'Médias:\nPopulação:\t{dados.Idade.mean()}\nAmostra:\t{amostras.mean().mean()}')
# + [markdown] colab_type="text" id="7lIWJCAKlPmA"
# O Teorema do Limite Central afirma que o **desvio padrão igual ao desvio padrão da variável original dividido pela raiz quadrada do tamanho da amostra**.
# +
print(f'\nDesvio-padrão:\nPopulação:\t{dados.Idade.std()}\nAmostra:\t{amostras.mean().std()}')
print(f'\nErro padrão da média: desvio padrão das médias amostrais\n{dados.Idade.std() / np.sqrt(n)}')
# + [markdown] colab_type="text" id="ku8CE67iG3Ge"
# ### 7.3. Níveis de confiança e significância
# + [markdown] colab_type="text" id="mk6ZsvgyG3Gf"
# **Nível de confiança** ($1 - \alpha$) é a probabilidade de acerto da estimativa.
# **Nível de significância** ($\alpha$) é a probabilidade de erro da estimativa.
#
# **Nível de confiança** é o grau de confiabilidade do resultado da estimativa estar dentro de determinado intervalo. Quando fixamos em uma pesquisa um **nível de confiança** de 95%, por exemplo, estamos assumindo que existe uma probabilidade de 95% dos resultados da pesquisa representarem bem a realidade, ou seja, estarem corretos. O *nível de confiança* de uma estimativa pode ser obtido a partir da área sob a curva normal.
# + [markdown] colab_type="text" id="tk-TwhhfG3Gi"
# **Erro inferencial** é definido pelo **desvio padrão das médias amostrais** $\sigma_\bar{x}$ e pelo **nível de confiança** determinado para o processo.
#
# $$e = z \frac{\sigma}{\sqrt{n}}$$
# + [markdown] colab_type="text" id="-r6EMnH-G3HT"
# **Intevalo de confiança para a média da população**
# - Com desvio padrão populacional conhecido:
#
# $$\mu = \bar{x} \pm z\frac{\sigma}{\sqrt{n}}$$
#
# - Com desvio padrão populacional desconhecido:
#
# $$\mu = \bar{x} \pm z\frac{s}{\sqrt{n}}$$
# + [markdown] colab_type="text" id="AgnrFjdMG3HT"
# **Exemplo:** Arroz
#
# Suponha que os pesos dos sacos de arroz de uma indústria alimentícia se distribuem aproximadamente como uma normal de *desvio padrão populacional igual a 150 g*. Selecionada uma *amostra aleatório de 20 sacos* de um lote específico, obteve-se um *peso médio de 5.050 g*. Construa um intervalo de confiança para a *média populacional* assumindo um *nível de significância de 5%*.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="aF2QQ0z3G3HU" outputId="31b20b10-a4f7-49fe-b113-f0fff1e2f30f"
# Texto fornece
media_amostra = 5050
n = 20
significancia = 0.05 # alfa
confianca = 1 - significancia
desvio_padrao = 150
# +
from scipy.stats import norm
import numpy as np
# Obtendo valores
# area sob a curva: 0.5 + (0.95 / 2)
z = norm.ppf(0.975)
sigma = desvio_padrao / np.sqrt(n)
e = z * sigma
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ebvikT95G3Hi" outputId="bfbc2f5c-e706-46a6-aa6c-e94632aed4fc"
intervalo_manual = (
media_amostra - e,
media_amostra + e
)
intervalo_manual
intervalo_pro = norm.interval(alpha = 0.95, loc = media_amostra, scale = sigma)
intervalo_pro
# -
# ### 7.4. Tamanho da amostra: População infinita
# + [markdown] colab_type="text" id="X1AxpKtIG3Gn"
# **Com desvio padrão conhecido**
#
# $$n = \left(z\frac{\sigma}{e}\right)^2$$
#
# **Com desvio padrão DESconhecido**
#
# $$n = \left(z\frac{s}{e}\right)^2$$
#
#
# Onde:
#
# $z$ = variável normal padronizada
#
# $\sigma$ = desvio padrão populacional
#
# $s$ = desvio padrão amostral
#
# $e$ = erro inferencial
#
# *Observações:**
# - O desvio padrão ($\sigma$ ou $s$) e o erro ($e$) devem estar na mesma unidade de medida.
# - Quando o erro ($e$) for representado em termos percentuais, deve ser interpretado como um percentual relacionado à média.
# + [markdown] colab_type="text" id="dcdzo7EgG3Go"
# **Exemplo:** Rendimento médio
#
# Estamos estudando o rendimento mensal dos chefes de domicílios no Brasil. Nosso supervisor determinou que o **erro máximo em relação a média seja de R$\$$ 100,00**. Sabemos que o **desvio padrão populacional** deste grupo de trabalhadores é de **R$\$$ 3.323,39**. Para um **nível de confiança de 95%**, qual deve ser o tamanho da amostra de nosso estudo?
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="XtJ5I_kaKDN3" outputId="a519e330-ddf0-492c-f42d-5cef36fed4c9"
# Dados do texto
z = norm.ppf(0.975)
sigma = 3323.39
e = 100
# Calculando n
n = (z * (sigma / e)) ** 2
print(f'Tamanho da amostra será de: {int(n.round())}')
# + [markdown] colab_type="text" id="FlAz5kycTyNQ"
# ### 7.5. Tamanho da amostra: população finita
# + [markdown] colab_type="text" id="UOe02McSTyNR"
# **Com desvio padrão conhecido**
#
# $$n = \frac{z^2 \sigma^2 N}{z^2 \sigma^2 + e^2(N-1)}$$
#
# **Com desvio padrão desconhecido**
#
# $$n = \frac{z^2 s^2 N}{z^2 s^2 + e^2(N-1)}$$
#
#
# Onde:
#
# $N$ = tamanho da população
#
# $z$ = variável normal padronizada
#
# $\sigma$ = desvio padrão populacional
#
# $s$ = desvio padrão amostral
#
# $e$ = erro inferencial
# + [markdown] colab_type="text" id="upv1mM7fTyNS"
# **Exemplo:** Indústria de refrigerante
#
# Em um lote de **10.000 latas** de refrigerante foi realizada uma amostra aleatória simples de **100 latas** e foi obtido o **desvio padrão amostral do conteúdo das latas igual a 12 ml**. O fabricante estipula um **erro máximo sobre a média populacional de apenas 5 ml**. Para garantir um **nível de confiança de 95%** qual o tamanho de amostra deve ser selecionado para este estudo?
# + [markdown] colab_type="text" id="_y3-3VIBTyNS"
# ### Obtendo $N$
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="gXWn9zTETyNS" outputId="4daa26ff-4a96-47fb-9f0e-53a4390d701e"
# Dados do texto
N = 1000
z = norm.ppf((0.5 + (0.95 / 2)))
s = 12
e = 5
# Obtendo n
n = ((z**2) * (s**2) * (N)) / (((z**2) * (s**2)) + ((e**2) * (N - 1)))
print(f'Tamanho da amostra será de: {int(n.round())}.')
| exercises_statistic/IBGE_project/analise-estatistica-pt4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="uKgZQAB_cS3X" outputId="131d5d34-91fc-40c8-f184-80b080f11385"
# Connect to Google Drive
from google.colab import drive
drive.mount('/content/drive')
# + id="MEFVARL0c4KA"
# Copy the dataset from Google Drive to local
# !cp "/content/drive/MyDrive/CBIS_DDSM.zip" .
# !unzip -qq CBIS_DDSM.zip
# !rm CBIS_DDSM.zip
cbis_path = 'CBIS_DDSM'
# + id="wJrsTUufdDY7"
# Import libraries
# %tensorflow_version 1.x
import os
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import itertools
from sklearn.metrics import confusion_matrix, roc_curve, auc, classification_report
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras import models
from tensorflow.keras import optimizers
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LearningRateScheduler, Callback
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras.regularizers import l2
from tensorflow.keras.utils import plot_model
from tensorflow.keras.utils import to_categorical
# + [markdown] id="OG3U3WgedKBo"
# # Data pre-processing
# + id="hqJCZw80dLIy"
def load_training():
"""
Load the training set (excluding baseline patches)
"""
images = np.load(os.path.join(cbis_path, 'numpy data', 'train_tensor.npy'))[1::2]
labels = np.load(os.path.join(cbis_path, 'numpy data', 'train_labels.npy'))[1::2]
return images, labels
def load_testing():
"""
Load the test set (abnormalities patches and labels, no baseline)
"""
images = np.load(os.path.join(cbis_path, 'numpy data', 'public_test_tensor.npy'))[1::2]
labels = np.load(os.path.join(cbis_path, 'numpy data', 'public_test_labels.npy'))[1::2]
return images, labels
def remap_label(l):
"""
Remap the labels to:
0 -> mass benign
1 -> mass malignant
2 -> calcification benign
3 -> calcification malignant
"""
if 1 <= l <= 4:
return l-1
else:
print("[WARN] Unrecognized label (%d)" % l)
return None
# + [markdown] id="YMNT6LKxdSHi"
# The data is prepared following these steps:
#
# 1. Import the training and testing data from numpy arrays
#
# 2. Remove the images and labels related to baseline patches (even indices in the arrays)
#
# 3. Adjust the labels for the classification problem, so that 0 corresponds to 'benign mass', 1 to 'malignant mass', 2 to 'bening calcification' and 3 to 'malignant calcification'
#
# 4. Convert the labels to categorical format, required by the categorical_crossentropy loss function
#
# 5. Normalize the pixels to be in the range (0-1) floating point
#
# 6. Shuffle the training set (and labels accordingly, of course)
#
# 7. Split the training data into 'training' and 'validation' subsets
#
# 8. Build Keras generators for training and validation data. Note that data augmentation is used from the beginning, as its value was proven in the previous notebook (Scratch_CNN_2_class)
# + colab={"base_uri": "https://localhost:8080/"} id="g3S8M8Pedd5n" outputId="e3a4bb55-a8ac-491a-8918-defc0179a989"
# Load training and test images (abnormalities only, no baseline)
train_images, train_labels= load_training()
test_images, test_labels= load_testing()
# Number of images
n_train_img = train_images.shape[0]
n_test_img = test_images.shape[0]
print("Train size: %d \t Test size: %d" % (n_train_img, n_test_img))
# Compute width and height of images
img_w = train_images.shape[1]
img_h = train_images.shape[2]
print("Image size: %dx%d" % (img_w, img_h))
# Remap labels
train_labels = np.array([remap_label(l) for l in train_labels])
test_labels = np.array([remap_label(l) for l in test_labels])
# Convert the labels to categorical format
train_labels = to_categorical(train_labels)
test_labels_raw = test_labels.copy()
test_labels = to_categorical(test_labels)
# Create a new dimension for color in the images arrays
train_images = train_images.reshape((n_train_img, img_w, img_h, 1))
test_images = test_images.reshape((n_test_img, img_w, img_h, 1))
# Convert from 16-bit (0-65535) to float (0-1)
train_images = train_images.astype('uint16') / 65535
test_images = test_images.astype('uint16') / 65535
# Shuffle the training set (originally sorted by label)
perm = np.random.permutation(n_train_img)
train_images = train_images[perm]
train_labels = train_labels[perm]
# Create a generator for training images
train_datagen = ImageDataGenerator(
validation_split=0.2,
rotation_range=180,
shear_range=10,
zoom_range=0.2,
horizontal_flip=True,
vertical_flip=True,
fill_mode='reflect'
)
# Fit the generator with some images
train_datagen.fit(train_images)
# Split train images into actual training and validation
train_generator = train_datagen.flow(train_images, train_labels, batch_size=128, subset='training')
validation_generator = train_datagen.flow(train_images, train_labels, batch_size=128, subset='validation')
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="2k7G_ZMGdenH" outputId="36542130-555b-412c-d33f-3c3401e746e9"
# Visualize one image from the dataset and its label, just to make sure the data format is correct
idx = 0
plt.imshow(train_images[idx][:,:,0], cmap='gray')
plt.show()
print("Label: " + str(train_labels[idx]))
# + [markdown] id="DeWPZez4ekuu"
# # Experiment 3
# + [markdown] id="UaAOfZLTeoUF"
#
# What about another convolutional block? Does it help distinguishing between malignant and benign bodies? The next experiment provides an answer to these question, by training a deeper model with an extra Conv2D layer with 512 features.
# + colab={"base_uri": "https://localhost:8080/"} id="Byf3ZaGHeRUs" outputId="b25687be-0a88-46de-b6fb-254cc2299ffc"
# Model 3
model_3 = models.Sequential()
model_3.add(layers.Conv2D(64, (3, 3), activation='relu', input_shape=(150, 150, 1)))
model_3.add(layers.MaxPooling2D((2, 2)))
model_3.add(layers.Conv2D(128, (3, 3), activation='relu'))
model_3.add(layers.MaxPooling2D((2, 2)))
model_3.add(layers.Conv2D(256, (3, 3), activation='relu'))
model_3.add(layers.MaxPooling2D((2, 2)))
model_3.add(layers.Conv2D(512, (3, 3), activation='relu'))
model_3.add(layers.MaxPooling2D((2, 2)))
model_3.add(layers.Flatten())
model_3.add(layers.Dense(64, activation='relu'))
model_3.add(layers.Dropout(0.5))
model_3.add(layers.Dense(4, activation='softmax'))
model_3.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="-H3cCynreqNR" outputId="89288e79-527c-409f-b877-e486c174b28a"
# Early stopping (stop training after the validation loss reaches the minimum)
earlystopping = EarlyStopping(monitor='val_loss', mode='min', patience=80, verbose=1)
# Callback for checkpointing
checkpoint = ModelCheckpoint('model_3_4cl_best.h5',
monitor='val_loss', mode='min', verbose=1,
save_best_only=True, save_freq='epoch'
)
# Compile the model
model_3.compile(optimizer=RMSprop(learning_rate=0.001, decay=5e-4), loss='categorical_crossentropy', metrics=['accuracy'])
# Train
history_3 = model_3.fit_generator(
train_generator,
steps_per_epoch=int(0.8*n_train_img) // 128,
epochs=500,
validation_data=validation_generator,
callbacks=[checkpoint, earlystopping],
shuffle=True,
verbose=1,
initial_epoch=0
)
# Save
models.save_model(model_3, 'model_3_4cl_end.h5')
# + id="B7fy4aoKeu5u"
# !cp model* "/content/drive/MyDrive/models/"
# + colab={"base_uri": "https://localhost:8080/"} id="HZESE_Yhe1ZM" outputId="489c579b-b639-4144-a28d-d7ae8dd3b308"
# History of accuracy and loss
tra_loss_3 = history_3.history['loss']
tra_acc_3 = history_3.history['acc']
val_loss_3 = history_3.history['val_loss']
val_acc_3 = history_3.history['val_acc']
# Total number of epochs training
epochs_3 = range(1, len(tra_acc_3)+1)
end_epoch_3 = len(tra_acc_3)
# Epoch when reached the validation loss minimum
opt_epoch_3 = val_loss_3.index(min(val_loss_3)) + 1
# Loss and accuracy on the validation set
end_val_loss_3 = val_loss_3[-1]
end_val_acc_3 = val_acc_3[-1]
opt_val_loss_3 = val_loss_3[opt_epoch_3-1]
opt_val_acc_3 = val_acc_3[opt_epoch_3-1]
# Loss and accuracy on the test set
opt_model_3 = models.load_model('model_3_4cl_best.h5')
test_loss_3, test_acc_3 = model_3.evaluate(test_images, test_labels, verbose=False)
opt_test_loss_3, opt_test_acc_3 = opt_model_3.evaluate(test_images, test_labels, verbose=False)
opt_pred_3 = opt_model_3.predict([test_images, test_labels])
pred_classes_3 = np.rint(opt_pred_3)
print("Model 3\n")
print("Epoch [end]: %d" % end_epoch_3)
print("Epoch [opt]: %d" % opt_epoch_3)
print("Valid accuracy [end]: %.4f" % end_val_acc_3)
print("Valid accuracy [opt]: %.4f" % opt_val_acc_3)
print("Test accuracy [end]: %.4f" % test_acc_3)
print("Test accuracy [opt]: %.4f" % opt_test_acc_3)
print("Valid loss [end]: %.4f" % end_val_loss_3)
print("Valid loss [opt]: %.4f" % opt_val_loss_3)
print("Test loss [end]: %.4f" % test_loss_3)
print("Test loss [opt]: %.4f" % opt_test_loss_3)
print(classification_report(test_labels, pred_classes_3, digits=4))
# + colab={"base_uri": "https://localhost:8080/", "height": 991} id="xYtAMb-Pe4vx" outputId="5249f39e-cecb-4941-b64b-6ee304500ac2"
# Model accuracy
plt.figure(figsize=(7, 7), dpi=80, facecolor='w', edgecolor='k')
plt.title('Model 3 accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.plot(epochs_3, tra_acc_3, 'r', label='Training set')
plt.plot(epochs_3, val_acc_3, 'g', label='Validation set')
plt.plot(opt_epoch_3, val_acc_3[opt_epoch_3-1], 'go')
plt.vlines(opt_epoch_3, min(val_acc_3), opt_val_acc_3, linestyle="dashed", color='g', linewidth=1)
plt.hlines(opt_val_acc_3, 1, opt_epoch_3, linestyle="dashed", color='g', linewidth=1)
plt.legend(loc='lower right')
# Model loss
plt.figure(figsize=(7, 7), dpi=80, facecolor='w', edgecolor='k')
plt.title('Model 3 loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.plot(epochs_3, tra_loss_3, 'r', label='Training set')
plt.plot(epochs_3, val_loss_3, 'g', label='Validation set')
plt.plot(opt_epoch_3, val_loss_3[opt_epoch_3-1], 'go')
plt.vlines(opt_epoch_3, min(val_loss_3), opt_val_loss_3, linestyle="dashed", color='g', linewidth=1)
plt.hlines(opt_val_loss_3, 1, opt_epoch_3, linestyle="dashed", color='g', linewidth=1)
plt.legend();
# + id="SstbNtcKltTI"
def plot_confusion_matrix(cm, classes, normalize=False, title='Confusion matrix', cmap=plt.cm.Blues):
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + colab={"base_uri": "https://localhost:8080/", "height": 311} id="rCyGwlSelXM7" outputId="c0207218-97b4-4767-b879-40b27ad7ed3a"
pred_3_classes = opt_model_3.predict_classes(test_images)
confusion_mtx = confusion_matrix(test_labels_raw, pred_3_classes)
plot_confusion_matrix(confusion_mtx, classes=range(4), title='Model 3 confusion matrix')
# + id="4s4ORNHnlm_i"
| Scratch_CNN_4_class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unit 5 - Financial Planning
#
# +
# Initial imports for the libraries we will be using to analyze our data
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import alpaca_trade_api as tradeapi
from MCForecastTools import MCSimulation
# %matplotlib inline
# -
# Loading .env enviroment variables
load_dotenv()
# ## Part 1 - Personal Finance Planner
# ### Collect Crypto Prices Using the `requests` Library
# +
# Setting the current amount of crypto assets in posession with variables
my_btc = 1.2
my_eth = 5.3
# -
# Listing the Crypto API URLs as variable
btc_url = "https://api.alternative.me/v2/ticker/Bitcoin/?convert=CAD"
eth_url = "https://api.alternative.me/v2/ticker/Ethereum/?convert=CAD"
# +
# Fetching the current BTC price using request
# Turned Fetched data into a readable Json file to see where the current CAD price is located in the dictionary
btc_data = requests.get(btc_url).json()
import json
print(json.dumps(btc_data, indent=10))
# -
#Declaring the btc CAD price location within the dictionary file we fetched from the API URL
btc_price = btc_data["data"]["1"]["quotes"]["CAD"]["price"]
# +
# Fetchting the current ETH price, going through the same steps as above with BTC.
eth_data = requests.get(eth_url).json()
import json
print(json.dumps(eth_data, indent=10))
# -
#Declaring the eth CAD price location within the dictionary file we fetched from the API URL
eth_price = eth_data["data"]["1027"]["quotes"]["CAD"]["price"]
# +
# Computing the current value of the crypto portfolio, printing the results to check if the values are correct
print(btc_price)
print(eth_price)
my_btc_value = btc_price * my_btc
my_eth_value = eth_price * my_eth
# Here we are calculating the combined value of the crypto portfolio by adding the ETH and BTC values
total_crypto_value = round((my_btc_value + my_eth_value),2)
# -
# Printing the current crypto wallet balances
print(f"The current value of your {my_btc} BTC is ${my_btc_value:0.2f} CAD")
print(f"The current value of your {my_eth} ETH is ${my_eth_value:0.2f} CAD")
print(f"The current combined value of your cryptos is ${total_crypto_value:0.2f} CAD")
# ### Collect Investments Data Using Alpaca: `SPY` (stocks) and `AGG` (bonds)
# setting the Current amount of shares as variables
my_spy = 200
my_agg = 50
# +
# Setting the Alpaca API key and secret-key inorder to fetch data from alpaca website.
alpaca_api_key = os.getenv("ALPACA_API_KEY")
alpaca_secret_key = os.getenv("ALPACA_SECRET_KEY")
# Created an Alpaca API object to tell alpaca what data we want to acesss, in this case its from the base URL
api = tradeapi.REST(
alpaca_api_key,
alpaca_secret_key,
base_url="https://paper-api.alpaca.markets",
api_version = "v2"
)
# +
# Formating thr current date as ISO format, so alpaca understands which dates we want to retrive
today = pd.Timestamp("2021-03-12", tz="America/New_York").isoformat()
# Setting the tickers we want to retrive for our portfolio data
tickers = ["AGG","SPY"]
# Setting the timeframe to '1D' for the Alpaca API- this brings us the daily data
timeframe = "1D"
# Getting current closing prices for SPY and AGG using a alpaca SDK
# we are also setting the start and end date as the same to get data on the latest date
df_ticker = api.get_barset(
tickers,
timeframe,
start=today,
end=today,
limit=1000
).df
# Preview DataFrame
df_ticker.head()
# +
# Picked AGG and SPY close prices
# here we are creating a new data frame to house only the closing prices
df_closing_prices = pd.DataFrame()
df_closing_prices["AGG"] = df_ticker["AGG"]["close"]
df_closing_prices["SPY"] = df_ticker["SPY"]["close"]
# -
# Here we are setting the values for the closing prices
agg_close_price = df_closing_prices.iloc[0][0]
spy_close_price = df_closing_prices.iloc[0][1]
# Printing AGG and SPY closing prices
print(f"Current AGG closing price: ${agg_close_price}")
print(f"Current SPY closing price: ${spy_close_price}")
# +
# Computing the current value of shares in the portfolio
# Here we are multiplying the closing price by the number of shares/bonds they currently hold.
my_spy_value = spy_close_price * my_spy
my_agg_value = agg_close_price * my_agg
# By adding both the total SPY value and AGG value we get the total value of the portfolio and rounding the output to 2 decimals.
total_share_value = round((my_spy_value + my_agg_value),2)
# Print current value of shares/bonds
print(f"The current value of your {my_spy} SPY shares is ${my_spy_value:0.2f}")
print(f"The current value of your {my_agg} AGG shares is ${my_agg_value:0.2f}")
print(f"The current combined value of your AGG & SPY shares is ${total_share_value:0.2f}")
# -
# ### Savings Health Analysis
# Setting the monthly household income variable
monthly_income = 12000.0
# +
# Creating a savings DataFrame to house the summary data
# I am setting the index column to display the two types of asset classes in our portfolio, which are shares and crypto
# As well as creating a column to display the total amount currently in each asset class
amount_data = {
"Amount":[total_share_value,total_crypto_value]
}
asset_class = ["Shares","Crypto"]
df_savings = pd.DataFrame(amount_data,index=asset_class)
# Displaying the savings DataFrame
display(df_savings)
# -
# Plotting the savings portfolio pie chart
df_savings.plot.pie(y="Amount", title="Portfolio Composition")
# +
# calculating the ideal emergency fund total
emergency_fund = monthly_income * 3
# Calculating the total amount of savings they currently have
total_savings = total_share_value + total_crypto_value
emergency_fund_diff = emergency_fund - total_savings
# Validating the person's savings health
# If their total savings is greater than the emergency fund amount, they have more money than their savings goal.
# If their total savings is equal to the emergency fund amount, they've reached their savings goal.
# If their total savings is less than the emergency fund ammount, they have not met their savings goal.
if total_savings > emergency_fund:
print(f"Congratulations! You have enough money in your emergency fund.")
elif total_savings == emergency_fund:
print(f"Congratulations! You have reached your emergency fund financial goal.")
elif total_savings < emergency_fund:
print(f"You are ${emergency_fund_diff} , away from your emergency savings goal.")
# -
# ## Part 2 - Retirement Planning
#
# ### Monte Carlo Simulation
# +
# Setting the start and end dates of five years back from today.
# Sample results may vary from the solution based on the time frame chosen
start_date = pd.Timestamp('2016-03-12', tz='America/New_York').isoformat()
end_date = pd.Timestamp('2021-03-12', tz='America/New_York').isoformat()
# +
# Getting 5 years' worth of historical data for SPY and AGG
# Was only able to fetch 1000 lines of data per symbol. Limited by alpaca api
df_stock_data = api.get_barset(
tickers,
timeframe,
start=start_date,
end=end_date,
limit=1000
).df
# Display sample data
df_stock_data.head()
# -
# Configuring a Monte Carlo simulation to forecast 30 years cumulative returns
# Setting the number of simulations to 500, with weights of 40% bonds, and 60% stocks
mc_portfolio = MCSimulation(
portfolio_data = df_stock_data,
weights = [.40,0.60],
num_simulation = 500,
num_trading_days = 252*30
)
# Printing the simulation input data
mc_portfolio.portfolio_data.head()
# Running a Monte Carlo simulation to forecast 30 years cumulative returns
mc_portfolio.calc_cumulative_return()
# Plotting simulation outcomes
porfolio_line_plot = mc_portfolio.plot_simulation()
# Plot probability distribution and confidence intervals
portfolio_dist_plot = mc_portfolio.plot_distribution()
# ### Retirement Analysis
# +
# Fetching summary statistics from the Monte Carlo simulation results
portfolio_stats = mc_portfolio.summarize_cumulative_return()
# Print summary statistics
print(portfolio_stats)
# -
# ### Calculate the expected portfolio return at the 95% lower and upper confidence intervals based on a `$20,000` initial investment.
# +
# Setting the initial investment amount
initial_investment = 20000.00
# Using the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $20,000 investment
ci_lower = round(portfolio_stats[8]*initial_investment,2)
ci_upper = round(portfolio_stats[9]*initial_investment,2)
# Printing the results of our investment
print(f"There is a 95% chance that an initial investment of ${initial_investment} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower} and ${ci_upper}")
# -
# ### Calculate the expected portfolio return at the `95%` lower and upper confidence intervals based on a `50%` increase in the initial investment.
# +
# Setting the initial investment amount to 1.5 times as before
initial_investment_v2 = 20000 * 1.5
# Using the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $30,000 invesment
ci_lower_2 = round(portfolio_stats[8]*initial_investment_v2,2)
ci_upper_2 = round(portfolio_stats[9]*initial_investment_v2,2)
# Printing the results for our investment
print(f"There is a 95% chance that an initial investment of ${initial_investment_v2} in the portfolio"
f" over the next 30 years will end within in the range of"
f" ${ci_lower_2} and ${ci_upper_2}")
# -
# ## Optional Challenge - Early Retirement
#
#
# ### Five Years Retirement Option
# +
# Configuring a Monte Carlo simulation to forecast 5 years cumulative returns
# Changing the weights of the portfolio to 20% bonds and 80% stocks
mc_portfolio_5yr = MCSimulation(
portfolio_data = df_stock_data,
weights = [.20,0.80],
num_simulation = 500,
num_trading_days = 252*5
)
# -
# Running a Monte Carlo simulation to forecast our 5 years cumulative returns
mc_portfolio_5yr.calc_cumulative_return()
# Plotting the simulation outcomes
porfolio_5yr_line_plot = mc_portfolio_5yr.plot_simulation()
# Plotting the probability distribution and confidence intervals
portfolio_5yr_dist_plot = mc_portfolio_5yr.plot_distribution()
# +
# Fetching the summary statistics from the Monte Carlo simulation results
portfolio_5yr_stats = mc_portfolio_5yr.summarize_cumulative_return()
# Printing summary statistics
print(portfolio_5yr_stats)
# +
# Setting our initial investment to $60000 dollars to see what difference it makes, if more money is added
initial_investment_5yr = 60000.00
# Using the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000 investment
ci_lower_five = round(portfolio_5yr_stats[8]*initial_investment_5yr,2)
ci_upper_five = round(portfolio_5yr_stats[9]*initial_investment_5yr,2)
# Printing the results
print(f"There is a 95% chance that an initial investment of ${initial_investment_5yr} in the portfolio"
f" over the next 5 years will end within in the range of"
f" ${ci_lower_five} and ${ci_upper_five}")
# -
# ### Ten Years Retirement Option
# +
# Configuring a Monte Carlo simulation to forecast 10 years of cumulative returns
# with the same portfolio weights as before, 20% bonds and 80% stocks
mc_portfolio_10yr = MCSimulation(
portfolio_data = df_stock_data,
weights = [.20,0.80],
num_simulation = 500,
num_trading_days = 252*10
)
# -
# Running a Monte Carlo simulation to forecast 10 years cumulative returns
mc_portfolio_10yr.calc_cumulative_return()
# Plotting the simulation outcomes
porfolio_10yr_line_plot = mc_portfolio_10yr.plot_simulation()
# Plotting the probability distribution and confidence intervals
portfolio_10yr_dist_plot = mc_portfolio_10yr.plot_distribution()
# +
# Fetching our summary statistics from the 10yr Monte Carlo simulation results
portfolio_10yr_stats = mc_portfolio_10yr.summarize_cumulative_return()
# Printing summary statistics
print(portfolio_10yr_stats)
# +
# Setting our 10 yr initial investment to $60000 the same as before
initial_investment_10yr = 60000.00
# Using the lower and upper `95%` confidence intervals to calculate the range of the possible outcomes of our $60,000 investment
ci_lower_ten = round(portfolio_10yr_stats[8]*initial_investment_10yr,2)
ci_upper_ten = round(portfolio_10yr_stats[9]*initial_investment_10yr,2)
# Printing our simulation results
print(f"There is a 95% chance that an initial investment of ${initial_investment_10yr} in the portfolio"
f" over the next 10 years will end within in the range of"
f" ${ci_lower_ten} and ${ci_upper_ten}")
| Starter_Code/financial-planner.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 5-2. Quantum Circuit learning
# **Quantum Circuit Learning** (量子回路学習, QCL)は、量子コンピュータを機械学習に応用するためのアルゴリズムである[[1]](https://arxiv.org/abs/1803.00745)。
# 前節で学んだVQE (Variational Quantum Eigensolver)と同様、**量子・古典ハイブリッドアルゴリズム**であり、誤り訂正機能を持たない中規模の量子コンピュータである**NISQ** (Noisy Intermediate-Scale Quantum Computer)での動作を念頭に設計されている。実際のNISQデバイスを用いた実験もすでに行われており、2019年3月にIBMの実験チームによるQCLの実機実装論文[[2]](https://www.nature.com/articles/s41586-019-0980-2)がNatureに掲載され話題となった。
#
# 以下では、まずアルゴリズムの概要と具体的な学習の手順を紹介し、最後に量子シミュレータQulacsを用いた実装例を提示する。
# ### QCLの概要
#
# 近年、機械学習の分野においてディープラーニングが脚光を浴びている。
# ディープラーニングにおいては深い**ニューラルネットワーク**を用いて複雑な関数の近似を行うことで、入力と出力の関係を学習し、新しいデータに対して予測を行う事ができる。
#
# QCLは、このニューラルネットワークを**量子回路**、すなわち量子コンピュータに置き換えた機械学習手法である。量子回路を用いることで、量子力学の重ね合わせの原理を生かして**指数関数的に多数の基底関数**を用いて学習できるため、モデルの表現力が向上する。さらに、量子回路の満たすべき条件(ユニタリ性)により、**自動的にoverfittingを防げる**と考えられている。これにより、古典コンピュータにおける機械学習を超える、さらなる高性能化が期待できる。
# (詳細は参考文献[1]を参照のこと)
#
# ニューラルネットワークにおいては、各層の重みパラメータ$W$を調整することで関数の近似を行っているが、QCLにおいても考え方は全く同様である。
# すなわち、QCLで用いる量子回路には複数の「回転ゲート」が含まれているが、この回転ゲートの回転角$\theta$を調整することで、関数の近似を行うのである。具体的な手順を以下に示す。
# ### 学習の手順
# 1. 学習データ $\{(x_i, y_i)\}_i$ を用意する($x_i$は入力データ、$y_i$は$x_i$から予測したい正解データ(教師データ))
# 2. $U_{in}(x)$という、入力$x$から何らかの規則で決まる回路を用意し、$x_i$の情報を埋め込んだ入力状態$\{|\psi_{\rm in}(x_i)\rangle\}_i = \{U_{in}(x_i)|0\rangle\}_i$ を作る
# 3. 入力状態に、パラメータ$\theta$に依存したゲート$U(\theta)$を掛けたものを出力状態$\{|\psi_{\rm out}(x_i, \theta)\rangle = U(\theta)|\psi_{\rm in}(x_i)\rangle \}_i$とする
# 4. 出力状態のもとで何らかのオブザーバブルを測定し、測定値を得る(例:1番目のqubitの$Z$の期待値$\langle Z_1\rangle = \langle \psi_{\rm out} |Z_1|\psi_{\rm out} \rangle$)
# 5. $F$を適当な関数(sigmoidとかsoftmaxとか定数倍とか何でもいい)として、$F(測定値_i)$をモデルの出力$y(x_i, \theta)$とする
# 6. 正解データ$\{y_i\}_i$とモデルの出力$\{y(x_i, \theta)\}_i$の間の乖離を表す「コスト関数$L(\theta)$」を計算する
# 7. コスト関数を最小化する$\theta=\theta^*$を求める
# 8. $y(x, \theta^*)$が、所望の予測モデルである
#
#
# 
# (QCLでは、入力データ $x$ をまず$U_{in}(x)$を用いて量子状態に変換し、そこから変分量子回路$U(\theta)$と測定等を用いて出力$y$を得る(図では出力は$\langle B(x,\theta)\rangle$)。出典:参考文献[1]の図1を改変)
# ### 量子シミュレータQulacsを用いた実装
# 以下では関数の近似のデモンストレーションとして、sin関数 $y=\sin(\pi x)$ のフィッティングを行う。
import numpy as np
import matplotlib.pyplot as plt
from functools import reduce
# +
######## パラメータ #############
nqubit = 3 ## qubitの数
c_depth = 3 ## circuitの深さ
time_step = 0.77 ## ランダムハミルトニアンによる時間発展の経過時間
## [x_min, x_max]のうち, ランダムにnum_x_train個の点をとって教師データとする.
x_min = - 1.; x_max = 1.;
num_x_train = 50
## 学習したい1変数関数
func_to_learn = lambda x: np.sin(x*np.pi)
## 乱数のシード
random_seed = 0
## 乱数発生器の初期化
np.random.seed(random_seed)
# -
# #### 学習データの準備
# +
#### 教師データを準備
x_train = x_min + (x_max - x_min) * np.random.rand(num_x_train)
y_train = func_to_learn(x_train)
# 現実のデータを用いる場合を想定し、きれいなsin関数にノイズを付加
mag_noise = 0.05
y_train = y_train + mag_noise * np.random.randn(num_x_train)
plt.plot(x_train, y_train, "o"); plt.show()
# -
# #### 入力状態の構成
# まず、入力値$x_i$を初期状態$|00\ldots0\rangle$に埋め込むためのゲート $U_{\rm in}(x_i)$を作成する。
# 参考文献[1]に従い、回転ゲート$R_j^X(\theta)=e^{-i\theta X_j/2}, R_j^Z(\theta)=e^{-i\theta Z_j/2}$を用いて$U_{\rm in}(x) = \prod_j R^Z_j(\cos^{-1} x^2) R^Y_j(\sin^{-1}x)$と定義する。
# 入力値$x_i$は、$|\psi_{\rm in}(x_i)\rangle =U_{\rm in}(x_i)|00\ldots0\rangle$ という量子状態に変換されることになる。
## Google Colaboratoryの場合・Qulacsがインストールされていないlocal環境の場合のみ実行してください
# !pip install qulacs
# +
# 初期状態の作成
from qulacs import QuantumState, QuantumCircuit
state = QuantumState(nqubit) # 初期状態 |000>
state.set_zero_state()
print(state.get_vector())
# -
# xをエンコードするゲートを作成する関数
def U_in(x):
U = QuantumCircuit(nqubit)
angle_y = np.arcsin(x)
angle_z = np.arccos(x**2)
for i in range(nqubit):
U.add_RY_gate(i, angle_y)
U.add_RZ_gate(i, angle_z)
return U
# 入力状態を試す
x = 0.1 # 適当な値
U_in(x).update_quantum_state(state) # U_in|000>の計算
print(state.get_vector())
# #### 変分量子回路$U(\theta)$の構成
# 次に、最適化すべき変分量子回路$U(\theta)$を作っていく。これは以下の3手順で行う。
#
# 1. 横磁場イジングハミルトニアン作成
# 2. 回転ゲート作成
# 3. 1.と2.のゲートを交互に組み合わせ、1つの大きな変分量子回路$U(\theta)$を作る
# ##### 1. 横磁場イジングハミルトニアン作成
# 4-2節で学んだ横磁場イジングモデルによる時間発展を行い量子回路の複雑性(エンタングルメント)を増すことで、モデルの表現力を高める。(本パートは、詳細を知りたい読者以外は読み飛ばしていただいて構わない。)
# 横磁場イジングモデルのハミルトニアンは以下の通りで、$U_{rand} = e^{-iHt}$という時間発展演算子を定義する。
#
# $$
# H = \sum_{j=1}^N a_j X_j + \sum_{j=1}^N \sum_{k=1}^{j-1} J_{jk} Z_j Z_k
# $$
#
# ここで係数$a$, $J$は$[-1, 1]$の一様分布である。
## 基本ゲート
from qulacs.gate import X, Z
I_mat = np.eye(2, dtype=complex)
X_mat = X(0).get_matrix()
Z_mat = Z(0).get_matrix()
## fullsizeのgateをつくる関数.
def make_fullgate(list_SiteAndOperator, nqubit):
'''
list_SiteAndOperator = [ [i_0, O_0], [i_1, O_1], ...] を受け取り,
関係ないqubitにIdentityを挿入して
I(0) * ... * O_0(i_0) * ... * O_1(i_1) ...
という(2**nqubit, 2**nqubit)行列をつくる.
'''
list_Site = [SiteAndOperator[0] for SiteAndOperator in list_SiteAndOperator]
list_SingleGates = [] ## 1-qubit gateを並べてnp.kronでreduceする
cnt = 0
for i in range(nqubit):
if (i in list_Site):
list_SingleGates.append( list_SiteAndOperator[cnt][1] )
cnt += 1
else: ## 何もないsiteはidentity
list_SingleGates.append(I_mat)
return reduce(np.kron, list_SingleGates)
# +
#### ランダム磁場・ランダム結合イジングハミルトニアンをつくって時間発展演算子をつくる
ham = np.zeros((2**nqubit,2**nqubit), dtype = complex)
for i in range(nqubit): ## i runs 0 to nqubit-1
Jx = -1. + 2.*np.random.rand() ## -1~1の乱数
ham += Jx * make_fullgate( [ [i, X_mat] ], nqubit)
for j in range(i+1, nqubit):
J_ij = -1. + 2.*np.random.rand()
ham += J_ij * make_fullgate ([ [i, Z_mat], [j, Z_mat]], nqubit)
## 対角化して時間発展演算子をつくる. H*P = P*D <-> H = P*D*P^dagger
diag, eigen_vecs = np.linalg.eigh(ham)
time_evol_op = np.dot(np.dot(eigen_vecs, np.diag(np.exp(-1j*time_step*diag))), eigen_vecs.T.conj()) # e^-iHT
# -
time_evol_op.shape
# qulacsのゲートに変換しておく
from qulacs.gate import DenseMatrix
time_evol_gate = DenseMatrix([i for i in range(nqubit)], time_evol_op)
# ##### 2. 回転ゲート作成、3. $U(θ)$の構成
#
# 先ほど構成したランダム横磁場イジングモデルによる時間発展$U_{rand}$と、
# $j \:(=1,2,\cdots n)$番目の量子ビットに回転ゲート
#
# $$
# U_{rot}(\theta_j^{(i)}) = R_j^X(\theta_{j1}^{(i)})R_j^Z(\theta_{j2}^{(i)})R_j^X(\theta_{j3}^{(i)})
# $$
#
# をかけたものを組み合わせて変分量子回路$U(\theta)$を構成する。 ここで$i$は量子回路の層を表す添字で、$U_{rand}$と上記の回転を合計$d$層繰り返す。
# つまり全体では、
#
# $$
# U \left( \{ \theta_j^{(i)} \}_{i,j} \right) = \prod_{i=1}^d \left( \left( \prod_{j=1}^n U_{rot}(\theta_j^{(i)})\right) \cdot U_{rand} \right)
# $$
#
# という変分量子回路を用いる。全部で $3nd$ 個のパラメータがあることになる。各$\theta$の初期値は$[0, 2\pi]$の一様分布にとっておく。
from qulacs import ParametricQuantumCircuit
# output用ゲートU_outの組み立て&パラメータ初期値の設定
U_out = ParametricQuantumCircuit(nqubit)
for d in range(c_depth):
U_out.add_gate(time_evol_gate)
for i in range(nqubit):
angle = 2.0 * np.pi * np.random.rand()
U_out.add_parametric_RX_gate(i,angle)
angle = 2.0 * np.pi * np.random.rand()
U_out.add_parametric_RZ_gate(i,angle)
angle = 2.0 * np.pi * np.random.rand()
U_out.add_parametric_RX_gate(i,angle)
# パラメータthetaの初期値のリストを取得しておく
parameter_count = U_out.get_parameter_count()
theta_init = [U_out.get_parameter(ind) for ind in range(parameter_count)]
theta_init
# 後の便利のため、$U(\theta)$のパラメータ$\theta$を更新する関数を作成しておく。
# パラメータthetaを更新する関数
def set_U_out(theta):
global U_out
parameter_count = U_out.get_parameter_count()
for i in range(parameter_count):
U_out.set_parameter(i, theta[i])
# #### 測定
# 今回は、0番目の量子ビットのpauli Zの、出力状態$|\psi_{\rm out}\rangle$での期待値をモデルの出力とする。
# すなわち、$y(\theta, x_i) = \langle Z_0 \rangle = \langle \psi_{\rm out}|Z_0|\psi_{\rm out}\rangle$である。
# オブザーバブルZ_0を作成
from qulacs import Observable
obs = Observable(nqubit)
obs.add_operator(2.,'Z 0') # オブザーバブル2 * Zを設定。ここで2を掛けているのは、最終的な<Z>の値域を広げるためである。未知の関数に対応するためには、この定数もパラメータの一つとして最適化する必要がある。
obs.get_expectation_value(state)
# #### 一連の流れを関数にまとめる
# ここまでの流れをまとめて、入力$x_i$からモデルの予測値$y(x_i, \theta)$を返す関数を定義する。
# 入力x_iからモデルの予測値y(x_i, theta)を返す関数
def qcl_pred(x, U_out):
state = QuantumState(nqubit)
state.set_zero_state()
# 入力状態計算
U_in(x).update_quantum_state(state)
# 出力状態計算
U_out.update_quantum_state(state)
# モデルの出力
res = obs.get_expectation_value(state)
return res
# #### コスト関数計算
# コスト関数 $L(\theta)$は、教師データと予測データの平均二乗誤差(MSE)とする。
# cost function Lを計算
def cost_func(theta):
'''
theta: 長さc_depth * nqubit * 3のndarray
'''
# U_outのパラメータthetaを更新
# global U_out
set_U_out(theta)
# num_x_train個のデータについて計算
y_pred = [qcl_pred(x, U_out) for x in x_train]
# quadratic loss
L = ((y_pred - y_train)**2).mean()
return L
# パラメータthetaの初期値におけるコスト関数の値
cost_func(theta_init)
# パラメータthetaの初期値のもとでのグラフ
xlist = np.arange(x_min, x_max, 0.02)
y_init = [qcl_pred(x, U_out) for x in xlist]
plt.plot(xlist, y_init)
# #### 学習(scipy.optimize.minimizeで最適化)
# ようやく準備が終わり、いよいよ学習を行う。ここでは簡単のため、勾配の計算式を与える必要のないNelder-Mead法を用いて最適化する。勾配を用いる最適化手法(例:BFGS法)を用いる場合は、勾配の便利な計算式が参考文献[[1]](https://arxiv.org/abs/1803.00745)で紹介されているので参照されたい。
from scipy.optimize import minimize
# %%time
# 学習 (筆者のPCで1~2分程度かかる)
result = minimize(cost_func, theta_init, method='Nelder-Mead')
# 最適化後のcost_functionの値
result.fun
# 最適化によるthetaの解
theta_opt = result.x
print(theta_opt)
# #### 結果のプロット
# U_outに最適化されたthetaを代入
set_U_out(theta_opt)
# +
# プロット
plt.figure(figsize=(10, 6))
xlist = np.arange(x_min, x_max, 0.02)
# 教師データ
plt.plot(x_train, y_train, "o", label='Teacher')
# パラメータθの初期値のもとでのグラフ
plt.plot(xlist, y_init, '--', label='Initial Model Prediction', c='gray')
# モデルの予測値
y_pred = np.array([qcl_pred(x, U_out) for x in xlist])
plt.plot(xlist, y_pred, label='Final Model Prediction')
plt.legend()
plt.show()
# -
# 確かにsin関数の近似に成功している事がわかる。
# ここでは入力・出力ともに1次元の関数の近似という極めてシンプルなタスクを扱ったが、より多次元の入出力をもつ関数の近似や分類問題にも拡張が可能である。
# 意欲のある読者は、コラム ``5.2c.Application of QCL to Machine Learning`` において、代表的な機械学習のデータセットの一つである[Irisデータセット](https://scikit-learn.org/stable/auto_examples/datasets/plot_iris_dataset.html)の分類に挑戦されたい。
# ### 参考文献
# [1] <NAME>, <NAME>, M. Kitagawa, and <NAME>, “Quantum circuit learning”, [Phys. Rev. A 98, 032309 (2018)](https://journals.aps.org/pra/abstract/10.1103/PhysRevA.98.032309), arXiv版:https://arxiv.org/abs/1803.00745
# [2] <NAME> _et al._, “Supervised learning with quantum-enhanced feature spaces”, [Nature 567, 209–212 (2019)](https://www.nature.com/articles/s41586-019-0980-2), arXiv版:https://arxiv.org/abs/1804.11326
| notebooks/5.2_Quantum_Circuit_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Normalization
# The `normalize` is a helper function to z-score your data. This is useful if your features (columns) are scaled differently within or across datasets. By default, hypertools normalizes *across* the columns of all datasets passed, but also affords the option to normalize columns *within* individual lists. Alternatively, you can also normalize each row. The function returns an array or list of arrays where the columns or rows are z-scored (output type same as input type).
# ## Import packages
# +
import hypertools as hyp
import numpy as np
# %matplotlib inline
# -
# ## Generate synthetic data
# First, we generate two sets of synthetic data. We pull points randomly from a multivariate normal distribution for each set, so the sets will exhibit unique statistical properties.
# +
x1 = np.random.randn(10,10)
x2 = np.random.randn(10,10)
c1 = np.dot(x1, x1.T)
c2 = np.dot(x2, x2.T)
m1 = np.zeros([1,10])
m2 = 10 + m1
data1 = np.random.multivariate_normal(m1[0], c1, 100)
data2 = np.random.multivariate_normal(m2[0], c2, 100)
data = [data1, data2]
# -
# ## Visualize the data
geo = hyp.plot(data, '.')
# ## Normalizing (Specified Cols or Rows)
# Or, to specify a different normalization, pass one of the following arguments as a string, as shown in the examples below.
#
# + 'across' - columns z-scored across passed lists (default)
# + 'within' - columns z-scored within passed lists
# + 'row' - rows z-scored
# ### Normalizing 'across'
# When you normalize 'across', all of the data is stacked/combined, and the normalization is done on the columns of the full dataset. Then the data is split back into separate elements.
norm = hyp.normalize(data, normalize = 'across')
geo = hyp.plot(norm, '.')
# ### Normalizing 'within'
# When you normalize 'within', normalization is done on the columns of each element of the data, separately.
norm = hyp.normalize(data, normalize = 'within')
geo = hyp.plot(norm, '.')
# ### Normalizing by 'row'
norm = hyp.normalize(data, normalize = 'row')
geo = hyp.plot(norm, '.')
| docs/tutorials/normalize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Apple Stock
#
# Check out [Apple Stock Exercises Video Tutorial](https://youtu.be/wpXkR_IZcug) to watch a data scientist go through the exercises
# ### Introduction:
#
# We are going to use Apple's stock price.
#
#
# ### Step 1. Import the necessary libraries
# +
import pandas as pd
import numpy as np
# visualization
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# ### Step 2. Import the dataset from this [address](https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/09_Time_Series/Apple_Stock/appl_1980_2014.csv)
# ### Step 3. Assign it to a variable apple
# +
url = 'https://raw.githubusercontent.com/guipsamora/pandas_exercises/master/09_Time_Series/Apple_Stock/appl_1980_2014.csv'
apple = pd.read_csv(url)
apple.head()
# -
# ### Step 4. Check out the type of the columns
apple.dtypes
# ### Step 5. Transform the Date column as a datetime type
# +
apple.Date = pd.to_datetime(apple.Date)
apple['Date'].head()
# -
# ### Step 6. Set the date as the index
# +
apple = apple.set_index('Date')
apple.head()
# -
# ### Step 7. Is there any duplicate dates?
# NO! All are unique
apple.index.is_unique
# ### Step 8. Ops...it seems the index is from the most recent date. Make the first entry the oldest date.
apple.sort_index(ascending = True).head()
# ### Step 9. Get the last business day of each month
# +
apple_month = apple.resample('BM').mean()
apple_month.head()
# -
# ### Step 10. What is the difference in days between the first day and the oldest
(apple.index.max() - apple.index.min()).days
# ### Step 11. How many months in the data we have?
# +
apple_months = apple.resample('BM').mean()
len(apple_months.index)
# -
# ### Step 12. Plot the 'Adj Close' value. Set the size of the figure to 13.5 x 9 inches
# +
# makes the plot and assign it to a variable
appl_open = apple['Adj Close'].plot(title = "Apple Stock")
# changes the size of the graph
fig = appl_open.get_figure()
fig.set_size_inches(13.5, 9)
# -
# ### BONUS: Create your own question and answer it.
| 09_Time_Series/Apple_Stock/Exercises-with-solutions-code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # ML Model Test: Ranking
#
# This is a follow up of the TensorFlow Recommenders tutorials. On this notebook, we will be focusing on the "Ranking" stage of a Recommender System. All the information is in the following [link](https://www.tensorflow.org/recommenders/examples/basic_ranking).<br>
# We strongly recommend creating a **virtual environment** before running the following code. Let's start by getting our dependencies.
# !pip install tensorflow-recommenders
# !pip install -q --upgrade tensorflow-datasets
# + [markdown] tags=[]
# ## Imports
#
# Next, let's invoke the necessary packages.
# +
import os
import pprint
from typing import Dict, Text
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
import tensorflow_recommenders as tfrs
# -
# ## Dataset
#
# This is included in the TensorFlow library. We intend to use the MovieLens ratings and movies dataset. All the data will be considered for the `train` split. This time we also consider the `rating`.
# + tags=[]
ratings = tfds.load("ccp2_capstone_ratings", split="train")
ratings = ratings.map(lambda x: {
"movie_title": x["media_title"],
"user_id": x["user_id"],
"user_rating": x["user_rating"]
})
# -
# Let's take a look at the data structure:
for x in ratings.take(1).as_numpy_iterator():
print("Rating: ")
pprint.pprint(x)
# Let's now split the set into `train` and `test` sets. This is for having ways of validation after training the model.
# +
tf.random.set_seed(42)
shuffled = ratings.shuffle(100_000, seed=42, reshuffle_each_iteration=False)
train = shuffled.take(80_000)
test = shuffled.skip(80_000).take(20_000)
# -
# Next, we will identify unique `user_id`s and `movie_title`s. This is for having the vocabulary necessary for embedding vectors mapping.
# +
movie_titles = ratings.batch(1_000_000).map(lambda x: x["movie_title"])
user_ids = ratings.batch(1_000_000).map(lambda x: x["user_id"])
unique_movie_titles = np.unique(np.concatenate(list(movie_titles)))
unique_user_ids = np.unique(np.concatenate(list(user_ids)))
# + [markdown] tags=[]
# ## Implementing
#
# Time to build the Ranking model.
#
# ### Architecture
#
# We don't have so many efficiency constraints here as in "Retrieval", so we can allow a Neural Network with some more Layers.
# -
class RankingModel(tf.keras.Model):
def __init__(self):
super().__init__()
embedding_dimension = 32
# Compute embeddings for users.
self.user_embeddings = tf.keras.Sequential([
tf.keras.layers.StringLookup(
vocabulary=unique_user_ids, mask_token=None),
tf.keras.layers.Embedding(len(unique_user_ids) + 1, embedding_dimension)
])
# Compute embeddings for movies.
self.movie_embeddings = tf.keras.Sequential([
tf.keras.layers.StringLookup(
vocabulary=unique_movie_titles, mask_token=None),
tf.keras.layers.Embedding(len(unique_movie_titles) + 1, embedding_dimension)
])
# Compute predictions.
self.ratings = tf.keras.Sequential([
# Learn multiple dense layers.
tf.keras.layers.Dense(256, activation="relu"),
tf.keras.layers.Dense(64, activation="relu"),
# Make rating predictions in the final layer.
tf.keras.layers.Dense(1)
])
def call(self, inputs):
user_id, movie_title = inputs
user_embedding = self.user_embeddings(user_id)
movie_embedding = self.movie_embeddings(movie_title)
return self.ratings(tf.concat([user_embedding, movie_embedding], axis=1))
# + [markdown] tags=[]
# ### Metrics and Loss
#
# This is the method we will use to measure the "accuracy" of our model. We will use the `Ranking` task object for wrapping together loss function and metric computation, together with `MeanSquaredError` Keras loss.
# -
task = tfrs.tasks.Ranking(
loss = tf.keras.losses.MeanSquaredError(),
metrics=[tf.keras.metrics.RootMeanSquaredError()]
)
# ### Full model
#
# Here we put all the pieces together for creating our model. There is a high level of abstraction in the following code for selecting the appropriate training loop that matches our model.
class MovielensModel(tfrs.models.Model):
def __init__(self):
super().__init__()
self.ranking_model: tf.keras.Model = RankingModel()
self.task: tf.keras.layers.Layer = tfrs.tasks.Ranking(
loss = tf.keras.losses.MeanSquaredError(),
metrics=[tf.keras.metrics.RootMeanSquaredError()]
)
def call(self, features: Dict[str, tf.Tensor]) -> tf.Tensor:
return self.ranking_model(
(features["user_id"], features["movie_title"]))
def compute_loss(self, features: Dict[Text, tf.Tensor], training=False) -> tf.Tensor:
labels = features.pop("user_rating")
rating_predictions = self(features)
# The task computes the loss and the metrics.
return self.task(labels=labels, predictions=rating_predictions)
# ## Fitting and Evaluating
#
# This makes use of `Keras` functionalities. Let's start by instantiating the model.
model = MovielensModel()
model.compile(optimizer=tf.keras.optimizers.Adagrad(learning_rate=0.1))
# Shuffle, batch, and cache the training and evaluation data.
cached_train = train.shuffle(100_000).batch(8192).cache()
cached_test = test.batch(4096).cache()
# Finally, train the model
model.fit(cached_train, epochs=3)
# We can now evaluate our model.
model.evaluate(cached_test, return_dict=True)
# The RMSE metric is low, meaning our model has a good accuracy.
#
# ## Predictions
#
# We can now make predictions for a set of movies and rank them based on those predictions.
# +
test_ratings = {}
test_movie_titles = ["<NAME>", "<NAME>", "Star Wars"]
for movie_title in test_movie_titles:
test_ratings[movie_title] = model({
"user_id": np.array(["150"]),
"movie_title": np.array([movie_title])
})
print("Ratings:")
for title, score in sorted(test_ratings.items(), key=lambda x: x[1], reverse=True):
print(f"{title}: {score}")
# -
# ## Model Serving
#
# We will pack our ranking model into a single exportable as a `SavedModel` so we can deploy it with `TensorFlow Serving`.
# +
# Export the query model.
path = os.path.join(os.getcwd(), "../models/ranking/00000123/")
# Save the index.
tf.saved_model.save(model, path)
# -
# The next step is to deploy our model in Docker.
| src/Ranking_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv("Simple_Linear_Reg.csv")
df
#help(df.iloc)
x = df.iloc[:,:-1]
x
y = df.iloc[:,-1:]
y
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
lin_model = linear_model.LinearRegression()
a = lin_model.fit(x,y)
a
a.score(x,y) # R square value
a.intercept_ # C value in mx + c
a.coef_ # m - slope value for id and price
y_pred = a.predict(x)
y_pred
| linear-regression-lab/Linear-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="515b43fc" outputId="5d3df113-3a59-42e5-9434-84cfbf629a0b"
# !python -m pip install seaborn
# + id="c8f9fd11"
# %load_ext autoreload
# %autoreload 2
# + id="29859942"
import seaborn as sns
# + id="389e8fb5"
df = sns.load_dataset('titanic')
# + colab={"base_uri": "https://localhost:8080/", "height": 439} id="dfbfbf1d" outputId="76bfd641-5c92-435d-ab8d-109b519f09ad"
df
# + colab={"base_uri": "https://localhost:8080/"} id="f7458ee6" outputId="20a34faf-9e7d-473e-b43f-b035701c87ba"
df.info()
# + [markdown] id="3ccd1545"
# survived,pclass,age,sibsp,parch,fare
# + id="b171bf68"
x = df[['pclass','sibsp','parch','fare']]
y = df['survived']
# + id="f8ab69eb"
from sklearn.model_selection import train_test_split
# + id="520feee2"
X_train, X_test, Y_train, Y_test = train_test_split(x,y)
# + colab={"base_uri": "https://localhost:8080/"} id="ea304de5" outputId="8f01f53e-6125-4e7f-a418-4a493a864e82"
X_train.shape ,X_test.shape ,Y_train.shape ,Y_test.shape
# + id="136d198a"
from sklearn.linear_model import LogisticRegression
# + id="8e15b7c9"
logi_r = LogisticRegression()
# + colab={"base_uri": "https://localhost:8080/"} id="033bd9a8" outputId="d310f6f7-17fb-4469-9c28-7c43c647d832"
logi_r.fit(X_train,Y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="a5479717" outputId="43c1abb1-0065-4b53-eba5-cdcdf516a9bd"
logi_r.classes_,logi_r.coef_,logi_r.intercept_
# + colab={"base_uri": "https://localhost:8080/"} id="78d89fd0" outputId="34fe8ceb-8e8d-468d-efa8-0b0482e340c8"
logi_r.score(x,y)
# + colab={"base_uri": "https://localhost:8080/"} id="823971ef" outputId="9d95d413-23fe-4ed3-8a84-c1a77fb2a203"
logi_r.predict(X_train)
# + [markdown] id="9c2000b0"
# logi_r.predict_proba(X_train)
# + colab={"base_uri": "https://localhost:8080/"} id="c59f505d" outputId="c59173aa-7fc5-4386-cd59-bf8fa8bdc7b9"
logi_r.predict_proba(X_train)
# + colab={"base_uri": "https://localhost:8080/"} id="E1eUfdjHrnyd" outputId="9fc40dd8-3171-4f5c-eef3-3a6ffa53392f"
print('Hello')
# + id="f25c00df"
from sklearn import metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="bfc3a736" outputId="6fe61409-965a-4860-a174-bcba9550080e"
metrics.confusion_matrics(X_train,Y_train)
# + id="11db4e02"
| titanic_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ricardoV94/ThinkBayesPymc3/blob/master/ThinkBayes_Chapter_10.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="_QFayNAHRAga" colab_type="code" colab={}
# %%capture
pip install arviz
# + id="xVPW-yWN8sm1" colab_type="code" colab={}
import numpy as np
import pymc3 as pm
import theano
import theano.tensor as tt
import arviz as az
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.stats as st
# + id="6blJjHa-te2n" colab_type="code" colab={}
def custom_posterior_plots(trace):
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(12,4))
sns.kdeplot(trace['mu_m'], trace['sigma_m'], n_levels=5, cmap='Blues', ax=ax[0])
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
sns.kdeplot(trace['mu_f'], trace['sigma_f'], n_levels=5, cmap='Blues', ax=ax[1])
plt.gca().ticklabel_format(axis='both', style='plain', useOffset=False)
for axi in ax.ravel():
axi.set_ylabel('Stddev (cm)')
axi.set_xlabel('Mean height (cm)')
ax[0].set_title('Posterior joint distribution - males')
ax[1].set_title('Posterior joint distribution - females')
plt.tight_layout()
plt.figure()
sns.kdeplot(trace['cv_m'], cumulative=True, label='male', lw=3 )
sns.kdeplot(trace['cv_f'], cumulative=True, color='lightblue', label='female', lw=3)
plt.xlabel('Coefficient of variation')
plt.ylabel('Probability')
# + [markdown] id="Q_t4c5pxiCCP" colab_type="text"
# ## 10.1 The Variability Hypothesis
# + id="bCMYGZWAbCL4" colab_type="code" colab={}
# Load data
import pickle
from urllib.request import urlopen
d = pickle.load(urlopen("https://raw.githubusercontent.com/ricardoV94/ThinkBayesPymc3/master/data/variability_data.pkl"))
# + id="vpQBSqixcHo1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1e841cef-6179-4674-b1e3-ed52bbd8dbdb"
np.mean(d[1]), np.mean(d[2])
# + id="UHbetFwzcNLa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fbdaeafc-06ea-48b5-f412-aaefa56fe65e"
np.std(d[1]), np.std(d[2])
# + id="Zeqqgrc5cPem" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f8018824-9747-4bb8-f794-124ef6628c2e"
np.std(d[1]) / np.mean(d[1]), np.std(d[2]) / np.mean(d[2])
# + [markdown] id="4hAZEdDNcFF0" colab_type="text"
# ### 10.4 The posterior distribution of CV
# + id="DD5-WBxNdpG1" colab_type="code" colab={}
def find_prior_range_mean(xs, num_stderrs=4):
n = len(xs)
m = np.mean(xs)
s = np.std(xs)
stderr_m = s / np.math.sqrt(n)
spread_m = stderr_m * num_stderrs
mu_range = [m - spread_m, m + spread_m]
stderr_s = s / np.math.sqrt(2 * (n-1))
spread_s = stderr_s * num_stderrs
sigma_range = [s - spread_s, s + spread_s]
return mu_range, sigma_range
# + id="9CvZ4ottQWvM" colab_type="code" outputId="50d19fe2-ba2b-4279-9089-7f23d2869910" colab={"base_uri": "https://localhost:8080/", "height": 121}
mu_range_m, sigma_range_m = find_prior_range_mean(d[1])
mu_range_f, sigma_range_f = find_prior_range_mean(d[2])
with pm.Model() as m_10_4:
mu_m = pm.Uniform('mu_m', lower=mu_range_m[0], upper=mu_range_m[1])
mu_f = pm.Uniform('mu_f', lower=mu_range_f[0], upper=mu_range_f[1])
sigma_m = pm.Uniform('sigma_m', lower=sigma_range_m[0], upper=sigma_range_m[1])
sigma_f = pm.Uniform('sigma_f', lower=sigma_range_f[0], upper=sigma_range_f[1])
like_m = pm.Normal('like_m', mu=mu_m, sigma=sigma_m, observed=d[1])
like_f = pm.Normal('like_f', mu=mu_f, sigma=sigma_f, observed=d[2])
cv_m = pm.Deterministic('cv_m', sigma_m / mu_m)
cv_f = pm.Deterministic('cv_f', sigma_f / mu_f)
trace_m_10_4 = pm.sample(1000)
# + id="PU7OnDDwf6wc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 559} outputId="eab6a482-788d-4700-b44c-73888bef6687"
custom_posterior_plots(trace_m_10_4)
# + [markdown] id="X8ziDlN4mtRI" colab_type="text"
# ### 10.8 ABC (Approximate Bayesian Computation)
#
# Faster inference based on likelihood of sample statistics and not of sample individual values.
#
# 40x faster: from 25 it/s to 1000 it/s
# + id="BI0GHT4oVHAz" colab_type="code" outputId="6f09fedb-0348-400f-b639-dcf460b431cc" colab={"base_uri": "https://localhost:8080/", "height": 121}
mu_range_m, sigma_range_m = find_prior_range_mean(d[1])
mu_range_f, sigma_range_f = find_prior_range_mean(d[2])
n_m = len(d[1])
n_f = len(d[2])
with pm.Model() as m_10_8:
mu_m = pm.Uniform('mu_m', lower=mu_range_m[0], upper=mu_range_m[1])
mu_f = pm.Uniform('mu_f', lower=mu_range_f[0], upper=mu_range_f[1])
sigma_m = pm.Uniform('sigma_m', lower=sigma_range_m[0], upper=sigma_range_m[1])
sigma_f = pm.Uniform('sigma_f', lower=sigma_range_f[0], upper=sigma_range_f[1])
stderr_mu_m = sigma_m / pm.math.sqrt(n_m)
stderr_mu_f = sigma_f / pm.math.sqrt(n_f)
like_mu_m = pm.Normal('like_mu_m', mu=mu_m, sigma=stderr_mu_m, observed=np.mean(d[1]))
like_mu_f = pm.Normal('like_mu_f', mu=mu_f, sigma=stderr_mu_f, observed=np.mean(d[2]))
stderr_sigma_m = sigma_m / pm.math.sqrt(2 * (n_m - 1))
stderr_sigma_f = sigma_f / pm.math.sqrt(2 * (n_f - 1))
like_sigma_m = pm.Normal('like_sigma_m', mu=sigma_m, sigma=stderr_sigma_m, observed=np.std(d[1]))
like_sigma_f = pm.Normal('like_sigma_f', mu=sigma_f, sigma=stderr_sigma_f, observed=np.std(d[2]))
cv_m = pm.Deterministic('cv_m', sigma_m / mu_m)
cv_f = pm.Deterministic('cv_f', sigma_f / mu_f)
trace_m_10_8 = pm.sample(5000)
# + id="qQO6YfT-oBul" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 559} outputId="e2bc9a68-43f2-4087-9a68-e402c08eb623"
custom_posterior_plots(trace_m_10_8)
# + [markdown] id="PV1D19aXrZYz" colab_type="text"
# ### 10.9 Robust estimation
#
# Note: Results and conclusions are different from book. Females are more similar with num_sigmas = 1, and less with num_sigmas = 2. Possible bug?
# + id="MbQ9lh-bo-aB" colab_type="code" colab={}
def median_ipr(xs, p):
median = np.median(xs)
alpha = (100-p) / 2
percentile = np.percentile(xs, q=[alpha, 100-alpha])
ipr = percentile[1] - percentile[0]
return median, ipr
def median_sigma(xs, num_sigmas):
half_p = st.norm().cdf(num_sigmas) - 0.5
median, ipr = median_ipr(xs, half_p * 2 * 100)
sigma = ipr / 2 / num_sigmas
return median, sigma
# + id="skt-PNu3viyU" colab_type="code" colab={}
def find_prior_range_median(xs, num_sigmas, num_stderrs=4):
n = len(xs)
m, s = median_sigma(xs, num_sigmas)
stderr_m = s / np.math.sqrt(n)
spread_m = stderr_m * num_stderrs
mu_range = [m - spread_m, m + spread_m]
stderr_s = s / np.math.sqrt(2 * (n-1))
spread_s = stderr_s * num_stderrs
sigma_range = [s - spread_s, s + spread_s]
return mu_range, sigma_range
# + id="p7C_J-5yms4B" colab_type="code" colab={}
def create_model_10_9(num_sigmas):
with pm.Model() as m_10_9:
# Extract sample estimates
mu_range_m, sigma_range_m = find_prior_range_median(d[1], num_sigmas)
mu_range_f, sigma_range_f = find_prior_range_median(d[2], num_sigmas)
sample_median_m, sample_sigma_m = median_sigma(d[1], num_sigmas)
sample_median_f, sample_sigma_f = median_sigma(d[2], num_sigmas)
n_m = len(d[1])
n_f = len(d[2])
# Model
mu_m = pm.Uniform('mu_m', lower=mu_range_m[0], upper=mu_range_m[1])
mu_f = pm.Uniform('mu_f', lower=mu_range_f[0], upper=mu_range_f[1])
sigma_m = pm.Uniform('sigma_m', lower=sigma_range_m[0], upper=sigma_range_m[1])
sigma_f = pm.Uniform('sigma_f', lower=sigma_range_f[0], upper=sigma_range_f[1])
stderr_mu_m = sigma_m / pm.math.sqrt(n_m)
stderr_mu_f = sigma_f / pm.math.sqrt(n_f)
like_mu_m = pm.Normal('like_mu_m', mu=mu_m, sigma=stderr_mu_m, observed=sample_median_m)
like_mu_f = pm.Normal('like_mu_f', mu=mu_f, sigma=stderr_mu_f, observed=sample_median_f)
stderr_sigma_m = sigma_m / pm.math.sqrt(2 * (n_m - 1))
stderr_sigma_f = sigma_f / pm.math.sqrt(2 * (n_f - 1))
like_sigma_m = pm.Normal('like_sigma_m', mu=sigma_m, sigma=stderr_sigma_m, observed=sample_sigma_m)
like_sigma_f = pm.Normal('like_sigma_f', mu=sigma_f, sigma=stderr_sigma_f, observed=sample_sigma_f)
cv_m = pm.Deterministic('cv_m', sigma_m / mu_m)
cv_f = pm.Deterministic('cv_f', sigma_f / mu_f)
return m_10_9
# + id="Ivrg1RHn2Iap" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 156} outputId="c8cae120-cd51-40b7-cefb-577a9c9357b3"
traces_m_10_9 = []
for num_sigmas in (1, 2):
with create_model_10_9(num_sigmas):
traces_m_10_9.append(pm.sample(5000, progressbar=False))
# + id="B1jp9PjVs7sV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 559} outputId="aec7acef-19da-4289-8141-2e7ff393f42a"
custom_posterior_plots(traces_m_10_9[0])
# + id="aq8f7oVt0rpv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 559} outputId="1fecfb71-21a2-46de-b76a-808f49b39f8a"
custom_posterior_plots(traces_m_10_9[1])
# + id="Zjg7k68c556v" colab_type="code" colab={}
| ThinkBayes_Chapter_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # PyUnitWizard in your library
# *-Instructions to work with PyUnitWizard inside your own library-*
#
# Here you can find the instructions to include PyUnitWizard in your projects. Following this indications, you don't need to worry about the requirements of the libraries supported by PyUnitWizard.
#
# To illustrate how to include PyUnitWizard let's see a very simple library you can find in [github repository (examples directory)](https://github.com/uibcdf/PyUnitWizard/tree/main/examples): `testlib`. This is its structure:
# ```bash
# tree --dirsfirst --charset=markdown testlib
# ```
# ```
# testlib
# |-- box
# | |-- __init__.py
# | |-- methods_a.py
# | |-- methods_b.py
# | `-- methods_c.py
# |-- _pyunitwizard
# | `-- __init__.py
# |-- __init__.py
# ```
# Make a directory named `_pyunitwizard` in your project top directory. And include in `_pyunitwizard` a `__init__.py` file such as:
# ```python
# ### testlib/_pyunitwizard/__init__.py ###
# import pyunitwizard as puw
#
# # In this case Pint and openmm.unit are loaded:
# puw.load_libraries(['pint', 'openmm.unit'])
#
# # And openmm.unit is defined as default form
# puw.set_default_form('openmm.unit')
# ```
# Now, let's define some methods using your `_pyunitwizard` module. The first ones in the file `main.py`:
# ```python
# ### testlib/main.py ###
# from ._pyunitwizard import puw
#
# def sum_quantities(a, b, form=None):
#
# aa = puw.string_to_quantity(a, to_form=form)
# bb = puw.string_to_quantity(b, to_form=form)
# output = aa+bb
#
# return output
#
# def get_form(quantity):
#
# return puw.get_form(quantity)
#
# def libraries_loaded():
#
# return puw.libraries_loaded()
# ```
# And in a directory named `box` let's include two methods to test your `_pyunitwizard` module:
# ```python
# ### testlib/box/methods_a.py
# from .._pyunitwizard import puw
#
# def get_default_form():
#
# return puw.get_default_form()
# ```
#
# ```python
# ### testlib/box/methods_b.py
# from .._pyunitwizard import puw
#
# def set_default_form(form):
#
# return puw.set_default_form(form)
# ```
#
# Finnally, let's writte the `__init__.py` files in the top directory and in `box`:
# ```python
# # testlib/box/__init__.py
# from .methods_a import get_default_form
# from .methods_b import set_default_form
# ```
#
# ```python
# # testlib/__init__.py
# from .main import sum_quantities, get_form, libraries_loaded
# from . import box
# ```
# This way we already have a simple library using PyUnitWizard. You can check how `testlib` works:
# ```ipython
# In [1]: import testlib
#
# In [2]: testlib.libraries_loaded()
# Out[2]: ['pint', 'openmm.unit']
#
# In [3]: q = testlib.sum_quantities('2cm','3cm')
#
# In [4]: testlib.get_form(q)
# Out[4]: 'openmm.unit'
#
# In [5]: testlib.box.get_default_form()
# Out[5]: 'openmm.unit'
#
# In [6]: testlib.box.set_default_form('pint')
#
# In [7]: q = testlib.sum_quantities('2cm','3cm')
#
# In [8]: testlib.get_form(q)
# Out[8]: 'pint'
# ```
# <div class="alert alert-block alert-info">
# <b>Tip:</b> Together with testlib, in the github repository, you can find testlib2
# where pyunitwizard is included using only absolut import paths -as suggested by <a href="https://www.python.org/dev/peps/pep-0008/#imports">PEP8</a>-</div>
| docs/contents/user/In_Your_Library.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Select and distribute additional Auditory Percetion articles for phrase annotation
# ## Goal is deeper annotation.
# - A good number of annotations on this topic.
# - Phrase based annotation so we can be most flexible and assess at higher levels.
#
# ## Secondary goal, more cross-validation.
# - 1/2 of new articles will be annotated by all.
# - 1/2 of new articles will be annotated by only 1 annotator.
dest_dir = '/Users/ccarey/Documents/Projects/NAMI/rdoc/pdfs/20160205_rdoc_project'
# %mkdir {dest_dir}
# %cd {dest_dir}
# ### Get a list of all our previous pubmed ids used for annotation (irrespective of Auditory perception)
# +
from __future__ import print_function
import glob
import os
import re
# annotated and processed previously
prev_annotated_pdfs_dir = '/Users/ccarey/Documents/Projects/NAMI/rdoc/pdfs/all_pdfs_annotated_pmid_names/*.pdf'
pdfs = glob.glob(prev_annotated_pdfs_dir)
pdfs = [os.path.basename(pdf) for pdf in pdfs]
# and those from round 4 annotations
prev_annotated_pdfs_dir2 = '/Users/ccarey/Documents/Projects/NAMI/rdoc/pdfs/20160122_rdoc_project/*.pdf'
pdfs2 = glob.glob(prev_annotated_pdfs_dir2)
pdfs.extend([os.path.basename(pdf) for pdf in pdfs2])
print('{} total PDFS'.format(len(pdfs)))
# -
pattern = '([0-9]{6,9})'
p = re.compile(pattern)
pmids = [p.search(pdf).group() for pdf in pdfs if p.search(pdf)]
print('{} Total exisiting PMIDS'.format(len(pmids)))
pmids = set(pmids)
print('{} Unique exisiting PMIDS'.format(len(pmids)))
#print(pmids)
prev_pmids = pmids
# 229 unique existing pmids:
#
# ['22686386', '23744445', '22715197', '23055094', '23929944', '24806675', '22575329', '24930577', '25107317', '22776995', '24870123', '24388670', '21849230', '24933663', '20497902', '25734385', '20532489', '23555220', '24770625', '25834059', '21854953', '23904684', '22378876', '21206465', '23647728', '23127585', '20410871', '25142564', '21203376', '21954087', '22805501', '22379245', '25261920', '22505867', '24512610', '22239924', '21463060', '21699821', '24453310', '23504052', '25160677', '22088577', '21278378', '21426626', '22285891', '22664396', '24333745', '22968207', '25051268', '22438994', '20955866', '24964082', '22233352', '23190433', '21826029', '23934417', '22911829', '25280468', '23074247', '25142762', '22169884', '21613467', '24529045', '25017671', '22832959', '21276977', '24466064', '22447249', '22314045', '25521352', '22294809', '24651580', '24015304', '23990240', '24933724', '24126129', '21957257', '22474609', '20584721', '22048839', '24156344', '25348125', '24259275', '24899756', '21034683', '21050743', '23503620', '25788679', '23709163', '21259270', '21477924', '23299717', '21909371', '22004270', '21397252', '21419826', '25759762', '24252875', '24293773', '24235891', '24231418', '24116095', '22146934', '22131608', '20685988', '22986355', '22414937', '24359877', '23928891', '22496862', '23060019', '25036222', '25424865', '20695690', '25036160', '24804717', '23786695', '25749431', '23326604', '22215928', '20049632', '21531705', '20842164', '20663220', '23770566', '21898707', '23899724', '23622762', '24740391', '25126029', '23088207', '25348131', '25456277', '22286850', '23940642', '25061837', '24705497', '25773639', '21623612', '20525011', '25197810', '25126038', '25154749', '24618591', '24470693', '21118712', '21179552', '24980898', '23957953', '22315106', '24571111', '22043127', '23707592', '24101292', '24285346', '20620104', '24511281', '21077571', '23954763', '22467988', '24376698', '22470524', '23810448', '22035386', '25898427', '22379238', '22379239', '23452958', '24266644', '24426818', '23941878', '21626350', '24260331', '25325584', '21278907', '23646134', '25136085', '21675365', '23731439', '22163262', '24457211', '22998925', '24505292', '23143607', '25774613', '23083918', '21120131', '20939652', '25740534', '24214921', '23643925', '21440905', '25883640', '20634711', '22377810', '25913552', '23997704', '24023823', '21731598', '24412227', '21376761', '24333377', '21550590', '24725811', '21319926', '24045586', '21335015', '25258728', '22215925', '22119526', '24127147', '23193115', '21280045', '25024660', '21677894', '21816115', '20815182', '20661292', '21886801', '20615239', '23558179', '25756280', '20857862', '21392554', '22665872', '25581922', '20450941', '24125792', '22959616']
# ## Get new lists of pmids specific to topic
# +
#from __future__ import print_function
from Bio import Entrez
from subprocess import check_call
from shutil import copy2
#import glob
import time
import imp
import os
url2p = imp.load_source('Url2PubmedPmcPdf', '/Users/ccarey/Documents/Projects/NAMI/rdoc/scripts/Url2PubmedPmcPdf.py')
Entrez.email = "<EMAIL>"
def narrow_id_list(found_ids, omit_ids):
found_but_omit = list(set(found_ids) & set(omit_ids))
found_and_keep = list(set(found_ids) - set(omit_ids))
print('Removed this many ids: {}'.format(len(found_but_omit)))
return(found_and_keep)
def pubmed_central_search_to_pubmed_id(search_string, retmax=20):
# verify how many records match
handle = Entrez.egquery(term=search_string)
record = Entrez.read(handle)
# maybe useful if we are dealing with 100s of ids and don't want to overwhelm server?
for row in record["eGQueryResult"]:
if row["DbName"] == "pubmed":
print(row["Count"])
# fetch the ids for those records
handle = Entrez.esearch(db="pubmed", retmax=retmax, term=search_string)
record = Entrez.read(handle)
pubmed_ids = record["IdList"]
return(pubmed_ids)
def fetch_pdfs(pubmed_ids, stub_name):
u = url2p.Url2PubmedPmcPdf(pubmed_ids)
urls = u.get_urls()
found = []
for url in urls:
if url['url'] is not None:
cmd = 'curl -L {} -o {}.pdf'.format(url['url'], stub_name + url['pubmed'])
# print(cmd)
check_call(cmd, shell = True)
time.sleep(10)
found.append(url['pubmed'])
return(found)
# def copy_pdf_append_initials(initials):
# pdfs = glob.glob('*.pdf')
# for i in initials:
# os.mkdir(i)
# for p in pdfs:
# pi = p.replace('.pdf', '_' + i + '.pdf')
# copy2(p, os.path.join(i, pi))
def search_and_summarize(search_name, query, omit_ids):
ids = pubmed_central_search_to_pubmed_id(query, retmax=1000000)
new_ids = narrow_id_list(ids, omit_ids)
print('{} search of pubmed found {} ids of which {} are new'.format(search_name, len(ids), len(new_ids)))
return(new_ids)
# -
# ### Desire to send out more articles. 30 in common to all, 30 unique to all
# - 30 common to all + 30 unique per annotator * 4 annotators
# - 120 articles total desired.
# again, these are from preliminary work by Aurelien etc.
AP00 = '"Auditory Perception"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND Humans[Mesh])'
AP01 = '"Auditory Perception"[All Fields] AND "mismatch negativity "[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP02 = '"Auditory Perception"[All Fields] AND (("social control, formal"[MeSH Terms] OR ("social"[All Fields] AND "control"[All Fields] AND "formal"[All Fields]) OR "formal social control"[All Fields] OR "regulation"[All Fields]) AND ("haemodynamic"[All Fields] OR "hemodynamics"[MeSH Terms] OR "hemodynamics"[All Fields] OR "hemodynamic"[All Fields]) AND components[All Fields] AND sensory[All Fields] AND response[All Fields] AND ("substance-related disorders"[MeSH Terms] OR ("substance-related"[All Fields] AND "disorders"[All Fields]) OR "substance-related disorders"[All Fields] OR "habituation"[All Fields])) AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP03 = '"Auditory Perception"[All Fields] AND "hemodynamic"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP04 = '"Auditory Perception"[All Fields] AND "spatial localization"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP05 = '"Auditory Perception"[All Fields] AND (Sensory[All Fields] AND ("evoked potentials"[MeSH Terms] OR ("evoked"[All Fields] AND "potentials"[All Fields]) OR "evoked potentials"[All Fields] OR "erp"[All Fields])) AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP06 = '"Auditory Perception"[All Fields] AND "auditory hallucinations"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP07 = '"Auditory Perception"[All Fields] AND "hyperacusis"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP08 = '"Auditory Perception"[All Fields] AND "McGurk"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP09 = '"Auditory Perception"[All Fields] AND (auditory[All Fields] AND scene[All Fields] AND ("perception"[MeSH Terms] OR "perception"[All Fields])) AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP10 = '"Auditory Perception"[All Fields] AND "auditory masking"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP11 = '"Auditory Perception"[All Fields] AND "Gating"[All Fields] AND ("loattrfree full text"[sb] AND "2010/06/24"[PDat] : "2015/06/22"[PDat] AND "humans"[MeSH Terms])'
AP00_ids = search_and_summarize(search_name='AP00', query=AP00, omit_ids=prev_pmids)
AP01_ids = search_and_summarize(search_name='AP01', query=AP01, omit_ids=prev_pmids)
AP02_ids = search_and_summarize(search_name='AP02', query=AP02, omit_ids=prev_pmids)
AP03_ids = search_and_summarize(search_name='AP03', query=AP03, omit_ids=prev_pmids)
AP04_ids = search_and_summarize(search_name='AP04', query=AP04, omit_ids=prev_pmids)
AP05_ids = search_and_summarize(search_name='AP05', query=AP05, omit_ids=prev_pmids)
AP06_ids = search_and_summarize(search_name='AP06', query=AP06, omit_ids=prev_pmids)
AP07_ids = search_and_summarize(search_name='AP07', query=AP07, omit_ids=prev_pmids)
AP08_ids = search_and_summarize(search_name='AP08', query=AP08, omit_ids=prev_pmids)
AP09_ids = search_and_summarize(search_name='AP09', query=AP09, omit_ids=prev_pmids)
AP10_ids = search_and_summarize(search_name='AP10', query=AP10, omit_ids=prev_pmids)
AP11_ids = search_and_summarize(search_name='AP11', query=AP11, omit_ids=prev_pmids)
# 1449
# Removed this many ids: 17
# AP00 search of pubmed found 1449 ids of which 1432 are new
#
# 56
# Removed this many ids: 3
# AP01 search of pubmed found 56 ids of which 53 are new
#
# 0
# Removed this many ids: 0
# AP02 search of pubmed found 0 ids of which 0 are new
#
# 9
# Removed this many ids: 2
# AP03 search of pubmed found 9 ids of which 7 are new
#
# 3
# Removed this many ids: 1
# AP04 search of pubmed found 3 ids of which 2 are new
#
# 70
# Removed this many ids: 1
# AP05 search of pubmed found 70 ids of which 69 are new
#
# 3
# Removed this many ids: 1
# AP06 search of pubmed found 3 ids of which 2 are new
#
# 6
# Removed this many ids: 3
# AP07 search of pubmed found 6 ids of which 3 are new
#
# 10
# Removed this many ids: 2
# AP08 search of pubmed found 10 ids of which 8 are new
#
# 50
# Removed this many ids: 0
# AP09 search of pubmed found 50 ids of which 50 are new
#
# 1
# Removed this many ids: 0
# AP10 search of pubmed found 1 ids of which 1 are new
#
# 17
# Removed this many ids: 2
# AP11 search of pubmed found 17 ids of which 15 are new
# +
import collections
# total found across all searches
ids = [AP00_ids, AP01_ids, AP02_ids, AP03_ids, AP04_ids, AP05_ids, AP06_ids, AP07_ids, AP08_ids, AP09_ids, AP10_ids, AP11_ids]
ids = [j for i in ids for j in i]
print('total new pmids found across all searches : {}'.format(len(ids)))
# excluding most general search.
ids = [AP01_ids, AP02_ids, AP03_ids, AP04_ids, AP05_ids, AP06_ids, AP07_ids, AP08_ids, AP09_ids, AP10_ids, AP11_ids]
ids = [j for i in ids for j in i]
dups = [item for item, count in collections.Counter(ids).items() if count > 1]
print('duplicate pmids found across all searches : {}'.format(dups))
ids = [AP01_ids, AP02_ids, AP03_ids, AP04_ids, AP05_ids, AP06_ids, AP07_ids, AP08_ids, AP09_ids, AP10_ids, AP11_ids]
ids = [set(id_list) for id_list in ids]
ids = set.union(*ids)
print('unique new pmids found across all searches : {}'.format(len(ids)))
# -
# ### Found plenty new articles
#
# - total new pmids found across all but most general searches :
# - 187
# - But actually retrieved and unique across Audtory Perception for annotation (see below):
# - 146
#
# ### Fetch the pdfs for each search term.
# %pwd
# +
# 5th batch, Auditory perception, subterm...
batch = '05'
fetch_pdfs(AP01_ids, batch + '_AP01_')
#fetch_pdfs(AP02_ids, batch + '_AP02_') # none available
fetch_pdfs(AP03_ids, batch + '_AP03_')
fetch_pdfs(AP04_ids, batch + '_AP04_')
fetch_pdfs(AP05_ids, batch + '_AP05_')
fetch_pdfs(AP06_ids, batch + '_AP06_')
fetch_pdfs(AP07_ids, batch + '_AP07_')
fetch_pdfs(AP08_ids, batch + '_AP08_')
fetch_pdfs(AP09_ids, batch + '_AP09_')
fetch_pdfs(AP10_ids, batch + '_AP10_')
fetch_pdfs(AP11_ids, batch + '_AP11_')
# -
# I then listed the possible duplicate IDs and removed the later occuring duplicates. i.e. if ID occurred in AP01 and AP04 I only kept the AP01 version.
all_found = glob.glob( './*.pdf')
len(all_found)
# +
import collections
def report_found(search_ids, glob_pattern, id_pattern):
'''Report the pmids found within the search set of files.'''
files = glob.glob(glob_pattern)
p = re.compile('[0-9]{8,8}')
found = [p.search(pdf).group() for pdf in files if p.search(pdf)]
not_found = set(search_ids).difference(found)
print()
#print('{} searched = {}'.format(len(search_ids), search_ids))
#print('{} found = {}'.format(len(found), found))
print('{} {} searched.'.format(len(search_ids), glob_pattern))
print('{} {} found.'.format(len(found), glob_pattern))
print('{} {} not_found = {}'.format(len(not_found), glob_pattern, not_found))
return(found)
all_found_d = {}
all_found_d['AP01'] = report_found(AP01_ids, glob_pattern='05_AP01_*', id_pattern='[0-9]{8,8}')
all_found_d['AP03'] = report_found(AP03_ids, glob_pattern='05_AP03_*', id_pattern='[0-9]{8,8}')
all_found_d['AP04'] = report_found(AP04_ids, glob_pattern='05_AP04_*', id_pattern='[0-9]{8,8}')
all_found_d['AP05'] = report_found(AP05_ids, glob_pattern='05_AP05_*', id_pattern='[0-9]{8,8}')
all_found_d['AP06'] = report_found(AP06_ids, glob_pattern='05_AP06_*', id_pattern='[0-9]{8,8}')
all_found_d['AP07'] = report_found(AP07_ids, glob_pattern='05_AP07_*', id_pattern='[0-9]{8,8}')
all_found_d['AP08'] = report_found(AP08_ids, glob_pattern='05_AP08_*', id_pattern='[0-9]{8,8}')
all_found_d['AP09'] = report_found(AP09_ids, glob_pattern='05_AP09_*', id_pattern='[0-9]{8,8}')
all_found_d['AP10'] = report_found(AP10_ids, glob_pattern='05_AP10_*', id_pattern='[0-9]{8,8}')
all_found_d['AP11'] = report_found(AP11_ids, glob_pattern='05_AP11_*', id_pattern='[0-9]{8,8}')
# print()
# print('------- Reporting Found: ----------')
# print()
# print('{} Total found'.format(sum([len(v) for v in all_found_d.values()])))
# print()
all_found_d = collections.OrderedDict(sorted(all_found_d.items()))
# for k,v in all_found_d.iteritems():
# print('{}: {}'.format(k, v))
# -
# ### Reported found or not found for batch 4
# 53 05_AP01_* searched.
# 38 05_AP01_* found.
# 15 05_AP01_* not_found = set(['23242351', '21248117', '26142650', '21624926', '22279229', '25447378', '25173955', '23076102', '23042732', '24992584', '23015429', '21566197', '24336606', '26223716', '23148350'])
#
# 7 05_AP03_* searched.
# 4 05_AP03_* found.
# 3 05_AP03_* not_found = set(['20861363', '23804092', '24401714'])
#
# 2 05_AP04_* searched.
# 2 05_AP04_* found.
# 0 05_AP04_* not_found = set([])
#
# 69 05_AP05_* searched.
# 42 05_AP05_* found.
# 27 05_AP05_* not_found = set(['23242351', '22171057', '24525177', '23042734', '23678108', '21500313', '22790547', '22815876', '22786953', '23954727', '25632143', '25379456', '23308266', '21823798', '26041921', '21625011', '21325518', '21368051', '20578033', '21068187', '23015429', '24336606', '25653372', '22508089', '21893681', '24055864', '21613485'])
#
# 2 05_AP06_* searched.
# 1 05_AP06_* found.
# 1 05_AP06_* not_found = set(['24360035'])
#
# 3 05_AP07_* searched.
# 1 05_AP07_* found.
# 2 05_AP07_* not_found = set(['23876286', '25387539'])
#
# 8 05_AP08_* searched.
# 5 05_AP08_* found.
# 3 05_AP08_* not_found = set(['23810539', '25447378', '21625011'])
#
# 50 05_AP09_* searched.
# 41 05_AP09_* found.
# 9 05_AP09_* not_found = set(['24158725', '24956028', '25948273', '22090489', '24771006', '24548430', '23076102', '21261633', '25259525'])
#
# 1 05_AP10_* searched.
# 1 05_AP10_* found.
# 0 05_AP10_* not_found = set([])
#
# 15 05_AP11_* searched.
# 11 05_AP11_* found.
# 4 05_AP11_* not_found = set(['23035108', '20633569', '22508089', '22464943'])
#
# ------- Reporting Found: ----------
#
# 146 Total found
#
# AP01: ['19929331', '20578033', '20665718', '20929535', '21368051', '21483666', '21750713', '21808660', '21823798', '22163029', '22213909', '22221004', '22551948', '22570723', '22815876', '22916282', '23028971', '23131615', '23241212', '23308266', '23585888', '23617597', '23708059', '23715097', '23825422', '23850664', '23886958', '23920129', '24143195', '24158725', '24366694', '24475052', '24548430', '24771006', '25178752', '25231619', '25342520', '25379456']
#
# AP03: ['21159322', '21500313', '22496909', '23777481']
#
# AP04: ['22786953', '25710328']
#
# AP05: ['20146608', '20493828', '20598152', '20633569', '21233780', '21261633', '21305666', '21380858', '21807011', '21958655', '22367585', '22464943', '22547804', '22592306', '22628458', '22735387', '22773777', '22885999', '23071654', '23145143', '23251704', '23281832', '23301775', '23316957', '23326548', '23616340', '23624493', '23647558', '23664703', '23827717', '23935931', '23988583', '24072639', '24119225', '24314010', '24423729', '24695717', '24956028', '25245785', '25259525', '25535356', '25948269']
#
# AP06: ['22803512']
#
# AP07: ['25231612']
#
# AP08: ['21909378', '22390292', '23664001', '24974346', '24996043']
#
# AP09: ['20541597', '20826671', '20844143', '20975559', '21196054', '21209201', '21355664', '21387016', '21428515', '21945789', '22036957', '22280585', '22371616', '22371619', '22371621', '22612172', '22753470', '22829899', '22844509', '23029426', '23145699', '23423817', '23516340', '23527271', '23691185', '23825404', '23926291', '24003112', '24052177', '24239869', '24475030', '24478375', '24681354', '24711409', '24788808', '24821552', '24841996', '25433224', '25654748', '25659464', '25726262']
#
# AP10: ['22240459']
#
# AP11: ['20649227', '21131368', '22087275', '22384211', '22896044', '23716244', '24298171', '24736181', '24801767', '25024207', '25544613']
# #### Confirming we removed all duplicates
all_found = [v for k,v in all_found_d.items()]
all_found = [item for sublist in all_found for item in sublist]
print(len(all_found) == len(set(all_found)))
print(len(all_found))
# +
# 32 for everyone.
# 38 unique to each of 3 annotators.
import random
files = glob.glob('05_AP*.pdf')
idxs = range(len(files))
random.shuffle(idxs)
everyone = [files[idx] for idx in idxs[0:32]]
mk = [files[idx] for idx in idxs[32:70]]
cc = [files[idx] for idx in idxs[70:108]]
tc = [files[idx] for idx in idxs[108:146]]
# +
def copy_to_annotator(files, annotator, suffix):
# %mkdir {annotator}
# %mkdir {annotator}/annotated
# %mkdir {annotator}/irrelevant
for f in files:
f_dest = f.split('.')[0] + suffix
# %cp {f} ./{annotator}/{f_dest}
copy_to_annotator(mk, 'mk_', '_mk.pdf')
copy_to_annotator(everyone, 'mk_', '_mk.pdf')
# oops... gave tara exact same set as mk
copy_to_annotator(mk, 'tc_', '_tc.pdf')
copy_to_annotator(everyone, 'tc_', '_tc.pdf')
# copy_to_annotator(cc, 'cc_', '_cc.pdf')
# copy_to_annotator(everyone, 'cc_', '_cc.pdf')
# -
# ## Add the new abstracts to our medic database
# Insert fails if any are already in db.
# Update will overwrite previous records.
print(len(all_found))
print(all_found)
# 146
# ['19929331', '20578033', '20665718', '20929535', '21368051', '21483666', '21750713', '21808660', '21823798', '22163029', '22213909', '22221004', '22551948', '22570723', '22815876', '22916282', '23028971', '23131615', '23241212', '23308266', '23585888', '23617597', '23708059', '23715097', '23825422', '23850664', '23886958', '23920129', '24143195', '24158725', '24366694', '24475052', '24548430', '24771006', '25178752', '25231619', '25342520', '25379456', '21159322', '21500313', '22496909', '23777481', '22786953', '25710328', '20146608', '20493828', '20598152', '20633569', '21233780', '21261633', '21305666', '21380858', '21807011', '21958655', '22367585', '22464943', '22547804', '22592306', '22628458', '22735387', '22773777', '22885999', '23071654', '23145143', '23251704', '23281832', '23301775', '23316957', '23326548', '23616340', '23624493', '23647558', '23664703', '23827717', '23935931', '23988583', '24072639', '24119225', '24314010', '24423729', '24695717', '24956028', '25245785', '25259525', '25535356', '25948269', '22803512', '25231612', '21909378', '22390292', '23664001', '24974346', '24996043', '20541597', '20826671', '20844143', '20975559', '21196054', '21209201', '21355664', '21387016', '21428515', '21945789', '22036957', '22280585', '22371616', '22371619', '22371621', '22612172', '22753470', '22829899', '22844509', '23029426', '23145699', '23423817', '23516340', '23527271', '23691185', '23825404', '23926291', '24003112', '24052177', '24239869', '24475030', '24478375', '24681354', '24711409', '24788808', '24821552', '24841996', '25433224', '25654748', '25659464', '25726262', '22240459', '20649227', '21131368', '22087275', '22384211', '22896044', '23716244', '24298171', '24736181', '24801767', '25024207', '25544613']
#
cmd = ' '.join(all_found)
# !medic update {cmd} 2> /dev/null
# !medic --format tsv write ALL 2> /dev/null | cut -f1 | wc # 3359
# ## Save these new ids to a file
batch_05_fname = '/Users/ccarey/Documents/Projects/NAMI/rdoc/tasks/task_data_temp/batch_05_AP_pmids'
with open(batch_05_fname, 'wb') as f:
for pmid in all_found:
f.write(pmid + '\n')
# confirm we have 146 new abstracts
# !medic --format tsv write --pmid-list {batch_05_fname} 2> /dev/null | cut -f1 | wc
# # Noted mistake, accidentally gave entirely same set to everyone.
# desired:
# - 32 for everyone.
# - 38 unique to each of 3 annotators.
#
# - don't change mk
# - change Taras (but keep those she has already done as part of the 'common' set.)
# - change Chucks
# - add in common set to Janet
#
# +
import re
import os
import glob
import collections
import random
random.seed(0)
pattern = '([0-9]{8,8})'
p = re.compile(pattern)
# +
random.seed(0)
pmids = ['19929331', '20578033', '20665718', '20929535', '21368051', '21483666', '21750713', '21808660', '21823798', '22163029', '22213909', '22221004', '22551948', '22570723', '22815876', '22916282', '23028971', '23131615', '23241212', '23308266', '23585888', '23617597', '23708059', '23715097', '23825422', '23850664', '23886958', '23920129', '24143195', '24158725', '24366694', '24475052', '24548430', '24771006', '25178752', '25231619', '25342520', '25379456', '21159322', '21500313', '22496909', '23777481', '22786953', '25710328', '20146608', '20493828', '20598152', '20633569', '21233780', '21261633', '21305666', '21380858', '21807011', '21958655', '22367585', '22464943', '22547804', '22592306', '22628458', '22735387', '22773777', '22885999', '23071654', '23145143', '23251704', '23281832', '23301775', '23316957', '23326548', '23616340', '23624493', '23647558', '23664703', '23827717', '23935931', '23988583', '24072639', '24119225', '24314010', '24423729', '24695717', '24956028', '25245785', '25259525', '25535356', '25948269', '22803512', '25231612', '21909378', '22390292', '23664001', '24974346', '24996043', '20541597', '20826671', '20844143', '20975559', '21196054', '21209201', '21355664', '21387016', '21428515', '21945789', '22036957', '22280585', '22371616', '22371619', '22371621', '22612172', '22753470', '22829899', '22844509', '23029426', '23145699', '23423817', '23516340', '23527271', '23691185', '23825404', '23926291', '24003112', '24052177', '24239869', '24475030', '24478375', '24681354', '24711409', '24788808', '24821552', '24841996', '25433224', '25654748', '25659464', '25726262', '22240459', '20649227', '21131368', '22087275', '22384211', '22896044', '23716244', '24298171', '24736181', '24801767', '25024207', '25544613']
tc_done = glob.glob('/Users/ccarey/Dropbox/20150828_rdoc_project/tc_/annotated/05_AP*.pdf')
tc_done = [os.path.basename(tc) for tc in tc_done]
tc_done_ids = [p.search(pdf).group() for pdf in tc_done if p.search(pdf)]
tc_all = glob.glob('/Users/ccarey/Dropbox/20150828_rdoc_project/tc_/05_AP*.pdf')
tc_all = [os.path.basename(tc) for tc in tc_all]
tc_all_ids = [p.search(pdf).group() for pdf in tc_all if p.search(pdf)]
# Tara did 11 already
print(len(tc_done_ids))
tc_not_done = [tc for tc in tc_all_ids if tc not in tc_done_ids]
random.shuffle(tc_not_done)
tc_more_common = tc_not_done[0:21]
tc_all_common = tc_more_common + tc_done_ids
everyone = tc_all_common
print(everyone)
# -
everyone = ['23071654', '23850664', '24801767', '23647558', '23777481', '23585888', '22464943', '24788808', '23827717', '22496909', '22803512', '23886958', '23617597', '21387016', '23825422', '22829899', '21909378', '22240459', '22896044', '21355664', '22367585', '19929331', '20578033', '20929535', '21368051', '21483666', '21808660', '22221004', '22551948', '22570723', '23028971', '23131615']
# fixing TC
# %pwd
# +
def copy_to_annotator(files, annotator, suffix):
# %mkdir {annotator}
# %mkdir {annotator}/annotated
# %mkdir {annotator}/irrelevant
for f in files:
f_dest = f.split('.')[0] + suffix
# %cp {f} ./{annotator}/{f_dest}
print(everyone)
print(len(everyone))
everyone_files = [glob.glob('*' + e + '.pdf') for e in everyone]
# -
# pmids # all pmids for batch 05
mk_files = glob.glob('/Users/ccarey/Documents/Projects/NAMI/rdoc/pdfs/20160205_rdoc_project/out_20160205/mk_/*.pdf')
mk_files = [os.path.basename(mk) for mk in mk_files]
mk_ids = [p.search(pdf).group() for pdf in mk_files if p.search(pdf)]
to_be_assigned = [pmid for pmid in pmids if pmid not in mk_ids and pmid not in everyone]
len(to_be_assigned)
random.shuffle(to_be_assigned)
# 76 unassigned pmids, assign 38 to TC, assign 38 to CC
# to_be_assigned
to_be_assigned_files = [glob.glob('*' + t + '.pdf') for t in to_be_assigned]
# flatten list of list (each inner list was a single pdf)
everyone_files = [item for e in everyone_files for item in e]
to_be_assigned_files = [item for t in to_be_assigned_files for item in t]
to_be_assigned_files = [item for t in to_be_assigned_files for item in t]
tc_uniq = to_be_assigned_files[0:38]
cc_uniq = to_be_assigned_files[38:76]
copy_to_annotator(everyone_files, 'tc_', '_tc.pdf')
copy_to_annotator(tc_uniq, 'tc_', '_tc.pdf')
# +
copy_to_annotator(everyone_files, 'cc_', '_cc.pdf')
copy_to_annotator(cc_uniq, 'cc_', '_cc.pdf')
copy_to_annotator(everyone_files, 'jl_', '_jl.pdf')
# -
dest_dir = '/Users/ccarey/Documents/Projects/NAMI/rdoc/pdfs/20160205_rdoc_project'
# %cd {dest_dir}
pdfs = glob.glob('out_20160205/*/*.pdf')
pdfs = [os.path.basename(p) for p in pdfs]
pattern = '([0-9]{8,8})'
p = re.compile(pattern)
temp_ids = [p.search(pdf).group() for pdf in pdfs if p.search(pdf)]
counter=collections.Counter(temp_ids)
# noting mistake, accidentally gave all the same to everyone.
counter
| jupyter_notebooks/9_1_round_5_annotations_out.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sentiment Analysis
#
#
# ---
#
# With the rise of online social media platforms like Twitter, Facebook and Reddit, and the proliferation of customer reviews on sites like Amazon and Yelp, we now have access, more than ever before, to massive text-based data sets! They can be analyzed in order to determine how large portions of the population feel about certain products, events, etc. This notebook shows an end-to-end sentiment classification system from scratch.
#
# ## Step 1: Exploring the data!
#
# The dataset we are going to use is very popular among researchers in Natural Language Processing, usually referred to as the [IMDb dataset](http://ai.stanford.edu/~amaas/data/sentiment/). It consists of movie reviews from the website [imdb.com](http://www.imdb.com/), each labeled as either '**pos**itive', if the reviewer enjoyed the film, or '**neg**ative' otherwise. Reference below.
#
# > Maas, <NAME>., et al. [Learning Word Vectors for Sentiment Analysis](http://ai.stanford.edu/~amaas/data/sentiment/). In _Proceedings of the 49th Annual Meeting of the Association for Computational Linguistics: Human Language Technologies_. Association for Computational Linguistics, 2011.
#
# +
import os
import glob
def read_imdb_data(data_dir='data/imdb-reviews'):
"""Read IMDb movie reviews from given directory.
Directory structure expected:
- data/
- train/
- pos/
- neg/
- test/
- pos/
- neg/
"""
# Data, labels to be returned in nested dicts matching the dir. structure
data = {}
labels = {}
# Assume 2 sub-directories: train, test
for data_type in ['train', 'test']:
data[data_type] = {}
labels[data_type] = {}
# Assume 2 sub-directories for sentiment (label): pos, neg
for sentiment in ['pos', 'neg']:
data[data_type][sentiment] = []
labels[data_type][sentiment] = []
# Fetch list of files for this sentiment
path = os.path.join(data_dir, data_type, sentiment, '*.txt')
files = glob.glob(path)
# Read reviews data and assign labels
for f in files:
with open(f) as review:
data[data_type][sentiment].append(review.read())
labels[data_type][sentiment].append(sentiment)
assert len(data[data_type][sentiment]) == len(labels[data_type][sentiment]), \
"{}/{} data size does not match labels size".format(data_type, sentiment)
# Return data, labels as nested dicts
return data, labels
data, labels = read_imdb_data()
print("IMDb reviews: train = {} pos / {} neg, test = {} pos / {} neg".format(
len(data['train']['pos']), len(data['train']['neg']),
len(data['test']['pos']), len(data['test']['neg'])))
# -
# Now that the data is loaded in, let's take a quick look at one of the positive reviews:
print(data['train']['pos'][2])
# And one with a negative sentiment:
print(data['train']['neg'][2])
# We can also make a wordcloud visualization of the reviews.
# +
# Installing wordcloud
# # !pip install wordcloud
# +
import matplotlib.pyplot as plt
# %matplotlib inline
from wordcloud import WordCloud, STOPWORDS
sentiment = 'pos'
# Combine all reviews for the desired sentiment
combined_text = " ".join([review for review in data['train'][sentiment]])
# Initialize wordcloud object
wc = WordCloud(background_color='white', max_words=50,
# update stopwords to include common words like film and movie
stopwords = STOPWORDS.update(['br','film','movie']))
# Generate and plot wordcloud
plt.imshow(wc.generate(combined_text))
plt.axis('off')
plt.show()
# +
sentiment = 'neg'
# Combine all reviews for the desired sentiment
combined_text = " ".join([review for review in data['train'][sentiment]])
# Initialize wordcloud object
wc = WordCloud(background_color='white', max_words=50,
# update stopwords to include common words like film and movie
stopwords = STOPWORDS.update(['br','film','movie']))
# Generate and plot wordcloud
plt.imshow(wc.generate(combined_text))
plt.axis('off')
plt.show()
# -
# They look about the same, but for the words GOOD and BAD
# +
# import random
from sklearn.utils import shuffle
x = ["Spears", "Adele", "NDubz", "Nicole", "Cristina"]
y = [1, 2, 3, 4, 5]
def shuffle_together( a, b):
# author: tj
combined = list( zip(a, b) )
random.shuffle(combined)
a[:], b[:] = zip(*combined)
return a, b
# -
x, y = shuffle(x,y)
print(x,y)
# ### Form training and test sets
#
# Now that we've seen what the raw data looks like, combine the positive and negative documents to get one unified training set and one unified test set.
# +
from sklearn.utils import shuffle
def shuffle_together( a, b):
# author: tj
combined = list( zip(a, b) )
shuffle(combined)
a[:], b[:] = zip(*combined)
return a, b
def prepare_imdb_data(data):
"""Prepare training and test sets from IMDb movie reviews."""
# TODO: Combine positive and negative reviews and labels
# for data_use in ['train','test']:
# list_pos[ data_use ] = data[ data_use ]['pos']
# labels[ data_use ]
data_train = data['train']['pos'] + data['train']['neg']
labels_train = ['pos']*len( data['train']['pos']) + ['neg']*len( data['train']['neg'])
data_test = data['test']['pos'] + data['test']['neg']
labels_test = ['pos']*len( data['test']['pos']) + ['neg']*len( data['test']['neg'])
# TODO: Shuffle reviews and corresponding labels within training and test sets
data_train, labels_train = shuffle( data_train, labels_train )
data_test, labels_test = shuffle( data_test, labels_test )
# Return a unified training data, test data, training labels, test labets
return data_train, data_test, labels_train, labels_test
data_train, data_test, labels_train, labels_test = prepare_imdb_data(data)
print("IMDb reviews (combined): train = {}, test = {}".format(len(data_train), len(data_test)))
# -
# ## Preprocessing
#
# Our raw data includes HTML tags that need to be removed. We also need to remove non-letter characters, normalize uppercase letters by converting them to lowercase, tokenize, remove stop words, and stem the remaining words in each document.
#
# ### Convert each review to words
#
# The following function review_to_words() take care of all of the above.
# +
# BeautifulSoup to easily remove HTML tags
from bs4 import BeautifulSoup
# RegEx for removing non-letter characters
import re
# NLTK library for the remaining steps
import nltk
nltk.download("stopwords") # download list of stopwords (only once; need not run it again)
from nltk.corpus import stopwords # import stopwords
from nltk.stem.porter import *
stemmer = PorterStemmer()
# +
def review_to_words(review):
"""Convert a raw review string into a sequence of words."""
# Remove HTML tags and non-letters,
# convert to lowercase, tokenize,
# remove stopwords and stem
soup = BeautifulSoup(review,"lxml")
text = soup.get_text()
text = text.lower()
text=re.sub(r"[^a-zA-Z0-9]", " ", text) #remove chararacters that aren't letters or digits
words = text.split() # tokenize - The following didn't work because Resource 'punkt' is missing
#now remove stopwords
words = [w for w in words if w not in stopwords.words('english')]
words = [stemmer.stem(w) for w in words]
# Return final list of words
return words
review_to_words("""This is just a <em>test</em>.<br/><br />
But if it weren't a test, it would make for a <b>Great</b> movie review!""")
# -
# With the function `review_to_words()` fully implemeneted, we can apply it to all reviews in both training and test datasets. This may take a while, so we need to build in a mechanism to write to a cache file and retrieve from it later.
# +
import pickle
cache_dir = os.path.join("cache", "sentiment_analysis") # where to store cache files
os.makedirs(cache_dir, exist_ok=True) # ensure cache directory exists
def preprocess_data(data_train, data_test, labels_train, labels_test,
cache_dir=cache_dir, cache_file="preprocessed_data.pkl"):
"""Convert each review to words; read from cache if available."""
# If cache_file is not None, try to read from it first
cache_data = None
if cache_file is not None:
try:
with open(os.path.join(cache_dir, cache_file), "rb") as f:
cache_data = pickle.load(f)
print("Read preprocessed data from cache file:", cache_file)
except:
pass # unable to read from cache, but that's okay
# If cache is missing, then do the heavy lifting
if cache_data is None:
# Preprocess training and test data to obtain words for each review
words_train = list(map(review_to_words, data_train))
words_test = list(map(review_to_words, data_test))
# Write to cache file for future runs
if cache_file is not None:
cache_data = dict(words_train=words_train, words_test=words_test,
labels_train=labels_train, labels_test=labels_test)
with open(os.path.join(cache_dir, cache_file), "wb") as f:
pickle.dump(cache_data, f)
print("Wrote preprocessed data to cache file:", cache_file)
else:
# Unpack data loaded from cache file
words_train, words_test, labels_train, labels_test = (cache_data['words_train'],
cache_data['words_test'], cache_data['labels_train'], cache_data['labels_test'])
return words_train, words_test, labels_train, labels_test
# Preprocess data
words_train, words_test, labels_train, labels_test = preprocess_data(
data_train, data_test, labels_train, labels_test)
# Take a look at a sample
print("\n--- Raw review ---")
print(data_train[1])
print("\n--- Preprocessed words ---")
print(words_train[1])
print("\n--- Label ---")
print(labels_train[1])
# -
# ## Extracting Bag-of-Words features
#
# Now that each document has been preprocessed, we can transform each into a Bag-of-Words feature representation. Note that we need to create this transformation based on the training data alone, as we can't peek at the testing data.
#
# The dictionary or _vocabulary_ $V$ (set of words shared by documents in the training set) used here will be the one on which we train our supervised learning algorithm. Any future test data must be transformed in the same way for us to be able to apply the learned model for prediction. Hence, it is important to store the transformation / vocabulary as well.
#
# > **Note**: The set of words in the training set may not be exactly the same as the test set. What if we encounter a word during testing that you haven't seen before? Unfortunately, we'll have to ignore it, or replace it with a special `<UNK>` token.
#
# ### Compute Bag-of-Words features
# +
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.externals import joblib
# joblib is an enhanced version of pickle that is more efficient for storing NumPy arrays
def extract_BoW_features(words_train, words_test, vocabulary_size=5000,
cache_dir=cache_dir, cache_file="bow_features.pkl"):
"""Extract Bag-of-Words for a given set of documents, already preprocessed into words."""
# If cache_file is not None, try to read from it first
cache_data = None
if cache_file is not None:
try:
with open(os.path.join(cache_dir, cache_file), "rb") as f:
cache_data = joblib.load(f)
print("Read features from cache file:", cache_file)
except:
pass # unable to read from cache, but that's okay
# If cache is missing, then do the heavy lifting
if cache_data is None:
# Fit a vectorizer to training documents and use it to transform them
vectorizer = CountVectorizer(max_features=vocabulary_size,
preprocessor=lambda x: x, tokenizer=lambda x: x) # already preprocessed
features_train = vectorizer.fit_transform(words_train).toarray()
# Apply the same vectorizer to transform the test documents (ignore unknown words)
features_test = vectorizer.transform(words_test).toarray()
# NOTE: Remember to convert the features using .toarray() for a compact representation
# Write to cache file for future runs (store vocabulary as well)
if cache_file is not None:
vocabulary = vectorizer.vocabulary_
cache_data = dict(features_train=features_train, features_test=features_test,
vocabulary=vocabulary)
with open(os.path.join(cache_dir, cache_file), "wb") as f:
joblib.dump(cache_data, f)
print("Wrote features to cache file:", cache_file)
else:
# Unpack data loaded from cache file
features_train, features_test, vocabulary = (cache_data['features_train'],
cache_data['features_test'], cache_data['vocabulary'])
# Return both the extracted features as well as the vocabulary
return features_train, features_test, vocabulary
# Extract Bag of Words features for both training and test datasets
features_train, features_test, vocabulary = extract_BoW_features(words_train, words_test)
# Inspect the vocabulary that was computed
print("Vocabulary: {} words".format(len(vocabulary)))
import random
print("Sample words: {}".format(random.sample(list(vocabulary.keys()), 8)))
# Sample
print("\n--- Preprocessed words ---")
print(words_train[5])
print("\n--- Bag-of-Words features ---")
print(features_train[5])
print("\n--- Label ---")
print(labels_train[5])
# -
# Let's try to visualize the Bag-of-Words feature vector for one of our training documents.
# Plot the BoW feature vector for a training document
plt.plot(features_train[5,:])
plt.xlabel('Word')
plt.ylabel('Count')
plt.show()
np.mean(np.mean(features_train[:], axis=1))
np.max(np.mean(features_train[:], axis=1))
np.min(np.mean(features_train[:], axis=1))
# #### Sparse Bag-of-Words feature representation
#
# On average, only 2.16% of the vocabulary is seen in a single document. The rest of the entries are zeros. The biggest vocabulary use is 22%
#
#
# ### Zipf's law
#
# [Zipf's law](https://en.wikipedia.org/wiki/Zipf%27s_law), named after the American linguist <NAME>, is an empirical law stating that given a large collection of documents, the frequency of any word is inversely proportional to its rank in the frequency table. So the most frequent word will occur about twice as often as the second most frequent word, three times as often as the third most frequent word, and so on. In the figure below we plot number of appearances of each word in our training set against its rank.
# +
# Find number of occurrences for each word in the training set
word_freq = features_train.sum(axis=0)
# Sort it in descending order
sorted_word_freq = np.sort(word_freq)[::-1]
# Plot
plt.plot(sorted_word_freq)
plt.gca().set_xscale('log')
plt.gca().set_yscale('log')
plt.xlabel('Rank')
plt.ylabel('Number of occurrences')
plt.show()
# -
sorted_word_freq[:5] #not quite Zipf on the first 2, but, then it looks like it
# #### Zipf's law
#
# The most frequent word occurs 51695 times. The second is 48190, not 25000 as predicted by Zipf's law. Perhaps because we threw out all stop words etc?
#
#
# ### Normalize feature vectors
#
# Bag-of-Words features are intuitive to understand as they are simply word counts. But counts can vary a lot, and potentially throw off learning algorithms later in the pipeline. So, before we proceed further, we normalize the BoW feature vectors to have unit length.
#
# This makes sure that each document's representation retains the unique mixture of feature components, but prevents documents with large word counts from dominating those with fewer words.
# +
import sklearn.preprocessing as pr
# Normalize BoW features in training and test set
# first checking that pr.normalize is indeed dividing by the L2 norm of the vector
row = features_train[0,:]
x = pr.normalize(features_train[:1,:], axis=1)
y = row / np.sqrt(np.sum([elem**2 for elem in row]))
[np.sum(x), np.sum(y)]
# -
# Normalize BoW features in training and test set
features_train = pr.normalize(features_train, axis=1)
features_test = pr.normalize(features_test, axis=1)
# ## Classification using BoW features
#
# Now that the data has all been properly transformed, we can feed it into a classifier. To get a baseline model, we train a Naive Bayes classifier from scikit-learn (specifically, [`GaussianNB`](http://scikit-learn.org/stable/modules/generated/sklearn.naive_bayes.GaussianNB.html)), and evaluate its accuracy on the test set.
# +
from sklearn.naive_bayes import GaussianNB
# Train a Guassian Naive Bayes classifier
clf1 = GaussianNB()
clf1.fit(features_train, labels_train)
# Calculate the mean accuracy score on training and test sets
print("[{}] Accuracy: train = {}, test = {}".format(
clf1.__class__.__name__,
clf1.score(features_train, labels_train),
clf1.score(features_test, labels_test)))
# -
# Tree-based algorithms often work quite well on Bag-of-Words as their highly discontinuous and sparse nature is nicely matched by the structure of trees. As your next task, you will try to improve on the Naive Bayes classifier's performance by using scikit-learn's Gradient-Boosted Decision Tree classifer.
#
# ### Gradient-Boosted Decision Tree classifier
#
# Using [`GradientBoostingClassifier`](http://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingClassifier.html) from scikit-learn to classify the BoW data. This model has a number of parameters.
#
# > We can use a model selection technique such as cross-validation, grid-search, or an information criterion method, to find an optimal value for the hyperparameter.
# +
from sklearn.ensemble import GradientBoostingClassifier
n_estimators = 32
def classify_gboost(X_train, X_test, y_train, y_test):
# Initialize classifier
clf = GradientBoostingClassifier(n_estimators=n_estimators, learning_rate=1.0, max_depth=1, random_state=0)
# Classify the data using GradientBoostingClassifier
clf.fit(X_train, y_train)
# Perform hyperparameter tuning / model selection
# Print final training & test accuracy
print("[{}] Accuracy: train = {}, test = {}".format(
clf.__class__.__name__,
clf.score(X_train, y_train),
clf.score(X_test, y_test) ) )
# Return best classifier model
return clf
clf2 = classify_gboost(features_train, features_test, labels_train, labels_test)
# -
# ### Adverserial testing
#
# Tricking the algorithm to expose the main weakness of BoW by creating a movie review with a clear positive or negative sentiment that the model will classify incorrectly.
#
# +
my_review = "Not the dreary bore that I expected it to be. In fact, it was packed with action and I was at the edge of my seat. The pace of this movie makes the Matrix look depressingly quiet."
true_sentiment = 'pos' # true sentiment
# Apply the same preprocessing and vectorizing steps as you did for your training data
my_words = review_to_words( my_review )
vectorizer = CountVectorizer(vocabulary=vocabulary,
preprocessor=lambda x: x, tokenizer=lambda x: x) # already preprocessed
# note: # This vocabulary came from the vectorizer during previous training
# features_train, features_test, vocabulary = extract_BoW_features(words_train, words_test)
my_features = vectorizer.fit_transform( [my_words] ).toarray()
# Then call your classifier to label it
print("my review: ", my_review )
print("my words: ", my_words)
print("model-predicted sentiment: ", clf2.predict( my_features ), " whereas true sentiment is: ", true_sentiment)
| sentiment-analysis/Sentiment-Analysis-Gradient-Boosting/Gradient-Boosted-Classifier_sentiment_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit (conda)
# name: python3
# ---
# +
import os
EXAMPLE_DIR = os.getcwd()
os.chdir(os.path.join('..', '..'))
PROJECT_DIR = os.getcwd()
EXAMPLE_NAME = os.path.basename(EXAMPLE_DIR)
print('EXAMPLE_DIR: ', EXAMPLE_DIR)
print('PROJECT_DIR: ', PROJECT_DIR)
# jupyter-notebook --notebook-dir=$PWD --port=8889 --ip=0.0.0.0
# +
# %load_ext autoreload
# %autoreload 2
import sys
import gym
import time
import dill
import numpy as np
import networkx as nx
from typing import List
from collections import defaultdict
sys.path.append(os.path.join(PROJECT_DIR, 'src'))
from src.graph import Graph
from src.graph import graph_factory
from src.config import ROOT_PATH
from src.strategy_synthesis.multiobjective_solver import MultiObjectiveSolver
from src.graph.trans_sys import FiniteTransSys
from src.spot.Parser import ANDExpression, SymbolExpression, NotSymbolExpression, TrueExpression
# TODO: Install Wombats to your local directory relative to regret_synthesis_toolbox
WOMBATS_EXPERIMENT_DIRECTORY = os.path.join(os.path.dirname(PROJECT_DIR), 'wombats_experiments')
sys.path.append(WOMBATS_EXPERIMENT_DIRECTORY)
from wombats.systems.minigrid import DynamicMinigrid2PGameWrapper, GYM_MONITOR_LOG_DIR_NAME, BoxPacking
Graph.graph_dir = EXAMPLE_DIR
# +
debug = True
dfa_config_yaml="config/DFA"
pdfa_config_yaml="config/PDFA"
strategy_config_yaml="config/Strategy"
# Graph Arguments
load_game_from_file = False
plot_minigrid = False
plot_pdfa = True
plot_product = False
finite = True
view = True
save_flag = True
format = 'png'
# Multi-Objective Optimizer Arguments
stochastic = False
adversarial = True
plot_strategies=False
plot_graph_with_strategy = False
plot_graph_with_pareto = False
plot_pareto = True
speedup = True
NUM_OBJ = 3
PLAYER_MAPPING = {'eve': 'sys', 'adam': 'env'}
ROS_PACKAGE_DIR = os.path.join(PROJECT_DIR, '..')
STRATEGY_PICKLE_FILENAME = os.path.join(ROS_PACKAGE_DIR, 'python_examples', 'strategy.pickle')
LOCATION_MAPPINGS_PICKLE_FILENAME = os.path.join(ROS_PACKAGE_DIR, 'python_examples', 'location_mappings.pickle')
ADD_SYS_WEIGHT = True
ADD_ENV_WEIGHT = True
NUM_LOCATION = 3
ENV_ID = 'MiniGrid-Franka-BoxPacking-v0'
# -
# # Construct DFA and Manually Specify PDFA from DFA
#
#
# PDFA
pdfa = graph_factory.get(
'PDFA',
graph_name="pdfa2",
config_yaml=pdfa_config_yaml,
save_flag=save_flag,
plot=plot_pdfa,
view=view,
format=format)
# # Define a Franka environment
# +
LOCATIONS = ['L0', 'L1', 'L2', 'L3']
OBJECT_LOCATIONS = {
'o0': 'L0',
'o1': 'L1',
'o2': 'L2'}
TARGET_LOCATIONS = ['L3', 'L3', 'L3']
LOCATION_MAPPINGS = {
'H': np.array([ 0, 0, 0.085]),
'L0': np.array([0.6, 0.6, 0.085]),
'L1': np.array([0.5, -0.5, 0.085]),
'L2': np.array([0.0, 0.3, 0.085]),
'L3': np.array([0.3, 0, 0.085]),
}
world_config_kwargs = {
'locations': LOCATIONS,
'object_locations': OBJECT_LOCATIONS,
'target_locations': TARGET_LOCATIONS,
'distance_mappings': BoxPacking.locations_to_distance_mappings(LOCATION_MAPPINGS)}
env = gym.make(ENV_ID, **world_config_kwargs)
env = DynamicMinigrid2PGameWrapper(
env,
player_steps={'sys': [1], 'env': [1]},
monitor_log_location=os.path.join(EXAMPLE_DIR, GYM_MONITOR_LOG_DIR_NAME))
# +
file_name = ENV_ID + 'Game'
filepath = os.path.join(EXAMPLE_DIR, 'config', file_name)
config_yaml = os.path.relpath(filepath, ROOT_PATH)
# Game Construction
start = time.time()
trans_sys = graph_factory.get('TwoPlayerGraph',
graph_name='TwoPlayerGame',
config_yaml=config_yaml,
minigrid=env,
save_flag=save_flag,
plot=plot_minigrid,
view=view,
format=format)
end = time.time()
# -
init_node = trans_sys.get_initial_states()[0][0]
# trans_sys.plot_graph(format=format, view=False, start_node=init_node, n_neighbor=5)
# trans_sys.plot_graph(format=format, view=False)
# # Manually Define Game
# Given a list of objects, construct a product graph
#
# Product Game Construction
start = time.time()
game = graph_factory.get('ProductGraph',
graph_name=f'{EXAMPLE_NAME}_ProductAutomaton',
config_yaml=None,
trans_sys=trans_sys,
automaton=pdfa,
save_flag=True,
prune=False,
debug=False,
absorbing=True,
finite=finite,
plot=plot_product,
integrate_accepting=True,
view=view,
format=format)
end = time.time()
print(f'Product Construction took {end-start:.2f} seconds')
# +
# game.plot_graph(view=view, format=format)
# -
solver = MultiObjectiveSolver(game,
epsilon=1e-5,
max_iteration=300,
stochastic=stochastic,
adversarial=adversarial)
solver.solve(plot_strategies=plot_strategies,
plot_graph_with_strategy=plot_graph_with_strategy,
plot_graph_with_pareto=plot_graph_with_pareto,
plot_pareto=plot_pareto,
speedup=speedup,
debug=debug,
view=view,
format=format)
# +
# strategies = solver.get_strategies()
# strategy = list(strategies.values())[0]
# strategy._config_yaml = "config/Strategy"
# strategy.plot_graph(view=view, format=format, save_yaml=True)
for i, strategy in enumerate(list(solver.get_strategies().values())):
strategy._config_yaml = f"config/Strategy{i}"
strategy.plot_graph(view=False, format=format)
STRATEGY_PICKLE_FILENAME = os.path.join(EXAMPLE_DIR, '3', f'Strategy{i}.pickle')
try:
with open(STRATEGY_PICKLE_FILENAME, 'wb') as f:
dill.dump(strategy, f)
print(f'File Saved as {STRATEGY_PICKLE_FILENAME}')
except Exception as e:
print(e)
raise e
# -
# # TODO:
# 1. Read the strategy file
# 2. The system agent takes an action
# 3. Let the env chooses its action
# 4. Transit to the next system state (Be able to Identify the current state from the object locations)
# 5. 2-3
# 6. Check whether the current state is in the accepting state
def run(filename: str):
with open(filename, 'rb') as f:
strategy = dill.load(f)
print('-'*100)
print(f"Evaluate for a pareto point {strategy._init_pareto_point}")
print('-'*100)
runs_per_pareto, actions_per_pareto, cost_per_pareto, obs_per_pareto = strategy.get_max_cost_runs()
for p, plays in actions_per_pareto.items():
print('='*100)
print(f'Pareto Point {p}')
print('='*100)
costs = cost_per_pareto[p]
obss = obs_per_pareto[p]
for i, play in enumerate(plays):
play_cost = costs[i]
obs = obss[i]
print(f'Play {i}: Cost={np.sum(play_cost, axis=0)}')
for actions, c, o in zip(play, play_cost, obs):
print('\t', actions, c, o)
filename = os.path.join(EXAMPLE_DIR, '3', 'Strategy0.pickle')
run(filename)
filename = os.path.join(EXAMPLE_DIR, '3', 'Strategy1.pickle')
run(filename)
# +
player = 'sys'
SYS_ACTIONS = []
for multiactions in env.player_actions[player]:
action_strings = []
for agent, actions in zip(env.unwrapped.agents, multiactions):
action_string = []
for action in actions:
if action is None or np.isnan(action):
continue
a_str = agent.ACTION_ENUM_TO_STR[action]
action_string.append(a_str)
action_strings.append(tuple(action_string))
action_strs = action_strings[0] if player == 'sys' else action_strings[1:]
SYS_ACTIONS.append(tuple(action_strs))
player = 'env'
ENV_ACTIONS = []
for multiactions in env.player_actions[player]:
action_strings = []
for agent, actions in zip(env.unwrapped.agents, multiactions):
action_string = []
for action in actions:
if action is None or np.isnan(action):
continue
a_str = agent.ACTION_ENUM_TO_STR[action]
action_string.append(a_str)
action_strings.append(tuple(action_string))
action_strs = action_strings[0] if player == 'sys' else action_strings[1:]
ENV_ACTIONS.append(tuple(action_strs))
print(SYS_ACTIONS)
print(ENV_ACTIONS)
# +
from src.simulation.simulator import Simulator
filename = os.path.join(EXAMPLE_DIR, '3', 'Strategy1.pickle')
with open(filename, 'rb') as f:
strategy = dill.load(f)
ts0 = SYS_ACTIONS[0]
ts1 = SYS_ACTIONS[1]
ts2 = SYS_ACTIONS[2]
tf0 = SYS_ACTIONS[3]
tf1 = SYS_ACTIONS[4]
tf2 = SYS_ACTIONS[5]
tf3 = SYS_ACTIONS[6]
waits = SYS_ACTIONS[7]
i0 = ENV_ACTIONS[0]
i1 = ENV_ACTIONS[1]
i2 = ENV_ACTIONS[2]
wait = ENV_ACTIONS[3]
ret = ENV_ACTIONS[4]
# env_actions = [wait, wait, wait, wait, i0, wait, ret, i1,
# wait, ret, wait, i2, wait, ret, wait, wait]
sim = Simulator(env, strategy._game)
# sim.run_turn_based_game(
# sys_strategy=strategy,
# env_actions=env_actions,
# render=False,
# record_video=False)
# sim.get_stats()
# +
sys_actions = [ts0, tf3, ts1, tf3, ts2, tf3]
env_actions = [wait, wait, wait, wait, wait, wait]
env_state, env_action, sys_state, cost = strategy._game.reset()
i = 0
costs = []
while True:
sys_action = sys_actions[i]
env_state, obs, cost, done = strategy._game.step(sys_state, sys_action)
costs.append(cost)
if i == len(env_actions):
state = env_state
break
env_action = env_actions[i]
sys_state, obs, cost, done = strategy._game.step(env_state, env_action)
costs.append(cost)
i += 1
if i == len(sys_actions):
state = sys_state
break
print(state)
next_states = list(strategy._game._graph.successors(state))
print(next_states)
print('Cost', np.sum(costs, axis=0))
# -
| examples/franka_box_packing_extended/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Ejys4qb9K4yp" colab_type="text"
# <table class="tfo-notebook-buttons" align="center">
# <td>
# <a
# target="_blank"
# href="https://colab.research.google.com/github/notebookexplore/notebookexplore/blob/master/audio-processing/pytorch/audio_processing_tutorial.ipynb"
# ><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run
# in Google Colab</a
# >
# </td>
# <td>
# <a
# target="_blank"
# href="https://github.com/notebookexplore/notebookexplore/blob/master/audio-processing/pytorch/audio_processing_tutorial.ipynb"
# ><img
# src="https://www.tensorflow.org/images/GitHub-Mark-32px.png"
# />View source on GitHub</a
# >
# </td>
# </table>
# + id="U8adtjNkog1-" colab_type="code" colab={}
# !pip install torch>=1.2.0
# !pip install torchaudio
# %matplotlib inline
# + [markdown] id="oHpAbYz2og2G" colab_type="text"
#
# torchaudio Tutorial
# ===================
#
# PyTorch is an open source deep learning platform that provides a
# seamless path from research prototyping to production deployment with
# GPU support.
#
# Significant effort in solving machine learning problems goes into data
# preparation. torchaudio leverages PyTorch’s GPU support, and provides
# many tools to make data loading easy and more readable. In this
# tutorial, we will see how to load and preprocess data from a simple
# dataset.
#
# For this tutorial, please make sure the ``matplotlib`` package is
# installed for easier visualization.
#
#
#
# + id="AFPA4DPUog2I" colab_type="code" colab={}
import torch
import torchaudio
import matplotlib.pyplot as plt
# + [markdown] id="4HhXSHmGog2P" colab_type="text"
# Opening a dataset
# -----------------
#
#
#
# + [markdown] id="QEbIBsDFog2R" colab_type="text"
# torchaudio supports loading sound files in the wav and mp3 format. We
# call waveform the resulting raw audio signal.
#
#
#
# + id="Uh7cF3wnog2S" colab_type="code" colab={}
import requests
url = "https://pytorch.org/tutorials//_static/img/steam-train-whistle-daniel_simon-converted-from-mp3.wav"
r = requests.get(url)
with open('steam-train-whistle-daniel_simon-converted-from-mp3.wav', 'wb') as f:
f.write(r.content)
filename = "steam-train-whistle-daniel_simon-converted-from-mp3.wav"
waveform, sample_rate = torchaudio.load(filename)
print("Shape of waveform: {}".format(waveform.size()))
print("Sample rate of waveform: {}".format(sample_rate))
plt.figure()
plt.plot(waveform.t().numpy())
# + [markdown] id="sLxzNge9og2X" colab_type="text"
# Transformations
# ---------------
#
# torchaudio supports a growing list of
# `transformations <https://pytorch.org/audio/transforms.html>`_.
#
# - **Resample**: Resample waveform to a different sample rate.
# - **Spectrogram**: Create a spectrogram from a waveform.
# - **MelScale**: This turns a normal STFT into a Mel-frequency STFT,
# using a conversion matrix.
# - **AmplitudeToDB**: This turns a spectrogram from the
# power/amplitude scale to the decibel scale.
# - **MFCC**: Create the Mel-frequency cepstrum coefficients from a
# waveform.
# - **MelSpectrogram**: Create MEL Spectrograms from a waveform using the
# STFT function in PyTorch.
# - **MuLawEncoding**: Encode waveform based on mu-law companding.
# - **MuLawDecoding**: Decode mu-law encoded waveform.
#
# Since all transforms are nn.Modules or jit.ScriptModules, they can be
# used as part of a neural network at any point.
#
#
#
# + [markdown] id="JLoCOqHvog2Z" colab_type="text"
# To start, we can look at the log of the spectrogram on a log scale.
#
#
#
# + id="T-JJqufHog2a" colab_type="code" colab={}
specgram = torchaudio.transforms.Spectrogram()(waveform)
print("Shape of spectrogram: {}".format(specgram.size()))
plt.figure()
plt.imshow(specgram.log2()[0,:,:].numpy(), cmap='gray')
# + [markdown] id="LTshMbr9og2i" colab_type="text"
# Or we can look at the Mel Spectrogram on a log scale.
#
#
#
# + id="tL_b8Oa5og2j" colab_type="code" colab={}
specgram = torchaudio.transforms.MelSpectrogram()(waveform)
print("Shape of spectrogram: {}".format(specgram.size()))
plt.figure()
p = plt.imshow(specgram.log2()[0,:,:].detach().numpy(), cmap='gray')
# + [markdown] id="0GNui3v-og2n" colab_type="text"
# We can resample the waveform, one channel at a time.
#
#
#
# + id="1eHZFUR8og2p" colab_type="code" colab={}
new_sample_rate = sample_rate/10
# Since Resample applies to a single channel, we resample first channel here
channel = 0
transformed = torchaudio.transforms.Resample(sample_rate, new_sample_rate)(waveform[channel,:].view(1,-1))
print("Shape of transformed waveform: {}".format(transformed.size()))
plt.figure()
plt.plot(transformed[0,:].numpy())
# + [markdown] id="1mT305QXog2s" colab_type="text"
# As another example of transformations, we can encode the signal based on
# Mu-Law enconding. But to do so, we need the signal to be between -1 and
# 1. Since the tensor is just a regular PyTorch tensor, we can apply
# standard operators on it.
#
#
#
# + id="o5hJAVEnog2u" colab_type="code" colab={}
# Let's check if the tensor is in the interval [-1,1]
print("Min of waveform: {}\nMax of waveform: {}\nMean of waveform: {}".format(waveform.min(), waveform.max(), waveform.mean()))
# + [markdown] id="vlk9qo4Rog2y" colab_type="text"
# Since the waveform is already between -1 and 1, we do not need to
# normalize it.
#
#
#
# + id="TI3yBBSGog2z" colab_type="code" colab={}
def normalize(tensor):
# Subtract the mean, and scale to the interval [-1,1]
tensor_minusmean = tensor - tensor.mean()
return tensor_minusmean/tensor_minusmean.abs().max()
# Let's normalize to the full interval [-1,1]
# waveform = normalize(waveform)
# + [markdown] id="3Yu7ubXZog22" colab_type="text"
# Let’s apply encode the waveform.
#
#
#
# + id="qjytiCNYog23" colab_type="code" colab={}
transformed = torchaudio.transforms.MuLawEncoding()(waveform)
print("Shape of transformed waveform: {}".format(transformed.size()))
plt.figure()
plt.plot(transformed[0,:].numpy())
# + [markdown] id="pgApI-D6og27" colab_type="text"
# And now decode.
#
#
#
# + id="Ciijz1Kgog28" colab_type="code" colab={}
reconstructed = torchaudio.transforms.MuLawDecoding()(transformed)
print("Shape of recovered waveform: {}".format(reconstructed.size()))
plt.figure()
plt.plot(reconstructed[0,:].numpy())
# + [markdown] id="fyJ02hGMog3C" colab_type="text"
# We can finally compare the original waveform with its reconstructed
# version.
#
#
#
# + id="4B6pyGd5og3D" colab_type="code" colab={}
# Compute median relative difference
err = ((waveform-reconstructed).abs() / waveform.abs()).median()
print("Median relative difference between original and MuLaw reconstucted signals: {:.2%}".format(err))
# + [markdown] id="ayNUB_keog3H" colab_type="text"
# Migrating to torchaudio from Kaldi
# ----------------------------------
#
# Users may be familiar with
# `Kaldi <http://github.com/kaldi-asr/kaldi>`_, a toolkit for speech
# recognition. torchaudio offers compatibility with it in
# ``torchaudio.kaldi_io``. It can indeed read from kaldi scp, or ark file
# or streams with:
#
# - read_vec_int_ark
# - read_vec_flt_scp
# - read_vec_flt_arkfile/stream
# - read_mat_scp
# - read_mat_ark
#
# torchaudio provides Kaldi-compatible transforms for ``spectrogram`` and
# ``fbank`` with the benefit of GPU support, see
# `here <compliance.kaldi.html>`__ for more information.
#
#
#
# + id="NiSS8s10og3I" colab_type="code" colab={}
n_fft = 400.0
frame_length = n_fft / sample_rate * 1000.0
frame_shift = frame_length / 2.0
params = {
"channel": 0,
"dither": 0.0,
"window_type": "hanning",
"frame_length": frame_length,
"frame_shift": frame_shift,
"remove_dc_offset": False,
"round_to_power_of_two": False,
"sample_frequency": sample_rate,
}
specgram = torchaudio.compliance.kaldi.spectrogram(waveform, **params)
print("Shape of spectrogram: {}".format(specgram.size()))
plt.figure()
plt.imshow(specgram.t().numpy(), cmap='gray')
# + [markdown] id="v5DauoCqog3M" colab_type="text"
# We also support computing the filterbank features from waveforms,
# matching Kaldi’s implementation.
#
#
#
# + id="ONLSeJfIog3N" colab_type="code" colab={}
fbank = torchaudio.compliance.kaldi.fbank(waveform, **params)
print("Shape of fbank: {}".format(fbank.size()))
plt.figure()
plt.imshow(fbank.t().numpy(), cmap='gray')
# + [markdown] id="BX519cRTog3S" colab_type="text"
# Conclusion
# ----------
#
# We used an example raw audio signal, or waveform, to illustrate how to
# open an audio file using torchaudio, and how to pre-process and
# transform such waveform. Given that torchaudio is built on PyTorch,
# these techniques can be used as building blocks for more advanced audio
# applications, such as speech recognition, while leveraging GPUs.
#
#
#
| audio-processing/pytorch/audio_preprocessing_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Delaying a connection with a node
#
# Nodes allow for all sorts of advanced behavior
# that is typically done by modifying the code of a neural simulator.
# In Nengo, the `Node` object allows us to run custom code.
#
# In this example, we will implement
# an `n`-timestep delayed connection by using a node.
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import nengo
from nengo.processes import WhiteSignal
# +
model = nengo.Network(label="Delayed connection")
with model:
# We'll use white noise as input
inp = nengo.Node(WhiteSignal(2, high=5), size_out=1)
A = nengo.Ensemble(40, dimensions=1)
nengo.Connection(inp, A)
# We'll make a simple object to implement the delayed connection
class Delay:
def __init__(self, dimensions, timesteps=50):
self.history = np.zeros((timesteps, dimensions))
def step(self, t, x):
self.history = np.roll(self.history, -1)
self.history[-1] = x
return self.history[0]
dt = 0.001
delay = Delay(1, timesteps=int(0.2 / 0.001))
with model:
delaynode = nengo.Node(delay.step, size_in=1, size_out=1)
nengo.Connection(A, delaynode)
# Send the delayed output through an ensemble
B = nengo.Ensemble(40, dimensions=1)
nengo.Connection(delaynode, B)
# Probe the input at the delayed output
A_probe = nengo.Probe(A, synapse=0.01)
B_probe = nengo.Probe(B, synapse=0.01)
# -
# Run for 2 seconds
with nengo.Simulator(model) as sim:
sim.run(2)
# Plot the results
plt.figure()
plt.subplot(2, 1, 1)
plt.plot(sim.trange(), sim.data[A_probe], lw=2)
plt.title("Input")
plt.subplot(2, 1, 2)
plt.plot(sim.trange(), sim.data[B_probe], lw=2)
plt.axvline(0.2, c="k")
plt.title("Delayed output")
plt.tight_layout()
| docs/examples/usage/delay-node.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Flux.pl
# The `Flux.pl` Perl script takes four input parameters:
#
# `Flux.pl [input file] [output file] [bin width (s)] [geometry base directory]`
#
# or, as invoked from the command line,
#
# `$ perl ./perl/Flux.pl [input file] [output file] [bin width (s)] [geometry directory]`
# ## Input Parameters
# * `[input file]`
#
# `Flux.pl` expects the first non-comment line of the input file to begin with a string of the form `<DAQ ID>.<channel>`. This is satisfied by threshold and wire delay files, as well as the outputs of data transformation scripts like `Sort.pl` and `Combine.pl` if their inputs are of the appropriate form.
#
# If the input file doesn't meet this condition, `Flux.pl` -- specifically, the `all_geo_info{}` subroutine of `CommonSubs.pl` -- won't be able to load the appropriate geometry files and execution will fail (see the `[geometry directory]` parameter below).
# * `[output file]`
#
# This is what the output file will be named.
# * `[binWidth]`
#
# In physical terms, cosmic ray _flux_ is the number of incident rays per unit area per unit time. The `[binWidth]` parameter determines the "per unit time" portion of this quantity. `Flux.pl` will sort the events in its input data into bins of the given time interval, returning the number of events per unit area recorded within each bin.
# * `[geometry directory]`
#
# With `[binWidth]` handling the "per unit time" portion of the flux calculation, the geometry file associated with each detector handles the "per unit area".
#
# `Flux.pl` expects geometry files to be stored in a directory structure of the form
#
# ```
# geo/
# ├── 6119/
# │ └── 6119.geo
# ├── 6148/
# │ └── 6148.geo
# └── 6203/
# └── 6203.geo
# ```
#
# where each DAQ has its own subdirectory whose name is the DAQ ID, and each such subdirectory has a geometry file whose name is given by the DAQ ID with the `.geo` extension. The command-line argument in this case is `geo/`, the parent directory. With this as the base directory, `Flux.pl` determines what geometry file to load by looking for the DAQ ID in the first line of data. This is why, as noted above, the first non-comment line of `[input file]` must begin with `<DAQ ID>.<channel>`.
# ## Flux Input Files
# As we mentioned above, threshold files have the appropriate first-line structure to allow `Flux.pl` to access geometry data for them. So what does `Flux.pl` do when acting on a threshold file?
# We'll test it using the threshold files `files/6148.2016.0109.0.thresh` and `files/6119.2016.0104.1.thresh` as input. First, take a look at the files themselves so we know what the input looks like:
# !head -10 files/6148.2016.0109.0.thresh
# !wc -l files/6148.2016.0109.0.thresh
# !head -10 files/6119.2016.0104.1.thresh
# !wc -l files/6119.2016.0104.1.thresh
# (remember, `wc -l` returns a count of the number of lines in the file). These look like fairly standard threshold files. Now we'll see what `Flux.pl` does with them.
# ## The Parsl Flux App
# For convenience, we'll wrap the UNIX command-line invocation of the `Flux.pl` script in a Parsl App, which will make it easier to work with from within the Jupyter Notebook environment.
# +
# The prep work:
import parsl
from parsl.config import Config
from parsl.executors.threads import ThreadPoolExecutor
from parsl.app.app import bash_app,python_app
from parsl import File
config = Config(
executors=[ThreadPoolExecutor()],
lazy_errors=True
)
parsl.load(config)
# -
# The App:
@bash_app
def Flux(inputs=[], outputs=[], binWidth='600', geoDir='geo/', stdout='stdout.txt', stderr='stderr.txt'):
return 'perl ./perl/Flux.pl %s %s %s %s' % (inputs[0], outputs[0], binWidth, geoDir)
# _Edit stuff below to use the App_
# ## Flux Output
# Below is the output generated by `Flux.pl` using the threshold files `6148.2016.0109.0.thresh` and `6119.2016.0104.1.thresh` (separately) as input:
# ```
# $ perl ./perl/Flux.pl files/6148.2016.0109.0.thresh outputs/ThreshFluxOut6148_01 600 geo/
# $ head -15 outputs/ThreshFluxOut6148_01
# #cf12d07ed2dfe4e4c0d52eb663dd9956
# #md5_hex(1536259294 1530469616 files/6148.2016.0109.0.thresh outputs/ThreshFluxOut6148_01 600 geo/)
# 01/09/2016 00:06:00 59.416172 8.760437
# 01/09/2016 00:16:00 63.291139 9.041591
# 01/09/2016 00:26:00 71.041075 9.579177
# 01/09/2016 00:36:00 50.374580 8.066389
# 01/09/2016 00:46:00 55.541204 8.469954
# 01/09/2016 00:56:00 73.624386 9.751788
# 01/09/2016 01:06:00 42.624645 7.419998
# 01/09/2016 01:16:00 54.249548 8.370887
# 01/09/2016 01:26:00 45.207957 7.641539
# 01/09/2016 01:36:00 42.624645 7.419998
# 01/09/2016 01:46:00 65.874451 9.224268
# 01/09/2016 01:56:00 59.416172 8.760437
# 01/09/2016 02:06:00 94.290881 11.035913
# ```
# ```
# $ perl ./perl/Flux.pl files/6119.2016.0104.1.thresh outputs/ThreshFluxOut6119_01 600 geo/
# $ head -15 outputs/ThreshFluxOut6119_01
# #84d0f02f26edb8f59da2d4011a27389d
# #md5_hex(1536259294 1528996902 files/6119.2016.0104.1.thresh outputs/ThreshFluxOut6119_01 600 geo/)
# 01/04/2016 21:00:56 12496.770860 127.049313
# 01/04/2016 21:10:56 12580.728494 127.475379
# 01/04/2016 21:20:56 12929.475588 129.230157
# 01/04/2016 21:30:56 12620.769827 127.678079
# 01/04/2016 21:40:56 12893.309222 129.049289
# 01/04/2016 21:50:56 12859.726169 128.881113
# 01/04/2016 22:00:56 12782.226815 128.492174
# 01/04/2016 22:10:56 12520.020666 127.167443
# 01/04/2016 22:20:56 12779.643503 128.479189
# 01/04/2016 22:30:56 12746.060449 128.310265
# 01/04/2016 22:40:56 12609.144924 127.619264
# 01/04/2016 22:50:56 12372.771894 126.417419
# 01/04/2016 23:00:56 12698.269181 128.069490
# ```
# `Flux.pl` seems to give reasonable output with a threshold file as input, provided the DAQ has a geometry file that's up to standards. Can we interpret the output? Despite the lack of a header line, some reasonable inferences will make it clear.
# The first column is clearly the date that the data was taken, and in both cases it agrees with the date indicated by the threshold file's filename.
#
# The second column is clearly time-of-day values, but what do they mean? We might be tempted to think of them as the full-second portion of cosmic ray event times, but we note in both cases that they occur in a regular pattern of exactly every ten minutes. Of course, that happens to be exactly what we selected as the `binWidth` parameter, 600s = 10min. These are the time bins into which the cosmic ray event data is organized.
# Since we're calculating flux -- muon strikes per unit area per unit time -- we expect the flux count itself to be included in the data, and in fact this is what the third column is, in units of $events/m^2/min$. Note that the "$/min$" part is *always* a part of the units of the third column, no matter what the size of the time bins we selected.
# Finally, when doing science, having a measurement means having uncertainty. The fourth column is the obligatory statistical uncertainty in the flux.
# ## An exercise in statistical uncertainty
# The general formula for flux $\Phi$ is
#
# $$\Phi = \frac{N}{AT}$$
#
# where $N$ is the number of incident events, $A$ is the cross-sectional area over which the flux is measured, and $T$ is the time interval over which the flux is measured.
#
# By the rule of quadrature for propagating uncertainties,
#
# $$\frac{\delta \Phi}{\Phi} \approx \frac{\delta N}{N} + \frac{\delta A}{A} + \frac{\delta T}{T}$$
# Here, $N$ is the raw count of muon hits in the detector, an integer with a standard statistical uncertainty of $\sqrt{N}$.
# In our present analysis, errors in the bin width and detector area are negligible compared to the statistical fluctuation of cosmic ray muons. Thus, we'll take $\delta A \approx \delta T \approx 0$ to leave
# $$\delta \Phi \approx \frac{\delta N}{N} \Phi = \frac{\Phi}{\sqrt{N}}$$
# Rearranging this a bit, we find that we should be able to calculate the exact number of muon strikes for each time bin as
#
# $$N \approx \left(\frac{\Phi}{\delta\Phi}\right)^2.$$
#
# Let's see what happens when we apply this to the data output from `Flux.pl`. For the 6148 data with `binWidth=600`, we find
# ```
# Date Time Phi dPhi (Phi/dPhi)^2
# 01/09/16 12:06:00 AM 59.416172 8.760437 45.999996082
# 01/09/16 12:16:00 AM 63.291139 9.041591 49.0000030968
# 01/09/16 12:26:00 AM 71.041075 9.579177 54.9999953935
# 01/09/16 12:36:00 AM 50.37458 8.066389 38.9999951081
# 01/09/16 12:46:00 AM 55.541204 8.469954 43.0000020769
# 01/09/16 12:56:00 AM 73.624386 9.751788 57.000001784
# 01/09/16 01:06:00 AM 42.624645 7.419998 33.0000025577
# 01/09/16 01:16:00 AM 54.249548 8.370887 41.999999903
# 01/09/16 01:26:00 AM 45.207957 7.641539 35.0000040418
# 01/09/16 01:36:00 AM 42.624645 7.419998 33.0000025577
# 01/09/16 01:46:00 AM 65.874451 9.224268 51.00000197
# 01/09/16 01:56:00 AM 59.416172 8.760437 45.999996082
# 01/09/16 02:06:00 AM 94.290881 11.035913 72.9999984439
# ```
# The numbers we come up with are in fact integers to an excellent approximation!
# ---
# ### Exercise 1
# **A)** Using the data table above, round the `(Phi/dPhi)^2` column to the nearest integer, calling it `N`. With $\delta N = \sqrt{N}$, calculate $\frac{\delta N}{N}$ for each row in the data.
# **B)** Using your knowledge of the cosmic ray muon detector, estimate the uncertainty $\delta A$ in the detector area $A$ and the uncertainty $\delta T$ in the time bin $T$ given as the input `binWidth` parameter. Calculate $\frac{\delta A}{A}$ and $\frac{\delta T}{T}$ for this analysis.
# **C)** Considering the results of **A)** and **B)**, do you think our previous assumption that $\frac{\delta A}{A} \approx 0$ and $\frac{\delta T}{T} \approx 0$ compared to $\frac{\delta N}{N}$ is justified?
# ---
# ### Additional Exercises
# * Do the number of counts $N$ in one `binWidth=600s` bin match the sum of counts in the ten corresponding `binWidth=60s` bins?
#
# * Considering raw counts, do you think the "zero" bins in the above analyses are natural fluctuations in cosmic ray muon strikes?
#
# * Do the flux values shown above reasonably agree with the known average CR muon flux at sea level? If "no," what effects do you think might account for the difference?
# ---
# We can dig more information out of the `Flux.pl` output by returning to the definition of flux
#
# $$\Phi = \frac{N}{AT}.$$
#
# Now that we know $N$ for each data point, and given that we know the bin width $T$ because we set it for the entire analysis, we should be able to calculate the area of the detector as
#
# $$A = \frac{N}{\Phi T}$$
#
# One important comment: `Flux.pl` gives flux values in units of `events/m^2/min` - note the use of minutes instead of seconds. When substituting a numerical value for $T$, we must convert the command line parameter `binWidth=600` from $600s$ to $10min$.
#
# When we perform this calculation, we find consistent values for $A$:
# ```
# Date Time Phi dPhi N=(Phi/dPhi)^2 A=N/Phi T
# 01/09/16 12:06:00 AM 59.416172 8.760437 45.999996082 0.0774199928
# 01/09/16 12:16:00 AM 63.291139 9.041591 49.0000030968 0.0774200052
# 01/09/16 12:26:00 AM 71.041075 9.579177 54.9999953935 0.0774199931
# 01/09/16 12:36:00 AM 50.37458 8.066389 38.9999951081 0.0774199906
# 01/09/16 12:46:00 AM 55.541204 8.469954 43.0000020769 0.0774200035
# 01/09/16 12:56:00 AM 73.624386 9.751788 57.000001784 0.0774200029
# 01/09/16 01:06:00 AM 42.624645 7.419998 33.0000025577 0.0774200056
# 01/09/16 01:16:00 AM 54.249548 8.370887 41.999999903 0.0774199997
# 01/09/16 01:26:00 AM 45.207957 7.641539 35.0000040418 0.0774200083
# 01/09/16 01:36:00 AM 42.624645 7.419998 33.0000025577 0.0774200056
# 01/09/16 01:46:00 AM 65.874451 9.224268 51.00000197 0.077420003
# 01/09/16 01:56:00 AM 59.416172 8.760437 45.999996082 0.0774199928
# 01/09/16 02:06:00 AM 94.290881 11.035913 72.9999984439 0.0774199983
# ```
# In fact, the area of one standard 6000-series QuarkNet CRMD detector panel is $0.07742m^2$.
# It's important to note that we're reversing only the calculations, not the physics! That is, we find $A=0.07742m^2$ because that's the value stored in the `6248.geo` file, not because we're able to determine the actual area of the detector panel from the `Flux.pl` output data using physical principles.
# ## Testing binWidth
# To verify that the third-column flux values behave as expected, we can run a quick check by manipulating the `binWidth` parameter. We'll run `Flux.pl` on the above two threshold files again, but this time we'll reduce `binWidth` by a factor of 10:
# ```
# $ perl ./perl/Flux.pl files/6148.2016.0109.0.thresh outputs/ThreshFluxOut6148_02 60 geo/
# ```
# !head -15 outputs/ThreshFluxOut6148_02
# ```
# $ perl ./perl/Flux.pl files/6119.2016.0104.1.thresh outputs/ThreshFluxOut6119_02 60 geo/
# ```
# !head -15 outputs/ThreshFluxOut6119_02
# In the case of the 6148 data, our new fine-grained binning reveals some sparsity in the first several minutes of the data, as all of the bins between the `2:30` bin and the `13:30` bin are empty of muon events (and therefore not reported). What happened here? It's difficult to say -- under normal statistical variations, it's possible that there were simply no recorded events during these bins. It's also possible that the experimenter adjusted the level of physical shielding around the detector during these times, or had a cable unplugged while troubleshooting.
#
#
#
| Exploration_Flux/5_Flux_script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Iterative methods for solving linear systems
#
#
#
# Recall the prototypal PDE problem introduced in the Lecture 08:
# $$
# -u_{xx}(x) = f(x)\quad\mathrm{ in }\ \Omega = (0, 1)
# $$
# $$
# u(x) = 0, \quad\mathrm{ on }\ \partial\Omega = \{0, 1\}
# $$
#
# For the numerical discretization of the problem, we consider a **Finite Difference (FD) Approximation**. Let $n$ be an integer, a consider a uniform subdivision of the interval $(0,1)$ using $n$ equispaced points, denoted by $\{x_i\}_{i=0}^n$ . Moreover, let $u_i$ be the FD approximation of $u(x_i)$, and similarly $f_i \approx f(x_i)$.
#
# The linear system that we need to solve is
# $$
# u_i = 0 \qquad\qquad\qquad\qquad i=0,
# $$
# $$
# \frac{-u_{i-1} + 2u_i - u_{i+1}}{h^2} = f_i \qquad\qquad\qquad i=1, \ldots, n-1,\qquad\qquad\qquad(P)
# $$
# $$
# u_i = 0 \qquad\qquad\qquad\qquad i=n.
# $$
# +
# %matplotlib inline
from numpy import *
from matplotlib.pyplot import *
n = 33
h = 1./(n-1)
x=linspace(0,1,n)
a = -ones((n-1,)) # Offdiagonal entries
b = 2*ones((n,)) # Diagonal entries
A = (diag(a, -1) + diag(b, 0) + diag(a, +1))
A /= h**2
f = x*(1.-x)
# Change first row of the matrix A
A[0,:] = 0
A[:,0] = 0
A[0,0] = 1
f[0] = 0
# Change last row of the matrix A
A[-1,:] = 0
A[:,-1] = 0
A[-1,-1] = 1
f[-1] = 0
# Solution by direct method
u = linalg.solve(A, f)
# -
# ## Jacobi
#
# $$
# x_i^{k+1} = \frac{1}{A_{ii}} \times \left(b_i - \sum_{j\neq i} a_{ij}x_j^k\right)
# $$
#
#
# +
def jacobi(A, b, nmax=10000, eps=1e-10):
pass # TODO
sol_jacobi = jacobi(A, f)
print(linalg.norm(sol_jacobi - u)/linalg.norm(u))
# -
# ## Gauss-Seidel
#
# $$
# x_i^{k+1} = \frac{1}{A_{ii}} \times \left(b_i - \sum_{j=0}^{i-1} a_{ij}x_j^{k+1} - \sum_{j=i+1}^{N} a_{ij}x_j^k\right)
# $$
# +
def gauss_seidel(A,b,nmax=10000, eps=1e-10):
pass # TODO
sol_gauss_seidel = gauss_seidel(A, f)
print(linalg.norm(sol_gauss_seidel - u)/linalg.norm(u))
# -
# ## Gradient method
# $$
# {\bf r}^k = {\bf b} - A {\bf x}^k
# $$
#
# $$
# \alpha^k = \frac{{\bf r}^{k^{T}} {\bf r}^k}{{\bf r}^{k^{T}} A{\bf r}^k}
# $$
#
# $$
# {\bf x}^{k+1} = {\bf x}^k + \alpha^k {\bf r}^k
# $$
#
# ### Preconditioned gradient method
# $$
# P{\bf z}^k = {\bf r}^k
# $$
#
# $$
# \alpha^k = \frac{{\bf z}^{k^{T}} {\bf r}^k}{{\bf z}^{k^{T}} A{\bf z}^k}
# $$
#
# $$
# {\bf x}^{k+1} = {\bf x}^k + \alpha^k {\bf z}^k
# $$
#
# $$
# {\bf r}^{k+1} = {\bf r}^k - \alpha^k A{\bf z}^k
# $$
# +
def gradient(A, b, P, nmax=8000, eps=1e-10):
pass # TODO
sol_gradient = gradient(A, f, identity(len(A)))
print(linalg.norm(sol_gradient - u)/linalg.norm(u))
sol_preconditioned_gradient = gradient(A, f, A)
print(linalg.norm(sol_preconditioned_gradient - u)/linalg.norm(u))
# -
# ## Conjugate gradient
#
# $$
# \alpha^k = \frac{{\bf p}^{k^{T}} {\bf r}^k}{{\bf p}^{k^{T}} A{\bf p}^k}
# $$
#
#
# $$
# {\bf x}^{k+1} = {\bf x}^k + \alpha^k {\bf p}^k
# $$
#
# $$
# {\bf r}^{k+1} = {\bf r}^k - \alpha^kA {\bf p}^k
# $$
#
# $$
# \beta^k = \frac{(A{\bf p}^{k})^{T}{\bf r}^{k+1}}{(A{\bf p}^{k})^{T} {\bf p}^k}
# $$
#
# $$
# {\bf p}^{k+1} = {\bf r}^{k+1} - \beta^k{\bf p}^k
# $$
#
#
# ### Preconditioned conjugate gradient
#
#
# $$
# \alpha^k = \frac{{\bf p}^{k^{T}} {\bf r}^k}{(A{\bf p}^{k})^{T}{\bf p}^k}
# $$
#
#
# $$
# {\bf x}^{k+1} = {\bf x}^k + \alpha^k {\bf p}^k
# $$
#
# $$
# {\bf r}^{k+1} = {\bf r}^k - \alpha^kA {\bf p}^k
# $$
#
# $$
# P{\bf z}^{k+1} = {\bf r}^{k+1}
# $$
#
# $$
# \beta^k = \frac{(A{\bf p}^{k})^{T}{\bf z}^{k+1}}{{\bf p}^{k^T}A {\bf p}^k}
# $$
#
# $$
# {\bf p}^{k+1} = {\bf z}^{k+1} - \beta^k{\bf p}^k
# $$
#
# +
def conjugate_gradient(A, b, P, nmax=len(A), eps=1e-10):
N=len(A)
x = zeros_like(b)
tol = eps + 1
it = 0
r = b - dot(A,x)
rho_old = 1.
p_old = zeros_like(b)
while (it < nmax and tol > eps):
it += 1
z = linalg.solve(P,r)
rho = dot(r,z)
if (it > 1):
beta = rho/rho_old
p = z + beta*p_old
else:
p = z
q = dot(A,p)
alpha = rho/(dot(p,q))
x += p*alpha
r -= q*alpha
p_old = p
rho_old = rho
tol = linalg.norm(r,2)
print(it, tol)
return x
sol_conjugate_gradient = conjugate_gradient(A, f, identity(len(A)))
print(linalg.norm(sol_conjugate_gradient - u)/linalg.norm(u))
| notes/06a_linear_systems_iterative.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Lightning Talk Panel
#
# ## Notebooks
# - simple panel png displayer ##MAYBE SKIP
# - display geojson from an AOI folder of geojsons
# - deploy this geojson viewer app on a $10 ten dollar machine
# - on any TCP/IP port - single command from makefile
# - **s3 bucket analyzer**
# - a NOD to rclone -
# - Colorado Fire App Meets Jupyter - Sentinel - Open Data Cube
# - google earth - better and fascinating - 3d Models over burnt hulls
#
# ## Panel Gallery Below
# + language="html"
# <a href="https://panel.holoviz.org/gallery/index.html">link</a>
# -
from IPython.display import IFrame
IFrame('https://panel.holoviz.org/gallery/index.html', width=900, height=800)
# # Pretty Cool- cause:
# - it means very little code
# - and repeatable
| 4_Jan_2022/0_talk-1-11-2022-python-panel-rough-agenda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Implement from scratch
class MinHeap:
def __init__(self):
self.heapList = [0]
self.currentSize = 0
def percUp(self, i):
while i // 2 > 0:
if self.heapList[i] < self.heapList[i // 2]:
self.heapList[i], self.heapList[i // 2] = self.heapList[i // 2], self.heapList[i]
print(self.heapList)
i = i // 2
def insert(self, k):
self.heapList.append(k)
self.currentSize = self.currentSize + 1
self.percUp(self.currentSize)
def percDown(self, i):
while (i * 2) <= self.currentSize:
mc = self.minChild(i)
if self.heapList[i] > self.heapList[mc]:
self.heapList[i], self.heapList[mc] = self.heapList[mc], self.heapList[i]
print(self.heapList)
i = mc
def minChild(self, i):
if i * 2 + 1 > self.currentSize:
return i * 2
else:
if self.heapList[i * 2] < self.heapList[i * 2 + 1]:
return i * 2
else:
return i * 2 + 1
def delMin(self):
retval = self.heapList[1]
self.heapList[1] = self.heapList[-1]
self.currentSize -= 1
self.heapList.pop()
self.percDown(1)
return retval
def buildHeap(self, alist):
i = len(alist) // 2
self.currentSize = len(alist)
self.heapList = [0] + alist[:]
while (i > 0):
self.percDown(i)
i = i - 1
print("\nBuild heap")
heap = MinHeap()
heap.buildHeap([9, 6, 5, 2, 3])
print("\nInsert small")
heap.insert(1)
print("\nInsert large")
heap.insert(4)
print("\nDelete min")
heap.delMin()
# ### Use Collection Type
### import heapq
from random import shuffle
# #### Generate a random list
arr = [i for i in range(10)]
shuffle(arr)
print(arr)
# #### Heapify: O(N)
help(heapq.heapify)
heapq.heapify(arr)
print(arr)
# #### Heappop: O(1)
help(heapq.heappop)
heapq.heappop(arr)
print(arr)
# #### Heappush: O(logN)
help(heapq.heappush)
heapq.heappush(arr, 10)
print(arr)
# #### Heapreplace: O(logN)
help(heapq.heapreplace)
heapq.heapreplace(arr, 2)
print(arr)
# #### Heappushpop: O(logN)
help(heapq.heappushpop)
heapq.heappushpop(arr, 11)
print(arr)
| data_structure/heap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#import cell
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import simpleaudio as sa
import scipy
from scipy import fftpack as fft
from scipy.io import wavfile as wav
from scipy import signal as sig
import decimal as dec
# +
#import wav file for the pure sine sweep, guitar sine sweep, and room sine sweep.
fs, sine_sweep = wav.read('Sine Sweep.wav')
fs, guitar_sweep = wav.read('Guitar IR Sweep.wav')
fs, room_sweep_r = wav.read('Room IR Sweep Right.wav')
fs, room_sweep_l = wav.read('Room IR Sweep Left.wav')
pad_sine = np.zeros(2**20 - len(sine_sweep))
pad_guitar = np.zeros(2**20 - len(guitar_sweep))
pad_room = np.zeros(2**20 - len(room_sweep_r))
sine_sweep = np.concatenate([sine_sweep, pad_sine])
guitar_sweep = np.concatenate([guitar_sweep, pad_guitar])
room_sweep_r = np.concatenate([room_sweep_r, pad_room])
room_sweep_l = np.concatenate([room_sweep_l, pad_room])
#Guitar_sweep = Y[n]
#Sine_sweep = X[n]
#Transforming both through the FFT will give us Y(jw) and X(jw) respectively.
#Through this we can try to obtain the impulse response transform H(jw) through Y(jw)/X(jw)
fn_fft = np.linspace(-fs/2, fs/2, len(sine_sweep))
sine_fft = fft.fft(sine_sweep)
guitar_fft = (fft.fft(guitar_sweep))
room_l_fft = (fft.fft(room_sweep_l))
room_r_fft = (fft.fft(room_sweep_r))
plt.plot(fn_fft, np.abs(fft.fftshift(sine_fft)))
plt.figure()
def find_IR(y, x):
for i in range(0, len(x)):
if(np.abs(x[i]) < 4.5): #Filter out near 0 frequencies on the pure sine sweep. This eliminates noise.
x[i] = 100000000
ir_fft = y/x
plt.figure()
plt.plot(fn_fft, np.abs(fft.fftshift(ir_fft)))
toreturn = fft.ifft(ir_fft)
return toreturn
guitar_ir = find_IR(guitar_fft, sine_fft)
room_l_ir = find_IR(room_l_fft, sine_fft)
room_r_ir = find_IR(room_r_fft, sine_fft)
#n = np.arange(0, abs(2**20 / fs), 1/fs)
wav.write('Guitar IR.wav', fs, guitar_ir.astype('float'))
wav.write('Room IR Left.wav', fs, room_l_ir.astype('float'))
wav.write('Room IR Right.wav', fs, room_r_ir.astype('float'))
# +
#All encapsulating function
def create_IR_wav(name_sweep, name_sine, name_out, sample_length=2**20):
'''Function will take in 3 names, and output
the impulse response derived from the deconvolution
Arguments:
name_sweep: the sweep signal of the impulse response that needs to be derived, y[n]
name_sine: name of the pure sine sweep, x[n]
name_out: the name of the output file that will be the impulse response derived
sample_length: desired sample length to extend (pad with 0s) the signal. Recommended to be powers of 2.
Returns nothing
'''
#Read the required wav files
fs, y_signal = wav.read(name_sweep)
fs, x_signal = wav.read(name_sine)
#pad the wav files
pad_y = np.zeros(sample_length - len(y_signal))
pad_x = np.zeros(sample_length - len(x_signal))
y_signal = np.concatenate([y_signal, pad_y])
x_signal = np.concatenate([x_signal, pad_x])
#Derive the fourier transforms using fft
y_fft = fft.fft(y_signal)
x_fft = fft.fft(x_signal)
for i in range(0, len(x_fft)):
if(np.abs(x_fft[i]) < 4.5): #Filter out near 0 frequencies on the pure sine sweep. This eliminates noise.
x_fft[i] = 100000000
ir_fft = y_fft/x_fft
ir_output = fft.ifft(ir_fft)
wav.write(name_out, fs, ir_output.astype('float'))
return
#test the function
create_IR_wav('Guitar IR Sweep.wav', 'Sine Sweep.wav', 'Guitar IR func.wav')
| .ipynb_checkpoints/Convoluter_Deconvoluter-checkpoint.ipynb |